Abstract
Single-cell proteomics confidently quantifies cellular heterogeneity, however quantification of post-translational modifications, such as those deposited on histone proteins, remains elusive. Here, we develop a robust mass spectrometry-based method for the unbiased analysis of single-cell histone post-translational modifications (sc-hPTM). sc-hPTM identifies both single- and combinatorial histone post-translational modifications (67 peptidoforms in total), which includes nearly all frequently studied histone post-translational modifications with comparable reproducibility to traditional bulk experiments. As a proof of concept, we treat cells with sodium butyrate, a histone deacetylase inhibitor, and demonstrate that our method can i) distinguish between treated and untreated cells, ii) identify sub-populations of cells with heterogeneous response to the treatment, and iii) reveal differential co-regulation of histone post-translational modifications in the context of drug treatment. The sc-hPTM method enables comprehensive investigation of chromatin heterogeneity at single-cell resolution and provides a further understanding of the histone code.
Similar content being viewed by others
Introduction
Epigenetic states are imprinted through chemical modifications to DNA and associated histone proteins that reflect the intricate relationship between the epigenome and cellular phenotypes1. The ‘histone code’ refers to the complex interplay between diverse post-translational modifications (PTMs) of histones2. The most prevalent histone PTMs (hPTMs) encompass acetylation (Ac), methylation (Me), ubiquitination (Ub), and phosphorylation (Ph). These hPTMs influence gene expression by modulating chromatin structure through their distinct chemical properties and their capacity to attract chromatin-modifying enzymes and binding proteins. For instance, histone 4 lysine 16 acetylation (H4K16ac) contributes to chromatin decondensation and thereby increases gene transcription3. In contrast, trimethylation of histone 3 lysine 27 (H3K27me3) is generally associated with transcriptional repression and has been implicated in epigenetic reprogramming4,5. Notably, loss of H4K16ac and H4K20me3 are common hallmarks of cancer, while several other hPTMs have been linked to neurodegenerative, autoimmune, and other diseases6,7,8,9.
Bulk analysis of hPTMs, in which histones are extracted from cells at high purity, has markedly improved our understanding of their pivotal role in health and disease5,10,11,12,13. While bulk methods often require substantial input material (typically around 1 million cells), recent advancements have enabled hPTM profiling by MS from as few as 1000 cells, including patient-derived samples14,15. However, established single-cell profiling methods, such as single-cell RNA-sequencing, have revealed that conventional analyses of bulk injections often mask the presence of different cell types, as well as the variability among cells within the same type. Thus, single-cell analysis has emerged as a powerful tool for uncovering critical insights across diverse biomedical fields. For example, it has revealed fibroblast clusters linked to cancer immunotherapy resistance, identified stem-cell programs in metastatic breast cancer, highlighted transcriptomic reprogramming in aging cardiovascular endothelial cells, and shown increased epigenetic variation with age16,17,18,19. These findings provide unprecedented resolution in understanding complex biological processes and disease mechanisms.
While several methods have been introduced to analyze hPTMs from single cells, such as single-cell chromatin immunoprecipitation sequencing, these have had limited use in the field due to sample throughput, data sparsity, technical noise, and lack of hPTM multiplexing20,21,22,23. Recently, Cheung et al. developed EpiTOF for simultaneous profiling of 60 epigenetic targets in single human immune cells using heavy metal isotope-labeled antibodies and mass cytometry by time-of-flight19. Interestingly, this method was able to provide insights into age-related changes of chromatin by demonstrating an increase in hPTM cell-to-cell variability during immune cell aging. However, a fundamental limitation of the aforementioned approaches is that they rely on antibodies, which suffer from (i) cross-reactivity, (ii) steric hindrance when targeting combinatorial hPTMs, (iii) limited number of targets, (iv) quantitative ambiguity resulting from differences in affinity, (v) lack of multiplexing, and (vi) limited detection of novel modifications24,25,26. In contrast, liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS) provides a comprehensive view of the histone code through the untargeted and quantitative measurement of a multitude of hPTMs27,28,29,30,31,32,33. Recent advances in MS instrumentation have greatly improved sensitivity, quantification accuracy, and throughput, making LC-MS/MS the gold standard for global single-cell proteomics34,35,36,37,38,39. Although histones are commonly identified in such workflows due to their abundance within the cell, a robust method for accurate quantification of hPTMs at single-cell resolution is currently lacking.
Here, we present a method that enables robust identification and quantification of single-cell histone post-translational modifications (sc-hPTM) across the canonical histones H1, H2A, H2B, H3, and H4. We combined single-cell dispensing and semi-automated sample preparation using a cellenONE dispensing platform coupled with a Bruker timsTOF Ultra mass spectrometer. Our sc-hPTM workflow identified 67 histone peptidoforms (i.e., distinct combinations of modified and unmodified amino acids per histone peptide), comprising 25 unique PTMs, per single-cell. We demonstrated high quantification accuracy and precision of our single-cell data through comparisons with bulk samples, previous data, and titration curves. Importantly, we propose a strategy to distinguish technical noise from biological noise (cell-to-cell variation) using histone standards. Finally, we treated cells with a histone deacetylase (HDAC) inhibitor to perturb hPTMs and identified cellular heterogeneity and differential hPTM co-regulation in response to the treatment.
In this work, we detail the development, validation, and application of sc-hPTM, demonstrating its robustness and utility in advancing our understanding of the histone code. This sc-hPTM method represents a significant leap forward in single-cell epigenomics, complementing existing single-cell genomics, transcriptomics, and proteomics approaches. By enabling unbiased, multiplexed quantification of hPTMs at single-cell resolution, sc-hPTM fills a critical gap in our ability to study chromatin states in heterogeneous cell populations. This method has the potential to uncover insights into epigenetic regulation in various biological processes, including cellular differentiation, disease progression, and response to therapeutic interventions.
Results
Identification of hPTMs at single-cell resolution using LC-MS/MS
To analyze hPTMs with bulk methods, histones are extracted from millions of cells using sulfuric acid, followed by trichloroacetic acid precipitation10,11,31,40. Due to the high content of lysine residues in histone proteins, extracted histones are subsequently derivatized with propionic anhydride at the protein and peptide level to modify N-termini and free lysine residues. Propionylation is critical to hPTM analysis by LC-MS/MS as it ensures a sufficient size and hydrophobicity of histone peptides required for LC-MS/MS analysis. Recently, protocols have demonstrated that these steps can be automated using liquid handling, which allowed for increased throughput when profiling hPTMs of chemical perturbations across multiple biological backgrounds33. Our sc-hPTM protocol builds upon previous single-cell proteomics protocols by utilizing the cellenONE (Cellenion), a robotic picolitre dispensing platform, to automate sample preparation, which includes single-cell isolation, cell lysis, propionylation, and digestion35,36,41,42.
The experimental workflow, depicted in Fig. 1A, illustrates the direct isolation of single cells to a 384-well plate (Supplementary Fig. 1A–D) pre-filled with 1 µL cell lysis buffer, consisting of 0.2% (w/v) n-Dodecyl β-D-maltoside in 1 M Triethylammonium bicarbonate (TEAB)43. We implemented the preparation in 384-well plates due to their reduced well diameter and depth relative to 96-well plates to lower the reaction volume and reduce protein loss. After each step, the plate was sealed with adhesive aluminum foil, vortexed for 5 s, and centrifuged at 500×g for 30 s to ensure thorough mixing. To minimize evaporation or incomplete sealing of the plate, the outermost wells were excluded from sample processing. Following cell lysis, histone proteins were propionylated by dispensing 100 nL of a 1:4 propionic anhydride to acetonitrile solution, followed by incubation for 30 min at room temperature (Supplementary Fig. 1E–G). Of note, dispensing of 100 nL of a low-viscosity derivatization reagent with the cellenONE required fine-tuning of the piezo dispensing capillary nozzle parameters to obtain a stable droplet (Supplementary Fig. 1E; see “Methods” section). Next, the plate seal was removed and the temperature of the cellenONE plate holder was increased to 50 °C, ensuring complete evaporation of the propionylation reagents, which removes any chemical interference during the next digestion step. Following this, 1 µL digestion buffer was added, consisting of 3 ng/µL trypsin, 10 U/µL Benzonase, and 1% ProteaseMAX in 1 M TEAB. The plate was then sealed and incubated for 2 h at 50 °C within the cellenONE for histone digestion. The free amino termini of the digested peptides were subsequently propionylated as described above to improve their chromatographic retention. Finally, injections were manually diluted with 0.1% TFA and stored at -80 °C prior to LC-MS/MS analysis. To serve as a reference and assess the accuracy of our workflow, we also prepared injections consisting of ~100 cells (100-cell bulk), which were processed identically to the single cells.
A Schematic overview of the method, from left-to-right: (1) suspensions of single cells or histone standards were isolated and prepared within the wells of a 384-well plate using the liquid handling capabilities of the cellenONE instrument, (2) histone peptidoforms were acquired using the Vanquish LC and TimsTof Ultra MS instruments, and (3) data analysis was performed with Skyline and custom R scripts. For details on cellenONE cell isolation and preparation, see Supplementary Fig. 1. Created in BioRender. Cutler, R. (2025) https://BioRender.com/hvhvfvu. B Extracted ion chromatograms (XICs) of MS1 precursor (top panels) and MS2 fragments (bottom panels) for four representative histone peptidoforms in a representative single-cell. Peaks are picked based on the criteria shown in the box on the right side. For additional XICs of isobaric peptides, see Supplementary Fig. 2A. Source data are provided as a Source Data file. C Overview of the histone peptidoforms identified by the sc-hPTM method. Detected peptide sequences are highlighted. PTMs are represented by symbols shown in the legend on the bottom-right. Single PTMs are shown on top of sequences, while combinatorial PTMs are shown by a connecting line underneath the sequence. Note that specific isoforms of histone H1, H1.2/3/4, and H1.5 are not shown. For proportions of each modification type, see Supplementary Fig. 3A. Source data are provided as a Source Data file.
The resulting MS data were analyzed in Skyline44 using a spectral library derived from 100-cell bulk injections, with manual adjustment of peak boundaries based on predetermined requirements (Fig. 1B and Supplementary Data 1). For confident peptide identification, we used a combination of filtering criteria: (1) clear distinction of peptide signal vs measurement noise in 0-cell injections, (2) MS1 and MS2 signal alignment, (3) presence of the C13 isotopes, (4) retention time difference relative to the unmodified peptide, (5) the presence of unique fragment ions, (6) alignment with 100-cell bulk and (7) alignment to histone standards (Fig. 1B, right box). Isobaric peptidoforms of the H4 N-terminal peptide H4(4-17) (Supplementary Fig. 2A) were distinguished and separately quantified based on unique combinations of their fragment ions (Supplementary Fig. 2B).
Following all these criteria, we identified peptides from H1, H2A, H2B, H3, and H4, as well as their variants H1.2/3/4, H1.5, H2A1B, H2AJ, and H2AX (Supplementary Data 2). We were able to identify peptidoforms that were acetylated (Ac), methylated (Me), dimethylated (Me2), trimethylated (Me3), ubiquitinylated (Ub), succinylated (Su), and hydroxyisobutyrylated (Hib) (Supplementary Fig. 3A). These were present either individually or in combination with other PTMs, resulting in 67 distinct peptidoforms. Notably, we achieved comprehensive coverage of the N-terminal tails of H3 and H4, in particular, of peptides H3(9–17) and H4(4–17) that were identified with multiple peptidoforms (Fig. 1C). Missing regions included the H2B N-terminus and H1 C-terminus, which lack arginine residues, resulting in long hydrophobic peptides. Identification of these peptides via LC-MS/MS is challenging due to their poor chromatographic behavior and the presence of several isobaric peptidoforms. Additionally, some well-studied marks, such as H3K4me2/me3 and H4K20me2/me3, were undetected at the single-cell level and only at the bulk level, presumably due to their relatively low abundance45.
Benchmarking of sc-hPTM quantification
To benchmark the quantification accuracy of our method, we prepared 3 batches of HeLa cells and histone standards on separate days (Supplementary Table 1), totaling 60 injections with varying amounts of cells as input (Supplementary Fig. 3B). A batch is defined as a single 384-well plate with blank, single-cell, and histone standard samples that were prepared and injected for LC-MS/MS analysis together. We consistently detected 67 histone peptidoforms across all injections with <0.1% missing values, highlighting the strength of the data-independent acquisition (DIA) method. MS1 intensities in our measurements were on average over 3× greater than MS2 intensities and generally suffered less from distortion at lower signal intensities (Supplementary Fig. 1C). We therefore quantified histone peptide abundances at the MS1 level and normalized the raw intensities using the histone peptide ratio method to obtain relative abundances, as is done for MS-based histone analysis with the EpiProfile 2.0 program46. This included resolving co-eluting isobaric acetylation patterns of the H4(4–17) histone peptide, where the MS1 peak area is deconvolved using a combination of MS2 fragment ion intensities specific to the modification (Supplementary Fig. 2A, B; see “Methods” section).
Next, to distinguish between a positive peptidoform signal and measurement noise, we compared the raw intensities of injections containing single cells to those containing no cells, which were injections that contained only reagents (i.e., negative controls). Despite the expectedly low signal from single cells, we found that the MS1 signal intensities in the single-cell injections were on average ~9.5× above the injections containing no cells (Fig. 2A and Supplementary Fig. 3D). Peptides from single cells that had similar MS1 signal intensities in injections with no cells were considered as background noise and excluded in subsequent analyses (Fig. 2B, Supplementary Fig. 3E, and Supplementary Data 2; see “Methods” section).
A Comparison of the raw MS1 intensity of empty wells (Cell #0) and wells containing single cells (Cell #1) for 4 representative histone peptidoforms for each batch. Each batch represents an independent experiment. For all peptidoforms see Supplementary Fig. 3D. Statistics: for the number of injections analyzed in each group per batch see Supplementary Fig. 3B. Boxplots show the median (center), the 25th and 75th percentiles (bounds), and whiskers extending to the most extreme data points within 1.5× IQR. B Distribution of MS1 intensities for all histone peptidoforms in all batches for empty wells and wells containing single cells after filtering out peptides with low signal (see “Methods” section). For each batch separately, see Supplementary Fig. 3E. C Titration curves for four representative histone peptidoforms of raw MS1 intensity, where various amounts of cells were prepared in a single well and then assessed for linearity. Statistics: the significance of Pearson R correlation coefficients for each batch was tested using a two-sided t-test. For H2AK15Ac, the correlations were R = 0.98 (p < 2.2 × 10⁻¹⁶), R = 0.99 (p = 1.2 × 10⁻¹¹), and R = 0.99 (p = 4.3 × 10⁻¹¹) for batches 1–3, respectively. For H3K23Ac, R = 0.98 (p < 2.2 × 10⁻¹⁶), R = 0.98 (p = 2 × 10⁻¹⁰), and R = 0.98 (p = 9.5 × 10⁻¹⁰). For H3K9Me2, R = 0.98 (p < 2.2 × 10⁻¹⁶), R = 0.99 (p = 6.6 × 10⁻¹²), and R = 0.99 (p = 2.5 × 10⁻¹¹). For H4K5AcK12Ac, the correlations were R = 0.92 (p = 4.8 × 10⁻¹³), R = 0.97 (p = 6.7 × 10⁻⁹), and R = 0.94 (p = 4.2 × 10⁻⁷) for batches 1–3, respectively. Shaded areas represent 95% confidence intervals of the fitted linear model. For all peptides, see Supplementary Fig. 4A. D Distribution of Pearson R correlation coefficients for titration curves of all histone peptidoforms in each batch. A dashed line is drawn at R = 0.8 to indicate the threshold used for filtering out histone peptidoforms deemed to be non-quantifiable. Statistics: the significance of Pearson R correlation coefficients for each batch was tested using a t-test. E Correlation of histone peptidoform relative abundance between bulk (~100-cell) and ‘pseudobulk’ (mean of single cells) samples in batches 1–3. Statistics: significance of Pearson R correlation coefficients for each batch was tested using a t-test; R = 0.95 (p < 2.2 × 10⁻¹⁶) for batch 1, R = 0.90 (p < 2.2 × 10⁻¹⁶) for batch 2, and R = 0.97 (p < 2.2 × 10⁻¹⁶) for batch 3. Shaded areas represent 95% confidence intervals of the fitted linear model. For correlation using unnormalized MS1 intensity, see Supplementary Fig. 7A. F Principal component analysis of histone peptidoform relative abundances for single-cell and 100-cell bulk samples. For plot without batch-correction see Supplementary Fig. 6C. For direct comparison of single-cell and bulk samples see Supplementary Fig. 7B, C. G Individual histone PTMs (single and combinatorial PTMs) relative abundances detected on histone H3 in single cells (top) and 100-cell bulk (bottom) samples from batches 1–3. For comparison to previous bulk studies, see Supplementary Figs. 7D, E. Statistics: for the number of injections analyzed in each group per batch, see Supplementary Fig. 3B. Data are presented as mean values, and error bars are ±1 SD. Source data are provided as a Source Data file.
To benchmark the quantification accuracy of our method, we assessed the linearity of signal intensities using our sc-hPTM workflow by preparing a 5-point titration curve of varying cell inputs (i.e. 0, 1, 2, 4, and 8 cells) and a commercially available bovine histone standard from calf thymus (i.e. 0, 10, 20, 40, and 80 pg; 1 cell contains ~6.5 pg of histone protein47). As expected, the MS1 intensity was proportional to the number of cells across most peptidoforms and yielded consistently high Pearson R values, which demonstrated high quantification accuracy of the method (Fig. 2C, D and Supplementary Fig. 4A, B). Similarly, the titration of the bovine histone standards revealed comparable quantification (Supplementary Fig. 5A–D). Peptidoforms with an average R value of less than 0.8 were therefore excluded from subsequent analysis to ensure accurate quantification (see “Methods” section). As accurate quantification can be influenced by ‘under’ and ‘over’ propionylation of evaluated histone peptides, we assessed the efficiency of peptide derivatization using our sc-hPTM workflow. Levels of ‘under’ and ‘over’ propionylation were extrapolated based on the representative H3.1(9–17) peptide, which consistently showed the highest signal in single cells (see “Methods” section). We found between ~5–30% of the H3.1(9–17) peptide was under-propionylated, which differed between batches, while ~40% consistently showed over-propionylation across batches. Based on this, cells with abnormal levels of under-/over-propionylation percentages were excluded from downstream analyses (Supplementary Fig. 2F; see “Methods” section).
To further assess the accuracy of hPTM quantification in single cells, we compared the 100-cell bulk to single-cell injections. After the aforementioned quality control filtering, normalization, and batch-correction (Supplementary Fig. 6A–D; see “Methods” section), we quantified 44 peptidoforms across 4 100-cell bulk and 16 single-cell injections. Data were normalized using the peptide ratio method as previously described in ref. 46, followed by a Log(1 + x) transformation. The peptidoform relative abundances were averaged across all single cells within each batch to create a cumulative ‘pseudobulk’ sample. We observed good correlation (R = 0.94) of the ‘pseudobulk’ peptidoform relative abundances to the average of the bulk injections within each batch (Fig. 2E; see “Methods” section). Similar results were also obtained without normalization (average R value of 0.89; Supplementary Fig. 7A). The high correlation between the single-cell pseudobulk and 100-cell bulk injections indicated accurate quantification, while the variation between samples reflected the single-cell precision of our data.
We then performed principal component analysis (PCA) on the relative abundances of all peptidoforms. The single-cell and 100-cell bulk injections separated along PC1 and accounted for 33.88% of the variance in the data (Fig. 2F). We confirmed that this difference was not due to large systematic differences as the average difference of hPTM relative abundances between single-cell and the mean of 100-cell bulk injections was only 1.28 × 10−3 Log2-fold (Supplementary Fig. 7B). Nonetheless, we noted small, but consistent, differences including higher relative abundance of H3K9me2 and lower relative abundance of H3K9meK14ac in single cells as compared to 100-cell bulk injections (Supplementary Fig. 7C). The signal of peptidoforms reflecting such differential abundance was above background levels (Supplementary Fig. 3D) with high quantification accuracy (Supplementary Fig. 4A). These observations therefore suggested that the sc-hPTM method offers greater accuracy than bulk injections with larger input, which may be due to less manual sample handling or fewer enrichment and clean-up steps prior to acquisition. Finally, we compared the relative abundances of individual PTMs on histone H3 between single cells, 100-cell bulk injections, and bulk HeLa cell injections from previous studies, and found high correspondence between the single-cell and bulk data (Fig. 2G and Supplementary Figs. 7D, E). Collectively, we validated the accuracy of our sc-hPTM quantification against traditional bulk methods.
Biological cell-to-cell variability of hPTMs
To distinguish between technical and biological variability in single-cell injections, we leveraged the technical replicates that were acquired, defined as separate injections of the same amounts of bovine histone standard. After quality control filtering, normalization, and batch-correction of the histone standard injections, we determined the coefficient of variation (CV) of the relative abundances of peptidoforms for the injections prepared within a batch (intra-) or across batches (inter-). For the 10 pg histone standard injections (roughly equivalent to the amount of histone in a single-cell48), both intra- and inter-CVs were below 25% on average (Fig. 3A, B). For data that were unnormalized and not batch-corrected, intra-CVs were still below 25%, while inter-CVs were approximately double, which highlighted the impact of normalization and batch-correction (Supplementary Fig. 8A, B). To further assess the technical variability of our method, we compared the 10 pg histone standard technical replicate injections to technical replicates of bulk HeLa cells49,50 and processing replicates of cell lines treated with DMSO in the NIH Library of Integrated Network-Based Cellular Signatures consortium (LINCS)33 (Supplementary Table 2). We found that our across batch CVs were comparable to the CVs of the bulk HeLa data49,50 (Fig. 3C, left and middle panels) and the cell lines from the LINCS dataset33 (Fig. 3C, right panel). Overall, this analysis established a baseline for the technical variation expected from single-cell injection input amounts both within batches (intra-batch) and across batches (inter-batch), which opens up the opportunity to study cell-to-cell variation.
A, B CV of histone peptidoform relative abundances for histone standard technical replicates within each batch (intra-) (A) and across the batches 1–3 (inter-) (B). For results using raw MS1 intensities see Supplementary Fig. 8A, B. Statistics: for number of injections analyzed in each group per batch see Supplementary Fig. 5B. Boxplots show the median (center), the 25th and 75th percentiles (bounds), and whiskers extending to the most extreme data points within 1.5× IQR; points beyond whiskers are plotted individually as outliers. C CV of histone peptidoform relative abundances for 10 pg histone standard technical replicates across batches 1–3 compared to bulk technical replicates of HeLa cells (Sidoli et al. A49; Sidoli et al. B50) and processing replicates of cell lines treated with DMSO in the NIH LINCS (Dele-Oni et al.33). Histone peptidoforms were filtered to match those quantified in each respective study (Supplementary Table 2). Statistics: histone standard 10 pg (all panels) n = 12, Sidoli et al. A49 Hela n = 5, Sidoli et al. B49 Hela n = 4, Dele-Oni et al. (LINCS)33 n = 3; boxplots show the median (center), the 25th and 75th percentiles (bounds), and whiskers extending to the most extreme data points within 1.5× IQR; points beyond whiskers are plotted individually as outliers. D CV of histone peptidoform relative abundances for 10 pg histone standard compared to single cells across batches 1-3 and matched by 39 peptidoforms. Statistics: histone standard 10 pg n = 12, single cells n = 22; two-sided paired Wilcoxon test, ****p ≤ 0.0001; boxplots show the median (center), the 25th and 75th percentiles (bounds), and whiskers extending to the most extreme data points within 1.5× IQR. E, F CV of histone peptidoform relative abundances ranked from highest to lowest amongst single-cells from batches 1–3, shown with technical noise (E) and after removal of technical noise (i.e., cell-to-cell variation) (F). Source data are provided as a Source Data file.
An important capability for single-cell analysis is the quantification of subtle cell-to-cell biological variation that can reveal cell states and regulatory relationships inaccessible by bulk analyses. Indeed, the CVs of the majority of peptidoforms across single cells using sc-hPTM were significantly higher than those amongst the technical replicates (Fig. 3D). To determine biological cell-to-cell variation for individual peptidoforms, we adjusted for technical noise (determined by the 10 pg histone standard injections) and ranked peptidoforms by their CV (Fig. 3E, F and Supplementary Fig. 8C; see “Methods” section). This analysis revealed H4K5acK12ac and H3K14ac as the most biologically variable peptidoforms in our cell model, perhaps due to variation in cell cycle phase51, whereas H2AJK5acK9ac and H3K9me3K14ac were found to be the least variable.
Distinct single-cell hPTM profiles following HDAC inhibitor treatment
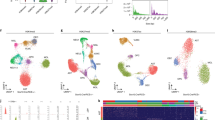
To determine if our sc-hPTM method can detect the change in abundance of biologically relevant hPTMs due to chemical perturbation, we treated HeLa cells with 5 mM HDAC inhibitor sodium butyrate (NaBut). NaBut competitively binds to the zinc sites of class I and II HDACs, resulting in hyperacetylation of several histone sites, particularly those located on histones H3 and H4. After a 24hour incubation, both control (PBS vehicle) and NaBut-treated cells were harvested and processed using our sc-hPTM workflow in 2 batches on separate days (Fig. 1A). Single cells were isolated from three different cell populations, control, treated, and a 1:1 mixture of control and treated (mixed) (Fig. 4A and Supplementary Table 3). We included a mixed population of cells to evaluate if these cells can be classified based on their hPTM profiles without prior knowledge of whether they originated from the control or treated groups. As in previous experiments, we also included 100-cell bulk injections for the control and treated groups. Following our quality control filtering, normalization, and batch-correction pipeline (Supplementary Fig. 9A–G; see “Methods” section), we were able to quantify 42 peptidoforms (Supplementary Data 2) in 5 controls, 11 treated, and 32 mixed single cells. Due to the low number of control cells that passed filtering criteria in this experiment, we further included 16 single HeLa cells and 4 100-cell bulk injections from the benchmarking experiments (as described above) in the analysis. It should be noted that these additional control cells did not receive any treatment and showed no substantial differences from the control cells in the HDACi experiment (Supplementary Fig. 9H). PCA of all analyzed injections revealed clear separation between control and treated cells along PC1, accounting for 35.46% of the variation, which was significantly correlated to the NaBut treatment variable (two-sided t-test, R = 0.53, p < 0.001). Mixed cells appeared to be uniformly distributed across PC1 and the two treatment groups, as expected (Fig. 4B). HDACi treatment did not impact global protein identification and quantification (Supplementary Fig. 10A-B). Importantly, global protein abundance did not correlate with hPTM quantities per single cell, which indicated that the treatment-specific clustering was driven by the epigenetic marks (Supplementary Fig. 10C).
A Schematic overview of experimental design. After HDAC inhibitor treatment, control and treated cell suspensions are mixed 1:1 to create the ‘mixed’ group, where the label of each cell was unknown. For plate layout, see Supplementary Table 2. Created in BioRender. Cutler, R. (2025) https://BioRender.com/hit8jl5. B PCA of histone peptidoform relative abundances for single-cell injections. Note that samples from benchmarking experiments were included to increase the number of control cells. For details, see Supplementary Fig. 9G. C PCA of single-cell injections as in (B), but using the summarized relative abundances of H4 peptide acetylation (i.e., mono, di, tri, and tetra) instead of individual hPTMs. These summarized relative abundances were used for classifying the mixed cells into control and treated groups as shown in (D). D Results of mixed cell classifcation shown as a heatmap with hierarchical clustering of scaled H4 peptide acetylation summarized relative abundances (Z-score). Cells are shown in columns, and summarized H4 PTMs are shown in rows. A description of the annotation rows at the top of the heatmap starting from the 1st row and moving down to the 4th row is as follows: 1st row: batch each cell was prepared in; 2nd row: labels of cells that are known used to train and test the k-nearest neighbors (KNN) classifier, as well as validate the accuracy of the predictions; 3rd row: random assignment of cells with known group (control or treated) to training (50%) and test (50%) groups to assess KNN classification accuracy; 4th row: predicted labels of mixed cells using KNN classification. E PCA, as shown in (B), with mixed cells now classified as control or treated as predicted by KNN classification as shown in (D).
Because acetylation of histone H4 is known to be regulated by NaBut treatment13, we summarized the 16 different combinations of acetylation on H4 (4–17) into five main types based on the number of acetylations (i.e., 0, mono, di, tri, and tetra) and calculated the summarized H4 acetylation relative abundances (see “Methods” section). PCA of summarize H4 acetylation relative abundances clearly separated the control and treated cells along PC1 and was significantly correlated to the NaBut treatment variable (two-sided t-test, R = 0.46, p < 0.001) (Fig. 4C). Importantly, focusing on only collapsed H4 acetylation patterns increased the explained variance of PC1 from 35.46% to 73.2%, compared to analyzing all peptidoforms. We therefore leveraged the collapsed H4 acetylation relative abundances to train a k-nearest neighbor (KNN) classifier. We used a random sample of 50% of cells with known groups (i.e., control or treated) for training and the other 50% of labeled cells for testing, which achieved an accuracy of 84.21% (see “Methods” section). We then used the trained classifier to assign cells from the mixed population to the control or treated groups (i.e., control = 15 cells and treated = 17 cells), which resolved two distinct clusters of cells separated based on the NaBut treatment (Fig. 4D, E). The successful cell classification was further supported through pathway analysis of the differentially regulated proteins in the treated cells, which highlighted induced cell cycle arrest and inhibited mitochondrial complex formation (Supplementary Fig. 10D)52,53. In summary, this analysis demonstrated cell classification without knowing the assigned condition a priori by using informed selection of hPTM subgroups. This is particularly critical in the analysis of complex cell populations in vivo that contain different cell types or different cellular states at distinct ratios.
HDAC inhibitor treatment induces histone hyperacetylation and cell-to-cell chromatin heterogeneity
We next performed differential abundance analysis to assess changes in the abundance of all peptidoforms using 35 control and 29 treated single cells (see “Methods” section). As expected, we found many of the acetylated peptidoforms of H2A.1, H3 and H4 to be significantly increased in the treated group relative to the controls (Fig. 5A and Supplementary Table 4). The largest increase in relative abundance was observed for the tri-acetylated H4 peptidoforms, H4K5acK8acK16ac and H4K5acK12acK16ac. In support of this, analysis of the collapsed H4 acetylation relative abundances revealed a significant increase in di-, tri-, and tetra-acetylated hPTMs, independent of the mono-acetylated hPTMs (Fig. 5B). Analysis of global PTM relative abundances of each modification type (i.e. acetylation, mono-methylation, di-methylation and tri-methylation), confirmed the significant increase in acetylation, while also revealing a corresponding increase in mono-methylation and a decrease in di-/tri-methylation (Fig. 5C). Together, the global increases in acetylation are indicative of an open chromatin state with higher levels of transcription in NaBut treated cells relative to the controls, which was additionally supported by the 100-cell bulk injections (Supplementary Fig. 11A–D and Supplementary Table 5).
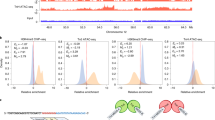
A Differential abundance of modified histone peptidoforms relative abundances in single cells for treated vs. control comparison. For the table of full results, see Supplementary Table 4. Statistics: control n = 35, treated n = 29; two-sided Wilcoxon test with Benjamini–Hochberg multiple testing correction. Red points are peptidoforms that are significantly differentially abundant (adjusted p-value < 0.05). B, C Summarized relative abundances of global PTMs (B) and H4 peptide acetylation (C) in single cells for control and treated groups. Statistics: control n = 35, treated n = 29; two-sided Wilcoxon test with Benjamini–Hochberg multiple hypothesis correction, adjusted p-values were 1.26 × 10⁻¹⁴ for 0-ac, 0.0534 for mono-ac, 2.16 × 10⁻¹⁶ for di-ac, 7.48 × 10⁻¹³ for tri-ac, and 1.59 × 10⁻¹¹ for tetra-ac, 1.07 × 10⁻⁷ for ac, 5.08 × 10⁻¹¹ for me, 0.0211 for me2, and 0.00018 for me3; p-value legend: *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Boxplots show the median (center), the 25th and 75th percentiles (bounds), and whiskers extending to the most extreme data points within 1.5× IQR; points beyond whiskers are plotted individually as outliers. D Heatmap and hierarchical clustering of modified histone peptidoform scaled relative abundances (Z-score) that were significantly differentially abundant (adjusted p-value < 0.05) between treated vs control groups as determined in (A). Single cells are shown in columns and histone peptidoforms are shown in rows. Peptidoforms are grouped by histone proteins (left side). E Sub-clustering results of treated group as shown in (D, right side), showing 2 subclusters (A and B) of cells identified within the treated group and the modified histone peptidoforms that are significantly differentially abundant (adjusted p-value < 0.1) between the subclusters. Subclusters were identified at the 2nd level of the hierarchical clustering tree shown in (D, top-right). Statistics: two-sided Wilcoxon test with Benjamini–Hochberg multiple testing correction. For full subcluster differential abundance results, see Supplementary Fig. 11E and Supplementary Table 6. Source data are provided as a Source Data file.
Interestingly, the relative abundance of H3 and H4 peptidoforms that were found to be significantly differentially abundant between control and treated groups (Fig. 5A, red points) was also observed to vary from cell-to-cell within the treated group, indicative of a heterogeneous response to the HDACi treatment (Fig. 5D, right side). This was supported by an increase in cell-to-cell heterogeneity in the treated group relative to the controls (Supplementary Fig. 12A). Thus, we sub-clustered the treated group and performed differential abundance analysis on the two resulting subclusters (Fig. 5E, Supplementary Fig. 11E, and Supplementary Table 6; see “Methods” section). Subcluster A was characterized by hyperacetylation of H4 tails, as well as high levels of H3K9acK14ac, which are known to be associated with active transcription. In contrast, cells within subcluster B showed relatively lower levels of H4 acetylation, as well as high levels of H3K9me3K14ac and H3K9me2, which are all associated with repressed transcription (Fig. 5E). Importantly, this differential response to the NaBut treatment was masked when analyzing the 100-cell bulk injections (Supplementary Fig. 11D). As it is possible that the classification of mixed cells into control or treated groups (as described in the previous section) may artificially induce intra-group variability, reanalysis without the mixed cells revealed comparable results (Supplementary Fig. 11F–I). In summary, our sc-hPTM method recapitulated the expected hyperacetylation effect of NaBut treatment using this analysis. Importantly, despite the relatively homogenous characteristics of HeLa cells within an in vitro system, the single-cell resolution revealed heterogeneous responses of individual cells to NaBut treatment.
Differential hPTM regulation induced by HDAC inhibitor treatment revealed by covariance analysis
The single-cell resolution of our sc-hPTM data has the potential to study the covariance of hPTM abundances and identify patterns of hPTM regulation in different biological contexts, such as modules of co-regulation that may indicate a shared function of a group of hPTMs. Importantly, this type of analysis is not possible in bulk data, as it is often masked by the averaging effect (Supplementary Fig. 11D). To perform hPTM covariance analysis, we correlated the relative abundance of each modified peptidoform with one another across the single cells and analyzed the resulting significant correlations (p-value < 0.05; see “Methods” section). It is important to note that because our MS-based bottom-up approach reflects the global profile of hPTMs within each cell, it does not offer single-molecule or single-nucleosome resolution to study the co-occurrence of hPTMs and is limited to the constraints of the proteolytic fragments produced by the digestion of histone tails. Thus, our covariance analysis is limited by the indirect correlations made between global hPTM levels. The resulting covariance matrices for the control (Fig. 6A, lower-left triangle; 125 significant correlations) and treated groups (Fig. 6A, upper-right triangle; 151 significant correlations) were hierarchically clustered relative to the control group (Supplementary Data 3). In the control group, this analysis revealed a module of positively correlated acetylated peptidoforms composed of H3K18ac and the majority of di-, tri-, and tetra-acetylated H4 peptides. Interestingly, these acetylated peptidoforms share a common association with open chromatin and gene expression3. In contrast, H3K9me and the H3K9me3K14ac bivalent mark were negatively correlated with the aforementioned cluster of acetylated peptidoforms, which corresponds to their known associations with inactive chromatin (Fig. 6A, lower-left triangle)54,55.
A Covariance matrix of modified histone PTM–PTM correlations derived from histone peptidoform relative abundances. PTM–PTM correlations for the control group are shown in the lower-left triangle, and the upper-right triangle are PTM–PTM correlations for the treated group. Colored boxes indicate Significant Pearson R correlation coeeficients (p-value < 0.05). Correlations of H3K23ac vs. H3K14ac and H4K5acK8acK12ac vs. H4K5ac, which are significantly altered due to NaBut treatment (adjusted p-value < 0.05), are highlighted in the control (blue dashed outline) and treated (red dashed outline) groups. Statistics: significance of Pearson R correlation coefficients was tested using a two-sided t-test; Significance of differential Pearson R correlation coefficients was tested using a permutation test with 10,000 permutations (see “Methods” section). For a table of Pearson R correlation coefficients p-values, see Supplementary Data 3. For full results and table of differential Pearson R correlation coefficient testing between treated vs. control groups, see Supplementary Fig. 12C, D and Supplementary Data 3. B Correlation of H3K23ac vs. H3K14ac histone peptidoform relative abundances in the control (left) and treated (right) groups was found to be significantly different (adjusted p-value < 0.0001) between the groups. Statistics: same as in (A). Control: R = 0.23, p = 0.19; Treated: R = 0.63, p = 0.00027. Shaded areas represent 95% confidence intervals of the fitted linear model. C Same as in (B), but for H4K5acK8acK12ac vs. H4K5ac whose correlation was significantly different (adjusted p-value < 0.0001) between the groups. Statistics: same as in (A). Control: R = 0.54, p = 0.00089; Treated: R = 0.13, p = 0.5. Shaded areas represent 95% confidence intervals of the fitted linear model. D, E Network diagrams derived from covariance matrices as shown in (A), showing the significant Pearson R correlation coefficients (p-value < 0.05) of modified histone PTM–PTM correlations (edges) for each of the modified peptidoforms (nodes). The control group is shown in (D) and treated in (E). Node size represents node degree (i.e., number of significant correlations), node color represents mean relative abundance amongst all cells in control and treated groups, edge color represents absolute correlation coefficient, and edge width represents the scaled correlation coefficient (see legend on left side). Statistics: same as in (A). Source data are provided as a Source Data file.
We then compared the covariance matrices of the control and treated group, which were found to be significantly different (Mantel’s test, p-value = 9.99 × 10−5) with a shift towards positive correlations in the treated group (Supplementary Fig. 12C). We checked if this was related to increased peptidoform variance, but found no difference (Supplementary Fig. 12B). Thus, the shift towards more positive correlations in the treated group suggests that under normal conditions, a role of HDACs are to suppress positive coordination between hPTMs. An example of this can be seen from the increased correlation between H3K23ac and H3K14ac in the treated group relative to the control group (Fig. 6B). This was orthogonally validated via flow cytometric immunostaining of intracellular H3, H3K23ac, and H3K14ac epitopes, providing single-cell measurements with robust statistical power derived of tens of thousands of cells (Supplementary Fig. 13; see “Methods” section). On the contrary, a significant loss of correlation was found between H4K5acK8acK12ac and H4K5ac in the treated group relative to the controls, which suggests that the coordinated acetylation of these mono- and tri-acetylated H4 peptidoforms is dependent on HDACs (Fig. 6C).
To obtain a complementary perspective of hPTM covariance, we visualized histone PTM–PTM correlations as networks for the control and treated groups, respectively (Fig. 6D, E). This revealed patterns of hPTM co-regulation both within (intra-) and across histone tails (inter). For example, an increase in intra-H3 co-regulation was observed in the treated groups as compared to the control group, which supports a regulatory role of HDACs to suppress intra-hPTM crosstalk (Fig. 6D). Interestingly, highly connected hPTM hubs potentially indicating central regulatory roles, such as H4K5acK16ac or H3K9meK14ac, were more prevalent in the treated group as compared to the control (Fig. 6E). Taken together, the analysis of how hPTMs indirectly interact or co-regulate, and how this changes across different biological contexts, will contribute to deciphering the histone code that was hypothesized over 20 years ago2.
Discussion
This study presents an innovative MS-based workflow that enables comprehensive and accurate profiling of hPTMs at single-cell resolution, termed sc-hPTM (Fig. 1A). We successfully identified 67 histone peptidoforms encompassing 25 individual hPTMs; of which at a minimum of 44 were confidently quantified. Comparatively, in bulk histone analysis via LC-MS/MS, approximately 200–300 histone peptidoforms are typically identified28,50,56,57. Our workflow constitutes a significant advance over previous antibody-based single-cell epigenomic approaches, which are limited in the number of targets and prone to cross-reactivity. Moreover, our methodology enables the identification and quantification of less common PTMs beyond currently available antibodies, such as H3K23su and H3K27hib, or previously unrecognized modifications (i.e., histidine methylation on H2A as demonstrated in bulk analysis58).
The single-cell resolution of our method allowed us to further dissect the heterogeneity of chromatin dynamics in response to the HDAC inhibitor NaBut. Although this treatment is known to induce histone acetylation to activate gene expression, we identified an unexpected subpopulation of cells with a hPTM profile suggestive of chromatin silencing (Fig. 5E). It will be interesting for future studies to elucidate what factors might lead to the development of this subpopulation of cells. In addition, differential hPTM covariance analysis revealed a strengthened positive correlation between H3K23ac and H3K14ac in response to the NaBut treatment (Fig. 6B and Supplementary Fig. 13F). These orthogonally validated results support previous findings implicating p300/CBP as the common lysine acetyltransferase to target these histone PTMs59, as well as results demonstrating that a positive cross-talk between these histone PTMs can be attributed to binding of the DPF subunit of the MORF complex to acylated H3K14, promoting H3K23ac60. Thus, differential hPTM covariance analysis has the potential to shed light on how epigenetic networks are rewired in development and disease.
In contrast to traditional bulk or low-input analysis, we utilize excess sample preparation reagents to account for the low sample concentration, bypass histone extraction, and prepare the entire cell contents to minimize adsorptive losses. Intuitively, histone extraction may lead to increased sensitivity by reducing background interference. However, introducing this step is infeasible for single-cell amounts due to the centrifugation step required32. In addition, bypassing histone extraction allowed for the analysis of other proteins within the same cell, providing simultaneous insights into cellular processes and their relation to the histone code (Supplementary Fig. 10). Advances in instrumentation have recently allowed the detection of protein PTMs using standard SCP workflows, including histone modifications due to their relatively high abundance in the cell61,62. While these publications are excellent proofs of concept for selected PTMs, they have intrinsic limitations, including the reduced coverage of detectable peptides due to trypsin digestion without derivatization. Although the use of propionylation in our sc-hPTM method provides good histone coverage, employing phenyl isocyanate for secondary histone propionylation could further enhance the detection of short hydrophilic peptides and low-abundance acetylations, such as H3K27Ac and H3K36Ac15,63. Moreover, detection of long, hydrophobic peptides may be improved by using RgpB (also known as Gingipain or GingisREX, GRX), an ArgC-specific enzyme that eliminates the need for lysine derivatization. However, GRX digestion fails to retain shorter H3 and H4 N-tail peptidoforms, which are biologically significant64. In addition to optimizing sample preparation, we aim to increase the analysis throughput by employing isotopic multiplexing and targeted MS acquisition, such as parallel reaction monitoring (PRM)35,65. We expect that very short gradients, such as 5 min, will be feasible while maintaining comparable sensitivity, potentially allowing the analysis of hundreds of cells per day35,65.
In contrast to existing single-cell methods for hPTM analysis, which are exclusively antibody-based, our MS-based method enables the detection of co-occurring histone modifications, or combinatorial histone marks. We anticipate that technological developments to reduce input requirements for middle-down and top-down MS approaches will enhance the detection of combinatorial PTMs by analyzing longer peptides or intact histone proteins and thus preserving spatial relationships between the modifications. Combinatorial hPTM patterns are thought to play a crucial role in establishing distinct chromatin states and recruiting chromatin-modifying enzymes. However, accurate quantification of combinatorial hPTMs from bulk injections is challenged by cell-to-cell heterogeneity. Our sc-hPTM method has demonstrated that specific histone peptidoforms, such as H3K9me2K14ac and H3K9me3K14ac (Supplementary Fig. 3D), are relatively abundant in single cells, which has the potential to shed light on the role of chromatin domains decorated by both silencing and activating marks. We identified H4K5acK12ac as having the largest variation amongst HeLa cells in culture, which is likely due to its known role in chromatin condensation during mitosis, as cells in these experiments were not synchronized by cell-cycle51,66. In contrast, the lack of variation in H2AJK5acK9ac is poorly studied, but may represent a stable epigenetic modification in proliferating cells in culture (Fig. 3F). Such insights into the cell-to-cell variability of the epigenetic landscape have important implications for understanding mechanisms of cellular proliferation, differentiation, adaptation, disease progression, and aging.
While the intricate combinatorial complexity of the histone code allows for billions of combinations of hPTMs, the comprehensive and reproducible fragmentation patterns generated with DIA enable the relative comparison of these hPTMs across large sample numbers. However, manual peak picking (e.g., due to identical mass shifts of certain PTMs that result in isobaric peptidoforms) is a major bottleneck as it is time-consuming and requires in-depth knowledge of histone peptides. At present, only a limited number of tools, such as EpiProfile46, which is specifically designed for data produced by a Thermo Fisher Scientific MS instrument, can automatically process histone DIA data. Thus, the development of dedicated computational tools tailored for the analysis of histone diaPASEF data from Bruker instruments will be an important step towards increasing the accessibility and scale of this approach. We anticipate that especially shorter gradients will benefit from current ion mobility prediction tools67,68,69, capable of accurately predicting highly modified peptidoforms to improve the confident and streamlined interpretation of convoluted sc-hPTM data70. While the present study was conducted on a single cultured cell model, future validation in primary tissues and heterogeneous samples (e.g., organoids and biopsies) will be needed to establish broader applicability and translational relevance.
In conclusion, this study presents a robust and versatile MS-based workflow, sc-hPTM, to profile the histone code at single-cell resolution. We provide unprecedented insights into the epigenetic heterogeneity within cell populations, and we envision that this method will open avenues for investigating the role of the histone code in diverse biological processes and diseases.
Methods
Cell culture
Low passage HeLa cells (ATCC CCL-2) were grown as adherent cultures in Eagle’s Minimum Essential Medium (ATCC 30-2003) supplemented with 10% fetal bovine serum (Thermo Fisher Scientific A5256801) and 1% penicillin-streptomycin (Thermo Fisher Scientific 15140122) in 10 cm dishes (Thermo Fisher Scientific 172931) at 5% CO2 and 37 °C. Cells were passaged at 70–80% confluence every 3 days. For drug treatment studies, fresh medium was supplemented with either 1× PBS (Thermo Fisher Scientific 10010023) as a vehicle control or sodium butyrate (Sigma-Aldrich 303410) at a concentration of 5 mM. The treatment was administered for 24 h, after which the cells were harvested for single-cell isolation and downstream sample preparation.
Cell isolation
Prior to harvesting, cells were washed with ice-cold 1× PBS. Cell detachment was achieved by treating the cells with 1 mL of trypLE (Thermo Fisher Scientific 12604013), followed by incubation at 37 °C for 3–5 min. The cells were washed two times with ice-cold 1× PBS, centrifuged at 500×g for 5 min at 4 °C, and the cell pellet was resuspended in ice-cold 1× PBS and strained using a 40 µm cell strainer (Millipore Sigma CLS431750), followed by centrifugation and washing. Cell concentration was determined using a Countess™ 3 Automated Cell Counter (Fisher Scientific AMQAX2000) to obtain a 200 cells/µL suspension. Cells were kept on ice prior to cell sorting.
Single-cell isolation and histone peptide preparation
All reagents were prepared fresh on the day of sample preparation. Each experiment (including sample preparation) were performed on separate days, which constituted a batch. For each batch, cells are isolated into a 384-well plate, which is then subject to histone peptide preparation. A type 1 piezo dispense capillary (Cellenion) was used for cell sorting and liquid handling within the cellenONE (Cellenion). All workflow steps outside of the cellenONE were done in a sterile cell culture hood with laminar air flow. Custom scripts were written within the cellenONE software for each step of the process (see ‘Code Availability’). One microliter of lysis reagent (0.2% dodecyl maltoside, Thermo Fisher Scientific 89902 in 1 M TEAB, Millipore Sigma 18597) was dispensed to respective wells of the 384-well plate (Thermo Fisher Scientific AB1384) using an electronic multichannel pipette (Sartorius LH-747321) and then placed in a 384-well plate holder in the cellenONE at 4 °C. A 1 pg/nL bovine histone standard (Sigma-Aldrich H9250) suspension in 1× PBS was degassed on ice for 10 min, and indicated amounts (i.e., 10, 20, 40, 80, and 100 pg) were dispensed using the ‘Reagent_Dispense’ program (Supplementary Table 1). Following this, the plate was sealed (Thermo Fisher Scientific AB0626), vortexed for 5 s, and centrifuged at 500×g for 30 s. To filter out dead cells during cell sorting, DAPI stain (Thermo Fisher Scientific D1306) was added to the cell suspension prior to sorting at a concentration of 1 µg/mL, followed by degassing on ice for 10 min. Live single cells (DAPI-) were sorted into the plate using the ‘cellenONE_Basic_WithscanFields’ program, where images of each isolated cell were acquired to verify successful isolation (Supplementary Fig. 1A–D and Supplementary Tables 1 and 3). For the 100-cell samples, the original cell suspension was resuspended in lysis reagent at a concentration of 100 cells/µL and manually dispensed 1 µL into the specified wells (Supplementary Tables 1 and 3) and were prepared identically to single-cell injections. The plate was then sealed, vortexed for 5 s, and centrifuged at 500×g for 30 s.
The 10× propionylation reagent (25% propionic anhydride, Sigma-Aldrich 240311 in acetonitrile Sigma-Aldrich 34851) was degassed on ice for 10 min. Nozzle parameters, i.e., voltage, pulse, and delay (400 ms), were tuned to account for the low viscosity of the propionylation reagent to obtain a stable drop (Supplementary Fig. 1E). The ‘Derivatization_Dispense’ program was then used to dispense 100 nL of propionylation reagent to obtain a final concentration of 1X in each well. The plate was then sealed, vortexed for 5 s, and centrifuged at 500×g for 30 s. Images of wells were acquired to confirm successful reagent dispense (Supplementary Fig. 1F, G). The plate was then incubated at room temperature for 30 min, followed by drying inside the cellenONE for 10–15 min at 50 °C. One microliter of master mix reagent (3 ng/µL trypsin Gold Promega V5280, 0.2 U/µL Benzonase® Nuclease Millipore E1014, 0.01% ProteaseMAX™ Promega V2071, and 1 M TEAB) was dispensed into wells. The plate was then sealed, vortexed for 5 s, and centrifuged at 500×g for 30 s. The sealed plate was placed back into the cellenONE and incubated for 2 hours at 50 °C using the ‘Digestion_Incubation’ program. Following the incubation, the plate was centrifuged at 500×g for 30 s to pull down any condensation. The plate was then placed back into the cellenONE for the 2nd round of derivatization, as described above. Finally, derivatized and digested histone peptides were diluted by dispensing 2.9 µL of dilution solution (0.1% Trifluoroacetic acid, Sigma-Aldrich T6508, 5% DMSO Thermo Fisher Scientific 85190, mass spec grade H2O, Sigma-Aldrich 900682) using an electronic multichannel pipette. The plate was then sealed, vortexed for 5 s, and centrifuged at 500×g for 30 s. Prepared plates were stored at −80 °C until LC-MS/MS analysis.
Intracellular immunohistochemistry
Isolated cells were incubated with 1:5000 fixable viability stain (BD Biosciences 565388) in 100 µL dPBS for 30 min on ice, followed by a PBS wash (5 min, 500×g). Then, cells were fixed and permeabilized using the Intracellular Flow Cytometry Kit (Methanol) (Cell Signaling 13593), followed by a wash in PEB buffer (PBS supplemented with 0.5% BSA and 2 mM EDTA; 8 min, 500×g).
For intracellular staining, cells were incubated with 1:50 anti-H3K23ac for 45 min in 300 µL PEB (Thermo Fisher Scientific, BS-3774R) and then washed with PEB (5 min, 500×g). This was followed by staining with 1:1000 Alexa Fluor 555–conjugated secondary antibody for 30 min in 300 µL PEB (Thermo Fisher Scientific A-21428) and then washed 2× with PEB (5 min, 500×g). Cells were then stained with 1:50 conjugated anti-H3 (Cell Signaling 82952) and 1:50 conjugated anti-H3K14ac (Cell Signaling 26828) in parallel for 45 min in 300 µL PEB, and then washed with PEB (5 min, 500×g). Finally, 30 min before flow cytometry analysis, cells were incubated with 1:1000 FxCycle (Invitrogen R37166).
For compensation and gating controls, single-stained controls were prepared in parallel, which included: unstained, viability only (using cells heated at 55 °C for 15 min to induce cell death), FxCycle only, anti-H3 only, anti-H3K14ac only, anti-H3K23ac only, and Alexa Fluor 555-conjugated secondary only. We also included a control where staining was performed with the conjugated antibodies anti-H3 and anti-H3K14ac with the Alexa Fluor 555-conjugated secondary to ensure no minimal cross-reactivity between the primary and secondary antibodies.
Flow cytometry analysis
Cells were analyzed on a Cytek Aurora spectral flow cytometer equipped with 405 nm, 488 nm, and 561 nm lasers. Acquisition was performed using SpectroFlo software (version 3.3.0) for spectral unmixing and autofluorescence extraction. Single-staining controls (as described above) were used to build a robust unmixing matrix, and single-cell level compensated parameters were exported in the CSV-channel format with the FlowJo software (version 11). Following this, downstream analysis was performed using a custom R script (version 4.5.1), including: (1) quality control to remove debris, cell doublets, dead cells, and outliers based on H3, H3K23ac, H3K14ac, and FxCycle; (2) cell cycle analysis; (3) differential abundance comparing control and treated populations; and (4) histone PTM–PTM covariance analysis (see ‘Code Availability’).
LC-MS/MS
Each batch of injections (i.e., a 384-well plate containing prepared histone peptides) was acquired using a Vanquish Neo UHPLC system (Thermo Fisher Scientific) coupled to a timsTOF Ultra (Bruker Daltonik GmbH). The contents of each well within the 384-well plate were defined as an injection, which consisted of blanks, cells, or histone standards. Peptides were separated on an Aurora Ultimate (25 cm × 75 µM, 1.7 µm particle size and 120 Å pore size IonOpticks AUR3-25075C18-CSI) integrated emitter column and eluted over a 30-min window using the following gradient: 5–23% Solvent B (0.1% formic acid Sigma-Aldrich F0507, acetonitrile) in 25.7 min, 23–35% Solvent B in 2.6 min and 35–45% Solvent B in 1.7 min, at a flow rate of 200 nL/min. The actively acquired sample plate was stored in the Vanquish Neo autosampler at 7 °C. To minimize evaporation, injections were unsealed for 12 h of measurement time, and the entire sample volume was injected for every analytical run.
For the dia-PASEF acquisition, full MS data were acquired in the range of 100–1700 m/z and 1.3 1/K0 [V-s/cm−2] to 0.6 1/K0 [V-s/cm−2]. DIA windows ranged from 300 m/z to 1000 m/z with 30 Th isolation windows and were acquired with ramp times of 200 ms. High sensitivity detection for low sample amounts was enabled without dia-PASEF data denoising. The collision energy was ramped as a function of increasing mobility, starting from 20 eV at 1/K0 [V-s/cm−2] to 59 eV at 1.6 1/K0 [V-s/cm−2].
Histone peptide identification and quantification
Peptide identification and quantification were performed using Skyline (version 22.2) as previously described in ref. 12. Briefly, ArgC was used as the digestion enzyme due to peptide derivatization. Charge states +1 and +2 were considered for precursor ions, and charge states +1 and +2 were considered for product ions. All sidechain modifications were set as variable modifications, and a maximum of four modifications were allowed to occur on any peptide. Propionyl on K and the N-terminus were set as fixed. Importantly, Skyline only allows one modification per amino acid. However, a first position K can present a propionyl on the N-terminus and a variable modification on the side chain. To allow for multiple modifications per amino acid, new modifications were defined by combining the mass of propionyl groups with all possible side chain modifications. Next, a FASTA database containing all histone variants and all possible modified histone ArgC peptides was used to create a target library in Skyline, as previously described in ref. 13. Shared peptides between different histone variants were given names of both histone variants. The staggered windows used for diaPASEF acquisition (as described in the previous section) were set as the isolation scheme. Raw diaPASEF data of the 100-cell and 100 pg histone standard injections were used to manually curate a reference for single-cell injections. This was done based on the alignment of MS1 and MS2 XICs, relative retention time differences expected between unmodified and modified peptides, and filtering for unique transitions (if applicable). Peptides that could not be confidently identified were filtered out for further analysis. One Skyline project containing 100-cell samples was used to create a spectral library to facilitate peak picking for all single-cell injections and additional separate projects per batch. Peak picking was limited to scans within 2 min of the MS/MS ID in the spectral library, with manual adjustment if needed. Each identified peptide was validated against signal absence in injections containing no cells and peak alignment with the histone standard and the 100-cell injections.
Isobaric histone peptide identification and quantification
Histone H4 contains isobaric peptides that do not separate chromatographically (Supplementary Fig. 3A). Thus, we deconvoluted the MS1 signal intensity using a combination of unique fragment ions for each peptidoform (Supplementary Fig. 3B) as has been previously described46. All fragment ion intensities unique for the isobaric peptide were summed, and the contributing proportion of that isobaric peptide to all fragments was calculated. The relative proportion of isobaric peptides that contain a unique combination of fragment ions is calculated through their proportion relative to the sum of all the fragment ions of that peptide. The relative contribution of all isobaric peptides totals to the MS1 intensity of the respective isobaric peptide. These calculations can be expressed as the following set of equations.
For a set of n isobaric peptides (P1, P2,…, Pn), where P1 has unique fragments and P2 to Pn have a unique combination of fragments:
Where
Thus, for i = 2 to n:
Given: MS1original = Original (convoluted) MS1 signal, MS1deconvoluted(Pi) = Deconvoluted MS1 signal for peptide I, Prop(Pi) = Proportion of signal attributed to peptide I, UF(Pi) = Sum of unique fragment intensities for peptide I, UCF(Pi) = Sum of unique combination fragment intensities for peptide I, and TF(Pi) = Sum of all fragment intensities for peptide i.
Derivatization efficiency analysis
We obtained estimates for propionylation derivatization efficiency for single and 100-cell injections by extracting various forms of the H3.1 9–17 peptide, which is easily detected in all injections56,71. To calculate this, we extracted the MS1 areas of the fully propionylated peptide (ArgC digest with propionylation on lysine and N-terminus), under propionylated in the 1st round (Trypsin digest with propionylation on lysine and N-terminus), under propionylated in the 2nd round (ArgC digest with propionylation on lysine), under propionylated in the 1st and 2nd rounds (Trypsin digest with no propionylation), and over propionylation (ArgC digest with propionylation on lysine, tyrosine, and N-terminus). Propionylation derivatization efficiency was then calculated as the percent of the peptide not correctly propionylated (i.e., the under- or overpropionylated forms) using the unnormalized MS1 intensities in the following equation:
Cell filtering
Our ‘one-pot’ histone derivatization protocol enabled the evaluation of the global ‘background’ proteome to determine sample preparation dropouts prior to dedicated hPTM quality control filtering (Supplementary Fig. 10A). To account for any errors that may have occurred during the histone peptide preparation, the distribution of unnormalized MS1 intensities for blank and single-cell injections are plotted and then used to calculate the median absolute deviation (MAD) for each injection relative to all cells in the given analysis. We then used a dynamic threshold to exclude cells, which was based on the MAD value that was sufficient to exclude all the injections containing no cells. This threshold varied from MAD > 0.8 in the analysis, which included injections from batches 1–3, to MAD > 1 in the analysis that included batches 1–5 (Supplementary Figs 6A, 9A).
To account for variation in derivatization efficiency that may have occurred during the histone peptide preparation, we calculated and plotted the propionylation derivatization efficiency (i.e., percent of histone peptides that were under- and overpropionylated) for single- and 100-cell injections as described in the previous section. We then used this to calculate the MAD for each injection relative to all cells in the given analysis, and then used a MAD threshold > 2 to exclude cells (Supplementary Figs. 3F, 9E).
Peptidoform filtering
Histone peptides that were only detected in their unmodified form were removed, as both unmodified and modified forms are needed to calculate hPTM relative abundance (described in the following section). For peptidoforms with multiple precursor charges (e.g., +2 and +3), the peptidoform with the charge state that showed a lower median MS1 signal intensity across all single cells in the analysis was removed. Peptidoforms that were not detected in greater than 50% of injections were removed due to inconsistent detection. Peptidoforms with a median MS1 intensity in single-cell injections that was not at least 150% greater than the median MS1 intensity in injections containing no cells were removed due to low signal intensity (Supplementary Fig. 3D). Peptidoforms that resulted in low (median R < 0.8) or insignificant correlations (p-value > 0.05) when calculating their respective titration curves were removed due to lack of quantifiability (Supplementary Fig. 4A). The peptidoforms and the reasons they were filtered out for each of the analyses can be found in Supplementary Data 2.
hPTM relative abundance normalization
To calculate the hPTM relative abundance, for each histone peptide, the MS1 intensity of the unmodified and modified peptidoforms was divided by the sum of unmodified and modified peptidoforms (e.g., for H2A1 4–11, H2A1K5unK9un and H2A1K5acK9un were used to create the peptide ratio), followed by a Log(1 + x) transformation (Supplementary Figs. 6B, 9B). Single PTM ratios were calculated by summing the peptide ratios for every unique modification at their respective amino acids within a peptide (e.g., H3K9ac = H3K9acK14un + H3K9acK14ac). Global PTM ratios were calculated by summing the PTM ratios for each unique modification (e.g., ac = H3K9ac + H3K14ac + H4K8ac + etc.). The summarized H4 acetylation PTM ratios were calculated by summing the peptide ratios of peptides containing 0 (Unmodified), 1 (Mono-Acetylation), 2 (Di-Acetylation), 3 (Tri-Acetylation) acetylations, or 4 (Tetra-Acetylation) (e.g., mono = H4K5acK8unK12unK16un + H4K5unK8acK12unK16un + H4K5unK8unK12acK16un + H4K5unK8unK12unK16ac).
Batch correction
The presence of batch effects was determined by performing PCA and then through qualitative examination of the first 2 principal components and their correlation to batch identities (Supplementary Figs. 6C, 9G). The removeBatchEffect function from the limma package72 was then used to correct for the observed batch effect while preserving the effects due to differences in treatment.
Single-cell vs 100-cell bulk comparisons
The 100-cell bulk and single-cell injections from batches 1–3 were processed through our quality control filtering, normalization, and batch-correction pipeline as described above. The mean hPTM relative abundance was calculated across all single cells to create ‘pseudobulk’ samples.
Technical and biological noise analysis
To define technical noise, bovine histone standard injections from batches 1–3 were processed through our quality control filtering as described above. The CVs of hPTM relative abundances were calculated within each batch (intra-) or across batches (inter-). Single-cell level 10 pg histone standards were compared to external datasets, including histone peptide ratio data of bulk technical replicates derived from HeLa cells (Sidoli et al. A49: 5 replicates, 16 peptidoforms; Sidoli et al. B50, 4 replicates, 12 peptidoforms) and level 3 normalized data (Light(PTM)/Heavy(PTM) divided by Light(NORM)/Heavy(NORM)) of processing replicates for cell lines in the NIH LINCS (3 replicates, 29 peptidoforms) (https://panoramaweb.org/lincs_pccse_gcp_2020.url33,73). For both external datasets, peptidoforms were matched to those identified in histone standard injections. CVs were then calculated for the matched peptidoforms (Supplementary Table 2) of the technical replicates in the external datasets relative to the inter-batch CV of the respective peptidoforms of the histone standards in the current dataset.
To define biological noise, single-cell injections from batches 1–3 were processed through our quality control filtering as described above, and the CVs of hPTM relative abundances were calculated. To remove technical noise, the variance of each peptidoform derived from the 10 pg bovine histone standard injections was subtracted from the respective variances of peptidoforms in the single cell injections. Following this, the square root of the adjusted variance of each peptidoform is divided by its respective mean to obtain the CV after technical noise removal (i.e., biological noise). Peptidoforms with greater technical noise than the sum of biological and technical noise were discarded from downstream analyses.
Cell classification analysis
Single-cell injections from batches 1–5 were processed through our quality control filtering pipeline as described above (Supplementary Fig. 9). PCA of single cells was performed using the PCAtools package74, where hPTM relative abundances were centered and scaled. Summarized H4 acetylation relative abundances (i.e., 0, mono-, di-, tri-, and tetra-acetylation) were calculated as described above. Following this, PCA was performed on the centered and scaled summarized H4 acetylation relative abundances. The heatmap of this was plotted using the ComplexHeatmap package75, where hierarchical clustering was performed on the rows (collapsed H4 acetylations) and columns (single-cell injections). To classify mixed cells (i.e. those where the group was unknown), either the control or treated groups, we trained and evaluated a K-nearest-neighbors (KNN) classifier based on the summarized H4 acetylation relative abundance Z-scores. To evaluate the KNN classifier, cells from the control and treated groups were randomly split 50:50 into training and test groups, where k = 2. Following this, all cells from the control and treated groups were used to train the classifier and this was then used to classify the mixed cells into the control or treated groups. The predicted labels (control or treated) of the mixed cells were then used in the downstream analyses.
Differential abundance analysis
Single-cell and 100-cell injections from batches 1–5 were processed through our quality control filtering pipeline as described above (Supplementary Fig. 9). For 100-cell injections, hPTM differential abundance (4 treated vs 8 control samples) was performed using an unpaired two-sided t-test followed by Benjamini–Hochberg multiple hypothesis correction. For single-cell injections, hPTM differential abundance (29 treated vs 35 control cells) was performed using an unpaired Wilcoxon rank sum test followed by Benjamini–Hochberg multiple hypothesis correction. Following this, the relative abundances of hPTMs that were identified to be significantly differentially abundant between the treated and control groups (adjusted p-value < 0.05) were centered and scaled (Z-scores) and then plotted as a heatmap using the ComplexHeatmap package75, where hierarchical clustering was performed on the rows (peptidoforms) and columns (single cells) within each group. Differential abundance analysis of relative abundances for global PTM and collapsed H4 acetylation PTM relative abundances for single-cells and 100-cell injections was performed using an unpaired Wilcoxon rank sum test.
Sub-clustering analysis
Following single-cell hPTM differential abundance analysis and hierarchical clustering of the significantly differentially abundant (adjusted p-value < 0.05) peptidoforms, 2 subclusters were identified at the 2nd level of the hierarchical clustering tree which were labeled ‘A’ and ‘B.’ hPTM differential abundance analysis (5 cells in subcluster A vs 24 cells in subcluster B) was performed using an unpaired Wilcoxon rank sum test followed by Benjamini–Hochberg multiple hypothesis correction. Following this, the relative abundances of hPTMs that were identified to be significantly differentially abundant between the A and B subclusters (adjusted p-value < 0.1) were scaled and centered (Z-scores) and then plotted as a heatmap as described above.
hPTM covariance analysis
For each treatment group (29 control cells and 35 treated cells), a covariance matrix was calculated for all pairs of modified peptidoforms to obtain Pearson R values of each PTM–PTM correlation, and the significance of the correlations was estimated using a two-sided t-test. The lower triangle of each covariance matrix was visualized using the corrplot package76, where only significant correlations (p-value < 0.05) were shown. Comparison of treated vs control covariance matrices at the global level was performed using Mantel’s permutation test with 10,000 permutations. Comparison of treated vs control PTM–PTM correlations at the peptidoform level was calculated as Log2 fold changes, and p-values were estimated using a permutation test followed by Benjamini–Hochberg multiple hypothesis correction. Following this, scatter plots of hPTM relative abundance for PTM–PTM correlations that were found to be significantly different in the treated vs control comparison were plotted, where Pearson R correlation coefficients were calculated, and p-values of these correlations were estimated using a two-sided t-test.
Cell–to-cell variation analysis
Single-cell injections from batches 1–5 were processed through our quality control filtering pipeline as described above (Supplementary Fig. 9). For each treatment group (29 control and 35 treated), pairwise Euclidean distance based on the relative abundance of all modified peptidoforms was calculated for all pairs of cells within each group. The distribution of pairwise Euclidean distances was then compared between treated vs control groups, and the significance of this comparison was estimated using a two-sided t-test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw mass spectrometry-based proteomics data generated in this study have been deposited in the MassIVE database under accession code MSV000094801. Source data is publicly available and has been deposited in figshare at https://doi.org/10.6084/m9.figshare.3040320177.
Code availability
The code used to develop the perform the analyses and generate results in this study is publicly available and has been deposited in Github at, https://github.com/cutleraging/single-cell-histone-ptm under an MIT license. The specific version of the code associated with this publication is archived in Zenodo and is accessible via https://doi.org/10.5281/zenodo.1728224078.
References
Bernstein, B. E., Meissner, A. & Lander, E. S. The mammalian epigenome. Cell 128, 669–681 (2007).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Shogren-Knaak, M. et al. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311, 844–847 (2006).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
van Mierlo, G. et al. Integrative proteomic profiling reveals PRC2-dependent epigenetic crosstalk maintains ground-state pluripotency. Cell Stem Cell 24, 123–137.e8 (2019).
Chen, K. et al. Neurodegenerative disease proteinopathies are connected to distinct histone post-translational modification landscapes. ACS Chem. Neurosci. 9, 838–848 (2018).
Kontopoulos, E., Parvin, J. D. & Feany, M. B. Alpha-synuclein acts in the nucleus to inhibit histone acetylation and promote neurotoxicity. Hum. Mol. Genet. 15, 3012–3023 (2006).
Fraga, M. F. et al. Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nat. Genet. 37, 391–400 (2005).
van der Kroef, M. et al. Histone modifications underlie monocyte dysregulation in patients with systemic sclerosis, underlining the treatment potential of epigenetic targeting. Ann. Rheum. Dis. 78, 529–538 (2019).
Creech, A. L. et al. Building the connectivity map of epigenetics: chromatin profiling by quantitative targeted mass spectrometry. Methods 72, 57–64 (2015).
Litichevskiy, L. et al. A library of phosphoproteomic and chromatin signatures for characterizing cellular responses to drug perturbations. Cell Syst. 6, 424–443.e7 (2018).
Strahl, B. D. et al. Methylation of histone H4 at arginine 3 occurs in vivo and is mediated by the nuclear receptor coactivator PRMT1. Curr. Biol. 11, 996–1000 (2001).
Stransky, S., Cutler, R., Aguilan, J., Nieves, E. & Sidoli, S. Investigation of reversible histone acetylation and dynamics in gene expression regulation using 3D liver spheroid model. Epigenetics Chromatin 15, 35 (2022).
Abshiru, N. A. et al. Targeted detection and quantitation of histone modifications from 1,000 cells. PLoS One 15, e0240829 (2020).
Noberini, R. et al. Spatial epi-proteomics enabled by histone post-translational modification analysis from low-abundance clinical samples. Clin. Epigenetics 13, 1–16 (2021).
Kieffer, Y. et al. Single-cell analysis reveals fibroblast clusters linked to immunotherapy resistance in cancer. Cancer Discov. 10, 1330–1351 (2020).
Lawson, D. A. et al. Single-cell analysis reveals a stem-cell program in human metastatic breast cancer cells. Nature 526, 131–135 (2015).
Gou, B. et al. Single-cell analysis reveals transcriptomic reprogramming in aging cardiovascular endothelial cells. Front Cardiovasc Med. 9, 900978 (2022).
Cheung, P. et al. Single-cell chromatin modification profiling reveals increased epigenetic variations with aging. Cell 173, 1385–1397.e14 (2018).
Rotem, A. et al. Single-cell ChIP-seq reveals cell subpopulations defined by chromatin state. Nat. Biotechnol. 33, 1165–1172 (2015).
Ku, W. L. et al. Single-cell chromatin immunocleavage sequencing (scChIC-seq) to profile histone modification. Nat. Methods 16, 323–325 (2019).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1–10 (2019).
Fuchs, S. M., Krajewski, K., Baker, R. W., Miller, V. L. & Strahl, B. D. Influence of combinatorial histone modifications on antibody and effector protein recognition. Curr. Biol. 21, 53–58 (2011).
Egelhofer, T. A. et al. An assessment of histone-modification antibody quality. Nat. Struct. Mol. Biol. 18, 91–93 (2011).
Peach, S. E., Rudomin, E. L., Udeshi, N. D., Carr, S. A. & Jaffe, J. D. Quantitative assessment of chromatin immunoprecipitation grade antibodies directed against histone modifications reveals patterns of co-occurring marks on histone protein molecules. Mol. Cell. Proteom. 11, 128–137 (2012).
Lu, C., Coradin, M., Porter, E. G. & Garcia, B. A. Accelerating the field of epigenetic histone modification through mass spectrometry-based approaches. Mol. Cell. Proteom. 20, 100006 (2021).
Verhelst, S. et al. Comprehensive histone epigenetics: a mass spectrometry based screening assay to measure epigenetic toxicity. MethodsX 7, 101055 (2020).
Garcia, B. A. et al. Chemical derivatization of histones for facilitated analysis by mass spectrometry. Nat. Protoc. 2, 933–938 (2007).
Knapp, A. R. et al. Quantitative profiling of histone post-translational modifications by stable isotope labeling. Methods 41, 312–319 (2007).
Govaert, E. et al. Extracting histones for the specific purpose of label-free MS. Proteomics 16, 2937–2944 (2016).
Sidoli, S., Bhanu, N. V., Karch, K. R., Wang, X. & Garcia, B. A. Complete workflow for analysis of histone post-translational modifications using bottom-up mass spectrometry: from histone extraction to data analysis. J. Vis. Exp. https://doi.org/10.3791/54112 (2016).
Dele-Oni, D. O. et al. Proteomic profiling dataset of chemical perturbations in multiple biological backgrounds. Sci. Data 8, 226 (2021).
Schoof, E. M. et al. Quantitative single-cell proteomics as a tool to characterize cellular hierarchies. Nat. Commun. 12, 3341 (2021).
Petelski, A. A. et al. Multiplexed single-cell proteomics using SCoPE2. Nat. Protoc. 16, 5398–5425 (2021).
Ctortecka, C. et al. Automated single-cell proteomics providing sufficient proteome depth to study complex biology beyond cell type classifications. Nat. Commun. 15, 5707 (2024).
Heil, L. R. et al. Evaluating the performance of the Astral mass analyzer for quantitative proteomics using data-independent acquisition. J. Proteome Res. 22, 3290–3300 (2023).
Petrosius, V. et al. Evaluating the capabilities of the Astral mass analyzer for single-cell proteomics. Preprint at bioRxiv https://doi.org/10.1101/2023.06.06.543943 (2023).
Guzman, U. H. et al. Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition. Nat. Biotechnol. 42, 1855–1866 (2024).
Rodriguez-Collazo, P., Leuba, S. H. & Zlatanova, J. Robust methods for purification of histones from cultured mammalian cells with the preservation of their native modifications. Nucleic Acids Res. 37, e81 (2009).
Matzinger, M., Müller, E., Dürnberger, G., Pichler, P. & Mechtler, K. Robust and easy-to-use one-pot workflow for label-free single-cell proteomics. Anal. Chem. 95, 4435–4445 (2023).
Ctortecka, C. et al. An automated nanowell-array workflow for quantitative multiplexed single-cell proteomics sample preparation at high sensitivity. Mol. Cell. Proteom. 22, 100665 (2023).
Tsai, C.-F. et al. Surfactant-assisted one-pot sample preparation for label-free single-cell proteomics. Commun. Biol. 4, 265 (2021).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Guo, Q., Sidoli, S., Garcia, B. A. & Zhao, X. Assessment of quantification precision of histone post-translational modifications by using an ion trap and down to 50 000 cells as starting material. J. Proteome Res. 17, 234–242 (2018).
Yuan, Z.-F. et al. EpiProfile 2.0: a computational platform for processing Epi-proteomics mass spectrometry data. J. Proteome Res. 17, 2533–2541 (2018).
Wiśniewski, J. R., Hein, M. Y., Cox, J. & Mann, M. A. ‘Proteomic ruler’ for protein copy number and concentration estimation without spike-in standards. Mol. Cell. Proteom. 13, 3497–3506 (2014).
Schachner, L. F. et al. Decoding the protein composition of whole nucleosomes with Nuc-MS. Nat. Methods 18, 303–308 (2021).
Sidoli, S., Lin, S., Karch, K. R. & Garcia, B. A. Bottom-up and middle-down proteomics have comparable accuracies in defining histone post-translational modification relative abundance and stoichiometry. Anal. Chem. 87, 3129–3133 (2015).
Sidoli, S., Simithy, J., Karch, K. R., Kulej, K. & Garcia, B. A. Low resolution data-independent acquisition in an LTQ-orbitrap allows for simplified and fully untargeted analysis of histone modifications. Anal. Chem. 87, 11448–11454 (2015).
Javasky, E. et al. Study of mitotic chromatin supports a model of bookmarking by histone modifications and reveals nucleosome deposition patterns. Genome Res. 28, 1455–1466 (2018).
Xue, S. & Rao, P. N. Sodium butyrate blocks HeLa cells preferentially in early G1 phase of the cell cycle. J. Cell Sci. 51, 163–171 (1981).
Zhang, K. et al. Butyrate inhibits the mitochondrial complex Ι to mediate mitochondria-dependent apoptosis of cervical cancer cells. BMC Complement. Med. Ther. 23, 212 (2023).
Padeken, J., Methot, S. P. & Gasser, S. M. Establishment of H3K9-methylated heterochromatin and its functions in tissue differentiation and maintenance. Nat. Rev. Mol. Cell Biol. 23, 623–640 (2022).
Price, A. J., Manjegowda, M. C., Kain, J., Anandh, S. & Bochkis, I. M. Hdac3, Setdb1, and Kap1 mark H3K9me3/H3K14ac bivalent regions in young and aged liver. Aging Cell 19, e13092 (2020).
Sidoli, S. et al. One minute analysis of 200 histone posttranslational modifications by direct injection mass spectrometry. Genome Res. 29, 978–987 (2019).
Sidoli, S. et al. A mass spectrometry-based assay using metabolic labeling to rapidly monitor chromatin accessibility of modified histone proteins. Sci. Rep. 9, 13613 (2019).
Hayashi, T., Daitoku, H., Uetake, T., Kako, K. & Fukamizu, A. Histidine Nτ-methylation identified as a new posttranslational modification in histone H2A at His-82 and H3 at His-39. J. Biol. Chem. 299, 105131 (2023).
Henry, R. A., Kuo, Y.-M. & Andrews, A. J. Differences in specificity and selectivity between CBP and p300 acetylation of histone H3 and H3/H4. Biochemistry 52, 5746–5759 (2013).
Klein, B. J. et al. Histone H3K23-specific acetylation by MORF is coupled to H3K14 acylation. Nat. Commun. 10, 4724 (2019).
Orsburn, B. C. Single cell proteomics by mass spectrometry reveals deep epigenetic insight and new targets of a class specific histone deacetylase inhibitor. Preprint at bioRxiv https://doi.org/10.1101/2024.01.05.574437 (2024).
Mun, D.-G. et al. Diversity of post-translational modifications and cell signaling revealed by single cell and single organelle mass spectrometry. Commun. Biol. 7, 884 (2024).
Maile, T. M. et al. Mass spectrometric quantification of histone post-translational modifications by a hybrid chemical labeling method. Mol. Cell. Proteom. 14, 1148–1158 (2015).
Daled, S. et al. Histone sample preparation for bottom-up mass spectrometry: a roadmap to informed decisions. Proteomes 9, 17 (2021).
Derks, J. et al. Increasing the throughput of sensitive proteomics by plexDIA. Nat. Biotechnol. 41, 50–59 (2023).
Hayashi-Takanaka, Y. et al. Histone modification dynamics as revealed by multicolor immunofluorescence-based single-cell analysis. J. Cell Sci. 133, jcs243444 (2020).
Zeng, W.-F. et al. AlphaPeptDeep: a modular deep learning framework to predict peptide properties for proteomics. Nat. Commun. 13, 7238 (2022).
Declercq, A. et al. TIMS 2 Rescore: a DDA-PASEF optimized data-driven rescoring pipeline based on MS 2 Rescore. Bioinformatics https://doi.org/10.1101/2024.05.29.596400 (2024).
Gessulat, S. et al. Prosit: proteome-wide prediction of peptide tandem mass spectra by deep learning. Nat. Methods 16, 509–518 (2019).
Fuller, C. N. et al. Bottom-up histone post-translational modification analysis using liquid chromatography, trapped ion mobility spectrometry, and tandem mass spectrometry. J. Proteome Res. 23, 3867–3876 (2024).
Searfoss, R. M., Karki, R., Lin, Z., Robison, F. & Garcia, B. A. An optimized and high-throughput method for histone propionylation and data-independent acquisition analysis for the identification and quantification of histone post-translational modifications. J. Am. Soc. Mass Spectrom. 34, 2508–2517 (2023).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Keenan, A. B. et al. The library of integrated network-based cellular signatures NIH program: system-level cataloging of human cells response to perturbations. 6, Cell Syst. 6, 13–24 (2018).
Blighe, K. PCAtools: everything principal components analysis. Github https://github.com/kevinblighe/PCAtools (2021).
Gu, Z. Complex heatmap visualization. Imeta 1, e43 (2022).
Taiyun. Corrplot: a visual exploratory tool on correlation matrix. Github https://github.com/taiyun/corrplot (2025).
Cutler, R. Mass spectrometry-based profiling of single-cell histone post-translational modifications to dissect chromatin heterogeneity. figshare https://doi.org/10.6084/m9.figshare.30403201.v1 (2025).
Cutler, R. Mass spectrometry-based profiling of single-cell histone post-translational modifications to dissect chromatin heterogeneity. zenodo https://doi.org/10.5281/zenodo.17282239 (2025).
Acknowledgements
We thank all members of the Sidoli and Vijg laboratories for helpful discussions. C.C. is a recipient of a SPARC Award from the Broad Institute of MIT & Harvard (#800444) that partially supported this work. L.C. acknowledges the Research Foundation Flanders – FWO for personal funding (1SF2622N) and for awarding a mobility grant to join the Sidoli Lab (V400623N). This work has been supported in part by grants: T32AG023475 to R.C.; U19AG056278, U01HL145560, U01ES029519, P01AG017242, P01AG047200, RF1AG068908, P30AG038072, US Department of Defense (BC180689P1) to J.V. This work was also supported in part by grants P01CA206978 to S.A.C from the NIH, U24CA270823, U01CA271402 to S.A.C., as well as a grant from the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation. This work was also supported by the Michael J. Fox Foundation and the Paul F. Glenn Center for the Biology of Human Aging at Albert Einstein College of Medicine. The Sidoli lab gratefully acknowledges funding from the Hevolution Foundation, the Einstein-Mount Sinai Diabetes Center, Merck, Relay Therapeutics, and the NIH Office of the Director (S10OD030286). We acknowledge Dr. Bernice Morrow and Dr. Jidong Shan in the Molecular Cytogenetics Core at Albert Einstein College of Medicine for maintaining the cellenONE instrument. Flow cytometry studies were carried out using resources of the FACS Core Facility of the Einstein Cancer Center, which is supported by NIH/NCI Cancer Center Service Grant P30 CA13330 and by Shared Instrumentation grants S100D026833 and S100D032169.
Author information
Authors and Affiliations
Contributions
R.C. and S.S. conceptualized the study. R.C., C.C., M.P., and S.S. designed the experiments. R.C., L.C., and J.C. designed the sample preparation method and prepared the injections. C.C., A.S.V.J., and M.P. designed and optimized MS acquisition methods. C.C. acquired the injections. F.L. performed the flow cytometry analysis. R.C., L.C., and C.C. analyzed the data and wrote the manuscript. D.D., J.V., S.S., and S.A.C. acquired funding. M.D., D.D., J.V., M.P., S.A.C., and S.S. supervised the research and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.A.C. is a member of the scientific advisory boards of Kymera, PTM BioLabs, Seer, and PrognomIQ. J.V. is a co-founder of SingulOmics Corp. and MutagenTech Inc. J.C. is an employee of Cellenion. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jeroen Krijgsveld and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cutler, R., Corveleyn, L., Ctortecka, C. et al. Mass spectrometry-based profiling of single-cell histone post-translational modifications to dissect chromatin heterogeneity. Nat Commun 16, 11100 (2025). https://doi.org/10.1038/s41467-025-66031-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66031-0