Abstract
Nonsense mutations change a sense codon into a premature termination codon (PTC) in mRNA and account for approximately 18.5% of human inherited retinal diseases (IRDs)-related mutation. Nonsense suppression therapies by small molecular drugs or suppressor tRNAs (sup-tRNAs) can introduce an amino acid at PTC, thereby promoting the production of full-length proteins. While sup-tRNA-based therapies have shown promising results in cell culture models, challenges remain for in vivo delivery, particularly regarding efficacy, stability, and safety. In this study, we engineer the body sequence of sup-tRNAArg to enhance readthrough efficiency at clinically relevant PTCs in the RPE65 and ABCA4 genes. By using the self-complementary adeno-associated virus (scAAV), we achieve restoration of RPE65 protein expression in up to 50.2% retinal pigment epithelium (RPE) cells in a mouse model carrying the RPE65-R44X nonsense mutation. Notably, the engineered sup-tRNAArg significantly restores retinal function in mice, with effects lasting for at least 36 weeks. Furthermore, scAAV8.sup-tRNAArg exhibits minimal retinal toxicity and had negligible effects on global readthrough at native termination codons as revealed by ribosome profiling. Our findings demonstrate scAAV-delivered sup-tRNAArg is a potential therapeutic intervention for IRDs caused by nonsense mutations in RPE65 gene.
Similar content being viewed by others
Introduction
Inherited retinal diseases (IRDs) are a heterogenous group of retinal degenerative diseases with at least 300 causative genes identified to date1. In-frame nonsense mutations account for approximately 18.5% of human IRDs-related mutations2. mRNAs bearing premature stop codon (PTC) are incapable of producing full-length proteins and frequently activate nonsense mediated decay (NMD) pathways3. Nonsense suppression enables the incorporation of an amino acid at the PTC. It is an appealing treatment for IRDs because it relies on the specific PTC sequence and wild-type codon rather than the underlying gene. To overcome PTCs, small-molecule readthrough drugs have been used to treat IRDs, including retinitis pigmentosa4, Usher syndrome5 and aniridia6. However, insertion of random amino acids, off-target effects and toxicity in long-term use have limited their clinical application7.
With advancements in gene therapy, adeno-associated virus (AAV)-delivered gene replacement therapy can be used to treat IRDs caused by nonsense mutations8. However, many IRDs causal genes, including ABCA4, whose pathogenic variants account for ~24.8% of IRDs-causing mutations, exceed the packaging capacity of AAV ( ~4.7 kb for ssDNA)2. Additionally, long-term follow-up studies have shown that patients who received RPE65 gene therapy exhibit persistent photoreceptor loss and a gradual decline in visual acuity improvement9,10. While the cause of this progressive retinal degeneration remains unclear, inadequate expression of RPE65 from the delivered transgene or from mutated endogenous RPE65 transcripts may contribute to the observed decline9,11,12. CRISPR-based genome editing holds promise; however, it faces significant challenges for in vivo delivery due to its large size and potential immunogenicity13.
The anticodon-engineered tRNA (suppressor tRNA, sup-tRNA) can precisely suppress PTCs through Watson-Crick interactions with all three nucleotides. The concept of sup-tRNA was first proposed by Sveda and colleagues in 197714. Many efforts have been made to expand the application of sup-tRNAs in various disease contexts15,16,17,18. Recent studies have successfully delivered sup-tRNAs in vivo via AAV19 or lipid nanoparticles (LNPs)20. The use of AAV for sup-tRNA delivery opens new possibilities for gene therapies, as the protein expression restored by sup-tRNA is derived from endogenous transcripts. Nevertheless, its low efficacy restricts the widespread application of sup-tRNA in clinically relevant monogenic model systems.
Multiple studies on tRNA biology and body sequence engineering suggest the possibility of designing more effective sup-tRNAs by optimizing the body sequences, especially the TΨC-stem and anticodon-stem20,21,22,23. We reason that by applying a similar strategy in which body sequences crucial for tRNA stabilization and decoding are modulated, we can efficiently suppress endogenous PTCs in vivo and correct clinically relevant disease phenotypes. We also leverage a self-complimentary adeno-associated viral vector to deliver engineered sup-tRNAArg (scAAV8.sup-tRNAArg). scAAV8.sup-tRNAArg achieves high efficacy in restoring the visual function in an inherited retinal disease mouse model harboring a homozygous nonsense mutation in the RPE65 gene (RPE65 p.R44X)24, which recapitulates the human condition and is analogous to that of human patients with LCA225,26. These data underscore the potential of sup-tRNA body sequence engineering and provide robust evidence for further applications of scAAV-delivered sup-tRNA in other IRDs caused by nonsense mutations.
Results
Rational design of sup-tRNAArg for high efficacy
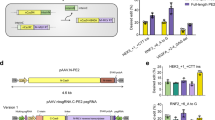
The body sequences of tRNAs are fine-tuned for efficient aminoacylation and decoding27,28. In Eukarya, tRNAArg A/C35 and A/U/G36 are indispensable for the aminoacylation process by arginine aminoacyl-tRNA synthetase (RARS)29. Therefore, to achieve high efficacy in decoding the UGA stop codon and introducing arginine precisely, we selected near-cognate human tRNAArg isodecoders (only one mismatch base with UGA), namely, tRNAArg-TCT-1 to tRNAArg-TCT-5 and tRNAArg-TCG-1 to tRNAArg-TCG-6 and mutated their anticodons to 5’-UCA-3’ (Fig. 1a and Supplementary Table 1). The U6 promoter was used to stably express sup-tRNA since it enhances suppression activity30. The codon-related translation velocity significantly influences on the efficacy of sup-tRNAs31. Therefore, to obtain the sup-tRNAArg with highest suppressive efficacy at mRPE65-R44X, we constructed an enhanced green fluorescence protein (EGFP) reporter containing the sequence context around the mRPE65-R44X PTC (Supplementary Fig. 1a). All sup-tRNAArg successfully induced readthrough and partially restored EGFP signal in HEK293 cells (Supplementary Fig. 1b, c). Among them, sup-tRNAArg-TCT-1, sup-tRNAArg-TCT-2, sup-tRNAArg-TCT-3, sup-tRNAArg-TCT-4 and sup-tRNAArg-TCG-1 were the top 5 most potent sup-tRNAs. To evaluate their efficacy in a more precise way, we constructed a dual-luciferase reporter containing the context sequence around mRPE65-R44X (Fig. 1b). G418, an efficient readthrough compound, induced only 2.5% readthrough while sup-tRNAArg-TCT-1 was more efficient and induced 26.6% readthrough (Fig. 1c). Therefore, we chose the body sequence of sup-tRNAArg-TCT-1 (TCT-1) for further optimization.
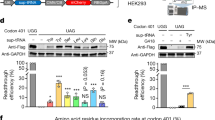
a Schematic showing that sup-tRNAArg engineered from near-cognate natural tRNAArg recognizes UGA stop codon by the mutated anticodon. Created in BioRender. Yang, Y. (2025) https://BioRender.com/sb411h1. b Schematic of the workflow used to evaluate the readthrough efficacy of sup-tRNAArg with plasmid constructs encoding the dual-luciferase reporter containing RPE65-R44X sequence context. Created in BioRender. Yang, Y. (2025) https://BioRender.com/sb411h1. c Readthrough efficacy of sup-tRNAArg calculated using the ratio of normalized Gaussia luciferase (Gluc) activity to firefly luciferase (Fluc) activity. Data are mean ± s.e.m. of eight biological replicates. d Nucleotide substitutions in the anticodon-stem (A1-A6 variants) or TΨC-stem (T1-T4 variants). e Comparison of PTC suppression efficiency of TCT-1 and its variants. The readthrough activity was calculated using the ratio of Gluc activity to Fluc activity. Data are mean ± s.e.m. of four biological replicates. f Comparison of PTC suppression efficiency of combined variants and TCT-1. The readthrough activity was calculated using the ratio of Gluc activity to Fluc activity. Data are mean ± s.e.m. of three biological replicates. g Schematic of the workflow used to evaluate the PTC suppression activity of sup-tRNAArg with plasmid constructs encoding the full-length mRPE65-R44X cDNA. Created in BioRender. Yang, Y. (2025) https://BioRender.com/sb411h1. h Representative western blot images (top) and quantification (bottom) of mouse RPE65 protein expression restored by sup-tRNAArg. Data are mean ± s.e.m. of five biological replicates. i qPCR quantification of mRPE65 cDNA in HEK293 cells received indicated treatment. Data are mean ± s.e.m. of five biological replicates. Statistical analysis was performed by two-sided t-test.
Inspired by previous works20,22,32, the rational design in this study focused on the anticodon-stem, which modulates the decoding ability and amino acylation by arginyl-tRNA synthetase (RARS), and the TΨC-stem, which determines the binding affinity to elongation factor33,34,35. To avoid disturbing the amino acylation by RARS of TCT-1, we designed various anticodon-stem (A1-A6) based on the anticodon-stem of sup-tRNAArg-TCG (Fig. 1d and Supplementary Table 2). The readthrough level of a specific sup-tRNA depends on the translation efficacy and competitive ability against translation termination complex at PTC. Stable interactions with eEF1A tend to confer higher competitive ability; however, it will impede the dissociation of eEF1A after sup-tRNA into the A site of ribosome, resulting in reduced rates of peptide bond formation34. Therefore, we designed various and TΨC-stem (T1-T4) sequences (Fig. 1d and Supplementary Table 2) according to the free energy (ΔΔG) contributions of different base pairs36. Compared with TCT-1, site specific engineering (T1, T2, A4, A5) increased the readthrough efficiency by 1.1- to 1.6-fold (Fig. 1e). Subsequent combining of mutations in TΨC-stem and anticodon-stem resulted in further enhancement in readthrough ability. The final version sup-tRNAArg-A4T1 (A4T1) yielded 2.2-fold readthrough compared with TCT-1 (A4T1: 49.8% vs TCT-1: 22.6%) (Fig. 1f). As the most efficient variant of TCT-1 which incorporated predominantly Arg at the UGA stop codon (Supplementary Fig. 2a, b), A4T1 induced 2-fold readthrough of TCT-1 at multiple dosages (Supplementary Fig. 3a, b). More importantly, the culture media of HEK293 cells transfected with a small amount of A4T1 plasmid (10 ng) presented increased Gluc luciferase activity compared with that of HEK293 cells transfected with 50 ng TCT-1 (Supplementary Fig. 3c). To further confirm the superiority of A4T1 when sup-tRNA is insufficiently transduced, we transfected HEK293 cells with 5 ng of sup-tRNA and 50 ng of the RPE65-R44X-EGFP reporter. The percentage of EGFP fluorescence restored by A4T1 was ~32.7% of that in cells transfected with RPE65-R44-EGFP, whereas the percentage restored by TCT-1 was approximately 12.5% (Supplementary Fig.3d-f). A4T1 induced EGFP expression in 19% more cells than TCT-1 did (Supplementary Fig. 3g). Moreover, we measured the sup-tRNA abundance by appending a self-cleaving ribozyme to the 3’ end of the sup-tRNA as previously described23 (Supplementary Fig. 3h). A4T1 showed a similar expression level compared with TCT-1 (Supplementary Fig. 3i), indicating the higher readthrough level was not due to increased expression.
When transfected as plasmid-encoded formulation, both TCT-1 and A4T1 showed toxicity similar to that of EGFP (Supplementary Fig. 4a). The unfolded protein response is activated when cells undergo global readthrough induced by the depletion of translation termination factors37. The mRNA levels of ATF4, BIP and CHOP were not significantly upregulated by TCT-1 or A4T1 treatment (Supplementary Fig. 4b), demonstrating the safety of sup-tRNAs.
sup-tRNAArg inhibits NMD and restores full-length proteins
The translational dynamics differ largely in full-length proteins and sequence context-containing luciferase reporters31. Therefore, we evaluated whether the full-length RPE65-R44X transcript could be restored via sup-tRNAArg (Fig. 1g). Results of western blot after co-transfection of sup-tRNAArg and full-length RPE65-R44X plasmids in HEK293 cells revealed that A4T1 restored 72.3% full-length RPE65 protein expression while TCT-1 restored 33.6% (Fig. 1h). Furthermore, G418 had no detectable effect on restoring full-length RPE65 protein expression. NMD substantially hinders the therapeutic outcome of nonsense suppression therapies38. The RPE65-R44X mRNA levels were substantially restored by treatment with TCT-1 and A4T1 and the restoration was dose-dependent (Fig. 1i). G418 has no significant effect on antagonizing NMD (Fig. 1i). A previous study reported that antagonism toward NMD is related to readthrough levels39. The lower restoration rate of A4T1 suggests that mechanisms other than readthrough ability affect the inhibition of NMD.
To test whether sequence engineering improved nonsense suppression efficacy for other disease relevant nonsense mutation, we constructed a plasmid expressing full-length human ABCA4 gene (hABCA4) containing the clinically common nonsense mutation p.R681X and co-transfected it with plasmids expressing TCT-1 or A4T1 (Supplementary Fig. 5a). The western blot results demonstrated that, compared with TCT-1, A4T1 restored the expression of the full-length ABCA4 protein by 1.8-fold (Supplementary Fig. 5b), indicating the sequence engineering strategy is able to yield greater efficiency in producing various full-length proteins. The nucleotide immediately following the PTC ( + 4 nucleotide) has a substantial influence on the readthrough capacity of readthrough drugs40 as well as sup-tRNAs41. We detected the readthrough capacity of the two sup-tRNAs on dual-luciferase reporters with either U, G, A, or C as +4 nucleotide. The results showed that A4T1 achieved higher readthrough level for all +4 nucleotides (U: 2.1-fold; G: 4.4-fold; A: 4.3-fold; C: 3.7-fold) (Supplementary Fig. 5c).
scAAV8.sup-tRNAArg suppresses RPE65-R44X nonsense mutation in vivo
To test whether sup-tRNAArg could rescue visual function in rd12 mice harboring RPE65-R44X nonsense mutation and evaluate the effectiveness of the sequence engineering strategy, we generated two scAAV vector constructs carrying sup-tRNAArg-TCT-1 and sup-tRNAArg-A4T1 and packaged each construct into an AAV8 capsid (scAAV8.sup-tRNAArg-TCT-1 and scAAV8.sup-tRNAArg-A4T1, hereafter scAAV8.TCT-1 and scAAV8.A4T1). The two vectors were injected into the subretinal space of 2-week-old rd12 mice at 1 × 108, 3 × 108, or 1 × 109 genome copies (GC) per eye (Supplementary Fig. 6a). Scotopic and photopic electroretinography (ERG) were used to evaluate the therapeutic outcomes. Typically, rd12 mice exhibit early extinguishing ERG amplitudes. Three weeks after injection, mice treated with two types of scAAV8.sup-tRNAArg presented dose-dependent increases in both a- and b-wave amplitudes in scotopic and photopic ERG examinations (Supplementary Fig. 6b). More specifically, mice treated with scAAV8.A4T1 exhibited an overall higher a- and b-wave amplitudes compared with those treated with scAAV8.TCT-1 at the same dosages (Supplementary Fig. 6c, d). Western blot analysis demonstrated that high-dose scAAV8.A4T1 treatment successfully restored an average of 16.4% (8.0-30.1%) RPE65 protein compared with that of the WT control (Supplementary Fig. 6e), highlighting the efficacy of scAAV8.A4T1 in retinal pigment epithelium (RPE). Photoreceptors are also a major target cell type for gene therapy in IRDs. To assess whether scAAV8.A4T1 exhibits higher readthrough efficacy in photoreceptors, we employed an in vivo reporter system incorporating the ABCA4-R681X mutation sequence context. Analogous to the RPE65-R44X-EGFP reporter used in our in vitro screening, the ABCA4-R681X sequence context was inserted downstream of the AUG start codon of EGFP, and plasmids encoding either R681X-EGFP or R681-EGFP were packaged into AAV vectors. A human RK promoter was used to drive expression of R681X-EGFP predominantly in photoreceptors (Supplementary Fig. 7a). The results showed that scAAV8.A4T1 restored stronger EGFP signals in photoreceptor cells compared with scAAV8.TCT-1 (Supplementary Fig. 7b). The overall lower efficacy observed in this reporter system compared with rd12 mice is likely attributable to several factors, including limited AAV transduction efficiency for photoreceptors, the dual-AAV strategy, and the intrinsically lower readthrough rate of ABCA4-R681X.
scAAV8.A4T1 rescues retinal function of rd12 mice
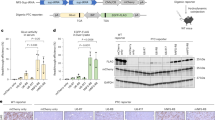
Encouraged by the promising short-term results, we aimed to conduct a comprehensive evaluation of the long-term efficacy and safety of scAAV8.sup-tRNAArg. scAAV8.A4T1 or scAAV8.TCT-1 was injected into the subretinal space at 1 × 109 GC per eye (Fig. 2a). We initially detected the RPE65 protein level at 12 weeks post injection. Results of western blot analysis showed a significant RPE65 band in lysates from scAAV8.A4T1-injected mouse eyes (Fig. 2b). The immunofluorescence on RPE flat-mount revealed an average of 36.0% RPE65 positive cell in scAAV8.A4T1-treated eyes and 4.5% in scAAV8.TCT-1-treated eyes (Fig. 2c). The mRNA level of RPE65 in rd12 mice was reported to be approximately 50%-60% of that in WT mice11, which was also confirmed by our results (Fig. 2d). This should be ascribed to a moderate level of NMD. scAAV8.A4T1 stabilized the expression of RPE65-R44X mRNA at levels similar to those of RPE65-R44 (Fig. 2d). We subsequently analyzed the retinal function by scotopic and photopic ERG. The results of scotopic and photopic ERG demonstrated that the a- and b-wave amplitudes in eyes treated with scAAV8.A4T1 were significantly increased in response to multiple intermittent flash stimuli compared with those in untreated eyes or scAAV8.TCT-1-treated eyes (Fig. 3a-f). Specifically, compared with those of the wild-type mice, the scAAV8.A4T1 treatment rescued 21.3% and 35.6% of the a- and b-wave amplitudes in scotopic ERG (Fig. 3c) and rescued 83.0% and 76.9% of the a- and b-wave amplitudes in photopic ERG (Fig. 3f). Since rd12 mice exhibited early cone cell degeneration, the significant improvement in photopic ERG results after scAAV8.A4T1 treatment might result from prolonged cone survival. To evaluate the survival rate of cones, retinal flat-mounts were stained with peanut agglutinin (PNA)-lectin and M-opsin. A total of 81.8% of the retinal area was M-opsin positive in eyes treated with scAAV8.A4T1 while 34.2% in untreated eyes and 45.7% in scAAV8.TCT-1-treated eyes (Fig. 3g). The survival of cones was also calculated by averaging the number of cones in four fields at the dorsal and ventral retina at 1 mm from the optic nerve in both treated and untreated eyes. PNA-lectin binds to the galactose-galactosamine disaccharide residues, which is specifically produced by living cones42. Compared with that in untreated eyes, the average number of PNA-lectin positive cells was significantly higher in scAAV8.A4T1 treated eyes (1608.8 vs 352.1). Moreover, compared with scAAV8.TCT-1, scAAV8.A4T1 had better protective effects (1608.8 vs 718.7) (Fig. 3h, i).
a Schematic of the scAAV vector system and the workflow used to examine the efficacy and safety of subretinal delivery of scAAV8.sup-tRNAArg in rd12 mice. Nucleotide sequences are shown in Supplementary Data 1. Created in BioRender. Yang, Y. (2025) https://BioRender.com/sb411h1. b Representative western blot images (left) and quantification (right) of mouse RPE65 protein expression in retinal pigment epithelium (RPE)/choroid/sclera complex of mice. Data are mean ± s.e.m. of four biological replicates. c Representative immunofluorescence images (left) and quantification (right) of RPE flatmounts of treated mice. Red indicates RPE65, green indicates ZO-1 and blue indicates DAPI. RPE65+ cells are calculated from four fields per RPE flatmount (n = 4 flatmounts). Mean ± s.e.m. is shown. Scale bar, 1000 μm for full field scan; 50 μm for magnified field. d Droplet digital PCR quantification of mRPE65 cDNA in RPE/choroid/sclera complex of rd12 mice. Data are mean ± s.e.m. of three biological replicates. Statistical analysis was performed by two-sided t-test.
Representative wave form of mice from each group upon light stimulus of 1.0 log (cd s/m2) (a), the average b-wave and a-wave amplitudes upon the indicated light stimulus (b), the average b-wave and a-wave amplitudes of mice from each group at the age of 5 weeks and 14 weeks upon light stimulus of 1.0 log (cd s/m2) (c) in scotopic electroretinography (ERG) assay (n = 6 in each group). Mean ± s.e.m. are shown. Representative wave form of mice from each group upon light stimulus of 1.5 log (cd s/m2) (d), the average b-wave and a- wave amplitudes upon the indicated light stimulus (e), the average b-wave and a-wave amplitudes of mice from each group at the age of 5 weeks and 14 weeks upon light stimulus of 1.5 log (cd s/m2) (f) in photopic ERG assay (n = 6 in each group). Mean ± s.e.m. is shown. g Representative immunofluorescence images (left) and quantification (right) of retina flatmounts in mice from each group. Red indicates M-opsin, green indicates peanut agglutinin (PNA)-lectin and blue indicates DAPI. The M-opsin positive area of each flatmount is calculated (n = 3 flatmounts). Mean ± s.e.m. is shown. Scale bar, 1000 μm. h Magnified view of M-opsin positive (red) and PNA-lectin positive (green) cells in the retina of mice from each group. Scale bar, 50 μm. i The PNA-lectin positive cells in four fields per flatmount were calculated (n = 4 flatmounts), each dot represents the quantification result of a field. Mean ± s.e.m. are shown. Statistical analysis was performed by one-way ANOVA followed by two-sided Dunnett’s multiple comparisons (c, f), two-sided t-test (g, i) and two-tailed Mann–Whitney U-test (b, e).
scAAV8.A4T1 restored vision-guided behavior in rd12 mice
To test whether the preserved retinal function leads to an improvement in vision, we first performed two modified visual cliff tests, which depend on animals’ ability to discriminate the deep side of a visual cliff field. In the first test, the mice were placed on a small platform that enabled them to choose to step into the safe or unsafe zone (Fig. 4a). In the 10 step-down test, the WT mice preferred to enter the safe zone (safe zone: 6.7 ± 0.8; unsafe zone: 2.5 ± 0.3; cliff 0.8 ± 0.5) while the untreated mice showed a preference for staying on the cliff (safe zone: 2.3 ± 0.6; unsafe zone: 2.5 ± 0.7; cliff 5.2 ± 1.1), indicating this test is suitable for evaluating the vision-guided behavior of rd12 mice (Supplementary Fig. 8a). We observed that the number of choices for staying on the cliff was significantly decreased in scAAV8.A4T1 treated mice (1.4 ± 0.7) and scAAV8.TCT-1 treated mice (2.0 ± 0.7) compared with the untreated mice (Supplementary Fig. 8a). More importantly, the scAAV8.sup-tRNAArg treated mice had an increased success rate compared with that of the untreated mice (46.2 ± 6.0% (scAAV8.TCT-1) and 61.3 ± 6.3% (scAAV8.A4T1) vs 23.3 ± 5.6% (untreated), Fig. 4b). Moreover, the mice that received scAAV8.sup-tRNAArg treatment spent less time making choices (Supplementary Fig. 8b).
a Schematic of the choice preference test on visual cliff apparatus. b The rate for choosing safe zone (success rate) of each mouse (n = 6 for WT and untreated group, n = 8 for scAAV8.sup-tRNAArg treated group). Data are shown as mean ± s.e.m. c Schematic of the open field test on visual cliff apparatus. Created in BioRender. Yang, Y. (2025) https://BioRender.com/sb411h1. d The representative heatmap representing time spent at each position related to the place preference of mice in each group. e The cumulative time spent in the unsafe zone of mice in each group (n = 12). Mean ± s.e.m. is shown. f Schematic of the modified Morris water maze. Created in BioRender. Yang, Y. (2025) https://BioRender.com/sb411h1. g Representative track plots of mice from each group at the age of 14 weeks. Created in BioRender. Yang, Y. (2025) https://BioRender.com/sb411h1. h Statistical results of success rate to locate the platform within 1 min for mice in each group (n = 6). i Statistical results of the escape latency from each attempt (n = 6 mice per group). Statistical results of the time that each mouse spent in every single test (n = 6). Each mouse complete four tests. Data are mean ± s.e.m. Statistical analysis was performed by two-sided t-test.
The second test was an open-field test. The mice could move freely in both safe and unsafe zones (Fig. 4c). The cumulative time that the mice spent in the safe or unsafe zone was recorded. Compared with the untreated mice, the WT mice spent significantly less time in the unsafe zone (154.5 ± 10.4 s vs 92.8 ± 14.7 s, Fig. 4d, e). Comparing with untreated or scAAV8.TCT-1-treated rd12 mice, the mice with scAAV8.A4T1 treatment spent significantly less time in unsafe zone (scAAV8.A4T1 vs untreated: 81.8 ± 11.5 s vs 154.5 ± 10.4 s; scAAV8.A4T1 vs scAAV8.TCT-1: 81.8 ± 11.5 s vs 132.2 ± 15.1 s). The mice with scAAV8.TCT-1 treatment showed only slight improvement compared with the untreated (132.2 ± 15.1 s vs 154.5 ± 10.4 s). Of note, results showed that the routes and the total path they moved were similar to those of WT mice (Supplementary Fig. 9a, b), indicating that the longer time spent by scAAV8.A4T1-treated mice in the safe zone is not due to them passively or randomly standing still, but rather because their ability to distinguish between the safe and unsafe zone improves during continuous movement.
To further confirm the therapeutic effects, we employed a modified Morris water maze test which directly depends on the visual acuity instead of a merged function in vision and memory (Fig. 4f). In the initial evaluation, the WT mice were able to find the visible platform when their first tryout, while the untreated mice were unable to locate the platform. Therefore, the test was suitable for evaluating the therapeutic effects on rd12 mice. We recorded the routes and success rates of each mouse in the four tests. The WT mice had a 91.7% success rate while the untreated mice had only a 12.5% success rate (Fig. 4g, h). Mice treated with scAAV8.TCT-1 (41.7%) and scAAV8.A4T1 (58.3%) had higher success rates compared with the untreated mice (Fig. 4h). The time that a mouse used to step on the platform was also recorded as “escape latency”. WT mice had the shortest escape latency as 19.5 ± 4.0 s (Fig. 4i). Compared with the untreated mice, the scAAV8.A4T1-treated mice had a significantly shorter escape latency (53.8 ± 3.5 s vs 36.8 ± 4.7 s, Fig. 4i). Taken together, these results indicated that scAAV8.A4T1 showed better therapeutic effects and was able to rescue the visual function of rd12 mice.
Safety and long-term evaluation of scAAV8.sup-tRNAArg treatment in mice
The retinal structure of eyes treated with scAAV8.sup-tRNAArg was examined by H&E staining. We did not observe significant structural disturbance following subretinal delivery of scAAV8.sup-tRNAArg (Fig. 5a). The thickness of outer nuclear layer (ONL) did not significantly change at 12 weeks after treatment (Fig. 5a). To evaluate the overall changes at the transcriptional level, we performed mRNA sequencing. In scAAV8.A4T1-treated mouse RPE/choroidal/sclera complex, we detected 26 upregulated and 178 downregulated transcripts compared with those in untreated rd12 mice (Fig. 5b). The upregulated genes were mainly enriched in terms of visual perception and eye development (Fig. 5c), while the downregulated genes were mainly enriched in terms related to immune responses (Fig. 5c), indicating that the inflammation related to absence of RPE6543 is mitigated following scAAV8.A4T1 treatment. Notably, 5 upregulated and 70 downregulated transcripts were detected in RPE/choroidal/sclera complex of mouse treated with scAAV8.A4T1 compared with those treated with scAAV8.TCT-1 (Fig. 5d). The upregulated transcripts were also enriched in term of visual perception (Fig. 5e). Only limited transcriptome changes were detected between scAAV8.TCT-1-treated and untreated mice (Supplementary Fig. 10a). The enriched GO terms did not indicate obvious cytotoxicity as well (Supplementary Fig. 10b). Subsequently, we analyzed whether scAAV8.A4T1 upregulates endogenous NMD substrates that have a UGA normal stop codon44. The mRNA sequencing data revealed that none of the upregulated transcripts were identified as NMD substrates (Fig. 5f), providing evidence against global UGA readthrough. The mRNA levels of BIP and CHOP in the RPE of mice treated with scAAV8.A4T1 were not increased compared with those in untreated mice (Fig. 5g). Transcripts associated with the UPR pathway in both scAAV8.TCT-1- and scAAV8.A4T1-treated group were not significantly altered either (Fig. 5h). To assess the potential readthrough of scAAV8.TCT-1 and scAAV8.A4T1 at natural termination codons (NTCs), we performed ribosome profiling on the retina/RPE of mice that received sup-tRNA administration. The results indicated that neither scAAV8.TCT-1 nor scAAV8.A4T1 induced significant readthrough at NTCs (Fig. 5i). Previous studies have shown that aminoglycosides such as G418 perturb ribosome elongation at various codons19,40. In contrast, we did not observe such loss of correlation in ribosome occupancy between scAAV.sup-tRNA-treated and untreated mice (Supplementary Fig. 11a, b).
a Representative H&E staining images of retina in untreated (n = 3), scAAV8.TCT-1 treated (n = 5) or scAAV8.A4T1 treated (n = 6) rd12 mice. Scale bar = 50 μm for retinal slices. b Volcano plot demonstrating differentially expressed transcripts revealed by RNA-seq in retinal pigment epithelium (RPE) of mice treated with scAAV8.A4T1 compared with untreated ones (n = 3 per group). Red and green dots denote significantly upregulated and downregulated genes (adjusted P < 0.05 and fold change > 2), respectively. The p values were calculated by two-tailed test and adjusted by Benjamini–Hochberg method. c Gene ontology (GO) enrichment analysis on upregulated and downregulated transcripts in (b). d Differentially expressed transcripts revealed by RNA-seq in RPE of mice treated with scAAV8.A4T1 compared with those treated with scAAV8.TCT-1 (n = 3 per group). The p values were calculated by two-tailed test and adjusted by Benjamini–Hochberg method. e GO enrichment analysis on upregulated and downregulated transcripts in (d). f Venn diagram showing the upregulated transcripts in (b) and published set of 1000 human putative NMD substrates. g qPCR quantification of BIP and CHOP cDNA in RPE of mice in each group (n = 3 mice per group). h Heatmap showing expression level of transcripts in UPR pathway. i Metagene plot (left) showing abundance of ribosome footprints (RFs) relative to the distance from the native termination codon at position 0. Magnified view (right) of the 3’UTR showing moderately increased RFs in this region in scAAV8.sup-tRNAArg treated mouse (n = 3 samples per group). Data are mean ± s.e.m. Statistical analysis was performed by two-sided t-test.
In a cohort of mice followed for 36 weeks post-treatment, the ONL thickness in the scAAV8.A4T1-treated group decreased at a slightly slower rate compared with untreated mice (Supplementary Fig. 12a). The expression level of RPE65 protein (Supplementary Fig. 12b) and the amplitudes of a- and b-waves in scotopic and photopic ERG remained relatively stable over the 36-week period (Supplementary Fig. 12c–e). These results demonstrate the overall safety and sustained effectiveness of subretinal delivery of scAAV8.A4T1 in rd12 mice.
Discussion
In this proof-of-concept study, we demonstrated the therapeutic potential of sup-tRNAArg in suppressing nonsense mutation that causes IRDs in preclinical settings. We screened eleven near-cognate sup-tRNAArg and identified the most efficient one, sup-tRNAArg-TCT-1. By applying a body sequence engineering strategy based on sup-tRNAArg-TCT-1, we generated an optimized variant, sup-tRNAArg-A4T1, which restored 72.3% of full-length RPE65 protein expression in vitro. Using an scAAV-based gene delivery platform, we successfully restored retinal function and vision-guided behavior in rd12 mice without significant toxicity, thus providing a compelling example for the development of sup-tRNA as a potential therapeutic approach for IRDs caused by nonsense mutations in RPE65 gene.
Despite being efficient in in vitro systems, the low suppression efficacy of endogenous PTCs in vivo remains a key challenge for the use of sup-tRNA as a therapeutic strategy. AAV vectors, known for their high transduction efficiency across various tissues, present an ideal delivery system for sup-tRNA19 or engineered tRNA-synthetase-tRNA pairs45. However, the multimeric design of sup-tRNA expression cassettes is required to meet therapeutic demands, albeit at risk of AAV genome truncation19. To overcome this limitation, we introduced a sequence engineering strategy that bypasses this risk and reaches clinically relevant therapeutic thresholds in rd12 mice. A recent study by Porter and co-workers screened the sup-tRNA library to optimize their function23. The results showed that specific mutations in TΨC-stem of sup-tRNAArg (C49-G65, C50-G64, G51-C63) enhanced readthrough activity, which is similar with our findings for sup-tRNAArg-A4T1 (G49-C65, C50-G64, G51-C63). Mechanistically, the TΨC-stem with high GC content had a lower ΔG36, representing a more stable tRNA-eEFIA complex46. This underscores the potential of sequence engineering in optimizing sup-tRNA functionality for therapeutic purposes.
Safety evaluation is critical for the clinical translation of sup-tRNA-based therapeutics for inherited human diseases. The sequence engineering strategy exhibited a good safety profile as no gross toxicity, NTC readthrough or induction of the UPR pathway was observed in rd12 mice treated with scAAV8.A4T1. The readthrough of NTC may elicit immune responses47. The results of RNA-seq showed no significant immune activation was detected in the RPE of mice treated with scAAV8.A4T1 or scAAV8.TCT-1. Two main factors confer a relatively safe nonsense suppression profile on sup-tRNA: 1. sup-tRNA only cognate to a single stop (i.e. UGA in our study), preventing readthrough of a major fraction of transcripts which employ multiple in-frame stop codons48,49; 2. The 3’-UTR region near the NTC can form specific secondary structure50 or bind proteins which promote translation termination51,52. While AAV vectors are generally non-pathogenic, high-dose applications can trigger immune responses53. This sequence engineering strategy allowed the use of reduced amounts of sup-tRNAArg (Supplementary Fig. 3c) and scAAV8.sup-tRNAArg (Supplementary Fig. 6c, d) to achieve similar efficacy, therefore enabling scAAV8.sup-tRNAArg to be administered at a safer dosage, especially for IRDs with higher therapeutic thresholds.
Although the sequence engineering strategy enabled sup-tRNAArg to restore visual function in rd12 mice efficiently and safely, the specific mutations we generated may not be universally applicable to sup-tRNAs carrying other amino acids. For example, similar engineering strategies targeting the TΨC-stem in sup-tRNAGly and sup-tRNALeu resulted in minimal improvements in readthrough efficiency20,23. This suggests that the role of TΨC-stem in nonsense mutation suppression may involve mechanisms beyond stability regulation, such as the recognition of sup-tRNA by aminoacyl-tRNA synthetase. Future strategies for optimizing tRNAs that do not benefit from TΨC-stem engineering should consider alternatives, such as optimizing the 5’-upstream control element and 3’-trailer sequences that govern tRNA transcription and processing23.
The anatomical accessibility and immune-privileged status of the eye make it an ideal target for gene therapy. Given the limitations observed in long-term follow-up studies of RPE65 gene therapy, considerable efforts have been directed toward developing alternative strategies for IRDs. Base editing and prime editing yield promising outcomes in various IRDs models54,55,56,57,58, though CRISPR-based genome editing requires careful evaluation of sgRNA/pegRNA specificity for each target site. Off-target effects, although rare, remain a major safety concern due to the irreversible nature of genome editing. In contrast, the scAAV8.A4T1 may avoid triggering immune responses or inducing permanent genomic alterations, offering an alternative to genome editing. Although initially developed for LCA2, scAAV8.A4T1 also restored R681X-EGFP expression in photoreceptors with higher efficacy than scAAV8.TCT-1 (Supplementary Fig. 7b). This cross-context activity underscores the potential of scAAV8.A4T1 as a versatile approach for treating inherited retinal diseases caused by UGA nonsense mutations, while also highlighting the need for further optimization to achieve robust efficacy across different retinal cell types.
Methods
Plasmid construction
To assess the effectiveness of sup-tRNAs in premature termination codon readthrough, we constructed an EGFP reporter plasmid containing the CMV promoter and SV40 polyA. We inserted 45 bp of RPE65 cDNA sequence of rd12 mice containing PTC site downstream of the start codon of EGFP cDNA (Supplementary Data 1). A plasmid containing the RPE65 cDNA sequence of wild-type mice was used as a control. The hU6-driven sup-tRNAArg plasmids were constructed with pU6-Sp-pegRNA-GG-acceptor (#132777), which was obtained from Addgene (Watertown, MA). The sup-tRNA oligos were ordered and ligated with vector by T4 DNA ligase (New England Biolabs, M0202T) as previously mentioned59. To test readthrough efficiency of sup-tRNAArg more accurately, we constructed a dual luciferase reporter plasmid expressing the firefly luciferase (Fluc, as transfection control) and Gaussia luciferase (Gluc). The sequences of two luciferase were separated by the P2A sequence and part of RPE65 cDNA sequence of wild-type or rd12 mice (Supplementary Data 1).
To express sup-tRNAArg in vivo, we constructed two AAV vector constructs (pAAVsc.sup-tRNAArg-TCT-1 and pAAVsc.sup-tRNAArg-A4T1). Specifically, the sequence of sup-tRNAArg was cloned into a plasmid containing the hU6 promoter and U6 terminator sequence flanked by scAAV2 ITRs. To express ABCA4-EGFP reporter in vivo, we inserted 45 bp of human ABCA4 cDNA sequence containing R681/R681X downstream of the start codon of EGFP cDNA. The sequences were cloned into a plasmid containing the human rhodopsin kinase promoter flanked by scAAV2 ITRs (Supplementary Data 1).
To assess the restoration of full-length protein expression by sup-tRNAs in different diseases caused by nonsense mutations, we constructed two plasmids expressing mouse RPE65 (mRPE65) cDNA carrying PTC site (TGA) at codon 44 and human ABCA4 (hABCA4) cDNA carrying PTC site (TGA) at codon 681.
All the constructed plasmids were verified via Sanger sequencing.
Cell culture and transfection
Human embryonic kidney 293 (HEK293) cells (ATCC, Manassas, USA, Cat. No. CRL-1573) were cultured with DMEM (Gibco, Cat. No. 1190592) supplemented with 10% fetal bovine serum (FBS) (Gibco, Cat. No. A567071) and 1% PS (100 U/mL penicillin/streptomycin) (Boster Biological Technology Co. Ltd., Cat. No. 17I07A16) at 37 °C with 5% CO2. HEK293 cells were transfected using TransIT-X2® (Mirus Bio, Cat. No. MIR 6000). G418 (Life Technologies, Cat. No. 10131027) was used at a final concentration of 0.1 mg/mL.
Dual luciferase assay
HEK293 cells were seeded in 96-well cell culture plates (Corning, Cat. No. 3300) at the density of 4 × 104 cells per well and grown in DMEM supplemented with 10% FBS and 1% PS at 37 °C with 5% CO2. 16—24 h later, the dual luciferase reporter plasmid and plasmid expressing sup-tRNAArg were co-transfected into cells using TransIT-X2®. At 24 h post-transfection, the cell lysates and culture media were collected. Fluc was measured with the Pierce Firefly Luciferase Glow Assay Kit (Thermo Scientific, Cat. No. 16177) and Gluc was measured with the Pierce Gaussia Luciferase Flash Assay kit (Thermo Scientific, Cat. No. 16159) using a plate reader (SpectraMax iD3). The ratio of Gluc over Fluc with the RPE65-R44X construct, normalized to that of the RPE65-R44 construct, reflects readthrough efficiency.
Flow cytometry
The cells were detached with trypsin and collected in flow cytometry tubes. The cell pellets were collected after centrifugation and resuspended in PBS (Gibco, Cat. No. 14190144). Flow cytometry was performed on an Acea NovoCyte Flow Cytometer (Agilent) and the data were analyzed using the Flowjo software. The signal of EGFP positive cells with the RPE65-R44X construct normalized to that of the RPE65-R44 construct reflects the effectiveness of sup-tRNAs in premature termination codon readthrough.
Immunoprecipitation and mass spectrometry
Plasmids expressing sup-tRNAs (TCT-1 or A4T1) and mRPE65 cDNA (R44 or R44X) fused to Flag-tag, respectively, were co-transfected in HEK293 cells. Cells were collected 48 h after transfection and lysed in IP lysis buffer (Thermo Scientific, Cat. No. 87787). mRPE65-Flag protein was purified with anti-Flag magnetic agarose (Thermo Scientific, Cat. No. A36797) according to the manufacturer’s protocol. The purified protein was subjected to SDS-PAGE and gels were stained by Coomassie blue to visualize the target bands. The bands were cut and in-gel digested with Pepsin, and the peptides were desalted using a C18 desalting column (Thermo Scientific, Cat. No 89873) and lyophilized in a SpeedVac and resuspended in 10 μl of 0.1% (v/v) formic acid for mass spectrometry analysis. Data were acquired using an Orbitrap Fusion Lumos Tribrid mass spectrometer (Thermo Fisher Scientific). Raw data were processed using Thermo Proteome Discoverer 3.0 (Thermo).
CCK8 assay
The cells were seeded in 96-well cell culture plates at 4 × 104 cells per well. 16–24 h later, the cells were transfected with sup-tRNAArg-TCT-1, sup-tRNAArg-A4T1 or EGFP. The cell culture medium was replaced with new medium containing CCK-8 (Yeasen, Cat. No. 40203ES76) and incubated in the dark. The value of optical density was measured at 460 nm using a plate reader (SpectraMax iD3). The ratio of the difference between the transfected plasmid-treated cells and blank well (only cell culture medium and CCK-8) to the difference between un-transfected plasmid cells and blank wells reflects the cell viability.
Western blot
The cells were lysed in 50 μL of ice-cold RIPA buffer (Beyotime, Cat. No. P0013B) containing protease inhibitors (Sigma, Cat. No. 539136) at 4 °C for 45 min with vortexing every 15 min to produce protein from the cell lysates. The RPE was added to 30 μL of RIPA buffer containing protease inhibitors before homogenization using a motor tissue grinder (Omni International Corp.), and the mixture was maintained with constant agitation for 1 h at 4 °C to prepare the protein from the posterior eye of mouse. Then, the lysates were centrifuged for 15 min at 17,000 × g and 4 °C. A BCA protein assay (Thermo Scientific, Cat. No. 23225) was used to quantify the total protein concentration. SDS-PAGE was used to separate the samples (20 μg of the cell lysate or the RPE), which were then transferred to PVDF membranes. The membranes were incubated with primary antibodies at 4 °C overnight after being blocked with 5% nonfat milk in TBST at RT for 2 h. Rabbit anti-RPE65 monoclonal antibody (1:1000; Abcam, Cat. No. ab231782), mouse anti-GAPDH polyclonal antibody (ABclonal, Cat. No. AC002), mouse anti-ABCA4 antibody (1:1000; Santa Cruz, Cat. No. SC-65672) and anti-β-Actin monoclonal antibody (ABclonal, Cat. No. AC026) were the primary antibodies used in this study. After being washed in TBST, the membranes were incubated with an HRP-conjugated secondary antibody at RT for 1 h. The secondary antibodies included goat anti-rabbit IgG (1:10,000; Zsbio, Cat. No. ZB2301) and goat anti-mouse IgG (1:10,000; Zsbio, Cat. No. ZB2305). The blots were imaged with an iBrightTM CL1000 imaging system (Thermo Scientific). The data were analyzed using the iBright analysis software version 5.4.0.
AAV vector production
All AAV8 vectors were produced by triple plasmid transfection of HEK293 cells as previously described60. Briefly, 10-layer cell stacks (Corning, Cat. No. 3270) were seeded with 3 × 108 HEK293 cells. PEI-based transfections were carried out when the cell confluence reached 75 ~ 90%. The plasmids were used at a ratio of 2:1:1. The PEI-Max/DNA ratio was maintained at 2:1 (w/w). For each cell mixture, the plasmids and PEI-Max were mixed, vortexed and incubated for 15 min at RT before being added to 1 liter of serum-free DMEM (SFM). The PEI-Max/DNA mixture was added to the stack in place of the culture medium, and it was then incubated at 37 °C with 5% CO2. Then, 500 mL of SFM was added at 72 h after transfection, and the mixture was incubated for an additional 48 h. The virus was subsequently harvested and purified via iodixanol gradient ultracentrifugation. Digital droplet polymerase chain reaction (ddPCR) was used to measure the genome titer (GC/mL) of the AAV8 vector, with the forward primers, reverse primers, and probes (Supplementary Data 2). All vectors used in this study passed the endotoxin assay via a QCL-1000 Chromogenic LAL Test Kit (Cambrex, East Rutherford, NJ).
Animals
Rd12 mice were obtained from the Jackson Laboratory (Bar Harbor). WT C57BL/6 J mice were obtained from Chengdu Dossy Experimental Animals Co., Ltd. (Chengdu, China). The animals were maintained under a 12 h light/dark cycle with unrestricted access to food and water at the temperature around 20–24 °C with 40–60% humidity. In this study, the mice were anesthetized intraperitoneally with xylazine (12 mg/kg) and ketamine (80 mg/kg) unless otherwise stated. The pupils were dilated with an eye drop that included 0.5% tropicamide and 0.5% phenylephrine hydrochloride. The Institutional Animal Care and Concern Committee of West China Hospital Sichuan University gave its approval for the animal experiments, and the committee’s rules were followed when caring for the animals.
Subretinal injection
At the age of two weeks, subretinal injections were performed on the mice under a stereomicroscope (Leica). A 31-gauge needle was used to create a limbal hole after isoflurane inhalation anesthesia and pupil dilatation. The subretinal space was targeted by a blunt 33-gauge needle (Hamilton Co., Cat. No. 80011&780305) inserted through the hole without lens damage. Each eye of a mouse was injected with 1 μL of scAAV8 in three doses (1 × 108 GC/eye, 3 × 108 GC/eye and 1 × 109 GC/eye). Immediately after injection, the fundus was observed via fundus imaging microscopy (Micron IV, Phoenix Research Labs). Successful injections were those that created subretinal blebs but no significant vitreous hemorrhage or subretinal bleeding. Following fundus observation, the cornea was treated with ofloxacin eye ointment.
Electroretinogram (ERG)
Scotopic and photopic ERG were used to evaluate the retinal function of the mice. ERG was recorded according to the manufacturer’s instructions for the Phoenix Ganzfeld ERG (Phoenix Research Labs, Pleasanton, CA). The mice were dark adapted for 16 hours, after which all preparations were carried out in dim red light. A heating pad was used to maintain the body temperature of the mice after anesthesia by intraperitoneal ketamine/xylazine. The ground electrode was positioned subcutaneously in the tail, whereas the reference electrode was positioned subcutaneously on the forehead between the ears. After pupil dilation with tropicamide, a corneal electrode with 2.5% hypromellose was placed on the cornea. Scotopic ERG was recorded at four stimulus intensities (0.1, 0.7, 1.0 and 1.3 log cd s/m2) to evaluate the retinal function of the mice. Photopic ERG was stimulated with green light (wavelength, 504 nm) and photopic recordings were performed after a 5 min light adaptation at an intensity of 1.3 log cd s/m2, which also served as background light for duration of the optical recording. Photopic ERG was recorded at three stimulus intensities: 0.3, 0.9 and 1.5 log cd s/m2.
Immunofluorescence of the RPE and retina
After enucleation, the whole RPE and the retina were carefully separated, and four radial cuts were performed toward the optic nerve head to flatten the RPE and retina. The RPE or retina was fixed in 4% paraformaldehyde in 0.1 M PBS for 1 hour at RT before being rinsed three times for 10 min each time. The samples were the blocked with 5% goat serum for 1 hour at RT before being stained with primary antibody for 24 h at 4 °C. The primary antibodies for the RPE included rabbit anti-RPE65 (1:500; Abcam, Cat No. ab231782) and mouse anti ZO-1 (1:1000; Thermo Scientific, Cat. No. 339100); those for the retina included rabbit anti-M opsin (1:1000 dilution; Novus Biologicals, Cat. No. NB11074730) and FITC-conjugated peanut agglutinin (1:200; Sigma, L7381). Following washing three times in PBST, the samples were incubated with secondary antibodies for 1 h at RT. Goat anti-rabbit IgG Alexa Fluor 594 (1:1000; Abcam, Cat. No. ab150080), goat anti-mouse IgG Alexa Fluor 488 (1:1000; Abcam, Cat. No. ab150113) and goat anti-rabbit IgG Alexa Fluor 488 (1:1000; Abcam, Cat. No. ab150077) were the secondary antibodies employed in this study. The samples were stained with DAPI and washed three times before being mounted. The samples were the photographed using a confocal laser microscope (Nikon).
For frozen cross sections, eyes were carefully enucleated, fixed in 4% paraformaldehyde in 0.1 M PBS for 1 h at RT before being rinsed three times for 10 min each time. The center of the cornea was cut off and eyes were cryoprotected in 30% sucrose/PBS overnight at 4 °C. Then the eye cups were embedded in O.C.T. (Sakura Finetek USA, Inc.) and were cut into retinal slices with a thickness of 10 μm. The retinal slices were rinsed with PBS for 5 min and were blocked with 5% normal goat serum and 0.2% Triton X-100 in PBS for 1 h at RT. After that, the retinal slices were incubated with primary antibodies (anti-GFP, Novus Biologicals, Cat. No. NB600) in blocking solution overnight at 4 °C. Following washing three times in PBST, the samples were incubated with secondary antibodies (1:1000; Abcam, Cat. No. ab150080) for 1 hour at RT. The samples were then stained with DAPI and photographed using a confocal laser microscope (Nikon).
Visual cliff test
A visual cliff test and a step-down trial were used to assess visual function as previously described. The apparatus consisted of two plastic boxes. Half of the top of the first box (length: 50 cm; width: 40 cm; height: 50 cm) is covered, whereas the other half is open. On the top of the covered half, a checkerboard pattern with black and white squares (2 cm × 2 cm) was settled, which continued to the bottom of the opened half. The second box had an open side facing upward, a transparent bottom, and black walls around it. The second box was placed on the top of the first box. In this setting, the second box represented an open field composed of two equal sections: the section with the checkerboard pattern on the top of the first box (length: 45 cm; width:20 cm), called the “safe zone” (or “shallow end”), and the section with a cliff between the transparent bottom of the top box and the checkboard-patterned bottom, called the “unsafe zone” (or “deep end”). The homogenous luminance in the apparatus was set at 10 lux at the height of the mouse eyes.
For the choice preference test, a rising platform was placed between the checkboard-patterned area and the transparent area to create the illusion of a cliff. Each mouse performed 10 trials, and if the mouse moved toward the checkboard-patterned area, the choice was safe (succeed); if the mouse moved toward the clear area, the choice was unsafe (failed). The time limit for each trial was 5 minutes. Each choice made by the mouse was recorded and analyzed comprehensively. Between each trial, the device was cleaned with water and 70% ethanol. We statistically analyzed the probability of each group of mice selecting the safe zone, unsafe zone and cliff within 5 minutes, as well as the time taken for ten selections.
For the open field test, a mouse was left free to roam the entire open field for 300 s after being set up in the corner of the checkboard pattern. The behavior was recorded by a video camera, and the time that the test animal spent in the safe or unsafe zone was analyzed by ANY maze software (Global Biotech Inc., Shanghai, China). Between each mouse, the device was cleaned with water and 70% ethanol. We statistically analyzed the cumulative time spent in the unsafe zone and the path length of each group of mice in this study.
Morris water maze
The Morris water maze was used to assess the cone-related function under photopic condition as described previously3, with slight modifications. The apparatus is a circular plastic water tank with a visible platform (10 cm in diameter, 0.5 cm above the water level) positioned under a dim-lit (10 lux) condition. The tank was filled with opacified water at 23 ± 2 °C. Briefly, a mouse was placed in the water from one of four equally spaced start locations. Each mouse performed 4 trials. Behavioral data were acquired as the latency to escape to the platform during the trials. The subjects were given up to 60 s to escape during each trial. If they did not escape within the allotted time, they were guided gently to the platform and their escape time was recorded as 60 s. The behavior was recorded by a video camera, and the time to escape was analyzed by ANY maze software (Global Biotech Inc., Shanghai, China). We statistically analyzed the rate of successful escape within 60 s and the average escape time of each group.
Droplet digital PCR
RNA from the mouse RPE was extracted using an AllPrep DNA/RNA Mini Kit (Qiagen, Cat. No. 80204). The RNA was RT using a cDNA Reverse Transcription kit (Takara, Cat. No. RR047A). Mouse RPE65 cDNA was quantified in a multiplexed reaction using Taqman reagents targeting mRPE65 (Azenta Life Sciences) and mGAPDH (Azenta Life Sciences). The primers and probes used are listed in Supplementary Data 2. Droplet digital PCR was performed with a QX200 droplet digital PCR system (Bio-Rad).
RT-qPCR
HEK293 cells were seeded into 12-well plates at 4 × 105 cells per well 24 h prior to transfection. The plasmid expressing full-length mRPE65-R44X cDNA and the plasmid expressing sup-tRNAArg were co-transfected. G418 was added to culture media to a final concentration of 0.1 mg/mL. The cells were lysed 24 hours after the transfection. 1 μg total RNA was reverse transcribed using reverse transcriptase (Takara, Cat. No RR047A) in a 20 μL total volume. Quantitative PCR was performed using a TB Green Premix Ex Taq II kit (Takara, Cat. No. RR820A) on LightCycler® 96 (Roche). The GAPDH transcript was used as a housekeeping expression control.
For the animal study, after enucleation, the whole RPE was carefully separated and homogenized via a motor tissue grinder (Omni International Corp.), and 100 ng total RNA was reverse transcribed using reverse transcriptase (Takara, Cat. No. RR047A) in 20 μL total volume. Quantitative PCR was performed using TB Green Premix Ex Taq II kit (Takara, Cat. No. RR820A) on LightCycler® 96 (Roche). GAPDH transcript was used as a house-keeping expression control.
The primers used are listed in Supplementary Data 2. The analysis was performed using the ΔΔCq approach. Technical duplicates of each biological replicate reaction were carried out for each sample.
Histology
Twelve or thirty-six weeks after injection, the eyes were enucleated and fixed in 4% PFA for 24 h and embedded in paraffin. Sectioning and haematoxylin and eosin (H&E) staining were performed. Images were acquired on an Olympus upright microscope. The thickness of the ONL was measured at various distances from optic nerve head by ImageJ software.
mRNA sequencing (RNA-seq)
Total RNA was extracted using RNAsimple Total RNA Kit (DP419, TIANGEN). Libraries were constructed using U-mRNAseq Library Prep Kit (AT4221, KAITAI-BIO with Ribo-off rRNA Depletion Kit (Bacteria) (N407, Vazyme). Libraries were pooled and sequenced using the Illumina NovaSeq machine as 150-bp paired-end sequencing reads. Raw reads were trimmed using fastp (v0.23.0,https://github.com/OpenGene/fastp), low-quality reads and reads with sizes less than 50 bp (--length_required 50) were removed. The reference genome and gene model annotation files were downloaded from the genome website directly (Homo sapiens, Ensembl v101). Clean data were mapped to the reference genome using HISAT2 (v2.2.1,http://daehwankimlab.github.io/hisat2/). Next, the mapped reads of each sample were assembled using StringTie v1.3.6 in a reference-based approach. All RNAs were quantified as fragments per kilobase million mapped reads (FPKM) by StringTie. The formula was defined as FPKM = 106×F/(NL×10−3), where F is the number of fragments assigned to a certain gene in a certain sample, N is the total number of mapped reads in the certain sample, and L is the length of the certain gene. Differentially expressed genes were determined by edgeR (https://bioconductor.org/packages/release/bioc/html/edgeR.html), and genes with FDR < 0.05 and |log2-fold change | >1 were identified as differentially expressed genes.
GO (Gene Ontology, http://geneontology.org/) enrichment analysis was performed for all differentially expressed mRNAs. GO terms with p < 0.05 were considered to be significantly enriched. GO enrichment analysis showed the biological functions of differentially expressed mRNAs. KEGG (Kyoto Encyclopedia of Genes and Genomes, http://www.kegg.jp/) is a database resource that contains a collection of manually drawn pathway maps representing our knowledge of the molecular interaction and reaction networks. Using the same method as GO enrichment analysis, significantly enriched KEGG pathways were identified.
Measurement of tRNA transcript abundance
The self-cleaving ribozyme drz-Bflo-2 was added immediately downstream of sup-tRNA (sup-tRNA-RZ) sequence as previously described23 for quantification of the transcript abundance of sup-tRNAArg-TCT-1 or sup-tRNAArg-A4T1. HEK293 cell were seeded into 12-well plate. After 24 h, 200 ng sup-tRNA-RZ expressing plasmid was transfected to the cell. Forty-eight hours post-transfection, RNA was extracted using an AllPrep DNA/RNA Mini Kit (Qiagen, Cat. No. 80204). The RNA was RT using a cDNA Reverse Transcription kit (Takara, Cat. No. RR047A). FAM-labeled RZ probe and FAM-labeled TATA-binding protein (TBP) probes were purchased from Azenta (Supplementary Data 2) and used with TaqMan™ Fast Advanced (Thermo Fisher, Cat. No. 4444556) on an Applied Biosystems QuantStudio 3 Real-Time PCR System.
Ribosome profiling and data analysis
Twelve weeks after injection, the eyes were enucleated and retina/RPE/choroidal/sclera complex was dissociated and immediately frozen in liquid nitrogen and ground to power in liquid nitrogen with a mortar and pestle, then dissolved in 400 μL of lysis buffer.
The ribosomal profiling technique was carried out as reported previously61, with a few modifications as described below. The resuspended extracts in lysis buffer were transferred to new microtubes, pipetted several times and incubated on ice for 10 min. Then cells were triturated ten times through a 26-G needle. The lysate was centrifuged at 20000 g for 10 min at 4 °C, and the supernatant was collected. To prepare RFs, 10 μL of RNase I (NEB, Ipswich, MA, USA) and 6 μL of DNase I (NEB, Ipswich, MA, USA) were added to 400 μL of lysate, which was then incubated for 45 min at room temperature with gentle mixing on a Nutator mixer. Nuclease digestion was stopped by adding 10 μL of SUPERase·In RNase inhibitor (Ambion, Austin, TX, USA). Size exclusion columns (Illustra MicroSpin S-400 HR Columns; GE Healthcare; Cat. No. 27- 5140-01) were equilibrated with 3 mL of polysome buffer by gravity flow and centrifuged at 600 g for 4 min at room temperature. 100 μL of digested RFs were added to the column and centrifuged at 600 g for 2 min. Next, 10 μL 10% (wt/vol) SDS was added to the elution, and RFs with a size greater than 17nt were isolated according to the RNA Clean and Concentrator-25 kit (Zymo Research; Cat. No. R1017). rRNA was removed using the method reported previously62. Briefly, short (50–80 bases) antisense DNA probes complementary to rRNA sequences were added to solution containing RFs, then RNase H (NEB, Ipswich, MA, USA) and DNase I (NEB, Ipswich, MA, USA) were added to digest rRNA and residual DNA probes. Finally, RFs were further purified using magnet beads (Vazyme, Nanjing, Jiangsu, China).
After obtaining ribosome footprints above, Ribo-seq libraries were constructed using NEBNext® Multiple Small RNA Library Prep Set for Illumina® (Cat. No. E7300S, E7300L). Briefly, adapters were added to both ends of RFs, followed by reverse transcription and PCR amplification. The 140–160 bp size PCR products were enriched to generate a cDNA library and sequenced using Illumina NovaSeq X Plus by Gene Denovo Biotechnology Co. (Guangzhou, China).
Low quality reads were filtered by fastp63. Raw reads containing over 50% of low-quality bases or over 10% of N bases were removed. Adapter sequences were trimmed. Reads with length between 20–40 bp were retained for subsequent analysis. Short reads alignment tool Bowtie264 was used for mapping reads to ribosome RNA (rRNA) database, GenBank, Rfam database. The reads mapped to rRNAs, transfer RNAs (tRNA), small nuclear RNAs (snRNA), small nucleolar RNAs (snoRNA), and miRNA were removed. Processed RNA-seq reads were aligned to the genome using whole genome alignment by STAR65 with 2- pass setting enabled. The reads mapped to classical noncoding RNAs (including long noncoding RNAs, microRNA precursors) were abandoned. Then remaining RFs were assigned to different genomic features (5’UTR, coding sequence (CDS), 3’UTR and intron) based on the position of the 5’ end of the alignment. Three-nucleotide periodicity was plotted using the riboWaltz R package66 as described. By using density data, relative 3′ UTR density was calculated by dividing 3′ UTR density by CDS density and plotted. Average codon occupancies were calculated using the ribosome densities of each codon divided by the average density for the CDS of each transcript.
Statistics and reproducibility
Blinding was not used for the study. The experiments were performed by different co-authors. Littermate mice were randomized among experimental groups. Animals were assigned randomly to treated and non-treated groups. Animals used in the same experiment were age and sex matched. Sample sizes were chosen based on other similar studies. No data were excluded from the analyses. The statistical tests used for each experiment are stated in the corresponding figure captions. All statistical analysis was performed by using GraphPad Prism 9.
Ethics statement
The Institutional Animal Care and Concern Committee of West China Hospital, Sichuan University gave its approval for the animal experiments under animal protocol no.20240809009, and the committee’s rules were followed when caring for the animals.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-seq data can be found in the Genome Sequence Archive (GSA) using accession number CRA024668. Mass spectrometry data can be found in ProteomeXchange with accession number PXD062957 [https:// iprox.cn/page/project.html?id=IPX0011710000]. The ribo-seq data are available from GSA under accession number CRA029266. Source data is available for Figs. 1–5 and Supplementary Figs. 1, 3-8, 12 in the associated source data file. Source data are provided with this paper.
References
Antas, P., Carvalho, C., Cabral-Teixeira, J., de Lemos, L. & Seabra, M. C. Toward low-cost gene therapy: mRNA-based therapeutics for treatment of inherited retinal diseases. Trends Mol. Med. 30, 136–146 (2024).
Schneider, N. et al. Inherited retinal diseases: linking genes, disease-causing variants, and relevant therapeutic modalities. Prog. Retin Eye Res. 89, 101029 (2022).
Maquat, L. E. Nonsense-mediated mRNA decay: splicing, translation and mRNP dynamics. Nat. Rev. Mol. Cell Biol. 5, 89–99 (2004).
Guerin, K. et al. Systemic aminoglycoside treatment in rodent models of retinitis pigmentosa. Exp. Eye Res 87, 197–207 (2008).
Goldmann, T., Overlack, N., Wolfrum, U. & Nagel-Wolfrum, K. PTC124-mediated translational readthrough of a nonsense mutation causing Usher syndrome type 1C. Hum. Gene Ther. 22, 537–547 (2011).
Gregory-Evans, C. Y. et al. Postnatal manipulation of Pax6 dosage reverses congenital tissue malformation defects. J. Clin. Investig. 124, 111–116 (2014).
Spelier, S., van Doorn, E. P. M., van der Ent, C. K., Beekman, J. M. & Koppens, M. A. J. Readthrough compounds for nonsense mutations: bridging the translational gap. Trends Mol. Med. 29, 297–314 (2023).
Botto, C. et al. Early and late stage gene therapy interventions for inherited retinal degenerations. Prog. Retin Eye Res. 86, 100975 (2022).
Bainbridge, J. W. et al. Long-term effect of gene therapy on Leber’s congenital amaurosis. N. Engl. J. Med. 372, 1887–1897 (2015).
Jacobson, S. G. et al. Improvement and decline in vision with gene therapy in childhood blindness. N. Engl. J. Med. 372, 1920–1926 (2015).
Wright, C. B. et al. The Rpe65 rd12 allele exerts a semidominant negative effect on vision in mice. Investig. Ophthalmol. Vis. Sci. 55, 2500–2515 (2014).
Gardiner, K. L. et al. Long-term structural outcomes of late-stage RPE65 gene therapy. Mol. Ther. 28, 266–278 (2020).
Wang, D., Zhang, F. & Gao, G. CRISPR-based therapeutic genome editing: strategies and in vivo delivery by AAV vectors. Cell 181, 136–150 (2020).
Capecchi, M. R., Haar, R. A., Capecchi, N. E. & Sveda, M. M. The isolation of a suppressible nonsense mutant in mammalian cells. Cell 12, 371–381 (1977).
Chang, J. C., Temple, G. F., Trecartin, R. F. & Kan, Y. W. Suppression of the nonsense mutation in homozygous beta 0 thalassaemia. Nature 281, 602–603 (1979).
Temple, G. F., Dozy, A. M., Roy, K. L. & Kan, Y. W. Construction of a functional human suppressor tRNA gene: an approach to gene therapy for beta-thalassaemia. Nature 296, 537–540 (1982).
Bordeira-Carrico, R. et al. Rescue of wild-type E-cadherin expression from nonsense-mutated cancer cells by a suppressor-tRNA. Eur. J. Hum. Genet. 22, 1085–1092 (2014).
Lueck, J. D. et al. Engineered transfer RNAs for suppression of premature termination codons. Nat. Commun. 10, 822 (2019).
Wang, J. et al. AAV-delivered suppressor tRNA overcomes a nonsense mutation in mice. Nature 604, 343–348 (2022).
Albers, S. et al. Engineered tRNAs suppress nonsense mutations in cells and in vivo. Nature 618, 842–848 (2023).
Ogawa, A., Hayami, M., Sando, S. & Aoyama, Y. A concept for selection of codon-suppressor tRNAs based on read-through ribosome display in an in vitro compartmentalized cell-free translation system. J. Nucleic Acids 2012, 538129 (2012).
Albers, S. et al. Repurposing tRNAs for nonsense suppression. Nat. Commun. 12, 3850 (2021).
Porter, J. J., Ko, W., Sorensen, E. G. & Lueck, J. D. Optimization of ACE-tRNAs function in translation for suppression of nonsense mutations. Nucleic Acids Res. 52, 14112–14132 (2024).
Pang, J. J. et al. Retinal degeneration 12 (rd12): a new, spontaneously arising mouse model for human Leber congenital amaurosis (LCA). Mol. Vis. 11, 152–162 (2005).
Shi, J. et al. Clinical features and natural history in a cohort of chinese patients with RPE65-associated inherited retinal dystrophy. J. Clin. Med. 10, https://doi.org/10.3390/jcm10225229 (2021).
Chung, D. C. et al. The natural history of inherited retinal dystrophy due to biallelic mutations in the RPE65 gene. Am. J. Ophthalmol. 199, 58–70 (2019).
Kompatscher, M. et al. Contribution of tRNA sequence and modifications to the decoding preferences of E. coli and M. mycoides tRNAGlyUCC for synonymous glycine codons. Nucleic Acids Res. 52, 1374–1386 (2024).
Ledoux, S., Olejniczak, M. & Uhlenbeck, O. C. A sequence element that tunes Escherichia coli tRNA(Ala)(GGC) to ensure accurate decoding. Nat. Struct. Mol. Biol. 16, 359–364 (2009).
Giege, R. & Eriani, G. The tRNA identity landscape for aminoacylation and beyond. Nucleic Acids Res. 51, 1528–1570 (2023).
Koukuntla, R., Ramsey, W. J., Young, W. B. & Link, C. J. U6 promoter-enhanced GlnUAG suppressor tRNA has higher suppression efficacy and can be stably expressed in 293 cells. J. Gene Med. 15, 93–101 (2013).
Bharti, N. et al. Translation velocity determines the efficacy of engineered suppressor tRNAs on pathogenic nonsense mutations. Nat. Commun. 15, 2957 (2024).
Geslain, R. & Pan, T. Functional analysis of human tRNA isodecoders. J. Mol. Biol. 396, 821–831 (2010).
Nguyen, H. A., Sunita, S. & Dunham, C. M. Disruption of evolutionarily correlated tRNA elements impairs accurate decoding. Proc. Natl. Acad. Sci. USA 117, 16333–16338 (2020).
Schrader, J. M., Chapman, S. J. & Uhlenbeck, O. C. Tuning the affinity of aminoacyl-tRNA to elongation factor Tu for optimal decoding. Proc. Natl. Acad. Sci. USA 108, 5215–5220 (2011).
Yarus, M., Cline, S., Raftery, L., Wier, P. & Bradley, D. The translational efficiency of tRNA is a property of the anticodon arm. J. Biol. Chem. 261, 10496–10505 (1986).
Uhlenbeck, O. C. & Schrader, J. M. Evolutionary tuning impacts the design of bacterial tRNAs for the incorporation of unnatural amino acids by ribosomes. Curr. Opin. Chem. Biol. 46, 138–145 (2018).
Huang, L. et al. Targeting translation termination machinery with antisense oligonucleotides for diseases caused by nonsense mutations. Nucleic Acid Ther. 29, 175–186 (2019).
Linde, L. et al. Nonsense-mediated mRNA decay affects nonsense transcript levels and governs response of cystic fibrosis patients to gentamicin. J. Clin. Investig. 117, 683–692 (2007).
Keeling, K. M. et al. Leaky termination at premature stop codons antagonizes nonsense-mediated mRNA decay in S. cerevisiae. RNA 10, 691–703 (2004).
Wangen, J. R. & Green, R. Stop codon context influences genome-wide stimulation of termination codon readthrough by aminoglycosides. Elife 9, https://doi.org/10.7554/eLife.52611 (2020).
Phillips-Jones, M. K., Hill, L. S., Atkinson, J. & Martin, R. Context effects on misreading and suppression at UAG codons in human cells. Mol. Cell Biol. 15, 6593–6600 (1995).
Blanks, J. C. & Johnson, L. V. Specific binding of peanut lectin to a class of retinal photoreceptor cells. A species comparison. Investig. Ophthalmol. Vis. Sci. 25, 546–557 (1984).
Montaser, A. B. et al. Retinal proteome profiling of inherited retinal degeneration across three different mouse models suggests common drug targets in retinitis pigmentosa. Mol. Cell Proteom. 23, 100855 (2024).
Colombo, M., Karousis, E. D., Bourquin, J., Bruggmann, R. & Muhlemann, O. Transcriptome-wide identification of NMD-targeted human mRNAs reveals extensive redundancy between SMG6- and SMG7-mediated degradation pathways. RNA 23, 189–201 (2017).
Shi, N. et al. Restoration of dystrophin expression in mice by suppressing a nonsense mutation through the incorporation of unnatural amino acids. Nat. Biomed. Eng. 6, 195–206 (2022).
LaRiviere, F. J., Wolfson, A. D. & Uhlenbeck, O. C. Uniform binding of aminoacyl-tRNAs to elongation factor Tu by thermodynamic compensation. Science 294, 165–168 (2001).
Porter, J. J., Heil, C. S. & Lueck, J. D. Therapeutic promise of engineered nonsense suppressor tRNAs. Wiley Interdiscip. Rev. RNA 12, e1641 (2021).
Brown, C. M., Dalphin, M. E., Stockwell, P. A. & Tate, W. P. The translational termination signal database. Nucleic Acids Res. 21, 3119–3123 (1993).
Dalphin, M. E., Brown, C. M., Stockwell, P. A. & Tate, W. P. The translational signal database, TransTerm, is now a relational database. Nucleic Acids Res. 26, 335–337 (1998).
Firth, A. E., Wills, N. M., Gesteland, R. F. & Atkins, J. F. Stimulation of stop codon readthrough: frequent presence of an extended 3’ RNA structural element. Nucleic Acids Res. 39, 6679–6691 (2011).
Biziaev, N. et al. The impact of mRNA poly(A) tail length on eukaryotic translation stages. Nucleic Acids Res. 52, 7792–7808 (2024).
Ivanov, A. et al. PABP enhances release factor recruitment and stop codon recognition during translation termination. Nucleic Acids Res. 44, 7766–7776 (2016).
Wang, D., Tai, P. W. L. & Gao, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat. Rev. Drug Discov. 18, 358–378 (2019).
Suh, S. et al. Restoration of visual function in adult mice with an inherited retinal disease via adenine base editing. Nat. Biomed. Eng. 5, 169–178 (2021).
She, K. et al. Dual-AAV split prime editor corrects the mutation and phenotype in mice with inherited retinal degeneration. Signal Transduct. Target Ther. 8, 57 (2023).
Muller, A. et al. High-efficiency base editing in the retina in primates and human tissues. Nat. Med. 31, 490–501 (2025).
Su, J. et al. In vivo base editing rescues photoreceptors in a mouse model of retinitis pigmentosa. Mol. Ther. Nucleic Acids 31, 596–609 (2023).
Choi, E. H. et al. In vivo base editing rescues cone photoreceptors in a mouse model of early-onset inherited retinal degeneration. Nat. Commun. 13, 1830 (2022).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Lock, M. et al. Rapid, simple, and versatile manufacturing of recombinant adeno-associated viral vectors at scale. Hum. Gene Ther. 21, 1259–1271 (2010).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Morlan, J. D., Qu, K. & Sinicropi, D. V. Selective depletion of rRNA enables whole transcriptome profiling of archival fixed tissue. PLoS One 7, e42882 (2012).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Lauria, F. et al. riboWaltz: optimization of ribosome P-site positioning in ribosome profiling data. PLoS Comput. Biol. 14, e1006169 (2018).
Acknowledgements
This study is supported by the National Natural Science Foundation of China (No.82222030, U22A20311, 82201212). National Key R&D Program of China (No.2023YFC3403300, 2023YFC3403302 and 2024YFC3406704). Sichuan Science and Technology Program (No. 2025ZNSFSC1710). China Postdoctoral Science Foundation (No. 2024M762227). Natural Science Foundation of Sichuan Province (No. 2023NSFSC1670). 1.3.5 Project for Disciplines of Excellence from West China Hospital of Sichuan University (No. ZYYC23009) and Postdoctor Research Fund of West China Hospital, Sichuan University (2024HXBH113). All vectors were produced by Chengdu Genevector Biotech (Chengdu, China). We thank Xuejin Lu, a full-time employee of Chengdu Genevector Biotech, for help with the animal studies.
Author information
Authors and Affiliations
Contributions
Y.Y. and F.L. conceived and supervised the study. C.R. and L.S. designed, performed, and analysed the in vivo experiments and generated figures. C.R., L.S., and M.H. designed, performed, and analysed the in vitro experiments and generated figures. X.J. performed the droplet digital PCR experiments. C.R., L.S., M.H., and L.L. performed water maze experiments. C.R., L.S., and M.H. wrote the manuscript; Y.W., F.L., and Y.Y. reviewed and edited the manuscript. Fanfei Liu, J.S., J.F., Yiliu Yang, Q.X., X.W., Q.Y., Man Liu, Min Luo, Q.L., Y.A., Manjun Li, K.S. provided technical support. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.R., F.L., and Y.Y. are inventors on a patent application to the China National Intellectual Property Office pertaining to the sequence engineered sup-tRNA of this work (application number 2025103935536) filed by West China Hospital. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ren, C., Song, L., Hu, M. et al. AAV-delivered engineered suppressor tRNA rescues visual function in mice with an inherited retinal disease. Nat Commun 16, 11185 (2025). https://doi.org/10.1038/s41467-025-66176-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66176-y