Abstract
DNA damage is a constant threat to genome integrity and function. Diminished capacity for DNA repair is linked to many human diseases, therefore, understanding the molecular pathways responding to DNA damage is key for developing novel therapies. Lack of unbiased probes to report DNA damage dynamics in living cells and animals limits our current efforts to completely understand DNA repair processes. In this study, we overcome these limitations by engineering protein probes containing the tandem-BRCT domain of MCPH1, which we show to have a specific affinity for the DNA-damage-associated histone mark γH2AX. We employ these probes to track DNA damage dynamics in living cells exposed to a panel of different genotoxic insults, to visualize DNA damage targeted to heterochromatinised satellite repeats, and to map DNA double strand breaks genome-wide. Finally, we highlight the versatility of our probe to visualize programmed double strand breaks during gametogenesis in C. elegans. Taken together, we present a novel protein probe with broad application potential for DNA damage research.
Similar content being viewed by others
Introduction
Genome stability is constantly threatened by DNA lesions triggered by endogenous and exogenous factors1. Double Strand Breaks (DSBs), one of the most deleterious types of DNA damage, trigger the activation of the DNA Damage Response (DDR). This comprises a set of coordinated signaling pathways that elicit the hierarchical recruitment of repair proteins to DNA damage sites2. In eukaryotic cells, DSB detection and repair occur in the context of chromatin3. In response to DSBs, chromatin is rearranged, and the C-terminal tail of histone H2A is phosphorylated. In mammals, the variant H2AX is phosphorylated (γH2AX) by phosphatidylinositol 3-kinase (PI3K)-like protein kinases, including ATM and DNA-PK specialized kinases3. γH2AX serves as a signaling platform that guides spatiotemporal recruitment of DNA repair factors to DSBs4.
Currently, monitoring of γH2AX distribution, quantification, and kinetics in a spatiotemporal manner is highly constrained by the availability of effective live-cell-compatible probes. Apart from some recent advances using nanobodies or antigen-binding fragments5,6,7, methods employing conventional antibodies directed against γH2AX in fixed/permeabilized cells are more commonly used. Although such approaches have proven very efficient in scoring γH2AX distribution on chromatin, their use as probes to follow the kinetics of DSB formation is very limited. Furthermore, detection of γH2AX in living organisms or other biological specimens that could potentially be employed as a biomarker for drug development, exposure to radiation, as well as for cancer chemo- and radio-therapy trials, has also been hampered by the lack of specific and efficient probes.
Here, we developed novel DNA damage-specific sensors that allow live monitoring of DSBs in living cells. Based on our previous work8, we developed engineered chromatin readers (eCRs) containing selected naturally occurring tandem BRCT domains and screened for eCRs highly selective for γH2AX. BRCT domains have been previously characterized as “readers” of phosphorylated H2AX histones9. We show that the stable expression of a specific Microcephalin 1 (MCPH1) tandem-BRCT-containing eCR allows scoring dynamics and distribution of γH2AX on chromatin by live imaging and genomics. This sensor provides a novel and reliable tool for DNA damage visualization in living cells and organisms exposed to different DNA damage insults.
Results
Generation and selection of engineered DNA damage sensors specific for γH2AX
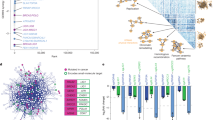
To develop an engineered chromatin reader (eCR) for the detection of DSBs in living cells, we explored the specificity and affinity of DDR proteins that are recruited to γH2AX sites in response to the presence of DSBs. BRCA1 C-terminal (BRCT) domains recognize and bind linear motifs phosphorylated by kinases that are activated following DNA damage10,11. These evolutionarily conserved domains are frequently present in proteins involved in the DDR and are found either as single units or in tandem copies, separated by a variable linker region11. A database search for mouse BRCT domains resulted in 49 singleton BRCT units grouped in six distinct branches based on amino acid sequence similarity (Fig. 1a). We narrowed down our selection to tandem BRCTs from proteins whose localization to sites of DNA damage is well documented. Our final selection comprised tandem BRCT domains present in the following proteins (Fig. 1b): MDC1 (BRCT 1-2)12, BRCA1 (BRCT 1-2)13, PAXI1 (BRCT 5-6)14, TP53BP1 (BRCT 1-2)15, MCPH1 (BRCT 2-3)16, DNLI4 (BRCT 1-2)17, and TOPBP1 (BRCT 5-6)18. Several of these domains have been characterized as modules responsible for recruiting DDR factors to γH2AX sites10. Amongst them are the tandem BRCTs of the mediator of DNA damage checkpoint (MDC1) that regulate the recognition and repair of DSBs19,20, and of BRCA1 itself, which play major roles in DSB repair21,22. The cDNAs encoding the selected tandem BRCTs were stably expressed as eGFP-fusion eCRs from a defined site in the mouse ES cell (mESC) genome by Recombinase-Mediated Cassette Exchange (RMCE), as previously described8 (Fig. 1c). The selected eCRs displayed stable and homogeneous expression, as assessed by measuring eGFP levels by flow cytometry (Supplementary Fig. S1a). Furthermore, eCR expression did not influence cell viability upon induction of DSBs (Supplementary Fig. S1b).
a Unrooted tree illustrating the conservation and relationship between individual BRCT domains. BRCT domains from Mus musculus were clustered based on multiple amino acid sequence alignments using the maximum-likelihood method. Support values for main branches (bootstrap test with 1000 replications) are shown. Colors are based on the clustering of singleton BRCTs (I to VI). Numbers shown after the underscore in protein names refer to the relative position of the singleton BRCT in each protein. Dots designate the selected individual BRCT domains used in the rest of this study. b Schematic representing the selected mouse proteins containing the tandem BRCT domains used in this study. Individual BRCT domains are highlighted and colored based on their clustering from the unrooted tree. c Schematics of the eCR expression cassette used in mESCs to generate stably expressed BRCTs fused with eGFP. The construct contains a biotin acceptor site (bio) and a nuclear localization signal (NLS) under the control of the CAGGS promoter (CAG). RMCE (Recombinase-Mediated Cassette Exchange), followed by double selection, was applied to generate stable integration of the expression construct at a defined site in the mouse genome with the aid of the Cre recombinase. d Representative live-cell images of mESCs stably expressing the indicated BRCTs fused to eGFP and the nuclear eGFP control (green). Cells were treated with ETO for 1 h. Cells were incubated with phenol-red-free medium containing Hoechst (magenta) and immediately imaged. Scale bar is: 50 µm. e Quantification of the number of eGFP foci per nucleus in mESCs from (d). Left panel (DMSO), right panel (ETO)-treated cells. The data represent mean ± SD from at least 52 cells of one of three independent biological replicates. f Percentage of cells with discernible eGFP foci per nucleus in mESCs from ETO-treated cells (d). Error bars represent mean ± SD from three individual technical replicates. Similar results were obtained in two independent experiments. g Representative live-cell images of mESCs expressing MCPH1-BRCT-eGFP. Cells were treated for 1 h with the different indicated DNA-damaging agents or exposed to UV (1 h post 10 mJ/cm2 irradiation) and incubated for 1 h after UV irradiation. Cells were incubated with phenol-red-free medium containing Hoechst (magenta) and immediately imaged. The scale bar is: 5 µm. h Quantification of the number of eGFP foci per nucleus in mESCs exposed to the indicated genotoxic conditions. The data represent mean ± SD from at least 80 cells of one of three independent biological replicates. Source data are provided as a Source Data file.
Live-cell imaging of the stable mESC lines expressing the selected BRCT variants revealed distinct levels of eGFP-detectable nuclear foci following incubation with etoposide (ETO), a topoisomerase II inhibitor that induces DSBs. Amongst the selected eCRs, the tandem BRCT domain of MCPH1 showed the largest number of distinct foci in undamaged and damaged chromatin (Fig. 1d, e). To assess the specificity of the selected eCRs for DSB sites, we quantified eCR-GFP foci by immunofluorescence and measured their colocalization with γH2AX foci in cells exposed to ETO. In agreement with live-cell imaging microscopy, MCPH1-BRCT showed the largest number of distinct foci specifically colocalizing with sites of DNA damage marked by γH2AX (Supplementary Fig. S1c–e). The BRCA1-BRCT-eCR also displayed an increased number of foci associated with DNA damage, while the additional eCRs showed reduced numbers of eGFP foci and varying degrees of colocalization to γH2AX sites in the presence of ETO (Fig. 1d–f and Supplementary Fig. S1c–e). This was not due to different levels of DNA damage in the analyzed cell lines since quantification of nuclear foci revealed comparable numbers of γH2AX signals in treated and control cell lines (Supplementary Fig. S1f).
Although recruitment of BRCT domain-containing full-length proteins to DSB sites has been previously established, our results indicate that the tandem BRCT domain of MCPH1, and to some extent the tandem BRCT domain of BRCA1, functions as a single structural unit and is sufficient for recruitment to γH2AX sites (Fig. 1d–f). The isolated tandem BRCT domains from the well-characterized γH2AX-binder MDC120 did not show a strong association with γH2AX in our assays, suggesting that additional domains present within the full-length protein are required, together with its tandem BRCT, for recruitment of MDC1 to DNA damage sites.
Given its efficient recruitment to γH2AX sites, we selected the MCPH1-BRCT-eCR and further examined its application as a DNA damage sensor. Besides ETO, damage-induced MCPH1-BRCT foci formation was also observed in mESCs treated with various DNA-damage or replication-stress conditions that substantially triggered γH2AX, including Camptothecin (CPT), Methylmethanesulfonate (MMS), Ultra Violet radiation (UV), and Cisplatin (CISP) (Fig. 1g, h and Supplementary Fig. S1g–i), further confirming that the selected eCR can relocate to sites of DNA damage triggered by distinct genotoxic conditions.
Functional evaluation of MCPH1-BRCT-eCR interactions with γH2AX
Next, we examined whether the observed localization of MCPH1-BRCT-eGFP indeed requires γH2AX. Inhibition of ataxia-telangiectasia-mutated (ATM), one of the kinases that phosphorylates H2AX in the presence of DSBs, significantly reduced the formation of MCPH1-BRCT-eGFP foci in mESCs exposed to CPT/Olaparib (OLAP) (Fig. 2a, b). To further test the requirement for H2AX phosphorylation for the eCR engagement with H2AX, we performed co-immunoprecipitation experiments in the presence of λ-phosphatase and phosphatase inhibitors. We observed that λ-phosphatase treatments abolish the MCPH1-BRCT-mediated pulldown of γH2AX, which is rescued by the presence of phosphatase inhibitors (Supplementary Fig. S2a), further supporting the requirement of the phosphorylation for MCPH1-BRCT-eCR binding to γH2AX. Finally, we introduced the MCPH1-BRCT-eCR and a background control probe into H2AX-deficient RPE-1 cells23 (Supplementary Fig. S2b) and tested its binding in the presence of ETO. While we observe a clear and specific colocalization of the MCPH1-BRCT-eCR probe to γH2AX sites and to sites marked by the DDR factor 53BP1 in ETO-treated wild-type RPE-1 cells, no signal could be detected in the absence of H2AX (Fig. 2c, d). Identical results were obtained in cells exposed to UV damage (Supplementary Fig. S2c, d). Furthermore, live-cell imaging showed that MCPH1-BRCT-eGFP failed to form discernible foci in H2AX-deficient RPE-1 cells subjected to DNA damage (Supplementary Fig. S2g and Supplementary Videos S1 and S2). Taken together, these experiments confirm the requirement of γH2AX for MCPH1-BRCT-eCR recruitment to DNA damage sites.
a Representative live-cell imaging of mESCs expressing MCPH1-BRCT-eGFP. Following treatment of mESCs with ATM inhibitor (ATMi) for 24 h, cells were treated with CPT-Olap for 1 h and immediately imaged. Scale bars: 5 µm. b Quantification of the number of discernible eGFP foci per nucleus from mESCs from (a). Data represent mean ± SD from at least 70 nuclei (segmented based on the nuclear eGFP background signal), of one of three independent biological replicates. c Representative immunofluorescence micrographs of fixed, wild-type and H2AXKO RPE-1 cells showing the nuclear localization of the stably expressed MCPH1-BRCT-eGFP or the nuclear eGFP control (green) and their colocalization with γH2AX (red). Co-staining of RPE-1 cells with TP53BP1 (gray) is also shown and indicates the presence of γH2AX-independent 53BP1 foci not detected by MCPH1-BRCT-eGFP. Nuclei were co-stained with DAPI. DAPI-based nuclear masks are outlined in yellow dashed lines. All scale bars are 5 µm. d Percentage colocalization of nuclear eGFP signal (green) with γH2AX (red). Box plots represent the median (50th percentile central line), the 25th and 75th percentiles (bounds of the box), and whiskers (extended from the smallest to the largest values) of 55 nuclei of one of three independent biological replicates.
To test if the individual BRCT domains of MCPH1 are sufficient for the recruitment to DSB sites, we next generated mESC lines stably expressing either the BRCT2 or the BRCT3 domain alone and examined their recruitment to γH2AX sites upon treatment of cells with ETO (Supplementary Fig. S2e). Recruitment of both individual BRCTs to γH2AX sites is partially compromised relative to the recruitment of the tandem MCPH1-BRCT (Supplementary Fig. S2e, f), further confirming that their association in tandem plays a crucial role in recruitment to DSB sites. To assess the specificity of the selected eCRs, we introduced point mutations known to disrupt the recruitment of full-length MCPH1 to DNA damage foci to the BRCT domains16. We generated cell lines expressing MCPH1 tandem BRCT harboring a single point mutation within the BRCT2 (MCPH1 BRCT2-W706R-BRCT3-WT) or BRCT3 (MCPH1 BRCT2-WT-BRCT3-W815R) domains. Recruitment to ETO-γH2AX sites was strongly reduced in both cell lines, much more so than the recruitment of individual wild-type BRCT2 or BRCT3 (Supplementary Fig. S2e, f). The results using a point mutation within a single BRCT, while keeping the correlate BRCT intact, corroborate the idea that their presence in tandem enhances their affinity toward γH2AX, as previously observed with other BRCT domains24. This result also indicates that both selected point mutations are weakening the cooperative affinity of both individual BRCTs toward γH2AX. Altogether, these data strongly suggest that the MCPH1-BRCT eCR is specifically recruited to DNA damage sites by binding γH2AX via its BRCT tandem domain.
Previous studies have designed probes specific to γH2AX5,7, among them, one study designed a bivalent nanobody targeting γH2AX6. We introduced the cDNA encoding this γH2AX-specific bivalent nanobody fused to eGFP into mESCs using RMCE and compared it side by side with our MCPH1-BRCT-eCR (Supplementary Fig. 3a). Overall, we observed a lower colocalization of the nanobody with DNA damage sites marked by γH2AX induced by ETO (Fig. 3a, b). We also observed a reduced number of RAD51 foci in the presence of the nanobody, when compared to cells expressing the MCPH1-BRCT-eCR or the eGFP background control eCR (Supplementary Fig. S3b, c). Furthermore, time-lapse live-cell imaging after DNA damage induction indicates slower localization kinetics of the nanobody-based probe (Supplementary Fig. S3d–g and Supplementary Videos S3–S5). This shows that our eCR probe has faster localization kinetics at DNA damage sites and does not interfere with the binding of endogenous proteins.
a Representative immunofluorescence images of fixed mESCs stably expressing MCPH1-BRCT-eGFP or the (C6) bivalent-γH2AX-nanobody fused to eGFP. Nuclear eGFP was used as a negative control. Cells were treated with either ETO or DMSO for 1 h and stained with antibodies against γH2AX (red). Nuclei were co-stained with DAPI. DAPI-based nuclear masks are outlined in yellow dashed lines. Scale bars: 5 µm. b Percentage colocalization of nuclear eGFP signal with γH2AX. Box plots represent the median (50th percentile central line), the 25th and 75th percentiles (bounds of the box), whiskers (extended from the smallest to the largest values), and outliers (individual data points) of 55 nuclei of one of three independent biological replicates. c Representative filmstrips (pre-bleached, bleached, and post-bleached (5 s)) from FRAP experiments performed in live U2OS cells stably expressing MCPH1-BRCT-eGFP. Cells were treated with ETO (within a damage foci) or DMSO (outside spontaneous damage foci), for ~1 h. Zoomed regions of interest (ROI) are depicted on the upper-right side of each image. Scale bars: 50 µm. d FRAP recovery curves of U2OS cells from (c). Data shows the mean intensity (pre-bleached and background-normalized ROIs) ± SD from at least 12 randomly selected ROIs per condition. The t=hl (time of the half-life) of each curve is indicated. Representative of two biological replicates.
This behavior is likely due to a short residency time of the MCPH1-BRCT-eCR on chromatin. To test this dynamic binding, we performed FRAP (Fluorescence Recovery after Photobleaching) in U2OS cells expressing the MCPH1-BRCT-eCR and calculated the recovery half-life of eGFP foci from independent nuclei upon exposure to ETO for ~1 h and cells treated with DMSO. The half-life of MCPH1-BRCT-eGFP in ETO-damaged nuclei (time-half life: t-hl = 1.34 s ± 0.86) was nearly identical to undamaged control cells (t-hl = 1.07 s ± 0.76), suggesting that the binding and diffusion behavior of MCPH1-BRCT is not affected when chromatin is exposed to DNA damage (Fig. 3c, d and Supplementary Fig. S3j). Furthermore, the averaged retention half-life of MCPH1-BRCT-eGFP was only twice as long when compared to the half-life of the diffuse NLS-eGFP, which served as a non-interacting protein control (t-hl = 0.83 s ± 0.49; t-hl = 0.74 s ± 0.51, undamaged and damaged chromatin, respectively) (Fig. 3c, d and Supplementary Fig. S3h–j). This suggests that the MCPH1-BRCT-eCR probe repeatedly binds to and dissociates from chromatin with fast on-off rates.
Expression of MCPH1-BRCT-eGFP is compatible with endogenous DSB formation in living cells
Since we have already shown that our sensor can be used in different cellular contexts, we generated additional MCPH1-BRCT eCRs-expressing clones of human osteosarcoma (U2OS) cells, which are better suited for assessing the effect of the eCR on endogenous DNA damage kinetics (Supplementary Fig. S4a, b). Consistent with the findings obtained in mESC, the MCPH1-BRCT eCR colocalized to γH2AX-marked damage sites after exposure of U2OS cells with ETO (Supplementary Fig. S4c, d) and required the ATM kinase for its localization (Supplementary Fig. S4e–g).
First, we tested if the presence and binding of the MCPH1-BRCT eCR would interfere with the binding of endogenous DNA repair factors. Since 53BP1 recruitment to damage sites occurs relatively early and directly downstream of γH2AX and is affected by the recruitment of MDC125, we argued that this would be the best readout to test potential interferences by our probe. By comparing the localization of 53BP1 in the presence of the MCPH1-BRCT probe and the eGFP-only control, we could not identify any differences in 53BP1 recruitment to DNA damage sites (Fig. 4a, b and Supplementary Fig. S4h). The same was true for the HR factor RAD51, whose recruitment remained unaffected by MCPH1-BRCT expression (Fig. 4a, c and Supplementary Fig. S4h). Altogether, these results suggest that the probe does not interfere with the recruitment of DNA damage factors and does not affect the global DNA repair pathway choice.
a Representative immunofluorescence micrographs of 53BP1, RAD51, and γH2AX (red) in fixed U2OS cells stably expressing MCPH1-BRCT-eGFP or nuclear eGFP control (green), treated with ETO for 1 h. Nuclei were co-stained with DAPI. The nuclear mask, based on DAPI signal, is shown as yellow dashed lines for anti-RAD51 and anti-γH2AX images. Scale bars: 10 µm. b left chart: Number of 53BP1 foci per nucleus, quantified across three independent experiments from cells from (a), Data represent mean ± SD, n = 60 nuclei per experiment. Chart to the right: Quantification of cells with >10 53BP1 foci as represented in (a). c Chart to the left: Number of RAD51 foci per nucleus, quantified across three independent experiments from cells from (a). Data represent mean ± SD, n = 50 nuclei per experiment. Chart to the right: Quantification of cells with >5 RAD51 foci as represented in (a). Percentage colocalization of (eGFP-anti-53BP1), (eGFP-anti-RAD51), and (eGFP-anti-γH2AX) can be found in Supplementary Fig. S4h. Source data are provided as a Source Data file.
Having shown that the probe does not interfere with recruitment of DNA repair factors to chromatin based on imaging in fixed cells, we further wanted to exclude the possibility that its expression interferes with global DNA repair kinetics and cellular function. Cells stably expressing the eCR showed similar sensitivity to different doses of UV radiation and different concentrations of CPT and ETO, as observed in cells expressing eGFP only, or wild-type U2OS cells (Supplementary Fig. S5a–c). Clonogenic survival assays of U2OS cells exposed to ETO and UV radiation further confirmed these results and suggest that expression of the eCR does not interfere with the ability of cells to grow and form colonies (Supplementary Fig. S5d–g).
By monitoring the kinetics of γH2AX in U2OS cells continuously exposed to ETO for 24 h or pulsed for three hours and then grown in medium without ETO for 24 h, we could test whether the presence of eCRs affects the ability of cells to recover from DNA damage alone (Supplementary Fig. S6a). Measurement of γH2AX levels by flow cytometry at different time points did not indicate differences between wild-type cells or cells expressing MCPH1-BRCT-eGFP or eGFP alone (Supplementary Fig. S6b and S7a, b). In all cases, a partial clearance of γH2AX was observed already after 12 h post-release from ETO, whilst an almost complete clearance was observed 24 h post-release, indicative of completed DNA repair. Levels of γH2AX within the cells continuously exposed to ETO kept increasing and plateaued at 24 h (Supplementary Fig. S6b and S7a, b). We confirmed the results obtained by flow cytometry by assessing γH2AX as well as 53BP1 resolution by immunofluorescence with cells fixed at different time points after release from ETO (Supplementary Fig. S6c–f). In addition, cell cycle profiling using bivariate DNA/PCNA flow cytometry analysis did not reveal differences in the cell cycle progression of U2OS cells expressing either MCPH1-BRCT-eGFP, eGFP, or wild-type cells (Supplementary Fig. S7c, d). Altogether, these results support that expression of the MCPH1-BRCT eCR does not interfere with the endogenous kinetics of DSBs repair processes.
MCPH1-BRCT-eCR is dynamically recruited to γH2AX sites and tracks DNA damage kinetics in living cells and animals
The short residency time of the probe on chromatin and the absence of interference with endogenous DSB repair suggest that it could be employed as a probe to follow the kinetics of DSB formation in living cells. This would overcome current limitations imposed by conventional antibodies directed against γH2AX, which can only be employed in fixed/permeabilized cells. Time-lapse confocal microscopy following addition of CPT was conducted in mESC and U2OS cell lines expressing the MCPH1 probe, including eGFP controls, over a period of 90 min, capturing images every 5 min (Supplementary Fig. S8a, b). Estimation of the number of foci per nucleus revealed a gradual increase in the number of MCPH1-BRCT-eGFP foci already after 15 min following addition of CPT in both cell lines (Supplementary Fig. S8c, d). Extended acquisition over a period of approximately 15 h in mESCs revealed a gradual increase in the number of foci post exposure to CPT, plateauing around 6 h, and remaining stable up until ~15 h (Fig. 5a, b; Supplementary Videos S6 and S7; eGFP controls: Supplementary Videos S8 and S9). This was different from results obtained in U2OS cells, where formation of eCR foci peaked at 9 h post CPT treatment and decreased over time (Fig. 5c, d; Supplementary Videos S10 and S11). This difference is likely due to the absence of a G1 checkpoint, which may allow mESC to enter S phase without fully repairing the DSBs26, while U2OS cells promote DSB repair before entering S phase. The specificity of the eCR sensor was confirmed by point mutations in both MCPH1-BRCT domains, which significantly reduced the number of fluorescent eGFP foci upon treatment of live mESCs and U2OS with CPT. Importantly, the point mutations also reduced the number of MCPH1-BRCT foci in undamaged mESC and U2OS cells (DMSO-treated), attesting to the sensitivity of the eCR as a γH2AX readout (Supplementary Fig. S8e–h).
a Representative filmstrips of time-lapse fluorescence microscopy obtained from proliferating mESCs expressing MCPH1-BRCT-eGFP at different time points and imaged after the indicated treatments (from Supplementary Videos S7 and S8). The ROI (white dashed line) around selected nuclear eGFP is shown. b Quantification of nuclear eGFP foci from (a). Cells were imaged every 5 min right after the addition of the indicated agents; quantification was performed with at least 50 nuclei from two independent time-lapse microscopy experiments performed with independent biological replicates, using images captured every 3 h. Data represent mean ± SD. c Representative filmstrips of time-lapse fluorescence microscopy obtained from proliferating U2OS cells expressing MCPH1-BRCT-eGFP and imaged after the indicated treatments (from Supplementary Videos S10 and S11). The ROI (white dashed line) around selected nuclear eGFP is shown. All scale bars are 5 µm. d Quantification of nuclear eGFP foci from U2OS cells from (c). eGFP foci per nuclei were quantified according to (b) from at least 50 cells obtained from two independent time-lapse microscopy experiments performed with independent biological replicates. Data represent mean ± SD. e Representative confocal images of MCPH1::NLS::eGFP in control (wild type) and spo-11-depleted live C. elegans. 2.5× zoomed images are also depicted. Scale bars: 50 µm. f Percentage of nuclei with the indicated amount of MCPH1::eGFP foci in wild-type and spo-11 animals. Data represent mean ± SE from 600 to 650 randomly selected nuclei from at least 11 biological replicates. Source data are provided as a Source Data file.
To further explore the applicability of the MCPH1-BRCT eCR as a damage sensor beyond induced DNA damage in cell culture conditions, we introduced a slightly modified construct expressing the eGFP-tagged MCPH1-BRCT eCR into the Caenorhabditis elegans genome to monitor programmed meiotic DNA DSBs induced by the topoisomerase-like protein Spo-11 (SPO-11). We detected the presence of distinct MCPH1-BRCT-eGFP foci in the nematode germline, which was significantly reduced upon depletion of spo-11 (Fig. 5e, f). Collectively, these results indicate that the MCPH1-BRCT eCR captures DSB repair kinetics and can be employed as a damage sensor in living cells and animals.
MCPH1-BRCT eCR allows visualization and genome-wide detection of targeted DNA damage sites
The DNA damage generated by the genotoxic conditions utilized above results in DSBs that are randomly distributed along the genome, and it remains unclear if our probe can bind to DNA damage sites that are experimentally induced at defined genomic sites and/or if the chromatin context could influence the recruitment. To evaluate whether our sensor can also bind to DNA damage sites triggered by a single DSB, we made use of available RPE-1 cells expressing an inducible Cas9 under transcriptional and proteasomal control27 and introduced our MCPH1-BRCT eCR probe in the presence of guide RNAs that target either one or four genomic loci (Fig. 6a). We reliably detected DNA damage sites following DOX and Shield-1 treatment, with sufficient sensitivity to visualize DSBs induced by sgRNAs targeting even a single genomic locus (Fig. 6b, c).
a Schematic representation of CRISPR-Cas9 with specific sgRNAs targeting different loci in human RPE-1 cells chromosomes (chr) and mouse Major Satellite DNA repeats (MSRs), which are present in all but Y chromosomes: 40, XX. A schematic depicting the recruitment of MCPH1-BRCT-eGFP to γH2AX (S139P – phospho-Serine 139) sites triggered by these sgRNAs is shown. Chromosome schematics (green) were retrieved and modified from https://www.ensembl.org/Multi/Tools/Blast. Arrows (1 copy – red arrow, 4 copies – black arrows, MSRs, orange arrows) indicate the chromosomal target of the sgRNAs. b Representative micrographs of fixed RPE-1 cells stably expressing MCPH1-BRCT-eGFP treated with Shield-1 and Doxycycline (Dox) for 12 h and transfected with sgRNAs targeting 1 or 4 genomic locations for 7–8 h. A plasmid without sgRNA was used as a control. Cells were stained with anti-Cas9 (gray) and anti-γH2AX (red), and nuclei were co-stained with DAPI. Nuclear masks, based on DAPI signal, are shown. Scale bars: 5 µm. c Quantification of the number of discernible nuclear γH2AX and eGFP foci per Cas9-positive nuclei (n = 55). Box plots represent the median (50th percentile central line), the 25th and 75th percentiles (bounds of the box), and whiskers (extended from the smallest to the largest values). d Representative immunofluorescence images of fixed mESCs expressing Cas9+sgRNA#3 showing the recruitment of Cas9 (anti-Cas9, red) and MCPH1-BRCT-eGFP (green) to DAPI-stained chromocenters. Cells expressing Cas9 without the guide were used as controls. Nuclei were co-stained with DAPI. Scale bars: 5 µm. e Quantification of the percentage of eGFP-positive chromocenters per nucleus in mESC expressing MCPH1-BRCT-eGFP transfected with Cas9+sgRNA#3. Only chromocenters stained by anti-Cas9 were considered. Box plots represent the median (50th percentile central line), the 25th and 75th percentiles (bounds of the box), and whiskers (extended from the smallest to the largest values) of 55 cells of one of three technical replicates performed independently. f Percentage colocalization between eGFP and anti-Cas9-positive nucleus in cells expressing MCPH1-BRCT-eGFP and transfected with Cas9+sgRNA#3. Box plots represent the median (50th percentile central line), the 25th and 75th percentiles (bounds of the box), and whiskers (extended from the smallest to the largest values) of 100 cells of one of two technical replicates performed independently. Source data are provided as a Source Data file.
To test whether our sensor can also bind to DNA damage targeted to less-accessible sites in the genome, we expressed Cas9 and individual guides targeting Cas9 to major satellite repeats (MSR) residing in the heterochromatin of mESCs28 (Fig. 6a). This system was previously used to target DSBs to MSRs in mESC, which can be visualized as dense chromocenters stained by DAPI or Hoechst29. We show that, as expected, in the presence of the guides, Cas9 is indeed targeted to these chromocenters, resulting in elevated levels of γH2AX (Supplementary Fig. S9a, b). By expressing the MCPH1-BRCT eCR in the presence of the MSR guides, we observe that the eCR reliably localizes to sites of DNA damage, colocalizing with chromocenters marked by γH2AX (Fig. 6d–f and Supplementary Fig. S9c–e). This localization to chromocenters was not observed when we introduced a catalytically dead dCas9, highlighting that the binding is dependent on the DSB, and not mediated by unspecific interactions with Cas9 or the guide RNA (Supplementary Fig. S9f). We further evaluated if the binding of the eCR would alter the chromocenter structure by measuring their sphericity, volume, and estimating their numbers in the presence or absence of the probe and Cas9. While we see no significant changes in chromocenter sphericity and numbers in any of the tested conditions, we observe that chromocenter volume is increased when active Cas9 is targeted to MSR (Supplementary Fig. S9g–i). This, however, is independent of the MCPH1-BRCT eCR, since the volume of chromocenters is unchanged between cells expressing the probe relative to cells expressing the eGFP control only (Supplementary Fig. S9h). This indicates that the eCR can detect γH2AX at sites targeted for DNA damage, even within densely packed chromatin.
Encouraged by these results, we wanted to test the applicability of the MCPH1-BRCT eCR for the detection of DNA damage genome-wide. Toward this, we introduced the probe to available U2OS DIvA cells expressing an inducible AsiSI restriction enzyme that creates DNA damage at defined sites in the genome30. Addition of 4-OHT to the medium results in accumulation of DSBs that can be visualized by the MCPH1 probe (Fig. 7a–c, Supplementary Fig. S10a–c, and Supplementary Video S12). Since in this system the DNA damage occurs at defined genomic locations, we evaluated the genome-wide localization of the MCPH1-BRCT eCR in relation to the previously identified BLESS DNA damage sites31. Toward this, we performed ChIP-seq enriching for MCPH1-BRCT-eGFP and γH2AX to directly identify the DSB sites in these cells in the presence of 4-OHT and DMSO. Sequencing and analysis of the material enriched in 4-OHT over DMSO conditions (Fig. 7d) revealed the genome-wide localization of the MCPH1 probe, resulting in accurate colocalization with γH2AX over known BLESS DSBs in the presence of 4-OHT (Fig. 7e and Supplementary Fig. S10d, e). This confirms the specificity of the MCPH1 probe for γH2AX at the induced sites, and furthermore, its suitability for genome-wide studies.
a Schematic of the DIvA (DSB Inducible via AsiSI) cell line, which expresses a restriction enzyme (AsiSI) fused to a modified estrogen receptor ligand-binding domain (AsiSI-ER). Addition of 4-hydroxytamoxifen (4-OHT) relocates AsiSI-ER to the nucleus, inducing multiple sequence-specific DSBs across the human genome. MCPH1-BRCT-eGFP is then targeted to these sites. b Representative snapshots of live U2OS cells stably expressing MCPH1-BRCT-eGFP (wild type or DIvA), incubated with 4-OHT and imaged over 4 h. Scale bars: 50 µm. c Quantification of the number of eGFP foci per nucleus in the U2OS DIvA after addition of 4-OHT. Data represent mean SD from ±25 nuclei retrieved from two independent time-lapse fluorescence microscopy experiments (see representative Supplementary Video S12). d Scatter plot showing the log2-fold change of γH2AX and MCPH1-BRCT-eGFP ChIP-seq signal in +4-OHT versus –OHT conditions across 1.280 in silico called AsiSI sites in the human genome. ChIP-seq enrichment was quantified in ±10 kb windows centered on the DSBs. Red stars highlight the curated set of 80 DBSs list of BLESS (Breaks Labeling, Enrichment on Streptavidin, and next-generation Sequencing)-defined DSB sites31. Pearson’s r = 0.79. e Representative genome browser view representing the ChIP-seq signal at a single DSB located on chromosome 1. Shown are log2-fold change (+4-OHT/−4-OHT) signals of CPM normalized read counts per 200 bp. A curated list of 80 DSB described in (d) is indicated at the bottom row. Source data are provided as a Source Data file.
Discussion
While many efforts to detect γH2AX in living cells have been made5,6,7, current methods to monitor its distribution rely mainly on endpoint measurements in cells subjected to fixation/permeabilization and antibodies, suffer from limited specificity and sensitivity, or from laborious production procedures. Although such methods allow the quantification of γH2AX foci over time, tracking the early kinetics of DSBs in a spatiotemporal manner is limited by the lack of effective and compatible probes. Tagging of endogenous full-length proteins such as 53BP132, or expression of a truncated version, has been used as a probe for DSBs in living cells33. However, recruitment of repair proteins to DSBs can be influenced by the specific repair pathway choice and therefore preclude their use as a neutral DSB reporter in living cells. In this study, we present a novel DNA damage sensor to track DNA damage in living cells. Toward this, we established engineered chromatin readers (eCRs) selective to γH2AX by screening selected tandem BRCT domains from well-characterized proteins associated with DNA damage response. These eCRs were introduced to mouse stem cells, as previously described8, and assessed for their recruitment to DNA damage sites marked by γH2AX. Amongst the selected candidates, the tandem BRCT domain of MCPH1 (BRCT2 and BRCT3 linked by a small unstructured linker) exhibited the highest affinity toward DSBs. Although single BRCT domains have also been previously shown to interact with γH2AX phosphopeptides34, our data indicates that single BRCT versions of the tandem MCPH1-BRCT were not sufficient to interact with γH2AX sites efficiently – in line with the model where recognition of γH2AX is achieved when the phosphorylated residue is sandwiched by two separated singleton BRCTs35. We show that the dynamic formation of MCPH1-BRCT-eGFP foci is entirely dependent on the presence of H2AX, and furthermore, on phosphorylation of H2AX by ATM, confirming that foci detection faithfully reports the H2AX phosphorylation status and DSBs.
By using several quantitative and functional approaches, we have validated the feasibility of the tandem BRCT domain of MCPH1 to cytologically monitor spatiotemporal dynamics and kinetics of DNA DSBs. Our results demonstrate that using natural BRCT reader domains allows a more physiological engagement with chromatin, resulting in a dynamic and less disruptive readout of DNA damage when compared to nanobody-based solutions. Furthermore, we show that the eCR performs well in both fixed and living cells and can be used as an antibody alternative in standard chromatin immunoprecipitation and immunofluorescence microscopy protocols. By targeting DNA damage to distinct genomic sites via Cas9, we could further show that our probe is applicable to detect single DSBs as well as DSBs in highly condensed regions of the nucleus marked by heterochromatin28, providing a potential tool to probe DNA damage mechanisms in different chromatin environments. We further show that the probe is suitable for mapping genome-wide DSBs, which we detected by ChIP-seq in available DIvA U2OS cell lines30.
Importantly, we show that the presence of the DNA damage sensor does not interfere with endogenous DNA repair pathways, since its interaction with γH2AX is highly dynamic, resulting in short retention time on chromatin. This allows the use of the sensor in living systems to profile DNA damage dynamics in various settings. Live imaging in ES and U2OS cells indicated that the presence of MCPH1-BRCT on chromatin was detected shortly after the induction of DSBs in the form of discrete nuclear foci and was resolved upon removal of the drug that triggered DSBs. The evolutionary conservation of H2A phosphorylation at DNA damage sites36 allowed us to apply the DNA damage sensor to detect meiosis-specific DSBs in the C. elegans germline. These findings highlight its applicability to follow the spatiotemporal resolution of DSBs and gain significant insights into DSB kinetics in different model systems ranging from cell lines to living animals.
We expect this probe to be applicable to a diverse set of conditions, ranging from its usage as a reliable reporter of DNA damage in biomedical and clinical settings to its application as a recombinant protein to replace antibodies, as well as in high-throughput omics studies aimed to identify the genomic and proteomic context associated with DNA damage in a wide range of living organisms.
Methods
Selection of BRCT domains
All singleton BRCT domains from Mus musculus (Fig. 1a) were manually retrieved from Uniprot. Tandem BRCT domains, comprising two or more closely located BRCT domains linked by a variable linker region, were manually retrieved, and their association in tandem was further confirmed based on the literature9,37. Amino acid sequences of singleton BRCTs were aligned using MAFFT38 (with the FFT-NS-I iterative method and default parameters) using the E-INS-I iterative method, recommended for sequences with conserved domains and long gaps. Amino acid sequences of the selected tandem BRCTs were also aligned using MAFFT using the E-INS-I iterative method, recommended for sequences with conserved domains and long gaps. Maximum-likelihood unrooted phylogenetic trees were inferred from multiple amino acid sequence alignments with PhyML version 3.0 with the WAG model39 and edited with the Interactive Tree of Life tool (iTOLv5)40. Supporting values for the inferred tree were generated using BOOSTER using 1000 Bootstraps replicates41.
Molecular cloning
The selected mouse tandem BRCT domains (Fig. 1b; Supplementary Data 1) were synthesized by Integrated DNA Technologies (IDT) based on available Uniprot annotations. The amino acid sequence of the bivalent nanobody raised against γH2AX (clone C6) was retrieved from ref. 6, reverse translated, and codon optimized for expression in mice. All these constructs were assembled according to ref. 8 into the Recombinase-mediated cassette exchange (RMCE) targeting vector partbit-v6 (Addgene 179392) by Gibson assembly. The final construct harbored the reader domains or the bivalent nanobody, N-terminally tagged with a biotin acceptor in frame to a cassette containing an NLS, an eGFP, followed by the puromycin-N-acetyltransferase or the Blasticidin (BlastR) resistance gene expressed through an Internal Ribosome Entry site (IRES). Single MCPH1 BRCTs, BRCT2, and BRCT3 were cloned into the same constructions without the linker. Point mutations in the BRCT2 and 3 (W706R and W815R) were based on ref. 16. Expression of all constructs is driven by the constitutive CAG promoter. To generate the construct expressing the catalytic dead Cas9 (dCas9), the DNA sequence from the plasmid pdCas9 (Addgene 46569) was Gibson assembled into the EcoRI and AgeI sites of the plasmid PX330 (Addgene 42230). Small guide RNAs (sgRNAs) sequences targeting murine major satellite repeats (MSRs) are from ref. 29 sgRNA#1:GGCGAGAAAACTGAAAATCA; sgRNA#2:GCGAGGAAAACTGAAAAAGG; sgRNA#3: GAAATGTCCACTGTAGGACG, and were cloned onto pX330 plasmid (Addgene 42230) or pX330-dCas9 according to ref. 42. sgRNAs sequences targeting single (GCCGATGGTGAAGTGGTAAG) - or 4 (TGGACTGCAGTACACAATCA) copy loci in the human iCut-RPE-derived cells are from ref. 27 and were cloned into the pX458-Dual-Guide-Donor-ShuttleOnly (Addgene 171098), as described above. All constructs were confirmed by Sanger Sequencing.
CRISPR/Cas9-mediated genome editing
The amino acid sequence of the mouse MCPH1-BRCT was reverse translated to C. elegans germline-optimized codons. This sequence, followed by DNA encoding a 12-amino acid germline-optimized linker (IGGGGSGGGGSA) and a germline-optimized EGL-13 NLS for optimal nuclear localization, was synthesized in a single gblock (IDT). This cassette was then PCR amplified to generate overhangs for CRISPR-Cas9 editing. For this, we used an existing C. elegans strain and placed this whole module between an eft-3 promoter for ubiquitous expression, and a germline-optimized GFP with smu-1 introns (to prevent germline silencing), followed by a tbb-2 3′ UTR. The he423 [peft-3::MCPH1-BRCT::NLS::eGFP(GLO)] allele in the strain SV2569: he423 [peft-3::MCPH1-BRCT::NLS::eGFP(GLO)] was generated using the homology-directed repair of CRISPR/ Cas9-induced DNA double-strand breaks, for which the reagents were micro-injected into the gonad of young adults. The allele was generated using the Alt-R CRISPR/Cas9 system (IDT), as previously described43. As a repair template, MCPH1-BRCT was amplified from a gBlock (IDT) using primers with 5′ SP9 modifications (IDT). MCPH1-BRCT was inserted into a pre-existing allele by generating double-stranded breaks using two sgRNAs. To verify the edit, the insertion was PCR amplified and sequenced by Sanger sequencing.
RNA-mediated interference (RNAi)
For RNAi of C. elegans, bacterial cultures of E. coli HT115 containing L4440 empty vector or vector with genomic gene inserts were grown overnight, induced with 1 mM IPTG for 1 h, 5 times concentrated, and seeded onto NGM plates containing 12.5 μg/mL Tetracycline, 100 μg/mL Ampicillin, and 2 mM IPTG. RNAi feeding initiated from the L4 stage, and the number of germline foci was analyzed in late L4/young adult animals of the following generation. RNAi clones for spo-11 were obtained from the Ahringer44 database and sequence verified.
Mammalian cell Culture and cell lines
Mouse embryonic stem cells (HA36CB1) were cultured as previously described8 and stable mESC lines were obtained via recombinase-mediated cassette exchange - RMCE45. Briefly, constructs were co-transfected using Lipofectamine 3000 reagents (Thermo Fisher Scientific), with a Cre-recombinase expression plasmid (Addgene 19131) into RMCE-competent mESC (HA36CB1), followed by selection with 3 mM ganciclovir (Selleckchem, S1878) and 2 μg/mL puromycin (InvivoGen, ant-pr-1). Individual clones were picked, and homogeneous and stable expression of proteins was confirmed by flow cytometry or Western blotting. Human U2OS cells46 were kindly provided by Dr. Susanne MA Lens, University Medical Center Utrecht, the Netherlands. The DIvA-AsiSI-ER-U2OS cell line30, originally generated in the laboratory of Gaëlle Legube (MCD, Centre de Biologie Intégrative (CBI), CNRS, Université de Toulouse, Toulouse, France). RPE-1 iCut cell line (RPE-1)27 and H2AXKO RPE-1 cell lines23, were kindly provided by Bas van Steensel (Division of Gene Regulation, Netherlands Cancer Institute, Amsterdam, the Netherlands) and René H. Medema (Division of Cell Biology, Netherlands Cancer Institute, Amsterdam, the Netherlands). These cells were cultured in Dulbecco’s modified Eagle’s medium – DMEM (Invitrogen, 11584486), supplemented with 1× non-essential amino acids (Thermo Fisher Scientific, 11140035), 1× Glutamax (Thermo Fisher Scientific, 11584466), 1 mM sodium pyruvate (Thermo Fisher Scientific, 11360), 1% penicillin–streptomycin antibiotics (Gibco, 15070-063) and 10% FCS (Westburg, CA FBS-11A) at 37 °C and 5% CO2. For the generation of stable U2OS expressing eGFP, parental wild-type U2OS cells were seeded in six-well plates at 0.1 × 106 cells/mL and lipotransfected using Lipofectamine 3000 reagent (Thermo Fisher Scientific) for 48 h with PvuI-linearized plasmids (Parbitv-6 constructs). Following incubation for 48 h, cells were selected with complete media containing 2 μg/mL puromycin (InvivoGen, ant-pr-1) for 10 days and maintained with 0.5 μg/mL Puromycin. Individual clones, obtained through limited dilution of a resistant polyclonal cell population in 96-well plates, were expanded, and construct integrations were confirmed by FACS and Western Blotting. For the generation of stable DIvA-AsiSI-ER-U2OS, RPE-1 iCut wild-type (clone#1) and H2AXKO RPE-1 (clone#2) cell lines stably expressing eGFP constructs, cells were seeded and transfected with FspI linearized plasmids (Parbitv-6 constructs harboring Blasticidin resistance marker), as described above. Cells were selected with complete media containing 7.5 μg/ml Blasticidin (InvivoGen - ant-bl-1) for 14 days and maintained with 2.5 μg/mL Blasticidin. Homogeneous GFP-positive cell populations were obtained by FACs sorting (FACSAria Fusion Flow Cytometer – BD Bioscience) and confirmed by FACS and Western Blotting. Where indicated, the following compounds were used at the indicated final concentrations, unless stated otherwise: 4-hydroxytamoxifen − 4-OHT (300 nM, Sigma–Aldrich – H6278), ATMi KU-55933 (10 μM, Selleckchem - S1092), Camptothecin-CPT (25 nM, Enzo Life Sciences - ALX-350-015), Cisplatin-CISP (10 μg/mL, Millipore − 232120), Etoposide-ETO (20 μM, MedChemExpress - HY-13629), Methylmethanesulfonate-MMS (0.25 mM, Sigma–Aldrich − 129925), Olaparib-OLAP (1 μM, Sigma–Aldrich - SML3705). For UV irradiation, culture media were removed, and cells were washed once with DPBS. Cells were then subjected to UV-C (254 nm) at different mJ/cm2 rates using the GS Gene Linker UV Chamber (Bio-Rad). Cells were then immediately incubated with fresh and warm media.
siRNA transfections
Individual siRNA transfections were performed using Ambion Silencer Select siRNAs and Lipofectamine RNAiMAX (Thermo Fisher Scientific), according to the manufacturer’s instructions. For depletion of ATM, 40 nM of siRNA (s1708) was used. The Silencer Select Negative Control number 1 (Thermo Fisher, 4390843) was used as a non-targeting control. Cells were fixed/collected 48 h post-transfection.
Transient transfections
Transient transfections were performed with Lipofectamine 3000 (Life Technologies) according to the manufacturer’s instructions. Briefly, cells were co-transfected with pX330-U6 Chimeric-BB-CBh-hspCas9 or pX330-dCas9 constructs cloned with small sgRNAs targeting MSRs. Cells were fixed or harvested 5–6 h post-transfection. For transfection of RPE-1 iCut cell lines stably expressing MCPH1-BRCT-eGP, Cas9 was activated by the addition of 1 μg/mL Doxycycline (Sigma–Aldrich) and 1 μM Shield-1 (MedChemExpress - HY-112210) 12 h prior to transfection with pX458-Dual-Guide-Donor-ShuttleOnly harboring the specified sgRNAs targeting the Human genome. Cells were fixed 7–8 h post-transfection. Plasmids lacking sgRNA were used as controls.
Flow cytometry
For measuring eGFP levels in live mESC and U2OS cell lines, cells were resuspended in DPBS containing 2% FCS and 1 mM EDTA and immediately analyzed on a FACSCanto II or FACSymphony A1 Cell Analyzer (BD Biosciences). For γH2AX measurements in U2OS cells, 1 × 106 cells were collected, resuspended in PBS, and permeabilized with PBS/0.1% Triton X-100/1% BSA for 10 min on ice. Cells were then fixed in ice-cold methanol for 3 min at −20 °C. Cells were spun down and pellets resuspended in PBS. Cells were blocked with 1% BSA in PBS for 1 h and then incubated with anti-γH2AX-Alexa Fluor 647 antibody (1:200) (Biolegend, 613408) or Anti PCNA-PE conjugate (1:200) (Invitrogen, 12-910-42). Cells were then washed and resuspended in DAPI staining solution (0.1% (v/v) Triton X-100-1mg/mL DAPI (Sigma–Aldrich, MBD0015)). Alternatively, cells not incubated with conjugated antibodies were fixed with 70% ice-cold ethanol at 4 °C for at least 30 min, washed with DPBS, and directly incubated with a DAPI-stained solution. Samples were acquired on a FACSCanto II or FACSymphony A1 Cell Analyzer (BD Biosciences), and data were analyzed using FlowJo software (version 10.9, Tree Star). The proportion of cells in each cell cycle stage was assessed using FlowJo with the Watson-pragmatic model, which fits the Gaussian DNA distribution curves to the stages of the cell cycle, or by gating the proportion of cells using the DNA content coupled to S phase distribution (PCNA staining). Quenching of eGFP fluorescence with Methanol fixation was controlled using single staining samples and fluorescence compensation to avoid potential bleed-through of fluorescent signals.
Western blotting
To measure the levels of eCRs, 10 μg of crude nuclear extracts, prepared as previously described47, were resolved by SDS-PAGE, transferred to nitrocellulose membranes (Amersham, GE10600001), stained with Ponceau-S (Sigma), and blocked in 5% wt/vol non-fat milk or BSA in TBS-0.1% Tween 20. To measure γH2AX levels, acidic Histone Extracts were obtained as described in ref. 8. Briefly, cells were lysed with NETN buffer (20 mM Tris, pH 8, 0.5% (vol/vol) NP-40, 100 mM NaCl, and 1 mM EDTA, pH 8), supplemented with 1× protease inhibitor cocktail (PIC) (Roche) and dithiothreitol (DTT) (Sigma–Aldrich). Cell lysates were centrifuged, and histones were extracted by resuspending pellets with 0.2 N HCl overnight. The acidic histone extract was centrifuged, and the histone extracts were then neutralized with 1 N NaOH. 3 μg of extracts was resolved by SDS-PAGE. Membranes were then probed with the following primary antibodies overnight at 4 °C: anti-GFP (Abcam, Ab290), anti-CRISPR/Cas9 (Abcam, ab191468), anti-Lamin-B1 (Santa Cruz Biotechnology, sc-6216), anti-γH2AX Phospho Ser. 139 (Biolegend, 613402), anti-Histone H3 (Abcam, ab1791), anti-Histone H1.2 (Abcam, Ab17677), and anti-Histone H2A.X (EPR895, Abcam, 124781). Detection was done with species-specific horseradish peroxidase-conjugated secondary antibodies and Pierce Peroxidase IHC Detection Kit (Thermo Fisher Scientific). Alternatively, membranes were also incubated with StarBright Blue 700 Fluorescent Secondary antibodies (Bio-Rad).
Nuclear extraction for GFP pull down
mESCs were treated with ETO for 1 h, collected in 50 mL Falcon tubes, washed once with PBS, and centrifuged at 400 × g for 5 min at 4 °C. Pellets were resuspended in 12 mL ice-cold PBS and transferred to 15 mL Falcon tubes. After harvesting, the supernatant was removed and resuspended in Buffer A (10 mM HEPES-KOH (pH 7.9), 1.5 mM MgCl₂, 10 mM KCl), according to the pellet volume. After incubation on ice for 10 min, cells were centrifuged, and the supernatant was aspirated. The pellet volume was determined again, and cells were resuspended in 2 volumes of Buffer A supplemented with 0.15% NP-40 and an EDTA-free protease inhibitor cocktail (Roche). The suspension was then homogenized using a glass Dounce homogenizer (Kimble Kontes) with a tight-fitting pestle (type B), applying 40 strokes. The homogenate was transferred back to a 15 mL Falcon tube and centrifuged for 15 min, 3200 × g at 4 °C. The cytoplasmic extract (supernatant) was discarded, and the pellet was washed with 10 volumes of PBS. The supernatant was then discarded, and the pellet volume was noted. Nuclei were resuspended in 2 volumes of Buffer C (420 mM NaCl, 20 mM HEPES-KOH (pH 7.9), 20% glycerol (v/v), 2 mM MgCl₂, 0.2 mM EDTA, 0.1% NP-40 (v/v), 1× EDTA-free protease inhibitor cocktail (Roche), 0.5 mM DTT). Resuspended nuclei were transferred to Eppendorf tubes, gently mixed by pipetting up and down 10 times, and rotated for 1 h at 4 °C. Samples were centrifuged (45 min, 21,400 × g, 4 °C), and the supernatant containing the nuclear extract was collected. Protein concentration was determined using the Qubit Protein Assay (Thermo Fisher).
Treatment of nuclear extract with lambda phosphatase
Nuclear extracts originated from mESCs treated with ETO were divided into separate tubes (250 µg of nuclear extract per condition), and the volumes were adjusted to 500 µl with Buffer B containing 1 mM MnCl₂ and fresh 1× EDTA-free protease inhibitor cocktail (Roche). The extracts were then either treated with 1 µl of λ-phosphatase (NEB, P0753S), treated with the same amount of λ-phosphatase together with 1× PhosSTOP (Roche, 4906845001), or left untreated, as a control. Nuclear extracts were treated for 30 min at 30 °C in a thermomixer. 0.1% NP-40 was added to the extracts, which were immediately used in pull-down experiments.
GFP Pull down
For GFP Pulldowns, 15 μl of GFP-trap magnetic beads (Chromotek, gtma-20) was washed according to the manufacturer’s protocol and incubated with Nuclear Extract. Pulldowns were performed by rotating the beads with the nuclear extracts for 90 min at 4 °C. The beads were bound to a magnetic rack and washed three times with 1 ml PBS with 0.5% NP-40, two times with 1 mL buffer C (0.1% NP-40), and two times with 1 mL PBS (without NP-40). Supernatant was then removed, and beads were resuspended in 2× SDS-sample buffer (Laemmli) and analyzed by western blotting with anti-GFP (Ab290), anti-γH2AX (Biolegend, 613402), anti-H2AX (Abcam, 124781), and anti-Lamin-B1 (Santa Cruz, sc-374015-B10) and detected as described above.
Immunofluorescence of fixed cells
mESC were grown on sterile coverslips (0.13–0.16 mm – Ted Pella) coated with 0.2% gelatine. U2OS and RPE-1 cells were grown on the same coverslips without gelatine. Alternatively, cells were grown on 12-well removable chambers (Ibidi, 81201) with or without 0.2% gelatine. mESCs were fixed in 3.7% formaldehyde (Sigma–Aldrich, 252549) in 1× PBS for 15 min at Room Temperature (RT). U2OS and RPE-1 cells were fixed in 2% methanol-free formaldehyde (Thermo Scientific, 28908) in 1× PBS for 10 min at RT. After fixation, cells were washed three times with 1× PBS and then permeabilized with 0.5% Triton X-100 (Sigma–Aldrich) for 5 min on ice, followed by 10 min at RT (mESC) or permeabilized with 0.1% Triton X-100 for 10 min at RT (U2OS and RPE-1 cells). Cells were washed 3× with 1× PBS for 4 min and blocked at RT for 1 h with sterile-filtered blocking buffer (1% BSA (Sigma–Aldrich) dissolved in 0.1% PBS-Tween 20). All primary and secondary antibodies were diluted in filtered blocking buffer. Primary antibody incubations were performed overnight at 4 °C in a humid chamber. Secondary antibodies were incubated for 1 h at RT. Following incubation with different primary antibodies, the coverslips were washed three times with 1×-PBS-0.1% Tween 20 and once with 1× PBS. Next, coverslips were mounted using antifading mounting medium containing DAPI (Vectashield, H2000). The following primary antibodies were used: anti-γH2AX (mouse, Biolegend − 613402, 1:1000), anti-γH2AX (rabbit, Cell Signaling – (20E3) 9718T, 1:2000), anti-CRISPR-Cas9 (Abcam, ab191468, 1:1000), anti 53BP1 (Abcam 172580,1:2000), anti-Histone H2AX (rabbit – Abcam – 124781, 1:500) and anti-RAD51 (14B4 Invitrogen MA1-23271, 1:500-1:1000). The following secondary antibodies were used: Alexa Fluor 488 Goat anti-Rabbit IgG H + L (Thermo Fisher Scientific, A11034, 1:500), Alexa Fluor 488 Goat anti-mouse IgG H + L (Thermo Fisher Scientific, 2465113, 1:500), Alexa Fluor 568 Goat anti-mouse IgG H + L (Thermo Fisher Scientific, A11004, 1:500), Alexa Fluor 647 Goat anti-Rabbit IgG H + L (Thermo Fisher Scientific, A21244, 1:500). Images of fixed cells were acquired with a Zeiss LSM 700 confocal laser scanning microscope (Biology Image Center, Utrecht University, Netherlands) with a Plan-Apochromat 63 × 1.4 NA oil immersion objective (Carl Zeiss, Germany) using 405, 488, 555, and 647 nm laser lines with a Zen imaging software (Zeiss) version 8.1.7.484. Images consist of a z-stack of 4–12 planes collected at 0.37 µm intervals and are shown at Maximum Z-projections. In Supplementary Fig. S6c, images from a single Z-plane were collected. It is worth noting that overexpression of MCPH1-BRCT-eGFP in U2OS or RPE-1 cells may interfere with detection after fixation. To avoid this, we recommend using clones with relatively lower GFP expression and avoiding transient transfection with plasmids where the eCR is driven by strong promoters. Overlap of the emission spectra (bleed-through) was corrected by automatic and manual adjustments of multiple channels, by using a negative control (wild-type untagged cells), and by performing sequential imaging.
Live-cell imaging microscopy
1.5 × 104 mESCs or 2.5 × 104 U2OS or RPE-1 cells expressing either eGFP-tagged eCRs or eGFP control cells were seeded ~16–20 h before imaging. Unless stated otherwise, mESC cells were seeded in 8-well Ibidi chambers (Ibidi, 80826) prior to being coated with 0.2% gelatine. U2OS and RPE-1 cells were seeded in uncoated 8-well Ibidi chambers (Ibidi, 80826) or 35 mm Quad-µ-Dishes (Ibidi, 80416). For single time point acquisitions (Fig. 1d, g), cells were incubated with specific phenol-red-free DMEM (Thermo Fisher Scientific, 31053028) containing Hoechst-33342 (Thermo Fisher Scientific, 62249) and immediately imaged. For time-lapse microscopy, environmental controllers were set up so that environmental chambers were stably operating under 5% CO2, at 37 °C, under 80% relative humidity prior to incubation of ibidi chambers. Chambers containing the cells incubated with phenol-red-free DMEM without genotoxic agents or vehicle control (half of the volume) were carefully transported to environmental chambers. Immediately after positioning of cells, establishment of data points, and additional imaging parameters, chambers were topped up with the other half of the media to reach the final drug concentration. After ensuring that the image fields remained in focus, time-lapsephotography was immediately started to prevent any delays. Time-lapse microscopy from Supplementary Videos S1–S4 and S12 and images shown in (Figs. 1d, g; 7b, c and Supplementary Fig. S3d–g) were acquired on a Nikon Eclipse Ti-E microscope equipped with a motorized stage (ASI), Nikon Plan Apo, driven by MetaMorph Microscopy Automation & Image Analysis Software (Molecular Devices) and equipped with a Yokogawa CSU-X1-A1 confocal head, using Plan Apo VC 60×/1.40, oil, WD = 0.15 (Nikon, Japan), 9.1 pix/μm or 109 nm/pix an Andor iXon DU-885 camera. A Z-stack of around 6–10 slices (2 μm increment, 0.5 μm step size and range of 6 around the centre). Images were obtained using the ET525/50 m (GFP) or ET460/50 m (DAPI) filters with exposure time settings ranging from 50 to 200 milliseconds (ms). For dual color imaging, sequential imaging was used with a maximum exposure of 0.5 s per channel. Time-lapse microscopy from Supplementary Videos S10 and S11 and images in (Figs. 2a and 5c and Supplementary Fig. S4e and S8a, b), were acquired on a DeltaVision RT imaging system - GE Healthcare equipped with a heated 37 °C chamber, Photometrics Cascade II EM-CCD camera, and four detection channels (Cell Microscopy Core, UMC Utrecht, Netherlands). Images were captured using an Olympus 60×-142 NA-UPLSApo/O oil objective and an exposure time setting of 50 ms. Appropriate 2-µm-spaced z-sections (from 4 to 8 planes) were collected, and images were further deconvolved using the SoftWoRx software 6.1, with default parameters. Time-lapse microscopy from Supplementary Videos S5–S9 and Images in (Fig. 5a) were acquired in an Olympus ScanR Screening System (ScanR Image Acquisition 3.01) (Center for Microscopy and Image Analysis, UZH, Switzerland) equipped with an inverted motorized Olympus IX83 microscope, a motorized stage, IR-laser hardware autofocus, a fast emission filter wheel, a CellVivo incubation system, and a Hamamatsu ORCA-FLASH 4.0 V2, high-sensitivity digital monochrome scientific cooled sCMOS camera. Images were captured using a 40× Phase PLAPON (NA 1.42), oil objective, and an exposure time setting of 100 ms.
Live-cell imaging of C. Elegans
Imaging of C. elegans was performed by mounting the larvae on a 5% agarose pad in 20 mM tetramisole solution in M9 (0.22 M KH2PO, 0.42 M Na2HPO4, 0.85 M NaCl, 0.001 M MgSO4). Spinning disk confocal imaging was performed using a Nikon Ti-U microscope equipped with a Yokogawa CSU-X1 spinning disk using a 60×-1.4 NA objective, and a Prime BSI Express Scientific CMOS camera. Spinning disk z-stacks were acquired using MetaMorph Microscopy Automation & Image Analysis Software.
Fluorescence recovery after photobleaching (FRAP)
For FRAP assays, U2OS cells were grown on 35 mm Quad-µ-Dishes (Ibidi, 80416) and medium containing either ETO or DMSO control was added as described above (live-cell imaging). Images were acquired on the spinning disk-based confocal scanner unit described above, with CO2 and temperature control - STXG-PLAMX-SETZ21L (TokaiHit, Japan), equipped with the Gataca iLas FRAP/Photoablation system. The 355 nm laser (Teem Photonics) of the iLAS pulse system was used to induce photodamage by targeting the laser on the spinning disk microscope in a circular region of interest (ROI) with a diameter of 7 pixels surrounding the selected foci in ETO-treated cells, the chromatin in undamaged cells and the background, at 50% laser power intensity, 2000 repetitions and 150 ms exposure. Post-bleach fluorescence was monitored in a time-lapse for a total of 7 s with a time interval of 0.100 s. Fluorescence intensities (mean per pixel) were saved for each ROI and used to determine the diffusion coefficients and t1/2 half-time of recovery using MATLAB (frap_analysis, version 1.0.0.0) according to ref. 48.
Image processing and data analysis
Image serial stacks were generated and processed with Fiji (ImageJ 64-bit; Version 2.00-rc-54/1.51 h), and appropriate Z-project planes were then selected for further image analysis. Unless stated otherwise, representative images and time-lapse microscopy videos represent a Maximum Intensity Z-projection of the adequately collected Z-planes. Quantifications under different genotoxic conditions included both S and non-S-phase cells. CellProfiler (Broad Institute, version 4.2.6)49 was used to estimate the number of γH2AX, TP53BP1, RAD51, or eGFP foci in fixed cells. Every image was corrected for illumination biases using the “Rescale Intensity” module prior to image analyses. For mESCs, images were also processed with a Gaussian filter to remove the foreground signal. Nuclei (DAPI or Hoechst stained) were segmented with the Otsu threshold and watershed modules to produce a binary image. Segmented nuclei were further identified using the IdentifyPrimaryobjects module. Subsequently, the resulting images were analyzed by a watershed algorithm with varying thresholds to detect foci outlines, depending on the cell line, using the “IdentifySecondaryobjects” function. Drug treatment did not bias the determination of parameters. Missegmented nuclei and nuclei within large colonies from mESCs were manually assessed and discarded from the analyses. The number of eGFP foci per nucleus from time-lapse microscopy of live eGFP-tagged mESCs, U2OS, or RPE-1 cells was estimated with the Imaris Software (Oxford Instruments, version 10.0.1). Firstly, a surface was created based on the GFP signal background. Generated surfaces were further segmented through a manual histogram-based thresholding. Missegmented nuclei, mitotic cell nuclei, and nuclei whose signal was oversaturated (based on assessment of out-of-range pixels) were further discarded. The number of foci per nucleus (green channel) was estimated using the “Spots” function from the Imaris software. The percentage of colocalization was based on pixel-intensity-correlation measurements using the coloc2 plugin (version 3.1.0) from Fiji – Manders’ coeficient50 with Costes method for statistical significance (PSF of 3.0 and Costes randomizations of 50) according to ref. 51. Colocalization was assessed within regions of interest defined by DAPI-based segmentation of maximum Z-projection images, after background correction and subtraction in both channels. Manders' split colocalization coefficients (m1 and m2) can vary from 0 to 1. In this work, colocalization data represents the determined Mander’s coefficient multiplied by 100. For experiments in Fig. 6 and Supplementary Fig. S9, only Cas9-positive chromocenters were included in the analysis, and chromocenters from Z-projection (Maximum Intensity) images were counted. Whenever required, cells were equally adjusted for brightness and contrast. The Fiji plugin NucleusJ 52 was used to segment nuclei and chromocenters from randomly pre-selected Cas9-positive nuclei. Nuclei with segmentation errors were excluded from further analysis. The plugin was also employed to quantify chromocenter volume, sphericity, and number from maximum intensity Z-projections, as described in ref. 52. Quantification of the number of GFP foci per nucleus in C. elegans was done according to ref. 53 using the Imaris Software (Oxford Instruments, version 10.0.1). Individual nuclei were identified as surface objects based on GFP signal background through manual thresholding. Poorly segmented nuclei were excluded from the analyses. GFP foci were then identified using the Spots function. To circumvent the lack of a marker to identify DSB transition zones, nuclei were randomly selected in different regions of multiple animals. All statistical analyses and data visualization were carried out with R and GraphPad-Prism (version 10.6.0).
Chromatin immunoprecipitation (ChIP)
ChIP assays were performed with U2OS DIvA cell lines stably expressing the MCPH1-BRCT eCR fused to eGFP. A total of 20 × 106 cells were incubated with medium containing either 300 nM 4-Hydroxytamoxifen (4-OHT) or DMSO (mock control) for 4 h prior to harvesting. Cells were crosslinked directly on plates with 1% methanol-free formaldehyde (Thermo Fisher, 28908) for 8 min at room temperature and quenched by adding glycine (Sigma–Aldrich) to a final concentration of 0.125 mM at 4 °C for 10 min. Cells were washed twice with ice-cold PBS, harvested by scraping in ice-cold PBS containing proteinase inhibitor cocktail (Roche, 05056489001), and pelleted at 600 × g for 7 min at 4 °C. Cells were incubated for 10 min on ice in buffer containing 10 mM EDTA, 10 mM Tris pH 8, 0.5 mM EGTA, and 0.25% Triton X-100. After centrifugation, cells were incubated in buffer containing 1 mM EDTA, 10 mM Tris pH 8, 0.5 mM EGTA, 200 mM NaCl, and passed 3 times through 26-G needles (BD, 303800), followed by incubation on ice for 10 min. Cells were centrifuged for 600 × g for 7 min at 4 °C. Chromatin was extracted in a high-salt buffer containing 50 mM HEPES, pH 7.5, 1 mM EDTA, 1% Triton X-100, 0.1% deoxycholate, 0.1% SDS, and 500 mM NaCl for 2 h at 4 °C. Chromatin was sheared using a Bioruptor Pico sonicator (Diagenode). Protein-G Dynabeads (Invitrogen, 10004D) were washed twice with TE buffer pH 8 for 5 min at room temperature and blocked with 1 mg BSA (Sigma–Aldrich), 100 ng yeast tRNA (R8508, Sigma) in TE buffer containing proteinase inhibitor cocktail. 100 µg of chromatin was pre-cleared with 20 μl of pre-blocked protein-G magnetic beads for 1 h at 4 °C. Then, 5% of cleared chromatin was stored at −20 °C as input until decrosslinked together with the IP material. For the IP, 1 μg of anti-phospho-Histone H2A.X (Ser139) Antibody, clone JBW301 (Sigma–Aldrich, 05636) or 7.5 μg anti-GFP (Abcam, ab290) was added to cleared chromatin and incubated overnight at 4 °C. The following day, 30 μl blocked protein-G beads were added to the chromatin and incubated for 3 h at 4 °C. Beads were washed twice with high-salt buffer, followed by another time with 50 mM LiCl, 0.5% NP-40, 0.5% deoxycholate, and twice with TE buffer at 4 °C. Next, chromatin was eluted in 1% SDS and 100 mM NaHCO3 containing 60 μg RNaseA (Roche, 10109169001) and incubated at 37 °C for 30 min. Elution was adjusted to 10 mM EDTA, 40 mM Tris pH 8.0, and 25 μg Proteinase K (Roche, 3115836001) and incubated for 3 h at 55 °C. Final decrosslinking was done overnight at 65 °C. Input chromatin was decrosslinked in parallel in TE buffer, pH 8, and 60 μg RNaseA was added and incubated at 37 °C for 30 min. Chromatin was adjusted to 1% SDS, 100 mM NaCl, and 60 μg Proteinase K and incubated for 3 h at 55 °C. Final decrosslinking was done overnight at 65 °C. Eluted material was cleaned up by phenol-chloroform extraction, and DNA was quantified using a Qubit 3.0 fluorometer. 10 ng of DNA was used for subsequent library preparation.
ChIP-seq library preparation and data analysis
For ChIP-seq, libraries were prepared using the NEBNext Ultra II DNA Library Prep Kit for Illumina (E7645) following the standard protocols. Samples with different index barcodes were combined at equal molar ratios, with the exception of input DNA (0.7× ratio), and sequenced as pools. Sequencing of library pools was performed on NextSeq2000 machines according to Illumina standards, with 50-bp paired-end sequencing. Library demultiplexing was performed following Illumina standards. The quality of each raw sequencing file (fastq) was verified with FastQC (v0.12.1) (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Samples were filtered for low-quality reads and adapter sequences using trim galore (v0.6.10). All files were aligned to the reference human genome assembly GRCh37 (hg19) using Bowtie2 (v2.5.1)54. Multiple-matching reads were eliminated using SAMtools (v1.20)55. Duplicated reads were removed by Picard (v3.1.1) (https://broadinstitute.github.io/picard/). DeepTools (v3.5.5) was used for read depth normalization and visualization56. Coverage tracks were generated by the BamCoverage module of deepTools after normalizing counts per million with a 200 bp bin size, read extension, and length smoothing to 400 bp. ChIP-seq coverage tracks were visualized using IGV (v2.18.4)57. Averaged ChIP-seq profiles were generated using the R package ggplot2. The x-axis represents genomic position relative to DSBs, and the y-axis represents the mean coverage at 1 kb bins, where data were smoothed using a 50 kb span. The in silico called AsiSI sites were identified genome-wide in the human reference genome (hg19) using the BSgenome.Hsapiens.UCSC.hg19 package in R. Genomic coordinates of the corresponding cut sites were extracted and exported in BED format for downstream analyses.
Cell viability assays
Cell viability assays were performed using the CellTiter-Glo Luminescent Cell Viability Assay (Promega, G7570), according to the manufacturer’s instructions. Briefly, 5000 mESCs or 7500 U2OS cells were seeded onto black-walled and flat-bottom 96-well plates (Greiner). For mESCs, plates were coated with 0.2% gelatine prior to seeding. Cells were incubated at different time points with different concentrations of DNA damage agents or UV-radiated and allowed to recover at 37 °C and 5% CO2. Cells treated with DMSO or non-UV-radiated cells were used as controls. Prior to measuring, the same volume of freshly prepared CellTiter-Glo reagent was added, and plates were incubated for 2 min on an orbital shaker, followed by 10 min incubation at RT. Plates were measured using the luminescence microplate reader CLARIOstar Plus (BMG Labtech). The raw data generated for control treatments, set to 100%, was used to estimate the percentage of viable cells.
Clonogenic assays
For colony survival assays, U2OS cells were seeded at a single-cell density onto 6-well plates overnight (1500 cells per well). The next day, cells were either UV irradiated and allowed to recover under normal growth conditions, changing the medium every 2 days, or treated with CPT or ETO for 24 h, washed three times with 37 °C U2OS medium, and allowed to recover. After 10 days, the medium was removed, and the cells were rinsed with PBS. Cells were then fixed with 2 mL of fixation solution (Methanol:Acetic acid 3:1) and incubated for 5 min at RT. After removing the fixation solution, colonies were stained with 0.5% crystal violet (VWR, 20.846.361) prepared in 20% methanol for 15 min. Plates were washed extensively with water and allowed to dry overnight. The number of individual colonies, as well as the signal intensity of the whole plate normalized to the background (region on plates devoid of colonies), was estimated with Fiji, and values were used to calculate plating efficiency and surviving fraction relative to non-UV-radiated/DMSO-treated cells. Clonogenic survival was estimated based on control plates (set to 100%).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The ChIP-seq data sets generated in this study have been deposited in NCBI GEO under the accession number GSE308329. Source data are provided with this paper.
References
Ciccia, A. & Elledge, S. J. The DNA damage response: making it safe to play with knives. Mol. Cell 40, 179–204 (2010).
Dantuma, N. P. & van Attikum, H. Spatiotemporal regulation of posttranslational modifications in the DNA damage response. EMBO J. 35, 6–23 (2016).
Kinner, A., Wu, W., Staudt, C. & Iliakis, G. γ-H2AX in recognition and signaling of DNA double-strand breaks in the context of chromatin. Nucleic Acids Res. 36, 5678–5694 (2008).
Paull, T. T. et al. A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr. Biol. 10, 886–895 (2000).
Rajan, M. et al. Generation of an alpaca-derived nanobody recognizing γ-H2AX. FEBS Open Bio 5, 779–788 (2015).
Moeglin, E. et al. A novel nanobody precisely visualizes phosphorylated histone H2AX in living cancer cells under drug-induced replication stress. Cancers 13, 3317 (2021).
Trakarnphornsombat, W. & Kimura, H. Live-cell tracking of γ-H2AX kinetics reveals the distinct modes of ATM and DNA-PK in the immediate response to DNA damage. J. Cell Sci. 136, jcs260698 (2023).
Villaseñor, R. et al. ChromID identifies the protein interactome at chromatin marks. Nat. Biotechnol. 38, 728–736 (2020).
Mesquita, R. D., Woods, N. T., Seabra-Junior, E. S. & Monteiro, A. N. A. Tandem BRCT domains. Genes Cancer 1, 1140–1146 (2010).
Manke, I. A., Lowery, D. M., Nguyen, A. & Yaffe, M. B. BRCT repeats as phosphopeptide-binding modules involved in protein targeting. Science 302, 636–639 (2003).
Woods, N. T. et al. Charting the landscape of tandem BRCT domain-mediated protein interactions. Sci. Signal. 5, rs6 (2012).
Lou, Z. et al. MDC1 maintains genomic stability by participating in the amplification of ATM-dependent DNA damage signals. Mol. Cell 21, 187–200 (2006).
Jeon, G. S. et al. Deregulation of BRCA1 leads to impaired spatiotemporal dynamics of γ-H2AX and DNA damage responses in Huntington’s disease. Mol. Neurobiol. 45, 550–563 (2012).
Jowsey, P. A., Doherty, A. J. & Rouse, J. Human PTIP facilitates ATM-mediated activation of p53 and promotes cellular resistance to ionizing radiation *. J. Biol. Chem. 279, 55562–55569 (2004).
Ward, I. M., Minn, K., Jorda, K. G. & Chen, J. Accumulation of checkpoint protein 53BP1 at DNA breaks involves its binding to phosphorylated histone H2AX *. J. Biol. Chem. 278, 19579–19582 (2003).
Wood, J. L., Singh, N., Mer, G. & Chen, J. MCPH1 Functions in an H2AX-dependent but MDC1-independent Pathway in Response to DNA Damage. J. Biol. Chem. 282, 35416–35423 (2007).
Grawunder, U., Zimmer, D. & Lieber, M. R. DNA ligase IV binds to XRCC4 via a motif located between rather than within its BRCT domains. Curr. Biol. 8, 873–879 (1998).
Wang, J., Gong, Z. & Chen, J. MDC1 collaborates with TopBP1 in DNA replication checkpoint control. J. Cell Biol. 193, 267–273 (2011).
Lee, M. S., Edwards, R. A., Thede, G. L. & Glover, J. N. M. Structure of the BRCT repeat domain of MDC1 and its specificity for the free COOH-terminal end of the γ-H2AX histone tail. J. Biol. Chem. 280, 32053–32056 (2005).
Stucki, M. et al. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell 123, 1213–1226 (2005).
Clapperton, J. A. et al. Structure and mechanism of BRCA1 BRCT domain recognition of phosphorylated BACH1 with implications for cancer. Nat. Struct. Mol. Biol. 11, 512–518 (2004).
Li, M. & Yu, X. Function of BRCA1 in the DNA damage response is mediated by ADP-ribosylation. Cancer Cell 23, 693–704 (2013).
Vergara, X. et al. Megabase-scale detachment of genome-nuclear lamina interactions is required for efficient repair of double stranded breaks. bioRxiv Preprint at https://doi.org/10.1101/2025.07.03.662902 (2025).
Williams, R. S., Lee, M. S., Hau, D. D. & Glover, J. N. M. Structural basis of phosphopeptide recognition by the BRCT domain of BRCA1. Nat. Struct. Mol. Biol. 11, 519–525 (2004).
Stewart, G. S., Wang, B., Bignell, C. R., Taylor, A. M. R. & Elledge, S. J. MDC1 is a mediator of the mammalian DNA damage checkpoint. Nature 421, 961–966 (2003).
Tichy, E. D. et al. Mouse embryonic stem cells, but not somatic cells, predominantly use homologous recombination to repair double-strand DNA breaks. Stem Cells Dev. 19, 1699–1711 (2010).
van den Berg, J. et al. A limited number of double-strand DNA breaks is sufficient to delay cell cycle progression. Nucleic Acids Res. 46, 10132–10144 (2018).
Peters, A. H. F. M. et al. Partitioning and plasticity of repressive histone methylation states in mammalian chromatin. Mol. Cell 12, 1577–1589 (2003).
Tsouroula, K. et al. Temporal and spatial uncoupling of DNA double strand break repair pathways within mammalian heterochromatin. Mol. Cell 63, 293–305 (2016).
Iacovoni, J. S. et al. High-resolution profiling of γH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 29, 1446–1457 (2010).
Clouaire, T. et al. Comprehensive mapping of histone modifications at DNA double-strand breaks deciphers repair pathway chromatin signatures. Mol. Cell 72, 250–262.e6 (2018).
Kilic, S. et al. Phase separation of 53BP1 determines liquid-like behavior of DNA repair compartments. EMBO J. 38, e101379 (2019).
Dimitrova, N., Chen, Y.-C. M., Spector, D. L. & de Lange, T. 53BP1 promotes non-homologous end joining of telomeres by increasing chromatin mobility. Nature 456, 524–528 (2008).
Sun, L. et al. Structural insight into BLM recognition by TopBP1. Structure 25, 1582–1588.e3 (2017).
Shao, Z. et al. Specific recognition of phosphorylated tail of H2AX by the tandem BRCT domains of MCPH1 revealed by complex structure. J. Struct. Biol. 177, 459–468 (2012).
Osakabe, A. & Molaro, A. Histone renegades: unusual H2A histone variants in plants and animals. Semin. Cell Dev. Biol. 135, 35–42 (2023).
Leung, C. C. Y. & Glover, J. M. BRCT domains. Cell Cycle 10, 2461–2470 (2011).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Lemoine, F. et al. Renewing Felsenstein’s phylogenetic bootstrap in the era of big data. Nature 556, 452–456 (2018).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Ghanta, K. S., Ishidate, T. & Mello, C. C. Microinjection for precision genome editing in Caenorhabditis elegans. STAR Protoc. 2, 100748 (2021).
Fraser, A. G. et al. Functional genomic analysis of C. elegans chromosome I by systematic RNA interference. Nature 408, 325–330 (2000).
Baubec, T., Ivánek, R., Lienert, F. & Schübeler, D. Methylation-dependent and -independent genomic targeting principles of the MBD protein family. Cell 153, 480–492 (2013).
Vader, G., Maia, A. F. & Lens, S. M. The chromosomal passenger complex and the spindle assembly checkpoint: kinetochore-microtubule error correction and beyond. Cell Div. 3, 10 (2008).
Manzo, M. et al. Isoform-specific localization of DNMT3A regulates DNA methylation fidelity at bivalent CpG islands. EMBO J. 36, 3421–3434 (2017).
Govindaraj, K., Hendriks, J., Lidke, D. S., Karperien, M. & Post, J. N. Changes in fluorescence recovery after photobleaching (FRAP) as an indicator of SOX9 transcription factor activity. Biochim. Biophys. Acta Gene Regul. Mech. 1862, 107–117 (2019).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinform. 22, 433 (2021).
Manders, E. M. M., Verbeek, F. J. & Aten, J. A. Measurement of co-localization of objects in dual-colour confocal images. J. Microsc. 169, 375–382 (1993).
Costes, S. V. et al. Automatic and quantitative measurement of protein-protein colocalization in live cells. Biophys. J. 86, 3993–4003 (2004).
Poulet, A. et al. NucleusJ: an ImageJ plugin for quantifying 3D images of interphase nuclei. Bioinformatics 31, 1144–1146 (2015).
Toraason, E., Adler, V. L. & Libuda, D. E. Aging and sperm signals alter DNA break formation and repair in the C. elegans germline. PLoS Genet. 18, e1010282 (2022).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Acknowledgements
We thank members of the Baubec laboratory for their input and criticism. Furthermore, we thank the Biology Imaging Center at Utrecht University, the Utrecht University FACS facilities at the Science and Veterinary Faculties, and the Cell Microscopy Core at UMC Utrecht for their support. We thank Dr. Susanne MA Lens (University Medical Center Utrecht, Netherlands) for kindly providing the Human U2OS cells. We also thank Dr. Gaëlle Legube (MCD, Centre de Biologie Intégrative (CBI), CNRS, Université de Toulouse, Toulouse, France) for providing the DIvA-AsiSI-ER-U2OS cell line. The RPE-1 iCut cell line (RPE-1) and H2AXKO RPE-1 cell lines were kindly provided by Dr. Bas van Steensel (Division of Gene Regulation, Netherlands Cancer Institute, Amsterdam, the Netherlands) and Dr. René H. Medema (Division of Cell Biology, Netherlands Cancer Institute, Amsterdam, The Netherlands). This work was supported by the European Research Council (865094 - ChromatinLEGO - ERC-2019-COG), the Dutch Research Council (NWO-Vici, VI.C.222.012), Utrecht University, and the EMBO Young Investigator program.
Author information
Authors and Affiliations
Contributions
R.C.S. and T.B. conceived and designed the study. K.E., D.C.R., V.P., and D.tB. developed tools and protocols and performed experiments. N.K., and N.P.V.dW. performed key technical experiments under direction. R.C.S., K.E., and T.B. analyzed data and interpreted results. X.V. and A.O. provided key reagents. V.P. performed experiments on C. elegans under the supervision of S.v.d.H. R.C.S., and T.B. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Robin Sebastian and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
da Silva, R.C., Eleftheriou, K., Recchia, D.C. et al. Engineered chromatin readers track damaged chromatin dynamics in live cells and animals. Nat Commun 16, 10127 (2025). https://doi.org/10.1038/s41467-025-65706-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65706-y