Abstract
ASC (Apoptosis-associated Speck-like protein containing a CARD) is a key adaptor protein that assembles inflammasomes by linking sensors such as NLRP3 to effectors like Caspase-1 via its PYD and CARD Death Domains. Due to ASC’s propensity to self-aggregate, most high-resolution structural studies focused on isolated PYD or CARD domains, leaving the atomic basis of full-length ASC assembly unknown. Here we determine atomic-resolution cryo-EM structures of PYD and CARD filaments from full-length ASC, revealing characteristic multitrack bundles composed of alternating ASCPYD and ASCCARD filaments that expose multiple interfaces for flexible assembly and efficient signaling. We further show that Caspase-1 filaments nucleate specifically from the B-end of ASCCARD filaments, and that the interdomain linker modulates bundle formation. The ASC isoform ASCb, with a four-residue linker, adopts a distinct architecture, correlating with reduced Caspase-1 activation efficiency. In ASC⁻/⁻ THP-1 cells, only wild-type ASC, not interface-disrupting mutants, restored ASC speck formation and Caspase-1 activation, underscoring the requirement for intact multitrack bundles. Cryo-electron tomography captures snapshots of higher-order inflammasome structures. These findings collectively define the structural and functional principles by which ASC organizes inflammasomes to amplify immune signaling.
Similar content being viewed by others
Introduction
Innate immune system senses endogenous and exogenous stimuli and elicits innate inflammatory responses, with the inflammasome serving as a central mediator of proinflammatory cytokine maturation1,2,3,4. Inflammasomes are cytoplasmic multimeric protein complexes assembled from sensor, adaptor, and effector proteins. Sensor proteins recognize pathogen-associated molecular patterns (PAMPs) and damage-associated molecular patterns (DAMPs), subsequently recruiting adaptor and effector partners to initiate a cascade of caspase activation, ultimately leading to inflammation and apoptotic or pyroptotic cell death5,6,7.
ASC (Apoptosis-associated Speck-like protein containing a CARD) functions as a primary adapter protein in the inflammasome complex with its two Death-fold Domains: PYrin Domain (PYD) and CAspase Recruitment Domain (CARD). ASC bridges upstream sensors, such as NACHT, LRR and PYD domains-containing protein 3 (NLRP3), and downstream effectors, including CARD-containing Caspase-11. NLRP3 activation leads to the relocation and oligomerization of NLRP3 to interact with ASC to form large, filamentous structures termed “specks” with ~1 μm in diameter visible under fluorescence microscopy8.
For its crucial importance in the body’s defense mechanisms and inflammatory diseases, if not properly regulated, various structural biology methods have been applied to study the ASC speck. Unfortunately, the strong tendency of ASC to aggregate presented a formidable challenge for conventional high-resolution structural studies. Therefore, the majority of prior efforts were focused on individual PYD or CARD domains of ASC in monomeric or filamentous states, based on which models of ASC-mediated inflammasome were proposed9,10,11,12,13,14,15, while structural studies of full-length ASC were limited to a solution NMR16, more recently, a study by negative-staining electron microscopy (ns-EM)17 and a lower-resolution in situ cryo-electron tomography (cryo-ET) study18. ASC speck induced by tagged NLRP3 was recently shown to progressively solidify into condensates, where no long-range ASC filaments were observed19.
Previous studies described full-length ASC as assemblies in which ASCPYD domains form central filaments with ASCCARD domains extending outward and mediating filament crosslinking11,20. Specifically, full-length ASC has been observed to form filamentous bundles by low-resolution ns-EM11,17, in which all filaments were presumed to consist of ASCPYD domains without direct experimental validation. In contrast, fluorescently tagged full-length ASC has also been reported to adopt single or lightly branched filaments based on ns-EM and cryo-ET analyses18,21. These discrepancies in ASC morphology likely stem from the absence of high-resolution atomic structures describing full-length ASC-mediated inflammasome assembly.
In this work, through a series of atomic-resolution cryo-EM structures of full-length ASC, its isoform b (ASCb), and full-length Caspase-1, we reveal the structural basis for the growth of Caspase-1 filaments from pre-existing full-length ASC or ASCb assemblies. Combined with further mutational, kinetic, cellular and cryo-ET studies, we establish the atomic mechanisms underlying full-length ASC-mediated inflammasome assembly. Together, these insights refine prevailing models, highlight principles of supramolecular signaling platform organization, and provide a robust framework for understanding ASC-dependent host defense and for developing precision therapies targeting inflammatory diseases.
Results
Native full-length ASC assembly is altered by its C-terminal protein tag
Considering the conflicting reports on the morphology of full-length ASC in the literature11,17,18,21, filamentous bundles versus single filaments, we hypothesized that the C-terminal fluorescence tag, although widely used to monitor ASC in cells20,22, may alter ASC assembly. To test this, we first performed cryo-EM studies on full-length ASC-HRV-3C-GFP-Strep (Supplementary Figs. 1a and 2a), a full-length ASC construct containing a cleavable C-terminal GFP tag (hereafter referred to as ASC-GFP for simplicity). ASC-GFP assembled into long filaments with irregular appearances (Supplementary Fig. 3). Helical refinement to a final resolution of 3.14 Å revealed that these filaments contained ASCPYD forming the filament core (Supplementary Fig. 3), while ASCCARD and GFP appeared as smear densities on the filament surface (2D classifications in Supplementary Fig. 3), thereby increasing filament thickness to 23.0 ± 3.4 nm, similar to the 22–30 nm diameter reported for ASC-mCerulean18. These findings are consistent with previous reports describing single filaments of full-length ASC with C-terminal mCherry or mCerulean tag as determined by ns-EM or cryo-ET18,21, supporting the conclusion that C-terminal protein tags promote single-filament assembly.
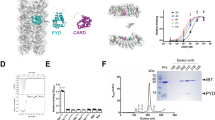
We then removed the C-terminal GFP tag from full-length ASC-GFP by HRV-3C protease treatment to generate tag-free full-length ASC (Supplementary Fig. 1a). Following tag removal, full-length ASC rapidly aggregated, and the resultant pellet was harvested and resuspended in solution. Previous studies reported that ASC polymerization is pH-dependent, with neutral pH favoring filamentous bundle formation17. In our cryo-EM grid preparations, neutral pH (7.8) was also used, and NaCl was found to better preserve a fraction of ASC bundles in solution, enabling high-resolution cryo-EM analysis (Supplementary Fig. 4a). Large aggregates contained overlapping filaments and lacked periodic features, making them unsuitable for tracing, whereas short bundles were well-dispersed and traceable (Supplementary Fig. 4a). Reference-free 2D classification of segments revealed two distinct classes corresponding to PYD and CARD filaments, respectively (2D classification in Supplementary Fig. 4a). Further 3D classification and helical refinement yielded high-resolution cryo-EM structures of ASCPYD and ASCCARD filaments at 3.21 and 2.91 Å, respectively, into which atomic models were built with high confidence (Fig.1a, b, Supplementary Fig. 4a, Supplementary Table 1). These results demonstrate that tag-free full-length ASC assembles into filamentous bundles composed of both ASCPYD and ASCCARD filaments.
a Helical reconstruction of full-length ASCPYD filament overlaid with atomic models shown in different colors; one PYD subunit (orange) is zoomed in, and the density map for helix α6 of PYD is shown. b Helical reconstruction of full-length ASCCARD filament overlaid with atomic models shown in different colors; one CARD subunit (purple) is zoomed in, and the density map for helix α6 of CARD is shown. c 2D classifications for 2-filament segments; inter-filament distances were measured using ImageJ; the assignment of filament was based on the distinct features of 2D averages for PYD and CARD filaments.
Our comparative structural analyses of ASC in the presence and absence of a C-terminal GFP tag clearly demonstrate that the tag disrupts bundle formation, favoring single-filament assembly. We conclude that discrepancies in ASC morphology reported in earlier studies likely result from the use of C-terminal protein tags in certain ASC constructs. Thus, caution is warranted when interpreting data from such studies. Importantly, our high-resolution cryo-EM analyses unequivocally identified the structural compositions of ASC filamentous bundles, achieving a level of detail that was not possible with previous ns-EM approaches11,17.
Structural comparisons between full-length ASC filaments with stand-alone filaments
We performed comprehensive structural analyses of ASCPYD and ASCCARD filaments within full-length ASC to determine whether neighboring filaments in ASC bundles influence filament conformation. When compared to previously published structures of ASCPYD-alone (PDB code: 3J63)11 and ASCCARD-alone filaments (PDB code: 6N1H)15, PYD and CARD filaments in full-length ASC bundles exhibited highly similar conformations, as judged by helical parameters, root mean square differences of Cα atoms (RMSD-Cα), and relative interface orientation (Supplementary Table 2). However, the total interface area for all three types of interfaces (Supplementary Figs. 5a and 6a) in full-length ASCPYD filament increased by 389.9 Å2, while it decreased by 386.6 Å2 in full-length ASCCARD filament (Table 1). These changes suggest tighter packing of PYD subunits and weaker interactions among CARD subunits in filaments formed by full-length ASC. Subtle structural differences were identified at individual interfaces (Supplementary Figs. 5 and 6). Notably, the ASCPYD and ASCCARD filaments of full-length ASC were resolved at higher resolutions than stand-alone filaments (3.2 versus 3.8 Å for ASCPYD filament and 2.9 versus 3.2 Å for ASCCARD filament), raising the possibility that resolution differences and sample preparation methods might have contributed to some of the observed conformational variations.
For the intra-strand type I interface, relative to the ASCPYD-alone structure, two pairs of electrostatic interactions, between the side chains of D6 and K26 and the side chains of E13 and K22, were re-oriented in the full-length ASCPYD structure, resulting in closer interactions between E13 and K22 (Supplementary Fig. 5b). At the inter-strand type II interface, the main chains of residues L78 and Q79 in the α5–α6 loop were rotated by ~180° in the full-length ASCPYD filament, leading to altered side-chain orientations (Supplementary Fig. 5c). These atomic positions were supported by strong electron density, and were consistent with other reported ASCPYD structures (Supplementary Fig. 5e). At the inter-strand type III interface, the α1–α2 loop was shifted by ~1.8 Å and a new salt bridge was formed between E19 and R41 (the shortest distance was 3.3 Å) in the full-length ASCPYD filament (Supplementary Fig. 5d).
In comparison with the ASCCARD-alone filament, no major differences were observed at the type II interface of the full-length ASCCARD filament (Supplementary Fig. 6c). However, the side chains of E130 at the type I interface and E144 at the type III interface of the full-length ASCCARD filament rotated by ~180°, forming new atomic interactions (Supplementary Fig. 6b, d). Moreover, the side chains of E130 and E144, as well as the α3-α4 loop (residues 151–155) at the type III interface, displayed large variations compared to ASCCARD-alone filaments, but aligned well with the 2.0 Å-resolution X-ray structure (PDB code: 6KI0)14 (Supplementary Fig. 6e).
Full-length ASC forms multitrack bundles of alternating PYD and CARD filaments
To determine how ASCPYD and ASCCARD filaments are organized within full-length ASC bundles, we performed 2D classification on segments containing two neighboring filaments. Only pairs of adjacent ASCPYD–ASCCARD filaments were observed, with an inter-filament distance of 10.8 ± 2.6 Å (Fig.1c), whereas no ASCPYD–ASCPYD or ASCCARD–ASCCARD pairs were detected. This finding was also confirmed by inspection of raw micrographs (Supplementary Fig. 4b). In most 2D classes, the orientation of ASCCARD filaments could be readily determined from their structural features, whereas ASCPYD filaments appeared as parallel stripes without directional details, indicating that their orientation relative to ASCCARD filaments may be bi-directional. However, parallel or anti-parallel arrangement of neighboring filaments could not be separated by 2D classification, possibly due to the flexible linker between ASCPYD and ASCCARD filaments that led to subtle variations in their relative orientations. These results support an alternating arrangement of ASCPYD and ASCCARD filaments within full-length ASC bundles, with flexible relative orientations.
To further resolve the organization of ASCPYD/ASCCARD filaments in full-length ASC bundles, we traced each segment back to the original micrographs and determined filament direction by comparing 2D classes with projections from 3D models (Supplementary Fig. 7). The A-end or B-end, corresponding to the end with Type a or Type b interfaces11,15,21, was used to label the filaments on raw micrographs (Supplementary Figs. 7 and 8). Analysis of two-, three- and four-filament full-length ASC bundles is summarized in Table 2. In 20 two-filament bundles, ASCPYD–ASCCARD pairs were aligned in either parallel or anti-parallel orientation with a ratio close to 1:1 (Supplementary Fig. 8, Table 2), indicating that the growth directions of adjacent filaments were not correlated. In 21 three-filament ASC bundles, filaments separated by one filament were always of the same type (ASCPYD or ASCCARD) and shared the same growth direction; notably, 18 bundles (86%) had a central ASCPYD filaments (Table 2). Similarly, in the 18 four-filament bundles, filaments separated by one filament were always of the same type (ASCPYD or ASCCARD), with 81% filaments sharing the same growth direction (Table 2). These observations confirm that side-by-side alignment is the predominant assembly pattern, whereas filament growth direction remains variable.
No densities were observed for the linker connecting the ASCPYD and ASCCARD domains (residues 91–113 in Supplementary Fig. 2a) in our cryo-EM reconstructions. To rule out the possibility that linker degradation during sample preparation caused this absence, we monitored the full-length ASC integrity during HRV-3C protease treatment over a time course and before and after grid preparation, and detected no substantial degradation (Supplementary Fig. 9). Therefore, the absence of densities for the 23-residue ASC linker likely reflects its intrinsic flexibility. This flexibility likely plays a functional role, as the covalent linker tethers the ASCPYD and ASCCARD domains within the same ASC molecule and promotes the local enrichment of one domain type (e.g., ASCCARD) around filaments formed by the other type (e.g., ASCPYD), thereby enabling rapid assembly of ASC condensates with variable filament orientations, generating abundant nucleation sites for efficient Caspase-1 activation.
We further examined the contributions of ASCPYD and ASCCARD domains to ASC bundle formation through targeted mutations disrupting specific filament interfaces: K21E/K22E (ASCPYD type Ib), F59E (ASCPYD type IIa), R41E (ASCPYD type IIIa), D134K (ASCCARD type Ib), R125E (ASCCARD type IIb), and D143K/E144K (ASCCARD type IIIa). Each mutation abolished one interface in the respective filament and resulted in the formation of a single filament in all cases (Supplementary Fig. 10), demonstrating that each interface is indispensable for the multitrack bundle assembly of full-length ASC. As anticipated, the K21E/K22E and R41E mutations, which disrupted type I and III interfaces in the ASCPYD filament, yielded ASCCARD filaments (Supplementary Figs. 11 and 12); conversely, the R125E mutation, which disrupted the type II interface in the ASCCARD filament, yielded an ASCPYD filament (Supplementary Fig. 13). Cryo-EM helical reconstructions confirmed the identity of these mutant filaments (Supplementary Figs. 11–13). Collectively, these results demonstrate that the multitrack architecture of full-length ASC depends on contributions from both ASCPYD and ASCCARD filaments; however, disrupting the interface in one filament type does not impede the formation of the other.
Structure of CARD filament formed by full-length Caspase-1
We determined the cryo-EM structure of the full-length Caspase-1 filament. To prevent self-cleavage, a C285A point mutant of full-length Caspase-1 was used throughout this study (referred to as full-length Caspase-1 hereafter for simplicity). The protein was purified to high purity (Supplementary Fig. 1b) and assembled into filaments that were either heavily or lightly decorated with surface bulges (Supplementary Fig. 14). 2D classification using large boxes to capture these bulges failed to resolve high-resolution details beyond the Caspase-1CARD filaments, indicating that the bulges, likely contributed by the p20 and p10 domains of Caspase-1, were flexible relative to the CARD filament core. Helical refinement of lightly decorated filaments yielded a cryo-EM reconstruction of 2.66 Å resolution (Supplementary Fig. 14, Supplementary Table 1), enabling construction of an atomic structure into the density map (Fig. 2, Supplementary Fig. 2c). Compared with a previously published 4.8 Å-cryo-EM structure of Caspase-1CARD-alone filament (PDB code: 5FNA)23, the full-length Caspase-1CARD filament exhibited identical helical parameters, similar interface orientations, an RMSD-Cα of 1.53 Å (Supplementary Table 2), and a smaller total interface area (decreased by 300.2 Å2 for all three types of interfaces, Table 1). The relatively high RMSD-Cα primarily resulted from the first 14 residues, where the Caspase-1CARD-alone structure adopted a different register compared to our high-resolution Caspase-1CARD structural model (Fig. 2b), as supported by strong electron density (Fig. 2c).
a Helical reconstruction of full-length Caspase-1CARD filament overlaid with atomic models shown in different colors; one CARD subunit (green, in the red dashed box) is zoomed in. b Superposition of full-length Caspase-1CARD subunit (green) with Caspase-1CARD-alone (PDB accession code: 5FNA, in grey), where the Cα traces of α1 and α1A helices have different registers (in the red dashed ellipsoid). c Zoom-in of the Cα traces of α1 and α1A helices (in the red dashed ellipsoid) and electron density for full-length Caspase-1CARD to support its atomic structure. d Location of trimeric interactions in full-length Caspase-1CARD filament; the trimeric interface highlighted in red solid circle is zoomed in with the closest inter-residue interactions labeled. e Superposition of full-length Caspase-1CARD (shown in color) and full-length ASCCARD (grey) filaments in the trimeric region to show the compatibility of the interfaces.
Our atomic-resolution model of full-length Caspase-1CARD filament allowed a detailed analysis of filament interfaces (Table 1, Supplementary Fig. 15, Supplementary Table 2) that were not possible with the earlier 4.8 Å-resolution structure23. Among the three types of interfaces, three residues (N23, E38, and R55) formed an interaction network at the junction of all three interfaces (Fig. 2d, highlighted by two red circles; Supplementary Fig. 15b–d, residues labeled in blue), underscoring their pivotal role in stabilizing the full-length Caspase-1CARD filament. Furthermore, superposition of the full-length ASCCARD filament with the full-length Caspase-1CARD filament revealed that this three-residue interaction network was largely conserved in both sequence and structure: the E130-E144-R160 network in full-length ASCCARD filament closely resembled the N23-E38-R55 network in full-length Caspase-1CARD filament (Fig. 2e). This structural similarity explains how the CARD filaments of ASC and Caspase-1 are inherently compatible at this trimeric interface, facilitating direct CARD-CARD interactions during inflammasome assembly.
Caspase-1 activation by full-length ASC bundles
Activated NLRP3 initiates ASC condensation, which subsequently recruits Caspase-1 to undergo filamentous polymerization and activation. This activation leads to cleavage of gasdermin D (GSDMD), driving pyroptotic cell death, and processing of IL-1β, resulting in the release of inflammatory cytokines. We therefore examined whether Caspase-1 can polymerize from ASCCARD filament within full-length ASC bundles through filamentous CARD-CARD interactions.
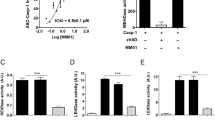
Full-length ASC bundles were prepared as described above and then added to the Caspase-1 monomer solution at different ratios. The pellet fraction obtained after HRV-3C protease cleavage, composed of ASC-Caspase-1 complexes (Supplementary Fig. 1c), was subjected to EM analysis. In ns-EM, full-length ASC bundles exhibited their characteristic multitrack architecture, and full-length Caspase-1 appeared as thicker filaments extending from one end of ASC bundles (Fig. 3a). Isolated full-length Caspase-1 filaments were also observed, arising from spontaneous polymerization, a phenomenon previously reported23.
a Negative staining EM image showing a multitrack full-length ASC bundle connected to a thicker and darker full-length Caspase-1 filament. b Cryo-EM image showing full-length ASCCARD filaments (in purple) connected to full-length Caspase-1CARD (in green) filament based on segment positions of helical reconstruction of corresponding filaments; direction is labelled for filaments where the transition between two filaments is clearly seen. Both negative staining EM and cryo-EM were performed for Caspase-1 samples seeded by different molar ratios of ASC (ASC:Caspase-1 ratio at 1:1, 1:3, 1:10, and 1:50). The representative micrographs displaying good dispersity of filaments are shown. c Superposition of atomic structural models for full-length ASCCARD and full-length Caspase-1CARD filaments.
We also collected cryo-EM data on the same sample. Reconstructions were obtained at 4.41 Å for ASCPYD filaments, and at 4.02 and 4.00 Å for the full-length ASCCARD and Caspase-1CARD filaments, respectively (Supplementary Fig. 16). Mapping segments back to the original micrographs (resolution better than 6 Å as determined by CTF estimation) revealed Caspase-1CARD filaments extending from the ends of full-length ASCCARD filaments (Fig. 3b), with additional representative micrographs shown in Supplementary Fig. 17. Thus, the observed connections between ASCCARD and Caspase-1CARD filaments are supported by sub-atomic resolution data, despite local low resolution or noise in certain regions. Orientation analysis of segments from 2D classification demonstrated that Caspase-1CARD filaments extended from the B-end of ASCCARD filaments (Fig. 3b, c). This observation agrees with a previous hypothesis based on interface area calculation and a CARD-octamer structure (four ASCCARD and four Caspase-1CARD monomers with mutations in the type II interface, connected by linkers)13,15. Our findings provide direct structural evidence that Caspase-1 polymerizes from the B-end of ASCCARD filaments.
To further assess the influence of ASC on Caspase-1 signaling at the cellular level, we generated ASC-/- THP-1 cells and reconstituted them with full-length ASC wild type or mutants carrying single disrupted interfaces in either the ASCPYD or ASCCARD domain (Fig. 4a). Similar experiments have been conducted in immortalized murine bone marrow-derived macrophages (BMDMs) using fluorescently tagged ASC20. In contrast, all our ASC constructs were tag-free, avoiding artifacts that could impair ASC bundle assembly (Supplementary Fig. 3). Differentiated macrophage-like cells were stimulated with LPS, nigericin, and ATP to activate the NLRP3 inflammasome. Caspase-1 activity was assessed by measuring GSDMD cleavage through lactate dehydrogenase (LDH) release and IL-1β processing via cytokine secretion assays by Enzyme-Linked ImmunoSorbent Assay (ELISA).
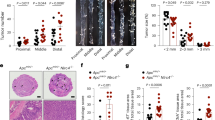
a ASC expression levels in different THP-1 cell lines. THP-1 cell lines were treated with LPS for 3 h to prime the inflammasome, followed by Nigericin/ATP stimulation for 30 minutes to trigger activation. Source data were provided as a Source Data file. Caspase-1 activation was assessed by two complementary assays: b LDH release assay to measure pyroptotic cell death, and c IL-1β ELISA to quantify mature cytokine secretion. Data in b and c are shown as mean ± SD from three independent experiments (n = 3), and source data were provided as Source Data files. d Representative immunofluorescence images of different THP-1 cell lines after stimulation, as those in b and c; nuclei were stained with DAPI (blue); ASCPYD and ASCCARD domains were visualized using different ASC antibodies (green for ASCPYD, red for ASCCARD). In all panels, ASC−/− refers to THP-1 cells with ASC knocked out at both loci, while “ASC−/− +” indicates where either full-length ASC or its various mutants were reintroduced into ASC−/− THP-1 cells.
In ASC-/- THP-1 cells, LDH release and IL-1β secretion were abolished, confirming that loss of ASC completely blocks Caspase-1 activation. Reconstitution with full-length ASC restored LDH and IL-1β release levels to ~67% and ~64% of WT levels, respectively (Fig. 4b, c). Immunofluorescence microscopy using two fluorophore-labeled antibodies against ASCPYD (green color) and ASCCARD (red color) revealed micrometer-sized ASC specks (yellow in merged images in both WT cells and ASC−/− cells reconstituted with full-length ASC (Fig. 4d).
In contrast, ASC−/− THP-1 cells expressing ASC mutants with disrupted interfaces, three in the ASCPYD filament (K21E/K22E, F59E, and R41E) and three in the ASCCARD filament (D134K, R125E, and D143K/E144K), had LDH and IL-1β release levels reduced to baseline (Fig. 4b, c). Correspondingly, immunofluorescence microscopy detected no ASC specks in these mutants. Instead, ASCPYD mutants displayed diffuse signals with filamentous projections, whereas ASCCARD mutants exhibited dispersed signals with tangled filament features (Fig. 4d). These patterns aligned well with our EM analyses of purified mutants (Supplementary Figs. S10–S13), which confirms that disruption of an interface in one filament type does not impair formation of the other filament type. However, the widespread perinuclear ASC signals in these mutants, as opposed to the condensed ASC specks in WT, indicate that correct ASC bundle assembly is essential for speck formation. We noted that ASCPYD and ASCCARD do not co-localize in certain regions of ASC mutant samples, which could arise from simultaneous staining of the sample by two antibodies during the experiment, where one antibody blocks the access of the other antibody to ASC.
Together, these findings support the conclusion that disruption of interfaces in either ASCPYD or ASCCARD filaments prevents ASC bundle consolidation, abolishing Caspase-1 activation.
Atomic cryo-EM structure of full-length ASC isoform b (ASCb)
Compared to full-length ASC, which contains a 23-residue linker between the PYD and CARD domains, its isoform b, ASCb, possesses the same PYD and CARD domains, but with a much shorter four-residue linker. Full-length ASCb was similarly prepared to high purity and subjected to EM study (Supplementary Fig. 1d). Strikingly, unlike the multitrack bundles of 70–80 Å in diameter observed for full-length ASC, full-length ASCb assembled as single filaments with a diameter of 130 Å (Supplementary Fig. 18). The ASCb filament structure was reconstructed to 2.66 Å by helical refinement, with helical parameters of 53.0° and 13.8 Å (Supplementary Fig. 18), which are very close to those of the full-length ASCPYD filament (Supplementary Table 2). The ASCb filament consisted of an inner core formed by the PYD domain, and an outer layer contributed by the CARD domain (Fig. 5a, Supplementary Fig. 2b). The outer layer can be described as six single-file CARD chains, in which each CARD domain interacts only longitudinally with its neighboring CARD domains along the filament axis. Importantly, the linker region in the ASCb filament is well-ordered and resolved, with clear density (Fig. 5b).
a Helical reconstruction of the ASCb filament overlaid with atomic models shown in different colors for different monomers (left panel) or different colors for the PYD domain (in orange), linker (in cyan), or CARD domain (in purple) (right panel). b One ASCb subunit is zoomed in with the linker in a red dashed box, for which the density map is shown. c Superposition of atomic structures of ASCb (PYD, linker, and CARD domain are shown in orange, cyan, and purple colors, respectively) with full-length ASCPYD and full-length ASCCARD domains (in grey). Electron densities for specific regions are shown. d Comparison of type II interface in ASCb filament (cyan for ASCb-1 subunit and yellow for ASCb-2 subunit), and full-length ASCPYD (in grey). The interactions of CARD-1 with PYD-2 and CARD-2 (in red dashed box) are zoomed in on the right. Double dashed lines mark the same interface in ASCb and full-length ASCPYD filaments.
The subunit structures of ASCb were readily superimposable with those of full-length ASCPYD and ASCCARD, except for two notable differences: (1) the N-terminal residue M1 in the PYD domain formed hydrophobic interactions with the side-chain atoms of Q91 in the ordered linker region, and (2) residue H94 in the linker was repositioned to form hydrophobic interactions with multiple residues (I96, W152, and K155) in the CARD domain of full-length ASCb (Fig. 5c).
Compared with the full-length ASCPYD filament, the ASCb filament exhibited a threefold increase in interface area at the type II interfaces (Fig. 5d, Table 1), while the interface areas at the type I or III interface changed only slightly (Supplementary Fig. 19, Table 1). This expansion of the type II interface was primarily due to contributions from the CARD domain, where the CARD domain from one ASCb (termed as CARD-1) interacts with both PYD and CARD domains (termed as PYD-2 and CARD-2, respectively) of an adjacent ASCb subunit (Fig. 5d). Specifically, residues Q160 and E164 in helix α5 in CARD-1 domain interact with R33 and Y36 of PYD-2, while residue K120 of CARD-1 engages in favorable hydrophobic interactions with T63 and H90 of PYD-2. Additionally, residues D115, Y118, and Y127 of CARD-1 wrap around residue Q98 of CARD-2 (Fig. 5d).
These findings reveal that the shortened linker in ASCb enables the CARD domain to establish extensive contacts with adjacent subunits, thereby stabilizing the filament through expanded type II interfaces.
Extension of full-length Caspase-1 filaments from ASCb filaments
To understand how ASCb initiates the polymerization of Caspase-1CARD filaments, we extracted a CARD tetramer structure from one helical turn of the full-length Caspase-1CARD filament determined in this study. This tetramer, stabilized by an intra-strand type III interface, likely represents the initiation phase of Caspase-1CARD filament assembly. The CARD tetramer structure was aligned to the ASCbCARD filament at three distinct positions, corresponding to the A-end, middle, and B-end of the ASCbCARD filament (Supplementary Fig. 20a). Steric clashes were observed when the tetramer was aligned at the A-end or middle position (Supplementary Fig. 20b, c). In contrast, no clash was found when the tetramer was aligned to the CARD domain at ASCb B-end (Supplementary Fig. 20d), indicating that Caspase-1 filaments could grow exclusively from the B-end of ASCb filaments.
We next prepared samples using ASCb filaments as seeds to initiate Caspase-1 filament growth (Supplementary Fig. 1e). Under ns-EM, ASCb filaments appeared as thin, smooth filaments with uniform width, whereas Caspase-1 filaments were thicker with uneven width and a rough surface (Supplementary Fig. 20e, another representative micrograph in Supplementary Fig. 21). In many cases, Caspase-1 filaments were found emerging from the ends of ASCb filaments at angles of ~60°–90° (Supplementary Fig. 20e). Importantly, Caspase-1 filaments were never observed connecting to the side of ASCb filaments, agreeing with our conclusion that Caspase-1 filament growth is restricted to the ASCb B-end.
Superposition of the full-length Caspase-1CARD filament onto the ASCb filament at the B-end predicted an ideal angle of 66.8° between the two filaments (Supplementary Fig. 22a), closely matching the measured angles of ~60° to 90° observed in ns-EM micrographs (Supplementary Fig. 20e). The two filament types are connected by a single ASCb CARD domain. Normal mode analysis of a hypothetical ASCb-Caspase-1CARD complex indicated high structural flexibility of the Caspase-1CARD filament (Supplementary Fig. 22b), suggesting that the angles between the ASCb and Caspase-1CARD filaments vary widely around the ideal value of 66.8°, consistent with our observations (Supplementary Fig. 20e). Moreover, steric hindrance between two Caspase-1CARD filaments growing from adjacent ASCbCARD domains (Supplementary Fig. 22c) likely favors a growth pattern in which Caspase-1CARD filaments extend from alternating ASCbCARD domains (Supplementary Fig. 22d).
Linker between PYD and CARD domains affects full-length ASC bundle formation
Full-length ASC and ASCb differ only in the linkers connecting the PYD and CARD domains, a 23-residue linker in full-length ASC versus a four-residue linker in the latter. Yet, they form distinct 3D architectures as revealed by our structural analyses. To further investigate how linker length and composition influence ASC bundle formation, we made 11 full-length ASC mutants in the linker region: two mutants reducing proline content (P103A/P104A and P97A/P103A/P104A/P110A), two mutants increasing the proline content (G94P/A95P/A96P and A107P/A108P/K109P), four mutants shortening the linker by 5, 10, and 15 residues (Δ95–99, Δ103–107, Δ95–104, and Δ96–110), and three mutants extending the linker by 5, 10, and 15 residues at various positions (A95-GGSGG, A95-GGSGG + A110-GSGGS, and A95-GGSGG + A110-GSGGS + P104-SGGSG).
Dynamic light scattering (DLS) analysis of polymerization revealed that only four mutants exhibited significant (>20%) reductions in bundle formation activity compared to wild-type ASC (Table 3). These included A107P/A108P/K109P that increases proline content near the CARD domain (60% activity), Δ95–104 or Δ96–110 that shorten the linker by 10 or 15 residues (60% or 10% activities, respectively), and A95-GGSGG + A110-GSGGS + P104-SGGSG that extends the linker by 15 residues (60% activities) (Table 3, Supplementary Fig. 23).
These results suggest that the length and composition of the ASC linker are tolerant to changes while maximizing bundle formation during inflammasome activation. Significantly shortening the linker caused a greater disruption of bundle formation function than either increasing its rigidity or length. Consistent with this, ASCb, with a four-residue linker, forms a structurally distinct filament (Fig. 5) and displays a polymerization rate nearly twice that of full-length ASC (Table 3, Supplementary Fig. 23).
Assembly of full-length ASC bundles
To better understand how multitrack full-length ASC bundles assemble into high-order structures, we further characterized full-length ASC samples by cryo-ET. Tomographic reconstructions revealed multitrack ASC bundles embedded in ice at various orientations (Fig. 6a, e). Filamentous bundles were readily annotated by a convolutional neural network-based method (Fig. 6b, c, f, g)24, which facilitated quantitative analysis of ASC bundle assembly. Cryo-ET analysis of neighboring filaments revealed that most were aligned parallel to each other, forming flat multitrack bundles. These flat bundles are stacked randomly at different angles, likely mediated by interactions between exposed ASCPYD or ASCCARD domains on their surfaces, eventually converting into higher-order condensates. Of note, the high-order assemblies from purified full-length ASC observed in this study should be further validated under a cellular context in the future.
a and e Two snapshots for tomography reconstructions of full-length ASC. b and f Segmentation by a convolutional neural network for filaments in reconstructions. c and g Zoom-in of the bundle. d and h Assembly of the bundles in more detail.
In one representative case (Fig. 6a–d, Supplementary Movie 1), three bundles, 2-filament bundle (gold), 3-filament bundle (crimson), and a short bundle (blue), were positioned adjacent to each other. A 4-filament bundle (cyan) stood almost vertically on the gold bundle. Green and magenta bundles are stacked parallel to the gold and crimson bundles, while a 4-filament bundle (salmon) is further stacked on the green bundle, contacting the cyan bundle to form a closed rectangular-prism-like structure.
In another case (Fig. 6e–h, Supplementary Movie 2), a 2-filament bundle (cyan) extended above the 3-filament bundle (crimson) located on one surface of the ice layer, whereas a lime-colored bundle lay in the same plane as the crimson bundle. Additional bundles (blue and salmon) extended from the green bundle, but in a different plane. Further, pink, magenta, tan and gray-colored bundles emanated from the green bundle, while gold and chocolate-colored bundles were located on the opposite ice surface, connected to the green bundle via the magenta bundle. Three additional examples of such assemblies were shown in Supplementary Fig. 24, Supplementary Movies 3–5.
Collectively, these cryo-ET snapshots vividly illustrate the rapid and versatile assembly of high-order inflammasome structures, driven by the unique ability of full-length ASC to form multitrack bundles of alternating ASCPYD and ASCCARD filaments. Based on these structural and functional insights gleaned from our multipronged analyses, we propose a hypothetical model for full-length ASC-mediated inflammasome assembly (Fig. 7 and Supplementary Movie 6).
a NEK7-NLRP3 oligomer nucleates the formation of the ASCPYD filament. b Full-length ASC (ASCPYD and ASCCARD filaments) extends from the central ASCPYD filament (highlighted by the red star (*)) along many possible directions, among which only two dashed arrows are shown as examples. c Amplification of full-length ASC bundle from ASCPYD or ASCCARD filament as indicated by dashed arrows. d Exposed B-ends of ASCCARD filaments provide many initiation sites for Caspase-1CARD filament polymerization. UCSF Chimera was used for figure preparation.
Discussion
Full-length ASC forms a prion-like assembly upon NLRP3 activation, inducing the polymerization of Caspase-1 filament to drive rapid immune responses17,20,21,25,26,27,28. Despite extensive prior efforts17,18,29,30, most structural studies focused on the isolated PYD or CARD domains of ASC in monomeric or filamentous states9,10,11,12,13,14,15, whereas the full-length protein has only been characterized by solution NMR16 and most recently by ns-EM17. Consequently, the atomic structural mechanisms by which full-length ASC mediates inflammasome assembly and amplifies signaling have remained elusive.
By extensively optimizing experimental conditions and EM grid preparations, we successfully imaged well-separated short bundles of full-length ASC by ns-EM and cryo-EM, enabling the determination of high-resolution structures for both ASCPYD and ASCCARD filaments. We discovered that full-length ASC assembles into distinct multitrack bundles composed of alternating ASCPYD and ASCCARD filaments. Notably, ASCPYD filaments are frequently positioned at the center of these bundles, flanked by ASCCARD filaments on each side in 86% of the 3-filament bundles analyzed (Table 2).
Collectively, our data support a hypothetical model for full-length ASC-mediated inflammasome assembly (Fig. 7). When sensor proteins such as NLRP3 polymerize, the ASCPYD domains of full-length ASC assemble into ASCPYD filaments, thereby initiating the subsequent formation of ASCCARD filaments. These ASCCARD filaments are contributed by ASCCARD domains that are either exposed on the surface of already-assembled ASCPYD filaments or from unpolymerized full-length ASC proteins. The elevated local concentration of ASCCARD domains on ASCPYD filament surfaces likely facilitates this process. Subsequently, exposed domains on the surfaces of ASCPYD or ASCCARD filaments may nucleate adjacent complementary filaments, promoting rapid inflammasome expansion and signal amplification. The overall result is that while some full-length ASC proteins contribute both their PYD and CARD domains to two separate filaments, thus maintaining inter-filament connectivity, other full-length ASC proteins incorporate into either ASCPYD or ASCCARD filaments alone. This model remains speculative and warrants further experimental validation.
We also solved the high-resolution cryo-EM structure of full-length Caspase-1CARD filament, revealing that it is structurally compatible with the B-end interface of full-length ASCCARD filament (Fig. 2d, e). Due to steric constraints, Caspase-1CARD filament growth appears restricted to the B-end of existing ASCCARD filaments. These structural insights support a revised model for ASC-mediated inflammasome assembly (Fig. 7).
The inter-domain linker also plays a crucial role in ASC function. Our data show that significantly shortening the linker severely disrupts bundle formation, whereas increasing its rigidity or length has a milder effect (Table 3, Supplementary Fig. 23), supporting earlier observations that linker length influences ASC assembly31,32,33. Consistent with this, the isoform ASCb, with a four-residue linker, loses the characteristic multitrack arrangement of full-length ASC bundles and instead forms tightly packed CARD layers along its inner PYD filament (Fig. 5). Structural modeling suggests that each ASCb filament contains three sites capable of seeding Caspase-1 filament formation (Supplementary Fig. 22d); however, each site provides only a single ASCb CARD domain, likely reducing nucleation efficiency. In marked contrast, full-length ASC not only provides an ordered oligomeric interface at the B-end of ASCCARD filaments for high-efficiency nucleation of Caspase-1CARD filaments, but also offers numerous nucleation sites (Fig.7). This explains why ASCb is less effective at activating Caspase-1 than full-length ASC34,35, despite its faster self-polymerization rate than full-length ASC (Table 3).
The conventional model of ASC speck formation posits that Caspase-1 is recruited to clustered ASCCARD domains decorating ASCPYD filament surface via the 23-residue linker, suggesting that larger ASC specks with more clustered ASCCARD domains would enhance Caspase-1 activity. However, this assumption conflicts with prior observations that ASC speck size inversely correlates with Caspase-1 activity10,36,37. Our findings offer an alternative explanation: the number of potential Caspase-1 nucleation sites depends on the number of accessible B-ends of ASCCARD filaments. As ASC specks enlarge, crowding near the center may drastically reduce the availability of these B-ends, thereby decreasing Caspase-1 activation.
Together, these high-resolution structural studies and functional analyses provide critical insights into the precise and efficient assembly of ASC specks, supporting a refined model for ASC-dependent host defense and inflammatory diseases. Furthermore, they establish a foundation for extracting detailed ultrastructural information from large-scale but intrinsically low-resolution approaches.
Methods
Cell culture
Sf9 cells (Invitrogen, cat. no. 11496-015) were cultured in Sf-900TM II SFM (Gibco) supplemented with 5% (v/v) heat-inactivated Fetal Bovine Serum (FBS) (Gibco). HEK 293S GnTI- cells (ATCC, cat. no. CRL-3022) were grown in Union cell culture medium (Union-Biotech) supplemented with 2% FBS (Gibco). HEK293FT cells (Invitrogen, cat. no. R70007) were cultured in Dulbecco’s Modified Eagle Medium high glucose (DMEM) (Gibco) supplemented with GlutaMax (Gibco), NEAA (Gibco), and 10% FBS. Human monocytic THP-1 cells (National Collection of Authenticated Cell Culture, cat. no. TCHu 57) were cultured in Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco) supplemented with 10% FBS and 0.05 mM β-mercaptoethanol (Sigma).
Cloning, protein expression and purification
Genes encoding human ASC (UniProt entry: Q9ULZ3) and ASCb (UniProt entry: Q9ULZ3-2) were cloned into the pEZT-BM vector, a kind gift from Ryan Hibbs (Addgene plasmid # 74099)38, using Not I and Nhe I restriction sites, with an HRV-3C-protease-cleavable GFP-Strep tag fused at the C-terminus. The resulting constructs were named “ASC-GFP” and “ASCb-GFP”, respectively. Point mutations were introduced into ASC-GFP construct using corresponding oligonucleotides as indicated in Supplementary Table 3.
The gene encoding human Caspase-1 (UniProt entry: P29466) containing the C285A mutation was cloned into the pEZT-BM vector with an HRV-3C-protease-cleavable Strep-GFP tag at the N-terminus (construct was named “GFP-Caspase-1”). The C285A mutation was introduced to prevent self-cleavage of Caspase-1. Bacmids and baculoviruses for all constructs were prepared using Bac-to-Bac system according to the manufacturer’s instructions (Invitrogen).
HEK 293S GnTI− cells grown in Union cell culture medium at a cell density of 1.2 × 106 cells/mL were transduced with 5–10% (v/v) baculovirus, where 10 mM sodium butyrate (Sangon) was supplemented after 12–16 h post-transduction. Cells were harvested after an additional 48 h. Harvested cells were washed twice with PBS, flash-frozen in liquid nitrogen, and stored at −80 °C.
For purification of ASC-GFP, cells were lysed by sonication in Buffer A containing 20 mM HEPES (pH 7.8), 500 mM NaCl, 1 mM TCEP, 4% glycerol, with protease inhibitor cocktail (GenDEPOT) and nuclease inhibitor (Vazyme). Cell lysate was then centrifuged at 40,000×g for 30 min; and the supernatant containing soluble fusion protein was incubated with Strep-Tactin Sepharose resin (IBA) for 1 h. Resin bound with ASC-GFP protein was washed in Buffer A; and ASC-GFP was eluted from resin with Buffer A containing 10 mM d-Desthiobiotin (Millipore Sigma). All purification steps were carried out at 4 °C. ASCb-GFP and GFP-Caspase-1 proteins were similarly purified.
Filament or bundle formation
For full-length ASC bundle formation, eluted ASC-GFP was incubated with HRV-3C protease for 1 h at room temperature with continuous rotation to cleave GFP-Strep tag. Aggregated ASC was collected by gentle centrifugation and washed with Buffer B containing 20 mM HEPES (pH 7.8), 150 mM NaCl and 1 mM TCEP three times. ASC was then resuspended in Buffer B with different salt concentrations. Lower salt concentration, 50–150 mM NaCl, gave a higher proportion of dispersed and regular bundles, where buffer containing KCl gave irregular bundles39. Therefore, a buffer with 150 mM NaCl was used to resuspend the ASC pellet for EM studies. ASCb filament formation followed a similar protocol.
For the formation of Caspase-1 filament, elution of GFP-Caspase-1 was concentrated to 500 μL with a concentrator (30 kDa-MW-cutoff, Millipore Sigma) and treated with HRV-3C protease at a molar ratio of 1:5 to remove the N-terminal Strep-GFP tag for 1 h at room temperature. The reaction mixture was subsequently loaded onto a Superdex 200 10/300 GL size-exclusion column (GE Healthcare) pre-equilibrated with Buffer B. Peak fractions of Caspase-1 were collected for ns-EM and cryo-EM studies.
Filament extension assay
Freshly prepared full-length ASC bundle or ASCb filament was resuspended in Buffer B, serving as the seed. The seed was added to GFP-Caspase-1 at different molar ratios (1:1, 1:3, 1:10, and 1:50), followed by incubation with HRV-3C protease for 1 h at room temperature. Aggregated proteins were collected by gentle centrifugation and washed with Buffer B three times. The sample was resuspended in Buffer B for EM studies. The sample with a ratio of 1:10 gave the best filament concentration for EM studies.
Negative-stain EM grid preparation and data acquisition
About 3 μL of sample was applied to a 200 mesh grid of thin carbon support film (Beijing Zhongjingkeyi Technology Co., Ltd), which was glow-discharged using Solarus II Model 955 (Gatan). After one minute of incubation, 3 μL of uranyl acetate solution (0.2% w/v) was applied to the grid to stain the sample. Solution was blotted away from the side of the grid, and the grid was air-dried for EM analysis. Micrographs of negative-stain grid were captured in a Talos L120C (Thermo Fisher Scientific Inc.) equipped with a 2048 × 2048 BM-ceta camera running at 120 kV.
Cryo-EM grid preparation and data acquisition
About 3 μL sample was applied to a glow-discharged Cryo Matrix amorphous alloy film R1.2/1.3 300 mesh grid (Zhenjiang Lehua Technology Co.) and vitrified using a Vitrobot (Thermo Fisher Scientific Inc.) at 4 °C and 100% humidity. Data collection was performed using a 300 kV Titan Krios G3i (Thermo Fisher Scientific) equipped with a BioQuantum-K3 Summit camera (Gatan) with a 20 eV slit and super-resolution mode. Micrographs were captured at a nominal magnification of 105,000 and a calibrated pixel size of 0.425 Å. Each movie stack received a total electron dosage of 50 e−/Å2 in 2.19-s exposure.
Cryo-EM data processing and model building
Movie frames were aligned with the patch motion correction module, and the contrast transfer function (CTF) for aligned micrographs was estimated using the patch CTF correction module in Cryosparc40. Micrographs with a resolution higher than 5 or 6 Å were kept for filament tracing. Segments were cleaned by 2D classification and heterogeneous classification using Cryosparc40. Helical reconstruction was used to refine filament models in Cryosparc40. Initial atomic models were from reported structures, including PDB codes: 3J63, 6N1H, and 5FNA. The atomic model was manually adjusted in Coot41 and further refined using PHENIX42. Structural models were visualized and prepared for presentation in PyMol43 and Chimera44. ImageJ45, DynDom46, InterProSurf47, and elNemo48 web servers are used for structural analysis.
For data processing of Caspase-1 filament growth seeded by full-length ASC bundles, a total of 1160 micrographs better than 6 Å resolution after patch motion correction were chosen for filament tracing. Several rounds of 2D classification were performed to remove junk in the dataset, and 377,345 segments were kept. By heterogeneous refinement using high-resolution models of ASCPYD, ASCCARD, and Caspase-1CARD as initial models (lowpass at 8 Å), segments were classified into three classes. The separation of segments of different filaments was confirmed by 2D classification: ASCPYD showed stripe-like features, ASCCARD showed spiky-caterpillar-like features, and Caspase-1CARD showed smooth-caterpillar-like features. The following 3D classification led to final helical reconstruction of full-length ASCPYD filament at 4.41 Å for 50,115 segments, full-length ASCCARD filament at 4.02 Å for 133,105 segments, and full-length Caspase-1CARD filament at 4.00 Å for 194,125 segments.
Determination of filament direction
For filament direction determination, the procedure was similar to that used for NLRP3-PYD filament21, with some modifications. For each filament after helical refinement in Cryosparc40, corresponding segments were converted to Relion49 using the program csparc2star.py50 followed by 2D classification. For a specific filament to be analyzed, the information, including segment coordinates, in-plane rotation angle (ψ), and 2D class to which the segment belongs, was retrieved from the file after 2D classification in Relion49. By comparing the 2D class with the 3D model, the A-end or B-end was determined by manual inspection based on the structural feature of the PYD or CARD filament. The 2D class was rotated by the in-plane rotation angle to illustrate the orientation of a specific segment. This orientation determination was performed on multiple segments of a filament in order to get a consistent filament direction. For some filaments, this method gave results of mixed filament directions, which may be caused by the inaccuracy in 2D classification or the low image quality of local regions. In this situation, the filament direction was labeled as a question mark in the raw micrograph and considered as an undetermined filament direction.
Cryo-electron tomography of full-length ASC
BSA-coated gold solution was mixed with the full-length ASC sample at a ratio of 1:3, and was loaded on a glow-discharged R1.2/1.3 300 mesh grid (Zhejiang Lehua Technology Co.), followed by vitrification with Vitrobot (Thermo Fisher Scientific Inc.) at 4 °C and 100% humidity. Tilt-series were collected using SerialEM51 at a nominal 42,000 magnification and calibrated pixel size of 2.17 Å, with a total dose of 100 e−/A2 over a tilt range of −50° to +50° with 2° increments, and a nominal defocus of −6 μm. Raw movies were aligned using MotionCor252, and reconstruction was performed by weighted back projection in AreTomo53, and was further smoothed in IMOD54 with Gaussian kernels (“clip smooth” command). Totally 54 sets of tilt data were collected, where 42 reconstructions showed clear features for multitrack bundles, and 12 reconstructions showed heavy aggregates or bad quality after manual inspection and were not used in further analyses. The 42 reconstructions were further processed by normalization and clamping in EMAN2 (“normalize” and “threshold.clampminmax” command). Convolutional neural networks in EMAN224 were used to annotate the filament in full-length ASC reconstructions. Specifically, each filament segment was considered as the feature of interest (FOI), and picked by a pen tool in EMAN2 with an appropriate size to cover the single filament. Several tens of FOIs were trained by the convolutional neural network method in EMAN2. The trained features were then applied to the whole tomogram. Filamentous bundles were further manually segmented and visualized in UCSF Chimera44 and ChimeraX55.
Measurement of full-length ASC polymerization kinetics in vitro
Protein samples were centrifuged at 24,000×g at 4 °C for 10 minutes before the experiment. Protein concentration was adjusted to 4 μM with buffer containing 20 mM HEPES (pH 7.8), 500 mM NaCl, 1 mM TCEP, and 4% glycerol. Polymerization was initiated by the addition of HRV-3C protease to cleave off the GFP-Strep tag at the C-terminus. Polymerization of 400 μL protein sample was monitored by dynamic light scattering with NanoBrook Omni instrument in a disposable UV-transparent plastic cuvette (LABGIC). Data were acquired with an interval of 60 s at 25 °C until the scattering intensity reached a plateau.
The measurement of count rate (scattering intensity in the unit of kcps) over time displayed a curve of three periods: (1) incubation period where residual or background intensity was measured when protein did not reach its critical concentration yet; (2) logistic growth period where increasing intensity was measured during the polymerization; and (3) plateau period where intensity did not increase significantly. The maximum intensity of different measurements for the same sample was found to be different, possibly due to bundles of various sizes. Data points of initial signal fluctuations (because of the system instability) and later plateau fluctuations (from non-uniform bundles and aggregates) were removed. Background intensity was subtracted from all data points to obtain the net intensity increase. The time-dependent intensity growth was fitted independently for each measurement by a logistic growth equation in Prism 10.2.3 (GraphPad Software), I(t) = I∞/(1 + e−k(t−ts)), where I(t) was the signal at time t, k was the growth rate or steepness of the curve, and ts was the sigmoid midpoint. Measurement for each sample was performed at least on two different days, and at least three times on each day, and the polymerization rate k was compared for different proteins.
Generation of ASC-/- THP-1 cells by CRISPR/Cas9 genome editing
The lentiCRISPRv2 plasmid, a kind gift from Dr. Feng Zhang56 (Addgene plasmid # 52961), was used to express Cas9 and guide RNA (gRNA). Three gRNA sequences were designed using CHOPCHOP, and the most efficient sequence was: 5’-TCTTGAGCTCCTCGGCGGTC-3’.
HEK293FT cells were transfected with the lentiCRISPRv2 plasmid, along with two helper plasmids, psPAX2 (Addgene plasmid # 12260) and pMD2.G (Addgene plasmid # 12259), in the presence of Lipofectamine 3000 (Invitrogen) to produce lentivirus. The cell culture medium containing lentiviral particles was collected at 48 and 72 h post-transfection. The viral supernatant was subsequently filtered through a 0.45 μm filter (PALL), and concentrated using 4×PEG8000/NaCl solution (Sigma), with the final concentrations of PEG8000 and NaCl at 10% (w/v) and 0.3 M, respectively. THP-1 cells were transduced by spinoculation with the concentrated lentivirus at 1200×g for 1 h in the presence of 8 μg/mL polybrene (Santa Cruz Biotechnology), followed by a three-day recovery period. Stably transduced cells were selected with 3 μg/mL of puromycin (Gibco) for a duration of 6 days and subsequently subjected to limiting dilution to isolate single-cell clones, which were expanded to establish clonal cell lines. The ASC−/− cells were validated through Sanger sequencing using oligonucleotides specified in Supplementary Table 3 and Western Blot analysis.
Transduction of ASC and its mutants into ASC−/− THP-1 cells
Both full-length ASC and its mutants were subcloned into the pLenti vector (Addgene plasmid # 52962) via Xba I and BamH I restriction sites. Recombinant lentiviruses were produced from these constructs and used to transduce ASC−/− THP-1 cells as described above. Stable cell lines were selected with blasticidin (Gibco) at 15 μg/mL for 6 consecutive days, and ASC expression levels were systematically verified by Western Blot analysis. Uncropped and unprocessed scans of blots were provided as Source-data-Fig.4a.pptx.
Inflammasome stimulation
Prior to THP-1 cells seeding, six-well cell culture plates were coated with 0.01% poly-l-ornithine solution (Sigma) overnight at 37 °C. Cells were plated at a density of 6 × 105 cells/mL (2 mL/well) in complete medium containing 100 ng/mL phorbol 12-myristate 13-acetate (PMA) (InvivoGen) and differentiated into macrophage-like cells in 48 h. For inflammasome stimulation, cells were primed with 200 ng/mL LPS (Sigma) in serum-free RPMI 1640 medium for 3 h, followed by co-treatment with 10 μM nigericin (GLPBIO) and 5 mM ATP (pH 7.4, Sigma) for 30 minutes. For immunofluorescence microscopy, cells were further treated with 20 μM Z-VAD-FMK (APExBio, Caspase-1 inhibitor) for 1 h.
Cell death and IL-1β ELISA assay
Different THP-1 ASC cells were seeded at 105 cells/mL into 96-well plates. After the indicated inflammasome stimulation treatments, culture supernatants were collected for analysis. Cell death was quantified using the LDH Cytotoxicity detection kit (Sigma), and IL-1β release was measured by ELISA (BD Biosciences). Source data were provided as Source-data-Fig.4b.xlsx and Source-data-Fig.4c.xlsx, respectively.
To normalize for spontaneous cell lysis, the percentage of cell death was calculated as follows:
Immunofluorescence microscopy
THP-1 cells were seeded onto round coverslips in 24-well culture plates. Following inflammasome stimulation, cells were washed twice with PBS and fixed with 4% paraformaldehyde (PFA) (Aladdin) for 20 min. Cells were then permeabilized and blocked in PBS containing 0.1% Triton X-100 and 5% BSA for 1 h at room temperature. Subsequently, cells were incubated overnight at 4 °C with ASCPYD antibody (F-9) Alexa Fluor 488 and ASCCARD antibody (B-3) Alexa Fluor 594 (Santa Cruz Biotechnology) diluted 1:100 in SignalUp primary antibody dilution buffer (Beyotime). Nuclei were counterstained with 4’,6-diamidino-2-phenylindole (DAPI) (Beyotime). Images were acquired using a Zeiss LSM980 confocal fluorescence microscope equipped with a Plan-Apochromat ×63/1.40 oil DIC M27 objective lens. Z-stack digital imaging was performed using ZEN imaging software 3.7 (Carl Zeiss, GmbH).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this paper are available from the corresponding authors upon reasonable request. The Cryo-EM density maps and atomic coordinates have been deposited in Electron Microscopy Data Bank (EMDB) and Protein Data Bank (PDB) under accession codes EMDB-66385 and 9WZ8 (ASC-GFP), EMDB-66381 and 9WZ4 (ASCPYD filament of full-length ASC), EMDB-66382 and 9WZ5 ASCPYD (ASCCARD filament of full-length ASC), EMDB-66387 and 9WZB (K21E/K22E mutant of full-length ASC), EMDB-66388 and 9WZC (R41E mutant of full-length ASC), EMDB-66391 and 9WZD (R125E mutant of full-length ASC), EMDB-66383 and 9WZ6 (full-length Caspase-1), EMDB-66384 and 9WZ7 (full-length ASCb), EMDB-66401 and 9WZG (ASCPYD filament of full-length ASC+full-length Caspase-1), EMDB-66402 and 9WZH (ASCCARD filament of full-length ASC+full-length Caspase-1), and EMDB-66403 and 9WZI (Caspase-1CARD filament of full-length ASC+full-length Caspase-1). Source data are provided with this paper.
Code availability
Custom code based on disparticle.py from the pyem program was used to display filament locations on the micrograph. The code has been deposited in Zenodo under entry 17234644.
References
Latz, E., Xiao, T. S. & Stutz, A. Activation and regulation of the inflammasomes. Nat. Rev. Immunol. 13, 397–411 (2013).
Zheng, D., Liwinski, T. & Elinav, E. Inflammasome activation and regulation: toward a better understanding of complex mechanisms. Cell Discov. 6, 36 (2020).
Lamkanfi, M. Emerging inflammasome effector mechanisms. Nat. Rev. Immunol. 11, 213–220 (2011).
Rathinam, V. A. & Fitzgerald, K. A. Inflammasome complexes: emerging mechanisms and effector functions. Cell 165, 792–800 (2016).
Fu, J. & Wu, H. Structural mechanisms of NLRP3 inflammasome assembly and activation. Annu. Rev. Immunol. 41, 301–316 (2023).
Swanson, K. V., Deng, M. & Ting, J. P. The NLRP3 inflammasome: molecular activation and regulation to therapeutics. Nat. Rev. Immunol. 19, 477–489 (2019).
de Alba, E. Structure, interactions and self-assembly of ASC-dependent inflammasomes. Arch. Biochem. Biophys. 670, 15–31 (2019).
Masumoto, J. et al. ASC, a novel 22-kDa protein, aggregates during apoptosis of human promyelocytic leukemia HL-60 cells. J. Biol. Chem. 274, 33835–33838 (1999).
Liepinsh, E. et al. The death-domain fold of the ASC PYRIN domain, presenting a basis for PYRIN/PYRIN recognition. J. Mol. Biol. 332, 1155–1163 (2003).
Yu, T. G. et al. Oligomeric states of ASC specks regulate inflammatory responses by inflammasome in the extracellular space. Cell Death Discov. 9, 142 (2023).
Lu, A. et al. Unified polymerization mechanism for the assembly of ASC-dependent inflammasomes. Cell 156, 1193–1206 (2014).
Gong, Q. et al. Structural basis for distinct inflammasome complex assembly by human NLRP1 and CARD8. Nat. Commun. 12, 188 (2021).
Robert Hollingsworth, L. et al. Mechanism of filament formation in UPA-promoted CARD8 and NLRP1 inflammasomes. Nat. Commun. 12, 189 (2021).
Xu, Z. et al. Homotypic CARD-CARD interaction is critical for the activation of NLRP1 inflammasome. Cell Death Dis. 12, 57 (2021).
Li, Y. et al. Cryo-EM structures of ASC and NLRC4 CARD filaments reveal a unified mechanism of nucleation and activation of caspase-1. Proc. Natl Acad. Sci. USA 115, 10845–10852 (2018).
de Alba, E. Structure and interdomain dynamics of apoptosis-associated speck-like protein containing a CARD (ASC). J. Biol. Chem. 284, 32932–32941 (2009).
Nambayan, R. J. T., Sandin, S. I., Quint, D. A., Satyadi, D. M. & de Alba, E. The inflammasome adapter ASC assembles into filaments with integral participation of its two death domains, PYD and CARD. J. Biol. Chem. 294, 439–452 (2019).
Liu, Y. et al. Cryo-electron tomography of NLRP3-activated ASC complexes reveals organelle co-localization. Nat. Commun. 14, 7246 (2023).
Wang, J. et al. Human NLRP3 inflammasome activation leads to formation of condensate at the microtubule organizing center. Preprint at bioRxiv https://doi.org/10.1101/2024.09.12.612739 (2024).
Dick, M. S., Sborgi, L., Ruhl, S., Hiller, S. & Broz, P. ASC filament formation serves as a signal amplification mechanism for inflammasomes. Nat. Commun. 7, 11929 (2016).
Hochheiser, I. V. et al. Directionality of PYD filament growth determined by the transition of NLRP3 nucleation seeds to ASC elongation. Sci. Adv. 8, eabn7583 (2022).
Soriano-Teruel, P. M. et al. Identification of an ASC oligomerization inhibitor for the treatment of inflammatory diseases. Cell Death Dis. 12, 1155 (2021).
Lu, A. et al. Molecular basis of caspase-1 polymerization and its inhibition by a new capping mechanism. Nat. Struct. Mol. Biol. 23, 416–425 (2016).
Chen, M. et al. Convolutional neural networks for automated annotation of cellular cryo-electron tomograms. Nat. Methods 14, 983–985 (2017).
Xiao, L., Magupalli, V. G. & Wu, H. Cryo-EM structures of the active NLRP3 inflammasome disc. Nature 613, 595–600 (2023).
Broz, P. & Dixit, V. M. Inflammasomes: mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 16, 407–420 (2016).
Gambin, Y. et al. Single-molecule fluorescence reveals the oligomerization and folding steps driving the prion-like behavior of ASC. J. Mol. Biol. 430, 491–508 (2018).
Lu, A. & Wu, H. Structural mechanisms of inflammasome assembly. FEBS J. 282, 435–444 (2015).
Gluck, I. M. et al. Nanoscale organization of the endogenous ASC speck. iScience 26, 108382 (2023).
Kuri, P. et al. Dynamics of in vivo ASC speck formation. J. Cell Biol. 216, 2891–2909 (2017).
Li, T., Gil Pineda, LI, Stevens, A. O. & He, Y. Key factors regulating the interdomain dynamics may contribute to the assembly of ASC. Biology 12, 796 (2023).
Diaz-Parga, P., Gould, A. & de Alba, E. Natural and engineered inflammasome adapter proteins reveal optimum linker length for self-assembly. J. Biol. Chem. 298, 102501 (2022).
Diaz-Parga, P. & de Alba, E. Inflammasome regulation by adaptor isoforms, ASC and ASCb, via differential self-assembly. J. Biol. Chem. 298, 101566 (2022).
Bryan, N. B. et al. Differential splicing of the apoptosis-associated speck like protein containing a caspase recruitment domain (ASC) regulates inflammasomes. J. Inflamm. 7, 23 (2010).
Matsushita, K. et al. A splice variant of ASC regulates IL-1beta release and aggregates differently from intact ASC. Mediat. Inflamm. 2009, 287387 (2009).
Nagar, A., Rahman, T. & Harton, J. A. The ASC speck and NLRP3 inflammasome function are spatially and temporally distinct. Front. Immunol. 12, 752482 (2021).
Nagar, A., DeMarco, R. A. & Harton, J. A. Inflammasome and caspase-1 activity characterization and evaluation: an imaging flow cytometer-based detection and assessment of inflammasome specks and caspase-1 activation. J. Immunol. 202, 1003–1015 (2019).
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. Manipulation of subunit stoichiometry in heteromeric membrane proteins. Structure 24, 797–805 (2016).
Martin-Sanchez, F. et al. ASC oligomer favors caspase-1CARD domain recruitment after intracellular potassium efflux. J. Cell Biol. 222, 202003053 (2023).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Schrodinger, L. L. C. The PyMOL Molecular Graphics System, Version 2.5 (Schrodinger, LLC, 2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Veevers, R. & Hayward, S. Methodological improvements for the analysis of domain movements in large biomolecular complexes. Biophys. Physicobiol. 16, 328–336 (2019).
Negi, S. S., Schein, C. H., Oezguen, N., Power, T. D. & Braun, W. InterProSurf: a web server for predicting interacting sites on protein surfaces. Bioinformatics 23, 3397–3399 (2007).
Suhre, K. & Sanejouand, Y. H. ElNemo: a normal mode web server for protein movement analysis and the generation of templates for molecular replacement. Nucleic Acids Res. 32, W610–W614 (2004).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, 42166 (2018).
Asarnow D., Palovcak, E. & Cheng, Y. UCSF pyem v0.5. Zenodo https://doi.org/10.5281/zenodo.3576630 (2019).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zheng, S. et al. AreTomo: an integrated software package for automated marker-free, motion-corrected cryo-electron tomographic alignment and reconstruction. J. Struct. Biol. X 6, 100068 (2022).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Acknowledgements
J.M. wants to thank the support from the National Key Research and Development Program of China (No. 2024YFA1307502, J.M.), the Science and Technology Innovation Plan of Shanghai Science and Technology Commission (No. 23JS1400200, J.M.), and the Research Fund for International Senior Scientists (No. W2431060, J.M.).
Author information
Authors and Affiliations
Contributions
Q.W. and J.M. designed and supervised research; D.X., S.L. and H.Y. performed research; D.X., F.N., Z.L., G.F., Q.W. and J.M. analyzed data; D.X., F.N., Q.W. and J.M. wrote the paper; all authors agreed on the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
Q.W. is an employee of Harcam Biomedicines who is bound by confidentiality agreements that prevent the disclosure of the competing interests in this work. All remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xue, D., Ni, F., Liu, S. et al. Atomic mechanisms of full-length ASC-mediated inflammasome assembly. Nat Commun 16, 10564 (2025). https://doi.org/10.1038/s41467-025-65578-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65578-2