Abstract
Extracellular osmotic pressure is a key modulator of intracellular biophysical properties and cellular functions. However, its impact on the cell nucleus remains elusive, largely due to the challenges in real-time measurement of local environmental properties and reaction kinetics at specific loci within the nucleus. Here, we employ the dCas9-SunTag system to investigate the biophysical response at target DNA loci to osmotic pressure alterations. We reveal that variations in extracellular osmotic pressure modulate the efficiency of dCas9-SunTag-mediated fluorescent labelling rapidly and reversibly, with hypoosmotic condition increasing and hyperosmotic condition decreasing the number and fluorescence intensity of foci for telomeres and genes. Strikingly, osmotic pressure also regulates gene transcription with the dCas9-SunTag system, mirroring its effects on fluorescent labelling, as evidenced by changes in mRNA burst frequency. The underlying mechanism is that osmotic pressure shifts the binding-unbinding equilibrium of specific proteins to dCas9-SunTag complex by altering intranuclear crowding. These findings not only highlight the role of mechanical cues in modulating DNA-related processes within the nucleus, but also establish the dCas9-SunTag system as a sensitive probe for intranuclear crowding in response to extracellular cues, notably osmotic pressure.
Similar content being viewed by others
Introduction
Cells are continually subjected to fluctuations of extracellular environment in vivo, which significantly impact intracellular biophysical properties, thereby influencing cellular functions1,2,3,4. Among diverse extracellular environmental factors, osmotic pressure is of great importance involved in such fundamental cellular behaviours as cell migration2,5, mitosis3 and differentiation4, as well as in tissue homoeostasis and development6,7. Previous biophysical studies have revealed that the osmotic pressure impacts intracellular properties in various aspects, including the modification of plasma membrane tension as well as lipid liquid-liquid phase separations8, the rearrangement of cytoskeletal networks and their elastic mechanics9, the alternation of cell volume and morphology10, and the modulation of intracellular molecular crowding and diffusion11,12,13. While the osmotic pressure has been extensively studied for its influence on the intracellular properties, its effect on the cell nucleus remains largely unclear, which is particularly significant as the nucleus is the central location for DNA-related functions14.
Alterations of extracellular osmotic pressure are known to modulate nuclear volume15,16,17 and intranuclear structural features18,19,20, and are implicated in DNA repair21, chromosome replication22 and gene expression9,23,24,25. The mechanism of osmotic pressure impacting DNA-related functions is proposed to involve changes in chromatin architecture and nuclear compartmentalisation due to altered intranuclear molecular crowding and induced phase separation18,19,20. Additionally, osmotic pressure may modulate crowding and thereby influence molecular diffusion dynamics and reaction kinetics19,26,27,28,29, directly affecting DNA-related processes. However, understanding the precise mechanisms by which osmotic pressure affects these processes within complicated nuclear environment remains elusive, primarily due to the challenge in real-time measurement of local environmental properties and reaction kinetics at specific genomic loci.
To address this gap, a sensitive method that probes molecular interactions at specific DNA sequences is essential. The CRISPR-Cas (clustered regularly interspaced short palindromic repeats-CRISPR associated) system is renowned for its ability to target specific DNA sequences with high precision30. By integrating the SunTag31, which presents multiple binding sites for various proteins, including fluorescence proteins (FPs) and transcription activation domains (ADs), we may be able to translate the effect of intranuclear environmental factors, such as molecular crowding, into measurable fluorescent signals or transcriptional responses. This combined system holds promise as a potential tool to assess how osmotic pressure influences specific DNA sites and their associated gene functions. Yet, the precise response of this system to intranuclear environment and osmotic pressure variations has not been characterised.
Here, we have investigated the impact of extracellular osmotic pressure on the dCas9-SunTag system in live cells, particularly its capacity in DNA labelling and gene transcription. We discover that the extracellular osmotic pressure variations modulate the efficiency of dCas9-SunTag-mediated fluorescent labelling in a rapid, reversible and repeatable fashion, with hypoosmotic condition raising and hyperosmotic condition diminishing the number and fluorescence intensity of foci, for both telomeres and protein-coding genes. Our experimental research combined with theoretical analysis suggests that osmotic pressure shifts the binding-unbinding equilibrium between fluorescence proteins and dCas9-SunTag complex, by altering intranuclear local crowding. More importantly, we find that osmotic pressure also regulates gene transcription with the dCas9-SunTag system in a manner that parallels its effects on fluorescent labelling. This is further supported by observations of the changes in the mRNA burst frequency under varying osmotic conditions, consistent with our ensemble-level measurements. Our results not only underscore the crucial role of osmotic pressure in regulating the dCas9-SunTag system for DNA labelling and gene expression, and but also highlight how mechanical cues influence intranuclear biological processes. Furthermore, the dCas9-SunTag system serves as a sensitive probe for investigating intranuclear crowding at specific genomic loci, offering insights into cellular responses to complex extracellular stimuli.
Results
Osmotic pressure regulates fluorescent labelling of target DNA with dCas9-SunTag system
The dCas9-SunTag system is a versatile platform for target DNA labelling and gene expression31. As illustrated in Fig. 1a (left), the SunTag consists of an array of repetitive peptide sequence (GCN4 peptide epitope), to which the cognate GCN4 single-chain variable fragment (scFv) antibodies can be attached. The SunTag system can recruit multiple copies of a protein that is fused to the scFv antibody, e.g. fluorescence protein or transcription activation domain, to achieve signal amplification. The complex of the dCas9 protein and the SunTag (dCas9-SunTag) is directed by a single guide RNA (sgRNA) to the target DNA locus with the complementary sequence for imaging or transcription regulation.
a Schematic of the target DNA fluorescence labelling with dCas-SunTag system (left) and the response to external osmotic alterations in a live cell (right). FP, fluorescence protein. b Snapshots of the cells with telomeres labelled with dCas9-SunTag-FP (green) and H2B labelled with mCherry (red) under different osmotic conditions. From 3D orthogonal view along z axis. Scale bars, 5 μm. c Number increment (top) and total fluorescence intensity increment (bottom) of telomere foci in a cell labelled with dCas9-SunTag system versus time during the hypoosmotic and hyperosmotic treatments. n = 10. AU, arbitrary units. d Number increment, total fluorescence intensity change percentage of telomere foci labelled with dCas9-SunTag system, and nuclear volume change percentage in a cell around 10 minafter adding water or PEG. 30% H2O, n = 54; 20% H2O, n = 55; 10% H2O, n = 62; ctrl (control), n = 42; 2.5% PEG, n = 39; 5% PEG, n = 60; 10% PEG, n = 37. For c and d, the increment or the change percentage is relative to the initial case right before the osmotic shock. The same cells are analysed before and after osmotic treatment. All statistics above are presented as mean ± SE (sample size n refers to cell number). Two-sample t-test is performed with p value annotated.
We first study the effect of extracellular osmotic pressure on the telomere fluorescence labelling with the dCas9-SunTag system in live cells. A bright green fluorescence protein fused with scFv, scFv-mNeonGreen32, is expressed in cells as the binding protein to SunTag. Under confocal laser scanning microscopy, we observe fluorescent spots in cell nuclei, as expected. The identity of these spots as telomere loci is confirmed through fluorescence in situ hybridisation (FISH) assays using a Cy5-tagged telomere probe32 (Supplementary Fig. 1). To decrease or increase the extracellular osmotic pressure, we add water or 200-Da polyethylene glycol (PEG) to cell culture medium, respectively4,10,16, with the actual extracellular osmotic pressure being calculated (Supplementary Fig. 2). Interestingly, as the hypoosmotic pressure is exerted by adding 30% water, both the number and the fluorescence intensity of telomere foci in a cell increase apparently (Fig. 1b, c). In contrast, the hyperosmotic pressure by the addition of 10% PEG decreases the number and the fluorescence intensity of telomere foci significantly (Fig. 1b, c).
Our results show that the telomere labelling by the dCas9-SunTag system is changed with the extracellular osmotic pressure, yet it could be resulted from two ways: the quantity of binding proteins on dCas9-SunTag, and the quantity of dCas9-SunTag on telomere. We thus repeat experiments on the telomere labelling with the dCas9 directly fused with a fluorescence protein33 for comparison. The results show that the number and the fluorescence intensity of telomere foci in a cell almost remain constant in this case (Supplementary Fig. 3), indicating that the extracellular osmotic pressure has a negligible influence on the binding between dCas9 and telomeres, but plays a critical role in the binding of fluorescence proteins to dCas9-SunTag. It also verifies our assumption that the dCas9-SunTag system is sensitive to the change of osmotic pressure.
Given the dynamic nature of extracellular osmotic pressure, which can fluctuate and return to isotonic conditions, we seek to determine if these variations are mirrored in the fluorescent labelling of telomeres. Notably, when the osmotic pressure returns to its initial isotonic state, the telomere foci, as indicated by fluorescence intensity and number, also revert to their initial levels (Fig. 1b, c). Noted that despite a slight decrease in the number and total intensity of telomere foci due to photobleaching, the effect of osmotic pressure remains clear and significant (Supplementary Fig. 4). Encouraged by the reversible response, we proceed to investigate the cyclic stability of this response. Cells are subjected to a non-isotonic medium for 15 min, followed by a recovery period in isotonic medium for 30 min, with this treatment regimen repeated three times. Throughout these cycles, the number and the fluorescence intensity of telomere foci exhibit a consistent and reversible response (Supplementary Fig. 5). These results not only confirm the reversibility of fluorescent labelling changes induced by osmotic pressure alterations but also establish the repeatability of these responses under controlled perturbations.
To determine the sensitivity of telomere fluorescence labelling to osmotic pressure, we vary extracellular osmolarity using water or PEG of different concentrations. We observe that nuclear volume changes inversely with extracellular osmotic pressures (Fig. 1d), confirming the presence of a gradual osmotic gradient. Our statistical analysis indicates that the telomere fluorescent labelling efficiency is inversely related to external osmotic pressure within a certain range (Fig. 1d). We notice that both the number and the intensity of telomere foci labelled with dCas9-SunTag-FP vary among cells under isotonic condition, reflecting cell-to-cell heterogeneity. We confirm that such variations do not affect the effect of osmotic pressure on telomere labelling, as the results remain consistent across cells with varying initial cases (Supplementary Fig. 6).
Having established the influence of extracellular osmotic pressure at telomeres, which are highly repetitive and chromosomally distal31,32,33,34, we seek to assess whether the response of dCas9-SunTag system to extracellular osmotic pressure is consistent with other genomic regions, especially protein-coding genes that contain low-repeat sequences. We choose MUC4 gene, an epithelial mucin gene with low-repeat characteristics35. Consistent with the telomere labelling results, the MUC4 gene labelling efficiency is elevated under hypoosmotic condition and reduced under hyperosmotic condition (Supplementary Fig. 7). We further confirm that the relationship between MUC4 labelling efficiency and osmotic pressure remains reliable, unaffected by the initial fluorescence intensity of foci (Supplementary Fig. 8). These results imply that the DNA labelling with the dCas9-SunTag system in response to osmotic pressure may not be dependent on the specific DNA sequence locus, indicating a potentially universal effect across various genomic elements.
Osmotic pressure modulates binding–unbinding equilibrium of fluorescence proteins to dCas9-SunTag
Our observation that osmotic pressure can dynamically modulate the fluorescent labelling of target DNA suggests a variable interaction between binding proteins and the dCas9-SunTag. We next aim to explore the underling mechanism of this modulation. We first perform fluorescence recovery after photobleaching (FRAP) experiments focusing on the telomere foci labelled with the dCas9-SunTag system, to examine whether these interactions are indeed dynamic. After photobleaching, the telomere foci reappear within half a minute, while FRAP experiments conducted in areas lacking telomere foci show a much quicker recovery of fluorescence intensity within a few seconds (Fig. 2a, b). This disparity suggests that the binding of fluorescence proteins to dCas9-SunTag is markedly slower compared to the diffusion of unbound fluorescence proteins. Furthermore, telomere spots labelled with dCas9-FP show no recovery within the duration of the FRAP experiments (Fig. 2a, b, and Supplementary Fig. 9). Collectively, these FRAP findings suggest the highly dynamic equilibrium of fluorescence proteins binding and unbinding to dCas9-SunTag, in contrast to a relatively stable interaction between telomeres and dCas9. This implies that the interaction between fluorescence proteins and dCas9-SunTag is more sensitive to environmental perturbations.
a 2D snapshots of FRAP experiments on the area with a telomere focus labelled with dCas9-SunTag-FP (top, green) as well as on the blank area without a telomere focus (middle) in a cell, and on the area with a telomere focus labelled with dCas9-FP (bottom, red, with blue nucleus stained by Hoechst 33342). The photobleached area is marked by a white circle. b Normalised fluorescence intensity inside the photobleached area (2D) over time. Statistics for cells with n = 23 for dCas9-SunTag-FP, n = 9 for dCas9-SunTag-FP (blank) and n = 11 for dCas9-FP. c Snapshots of the fluorescence condensation of telomere spots labelled with dCas9-SunTag-FP in a cell upon hypoosmotic shock and the fluorescence dispersion upon hyperosmotic shock. From 3D orthogonal view along z axis. The region of interest marked by a white square is enlarged and shown below the image. d 2D radial distribution of normalised fluorescence intensity of a telomere focus during the condensation upon hypoosmotic shock and during the dispersion upon hyperosmotic shock. Statistics for telomere foci in the same cells shown in c with n = 14 for 30% H2O and n = 18 for 10% PEG. Apparent binding rate of fluorescence protein to dCas9-SunTag upon hypoosmotic shock (e) and apparent unbinding rate upon hyperosmotic shock (f). The apparent binding rate is obtained over 6–10 min after adding H2O into medium (n = 10 cells) while the apparent unbinding rate is calculated over 1–5 min after adding PEG (n = 7 cells). Scale bars, 5 μm. All statistics above are presented as mean ± SE.
Then, we reason that the osmotic shock could disrupt the equilibrium between the binding and the unbinding of fluorescence proteins to the dCas9-SunTag. Our hypothesis is that hypoosmotic shock favours binding over unbinding, leading to macroscopic fluorescence condensation, whereas hyperosmotic shock promotes the opposite effect, resulting in fluorescence dispersion. To explore this, we meticulously monitor the telomere foci upon osmotic shock, to directly quantify the real-time effect of osmotic pressure on the binding dynamics of fluorescence proteins to dCas9-SunTag. Upon hypoosmotic shock, existing telomere foci intensify gradually and new foci may emerge, while hyperosmotic shock leads to a dimming and potential disappearance of these telomere foci (Fig. 2c, d, and Supplementary Figs. 10, 11).
To further understand the condensation or dispersion of fluorescence proteins, we develop a physical model to describe these processes. Hypothesised for simplicity that the diffusion of fluorescence proteins and the binding or unbinding of them to dCas9-SunTag are uncoupled, the spatiotemporal evolution of fluorescence proteins is depicted as:
where r is the position vector, t is the time, cm is the concentration of mobile fluorescence protein, cf is the concentration of fixed fluorescence protein which is bound to the dCas9-SunTag on a target locus, D is the diffusion coefficient of mobile fluorescence protein, kb is the binding rate and ku is the unbinding rate. Noted that in a live cell, the movement of fluorescence proteins is enhanced by active fluctuations36. Thus, the parameter D is interpreted as the apparent diffusion coefficient which integrate the effect of active fluctuations. In an equilibrium state under isotonic condition, kbicm − kuicf = 0 (the subscript ‘i’ refers to ‘isotonic’).
Our experiment results reveal three key insights: 1) diffusion is significantly faster than binding-unbinding dynamics, 2) the fluorescence intensity of a target focus approximates a Gaussian distribution, and 3) kb = kbi + Δkb and Δkbcm ≫ ∣kbicm − kuicf∣ upon hypoosmotic shock, while ku = kui + Δku and Δkucf ≫ ∣kbicm − kuicf∣ upon hyperosmotic shock. Δkb is the apparent binding rate and Δku is the apparent unbinding rate, reflecting the effective rate of reaction as it transitions from one equilibrium state to another due to environmental changes. Based on these premises, we derive a simplified solution to Equation (1), which effectively interprets our experimental outcomes. The fluorescence intensity of a bright target focus with background subtraction is
where r is the distance from a focus centre, Δt is the relative time, I0 represents the fluorescence intensity at r = 0 and Δt = 0, R0 represents the standard deviation of the fluorescence radial distribution at Δt = 0, Im0 represents the intensity corresponding to mobile fluorescence proteins at Δt = 0, and v is the volume ratio of target spot to nucleus.
We then employ Eq. (2) to fit the experimental fluorescence distribution of foci within cells, as depicted in Supplementary Fig. 12. This approach allows us to determine the apparent binding rate Δkb during hypoosmotic shock and the apparent unbinding rate Δku during hyperosmotic shock. The apparent binding rate increases as osmotic pressure decreases (Fig. 2e), and the apparent unbinding rate correlates positively with osmotic pressure (Fig. 2f), in agreement with the observation. The close alignment between our experiment data and our theoretic model indicates that the macroscopic changes in telomere focus fluorescence are probably attributed to the microscopic perturbation of the binding-unbinding equilibrium between fluorescence proteins and dCas9-SunTag in response to osmotic shock. This suggests that osmotic pressure may modulate the availability of binding sites on the SunTag, likely by affecting intracellular molecular crowding37. Our simplified model also underscores the persistence of fundamental physical and chemical principles in the complicated nuclear environment.
Osmotic pressure alters intranuclear dynamics and crowding to modulate binding–unbinding equilibrium of fluorescence proteins to dCas9-SunTag
Next, we seek to elucidate the mechanism underlying the effect of extracellular osmotic pressure on the intranuclear binding-unbinding equilibrium. We hypothesise that changes in osmotic pressure would alter the intranuclear molecular crowding, which in turn would affect the interactions between fluorescent proteins and dCas9-SunTag. Our observations of nuclear volume alterations under various osmotic conditions has supported this (Fig. 1d), as nuclear volume is a reflection of the overall intranuclear molecular crowding, given that the amount of intranuclear macromolecules remains relatively constant over a short period of osmotic shock15,17,38,39. Yet considering the heterogeneous and complicated environment in nucleus, we need to further assess the impact of extracellular osmotic pressure on intranuclear molecular crowding.
Recognising that molecular crowding can influence chromatin dynamics, we observe the motion of chromatin in the nucleus as an indicator. Using particle image velocimetry (PIV) algorithms40 to analyse the videos of nucleus with H2B-mCherry expression at a 1-s time interval, we map the velocity of chromatin motion within the nucleus (Fig. 3a). Our findings reveal that hypoosmotic pressure enhances the overall motion of chromatin, suggesting a reduction in global crowding within the nucleus, while hyperosmotic pressure has the opposite effect (Fig. 3a, b).
a 2D velocity map of chromatin motion in cells. b Average chromatin velocity (2D) over 1 min. Statistics for cells with n = 35 for ctrl, n = 19 for 30% H2O and n = 16 for 10% PEG. Two-sample t-test is performed with p value annotated. c Overlay (2D) of telomeres labelled with dCas9-SunTag-FP at 60-s interval under initial isotonic condition, with the enlarged telomere focus marked by the white arrow on the image at 0 s and the 2D trajectory of it over 60 s. d 2D trajectories of 10 telomere foci in a cell over 40 s. The trajectory of each telomere focus is plotted in an individual colour with starting point shifted to the origin. e Mean squared displacement (MSD) of a telomere focus versus time lag. MSD in fixed cells for comparison. Ctrl, n = 196; 30% H2O, n = 115; 10% PEG, n = 72; fixed cell, n = 200. f Scatters of the fluorescence intensity of each telomere focus versus the diffusion exponent α of it. g Correlation between the average α and the total fluorescence intensity change percentage of telomere foci in a cell. Statistics for cells. Ctrl, n = 22; 30% H2O, n = 12; 10% PEG, n = 10. h Left, 2D snapshots of a cell with telomeres labelled with dCas9-SunTag-FP (green) and H2B labelled with mCherry (red). Right, the fluorescence intensity profiles along the white line drawn in the left image. i Change percentage of H2B-mCherry intensity within the regions of the telomere foci in a cell (3D) under non-isotonic conditions relative to the initial isotonic case. Statistics for cells with n = 49 for 30% H2O and n = 20 for 10% PEG. One-sample t-test with test mean of zero is performed with p value annotated. All statistics above are presented as mean ± SE.
After establishing the influence of extracellular osmotic pressure on overall chromatin dynamics, we further investigate if the dynamics of DNA loci is altered by the local crowding variations. We employ single-particle tracking methods41,42 to analyse the movement of telomeres labelled with dCas9-SunTag system (Fig. 3c). Our analysis reveals that telomere trajectories in a cell under hypoosmotic pressure exhibit reduced confinement and increased mobility, compared to their initial state, while hyperosmotic pressure restricts their movement (Fig. 3d). The histograms of telomere displacement from the mean position show a broader spread under hypoosmotic pressure, indicating less local crowding, and a narrower distribution under hyperosmotic pressure, suggesting increased local crowding (Supplementary Fig. 13). The mean squared displacement (MSD) curves further substantiate these findings, demonstrating an increase in telomere motion under hypoosmotic pressure and a decrease under hyperosmotic pressure (Fig. 3e). Collectively, these results indicate that hypoosmotic pressure promotes motion of telomeres, possibly due to reduced local crowding, while hyperosmotic pressure constrains telomere movement, likely due to enhanced local crowding.
To quantify the correlation between local crowding and fluorescent labelling with the dCas9-SunTag system, we calculate the diffusion exponent α for each telomere trajectory from MSD curves, which characterises the motion type and is correlated with the crowding41,42. We observe a positive correlation between α and the fluorescence intensity of a telomere focus under isotonic condition (Fig. 3f), suggesting that less crowded loci tend to bind more fluorescence proteins. More importantly, we find that this positive correlation is maintained under non-isotonic conditions (Fig. 3f), with both the average α and the total fluorescence intensity of telomere foci in a cell synchronously increasing under hypoosmotic pressure and decreasing under hyperosmotic pressure (Fig. 3g). Collectively, our experimental results of intranuclear dynamics support the notion that osmotic pressure regulates the binding of fluorescence proteins to dCas9-SunTag by altering intranuclear local crowding.
Given that chromatin is the main component of intranuclear volume43, intranuclear molecular crowding is largely attributed to chromatin density. To assess the chromatin density, we utilise H2B fluorescently labelled with mCherry as a proxy. Our observations reveal that hypoosmotic pressure leads to a decrease in H2B-mCherry signal intensity coincident with an increase in telomere-associated fluorescence intensity at corresponding loci, while hyperosmotic pressure yields the converse results (Fig. 3h, i). These findings suggest that the local modulation of chromatin crowding around telomeres plays a role in the observed variations in telomere focus fluorescence intensity labelled with dCas9-SunTag system within the nucleus.
Furthermore, we examine whether alternative methods of modulating intranuclear crowding could similarly regulate dCas9-SunTag labelling efficiency. By adding trypsin-EDTA to detach adherent cells under isotonic condition, we induce cellular rounding and nuclear expansion (Supplementary Fig. 14)44. Consistent with hypoosmotic effect, this decreased crowding increases both the number and the total intensity of telomere foci (Supplementary Fig. 14), supporting our hypothesis that reduced crowding enhances the binding of fluorescence proteins to dCas9-SunTag. Moreover, the modulation of intranuclear molecular crowding is concomitant with changes in ion concentration. The ions are known to modulate protein interaction through electrostatic45 or Hofmeister46 effects. Notably, the total intracellular ion concentration decreases under hypoosmotic condition15, while it increases under trypsin treatment due to ion influx through ion channels and pumps on plasma membranes44. Therefore, the elevation of telomere labelling efficiency under both conditions suggests that the crowding alternations rather than ion changes primarily drive the observed effect.
Osmotic pressure regulates transcription of target genes with dCas9-SunTag system
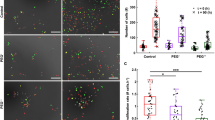
We have demonstrated that extracellular osmotic pressure modulates the binding of fluorescence proteins to dCas9-SunTag, thereby regulating target DNA labelling. Extending this observation, we question whether osmotic pressure could also influence gene transcription in a similar manner. We hypothesise that hypoosmotic pressure would enhance transcription efficiency by increasing the recruitment of transcription activation domains, while hyperosmotic pressure would have the opposite effect. To test this, we utilise sgRNA targeting the promoter of endogenous gene HBG1, and fuse synthetic transcription activation domain VPR47 or VP6448 to scFv-FP for transcription activation with the dCas9-SunTag system (Fig. 4a, and Supplementary Fig. 15). After 1-h osmotic treatment, the mRNA expression of HBG1 is measured through quantitative reverse transcription polymerase chain reaction (RT-qPCR). Consistent with our hypothesis, the relative mRNA expression of HBG1 increases under hypoosmotic treatment and decreases under hyperosmotic condition, with mRNA expression gradients corresponding to the extent of water or PEG addition within a certain range (Fig. 4b).
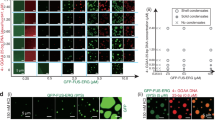
a Schematic of the transcription activation of a target gene with dCas9-SunTag-FP-AD. AD, activation domain. b Relative mRNA expression of HBG1 with the dCas9-SunTag recruiting the transcription activation domains VPR or VP64 under different osmotic conditions. Statistics for biological replicates with n = 4 for H2O and n = 6 for PEG. c Schematic of CRISPRTag-TRE-luciferase-24×MS2 DNA locus labelled and activated by dCas9-SunTag-FP-AD, with nascent MS2-tagged luciferase mRNA labelled by MCP-mCherry. d Relative luciferase expression with the dCas9-SunTag-FP-AD under different osmotic conditions. n = 3 independent experiments. e Left, snapshot showing the CRISPRTag-TRE-luciferase-24×MS2 locus labelled with dCas9-SunTag-sfGFP-AD. From 3D orthogonal view along z axis. Scale bar, 5 μm. Right, enlarged 2D snapshots at the target locus (marked by a white square on the left image) over time, indicating the transcription burst. Scale bar, 500 nm. 2D radial distributions of sfGFP and mCherry intensity are shown at the bottom. f Frequency of mRNA burst under various osmotic treatments. Statistics for target loci over 20 min. VPR, ctrl, n = 8; 20% H2O, n = 4; 10% PEG, n = 4. VP64, ctrl, n = 10; 20% H2O, n = 7; 10% PEG, n = 3. All statistics above are presented as mean ± SE.
With the findings that osmotic pressure regulates transcription, we next investigate whether this regulation would further impact downstream protein expression. We employ a CRISPRTag-TRE-luciferase-24×MS2 reporter cell line49, where the expression of exogenous luciferase serves as a more accurate and accessible proxy for protein expression than endogenous proteins. In this reporter cell line, a DNA fragment consisting of a CRISPR targetable sequence (CRISPRTag) and a Tet-inducible promoter (TRE)-driven luciferase-24×MS2 expression cassette is inserted into the ACTB site in an HEK293T cell49 (Fig. 4c). The specific targeting and efficient transcription activation are verified by the luciferase expression observed exclusively with the complete dCas9-SunTag-FP-AD/sgRNA system, as robust transcription activation requires precise spatial assembly of the dCas9-SunTag-FP-AD complex at the target site (Supplementary Fig. 16). Then the osmotic effect is investigated. As anticipated, the hypoosmotic pressure enhances the luciferase expression, while the hyperosmotic pressure has a repressive consequence (Fig. 4d). And the luciferase expression level exhibits a gradient with the concentration gradient of water or PEG within a certain span (Fig. 4d). These results confirm that the regulation of transcription with dCas9-SunTag system by osmotic pressure subsequently affects the downstream protein expression.
After establishing the regulatory effects of osmotic pressure on gene expression at ensemble level, we delve deeper into the transcription process at the molecular level. We label nascent MS2-tagged luciferase mRNA with MCP-mCherry, enabling real-time visualisation of transcription events (Fig. 4c). Under isotonic condition, the MCP-mCherry fluorescence intensity at the target locus shows mRNA synthesis occurring in a bursting mode (Fig. 4e), with a burst frequency of approximately 0.1 min−1 (Fig. 4f), consistent with previous studies50. Notably, the frequency of transcription burst is found to increase under hypoosmotic condition and decrease under hyperosmotic condition (Fig. 4f), aligning with our ensemble-level findings. These findings indicate that osmotic pressure modulates transcription activation frequency, thereby affecting gene expression with the dCas9-SunTag system.
Discussion
In this paper, we reveal the impact of extracellular osmotic pressure on the intranuclear environment and its subsequent effects on DNA labelling and transcription with the dCas9-SunTag system. We demonstrate that osmotic pressure modulates the binding-unbinding equilibrium of fluorescence proteins with the dCas9-SunTag complex, which in turn reversibly and repeatedly affects the fluorescent labelling of target DNA sequences. Specifically, hypoosmotic condition reduces intranuclear crowding, enhancing the binding of fluorescence proteins to dCas9-SunTag and increasing the number and the fluorescence intensity of target DNA foci. In contrast, hyperosmotic condition increases intranuclear crowding, promoting the unbinding of fluorescence proteins from dCas9-SunTag and decreasing the number and the fluorescence intensity of target DNA foci. More importantly, we discover that osmotic pressure regulates target gene transcription with the dCas9-SunTag system by observing changes in mRNA burst frequency directly, which has a further influence on downstream protein expression.
Our study enhances our understanding of how nuclei sense and respond to extracellular osmotic pressure changes, a phenomenon with implications for cell differentiation and development14,15,19. By leveraging the dCas9-SunTag system for fluorescent imaging and theoretical analysis, we directly quantify the correlations between protein binding kinetics, chromatin movements and gene expression at specific DNA loci within live-cell nuclei. We find that extracellular osmotic pressure modulates intranuclear crowding, perturbing protein binding-unbinding equilibrium, which in turn directly influences DNA-related processes. This is consistent with reports that molecular crowding reduces gene transcription in cells9,23,24, and our findings further highlight the role of nucleus as a mechanosensor14 by demonstrating that osmotic pressure changes directly affect protein binding kinetics. Furthermore, our results show that proteins condense in less crowded environments due to enhanced association with dCas9-SunTag, contrasting with previous studies that observed protein condensation in more crowded environments through phase separation51,52,53. The colocalization in the labelling specificity assessment (Supplementary Fig. 1) suggests that most of the foci correspond to target DNA loci rather than non-specific phase-separated aggregates. The association is possibly a reflection of the accessibility of binding sites on the SunTag. In a sparse environment, the SunTag peptide chain might expose more sites for the specific scFv-fused proteins to bind on. On the contrary, in a more crowded environment, the binding sites might be less accessible for the specific proteins due to spatial confinement of the SunTag chain. This suggests an alternative mechanism of nuclear response to osmotic pressure changes and emphasises the complexity of intranuclear environment.
The ability of the dCas9-SunTag system to respond to osmotic pressure offers an avenue for investigating and manipulating cellular functions. Compared to traditional control of the dCas9 system with chemically54 or optically55 induced dimerisation, the mechanical strategy using extracellular osmotic pressure modulations with dCas9-SunTag system provides a swift and reversible alternative way to regulate target DNA labelling and gene expression. More importantly, our study uncovers that the sensitivity of SunTag system to osmotic pressure offers an approach for investigating the response of intracellular microenvironment to extracellular stimuli. Combined with dCas9, the dCas9-SunTag system serves as a powerful probe for exploring the influence of osmotic pressure on nucleic at single-genomic-loci level. In addition to monitoring intranuclear crowding, the integration of the SunTag system with proteins specific to cellular organelles31 can be used to probe the response of various cellular compartments to osmotic pressure changes, providing a comprehensive understanding of how mechanical cues influence cellular functions across different organelles.
Methods
Plasmids, cell culture and transient transfection
The plasmids used in this work and the corresponding sources are listed in Supplementary Table 1.
HEK293T (including the variant cell lines, H2B-mCherry reporter cell line and CRISPRTag-TRE-luceferase-24×MS2 reporter cell line49) cells were cultured in Dulbecco’s modified Eagle medium (Gibco) with high glucose supplemented with 10% foetal bovine serum (Gibco) and 1% penicillin-streptomycin (Gibco). They were incubated at 37 °C and 5% CO2.
The cells were transfected with plasmids using LipofectamineTM 3000 (Invitrogen). About 24 h after the transfection, the transfection mixture was substituted with fresh growth medium. Around 48 h after the transfection, the cells were observed and applied in the experiments of osmotic effect.
Microscopy
Adherent cells were grown in glass-bottom dishes (Cellvis) and imaged on Leica TCS SP8 laser scanning confocal microscope system equipped with a chamber maintained at 37 °C and 5% CO2. The accompanying software was Leica Application Suite X. Images were acquired using a 40 × oil immersion objective (1.30 NA) and excitation lasers of 405 nm for Hoechst 33342, 488 nm for green fluorescence proteins (mNeonGreen and sfGFP) and 552 nm for red fluorescence proteins (mCherry and 3 × mRuby3). The scanning format was 512 × 512 and the pixel size was 253 nm × 253 nm. For z-stack scanning, the step size along z axis was 0.60 μm. For intranuclear 2D dynamics tracking, the time interval was 1 s and the recording duration was 1 min. Imaging positions were accurately recorded by the system to ensure that the same individual cells were measured before and after osmotic treatment and other drug treatment. Subsequent image visualisation was performed on ImageJ or Imaris.
Fluorescence in situ hybridisation (FISH)
Cells were fixed with 4% paraformaldehyde, permeabilized with 0.7% Triton X-100 and 0.1% saponin in 2× SSC (sodium saline citrate) buffer, and treated with RNase A at 37 °C for 1 h. Then, cells underwent dehydration in a graded ethanol series (70%, 85% and 100%) for 5 min each. Subsequently, cells were treated with 70% formamide in 2× SSC at 85 °C for 30 min. Then cells were sequentially dehydrated again in 70%, 80% and 95% ethanol for 5 min each in an ice bath. Next, Cells were incubated with 2 ng/mL Cy5-tagged telomere probe32 in a hybridisation mixture (10% dextran sulfate, 50% formamide and 500 ng/mL salmon sperm DNA in 2× SSC) at 37 °C overnight. Finally, cells were washed with 2× SSC three times.
Osmotic treatment and other drug treatment
For the hypoosmotic or hyperosmotic treatment, the initial isotonic cell culture medium was substituted with a mixture of culture medium and water or 200-Da PEG (Sigma-Aldrich), respectively, at a certain volumetric ratio. The osmotic pressure applied to cells was estimated by diluting the osmolarity due to water or adding the osmolarity due to PEG (5.62 Osm/L) to that of isotonic medium (300 mOsm/L)4,10,16 (Supplementary Fig. 2). For the recovery procedure, the non-isotonic solution was removed and fresh medium was added.
For the procedure of detaching adherent cells from glass substrate under isotonic condition, 0.05% trypsin-EDTA (Gibco) was added into the dish right after the initial medium was discarded.
During imaging experiments, all the operations of aspirating and adding solutions were meticulously performed on the sample stage of the microscope without moving the dishes, ensuring that the same individual cells were observed before and after drug treatment. For all the data analysis in the imaging experiments of osmotic effect or trypsin effect, the same individual cells were analysed across the treatment.
3D image processing and analysis
Nuclei in 3D images (identified by Hoechst 33342 or H2B-mCherry) were detected by the Surface tool on Imaris. Target fluorescence foci in 3D images were detected by the Spots tool on Imaris, where the estimated focus was modelled as a prolate spheroid with xy diameter of 0.7 μm and z diameter of 1.4 μm, and the filter criterion was the quality of a focus, defined as the centre fluorescence intensity with background subtraction (the intensity of the channel Gaussian filtered by 3/4 of spot radius minus the intensity of the original channel Gaussian filtered by 8/9 of spot radius). To quantify the intensity of target foci in a nucleus, the intensity of nuclear overall background (outside the identified focus spheroids in the nucleus) was firstly subtracted from the original signal to extract the background-corrected intensity. The total fluorescence intensity of target foci in a cell was calculated as the background-corrected intensity sum of all the identified focus spheroids. The quantification was performed on Matlab.
Analysis for fluorescence recovery after photobleaching (FRAP)
The normalised fluorescence intensity inside the photobleached area over time t (Fig. 2b) was calculated as
where I(t) was the original intensity at time t, \({I}_{{\rm{pre}}}\) was the original intensity right before photobleaching while I0 was the one at the first time point after photobleaching.
Particle image velocimetry (PIV)
PIV40 for 2D images of H2B-mCherry was carried out with PIVlab on Matlab. The algorithm of FFT window deformation was employed. Three successive passes were run, with the respective interrogation areas of 16, 8, 4 and the respective steps of 8, 4, 2. Then 2D Gaussian function was used to find a displacement peak with sub-pixel precision. During post-processing, velocity limits were set with standard deviation filter threshold of 8 and local median filter threshold of 3. Data smoothing of the least strength was performed to obtain the final reliable results.
Analysis for single-particle tracking
The 2D trajectories of telomere foci in a cell were tracked by the Spots tool on Imaris, where the estimated focus was modelled as a circle with a diameter of 0.7 μm and the filter criterion was the quality of a focus with background subtraction. The tracking algorithm was Brownian motion with a maximum distance of 0.7 μm and a maximum time gap of three frames. The translational and rotational drift was subtracted to obtain the final corrected trajectories, with main steps as follows: firstly, the mass centre of all telomere foci in each frame is subtracted from the trajectories, which corrects for translational drift; next, the average rotation matrix in the xy plane for all foci around the mass centre is calculated for each time point; finally, each trajectory is multiplied by the inverse rotation matrix.
The MSD of a telomere trajectory as a function of time lag τ was calculated as
where t was the acquisition time and r was the 2D coordinate. The MSD curve within a range of 0 < τ ≤15 s was fitted to
where D was the diffusion coefficient and α was the diffusion exponent41,42. The MSD calculation as well as the fitting was performed on Matlab.
Analysis for nascent mRNA at target loci
For the analysis on nascent mRNA tagged with MS2-MCP-mCherry, the target site was identified by the target sfGFP focus. The region inside a prolate spheroid with xy diameter of 1 μm and z diameter of 2 μm was denoted as Ω1. The region inside a prolate spheroid with xy diameter of 4 μm and z diameter of 8 μm and within a distance of 1 μm from the centre along z axis was denoted as Ω2. The MCP-mCherry effective intensity at the target sfGFP focus was calculated as
where Ioriginal was the original intensity mean in Ω1, Ibackground was the original intensity mean in Ω2 − Ω1. If Ieffective at a time point was greater than 10%, it was considered that an mRNA burst was occurring.
Measurement of mRNA expression: quantitative reverse transcription polymerase chain reaction (RT-qPCR)
RNA was isolated using Trizol Reagent (Vazyme, R411-02-AA) and reverse-transcribed into cDNA using HiScript II Q RT SuperMix for qPCR (+gDNA wiper) (Vazyme, R223-01). Quantitative real-time PCR (qPCR) was performed on LightCycler 96 System using 2xTaq pro universal SYBR qPCR Master Mix (Vazyme, R712-02-AA). The ΔΔCt method was employed to obtain the relative gene expression normalised by the glyceraldehyde phosphate dehydrogenase (GAPDH) expression. For the qPCR primers, please refer to the source literature49.
For the experiment of osmotic effect, mRNA expression was measured after the cells were treated under the corresponding osmotic condition for about 1 h. The relative mRNA expression for each osmotic condition shown in Fig. 4b was normalised by the mRNA expression of blank group under the same osmotic condition. The blank group referred to the cells transfected with empty plasmid vectors (pBluescript SK+).
Luciferase assay
HEK293T CRISPRTag-TRE-luceferase-24×MS2 reporter cells were seeded in a 12- or 24-well plate. 45 μg/mL D-luciferin (Beyotime) was added to each well, and then the cells were incubated at 37 °C and 5% CO2 for 5 min. Subsequently, the plate was imaged on IVIS Lumina II In Vivo Imaging System. Bioluminescence analysis was performed on Living Image. The luciferase expression was represented by the detected bioluminescence in a well, normalised by the blank well where the cells were transfected with empty plasmid vectors.
For the experiment of osmotic effect, luciferase assay was performed after the cells were treated under the corresponding osmotic condition for about 3 h. The relative luciferase expression for each osmotic condition shown in Fig. 4d was obtained by subtracting the luciferase expression of blank group under the same osmotic condition.
Statistics and reproducibility
Detailed information on study design, experimental implementations and statistics analyses is provided in the corresponding subsections of Methods and figure captions. No statistical method was used to predetermine sample size. Cell sampling was random. Investigators were blinded to allocation during experiments and outcome assessment. Data analyses were performed on Matlab or Origin, unless stated otherwise. For the calculation of change percentage, values of infinity were not included in the statistics. For two-sample t-test, equal variance was not assumed and Welch correction was employed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data generated in this study are provided in the Source Data file with the paper. Source data are provided with this paper.
References
Miermont, A. et al. Severe osmotic compression triggers a slowdown of intracellular signaling, which can be explained by molecular crowding. Proc. Natl Acad. Sci. 110, 5725–5730 (2013).
Charras, G. & Sahai, E. Physical influences of the extracellular environment on cell migration. Nat. Rev. Mol. Cell Biol. 15, 813–824 (2014).
Stewart, M. P. et al. Hydrostatic pressure and the actomyosin cortex drive mitotic cell rounding. Nature 469, 226–230 (2011).
Guo, M. et al. Cell volume change through water efflux impacts cell stiffness and stem cell fate. Proc. Natl Acad. Sci. 114, E8618–E8627 (2017).
Jiang, C. et al. Switch of cell migration modes orchestrated by changes of three-dimensional lamellipodium structure and intracellular diffusion. Nat. Commun. 14, 5166 (2023).
Chan, C. J. & Hiiragi, T. Integration of luminal pressure and signalling in tissue self-organization. Development 147, dev181297 (2020).
Liu, F. et al. Intercellular flow dominates the poroelasticity of multicellular tissues. Nat. Phys. 21, 1311–1318 (2025).
Colom, A. et al. A fluorescent membrane tension probe. Nat. Chem. 10, 1118–1125 (2018).
Chao, P.-hG., West, A. C. & Hung, C. T. Chondrocyte intracellular calcium, cytoskeletal organization, and gene expression responses to dynamic osmotic loading. Am. J. Physiol.-Cell Physiol. 291, C718–C725 (2006).
Zhou, E. H. et al. Universal behavior of the osmotically compressed cell and its analogy to the colloidal glass transition. Proc. Natl Acad. Sci. 106, 10632–10637 (2009).
Delarue, M. et al. mtorc1 controls phase separation and the biophysical properties of the cytoplasm by tuning crowding. Cell 174, 338–349.e20 (2018).
Li, B. et al. Intracellular transport is accelerated in early apoptotic cells. Proc. Natl Acad. Sci. 115, 12118–12123 (2018).
Zhang, M.-L. et al. Interplay between intracellular transport dynamics and liquid-liquid phase separation. Adv. Sci. 11, 2308338 (2024).
Kirby, T. J. & Lammerding, J. Emerging views of the nucleus as a cellular mechanosensor. Nat. Cell Biol. 20, 373–381 (2018).
Finan, J. D. & Guilak, F. The effects of osmotic stress on the structure and function of the cell nucleus. J. Cell. Biochem. 109, 460–467 (2010).
Roffay, C. et al. Passive coupling of membrane tension and cell volume during active response of cells to osmosis. Proc. Natl Acad. Sci. 118, e2103228118 (2021).
Deviri, D. & Safran, S. A. Balance of osmotic pressures determines the nuclear-to-cytoplasmic volume ratio of the cell. Proc. Natl Acad. Sci. 119, e2118301119 (2022).
Irianto, J. et al. Osmotic challenge drives rapid and reversible chromatin condensation in chondrocytes. Biophys. J. 104, 759–769 (2013).
Lebeaupin, T., Smith, R. & Huet, S. The Multiple Effects of Molecular Crowding in the Cell Nucleus: from Molecular Dynamics to the Regulation of Nuclear Architecture, vol. 2, 209–232 (Academic Press, 2018).
Amat, R. et al. Rapid reversible changes in compartments and local chromatin organization revealed by hyperosmotic shock. Genome Res. 29, 18–28 (2019).
Murr, R. et al. Histone acetylation by trrap-tip60 modulates loading of repair proteins and repair of DNA double-strand breaks. Nat. Cell Biol. 8, 91–99 (2006).
Akabayov, B., Akabayov, S. R., Lee, S.-J., Wagner, G. & Richardson, C. C. Impact of macromolecular crowding on DNA replication. Nat. Commun. 4, 1615 (2013).
Kültz, D. Osmotic Regulation of DNA Activity and the Cell Cycle, Vol. 1, 157–179 (Elsevier, 2000).
Vaňková Hausnerová, V. & Lanctôt, C. Chromatin decondensation is accompanied by a transient increase in transcriptional output. Biol. Cell 109, 65–79 (2017).
Zhang, X. et al. Compression drives diverse transcriptomic and phenotypic adaptations in melanoma. Proc. Natl Acad. Sci. 120, e2220062120 (2023).
Zimmerman, S. B. & Minton, A. P. Macromolecular crowding: biochemical, biophysical, and physiological consequences. Annu. Rev. Biophys. 22, 27–65 (1993).
Bancaud, A. et al. Molecular crowding affects diffusion and binding of nuclear proteins in heterochromatin and reveals the fractal organization of chromatin. EMBO J. 28, 3785–3798 (2009).
Bounedjah, O. et al. Macromolecular crowding regulates assembly of mRNA stress granules after osmotic stress. J. Biol. Chem. 287, 2446–2458 (2012).
Wang, H. et al. Synthesis of abiotic supramolecular polymers inside living cells via organocatalysis-mediated self-assembly. Angew. Chem. Int. Ed. 64, e202500998 (2025).
Wright, A., Nuñez, J. & Doudna, J. Biology and applications of CRISPR systems: harnessing nature’s toolbox for genome engineering. Cell 164, 29–44 (2016).
Tanenbaum, M., Gilbert, L., Qi, L., Weissman, J. & Vale, R. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Ye, H., Rong, Z. & Lin, Y. Live cell imaging of genomic loci using CRISPR-Cas9 system and a bright fluorescent protein. Protein Cell 8, 853–855 (2017).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Kong, J.-W., Dou, S.-X., Li, W., Li, H. & Wang, P.-Y. Parallel dna g-quadruplex induced and stabilized by curaxin cbl0137. Chin. Phys. Lett. 40, 078701 (2023).
Moniaux, N. et al. Complete sequence of the human mucin muc4: a putative cell membrane-associated mucin. Biochem. J. 338, 325–333 (1999).
Guo, M. et al. Probing the stochastic, motor-driven properties of the cytoplasm using force spectrum microscopy. Cell 158, 822–832 (2014).
Zosel, F., Soranno, A., Buholzer, K. J., Nettels, D. & Schuler, B. Depletion interactions modulate the binding between disordered proteins in crowded environments. Proc. Natl Acad. Sci. 117, 13480–13489 (2020).
Tzur, A., Kafri, R., LeBleu, V. S., Lahav, G. & Kirschner, M. W. Cell growth and size homeostasis in proliferating animal cells. Science 325, 167–171 (2009).
Cooper, K. L. et al. Multiple phases of chondrocyte enlargement underlie differences in skeletal proportions. Nature 495, 375–378 (2013).
Thielicke, W. & Stamhuis, E. J. Pivlab—towards user-friendly, affordable and accurate digital particle image velocimetry in MATLAB. J. Open Res. Softw. (2014).
Dix, J. A. & Verkman, A. Crowding effects on diffusion in solutions and cells. Annu. Rev. Biophys. 37, 247–263 (2008).
Jiang, C., Dou, S.-X., Wang, P.-Y. & Li, H. Quantifying transport dynamics with three-dimensional single-particle tracking in adherent cells. STAR Protoc. 3, 101790 (2022).
Rouquette, J. et al. Revealing the high-resolution three-dimensional network of chromatin and interchromatin space: a novel electron-microscopic approach to reconstructing nuclear architecture. Chromosome Res. 17, 801–810 (2009).
Lordon, B., Campion, T., Gibot, L. & Gallot, G. Impact of trypsin on cell cytoplasm during detachment of cells studied by terahertz sensing. Biophys. J. 123, 2476–2483 (2024).
Zhou, H.-X. & Pang, X. Electrostatic interactions in protein structure, folding, binding, and condensation. Chem. Rev. 118, 1691–1741 (2018).
Gregory, K. P. et al. Understanding specific ion effects and the Hofmeister series. Phys. Chem. Chem. Phys. 24, 12682–12718 (2022).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Beerli, R. R., Segal, D. J., Dreier, B. & Barbas, C. F. Toward controlling gene expression at will: specific regulation of the erbb-2/her-2 promoter by using polydactyl zinc finger proteins constructed from modular building blocks. Proc. Natl Acad. Sci. 95, 14628–14633 (1998).
Ma, S. et al. Phase-separated dropcrispra platform for efficient gene activation in mammalian cells and mice. Nucleic Acids Res. 51, 5271–5284 (2023).
Wang, Y., Ni, T., Wang, W. & Liu, F. Gene transcription in bursting: a unified mode for realizing accuracy and stochasticity. Biol. Rev. 94, 248–258 (2019).
Hancock, R. A role for macromolecular crowding effects in the assembly and function of compartments in the nucleus. J. Struct. Biol. 146, 281–290 (2004).
Jalihal, A. P. et al. Multivalent proteins rapidly and reversibly phase-separate upon osmotic cell volume change. Mol. Cell 79, 978–990.e5 (2020).
Gao, C. et al. Hyperosmotic-stress-induced liquid-liquid phase separation of ALS-related proteins in the nucleus. Cell Rep. 40, 111086 (2022).
Zetsche, B., Volz, S. E. & Zhang, F. A split-cas9 architecture for inducible genome editing and transcription modulation. Nat. Biotechnol. 33, 139–142 (2015).
Polstein, L. R. & Gersbach, C. A. A light-inducible crispr-cas9 system for control of endogenous gene activation. Nat. Chem. Biol. 11, 198–200 (2015).
Acknowledgements
This study is supported by the National Natural Science Foundation of China (12074043 & 12122402 to H.L., 82070002 & 82370078 to Z.R., 12047503 & 12275332 to F.M.), the Major Project of Guangzhou National Laboratory (GZNL2023A03004 to Z.R.). F.M. acknowledges support from Chinese Academy of Sciences (XDPB15 to F.M.), Max Planck Society (Max Planck Partner Group), Wenzhou Institute (WIUCASICTP2022 and WIUCASQD2023009) and Beijing National Laboratory for Condensed Matter Physics (2023BNLCMPKF005).
Author information
Authors and Affiliations
Contributions
H.L. supervised the project. H.L., Z.R. and S.X.D. conceived the project. T.W., X.Y., C.J. and H.Y. performed the experiments. T.W. and F.M. developed the theoretical model. T.W., X.Y., P.Y.W., F.M., S.X.D., Z.R. and H.L. analysed and interpreted the data. K.L., H.Y.L., Y.R.L. and P.Y.W. contributed the new samples, agents, equipment or analytic tools. T.W., S.X.D., Z.R. and H.L. wrote the manuscript. All authors edited and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xiakun Chu, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wei, T., Yang, X., Jiang, C. et al. Osmotic pressure regulates DNA labelling and transcription with dCas9-SunTag system in live cells. Nat Commun 17, 773 (2026). https://doi.org/10.1038/s41467-025-66160-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66160-6