Abstract
Metazoans utilize the small RNA pathway to regulate gene expression and maintain genome integrity. This pathway directs histone H3 lysine 9 tri-methylation (H3K9me3) or histone H3 lysine 27 tri-methylation (H3K27me3) at target loci to induce transcriptional gene silencing. Interestingly, some small RNAs are generated from genomic loci enriched in H3K9me3 or H3K27me3. However, the transcription mechanism of small RNA precursors from these heterochromatic regions remains unclear. In C. elegans, piRNAs originate from two genomic clusters enriched with H3K27me3 marks, which recruit the H3K27me3 reader UAD-2 and the upstream sequence transcription complex (USTC). Here, we demonstrate that piRNA transcription in C. elegans relies on TAF-12, a subunit of the basal transcription factor IID (TFIID). Depletion of TAF-12 reduces the production of both piRNA precursors and mature piRNAs. TAF-12 interacts with UAD-2 and facilitates piRNA focus formation in germ cell nuclei. We further show that TAF-12 triggers piRNA transcription by recruiting the RNA polymerase II subunit RPB-5, the Mediator complex subunit MDT-8, and the general transcription factors GTF-2F2 and GTF-2H2C. Thus, piRNA transcription within heterochromatic regions depends on the collaboration between histone modification readers, piRNA-specific transcription factors, and core transcription machinery.
Similar content being viewed by others
Introduction
PIWI-interacting RNAs (piRNAs) are functionally conserved in metazoans, while vary in length and sequence among different species. piRNAs associate with Argonaute proteins of the PIWI clade to safeguard genome integrity by suppressing foreign genetic elements at both transcriptional and post-transcriptional levels1,2,3,4,5,6,7,8,9. Besides, the piRNA pathway is involved in fertility, sex determination, viral defense, and transgenerational inheritance10,11,12,13,14,15,16,17,18. In C. elegans, piRNAs are also named 21U-RNAs, given that they are predominantly 21 nucleotides in length and bear a 5’ terminal uridine residue2,19,20,21,22. There are two types of piRNA genes. Type-I piRNA genes are located in two piRNA clusters on chromosome IV and possess the Ruby motif upstream of their transcription start sites. In contrast, type-II piRNAs are much less abundant, often transcribed bidirectionally from the transcription start sites of coding genes, and likely lack the Ruby motif21,22,23.
C. elegans piRNA genes are individually transcribed by RNA polymerase II (Pol II). The transcription of piRNA precursors relies on the upstream sequence transcription complex (USTC), which is composed of PRDE-1, SNPC-4, TOFU-4, and TOFU-524,25,26. The USTC complex coats two piRNA clusters and forms distinct piRNA foci in germ cells. Casein kinase II (CK2) directly phosphorylates the TOFU-4 protein to promote the USTC assembly27. Interestingly, piRNA clusters exhibit signatures of facultative heterochromatin mark histone 3 lysine 27 tri-methylation (H3K27me3)28. H3K27me3 is catalyzed by the Polycomb repressive complex 2 (PRC2), which in C. elegans consists of MES-2, MES-3, and MES-629,30,31. Knockdown of MES-2, MES-3, or MES-6 results in dispersed piRNA foci of the USTC complex and reduced piRNA levels32. UAD-2 recognizes and binds to the H3K27me3 mark via its chromodomain, enhancing the USTC complex’s binding to piRNA clusters32,33. UAD-2 accumulates in nuclear piRNA foci and exhibits properties of liquid-liquid phase separation to facilitate the assembly of piRNA transcription machinery34. Additionally, the chromatin-remodeling factor ISW-1 recruits the USTC complex to organize the local nucleosome environment upstream of individual piRNA genes and maintains high nucleosome density across piRNA clusters32,35. However, the mechanism by which H3K27me3 promotes piRNA production and how piRNA precursors are transcribed from heterochromatic clusters remains mysterious.
After piRNA transcription initiation, the RNA polymerase II subunit RPB-9 recruits the integrator complex to terminate the transcription of piRNA precursors36,37. These precursors are exported from the nucleus and bound by the PICS/PETISCO complex, which is enriched in the E granule, to stabilize piRNA precursors38,39,40,41,42. piRNA precursors are then cleaved at their 5’-terminal by the PUCH complex43. After the removal of the 5’ m7G-cap and the first two nucleotides, piRNA precursors are loaded onto the PIWI protein PRG-12. The 3’-terminal of piRNA precursors is trimmed by the conserved exonuclease PARN-1 and 2’-O-methylated by the RNA methyltransferase HENN1 to generate mature piRNAs44,45,46,47,48,49. The processing and maturation of piRNAs may occur sequentially in distinct perinuclear germ granules42.
In Drosophila melanogaster, most piRNA genes are transcribed from dual-strand piRNA clusters enriched in the heterochromatic H3K9me3 mark50. Rhino, a paralogue of heterochromatin protein-1 (HP1), specifically binds to H3K9me3-enriched chromosomal regions through its chromodomain51,52. The zinc finger protein Kipferl recruits Rhino to guanosine-rich DNA motifs present at piRNA source loci and stabilizes it on chromatin53. Rhino interacts with Deadlock and Cutoff (known as the RDC complex) to promote non-canonical transcription from these loci54,55,56,57. The RDC complex further recruits the germline-specific TFIIA-L paralogue Moonshiner, the transcription factor TFIIA-S, and the TATA-box binding protein (TBP)-related factor TRF2 to initiate piRNA transcription58. Interestingly, the H3K27 methyltransferase E(z) guides Rhino to Kipferl-independent piRNA source loci to regulate transposable element expression and piRNA production in Drosophila germ cells59, suggesting that the mechanism by which heterochromatin marks promote piRNA transcription is evolutionarily conserved across species.
Pol II initiates transcription as part of the pre-initiation complex (PIC), which consists of Pol II, the Mediator complex, and a set of general transcription factors (GTFs), including TFIIA, TFIIB, TFIID, TFIIE, TFIIF, and TFIIH60,61,62. Among these GTFs, the TATA binding protein (TBP), a subunit of TFIID, binds directly to core promoters, facilitating promoter melting and transcription initiation. This process is stabilized by TFIIA and TFIIB through direct contacts with TBP. TFIIB further recruits Pol II, TFIIE, TFIIF, and TFIIH. Finally, Mediator binds to GTFs that occupy the enhancer to yield a functional PIC. However, in C. elegans, most piRNA source loci are located in facultative heterochromatin regions, suggesting that the initiation mechanism of piRNA transcription may differ from that of protein-coding genes.
Here, we identify that the H3K27me3 reader UAD-2 directly interacts with the TFIID transcription factor TAF-12 to drive piRNA transcription. UAD-2 utilizes its C-terminal domain to bridge TAF-12 and TOFU-4. TAF-12 accumulates in piRNA foci, and its localization is dependent on UAD-2 and the USTC complex. TAF-12 contains a histone-fold domain and a conserved C-terminal domain, both of which are essential for TAF-12’s function in piRNA production. Additionally, the RNA Pol II subunit RPB-5, the Mediator complex subunit MDT-8, the TFIIF subunit GTF-2F2, and the TFIIH subunit GTF-2H2C promote piRNA production by forming a transcription complex with TAF-12. Overall, our data provide key insights into the transcription mechanism of piRNAs within heterochromatic regions.
Results
Identification of TAF-12 interacting with UAD-2 to drive piRNA transcription
To elucidate the molecular mechanism of UAD-2 in piRNA transcription, we performed immunoprecipitation (IP) experiments from whole-worm lysates of uad-2::gfp::3xflag animals and identified putative UAD-2-interacting partners through quantitative mass spectrometry (MS). The most enriched protein was TAF-12, a TBP-associated transcription factor (Fig. 1a and Supplementary Fig. 1a). Using FLAG-tagged TAF-12 as bait, we conversely substantiated the interaction between UAD-2 and TAF-12 via the IP-MS method (Fig. 1b). TAF-12 contains a histone-fold domain and is highly conserved in metazoans as a component of the general transcription factor complex TFIID (Fig. 1c), which plays a central role in RNA polymerase II (Pol II)-dependent transcription initiation63,64,65. In contrast to uad-2 mutants, which are viable with developed gonads, taf-12 mutants are embryonically lethal32. Thus, we utilized the auxin-inducible degron (AID) system to specifically deplete TAF-12 proteins in the germline (Fig. 1d). The knockdown of TAF-12 dramatically reduced UAD-2::GFP fluorescence intensity and abolished piRNA focus formation of UAD-2 (Fig. 1e). This also resulted in the reduction of uad-2 mRNA and UAD-2::GFP protein levels (Fig. 1f and Supplementary Fig. 1b, c). The TAF-12::tagRFP (LG II) rescue experiment confirmed that TAF-12 is essential for the formation of UAD-2 condensates (Fig. 1e).
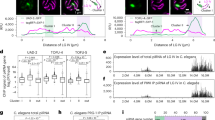
Scatter plots showing (a) GFP-tagged UAD-2 and b 3xFLAG-tagged TAF-12 interaction partners by immunoprecipitation followed by mass spectrometry (IP-MS). Y-axis, Log10 (WD score). X-axis, Molecular weight (kDa) of UAD-2 or TAF-12 co-immunoprecipitating proteins. c Phylogenetic tree of TAF-12 in indicated species. TAF-12 encodes a disordered region (gray) and a histone-fold domain (purple). d Schematic of the taf-12::AID knock-in allele (Chr III, in situ) generated by CRISPR-Cas9 technology in ieSi38 [sun-1p::TIR1::mRuby::sun-1_3’UTR + Cbr-unc-119(+)] (Chr IV) background. e Widefield fluorescence microscopy analysis of an adult hermaphrodite expressing UAD-2::GFP and TAF-12::AID with or without auxin treatment, indicating localization and fluorescence intensity of UAD-2::GFP. TAF-12::tagRFP (LG II) rescued UAD-2::GFP focus formation under auxin treatment. Germlines are outlined by white dashed lines. Scale bar, 50 μm. The zoom-in view showed UAD-2::GFP in meiotic germ cells. Scale bar, 10 μm. n = 6 independently imaged worms over three independent experiments with similar results. f Western blotting analysis of the expression levels of UAD-2::GFP::3xFLAG and β-actin with or without auxin treatment using anti-FLAG and anti-β-actin antibodies. β-actin was used as a loading control. Each experiment has been repeated three times with similar results. Source data are provided as a Source Data file. The relative abundance of g mature piRNAs from individual type-I (n = 4434) and type-II (n = 23,785) piRNA loci and h piRNA precursors (n = 28,219) in young adult hermaphrodites expressing UAD-2::GFP and TAF-12::AID with or without auxin treatment. The central horizontal line within the box represents the median piRNA abundance. Each box displays the interquartile range of the data (between the 25th and 75th percentile). Unpaired Wilcoxon test was used to derive p values. ****p value < 2.2 × 10−16.
In C. elegans, PRG-1 is the only functional PIWI clade Argonaute protein and can be loaded with piRNAs to silence transposable elements2. The expression level and perinuclear localization of PRG-1 depend on piRNA accumulation. Loss of piRNA biogenesis factors, such as PRDE-1, KIN-3, and TOFU-6, dramatically reduces the expression level of PRG-125,27,38. Thus, we utilized gfp::prg-1 animals as a piRNA reporter. The knockdown of TAF-12 disrupted the perinuclear localization of PRG-1 and caused a remarkable reduction in PRG-1 protein levels (Supplementary Fig. 1d, e). Small RNA sequencing further revealed a decrease in both piRNA precursors and mature piRNAs following germline-specific knockdown of TAF-12 (Fig. 1g, h and Supplementary Fig. 1f). Thus, TAF-12 interacts with UAD-2 and is crucial for piRNA production.
piRNA focus formation of TAF-12, UAD-2, and the USTC complex is interdependent
Given its interaction with UAD-2, we hypothesized that TAF-12 is enriched at the nuclear piRNA foci in germ cells, similar to UAD-2 and the USTC complex. To test this, we then inserted a gfp::3xflag tag to the taf-12 genomic locus and examined the expression pattern of TAF-12. In embryos, somatic cells, and germ cells, TAF-12 is constitutively expressed and colocalizes with the histone protein HIS-58 (Supplementary Fig. 2a–c). As expected, TAF-12 colocalizes with UAD-2 at the piRNA foci in germ cells (Fig. 2a).
a Pachytene germ cells from animals expressing UAD-2::GFP and TAF-12::tagRFP. TAF-12 aggregates in piRNA foci and colocalizes with UAD-2. n = 6 independently imaged worms over three independent experiments with similar results. b, c Pachytene germ cells from animals expressing TAF-12::GFP and tagRFP::SYP-1 in the indicated genetic backgrounds. SYP-1 is a component of the synaptonemal complex and is used as a marker to label the chromosomes. n = 6 independently imaged worms over three independent experiments with similar results. d Fluorescence images of representative pachytene germ cells of the indicated animals expressing TOFU-4::GFP, SNPC-4::GFP or PRDE-1::GFP and TAF-12::AID (with or without auxin treatment). Scale bar, 5 μm. n = 4 independently imaged worms over three independent experiments with similar results.
We used the synaptonemal complex protein SYP-1 as a marker to label chromosomes in pachytene cells66 and examined the cellular localization of TAF-12::GFP in uad-2 and ustc mutants. In the absence of UAD-2, TOFU-4, or PRDE-1, TAF-12 failed to form piRNA foci (Fig. 2b, c). The knockdown of TAF-12 also prevented TOFU-4, SNPC-4, and PRDE-1 from forming piRNA foci (Fig. 2d and Supplementary Fig. 2d). The protein level of TAF-12 remained largely unchanged in uad-2 and tofu-4 mutants, but was reduced to approximately 37% in prde-1 mutants (Supplementary Fig. 2e, f). Together, these results reveal that the enrichment of TAF-12, UAD-2, and the USTC complex at piRNA foci is mutually dependent.
UAD-2 binds TAF-12 and the USTC complex through its C-terminal domain
To assess the molecular connections between TAF-12, UAD-2, and the USTC complex, we examined protein-protein interactions among these factors using yeast two-hybrid (Y2H) assays. Consistent with previous work, TOFU-4 interacts with PRDE-1 in the Y2H assay24. In addition, we identified that TAF-12 interacts with UAD-2 as well (Fig. 3a and Supplementary Fig. 3a). Notably, UAD-2 also interacts with TOFU-4 (Fig. 3a and Supplementary Fig. 3a), which supports with the mutual dependency between UAD-2 and the USTC complex for piRNA focus formation.
a Yeast two-hybrid assay to probe protein-protein interactions between UAD-2, TAF-12, and the USTC complex. On nonselective medium (left) all constructs allow growth equally, under selection (right) only strains expressing proteins that interact can grow. Due to the self-activation activity of the DNA-binding domain (BD) tethered TOFU-5 protein, TOFU-5 is excluded from the Y2H experiments. b IUPred3 prediction of UAD-2 using the IUPred3 long disorder option. UAD-2 contains a chromodomain and a C-terminal domain. c The AlphaFold 3-predicted structure of TAF-12 of the indicated species. TAF-12 encodes a histone-fold domain and a C-terminal domain. d, e Yeast two-hybrid assays are shown probing protein-protein interactions between TOFU-4, the UAD-2 chromodomain (UAD-2 chromo), the UAD-2 C-terminal domain (UAD-2 CTD), the TAF-12 histone-fold domain (TAF-12 HFD), and the TAF-12 C-terminal domain (TAF-12 CTD). TOFU-4 lacks of recognizable domains. f Summary of protein–protein interaction experiments between TAF-12, UAD-2, TOFU-4, PRDE-1, SNPC-4, and TOFU-5. g Cartoon of observed protein-protein interactions between TAF-12, UAD-2, and TOFU-4. Data are based on yeast two-hybrid assays using truncation constructs to map interacting domains. CTD C-terminal domain, HFD histone-fold domain.
Besides the chromodomain that recognizes H3K27me3, UAD-2 contains a C-terminal domain (CTD) that may facilitate protein-protein interactions, similar to the chromo-shadow domain (Fig. 3b)54,67. TOFU-4 lacks any recognizable domains. Protein sequence alignment revealed that TAF-12 harbors a histone-fold domain (HFD) and a C-terminal domain (CTD), both of which are highly conserved across diverse species (Supplementary Fig. 3b). The AlphaFold 3-predicted structures of TAF-12 demonstrated that the histone-fold domain and the C-terminal domain of these homologous proteins adopt similar overall structure. The histone-fold domain comprises three tandem α-helices, while the C-terminal domain features a 310 helix (designated as η1), followed by an α-helix (Fig. 3c). The TAF-12 histone-fold domain is crucial for its interaction with the TAF-4 histone-fold domain, which regulates transcriptional activity in germline blastomeres68. Domain mapping experiments showed that the UAD-2 C-terminal domain interacts with both TOFU-4 and the TAF-12 histone-fold domain (Fig. 3d–g and Supplementary Fig. 3c). Thus, UAD-2 serves as a bridge connecting TAF-12 with the USTC complex.
The TAF-12 C-terminal domain is essential for piRNA production
TAF-12 contains an intrinsically disordered region (IDR, residues 1-231), a histone-fold domain (HFD, residues 225-292), and a C-terminal domain (CTD, residues 300-342) (Fig. 4a). Using CRISPR-Cas9 technology, we generated taf-12(ust692/ΔHFD) and taf-12(ust550/ΔCTD) animals (Fig. 4a and Supplementary Fig. 4a). The homozygous taf-12(ΔHFD) animals were embryonically lethal, indicating an essential role of the histone-fold domain. The AlphaFold 3-predicted protein-protein complex structure revealed that the long α4 helix of the TAF-12 histone-fold domain inserts into the angle formed by several α-helices of the UAD-2 C-terminal domain (Fig. 4b, left). The short α3 and α5 helices of the TAF-12 histone-fold domain pack against the α8 helix of the UAD-2 C-terminal domain. Further analysis using PDBsum identified E249, D253, and D260 within the TAF-12 histone-fold domain as key potential interaction residues with the UAD-2 C-terminal domain (Fig. 4b, right). We engineered a taf-12 mutant with arginine substitutions at these three residues and successfully obtained a heterozygous taf-12(E249R;D253R;D260R) allele. However, no homozygous taf-12(E249R;D253R;D260R) mutants were isolated from the progeny of heterozygous animals. These data suggest that the mutation may cause embryonic lethality and that the three residues are essential for TAF-12. Y2H assay revealed that the introduction of E249R;D253R;D260R triple mutations into the TAF-12 histone-fold domain completely blocked its interaction with UAD-2 and the UAD-2 C-terminal domain (Fig. 4c). We also mutated potential interaction residues in UAD-2 to alanine or aspartic acid. Both UAD-2(R569A;R618A;H619A) and UAD-2(R569D;R618D;H619D) retained their interaction with TOFU-4 (Fig. 4d). Yet the UAD-2(R569D;R618D;H619D) mutant disrupted its interaction with TAF-12, whereas UAD-2(R569A;R618A;H619A) remained competent for this interaction. These results suggest that E249, D253, and D260 of TAF-12, along with R569, R618, and H619 of UAD-2, are indispensable for their protein-protein interaction.
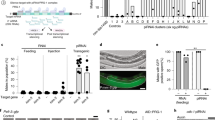
a Top: IUPred3 prediction of TAF-12 using the IUPred3 long disorder option. TAF-12 encodes an intrinsically disordered region (1-231 aa). Bottom: schematic of the taf-12 mutant alleles generated by CRISPR-Cas9 technology. The taf-12(ust692/ΔHFD) allele deletes 1275 bp and triggers embryonic lethality. The taf-12(ust550/ΔCTD) allele deletes 917 bp and the mutants are viable. b The AlphaFold 3-predicted structure of a TAF-12-UAD-2 complex. The interaction active-site residues of the TAF-12 histone-fold domain and the UAD-2 C-terminal domain are magnified in the bottom right. Salt bridges (yellow) and hydrogen bonds (blue) are depicted with dashed lines. Yeast two-hybrid assay to probe protein-protein interactions between UAD-2, TAF-12, the UAD-2 C-terminal domain, the TAF-12 histone-fold domain, and the TAF-12 histone-fold domain(E249R;D253R;D260R) (c) and between UAD-2, TAF-12, TOFU-4, UAD-2(R569A;R618A;H619A), and UAD-2(R569D;R618D;H619D) (d). On nonselective medium (left) all constructs allow growth equally, under selection (right) only strains expressing proteins that interact can grow. e Brood size of the indicated animals at 20 °C and 25 °C, respectively. Bleached embryos were hatched and grown at 20 °C or 25 °C. Then, L4 stage worms were transferred individually onto fresh NGM plates. Wild-type (n = 16) and taf-12(ΔCTD) mutant worms (n = 26) at 20 °C. Wild-type (n = 23) and taf-12(ΔCTD) mutant worms (n = 18) at 25 °C. Data are presented as mean values ± SEM. A two-tailed t-test was performed to determine statistical significance. f Fluorescence images showing localization of UAD-2 in mitotic and meiotic germ cells in indicated backgrounds. Scale bar, 5 μm. n = 6 independently imaged worms over three independent experiments with similar results. g, h Mitotic and meiotic germ cells of animals that express TOFU-4::GFP or PRDE-1::GFP in control and taf-12(ΔCTD) backgrounds. Scale bar, 5 μm. n = 6 independently imaged worms over three independent experiments with similar results. i Boxplots showing type-I (n = 4434) and type-II (n = 23,785) mature piRNA abundance (log10 (reads per million + 1)) in wild type and taf-12(ΔCTD) mutant worms. The central horizontal line within the box represents the median piRNA abundance. Each box displays the interquartile range of the data (between the 25th and 75th percentile). Unpaired Wilcoxon test was used to derive p values. ****p value < 2.2 × 10−16.
Unlike taf-12(ΔHFD) and taf-12(E249R;D253R;D260R) mutants, taf-12(ΔCTD) were viable. taf-12(ΔCTD) animals were fertile but displayed reduced brood sizes at both 20 °C and 25 °C compared to wild-type animals (Fig. 4e). Deletion of the TAF-12 C-terminal domain completely suppressed piRNA focus formation of UAD-2, TOFU-4, and PRDE-1 in meiotic cells and weakened piRNA foci in mitotic cells (Fig. 4f–h). In taf-12(ΔCTD) mutants, uad-2 mRNA levels remained largely unchanged, whereas UAD-2::GFP protein levels were significantly reduced (Supplementary Fig. 4b, c). We constructed animals expressing TAF-12(ΔCTD)::GFP or TAF-12(ΔIDR)::GFP and examined their cellular localization. TAF-12(ΔIDR)::GFP still accumulated at piRNA foci, while TAF-12(ΔCTD)::GFP did not enrich at piRNA foci throughout the germline (Supplementary Fig. 4d–f). Thus, the TAF-12 C-terminal domain is crucial for piRNA focus formation and UAD-2 expression.
To test whether the C-terminal domain and the IDR region are essential for TAF-12’s function in piRNA production, we sequenced piRNA populations from wild-type and taf-12 mutant animals. taf-12(ΔCTD) animals showed a significant reduction in both type-I and type-II mature piRNAs (Fig. 4i and Supplementary Fig. 4g). Similarly, mature piRNA levels were severely reduced in TAF-12(ΔCTD)::GFP mutants (Supplementary Fig. 4h). In TAF-12(ΔIDR)::GFP mutants, piRNA levels were modestly reduced (Supplementary Fig. 4i).
Taken together, these results suggest that TAF-12 may stabilize the piRNA transcription condensate and promote piRNA transcription through its C-terminal domain.
TAF-12 collaborates with MDT-8, RPB-5, GTF-2H2C, and GTF-2F2 to promote piRNA transcription
To further investigate the mechanism underlying piRNA transcription, we performed a candidate-based RNAi screen. These candidates included subunits of the Pol II complex, the Mediator complex, the P-TEFb complex, and a set of general transcription factors (TFIIA, -B, -D, -E, -F, and -H) necessary for Pol II-mediated transcription initiation or regulation (Supplementary Fig. 5a). Among these sixty-five factors, depletion of MDT-8, RPB-5, GTF-2H2C, and GTF-2F2 led to the dispersion of UAD-2 and TAF-12 foci in the nucleus (Fig. 5a, b).
a, b A candidate-based RNAi screen identified transcription factors required for piRNA focus formation of UAD-2 and TAF-12. These candidates include factors of the RNA Pol II, the Mediator complex, the P-TEFb complex and general transcription factors TFIIA, -B, -D, -E, -F, and -H. n > 30. Source data are provided as a Source Data file. c Fluorescence images of representative meiotic germ cells of the indicated animals expressing UAD-2::GFP and AID::RPB-5, AID::GTF-2F2, AID::GTF-2H2C, or AID::MDT-8 (with or without auxin treatment). Scale bar, 5 μm. n = 4 independently imaged worms over three independent experiments with similar results. d The relative abundance of total mature piRNAs (n = 28,219) in young adult hermaphrodites expressing UAD-2::GFP and AID::RPB-5, AID::GTF-2F2, AID::GTF-2H2C, or AID::MDT-8 (with or without auxin treatment). The central horizontal line within the box represents the median piRNA abundance. Each box displays the interquartile range of the data (between the 25th and 75th percentile). Unpaired Wilcoxon test was used to derive p values. ****p value < 2.2 × 10−16. e Meiotic germ cells of animals that express mCherry::PRDE-1 and GFP::GTF-2H2C, GFP::MDT-8, GFP::GTF-2F2, or GFP::RPB-5. GTF-2H2C and MDT-8 aggregate at piRNA foci and colocalize with PRDE-1. The white arrows indicate piRNA foci, the purple arrow indicates an unidentified focus of GFP::GTF-2H2C in germ cells. Scale bar, 5 μm. n = 6 independently imaged worms over three independent experiments with similar results. f Yeast two-hybrid assay to probe for protein-protein interactions between UAD-2, TAF-12, MDT-8, GTF-2F2, GTF-2H2C, and RPB-5. On nonselective medium (left) all constructs allow growth equally, under selection (right) only strains expressing proteins that interact can grow.
MDT-8, RPB-5, GTF-2H2C, and GTF-2F2 are crucial for transcription initiation and PIC assembly69,70,71,72,73,74,75. To verify the function of these factors in piRNA focus formation, we generated in situ AID-tagged mdt-8, rpb-5, gtf-2H2C, and gtf-2F2 animals and confirmed that UAD-2 is unable to form piRNA foci after germline-specific degradation of these factors (Fig. 5c).
We sequenced small RNA populations from control and auxin-treated animals. Germline-specific degradation of MDT-8, RPB-5, GTF-2H2C, and GTF-2F2 resulted in a significant reduction of mature piRNAs (Fig. 5d and Supplementary Fig. 5b–e). Meanwhile, RNAi-knockdown of mdt-8, rpb-5, and gtf-2H2C also disrupted the perinuclear localization of PRG-1 (Supplementary Fig. 5f). We then generated animals expressing GFP-tagged MDT-8, RPB-5, GTF-2H2C, or GTF-2F2 and examined their cellular localization. These proteins were broadly expressed in somatic nuclei (Supplementary Fig. 6a). In germ cells, GTF-2F2 and RPB-5 did not exhibit strong focus formation, whereas MDT-8 and GTF-2H2C accumulated in piRNA foci and colocalized with PRDE-1 (Fig. 5e). Notably, GTF-2H2C formed an additional, unidentified focus in germ cells, which was distinct from the piRNA foci.
Using yeast two-hybrid assays, we examined protein-protein interactions among UAD-2, TAF-12, MDT-8, RPB-5, GTF-2H2C, and GTF-2F2. The analyses revealed that TAF-12, MDT-8, and RPB-5 interact with each other (Fig. 5f and Supplementary Fig. 6b). We also detected the interaction between the TAF-12 histone-fold domain and RPB-5 (Supplementary Fig. 6c). Although MDT-8 interacts with full-length TAF-12, we could not detect MDT-8 interactions with either the histone-fold domain or the C-terminal domain of TAF-12. Notably, MDT-8 showed a moderate interaction with UAD-2 (Fig. 5f and Supplementary Fig. 6b). We concluded that TAF-12 bridges UAD-2 and the core transcription machinery to promote piRNA transcription.
Discussion
In this study, we demonstrated that TAF-12 serves as a crucial molecular link between the USTC complex, UAD-2, and the general transcription machinery at piRNA clusters, which are typically enriched with the facultative histone mark H3K27me328,35. TAF-12 forms piRNA foci in germ cells, and this aggregation relies on both UAD-2 and the USTC complex. The knockdown of TAF-12 results in the loss of both piRNA precursors and mature piRNAs. TAF-12 may initiate piRNA transcription by recruiting MDT-8, RPB-5, GTF-2H2C, and GTF-2F2. Our findings indicate a large piRNA transcription complex comprised of TAF-12, UAD-2, the USTC complex, three general transcription factors, and a subunit of the RNA Pol II (Fig. 6).
(Top) In C. elegans, the transcription of piRNA genes from piRNA clusters is facilitated by the recognition of the heterochromatic histone mark H3K27me3 via the chromodomain protein UAD-2. UAD-2 interacts with the upstream sequence transcription complex (USTC) and recruits the TFIID component TAF-12 to initiate the transcription of piRNA precursors. (Bottom) in D. melanogaster, piRNA transcription from dual-strand piRNA clusters relies on the reader protein of the heterochromatic histone mark H3K9me3 and H3K27me3, Rhino. Deadlock interacts with Rhino and recruits Moonshiner, TRF2, and TFIIA-S to facilitate piRNA transcription. Kipferl binds to G-rich DNA motifs and interacts with Rhino, thereby influencing Rhino’s binding profile.
Although RNAi-mediated knockdown of taf-12 resulted in embryonic lethality, taf-12(ΔCTD) mutants were viable and fertile. The deletion of the TAF-12 CTD disrupted its localization to piRNA foci and reduced mature piRNA levels. Meanwhile, while uad-2 mRNA remained largely unchanged in taf-12(ΔCTD) mutant, the UAD-2 protein level decreased to 40% of control animals. These results suggest that the TAF-12 CTD may indirectly facilitate piRNA production by stabilizing the UAD-2 proteins, and TAF-12 CTD deletion selectively impairs TAF-12 function in the piRNA pathway, while preserving its other functions. Notably, taf-12(ΔCTD) mutants exhibit near-complete sterility at 25 °C, suggesting that these mutants could serve as a useful separation-of-function allele for dissecting CTD-specific roles of TAF-12, particularly those required under elevated temperature conditions.
Our transcriptome and proteome studies indicate that both uad-2 mRNA and UAD-2 protein levels are significantly downregulated upon TAF-12 depletion. In addition, TAF-12 interacts with UAD-2, and their localization to piRNA foci is mutually dependent. Given that TAF-12 is a component of the general transcription machinery, these findings suggest that TAF-12 may act as a transcriptional co-activator of uad-2 gene. Meanwhile, in taf-12(ΔCTD) mutants, the protein level of UAD-2 was significantly reduced, yet the mRNA level of uad-2 remained largely unchanged, suggesting that the interaction between TAF-12 with UAD-2 might also stabilize UAD-2 protein. This dual role raises the possibility that TAF-12 serves as a molecular bridge coupling transcriptional activation of UAD-2 with the spatial assembly of the piRNA transcription machinery. Intriguingly, TAF-12 plays additional functions beyond the piRNA pathway. In germ cell nuclei, TAF-12 is enriched in piRNA foci but also exhibits distinct chromatin-associated localization that appears largely non-overlapping with UAD-2. Consistently, unlike uad-2 mutants, taf-12 null mutants show severe developmental defects, including embryonic lethality. Furthermore, the depletion of TAF-12 induces broad dysregulation of transcription. Previous work reported that in oocytes and early embryos, TAF-12 interacts with TAF-4, also a component of the TFIID complex, via its histone-fold domain, thereby promoting proper nuclear localization of TAF-4 and the release of transcriptional repression in germline precursors68.
The localization of TAF-12, UAD-2, and the USTC complex at piRNA foci occurs in an interdependent manner. Our work cumulatively suggested a working model regarding the mechanism of piRNA focus formation. First, TAF-12 is essential for the expression of uad-2 gene. The auxin-induced TAF-12 degradation strongly reduced the mRNA and protein levels of UAD-2. Similarly, other core transcription factors, including MDT-8, RPB-5, GTF-2H2C, and GTF-2F2, will also be required for the expression of uad-2 gene per se. Therefore, these transcription factors are required for the expression of UAD-2 protein and consequently the accumulation of UAD-2 in piRNA foci. Second, UAD-2 recognizes the H3K27me3 modification and then recruits TAF-12 and other core transcription factors through protein-protein interactions. Thus, in the absence of UAD-2 and the piRNA foci, TAF-12 and the core transcription machinery are unable to be recruited to piRNA foci. Third, our Y2H assay indicated that UAD-2 and TOFU-4 could interact with each other. Although it has not been demonstrated that the USTC complex could directly bind to Ruby motif, the components of the USTC complex contain DNA-binding domains and previous ChIP-seq assay also revealed the association of USTC complex with piRNA genome24. We speculate that UAD-2 and the USTC complex may act coordinately and promote each other to associate with both the H3K27me3- and piRNA-rich genome, which consequently leads to the formation of piRNA foci. Besides, UAD-2 and TAF-12 both contain a large intrinsically disordered region (IDR), which is often associated with liquid-liquid phase separation (LLPS) and the formation of membraneless condensates. Our recent study has identified that UAD-2 may account for the mobility and phase separation ability in the condensation of piRNA transcription machinery34. Whether and how LLPS properties of UAD-2 and TAF-12 facilitate their accumulation at piRNA foci require further investigation.
Our data reveal that piRNA transcription in heterochromatic regions relies on histone modification readers to recruit a specific transcription machinery. Similarly, in S. pombe, the transcription of siRNAs from heterochromatic loci depends on the chromodomain protein Chp176,77,78. Chp1 binds to methylated H3K9 via its chromodomain and tethers the RNAi-induced transcriptional silencing complex (RITS) to heterochromatin, thereby promoting siRNA production79,80,81. In Arabidopsis thaliana, the initial transcription of 24-nt siRNAs is mediated by SHH182,83. SHH1 binds to H3K9me2 via its SAWADEE domain and recruits Pol IV to transcribe precursor RNAs84,85,86. In Drosophila ovarian germ cells, the chromodomain protein Rhino specifically binds to piRNA source loci enriched with H3K9me3 and collaborates with Deadlock, Cutoff, and a Moonshiner-dependent transcription machinery to initiate non-canonical transcription87. These findings suggest that eukaryotes may employ a universally conserved strategy to facilitate small RNA production in heterochromatic regions.
It is striking that TAF-12 in C. elegans and Moonshiner in D. melanogaster exhibit analogous functions during piRNA transcription, although we did not identify sequence similarity between these two proteins. It is generally known that the genome structure of piRNA genes is significantly different among nematodes, flies, and mammals, which hurdled the understanding of the mechanism of piRNA transcription. However, in the past 10 years, the identification of the piRNA transcription machinery of C. elegans and D. melanogaster has gradually suggested a gross similarity between these two organisms. First, piRNA genes in both C. elegans and D. melanogaster are clustered on the genome, which are enriched with heterochromatin marks28,54,88. In C. elegans, most piRNA transcripts originate from two piRNA clusters on chromosome IV, which are enriched with H3K27me389. In Drosophila ovarian germ cells, the majority of piRNA populations are derived from dual-strand piRNA clusters enriched with H3K9me3. A recent study has reported that several dual-strand clusters, such as piRNA clusters 42AB and 38C, are decorated with both H3K9me3 and H3K27me359. Second, the heterochromatin readers play critical roles to recruit core transcription factors to initiate piRNA transcription. In C. elegans, the H3K27me3 reader protein UAD-2, the USTC complex, and the H3K27me3 methyltransferase PRC2 are required for piRNA biogenesis24,25,26,32. Here, we uncovered that UAD-2 recruits the TFIID transcription factor TAF-12 to promote piRNA gene transcription, which serves as a bridge between the upstream piRNA transcription complex and the core transcription machinery through its interactions with the RNA polymerase II subunit RPB-5 and the Mediator complex subunit MDT-8, therefore providing a model how pol II is transcribed from the H3K27me3-enriched heterochromatin region. In Drosophila, the chromodomain protein Rhino is a germline-specific factor crucial for piRNA production54. The distribution pattern of Rhino on piRNA clusters relies on both the zinc-finger domain protein Kipferl and the H3K27me3 methyltransferase E(z)53. Rhino interacts with Deadlock and Cutoff to form the RDC complex. Meanwhile, Deadlock interacts with Moonshiner and recruits it to piRNA clusters58. Moonshiner, a paralog of TFIIA, further recruits other general transcription factors TRF2 and TFIIA-S to form a distinct piRNA transcription machinery and promote bidirectional piRNA transcription. In these two models, UAD-2 and Rhino both encode chromodomain to recognize histone modifications, and are involved in recruiting the core transcription factors to initiate piRNA transcription within heterochromatic regions. In addition, in C. elegans, UAD-2 interacts with TOFU-4 via its uncharacterized C-terminal domain to form a distinct complex with the USTC. In Drosophila, Rhino interacts with Deadlock by its C-terminal chromo-shadow domain to form the RDC complex. Intriguingly, both TOFU-4 and Deadlock are rapidly evolving proteins and lack any recognizable domains. Based on the physical interactions and functions of these piRNA biogenesis factors, we speculate that the UAD-2/USTC complex is functionally analogous to the RDC complex, and TAF-12 is functionally equivalent to Moonshiner. Whether and how much the piRNA transcription mechanism is conserved in other organisms, especially in mammals, require further investigation.
In C. elegans, type-I piRNA genes are mostly transcribed from two discrete piRNA clusters enriched with H3K27me328. Type-II piRNA genes arise from annotated coding gene promoters throughout the genome21. Although PRDE-1 and SNPC-4 are specifically required for type-I piRNA production, UAD-2, TOFU-4, TOFU-5, TAF-12, and the H3K27me3 methyltransferase complex PRC2 are essential for both types of piRNA biogenesis24,25,26,32. Recent studies in Drosophila have shown that the ZAD zinc-finger protein Kipferl interacts with the Rhino chromodomain to define Rhino’s binding profile within H3K9me2- and H3K9me3-enriched piRNA clusters53. And at Kipferl-independent piRNA loci, the H3K27me3 methyltransferase E(z) guides Rhino binding to several piRNA clusters decorated by both H3K9me3 and H3K27me3 modifications59. These findings raise the possibility that additional histone modifications may contribute to specify type-I and type-II piRNA genes and that unidentified UAD-2 cofactors may facilitate its recruitment to dedicated piRNA genes. Determining whether type-II piRNA gene expression depends on particular chromatin modifications, and whether ZnF proteins in C. elegans employ a similar specialized mechanism to regulate the binding pattern of UAD-2 or the USTC complex, will provide vital insights into piRNA production.
Using a biotin ligase-based proximity labeling approach, a recent study identified the AT-hook transcription factor ATTF-6 as crucial for the enrichment of USTC to piRNA clusters and piRNA production90. ATTF-6 also aggregates in piRNA foci and is essential for piRNA focus formation. Based on these findings, we speculate that the C. elegans piRNA genes across piRNA clusters are transcribed as the following steps: UAD-2 recognizes the histone mark H3K27me3 at piRNA clusters via its chromodomain and recruits TAF-12 and TOFU-4 via its C-terminal domain. The USTC complex, containing TOFU-4 and PRDE-1, recognizes piRNA genome, cooperates with ATTF-6, the chromatin remodeling factors ISW-1, and MRG-1 to regulate chromatin accessibility near the transcription start site of piRNA genes35. TAF-12 further recruits other transcription factors, including RPB-5, MDT-8, GTF-2H2C, and GTF-2F2 to promote piRNA transcription.
Although our study has advanced the understanding of piRNA transcription machinery, the method employed to identify factors involved in piRNA transcription still exhibits certain limitations. First, our RNAi-based screen assay was designed to identify factors specifically required for piRNA focus formation and thereby affect piRNA transcription. Consequently, it will miss the factors that are essential for downstream activities in piRNA transcription, other than for piRNA focus formation. For example, the RNA Pol II subunit RPB-9 physically recruits the Integrator complex at piRNA clusters to promote efficient transcription termination, while it is not required for piRNA focus formation of the USTC complex36,37. Second, for some genes, RNAi is not effectively enough to reduce the mRNA and protein levels. To overcome these limitations, we are trying to systematically label transcription factors with fluorescent tags to identify additional piRNA focus-localized transcription factors in germ cells, as well as with minimal AID tags to detect other transcription factors required for piRNA transcription via CRISPR-Cas9 technology.
Methods
Strains
The Bristol strain N2 was used as the standard wild-type strain. All strains were grown at 20 °C unless otherwise specified. The strains used in this study are listed in Supplementary Data 1.
Construction of plasmids and transgenic strains
For the transgenes of transcription factors, endogenous promoter sequences, UTRs, and ORFs of transcription factors were PCR-amplified with the primers listed in Supplementary Data 2. A gfp::3xflag fused to a linker sequence (GGAGGTGGAGGTGGAGCT) was PCR-amplified with the primers 5′- GGAGGTGGAGGTGGAGCTAT -3′ and 5′- CTTGTCATCGTCATCCTTGTAATCGA -3′ from SHG1093 genomic DNA. A tagRFP::3xha fused to a linker sequence (GGAGGTGGAGGTGGAGCT) was PCR-amplified with the primers 5′- GGAGGTGGAGGTGGAGCTATG -3′ and 5′- GTAATCTGGAACATCGTATGGGTAAGCGTAATCTGGAACATCGTATGGGTAGTTGAGCTTGTGCCCG -3′ from SHG2673 (rrf-1(ust364[rrf-1::tagRFP]) I) genomic DNA42. ClonExpress MultiS One Step Cloning Kit (Vazyme C113-02, Nanjing) was used to connect these fragments with vector, which is amplified with 5′- TGTGAAATTGTTATCCGCTGGT -3′ and 5′- CACACGTGCTGGCGTTAC -3′ from pCFJ151. The injection mix contained PDD162 (50 ng/µl), transcription factors repair plasmid (50 ng/µl), pCFJ90 (5 ng/µl), and three sgRNAs (30 ng/µl). The mix was injected into young adult N2 animals. The transcription factors’ transgenes were integrated into the C. elegans genome locus of each gene in situ via a multiple sgRNA-based CRISPR-Cas9 gene editing system91.
Construction of deletion mutants
For gene deletions, triple-sgRNA-guided chromosome deletion was conducted as previously described91. To construct sgRNA expression vectors, the 20 bp unc-119 sgRNA guide sequence in the pU6::unc-119 sgRNA(F + E) vector was replaced with different sgRNA guide sequences. Addgene plasmid #47549 was utilized to express the Cas9 II protein. Plasmid mixtures containing 30 ng/μl of each of the three or four sgRNA expression vectors, 50 ng/μl of the Cas9 II expressing plasmid, and 5 ng/μl pCFJ90 were co-injected into N2 animals. Deletion mutants were screened by PCR amplification and confirmed by sequencing. The sgRNA sequences are listed in Supplementary Data 3.
Microscopy
To image somatic cells, L3 or L4 larvae were immobilized in 0.5 M sodium azide and mounted on 1.5% agarose pads. To image embryos and germ cells, six to ten gravid adults were dissected in 2 μl of 0.4× M9 buffer containing 0.1 M sodium azide on a coverslip and then mounted on freshly made 1.1% agarose pads. Imaging was performed using a Leica THUNDER Imaging System equipped with a K5 sCMOS camera, an HC PL FLUOTAR 100×/1.40-0.70 oil objective, an HC PL FLUOTAR 40×/0.80 objective, and an HC PL FLUOTAR 20×/0.80 objective. Images were taken using Leica Application Suite X software (version 3.7.4.23463). All images in this study are representative of more than 10 animals.
Immunoprecipitation followed by mass spectrometry analysis
IP-MS was conducted as previously reported38. Mixed-stage transgenic worms expressing UAD-2::GFP were collected and resuspended in equal volumes of 2× lysis buffer (50 mM Tris-HCl [pH 8.0], 300 mM NaCl, 10% glycerol, 1% Triton X-100, Roche ®cOmplete EDTA-Free Protease Inhibitor Cocktail, 10 mM NaF, and 2 mM Na3VO4) and lysed using a FastPrep-24 5 G homogenizer. The lysate supernatant was incubated with in-house-prepared anti-GFP beads for 1 h at 4 °C. The beads were then washed three times with cold lysis buffer. The GFP immunoprecipitates were eluted with chilled elution buffer (100 mM glycine-HCl [pH 2.5]). Approximately 1/8 of each eluate was subjected to western blot analysis, while the remainder was precipitated with TCA or cold acetone and dissolved in 100 mM Tris (pH 8.5) containing 8 M urea. Proteins were reduced with TCEP, alkylated with 10 mM IAA, and finally digested with trypsin at 37 °C overnight. LC-MS/MS analysis of the resulting peptides and MS data processing were performed as previously described92. A WD scoring matrix was employed to identify high-confidence candidate interacting proteins92.
Western blotting
Synchronized young adult worms incubated at 20 °C were collected and washed three times with 1× M9 buffer. Samples were stored at −80 °C until use. The worms were suspended in 1× SDS loading buffer and heated in a metal bath at 95 °C for 5–10 min. The suspensions were then centrifuged at 13,000 × g, and the supernatants were collected. Proteins were resolved by SDS-PAGE on gradient gels (10% separation gel, 5% spacer gel) and transferred to a Hybond-ECL membrane. After washing with 1× Tris-buffered saline with Tween-20 (TBST) buffer and blocking with 5% milk-TBST, the membrane was incubated overnight at 4 °C with antibodies. The membrane was washed three times for 10 min each with 1× TBST and then incubated with secondary antibodies at room temperature for 2 h. The membrane was washed thrice for 10 min with 1× TBST and then visualized. Antibodies used for western blotting: anti-FLAG (Mouse monoclonal, Sigma, F1804), 1:1000; anti-β-actin (Rabbit monoclonal, Beyotime, AF5003), 1:5000. Secondary antibodies: HRP-labeled goat anti-rabbit IgG (H + L) (Abcam, ab205718), 1:15,000; HRP-labeled goat anti-mouse IgG (H + L) (Beyotime, A0216), 1:1000.
RNA isolation
Young adult synchronized populations of worms were grown and washed three times with 1× M9 to remove bacterial residues. For RNA extraction, 400 µl of TRIzol Reagent (Ambion, 15596026) was added to a 100 µl worm aliquot, followed by 7–8 cycles of freezing in liquid nitrogen and thawing in a 42 °C water bath. Afterward, 100 µl of DNA/RNA Extraction Reagent (Solarbio LIFE SCIENCES, P1014) was added to the samples, which were then centrifuged for 15 min at 13,000 × g at 4 °C. The supernatant was further treated with 400 µl isopropanol, 400 µl pre-chilled 75% ethanol, and subjected to DNase I digestion (Thermo Fisher Scientific, EN0521). RNA was eluted into 20 µl of nuclease-free water, and each sample was divided into three aliquots for piRNA precursor, mature piRNA, and mRNA library preparation.
FastAP/RppH treatment for piRNA precursors
FastAP treatment of 2 µg of isolated RNA was performed in FastAP buffer using 2 µl of FastAP (Thermo Fisher ScientificTM, EP0654) in a 40 µl reaction. The reaction was incubated at 37 °C for 10 min, followed by heat inactivation for 5 min at 75 °C. The FastAP-treated RNA was extracted and precipitated overnight, following the protocol for decapping treatment with RppH (NEB, #M0356) according to the vendor’s instructions. The resulting RNA was used as input for small RNA library preparation.
Library preparation and sequencing
Small RNAs were subjected to RNA deep sequencing using an Illumina platform (Novogene Bioinformatic Technology Co., Ltd.). Briefly, small RNAs ranging from 15 to 40 nucleotides were gel-purified and ligated to a P7 adapter (5′- AGATCGGAAGAGCACACGTCTGAACTCCAGTCAC -3′) and a P5 adapter (5′- AGATCGGAAGAGCGTCGTGTAGGGAAAGAGTGT -3′). The ligation products were gel-purified, reverse transcribed, amplified, and sequenced using an Illumina Novaseq platform.
Small RNA-seq analysis
The Illumina-generated raw reads were first filtered to remove adapters, low-quality tags, and contaminants to obtain clean reads using fastp. For mature piRNA and pre-piRNA analysis, clean reads ranging from 17 to 35 nt were respectively mapped to mature piRNA regions, pre-piRNA regions, and the C. elegans transcriptome assembly WS243 using Bowtie2 v.2.2.5 with default parameters. The number of reads targeting each transcript was counted using custom Perl scripts. The number of total reads mapped to the transcriptome minus the number of total reads corresponding to sense ribosomal RNA (rRNA) transcripts (5S, 5.8S, 18S, and 26S), and sense protein-coding mRNA reads were used as the normalization number to exclude the possible degradation fragments of sense rRNAs and mRNAs.
piRNA gene annotations
piRNA annotations were downloaded from the piRBase online database (http://www.regulatoryrna.org/database/piRNA). Genomic coordinates of piRNA genes were obtained by SAMtools against the C. elegans ce11 genome assembly. Type II piRNA genes were obtained from a previous publication21. Type I piRNA gene lists were created by filtering the piRBase annotations with type II piRNA genes.
mRNA-seq analysis
The Illumina-generated raw reads were first filtered to remove adapters, low-quality tags, and contaminants to obtain clean reads at Novogene. The clean reads were mapped to the reference genome of WBcel235 via HISAT2 software (version 2.1.0)93. Differential expression analysis was performed using custom R scripts. A twofold-change cutoff was applied when filtering for differentially expressed genes. All plots were drawn using custom R scripts.
RT-qPCR
Synchronized late young adult worms were washed in M9 medium and ground with homogenizer lysis buffer (20 mM Tris-HCl pH 7.5, 200 mM NaCl, 2.5 mM MgCl2 and 0.5% Nonidet P-40). The eluates were incubated with TRIzol reagent (Invitrogen) followed by isopropanol precipitation and DNase I digestion (Qiagen). Complementary DNAs (cDNAs) were generated from RNAs using the HiScript III RT SuperMix for qPCR (Vazyme, R323) according to the vendor’s protocol. qPCR was performed using a LightCycler 96 real-time PCR system (Roche) with LightCycler 480 SYBR green I master (Roche, 04707516001). Levels of eft-3 mRNA were used as internal controls for sample normalization. The data analysis was performed using a ΔΔCt approach.
Candidate-based RNAi screen
RNAi experiments were performed at 20 °C by placing synchronized embryos on feeding plates as previously described32. HT115 bacteria expressing the empty vector L4440 were used as negative controls. Bacterial clones expressing double-stranded RNAs (dsRNAs) were obtained from the Ahringer RNAi library and were sequenced to verify their identity. All feeding RNAi experiments were performed for two generations except for sterile worms, which were RNAi-treated for one generation. Images were collected using a Leica DM4 B microscope.
Yeast two-hybrid assay
The yeast two-hybrid assay was performed according to the manufacturer’s protocol (Clontech user manual 630489). Briefly, the Saccharomyces cerevisiae strain AH109 was grown in YPDA selective medium at 30 °C. Assayed proteins were fused to the activation domain (AD) and DNA-binding domain (BD) of the Gal4 transcription factor and transformed into pGADT7 (AD) and pGBKT7 (BD). Individual colonies of transformed haploids were selected, picked and mated. Upon interaction of two fusion proteins the Gal4 transcription factor is reconstituted and will activate the reporter genes (ADE2, HIS3), which will allow growth on synthetic defined medium lacking Ade, His, Leu and Trp.
Brood size
L4 hermaphrodites were singled onto plates and transferred daily as adults until embryo production ceased and the progeny numbers were scored.
Statistics and reproducibility
Bar graph with error bar in Fig. 4e is presented with mean and standard error of the mean (SEM). Bar graph with error bar in Supplementary Fig. 1c is presented with mean and standard deviation (SD). All of the experiments were conducted with independent C. elegans animals for the indicated number (n) of replicates. No statistical method was used to predetermine sample size. No data were excluded from the analyses. Statistical analysis was performed with the two-tailed Student’s t test or unpaired Wilcoxon tests as indicated.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data reported in this study have been deposited in the Genome Sequence Archive in the National Genomics Data Center, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA: CRA022192). The mass spectrometry proteomics data reported in this paper have been deposited in the iProX repository under accession codes IPX0013654000. Source data are provided with this paper.
References
Mao, H. et al. The Nrde pathway mediates small-RNA-directed histone H3 lysine 27 trimethylation in Caenorhabditis elegans. Curr. Biol. 25, 2398–2403 (2015).
Batista, P. J. et al. PRG-1 and 21U-RNAs interact to form the piRNA complex required for fertility in C. elegans. Mol. Cell 31, 67–78 (2008).
Lee, H. C. et al. C. elegans piRNAs mediate the genome-wide surveillance of germline transcripts. Cell 150, 78–87 (2012).
Shen, E. Z. et al. Identification of piRNA binding sites reveals the argonaute regulatory landscape of the germline. Cell 172, 937–951 (2018).
Vagin, V. V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006).
Aravin, A. et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442, 203–207 (2006).
Girard, A., Sachidanandam, R., Hannon, G. J. & Carmell, M. A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202 (2006).
Zhang, D. et al. The piRNA targeting rules and the resistance to piRNA silencing in endogenous genes. Science 359, 587–592 (2018).
Wu, W. S. et al. Transcriptome-wide analyses of piRNA binding sites suggest distinct mechanisms regulate piRNA binding and silencing in C. elegans. RNA 29, 557–569 (2023).
Kiuchi, T. et al. A single female-specific piRNA is the primary determiner of sex in the silkworm. Nature 509, 633–636 (2014).
Schnettler, E. et al. Knockdown of piRNA pathway proteins results in enhanced Semliki Forest virus production in mosquito cells. J. Gen. Virol. 94, 1680–1689 (2013).
Grentzinger, T. et al. piRNA-mediated transgenerational inheritance of an acquired trait. Genome Res. 22, 1877–1888 (2012).
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012).
Ashe, A. et al. piRNAs can trigger a multigenerational epigenetic memory in the germline of C. elegans. Cell 150, 88–99 (2012).
Luteijn, M. J. et al. Extremely stable Piwi-induced gene silencing in Caenorhabditis elegans. EMBO J. 31, 3422–3430 (2012).
Brennecke, J. et al. An epigenetic role for maternally inherited piRNAs in transposon silencing. Science 322, 1387–1392 (2008).
Wang, G. & Reinke, V. A C. elegans Piwi, PRG-1, regulates 21U-RNAs during spermatogenesis. Curr. Biol. 18, 861–867 (2008).
Yigit, E. et al. Analysis of the argonaute family reveals that distinct argonautes act sequentially during RNAi. Cell 127, 747–757 (2006).
Das, P. P. et al. Piwi and piRNAs act upstream of an endogenous siRNA pathway to suppress Tc3 transposon mobility in the Caenorhabditis elegans germline. Mol. Cell 31, 79–90 (2008).
Bagijn, M. P. et al. Function, targets, and evolution of Caenorhabditis elegans piRNAs. Science 337, 574–578 (2012).
Gu, W. et al. CapSeq and CIP-TAP identify Pol II start sites and reveal capped small RNAs as C. elegans piRNA precursors. Cell 151, 1488–1500 (2012).
Ruby, J. G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127, 1193–1207 (2006).
Billi, A. C. et al. A conserved upstream motif orchestrates autonomous, germline-enriched expression of piRNAs. PLoS Genet. 9, e1003392 (2013).
Weng, C. et al. The USTC co-opts an ancient machinery to drive piRNA transcription in C. elegans. Gene Dev. 33, 90–102 (2019).
Weick, E. M. et al. PRDE-1 is a nuclear factor essential for the biogenesis of Ruby motif-dependent piRNAs in C. elegans. Gene Dev. 28, 783–796 (2014).
Kasper, D. M., Wang, G. L., Gardner, K. E., Johnstone, T. G. & Reinke, V. The SNAPc component SNPC-4 coats piRNA domains and is globally required for piRNA abundance. Dev. Cell 31, 145–158 (2014).
Zhang, G., Zheng, C., Ding, Y. H. & Mello, C. C. Casein kinase II promotes piRNA production through direct phosphorylation of USTC component TOFU-4. Nat. Commun. 15, 2727 (2024).
Beltran, T. et al. Comparative epigenomics reveals that RNA polymerase II pausing and chromatin domain organization control nematode piRNA biogenesis. Dev. Cell 48, 793–810 (2019).
Gaydos, L. J., Wang, W. C. & Strome, S. H3K27me and PRC2 transmit a memory of repression across generations and during development. Science 345, 1515–1518 (2014).
Ketel, C. S. et al. Subunit contributions to histone methyltransferase activities of fly and worm polycomb group complexes. Mol. Cell Biol. 25, 6857–6868 (2005).
Yuzyuk, T., Fakhouri, T. H. I., Kiefer, J. & Mango, S. E. The polycomb complex protein promotes the transition from developmental plasticity to differentiation in embryos. Dev. Cell 16, 699–710 (2009).
Huang, X. et al. A chromodomain protein mediates heterochromatin-directed piRNA expression. Proc. Natl. Acad. Sci. USA 118, e2103723118 (2021).
Hou, X. et al. Systematic characterization of chromodomain proteins reveals an H3K9me1/2 reader regulating aging in C. elegans. Nat. Commun. 14, 1254 (2023).
Zhu, C. et al. piRNA gene density and SUMOylation organize piRNA transcriptional condensate formation. Nat. Struct. Mol. Biol. 32, 1503–1516 (2025).
Paniagua, N., Roberts, C. J., Gonzalez, L. E., Monedero-Alonso, D. & Reinke, V. The upstream sequence transcription complex dictates nucleosome positioning and promoter accessibility at piRNA genes in the germ line. PLoS Genet. 20, e1011345 (2024).
Berkyurek, A. C. et al. The RNA polymerase II subunit RPB-9 recruits the integrator complex to terminate Caenorhabditis elegans piRNA transcription. EMBO J. 40, e105565 (2021).
Beltran, T., Pahita, E., Ghosh, S., Lenhard, B. & Sarkies, P. Integrator is recruited to promoter-proximally paused RNA Pol II to generate piRNA precursors. EMBO J. 40, e105564 (2021).
Zeng, C. et al. Functional proteomics identifies a PICS complex required for piRNA maturation and chromosome segregation. Cell Rep. 27, 3561–3572 (2019).
Rodrigues, R. J. C. et al. PETISCO is a novel protein complex required for 21U RNA biogenesis and embryonic viability. Gene Dev. 33, 857–870 (2019).
Wang, X. et al. Molecular basis for PICS-mediated piRNA biogenesis and cell division. Nat. Commun. 12, 5595 (2021).
Perez-Borrajero, C. et al. Structural basis of PETISCO complex assembly during piRNA biogenesis in C. elegans. Genes Dev. 35, 1304–1323 (2021).
Huang, X. et al. Compartmentalized localization of perinuclear proteins within germ granules in C. elegans. Dev. Cell 60, 1–20 (2024).
Podvalnaya, N. et al. piRNA processing by a trimeric Schlafen-domain nuclease. Nature 622, 402–409 (2023).
Tang, W., Tu, S., Lee, H. C., Weng, Z. P. & Mello, C. C. The RNase PARN-1 trims piRNA 3’ ends to promote transcriptome surveillance in C. elegans. Cell 164, 974–984 (2016).
Montgomery, T. A. et al. PIWI associated siRNAs and piRNAs specifically require the Caenorhabditis elegans HEN1 ortholog henn-1. PLoS Genet. 8, e1002616 (2012).
Billi, A. C. et al. The Caenorhabditis elegans HEN1 ortholog, HENN-1, methylates and stabilizes select subclasses of germline small RNAs. PLoS Genet. 8, e1002617 (2012).
Kamminga, L. M. et al. Differential impact of the HEN1 homolog HENN-1 on 21U and 26G RNAs in the germline of Caenorhabditis elegans. PLoS Genet. 8, e1002702 (2012).
Pastore, B., Hertz, H. L., Price, I. F. & Tang, W. pre-piRNA trimming and 2’-O-methylation protect piRNAs from 3’ tailing and degradation in C. elegans. Cell Rep. 36, 109640 (2021).
Pastore, B., Hertz, H. L. & Tang, W. Pre-piRNA trimming safeguards piRNAs against erroneous targeting by RNA-dependent RNA polymerase. Cell Rep. 43, 113692 (2024).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007).
Klattenhoff, C. et al. The HP1 homolog rhino is required for transposon silencing and piRNA production by dual-strand clusters. Cell 138, 1137–1149 (2009).
Vermaak, D. & Malik, H. S. Multiple roles for heterochromatin protein 1 genes in Drosophila. Annu. Rev. Genet. 43, 467–492 (2009).
Baumgartner, L. et al. The Drosophila ZAD zinc finger protein Kipferl guides Rhino to piRNA clusters. Elife 11, e80067 (2022).
Mohn, F., Sienski, G., Handler, D. & Brennecke, J. The rhino-deadlock-cutoff complex licenses noncanonical transcription of dual-strand piRNA clusters in Drosophila. Cell 157, 1364–1379 (2014).
Pane, A., Jiang, P., Zhao, D. Y., Singh, M. & Schupbach, T. The Cutoff protein regulates piRNA cluster expression and piRNA production in the Drosophila germline. EMBO J. 30, 4601–4615 (2011).
Chen, Y. A. et al. Cutoff suppresses RNA polymerase II termination to ensure expression of piRNA precursors. Mol. Cell 63, 97–109 (2016).
Zhang, Z. et al. The HP1 homolog rhino anchors a nuclear complex that suppresses piRNA precursor splicing. Cell 157, 1353–1363 (2014).
Andersen, P. R., Tirian, L., Vunjak, M. & Brennecke, J. A heterochromatin-dependent transcription machinery drives piRNA expression. Nature 549, 54–59 (2017).
Akkouche, A. et al. Binding of heterochromatin protein Rhino to a subset of piRNA clusters depends on a combination of two histone marks. Nat. Struct. Mol. Biol. 32, 1517–1527 (2025).
Sainsbury, S., Bernecky, C. & Cramer, P. Structural basis of transcription initiation by RNA polymerase II. Nat. Rev. Mol. Cell Biol. 16, 129–143 (2015).
Papai, G. et al. TFIIA and the transactivator Rap1 cooperate to commit TFIID for transcription initiation. Nature 465, 956–960 (2010).
Buratowski, S., Hahn, S., Guarente, L. & Sharp, P. A. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell 56, 549–561 (1989).
Malik, S. & Roeder, R. G. Regulation of the RNA polymerase II pre-initiation complex by its associated coactivators. Nat. Rev. Genet. 24, 767–782 (2023).
Thomas, M. C. & Chiang, C. M. The general transcription machinery and general cofactors. Crit. Rev. Biochem. Mol. Biol. 41, 105–178 (2006).
Naar, A. M., Lemon, B. D. & Tjian, R. Transcriptional coactivator complexes. Annu. Rev. Biochem. 70, 475–501 (2001).
Bilgir, C., Dombecki, C. R., Chen, P. F., Villeneuve, A. M. & Nabeshima, K. Assembly of the synaptonemal complex is a highly temperature-sensitive process that is supported by PGL-1 during Caenorhabditis elegans meiosis. G3 3, 585–595 (2013).
Smothers, J. F. & Henikoff, S. The HP1 chromo shadow domain binds a consensus peptide pentamer. Curr. Biol. 10, 27–30 (2000).
Guven-Ozkan, T., Nishi, Y., Robertson, S. M. & Lin, R. Global transcriptional repression in C. elegans germline precursors by regulated sequestration of TAF-4. Cell 135, 149–160 (2008).
Feaver, W. J., Svejstrup, J. Q., Henry, N. L. & Kornberg, R. D. Relationship of CDK-activating kinase and RNA polymerase II CTD kinase TFIIH/TFIIK. Cell 79, 1103–1109 (1994).
Shiekhattar, R. et al. Cdk-activating kinase complex is a component of human transcription factor Tfiih. Nature 374, 283–287 (1995).
Sun, Z. W. & Hampsey, M. Identification of the gene (SSU71/TFG1) encoding the largest subunit of transcription factor TFIIF as a suppressor of a TFIIB mutation in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 92, 3127–3131 (1995).
Svejstrup, J. Q. et al. Different forms of TFIIH for transcription and DNA repair: holo-TFIIH and a nucleotide excision repairosome. Cell 80, 21–28 (1995).
Zaros, C. et al. Functional organization of the Rpb5 subunit shared by the three yeast RNA polymerases. Nucleic Acids Res. 35, 634–647 (2007).
Schilbach, S. et al. Structures of transcription pre-initiation complex with TFIIH and Mediator. Nature 551, 204–209 (2017).
Nguyen, V. Q. et al. Spatiotemporal coordination of transcription preinitiation complex assembly in live cells. Mol. Cell 81, 3560–3575 (2021).
Kloc, A., Zaratiegui, M., Nora, E. & Martienssen, R. RNA interference guides histone modification during the S phase of chromosomal replication. Curr. Biol. 18, 490–495 (2008).
Reinhart, B. J. & Bartel, D. P. Small RNAs correspond to centromere heterochromatic repeats. Science 297, 1831 (2002).
Cam, H. P. et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 37, 809–819 (2005).
Motamedi, M. R. et al. Two RNAi complexes, RITS and RDRC, physically interact and localize to noncoding centromeric RNAs. Cell 119, 789–802 (2004).
Schalch, T. et al. High-affinity binding of Chp1 chromodomain to K9 methylated histone H3 is required to establish centromeric heterochromatin. Mol. Cell 34, 36–46 (2009).
Partridge, J. F., Scott, K. S., Bannister, A. J., Kouzarides, T. & Allshire, R. C. cis-acting DNA from fission yeast centromeres mediates histone H3 methylation and recruitment of silencing factors and cohesin to an ectopic site. Curr. Biol. 12, 1652–1660 (2002).
Law, J. A., Vashisht, A. A., Wohlschlegel, J. A. & Jacobsen, S. E. SHH1, a homeodomain protein required for DNA methylation, as well as RDR2, RDM4, and chromatin remodeling factors, associate with RNA polymerase IV. PLoS Genet. 7, e1002195 (2011).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Haag, J. R. & Pikaard, C. S. Multisubunit RNA polymerases IV and V: purveyors of non-coding RNA for plant gene silencing. Nat. Rev. Mol. Cell Biol. 12, 483–492 (2011).
Haag, J. R. et al. In vitro transcription activities of Pol IV, Pol V, and RDR2 reveal coupling of Pol IV and RDR2 for dsRNA synthesis in plant RNA silencing. Mol. Cell 48, 811–818 (2012).
Law, J. A. et al. Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1. Nature 498, 385–389 (2013).
Le Thomas, A. et al. Transgenerationally inherited piRNAs trigger piRNA biogenesis by changing the chromatin of piRNA clusters and inducing precursor processing. Genes Dev. 28, 1667–1680 (2014).
Pastore, B., Hertz, H. L. & Tang, W. Comparative analysis of piRNA sequences, targets and functions in nematodes. RNA Biol. 19, 1276–1292 (2022).
Beltran, T. et al. Comparative epigenomics reveals that RNA polymerase II pausing and chromatin domain organization control nematode piRNA biogenesis. Dev. Cell 48, 793–810.e796 (2019).
Wang, Y. H., Hertz, H. L., Pastore, B. & Tang, W. An AT-hook transcription factor promotes transcription of histone, spliced-leader, and piRNA clusters. Nucleic Acids Res. 53, https://doi.org/10.1093/nar/gkaf079 (2025).
Chen, X. et al. Dual sgRNA-directed gene knockout using CRISPR/Cas9 technology in Caenorhabditis elegans. Sci. Rep. 4, 7581 (2014).
Feng, G. et al. Hippo kinases maintain polarity during directional cell migration in Caenorhabditis elegans. EMBO J. 36, 334–345 (2017).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Acknowledgements
We are grateful to the members of the Guang laboratory for their comments and to Dr. Yong Ding’s laboratory for yeast strains. We are grateful to the International C. elegans Gene Knockout Consortium and the National Bioresource Project for providing the strains. Some strains were provided by the CGC, which is funded by the NIH Office of Research Infrastructure Programs (P40 OD010440). This work was supported by grants from the National Natural Science Foundation of China (32230016 to S.G., 32270583 to C.Z., 32300438 to X.H., 32400435 to X.C., and 32470633 to X.F.), the National Key R&D Program of China (2022YFA1302700 to S.G.), the Research Funds of Center for Advanced Interdisciplinary Science and Biomedicine of IHM (QYPY20230021 to S.G.), and the Fundamental Research Funds for the Central Universities awarded to S.G. This study was also supported in part by the China Postdoctoral Science Foundation under Grant Number 2023M733425 awarded to X.H.
Author information
Authors and Affiliations
Contributions
S.G., C.Z., and X.H. conceptualized the research; X.F., X.C., Y.Z., C.Z., and S.G. designed the research; X.H., Y.K., M.L., M.H., J.C., W.W., T.H., and X.S. performed the research; X.H., Y.K., M.L., J.C., T.H., and X.S. contributed new reagents; X.H., Y.K., M.H., and D.X. provided analytic tools and performed bioinformatics analysis; X.H., X.C., X.F., and S.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Heng-Chi Lee and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, X., Kuang, Y., Li, M. et al. The H3K27me3 reader UAD-2 recruits a TAF-12-containing transcription machinery to initiate piRNA expression within heterochromatic clusters. Nat Commun 16, 10538 (2025). https://doi.org/10.1038/s41467-025-65566-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65566-6