Abstract
The CPR and DPANN superphyla are globally distributed in anoxic habitats including extreme environments. However, the biogeography and potential ecological functions of their viruses remain unexplored. Here, we recover diverse CPR/DPANN metagenomic viral genomes from 90 acid mine drainage (AMD) sediments sampled across southeast China. Our data reveal deterministic processes as the primary driver of virome assembly shaping the distinct distribution patterns of CPR and DPANN viruses. While lifestyle prediction shows higher lytic virus diversity associated with DPANN, both CPR/DPANN viruses likely use the Piggyback-the-winner (PtW) strategy to co-exist with hosts in AMD sediments, with CPR viromes exhibiting increased lysis in low host-density regimes under intensive acidity/salinity conditions. A subsequent metatranscriptomic analysis uncovers diverse functional genes encoded by CPR and DPANN viruses actively expressed in situ, potentially supplementing host metabolisms yet diverging in replication, transcription, and translation-related functions. Furthermore, partial correlation network analysis suggests that putative symbiotic hosts of the CPR/DPANN may confer protection against viral infection through enhanced antiviral defense. Our results highlight the complex interplays between viruses, DPANN and CPR organisms, and their symbiotic hosts.
Similar content being viewed by others
Introduction
Recent genomic exploration has uncovered diverse CPR bacteria and DPANN archaea as major microbial dark matter lineages in a wide array of anoxic habitats including extreme environments (e.g., acid mine drainage (AMD)1,2, terrestrial geothermal springs3, hypersaline habitats4, and subsurface5,6,7). These uncultivated microorganisms are characterized by ultra-small (sub-micrometer-sized) cells, reduced genomes, and limited metabolic capacities, and constitute a remarkable position of the tree of life8,9. Due to their inability to de novo synthesize most nucleotides, amino acids and lipids, CPR and DPANN organisms have been speculated to adopt a symbiotic lifestyle relying on specific hosts for these necessary biomolecules10. Such symbiotic relationships have subsequently been supported by several co-culture studies11,12,13,14,15,16,17. Despite their extraordinarily high phylogenetic diversity, the community patterns and underlying ecological and evolutionary processes of CPR and DPANN remain poorly understood18.
Viruses are the most abundant biological entities on the planet19. They can infect all forms of life especially bacteria and archaea, and consequently significantly impact the prokaryotic community composition, functions and dynamics. Specifically, viruses may regulate host abundance by cell lysis, reprogram host metabolisms via virally encoded auxiliary metabolic genes (AMGs), and drive host evolution by horizontal gene transfer (HGT). Several recent studies have uncovered diverse CRISPR-Cas systems in metagenome-assembled genomes (MAGs) of specific DPANN and CPR lineages20,21,22,23,24, suggesting a rich virome associated with these episymbionts in the environment. Three-dimensional cryogenic electron tomography has demonstrated Thermoplasmatales-associated DPANN cells in acid mine drainage (AMD) biofilms as frequent targets of at least two types of viruses25. Metagenomics and virus-targeted direct-geneFISH have revealed lytic viruses infecting Candidatus Altiarchaeum in Earth’s crust21 and the general susceptibility of CPR and DPANN organisms to lytic viruses in groundwater26. Using a CRISPR screening approach, Wu et al. identified 97 globally distributed viruses and unclassified mobile genetic elements putatively associated with members across multiple DPANN phyla27. Despite these pioneering works, systematic comparative analyses of the viruses targeting CPR bacteria and DPANN archaea are lacking, especially in terms of their diversity patterns and potential influence on host prokaryotes over large-scale ecological ranges.
Here we report a comprehensive analysis of the biogeographical distribution and drivers of viruses infecting CPR and DPANN organisms in the AMD model system. Our study leveraged a massive collection of prokaryotic and viral genomes generated in a previous metagenomic survey of anoxic sediments sampled from geochemically diverse AMD environments across southeast China28. The prevalence and simultaneous enrichment of both lineages across a large number of samples from a single habitat (AMD sediment) have provided a unique opportunity to resolve the diversity patterns, functional repertoire and host interactions of their respective viruses along the explicit geographic and geochemical gradients.
Results
Characteristics of DPANN and CPR viromes in AMD sediments across southeast China
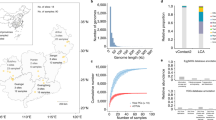
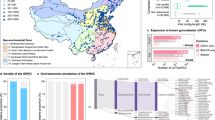
A total of 5678 viral population genomes (viral operational taxonomic units, vOTUs), which represented 11,112 viral genomes that ranged between 10-350 kb in size, were recovered from the metagenomes of 90 sediments sampled from diverse AMD environments across southeast China28 (Supplementary Data 1). Multiple host prediction approaches, including CRISPR-match, prophage in genome, genome homology match and integrated machine learning methods, were employed to screen viral genomes putatively infecting CPR or DPANN organisms. Finally, 241 CPR-associated vOTUs and 97 DPANN-associated vOTUs were identified (Fig. 1a and Supplementary Data 2, 3). Rarefaction curves of viral richness of both lineages showed a plateau, suggesting a sufficient coverage of our samples (Supplementary Fig. 1). The identified CPR vOTUs formed 162 viral clusters (VCs) and the DPANN vOTUs formed 71 VCs using vConTACT2 (Fig. 1b), with only a few clustering with previously reported groundwater and reference viruses. The clustering results based on 95% average nucleotide identity (ANI) further supported the rarity of these phages in both the Groundwater Virome Catalog (GWVC) and the IMG/VR database (Fig. 1c). Compared to groundwater CPR/DPANN viruses, those in AMD sediments exhibit hallmark proteins with significantly lower isoelectric points (pI), particularly in the DPANN lineages (Supplementary Fig. 2). This amino acid compositional bias indicates an adaptation to the high salinity in AMD habitats29. Taxonomic classification using PhaGCN2 revealed that 48.6% of the CPR vOTUs were assigned to the Caudoviricetes class, with Chaseviridae (14.5%), Mesyanzhinovviridae (12.8%) and Salasmaviridae (10.3%) being the most abundant families (Fig. 1d). While the majority of the DPANN viruses were also grouped within Caudoviricetes class, they were most frequently affiliated with Drexlerviridae (20.8%), Chaseviridae (18.9%), and Zobellviridae (18.9%). Resolving taxonomy of the infected hosts revealed that the CPR viruses mostly targeted the Paceibacteria (n = 94 virus-host linkages), while the DPANN viruses were mainly associated with members from the Micrarchaeales (85) and Parvarchaeales (48) (Fig. 1e). Notably, the viruses infecting the two lineages were predicted to adopt contrasting lifestyles: CPR viruses were primarily lysogenic, while DPANN viruses were largely lytic.
a Geographical distribution of DPANN and CPR viral richness. The provinces from which AMD sediments were sampled are depicted in gray. Sample sites are masked by pie charts. The pie size represents the viral richness at each site. The latitude and longitude grid and the scale bar are provided to precisely geo-reference the sampling area. b UpSet plot shows clustering result of CPR/DPANN viruses identified in this study and their overlap with Groundwater Virome Catalog (GWVC) and/or IMG/VR database. Vertical bars of upper plot show number of viral clusters of CPR, DPANN, and NCBI Viral RefSeq, denoted by circles below the histogram. The horizontal bars reflect the number of VCs in each viral datasets across all their possible combinations. Reference, NCBI Viral RefSeq. DPANN-AMD, DPANN viruses identified in this study. CPR-AMD, CPR viruses were identified in this study. c Shared and unique vOTUs between our dataset and public databases. d CPR (top) and DPANN (bottom) classification based on PhaGCN. Left: Bar charts represent the percentages of unclassified and classified vOTU number. Right: Pie charts show the classified viral proportion at family level. Classified vOTUs were further distributed to Caudoviricetes based on ICTV taxonomy. e The number of identified viral-host links in different taxa is illustrated in the alluvial plot.
Distribution of DPANN and CPR viruses in AMD sediments
A distance-decay relationship (DDR) was observed within the CPR and DPANN viral communities (p < 0.01, Supplementary Fig. 3a and b). However, the correlations are relatively weak (R2 < 0.16), especially at the local scale (R2 = 0.03 for CPR viruses and non-significant for DPANN viruses). Normalized stochasticity ratios were calculated to further investigate the drivers of distribution. The results revealed a great significance of deterministic processes primarily influenced by environmental selection, which accounted for 48.4–78.6% of the total variations in the community (Supplementary Fig. 3c). Random forest (RF) analysis further revealed that the CPR viral evenness (Pielou and Shannon index) and richness (vOTU number and Chao index) exhibited significant decreases with increasing electrical conductivity (EC) value and increases with rising pH value (p < 0.01), indicating a sensitivity to the extreme environmental conditions (Supplementary Fig. 4). Similarly, the abundances of both CPR viruses and their hosts declined as environmental conditions became more extreme (e.g., lower pH, higher EC) (Fig. 2a). In contrast, DPANN and their viruses responded inversely to pH and increased with EC. Compared to CPR, DPANN were enriched in more extreme conditions, such as pH levels below 3 and EC exceeding 3 dS/m (Supplementary Fig. 5).
a Associations of environmental factors and abundances based on correlation and random forest regression model. The circle size represents the variable importance (i.e., proportion of explained variability), and the color gradient in the heatmap denotes Pearson’s correlation coefficients. The bar chart shows the explanation of the response variable by the best model. C-host, the abundance of CPR. C-virus, the abundance of CPR virus. D-host, the abundance of DPANN. D-virus, the abundance of DPANN virus. b Distribution of sulfate (g/kg), Fe (g/kg), Pb (mg/kg), EC (mS/cm), and pH levels across MRT-based groups (A-F). For each box plot, the center line represents the median, the box limits represent the 25th and 75th percentiles (IQR), and the whiskers extend to 1.5×IQR. All individual data points are overlaid (Group A: n = 39; Group B: n = 34; Group C: n = 3; Group D: n = 5; Group E: n = 2; Group F: n = 6). Sulfate, Fe, and Pb concentrations are displayed on a log2 scale. c the relative abundance of CPR and DPANN in different groups. Colors were used to represent different taxa. d Viral composition in different groups. e, f NMDS results of CPR (e) and DPANN (f) viral structure colored by MRT-derived groups. Boxplots show distributions of different groups in NMDS1 and NMDS2. For each box plot, the center line represents the median, the box limits represent the 25th and 75th percentiles (IQR), and the whiskers extend to 1.5×IQR. Group differences were tested using PERMANOVA (adonis) and ANOSIM with 999 permutations based on Bray-Curtis distances.
We further examined the differential environmental responses of CPR and DPANN viruses through multivariate regression tree (MRT) analysis, modeling viral community composition against physicochemical gradients (pH, EC, etc.) with sum of squares partitioning (Supplementary Fig. 6a, b). Results showed that the viral communities could be distinctly classified into six groups (Groups A-F) based on environmental factors (Fig. 2b–d), which was also supported by non-metric multidimensional scaling (NMDS) analyses and multivariate analysis of variance based on distances and Permutations (PERMANOVA, p < 0.01) (Fig. 2e, f). Most of the samples were categorized as having high sulfate concentrations, which is a major feature typical of acidic mine environments. In the AMD sediments with relatively low iron concentrations, EC was identified as the primary determinant of viral community structure, and DPANN viruses dominated the high-EC samples (Group B). Conversely, under iron-replete conditions, pH became the dominant driver of viral differentiation, with DPANN viruses prevailing in low-pH niches (Group C), while CPR viruses showing an inverse distribution pattern. These distribution patterns were further corroborated by principal coordinate analysis based on Bray-Curtis dissimilarity, where principal component 1 (PC1) significantly correlated with pH, EC, and ferric iron (Supplementary Fig. 6c, d).
The variations of virus-host interaction across environmental gradients
To investigate the possible relationships between viral infection strategies and niche differentiation of CPR and DPANN in AMD sediments, we examined virus-host dynamics across environmental gradients. The lysogenic proportions of both DPANN and CPR viruses were significantly positively correlated with host abundance, and virus-host abundance ratios (VHRs) were significantly negatively correlated with host abundance, indicating that these viruses tend to adopt the Piggyback-the-Winner (PtW) life strategy (Fig. 3a). Robust Linear Regression (RLM) and Aggregated Boosted Tree (ABT) analysis were used to explore the environmental drivers of the VHRs (log10 scale) and lysogenic proportions, revealing EC as one of most significant regulators (explaining 7.8% − 15.6% variation) (Supplementary Fig. 7, 8). Specifically, there was a positive correlation between EC and the VHR of CPR viruses (Fig. 3b). However, for DPANN viruses, the VHR decreased with increasing EC. Moreover, the lysogenic proportion of CPR viruses exhibited a negative correlation with EC, whereas DPANN viruses displayed an opposite trend (Fig. 3b), indicating a viral-host dynamic pattern akin to that observed in the corresponding VHRs. Notably, the viral-host dynamics of CPR were also significantly influenced by pH (Fig. 3c and Supplementary Fig. 7), which exerted suppressive effects on both CPR bacteria and their associated viruses (Fig. 2a). In contrast, no significant pH-dependent differentiation was found among the DPANN viruses. Thus, our results demonstrate that the differences in VHRs and lysogenic proportions between DPANN and CPR viruses were significantly modified by EC and pH (which specifically affected CPR) (Supplementary Fig. 9), with viral life strategies covarying with the differential distribution of CPR and DPANN lineages.
a Host abundance correlation with virus-host abundance ratios (VHRs, log10 scale) and lysogenic proportion. The blue lines indicate the regression lines. The linear correlations are determined using two-tailed Pearson tests. Pearson correlation coefficients and significance levels are displayed in each panel. b VHR and lysogenic proportion correlations with EC. Pearson correlation coefficients and significance levels are displayed in each panel. c The differences in VHR (top) and lysogenic proportion (bottom) between extreme acidic environments (pH ≤3.375) and non-extreme acidic environments. Box plots show the median (center line), 25th-75th percentiles (box limits), and 1.5×IQR whiskers. Statistical significance was assessed using Wilcoxon rank-sum tests (two-sided) with sample sizes of n = 11 for low pH groups and n = 67 for high pH groups. Significance levels: *p < 0.05, **p < 0.01, ***p < 0.001. d The relationship between the microdiversity of DPANN/CPR viruses and environmental factors is illustrated in the heatmap. The color gradient represents Pearson’s correlation coefficients, while the asterisk signifies the statistical significance from a two-tailed Pearson test, adjusted for multiple comparisons using the Benjamini and Hochberg false discovery rate control method. Significance levels: *p < 0.05, **p < 0.01, ***p < 0.001. e The number of CPR/DPANN viral genes undergoing positive selection (pN/pS > 2.5). In the column chart, each column represents different functional annotation categories respectively. To better display the functional annotations with a smaller quantity, the corresponding columns have been enlarged.
Evolution of CPR/DPANN viruses shaped by environmental variables
Considering that virus-host antagonistic interactions may exert evolutionary pressures on viral populations, we investigated the nucleotide diversities (pi and theta) and the ratio of non-synonymous to synonymous mutations (pN/pS) in CPR/DPANN viruses inhabiting the heterogeneous AMD sediments. Results showed that nucleotide diversities were significantly higher in the lytic viruses (Supplementary Fig. 10). Moreover, the nucleotide diversities of CPR/DPANN viruses positively correlated with mean annual temperature (MAT) and extreme environmental conditions, such as high electrical conductivity (EC) and heavy metal concentrations (Fig. 3d). Genes with a pN/pS ratio exceeding 2.5 are identified as being influenced by selective pressure. As expected, the majority of DPANN viruses that adopt a lytic lifestyle were found to possess more than twice the number of selected genes as compared to their CPR counterparts (Supplementary Data 4). Most of the viral genes putatively undergoing positive selection were unannotated. Of the 35 viral genes annotated, 16 encoded virus structural proteins, such as baseplate protein, portal protein, capsid protein, etc. (Fig. 3e). Notably, five infection-related genes (tail lysin and soluble lytic murein transglycosylase) and one defense-related gene (type I restriction enzyme M protein) were also found undergoing selection.
Viruses contribute to host’s environmental resistance
To explore the role of DPANN and CPR phages in countering AMD-induced perturbations and promoting host homeostasis, we conducted a comprehensive screening of viral genes against KEGG pathways and multifunctional databases (Fig. 4a) (see Methods for details). The identified AMGs primarily originate from lysogenic viruses, with the number of AMGs carried by this virus type being twice that of lytic viruses (Supplementary Data 5). Interestingly, two of the DPANN AMGs were annotated as encoding rpoS, a sigma factor responsible for conferring resistance to acidic and hyperosmotic resistance in cells30. This transcriptional regulator has not been previously reported in viral genomes to the best of our knowledge. Phylogenetic analysis revealed that these viral-derived rpoS homologs exhibited significant divergence from known rpoS sequences (bootstrap support >90%) (Supplementary Fig. 11), even though one of them was derived from a DPANN prophage (WY1_virus228) (Fig. 4b, Supplementary Fig. 12, Supplementary Data 6). Ectoine serves as a crucial cellular defense mechanism against hyperosmotic in bacteria. Strikingly, we found AMGs encoding ectoine synthase in two CPR phages (Previously, ectoine synthase genes have been found only on prokaryotic genomes). Furthermore, we found evidence that the lysogenic phages carrying the ectoine synthase AMG (WH2_virus31) along with its host bacterium (MAG WH2.bin7) exhibited a hyperosmotic resistance potential. Specifically, there was a significant negative correlation between the abundance of WH2.bin7 and EC level in the AMD sediments (Supplementary Fig. 13). However, this relationship was disrupted in WH2_virus31-infected samples. The Wilcoxon rank-sum test results were validated using bootstrap resampling approach, which demonstrated that the abundance of WH2.bin7 population was significantly higher in the WH2_virus31-infected samples exhibiting elevated EC levels (Supplementary Fig. 14). Furthermore, WH2_virus31 predominantly occurred at the MAS mine site (EC > 3) and the abundance of which showed a highly positive correlation with the local EC levels (Supplementary Fig. 15; R = 0.98; p < 0.001), indicating its remarkable success under hyperosmotic conditions.
a AMGs associated with adaptation, host characteristics and biochemical cycling. Gene names are labeled in different colors to indicate their involvement in various functions. b Genomic map of the CPR/DPANN virus featuring novel AMGs in resistance functions. Different functional categories are distinguished using various colors. c A cellular diagram illustrating segments of the sulfur metabolism (map00920), Cysteine and methionine metabolism (map00270), glycolysis / gluconeogenesis (map00010), Citrate cycle (map00020), and fatty acid biosynthesis (map00061) KEGG pathways. Genes detected in hosts and viruses are colored according to pathway type. The asterisk mark represents AMG. The prohibited symbol represents the absence of the gene in the corresponding host. Undetected genes and related metabolites (unmeasured) are grayed out.
Heavy metals are significant factors that could induce oxidative damage in AMD environments. Notably, we identified 10 CPR viral AMGs and seven DPANN viral AMGs involved in the biosynthesis of folate (DHFR, folE, moaA, queC, queD and queE) (Fig. 4a), which could facilitate the mitigation of reactive oxygen species (ROS) damage31, apart from another role in DNA modification that protect phage DNA from host restriction systems32. Meanwhile, AMGs related to nicotinate and nicotinamide metabolism (e.g., NAMPT, pncA, pncB, nadM, and nadE) were identified in both DPANN and CPR viruses (Fig. 4a). These genes have previously been implicated to enhance microbial resistance to oxidative damage31. Additionally, several AMGs encoding metalloregulatory proteins were identified in some CPR viruses. These include SmtB (ziaR), which is known for its ability to resist Cd concentration33. This AMG was predominantly found in Cd-enriched sediment samples, especially from the SKS, DBS and WY mine sites (Supplementary Fig. 16; Wilcoxon rank-sum test’s p < 0.001). The above-mentioned resistance associated AMGs were further validated using Phyre2 with a confidence level exceeding 99.7% and AlphaFold2 with a Predicted Local Distance Difference Test (pLDDT) value over 70 (Supplementary Data 5), demonstrating that each of the AMG sequences is likely to encode a structurally intact protein capable of performing its biological functions.
Complementing host metabolic potential and biochemical pathways
Our analyses also uncovered that the diverse AMGs may contribute to the host’s metabolic capabilities. Notably, a CPR phage encoding accA was identified as catalyzing the initial and rate-limiting step of fatty acid biosynthesis, which was absent in its host (Fig. 4a, c). Meanwhile, the ComEB annotated in both CPR viruses and DPANN viruses may facilitate DNA uptake and nucleotide scavenging. The comEB gene is a component of the comE operon, an important DNA uptake system in these symbionts10. Interestingly, two AMGs named purD were discovered in CPR viruses (Fig. 4c), which could contribute to de novo purine nucleotide synthesis. Phylogenetic analysis indicates that these viral-encoded purD likely originated from non-CPR phyla (Supplementary Fig. 17). In particular, one of purD from the CPR prophage shows the closest phylogenetic relationship with the Bacillota. For surface attachment function, AMGs involved in lectin biosynthesis were exclusively identified in CPR phages, whereas glycosyltransferase and nucleotide sugar biosynthesis genes were detected in both CPR viruses and DPANN viruses.
Abundant AMGs were found to impact biogeochemical processes (Fig. 4a). In terms of sulfur metabolism, a variety of AMGs (cysW, cysC, cysH, dmsA, and dmsC) were identified as complementing the hosts’ assimilatory sulfate reduction and DMSO degradation pathways. Moreover, cysH in viruses may also be involved in cysteine synthesis for potential disulfide bond stabilization of viral proteins34. Regarding iron cycling, a total of five AMGs were annotated, with four deriving from CPR viruses. Two of these genes showed positive correlations with Fe (Supplementary Fig. 18), which could assist the hosts in iron utilization, particularly in the case of WY3_virus73, which harbored two genes (one encoding PchH and the other encoding a permease) associated with the ABC transporter. For carbon cycling, a CPR phage was found to carry B-glucosidase (celB), which may facilitate cellulose hydrolysis by the host. Additionally, eight CPR viral AMGs (gpmB, gpmI, mdh and pgl) along with one DPANN viral AMG (pgl) were associated with glycolysis, gluconeogenesis, the citrate cycle, and the pentose phosphate pathway (TCA cycle). PhoH was identified as the major AMG related to phosphorus metabolism, with five out of the six sequences originating from DPANN viruses. Furthermore, phoH was predominantly found in lysogenic viruses, and the abundance of related viruses was observed to be highest in samples exhibiting low total phosphorus (TP) levels (Supplementary Fig. 19), suggesting viral regulation of hosts under phosphorus starvation.
Differences in replication, transcription and translation functions between the CPR viruses and DPANN viruses
To elucidate the underappreciated differentiation between viruses infecting CPR and DPANN, we examined the differences in their replication, transcription and translation functions. A core set of DNA replication proteins were identified, such as primase and DNA polymerase (Supplementary Data 7). Despite the identification of several subunits of bacterial DNA polymerase III in 20 CPR viruses and six DPANN viruses, an archaeal D-family DNA polymerase small subunit (K02323) was identified in TL1-5_virus16, a virus associated with the DPANN archaeon based on multiple methods. Notably, six out of the 12 proteins in TL1-5_virus16 are homologs of those found in giant viruses, including DNA topoisomerase I, an enzyme previously thought to be exclusive to eukaryotic viruses35. Both the CPR viruses and DPANN viruses recovered in this study possess a variety of transcription factors. Strikingly, the eukaryotic-like general transcription factor TBP, which has never been observed in viruses, was identified in 2 DPANN viruses. This transcription factor is an integral component of the archaeal transcription apparatus, which can be regarded as a simplified analogue of the eukaryotic Pol II counterpart36. Additionally, we found several aminoacyl-tRNA synthetases (arginyl-tRNA synthetase, aspartyl-tRNA synthetase, glutamyl-tRNA, and synthetase methionyl-tRNA synthetase) exclusively in the DPANN viruses (TL1-5_virus16, YP3_provirus18 and WH2_virus72), which were rich in transcriptional and translational genes. These enzymes have been identified in cellular organisms and giant viruses previously37,38, where they are responsible for attaching cognate amino acids to specific tRNAs during translation. Notably, one gene encoding acetylation lowers binding affinity (Alba) protein was identified adjacent to the aspartyl-tRNA synthetase gene. This protein may function as an RNA chaperone to protect RNA from degradation or as an archaeal component involved in the translation pathway for leaderless mRNA39,40. While conserved in many archaea, including DPANN, it has never been found in archaeal viruses.
Transcriptional activities of CPR/DPANN viruses in AMD sediments
To further investigate the metabolic activities of CPR/DPANN viruses in AMD sediments, we conducted a metatranscriptomic analysis on additional sediment samples collected from the DBS, DX, and YS mine sites with varying pH and EC values (Supplementary Data 8). Results revealed that seven DPANN viruses and five CPR viruses were actively expressed in situ. The detected transcriptions of genes related to replication and structure in the DPANN and CPR viruses highlighted their virion production in the AMD sediments. Notably, the activity of CPR viruses was only detected in samples with a pH value greater than 3, indicating their inability to thrive in extremely acidic conditions. The YS10-S-2_k141_1018658 (YP3_provirus18) associated with the DPANN host (MAS5.bin1) (through spacer match) demonstrated a robust activity in the YS10-S-2 sample, with 10 viral genes being actively expressed (Supplementary Fig. 20). One of these genes was identified as a glycosyltransferase. Structural model prediction using Phyre2 showed a 99.8% identity and 95% coverage with an O-antigen biosynthesis glycosyltransferase (wbnH), which is involved in the formation of surface lipopolysaccharide (LPS). We further analyzed the transcriptional profile of MAS5.bin1, revealing the expression of the S-layer protein. The S-layer protein necessitates LPS containing an O-antigen polymer for its attachment to the outer membrane surface, crucial for facilitating cell-to-cell interactions between DPANN archaea and their symbiotic hosts41,42.
Interactions between viruses and symbiotic hosts of DPANN/CPR
We initially identified the abundance-associated and metabolically complementary community members of the virus-targeted DPANN/CPR (see the “Methods” section) as putative symbiotic hosts (SHs). An additional horizontal gene transfer (HGT) analysis searching for signatures of past symbiosis was conducted. The yielded HGT events identified 12 reliable SHs and demonstrated a strong domain-level specificity (Archaea for DPANN, Bacteria for CPR) (Supplementary Data 9). Thus, for each virus-targeted DPANN/CPR, candidates were selected as SHs based on any of the following criteria: HGT evidence, highest abundance correlation, or highest MIP value within their respective domains. With these stringent criteria, a total of 152 MAGs were identified as SHs in our 2625-MAG dataset (Supplementary Data 10). Notably, 28 of the identified SHs were annotated as sulfate-reducing microorganisms (SRMs), indicating a prevailing association between the DPANN/CPR viruses and sulfur cycling SHs in the AMD sediments. Mantel tests further revealed significant correlations between DPANN/CPR viruses and these sulfate-reducing SHs (R² = 0.43 and 0.38; p < 0.001). These correlations remained significant (R² = 0.28 and 0.21; p < 0.001) even after accounting for their DPANN/CPR hosts as a covariate, supporting a strong relationship.
We subsequently constructed a partial correlation network focusing on significant associations between the SHs and viruses identified for each CPR/DPANN, exposing direct linkages that persist after controlling for confounding factors, including the CPR/DPANN itself and other co-occurring SHs and viruses (Fig. 5a). A total of 90 positive correlation and 45 negative correlation linkages were identified (Fig. 5b), indicating complex interactions, which are also present in HGT-related groups (Supplementary Data 11). Furthermore, in positive linkages, we observed a correlation between the gpmB-carrying virus MAS4_virus85 and its sulfate-reducing SH, consistent with a mutualistic carbon exchange relationship. The negative effect was exemplified by the YP4_provirus51 related linkages, which are dominated by negative correlations. The host (YP2.bin58) CRISPR-Cas system harbored a spacer matching the viral genome (with a 1-nt mismatch), and retained complete adaptation and interference modules (Fig. 5b). Correspondingly, an anti-CRISPR operon was identified in the viral genome, as indicated by a prediction score of 6.878 using AOminer43, suggesting an arms race relationship. Intriguingly, while the abundance of YP4_provirus51 positively correlated with both YP2.bin58 (Supplementary Fig. 21) and associated SHs (Fig. 5c), it exhibited a negative correlation with the availability of associated SHs, as reflected by the SHs-YP2.bin58 abundance ratio (Fig. 5d). This “source-sink” dynamic suggests SHs may protect DPANN/CPR symbionts by suppressing viral replication44. Genomic evidence further supports functional complementarity: the SH (MAS2.bin6) encoded complete antiviral systems (e.g., CBASS, AbiE), compensating for the YP2.bin58’s partial defenses in its genome including the defense island (NODE_67_length_192760_cov_10.863108; Supplementary Data 12–13). The AbiEi antitoxin obtained from the SH likely suppresses the expression of AbiEii toxin, and thus facilitates YP2.bin58 survival45.
a SH-virus association network based on partial correlation analysis. Node shapes distinguish SHs (circles) from viruses (triangles). Edge width represents the absolute value of partial correlation coefficients, and edge color indicates positive (red) or negative (blue) associations. Only SHs identified by horizontal gene transfer (HGT) events are labeled. Bar chart shows the distribution of positive and negative associations in the network. Boxplot displays the distribution of absolute partial correlation coefficients for all significant associations, providing insight into the strength of host-virus relationships. Boxplot show the median (center line), 25th-75th percentiles (box limits), and 1.5×IQR whiskers. All individual data points are overlaid. b Linkages of YP2.bin58 (one populations from DPANN) and the infecting phage. The phage is linked by CRISPR-Cas system. Functional modules of I-G CRISPR-Cas system are masked by different colors. c Scatter plot of abundance correlation between the SHs of YP2.bin58 and YP4_provirus51. The linear correlations are determined using two-tailed Pearson tests. Pearson correlation coefficients and significance levels are displayed in each panel. Gray points: viral abundance outliers (1.5×IQR, n = 5). Purple line: significant Pearson correlation after outlier exclusion; gray line: overall trend. d Scatter plot of YP4_provirus51 abundance versus SHs-YP2.bin58 abundance ratio (log10 scale). Pearson correlation coefficients and significance levels are displayed in each panel.
Discussion
The viruses residing the AMD model system have only recently been explored through metagenomics28. Our extensive sampling and metagenomic analysis have uncovered diverse and novel viruses putatively infecting CPR bacteria and DPANN archaea in the AMD sediments. This may be attributable to the wide range of geochemistry of the sediments and the diversified CPR and DPANN organisms therein. The large collection of CPR and DPANN associated viral genomes has enabled first insights into the biogeographic patterns of these viruses in the extreme environment, with EC and pH as the primary driving factors. Notably, contemporary environmental variation especially pH has been previously found to shape the regional and global distribution of microorganisms in AMD solutions46, and fluorescence in situ hybridization analysis has documented significant correlations between bacterial and archaeal population abundances and conductivity and rainfall in the Richmond Mine at Iron Mountain, Calif47. Thus, the biogeographic signals detected in the sediment CPR and DPANN viral assemblies resemble those in the AMD prokaryotic communities, in agreement with the obligate parasitic nature of viruses.
The microorganisms populating AMD environments encounter multiple extreme physiochemical conditions48,49. While previous studies have investigated the adaptation strategies of acidophiles thriving in these hostile conditions50,51,52, the potential role of their associated viruses has yet to be considered. This issue is particularly important in the current study as diverse and abundant DPANN and CPR organisms were detected in many of the AMD sediments analyzed and genome streamlining is a defining feature of these ultrasmall prokaryotes. Our results suggest that the CPR/DPANN viruses may enhance their hosts' resilience to extreme conditions, including extreme acidity, hyperosmolarity, and toxic heavy metals. The observed enrichment of DPANN viruses encoding phoH in phosphorus-deprived samples further indicates a viral potential to enhance the host’s adaptability to the oligotrophic AMD environments, although virally encoded phoHs have also been implicated for broader, environment-independent functions in the viral life cycle53. Similarly, recent studies have demonstrated the role of viruses in facilitating the detoxification processes of their hosts in soils contaminated with toxic organic compounds31,54. Meanwhile, the dependence of CPR and DPANN on symbiotic hosts is well recognized due to their inability to de novo synthesize key biomolecules (e.g., nucleotides, amino acids and lipids). Our analyses revealed that the CPR/DPANN viruses harbor the capability to compensate for their host’s metabolic processes and surface attachment, thereby emphasizing the viral contribution towards synthesizing essential materials and establishing symbiotic relationships. Collectively, these different lines of evidence implicate that the virus may represent an important yet overlooked factor leading to the ecological success of CPR and DPANN in our AMD sediments.
The iron and sulfur cycles are central to the functioning of AMD ecosystems. While Acidithiobacillus and Leptospirillum spp. as iron and/or sulfur oxidizers have been widely studied and implicated for their role in the formation of AMD (which is a significant environmental problem worldwide), recent genomic studies have highlighted the potential involvement of CPR and DPANN organisms in these key biogeochemical processes2,55. Our analyses further suggest a role of viruses by interacting with these ultrasmall, uncultivated prokaryotes in the AMD sediments. Specifically, viral infection may transform metabolisms of CPR and DPANN, especially through the identified virus-encoded genes associated with iron utilization, sulfate reduction, and DMSO degradation. Furthermore, the viral top-down control may not only regulate CPR and DPANN population dynamics, but also have an impact on their symbiotic interactions with free-living hosts and thus the assembly of the overall community. The former is particularly important because many of the predicted SHs of the virus-targeted CPR and DPANN are functional species from Leptospirillum, Acidithiobacillus, and Cuniculiplasma. It should be noted that Cuniculiplasma belong to the archaeal order Thermoplasmatales, members of which have either been implicated in acid generation in mine environments, or been demonstrated as episymbiotic hosts of DPANN through co-culture and enrichment cultures56,57. Additionally, many of the SHs (roughly 20%) predicted for the virus-targeted CPR and DPANN in our AMD sediments are potential sulfate-reducing microorganisms, including the Candidatus Acidulidesulfobacterium, which we discovered recently in an artificial AMD system58. The incomplete carbon degradation pathways in the CPR and DPANN may lead to the accumulation of pyruvate or acetyl-CoA10, which can serve as favorable carbon sources for their sulfate-reducing symbiotic hosts. Strikingly, some viruses are involved in the CPR’s glycolysis pathway, potentially enhancing the metabolic interdependence between these symbionts and their sulfate-reducing SHs. These findings are important as microbial sulfate reduction represents a promising technology for the bioremediation of AMD and associated environments. It should be reminded that the putative SHs in the present study were identified primarily through computational predictions (metabolic complementarity, co-abundance, HGT). While these represent ‘Standard’ methods in previous studies of CPR and DPANN symbionts59,60,61, further experiments integrating cultivation, high-resolution microscopy, and meta-omic approaches are needed to validate our predicted symbiotic interactions and their physical and ecological consequences.
The nature of the underlying processes governing the distinct ecological trajectories, as well as the evolutionary dynamics, of DPANN and CPR remains elusive. This is even more true for their little-studied viruses. Our collections of phylogenetically rich DPANN/CPR and their associated viral genomes recovered from the geographically separated and geochemically diverse AMD sediments provide a unique opportunity to address this knowledge gap. Our data suggest that the extreme environmental condictions may drive the evolution of DPANN/CPR viruses in the AMD sediments, particularly the genes related to infection and defense mechanisms in the DPANN viruses. This process may result in an arms race between the viruses and hosts, thereby driving their co-evolution. DPANN and CPR often evolve faster than free-living microorganisms, as their host-dependent lifestyle may lead to elevated mutation rates and loss of core functional gene62,63. Our results suggest that viruses may play a crucial role in this process, as exemplified by the functional genes phylogenetically related to those of other phyla identified in the prophages of the CPR and DPANN genomes. Interestingly, the DPANN viruses exhibit characteristics reminiscent of eukaryotic viruses. While this may indicate deep evolutionary roots potentially antedating eukaryote-archaeal divergence, convergent evolution cannot be ruled out. Additionally, our analyses revealed that DPANN and their viruses are more inclined to survive in extremely acidic and high-salt conditions. This is consistent with the ecological distribution of Thermoplasmatales (the major hosts of DPANN) in AMD systems49,64. A high proportion of the DPANN viruses exhibits a lytic lifestyle, aligned with a common feature of archaeal viruses in general65. In contrast, the CPR viruses are dominated by a lysogenic lifestyle and largely harbor a more diverse array of AMGs. This is important because, compared to DPANN genomes, CPR genomes are relatively streamlined in AMD sediments (Supplementary Fig. 22), resulting in their more constrained metabolic capabilities and thus likely a greater reliance on viral assistance. Of note, both CPR and DPANN viruses tend to adopt PtW strategy in AMD sediments. However, their lifestyle covaried with distinct host distributions under environmental gradients: the detected CPR viruses are predicted to exhibit increased lysis in low host-density regimes under intensive acidity/salinity condictions, while DPANN viruses with higher lytic diversity tend to adopt a relatively mutualistic dynamic. These covarying trends can be attributed to the fact that pre-existing host adaptations to ecological niches act as a selective force for the compatible virus-host dynamic. Meanwhile, viral predation may amplify distribution differences by suppressing host populations in extreme samples (e.g., CPR lysis under low-pH/high-EC conditions). Although this outcome is detrimental to the host, it presents an opportunity for the virus to disseminate to other suitable environments. Of note, inferences regarding virus-host dynamics at high pH/EC values should be interpreted with caution due to limited sample density in these ranges. Overall, the lifestyle differentiation between CPR and DPANN viruses reflects the intricate viral influence on microbial community assembly.
Resolving the spatiotemporal patterns of CPR/DPANN viruses is inherently challenging due to the complex and dynamic interplays between these viruses, the CPR/DPANN organisms and their free-living hosts. The reduced-complexity AMD ecosystem has been extensively studied through omics approaches to reveal their microbial community structure, dominant metabolic processes and inter-organism interactions48. Our current study is further benefit from the prevalence and abundance of both CPR and DPANN in the heterogeneous AMD sediments, revealing viruses as a pivotal role in their thriving and assembly. Nonetheless, the mechanisms generating and maintaining these ecological patterns remain largely unknown. To this end, AMD sediments as hot spots of CPR and DPANN could serve as ideal targets for designing further experiments to enrich our knowledge of the ecology and evolution of these uncultivated and enigmatic episymbionts and their associated viromes.
Methods
Sampling, metagenomic sequencing and physicochemical analyses of AMD sediments
From August to October in 2017, we collected > 100 surface sediments (0 ~ 10 cm) from geochemically diverse AMD environments in 18 mine sites across southeast China (22.96° − 31.68°N, 105.73° − 118.63°E) (Supplementary Data 1). Detailed procedures for sample collection and processing were described previously28,66. Total community genomic DNA extraction was successful (in terms of the quantity and quality of extracted DNA required for metagenomic sequencing) for 90 samples, for which metagenomic sequencing was conducted. The Illumina MiSeq sequencing (MiSeq Reagent Kit v3, 150-bp paired end reads) generated totally ~7 Tb raw reads data. A comprehensive dataset associated with these samples, which contains information on physicochemical properties, geographical distances, and climatic variables has been reported previously28,66. Specifically, the measured physicochemical parameters encompass pH, electrical conductivity (EC), total nitrogen (TN), total phosphorus (TP), total organic carbon (TOC), irons and heavy metals (Fe2+, Fe3+, total Fe, and Pb, Zn, Cu, Cd, Mn), as well as sulfate levels. The geographic and climate data consist of longitude and latitude coordinates, mean annual temperature (MAT), and mean annual precipitation (MAP).
Reconstruction of microbial and viral genomes and identification of DPANN and CPR viruses
Microbial and viral genomes were recovered and identified from the 90 AMD sediment metagenomes28. It should be noted that although viral genomes longer than 10 kb were used for ecological analyses in accordance with MIUViG standards67, they do not represent complete viral genomes. Taxonomic classification of CPR/DPANN genomes was performed using GTDB-Tk v2.3.0 (Genome Taxonomy Database r214)68 (Supplementary Data 14). The reads from each of the 90 sediment metagenomes were aligned to the viral representative genomes using BamM ‘make’ v1.7.3 (http://ecogenomics.github.io/BamM) with default parameters. Subsequently, the coverage of each sequence was determined using BamM ‘parse’ v1.7.3 in ‘tpmean’ coverage mode, which involved excluding the highest 5% and lowest 5% coverage regions, requiring a minimum nucleotide identity of 95%, and a minimum aligned length equivalent to at least 75% of each read.
To investigate linkages between DPANN/CPR and viruses, four methods were applied: (1) prophage detection within host metagenome-assembled genomes (MAGs); (2) alignment of host CRISPR spacers (predicted using metaCRT) against viral scaffolds via BLASTn (E-value ≤ 10⁻¹⁰, coverage > 95%, ≤1 mismatch); (3) sequence similarity analysis between viral and host genomes using BLASTn (E-value ≤ 10⁻³, bit score ≥50, alignment length ≥2.5 kb, identity ≥70%)69; and (4) machine learning-based host prediction with iPHoP70, using a custom database including MAGs from this study and a confidence score cutoff of 90. Prophages are identified as viral genomes that are precisely integrated within the scaffold of the host genome. Although these methods are reliable for genus-level host prediction70,71, more stringent criteria were applied for in-depth analyses. Only predictions supported by multiple lines of evidence—including tRNA matching (BLASTn with 100% identity and coverage) and k-mer similarity analyses—were retained for case study analysis. For k-mer-based predictions, significant associations were defined as follows: WIsH72 (p < 0.05), PHP73 (top-scoring host), and VirHostMatcher74 (score ≤0.25). tRNAs were identified in viral sequences using tRNAscan-SE75 (bacterial/archaeal mode) prior to matching with host genomes.
Viral classification and lifestyle prediction
Taxonomic annotation of the vOTUs was performed by PhaGCN2.02 with default parameters. Virus Metadata Resource of ICTV (International Committee on Taxonomy of Viruses) was used for the order level classification manually76. Viral lifestyle (lytic/lysogenic) was initially predicted by Deephage (v1.0)77 using a cutoff of 0.5. These predictions were subsequently refined by detecting lysogeny signals with CheckV78 and through manual detection of lysogeny-specific genes, including transposase, integrase, excisionase, resolvase, and recombinase as previously described79. In instances where discrepancies arose between DeePhage predictions and the lysogeny signals identified, the conflicting classifications were resolved based on the signal detection results. Open reading frames (ORFs) in each viral sequence were predicted using Prodigal v2.6.380 and compared against a custom set from Pfam81 using Hmmscan v3.3.282. The relative abundance of lysogenic viruses, referred to as the lysogenic proportion31, is used together with the VHR to infer viral life strategy (i.e., Kill-the-Winner or Piggyback-the-Winner).
Functional annotation and AMGs identification
Viral protein functions were annotated by VirSorter283, DRAM-v84 and VIBRANT85 with default parameters, and multifunctional databases (KEGG86, FeGenie55, SCycDB87, CCycDB88 and BacMet89). VirSorter2 (--prep-for-dramv) were used to generate the input files required for DRAM-v. The default parameters were employed for AMG identification in DRAM-v, resulting in the acquisition of AMG flag and auxiliary score. Genes with auxiliary score of 1–3 and AMG flag of –M and/or –F were considered as AMGs. Unannotated genes with auxiliary scores of 1–3 were assigned to multifunctional databases using BLASTp (with a threshold of 60 for the bit score and 10 − 5 for the E-value). The KEGG annotation was performed using KofamScan v1.2.06590, and the FeGenie annotation was conducted with default parameters in FeGenie.py. The genomic content of viral contigs containing AMGs was manually examined, and only the AMGs flanked by two viral genes were retained. To assess the genomic context, genome maps for viral contigs with AMGs were visualized based on DRAM-v and VirSorter2 annotations. Phyre2 (http://www.sbg.bio.ic.ac.uk/phyre2/) and Alphafold2 were employed for protein structural homology searches and prediction91,92.
Metatranscriptomic samples collection, sequencing and data processing
To investigate the potential activity and gene expression profiles of CPR/DPANN viruses in situ, AMD sediments (0 ~ 10 cm) were collected in 2023 from three (DBS, DX and YS) of the 18 mine sites. Each sediment was well mixed and divided into two fractions: one fraction for nucleic acid extraction (subsequently stored in liquid nitrogen) and the other for physicochemical measurements (air-dried). Total DNA and RNA was extracted using the RNeasy PowerSoil DNA Elution Kit (QIAGEN, Germany) and RNeasy PowerSoil Total RNA Kit (QIAGEN, Germany) according to the manufacturer’s instructions. The quality of DNA and RNA extracts was examined by an Agilent 4200 TapeStation System (Agilent, Germany). The rRNA transcripts were removed using Ribo-off rRNA Depletion Kit V2 (Bacteria) (Nanjing Vazyme Biotech Co.,Ltd.) in strict accordance with the provided instructions. Metagenomic and metatranscriptomic sequencing were conducted by the Magigene Company (Guangzhou, China) using an Illumina NovaSeq 6000 PE150 platform. Sequencing reads were quality-trimmed and assembled using the read_qc and assembly module of the metaWRAP software with default parameters93. Assembled transcriptomes were aligned to viral genes using BLASTn (E-value < 10 − 5, identity ≥95%, coverage = 100%). Viruses with at least three matched genes were considered to be active in the corresponding sample. Contaminated rRNA sequences were identified and eliminated using SortMeRNA with default parameters94. Relevant metatranscriptome reads were aligned to the viral genes using Bowtie2 (--very-sensitive). Subsequently, the expression level of each gene in every metatranscriptome was standardized as reads per kilobase per million mapped reads (RPKM) value.
Isoelectric point (pI) analysis of viral hallmark proteins
Viral hallmark genes were extracted from the results of Virsorter2. Protein sequences containing ‘X’ were excluded, resulting in the exclusion of three sequences out of a total pool comprising 11588 sequences. The pI of each protein was predicted in the Expasy ProtParam web server (https://web.expasy.org/protparam/).
Comparison of DPANN/CPR viral genomes to public databases
The identified vOTUs were subjected to comparative analysis with the viral genome databases of Groundwater Virome Catalog26 and IMG/VR v.4.095. Given the extensive scale of these databases, we initially employed the BLASTn algorithm to retrieve sequences from public databases that exhibit significant similarity to our viral genomes (coverage ≥50% and identity ≥95%). Subsequently, the retrieved sequences were combined with our viral genomes and all sequences were clustered using CD-HIT96 at a threshold of 95% sequence identity over 85% coverage. Viral clusters (VCs) were constructed using vConTACT v2.0197, integrating the genomes of vOTU identified in this study and prokaryotic viruses sourced from NCBI RefSeq database (release 211).
Symbiotic host identification of DPANN and CPR genomes
The genome-scale metabolic model of each DPANN and CPR MAG was reconstructed from its protein file using CarveMe (v.1.6.0) with default parameters98. Next, the metabolic interaction potential (MIP) of each pair of unique models was assessed using the global mode of SMETANA99. After removing NA results, a total of 215,151 MIP values were obtained representing the cooperative potential of genomic pairs. According to the MIP distribution, the 90th percentile (MIP = 5) was selected as the threshold score to filter out the pairs with relatively high interaction potential. 2482 putative symbiotic pairs were initially identified, which exhibit high cooperation potential (MIP ≥5) and positive abundance correlation (r > 0, p < 0.05). To provide more direct evidence supporting reliable host relationships (searching for signatures of past symbiosis as Li et al.59 described)), MetaCHIP100 was used to detect HGT events in these putative symbiotic pairs under default parameters. Finally, for each virus-targeted DPANN/CPR population, SHs were selected based on the following criteria: (1) evidence of HGT, and (2) both the candidate with the highest abundance correlation and the one with the highest MIP value within their respective domains (Archaea for DPANN and Bacteria for CPR). The average number of the targeted phyla associated with each CPR/DPANN genome is less than 2 (Supplementary Fig. 23). Symbiotic hosts containing DsrAB-type dissimilatory (bi)sulfite reductase were annotated as putative sulfate-reducing microorganisms (SRMs).
Microdiversity calculation
The metapop software utilized BAM files obtained from BamM v1.7.3 to identify single-nucleotide variants (SNVs) using local scales101. Viral populations were considered only if at least 70% of representative contigs had a minimum average depth of 10X coverage. SNVs meeting the criteria of QUAL score > 30 (phred-scaled), alternative allele frequency > 1%, and supported by at least four reads were retained as SNV loci. To address sequencing errors and coverage variations, genome-wide coverage was randomly subsampled to achieve a uniform 10X coverage per locus. To determine the average microdiversity in each sample, we computed mean π and Watterson’s θ values using 1000 bootstraps of 100 randomly subsampled π and θ values from the DPANN and CPR viruses (with replacement).
Partial correlation network analysis
To infer association networks between hosts and viruses, we calculated partial correlation coefficients based on microbial abundance data and constructed partial correlation networks as described by Fabbri et al.102. Specifically, for each CPR/DPANN population, we extracted the abundance profile of the population itself, its predicted symbiotic hosts (SHs), and its associated viruses (Supplementary Data 2, 14, 15). Pairwise partial correlation coefficients were computed using the pcor.shrink function from the corpcor R package. For networks with a small number of potential edges (≤10), we set the shrinkage intensity λ = 0, which is equivalent to traditional partial correlation estimation. For larger networks (> 10 edges), the shrinkage intensity λ was computed analytically by corpcor, with values between 0 and 1. Local false discovery rates (local FDR) were estimated from the resulting partial correlation coefficients using the fdrtool R package. We retained only associations with posterior probabilities (1 - local FDR) greater than 0.8 as significant edges. Finally, all results were pooled together, and the networks were constructed and visualized using the tidygraph and ggraph R packages, respectively.
Analysis of environmental drivers of virus-host dynamics
An initial screening of potential drivers was performed using univariate Robust Linear Regression (RLM) to assess the individual, linear effect of each environmental variable. This method is resistant to the influence of outliers, ensuring a more reliable assessment. For each predictor variable, a separate model was fitted against the response variable using the lmrob function (from the robustbase package in R) with the KS2014 setting for optimal robustness. To capture potential non-linear effects and complex interactions among environmental variables that cannot be detected by linear models, we employed a multivariate Aggregated Boosted Tree (ABT) model. This machine learning technique builds an ensemble of decision trees sequentially, where each new tree corrects the errors of the previous one, resulting in a powerful predictive model. The model was implemented using the gbm package in R. The model was trained using 5000 trees to ensure sufficient learning, and a 10-fold cross-validation procedure was applied to determine the optimal number of trees that minimized the prediction error and prevent overfitting.
Statistical analyses
Statistical analyses were conducted using different packages within the software R v4.2.270. Alpha-diversity indexes were calculated by employing the ‘diversity’ function within the vegan package. Bray–Curtis (abundance-based) distances were calculated to represent the viral and microbial beta-diversity using the vegan R package. The relative importance of deterministic processes was assessed using the NST package to determine the proportion of Modified Stochasticity Ratio (a special form of Normalized Stochasticity Ratio) below 50%103. The DDR rate was determined by calculating the gradient of a linear regression analysis using ln-transformed geographical distance and ln-transformed similarity in the microbial community composition. To select important indicators of the biotic matrices, we first conducted multicollinearity analysis of abiotic matrices using the ‘varclus’ function in the Hmisc package and set a threshold of 0.6. We utilized the randomForest R package to employ a machine-learning model, random forest, in order to assess the correlations between environmental factors and the compositional data. The rfPermute package was used to estimate the significance of importance metrics for a Random Forest model by permuting the response variable. The mvpart package was utilized to perform a sum of squares multivariate regression tree (MRT)104 (with default parameters) in order to establish the relationship between DPANN and CPR viral distribution with site characteristics. Non-parametric multivariate analysis methods, including ANOSIM and ADONIS, were employed to conduct significance analysis of the differences between different groups in non-metric multidimensional scaling (NMDS). All pairwise comparisons of distributed variables were performed using the Wilcoxon rank-sum test (alternative hypothesis = “greater” or “less”). As both CPR/DPANN and their viruses were scarce in the most extreme samples (pH <2 or EC > 7), these samples were excluded from analyses of virus-host dynamic differences. The Sankey diagrams were visualized using the RAWGraphs2.0 website (https://app.rawgraphs.io/).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw data from metagenomes, metatranscriptomes, and the assembled genomes of prokaryotic populations have been submitted to the NCBI BioProject database with accession code PRJNA666025. Reconstructed viral genomes can be accessed in the NCBI BioProject database using the accession code PRJNA648034 Source data are provided with this paper.
Code availability
The scripts and relevant data utilized for generating the figures in this study are included within this paper and can be accessed publicly on GitHub via the following link: (https://github.com/linzhl1/ultrasmall_phage_in_AMD).
References
Baker, B. J. et al. Lineages of acidophilic archaea revealed by community genomic analysis. Science 314, 1933–1935 (2006).
Chen, L. X. et al. Metabolic versatility of small archaea Micrarchaeota and Parvarchaeota. Isme j. 12, 756–775 (2018).
Qi, Y. L. et al. Analysis of nearly 3000 archaeal genomes from terrestrial geothermal springs sheds light on interconnected biogeochemical processes. Nat. Commun. 15, 4066 (2024).
Narasingarao, P. et al. De novo metagenomic assembly reveals abundant novel major lineage of Archaea in hypersaline microbial communities. Isme j. 6, 81–93 (2012).
Brown, C. T. et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523, 208–211 (2015).
Castelle, C. J. et al. Genomic expansion of domain archaea highlights roles for organisms from new phyla in anaerobic carbon cycling. Curr. Biol. 25, 690–701 (2015).
Probst, A. J. et al. Differential depth distribution of microbial function and putative symbionts through sediment-hosted aquifers in the deep terrestrial subsurface. Nat. Microbiol 3, 328–336 (2018).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol 1, 16048 (2016).
Castelle, C. J. et al. Biosynthetic capacity, metabolic variety and unusual biology in the CPR and DPANN radiations. Nat. Rev. Microbiol 16, 629–645 (2018).
He, X. et al. Cultivation of a human-associated TM7 phylotype reveals a reduced genome and epibiotic parasitic lifestyle. Proc. Natl. Acad. Sci. USA 112, 244–249 (2015).
Cross, K. L. et al. Targeted isolation and cultivation of uncultivated bacteria by reverse genomics. Nat. Biotechnol. 37, 1314–1321 (2019).
Sakai, H. D. et al. Insight into the symbiotic lifestyle of DPANN archaea revealed by cultivation and genome analyses. Proc. Natl. Acad. Sci. USA 119, e2115449119 (2022).
Krause, S. et al. The importance of biofilm formation for cultivation of a Micrarchaeon and its interactions with its Thermoplasmatales host. Nat. Commun. 13, 1735 (2022).
Golyshina, O. V. et al. ARMAN’ archaea depend on association with euryarchaeal host in culture and in situ. Nat. Commun. 8, 60 (2017).
Batinovic, S., Rose, J. J. A., Ratcliffe, J., Seviour, R. J. & Petrovski, S. Cocultivation of an ultrasmall environmental parasitic bacterium with lytic ability against bacteria associated with wastewater foams. Nat. Microbiol 6, 703–711 (2021).
Hamm, J. N. et al. Unexpected host dependency of Antarctic Nanohaloarchaeota. Proc. Natl. Acad. Sci. USA 116, 14661–14670 (2019).
He, C. et al. Genome-resolved metagenomics reveals site-specific diversity of episymbiotic CPR bacteria and DPANN archaea in groundwater ecosystems. Nat. Microbiol 6, 354–365 (2021).
Chevallereau, A., Pons, B. J., van Houte, S. & Westra, E. R. Interactions between bacterial and phage communities in natural environments. Nat. Rev. Microbiol 20, 49–62 (2022).
Burstein, D. et al. New CRISPR-Cas systems from uncultivated microbes. Nature 542, 237–241 (2017).
Rahlff, J. et al. Lytic archaeal viruses infect abundant primary producers in Earth’s crust. Nat. Commun. 12, 4642 (2021).
Liu, J., Jaffe, A. L., Chen, L., Bor, B. & Banfield, J. F. Host translation machinery is not a barrier to phages that interact with both CPR and non-CPR bacteria. mBio 14, e0176623 (2023).
Esser, S. P. et al. A predicted CRISPR-mediated symbiosis between uncultivated archaea. Nat. Microbiol 8, 1619–1633 (2023).
Crits-Christoph, A. et al. Functional interactions of archaea, bacteria and viruses in a hypersaline endolithic community. Environ. Microbiol 18, 2064–2077 (2016).
Comolli, L. R., Baker, B. J., Downing, K. H., Siegerist, C. E. & Banfield, J. F. Three-dimensional analysis of the structure and ecology of a novel, ultra-small archaeon. Isme j. 3, 159–167 (2009).
Wu, Z. et al. Unveiling the unknown viral world in groundwater. Nat. Commun. 15, 6788 (2024).
Wu, Z., Liu, S. & Ni, J. Metagenomic characterization of viruses and mobile genetic elements associated with the DPANN archaeal superphylum. Nat. Microbiol 9, 3362–3375 (2024).
Gao, S. et al. Patterns and ecological drivers of viral communities in acid mine drainage sediments across Southern China. Nat. Commun. 13, 2389 (2022).
Oren, A. Life at high salt concentrations, intracellular KCl concentrations, and acidic proteomes. Front Microbiol 4, 315 (2013).
Battesti, A., Majdalani, N. & Gottesman, S. The RpoS-mediated general stress response in Escherichia coli. Annu Rev. Microbiol 65, 189–213 (2011).
Xia, R. et al. Benzo[a]pyrene stress impacts adaptive strategies and ecological functions of earthworm intestinal viromes. Isme j. 17, 1004–1014 (2023).
Hutinet, G. et al. 7-Deazaguanine modifications protect phage DNA from host restriction systems. Nat. Commun. 10, 5442 (2019).
Busenlehner, L. S., Pennella, M. A. & Giedroc, D. P. The SmtB/ArsR family of metalloregulatory transcriptional repressors: Structural insights into prokaryotic metal resistance. FEMS Microbiol Rev. 27, 131–143 (2003).
Wang, L. et al. Potential metabolic and genetic interaction among viruses, methanogen and methanotrophic archaea, and their syntrophic partners. ISME Commun. 2, 50 (2022).
Kazlauskas, D., Krupovic, M. & Venclovas, Č. The logic of DNA replication in double-stranded DNA viruses: insights from global analysis of viral genomes. Nucleic Acids Res 44, 4551–4564 (2016).
Wenck, B. R. & Santangelo, T. J. Archaeal transcription. Transcription 11, 199–210 (2020).
Abergel, C., Rudinger-Thirion, J., Giegé, R. & Claverie, J. M. Virus-encoded aminoacyl-tRNA synthetases: structural and functional characterization of mimivirus TyrRS and MetRS. J. Virol. 81, 12406–12417 (2007).
Moniruzzaman, M. et al. Virologs, viral mimicry, and virocell metabolism: the expanding scale of cellular functions encoded in the complex genomes of giant viruses. FEMS Microbiol. Rev 47, fuad053 (2023).
Sanders, T. J., Marshall, C. J. & Santangelo, T. J. The role of archaeal chromatin in transcription. J. Mol. Biol. 431, 4103–4115 (2019).
Laursen, S. P., Bowerman, S. & Luger, K. Archaea: the final frontier of chromatin. J. Mol. Biol. 433, 166791 (2021).
Awram, P. & Smit, J. Identification of lipopolysaccharide O antigen synthesis genes required for attachment of the S-layer of Caulobacter crescentus. Microbiol. (Read.) 147, 1451–1460 (2001).
Johnson, M. D. et al. Cell-to-cell interactions revealed by cryo-tomography of a DPANN co-culture system. Nat. Commun. 15, 7066 (2024).
Yang, B., Khatri, M., Zheng, J., Deogun, J. & Yin, Y. Genome mining for anti-CRISPR operons using machine learning. Bioinformatics 39, btad309 (2023).
Zhong, Q. et al. Episymbiotic Saccharibacteria TM7x modulates the susceptibility of its host bacteria to phage infection and promotes their coexistence. Proc. Natl. Acad. Sci. USA 121, e2319790121 (2024).
Hampton, H. G. et al. AbiEi binds cooperatively to the type IV abiE toxin-antitoxin operator via a positively-charged surface and causes DNA bending and negative autoregulation. J. Mol. Biol. 430, 1141–1156 (2018).
Kuang, J. L. et al. Contemporary environmental variation determines microbial diversity patterns in acid mine drainage. Isme j. 7, 1038–1050 (2013).
Edwards, K. J., Gihring, T. M. & Banfield, J. F. Seasonal variations in microbial populations and environmental conditions in an extreme acid mine drainage environment. Appl Environ. Microbiol 65, 3627–3632 (1999).
Denef, V. J., Mueller, R. S. & Banfield, J. F. AMD biofilms: using model communities to study microbial evolution and ecological complexity in nature. Isme j. 4, 599–610 (2010).
Shu, W. S. & Huang, L. N. Microbial diversity in extreme environments. Nat. Rev. Microbiol 20, 219–235 (2022).
Baker-Austin, C. & Dopson, M. Life in acid: pH homeostasis in acidophiles. Trends Microbiol 15, 165–171 (2007).
Dopson, M., Baker-Austin, C., Koppineedi, P. R. & Bond, P. L. Growth in sulfidic mineral environments: metal resistance mechanisms in acidophilic micro-organisms. Microbiol. (Read.) 149, 1959–1970 (2003).
Dopson, M., Ossandon, F. J., Lövgren, L. & Holmes, D. S. Metal resistance or tolerance? Acidophiles confront high metal loads via both abiotic and biotic mechanisms. Front Microbiol 5, 157 (2014).
Goldsmith, D. B. et al. Development of phoH as a novel signature gene for assessing marine phage diversity. Appl Environ. Microbiol 77, 7730–7739 (2011).
Zheng, X. et al. Organochlorine contamination enriches virus-encoded metabolism and pesticide degradation associated auxiliary genes in soil microbiomes. Isme j. 16, 1397–1408 (2022).
Garber, A. I. et al. FeGenie: a comprehensive tool for the identification of iron genes and iron gene neighborhoods in genome and metagenome assemblies. Front Microbiol 11, 37 (2020).
Korzhenkov, A. A. et al. Archaea dominate the microbial community in an ecosystem with low-to-moderate temperature and extreme acidity. Microbiome 7, 11 (2019).
Golyshina, O. V. et al. Diversity of “Ca. Micrarchaeota” in Two Distinct Types of Acidic Environments and Their Associations with Thermoplasmatales. Genes (Basel) 10, 461 (2019).
Tan, S. et al. Insights into ecological role of a new deltaproteobacterial order Candidatus Acidulodesulfobacterales by metagenomics and metatranscriptomics. Isme j. 13, 2044–2057 (2019).
Li, Y. X. et al. Deciphering Symbiotic Interactions of “Candidatus Aenigmarchaeota” with Inferred Horizontal Gene Transfers and Co-occurrence Networks. mSystems 6, e0060621 (2021).
Zhang, Z. F., Liu, L. R., Pan, Y. P., Pan, J. & Li, M. Long-read assembled metagenomic approaches improve our understanding on metabolic potentials of microbial community in mangrove sediments. Microbiome 11, 188 (2023).
Peng, S. X. et al. Biogeography and ecological functions of underestimated CPR and DPANN in acid mine drainage sediments. mBio 16, e0070525 (2025).
Dombrowski, N., Lee, J. H., Williams, T. A., Offre, P. & Spang, A. Genomic diversity, lifestyles and evolutionary origins of DPANN archaea. FEMS Microbiol Lett 366 (2019).
Bokhari, R. H. et al. Bacterial origin and reductive evolution of the CPR group. Genome Biol. Evol. 12, 103–121 (2020).
Baker, B. J. et al. Enigmatic, ultrasmall, uncultivated Archaea. Proc. Natl. Acad. Sci. USA 107, 8806–8811 (2010).
Yi, Y. et al. A systematic analysis of marine lysogens and proviruses. Nat. Commun. 14, 6013 (2023).
Hao, Y. Q. et al. Microbial biogeography of acid mine drainage sediments at a regional scale across southern China. FEMS Microbiol. Ecol. 98,fiac002 (2022).
Roux, S. et al. Minimum Information about an Uncultivated Virus Genome (MIUViG). Nat. Biotechnol. 37, 29–37 (2019).
Chaumeil, P. A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Roux, S. et al. Ecogenomics and Potential Biogeochemical Impacts of Globally Abundant Ocean Viruses. Nature 537, 689–693 (2016).
Roux, S. et al. iPHoP: An integrated machine learning framework to maximize host prediction for metagenome-derived viruses of archaea and bacteria. PLoS Biol. 21, e3002083 (2023).
Edwards, R. A., McNair, K., Faust, K., Raes, J. & Dutilh, B. E. Computational approaches to predict bacteriophage-host relationships. FEMS Microbiol Rev. 40, 258–272 (2016).
Galiez, C., Siebert, M., Enault, F., Vincent, J. & Söding, J. WIsH: who is the host? Predicting Prokaryotic Hosts From Metagenomic Phage Contigs. Bioinformatics 33, 3113–3114 (2017).
Lu, C. et al. Prokaryotic virus host predictor: a Gaussian model for host prediction of prokaryotic viruses in metagenomics. BMC Biol. 19, 5 (2021).
Ahlgren, N. A., Ren, J., Lu, Y. Y., Fuhrman, J. A. & Sun, F. Alignment-free $d_2^*$ oligonucleotide frequency dissimilarity measure improves prediction of hosts from metagenomically-derived viral sequences. Nucleic Acids Res 45, 39–53 (2017).
Chan, P. P. & Lowe, T. M. tRNAscan-SE: Searching for tRNA genes in genomic sequences. Methods Mol. Biol. 1962, 1–14 (2019).
Siddell, S. G. et al. Virus taxonomy and the role of the International Committee on Taxonomy of Viruses (ICTV). J. Gen. Virol. 104, 001840 (2023).
Wu, S. et al. DeePhage: distinguishing virulent and temperate phage-derived sequences in metavirome data with a deep learning approach. Gigascience 10, giab056 (2021).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Cook, R. et al. Hybrid assembly of an agricultural slurry virome reveals a diverse and stable community with the potential to alter the metabolism and virulence of veterinary pathogens. Microbiome 9, 65 (2021).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 11, 119 (2010).
Mistry, J. et al. Pfam: The protein families database in 2021. Nucleic Acids Res 49, D412–d419 (2021).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput Biol. 7, e1002195 (2011).
Guo, J. et al. VirSorter2: a multi-classifier, expert-guided approach to detect diverse DNA and RNA viruses. Microbiome 9, 37 (2021).
Shaffer, M. et al. DRAM for distilling microbial metabolism to automate the curation of microbiome function. Nucleic Acids Res 48, 8883–8900 (2020).
Kieft, K., Zhou, Z. & Anantharaman, K. VIBRANT: automated recovery, annotation and curation of microbial viruses, and evaluation of viral community function from genomic sequences. Microbiome 8, 90 (2020).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28, 27–30 (2000).
Wang, B. et al. Metagenomic insights into the effects of submerged plants on functional potential of microbial communities in wetland sediments. Mar. Life Sci. Technol. 3, 405–415 (2021).
Zhou, J. CCycDB: an integrative knowledgebase to fingerprint microbial carbon cycling processes (Version 2.0). Zenodo https://doi.org/10.5281/zenodo.10045943 (2023).
Pal, C., Bengtsson-Palme, J., Rensing, C., Kristiansson, E. & Larsson, D. G. BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res 42, D737–D743 (2014).
Aramaki, T. et al. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 36, 2251–2252 (2020).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Powell, H. R., Islam, S. A., David, A. & Sternberg, M. J. E. Phyre2.2: a community resource for template-based protein structure prediction. J. Mol. Biol. 437, 168960 (2025).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP-a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
Camargo, A. P. et al. IMG/VR v4: an expanded database of uncultivated virus genomes within a framework of extensive functional, taxonomic, and ecological metadata. Nucleic Acids Res 51, D733–d743 (2023).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Bin Jang, H. et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat. Biotechnol. 37, 632–639 (2019).
Machado, D., Andrejev, S., Tramontano, M. & Patil, K. R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res 46, 7542–7553 (2018).
Zelezniak, A. et al. Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc. Natl. Acad. Sci. USA 112, 6449–6454 (2015).
Song, W., Wemheuer, B., Zhang, S., Steensen, K. & Thomas, T. MetaCHIP: community-level horizontal gene transfer identification through the combination of best-match and phylogenetic approaches. Microbiome 7, 36 (2019).
Gregory, A. C. et al. MetaPop: a pipeline for macro- and microdiversity analyses and visualization of microbial and viral metagenome-derived populations. Microbiome 10, 49 (2022).
Fabbri, L. et al. Childhood exposure to non-persistent endocrine disrupting chemicals and multi-omic profiles: A panel study. Environ. Int 173, 107856 (2023).
Ning, D., Deng, Y., Tiedje, J. M. & Zhou, J. A general framework for quantitatively assessing ecological stochasticity. Proc. Natl. Acad. Sci. USA 116, 16892–16898 (2019).
Glenn, D. A. Multivariate regression trees: a new technique for modeling species-environment relationships. Ecology 83, 1105–1117 (2002).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 31870111) the Natural Science Foundation of Guangdong Province (No. 2022A1515010625) the Basic and Applied Basic Research Foundation of Guangdong Province (No. 2024B1515040015).
Author information
Authors and Affiliations
Contributions
W.S.S. and L.N.H. designed the research; S.M.G. and Z.H.L. collected the data; Z.L.L. conducted the bioinformatic and statistical analysis with the help from S.M.G., S.X.P., L.Y.T., X.W.L. and S.Y.Z; Z.L.L. wrote the paper; S.M.G., F.G.M. and L.N.H. reviewed and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lin, ZL., Gao, SM., Peng, SX. et al. Biogeography and host interactions of CPR and DPANN viruses in acid mine drainage sediments. Nat Commun 16, 10492 (2025). https://doi.org/10.1038/s41467-025-65461-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65461-0