Abstract
Animals sense their metabolic needs to guide adaptive behaviors partly through serotonin, a neurotransmitter associated with feeding in many species. Here we investigate the ability of the serotonin system to evaluate and interpret diverse diets by studying long-term foraging behaviors of the nematode C. elegans on bacteria. Behavioral screens on a genome-wide collection of E coli strains identified 22 metabolic mutants that induce behavioral aversion and stress responses in C. elegans. We show that different classes of serotonergic neurons promote aversion to non-preferred E. coli diets and retention on preferred E. coli diets, respectively, through different serotonin receptors. Serotonin is integrated with dopamine and octopamine signals across distributed circuits to direct opposing behavioral responses to preferred and aversive diets. These results reveal interacting neuromodulatory circuits that guide context-dependent evaluation of dietary quality.
Similar content being viewed by others
Introduction
Animals discriminate between high-quality and low-quality food to optimize their development, reproduction, and physiology. While food quality can be sensed through olfactory and gustatory cues, internally-sensed metabolic needs also drive food preferences. For example, an interoceptive dopaminergic circuit allows Drosophila to reject an incomplete diet lacking essential amino acids1. Preferences can also change based on an animal’s physiology: newly eclosed female mosquitos feed on nectar, but after mating they prefer blood meals that support the growth of offspring2,3. The full extent and nature of metabolic factors that influence food preference in animals are not known4,5,6.
Many behaviors of the nematode Caenorhabditis elegans are regulated by its bacterial food. In addition to rapid chemotaxis to food-related metabolites and food ingestion through its pharynx7, it has longer-term foraging behaviors that reflect its evaluation of food quality and quantity over minutes to hours8. For example, animals display low-activity dwelling behaviors in high-quality food, and higher-activity roaming behaviors when food is limited9,10,11,12. At the extreme, animals will abandon a bacterial food source that is hard to eat, toxic, or pathogenic9,13,14,15,16,17,18. In these aversive conditions, C. elegans will initially enter the bacterial lawn, but after several hours begins to exit the lawn and accumulate in food-free regions. The animals also learn to avoid the odor of toxic bacteria in olfactory choice assays19,20. These slow acquired aversion behaviors can extend survival by limiting pathogen ingestion and toxin exposure, and may allow animals to select high-quality food sources in their heterogeneous natural environments. They are under extensive neuromodulatory regulation by the biogenic amines serotonin11,21,22, dopamine23,24, tyramine25,26, and octopamine27, as well as the neuroendocrine cytokine DAF-718,28, and the neuropeptides PDF-1 and FLP-111,13,16, suggesting that long-term behaviors are guided by slow-acting neuromodulatory systems.
Serotonin links physiology and behavior in many animals. In mammals, serotonin is synthesized by neurons in the brainstem and by enteric neurons and enteroendocrine cells in the digestive system; it regulates locomotion, anxiety, and food responses including appetite, satiety, obesity, gut motility, and nausea29,30. Similarly, the serotonergic system in C. elegans is implicated in a variety of food-related behaviors. The primary sources of serotonin are three pairs of neurons called NSM, ADF, and HSN. The NSM enteric neurons reside in the pharynx, detect food through proprioceptive endings, and release serotonin to stimulate feeding, locomotor slowing, and dwelling behaviors31; NSM also has a key role in acquired aversion to bacteria spiked with mitochondrial toxins32. The ADF sensory neurons detect bacterial metabolites and regulate both acute feeding behaviors and learned avoidance of bacterial pathogens20,33. The HSN neurons reside in the midbody and regulate egg-laying, locomotion, and dwelling behaviors34. Serotonin acts on six receptors with different signaling mechanisms that are collectively expressed on nearly half of all C. elegans neurons; several receptors contribute to each serotonin-regulated behavior11,35. The general organization of serotonin systems with a few serotonin-expressing neurons, multiple serotonin receptors, and widely distributed serotonin receptor-expressing neurons appears to be shared between mammals and C. elegans. As serotonin and other neuromodulators act extrasynaptically, these distributed systems create a challenge for circuit mapping. Understanding information flow requires detailed identification of the functional connections between serotonin- and serotonin-receptor expressing neurons in specific behavioral contexts35,36,37.
Caenorhabditis elegans feeds on a variety of bacteria in its natural environment38, but is typically cultivated on E. coli in the laboratory39. The influences of diet on C. elegans growth and physiology have been studied using classical methods40,41,42,43,44 and extended using the 3983 E. coli gene knockout strains in the Keio collection45, identifying E. coli genes required for rapid C. elegans development46, longevity47, survival under mitochondrial stress48, and resistance to the toxins 5-Fluoruracil, FUDR, camptothecin, and Aflatoxin B149,50,51. Here, we use the Keio E. coli collection to systematically identify bacterial diets that elicit aversion behavior in C. elegans, which we call “mediocre” diets based on this property. Pathway enrichment of these E. coli mutants suggests that C. elegans monitors dietary quality using a few primary bacterial signals: global bacterial metabolism (crp), cysteine biosynthesis, vitamin B6 biosynthesis, ferric iron transport, and bacterial membrane integrity. Using C. elegans genetics and circuit mapping, we find that serotonin has a dual role in assessing dietary quality: it promotes aversion from a mediocre diet and suppresses aversion from a high-quality diet through context-dependent roles in serotonin-producing neurons and multiple serotonin receptors. Serotonin acts together with dopamine to modulate octopaminergic and tyraminergic neurons, and serotonin also acts on other neuronal targets to regulate behavior. Our results reveal a distributed neuromodulatory network of biogenic amines, receptors, and target neurons that interpret dietary quality through cooperative and antagonistic effects on behavior.
Results
A genome-wide screen for E. coli mutants that elicit aversion behavior in C. elegans
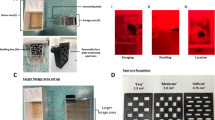
We evaluated C. elegans behavior using a long-term foraging assay that has been widely used to study responses to toxic or pathogenic bacteria15,16,18,19,52. 15–20 C. elegans L4 larvae are placed on a plate seeded with a dense patch of bacterial food and the location of the animals is tracked by video recording for 20 h (Fig. 1a). Over 80% of animals remain within a patch of the standard E. coli bacterial food throughout a 20 h assay, whereas fewer than 10% of animals remain on patches of pathogenic Pseudomonas aeruginosa bacteria after 16–20 h (Fig. 1b). An aversion ratio is defined as the steady-state fraction of animals off the food patch 16–20 h after the beginning of the assay (Fig. 1c).
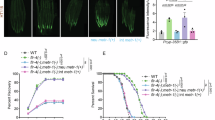
a Schematic depiction of aversion behavior screen. b Behavioral responses of wild-type C. elegans to different E. coli strains and to the pathogenic bacteria Pseudomonas aeruginosa PA14. Each trace represents the average of at least 8 individual assays per strain across 20 h. c Comparison of pooled behavioral responses of wild-type C. elegans on BW25113 (WT) and the 22 Keio mutants identified from the whole-genome screen. The mean values for each group are shown. Black dashed line indicates mean value from WT group (n = 74) and red dashed line indicates mean value from pooled positive hits (n = 229). d Simplified depiction of identified metabolic pathways in E. coli, full aversion time course for one mutant from each metabolic pathway, and averaged aversion ratios at 16–20 h. Complete metabolic pathways and additional aversion assays are shown in Supplementary Fig. 1; bacterial growth properties are reported in Supplementary Data 1. Genes identified in this study are highlighted in color. cAMP, cyclic AMP. CRP, cAMP Receptor Protein. ECA, Enterobacterial Common Antigen. PLP, Pyridoxal 5’-phosphate. Ent, Enterobactin. IM, Inner Membrane. OM, Outer Membrane. Each time course trace shows the mean of 9-15 biological replicates; shaded region represents the 95% confidence interval. Each data point in the box plot represents an individual assay (n = 9–15), with median ± quartiles in boxes and Min to Max whiskers. ***P < 0.001 by one-way ANOVA with Dunnett’s multiple correction. All sample sizes, statistical tests used, and exact P values are provided in Supplementary Data 4. Source data are provided as a Source Data file.
To perform a systematic investigation of bacterial effects on C. elegans aversion behavior, we screened the E. coli Keio knockout collection45 of 3983 deletion mutants. Following a prescreen that yielded 262 potentially aversive strains, quantitative video recording identified 22 E. coli mutants that elicited significant aversion behavior (Table 1), with an average aversion ratio of 0.4 compared to 0.18 for the parental BW25113 strain (henceforth, wild-type) (Fig. 1c). These 22 genes included multiple hits in five metabolic pathways: regulation of global transcription, biosynthesis of the enterobacterial common antigen (ECA), biosynthesis of the essential nutrients cysteine and vitamin B6, and ferric iron transport (Fig. 1d, and Supplementary Fig. 1). Aversion ratios for representative genes from each pathway are show in Fig. 1d; the enriched pathways are described briefly below.
In response to reduced availability of a preferred carbon source such as glucose, the E. coli adenylate cyclase CyaA catalyzes the production of cyclic AMP (cAMP), which is recognized and bound by CRP, the cAMP receptor protein transcription factor53,54,55,56. ∆crp and ∆cyaA, but not other mutants in this pathway, elicited significant aversion in C. elegans (Fig. 1d, Supplementary Fig. 1a).
E. coli expresses enterobacterial common antigen (ECA) on the outer membrane, where it regulates bacterial cell shape and stress responses. Four ECA biosynthetic mutants, ∆wzxE, ∆rffT, ∆rffA and ∆rffC, elicited aversive responses in C. elegans. The ECA flippase WzxE flips the resulting ECA unit across the inner membrane prior to ECA maturation57 (Fig. 1d). No aversion was induced by mutants in earlier or later steps of ECA synthesis, suggesting that accumulation of an inner membrane intermediate, rather than the loss of ECA, drives C. elegans aversion behavior (Supplementary Fig. 1b).
In E. coli, cysteine is synthesized from serine through the serine acetyltransferase CysE, which converts L-serine to O-acetyl-L-serine (OAS), and the O-acetylserine sulfhydrylase CysK. CysE and CysK form a complex that allosterically activates serine to cysteine conversion58 (Fig. 1d, and Supplementary Fig. 1c). ∆cysE and ∆cysK bacteria were moderately aversive to C. elegans. Cysteine is not an essential amino acid for C. elegans growth40, but cysteine limitation reduces synthesis of the protective redox metabolite glutathione59,60.
Four E. coli mutants affecting vitamin B6 synthesis elicited C. elegans aversion behavior (Fig. 1d, and Supplementary Fig. 1d). Vitamin B6 is an essential nutrient for C. elegans40 and is required for synthesis of serotonin, dopamine, tyramine and octopamine transmitters61. In a possible overlap with other aversive bacteria, vitamin B6 is required for bacterial cysteine synthesis (cysK) and ECA synthesis (rffA/wecE)62. E. coli vitamin B6 biosynthetic genes are essential for C. elegans growth in certain nutrient-limited conditions63 and can confer C. elegans resistance to the toxin 5-fluorouracil49.
Fifteen enzymes support E. coli ferric iron uptake and release through the small-molecule siderophore enterobactin. The four mutants that elicited aversion in C. elegans, ∆fepB, ∆fepD, ∆fepG, and ∆fes (Fig. 1d, and Supplementary Fig. 1e) accumulate a ferric iron-enterobactin complex in the E. coli cytoplasm or periplasm that has been proposed to be toxic to C. elegans mitochondria46,48. These mutants also elicit production of reactive oxygen species (ROS), but other ROS-producing mutants did not elicit aversion (Supplementary Fig. 1e, f).
Individual mutants that can drive C. elegans aversion include ∆fabH (fatty acid biosynthesis), ∆tatB (twin arginine translocation component, ∆ynhG (L,D-transpeptidase), ∆alaS (alanine-tRNA ligase), ∆aceF (pyruvate dehydrogenase E2 subunit), and ∆dnaJ (a chaperone protein) (Supplementary Fig. 1g, and Table 1).
In summary, C. elegans shows aversion to several E. coli diets with altered metabolic properties. The relatively small number of genes and pathways represented suggests that aversion is an active, relatively specific behavior; in particular, over 200 of the E. coli deletion mutants that result in slow C. elegans growth did not elicit aversion in this assay40.
E. coli mutants that elicit C. elegans aversion induce stress responses
We selected one E. coli mutant from each of the five biological pathways (∆crp, ∆wzxE, ∆pdxJ, ∆cysE and ∆fepB) to characterize in more detail. Each selected E. coli mutant was tested for effects on C. elegans developmental rate, brood size, feeding rate, colonization (a marker of bacterial pathogenesis), and lifespan (Supplementary Fig. 2a). The strongest defects were observed on ∆fepB, which caused substantial developmental delay, reduced brood size, and increased colonization of C. elegans, in agreement with the toxicity of this strain observed in previous screens46. ∆crp caused reduced lifespan, reduced feeding, and a slight developmental delay; ∆wzxE also caused reduced feeding and a slight developmental delay; ∆pdxJ caused a reduced brood size (Supplementary Fig. 2a). These results indicate that aversive bacterial strains can affect C. elegans physiology.
To refine this insight, we examined the effects of bacterial mutants on C. elegans molecular reporters of physiological stress14. Expression of a daf-7::GFP reporter in ASJ sensory neurons is induced by bacterial pathogens and by food that is physically difficult for C. elegans to ingest18,28. This reporter gene was significantly induced by ∆crp, ∆wzxE, and ∆fepB bacteria, but not by ∆cysE or ∆pdxJ bacteria (Fig. 2a). An hsp-6::GFP reporter for mitochondrial stress was also significantly induced by ∆crp and ∆fepB bacteria46 (Fig. 2b). A gst-4::GFP reporter for oxidative stress was induced by ∆cysE and ∆wzxE bacteria (Fig. 2c), but not by ∆crp, ∆fepB, or ∆pdxJ bacteria. These results suggest that aversive diets can selectively activate C. elegans stress responses. Several other C. elegans stress reporters were also induced by one or more of the aversive diets (Supplementary Fig. 2b-f).
a–c Effects of mediocre bacterial diets on C. elegans stress reporters. a Quantification of daf-7::gfp marker in ASJ sensory neurons. b Quantification of the mitochondrial UPR marker hsp-6::gfp. c Quantification of the oxidative stress marker gst-4::gfp. GFP fluorescence intensities quantified as Arbitrary Units (A.U.). Each data point indicates fluorescence from an individual animal (n = 20-45). Error bars indicate mean ± SD. ns, not significant, **P < 0.01, ***P < 0.001 by one-way ANOVA corrected by Dunnett’s multiple comparisons. d Effects of chemical supplementation on aversion behavior to wild-type, ∆crp, or ∆cysE bacteria. Top, schematic diagram of chemical supplementation. Pre-adding indicates chemicals provided during bacteria growth; post-adding indicates chemicals supplied one hour before the behavioral assay. Final concentrations in assay plates were 0.4% glucose, 200 μM cysteine (pre-adding) or 50 μM cysteine (post-adding), and 200 μM OAS. Each data point indicates individual assay (n = 8–18 assays). Results are shown with median ± quartiles in boxes and Min to Max whiskers. ns, not significant, *P < 0.05, **P < 0.01, ***P < 0.001 by two-tailed, unpaired t test or one-way ANOVA corrected by Dunnett’s multiple comparisons. e Aversion behaviors at different bacterial densities. Wild-type, ∆crp, or ∆cysE were seeded at different densities and CFU from control plates were counted at the beginning of the behavioral assay for most assays; for pre- and post- added cysteine, CFU from control plates were counted at the end of the assay. Each data point indicates individual assay (n = 10-36). Results are shown with median ± quartiles in boxes and Min to Max whiskers. ns, not significant, *P < 0.05, ***P < 0.001 by one-way ANOVA corrected by Dunnett’s multiple comparisons. Additional stress reporters are shown in Supplementary Fig. 2, and additional assays and bacterial counts are shown in Supplementary Fig. 3. All sample sizes, statistical tests used, and exact P values are provided in Supplementary Data 4. Source data are provided as a Source Data file.
The metabolic effects of the bacterial mutants were confirmed by chemical supplementation either prior to bacterial growth (pre-adding) or immediately before the aversion assay (post-adding) (Fig. 2d). Addition of glucose to the assay plates during bacterial growth, which should bypass the metabolic function of CRP, fully suppressed C. elegans aversion to the ∆crp diet (Fig. 2d). Addition of cysteine or its precursor O-acetyl-serine either before or after bacterial growth suppressed C. elegans aversion to the ∆cysE diet, consistent with a cysteine-related dietary deficiency (Fig. 2d). Addition of vitamin B6 to ∆pdxJ and related bacterial mutants suppressed C. elegans aversion if the vitamin was provided before bacterial growth, but did not suppress aversion if only provided during the assay (Supplementary Fig. 2g). This pattern suggests that aversion results in part from altered bacterial metabolism49.
Foraging in C. elegans is sensitive to bacterial density64,65. To control for effects of differential bacterial growth, we measured and varied the density of wild-type, ∆crp, and ∆cysE bacterial strains and assessed effects on C. elegans behavioral responses (Fig. 2e and Supplementary Fig. 3). Attraction to wild-type bacteria was stable across a 10-fold range of cell densities. Aversion to Δcrp was observed across cell densities, consistent with a stress response. Aversion to ΔcysE was rescued at high bacterial densities, and was also rescued by acutely supplementing low ΔcysE bacterial densities with cysteine (Fig. 2e and Supplementary Fig. 3). These results are consistent with a cysteine-related metabolic deficit.
The results of these experiments suggest that at least two different bacterial qualities can drive C. elegans aversion: a metabolic change that induces mitochondrial stress (∆crp, ∆fepB) and a metabolic change that induces oxidative stress (∆cysE); both processes may contribute to aversion from ∆wzxE. ∆crp and ∆cysE were used as representative examples of aversive bacteria of two different classes. We refer to these as “mediocre” diets based on their aversive qualities and their activation of C. elegans stress reporters.
C. elegans evaluates opposing food qualities with distinct classes of serotonergic neurons
To ask how C. elegans distinguishes between bacteria in the aversion assay, we began with serotonin, a neuromodulator that regulates multiple food-related behaviors as well as avoidance of pathogenic bacteria11,14,20,32,66. Animals mutant for the tryptophan hydroxylase gene tph-1 lack all neuronally-synthesized serotonin67,68. We found that two independent tph-1 mutants had reduced aversion to both ∆crp and ∆cysE mediocre diets, while retaining apparently normal responses to wild-type bacteria (Fig. 3a and Supplementary Fig. 4a).
a Aversion behaviors of tph-1 null mutants and neuron-specific knockout mutants on three diets. tph-1 is expressed in three major serotonergic neuron classes, ADF, NSM and HSN, and encodes a tryptophan hydroxylase that catalyzes the first step in serotonin biosynthesis. Cell-specific knockout of a single-copy genomic tph-1 insertion was achieved by Cre-lox recombination11 (n = 9–15 assays). All figures show the tph-1(mg280) null allele, other than Supplementary Fig. 4, which shows similar aversion behaviors in the tph-1(n4622) null allele. b ADF silencing with a HisCl transgene and histamine induces aversion to wild-type E. coli BW25113 (n = 9 assays). c NSM silencing with a HisCl transgene and histamine suppresses aversion to ∆crp and ∆cysE diets (n = 9–11 assays). d HSN-deficient egl-1(gain-of-function) mutants show decreased aversion to the ∆crp diet (n = 9–12 assays). e Schematic model of serotonergic neuron functions. For all panels, each data point indicates individual assay. Results are shown with median ± quartiles in boxes and Min to Max whiskers. ns, not significant, *P < 0.05, **P < 0.01, ***P < 0.001 by two-tailed, unpaired t test (panels a, c, d) or one-way ANOVA corrected by Dunnett’s multiple comparisons (b). Full genotypes of all C. elegans strains are provided in Supplementary Data 3. All sample sizes, statistical tests used, and exact P values are provided in Supplementary Data 4. Source data are provided as a Source Data file.
Three classes of tph-1-expressing neurons synthesize serotonin in C. elegans hermaphrodites. To ask which neurons regulate aversion, we eliminated tph-1 in individual neuron classes using Cre/lox recombination25, and tested the resulting animals on wild-type, ∆crp and ∆cysE diets. Remarkably, each of the three serotonin-producing neurons affected aversion behaviors (Fig. 3a). Depletion of serotonin synthesis in ADF neurons resulted in a small but significant increase in aversion from wild-type bacteria; depletion of serotonin synthesis in NSM neurons decreased aversion from both ∆crp and ∆cysE mediocre diets; and depletion of serotonin synthesis in HSN neurons decreased aversion from the ∆crp diet. These results suggest that serotonin can either increase or decrease aversion behavior, depending on the neuronal serotonin source and the dietary context.
To validate these results with an orthogonal approach, we manipulated individual serotonergic neurons and tested the resulting strains on the three bacterial diets. Acute chemogenetic silencing of ADF neurons using a histamine-gated chloride channel (HisCl1) transgene69 and exogenous histamine significantly increased aversion from wild-type bacteria (Fig. 3b), resembling the tph-1 knockout in ADF. Acute chemogenetic silencing of NSM neurons suppressed aversion to the mediocre ∆crp and ∆cysE diets, recapitulating the effects of the tph-1 knockout in NSM (Fig. 3c). Finally, a strain in which the HSN neurons were genetically ablated (egl-1(gf)) had diminished aversion to the mediocre ∆crp diet, resembling animals in which HSN did not synthesize serotonin (Fig. 3d). These results confirm that serotonergic ADF neurons prevent aversion from wild-type diets, whereas serotonergic NSM and to a lesser extent HSN neurons promote aversion from mediocre ∆crp and ∆cysE diets (Fig. 3e).
The serotonin receptor SER-5 suppresses aversion from a wild-type diet together with the biogenic amine octopamine
C. elegans has six known serotonin receptors: two serotonin-gated ion channels (MOD-1, a chloride channel, and LGC-50, a cation channel), and four G-protein coupled receptors (SER-5 (Gs-coupled), SER-7 (Gs-coupled), SER-1 (Gq-coupled) and SER-4 (Gi/o-coupled)) (Fig. 4a)67. Collectively, these receptors are expressed in almost half of all neuronal classes, and typically act cooperatively in foraging and feeding behaviors35. We tested mutants for each serotonin receptor for aversion on the wild-type diet as well as mediocre ∆crp and ∆cysE diets; for clarity, the results are discussed separately on each diet below.
a Schematic diagram of six serotonin receptors in C. elegans, with relevant G-protein for GPCRs indicated. Serotonin receptor mutations are indicated for each gene. b Behavioral responses of wild-type, tph-1, and serotonin receptor mutant animals on the wild-type BW25113 diet. ser-4 and ser-5 mutants showed significant aversion (n = 9–12 assays). c Aversion behavior of ser-5 mutants expressing ser-5 cDNA from its endogenous promoter (ser-5) or promoters for RIM and RIC (tdc-1), RIC (tbh-1), or enteric neurons of the pharynx (glr-7). Two transgenic lines were tested for each rescue plasmid (n = 10–17 assays). d Aversion behavior of tdc-1 and tbh-1 null mutants on BW25113 (n = 11–12 assays). TDC-1 tyrosine decarboxylase is expressed in RIM and RIC neurons and uv1 neuroendocrine cells, and converts tyrosine to tyramine (TA). TBH-1 tyramine b-hydroxylase converts tyramine to octopamine (OA) in RIC neurons and gonadal sheath cells. e Expression of the tetanus toxin light chain in RIM and/or RIC neurons to block synaptic release induced aversion to BW25113 (n = 14–17 assays). f Rescue of tdc-1 using an inverted Cre-lox strategy. Cre expression was driven by the rimb-1 promoter (intersection in RIM, RIC, and uv1 cells), or dmsr-7 promoter (intersection in RIM and RIC neurons) (n = 12-14 assays). g Rescue of tdc-1 mutants by expression of tdc-1 cDNA from the tdc-1 promoter (RIM, RIC and uv1 cells) or the tbh-1 promoter (RIC and gonadal sheath cells) (n = 9–12 assays). h Schematic model of SER-5 function. For b–g each data point indicates individual assay. Results are shown with median ± quartiles in boxes and Min to Max whiskers. ns, not significant, *P < 0.05, **P < 0.01, ***P < 0.001 by one-way ANOVA corrected by Dunnett’s (b–e) or Tukey’s (f, g) multiple comparisons. Full genotypes of all C. elegans strains are provided in Supplementary Data 3. All sample sizes, statistical tests used, and exact P values are provided in Supplementary Data 4. Source data are provided as a Source Data file.
Two of the six serotonin receptor mutants, ser-4 and ser-5, showed strikingly enhanced aversion to the wild-type diet, while the mutants in the other four receptor genes were normal (Fig. 4b). Previous studies suggested that serotonin release from ADF neurons activates the excitatory SER-5 receptor in a variety of neurons to regulate sensory responses, food intake, and longevity70,71,72. ser-5 is expressed in RIC neurons35,73, and serotonin released from ADF can activate RIC74,75. We found that a ser-5 cDNA rescued behavior on the wild-type diet when expressed from its endogenous promoter or from promoters for tdc-1 (expressed in RIM and RIC neurons) or tbh-1 (expressed in RIC neurons) (Fig. 4c). Expression in ser-5-expressing enteric neurons did not rescue behavior on the wild-type diet (Fig. 4c).
The RIM and RIC neurons express tyrosine decarboxylase (tdc-1) to produce the neurotransmitter tyramine, and RIC also expresses tyramine beta-hydroxylase (tbh-1) that converts tyramine into octopamine. tdc-1 mutants lack both tyramine and octopamine, whereas tbh-1 mutants lack octopamine and accumulate excess tyramine33. We examined the behaviors of tdc-1 and tbh-1 mutant animals on wild-type diets. Like ser-5 mutant animals, both tdc-1 and tbh-1 mutants had substantial aversion from the wild-type diet (Figs. 4d, 7g). Expressing the tetanus toxin light chain to block synaptic release from RIC neurons or from RIM and RIC neurons also induced aversion (Fig. 4e). Normal behavior was restored to tdc-1 mutants by restoring expression in RIC neurons or in RIC and RIM neurons (Fig. 4f, g).
Together, these results suggest that serotonin from ADF acts on SER-5 to enhance the activity of tyramine and octopamine-producing cells that suppress aversion from a high-quality diet (Fig. 4h). The combination of strong ser-5 rescue in RIC alone (Fig. 4c) and equivalent levels of aversion induced by tbh-1 and tdc-1 mutations (Fig. 4d) suggest that octopamine from RIC neurons is the primary signal that suppresses aversion from wild-type bacteria (Fig. 4h) but tyramine may also have a role (see below).
The serotonin receptor SER-7 and dopamine promote aversion from a mediocre ∆crp diet
The serotonin receptor mutants ser-7, mod-1, and lgc-50 had reduced aversion from the mediocre ∆crp diet (Fig. 5a). Previous studies showed that the excitatory SER-7 receptor acts in enteric neurons of the pharynx to regulate feeding10. We found that a ser-7 cDNA rescued aversion on the mediocre ∆crp diet when expressed from its endogenous promoter or a flp-21 promoter that is expressed in overlapping enteric neurons (Fig. 5b)10. Among these neurons, ser-7 expression in the I1 enteric neuron partly rescued aversion from the mediocre ∆crp diet, but expression in several other feeding-related neurons did not rescue (Fig. 5c). These results suggest that the I1 neuron, likely responding to serotonin released from the NSM neurons, is part of an enteric circuit that promotes aversion from the mediocre ∆crp diet (Fig. 5d). ser-7 acts in the MC and M4 enteric neurons to promote feeding or recognition of familiar food10,21, but expression in these neurons did not rescue aversion to the mediocre ∆crp diet, nor did expression in I1 neuron rescue ser-7 feeding, indicating that aversion and feeding are separable functions of ser-7 (Fig. 5c, Supplementary Fig. 4b). Additional ser-7, mod-1, and lgc-50 -expressing neurons that affect aversion from ∆crp remain to be identified.
a Behavioral responses of wild-type, tph-1, and serotonin receptor mutant animals on the mediocre ∆crp diet. Aversion is diminished in two ser-7 null mutants and in mod-1 and lgc-50 mutants (n = 8–24 assays). The ser-7(tm1325) allele was used in all subsequent experiments. b Aversion behavior of ser-7 mutants expressing ser-7 cDNA from its endogenous promoter (ser-7), or a promoter for overlapping neurons (flp-21). Two transgenic lines were tested for each plasmid (n = 9–27 assays). c Aversion behavior of ser-7 mutants with ser-7 rescue in the I1 neuron and other sets of enteric neurons. Two transgenic lines were tested for each rescue plasmid (n = 9–73 assays). d Schematic model of SER-7 serotonin signaling in aversion from the mediocre ∆crp diet. For (a–c), each data point indicates individual assay. Results are shown with median ± quartiles in boxes and Min to Max whiskers. ns, not significant, *P < 0.05, **P < 0.01, ***P < 0.001 by one-way ANOVA corrected by Dunnett’s (a, b) or Tukey’s (c) multiple comparisons. Full genotypes of all C. elegans strains are provided in Supplementary Data 3. All sample sizes, statistical tests used, and exact P values are provided in Supplementary Data 4. Source data are provided as a Source Data file.
Further insight into aversion from the ∆crp diet came from examination of the neurotransmitter dopamine. Animals mutant for the dopamine biosynthetic enzyme cat-2 (tyrosine-3-monooxygenase) had diminished aversion to the ∆crp mediocre diet (Fig. 6a). cat-2 is expressed in three classes of ciliated sensory neurons, CEP, ADE, and PDE. Expression of a cat-2 cDNA from its own promoter or a promoter that was selectively expressed in CEP sensory neurons rescued aversion on the ∆crp diet (Fig. 6a).
a Aversion behavior of cat-2 mutants expressing cat-2 genomic DNA from its the endogenous promoter (cat-2), or promoters for all dopaminergic neurons (dat-1), CEP neurons (p27), CEP sheath glial cells (hlh-17), ADE and PDE neurons (dat-1p19), or intestine (elt-2) on the mediocre ∆crp diet (n = 9–18 assays). b Aversion behavior of WT and dopamine receptor mutants on the mediocre ∆crp diet (n = 9–18 assays). c Aversion behavior of dop-6 mutants with dop-6 rescue in tdc-1- or tbh-1- expressing cells on the mediocre ∆crp diet (n = 9-10 assays). d Genetic interaction of cat-2 with tdc-1 or tbh-1 (tyramine and/or octopamine biosynthesis mutants) on the mediocre ∆crp diet (n = 7-9 assays). e Genetic interaction of dop-6 with tdc-1 or tbh-1 on the mediocre ∆crp diet, and schematic model of dopamine signaling. Stopped and dotted lines represent inhibition of RIC and octopamine (n = 10–15 assays). f Scanning electron micrographs of wild-type BW25113 bacteria, ∆crp bacteria, and ∆wxzE bacteria (left) and quantification of bacterial length and width (right, with SEM). For length, WT, n = 58; ∆crp, n = 41; ∆wzxE, n = 72. For width, WT, n = 59; ∆crp, n = 54; ∆wzxE, n = 60. Scale bar, 2 μm. For (a–e), each data point indicates individual assay. Results are shown with median ± quartiles in boxes and Min to Max whiskers. ns, not significant, *P < 0.05, **P < 0.01, ***P < 0.001 by one-way ANOVA corrected by Dunnett’s multiple comparisons (a–c) or Tukey’s multiple comparisons (d, e), or by two-tailed, unpaired t test (b, dop-6). For (f), each data point indicates individual bacterial cell. ***P < 0.001 by two-tailed Mann-Whitney test. Full genotypes of all C. elegans strains are provided in Supplementary Data 3. All sample sizes, statistical tests used, and exact P values are provided in Supplementary Data 4. Source data are provided as a Source Data file.
Among dopamine receptors, animals mutant for the inhibitory (Gi/o-coupled) dopamine receptor DOP-6, but not other dopamine receptors, had reduced aversion to the ∆crp mediocre diet, like cat-2 mutants (Fig. 6b). dop-6 is expressed in RIC neurons35,73, and previous studies showed that dop-6 can inhibit RIC activity72. We found that expression of a dop-6 cDNA in the RIC neuron, or in RIM and RIC neurons, rescued aversion from the mediocre ∆crp diet (Fig. 6c).
To further explore the dopamine circuit, we used double mutants to ask how dopamine interacts with RIM and RIC neurotransmitters. Animals mutant for the RIM/RIC neurotransmitter enzymes tdc-1 or tbh-1 avoided the mediocre ∆crp diet (Figs. 6d, 7h), as did cat-2 tdc-1 and cat-2 tbh-1 double mutants, unlike cat-2 single mutants (Fig. 6d). tdc-1 dop-6 or tbh-1 dop-6 double mutants also avoided the mediocre ∆crp diet, as expected if DOP-6 inhibits tyramine and octopamine function from RIM and RIC (Fig. 6e). As RIC expression was sufficient to rescue dop-6, and cat-2 aversion defects were fully suppressed by tbh-1, it is likely that octopamine from RIC neurons can suppress aversion from the ∆crp diet but is inhibited by dopamine.
a Behavioral responses of wild-type, tph-1, and serotonin receptor mutant animals on the mediocre ∆cysE diet. Aversion is diminished in ser-1 and mod-1 mutants (n = 8–18 assays). b Genetic interaction of mod-1 with ser-1 on the mediocre ∆cysE diet (n = 9-11 assays). c Aversion behavior of ser-1 mutants expressing ser-1 cDNA from its endogenous promoter (ser-1) or promoters for RIA (glr-3) or RIC neurons (tbh-1). Two transgenic lines were tested for each plasmid (n = 9–15 assays). d Aversion behavior of mod-1 rescued in RIM and/or RIC neurons. Two transgenic lines were tested for each plasmid (n = 12–16) assays. e Genetic interaction of mod-1 with tdc-1 or tbh-1 on the ∆cysE diet and schematic model of MOD-1 action. Stopped and dotted lines represent inhibition of RIC and octopamine (n = 10-13 assays). f Genetic interaction of ser-1 with tdc-1 on the mediocre ∆cysE diet (n = 10-12 assays). Genetic interactions of tph-1, tdc-1, and tbh-1 across bacterial diets. (g) wild-type diet (n = 12 assays); (h) ∆crp diet (n = 9-15 assays); (i) ∆cysE diet (n = 12-15 assays). g–i use the tdc-1(n3420) allele; all other figures and panels use tdc-1(n3419). For (a–f), each data point indicates individual assay. Results are shown with median ± quartiles in boxes and Min to Max whiskers. ns, not significant, *P < 0.05, **P < 0.01, ***P < 0.001 by one-way ANOVA corrected by Dunnett’s multiple comparisons (a, c, d) or Tukey’s multiple comparisons (b, e–i). Full genotypes of all C. elegans strains are provided in Supplementary Data 3. All sample sizes, statistical tests used, and exact P values are provided in Supplementary Data 4. Source data are provided as a Source Data file.
The CEP neurons that produce dopamine are implicated in mechanosensory detection of bacteria76. To ask whether ∆crp bacteria might have unusual mechanosensory features, we examined these bacteria by scanning electron microscopy. Under our growth conditions, ∆crp cells were significantly larger than wild-type E. coli (Fig. 6f). A second bacterial strain with morphological alterations was ∆wzxE, the ECA biosynthesis mutant (Fig. 6f). These defects may be related to the defective outer membrane structures of ∆wzxE mutants. Like aversion to the ∆crp diet, aversion from the ∆wzxE diet required tph-1, cat-2, and ser-7 genes (Supplementary Fig. 4c).
In summary, aversion from the mediocre ∆crp diet involves antagonistic interactions between serotonin and dopamine, which promote aversion, and octopamine, which suppresses aversion. Cell-specific rescue experiments identify NSM, HSN, CEP, I1, and RIC neurons as regulators of ∆crp aversion.
Two serotonin receptors, SER-1 and MOD-1, promote aversion from a mediocre ∆cysE diet
The serotonin receptor mutants ser-1 and mod-1 had reduced aversion from the mediocre ∆cysE diet, like tph-1 mutants (Fig. 7a, b). The excitatory receptor ser-1 is expressed in RIA and RIC neurons, among other locations35,73,74,77, and regulates nociception by acting in RIC neurons74. We found that a ser-1 cDNA rescued aversion from the ∆cysE diet when expressed from its endogenous promoter or when expressed specifically in RIA neurons, but did not rescue when expressed in RIC neurons (Fig. 7c).
Serotonin released from NSM can inhibit RIM and RIC neurons through the MOD-1 receptor to modulate locomotion or feeding78,79,80. Expressing mod-1 in RIC neurons or in RIM and RIC neurons rescued aversion of mod-1 mutants on the mediocre ∆cysE diet (Fig. 7d), Double mutants between mod-1 and the RIM/RIC neurotransmitter enzymes tdc-1 or tbh-1 restored aversion on the mediocre ∆cysE diet (Fig. 7e). The combination of mod-1 rescue in RIC neurons and aversion in mod-1 tbh-1 double mutants indicates that octopamine from RIC can suppress aversion from the ∆cysE diet, but is inhibited by mod-1.
Notably, different combinations of genes, neurons, and bacterial diets did not always yield simple interpretations. For example, tdc-1 ser-1 double mutants had strong aversion to the ∆cysE diet, indicating that tdc-1 can suppress ser-1, even though ser-1 aversion was not rescued in the RIC neurons (Fig. 7f). The dopamine-deficient cat-2 mutant had reduced aversion to the mediocre ∆cysE diet, but CEP neurons and the dopamine receptor dop-6 were not specifically required for this behavior (Supplementary Fig. 5a, b), further distinguishing the requirements for aversion from ∆crp and ∆cysE mediocre diets.
Finally, genetic interactions between tph-1, tdc-1, and tbh-1 highlighted the context-dependent contributions of serotonin, tyramine, and octopamine on different bacterial diets (Fig. 7g-i). Double mutants between tph-1 and tbh-1 resembled tph-1 mutants on all diets, pointing to a central role of serotonin across conditions, and a secondary role for octopamine. However, on a wild-type diet tyramine may have at least some serotonin-independent functions, as tdc-1 tph-1 double mutants had intermediate phenotypes compared to either single mutant (Fig. 7g).
Discussion
Ingestion of toxic foods that induce sickness, nausea, or gastrointestinal dysfunction is followed by aversion behaviors that prolongs an animal’s survival1,81. C. elegans initially enters a bacterial lawn using a simple rule-of-thumb based on bacterial cell density64 but after several hours of exposure animals avoid lethal bacterial foods such as certain RNAi-expressing bacteria14, food spiked with exogenous toxins14,32, or pathogens15,16,18,82. Here we show that a less extreme, but related aversion behavior is induced by a small number of mutant E. coli diets that induce mitochondrial or redox stress markers. We find that aversion is regulated by interacting serotonin, dopamine, and octopamine neurotransmitters and receptors.
Our genome-wide screen identified 22 E. coli mutants that induced aversive responses, a small fraction of the 244 E. coli mutants that result in C. elegans developmental delay46. Aversion is a stringent assay that requires animals to spend a substantial fraction of their time away from the only available food, and thus it may not be induced by simple nutrient limitation. In addition, the aversion assay begins at the L4 larval stage, when animals may have sufficient reserves of many nutrients. In this setting, we suggest that aversion represents a behavioral response to diets that induce metabolic stress. In agreement with this interpretation, the genes identified here overlap with those that elicit avoidance of mitochondrial toxins or bacterial pathogens14,16,32,83.
We focused our studies on two bacterial genes whose effects on C. elegans had not been examined previously, the catabolite regulator CRP and the cysteine biosynthesis gene CysE. CRP is a global regulator of bacterial metabolism on low-carbohydrate media. On nematode growth medium, ∆crp bacteria were abnormally large, and induced expression of C. elegans stress reporters associated with toxic or inedible bacteria (daf-7) and the mitochondrial unfolded protein response (hsp-6)84. These two reporters were also induced by the ∆fepB ferric enterobactin transport mutant that elicited aversion. Interestingly, our screen yielded the same four fes/fep genes as screens for C. elegans developmental delay46, poor development on a nutrient-limited food source63, and synthetic lethality with the C. elegans mitochondrial mutant spg-748. The ferric-enterobactin complex that accumulates in these strains appears to cause mitochondrial toxicity48. As mitochondrial toxins can drive food aversion in C. elegans14, we suggest that mitochondrial dysfunction contributes to aversion to ∆fes/fep and ∆crp diets. Aversion to the mediocre ∆crp diet may also reflect an abnormal size or shape that makes the bacteria harder to ingest; hard-to-ingest food can also drive behavioral aversion and daf-7 stress reporters28. The identification of cysE as a mediocre food that elicits the oxidative stress reporter gst-4 (i.e., glutathione S-transferase) is consistent with the role of cysteine in synthesis of the antioxidant glutathione. C. elegans synthesizes glutathione from cysteine and other precursors, and this pathway requires more cysteine than protein synthesis. Either an excess or a deficiency of dietary thiols has effects on redox states that are disadvantageous for C. elegans85, explaining why cysteine, an amino acid that C. elegans can synthesize on its own, nonetheless can be limited by bacterial production and disruption of glutathione homeostasis.
These experiments were conducted on BW25113, an E. coli K12 strain that has been widely used to study metabolism in C. elegans, and not on the typical OP50 E. coli B strain. While these bacteria are approximately equal in attractiveness, they are not metabolically identical19. For example, although animals develop quickly and feed well on K12 strains compared to the OP50 B strains, they have reduced lipid stores86,87,88 and altered lifespan regulation caused by changes in bacterial lipopolysaccharide (LPS)89. Conversely, OP50 is a poor source of the essential vitamin B12 and can result in slower larval development of C. elegans9,42,86 and thus may engage different behavioral and stress responses than K12 bacteria.
The neurotransmitter serotonin is required for C. elegans to distinguish between high-quality and mediocre E. coli diets, with bidirectional roles in guiding behavior (Fig. 8a, b). Serotonin has long been associated with positive aspects of food quality in C. elegans, and stimulates behaviors such as feeding, egg-laying, and dwelling that are observed in beneficial conditions. However, serotonin has a prominent role in behavioral and physiological responses to toxic foods in mammals90, and in C. elegans, serotonin is required for the avoidance of toxic or pathogenic bacteria14,32 and for the learned avoidance of bacterial pathogens20. Using cell-specific knockouts and acute silencing experiments, we found that high-quality and mediocre diets were interpreted by different classes of serotonergic neurons. The serotonergic ADF neurons supported retention on a high-quality bacterial diet, while serotonergic NSM neurons and to a lesser extent HSN neurons drove aversion from mediocre diets (Fig. 8b).
a Neurotransmitters and receptors that regulate aversion on the wild-type diet and mediocre ∆crp or ∆cysE diets. Mean aversion ratios across all assays were used to generate the bubble map; dotted lines represent the mean aversion ratio of wild-type animals on each diet; statistical comparisons appear in the main figures. b Preliminary neural circuits for aversion behavior; additional neurons remain to be defined. Arrows represent activation of neurons, transmitters, or behaviors; stopped and dotted lines represent inhibition.
These results add to existing evidence that serotonin and individual serotonergic neurons have context-dependent functions. For example, ADF serotonin drives increased feeding on familiar bacteria21,80, possibly aligning with the appetitive retention on the (familiar) high-quality diet observed here and attraction to bacterial metabolites33. However, ADF serotonin can also drive learned avoidance of pathogenic bacteria, an aversive behavior20,91. Conversely, serotonin from NSM neurons supports dwelling behavior on food and slow locomotion upon food encounters, which are appetitive behaviors11,31, but our results indicate that NSM serotonin can drive avoidance of mediocre foods. While this finding appears surprising, it matches prior results showing that NSM serotonin drives slow avoidance of food spiked with the mitochondrial toxin antimycin A17. Moreover, antimycin exposure causes NSM to change its properties such that calcium levels increase after food removal rather than food addition17. Together, these results argue that serotonergic neurons respond to metabolic history and not just immediate sensory inputs from food. In agreement with this hypothesis, serotonin synthesis in both ADF and NSM is regulated by food history80,92, serotonin levels in ADF are increased by pathogen exposure20, and serotonin synthesis in NSM is induced by mitochondrial toxins32.
By screening for aversion behavior on three different bacterial diets, we identified roles for all six known serotonin receptors in the aversion assay. Two of the serotonin receptors, ser-4 and ser-5, were required to suppress aversion from a high-quality diet; the other four receptors, ser-7, lgc-50, mod-1, and ser-1, supported aversion from two mediocre diets (Fig. 8a). Some serotonin receptors distinguished among mediocre foods, such that ser-7 only affected aversion to ∆crp, whereas ser-1 only affected aversion to ∆cysE. This double dissociation indicates that the nervous system can discriminate among aversive contexts. All six serotonin receptors also participate in serotonin-dependent locomotor slowing when C. elegans enters a bacterial lawn35. Slowing on lawn entry relies on NSM and HSN serotonergic neurons, like aversion from mediocre diets, but the contributions of receptors are different: mod-1, ser-4, and lgc-50 have partly redundant functions that promote slowing behavior, while ser-1, ser-5, and ser-7 have modulatory roles35. In another assay, food-dependent regulation of nociception, the serotonin receptors ser-1, ser-5, and mod-1 have congruent but non-redundant roles93. These comparisons highlight the distinctiveness of serotonin signaling in different contexts. We note that serotonin-deficient tph-1 mutants move rapidly on food, but are largely normal in their locomotion32,35,94, as are the single-cell serotonin knockouts that elicit aversion behavior11. The six serotonin receptor mutants, as well as dopamine and octopamine mutants, also have largely normal locomotion on food32,35,94,95.
In addition to serotonin, the monoamine transmitters dopamine and octopamine contribute to aversion behavior. Dopamine promotes aversion from both mediocre diets, while octopamine suppressed aversion from all diets. Dopamine is also required for aversion from bacteria spiked with the mitochondrial toxin antimycin83, but the ∆crp diet and antimycin engage different dopamine receptors83. Like serotonergic neurons, dopamine neurons can reshape their properties based on context: exposure to a low-quality diet increases dopaminergic neuron responses to bacterial conditioned medium72.
The octopaminergic RIC neurons oppose the action of serotonin from NSM, suppressing aversion from all bacterial lawns. Octopamine and tyramine also suppress aversion from the bacterial pathogen Pseudomonas aeruginosa PA14, which downregulates expression of the biosynthetic enzyme tdc-1 in RIM and RIC neurons to reduce tyramine and octopamine signaling and enhance aversion16. We found that the serotonin and dopamine receptors ser-5, mod-1, and dop-6 could all modulate aversion when expressed in the octopaminergic RIC neurons, alone or together with the tyraminergic RIM neurons. Other serotonin targets such as RIA and I1 neurons appear selective for particular bacterial contexts, although future studies may reveal additional connections among them. We did not test all known sites of serotonin receptor expression, so the neurons identified here may not be the only ones that can affect aversion.
Serotonin and tyramine/octopamine have antagonistic functions in C. elegans egg-laying, feeding regulation, and nociception74,79,80,96. We observe similar antagonism on mediocre diets, but our results on the high-quality diet show that serotonin can potentiate RIC and octopamine signaling in certain conditions. Tracing the interactions between these neurons and transmitters will provide more insight into these neuromodulatory circuits and their responses to different bacterial diets. More generally, the parallels and contrasts between these wireless circuits for aversion provide insight into the logic of neuromodulatory integration across molecules and contexts.
Methods
A list of Reagents and Resources used in this study is available in Supplementary Data 2.
The full genotypes of strains shown in each figure are provided in Supplementary Data 3.
C. elegans strains
All C. elegans strains were maintained on nematode growth medium (NGM) agar plates seeded with bacterial diet E. coli BW25113 at 20 °C. CGC1 (formerly known as PD1074) was provided by the Caenorhabditis Genetics Center (CGC) and used as the wild-type strain. All strains used in this study are listed in Supplementary Data 2.
Bacterial strains
E. coli strains OP50, DA837, HB101, BW25113, and Comamonas aquatica strain DA1877 were cultured in Lysogeny Broth (LB) liquid medium at 37 °C. Pseudomonas aeruginosa strain PA14 was cultured in Lysogeny Broth (LB) liquid medium at 25 °C. E. coli Keio deletion mutants45 were purchased from the National Bioresource Project of Japan (NBRP) and were grown at 37 °C in LB medium with 25 μg/mL kanamycin.
Generation of transgenes
To construct plasmids for expression of ser-5 in C. elegans neurons, promoter sequences of ser-5 (2001bp), glr-7 (2000bp), tdc-1 (4550 bp) or tbh-1 (4537 bp) were inserted into the pSM::SL2::GFP::short unc-54 3’ UTR vector along with a ser-5 cDNA. Injection mixtures containing Pser-5::ser-5::SL2::GFP (30 ng/uL), Pglr-7::ser-5::SL2::GFP (30 ng/uL), Ptdc-1::ser-5::SL2::GFP (30 ng/uL), or Ptbh-1::ser-5::SL2::GFP (30 ng/uL) together with the co-injection marker Pmyo-3::mCherry (3 ng/uL) were injected into CX13569 ser-5(tm2647) I.
To construct plasmids for expression of ser-7 in C. elegans neurons, promoter sequences of ser-7 (2118 bp), flp-21 (2003bp), del-7 (839 bp), ceh-28 (1981bp), ceh-19 (1496 bp), ceh-53 (1476 bp), ceh-45 (2022 bp), lgc-8 (1999bp), or flp-15 (2402 bp) were inserted into the pSM::SL2::GFP::short unc-54 3’ UTR vector along with a ser-7 cDNA. Injection mixtures containing Pser-7::ser-7::SL2::GFP (30 ng/uL), Pflp-21::ser-7::SL2::GFP (30 ng/uL), Pdel-7::ser-7::SL2::GFP (30 ng/uL), Pceh-28::ser-7::SL2::GFP (30 ng/uL), Pceh-19::ser-7::SL2::GFP (30 ng/uL), Pceh-53::ser-7::SL2::GFP (30 ng/uL), Pceh-45::ser-7::SL2::GFP (30 ng/uL), Plgc-8::ser-7::SL2::GFP (30 ng/uL), or Pflp-15::ser-7::SL2::GFP (30 ng/uL) together with the co-injection marker Pmyo-3::mCherry (3 ng/uL) were injected into DA2100 ser-7(tm1325) X.
To construct plasmids for expression of ser-1 in C. elegans neurons, promoter sequences of ser-1 (2103 bp), tbh-1 (4537 bp), or glr-3 (2838 bp) were inserted into the pSM::SL2::GFP::short unc-54 3’ UTR vector along with a ser-1 cDNA. Injection mixtures containing Pser-1::ser-1::SL2::GFP (30 ng/uL), Ptbh-1::ser-1::SL2::GFP (30 ng/uL), or Pglr-3::ser-1::SL2::GFP (30 ng/uL) together with the co-injection marker Pmyo-3::mCherry (3 ng/uL) were injected into DA1814 ser-1(ok345) X.
To construct plasmids for expression of histamine-gated chloride channels in order to chemically silence C. elegans neurons, promoter sequence of srh-142 (3976 bp), or del-7 (839 bp) were inserted into the pSM::SL2::mCherry::short unc-54 3’ UTR vector along with the HisCl1 coding sequence. Injection mixtures containing Psrh-142::HisCl1::SL2::mCherry (20 ng/uL) together with the co-injection marker Pelt-2::NLS::GFP (2.5 ng/uL), or Pdel-7::HisCl1::SL2::mCherry (20 ng/uL) (no coinjection marker) were injected into PD1074.
To construct plasmids for expression of cat-2 in C. elegans neurons, promoter sequences of cat-2prom3 (1143 bp), dat-1 (786 bp), hlh-17 (3008 bp), or elt-2 (5049 bp) were inserted into the pSM::SL2::GFP:: unc-54 3’ UTR vector along with the cat-2 genomic coding region. Injection mixtures containing Pcat-2prom3::cat-2::SL2::GFP (30 ng/uL), Pdat-1::cat-2::SL2::GFP (30 ng/uL), Phlh-17::cat-2::SL2::GFP (30 ng/uL), or Pelt-2::cat-2::SL2::GFP (30 ng/uL) together with the co-injection marker Pmyo-3::mCherry (5 ng/uL) were injected into CX11078 cat-2(e1112) II.
To construct plasmids for expression of dop-6 in C. elegans neurons, promoter sequences of tdc-1 (4550 bp) or tbh-1 (4537 bp) were inserted into the pSM::SL2::GFP::short unc-54 3’ UTR vector along with a dop-6 cDNA. Injection mixtures containing Ptdc-1::dop-6::SL2::GFP (30 ng/uL), or Ptbh-1::dop-6::SL2::GFP (30 ng/uL) together with the co-injection marker Pmyo-3::mCherry (3 ng/uL) were injected into RB1680 dop-6(ok2090) X.
To construct plasmids for expression of mod-1 in C. elegans neurons, promoter sequences of tdc-1 (4550 bp) or tbh-1 (4537 bp) were inserted into the pSM::SL2::GFP::short unc-54 3’ UTR vector along with a mod-1 cDNA. Injection mixtures containing Ptdc-1::mod-1::SL2::GFP (30 ng/uL), or Ptbh-1::mod-1::SL2::GFP (30 ng/uL) together with the co-injection marker Pmyo-3::mCherry (3 ng/uL) were injected into MT9668 mod-1(ok103) V.
To construct plasmids for inverted rescue of tdc-1 in C. elegans neurons, the promoter sequence of tdc-1 was assembled with the pSM vector together with the inverted/floxed tdc-1 cDNA-sl2GFP. Injection mixtures containing Ptdc-1::inverted/floxed tdc-1 cDNA-sl2GFP (60 ng/uL) together with the co-injection marker Pmyo-3::mCherry (3 ng/uL) were injected into MT13113 tdc-1 (n3419) II, to generate the inverted tdc-1 rescue background line. This line with ‘No Cre’ was identified as tdc-1; kyIs837. kyIs837 was a spontaneously integrated allele that was backcrossed three times. To generate rescue lines for neurons, transgenic lines containing Primb-1::nCre (5 ng/uL) or Pdmsr-7::nCre (3 ng/uL) together with the co-injection maker myo-2::mCherry (1 ng/uL) were crossed with the inverted tdc-1 rescue background line.
Screen for C. elegans aversion to E. coli Keio knockout mutants
To prepare bacteria for the primary behavioral assay, individual E. coli deletion mutants were grown in LB medium with 25 μg/mL kanamycin at 37 °C for about 16 h. 12.5 μL overnight cultures were seeded onto 12-well plates containing 3.5 mL standard NGM agar in each well, dried, grown at 37 °C for one day and incubated at room temperature for another day. Animals were maintained on the standard wild-type bacteria BW25113 for at least two generations before the behavioral assay. About 15-20 L4 animals were placed in each well of 12-well plates that contained BW25113_WT or individual E. coli mutants and incubated at 21 °C for 20 h. Plates were then visually screened for the fraction of animals that exhibited aversive behavior, defined as (animals outside the bacterial lawn)/(total number of animals). Each E. coli deletion mutant was screened at least twice. A secondary screen was performed for identified positive hits from the primary screen, using the same method. Positive hits identified from the secondary screen were further verified in assays on 6-well plates that were video-tracked for 20 h (recording rate 1 frame per minute, 1200 frames per video) with at least three independent replicates. Behavioral rigs were equipped with four 15 MP cameras (PL-D7715, Pixelink), one per six-well plate, and videos were cropped to individual wells and analyzed every six frames (200 data points per condition) using published MatLab codes (https://doi.org/10.5281/zenodo.10723701)16. Wild-type C. elegans on wild-type BW25113 bacteria were defined as having low aversive behavior (aversion ratio less than 0.2). Bacterial mutants that induced significantly higher aversion than BW25113_WT were chosen for subsequent studies.
Genotyping identified E. coli mutants
Bacterial colonies from individual bacterial mutants were streaked on LB agar plates, grown at 37 °C, and genotyped by PCR using the kanamycin-cassette-specific primers (forward primer 5’-CGGTGCCCTGAATGAACTGC-3’, reverse primer 5’-CAGTCATAGCCGAATAGCCT-3’) and genomic primers for individual bacterial mutants, which were designed at starting 100-300 bases upstream of start-codon (forward primer) or downstream of stop codon (reverse primer). The correct fragment sizes were verified.
Quantification of Pdaf-7::gfp
Pdaf-7::gfp animals at the L4 stage were transferred onto behavioral assay plates seeded with E. coli wild-type (BW25113) or test bacteria and incubated at 21˚ C for 16 h. Animals mounted on 2% agarose pads and 20 mM sodium azide were imaged under the same parameters (63x objective, Z-stack and exposure time) on the Zeiss Axioimager Z1. Quantification of mean GFP intensity was performed on maximum fluorescence intensity projections of ASI and ASJ cell bodies in FIJI.
Fluorescent reporter quantification assays
All animals containing fluorescent markers (hsp-6p::gfp, hsp-4p::gfp, gst-4p::gfp, clec-60p::gfp, irg-1p::gfp, or irg-5p::gfp) were picked onto behavioral assay plates seeded with E. coli test bacteria at the L4 stage and incubated at 21 °C for 16 h, except animals with the stress reporter gst-4::gfp, which were incubated for 8 h before imaging. Animals mounted on 2% agarose pads and 20 mM sodium azide were imaged on the Zeiss Axioimager Z1. As these reporters are broadly expressed, mean GFP fluorescence per pixel was quantified across the body using FIJI. Control experiments showed that the body sizes of hsp-6p::gfp and gst-4p::gfp animals grown on each bacterial strain were comparable.
Bacterial colonization assay
Animals were treated under behavioral assay condition with indicated bacterial mutants for 12 h, picked into tubes with M9 buffers, washed three times, and placed on empty NGM plates containing 100 μg/mL carbenicillin for 1 hour. At least five animals were then individually picked into a new Eppendorf tube with M9 buffer and homogenized using a motorized tissue grinder (Fisher Scientific) coupled with a 1.5 mL plastic pestle (Fisher Scientific). The lysate was plated onto LB plates and incubated at 37 °C for 16 h. Bacterial colonies were counted manually. Three independent biological repeats were performed for E. coli wild-type or mutants.
Pharyngeal pumping (feeding) assay
Animals were incubated as above for 12 h on plates seeded with indicated bacteria, then recorded using a LEICA MZ6 microscope coupled with cameras for 30 s and the number of pharyngeal contractions per animal was counted manually. At least six animals were recorded in each of two or three independent repeats.
Brood size and development assay
For the brood size assay, an individual animal at the L4 stage was picked onto one NGM plate seeded with 200 μL bacteria grown under the conditions used in the behavioral assay. Animals were picked onto new plates every day and maintained at 20 °C. Brood size was determined by counting the total number of progeny across all plates for each animal. At least three independent biological replicates (6–8 animals for each repeat) were performed.
For the development assay, five synchronized adult worms were allowed to lay eggs for 3 h at 20 °C, on NGM plates with bacteria grown under the conditions used in the behavioral assay. Eggs were allowed to develop at 20 °C for 72 h, and scored for the number of progeny that had reached the L4 or adult stage as a fraction of all progeny.
Lifespan assay
Seeded NGM plates used for lifespan assays were prepared with bacteria grown under the conditions used in the behavioral assay. About 20 synchronized L4 worms were picked onto assay plates and maintained at 25 °C. To separate parents from offspring, adult animals were picked onto freshly prepared assay plates daily until no further progeny were observed. Dead animals were identified by non-response to gentle prodding. Animals that were lost or crawled off agar were censored.
Chemical supplementation
50% glucose (Millipore Sigma), 0.2 M cysteine, 0.2 M O-acetyl-serine and 40 mM PLP were prepared by dissolving powders in deionized water, filtered and stored at −20 °C. For behavioral analysis with pre-adding chemicals, stock solutions were supplemented into NGM agar immediately before pouring to desired final concentrations (glucose, 0.4%; Cys, 200 μM; OAS, 200 μM; PLP, 40 μM). Bacterial seeding and behavioral assay were performed as described in the section ‘Screen for C. elegans aversion to E. coli Keio knockout mutants’. For behavioral analysis with post-adding chemicals, assay plates were prepared by first seeding 12.5 μL indicated bacterial cultures onto NGM agar for growth, as described in the section ‘Screen for C. elegans aversion to E. coli Keio knockout mutants’. One hour before the behavioral assay, chemicals were directly added onto the bacterial spots to reach the final concentration under a total volume of 6 mL NGM agar (Cys, 50 μM; OAS, 200 μM; PLP, 40 μM). NGM plates without any added chemicals were assayed in parallel.
Bacterial density experiments
Overnight bacterial cultures were grown in LB medium at 37 °C and seeded onto behavioral assay plates (Supplementary Fig. 3). Standard NGM agar plates contained 0.3% NaCl, 0.25% peptone, 25 mM potassium phosphate, 1 mM CaCl2, 1 mM MgSO4, 5 μg/mL cholesterol, 2.2% agar. To increase bacterial density of BW25113_WT, ∆crp or ∆cysE bacteria, 12.5 μL of overnight bacterial cultures (1x) or different concentrated suspensions of those cultures (10x, 20x, or 40x) were seeded onto NGM agar and allowed to grow for two days as indicated before behavioral tests. To decrease the bacterial density of BW25113 (WT), the behavioral assay plates contained NGM agar with low amounts of peptone (0.125%, 0.05%, or 0.025%) and were seeded with 12.5 μL of overnight bacterial culture as above. For pre-adding conditions, 200 μM cysteine or OAS were added to NGM agar when the plates were made, prior to bacterial seeding. For post-adding cysteine or OAS, assay plates were prepared by first seeding 12.5 μL of the indicated bacterial cultures onto NGM agar for two days’ growth, and then cysteine or OAS was administered to the final 50 μM or 200 μM concentration in a total volume of 6 mL NGM agar, respectively. Matched pairs of agar plates were used for behavioral testing (with animals) or for determining bacterial density. Bacteria were harvested at the beginning of the behavioral assay, except for supplementation experiments (Fig. 2e and Supplementary Fig. 3i and j), in which bacteria were counted at the end of the behavioral assay. Bacteria were washed off the agar with 2 mL M9 buffer, subjected to OD600 measurement using a DS-11 FX+ spectrophotometer (DeNovix), and bacterial density determined by serial dilution, plating on agar plates, and counting colony-forming units (CFU)64. We did not observe any significant differences in CFU numbers after plating on LB or NGM agar plates.
Neuron silencing by histamine assay
1 M histamine dihydrochloride (Millipore Sigma) was prepared by dissolving powder in deionized water, filtered and stored at -20 °C. Histamine stock solution was added to NGM agar at a final concentration of 10 mM prior to seeding with bacteria. For behavioral analysis, NGM plates with or without histamine and animals with or without expression of HisCl1 channels were assayed in parallel.
Scanning electron microscopy
Bacteria were gently eluted from assay plates and fixed in 2% glutaraldehyde, 4% formaldehyde in 0.1 M sodium cacodylate buffer, pH 7.2, for more than 1 hour at room temperature followed by overnight fixation at 4 °C. After post-fixation with 1% osmium tetroxide in 0.1 M sodium cacodylate buffer, pH 7.2 for 1 h at room temperature, samples were dehydrated in a graded series of ethanol followed by hexamethyldisilazane. Bacteria were spread onto glass coverslips and let air-dry overnight. The next morning, samples were sputter-coated with 10 nm of iridium using a Leica ACE600 sputter coater. Images were acquired in a scanning electron microscope JEOL JSM-IT500HR at 5.0 kV (JEOL USA, Inc).
Statistical analysis
Statistical analyses were performed using GraphPad Prism and are detailed in the figure legends. Detailed statistical summaries are provided in Supplementary Data 4.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated in this study are provided in this paper, Supplementary Information, Supplementary Data, and Source Data file. Source data are provided with this paper. All information and requests for reagents or strains should be directed to the lead contact Cornelia Bargmann (cori@rockefeller.edu). Source data are provided with this paper.
References
Bjordal, M., Arquier, N., Kniazeff, J., Pin, J. P. & Léopold, P. Sensing of amino acids in a dopaminergic circuitry promotes rejection of an incomplete diet in Drosophila. Cell 156, 510–521 (2014).
Barredo, E. & DeGennaro, M. Not just from blood: mosquito nutrient acquisition from nectar sources. Trends Parasitol. 36, 473–484 (2020).
Tung, G. A. & Fonseca, D. M. Internal and external drivers interact to create highly dynamic mosquito blood-feeding behaviour. Proc. Biol. Sci. 291, 20241105 (2024).
Richter, C. P., Holt, L. E. & Barelare, B. Vitamin B1 craving in rats. Science 86, 354–355 (1937).
Koyama, T. et al. A nutrient-responsive hormonal circuit mediates an inter-tissue program regulating metabolic homeostasis in adult Drosophila. Nat. Commun. 12, 5178 (2021).
Berthoud, H. R., Münzberg, H. & Morrison, C. D. Blaming the brain for obesity: Integration of hedonic and homeostatic mechanisms. Gastroenterology 152, 1728–1738 (2017).
Avery, L. & Shtonda, B. B. Food transport in the C. elegans pharynx. J. Exp. Biol. 206, 2441–2457 (2003).
Katzen, A. et al. The nematode worm C. elegans chooses between bacterial foods as if maximizing economic utility. eLife 12, e69779 (2023).
Shtonda, B. B. & Avery, L. Dietary choice behavior in Caenorhabditis elegans. J. Exp. Biol. 209, 89–102 (2006).
Song, B. M. & Avery, L. Serotonin activates overall feeding by activating two separate neural pathways in Caenorhabditis elegans. J. Neurosci. 32, 1920–1931 (2012).
Flavell, S. W. et al. Serotonin and the neuropeptide PDF initiate and extend opposing behavioral states in C. elegans. Cell 154, 1023–1035 (2013).
Kim, D. H. & Flavell, S. W. Host-microbe interactions and the behavior of Caenorhabditis elegans. J. Neurogenet. 34, 500–509 (2020).
Scheer, E. & Bargmann, C. I. Sensory neurons couple arousal and foraging decisions in Caenorhabditis elegans. eLife 12, e88657 (2023).
Melo, J. A. & Ruvkun, G. Inactivation of conserved C. elegans genes engages pathogen- and xenobiotic-associated defenses. Cell 149, 452–466 (2012).
Pradel, E. et al. Detection and avoidance of a natural product from the pathogenic bacterium Serratia marcescens by Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 104, 2295–2300 (2007).
Marquina-Solis, J. et al. Antagonism between neuropeptides and monoamines in a distributed circuit for pathogen avoidance. Cell Rep. 43, 114042 (2024).
Tsai, S. H., Wu, Y. C., Palomino, D. F., Schroeder, F. C. & Pan, C. L. Peripheral peroxisomal β-oxidation engages neuronal serotonin signaling to drive stress-induced aversive memory in C. elegans. Cell Rep. 43, 113996 (2024).
Meisel, J. D., Panda, O., Mahanti, P., Schroeder, F. C. & Kim, D. H. Chemosensation of bacterial secondary metabolites modulates neuroendocrine signaling and behavior of C. elegans. Cell 159, 267–280 (2014).
Liao, C. P. et al. Neurophysiological basis of stress-induced aversive memory in the nematode Caenorhabditis elegans. Curr. Biol. 32, 5309–5322.e6 (2022).
Zhang, Y., Lu, H. & Bargmann, C. I. Pathogenic bacteria induce aversive olfactory learning in Caenorhabditis elegans. Nature 438, 179–184 (2005).
Song, B. M., Faumont, S., Lockery, S. & Avery, L. Recognition of familiar food activates feeding via an endocrine serotonin signal in Caenorhabditis elegans. eLife 2, e00329 (2013).
Ji, N. et al. A neural circuit for flexible control of persistent behavioral states. eLife 10, e62889 (2021).
Sawin, E. R., Ranganathan, R. & Horvitz, H. R. C. elegans locomotory rate is modulated by the environment through a dopaminergic pathway and by experience through a serotonergic pathway. Neuron 26, 619–631 (2000).
Oranth, A. et al. Food sensation modulates locomotion by dopamine and neuropeptide signaling in a distributed neuronal network. Neuron 100, 1414–1428.e10 (2018).
Alkema, M. J., Hunter-Ensor, M., Ringstad, N. & Horvitz, H. R. Tyramine functions independently of octopamine in the Caenorhabditis elegans nervous system. Neuron 46, 247–260 (2005).
Özbey, N. P. et al. Tyramine acts downstream of neuronal XBP-1s to coordinate inter-tissue UPRER activation and behavior in C. elegans. Dev. Cell 55, 754–770.e6 (2020).
Churgin, M. A., McCloskey, R. J., Peters, E. & Fang-Yen, C. Antagonistic serotonergic and octopaminergic neural circuits mediate food-dependent locomotory behavior in Caenorhabditis elegans. J. Neurosci. 37, 7811–7823 (2017).
Boor, S. A., Meisel, J. D. & Kim, D. H. Neuroendocrine gene expression coupling of interoceptive bacterial food cues to foraging behavior of C. elegans. eLife 12, e91120 (2024).
Buckley, N. A., Dawson, A. H. & Isbister, G. K. Serotonin syndrome. BMJ 348, g1626 (2014).
Mawe, G. M. & Hoffman, J. M. Serotonin signalling in the gut-functions, dysfunctions and therapeutic targets. Nat. Rev. Gastroenterol. Hepatol. 10, 473–486 (2013).
Rhoades, J. L. et al. ASICs mediate food responses in an enteric serotonergic neuron that controls foraging behaviors. Cell 176, 85–97.e14 (2019).
Chiang, Y. C., Liao, C. P. & Pan, C. L. A serotonergic circuit regulates aversive associative learning under mitochondrial stress in C. elegans. Proc. Natl. Acad. Sci. USA 119, e2115533119 (2022).
Brissette, B. et al. Chemosensory detection of polyamine metabolites guides C. elegans to nutritive microbes. Sci. Adv. 10, eadj4387 (2024).
Huang, Y. C. et al. A single neuron in C. elegans orchestrates multiple motor outputs through parallel modes of transmission. Curr. Biol. 33, 4430–4445.e6 (2023).
Dag, U. et al. Dissecting the functional organization of the C. elegans serotonergic system at whole-brain scale. Cell 186, 2574–2592.e20 (2023).
Ren, J. et al. Anatomically defined and functionally distinct dorsal raphe serotonin sub-systems. Cell 175, 472–487.e20 (2018).
Ren, J. et al. Single-cell transcriptomes and whole-brain projections of serotonin neurons in the mouse dorsal and median raphe nuclei. eLife 8, e49424 (2019).
Dirksen, P. et al. The native microbiome of the nematode Caenorhabditis elegans: gateway to a new host-microbiome model. BMC Biol. 14, 38 (2016).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Zečić, A., Dhondt, I. & Braeckman, B. P. The nutritional requirements of Caenorhabditis elegans. Genes Nutr. 14, 15 (2019).
Ferris, H., Hieb, W. F. & Ellsworth, C. Dougherty: a pioneer in the selection of Caenorhabditis elegans as a model organism. Genetics 200, 991–1002 (2015).
MacNeil, L. T., Watson, E., Arda, H. E., Zhu, L. J. & Walhout, A. J. Diet-induced developmental acceleration independent of TOR and insulin in C. elegans. Cell 153, 240–252 (2013).
Watson, E., MacNeil, L. T., Arda, H. E., Zhu, L. J. & Walhout, A. J. M. Integration of metabolic and gene regulatory networks modulates the C. elegans dietary response. Cell 153, 253–266 (2013).
Revtovich, A. V., Lee, R. & Kirienko, N. V. Interplay between mitochondria and diet mediates pathogen and stress resistance in Caenorhabditis elegans. PLoS Genet 15, e1008011 (2019).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Zhang, J. et al. A delicate balance between bacterial iron and reactive oxygen species supports optimal C. elegans development. Cell Host Microbe 26, 400–411.e3 (2019).
Han, B. et al. Microbial genetic composition tunes host longevity. Cell 169, 1249–1262.e13 (2017).
Govindan, J. A., Jayamani, E. & Ruvkun, G. ROS-based lethality of Caenorhabditis elegans mitochondrial electron transport mutants grown on Escherichia coli siderophore iron release mutants. Proc. Natl. Acad. Sci. USA 116, 21651–21658 (2019).
Scott, T. A. et al. Host-microbe co-metabolism dictates cancer drug efficacy in C. elegans. Cell 169, 442–456.e18 (2017).
García-González, A. P. et al. Bacterial metabolism affects the C. elegans response to cancer chemotherapeutics. Cell 169, 431–441.e8 (2017).
Tang, B., Xue, K. S., Wang, J. S., Williams, P. L. & Tang, L. Host-microbiota affects the toxicity of Aflatoxin B in Caenorhabditis elegans. Food Chem. Toxicol. 176, 113804 (2023).
Hao, Y. et al. Thioredoxin shapes the C. elegans sensory response to Pseudomonas produced nitric oxide. eLife 7, e36833 (2018).
Sutrina, S. L. et al. Biofilm growth of Escherichia coli is subject to cAMP-dependent and cAMP-independent inhibition. J. Mol. Microbiol Biotechnol. 25, 209–225 (2015).
Wolfgang, M. C., Lee, V. T., Gilmore, M. E. & Lory, S. Coordinate regulation of bacterial virulence genes by a novel adenylate cyclase-dependent signaling pathway. Dev. Cell 4, 253–263 (2003).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Microbiol. 6, 613–624 (2008).
Zheng, D., Constantinidou, C., Hobman, J. L. & Minchin, S. D. Identification of the CRP regulon using in vitro and in vivo transcriptional profiling. Nucleic Acids Res. 32, 5874–5893 (2004).
Rai, A. K. & Mitchell, A. M. Enterobacterial Common Antigen: Synthesis and function of an enigmatic molecule. mBio 11, 01914–01920 (2020).
Benoni, R. et al. Modulation of Escherichia coli serine acetyltransferase catalytic activity in the cysteine synthase complex. FEBS Lett. 591, 1212–1224 (2017).
Poltorack, C. D. & Dixon, S. J. Understanding the role of cysteine in ferroptosis: progress & paradoxes. FEBS J. 289, 374–385 (2022).
Yu, X. & Long, Y. C. Crosstalk between cystine and glutathione is critical for the regulation of amino acid signaling pathways and ferroptosis. Sci. Rep. 6, 30033 (2016).
Hare, E. E. & Loer, C. M. Function and evolution of the serotonin-synthetic bas-1 gene and other aromatic amino acid decarboxylase genes in Caenorhabditis. BMC Evol. Biol. 4, 24 (2004).
Tramonti, A. et al. Knowns and unknowns of vitamin B6 metabolism in Escherichia coli. EcoSal Plus. 9, ESP-0004–2021 (2021).
Feng, M., Gao, B., Ruiz, D., Garcia, L. R. & Sun, Q. Bacterial vitamin B6 is required for post-embryonic development in C. elegans. Commun. Biol. 7, 367 (2024).
Madirolas, G. et al. Caenorhabditis elegans foraging patterns follow a simple rule of thumb. Commun. Biol. 6, 841 (2023).
Lee, K. S. et al. Serotonin-dependent kinetics of feeding bursts underlie a graded response to food availability in C. elegans. Nat. Commun. 8, 14221 (2017).
Salvan, P. et al. Serotonin regulation of behavior via large-scale neuromodulation of serotonin receptor networks. Nat. Neurosci. 26, 53–63 (2023).
Chase, D. L. & Koelle, M. R. Biogenic amine neurotransmitters in C. elegans. WormBook, ed. The C. elegans Research Community, https://doi.org/10.1895/wormbook.1.132.1 (2007).
Yu, J. et al. Parallel pathways for serotonin biosynthesis and metabolism in C. elegans. Nat. Chem. Biol. 19, 141–150 (2023).
Pokala, N., Liu, Q., Gordus, A. & Bargmann, C. I. Inducible and titratable silencing of Caenorhabditis elegans neurons in vivo with histamine-gated chloride channels. Proc. Natl. Acad. Sci. USA 111, 2770–2775 (2014).
Guo, M. et al. Reciprocal inhibition between sensory ASH and ASI neurons modulates nociception and avoidance in Caenorhabditis elegans. Nat. Commun. 6, 5655 (2015).
Cunningham, K. A. et al. AMP-activated kinase links serotonergic signaling to glutamate release for regulation of feeding behavior in C. elegans. Cell Metab. 16, 113–121 (2012).
Zhang, B., Jun, H., Wu, J., Liu, J. & Xu, X. Z. S. Olfactory perception of food abundance regulates dietary restriction-mediated longevity via a brain-to-gut signal. Nat. Aging 1, 255–268 (2021).
Hammarlund, M., Hobert, O., Miller, D. M. & Sestan, N. The CeNGEN project: The complete gene expression map of an entire nervous system. Neuron 99, 430–433 (2018).
Guo, M. et al. Dissecting molecular and circuit mechanisms for inhibition and delayed response of ASI neurons during nociceptive stimulus. Cell Rep. 25, 1885–1897.e9 (2018).
Harris, G. et al. Dissecting the serotonergic food signal stimulating sensory-mediated aversive behavior in C. elegans. PLoS One 6, e21897 (2011).
Goodman, M. B. Mechanosensation. WormBook, ed. The C. elegans Research Community, https://doi.org/10.1895/wormbook.1.62.1 (2006).
Dernovici, S., Starc, T., Dent, J. A. & Ribeiro, P. The serotonin receptor SER-1 (5HT2ce) contributes to the regulation of locomotion in Caenorhabditis elegans. Dev. Neurobiol. 67, 189–204 (2007).
Bechtel, W. & Bich, L. Using neurons to maintain autonomy: learning from C. elegans. Biosystems 232, 105017 (2023).
Li, Z. et al. Dissecting a central flip-flop circuit that integrates contradictory sensory cues in C. elegans feeding regulation. Nat. Commun. 3, 776 (2012).
Liu, H. et al. Reciprocal modulation of 5-HT and octopamine regulates pumping via feedforward and feedback circuits in C. elegans. Proc. Natl. Acad. Sci. USA 116, 7107–7112 (2019).
Xie, Z. et al. The gut-to-brain axis for toxin-induced defensive responses. Cell 185, 4298–4316.e21 (2022).
Filipowicz, A., Lalsiamthara, J. & Aballay, A. TRPM channels mediate learned pathogen avoidance following intestinal distention. eLife 10, e65935 (2021).
Chou, S. H., Chen, Y. J., Liao, C. P. & Pan, C. L. A role for dopamine in C. elegans avoidance behavior induced by mitochondrial stress. Neurosci. Res 178, 87–92 (2022).
Chen, L. T. et al. Neuronal mitochondrial dynamics coordinate systemic mitochondrial morphology and stress response to confer pathogen resistance in C. elegans. Dev. Cell 56, 1770–1785.e12 (2021).
Gusarov, I. et al. Dietary thiols accelerate aging of C. elegans. Nat. Commun. 12, 4336 (2021).
Brooks, K. K., Liang, B. & Watts, J. L. The influence of bacterial diet on fat storage in C. elegans. PLoS One 4, e7545 (2009).
Neve, I. A. A. et al. Metabolite profiling leads to the development of an RNA interference strain for Caenorhabditis elegans. G3 (Bethesda) 10, 189–198 (2020).
Gao, A. W. et al. A sensitive mass spectrometry platform identifies metabolic changes of life history traits in C. elegans. Sci. Rep. 7, 2408 (2017).
Maier, W., Adilov, B., Regenass, M. & Alcedo, J. A neuromedin U receptor acts with the sensory system to modulate food type-dependent effects on C. elegans lifespan. PLoS Biol. 8, e1000376 (2010).
Spiller, R. Serotonin and GI clinical disorders. Neuropharmacology 55, 1072–1080 (2008).
Ha, H. I. et al. Functional organization of a neural network for aversive olfactory learning in Caenorhabditis elegans. Neuron 68, 1173–1186 (2010).
Entchev, E. V. et al. A gene-expression-based neural code for food abundance that modulates lifespan. eLife 4, e06259 (2015).
Harris, G. P. et al. Three distinct amine receptors operating at different levels within the locomotory circuit are each essential for the serotonergic modulation of chemosensation in Caenorhabditis elegans. J. Neurosci. 29, 1446–1456 (2009).
Stern, S., Kirst, C. & Bargmann, C. I. Neuromodulatory control of long-term behavioral patterns and individuality across development. Cell 171, 1649–1662.e10 (2017).
Ben Arous, J., Laffont, S. & Chatenay, D. Molecular and sensory basis of a food related two-state behavior in C. elegans. PLoS One 4, e7584 (2009).
Horvitz, H. R., Chalfie, M., Trent, C., Sulston, J. E. & Evans, P. D. Serotonin and octopamine in the nematode Caenorhabditis elegans. Science 216, 1012–1014 (1982).
Acknowledgements
We thank Steven Flavell, Cassi Estrem, Yun Zhang, Chun-Liang Pan, Luciano Marraffini, Dennis Kim, and members of the Bargmann lab for their comments on the manuscript. We thank Robert Horvitz, William Schafer, Sreekanth Chalasani, and the Caenorhabditis Genetics Center (P40 OD010440) for sharing strains. We thank Dr. Hilda Amalia Pasolli at the Rockefeller University Electron Microscopy Facility (RRID:SCR_017792) for conducting electron microscopy. L.F. was supported by a postdoctoral fellowship from the Kavli Institute for Neural Systems. A.H. was supported by an NIH Predoctoral Fellowship. Y.G. was supported by an NSF Predoctoral Fellowship. This work was supported by a grant from the Chan Zuckerberg Initiative to C.I.B.
Author information
Authors and Affiliations
Contributions
L.F. and C.I.B. designed experiments. L.F. conducted the screen, behavioral, genetic and molecular experiments. J. M-S. and L.Y. conducted behavioral experiments and generated transgenic strains. A.H. generated and validated tph-1 knockout strains. Y.G. generated NSM::HisCl1 transgenic strains. L.F. and C.I.B. analyzed and interpreted results and wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Feng, L., Marquina-Solis, J., Yue, L. et al. Context-dependent serotonin signaling links dietary quality to foraging decisions. Nat Commun 16, 10479 (2025). https://doi.org/10.1038/s41467-025-65491-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65491-8