Abstract
Meiotic recombination generates new genetic combinations and ensures proper chromosome segregation, yet its regulation remains unclear in many species. Here, we show that Mediator of paramutation1 (Mop1), a regulator of DNA methylation, affects the recombination landscape in maize. High-resolution crossover mapping reveals that loss of MOP1 reduces crossover numbers during male meiosis, while the total number in female meiosis remains largely unchanged despite local shifts in crossover distribution. We find that new crossovers occur on miniature inverted-repeat transposable elements (MITEs), particularly in regions with greater genetic diversity. Analysis of DNA methylation and histone modifications, together with gene expression, shows that loss of MOP1 reduces methylation at MITEs near genes enriched with histone marks linked to open and active chromatin, and downregulates a gene required for chromosome organization. Together, these findings suggest that both chromosome structure and transposon methylation contribute to recombination control, providing insights into epigenetic control of genetic variation.
Similar content being viewed by others
Introduction
Most sexually reproducing organisms rely on a specialized cell division called meiosis to produce gametes or spores. A crucial step during meiosis is meiotic recombination, which ensures the formation of bivalents between homologous chromosomes and their proper segregation. This process begins with the formation of double-strand breaks (DSBs) by topoisomerase-like proteins, SPO11-1 and SPO11-2, along with other proteins1,2,3. ASYNAPTIC1 (ASY1), a HORMA-domain protein, coats chromosome axes and is required for homolog recognition and synapsis4,5,6. ASYNAPTIC3 (ASY3) functions as a structural axis protein that anchors ASY1 to chromosomes7, while ASYNAPTIC4 (ASY4) stabilizes both ASY1 and ASY3, ensuring proper axis assembly8. ZIPPER1 (ZYP1), the transverse filament protein of the SC, bridges homologous chromosomes during synapsis and is critical for crossover (CO) interference and the assurance of at least one obligate CO per homologous pair9.
Following DSB formation, the 5’-ends of the broken DNA are resected to generate 3’-single-stranded DNA tails, which invade the non-sister chromatid of a homologous chromosome with the assistance of RAD51 and DMC1 recombinase proteins, eventually forming a stable intermediate known as the double Holliday junction (dHJ). These dHJs serve as key intermediates in homologous recombination10,11. Depending on the resolution pathway, they can be resolved into COs or, less commonly, non-crossovers (NCOs), although many NCOs are thought to result from synthesis-dependent strand annealing events that never form a dHJ12. The majority of DSBs are resolved as NCOs, with a smaller fraction resulting in COs. For example, in Arabidopsis, ~ 200 DSBs occur during meiosis, with only around 10 resolved as COs13, whereas in maize, 11–20 COs are observed out of 200 to 500 DSBs14. The mechanisms that determine which DSBs are resolved as COs remain poorly understood. However, at least three mechanisms play a role: CO assurance ensures the formation of at least one obligatory CO per homologous chromosome pair for proper orientation and segregation15; CO interference influences CO distribution by preventing multiple COs in close proximity16; and CO homeostasis regulates the overall number of COs to maintain genomic stability17. However, the effects of CO homeostasis appear to be limited in maize14.
Meiotic recombination varies between species and even sexes. In mammals, recombination hotspots are often associated with histone 3 lysine 4 trimethylation (H3K4me3) peaks near PRDM9 binding sites, a key determinant of recombination hotspots18,19. In humans, males exhibit lower recombination rates than females across most of the genome, but showed elevated rates near the telomeres20. Similar patterns are observed in many other eukaryotes, such as sticklebacks21,22. In plants, which lack PRDM9 homologs, recombination hotspots are associated with genes, particularly near transcription start sites (TSSs) and transcription termination sites (TTSs), which are characterized by reduced nucleosome density, DNA hypomethylation, low histone 3 lysine 9 dimethylation (H3K9me2), and increased levels of H3K4me3 and H2A.Z23,24,25. Recombination landscapes in plants also differ between sexes. In Arabidopsis thaliana, male meiosis shows higher CO rates than female meiosis, particularly at the ends of chromosomes26. In barley (Hordeum vulgare), male meiosis exhibits higher recombination frequencies than female meiosis at low temperatures, and this difference becomes more pronounced at higher temperatures27. In maize (Zea mays), more COs were reported in male meiosis than in female meiosis in hybrids of the Zheng58 and SK lines28. However, in B73 × Mo17 hybrids, CO numbers and distributions are similar between the sexes, though local differences are observed25, suggesting that genetic background influences recombination landscapes.
COs are unevenly distributed along chromosomes and are generally enriched in gene-rich euchromatic regions but suppressed in heterochromatin, which contains heavily methylated and transcriptionally silenced transposable elements (TEs) with high H3K9me2 levels. DNA methylation in plants occurs in three different cytosine contexts, CG, CHG (where H = A, T, or C), and CHH29,30. In Arabidopsis, de novo methylation in all these three cytosine contexts is catalyzed by DOMAINS REARRANGED METHYLTRANSFERASE 2 (DRM2) through the RNA-directed DNA methylation (RdDM) pathway. CG cytosine methylation (mCG) is maintained by METHYTRANSFERASE 1 (MET1), and maintenance of CHG methylation (mCHG) is catalyzed by CHROMOMETHYLASE 3 (CMT3), a DNA methyltransferase that contains a chromodomain that recognizes H3K9me2 through a self-reinforcing loop between DNA methylation and H3K9me230,31,32,33. mCHH is maintained by persistent de novo methylation by DRM2 via the RdDM pathway, which requires the presence of small RNAs and relatively open chromatin, particularly on young and short transposons and the edges of long transposons. In addition, mCHH can also be maintained by CHROMOMETHYLASE 2 (CMT2), in conjunction with H3K9me2 to methylate internal regions of long transposons within deep heterochromatic regions34,35, similar as mCHG.
In Arabidopsis, the loss of methylation in the mutants of MET1 or DECREASED DNA METHYLATION 1 (DDM1) results in increased meiotic CO rates in chromosomal arms but decreased or unchanged CO rates in pericentromeric regions, despite these regions being highly methylated and expected to be more affected by the loss of methylation36,37,38. In contrast, disruption of non-CG methylation in cmt3 mutants or mutants of histone methyltransferase KRYPTONITE (KYP, also known as SUVH4, SUPPRESSOR OF VARIEGATION 3–9 HOMOLOG 4) increases meiotic recombination in the pericentromeric regions39, highlighting the complex role of DNA methylation and histone methylation in meiotic recombination. Unlike Arabidopsis, where mutants that have global and significant effects on DNA methylation are viable, such mutants are lethal in maize40, likely due to the extensive CG methylation in the maize genome41. In our previous study using maize mop1 (mediator of paramutation1) mutants, we demonstrated that the loss of CHH methylation increases recombination in chromosomal arms and decreases it in pericentromeric regions42. MOP1 is a putative ortholog of RNA DEPENDENT RNA POLYMERASE 2 (RDR2) in Arabidopsis, a major component in the RdDM pathway30,43. In mop1 mutants, 24-nucleotide (nt) small RNAs are dramatically reduced44, leading to a near complete loss of CHH methylation at the edges of TEs near transcriptionally active genes41,45. While our earlier research provided valuable insights, its resolution was limited by the analysis of large genomic regions using a few genetic markers and the small number of backcrossed (BC1) individuals, making it challenging to identify the specific genomic and epigenomic features influencing meiotic COs in mop1 mutants.
In this study, we perform whole-genome resequencing for 423 BC1 individuals from male and female mop1 mutant plants and their wild-type (WT) controls and identify a total of 4048 COs, 60% of which are mapped to less than 5 kb. Our data reveal that overall CO numbers are reduced in male mop1 mutants compared to female mop1 mutants. Redistribution of COs occurs in both sexes but is more pronounced in female meiosis. mop1 preferentially introduces new CO sites in females within regions of higher genetic diversity, particularly on miniature inverted repeat transposable elements (MITEs). These MITEs, which overlap with differentially methylated regions (DMRs) and CO sites along with their flanking regions, are enriched in open chromatin regions, exhibit higher levels of H2A.Z and histone acetylation in WT, and show reduced CHH methylation levels in mop1 mutants compared to other MITEs. Our study provides insights into how DNA methylation and TEs influence recombination landscapes in maize, with potential implications for other species.
Results
Reduced COs in male mop1 mutants
To determine the effects of the mop1 mutation on meiotic recombination, we performed whole genome resequencing of four backcrossed (BC1) populations derived from male and female mop1 mutant plants (Mo17/B73;mop1/mop1) and their male and female WT sibling controls (Mo17/B73;+/+) in the hybrid background of B73 and Mo17 (Fig. 1a). We obtained more than 4 million single nucleotide polymorphisms (SNPs) from each of the four populations (Supplementary Fig. 1 and Supplementary Table 1). With the exception of a few regions lacking polymorphisms, the SNPs covered nearly the entire genome, with an average density of 1.9 SNPs per kilobase (kb) (Supplementary Figs. 1–3). We used these SNPs as genetic markers and identified 1069 COs in female WT, 968 COs in female mop1, 914 COs in male WT, and 1097 COs in male mop1 in 110, 97, 94, and 122 BC1 individuals, which translates to 19.45, 19.75, 19.44, and 17.98 COs per meiosis, respectively (Fig. 1b, Supplementary Fig. 4, Supplementary Table 2, and Supplementary Data 1). Of these COs, 45% were mapped to less than 2 kb of two SNP makers, and 60% were within a 5 kb interval, demonstrating high-resolution CO mapping (Fig. 1b, Supplementary Fig. 5, and Supplementary Data 1). In both WT and mop1 genomes, COs were primarily confined to gene-dense chromosomal regions (Fig. 1c and Supplementary Fig. 6), consistent with patterns previously reported in other species, including potatoes46, tomatoes47, and maize25.
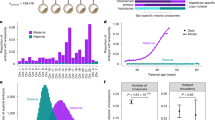
a Genetic strategy used to construct the backcross (BC1) populations. b COs and their interval sizes. c Distribution of COs across the ten maize chromosomes. The gray-shaded areas indicate the pericentromeric regions of each chromosome. d Average COs per meiosis in the four populations. Individual points represent data values, and black dashed lines indicate the mean. Statistical analysis was performed using a two-tailed Kruskal-Wallis test, followed by Dunn’s post hoc test with Bonferroni correction. *, P < 0.05; **, P < 0.01; ns, not significant. e Cumulative genetic distance and recombination rate of chromosome 1. Gray-shaded areas indicate the pericentromeric regions. Source data are provided as a Source Data file.
When comparing COs between WT and mop1, we found that very few COs were shared. For instance, of the 484 COs mapped within 2 kb intervals in female WT, only 21 (4.3%) overlapped with CO intervals in female mop1. Even after expanding the CO intervals by 2 kb on each side, the number of shared COs did not increase significantly. A similar pattern was observed between male WT and male mop1 (Supplementary Fig. 7). Despite these differences in CO positions, the average number of COs per female meiosis did not differ significantly between WT and mop1. In contrast, COs per male meiosis were nearly significantly reduced in male mop1 mutants compared to the male WT control (Fig. 1d, P = 0.058). While WT plants showed no significant differences in CO numbers between male and female meiosis, as previously observed in B73 and Mo17 hybrids25, COs were significantly reduced in male mop1 mutants compared to female mop1 mutants (Fig. 1d). These findings suggest that MOP1 plays a role in regulating recombination differences between male and female meiosis.
In our previous study, we noted that although the overall number of COs remained unchanged, mop1 mutants exhibited a shift in CO frequency toward chromosomal arms42. To further explore this, we separated the COs based on their locations in chromosomal arms and pericentromeric regions. However, this shift was not significantly detected in either female or male mop1 using our CO maps (Supplementary Fig. 8). Despite this, a closer examination of individual chromosomes revealed a distinct pattern of CO redistribution in female mop1 mutants, with increased COs in chromosomal arms and decreased COs in pericentromeric regions for chromosomes 1, 4, 5, 6, 7, 8, and 9 (Supplementary Figs. 9 and 10). A similar pattern was observed in male mop1 mutants for chromosomes 1, 2, 3, 5, 6, and 10 (Supplementary Figs. 9 and 11).
To further shed light on the role of MOP1 in meiotic recombination, we generated genetic maps using the Marey maps package48. Genome-wide distributions of genetic distances (cM) and recombination rates (cM/Mb) were similar between WT and mop1 mutants and between female and male meiosis (Fig. 1e, Supplementary Figs. 12–15, Supplementary Table 3, and Supplementary Data 2). Nevertheless, we still observed many distinct recombination peaks in different regions of chromosomes in both WT and mop1 mutants, as well as between the two sexes, indicating the local variations in recombination patterns across different genetic and sex-specific contexts.
Motif shifts in male and female mop1 COs
To understand the genomic sequences associated with COs in WT maize and determine whether they differ in mop1, we investigated the sequence motifs linked to COs. To enhance the accuracy of our analysis, we focused exclusively on 484, 417, 431, and 503 COs mapped within a 2 kb interval in female WT, female mop1, male WT, and male mop1, respectively. These COs were hereafter referred to as “high-resolution COs”. Sequences from these CO regions were extracted and merged to create a common dataset, and potential motifs were identified using STREME49. The top five most frequent motifs primarily consisted of C/G and A/T polymers (Fig. 2), consistent with previous reports associating such motifs with CO sites in maize25, wheat50, and rice51. All CO intervals contained at least one of these top five motifs. Interestingly, the frequencies of the top three C/G motifs were reduced in male mop1, compared to male WT. For instance, the top one C/G motif appeared at a frequency of 36.5% in male WT, but only 31.2% in male mop1. In contrast, the frequencies of the two A/T polymers were markedly increased in female mop1 compared to female WT. Specifically, the frequencies of the two A/T motifs in female WT were 30.6% and 37.6%, whereas in female mop1, they increased to 37.1% and 43.1%, respectively. However, these changes were not statistically significant based on the Q values (the adjusted P value under the Benjamini–Hochberg correction of the binomial tests), although the raw P-values for all five comparisons were significant (Supplementary Table 4). A/T polymers are known to be associated with regions of low nucleosome occupancy, which makes the DNA more accessible and facilitates the bindings of various molecular machineries52,53. These observations suggest that in female mop1 mutants, A/T enriched regions may transition to open chromatin states, potentially introducing new COs, whereas in male mop1, COs may tend to occur in regions with lower C/G content.
Blue-highlighted percentages indicate reduced frequencies of G/C-associated motifs in male mop1, whereas red-highlighted percentages denote increased frequencies of A/T-associated motifs in female mop1 mutants.
mop1 introduces new COs in regions of higher genetic diversity
Local sequence polymorphisms and structural variation, including SNPs, insertions, and deletions, are known to influence CO activity54,55,56. To explore whether mop1 preferentially introduces new CO sites in certain genomic regions, we compared SNP and InDel (insertions + deletions) density (number of SNPs and InDels between B73 and Mo17 per kb) in CO regions identified in mop1 mutants and wild types. Our analysis revealed that SNP density was significantly higher in CO regions identified in female mop1 mutants compared to those COs identified in female WT (Fig. 3a). A similar pattern was observed between male mop1 and male WT, but the difference was not statistically significant. Next, we analyzed SNP density around CO midpoints and their flanking regions. In both female and male WT and mop1, SNP density showed a pronounced dip at the CO sites (the midpoints of CO intervals), forming a clear valley. Interestingly, this valley was shallower in both female and male mop1 mutants compared to their WT counterparts (Fig. 3b), suggesting that mop1 COs may tolerate or even favor more local sequence diversity. To further investigate this, we examined CO rates across a range of SNP densities. In female mop1, CO rates peaked at approximately 13 SNPs/kb, whereas in female WT, the peak occurred at around 7 SNPs/kb (Fig. 3c). This distinct difference was not observed between male mop1 and male WT. Interestingly, we also noticed that when the SNP density was up to approximately 20 SNPs/kb, CO rates were near zero, suggesting that there is a threshold between sequence similarity and recombination outcomes.
a Average SNP density within CO intervals. Boxes represent the interquartile range (IQR) between the first (Q1) and third (Q3) quartiles, with the horizontal line inside each box indicating the median (Q2). Whiskers extend to the smallest and largest data points within 1.5 × IQR. Statistical analysis was performed using a two-tailed Kruskal-Wallis test, followed by Dunn’s post hoc test with Bonferroni correction. *, P < 0.05; ns, not significant. b Distribution of SNP density within 2 kb regions of CO midpoints. c Relationship between CO rates and SNP density across the whole genome and within the 2 kb flanking regions of genes containing TEs. Source data are provided as a Source Data file.
To better understand the genomic context of these CO regions, we categorized COs into four groups: 2 kb flanking regions of genes with TEs, 2 kb flanking regions of genes without TEs, TEs outside 2 kb flanking regions of genes, and unclassified regions. Our data showed that only COs located in the 2 kb flanking regions of genes with TEs exhibited significantly higher SNP density in female mop1 compared to female WT (Fig. 3c and Supplementary Fig. 16). This suggests that mop1 preferentially introduces new CO sites in regions with higher genetic diversity, specifically in TEs near genes.
Next, we compared InDels in CO regions between mop1 mutants and wild types. No significant differences were observed in either the number or length of InDels between CO regions in mop1 mutants and those in wild types in both female and male meioses (Supplementary Fig. 17). For example, the average InDel length was 4.9 bp in female and 4.4 bp in male mop1 mutants, which was not significantly different from 4.4 bp in female and 4.1 bp in male WT plants. Interestingly, recombination exhibited a much lower tolerance to InDels than to SNPs. CO rates peaked at approximately 4 InDels/kb but dropped to zero when the InDels reached 12 (Supplementary Fig. 18), indicating that InDels are more disruptive to homologous recombination.
COs are associated with MITEs, particularly in mop1 mutants
To identify local differences in CO patterns between WT and mop1 and between female and male meioses, we compared the 1835 high-resolution COs with genes and TEs. Approximately 69% of these COs were located within the 2 kb flanking regions of genes or within gene bodies, and 44.3% overlapped with TEs. Of these TEs, on average, 11.1% were located outside 2 kb flanking regions of genes, 24.7% were in the 2 kb flanking regions, and 8.6% were within gene bodies (Fig. 4a). Given that the mop1 mutation largely reduces CHH methylation at the edges of TEs near transcriptionally active genes and redistributes CO landscape41,42,45, we sought to determine whether specific types of TEs are associated with COs. We first performed correlation analysis between CO numbers and the density of various TE superfamilies, including long terminal repeat (LTR) retrotransposons, long interspersed nuclear elements (LINEs), terminal inverted repeat (TIR) DNA transposons, Helitrons, and miniature inverted repeat elements (MITEs). CO numbers were negatively correlated with LTR retrotransposons, particularly LTR/Gypsy concentrated in pericentromeric regions (Fig. 4b and Supplementary Figs. 19–22). In contrast, CO numbers were positively correlated with LINEs, TIR DNA transposons, Helitrons, and MITEs, particularly in chromosome arm regions (Fig. 4c and Supplementary Figs. 19–22). These correlations were consistent across four populations (female WT, female mop1, male WT, and male mop1) (Supplementary Figs. 21 and 22). Among these TEs, MITEs exhibited the strongest positive correlation with CO numbers (Fig. 4c). For example, of the randomly selected 1835 regions matched in length to the 1835 COs, only 1.5% overlapped with MITEs (Fig. 4d). In contrast, 33.8% of COs within 2 kb intervals in female WT overlapped with MITEs and this proportion increased to 39.3% in female mop1. A similar trend was observed in male WT (31.8%) and male mop1 (34.7%), although the increase was less pronounced compared to females (Fig. 4d). This pattern was true for MITEs both within and outside 2 kb flanking regions of genes, but the effect was stronger within these regions, where the majority of COs overlapped with MITEs (Supplementary Fig. 23). Those MITEs overlapping with COs are highly enriched near genes, especially in mop1 mutants (Supplementary Fig. 24). Furthermore, we examined MITE subtypes and found that DTA (derived from hAT elements) and DTH (Tourist-like, derived from PIF-Harbinger elements) were the two most enriched in COs, particularly in female mop1 mutants (Fig. 4e). These findings suggest that the mop1 mutation has a greater impact on CO patterns in females than in males, particularly through its association with MITEs near genes.
a Genomic locations of COs within 2 kb intervals. b COs per meiosis across the four populations and relative TE density across chromosome 7. The gray-shaded area represents the pericentromeric region. c Spearman’s rank correlation between crossover numbers (from all four populations) and MITEs in 1 Mb windows across the genome. d CO locations relative to TEs. A total of 1835 random regions were selected, each matched in length to one of the 1835 high-resolution COs. The midpoints of CO intervals and random regions were used to compare their locations with TEs. LTR, long terminal repeat retrotransposons; LINEs, long interspersed nuclear elements; DNA, DNA transposons; MITEs, miniature inverted-repeat transposable elements. e Classification of MITEs overlapping with COs. Numbers in parentheses indicate the raw counts of the COs overlapping with each MITE family. Source data are provided as a Source Data file.
CO hotspots are enriched at MITEs, coldspots at LTR retrotransposons
To further explore local differences in the CO landscape between WT and mop1 genomes, we identified CO hotspots and coldspots. Hotspots were defined as a 5 kb region with a recombination rate (cM/Mb) at least five times higher than the genome-wide average25. We identified 480, 391, 429, and 525 total CO hotspots in female WT, female mop1, male WT, and male mop1, respectively, which translates to an average of 4.4, 4.0, 4.6, and 4.3 hotspots per individual (Fig. 5a). Only a small portion of these hotspots were shared across populations. For example, only three hotspots were common to all four populations, and 33 were shared by at least three. Hotspots were predominantly unique between sexes and genotypes. Of the 871 hotspots identified in females and 954 hotspots in males, only 128 were shared between the two sexes. Comparing hotspots between WT and mop1, we observed that 58 (12.1%) hotspots in female WT overlapped with hotspots in female mop1 (14.8%), and 44 hotspots were shared between male WT (10.3%) and male mop1 (8.4%) (Fig. 5a).
a Number of CO hotspots. b Distribution of CO hotspots across the ten maize chromosomes. Gray vertical bars indicate chromosome boundaries, and red vertical bars represent the number of hotspots within 10 Mb windows. c Proportions of different types of transposable elements (TEs) within the 2 kb flanking regions of CO hotspots. Random regions, equal in number and matched in length to CO hotspots, were used as controls. d Number of coldspots. e Distribution of coldspots on chromosome 7. Blue regions mark coldspots, and red vertical dotted lines indicate the boundaries between chromosomal arms and pericentromeric regions. f Proportions of different types of TEs within coldspots across populations. Random regions, equal in number and matched in length to CO coldspots, were used as controls. LTR, long terminal repeat retrotransposons; LINE, long interspersed nuclear elements; DNA, DNA transposons; MITE, miniature inverted-repeat transposable elements. Source data are provided as a Source Data file.
Next, we examined the location of these hotspots. Our data showed that most of the hotspots were located at the chromosomal ends, with 713 (39.1%) and 1166 (63.9%) confined to the first 10 Mb and 20 Mb regions, respectively (Fig. 5b). To further understand these hotspots, we compared their locations with TEs. Approximately 64.9% were near MITEs. In addition, around 15.3% were associated with other DNA transposons (TIRs and Helitrons), while a relatively small proportion (17.2%) were near LTR retrotransposons, despite LTRs making up over 73.2% of the maize genome (Fig. 5c).
To identify enriched gene functions associated with COs and CO hotspots, we performed Gene Ontology (GO) analysis on genes located near COs and CO hotspots. Of the 210 genes identified within 10 kb of the high-resolution COs in female WT, 16 GO terms were significantly enriched, the majority of which were associated with transcription regulation and biosynthetic processes. In contrast, few enriched GO terms were found in female mop1, male WT, and male mop1 (Supplementary Fig. 25a). Extending this analysis to genes within 10 kb of CO hotspots, we identified 543, 434, 466, and 508 genes in the four populations. These genes were predominantly involved in binding activities, including small molecules, anion, ion, compound, and nucleotide binding (Supplementary Fig. 25b).
Next, we focused on CO coldspots, which were defined as genomic regions of at least 1 Mb in length devoid of any CO. We identified a total of 1167, 1310, 1191, and 1278 coldspots in female WT, female mop1, male WT, and male mop1, respectively (Fig. 5d). Unlike hotspots, coldspots were largely conserved across populations. For instance, 605 coldspots were common to all four populations, and an additional 413 were shared by at least three populations. These coldspots were primarily concentrated in the pericentromeric regions of chromosomes (Fig. 5e and Supplementary Fig. 26). It is not surprising to us that the majority of these coldspots overlapped with LTR retrotransposons, aligning with their predominance in pericentromeric regions (Fig. 5f). GO terms enriched in coldspot-associated regions were largely consistent across populations, with many terms related to metabolic processes, catalytic activity, and compound and small molecule binding (Supplementary Figs. 27–29).
mop1 likely introduces new COs by reducing methylation on MITEs in open chromatin
Our previous study hypothesized that the redistribution of the recombination landscape in mop1 mutants results from the removal of CHH methylation near transcriptionally active genes42. In this study, we were able to test this hypothesis using our high-resolution CO maps. By reanalyzing the DNA methylation data from immature anthers, which contain meiocytes undergoing meiosis I, of WT and mop1 mutants, we examined the DNA methylation levels at CO sites and their 2 kb flanking regions. Our data showed that average CG and CHG methylation levels at CO sites and their flanking regions were significantly lower than at random genomic regions (CG: 23.8% versus 78.3%; CHG: 12.4% versus 60.4%) (Fig. 6a and Supplementary Figs. 30 and 31). No significant differences in CG and CHG DNA methylation levels were observed at CO regions in mop1 compared to WT. In contrast, CHH methylation was significantly reduced at CO regions in mop1 mutants. Although CHH methylation levels at CO regions in WT were comparable to those in random regions, they were further reduced in mop1 mutants (Fig. 6a and Supplementary Figs. 30 and 31).
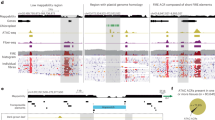
a DNA methylation levels around CO sites between mop1 mutants and WT controls for COs identified in male mop1 samples. Methylation data were obtained from immature anthers42. b Numbers of hypomethylated (hypo) and hypermethylated (hyper) DMRs, and overlap of hypo DMRs with CO sites ± 2kb. c Genomic distribution of DMRs overlapping or not overlapping with CO sites ± 2kb. d Proportion of TEs overlapping with both DMRs and CO sites ± 2kb, and with DMRs outside CO sites ± 2kb. DNA, DNA transposons; LTR, long terminal repeat retrotransposons; LINEs, long interspersed nuclear elements; MITEs, miniature inverted-repeat transposable elements. e DNA methylation levels of MITEs overlapping with DMRs within and outside CO sites ± 2 kb in WT and mop1. f Histone modification levels among MITEs in four categories. Histone data were obtained from leaves58. For panels (e and f), boxes represent the interquartile range (IQR) between the first (Q1) and third (Q3) quartiles, with the horizontal line inside each box indicating the median (Q2). Whiskers extend to the smallest and largest data points within 1.5 × IQR. Statistical significance was assessed using a two-tailed Kruskal-Wallis test followed by Dunn’s post hoc test with Bonferroni correction. *, P < 0.05; **, P < 0.01; ***, P < 0.001; ns, not significant. g Fine-scale visualization of a CO site and its DNA methylation and histone modification profiles in IGV88. The red box highlights hypo CHH and CHG DMRs on a MITE located within a CO. The blue box marks a CHH hypo DMR on a MITE outside CO regions. Source data are provided as a Source Data file.
To understand the effects of local methylation changes on COs, we next identified differentially methylated regions (DMRs) between WT and mop1 mutants in immature anthers42. In total, we identified 3189 CG, 4800 CHG, and 13,864 CHH DMRs in male mop1 mutants compared to the male WT control (Fig. 6b). Of these DMRs, 2749 (86.2%) CG, 4,140 (86.3%) CHG, and 12,162 (87.7%) CHH were hypomethylated (hypo), indicating that mop1 primarily removes DNA methylation. Interestingly, the numbers of both hypermethylated (hyper) and hypo CHH DMRs formed two distinct peaks near TSSs and TTSs, completely overlapping with the mCHH islands (Supplementary Fig. 32). This suggests that the mop1 mutation primarily affects these island regions. We next compared these DMRs with high-resolution COs. Surprisingly, less than 1% of the hypomethylated DMRs overlapped with CO sites and their 2 kb flanking regions (Fig. 6b). For example, 9 (0.3%) CG, 30 (0.7%) CHG, and 101 (0.8%) CHH DMRs overlapped with CO sites ± 2 kb, suggesting that the removal of DNA methylation alone is insufficient to create new CO sites in most regions.
Next, we investigated why some DMRs overlapped with COs while others did not by comparing their genomic locations. DMRs that did not overlap with CO sites ± 2 kb were predominantly located in TEs outside 2 kb flanking regions of genes and were enriched in LTR retrotransposons (Fig. 6c, d). In contrast, DMRs overlapping with CO sites ± 2 kb were primarily located in the 2 kb up and downstream regions of genes with TEs, the majority of which were MITEs (Fig. 6c, d). Among the nine CG DMRs overlapping with CO sites ± 2 kb, six overlapped with TEs, all of which were MITEs. For CHG DMRs, 23 out of 30 DMRs overlapped with TEs, with 15 (65.2%) being MITEs. Similarly, 97 out of the 101 CHH DMRs overlapped with TEs, of which 65 (67.0%) were MITEs (Fig. 6c, d). We further examined the methylation changes in these MITEs in mop1 and found that their methylation levels were significantly reduced in mop1, both within CO sites and their 2 kb flanking regions, as well as outside these regions. CHG and CHH methylation showed a greater reduction in MITEs overlapping with CO sites ± 2 kb (Fig. 6e). These findings suggest that mop1 is likely to introduce new CO sites by locally reducing DNA methylation at MITEs within 2 kb flanking regions of genes.
DNA methylation and histone modifications act as complementary defense mechanisms to maintain TE silencing57. In plants, recombination hotspots are often associated with reduced nucleosome density, DNA hypomethylation, low H3K9me2, and increased levels of H3K4me3 and H2A.Z23. To further demonstrate the relationship between MITEs and COs, we analyzed one histone variant (H2A.Z) and seven histone marks (H3K27ac, H3K56ac, H3K9ac, H3K27me3, H3K36me3, H3K4me1, and H3K4me3) using public histone data from maize leaves58. All these histone marks were higher at CO sites and their flanking regions compared to random regions, with H2A.Z, H3K27ac, H3K56ac, H3K9ac, and H3K4me3 peaking at CO sites (Supplementary Fig. 33). Next, we classified MITEs into four categories: i) MITEs overlapping with both DMRs and CO sites ± 2 kb, ii) MITEs overlapping with DMRs but not CO sites ± 2 kb, iii) MITEs outside DMRs but overlapping with CO sites ± 2 kb, and iv) MITEs outside both DMRs and CO sites ± 2 kb. Our data revealed that MITEs overlapping with both DMRs and CO sites ± 2 kb exhibited the highest levels of H2A.Z, H3K27ac, H3K56ac, and H3K9ac compared to the other three categories in WT leaves. MITEs overlapping with either DMRs or CO sites ± 2 kb displayed similar levels of these four histone marks, whereas MITEs outside both DMRs and CO sites ± 2 kb showed the lowest levels (Fig. 6f). In contrast, H3K27me3, H3K36me3, H3K4me1, and H3K4me3 did not differ significantly among the four categories (Supplementary Fig. 34).
To view COs and their relationship with MITEs and epigenetic status, we visualized a 49 kb region on chromosome 9. Within this region, we identified two CHH hypo DMRs in the mop1 mutant, with one overlapping with a CO and a MITE (highlighted in red), and another overlapping with MITEs but lacking a CO (highlighted in blue) (Fig. 6g). In the CHH hypo DMR that overlapped with a MITE and a CO, we observed elevated levels of H2A.Z and H3 acetylation, along with reduced H3K27me3 levels in this region in WT leaves. Interestingly, in this region, the loss of CHH methylation in mop1 was also accompanied by decreased levels of CG and CHG methylation. In contrast, the second CHH hypo DMR, which overlapped with a MITE but did not coincide with a CO, showed consistently high levels of CG and CHG methylation in both WT and mop1. This region also exhibited high levels of H3K27me3 and low levels of H3 acetylation in WT leaves. Together, these results suggest that new CO sites in mop1 mutants are associated with MITEs located at open chromatin regions marked by active histone modifications and characterized by reduced DNA methylation.
ASY4 is downregulated in mop1
To further explore the mechanisms underlying the shift in COs in mop1 and whether the loss of CHH methylation affects the expression of flanking genes, we performed transcriptomic analysis of immature anthers from mop1 mutants and their sibling WT controls. In total, we identified 186 upregulated and 199 downregulated differentially expressed genes (DEGs) (Supplementary Data 3). We then examined the expression of 41 genes involved in the meiosis pathway. Among them, only ASY4, a chromosome axis gene8, was significantly downregulated in mop1. Two additional genes, ZYP1 and ZYP1-interacting protein 4 (ZIP4), which are essential for proper chromosome synapsis and play key roles in obligate CO formation and CO interference59,60,61, were nearly significantly downregulated (Padj = 0.0991 and Padj = 0.1574, respectively; Supplementary Fig. 35 and Supplementary Data 4). This suggests that the altered CO landscape in mop1 may result from a combination of weakened axis/ SC structure due to reduced ASY4 and ZIP4 expression together with CHH hypomethylation, which collectively modulate chromatin configuration and recombination positioning.
Next, we assessed whether these DEGs harbored DMRs and COs within their bodies and 2 kb flanking regions. Only a small proportion of DEGs contained nearby DMRs. Specifically, 15 (3.9%) and 41 (10.6%) DEGs had CG and CHG DMRs, respectively, in these regions. A total of 83 (21.6%) DEGs harbored CHH DMRs nearby, the majority of which were hypomethylated. We then examined whether MITEs and COs overlapped with the DMRs of DEGs and found no COs near any of these 139 DEGs. These results suggest that the removal of CHH methylation at MITEs that introduce COs near genes does not necessarily alter gene expression.
CO interference is unchanged in mop1
CO interference regulates CO distribution by preventing multiple COs from occurring in close proximity16. We hypothesized that the altered CO landscape in mop1 mutants could be influenced by changes in CO interference. To test this, we measured CO interference using the crossover coefficient (CoC), a negative indicator of CO interference16. Our data revealed that no significant differences in CO interference in either female or male mop1 mutants, although female mop1 exhibited a trend toward lower CO interference compared to their WT controls (Supplementary Fig. 36), consistent with the non-significant changes yet reduced ZYP1 expression in mop1 mutants (Supplementary Fig. 35). Together, these findings suggest that the loss of CHH methylation does not result in significant changes in CO interference and that CO interference is unlikely to be the primary factor driving the CO changes in mop1.
Discussion
In this study, we investigated the recombination landscapes of both female and male meiosis in a mutant of a component that is essential in the RdDM pathway. Using high-resolution CO maps, we confirmed that the mop1 mutation exerts relatively subtle effects on global recombination rates, likely because it primarily removes CHH methylation at mCHH islands. Although CHH methylation comprises only 2.4% of genome-wide methylation, it is highly enriched near transcriptionally active genes and TEs, particularly in regions associated with recombination hotspots45,62. Our data demonstrate that mop1 influences recombination in a sex-specific manner at both global and local scales. Globally, CO numbers were reduced in mop1 males but remained unchanged in females (Fig. 1d). Locally, only ~4% of COs were shared between WT and mop1 (Supplementary Fig. 7). However, we do not believe these represent entirely new COs triggered by mop1. Rather, the global reduction of CHH methylation in mop1 may alter chromatin structure or positions, leading to small shifts in the apparent locations of COs along the reference genome. In other words, while the underlying CO events may be the same, they can be detected at slightly different genomic coordinates depending on chromatin context and mapping resolution. Interestingly, mop1 targets lower G/C enriched regions in males and more A/T enriched regions in females (Fig. 2). These findings highlight that mop1 has distinct and sex-specific influences on meiotic recombination. However, the mechanisms underlying these differences remain to be fully elucidated. Fine-scale recombination differences between sexes have been observed in maize hybrids of B73 and Mo17, despite the overall similarities in CO number and distribution25. Particularly, male and female CO sites exhibit differences in H3K4me3 and nucleosome occupancy patterns25. In mop1 mutants, the loss of CHH methylation is often accompanied by additional loss of CG and CHG methylation41,42,45, which may lead to changes in histone modifications and chromatin accessibility. It is possible that the degree of these changes differs between the sexes, resulting in distinct recombination patterns observed in mop1 mutants.
One of the most striking findings of this study is that MITEs are associated with the changes in the recombination landscape in mop1 mutants. The enrichment of Stowaway-like MITEs in CO regions has been previously observed in potato46, wheat50, and rice51. Here, we demonstrated that the reduction of DNA methylation in mop1 mutants can induce new COs at MITEs, particularly hAT-derived and Tourist-like MITEs (Figs. 4 and 6). MITEs preferentially insert into nucleosome-depleted, open chromatin regions, particularly gene promoters63,64. Tourist-like MITEs also exhibit a preference for TAA or TTA-rich sequences, which are commonly observed in motifs at CO sites25,65 (Fig. 2). These MITEs are located in regions enriched with active histone modifications, such as H2A.Z and histone acetylation (Fig. 6f). This makes them ideal targets in mop1 mutants, where CHH methylation is removed, facilitating the deposition of active histone marks and ultimately promoting the formation of new CO sites. It is worth noting that less than 1% of DMRs in mop1 mutants overlapped with COs (Fig. 6b), suggesting that while DNA hypomethylation and open chromatin are necessary for CO formation, they are not sufficient on their own. Previous studies have suggested that global reductions in DNA methylation are likely to influence recombination patterns via changes in chromatin structure chromosome-wide25, and thus the effect of DNA hypomethylation on recombination is indirect. In met1 and ddm1 mutants36,37,38, it has been proposed that CO remodeling is driven by the loss of CG methylation in the centromeric regions, which alters chromatin structure and leads to the inhibition of centromere-proximal DSBs38. However, we do not believe this is the case for mop1. Both CG and CHG methylation are not dramatically reduced in the centromeric regions in mop1 (Supplementary Fig. 31)42. Nevertheless, mop1 does reduce CG and CHG methylation in many genomic regions (Fig. 6b). Therefore, the effects of mop1 are likely confined to specific regions. Using high-resolution CO maps, we showed that mop1 precisely targeted TEs, particularly MITEs near genes, introducing some new CO sites at these regions. We propose that, rather than altering the overall chromatin structure of chromosomes, mop1 specifically modifies the chromatin state of particular regions, such as MITEs near genes. This localized change in chromatin state ultimately leads to alterations in recombination rates at these sites. Future investigations focusing on chromatin accessibility and histone modification profiles directly in meiocytes from mop1 mutants will be crucial to further elucidate the mechanisms underlying these observations.
Another important finding is that new CO sites in mop1 mutants were introduced in regions with higher genetic diversity, particularly during female meiosis (Fig. 3). Our data suggest that there is a threshold for sequence diversity between homologous alleles, and this threshold is larger in female mop1 mutants compared to female WT (13 versus 7 SNPs/kb), particularly in the 2 kb flanking regions of genes containing TEs (Fig. 3b, c). Beyond this threshold, CO rates drop dramatically, reaching near zero when SNP density exceeds 20 SNPs/kb (Fig. 3c). This parabolic relationship between CO frequency and SNP density has also been observed in Arabidopsis, where increasing SNP density is positively associated with COs until a threshold is reached, after which the relationship becomes negative66. CO formation is known to associate with elevated sequence polymorphisms66,67, which is mediated by MutS homolog 2 (MSH2), a key component that detects base mismatches in recombining molecules via the Class I CO formation pathway54,66,68. However, excessive sequence divergence between homologous chromosomes can inhibit normal homologous pairing, ultimately suppressing recombination. The observations in our study suggest that the RdDM pathway may have some direct or indirect effects on modulating MSH2 activity. We hypothesize that the loss of CHH methylation at TEs, particularly MITEs near genes, enhances chromatin accessibility in regions with moderate sequence divergence. This increased chromatin accessibility may allow MSH2 to tolerate greater sequence divergence (a higher polymorphism threshold) by creating a more “open” chromatin state. This compensates for the additional challenges posed by sequence mismatches during homologous recombination. Furthermore, we propose that this increased tolerance to sequence mismatches due to enhanced chromatin accessibility is more pronounced in female meiosis than in male meiosis. This hypothesis is supported by our observations that A/T-rich motifs, which are commonly associated with low nucleosome occupancy and open chromatin regions52,53, were enriched at CO sites in females (Fig. 2). In contrast, no significant differences in SNP density were observed at CO regions in male mop1 compared to male WT (Fig. 3). In mammals, female meiosis is frequently characterized by a less condensed chromatin state and more permissive recombination regulation compared to the highly compacted chromatin environment in male meiosis69. Whether this phenomenon is conserved in maize remains uncertain, as the technical challenges of analyzing female meiosis and embryo sac development in maize have limited direct examination of chromatin states during female meiosis.
Importantly, we observed that ASY4 is significantly downregulated in mop1 mutants (Supplementary Fig. 35 and Supplementary Data 4). ASY4 together with ASY1 and ASY3 form critical components of the meiotic chromosome axis and synaptonemal complex. In Arabidopsis, depletion of axis/SC components such as ASY1 and ZYP1 leads to redistribution of COs toward subtelomeric regions4,5,59,60, a pattern reminiscent of the CO shifts observed in mop1 in this and our previous study42. These observations suggest that the altered CO landscape in mop1 may result from the combined effects of reduced axis/SC component expression and CHH hypomethylation. This highlights a potential dual mechanism in which epigenetic changes and structural protein depletion collaboratively influence meiotic recombination.
Overall, our findings highlight the intricate role of mop1 in regulating meiotic recombination, with sex-specific effects, targeted alterations at MITE-associated regions, and tolerance for increased sequence diversity. These results provide valuable insights into the interplay between DNA methylation, chromatin remodeling, and recombination in plants. Importantly, our study suggests that manipulating epigenetic regulators like mop1 could provide an alternative approach to increase recombination in genomic regions near TEs or those with moderate sequence divergence. This has direct applications in crop improvement by enabling more efficient introgression of desirable traits, breaking linkage drag, and unlocking hidden genetic variation. Such strategies could be particularly valuable in maize and other crops whose genomes are largely composed of TEs.
Methods
Plant materials
The mop1 mutation was introgressed into the B73 and Mo17 backgrounds for at least seven generations. Heterozygous mop1 plants in the Mo17 background were crossed with heterozygous mop1 plants in the B73 background (Mo17;mop1/+ × B73;mop1/+) to generate F1 hybrid mop1 mutants (Mo17/B73;mop1/mop1) and their hybrid WT siblings (Mo17/B73;+/+). These F1 hybrid mop1 mutants and WT plants were reciprocally backcrossed with B73 to produce four BC1 populations derived from male mop1, female mop1, male WT, and female WT (Fig. 1a). These BC1 populations were used to detect COs in F1 meiocytes. The genetic stocks were constructed in the Ecology Research Center at Miami University (Oxford, Ohio).
Whole genome sequencing and SNP identification
Genomic DNA was isolated from 423 BC1 individuals and subjected to whole genome sequencing on the NovaSeq PE150 platform at Novogene, with a sequencing depth of ~ 3X per sample. We also downloaded the raw reads for parental genomes B73 and Mo17 (accession numbers ERP114875 and SRS302451) for comparative analysis. The raw reads were quality controlled by FastQC v0.11.7 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and trimmed using Trimmomatic v0.39 to remove adapters and low-quality reads70. Clean reads were mapped to the maize B73 reference genome version 5 using Bowtie 271,72. Reads with mapping quality scores < 10 or mapped to multiple locations were discarded. Duplicate reads were removed with PicardTools (version 1.98), and SNP calling was performed using the Genome Analysis Toolkit (GATK, ver. 4.4.0.0)73. Variant detection and genotype calling were performed with the HaplotypeCaller tool in GATK using default parameters. Variants were further filtered using VCFtools74 with the following criteria: minor allelic frequency between 0.1–0.4, minimum quality ≥ 30, minimum read depth ≥ 1, and ≤ 80% missing data per SNP site. SNP sites identified as heterozygous in either parental line (B73 or Mo17) were excluded from the analysis.
Determination of CO positions
The genome was divided into 200 kb windows, and the frequency of Mo17 alleles in each window was calculated as: Mo17 allele frequency = no. of Mo17 alleles / (no. of Mo17 alleles + no. of B73 alleles). Genotype breakpoints were determined using the mean shift method implemented in the R package ‘cumSeg’75. SNPs within 5 Mb upstream and downstream of each breakpoint were extracted, and a Hidden Markov Model (HMM) was applied to compute the posterior probability of B73/B73 homozygous or B73/Mo17 heterozygous regions76 (Supplementary Fig. 37).
CO landscape analysis
Genetic maps and recombination rates were developed using TASSEL77, R/qtl78, ASMap79, and MareyMap48. The VCF files were first converted to HapMap format and filtered in TASSEL 5 using the Filter Genotype Taxa Table function, applying a 50% missing data threshold per sample77. The filtered data were then reformatted into the ABH format, where homozygous alleles from the reference parent were coded as ‘A’, alternate homozygous alleles as ‘B’, and heterozygous alleles as ‘H.’ Given that this study involved BC1 populations, all homozygous alternate alleles (‘B’) were replaced with missing data (‘NA’). This formatted dataset was subsequently used for linkage mapping and recombination analysis using R/qtl, where additional quality control steps were performed, including error correction and missing data imputation using a Hidden Markov Model (fill.geno(method = “argmax”))78. Linkage groups were assigned using ASMap79, leveraging the MSTmap algorithm for efficient marker ordering. The final genetic map was constructed using the Kosambi mapping function (est.map) in R/qtl, and CO events were identified with locateXO. The recombination rates were estimated using MareyMap, based on genetic and physical distances48.
CO hotspots were identified by partitioning the genome into 5 kb windows, with windows having recombination rates five times higher than the genome-wide average considered hotspots. CO coldspots were defined as 1 Mb genomic regions devoid of COs, and adjacent coldspots were merged.
To determine COs in different genomic locations, we compared the locations of the COs with gene and TE annotations, which were downloaded from MaizeGDB using the intersect function in BEDTools80. If a CO overlapped with multiple annotations, the priority order was gene bodies, 2 kb upstream of genes, 2 kb downstream of genes, TEs, and unclassified regions. DMRs in the 2 kb flanking regions of genes were further classified based on the presence or absence of TEs.
Analysis of WGBS data and identification of DMRs
Raw reads of WGBS data were retrieved from a previous publication and processed following established bioinformatic pipelines41,42. Briefly, raw reads were trimmed with Trimmomatic v0.3970 and mapped to the B73 reference genome v5 using Bismark under the following parameters (-I 50, -N 1)71,81. PCR duplicates were removed using Picard tools. Alignments were performed to identify the methylated cytosines using the Bismark methylation extractor with default parameters. The numbers of methylated and unmethylated reads were counted using bismark2bedGraph and coverage2cytosine in Bismark.
To identify DMRs between WT and mop1, we first filtered out the cytosines with less than two mapped reads. The software “metilene” was used for DMR calling82. Specially, a DMR was determined as containing at least eight cytosine sites with the distance of 2 adjacent cytosine sites < 300 bp and with the average methylation differences in CG and CHG > 0.4 and in CHH > 0.2 between WT and mop141. These DMRs were compared with CO positions using the intersect function in BEDTools80.
Analysis of various histone modifications
ChIP-seq data from maize leaves, including the peak information for different histone modifications, were collected from a previously published source58. Because the peak positions were based on genome version 4, sequences of CO intervals in version 5 were remapped to version 4 using BLAST to determine their positions in version 4. The signals of each histone were measured in the CO regions following the formula (treat reads+1)/(input reads + 1) or log2(treat reads + 1)/(input reads + 1).
RNA sequencing and data analysis
The 1.5-2.0 mm long immature anthers were collected from the F1 hybrid plants of Mo17/B73;mop1/mop1 and Mo17/B73;+/+ for three biological replicates. To ensure the correct meiosis I stage, we first examined the expression of DMC1 (Disrupted Meiotic cDNA1) and ZYP1 using quantitative reverse transcription PCR (qRT-PCR)83. Total RNA was extracted from the immature anthers using the Qiagen RNeasy Plant Mini-Kit (Qiagen), according to the manufacturer’s instructions. Subsequently, library preparation and RNA sequencing were conducted at Biomarker Technologies (BMKGENE).
Raw RNA-seq reads were quality controlled by FastQC v0.11.7 and trimmed using Trimmomatic v0.39 to remove adapters and low-quality reads70. The trimmed reads were then aligned to the B73 reference genome version 5, using HISAT v2.2.184. Only uniquely mapped reads were retained. The number of reads per gene for each sample was quantified using HTSeq v0.11.285. DEGs between the WT and mop1 were identified using the R package DESeq2 v1.12.386, and significance was determined based on adjusted P-values from Wald tests with FDR correction.
Analysis of CO interference
CO interference was calculated using the crossover coefficient (CoC) based on the Beam Film (BF) model implemented in MADpatterns16,87. Genetic distances for each chromosome were converted to a scale of 0 to 1 as input data, and then the values were predicted for each CO site using linear interpolation. Further analysis followed the user manual of MADpatterns available on GitHub (https://github.com/mwhite4/MADpatterns).
Statistical analysis
To assess data distribution, normality was tested using the Shapiro-Wilk test for small sample sizes (n < 50) and the Kolmogorov-Smirnov test for larger sample sizes (n ≥ 50). Statistical analyses comparing the average number of COs per meiosis, SNP density within CO intervals, InDel density within CO intervals, DNA methylation levels, and histone modification levels among the four populations were performed using the Kruskal-Wallis test, followed by Dunn’s post hoc test with Bonferroni correction. Motif enrichment between WT and mop1 was analyzed using binomial tests with Benjamini–Hochberg correction. The correlation between CO density and TE density was evaluated using Spearman’s rank correlation. Differences in the distance at which the CoC value reaches 1 between WT and mop1, for both female and male populations, were assessed using Student’s t test. All statistical analyses were performed in Python using the SciPy, statsmodels, pingouin, and scikit-posthocs packages. Data processing and visualization were conducted using pandas, numpy, matplotlib, seaborn, and IGV88. Statistical significance was defined as P < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The whole genome sequencing data for 423 BC1 individuals, derived from female and male mop1 mutants and their wild-type controls, have been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive under the associate PRJNA1198964. The RNA-seq data for immature anthers from mop1 mutants and their WT siblings are available in the NCBI Gene Expression Omnibus under the accession GSE304481. The genome assembly, along with gene and transposon annotations, was downloaded from MaizeGDB (https:// www.maizegdb.org/). Data are available at Figshare (https://doi.org/10.6084/m9.figshare.30223450). Source data are provided with this paper.
References
Keeney, S. & Neale, M. J. Initiation of meiotic recombination by formation of DNA double-strand breaks: mechanism and regulation. Biochem. Soc. Trans. 34, 523–525 (2006).
Oger, C. & Claeys Bouuaert, C. SPO11 dimers are sufficient to catalyse DNA double-strand breaks in vitro. Nature 639, 792–799 (2025).
Tang, X. et al. In vitro reconstitution of meiotic DNA double-strand-break formation. Nature 639, 800–807 (2025).
Lambing, C., Kuo, P. C., Tock, A. J., Topp, S. D. & Henderson, I. R. ASY1 acts as a dosage-dependent antagonist of telomere-led recombination and mediates crossover interference in Arabidopsis. Proc. Natl. Acad. Sci. USA 117, 13647–13658 (2020).
Pochon, G. et al. The Arabidopsis Hop1 homolog ASY1 mediates cross-over assurance and interference. PNAS Nexus 2, pgac302 (2023).
Armstrong, S. J., Caryl, A. P., Jones, G. H. & Franklin, F. C. Asy1, a protein required for meiotic chromosome synapsis, localizes to axis-associated chromatin in Arabidopsis and Brassica. J. Cell Sci. 115, 3645–3655 (2002).
Ferdous, M. et al. Inter-homolog crossing-over and synapsis in Arabidopsis meiosis are dependent on the chromosome axis protein AtASY3. PLoS Genet. 8, e1002507 (2012).
Chambon, A. et al. Identification of ASYNAPTIC4, a Component of the meiotic chromosome axis. Plant Physiol. 178, 233–246 (2018).
Higgins, J. D., Sanchez-Moran, E., Armstrong, S. J., Jones, G. H. & Franklin, F. C. The Arabidopsis synaptonemal complex protein ZYP1 is required for chromosome synapsis and normal fidelity of crossing over. Genes Dev. 19, 2488–2500 (2005).
Kurzbauer, M. T., Uanschou, C., Chen, D. & Schlogelhofer, P. The recombinases DMC1 and RAD51 are functionally and spatially separated during meiosis in Arabidopsis. Plant Cell 24, 2058–2070 (2012).
Brown, M. S., Grubb, J., Zhang, A., Rust, M. J. & Bishop, D. K. Small Rad51 and Dmc1 complexes Often co-occupy both ends of a meiotic DNA double strand break. PLoS Genet. 11, e1005653 (2015).
Heyer, W. D. Recombination: Holliday junction resolution and crossover formation. Curr. Biol. 14, R56–R58 (2004).
Modliszewski, J. L. et al. Elevated temperature increases meiotic crossover frequency via the interfering (Type I) pathway in Arabidopsis thaliana. PLoS Genet. 14, e1007384 (2018).
Sidhu, G. K. et al. Recombination patterns in maize reveal limits to crossover homeostasis. Proc. Natl. Acad. Sci. USA 112, 15982–15987 (2015).
Jones, G. H. & Franklin, F. C. Meiotic crossing-over: obligation and interference. Cell 126, 246–248 (2006).
Zhang, L., Liang, Z., Hutchinson, J. & Kleckner, N. Crossover patterning by the beam-film model: analysis and implications. PLoS Genet. 10, e1004042 (2014).
Martini, E., Diaz, R. L., Hunter, N. & Keeney, S. Crossover homeostasis in yeast meiosis. Cell 126, 285–295 (2006).
Baudat, F. et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327, 836–840 (2010).
Diagouraga, B. et al. PRDM9 methyltransferase activity is essential for meiotic DNA double-strand break formation at its binding sites. Mol. Cell 69, 853–865 (2018).
Bherer, C., Campbell, C. L. & Auton, A. Refined genetic maps reveal sexual dimorphism in human meiotic recombination at multiple scales. Nat. Commun. 8, 14994 (2017).
Sardell, J. M. & Kirkpatrick, M. Sex differences in the recombination landscape. Am. Nat. 195, 361–379 (2020).
Sardell, J. M. et al. Sex differences in recombination in sticklebacks. G3 8, 1971–1983 (2018).
Choi, K. et al. Arabidopsis meiotic crossover hot spots overlap with H2A.Z nucleosomes at gene promoters. Nat. Genet. 45, 1327–1336 (2013).
Rodgers-Melnick, E. et al. Recombination in diverse maize is stable, predictable, and associated with genetic load. Proc. Natl. Acad. Sci. USA 112, 3823–3828 (2015).
Kianian, P. M. A. et al. High-resolution crossover mapping reveals similarities and differences of male and female recombination in maize. Nat. Commun. 9, 2370 (2018).
Giraut, L. et al. Genome-wide crossover distribution in Arabidopsis thaliana meiosis reveals sex-specific patterns along chromosomes. PLoS Genet. 7, e1002354 (2011).
Phillips, D. et al. The effect of temperature on the male and female recombination landscape of barley. N. Phytol. 208, 421–429 (2015).
Luo, C., Li, X., Zhang, Q. & Yan, J. Single gametophyte sequencing reveals that crossover events differ between sexes in maize. Nat. Commun. 10, 785 (2019).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Matzke, M. A. & Mosher, R. A. RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nat. Rev. Genet. 15, 394–408 (2014).
Kim, J., Kim, J. H., Richards, E. J., Chung, K. M. & Woo, H. R. Arabidopsis VIM proteins regulate epigenetic silencing by modulating DNA methylation and histone modification in cooperation with MET1. Mol. Plant 7, 1470–1485 (2014).
Law, J. A. et al. Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1. Nature 498, 385–389 (2013).
Du, J. et al. Dual binding of chromomethylase domains to H3K9me2-containing nucleosomes directs DNA methylation in plants. Cell 151, 167–180 (2012).
Stroud, H. et al. Non-CG methylation patterns shape the epigenetic landscape in Arabidopsis. Nat. Struct. Mol. Biol. 21, 64–72 (2014).
To, T. K. et al. RNA interference-independent reprogramming of DNA methylation in Arabidopsis. Nat. Plants 6, 1455–1467 (2020).
Melamed-Bessudo, C. & Levy, A. A. Deficiency in DNA methylation increases meiotic crossover rates in euchromatic but not in heterochromatic regions in Arabidopsis. Proc. Natl. Acad. Sci. USA 109, E981–E988 (2012).
Mirouze, M. et al. Loss of DNA methylation affects the recombination landscape in Arabidopsis. Proc. Natl. Acad. Sci. USA 109, 5880–5885 (2012).
Yelina, N. E. et al. DNA methylation epigenetically silences crossover hot spots and controls chromosomal domains of meiotic recombination in Arabidopsis. Genes Dev. 29, 2183–2202 (2015).
Underwood, C. J. et al. Epigenetic activation of meiotic recombination near Arabidopsis thaliana centromeres via loss of H3K9me2 and non-CG DNA methylation. Genome Res. 28, 519–531 (2018).
Li, Q. et al. Genetic perturbation of the maize methylome. Plant Cell 26, 4602–4616 (2014).
Liu, B. et al. Heritable changes of epialleles near genes in maize can be triggered in the absence of CHH methylation. Plant Physiol. 194, 2511–2532 (2024).
Zhao, M. et al. The mop1 mutation affects the recombination landscape in maize. Proc. Natl. Acad. Sci. USA 118 (2021).
Alleman, M. et al. An RNA-dependent RNA polymerase is required for paramutation in maize. Nature 442, 295–298 (2006).
Nobuta, K. et al. Distinct size distribution of endogeneous siRNAs in maize: Evidence from deep sequencing in the mop1-1 mutant. Proc. Natl. Acad. Sci. USA 105, 14958–14963 (2008).
Li, Q. et al. RNA-directed DNA methylation enforces boundaries between heterochromatin and euchromatin in the maize genome. Proc. Natl. Acad. Sci. USA 112, 14728–14733 (2015).
Marand, A. P. et al. Meiotic crossovers are associated with open chromatin and enriched with Stowaway transposons in potato. Genome Biol. 18, 203 (2017).
Fuentes, R. R. et al. A catalogue of recombination coldspots in interspecific tomato hybrids. PLoS Genet. 20, e1011336 (2024).
Rezvoy, C., Charif, D., Gueguen, L. & Marais, G. A. MareyMap: an R-based tool with graphical interface for estimating recombination rates. Bioinformatics 23, 2188–2189 (2007).
Bailey, T. L. STREME: accurate and versatile sequence motif discovery. Bioinformatics 37, 2834–2840 (2021).
Darrier, B. et al. High-resolution mapping of crossover events in the hexaploid wheat genome suggests a universal recombination mechanism. Genetics 206, 1373–1388 (2017).
Marand, A. P. et al. Historical meiotic crossover hotspots fueled patterns of evolutionary divergence in rice. Plant Cell 31, 645–662 (2019).
Kaplan, N. et al. The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 458, 362–366 (2009).
Struhl, K. & Segal, E. Determinants of nucleosome positioning. Nat. Struct. Mol. Biol. 20, 267–273 (2013).
Szymanska-Lejman, M. et al. The effect of DNA polymorphisms and natural variation on crossover hotspot activity in Arabidopsis hybrids. Nat. Commun. 14, 33 (2023).
Dooner, H. K. & He, L. Maize genome structure variation: interplay between retrotransposon polymorphisms and genic recombination. Plant Cell 20, 249–258 (2008).
Ziolkowski, P. A. & Henderson, I. R. Interconnections between meiotic recombination and sequence polymorphism in plant genomes. N. Phytol. 213, 1022–1029 (2017).
Liu, B. & Zhao, M. How transposable elements are recognized and epigenetically silenced in plants?. Curr. Opin. Plant Biol. 75, 102428 (2023).
Ricci, W. A. et al. Widespread long-range cis-regulatory elements in the maize genome. Nat. Plants 5, 1237–1249 (2019).
Capilla-Perez, L. et al. The synaptonemal complex imposes crossover interference and heterochiasmy in Arabidopsis. Proc. Natl. Acad. Sci. USA 118 (2021).
France, M. G. et al. ZYP1 is required for obligate cross-over formation and cross-over interference in Arabidopsis. Proc. Natl. Acad. Sci. USA 118 (2021).
Chelysheva, L. et al. Zip4/Spo22 is required for class I CO formation but not for synapsis completion in Arabidopsis thaliana. PLoS Genet. 3, e83 (2007).
Gent, J. I. et al. CHH islands: de novo DNA methylation in near-gene chromatin regulation in maize. Genome Res. 23, 628–637 (2013).
Bureau, T. E. & Wessler, S. R. Stowaway: a new family of inverted repeat elements associated with the genes of both monocotyledonous and dicotyledonous plants. Plant Cell 6, 907–916 (1994).
Feschotte, C., Swamy, L. & Wessler, S. R. Genome-wide analysis of mariner-like transposable elements in rice reveals complex relationships with stowaway miniature inverted repeat transposable elements (MITEs). Genetics 163, 747–758 (2003).
Zhang, X., Jiang, N., Feschotte, C. & Wessler, S. R. PIF- and Pong-like transposable elements: distribution, evolution and relationship with Tourist-like miniature inverted-repeat transposable elements. Genetics 166, 971–986 (2004).
Blackwell, A. R. et al. MSH2 shapes the meiotic crossover landscape in relation to interhomolog polymorphism in Arabidopsis. EMBO J. 39, e104858 (2020).
Roselius, K., Stephan, W. & Stadler, T. The relationship of nucleotide polymorphism, recombination rate and selection in wild tomato species. Genetics 171, 753–763 (2005).
Dluzewska, J. et al. MSH2 stimulates interfering and inhibits non-interfering crossovers in response to genetic polymorphism. Nat. Commun. 14, 6716 (2023).
Cahoon, C. K. & Libuda, D. E. Leagues of their own: sexually dimorphic features of meiotic prophase I. Chromosoma 128, 199–214 (2019).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Hufford, M. B. et al. De novo assembly, annotation, and comparative analysis of 26 diverse maize genomes. Science 373, 655–662 (2021).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Jia, S. & Shi, L. Efficient change-points detection for genomic sequences via cumulative segmented regression. Bioinformatics 38, 311–317 (2022).
Andolfatto, P. et al. Multiplexed shotgun genotyping for rapid and efficient genetic mapping. Genome Res. 21, 610–617 (2011).
Bradbury, P. J. et al. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23, 2633–2635 (2007).
Broman, K. W., Wu, H., Sen, S. & Churchill, G. A. R/qtl: QTL mapping in experimental crosses. Bioinformatics 19, 889–890 (2003).
Taylor, J. & Butler, D. R Package ASMap: Efficient genetic linkage map construction and diagnosis. J. Stat. Softw. 79, 1–29 (2017).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Juhling, F. et al. metilene: fast and sensitive calling of differentially methylated regions from bisulfite sequencing data. Genome Res. 26, 256–262 (2016).
Begcy, K. & Dresselhaus, T. Tracking maize pollen development by the Leaf Collar Method. Plant Reprod. 30, 171–178 (2017).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Putri, G. H., Anders, S., Pyl, P. T., Pimanda, J. E. & Zanini, F. Analysing high-throughput sequencing data in Python with HTSeq 2.0. Bioinformatics 38, 2943–2945 (2022).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
White, M. A., Wang, S., Zhang, L. & Kleckner, N. Quantitative modeling and automated analysis of meiotic recombination. Methods Mol. Biol. 1471, 305–323 (2017).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 14, 178–192 (2013).
Acknowledgements
We thank the University of Florida HiPerGator Supercomputer for providing computational resources for this analysis. We are grateful to Dr. Gwonjin Lee and Akwasi Yeboah for their technical assistance with CO interference analysis and for valuable discussions on the entire project. We also thank Dr. Jin Zhang from the Department of Statistics at Miami University for her consultation on the statistical analysis. This work was supported by the National Institute of General Medical Sciences of the National Institutes of Health under Award Number R15GM135874, the National Science Foundation under Award Number IOS2306220, startup funds from Miami University and the University of Florida, and the University of Florida Archer Early Career Seed Grant awarded to M.Z.
Author information
Authors and Affiliations
Contributions
M.Z. conceived and designed the experiments and generated the mapping populations. M.M.H. and L.Y. conducted plant genotyping. M.M.H., L.Y., M.W., and M.Z. analyzed the data. M.M.H. and M.Z. prepared the original draft, and W.P.P. provided critical review and edits to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kyuha Choi, Natasha Yelina, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hasan, M.M., Yin, L., Wang, M. et al. mop1 affects maize recombination landscapes by modulating methylation of MITEs near genes in open chromatin. Nat Commun 16, 10476 (2025). https://doi.org/10.1038/s41467-025-65469-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65469-6