Abstract
DNA molecules can be used to build “neural networks” that function like the brain, enabling them to perform complex computational tasks. However, a fundamental limitation of existing DNA networks is that their most basic computing units cannot perform true continuous and precise analog calculations, which restricts their ability to process complex information effectively. To address this, here we develop a DNA computing unit called CALCUL. This system successfully achieves fully analog computation, where all inputs, weighting parameters, and outputs are continuous and precise values. It performs the core operations of a neural network rapidly with high accuracy and is reusable. By integrating magnetic bead technology, we also enable modular operations and the construction of multilayer networks. Ultimately, we use this technology to construct a deep DNA neural network that correctly identifies complex color images with 100% accuracy. These developments provide a robust foundation for building more powerful and precise molecular computers.
Similar content being viewed by others
Introduction
Currently, DNA has become a widely used macromolecule material for molecular computing, capable of storing and processing information at the molecular scale1,2,3,4,5,6,7. Numerous DNA logic circuits with specific functions have been established8,9,10,11, such as binary square root circuit, half adder and full adder. Although the field of DNA logic circuits has been developing rapidly, it is primarily unsuitable for dealing with complex and analog input information, such as recognizing images with numerous pixels. Under such scenario, neural networks are much better options as they possess the advantages of large-scale parallel computing and weighted processing of input data12,13,14,15,16,17,18,19. Therefore, in recent years, researchers have been endeavoring to use DNA to construct the foundational element of a neural network: the weighted summation unit, and based on which, a variety of DNA neural networks were further developed. Cherry and Qian proposed a “winner take all” neural network to recognize nine types of digital figures, showcasing the feasibility of using DNA to construct complex neural networks13. Okumura et al. invented the enzyme driven DNA neural networks, which showed much faster computing speed and thereby implemented spatial partitioning of nonlinear separable regions20. Xiong et al. further integrated the weighted summation operations into convolutional networks and implemented recognition of 32 12×12 images with 100% correctness, which revealed the computational potential of DNA neural networks21. For biomedical applications, Zhang et al. developed a DNA neural network based miRNA classifiers for cancer diagnosis in serum samples22. Overall, due to the capability of highly paralleled computing and processing complex analog information, DNA neural networks are promising and powerful systems that might bring molecular computing to a much more diverse and functionalized level.
However, none of the current DNA neural networks are fully analog. This is because the fundamental elements of DNA neural networks—the weighted summation units—cannot achieve complete analog computing. In other words, their inputs, outputs, and weights cannot simultaneously be continuous and accurate. For instance, in the DNA weighted summation unit developed by Xiong et al., the input can only be a discrete value of 0 or 121. In another weighted summation unit developed by Zhang et al., the input and output can both be continuous. However, the weights are discrete and finite integer values (1, 2, 3, 4 and 5)22. In the weighted summation units developed by Genot et al., the inputs, outputs, and weights could theoretically be continuous, but their accuracy has only been characterized in the simplest case of two-input weighted summation. The accuracy of more complex weighted summation has not been tested, let alone that of convolution or deep neural networks23. More importantly, we believe that their sequence and structural design cannot ensure accurate analog computation due to the stochastic nature of their designed chemical reactions.
Unfortunately, the foundation of DNA circuits is concentration-based DNA chemical reactions, which are naturally continuous/analog. This is fundamentally different from electronic circuits, whose basic reactions are inherently digital. The absence of suitable analog computing units necessitates quantization of the analog DNA concentrations during weighted summation. This not only reduces the computational accuracy of the neural network but also necessitates the design of additional nodes to maintain the overall performance of the neural network. Furthermore, it requires the addition of extra quantization modules between nodes, which in turn causes the number of DNA strands in the network to proliferate exponentially with depth. As a result, constructing complex DNA neural networks, especially those with deep layers, becomes challenging.
Given these challenges, none of the current DNA weighted summation units can achieve fully analog computations where the inputs, weights and outputs are not only continuous but also accurate. The lack of such operation units blocks the construction of complex and deep DNA neural networks and limits the field’s development.
To overcome these fundamental limitations, here we develop the CALCUL unit. Our system represents an advance in DNA-based neural computing, demonstrating high-performance weighted summation operations for multiple inputs within short timeframes. The system successfully implements both full-connection and convolution operations with high accuracy while maintaining functional reusability. Through integration of magnetic-bead techniques, we expand the system’s computational capabilities to enable multi-layer neural network operations. This enhanced CALCUL platform incorporates weighted summation with ReLU activation, negative weight implementation, and operation concatenation - all while preserving computational precision. The system’s capabilities are rigorously validated through image recognition tasks, including successful classification of handwritten digits using convolutional network architectures.
Building upon these capabilities, we construct a deep DNA neural network architecture comprising two convolutional layers and one fully-connected layer, demonstrating robust performance in classifying pixel-based images with continuous value inputs. Collectively, these achievements establish CALCUL units as a versatile and powerful platform for implementing DNA-based computing systems and neural network operations.
Results and discussion
Working principles
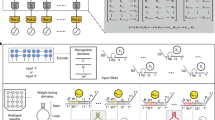
Herein, we have established a Classified allosteric-toehold based continuous and ultra-accurate (CALCUL) computing unit for weighted summation, in which the inputs, outputs and weights all present as continuous and accurate values. Based on the CALCUL unit, analog convolutional and fully-connected DNA neural networks were further established. As shown in Fig. 1A, B, we have designed two modes of CALCUL units: the cis-CALCUL mode for fully-connected operations and the trans-CALCUL mode for convolution operations. The difference is that in the cis-mode, the P-strand containing branch-migration domain B serves as the input and the Q-strand containing toehold domain A* serves as the weight, and vice versa for the trans-mode. Taking cis-CALCUL mode as an example, the CALCUL unit can be divided into two sections: the weight multiplication section and the summation section. The input strands (denoted as P-strand) are all single-stranded DNAs consisting of domain A and B. Although the sequences of domain A in different input strands are different, the sequences of their domain B remain consistent.
A Schematic illustration of the weighted summation operation of the cis-CALCUL unit. Multiplexed classified allosteric-toehold (CAT) annealing reactions are undergoing within one tube, realizing “real” weighted summation operation with the inputs, weights and outputs all being continuous and accurate values. B Schematic illustration of the weighted summation operation of the trans-CALCUL unit. Similarly, based on Multiplexed classified allosteric-toehold (CAT) annealing reactions, but with different roles played by the domains compared to cis-CALCUL units, which helps it to be more suitable for convolution operations. C Schematic illustration of the origin of the high accuracy analog computing of the CALCUL units. The temperature-annealing protocol and detection temperature settings used throughout this study are shown in the figure. [Created in BioRender. Main, T. (2025) https://BioRender.com/x7x6crx]. Source data are provided as a Source Data file.
In fact, the primary driving force in our computational reactions originates from the temperature-annealing process. For each input, it will undergo a classified allosteric-toehold (CAT) annealing process as illustrated in Fig. 1A: there are two corresponding weight strands; When one weight strand (Q-strand), the input P-strand, and the T-strand/O-strand duplex are present together, thermal annealing leads to the formation of an effective PQT triplex (fluorescent); When another weight strand (Q’-strand), the input P-strand, and the T’-strand/O-strand duplex are present together, thermal annealing leads to the formation of an effective PQ’T’ triplex (non-fluorescent). Notably, during the isothermal process following annealing, the classified allosteric-toehold design in this system facilitates the convergence of the reaction outcome toward a stable state of complete reaction (the reaction mechanism is illustrated in Supplementary Fig. 2). Then, we can flexibly adjust the percentage of P-strand entering the effective downstream pathway at any value between 0 and 1 by setting the ratio of [Q-strand]/([Q-strand]+[Q’-strand]) at the corresponding value. As we know, the amount of P-strand entering into effective downstream pathway is exactly Iiwi, wherein the [P-strand] is Ii and the [Q-strand]/([Q-strand]+[Q’-strand]) ratio is wi. So far, the CALCUL unit has finished the weight multiplication operation (Iiwi) with the Ii and wi at any designated value. Then, to fully accomplish weighted summation, i.e. ∑Iiwi, we need a summation section that adds up all the Iiwi. As we have described above, in our sequence design, the domain B of P-strand and the domain D of Q-strand will cooperatively bind to T-strand, and the domain B of all inputs are the same and so are the domain D of all Q-strands. Therefore, all the input strands bound to their corresponding Q-strands will react with the same T-strand/O-strand duplex during the annealing process, and such integration of CAT reactions is exactly the summation operation of ∑Iiwi. Meanwhile, all the input strands that bound to corresponding Q’-strands will react with the same T’-strand/O-strand duplex during the annealing process, producing non-fluorescent PQ’T’ triplex. So far, the CALCUL unit has realized its inputs, weights and outputs to be continuous simultaneously.
While, as shown in Fig. 1B, in the trans-CALCUL unit, Q-strands serve as inputs and P-strands serve as weights. Alongside P-strand, a P’-strand labeled with a quencher is designed, and the weight value equals to the percentage of [P-strand]/([P-strand]+[P’-strand]). Different to the cis-CALCUL, in which a pair of effective module and void module is designed, there is only one module in the trans-CALCUL. Since the P’-strand is labeled with a quencher, the P’QT triplex will not emit signal and is classified as void output; the PQT triplex will emit signal and is classified as effective output, achieving the weight multiplication operations. Similar to the cis-CALCUL, all P-strands and Q-strands will react with the same OT duplex, realizing accumulated weight multiplication (weighted summation). We would like to note that the trans-CALCUL units omit the OT’ duplex, so the reaction system is even more concise than the cis-CALCUL units, this is particularly crucial for constructing ConvNet (Convolutional neural Network).
However, aside from continuity, the weighted summation unit, as the foundation of DNA neural networks, must also be highly accurate. As for this, the computing accuracy of previously reported weighted summation units was – to the best of our knowledge - never thoroughly demonstrated13,20,21,22,23. They focused on the units’ linear responses to inputs and the correctness in classifications, and ignored the absolute errors of the weighted summation operation itself. This will not cause conspicuous computing deviations when they are only one-layer networks. Yet with the neural network goes deeper, the deviation would accumulate and greatly affect the network’s functionality. We believe the previously reported weighted summation units require too many DNA strands and involve multiple pathways of strand displacement reactions, so the thermodynamics, kinetics and crosstalk cannot be robustly controlled, leading to inevitable and considerable deviations.
Most DNA circuits aim to optimize thermodynamics for better performance, but kinetic barriers often cause output signals to deviate from theoretical values. For instance, basic toehold reactions may plateau prematurely, and more complex reactions are similarly affected. Thus, addressing kinetic barriers is crucial for DNA analog circuit construction (validation experiments are shown in Supplementary Fig. 5). Our designs consider these barriers, ensuring compatibility with thermal annealing. To achieve temperature-annealing-based computational reactions with high uniformity and accuracy, our mechanism incorporates sophisticated designs in both the reaction process and structural architecture. As shown in Fig. 1C, the CALCUL units utilize an identical strand displacement pathway for all inputs, ensuring uniform reaction thermodynamics and enhancing computing accuracy. To bolster reaction robustness, three key designs were implemented. Firstly, a 18-nt toehold long domain D was employed to favor the forward reaction, which means that the Q strand will bind to the T strand with robust and sufficient stability, ensuring near-complete conversion and reproducibility. Secondly, an allosteric domain X/X* was incorporated into the modules to prevent leakage between the P-strand and modules in the absence of the Q-strand, safeguarding against common issues in strand displacement systems that compromise accuracy. It is particularly noteworthy that the allosteric domain X/X* design differs significantly from the dissociation region in conventional toehold-mediated strand displacement reactions. This architecture not only effectively suppresses signal leakage but also ensures thermodynamic irreversibility - a critical feature for implementing temperature-annealing-based computational reactions (details are shown in Supplementary Fig. 2). Lastly, the longer T-strand was designated as the output strand to mitigate crosstalk between OT duplex and OT’ duplex, ensuring that fluorescent signals remain unchanged despite potential crosstalk, thereby preserving computing accuracy. In the fluorescent detection system design, we employed a modification scheme with the fluorophore conjugated to the long T-strand and the quencher attached to the short O-strand. By maintaining the quencher strand concentration at a 5% excess over the fluorophore strand, this strategy not only compensates for errors introduced during concentration measurement and sample loading, but also minimizes the impact of crosstalk reactions on signal output (This scheme is also applicable to magnetic bead assisted computational reactions). Notably, using the longer T-strand as output not only reduces crosstalk but also aids annealing, as annealing can accelerate crosstalk, but our design prevents it from affecting the output signal. The common approach of using the shorter O-strand as output is not suitable for annealing and cannot achieve precise analog computing, the experimental validation results for this section are presented in Supplementary Fig. 14. More detailed elaboration of the origin of the high accuracy analog computing of the CALCUL units could be found in Supplementary Section S1 of the Supplementary Information.
Integration of multiple pathways of allosteric toehold strand displacement
From the working principle described above, the Integration of multiple pathways of allosteric-toehold strand displacement reaction serves as the foundation for both cis- and trans- CACUL units: In cis-CALCUL units, weighted summation is achieved by two parallel integrations (effective and ineffective pathways) sharing the same input strands; in trans-CALCUL units, weighted summation corresponds to the integration with the input/output strand roles swapped. We therefore prioritized testing the feasibility and accuracy of such integration (Fig. 2A). We synthesized 8 inputs (P-strand-1 to 8) and their corresponding weights (Q-strand-1 to 8). According to the sequence design, all P-strands are led to invade the same T-strand-1/O-strand-1 duplex. To simplify, we set the P-strands to be binary (150 nM of input represented 1; no input represented 0) and the Q-strands to be analog values between 0 and 1, normalized by 150 nM. Then, we performed the operations of integrating 2, 4 and 8 inputs/weights, respectively. As shown in Fig. 2A, for each experimental group, we firstly conducted the operations on 4 designated standard sets of inputs/weights to obtain the standard curve, and the linearities in all groups were over 0.999. Then, we conducted the computation on two testing sets of inputs/weights, and based on the standard curves, we could transform the fluorescent signals to output values. The experimental results closely matched to the theoretical values, with all deviations remaining below 2.5%. It was worth noting that we could add strands (denoted as c-Q-strand) complementary to the Q-strands to reset the whole computing process and then restart it by adding W-strands again, and by doing so, we implemented 2 rounds of calculation with the computing accuracies ranging from 90.7% to 100.0%, median 98.6% (Supplementary Figs. 4, 6 and 7).
The output value was calculated from the fluorescence intensity using a standard curve. The dashed line beside the output curve indicated the theoretical output and its affiliated zone in light color represents a deviation of ±5%. A Schematic illustration and experimental results of integrating multiple pathways of allosteric-toehold strand displacement into one T-strand/O-strand duplex. From top to bottom: Row 1 displays the fluorescence intensity responses on four designated standard sets of inputs/weights, the standard curve, and the results of two summation operations of integrating two allosteric-toehold strand displacement reactions. Row 2 and 3 display the above data of integrating four and eight allosteric-toehold strand displacement reactions. The digits in red are experimental values and those in black are theoretical values. B Schematic illustration of parallelly integrating multiple pathways of allosteric-toehold strand displacement. n equals the total number of input strands (all weight strands that interact with the nth input strand are labeled n1, n2 through nm), while m equals the number of output pathways (all weight strands that direct to the mth output pathway are labeled 1 m, 2 m through nm). C Experimental results of multiple inputs being integrated into multiple T-strand/O-strand duplex, simultaneously and respectively. The digits out of the brackets are experimental values and those in the brackets are theoretical values. D The box plots display the distribution of relative errors between experimental and theoretical values for all integration experiments shown in Fig. 2A, C (n = 3 technical replicates). The center line represents the median (50th percentile), the box bounds indicate the interquartile range (IQR; 25th to 75th percentiles), and the whiskers extend to the minimum and maximum data points within 1.5 × IQR from the quartiles. Any values beyond the whiskers are considered outliers. Here and below, all of the experiments were technically repeated for three times. [Created in BioRender. Main, T. (2025) https://BioRender.com/x7x6crx]. Source data are provided as a Source Data file.
To further demonstrate the feasibility and accuracy, we tested multiplexed and paralleled integration (Fig. 2B). Firstly, two P-strands were designed to be integrated into two different T-strand/O-strand duplexes, simultaneously and respectively. Similarly, P-strands were binary and the weights were values between 0 and 1, normalized by 120 nM. Experimental results in Fig. 2C showed that paralleled integration of two allosteric toehold strand displacement reactions in one tube was completely feasible: the standard curves of two reaction pathways were both highly linear (Supplementary Fig. 8) and the computing accuracies were 98.9% and 98.0%. Furthermore, we tried integrating three paralleled reactions, simultaneously and respectively. The standard curves were highly linear (Supplementary Fig. 10) and the computation results were all highly accurate, with the deviations less than 2.9% (Fig. 2C). Notably, we also tested recycling of the paralleled multiplexed integration. Shown in Supplementary Figs. 9, 11, the above duplex and triplex integration were recycled for 2 rounds, and the linearity of standard curves remained higher than 0.99 and the computing accuracy kept over 95.3%. Overall, we have demonstrated that integration of multiple allosteric toehold pathways and paralleling such integration are both feasible and accurate (Fig. 2D).
Construction of the CALCUL units
Having demonstrated the feasibility and accuracy of integrating multiple pathways of allosteric toehold strand displacement, we then constructed complete cis- and trans- CALCUL units for weighted summation (∑Ii wi). The Ii were analog values between 0 and 1, derived from normalizing the concentration of the corresponding input strand by 120 nM. The wi was equal to [Weight-strand]/([Weight-strand]+[Weight-strand’]). As shown in Fig. 1A, B, in the cis-CALCUL units, P-strands serve as inputs, Q-strands and Q ‘- strands serve as weights, while in the trans-CALCUL units, Q-strands serve as inputs, P-strands and P’- strands serve as weights. We then conducted the weighted summation operations on 2, 4 and 8 inputs (n = 2, 4, 8) (Supplementary Figs. 12–17), respectively. For each n, we designed a set of higher weights and a set of lower weights so that we could examine the computing accuracy at a wide range. As was shown in Fig. 3, for all tests, both the cis- and trans- CALCUL units could implement the weighted summation operations within 40 min. And the computing accuracy was from 96.0% to 100.0% (median 98.9%) and 94.7% to 99.4% (median 98.0%) for cis- and trans- CALCUL units respectively. In addition, both the two types of CALCUL units could be recycled. The computing accuracy remained over 96.6% and 90.2% for cis- and trans- CALCUL units after 2 rounds of computations (Supplementary Figs. 18, 19). To the best of our knowledge, the performance of previously reported DNA weighted summation tools has been primarily evaluated through qualitative assessments. While these studies successfully demonstrated functional classification capabilities, a quantitative characterization of computing accuracy - specifically, the direct comparison between theoretical and experimental values - remains to be fully established. Our work addresses this gap by providing comprehensive data that rigorously validates the computational precision of the CALCUL unit. Considering its fast speed, distinct simplicity and high accuracy, we believe the CALCUL unit was a leading DNA tool for weighted summation in the field of DNA neural network (A systematic and comprehensive comparison is provided in Supplementary Table 27).
The output value was calculated from the fluorescence intensity using a standard curve. The dashed line beside the output curve indicated the theoretical output and its affiliated zone in light color represents a deviation of ±5%. The digits in red are experimental values and those in black are theoretical values. In practical applications, the resetting process for both cis- and trans- CALCUL units is temperature-annealing-based, while strand displacement reactions are depicted here for clarity. A The output values of the cis-CALCUL unit in weighted summation and the schematic illustration and experimental results of reset and recycling of the cis-CALCUL unit. B The output values of the trans-CALCUL unit in weighted summation and the schematic illustration and experimental results of reset and recycling of the trans-CALCUL unit. [Created in BioRender. Main, T. (2025) https://BioRender.com/x7x6crx]. Source data are provided as a Source Data file.
The cis-CALCUL units based full-connection and the trans-CALCUL units based convolution operations
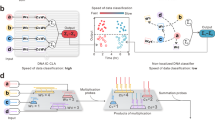
As shown in Fig. 4A, for the full-connection operations based on cis-CALCUL units, we designed fully connected layers with 2 inputs/2 outputs, 3 inputs/3 outputs and 4 inputs/3 outputs. Experimental results showed that the cis-CALCUL unit based full-connection network successfully implemented the operations with the computing accuracy ranging from 96.1% to 99.3% (median 97.7%) and the operation time was less than 30 min. It is worth noting that for full connection operations, the cis-CALCUL unit is more compatible because full connection system requires assigning separate weights to each input in different pathways.
The output value was calculated from the fluorescence intensity using a standard curve. The dashed line beside the output curve indicated the theoretical output and its affiliated zone in light color represents a deviation of ±5%. A Schematic illustration and experimental results of the full-connection operations by the cis-CALCUL unit. The digits out of the brackets are experimental values and those in the brackets are theoretical values. B Schematic illustration of the convolution operation based on the trans-CALCUL unit. The convolution operation uses an equal stride design (s = k) for an i × j input matrix. According to discrete convolution principles, the number of weighted summation operations (n) satisfies: n = ((i + 1)/k)×((j + 1)/k), (where k is the kernel width). C The experimental results of recognition of 16-pixel color images depicting parallel lines in different directions by the trans-CALCUL unit based ConvNet displayed in the lower half. Results are shown as mean ± SD with overlaid scatter plots of individual technical replicates (n = 3) [Created in BioRender. Main, T. (2025) https://BioRender.com/x7x6crx]. Source data are provided as a Source Data file.
We then wanted to utilize our trans-CALCUL units to implement convolution. The schematic illustration of the convolution operation with a 2×2 stride 2 kernel by the trans-CALCUL unit was shown in Fig. 4B. In each calculation, m×n inputs are split into k×k regions of interest (with the same size as the kernel). Each input cell is represented by Qij strand, and each kernel cell by a combination of Pij and P’ij. Element-wise multiplication and summation (the convolution operation) is performed to give the output cell. In our case, stride s is identical with the kernel size k, the output matrix size is thus [(n − k)/s + 1] × [(m − k)/s + 1]. It is worth noting that in the trans-CALCUL unit, the sequences of the A domains in different P and P’ strand pairs exhibit diversity, serving to determine the specific positions of the weights within the convolutional kernel. Meanwhile, the sequences of the B domains remain identical across all P and P’ strand pairs. This arrangement allows different convolutional kernels to share the same weight strands, thereby significantly reducing the number of strands required for a convolutional operation. For experimental verification, we designed the trans-CALCUL unit based ConvNet to recognize two 16-pixel (4×4) images depicting parallel lines in opposite directions by 2×2 kernels. The four modules were labeled with the same quencher (BHQ-2) but with four different fluorophores (FAM, HEX, TAMRA, ROX), so the four values of the 2×2 output feature map could be reported individually and simultaneously. We firstly ran the ConvNet on four standard sets of inputs to obtain four standard curves for all channels and then on the two images. As shown in Fig. 4B and Supplementary Fig. 24, the four channels all exhibited high linearity (R2 > 0.99). The system achieved perfect recognition, with the output feature maps showing high accuracy (95.2–100%; median 96.5%) relative to theoretical results.
The trans-CALCUL unit based ConvNet for image recognition
To be more convincing, we further used the trans-CALCUL unit based ConvNet to recognize six 64-pixel (8×8) images depicting digit 8 and 9 by 4×4 kernels (Fig. 5). The output 2×2 feature map was further pooled by summating each of the columns to give 2 outputs indicating the result for each category. Similarly, the linearities were over 0.99 (Supplementary Fig. 26), the recognition was fully correct, and the final output values were highly accurate (90.3% to 99.4%, median 94.7%). We would like to note that each of the recognition/convolution process was implemented within a single tube, without segmenting into paralleled tubes at all.
Schematic illustration and experimental results of recognition of 64-pixel color images depicting digit 8 or 9 by the trans-CALCUL unit based ConvNet. The digits in blue and green are experimental output values and those in black are theoretical values. Results are shown as mean ± SD with overlaid scatter plots of individual technical replicates (n = 3). Throughout this study, two-digit numbers enclosed in circles represent the fractional part of input values (i.e., the first two decimal places), whereas standalone single-digit numbers specifically denote integers (0 or 1). Source data are provided as a Source Data file.
Magnetic bead assisted CALCUL units
In the above experiments, we have thoroughly characterized the CALCUL unit and the DNA neural networks based on it. However, we must emphasize that if deep DNA neural networks were to be constructed exclusively using the current CALCUL units, the interlayer reactions would remain isolated. Under such conditions, the layer-to-layer connections would operate through abstract numerical transfers rather than physical DNA strand, which to some extent weakened its practicability. Therefore, we would like to further improve the CALCUL unit so that the output strands enter into the downstream layer and thereby the whole computing process could be completely implemented by DNA strands. As we know, the inevitable DNA crosstalk and leakage has long been the restraint of the field; if adding all of the upstream strands into the downstream layer, the undesired side reactions would be significant24,25. To overcome this, we introduced magnetic bead to “purify” the upstream strands and only allow the effective output strands to enter into the downstream layer, and by doing this, we have further extended the capability of the CALCUL units.
The first improvement was the incorporation of the cis-CALCUL unit and trans-CALCUL unit based ReLU module, which made our CALCUL unit more compatible for building neural networks. For the cis-CALCUL unit based ReLU module was shown in Fig. 6A, in the upstream CALCUL unit, the input P-strand had four domains: domain A and B were for allosteric toehold reaction, domain M and domain block were for ReLU. The O-strand was labeled with a biotin instead of a quenching group, and the T-strand was extended with an extra K* domain while the T’-strand remained unchanged. After weighted summation, magnetic beads coated with streptavidin were added and the magnetic field was applied. Consequently, the biotin labeled free O-strand, OWT triplex and OQ’T’ triplex would be absorbed onto the magnetic bead and isolated from the solution. The supernatant comprising of PQT triplex and PQ’T’ triplex was then transferred to another tube (Box 1), and the amount of PQT triplex was denoted as x. Then, Tr-strand and TrPDR triplex were added. The Tr-strand contained a domain K that was fully complementary to the domain K* of PQT triplex. The PN-strand was the input strand of the downstream CALCUL unit and the R-strand was labeled with a biotin. Since Tr-strand and TrPDR triplex contained the domain K, they both can hybridize with PQT triplex. However, the Tr-strand had much higher reaction priority with PQT because it was free and fully exposed whereas the domain K within TrPDR triplex was relatively blocked. Therefore, the PQT triplex would firstly bind to Tr-strand and produced PQTTr quadruplex. The amount of free Tr-strand added was denoted as b, and we could deduce that the amount of residual free PQT triplex was x-b. Then, the residual PQT triplex would react with the TrPDR triplex and produce identical amount (x-b) of PDR duplex. Overall, after reaction, the system consisted of PQTTr quadruplex, PQ’T’ triplex, excess TrPDR triplex and PDR duplex of amount x-b (Box 2). Then, the magnetic separation was applied and the supernatant was discarded. After resuspending the precipitate, the system contained excess TrPDR triplex and IDR duplex of amount x-b (Box 3). The last step was adding c-R-strand that was fully complementary to the R-strand and applied magnetic separation. Finally, the supernatant was transferred to another tube (Box 4) and the solution consisted of TrPD duplex and free ID-strand of amount x-b. Since the TrPD duplex was inert for downstream CALCUL unit, the final output of the upstream CALCUL unit was PN-strand of amount x-b, and the concentration of PN-strand was a×(x-b), where a was the dilution factor determined by the final solution volume. Overall, throughout the above process, we have achieved a ReLU module with the following expression:
The output value was calculated from the fluorescence intensity using a standard curve. The dashed line beside the output curve indicated the theoretical output and its affiliated zone in light color represents a deviation of ±5%. The digits in red are experimental output values and those in black are theoretical values. A Schematic illustration of the cis-CALCUL unit based ReLU module with a bias of “b”. B Schematic illustration and experimental verification of attaching the cis-CALCUL unit based ReLU to a weighted summation operation with 4 inputs. The right panel shows the fluorescence results from the 4 standard groups used to generate the standard curve, and the converted output value for the experimental group as determined by this curve. C Schematic illustration of the trans-CALCUL unit based ReLU module with a bias of “b”. D Schematic illustration and experimental verification of attaching the trans-CALCUL unit based biased ReLU to a weighted summation operation with 5 inputs. The right panel shows the fluorescence results from the 4 standard groups used to generate the standard curve, and the converted output value for the experimental group as determined by this curve. [Created in BioRender. Main, T. (2025) https://BioRender.com/x7x6crx]. Source data are provided as a Source Data file.
The ReLU and subtraction operations are considerably simpler when implemented with the magnetic bead-assisted trans-CALCUL unit than with the cis-CALCUL unit, due to the omission of the OT’ duplex. As shown in Fig. 6C, similarly, the O-strand and P’-strand are labeled with biotin instead of quenching group, and the T-strand is extended with an additional K* domain. The subsequent reaction principle and magnetic separation operation are consistent with the ReLU based on the cis-CALCUL unit, and finally, the ReLU module based on the trans-CALCUL unit can also be constructed.
For experimental verification, we implemented the weighted summation operation on 4 standard sets of inputs/weights to obtain the standard curve and 1 testing set of inputs/weights. For the cis-CALCUL unit based ReLU module the constant b in the ReLU was set at 0.2 by controlling the amount of Tr-strand. As shown in Fig. 6B, the standard curve demonstrated high linearity (R2 = 0.9977), and the computing result closely matched the theoretical value, with an accuracy of 98.9%. For the trans-CALCUL unit based ReLU module was shown in Fig. 6D, the constant b in the ReLU was set at 0.3 by controlling the amount of Tr-strand. The standard curve demonstrated high linearity (R2 = 0.9903), and the computed results closely matched the theoretical values, with accuracies of 96.2% and 99.4%. The computing results for both modes collectively demonstrate the feasibility and accuracy of incorporating a magnetic bead-assisted ReLU into the CALCUL unit.
The second improvement was the introduction of negative weights. The subtraction operation principle based on cis- and trans-CALCUL modes is the same. For the convenience of expression, IWT is uniformly used to represent PQT triplex in the cis-CALCUL mode and QPT triplex in the trans-CALCUL mode, and IWNTN is uniformly used to represent PQNTN triplex in the cis-CALCUR mode and QPNTN triplex in the trans-CALCUL mode.
As was shown in Fig. 7A, the inputs multiplied by positive weights would be integrated into T-strand/O-strand duplex by the weight strands, producing IWT triplex and biotinylated O-strand, whereas the inputs multiplied by negative weights would be into TN-strand/O-strand duplex by WN-strand, producing IWNTN triplex and biotinylated O-strand. It was worth noting that the T-strand was extended with overhang T+ domain and S+ domain, and the TN-strand was also extended with overhang T- domain and S- domain. After adding magnetic bead and applying magnetic field, the supernatant comprising of IWT triplex and IWNTN triplex was transferred to the downstream tube, wherein the overhang 10-nt T- domain of IWNTN triplex and the overhang 6-nt T+ domain of IWT triplex would cooperatively lead the two triplexes to hybridize with U+-strand/U--strand duplex, producing IWTU+ quadruplex and IWNTNU- quadruplex. According the sequence design, unless united, IWT triplex or IWNTN triplex alone could not stably bind to the U+U- duplex. Therefore, the U+U- duplex served as an annihilator (Supplementary Fig. 1C, 3), consuming IWT triplex by the amount of IWNTN triplex, which, in all, presented as subtracting IWNTN from IWT (Fig. 7B and Supplementary Fig. 27A). Finally, the biotinylated U+-strand and U--strand would isolate the IWTU+ quadruplex and IWNTNU- quadruplex from the system and leave free IWT triplex, as the final output, to enter into the downstream layer. We then implemented the above weighted summation and subtraction operations on 4 standard sets of inputs/weights to obtain the standard curve and 2 testing sets of inputs/weights. As shown in Fig. 7C, Supplementary Figs. 27B, 28, the standard curves for both CALCUL modes exhibited high linearity (0.9999 and 0.9937), and the accuracies of the computing results were favorable (99.3% and 98.8% for cis-CALCUL; 97.0% and 95.0% for trans-CALCUL). Notably, the use of negative weights expanded the available weight range to −1 ≤ w ≤ 1. While the weighted summation result could be negative, such negative values have limited utility in multi-layer networks, as they would not be propagated through the subsequent ReLU process (Y = X for X ≥ b; Y = 0 for X < b).
The digits out of the brackets are experimental values and those in the brackets are theoretical values. A Schematic illustration of the magnetic bead assisted CALCUL unit based negative-weight subtraction operation. The U+U- duplex annihilator is introduced to implement subtraction. B Experimental verification of the subtraction operation alone. C Schematic illustrations and experimental verification of a weighted summation operations of 8 inputs/weights by the cis-CALCUL units and trans-CALCUL units, wherein 4 weights are negative values. The digits in orange are experimental output values and those in black are theoretical values. D Schematic illustrations and experimental verification of a two-layer convolutional network based on magnetic bead assisted cis-CALCUL units and trans-CALCUL units. The network consists of both ReLU module and negative weights. E Schematic illustration of the magnetic bead assisted CALCUL unit based deep DNA neural network in recognizing color images depicting handwritten letters. The network consists of two convolutional layers and one fully-connected layer. Source data are provided as a Source Data file.
Magnetic bead assisted CALCUL unit based deep DNA neural networks
Having demonstrated the functionality and accuracy of magnetic bead assisted CALCUL unit, we next wanted to cascade the units to further build deep DNA neural networks. As shown in Fig. 7D, we firstly cascade two convolutional layers: the 16 inputs in a 4×4 pattern were convolved by a 2×2 kernel and activated by ReLU to produce a 2×2 feature map. The feature map was then convolved by a 2×2 kernel containing 2 positive weights and 2 negative weights and then subtracted to produce the final output. We ran the ConvNet on 3 standard sets of inputs/kernels to obtain the standard curve and 2 testing sets of inputs/kernels. As was shown in Fig. 7D, Supplementary Figs. 29, 30, the linearity of the standard curves of two CALCUL modes were 0.9928 and 0.9980. The computing accuracies were 88.8% and 91.4% for the cis-CALCUL, while 95.6% and 100% for the trans-CALCUL, demonstrating the cascading potential of magnetic bead assisted CALCUL units.
Finally, we built another deep DNA neural network to recognize and classify 144-pixel color images in the 12×12 pattern. As shown in Fig. 7E, the network consisted of a trans-CALCUL based convolution layer with 3×3 kernel and trans-ReLU, a trans-CALCUL based convolution layer with 2×2 kernel and cis-ReLU and one cis-CALCUL based fully connected layer with 4 inputs/2 outputs. This network is constructed on the subset of extended MNIST (EMNIST26) dataset, classifying handwritten letters A and B (Supplementary Fig. 31). At last, we used the CALCUL based deep neural network to recognize 12 144-pixel images depicting either “A’ or “B”. Experimental results in Fig. 7E showed that our ConvNet classified all the tested color images with 100% correctness, exhibiting the feasibility and accuracy of building CALCUL based deep DNA neural network.
We explicitly recognize that our design of setting the stride length equal to the convolutional kernel width imposes inherent limitations on DNA-based neural network operations. Unlike electronic convolutional neural networks, where adjustable stride lengths enhance model flexibility and performance, DNA-based implementations face fundamental biochemical constraints. These limitations necessitate careful optimization of reaction complexity, DNA strand design, and structural stability to ensure reliable computation. Currently, achieving freely stride lengths in DNA neural networks remains a significant technical challenge. After thorough evaluation, we adopted a fixed stride-kernel equivalence to guarantee complete input coverage, which is a strategy aligned with prior work on DNA convolutional neural networks21. While this approach simplifies system design and improves operational robustness, it embodies a deliberate trade-off between DNA reaction complexity and convolutional computing capability. We also would like to admit that although we have successfully cascaded the CALCUL units to implement deep neural network, the magnetic bead based cascading strategy is cumbersome and time consuming, and we anticipate that the microfluidic based devices, which are more convenient and autonomous, may significantly facilitate our computing system.
We have developed a DNA computing unit for weighted summation, whose inputs, weights and output are all continuous and accurate values. Both the cis- and trans- CALCUL units demonstrated high accuracy (≥94.7%) in weighted summation of up to 8 inputs within 40 min, and their computing accuracy remained above 90.7% after one round of reset and re-run. By scaling up and paralleling the weighted summations, we implemented the cis-CALCUL unit based full connection and the trans-CALCUL unit based convolution operations with the computing accuracies of over 96.1% and 95.2%, respectively. Using the trans-CALCUL unit based ConvNet, we recognized six 64-pixel (8×8) images depicting digit 8 and 9 by 4×4 kernels, the recognition was fully correct, and the final output values were highly accurate (90.3–99.4%, median 94.7%). Furthermore, by using magnetic beads to “purify” the system, we could cascade multiple layers and implement the whole computing process completely by DNA strands. Weighted summation followed by ReLU activation, weighted summation followed by subtraction and deep neural networks consisting of two convolutional layers and one fully-connected layer were built, all of which showed exceedingly high accuracy in computing and image recognition.
To preserve experimental simplicity, we implemented a minimalist deep network incorporating all essential convolutional neural network (CNN) functionalities. Our letter/digit recognition networks employed only 23 and 16 parameters respectively - orders of magnitude smaller than conventional EMNIST-performing CNNs requiring thousands of parameters. Crucially, this study focuses not on recognition accuracy (sub-30-parameter CNNs inherently underperform), but on verifying our chemical system’s capacity to replicate in silico predictions. Through multi-experiment validation on model-discernible samples (e.g., digits 8/9, letters A/B), we demonstrate precise biomolecular emulation of computational outcomes.
Overall, our established CALCUL unit integrates multiple advantages including concision, high speed, accuracy, and scalability, positioning it as a powerful computing unit for weighted summation. Based on this unit, we have developed an analog convolutional and fully-connected DNA neural network. These systems fully embody the key characteristics of DNA computing—such as intrinsic parallelism, molecular-level analog operation, noise tolerance, and biocompatibility—which endow them with exceptional capability in information processing tasks like pattern recognition. These features not only demonstrate the potential of DNA-based architectures as a robust framework for molecular intelligence but also highlight their promising translational value in clinical settings. We believe that the CALCUL unit and the deep neural networks built upon it will advance molecular computing systems and hold strong potential for biomedical applications including molecular diagnostics, gene expression profiling, and precision medicine16,22,27,28,29,30,31,32,33.
Methods
Sequence design
The detailed structural design of the strands and complexes used in this work was illustrated in Supplementary Figs. 1–3, and their sequences were shown in Supplementary Tables 4–26. To reduce synthesis errors, no more than four consecutive ‘A’s or ‘T’s (excluding poly A used for connecting biotin) and no more than three consecutive “C“s were designed. The CG% was set to approximately 40% to ensure appropriate melting temperatures. DNA strands used in the same reaction do not contain obvious secondary structures and undesired crosstalks at 37 °C. The sequence design was verified by NUPACK34 to confirm their binding energy and specificity.
Beyond NUPACK simulations, the designed DNA strands were experimentally screened using multiplication operations performed in both cis and trans configurations of the CALCUL unit (see Supplementary Figs. 36–39 and Supplementary Tables 124–135). Based on quantitative fluorescence measurements, candidate strand groups with fluorescence signals within ±5% of the median value were selected. Experimental groups falling outside this range were excluded.
DNA oligonucleotide synthesis
Based on the design, unlabeled oligonucleotides purified by high affinity purification (HAP) and labeled oligonucleotides purified by high-performance liquid chromatography (HPLC) were provided by Sangon Biotech and used without further purification. All strands were shipped lyophilized and resuspended at 20 μM in 1×Tris-EDTA buffer, and stored at 4 °C for further use.
Annealing protocol and buffer condition
The buffer used in all the experiments was 1×ThermoPol reaction buffer with 2 mM Mg2+ at pH 8.8 (New England Biolabs). All DNA strands were added to the target concentration according to experimental setting and mixed in 1× buffer solution. The strands were annealed by heating to 85 °C for 10 min, cooling to 65 °C, 55 °C and 45 °C at a rate of 4 °C per second, holding for 1 min at each temperature, repeating this entire cycle twice, and finally cooling to 37 °C.
Detailed experimental procedures
Single-layer computation experiments
Premixes of inputs, weights and modules were first prepared in accordance with the computational design. The solutions were then combined, subjected to an annealing procedure, and finally placed into a qPCR machine for data acquisition. Comprehensive details of the experimental configurations are systematically provided in Supplementary Tables 28–82.
Reset and recycling of the CALCUL units
Upon completion of a computational cycle, an equimolar amount of complementary strands relative to the weight strands was introduced into the reaction mixture. After annealing, the CALCUL unit was reset to its initial state. The premix weights solution was then reintroduced, and the system underwent another annealing process prior to fluorescence detection. A detailed description of the experimental setups can be found in Supplementary Tables 52–55, 65–68.
Multi-layer computation experiments
In multi-layer DNA neural networks, to facilitate the use of upstream DNA outputs as inputs for downstream layers, streptavidin-coated magnetic beads were employed to purify upstream DNA strands via affinity-based capture. The hierarchical interconnection protocol consists of the following steps:
-
(1)
weighted summation;
-
(2)
introduction of magnetic beads and incubation;
-
(3)
collection of the supernatant for activation function;
-
(4)
a second bead-based purification to isolate DNA inputs for downstream processing.
Detailed experimental procedures are systematically provided in Supplementary Fig. 40, Supplementary Tables 83–123.
Fluorescence acquisition
The output signal value were recorded by QuantReady K9600 qPCR machine and LineGene4800 qPCR machine at 37 °C with the fluorescence intensity measured. Four Fluorophore-Quencher pair were used: FAM (excitation 494 nm, emission 518 nm) - BHQ1 (quenching range 480–580 nm), HEX (excitation 535 nm, emission 556 nm) - BHQ1 (quenching range 480–580 nm), ROX (excitation 585 nm, emission 605 nm) - BHQ2 (quenching range 550–640 nm) and TAMRA (excitation 565 nm, emission 580 nm) - BHQ2 (quenching range 550–640 nm).
Considering efficiency (rapid computation completion), consistency (synchronized termination across replicates), and operational feasibility, we evaluated fluorescence signals at 10-min intervals for plateau verification. The termination criteria were applied as follows:
-
1)
For all curves, calculate the fluorescence increase magnitude during the 5-min window preceding each check timepoint. If none exceed 1% growth, the experiment terminates.
-
2)
In a set of experiments, if any curve fails to meet Criterion (1), the experiment will be reassessed after 10 min using the same standard. This iterative process is repeated until all curves satisfy the 1% threshold to determine a unified experimental termination time.
We also would like to note that in our experiment, we indeed observed some high-fluorescence curves (e.g., the full-connection operation with 4 inputs/3 outputs shown in Fig. 4A) continuing to exhibit exceptionally slow signal increases even after meeting the aforementioned experimental termination criteria. This phenomenon is likely caused by slowed reaction kinetics near thermodynamic equilibrium. Importantly, this behavior occurred synchronously in high-fluorescence standard curves. Since we convert fluorescence signals to computational results based on standard curves, the observed effect does not significantly impact the accuracy of CALCUL units. Experimental results confirm this conclusion (shown in Supplementary Figs. 32–35).
Statistics and reproducibility
Fluorescent signal data were exported from the qPCR instrument and analyzed using ORIGIN software. Quantitative outputs were derived from fluorescence signals using a pre-calibrated standard curve. For each weighted summation experiment, a standard curve was constructed by processing a minimum of three input/weight combinations designed to yield a gradient of output signals, under the same reaction conditions as the experimental samples. A scatter plot was generated with fluorescence intensity on the y-axis and the theoretical weighted summation value on the x-axis for these standards. The standard curve was then generated by performing linear regression (requiring R² > 0.99) on these data points. The equation derived from this regression was applied to convert the fluorescence intensity of experimental samples into quantitative output values. No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized, and the investigators were not blinded to allocation during experiments and outcome assessment. All data generated in this study are provided in the Source Data file.
Streptavidin coated magnetic beads
The streptavidin coated magnetic beads were provided by Beaver Biosciences Inc, with a diameter of 300 nm. In the experiment, the concentration of magnetic beads was set at 6.7 mg per μmol of biotinylated DNA, and they were incubated with shaking at 37 °C for 2 h, and magnetic separation was performed using a magnetic separator provided by Beaver Biosciences Inc. Notably, a conservative bead concentration (3× the recommended amount) was intentionally used to minimize incubation time. The cost can be further reduced by decreasing bead usage and extending incubation time without compromising separation efficiency.
Training and testing neural networks
We trained the multilayered convolution and full-connection network for image classification task among 4 categories of symbols: handwritten digits 8, 9 and handwritten English letters A and B. The Arabic numerals are from the corresponding subset of Modified National Institute of Standards and Technology (MNIST26) dataset, and English alphabets from the extended MNIST (EMNIST35) dataset.
Since the original image is 28×28 in both of the dataset, we resized them with bilinear interpolation to 8×8 (for 8 and 9) and 12×12 (for letter A and B). Since the total parameter counts are rather small (a total of 18 for convolution layer distinguishing 8 and 9, and 21 for distinguishing A and B), the size of each training subset in the MNIST/EMNIST is sufficient to saturate network performance (about 4000 images per category in the training subset). The testing set is the subset of the MNIST/EMNIST test set and contains about 800 images per category. These input cells are continuous floating point values between [0, 1].
The classification of 8 and 9 is achieved by a simple convolution operation followed by summation of columns before soft-max. The weight table is shown as below:
The network shown in Fig. 7E consists of 4 layers. Since the inputs of each layer is constrained within [0, 1], we scaled the output by a factor before entering the next layer of network.
We ensured \({K}_{A},{K}_{B},{W}_{C},{W}_{D},{b}_{A},{b}_{B},{b}_{C},{b}_{D}\) and \(C\) to be within [0, 1]. \({K}_{B}\) is a 2×2 convolution kernel with stride 2. For the image in the 8×8 pattern, \({K}_{A}\) is of size 2×2 with stride of 2. For the image in the 12×12 pattern, \({K}_{A}\) is of size 3×3 with stride of 3. We reduce the number of the fully connected layer in the classification task between letter A and B.
Since the direct output \(D\) or \(E\) are not necessarily between 0 and 1, we apply the softmax function before the training loss, after multiplying back the scaling factor that we have divided previously.
Or in the case of classification of English letters,
We use the sparse categorical cross-entropy loss function
The training is performed on PyTorch36 platform, learning rate is set to be 0.0001 with stochastic gradient descent (SGD) with a weight decay of 0.0001 and a momentum of 0.5. The network is trained for 100k iterations, and after each iteration, the weights and biases are clamped within [0, 1]. The weight matrices are initialized by Gaussian distribution with a mean of 0.5 and a standard deviation of 0.15. The performance of the network on the training and testing set is similar, indicating no significant overfitting. All the parameters are provided in the figures (Figs. 5 and 7).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the data supporting the findings of this study are available in the Source Data file. Source data are provided with this paper.
Code availability
The source code for this study is available at the GitHub repository: https://github.com/yang-z-03/dnaconv. A permanent, archived version is also hosted on Mendeley Data37 with the https://doi.org/10.17632/45vsfrghdr.1. All training result and parameters is demonstrated in the figures, and all testing and experimental accuracies are shown in Supplementary Information.
References
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Woods, D. et al. Diverse and robust molecular algorithms using reprogrammable DNA self-assembly. Nature 567, 366–372 (2019).
Genot, A. J., Bath, J. & Turberfield, A. J. Reversible logic circuits made of DNA. J. Am. Chem. Soc. 133, 20080–20083 (2011).
Song, X. & Reif, J. Nucleic acid databases and molecular-scale computing. ACS Nano 13, 6256–6268 (2019).
Wang, F. et al. Implementing digital computing with DNA-based switching circuits. Nat. Commun. 11, 121 (2020).
Fu, T. et al. DNA-based dynamic reaction networks. Trends Biochem. Sci. 43, 547–560 (2018).
Lim, C. K., Nirantar, S., Yew, W. S. & Poh, C. L. Novel modalities in DNA data storage. Trends Biotechnol. 39, 990–1003 (2021).
Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011).
Song, T. et al. Fast and compact DNA logic circuits based on single-stranded gates using strand-displacing polymerase. Nat. Nanotechnol. 14, 1075–1081 (2019).
Liu, L. et al. Multifunctional clip strand for the regulation of DNA strand displacement and construction of complex DNA nanodevices. ACS Nano 15, 11573–11584 (2021).
Chen, X. Expanding the rule set of DNA circuitry with associative toehold activation. J. Am. Chem. Soc. 134, 263–271 (2012).
Qian, L., Winfree, E. & Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 475, 368–372 (2011).
Cherry, K. M. & Qian, L. Scaling up molecular pattern recognition with DNA-based winner-take-all neural networks. Nature 559, 370–376 (2018).
Evans, C. G., O’Brien, J., Winfree, E. & Murugan, A. Pattern recognition in the nucleation kinetics of non-equilibrium self-assembly. Nature 625, 500–507 (2024).
Sahraeian, S. M. E. et al. Deep convolutional neural networks for accurate somatic mutation detection. Nat. Commun. 10, 1041 (2019).
Lopez, R., Wang, R. & Seelig, G. A molecular multi-gene classifier for disease diagnostics. Nat. Chem. 10, 746–754 (2018).
Kim, S. et al. Nanoparticle-based computing architecture for nanoparticle neural networks. Sci. Adv. 6, eabb3348 (2020).
Soltoggio, A., Stanley, K. O. & Risi, S. Born to learn: the inspiration, progress, and future of evolved plastic artificial neural networks. Neural Netw. 108, 48–67 (2018).
Stanley, K. O., Clune, J., Lehman, J. & Miikkulainen, R. Designing neural networks through neuroevolution. Nat. Mach. Intell. 1, 24–35 (2019).
Okumura, S. et al. Nonlinear decision-making with enzymatic neural networks. Nature 610, 496–501 (2022).
Xiong, X. W. et al. Molecular convolutional neural networks with DNA regulatory circuits. Nat. Mach. Intell. 4, 625 (2022).
Zhang, C. et al. Cancer diagnosis with DNA molecular computation. Nat. Nanotechnol. 15, 709–715 (2020).
Genot, A. J., Bath, J. & Turberfield, A. J. Combinatorial displacement of DNA strands: application to matrix multiplication and weighted sums. Angew. Chem. Int. Ed. Engl. 52, 1189–1192 (2013).
Lv, H. et al. DNA-based programmable gate arrays for general-purpose DNA computing. Nature 622, 292–300 (2023).
Xie, N. et al. Scaling up multi-bit DNA full adder circuits with minimal strand displacement reactions. J. Am. Chem. Soc. 144, 9479–9488 (2022).
Cohen, G., Afshar, S., Tapson, J. & Schaik, A.v. EMNIST: Extending MNIST to handwritten letters. In 2017 International Joint Conference on Neural Networks (IJCNN) 2921–2926 (IEEE, 2017). https://doi.org/10.1109/IJCNN.2017.7966217.
McDermott, A. M. et al. Identification and validation of oncologic miRNA biomarkers for luminal A-like breast cancer. PLoS One 9, e87032 (2014).
Benenson, Y., Gil, B., Ben-Dor, U., Adar, R. & Shapiro, E. An autonomous molecular computer for logical control of gene expression. Nature 429, 423–429 (2004).
Lancashire, L. J. et al. A validated gene expression profile for detecting clinical outcome in breast cancer using artificial neural networks. Breast Cancer Res Treat. 120, 83–93 (2010).
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012).
Xiao, M. et al. Programming drug delivery kinetics for active burst release with DNA toehold switches. J. Am. Chem. Soc. 141, 20354–20364 (2019).
Ma, Q. et al. An automated DNA computing platform for rapid etiological diagnostics. Sci. Adv. 8, eade0453 (2022).
Zhang, C. et al. Logical analysis of multiple single-nucleotide-polymorphisms with programmable DNA molecular computation for clinical diagnostics. Angew. Chem. Int Ed. Engl. 61, e202117658 (2022).
Zadeh, J. N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput Chem. 32, 170–173 (2011).
Deng, L. The MNIST database of handwritten digit images for machine learning research [best of the web]. IEEE Sign. Process. Mag. 29, 141–142 (2012).
Paszke, A. et al. PyTorch: An imperative style, high-performance deep learning library. In Advances in Neural Information Processing Systems, vol. 32, 8024–8035 (Curran Associates, Inc., 2019). https://proceedings.neurips.cc/paper_files/paper/2019/file/bdbca288fee7f92f2bfa9f7012727740-Paper.pdf.
Yang, Z. Code for DNA circuit based implementation of a convolutional neural network (Mendeley Data, 2025). https://doi.org/10.17632/45vsfrghdr.1.
Acknowledgements
This work was financially supported by the National Key Research and Development Program of China (Grant No. 2023YFE0210200 to X.X.), the National Natural Science Foundation of China (Grant Nos. 22104040 to J.D.; 82402466 to X.X.), the Natural Science Foundation of Wuhan City (Chenguang Project) (Grant No. 2024040801020331 to L.L.), and Jilin Provincial Science and Technology Foundation (YDZJ202401191ZYTS to X.X.).
Author information
Authors and Affiliations
Contributions
X. L. (Xiao Liu), Z.Z., and P.L. designed the overall research strategy, performed the experiments, and coordinated data collection and manuscript writing. X.X., J.D. and L.L. were the lead researchers for conceptualization of the project, data analysis and manuscript writing. Z.Y. performed the computational analysis, developed the code, and contributed to manuscript writing. H.H., M.W., X.L. (Xiaoding Lou), X.F. and K.T. participated in the experimental discussions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Chunhai Fan, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, X., Zheng, Z., Liu, P. et al. Deep convolutional and fully-connected DNA neural networks. Nat Commun 16, 10629 (2025). https://doi.org/10.1038/s41467-025-65618-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65618-x