Abstract
ADP-ribosylation is a highly dynamic and fully reversible post-translational modification performed by PARP enzymes that modulates protein function, abundance, localization, and turnover. Here we show that PARPs mount an antiviral response to influenza A virus infection causing a rapid and dramatic upregulation of global ADP-ribosylation that inhibits viral replication. Mass spectrometry analyzes define the global ADP-ribosylome during infection, creating an infection-specific profile with almost 4000 modification sites on ~1000 host proteins, as well as over 100 modification sites on viral proteins. Our data suggest that the global increase reflects a change in the form of ADP-ribosylation rather than modification of new targets. Functional assays demonstrate that modification of the viral replication machinery antagonizes its activity. We further show that the influenza A virus protein NS1 counteracts the anti-viral activity of PARPs and ADP-ribosylation, assigning a new activity to the primary viral antagonist of innate immunity. We identify PARP1 as the enzyme producing the majority of poly(ADP-ribose) present during infection. Influenza A virus replicates faster in cells lacking PARP1, linking PARP1 and ADP-ribosylation to the anti-viral phenotype. Together, these data establish ADP-ribosylation as an anti-viral innate immune-like response to viral infection antagonized by a previously unknown activity of NS1.
Similar content being viewed by others
Introduction
Diverse cellular stresses, including infection, reprogram the post-translational modifications of proteins, rapidly altering existing cellular landscapes to promote survival of the host. These modifications dynamically modulate target protein function, abundance, turnover, and localization, playing important roles in immune activation during infection1,2,3,4. Multiple lines of evidence have recently implicated ADP-ribosylation, along with ADP-ribosyltransferases (commonly known as PARPs5), as an important part of this anti-pathogen response6,7,8,9,10. Many PARP genes are induced by interferon (IFN) signaling, suggesting a role in the response to pathogens11,12,13,14,15. Moreover, multiple PARP genes show signs of strong positive selection and other hallmarks of evolutionary conflicts at the virus:host interface16,17. Successful viruses, therefore, need to antagonize, evade, or co-opt PARP-mediated cellular countermeasures to ensure their fitness. Consequently, multiple positive-sense RNA viruses are known to encode enzymes that remove ADP-ribose (ADPr) from proteins6,8. PARPs and ADP-ribosylation are an under-appreciated arm of cellular anti-viral responses.
PARPs affect diverse biological functions, including DNA repair, gene regulation, and modulation of immune responses18. Of the 17 members of the human PARP family, up to 15 are catalytically active and transfer ADPr moieties from nicotinamide adenine dinucleotide onto target proteins or nucleic acids19,20. The extent and form of ADP-ribosylation can exert differential effects on the modified protein18,21,22. PARPs can attach a single ADPr to a target protein [mono(ADP-ribosyl)ation or MARylation] or create multimeric chains of ADPr [poly(ADP-ribosyl)ation or PARylation]. ADP-ribosylation is dynamic and reversible. PARPs add ADPr to targets while ADP-ribosylhydrolases reduce PAR chain lengths or completely remove ADPr23,24,25.
Influenza is a recurring global threat to animals, including humans. Viral replication is entirely dependent upon the infected cell, with influenza viruses usurping host factors to support replication while avoiding or evading anti-viral factors. Here, we show that influenza A virus infection causes a dramatic upregulation of PARylation that inhibits viral replication. We define the global ADP-ribosylome during influenza A virus infection, mapping thousands of modifications on viral and host proteins at single amino acid resolution. Infection induces a significant increase in PARylation to already modified proteins. Modifications on the viral replication machinery disable its activity, whereas the anti-viral activity of ADP-ribosylation is suppressed by the influenza A virus protein NS1. PARP1 is identified as the primary enzyme responsible for infection-induced PARylation and its anti-viral activity. Thus, ADP-ribosylation tempers viral replication as a key component of cellular anti-viral responses to influenza A virus, a process that is counteracted by the viral NS1 protein.
Results
Influenza A virus infection triggers PARylation with modification of viral proteins
We had previously performed a gene correlation analysis that linked expression of cellular genes to susceptibility to infection26. Among the putative anti-viral factors with the strongest negative correlation scores, we identified a PARP family member clustering with the interferon-induced transmembrane proteins (IFITMs) 1, 2, and 3, known inhibitors of several RNA viruses26,27. We therefore investigated whether influenza virus infections triggered an ADP-ribosylation response. Basal levels of ADP-ribosylation were detected in mock-infected lung cells, whereas infection with influenza A virus increased total ADP-ribosylation, as stained by a pan-ADPr reagent (Fig. 1A). The increase is typified by the appearance of discrete ADP-ribosylated proteins on blots as well as higher-molecular weight smears. As a positive control, lung cells were also treated with hydrogen peroxide (H2O2), a known inducer of ADP-ribosylation, resulting in robust detection of ADPr. Blotting was repeated using reagents specific for MARylation or PARylation, revealing that infection increases PARylation whereas MARylation was largely unchanged, if not slightly reduced.
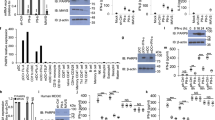
A Human lung A549 cells were infected with WSN, mock-infected, or treated with hydrogen peroxide (H2O2; positive control). Cell lysates were blotted with reagents specific for total ADP-ribosylation (PAN), MARylation (MAR), or PARylation (PAR) modifications or the indicated proteins. * = unknown protein in infected cells migrating at ~55 kDa, the molecular weight of influenza A virus NP. Lysates were probed for NP as a marker of infection and β-actin as a loading control. B PARylation blots of lysates from A549 cells infected or mock-infected with WSN with or without the PARG inhibitor PDD 00017273. Lysates were probed for NP as a marker of infection and β-actin as a loading control. C Chicken UMNSAH/DF-1 fibroblasts, pig kidney epithelial PK(15) cells, and Brazilian free-tailed bat lung epithelial Tb 1 Lu cells were mock-infected or infected with WSN or a bat-adapted WSN for Tb 1 Lu cells. PARylation and tubulin were detected by blotting. D A549 cells were inoculated with infectious or UV-inactivated virus, or mock-treated. PARylation was detected in whole cell lysates by blotting. E A549 cells were transfected with the indicated amounts of viral genomic RNA (vRNA), poly(I:C), or transfected without nucleic acid (“empty”). Cells were also subjected to interferon β (IFN-β)-, mock-, or H2O2-treatment as controls. Total ADP-ribosylation, viral NP and PA, and β-actin were detected by blotting. F Lysates from infected or mock-treated A549 cells were subject to immunoprecipitation with anti-NP of IgG control antibodies. PARylation was detected by blotting whole cell lysate (left) and immunoprecipitates (right). * = IgG heavy chain. Migration positions of molecular weight markers (kDa) are indicated. See also Supplementary Fig. 1. Source data are provided as a Source Data file.
ADP-ribosylation is highly dynamic and reversible through the activity of various enzymes such as poly(ADP-ribose) glycohydrolase (PARG)25. Therefore, we repeated infections in the presence of the PARG inhibitor PDD 00017273. PDD treatment increased the PARylation signal from infected cells compared to untreated infected cells, whereas ADP-ribosylation was largely unchanged in all conditions for mock-infected cells (Fig. 1B). These data confirm that infection causes a marked induction of PARylation. PDD treatment was included in subsequent experiments to facilitate detection of PARylated proteins. Infection with influenza virus also upregulated ADP-ribosylation in cells derived from chickens, pigs, and bats (Fig. 1C), all of which are natural hosts of influenza A virus28. Next, we exposed cells to infectious virus, or the same dose of UV-inactivated particles. Only cells exposed to replication-competent virus showed an increase in ADP-ribosylation (Fig. 1D). Finally, we asked whether viral RNA alone or innate immune activation is sufficient to induce ADP-ribosylation. Genomic RNA was isolated from influenza virions and transfected into A549 cells. Cells were also treated with poly(I:C), a dsRNA mimetic that stimulates innate immune responses, and with interferon. We confirmed that viral RNA and poly(I:C) triggered immune sensing, indicating successful introduction of these nucleic acids (Supplementary Fig. 1A, B). Nonetheless, while infection again induced PARylation but not MARylation, the innate-immune stimulators did not (Fig. 1E and Supplementary Fig. 1C). Thus, global PARylation changes are a common response to replication of influenza A virus.
To specifically test whether infection-induced ADP-ribosylation results in modification of viral proteins, we immunopurified NP from infected cells (capturing NP, as well as NP assembled into RNPs with the viral polymerase). Blotting of NP purified from infected cells revealed robust ADP-ribosylation indicative of PARylation, which was absent in control conditions (Fig. 1F). In an orthogonal approach, we first enriched ADP-ribosylated proteins from cell lysates using the recombinant macrodomain protein Af1521, which has high affinity for ADPr, and then probed for NP. NP was selectively captured by Af1521, but not the ADPr-binding mutant Af1521 G42E (Supplementary Fig. 1D). To further confirm the specificity of blotting and ensure detection of bona fide ADP-ribosylation events, we immuno-purified RNPs from infected cells and split samples into parallel blots. For the control membrane, western blots revealed ADP-ribosylation consistent with modifications to NP and the viral polymerase (Supplementary Fig. 1E, left). The other membrane was treated with hydroxylamine (HAM) (Supplementary Fig. 1E, right), which releases ADPr conjugates, primarily from modified aspartates, glutamates, and arginines29. HAM treatment markedly reduced detection of ADP-ribosylation, confirming that viral proteins are direct targets of PARP activity. HAM treatment did not completely eliminate the signal; fainter signal was still detected at the expected molecular weight of NP, suggesting that it may be modified at residues resistant to HAM treatment (e.g., histidines, cysteines, serines, or threonines). Together, these data show that influenza A virus infection induces a global upregulation of PARylation, including the modification of viral proteins.
Global identification of ADP-ribosylation sites during infection includes functionally important modification of viral proteins
Despite the broad impact of ADP-ribosylation on viral infections7, global analyzes of ADP-ribosylated proteins have largely been studied in the context of DNA damage and oxidative stress30,31. Therefore, we evaluated how influenza A virus infection alters the ADP-ribosylated proteome using enzymatic labeling of terminal ADPr coupled to mass spectrometry (ELTA-MS)31,32. ELTA selectively modifies ADP-ribosylated peptides with “clickable” N-azido-labeled tags, thereby enabling highly selective enrichment followed by MS for site identification. We defined the ADP-ribosylome in two biological replicates of A549 lung cells infected with the WSN strain of influenza A virus, mock-infected cells, or cells treated with H2O2 (positive control). Each sample was analyzed in technical triplicate. A peptide counted as ADP-ribosylated when it was detected in at least two technical replicates from the same biological replicate, one of which was by direct identification via tandem mass spectrometry (MS/MS). For increased stringency, we only considered modified peptides with site localization greater than 0.9. These results demonstrate high reproducibility across both technical and biological replicates, along with consistent identification of modification sites across ELTA-MS experiments at a range of amino acids, some of which were further validated through manual inspection (Supplementary Figs. 2 and 3 and Supplementary Data 1). For each condition, even with very strict cutoffs, we identified between ~850 and 1300 ADP-ribosylation sites on 500–700 proteins (Fig. 2A). When only considering those present in both biological replicates, we still identify ~800–1100 sites in ~440–530 proteins (Fig. 2B and Supplementary Fig. 2A). Interestingly, while infection produces a distinct ADP-ribosylation profile in A549 cells, there was a high degree of overlap among experimental conditions (i.e., mock, infected and H2O2-treated). Over half of the modified proteins, and almost three-quarters of all modified sites, were shared across all three conditions with a similar distribution of amino acid residues that were modified (Fig. 2B, C). This collection of shared proteins was highly enriched in gene ontology (GO) terms for protein translation (Supplementary Fig. 3B).
Human lung A549 cells were infected with WSN, mock infected, or treated with H2O2. ADP-ribosylation sites were identified by ELTA-MS. A Heat map clustering of unique ADP-ribosylation sites of human and viral proteins in two biological replicates. B Venn diagrams of modified proteins (left) and sites (right) identified in (A). C Distribution of ADP-ribosylation detected on the specified amino acid residues of human (left) or viral (right) proteins. Note that there were no modified cysteines in viral proteins. D Heatmap of the number of ADP-ribosylation sites identified on viral proteins during infection. E ADP-ribosylation sites mapped on to a concatenated influenza A virus proteome. Sites were mapped for biological replicate 1 on top and replicate 2 below. Sites identified in both biological replicates are marked in orange, whereas those unique to a replicate are in blue. F ELTA labeling of total ADPr isolated from A549 cells infected with WSN, mock infected, or treated with H2O2. ADPr was labeled with 32P-dATP by OAS1, separated by 15% urea-PAGE, and detected by phosphorimaging. * = non-specific band from 32P-dATP stock. G Ablation of ADP-ribosylation sites alters NP function. Polymerase activity assays performed in 293T cells with WT NP or ADP-ribosylation mutants. Expression of NP and mutants was confirmed by immunoblotting, along with β-actin as a loading control. For each condition, ELTA-MS was performed in technical triplicate on two independent biological replicates. Data in G are mean of n = 3 ± SD. *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001 as determined by a one-way ANOVA with Dunnett’s multiple comparisons test when compared to WT. See also Supplementary Figs. 2–4 and Supplementary Data 1. Source data are provided as a Source Data file.
We also identified ADP-ribosylation sites on eight of the ten canonical influenza A virus proteins (Fig. 2D, E). The distribution of amino acids modified in viral proteins was similar to that of host proteins, with the exception that there were no modified cysteines detected in viral proteins (Fig. 2C). NP repeatedly had the highest number of modifications, consistent with our biochemical approaches that detected ADP-ribosylated NP (Fig. 1F and Supplementary Fig. 1D). It is unclear whether this is because NP is the most frequently modified viral protein, or because it is one of the most abundant proteins in infected cells. Together, these findings suggest the presence of a core ADP-ribosylome in A549 cells and show that influenza A virus induces a unique profile of ADP-ribosylated proteins and modification sites, including viral proteins.
Although blotting revealed significant increases in PARylation, ELTA-MS showed that the total number of modified proteins or sites does not change between mock and infected cells (Fig. 1A–C versus Fig. 2A, B). ELTA-MS uses a phosphodiesterase to cleave the pyrophosphate bond in ADPr to leave a single phosphoribosyl moiety that enables an unambiguous identification of ADP-ribosylation sites31. However, this cleavage eliminates the length information originally associated with the modified site (e.g., MAR versus PAR). To directly detect changes in polymer length, we used ELTA to label ADPr isolated from cells that were infected, mock-infected, or exposed to H2O2 as a positive control. Infection dramatically increased PAR chain length, whereas PARylation was below the level of detection on mock-infected samples (Fig. 2F). H2O2 treatment induced higher levels of PARylation consistent with our blotting. These results help resolve the apparent contradiction between our blotting and MS data and suggest that infection does not alter which sites are modified, but instead primarily changes the ADP-ribosylation form from MAR to PAR or increases the frequency of modification at a particular site.
A panel of 18 NP variants representing high-confidence modification sites was used to determine the functional effects of ADP-ribosylation (Supplementary Data 1). ADP-ribosylation sites were found in the head and body domains of NP, the tail loop that mediates NP oligomerization, and flanking the positively-charged RNA binding groove (Supplementary Fig. 4A, B). ADP-ribosylation sites were ablated by changing the modified residues to alanine, and NP function was assessed in a polymerase activity assay. Polymerase activity increased for several of the NP ADPr-site variants, with the most pronounced effects observed for variants D34A, S69A, and E294A (Fig. 2G). In some cases like NP T92A or E107A, the variants supported activity approaching that in control conditions with wild-type NP even though the amino acid substitutions appeared to affect the overall stability or expression of NP, perhaps masking a net increase in NP activity (Fig. 2G). These data suggest that ADP-ribosylation at certain sites on NP confers an anti-viral phenotype. Insights from deep mutational scanning of NP revealed selective pressure favoring substitution at residue 294 that cannot undergo ADP-ribosylation, reinforcing that ADP-ribosylation may attenuate NP function (Fig. 2G)33. Curiously, the E252A variant resulted in a complete loss of polymerase activity (Fig. 2G), in agreement with prior work33.
Influenza A virus NS1 suppresses PARylation
Our data indicated that ADP-ribosylation is part of the anti-viral response to influenza A virus infection. Influenza virus NS1 is the primary viral protein that antagonizes host anti-viral responses and was previously shown to reduce ADP-ribosylation of argonaute RISC catalytic component 2 (AGO2) complexes34,35. Therefore, we investigated the impact of NS1 on global ADP-ribosylation by infecting cells with two closely related strains of influenza A virus encoding NS1 proteins of different potency: WSN, which was used in experiments described above, and A/Puerto Rico/8/1934 (H1N1; PR8). Both strains induced ADP-ribosylation, although PARylation was lower in cells infected with PR8 compared to WSN, despite similar infection levels (Fig. 3A). PARylation was transiently induced regardless of the strain, as indicated by the ability of PARG inhibitor to preserve the signal. PARP1, in addition to functioning as an ADP-ribosyltransferase, is a target for caspases and its cleavage is frequently used as a marker for the activation of apoptosis36,37. Influenza virus infection triggers apoptosis, which plays dual pro- and anti-viral roles during infection, and the regulation of apoptosis by influenza A virus is important for successful replication38,39,40. PARP1 cleavage was more pronounced in cells infected with WSN compared to PR8 (Fig. 3A). Curiously, WSN-infected cells had higher levels of PARylation despite also exhibiting higher levels of PARP1 cleavage.
A Strain-specific differences in ADP-ribosylation. Human lung A549 cells were infected with WSN or PR8, or mock infected, and treated with the PARG inhibitor PDD 00017273 where indicated. Proteins and ADP-ribosylation were detected by blotting. Lysates were probed for NP as a marker of infection and β-actin as a loading control. B NS1 counters ADP-ribosylation. ADP-ribosylation was detected in whole cell lysates from A549 cells infected with WSN or WSN lacking NS1 (WSN∆NS1). Lysates were probed for NP as a marker of infection. C ELTA labeling of total ADPr isolated from A549 cells infected with WSN, WSN∆NS1, mock infected, or treated with H2O2. ADPr was labeled with 32P-dATP by OAS1, separated by 15% urea-PAGE, and detected by phosphorimaging. * = non-specific band from 32P-dATP stock. D Influenza virus NS1 protein suppresses PARylation of viral proteins. Influenza A virus RNPs were assembled in 293T cells in the presence or absence of NS1. RNPs were immunopurified with anti-NP antibody and probed for PARylation or NP. Migration positions of molecular weight markers (kDa) are indicated. Source data are provided as a Source Data file.
Next, we tested a direct role of NS1 in regulating PARylation by infecting cells with a virus that lacks the NS1 gene (PR8∆NS1). PARylation was dramatically increased in cells infected with PR8∆NS1 compared to PR8, even though PR8∆NS1 virus is highly attenuated as revealed by lower levels of NP (Fig. 3B). As before, ELTA labeling revealed that infection caused a dramatic increase in ADPr polymer length and the amount of ADPr polymers was even more pronounced in cells infected with ∆NS1 virus (Fig. 3C). NS1 expression also reduced viral protein PARylation. Influenza A virus RNPs were assembled by expression in cells in the presence or absence of NS1. RNPs were immunopurified with antibodies against NP and probed for PARylation, revealing much less PARylation of viral proteins when NS1 was present (Fig. 3D). These data show that NS1 antagonizes global PARylation during infection while also reducing the PARylation intensity associated with viral proteins.
To further investigate the kinetics of PARylation and the associated role of NS1, we performed single-cycle infections in A549 cells with PR8 or PR8∆NS1. For PR8, a slight increase in PARylation was first detectable at 8 hpi, i.e., in a later stage of infection (Fig. 4A). PARylation levels then continued to increase over the time course. In contrast, PARylation was apparent as early as 4 hpi in cells inoculated with PR8∆NS1. As before, PARylation continued to increase over the time course, but at much higher levels. The increase in PARylation with PR8∆NS1 was associated with higher levels of PARP1 cleavage (Fig. 4A). This is consistent with the ability of NS1 to downregulate apoptosis41. NP is also a target for caspase cleavage, resulting in the removal of ~16 amino acids from the N-terminus42. Blotting revealed earlier NP cleavage in PR8∆NS1-infected cells compared to PR8-infected cells, again linking increased PARylation with apoptotic activity (Fig. 4A).
A NS1 suppresses infection-induced ADP-ribosylation throughout infection. Proteins and PARylation were detected by blotting lysates from human lung A549 cells infected with PR8 (MOI = 5) or PR8∆NS1 (MOI = 5) or mock infected. B Virus-specific differences in ADP-ribosylation map to NS1 and require its RNA-binding activity. A549 cells were infected with PR8∆NS1 (MOI = 4.5) and co-infected with virus encoding the indicated heterotypic NS1 or the RNA-binding mutant NS1 K38A/R41A (MOI = 0.5). Infections proceeded for 8 h prior to lysis and blotting. Migration positions of molecular weight markers (kDa) are indicated. Source data are provided as a Source Data file.
We capitalized on the robust ADP-ribosylation during PR8∆NS1 infection to test functions of NS1 that are important for controlling PARP activity. We induced PARylation by inoculating cells with PR8∆NS1 and then attempted to rescue the ∆NS1 phenotype by providing NS1 in trans by co-infection with virus encoding different NS1 proteins. Paralleling prior results, PR8 NS1 completely suppressed PARylation while WSN NS1 reduced, but did not eliminate it (Fig. 4B). We confirmed that this difference between PR8 and WSN was due solely to NS1 by swapping this gene between each virus. WSN encoding NS1PR8 (WSN NS1PR8) was now able to potently suppress PARylation, whereas PR8 encoding NS1WSN (PR8 NS1WSN) caused less suppression (Fig. 4B). One of the major mechanisms by which NS1 antagonizes host responses is through binding double-stranded RNA to prevent its sensing by innate immune activators43,44. We therefore generated virus encoding a variant that no longer binds double-stranded RNA (NS1 R38A/K41A)43. NS1 R38A/K41A lost its ability to suppress PARylation, with results indistinguishable from infection by PR8∆NS1 alone (Fig. 4B). These data reveal that NS1 counteracts the anti-viral activity of PARPs and ADP-ribosylation, possibly through its ability to bind dsRNA.
NS1 suppresses PARylation but does not reduce the number of ADP-ribosylation sites
As NS1 suppresses global PARylation, we performed ELTA-MS to compare infections in the presence or absence of NS1. In one set of experiments, we compared infections with WSN, PR8, and PR8∆NS1 (ELTA-MS #2 in Fig. 5A, C, E). In the other set, we also included WSN∆NS1 (ELTA-MS #3 in Fig. 5 B, D, F). Because viruses encoding NS1 deletions are attenuated, infection conditions were optimized for each virus to ensure equivalent expression levels of viral proteins. Each condition identified between ~1050 and 2400 modified sites on ~380–660 proteins (Fig. 5A–D and Supplementary Data 2 and 3). ADP-ribosylation sites were identified on up to eight different viral proteins, where again NP was the viral protein with the greatest number of ADP-ribosylation sites (Fig. 5E, F).
ADP-ribosylation sites were identified by ELTA-MS in two independent experiments. For ELTA-MS #2, human lung A549 cells were infected with PR8, PR8∆NS1, WSN, mock infected, or treated with H2O2. For ELTA-MS #3, infection with WSN∆NS1 was also included. Heat map clustering of unique ADP-ribosylation sites of human and viral proteins present in two biological replicates for ELTA-MS #2 (A) and ELTA-MS #3 (B). Venn diagram of the number of modified proteins (left) and sites (right) identified in ELTA-MS #2 (C) and ELTA-MS #3 (D). Heatmap of the number of ADP-ribosylation sites identified on viral proteins during infections in ELTA-MS #2 (E) and ELTA-MS #3 (F). For each condition, ELTA-MS was performed in technical triplicate on two independent biological replicates. See also Supplementary Fig. 5 and Supplementary Data 2–4. Source data are provided as a Source Data file.
Across all datasets, we detected ADP-ribosylation of proteins involved in diverse cellular functions, including consistent enrichment of proteins involved in protein binding, RNA binding, cadherin binding, and cell adhesion molecule binding (Supplementary Data 4). Infections induced a unique profile of ADP-ribosylation, while significant overlap among conditions reinforced the idea of a core ADP-ribosylome in A549 cells. Most of the modifications were shared across independent experiments, highlighting the reproducibility of ELTA-MS (Supplementary Fig. 5A). There was a high degree of overlap among modified proteins and sites for cells infected with WSN or PR8 with no notable change in the distribution of residues that were modified (Supplementary Fig. 5B, C). When considering all modified proteins identified for each condition in ELTA-MS#1-3, we again see a core ADP-ribosylome with most modified proteins are shared across conditions (Supplementary Fig. 5D). Remarkably, despite large differences in the intensity of PARylation revealed by blotting, cells infected with WT and ∆NS1 mutant virus had comparable numbers of ADP-ribosylation sites and modified proteins, with the majority of sites and proteins found in both conditions (Fig. 5C, D). This result was consistent among WSN and PR8 strains.
PARP1 directs anti-viral PARylation
Our data show that infection primarily changes PARylation. PARylation is performed by a limited subset of PARPs – PARP1, PARP2, PARP5a (TNKS) and PARP5b (TNKS2)19. ELTA-MS identified ADP-ribosylation at two canonical auto-modification sites in PARP1, E488 and E491, indicating that at least PARP1 was active in our infected cells (Supplementary Fig. 2 and Supplementary Data 2 and 3)45. To specifically test which PARPs are involved in the response to influenza virus infection, we measured PARylation in infected cells treated with a series of inhibitors possessing different specificities: PJ34 is a broad-specificity PARP inhibitor; Olaparib and Rucaparib target PARP1 and PARP2 with some off-target activity against other PARPs; AG14361 selectively inhibits PARP1; and, XAV939 primarily targets PARP5a/TNKS and PARP5b/TNKS218,46. ADP-ribosylation was unaffected by XAV939 in cells infected with either WSN or PR8, eliminating PARP5a/TNKS and PARP5b/TNKS2 as candidates (Fig. 6A). By contrast, PARylation was inhibited by Olaparib, Rucaparib, and the PARP1-specific AG14361, implicating PARP1 as the dominant enzyme mediating PARylation. Interestingly, caspase cleavage of NP still occurred under all conditions, indicating that infection-induced apoptosis does not require PARylation (Fig. 6A). Next, we purified viral RNPs from infected cells and probed them for MARylation or PARylation. Treating infected cells with Olaparib did not affect MARylation of the purified RNPs, but dramatically reduced the degree of PARylation (Fig. 6B). We generated two independent PARP1 knockout A549 cell lines to directly test its activity during infection (Fig. 6C). We infected these cells with PR8∆NS1 to create a highly sensitized setting to observe changes in ADP-ribosylation. Infection-induced PARylation was almost completely absent in both PARP1 knockout cell lines (Fig. 6D). We repeated these experiments using WT PR8 and WSN. Again, the global increase was absent in PARP1 knockouts (Fig. 6E). Together, these data show that PARP1 mediates the majority of PARylation on viral proteins as well as a global increase in the cell, whereas viral proteins are MARylated by an unknown PARP.
A Chemical inhibitors identify PARP1 as the primary PARP active during influenza A virus infection. PARylation was measured by blotting lysates from mock-treated or infected human lung A549 cells treated with the indicated PARP inhibitors or a DMSO control. B PARP1 modifies viral proteins. A549 cells were infected with WSN PB2-FLAG and treated with Olaparib or DMSO. Influenza A virus RNPs were immunopurified from infected cell lysate. Samples were probed with reagents specific for PAR or MAR. C Validation of PARP1 knockout in two clonal A549 cell lines by western blotting. β-actin was targeted as a loading control. PARP1 mediates ADP-ribosylation in IAV-infected cells. WT A549 or PARP1 knockout (KO) lines were infected with D PR8∆NS1 or E WT PR8 and WSN. ADP-ribosylation and proteins were detected by blotting whole-cell lysate. Lysates were probed for NP as a marker of infection and β-actin as a loading control. Migration positions of molecular weight markers (kDa) are indicated. Source data are provided as a Source Data file.
PARP1 both initiates ADP-ribosylation via MARylation and extends chains via PARylation47. We investigated these activities by complementing our knockout cells with PARP1, PARP1 E988A (inactive catalytic site), or PARP1 E988K (only able to MARylate)47,48. Complementation with WT PARP1 restored PARylation in cells infected with WSN or PR8 as revealed by blotting for PAR and pan-ADPr (Fig. 7A and Supplementary Fig. 6A). This result confirms that the knockout cell phenotype was due to the loss of PARP1, rather than off-target effects. Expression of the catalytically inactive PARP1 E988A did not restore PARylation, nor did expression of the MARylating PARP1 E988K change MARylation patterns, but some of this may be due to its lower enzymatic activity. Complementation with WT PARP1 resulted in an apparent minor increase in MARylation compared to the parental cells, overlapping with the high-molecular smearing in the PAR blot. Whether this represents actual MARylation by PARP1, or results from the low levels of cross-reactivity towards PAR reported for the MAR detection reagent, needs to be determined49. Curiously, complementation with a PARP1 variant D214A, which cannot undergo caspase cleavage, failed to restore PARylation in infected cells (Fig. S6B). These data indicate that PARP1 drives global PARylation during infection, but that other PARPs likely contribute the majority of MARylation.
A PARylating activity of PARP1 is required for infection-induced ADP-ribosylation. Human lung A549 PARP1 knockout cells were complemented with PARP1-V5 or the catalytically inactive PARP1 E988A or the MARylating mutant PARP1 E988K. Cells were infected or mock treated and whole cell extracts were used for blotting. Lysates were probed for NP as a marker of infection and β-actin as a loading control. B, C PARP1 suppresses influenza A virus replication. B Multi-cycle and C single-cycle replication of PR8 in parental A549 cells or A549 PARP1 knockout cells. Viral titers were determined by plaque assay for the timepoints indicated in the multicycle assay or 8 hpi for the single-cycle assay. Migration positions of molecular weight markers (kDa) are indicated. Data are mean of n = 3 ± SD. *p < 0.05 and **p < 0.01 as determined by two-sided t-test when compared to WT. See also Supplementary Fig. 6. Source data are provided as a Source Data file.
Finally, we tested the effect of PARP1 on viral replication by infecting PARP1 knockout cells. Multi-cycle replication assays were initiated in A549 and PARP1 knockout cells. Virus replicated to ~fivefold higher titers in cells lacking PARP1 during the early stages of infection (Fig. 7B). Viral titers ultimately reached the same plateau in both cell lines. In a single-cycle replication assay, PARP1 knockout cells again produced fivefold more virus than parental A549 cells (Fig. 7C). Thus, PARP1 orchestrates a global increase in PARylation during infection, establishing an anti-viral activity that slows the replication of influenza A virus.
Discussion
Viral infections trigger innate immune responses that direct anti-viral activity. Here we show that influenza A virus infection induces PARylation that antagonizes viral replication. Host and viral proteins are modified during infection, characterized by an increase in PARylation as opposed to a change in the global repertoire of ADP-ribosylated proteins. Infection appears to increase PARylation at sites that were already modified or increase the frequency at which any particular site is modified. Polymerase activity assays showed that specific modifications on the viral nucleoprotein disrupt its function, suggesting that ADP-ribosylation directly impacts the viral replication machinery. The degree of ADP-ribosylation was strain-dependent, with the differences mapping to the viral NS1 protein that suppresses PARylation. PARP1 was identified as the cellular enzyme responsible for this PARylation and conferring the anti-viral phenotype. These data show that ADP-ribosylation is an important component of the anti-viral response to influenza A virus.
Our results reveal a core ADP-ribosylome that is modified to create an infection-specific profile. Across all of our conditions, we identified 4542 unique modification sites on 1135 proteins (Supplementary Data 5). Stringent cutoffs ensured we only reported highly reproducible site identifications. Manual inspection of representative spectra showed the presence of fragment ions containing phospho-ribose that definitively identify modified residues (Supplementary Fig. 2). A potential limitation is that we may under-report modified residues as Asp-ADPr and Glu-ADPr adducts can be labile in MS workflows or during heat denaturing prior to SDS-PAGE, both of which could lead to signal loss50,51,52. ELTA-MS is also best suited for ADPr site identification, and alternative approaches will be needed to quantify the abundance of any particular modified peptide or how frequently the site in that peptide is modified. Although infection causes new modifications on a small number of proteins, most changes appear to involve PARylation at existing modification sites.
The smaller proteome of influenza A virus has enabled us to define the functional consequences of ADP-ribosylation at specific sites on viral proteins. A total of 27 ADP-ribosylation sites were identified in NP across all of our replicates, including 18 high-confidence sites that we investigated in polymerase activity assays (Figs. 2 and 5, Supplementary Figs. 4, 7, and 8, and Supplementary Data 1–3). NP binds RNA and assembles into homo-oligomers along the length of the viral genome, forming the scaffold on which viral transcription and replication take place53. NP binds RNA in a sequence-independent fashion using a large basic face to interact with the phosphate backbone54,55,56. Recent crystal structures identified specific residues in NP that contact the RNA, including S69, T92, and S36754, which we have also identified as high-confidence ADP-ribosylation sites. Mutating these residues to prevent ADP-ribosylation enhanced polymerase activity, indicating that ADP-ribosylation at these positions disrupts function, possibly by precluding binding to the viral genome. This finding also raises the intriguing possibility that the negatively charged phosphate backbone of longer ADPr chains may compete for binding not just at the residues that are modified, but across the larger positively charged RNA-binding surface of NP. Thus, at these sites, ADP-ribosylation, especially the polymeric form, may mimic nucleic acids to disrupt assembly of the replication machinery.
Different forms of ADP-ribosylation are critical for directing cellular functions. We observed a significant induction of PARylation during influenza infection that is dependent on PARylation-competent PARP1. PARP1 conferred anti-viral activity during influenza A virus infection, yet the PARP family possesses dual pro- and anti-viral functions. PARP1, PARP7, and PARP11 have each been reported to increase viral replication by reducing type I interferon production or signaling57,58,59. PARP1 also binds the influenza virus polymerase and may aid its function, although whether this activity requires ADP-ribosylation remains unclear60,61,62. Conversely, PARP7 and PARP11, along with PARP1, PARP9, PARP10, PARP12 and PARP13/ZC3HAV1 all antagonize infection6,7. PARP14 also possesses dual pro- and anti-viral activity, depending on the virus under investigation63. An anti-viral phenotype is not always dependent on catalytic activity, as PARP13/ZC3HAV1 exerts broad-spectrum anti-viral activity despite being catalytically inactive7,19. Both the long and short isoforms of PARP13/ZC3HAV1 inhibit influenza A virus replication64,65. PARP13L coordinates PARylation of the viral polymerase by an unknown PARP(s) and subsequent ubiquitin-mediated degradation64. As both PARP1 and PARP13/ZC3HAV1 interact with the viral polymerase, it is possible that the anti-viral activity of PARP1 we defined here may be contributing the ADP-ribosylation coordinated by PARP13L61,64. This contrasts with PARP13S, whose anti-viral activity appears to be independent of ADP-ribosylation65. Thus, the ultimate effect of PARPs and ADP-ribosylation during infection is likely contextual, depending on the virus, the cell, and the host.
Given the inhibitory effects of specific PARPs on viral replication, it is unsurprising that viruses have evolved mechanisms by which they combat PARPs and ADP-ribosylation. Diverse positive-sense RNA viruses including alphaviruses, coronavirids, rubella virus, and hepatitis E virus encode macrodomain proteins that remove ADP-ribosylation modifications from proteins. Viral macrodomains are necessary for successful replication6,8. Indeed, small molecule inhibitors of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) macrodomain Mac1 inhibit viral replication66. The influenza A virus genome does not encode a macrodomain, but it does encode the potent immune antagonist NS167. NS1 is not known to bind ADPr or possess ADP-ribosylhydrolase activity. Nonetheless, NS1 suppresses PARylation, with the potency of this activity varying between strains (Figs. 3 and 4)35. ELTA labeling showed similar ADPr polymer lengths in cells infected with WT or ∆NS1 virus and ELTA-MS showed that NS1 does not dramatically alter which sites are modified. However, blotting showed NS1 decreased global PARylation. Although ELTA-MS can identify modified sites, it cannot determine how frequently these sites are modified. Combined, our data suggest that NS1 functions primarily to reduce how often any particular site is PARylated, although a definitive answer requires further experimentation. Our data show that NS1 RNA-binding mutants no longer suppress ADP-ribosylation. This finding agrees with prior work using an NS1 phosphomimetic that prevents RNA binding35. As the RNA binding activity of NS1 has pleiotropic effects, it is not clear if dsRNA binding directly antagonizes ADP-ribosylation. Rather, the fact that treating cells with the dsRNA mimetic poly(I:C) does not induce ADP-ribosylation (Fig. 1E and Supplementary Fig. 1C) argues that NS1:dsRNA interactions likely play an indirect role. Influenza A virus appears to possess a unique strategy to combat PARPs and ADP-ribosylation.
The molecular signatures that activate PARPs during influenza A virus infection are unknown. The inhibitory effects of PARPs have been established in the context of RNA viruses, suggesting that PARP family members are players in the anti-viral RNA-sensing pathway, and hence could be activated indirectly by known RNA sensors, such as RIG-I, MDA5 or PKR68. However, it is unlikely that RIG-I is a major player in this process because PARylation was detected in chicken cells, which lack RIG-I (Fig. 1D)69. A more provocative hypothesis is that PARPs directly sense foreign RNAs70. PARP1 is activated by binding cellular small nucleolar RNAs (snoRNAs)71,72. While influenza genomes did not trigger PARylation, whether other certain species of viral RNAs can trigger PARP1 activation, or even host RNAs that are expressed in response to infection, remains to be determined. In summary, we established PARPs and ADP-ribosylation as new players in the battle against influenza A virus and highlight how the virus parries this attack.
Methods
Cell lines
Authenticated stocks of the following cell lines were purchased from the American Type Culture Collection: human embryonic kidney 293T cells (CRL-3216), human lung A549 cells (CCL-185), Madin-Darby canine kidney cells (MDCK; CCL-34), chicken UMNSAH/DF-1 cells (CRL-3586), pig kidney PK(15) cells (CCL-33), and Brazilian free-tailed bat lung Tb 1 Lu cells (CCL-88). The A549 innate immune reporter cell line encodes an integrated IFN-stimulated response element (ISRE) promoter expressing nanoluciferase-2A-GFP fusion protein. All cell lines were grown at 37 °C with 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% heat-inactivated fetal bovine serum. Media for the A549 ISRE reporter cells was supplemented with 5 μg/ml blasticidin. Cells were regularly tested and verified as free of Mycoplasma contamination using MycoAlert (Lonza LT07-318).
A549 PARP1 knockout cells were generated by co-transfecting the Cas9-expressing plasmid pMJ920 (Addgene 42234) and two different sgRNA-expressing plasmids targeting CCACCTCAACGTCAGGGTGC and TGGGTTCTCTGAGCTTCGGT creating a ~35 bp deletion. Cells were transfected with TransIT-X2 (Mirus MIR6005) in 6-well plates using the recommended protocols. 24 h after transfection, clonal cell lines were generated by seeding on average 0.5 cells per well in a 96-well plate, expanding clones, and confirming the loss of PARP1 by blotting.
Antibodies and ADPr detection reagents
Immunoblotting was performed with mouse α-tubulin DM1A (Sigma T9026), goat α-RNP (BEI Resources NR-3133), α-β actin (Proteintech 66009-1-Ig), and α-PARP1 (Cell Signaling 9542S). Immunoprecipitations were performed with mouse α-FLAG M2 (Sigma F1804) or α-NP H16-L10 (Bio X Cell BE0159) that was then captured on protein A agarose beads. ADP-ribosylation was detected by blotting with reagents specific for MAR (Bio-Rad AbD43647 and EMD Millipore MABE1076), PAN (EMD Millipore MABE1016) and PAR (EMD Millipore MABE1031)49,73. ADP-ribosylated proteins were enriched with GST-Af1521 that was expressed and purified from bacteria as described30. Where indicated, membranes were incubated overnight at room temperature with 1 M hydroxylamine (HAM; NH2OH) diluted in Tris-buffered saline and 0.5% Tween 20 before blotting for PAR.
Stable expression of PARP1
A549 PARP1 knockout cells were complemented using the Sleeping Beauty transposon system74. Briefly, A549 PARP1 knockout cells were co-transfected with 0.2 μg of pSB100 and 1.8 μg of pSBbiBP-PARP1-V5 or PARP1 mutant plasmids. Cells were transfected with TransIT-X2 (Mirus MIR6005) in 6-well plate using the recommended protocols. 24 h after transfection, cells were selected with to 1 μg/ml puromycin for 7 d. Expression of PARP1 or mutants was confirmed by blotting with anti-PARP1 or anti-V5-tag antibody.
Viruses and infections
Influenza A virus A/WSN/33 (H1N1; WSN), WSN encoding FLAG-tagged PB2 polymerase subunit (WSN-PB2-FLAG), and A/Puerto Rico/8/1934 (H1N1; PR8) were generated using the influenza A virus reverse genetics system75,76. PR8 that lacks NS1 (PR8ΔNS1) or encodes the RNA-binding mutant (NS1 K38A/R41A), as well as WSN that does not encode NS1 (WSN∆NS1), were propagated and titered on MDCK cells overexpressing NS1-GFP.
293T cells were infected in virus growth medium (VGM; DMEM supplemented with penicillin/streptomycin, 25 mM HEPES, 0.3% BSA). A549 cells were infected in VGM with 0.5 μg/ml TPCK-trypsin or Opti-MEM I medium with 2% FBS and 0.5 μg/ml TPCK-trypsin. For the detection of global ADP-ribosylation, A549 cells were infected at MOI 0.1, UMNSAH/DF-1 cells were infected in OptiMEM + 0.2% heat-inactivated FBS at MOI 1, and PK(15) and Tb 1 Lu cells were infected in OptiMEM + 2% FBS at MOI 1.
Multicycle replication infections were performed by inoculating cells with PR8 at MOI 0.01 in VGM with 1.0 μg/ml TPCK-trypsin and supernatants were collected from 8 to 72 hpi. Single cycle infections were performed by inoculating cells with PR8 at MOI 5 in VGM with 1 μg/ml TPCK-trypsin and progeny virus was collected at 8 hpi. Viruses were titered by plaque assay on MDCK cells.
Detection of ADP-ribosylation
Cells were mocked-treated or infected with the indicated viruses. In some experiments, the PARG inhibitor PDD 00017273 (MedChemExpress HY-108360) was added to a final concentration of 3 μM for 24 h before lysis. As a positive control, uninfected cells were exposed to 1 mM H2O2 for 10 min immediately before lysis. Cells were lysed in cold MOPS-RIPA buffer (16 mM MOPS pH 7.5, 150 mM NaCl, 1% deoxycholic acid, 0.2% SDS, 2% NP-40 with protease inhibitors, 1 μM ADP-HPD [Sigma 118415], and 40 μM PJ-34 [MilliporeSigma 528150]) or Tris-RIPA buffer (50 mM Tris pH 7.5, 150 mM NaCl, 0.5% deoxycholic acid, 0.1% SDS, 1% NP-40 with protease inhibitors and 1 μM ADP-HPD). The generic PARP inhibitor PJ-34 was included to prevent spurious ADP-ribosylation that can occur in lysates50. ADP-ribosylation was detected by blotting with purified MAR, PAN or PAR detection reagents49.
ADP-ribosylation was measured in the presence of the PARP inhibitors Olaparib (5 μM; MedChemExpress HY-10162), rucaparib (5 μM; MedChemExpress HY-10617A), AG14361 (10 μM; Sigma SML3081), XAV939 (5 μM; TOCRIS 3748), or PJ34 (4 μM; MedChemExpress HY-13688A). Stocks were prepared by dissolving inhibitors in DMSO. Cells were treated 1 h after infection.
Innate immune activation and ADP-ribosylation
To test the ability of influenza virus genomes to induce ADP-ribosylation, viral RNA was purified from virions using Trizol (Invitrogen) and transfected into A549 ISRE reporter cells using TransIT-X2 (Mirus, MIR 6005) according to the manufacturer’s instructions. As controls, cells were either mock treated, transfected without nucleic acid, transfected with high-molecular weight poly(I:C) (Invivogen tlrl-pic), incubated with 250 U/mL IFN-β, treated with 1 mM hydrogen peroxide (H₂O₂) for 10 min, or infected with WSN ΔNS1 at an MOI of 1. Cells were analyzed at 24 h post transfection or infection. ADP-ribosylation was detected by blotting as described above. ISRE activation was measured by imaging GFP fluorescence using EVOS FL Auto Imaging System (Thermo Fisher) under consistent exposure settings or by using the Nano-Glo Luciferase Reporter Assay System (Promega, N1120) according to the manufacturer’s instructions.
RNA sequencing
RNA-sequencing was previously reported77. Briefly, A549 cells were mock-infected or inoculated with WSN at an MOI of 0.02 in VGM with 0.25 μg/ml TPCK-trypsin. Infections were allowed to proceed for 24 h before RNA was harvested. In a separate condition, cells were mock-infected for 16 h followed by 8 h of interferon β (IFN-β) treatment (250 μ/ml). Each condition was repeated in biological triplicate. RNA was extracted from cells using TRIzol (Invitrogen 15596026) per manufacturer’s specifications. RNA samples were submitted to Novogene and prepared for mRNA sequencing using NEBNext Ultra RNA Library Prep Kit for Illumina (New England Biolabs E7530). Reads were trimmed and mapped to a concatenated hg19-WSN genome using Spliced Transcripts Alignment to a Reference (STAR)78. Differential expression was evaluated with DESeq279.
Polymerase activity assay
293T cells were transfected with plasmids encoding WSN PA, PB1, PB2, and NP, a viral RNA-like firefly luciferase reporter, and a Renilla luciferase internal control reporter using TransIT-2020 (Mirus MIR 5400). 18 sites in NP above or near our 0.9 localization cutoff were selected for mutagenesis. Plasmids encoding mutant versions of viral proteins were created by PCR-mediated mutagenesis and verified by sequencing. Firefly and Renilla luciferase activities were assayed ~24 h post transfection. Firefly luciferase was normalized to Renilla luciferase within each sample.
ELTA labelling of ADPr
ADPr isolation and labeling was performed following prior approaches31. For WT virus samples, A549 cells were infected with WSN at an MOI of 2 for 24 h with PDD added to 3 μM 10 h before harvest. A mock-infected control was performed in parallel. For NS1 mutant virus samples, A549 cells were infected with WSN∆NS1 at an MOI of 2 for 10 h in the presence of 3 μM PDD. For H2O2-treated samples, cells were treated with 3 μM PDD for 10 h and then 1 mM H2O2 for 10 m. Incubations were terminated by washing cells with cold PBS and lysing them in 20% (w/v) cold trichloroacetic acid for 15 m. Proteins were precipitated, washed in 70% ethanol, dried, and resuspend in 0.5 M KOH with shaking at 37 °C for 90 m to release ADPr from proteins. Sample were neutralized by the addition of MOPS to 0.8 M. Nucleic acids and proteins were digested by the addition of 25 μg DNase I (Roche 10104159001), 25 μg RNase A (Thermo EN0531) and 100 μg proteinase K (part of Roche 05080576001) with overnight incubation at 37 °C with shaking. ADPr was isolated from each samples using the High Pure miRNA Isolation Kit (Roche 05080576001) following the manufacturer’s instruction eluting samples with 100 μl distilled water. ADPr samples were labeled in 10 μl reactions with 0.5 μg of OAS1, 5 μCi α-32P-dATP (Revity BLU512H250UC), and 50 μg/ml low molecular weight poly(I:C) (Invivogen tlrl-picw) in 20 mM Tris pH 7.5, 20 mM magnesium acetate, and 2.5 mM DTT at 37 °C for 2 h. 6% of total isolated ADPr was used in each labeling reaction from infected and mock-treated samples. Given the extremely high degree of ADP-ribosylation caused by H2O2 treatment, only 1–1.5 % of the sample was used for labeling to allow for comparison on the same gel. Samples were denatured at 70 °C for 10 m in 25% urea, 12.5 mM NaCl, and 1 mM EDTA before separation on a 7 M urea 15% acrylamide gel using 0.5x TBE. Gels were analyzed by phosphorimaging.
Sample preparation for proteomic analyses
Three distinct ELTA-MS experiments were performed. In each experiment, each condition was performed in biological duplicate and samples were analyzed in technical triplicate. A549 cells were infected with WSN or PR8 at an MOI of 1 for 8 h and then treated with 1 mM H2O2 for 10 m or mock treated. In experiments with ∆NS1 viruses, infections with WSN∆NS1 or PR8∆NS1 were allowed to proceed for 16 h such that viral protein levels were roughly equivalent with that in WT-infected cells. PDD was added to the cell culture prior to the end of the incubation. Cells were then placed on ice and quickly washed with cold PBS, which was then fully removed. Cells were lysed with freshly made guanidine-hydrochloride (GdnHCl) buffer heated to 99 °C (6 M GdnHCl, 100 mM HEPES pH 8.0, 5 mM tris(2-carboxyethyl)phosphine [TCEP], 10 mM 2-chloroacetamide). Cells were scraped from the dish and transferred to an Eppendorf tube where they were incubated for an additional 10 min at 99 °C. Samples from the same biological replicate were pooled, flash frozen in liquid nitrogen, and stored at –80 °C until processing. Samples were thawed, and total protein concentration of lysates was determined by Bradford assay. Protein digestion reactions were prepared by diluting lysates containing 10 mg total protein sixfold in 100 mM Tris-HCl pH 8.0 and adding 100 µg Trypsin (Thermo Fisher 90058) and 100 µg LysC (Wako 129-02543). Samples were incubated overnight at 37 °C while shaking at 200 rpm. Next, 1 M triethylammonium acetic acid (TEAA) pH 7.5 was added to peptide solutions to a final concentration of 100 mM before clarifying peptide solutions by centrifugation for 30 min at 4 °C. 1 mg SepPak t-C18 Classic cartridges (Waters) were conditioned with 80% acetonitrile (ACN) and equilibrated with 100 mM TEAA pH 7.5 before loading peptide solutions. Peptides were washed with 100 mM TEAA pH 7.5 and eluted with 40% ACN. Samples were dried down to completion by vacuum centrifugation.
Enrichment of ADP-ribosylated peptides
ADP-ribosylated peptides were resuspended in MilliQ H2O and enriched using the ELTA-MS proteomics workflow31,32. First, peptides were incubated with 100 µM N6-(6-Azido)hexyl-dATP (Jena Bioscience CLK-NU-002), 20 µg/mL low molecular weight poly(I:C) (Invivogen tlrl-picw), and 20 µg/mL human 2´-5´-oligoadenylate synthase 1 (OAS1; prepared in-house as in ref. 31) in buffer containing 100 mM Tris-HCl pH 7.5, 20 mM magnesium acetate, and 2.5 mM dithiothreitol (DTT) for 1 h at 37 °C while shaking at 800 rpm. 100 µL of 50% dibenzocyclooctyne (DBCO)-agarose (Click Chemistry Tools 1034) equilibrated in 1× PBS was added to each OAS1 reaction before rotating end-over-end at room temperature for 1 h. The samples were centrifuged to pellet the agarose resin and the supernatant was discarded. The agarose resin was washed three times with 5 M NaCl, three times with 20% ACN, and three times with 1× PBS. After washing the resin twice in phosphodiesterase buffer containing 100 mM HEPES pH 8.0 and 15 mM MgCl2, the samples were transferred to new Eppendorf tubes. Human nudix 16 (NudT16) was prepared as described80, 5 μg was added to each sample, and the volume was brought up to 200 µL with phosphodiesterase buffer. Samples were incubated for 2 h at 37 °C while shaking at 1400 rpm. The samples were centrifuged to pellet the agarose resin and 150 µL of the eluate was transferred to new Eppendorf tubes. 10% trifluoroacetic acid (TFA) was added to samples to a final concentration of 1% TFA, and the samples were loaded onto homemade Stage-Tips that were conditioned with 80% ACN/0.1% TFA and equilibrated with 5% ACN/0.1% TFA. Peptide were washed with 0.1% TFA, eluted with 40% ACN/0.1% TFA, and dried down to completion by vacuum centrifugation.
LC-MS/MS analysis of enriched peptides
All enriched peptide samples were analyzed on a Thermo Orbitrap Fusion Lumos mass spectrometer coupled to an EASY-nLC 1200 system (Thermo) with a column type and stationary phase. LC Buffer A was 0.1% formic acid (FA), LC Buffer B was 0.1% FA/95% ACN, and the column oven was maintained at 40 °C. Peptides were separated with a 95-min gradient from 8 to 27% Buffer B, followed by an increase to 50% Buffer B over 5 min, another increase to 90% Buffer B over 5 min, and finally a 10-min washing block. MS data were collected with a spray voltage of 2.4 kV, capillary temperature of 200 °C, and RF lens of 30%. MS1 scans were performed at a resolution of 120 k, a scan range of 300 to 1800 m/z, a maximum injection time of 50 ms, and an automatic gain control (AGC) target of 1,000,000 charges. Precursor ions were isolated with a width of 1.6 m/z and an AGC target of 50,000 charges. Higher-energy collisional dissociation (HCD) fragmentation was performed with a normalized collision energy of 25% and a resolution of 30 k.
MS data analysis
Thermo.RAW files were converted to.mzML files using MSConvert from the ProteoWizard library81. Database searching was performed in FragPipe (v 20.0) with MSFragger (v 3.8), IonQuant (v 1.9.8), and Philosopher (v 5.0.0)82,83,84. The UniProt FASTA file for PR8 proteins PA, PA-X, PB1, PB1-F2, PB2, NP, HA, NA, M1, M2, NS1, and NEP/NS2 were downloaded from influenza A virus (strain A/Puerto Rico/8/1934 H1N1; Taxon ID 211044). The UniProt FASTA file for WSN proteins PA, PB1, PB2, NP, HA, NA, M1, M2, and NS1 were downloaded from influenza A virus (A/WSN/1933 H1N1; Taxon ID 382835) and WSN proteins PA-X and NEP/NS2 from influenza A virus (strain A/Wilson-Smith/1933 H1N1; Taxon ID 381518). The UniProt FASTA file for the human proteome was downloaded from UniProt Proteome ID UP000005640. The four FASTA files were manually concatenated in Python, and then decoys and contaminant proteins were added in FragPipe. A closed search in nonlabile mode was performed using the default FragPipe parameters with exceptions specified below. Fully enzymatic digestion used Trypsin and LysC rules. Variable modifications included 212.0086 (phospho-ribosylation) on DESKTYRCH (max 2 per peptide), 15.9949 (oxidation) on M (max 3 per peptide), 42.0106 (acetylation) on protein N-terminus (max 1), and –0.98401 (amidation) on protein C-terminus (max 1). The fixed modification was indicated as 57.02146 (carbamidomethylation) on C. The activation type filter was HCD and fragment ion series included b and y. Peptide-spectrum match (PSM) validation was performed with Percolator, PTM site localization was performed with PTMProphet, and protein inference was performed via ProteinProphet85,86,87. “Match between runs” (MBR) was turned on and the MBR ion false discovery rate (FDR) was set to 1%. “Normalize intensity across runs” was turned on and “peptide-protein uniqueness” was set to unique and razor peptides.
ELTA-MS uses high-energy collisional dissociation (HCD) for ADPr site localization, which can lead to mislocalization in some instances, especially on aspartate and glutamate residues88. We therefore also manually confirmed localization of representative spectra from host and viral proteins (Supplementary Fig. 2). Nonetheless, we cannot fully exclude the potential loss of Asp-ADPr and Glu-ADPr adducts under our experimental conditions that could lead to rare underreporting of modifications at these residues.
MS data filtering and visualization
All downstream analyzes of the DESKTYRCH_212.0086.tsv and combined_ion.tsv output files of the FragPipe search and plotting of the processed data were performed with homemade scripts in the statistical environment R (v. 4.3.2). Incorrectly assigned phospho-ribose sites on C were removed from the DESKTYRCH_212.0086 dataframe by comparing it to the carbamidomethylation sites on C in the combined_ion dataframe. Site indices [protein ID_amino acid_residue] were calculated for the combined_ion data frame. Peptides with one phospho-ribose were labeled as “pep_one-mod” and peptides with two phospho-riboses were labeled as “pep_multi-mod” and then two separate index rows were made for each multiply-modified peptide. The MS data (RT, m/z, charge, PSMs, etc.) were copied to each separate index row associated with the multiply-modified peptide. To ensure reproducibility across technical replicates and filter for modified peptide identifications with direct MS evidence, we developed a formula to calculate a score for each site index. MS/MS identifications were assigned an arbitrary value of 3 and MBR identifications were assigned an arbitrary value of 1. For each biological replicate, we scored all cases in which the site was identified in at least two technical replicates. The score equaled the sum of MS/MS identifications multiplied by 3 plus the sum of MBR identifications multiplied by 1. Modified peptide identifications were filtered for scores greater than or equal to 4, indicating at least one technical replicate containing direct MS/MS evidence of the modified peptide and another replicate with either matching or direct evidence. For plotting, modified peptide identifications were condensed into unique site indices. For PSMs of site indices derived from multiple modified peptide sequences, the PSMs for each modified peptide associated with that site index were summed. For PSMs of modified proteins, the PSMs for every modified peptide/site index associated with that protein were summed, excluding the ones labeled “pep_multi-mod” to avoid double counting.
For the clustering heatmaps, unique site indices were filtered for localization probabilities greater than or equal to 0.9. Then, identifications with scores greater than or equal to 4 were assigned a value of one (present) and identifications that were unmatched or had a score of less than 4 were assigned a value of zero (absent). Clustering heatmaps were generated using the pheatmap function in R (v. 4.3.2) and the Jaccard (“binary”) clustering distance for categorical data. Heatmaps showing the number of sites, max score, or number of PSMs were generated using ggplot2 in R.
For Venn diagrams, modified site indices were filtered for localization probabilities greater than or equal to 0.9, whereas modified proteins were filtered less stringently for localization probabilities greater than or equal to 0.6 in order to increase the number of modified peptide identifications, since ADPr is a labile modification during HCD fragmentation and thus can be difficult to localize. Venn diagrams were generated using InteractiVenn89.
Statistics and reproducibility
All experiments were repeated with at least three independent biological replicates, with at least three technical replicates for each experiment. Representative replicates are shown as the mean ± standard deviation. Pairwise comparisons were made using a two-tailed Student’s t-test. Multiple comparisons were performed with an ANOVA followed by an ad hoc Dunnett’s multiple comparisons test. GO enrichment was performed in gProfiler (2.0) and ShinyGO (0.8)90,91.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper as a Source Data File. RNA-seq data are available as part of BioProject PRJNA667475. Mass spectra raw files are accessible on the MassIVE database under accession code MSV000095351 [https://massive.ucsd.edu/ProteoSAFe/result.jsp?task=8964ee49fd274d3a9ca11a7129a6793c&view=advanced_view] with a ProteomeXchange ID PXD053989. Other raw data are available in the source data files and Supplementary Data 1-5. Source data are provided with this paper.
References
Liu, J., Qian, C. & Cao, X. Post-translational modification control of innate immunity. Immunity 45, 15–30 (2016).
Dawson, A. R. & Mehle, A. Flu’s cues: exploiting host post-translational modifications to direct the influenza virus replication cycle. PLoS Pathog. 14, e1007205 (2018).
Dawson, A. R., Wilson, G. M., Coon, J. J. & Mehle, A. Post-translation regulation of influenza virus replication. Annu. Rev. Virol. 7, 167–187 (2020).
Cao, X. Self-regulation and cross-regulation of pattern-recognition receptor signalling in health and disease. Nat. Rev. Immunol. 16, 35–50 (2016).
Lüscher, B. et al. ADP-ribosyltransferases, an update on function and nomenclature. FEBS J. 289, 7399–7410 (2022).
Leung, A. K. L., McPherson, R. L. & Griffin, D. E. Macrodomain ADP-ribosylhydrolase and the pathogenesis of infectious diseases. PLoS Pathog. 14, e1006864 (2018).
Fehr, A. R. et al. The impact of PARPs and ADP-ribosylation on inflammation and host–pathogen interactions. Genes Dev. 34, 341–359 (2020).
Alhammad, Y. M. O. & Fehr, A. R. The viral macrodomain counters host antiviral ADP-ribosylation. Viruses 12, 384 (2020).
Kuny, C. V. & Sullivan, C. S. Virus–host interactions and the ARTD/PARP family of enzymes. PLoS Pathog. 12, e1005453 (2016).
Lüscher, B., Verheirstraeten, M., Krieg, S. & Korn, P. Intracellular mono-ADP-ribosyltransferases at the host-virus interphase. Cell Mol. Life Sci. 79, 288 (2022).
Atasheva, S., Akhrymuk, M., Frolova, E. I. & Frolov, I. New PARP gene with an anti-alphavirus function. J. Virol. 86, 8147–8160 (2012).
Zhang, Y. et al. PARP9-DTX3L ubiquitin ligase targets host histone H2BJ and viral 3C protease to enhance interferon signaling and control viral infection. Nat. Immunol. 16, 1215–1227 (2015).
Eckei, L. et al. The conserved macrodomains of the non-structural proteins of Chikungunya virus and other pathogenic positive strand RNA viruses function as mono-ADP-ribosylhydrolases. Sci. Rep. 7, 41746 (2017).
Ryman, K. D. et al. Sindbis virus translation is inhibited by a PKR/RNase L-independent effector induced by Alpha/Beta interferon priming of dendritic cells. J. Virol. 79, 1487 (2005).
Liu, S.-Y., Sanchez, D. J., Aliyari, R., Lu, S. & Cheng, G. Systematic identification of type I and type II interferon-induced antiviral factors. Proc. Natl Acad. Sci. USA 109, 4239–4244 (2012).
Daugherty, M. D., Young, J. M., Kerns, J. A. & Malik, H. S. Rapid evolution of PARP genes suggests a broad role for ADP-ribosylation in host-virus conflicts. PLoS Genet. 10, e1004403 (2014).
Kerns, J. A., Emerman, M. & Malik, H. S. Positive selection and increased antiviral activity associated with the PARP-containing isoform of human zinc-finger antiviral protein. PLoS Genet. 4, e21 (2008).
Gupte, R., Liu, Z. & Kraus, W. L. Parps and adp-ribosylation: recent advances linking molecular functions to biological outcomes. Genes Dev. 31, 101–126 (2017).
Vyas, S. et al. Family-wide analysis of poly(ADP-ribose) polymerase activity. Nat. Commun. 5, 4426 (2014).
Weixler, L. et al. ADP-ribosylation of RNA and DNA: from in vitro characterization to in vivo function. Nucleic Acids Res. 49, 3634–3650 (2021).
Palazzo, L., Mikoč, A. & Ahel, I. ADP-ribosylation: new facets of an ancient modification. FEBS J. 284, 2932–2946 (2017).
Dasovich, M. & Leung, A. K. L. PARPs and ADP-ribosylation: deciphering the complexity with molecular tools. Mol. Cell 83, 1552–1572 (2023).
Jankevicius, G. et al. A family of macrodomain proteins reverses cellular mono-ADP-ribosylation. Nat. Struct. Mol. Biol. 20, 508–514 (2013).
Rosenthal, F. et al. Macrodomain-containing proteins are new mono-ADP-ribosylhydrolases. Nat. Struct. Mol. Biol. 20, 502–507 (2013).
Slade, D. et al. The structure and catalytic mechanism of a poly(ADP-ribose) glycohydrolase. Nature 477, 616–620 (2011).
Larson, G. P. et al. EPS8 facilitates uncoating of influenza A virus. Cell Rep. 29, 2175–2183 (2019).
Brass, A. L. et al. The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile Virus, and dengue virus. Cell 139, 1243–1254 (2009).
Long, J. S., Mistry, B., Haslam, S. M. & Barclay, W. S. Host and viral determinants of influenza A virus species specificity. Nat. Rev. Microbiol. 17, 67–81 (2019).
Cervantes-Laurean, D., Jacobson, E. L. & Jacobson, M. K. Preparation of low molecular weight model conjugates for ADP-ribose linkages to protein. in Methods in Enzymology Vol. 280 275–287 (Academic Press, 1997).
Jungmichel, S. et al. Proteome-wide identification of poly(ADP-Ribosyl)ation targets in different genotoxic stress responses. Mol. Cell 52, 272–285 (2013).
Ando, Y. et al. ELTA: enzymatic labeling of terminal ADP-ribose. Mol. Cell 73, 845–856.e5 (2019).
McPherson, R. L., Ong, S.-E. & Leung, A. K. L. Ion-pairing with triethylammonium acetate improves solid-phase extraction of ADP-ribosylated peptides. J. Proteome Res. 19, 984–990 (2020).
Bloom, J. D. An experimentally determined evolutionary model dramatically improves phylogenetic fit. Mol. Biol. Evol. 31, 1956–1978 (2014).
Klemm, C., Boergeling, Y., Ludwig, S. & Ehrhardt, C. Immunomodulatory nonstructural proteins of influenza A viruses. Trends Microbiol. 26, 624–636 (2018).
Seo, G. J. et al. Reciprocal inhibition between intracellular antiviral signaling and the RNAi machinery in mammalian cells. Cell Host Microbe 14, 435–445 (2013).
Lazebnik, Y. A., Kaufmann, S. H., Desnoyers, S., Poirier, G. G. & Earnshaw, W. C. Cleavage of poly(ADP-ribose) polymerase by a proteinase with properties like ICE. Nature 371, 346–347 (1994).
Nicholson, D. W. et al. Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature 376, 37–43 (1995).
Tran, V. et al. Influenza virus repurposes the antiviral protein IFIT2 to promote translation of viral mRNAs. Nat. Microbiol. 5, 1490–1503 (2020).
Wurzer, W. J. et al. Caspase 3 activation is essential for efficient influenza virus propagation. EMBO J. 22, 2717–2728 (2003).
Mühlbauer, D. et al. Influenza virus-induced caspase-dependent enlargement of nuclear pores promotes nuclear export of viral ribonucleoprotein complexes. J. Virol. 89, 6009–6021 (2015).
Zhirnov, O. P., Konakova, T. E., Wolff, T. & Klenk, H.-D. NS1 Protein of influenza A virus down-regulates apoptosis. J. Virol. 76, 1617–1625 (2002).
Zhirnov, O. P., Konakova, T. E., Garten, W. & Klenk, H. Caspase-dependent N-terminal cleavage of influenza virus nucleocapsid protein in infected cells. J. Virol. 73, 10158–10163 (1999).
Wang, W. et al. RNA binding by the novel helical domain of the influenza virus NS1 protein requires its dimer structure and a small number of specific basic amino acids. RNA 5, 195–205 (1999).
Lu, Y., Wambach, M., Katze, M. G. & Krug, R. M. Binding of the influenza Virus NS1 protein to double-stranded RNA inhibits the activation of the protein kinase that phosphorylates the eIF-2 translation initiation factor. Virology 214, 222–228 (1995).
Tao, Z., Gao, P. & Liu, H. Identification of the ADP-ribosylation sites in the PARP-1 automodification domain: analysis and implications. J. Am. Chem. Soc. 131, 14258–14260 (2009).
Calabrese, C. R. et al. Anticancer chemosensitization and radiosensitization by the novel poly(ADP-ribose) polymerase-1 inhibitor AG14361. J. Natl Cancer Inst. 96, 56–67 (2004).
Marsischky, G. T., Wilson, B. A. & Collier, R. J. Role of glutamic acid 988 of human poly-ADP-ribose polymerase in polymer formation: evidence for active site similarities to the ADP-ribosylating toxins (∗). J. Biol. Chem. 270, 3247–3254 (1995).
Rolli, V., O’Farrell, M., Ménissier-de Murcia, J. & de Murcia, G. Random mutagenesis of the poly(ADP-ribose) polymerase catalytic domain reveals amino acids involved in polymer branching. Biochemistry 36, 12147–12154 (1997).
Gibson, B. A., Conrad, L. B., Huang, D. & Kraus, W. L. Generation and characterization of recombinant antibody-like ADP-ribose binding proteins. Biochemistry 56, 6305–6316 (2017).
Weixler, L. et al. Protein and RNA ADP-ribosylation detection is influenced by sample preparation and reagents used. Life Science Alliance 6, e202201455 (2023).
Tashiro, K. et al. Chemoenzymatic and synthetic approaches to investigate aspartate- and glutamate-ADP-ribosylation. J. Am. Chem. Soc. 145, 14000–14009 (2023).
Longarini, E. J. & Matić, I. Preserving ester-linked modifications reveals glutamate and aspartate mono-ADP-ribosylation by PARP1 and its reversal by PARG. Nat. Commun. 15, 4239 (2024).
Zhu, Z., Fodor, E. & Keown, J. R. A structural understanding of influenza virus genome replication. Trends Microbiol. 31, 308–319 (2023).
Tang, Y.-S., Xu, S., Chen, Y.-W., Wang, J.-H. & Shaw, P.-C. Crystal structures of influenza nucleoprotein complexed with nucleic acid provide insights into the mechanism of RNA interaction. Nucleic Acids Res. 49, 4144–4154 (2021).
Ye, Q., Krug, R. M. & Tao, Y. J. The mechanism by which influenza A virus nucleoprotein forms oligomers and binds RNA. Nature 444, 1078–1082 (2006).
Ng, A. K. et al. Structure of the influenza virus A H5N1 nucleoprotein: implications for RNA binding, oligomerization, and vaccine design. FASEB J. 22, 3638–3647 (2008).
Wang, F. et al. Cytoplasmic PARP1 links the genome instability to the inhibition of antiviral immunity through PARylating cGAS. Mol. Cell 82, 2032–2049.e7 (2022).
Guo, T. et al. ADP-ribosyltransferase PARP11 modulates the interferon antiviral response by mono-ADP-ribosylating the ubiquitin E3 ligase β-TrCP. Nat. Microbiol. 4, 1872–1884 (2019).
Yamada, T. et al. Constitutive aryl hydrocarbon receptor signaling constrains type I interferon–mediated antiviral innate defense. Nat. Immunol. 17, 687–694 (2016).
Bortz, E. et al. Host- and strain-specific regulation of influenza virus polymerase activity by interacting cellular proteins. mBio 2, e00151-11 (2011).
Mayer, D. et al. Identification of cellular interaction partners of the influenza virus ribonucleoprotein complex and polymerase complex using proteomic-based approaches. J. Proteome Res. 6, 672–682 (2007).
Westera, L. et al. Poly-ADP ribosyl polymerase 1 (PARP1) regulates influenza A virus polymerase. Adv. Virol. 2019, 8512363 (2019).
Parthasarathy, S. et al. PARP14 is an interferon-induced host factor that promotes IFN production and affects the replication of multiple viruses. mBio 16, e0229925 (2025).
Liu, C. H., Zhou, L., Chen, G. & Krug, R. M. Battle between influenza A virus and a newly identified antiviral activity of the PARP-containing ZAPL protein. Proc. Natl Acad. Sci. USA 112, 14048–14053 (2015).
Tang, Q., Wang, X. & Gao, G. ZAPS inhibits influenza A viral protein expression and is antagonized by the virus encoded NS1. J. Virol. 91, e01909–e01916 (2017).
Wazir, S. et al. Discovery of 2-amide-3-methylester thiophenes that target SARS-CoV-2 Mac1 and repress coronavirus replication, validating Mac1 as an antiviral target. J. Med. Chem. 67, 6519–6536 (2024).
Krug, R. M. Functions of the influenza A virus NS1 protein in antiviral defense. Curr. Opin. Virol. 12, 1–6 (2015).
Hur, S. Double-stranded RNA sensors and modulators in innate immunity. Annu. Rev. Immunol. 37, 349–375 (2019).
Barber, M. R. W., Aldridge, J. R., Webster, R. G. & Magor, K. E. Association of RIG-I with innate immunity of ducks to influenza. Proc. Natl Acad. Sci. USA 107, 5913–5918 (2010).
Xing, J. et al. Identification of poly(ADP-ribose) polymerase 9 (PARP9) as a noncanonical sensor for RNA virus in dendritic cells. Nat. Commun. 12, 2681 (2021).
Kim, D. S. et al. Activation of PARP-1 by snoRNAs controls ribosome biogenesis and cell growth via the RNA helicase DDX21. Mol. Cell 75, 1270–1285.e14 (2019).
Huang, D., Kim, D.-S. & Kraus, W. L. Specific binding of snoRNAs to PARP-1 promotes NAD+-dependent catalytic activation. Biochemistry 59, 1559–1564 (2020).
Longarini, E. J. et al. Modular antibodies reveal DNA damage-induced mono-ADP-ribosylation as a second wave of PARP1 signaling. Mol. Cell 83, 1743–1760.e11 (2023).
Kowarz, E., Löscher, D. & Marschalek, R. Optimized Sleeping Beauty transposons rapidly generate stable transgenic cell lines. Biotechnol. J. 10, 647–653 (2015).
Neumann, G., Fujii, K., Kino, Y. & Kawaoka, Y. An improved reverse genetics system for influenza A virus generation and its implications for vaccine production. Proc. Natl Acad. Sci. USA 102, 16825–16829 (2005).
Kirui, J., Mondal, A. & Mehle, A. Ubiquitination upregulates influenza virus polymerase function. J. Virol. 90, 10906–10914 (2016).
Baker, S. F. et al. Alternative splicing liberates a cryptic cytoplasmic isoform of mitochondrial MECR that antagonizes influenza virus. PLoS Biol. 20, e3001934 (2022).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Daniels, C. M., Thirawatananond, P., Ong, S.-E., Gabelli, S. B. & Leung, A. K. L. Nudix hydrolases degrade protein-conjugated ADP-ribose. Sci. Rep. 5, 18271 (2015).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536 (2008).
Kong, A. T., Leprevost, F. V., Avtonomov, D. M., Mellacheruvu, D. & Nesvizhskii, A. I. MSFragger: ultrafast and comprehensive peptide identification in mass spectrometry–based proteomics. Nat. Methods 14, 513–520 (2017).
da Veiga Leprevost, F. et al. Philosopher: a versatile toolkit for shotgun proteomics data analysis. Nat. Methods 17, 869–870 (2020).
Yu, F., Haynes, S. E. & Nesvizhskii, A. I. IonQuant enables accurate and sensitive label-free quantification with FDR-controlled match-between-runs. Mol. Cell. Proteom. 20, 100077 (2021).
Käll, L., Canterbury, J. D., Weston, J., Noble, W. S. & MacCoss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 4, 923–925 (2007).
Shteynberg, D. D. et al. PTMProphet: fast and accurate mass modification localization for the trans-proteomic pipeline. J. Proteome Res. 18, 4262–4272 (2019).
Nesvizhskii, A. I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003).
Larsen, S. C., Hendriks, I. A., Lyon, D., Jensen, L. J. & Nielsen, M. L. Systems-wide analysis of serine ADP-ribosylation reveals widespread occurrence and site-specific overlap with phosphorylation. Cell Rep. 24, 2493–2505.e4 (2018).
Heberle, H., Meirelles, G. V., da Silva, F. R., Telles, G. P. & Minghim, R. InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 16, 169 (2015).
Kolberg, L., Raudvere, U., Kuzmin, I., Vilo, J. & Peterson, H. gprofiler2 - an R package for gene list functional enrichment analysis and namespace conversion toolset g:Profiler. F1000Res 9, ELIXIR–709 (2020).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Acknowledgements
We thank B. tenOever for reagents and Anya Crane (Integrated Research Facility at Fort Detrick/National Institute of Allergy and Infectious Diseases/National Institutes of Health, Fort Detrick, Frederick, MD, USA) for critically editing the manuscript. This work was supported by: the National Institutes of Health grants AI160779 to AM and AKLL, AI164690 to AM, AI125271 to AM, GM104135 to AKLL for the development of ELTA-MS, AI078985 to GPL, GM07215 to VT, and F31GM143918 and T32GM149382 to IU; the National Science Foundation GRFP DGE-1747503 to MPL; an H.I. Romnes Faculty Fellow funded by the Wisconsin Alumni Research Foundation and provided to AM; and a Vilas Faculty Mid-Career Investigator Award to AM. AM was a Burroughs Wellcome Fund Investigator in the Pathogenesis of Infectious Disease. We acknowledge support of the shared instrumentation grant S10OD021844 from the Center for Proteomics Discovery at the Johns Hopkins University School of Medicine. This work was supported in part through Battelle Memorial Institute’s former prime contract with the U.S. National Institute of Allergy and Infectious Diseases (NIAID) under Contract No. HHSN272200700016I and Laulima Government Solutions, LLC current prime contract with NIAID under Contract No. HHSN272201800013C. SY, YC and JHK performed this work as former employees of Battelle Memorial Institute and current employees of Tunnel Government Services (TGS), a subcontractor of Laulima Government Solutions, LLC under Contract No. HHSN272201800013C. All experiments were approved by the University of Wisconsin-Madison Institutional Biosafety Committee (IBC). The NIAID grant AI125271 that supported this work was reviewed by the University of Wisconsin-Madison Dual Use Research of Concern (DURC) Subcommittee in accordance with the United States Government September 2014 DURC Policy and determined to meet the criteria of DURC. The University of Wisconsin-Madison Institutional Contact for Dual Use Research reviewed this manuscript and confirmed that the studies described herein do not meet the criteria of DURC. The views and conclusions contained in this document are those of the authors and should not be interpreted as necessarily representing the official policies, either expressed or implied, of the U.S. Departments of the Army, Department of Defense, and Department of Health and Human Services, or of the institutions and companies affiliated with the authors, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government.
Author information
Authors and Affiliations
Contributions
Z.Z., K.A.D., G.P.L., V.T., J.H.K., A.K.L.L., and A.M. contributed to the conceptualization. Z.Z., I.U., K.A.D., R.L.M., G.P.L., P.C., A.K.L.L., and A.M. developed methodology. M.B., V.T., M.P.L., C.C., X.Y., Z.M., A.K.L.L., and A.M. wrote or executed software. Z.Z., I.U., K.A.D., M.B., E.M.F., and A.M. performed validation. Z.Z., I.U., R.L.M., M.B., V.T., M.P.L., C.C., X.Y., Z.M., J.H.K., A.K.L.L., and A.M. performed formal analysis. Z.Z., I.U., K.A.D., R.L.M., G.P.L., M.B., V.T., M.P.L., E.M.F., S.Y., Y.C., C.C., X.Y., Z.M., and A.M. did the investigation. Z.Z., I.U., and A.M. were responsible for data curation. Z.Z., I.U., A.K.L.L., and A.M. wrote the original draft, with all authors reviewing and editing. Z.Z., I.U., K.A.D., G.P.L., C.C., X.Y., Z.M., A.K.L.L., and A.M. created visualizations. J.H.K., A.K.L.L., and A.M. supervised the work. I.U., G.P.L., M.P.L., V.T., A.K.L.L., and A.M. acquired funding.
Corresponding authors
Ethics declarations
Competing interests
P.C. is an employee of ARase Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ivan Matić, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Z., Uribe, I., Davis, K.A. et al. Global remodeling of ADP-ribosylation by PARP1 suppresses influenza A virus infection. Nat Commun 16, 11176 (2025). https://doi.org/10.1038/s41467-025-66136-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66136-6