Abstract
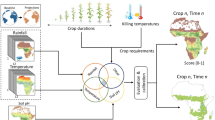
In an era marked by climatic uncertainties and burgeoning food security challenges, climate-crop models emerge as important tools for guiding investments in climate-resilient agriculture. Here, we construct climate-crop model applications for 24 crops spanning the African food basket, including cereals, legumes, oilseeds, roots/tubers, and vegetables, in support of the Vision for Adapted Crops & Soils (VACS) program. Notably, we expand the modeling framework for 19 “opportunity” crops that have hitherto been underrepresented or overlooked but hold potential to bolster agricultural resilience across the continent. We calibrate and parameterize our process-based models using field observations and comprehensive literature reviews. We evaluate the model outputs spatially and temporally against national crop statistics across Africa, showing acceptable reproduction of yields and response to reported extreme droughts and heat waves. This work offers a scalable framework for climate-crop modeling and impact assessments related to food security in Africa and beyond.
Similar content being viewed by others
Introduction
Africa is facing complex and multifaceted food security issues, stemming from a combination of past and present environmental, economic, and social factors1,2,3. One of the primary concerns is the continent’s vulnerability to climate change, which exacerbates existing challenges in agricultural production4,5,6. Africa is experiencing increasingly unpredictable weather patterns, including frequent heat waves, prolonged droughts, and erratic floods, which directly affect major crops such as maize, wheat, and rice6,7,8. These staple crops, vital for the sustenance of the African population, are highly vulnerable to climate change in many African regions7,9. This vulnerability is a critical concern, particularly in light of the projected population increase in Africa, which is expected to double by 205010. Such a demographic surge will inevitably escalate food demand, placing additional pressure on already strained food systems. In response to these challenges, the potential of traditional African crops emerges as a potential viable adaptation strategy11,12,13,14,15. Indigenous crops, such as sorghum, millet, and teff, are often more resilient to harsh weather conditions and poor soil structures (i.e., poor soil water dynamics) and offer a sustainable alternative to conventional staples16,17,18,19. These neglected and underutilized crops, henceforth referred to as “opportunity” crops, have historically been integral to local diets and agricultural practices20,21,22. Their inherent resilience to climate extremes makes them particularly suitable for cultivation in the changing climate23,24. Embracing these opportunity crops can not only contribute to food security, but also promote biodiversity and preserve cultural food practices. Motivated by the potential of opportunity crops, the U.S. Department of State, in partnership with the African Union and the Food and Agriculture Organization (FAO), launched the Vision for Adapted Crops & Soils (VACS, https://www.state.gov/the-vision-for-adapted-crops-and-soils) project. VACS seeks to adapt the world’s agricultural systems to the anticipated challenges of climate change, with an initial focus on the African continent.
Climate-crop models have emerged as indispensable tools in guiding investments towards climate-resilient agriculture, playing a crucial role in adapting agricultural practices to the changing climate9,25. The VACS pilot investigation relies on crop and climate modeling conducted by the Agricultural Model Intercomparison and Improvement Project (AgMIP, https://agmip.org). Since its launch in 2010, AgMIP’s accomplishments include the development of harmonized tools and protocols for analyzing agricultural systems using state-of-the-art models, integrating stakeholder-informed scenarios into global and regional assessments, and providing valuable resources for stakeholders and researchers26. AgMIP has significantly improved agricultural models, and scientific and technological capabilities, for assessing impacts of climate variability and change and other driving forces on agriculture, food security, and poverty at local to global scales27.
Modeling opportunity crops is a challenging task due to the limitations in model and data availability28,29. Although current mainstream process-based crop models, such as the Decision Support System for Agrotechnology Transfer (DSSAT)30 have been developed for dozens of major crops, many African opportunity crops are still overlooked. Input and reference data scarcity limits the development of new crop models in complex model platforms like DSSAT that require a large amount of genotype specific parameters30. Unlike major global crops, opportunity crops in Africa often lack systematic management and yield data collection efforts11. This includes limited field trials, phenotypic data, and genetic information, which are crucial for understanding crop behavior under different environmental conditions. Additionally, historical yield data, which is essential for calibrating and validating models, is often sparse or non-existent11. The situation is further complicated by the diverse agro-ecological zones in Africa and the complex management system, where these crops are grown, requiring region-specific data for accurate modeling31. This lack of data hinders the development of robust climate-crop models, limiting the ability to predict how these crops will respond to various climatic and environmental changes.
In this study, we adapt a generic, but simple dynamic process-based crop simulation model called SIMPLE32 to simulate opportunity crops in Africa. The SIMPLE model employs universally recognized climate-crop-soil processes, necessitating minimal parameters and data, and eschews crop-specific procedures, thereby facilitating broad applicability across a diverse array of crops. We undertake a comprehensive literature review to gather field data related to opportunity crops for model input. The primary objectives of our study are to (1) provide climate-crop model applications for 19 opportunity crops (in addition to 5 staple crops) and (2) develop a robust approach for calibrating genotype-specific parameters of opportunity crops, with focus on fine-tuning the parameters that define the unique genetic characteristics of these crops, tailored to specific environmental interactions and growth patterns. By honing in on the genotype-specific parameters, our approach aims to unlock the agricultural potential of opportunity crops, paving the way for their increased utilization and recognition in the African food system for climate change adaptation.
Results and discussion
Model calibration, validation, and evaluation
We constructed and expanded climate-crop model applications for 24 crops within SIMPLE including cereals, legumes, oilseeds, roots/tubers, and vegetables (e.g., SIMPLE-Fonio). Five of these crops—maize, soybean, groundnut, cassava, and tomato are considered reference crops, and the remaining 19 are opportunity crops33 (Table S1). SIMPLE uses 4 cultivar parameters and 9 species parameters to simulate the phenology, heat and drought tolerance, and photosynthesis rate for a specified crop. The definition of cultivar and species parameters are provided in Table S2. We simulate the period from 2000 to 2019 as the baseline for comparison.
Due to the lack of public opportunity crop observational datasets, we perform a thorough literature review to obtain as many details of crop parameters and field observations as possible. The literature review and crop cultivar and species parameter calibration and validation include the following four steps (Fig. 1):
-
1.
We first search for cultivar and species parameters of SIMPLE/DSSAT from the literature for the opportunity crops in Africa. If the parameters exist, we approximate SIMPLE parameters using the literature data, if not, we search for other existing crop models (e.g., AquaCrop) for the opportunity crops and/or expand to opportunity crop experiments from countries with similar climate to African agricultural regions34 (e.g., lablab in Central India is comparable to Central Africa given the same tropical savanna Köppen-Geiger climate classification35,36,37). If the existing crop model parameters correspond to the SIMPLE parameters, we approximate the SIMPLE parameters using literature data.
-
2.
If no existing cultivar and species parameters are available for the opportunity crops, we search for non-nutrient-stressed field experiment data from the literature that include crop phenology, biomass, and yield. Non-nutrient limiting experiments serve as an excellent source for calibration as SIMPLE does not consider nutrient stresses. We then use the observed data to calibrate the SIMPLE cultivar and species parameters by optimizing Root Mean Square Error (RMSE) and Relative RMSE (RRMSE) across phenology, biomass, and yield. Acknowledging that an experimental site is one example system within a more heterogeneous region, we further validate the model using country-level FAO crop yield data38 (see Methods).
-
3.
If a crop has neither existing cultivar and species parameters nor field-level observed data, we use FAO country-level yield data to calibrate the SIMPLE cultivar and species parameters to optimize the RMSE and RRMSE for the yield time series. We compare the simulated baseline yield to FAO yield data across multiple countries. Given that the FAO reported yield often include nutrient stresses, we acknowledge that the simulated yield may be higher than the FAO amount to represent potential yield and avoid overfitting the model parameters.
-
4.
If none of the above steps are applicable, expert knowledge from opportunity crop breeders can be used to estimate parameters based on analogue crop parameters, and clarify that parameters are theoretical and need to be tested/verified in future experiments. However, in this study, all crops investigated had either field experiment data and/or FAO country level data available.
The decision tree follows the logic of descending reliability to minimize uncertainties from existing model parameters, field experiment data, FAO data, and expert knowledge. a expert knowledge was not the sole justification for the parameters since we have either field experiment data and/or FAO country level data available for all crops investigated, but we still include this panel to make the decision tree complete, b this step is bypassed if there is no existing FAO country/continent level data for a specific crop.
Here we select a representative crop in each crop category (except oilseeds which has only two crops) to demonstrate the model calibration and validation process (Fig. 2), while the remaining crop results are shown in Table S3. The representative crops are chosen based on their high potential as opportunity crops from local experts (see Methods). Overall, the SIMPLE model generally demonstrates satisfactory performance (RRMSE < 30%) for rainfed crop productivity in Africa. Its flexibility, due to the lower required amount of data to run, allows it to handle diverse agricultural systems across the continent’s varied landscapes. The index of agreement (d-index) between the simulated and FAO continent aggregated yields when averaged across all crops is 0.41, indicating moderate agreement (Table S4). This agreement also remains valid when examining the separate crop types, with a d-index of 0.45, 0.43, 0.34, 0.44, and 0.34 for cereals, legumes, oilseeds, roots/tubers, and vegetables, respectively. Additionally, the model effectively captures broad patterns in crop yields and land use (Figs. S1–S6), providing valuable insights into the continent’s agricultural production dynamics. This may help policymakers and researchers understand potential impacts of climate change on food security and further socioeconomic influence and adaptation assessments. Despite challenges posed by limited data availability and the continent’s complex agroecological conditions, the model projections are generally consistent with spatiotemporal patterns (Fig. S7). This reinforces its utility as a decision-making tool for assessing regional agricultural resilience and planning adaptation strategies.
Figure showing the calibration results for four representative crops – fonio, cowpea, yams, and tomato in terms of phenology, biomass, and yield based on existing parameters and field experiments from the literature, and validation based on FAO country-level yield data. The treatments refer to different management practices such as planting date (detailed in Methods). The reason that the simulated results are generally higher than FAO in the validation is mainly attributed to the use of a non-nutrient stressed model, while FAO data is collected from in situ surveys with various environmental stresses including nutrient stress, pest and diseases, etc. and uncertainties in agricultural management38. The calibration results for all crops with field-level data available are shown in the Supplementary Figs. S1–S5. The validation results for all crops with country-level data available are shown in the Supplementary Fig. S6 and Table S4. All yields shown at 0% moisture content. Error bars are not available for the observations.
Analysis of crop cultivar and species parameters
This study quantifies model parameters that reflect the properties of many African opportunity crops. Here we examine in detail three of the most critical SIMPLE parameters for crop growth (Fig. 3): the thermal time requirement from sowing to maturity in daily mean temperature (Tsum), threshold for daily maximum temperature to start accelerating senescence due to heat stress (Tmax), and sensitivity of radiation-use efficiency (RUE) to drought stress (Swater). The comparison of the remaining parameters is shown in Figs. S7 and S8.
Radar charts showing the SIMPLE model cultivar parameter for the thermal time requirement from sowing to maturity (Tsum), and the species parameters for the heat stress threshold (Tmax) and the sensitivity of radiation use efficiency to water stress (Swater) for all 24 crops. The red points denote above average values while the blue points denote below average. The dashed gray line is the mean parameter value of all crops.
Tsum indicates the growth duration for a certain crop via thermal time accumulation, which is a significant index for agricultural planning and management, such as the planting time adjustment and crop rotation. Tsum values for all cereal crops fall below the average, suggesting that cereals generally exhibit shorter growth periods. This characteristic points to a faster development cycle for cereals compared to the other crop types. Conversely, all roots and tubers, except for sweet potatoes, have Tsum values above the average, indicating longer growth durations for these crops. Such extended periods are indicative of a slower maturation process, which may influence planting and harvesting strategies. Notably, among the variety of crops analyzed, cassava records the highest Tsum, showcasing its particularly lengthy growth cycle39, while teff registers the lowest Tsum, confirming its rapid maturation cycle40.
Tmax reflects the crop heat stress tolerance. Crops with high Tmax may have stronger resilience to elevated temperature impacts from climate change. Nearly all cereal crops (except for maize) have an above average Tmax value, suggesting that cereals generally exhibit a strong heat stress tolerance. In contrast, all legume and vegetable crops fall below average, indicating that these crops are more vulnerable to heat stress. Fonio, finger millet, pearl millet, and sesame exhibit the highest Tmax value, indicating that these crops may be particularly well-suited as opportunity crops in the context of global warming41,42,43.
Swater signifies the crop water stress sensitivity, which can be used for assisting cropland allocation and irrigation optimization. Most cereals and legumes have Swater close to the average. A low Swater for crops like teff, okra, and amaranth suggests that these crops can be allocated to relatively arid regions44, while crops with high Swater such as tomato and groundnut may be allocated to humid regions or prioritized with irrigation infrastructure45.
To further validate the genetic differences between the SIMPLE parameterization of the crop types, we show the crop Euclidean distance based on the normalized 9 species parameters (Fig. 4). A small crop distance indicates a strong similarity while a large distance indicates a strong difference (e.g., crop distance of 1 means the two crops are near identical, crop distance of 7 means the two crops are very different). Physically, crops belonging to the same crop type should be more identical than those disparate crops in terms of plant properties. This is reflected in Fig. 4, which demonstrates that crops are more likely to be closer to others belonging to the same crop type (black boxes are mostly filled by green and yellow pixels), e.g., pearl millet is very similar to fonio but very different to cocoyam. This consolidates the reliability of the model parameterization and calibration process in not only capturing the crop characterization such as phenology, biomass, and yield, but also in retaining the physical cohesion across crops.
The distance (unitless) is defined as the Euclidean distance based on the normalized 9 species parameters. A small distance between two crops means the two crops are similar to each other in terms of species parameters, and vice versa. The black boxes show the crops in the same crop type: cereals, legumes, oilseeds, roots/tubers, and vegetables. Green and yellow are the dominant pixel color within the black boxes, suggesting crops in the same crop type have the most identical species parameters.
Model performance and response to extreme events
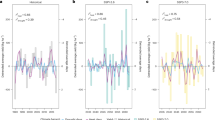
As a test of model response and coherence to observed yield sensitivity, we compare SIMPLE model outputs with observed data and disaster reports (Fig. 5). The International Disaster Database EM-DAT46 records severe drought and heat wave events that affect global agriculture (https://www.emdat.be/). When comparing the detrended simulated and FAO lowest yielding years (lowest 5 years within the 19-year period) with the reported EM-DAT extreme events during the 2000–2018 period, the simulations outperform FAO in 13 of the 16 crops for drought events and 4 of the 14 crops for heat wave events (when teal bars are above red bars in Fig. 5). The gray dashed line shows a hit rate demonstrating skill of a random probability of extremely low yield. For drought, 15 of the 16 crops are above the hit rate demonstrating skill for both simulations and FAO data (lowest 5 years within the 19-year period). For heat waves, 9 of the 14 crop simulations are above the hit rate demonstrating skill, while 11 of the 14 FAO reported crops are above. The difference between the number of crops in the drought and heat wave events is because two crops (Bambara groundnut and Pigeon pea) were not planted in the regions where heat waves were recorded. Heat waves are more widespread and long-lasting, while some droughts may be more localized and/or may have occurred at times of the year outside of the growing season (which is why 100% correspondence is not expected). This shows the model creditably captures the extreme drought and heat wave events in Africa and is slightly more responsive to the extreme events than the FAO data.
FAO (red bars) and simulated (teal bars) lowest yielding years (lowest 5 years within 2000–2018) compared to the EM-DAT database for extreme drought (top) and heat wave (bottom) events. Of the years when EM-DAT indicates an extreme event, simulated yield indicated that this was one of the worst 5 years for a specific crop in countries affected by the extreme event for a specified percentage of the time (30% means 30% of the extreme events fall in the worst 5 years for this crop). This indicates that simulations are as good as, and in some cases better, at capturing extreme events, although both FAO and EM-DAT are limited in capturing what may be local events at a country-scale reporting level. EM-DAT documented 133 drought events and 5 heat wave events during 2000–2018 in Africa. The difference between the number of crops shown in the drought events plots and the heat wave plots is because two crops were not planted in the regions when affected by heat waves. Gray dashed line shows 5 years out of 19-year threshold (26.3%), a hit rate indicating a random probability of extremely low (bottom five years) yield. Bambara groundnut and pigeon pea are not planted in the regions where heat waves are recorded by EM-DAT.
The EM-DAT database, which provides data at a national scale, is insufficient for capturing the complete influence of crop patterns beyond just the impacts of climate. Here, we analyze the correlation between simulated yield at both grid and country levels, FAO yield at country-level, and the Number of Heat Days (NHD) and Drought Days (NDD). Considering the independence of the model, we calculated three commonly used drought indices including the Standardized Precipitation Index (SPI)47, Standardized Precipitation Evapotranspiration Index (SPEI)48, and Self-Calibrated Palmer Drought Severity Index (sc-PDSI)49 for the crop growing season. Using maize and cowpea as examples, all indices at country-level show higher correlation with model simulation than FAO (except for NDD for cowpea) (Table 1). This indicates that the model is responsive to the heat and drought events.
In this study, we constructed crop model applications for 24 crops, of which 19 are opportunity crops, and demonstrated the feasibility of utilizing SIMPLE for opportunity crop model development. As the first endeavor in exploring groups of opportunity crops in physical-based models, this work involves a thorough literature review and state-of-the-art model configuration. This work enables VACS projections of future productivity and climate vulnerability (ref. 50), lays the foundation for modeling opportunity crop research and adaptation potential (ref. 51), investigating the role of opportunity crops towards a food and socioeconomic secure future under climate change at a national level (ref. 52), exploring climate impacts on nutrients and public health (ref. 53), and more future studies relevant to food and agriculture.
A significant challenge to enhancing agricultural models like SIMPLE in Africa is the scarcity of comprehensive and reliable crop observational data28,29. This shortage stems from various factors, including the high cost of data collection, infrastructural limitations, and inadequate investment in research, particularly for these crops. In many regions, farm-level data on crop yields, management practices, and soil conditions are sparse or nonexistent. While a high correlation between simulations and FAO crop anomalies would be ideal, it is important to recognize that country-level production anomalies in many crops are influenced by a complex interplay of factors beyond climate. These include changes in the cultivated area, socioeconomic and cultural transformations, geopolitical conflicts, migration patterns, and the availability of labor and agricultural equipment. Such multifaceted influences make it challenging to precisely match FAO crop yield data for certain crops. Despite this limitation, the present study serves as an effort to enhance our understanding of the potential and performance of various crops in Africa and provides a foundational framework for future investigations. Furthermore, logistical barriers often prevent accurate assessment of different agroecological zones, leaving many local variations unaccounted for. This gap hinders model calibration and validation, limiting the accuracy and robustness of projections. Consequently, the lack of comprehensive observational data prevents a nuanced understanding of specific regional crop performance, reducing the confidence in relying on the model outputs to inform policy recommendations. Addressing this data deficit requires concerted efforts to build more reliable and accessible databases through collaborations between governments, research institutions, and local communities.
While the SIMPLE model provides a valuable framework for analyzing agricultural trends, its inherent simplicity can lead to significant uncertainties. It does not fully account for some key processes that affect crop productivity, such as nutrient limitations and pest and disease pressures. This results in a somewhat coarse representation of crop responses to varying environmental and management conditions. Such limitations could lead to inaccuracies, especially in areas where nutrient deficiencies or specific farming practices significantly affect yields. To enhance model predictive power, incorporating more detailed data on soil health, nutrient management, and pest dynamics would be essential. Despite these uncertainties, the SIMPLE model remains a useful tool for broader initial regional analysis, provided its limitations are acknowledged, and findings are supplemented with field data and expert knowledge.
Looking forward, there is substantial scope to expand this foundational work to include simulations of additional crop types using the SIMPLE model. By extending coverage across a wider variety of crops, we can generate a more comprehensive picture of agricultural responses across different ecosystems and management practices. This expansion would not only refine our understanding of individual crop responses but also enhance our ability to make generalized predictions across agricultural systems.
Moreover, there is a keen interest in developing more sophisticated models for opportunity crops. One option is to follow in the footsteps of established agricultural modeling systems like DSSAT, which has been instrumental in simulating a multitude of crop and environmental interactions and includes some opportunity crops such as cowpea and teff. The ambition is to create models that further advance our understanding of intricate dynamics of soil fertility, management practices, and crop genetics, thereby offering more precision for modeling crop performance under various climatic and soil conditions once reliable soil nutrient data is available. Continued model improvement allows for a more nuanced exploration of adapted management strategies and seed varieties, tailored to the specific needs and limitations of different agricultural zones. Future work in this field will involve extensive collaboration across disciplines, bringing together agronomists, geneticists, climatologists, and computer scientists. By fostering an integrated approach, the research community can better address the multifaceted challenges of modern agriculture and contribute to the evidence needed to create resilient food systems that can withstand the uncertainties of future climate scenarios.
Methods
Introduction to the SIMPLE model
The SIMPLE model32 is based on one of the most widely used crop models - DSSAT, retaining the key hydrobiological components to crop growth while simplifying the computational process. Compared to many other crop models, SIMPLE has the advantage of easy interpretation and implementation, making it a suitable tool for exploring and evaluating opportunity crops. SIMPLE uses cumulative temperature accumulation (degree days) to determine crop phenological development from sowing to physiological maturity. Crop yield is calculated based on the daily biomass accumulation and harvest index (HI), where daily biomass accumulation is determined by photosynthesis rate, which is a function of canopy intercepted solar radiation \(({fSolar})\), radiation use efficiency (RUE), CO2 effect \((f\left({{{\mathrm{CO}}}}_{2}\right))\), temperature effect \((f\left({Temp}\right))\), and heat and drought stresses \((\min (f\left({{\mathrm{Heat}}}\right),f\left({{\mathrm{Water}}}\right)))\) (Eqs. 1 and 2). Detailed model computational equations for each component can be found in ref. 32. As a simplistic crop model, SIMPLE does not come with the same process-depth as other models. Such processes include the response to vernalization, photoperiod effect on phenology, and nutrient dynamics.
SIMPLE model configuration
The SIMPLE model can be setup at a single site or in a gridded way across larger regions, using the R programming language (version 4.3.0). We developed an automatic gridded input generation tool, and setup the model on the NASA Discover high performance supercomputer using a country-rolling computational approach. This allows the model to be run in any prescribed country/countries or the whole of Africa automatically with a high computational efficiency. The SIMPLE model is set up at 0.5° × 0.5° spatial resolution for the whole African continent from 2000 to 2019, considered as the baseline period, while the year 2019 is excluded from the analysis due to some crops in some regions not reaching maturity until outside of the time period examined. Similar to DSSAT, SIMPLE requires climate, soil, management, and genetic data as model inputs. The following subsections introduce the data sources for each category.
Climate data
The baseline daily climate data, including solar radiation (MJ m−2), 2-m maximum and minimum temperature (°C), and precipitation (mm) from 2000 to 2019, are obtained from European Centre for Medium-Range Weather Forecasts Reanalysis v5 (ECMWF-ERA5)54 W5E5 v1.0 reanalysis dataset. ERA5 provides high-resolution data across various meteorological variables, such as temperature, radiation, precipitation, and more from 1950 to the present, updated in near-real-time. With its fine spatial resolution of approximately 31 km and hourly temporal granularity, ERA5 facilitates detailed climate analyses and improves the accuracy of weather forecasting and climate modeling and is crucial for understanding past, present, and future climate dynamics globally. The ERA5 data used in this study are downscaled and centrally bias-adjusted following the Inter-Sectoral Impact Model Intercomparison Project (ISIMIP) framework55 and previous AgMIP global gridded impact assessments9.
Soil data
Soil data are from the gridded Global Soil Dataset for Earth System Models (GSDE)56, which has been shown to be comparable to other well-known soil datasets such as SoilGrids57. Soil data include soil available water capacity (defined as drained upper limit - lower limit), runoff curve number, and deep drainage coefficient. Root zone depth (mm) is also added from the Africa Soil Information Service (AfSIS)58. These parameters are all provided at a spatial resolution of 0.5° × 0.5° for integration into the crop model. SIMPLE incorporates these inputs within its water budget routine to determine the daily water balance and assess cumulative drought stress using the ARID index and ARID sensitivity parameter for each crop type58. In this modeling framework, soil moisture levels are reset at the start of each growing season.
Management data
The gridded crop calendar data, including planting and maturity dates for each crop, are from AgMIP’s Global Gridded Crop Model Intercomparison (GGCMI) repository9. The GGCMI crop calendar, which integrates various observational data sources, offers information on planting and maturity dates for five specific crops in this study—cassava, groundnut, maize, sorghum, and soybean—at a resolution of 0.5°. For crops that are not covered by the GGCMI crop calendar, we employed the calendar of a similar crop as a substitute where possible (for example, using the millet calendar in place of the finger millet calendar, or the peas calendar for the grass pea calendar). In cases where no analogous crop calendar was available, we developed a crop calendar tailored to defined agroecological zones. This involved populating each agroecological region34 with the predominant sowing dates derived from the FAO crop calendar, the International Fertilizer Association’s Fertilizer Use by Country crop calendar59, and relevant literature references60,61,62.
The gridded planting area data is obtained from CROPGRIDS63, except for Ghana where SPAM201064 was used since we had it validated against several sites. CROPGRIDS is a global geo-referenced harvested area dataset for 173 crops at a resolution of 0.05° through data merging from multiple independent sources. For crops not included in CROPGRIDS, we use an analog crop from CROPGRIDS as proxy (Table S2). African island countries Cabo Verde, Mauritius, and Seychelles are not included in this study due to their limited agricultural data. Grid cells with harvest area <5 ha are excluded from the modeling to balance accuracy and computational runtime efficiency. Given that Africa has a very low overall cropland irrigation rate (about 6%)65, we exclude irrigation in this study and all crops are simulated under rainfed conditions.
Metrics for skill assessment
Root Mean Square Error (RMSE), Relative Root Mean Squared Error (RRMSE), Coefficient of Determination (R²), and Index of Agreement (d-index) are chosen as skill metrics in modeling due to their effectiveness in evaluating different aspects of model performance (Eqs. 3–6). Each metric offers unique insights and justifications for its use.
RMSE is a standard statistical measure used to evaluate the accuracy of a model by quantifying the difference between predicted values and the actual observed values. RMSE is particularly useful because it is expressed in the same units as the simulated variable, making interpretation intuitive and direct. RRMSE is a normalized statistical measure used to assess the accuracy of predictive models, similar to RMSE but adjusted relative to the scale of the data. RRMSE is particularly advantageous in contexts where model outputs vary significantly in magnitude, allowing comparisons that are fair and contextually relevant. R² measures the proportion of variance in the dependent variable that is predictable from the independent variables in a regression model. R² is essential for evaluating the overall effectiveness of a model in explaining the variability in the data, offering a statistical summary of how well predictions conform to actual outcomes. The d-index provides a comprehensive assessment of model performance by accounting for both systematic and random disparities between observed and simulated data, which identifies additive and proportional differences in the means and variances of the observed and modeled values.
where N is the number of data points, \(y\left(i\right)\) is the i-th observation, \(\hat{y}(i)\) is its corresponding simulation, and \(\bar{y}\) is the observational mean.
Model calibration and validation
Cereals
Using fonio as the representative crop for cereals (Fig. 2), we obtained a field experiment consisting of a total of 7 varying planting date treatments with phenology, biomass, and yield data from Senegal covering 2 years in 2 fields in Bandafassi and Sinthiou Maleme66. We first calibrated the SIMPLE Tbase, Tsum, HI, Topt, RUE, and Tmax parameters using ref. 66, and then manually tuned the other parameters through comparisons with field data to minimize RMSE and RRMSE. We then use the FAO yield data of the median country across all country FAO yield data, Benin, as a reference to validate the model38. Similar calibration and validation procedures are followed for other crops described in the next sections.
Across all treatments, the calibration accuracy and effectiveness were quantitatively assessed using RMSE and RRMSE. For the crop phenology of fonio, the model achieved a RMSE of 2 days and an RRMSE of 3%, indicating a relatively precise simulation of the crop seasonal development compared to observed data. The RMSE and RRMSE are 1023 kg ha−1 and 26% respectively for the simulated biomass indicating satisfactory model performance although some challenges exist due to variability in biomass production. Similarly, yield simulations also presented a RMSE of 246 kg ha−1 and a RRMSE of 26%, underscoring an acceptable model performance in simulating fonio yield. The validation against FAO yield in Benin shows a RMSE of 222 kg ha−1 and RRMSE of 36%, suggesting a decent model performance at the country-level.
Legumes
Using cowpea as the representative crop for legumes (Fig. 2), we obtained a field experiment with a total of 3 varying planting date treatments with phenology, biomass, and yield data from Kpong, Ghana covering 1 year67. We start with the parameters Topt from ref. 68, Tmax from ref. 69, and SCO2 from ref. 70, then manually tuned other parameters through comparisons with field data to minimize RMSE and RRMSE. We then use the FAO yield data also from Ghana to validate the model. The FAO yield was adjusted for a moisture content of 12% to convert to dry weight.
For the growth duration of cowpea, the model achieved an RMSE of 2 days and an RRMSE of 3%, indicating a relatively precise simulation of the crop developmental timeline compared to observed data. In terms of biomass, the RMSE was 371 kg ha−1 and the RRMSE was 12%, reflecting an acceptable simulation of biomass production. Similarly, yield simulations presented a RMSE of 212 kg ha−1 and a RRMSE of 12%, underscoring an accurate model performance in simulating cowpea yield. The validation against the FAO yield in Ghana shows a RMSE of 137 kg ha−1 and RRMSE of 12%, suggesting a satisfactory model performance.
Roots/tubers
Using yams as the representative roots/tubers crop (Fig. 2), we obtained a field experiment in Nigeria covering 2 years with 5 planting date treatments in each year71. The crop duration was used to estimate Tsum whilst other phenology and growth parameters were obtained from literature and manually tuned through comparisons with field data to minimize RMSE and RRMSE. We then use the FAO yield data also in Nigeria to validate the model.
For the crop phenology of yams, the model achieved a RMSE of 28 days and an RRMSE of 12%, indicating adequate simulation of the longer growing seasons for yams. In terms of biomass, the RMSE was higher at 6053 kg ha−1, but had an acceptable RRMSE of 24%, reflecting greater variability in accurately modeling biomass production. Similarly, yield simulations presented a RMSE of 4327 kg ha−1 and an RRMSE of 22%, underscoring an acceptable model performance in simulating the higher observed yields of yams. The validation against the FAO yield in Nigeria shows a RMSE of 4503 kg ha−1 and RRMSE of 46%, suggesting fair model performance.
Vegetables
Using tomato as the representative crop for vegetables (Fig. 2), we obtained a greenhouse experiment with a total of 3 high fertigation treatments with phenology, biomass, and yield data from Tamale, Ghana covering 2 years with 2 cultivars, cv. Jalila and Yetty72. The model was calibrated for both cultivars and the better performing cultivar, Jalila, was used for the gridded simulations. We started with the default SIMPLE tomato parameters and adjusted Tsum from the FAO database73, then checked the other parameters through comparisons with field data to minimize RMSE and RRMSE, where the default parameters proved to provide the best fit. We then used the FAO yield data of the median country across all country FAO yield data, Liberia, as a reference to validate the model. The FAO yield was adjusted for a moisture content of 85% to convert to dry weight73.
For the phenology of tomato, the model achieved an RMSE of 10 days and a RRMSE of 6%, indicating a relatively precise simulation of the crop seasonal development compared to observed data. In terms of biomass, the RMSE was 1941 kg ha−1, with a RRMSE of 29%, reflecting decent model performance. Similarly, yield simulations presented a RMSE of 701 kg ha−1 and a RRMSE of 18%, underscoring an acceptable model performance in simulating tomato yield. The validation against the FAO yield in Liberia shows a RMSE of 1003 kg ha−1 and RRMSE of 60%, suggesting fair model performance. The higher RRMSE may be explained by below average yields between 2000–2011 (see Fig. 2 timeseries) or high variability in the FAO reported yield.
Following similar procedures, we configured the cereal crops including maize, teff, finger millet, sorghum, and pearl millet; legume crops including soybean, grass pea, mung bean, pigeon pea, lablab, and bambara groundnut; root/tuber crops including cassava, groundnut, sesame, cocoyam, taro, and sweet potato; and vegetable crops including African eggplant, okra, and amaranth. The detailed cultivar and species parameters are shown in Table S5.
Model evaluation at the continent-level
To justify that the model is applicable in the whole Africa continent rather than only in several sites and countries, we further evaluate the calibrated models for Africa, where Egypt and Djibouti are excluded due to the high share of cultivated area equipped for irrigation (nearly 100%)74.
Spatiotemporal evaluation
Here we demonstrate the comparison of the simulated yield of the four representative crops with the FAO average yield38 across the whole of Africa (Fig. 6). The comparison is made based on the country-year yield from both datasets, namely, each data point represents one country in one year. The mean yield, along with the density distribution show a good consistency between modeling results and FAO for fonio, yams, and tomato. The three crops have the mean yield and the most concentrated yield range at around 900 kg ha−1, 9000 kg ha−1, and 8000 kg ha−1 respectively. Cowpea, although showing a relatively different mean yield (about 500 kg ha−1 for FAO and 1200 kg ha−1 for simulation) and the most concentrated yield range, still has a large overlap between the two datasets. This provides the confidence that the calibrated models can represent the observed yield across the whole of Africa with acceptable fidelity in both spatial and temporal dimension. Meanwhile, the FAO data has much more outliers than the simulated results for every crop, showing large spatiotemporal variability and uncertainty. Such variability and uncertainty could originate from the abrupt changes in technology and seed variety in a certain year for some countries that significantly increased yield, or natural and socioeconomic shocks such as flooding and conflicts that significantly reduced yield, which are not capable of being captured by the model. Nevertheless, the models can still provide reliable results that reflect average yield conditions in Africa.
Each data point is a crop yield at a country-year for the baseline period (2000 to 2018) for FAO38 (red) and the SIMPLE model (teal). Box plots show the 90th percentile, 75th percentile, median, 25th percentile, and 10th percentile (from top to bottom) and outliers of observations and simulations. The legend refers to all panels. Note that the FAO reported tomato fresh weight is adjusted to 15% to represent the dry weight73. The comparison for all crops with FAO data available are shown in the Supplementary Fig. S9.
Spatial distribution evaluation
The spatial distribution comparisons of the four representative crops with FAO are shown in Fig. 7. The spatial pattern of the simulated yield for fonio is highly consistent with the FAO data. Fonio is found to be only planted in Western Africa. Countries including Guinea, Mali and Côte d’Ivoire have the highest yield of about 1000 kg ha−1. The simulated cowpea yield is also consistent with FAO in terms of spatial distribution. Cowpea is most productive in East African countries such as Tanzania, Uganda, and Madagascar and West African countries including Ghana and Nigeria, with the highest yield of around 2000 kg ha−1. Yams also show a good consistency in the spatial pattern with the FAO data, with West Africa being the most productive region. Countries such as Ghana, Togo, Benin, and Nigeria have the highest yields of about 15,000 kg ha−1. Central African Republic in Central Africa and Ethiopia in East Africa are also found to be very productive for yams. There are discrepancies between the spatial distribution of simulated tomato yield and the FAO data. We find that tomato has the highest simulated yield in Central and West Africa of 10,000 kg ha−1, while the FAO data signifies that Northern and Southern Africa have the highest yields. This may indicate a potential of planting tomato in Central and West Africa where there is less current cultivated area. Nevertheless, the calibrated model also shows high tomato yields in areas within South Africa, Morrocco, Algeria, and Tunisia along the coastal area.
It is worth noting that FAO does not provide subnational yield data. Hence, products from this study provide a higher resolution perspective on crop yields. Differences between simulations and FAO yield and production may also reflect missed opportunities for crop expansion or could indicate socioeconomic factors that diminish that crop use (e.g., a higher value crop outperforms a lower value crop). This suggests that future land use management should not be determined by the best place to grow a given crop but determined by the best crop that can be grown in a given place.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The daily climate data are from European Centre for Medium-Range Weather Forecasts Reanalysis v5 (ECMWF-ERA5) W5E5 v1.0 reanalysis dataset (https://www.ecmwf.int/en/forecasts/dataset/ecmwf-reanalysis-v5).Soil data are from the gridded Global Soil Dataset for Earth System Models (GSDE) (https://cmr.earthdata.nasa.gov/search/concepts/C1214604044-SCIOPS.html#). The gridded crop calendar data are from AgMIP’s Global Gridded Crop Model Intercomparison (GGCMI) repository (https://zenodo.org/records/5062513), FAO crop calendar (https://cropcalendar.apps.fao.org/#/home), and the International Fertilizer Association’s Fertilizer Use by Country crop calendar (https://www.ifastat.org/). The gridded planting area data is obtained from CROPGRIDS and SPAM2010 (https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/PRFF8V). Model results data is publicly available at https://doi.org/10.7910/DVN/MLRBYD.
Code availability
The code for SIMPLE crop model, data analysis and figure plotting is publicly available at https://doi.org/10.7910/DVN/MLRBYD.
References
FAO, AUC, ECA, & WFP. Africa - Regional Overview of Food Security and Nutrition 2023. (Statistics and trends. Accra, FAO, 2023).
Sibanda, F. Rastafari, climate change and crop failure in Africa. in Religion, Climate Change, and Food Security in Africa (eds Maseno, L., Omona, D. A., Chitando, E. & Chirongoma, S.) 99–111. https://doi.org/10.1007/978-3-031-50392-4_6 (Springer Nature Switzerland, 2024).
Oyelami, L. O., Edewor, S. E., Folorunso, J. O. & Abasilim, U. D. Climate change, institutional quality and food security: Sub-Saharan African experiences. Sci. Afr. 20, e01727 (2023).
Rezaei, E. E. et al. Climate change impacts on crop yields. Nat. Rev. Earth Environ. 4, 831–846 (2023).
Adesete, A. A., Olanubi, O. E. & Dauda, R. O. Climate change and food security in selected Sub-Saharan African Countries. Environ. Dev. Sustain. 25, 14623–14641 (2023).
Dasgupta, S. & Robinson, E. J. Z. Attributing changes in food insecurity to a changing climate. Sci. Rep. 12, 4709 (2022).
Lesk, C. et al. Compound heat and moisture extreme impacts on global crop yields under climate change. Nat. Rev. Earth Environ. 3, 872–889 (2022).
Simanjuntak, C. et al. Impact of climate extreme events and their causality on maize yield in South Africa. Sci. Rep. 13, 12462 (2023).
Jägermeyr, J. et al. Climate impacts on global agriculture emerge earlier in new generation of climate and crop models. Nat. Food 2, 873–885 (2021).
United Nations. Population. United Nations https://www.un.org/en/global-issues/population.
McMullin, S. et al. Determining appropriate interventions to mainstream nutritious orphan crops into African food systems. Glob. Food Secur. 28, 100465 (2021).
Hossain, A. et al. Neglected and underutilized crop species: are they future smart crops in fighting poverty, hunger and malnutrition under changing climate? in Neglected and Underutilized Crops - Towards Nutritional Security and Sustainability (eds Zargar, S. M., Masi, A. & Salgotra, R. K.) 1–50. https://doi.org/10.1007/978-981-16-3876-3_1 (Springer, Singapore, 2021).
Akinola, R., Pereira, L. M., Mabhaudhi, T., de Bruin, F.-M. & Rusch, L. A review of indigenous food crops in Africa and the implications for more sustainable and healthy food systems. Sustainability 12, 3493 (2020).
Leakey, R. R. B. et al. The future of food: domestication and commercialization of indigenous food crops in Africa over the Third Decade (2012–2021). Sustainability 14, 2355 (2022).
Mudau, F. N., Chimonyo, V. G. P., Modi, A. T. & Mabhaudhi, T. Neglected and underutilised crops: a systematic review of their potential as food and herbal medicinal crops in South Africa. Front. Pharmacol. 12, 809866 (2022).
Chaturvedi, P., Govindaraj, M., Govindan, V. & Weckwerth, W. Editorial: Sorghum and pearl millet as climate resilient crops for food and nutrition security. Front. Plant Sci. 13, 851970 (2022).
Khalifa, M. & Eltahir, E. A. B. Assessment of global sorghum production, tolerance, and climate risk. Front. Sustain. Food Syst. 7, 1184373 (2023).
Wilson, M. L. & VanBuren, R. Leveraging millets for developing climate resilient agriculture. Curr. Opin. Biotechnol. 75, 102683 (2022).
Umesh, M. R., Angadi, S., Gowda, P., Ghimire, R. & Begna, S. Climate-resilient minor crops for food security. in Agronomic Crops: Volume 1: Production Technologies (ed. Hasanuzzaman, M.) 19–32. https://doi.org/10.1007/978-981-32-9151-5_2 (Springer, 2019).
Imathiu, S. Neglected and underutilized cultivated crops with respect to indigenous African leafy vegetables for food and nutrition security. J. Food Secur. 9, 115–125 (2021).
Mondo, J. M. et al. Neglected and underutilized crop species in Kabare and Walungu territories, Eastern D.R. Congo: identification, uses and socio-economic importance. J. Agric. Food Res. 6, 100234 (2021).
Muthoni, J. & Nyamongo, D. O. Traditional food crops and their role in food and nutritional security in Kenya. J. Agric. Food Inf. 11, 36–50 (2010).
Chivenge, P., Mabhaudhi, T., Modi, A. T. & Mafongoya, P. The potential role of neglected and underutilised crop species as future crops under water scarce conditions in Sub-Saharan Africa. Int. J. Environ. Res. Public. Health 12, 5685–5711 (2015).
Farooq, M., Rehman, A., Li, X. & Siddique, K. H. M. Neglected and underutilized crops and global food security. in Neglected and Underutilized Crops (eds Farooq, M. & Siddique, K. H. M.) 3–19. https://doi.org/10.1016/B978-0-323-90537-4.00001-6 (Academic Press, 2023).
Alvar-Beltrán, J., Dibari, C., Ferrise, R., Bartoloni, N. & Marta, A. D. Modelling climate change impacts on crop production in food insecure regions: the case of Niger. Eur. J. Agron. 142, 126667 (2023).
Rosenzweig, C. et al. The Agricultural Model Intercomparison and Improvement Project (AgMIP): protocols and pilot studies. Agric. Meteorol. 170, 166–182 (2013).
Ruane, A. C. et al. An AgMIP framework for improved agricultural representation in integrated assessment models. Environ. Res. Lett. 12, 125003 (2017).
Adiku, S. G. K., MacCarthy, D. S. & Kumahor, S. K. A conceptual modelling framework for simulating the impact of soil degradation on maize yield in data-sparse regions of the tropics. Ecol. Model. 448, 109525 (2021).
Diancoumba, M. et al. Scientific agenda for climate risk and impact assessment of West African cropping systems. Glob. Food Secur. 38, 100710 (2023).
Hoogenboom, G. et al. The DSSAT crop modeling ecosystem. in Advances in crop modelling for a sustainable agriculture (Burleigh Dodds Science Publishing, 2019).
van Zonneveld, M. et al. Forgotten food crops in sub-Saharan Africa for healthy diets in a changing climate. Proc. Natl. Acad. Sci. USA 120, e2205794120 (2023).
Zhao, C. et al. A SIMPLE crop model. Eur. J. Agron. 104, 97–106 (2019).
Karl, K. A. et al. Opportunity Crop Profiles for the Vision for Adapted Crops and Soils (VACS) in Africa. https://doi.org/10.7916/7msa-yy32 (2024).
HarvestChoice & Institute (IFPRI), I. F. P. R. Agro-Ecological Zones for Africa South of the Sahara. Harvard Dataverse https://doi.org/10.7910/DVN/M7XIUB (2016).
Kharbamon, B. et al. Response of planting time and phosphorus dosage on yield and nutrient uptake in Dolichos Bean (Lablab Purpureus L.). Indian J. Hill Farming 17, 28–34 (2017).
Choudhary, R., Kushwah, S. S., Sharma, R. K. & Kachouli, B. K. Effect of sowing dates on growth, flowering and yield of indian bean varieties under agroclimatic conditions of Malwa Plateau in Madhya Pradesh. Legume Res. https://arccjournals.com/journal/legume-research-an-international-journal/LR-4212 (2020).
Beck, H. E. et al. Present and future Köppen-Geiger climate classification maps at 1-km resolution. Sci. Data 5, 180214 (2018).
FAOSTAT. https://www.fao.org/faostat/en/#home.
EL-Sharkawy, M. A. Cassava biology and physiology. Plant Mol. Biol. 53, 621–641 (2003).
Ruggeri, R. et al. Potential of teff as alternative crop for Mediterranean farming systems: Effect of genotype and mowing time on forage yield and quality. J. Agric. Food Res. 17, 101257 (2024).
Animasaun, D. A., Adedibu, P. A., Olawepo, G. K. & Oyedeji, S. Fonio millets: an underutilized crop with potential as a future smart cereal. in Neglected and Underutilized Crops (eds Farooq, M. & Siddique, K. H. M.) 201–219. https://doi.org/10.1016/B978-0-323-90537-4.00028-4 (Academic Press, 2023).
Gupta, S. M. et al. Finger millet: a “Certain” Crop for an “Uncertain” Future and a solution to food insecurity and hidden hunger under stressful environments. Front. Plant Sci. 8, 643 (2017).
ATLAS. Impact of Climate Change on Select Value Chains in Mozambique. https://weadapt.org/wp-content/uploads/2023/05/170110_ftf_climate_risks_to_value_chains_in_mozambique.pdf (2017).
Conti, V., Parrotta, L., Romi, M., Del Duca, S. & Cai, G. Tomato biodiversity and drought tolerance: a multilevel review. Int. J. Mol. Sci. 24, 10044 (2023).
Degu, H. D., Ohta, M. & Fujimura, T. Drought tolerance of eragrostis tef and development of roots. Int. J. Plant Sci. 169, 768–775 (2008).
Delforge, D. et al. EM-DAT: the Emergency Events Database. Int. J. Disaster Risk Reduct. 124, https://doi.org/10.1016/j.ijdrr.2025.105509 (2025).
McKee, T. B., Doesken, N. J. & Kleist, J. The relationship of drought frequency and duration to time scales. Proc. 8th Conf. Appl. Climatol. 179, 184 (1993).
Vicente-Serrano, S. M., Beguería, S. & López-Moreno, J. I. A multiscalar drought index sensitive to global warming: the standardized precipitation evapotranspiration index. https://doi.org/10.1175/2009JCLI2909.1 (2010).
Wells, N., Goddard, S. & Hayes, M. J. A. Self-Calibrating. Palmer Drought Severity Index https://journals.ametsoc.org/view/journals/clim/17/12/1520-0442_2004_017_2335_aspdsi_2.0.co_2.xml (2004).
Guarin, J. R. et al. Modeling the productivity of opportunity crops across Africa under climate change in support of the vision for adapted crops and soils. Nat. Plants. https://doi.org/10.1038/s41477-025-02157-9 (2025).
Fredenberg, E. et al. Vision for Adapted Crops and Soils (VACS) Research in Action: Opportunity Crops for Africa. 37 (Columbia University, Columbia Academic Commons, 2024).
Omotayo, A. O., Omotoso, A. B. & Asong, J. A. Leveraging Africa’s underutilized crops to combat climate change, water scarcity, and food insecurity in South Africa. Scientific Reports 15, 19404 (2025).
Carducci, B. et al. Anticipating climate impacts on nutrition: the role of climate-crop nutrient modeling. Nat. Clim. Chang. 15, 1165–1172 (2025).
Hersbach, H. et al. The ERA5 global reanalysis. Q. J. R. Meteorol. Soc. 146, 1999–2049 (2020).
Lange, S. Trend-preserving bias adjustment and statistical downscaling with ISIMIP3BASD (v1.0). Geosci. Model Dev 12, 3055–3070 (2019).
Shangguan, W., Dai, Y., Duan, Q., Liu, B. & Yuan, H. A global soil data set for earth system modeling. J. Adv. Model. Earth Syst. 6, 249–263 (2014).
Dai, Y. et al. A review of the global soil property maps for Earth system models. SOIL 5, 137–158 (2019).
Leenaars, J. G. B. et al. Mapping rootable depth and root zone plant-available water holding capacity of the soil of sub-Saharan Africa. Geoderma 324, 18–36 (2018).
Ludemann, C. I., Gruere, A., Heffer, P. & Dobermann, A. Global data on fertilizer use by crop and by country. Sci. Data 9, 501 (2022).
Malherbe, S. & Marais, D. Economics, yield and ecology: a case study from the South African tomato industry. Outlook Agric 44, 37–47 (2015).
Robinson, E. & Kolavalli, S. The case of tomato in Ghana: productivity. GSSP Working Paper No 19, 1–19 (2010).
Kamga, R. T., Some, S., Tenkouano, A., Issaka, Y. B. & Ndoye, O. Assessment of traditional African vegetable production in Burkina Faso. J. Agric. Ext. Rural Dev. 8, 141–150 (2016).
Tang, F. H. M. et al. CROPGRIDS: a global geo-referenced dataset of 173 crops. Sci. Data 11, 413 (2024).
Institute (IFPRI), I. F. P. R. Global Spatially-Disaggregated Crop Production Statistics Data for 2010 Version 2.0. Harvard Dataverse https://doi.org/10.7910/DVN/PRFF8V (2025).
You, L. et al. What is the irrigation potential for Africa? A combined biophysical and socioeconomic approach. Food Policy 36, 770–782 (2011).
Gueye, M. et al. Effect of planting date on growth and grain yield of fonio millet (Digitaria exilis Stapf) in the Southeast of Senegal. Int. J. Biol. Chem. Sci. 9, 581 (2015).
Tei-Labi, V.K. Evaluation of cowpea genotypes for yield and genotype by environment stability on the vertisols of the Accra plains. Master's thesis, University of Ghana, Legon (2015).
Department of Agriculture, Forestry and Fisheries, Republic of South Africa. Production Guidelines for Cowpea. https://www.nda.gov.za/phocadownloadpap/Brochures_and_Production_Guidelines/COWPEA%20PRODUCTION%20GUIDELINES%202014.pdf (2014).
Barros, J. R. A. et al. Selection of cowpea cultivars for high temperature tolerance: physiological, biochemical and yield aspects. Physiol. Mol. Biol. Plants 27, 29–38 (2021).
Adusei, G., Aidoo, M. K., Srivastava, A. K., Asibuo, J. Y. & Gaiser, T. The impact of climate change on the productivity of cowpea (Vigna unguiculata) under three different socio-economic pathways. Ital. J. Agron. 17, 2118 (2022).
Tobih, F. Planting Dates Effects on Growth and Yield of Yams (Dioscorea spp.) in Asaba Area, Delta State, Nigeria. J. Biol. Agric. Healthc. https://www.semanticscholar.org/paper/Planting-Dates-Effects-on-Growth-and-Yield-of-Yams-Tobih/f6520e1613bdbb5a4d7ec0137e9db3b8ffe6820f (2016).
Agbemabiese, Y. K., Abubakari, A.-H., Dzomeku, I. K. & Abdul-Ganiyu, S. Water and nutrient use efficiency of soilless grown greenhouse tomato (Lycopersicon esculentum L.). Cogenit Food Agric 9, 2275415 (2023).
FAO Database. Food and Agriculture Organization of the United Nations Land & Water Crop Information - Tomato. https://www.fao.org/land-water/databases-and-software/crop-information/tomato/en/ (2024).
You, L. et al. What is the irrigation potential for Africa? A combined biophysical and socioeconomic approach. IFPRI Discussion Paper 00993. Washington, D.C.: International Food Policy Research Institute. (2010).
Woli, P., Jones, J. W., Ingram, K. T. & Fraisse, C. W. Agricultural Reference Index for Drought (ARID). Agron. J. 104, 287–300 (2012).
Acknowledgements
This work is funded by the Rockefeller Foundation (grant number 2023 FCW 004) and is conducted in support of the U.S. Department of State Vision for Adapted Crops and Soils (VACS). M.Y., J.R.G., K.K., J.J., A.C.R., and C.E.R. were enabled by the NASA Earth Sciences Division support of the NASA GISS Climate Impacts Group.
Author information
Authors and Affiliations
Contributions
M.Y. and J.R.G. conducted the crop model simulations and data analysis and developed the manuscript and figures. D.S.M., B.S.F., G.O.W., and S.N. assisted in crop model calibration and regional crop information. D.S.M., K.K., and C.E.R. conceived the idea and coordinated integration with VACS. A.C.R. and C.E.R. coordinated integration with AgMIP. J.J. provided global modeling guidance and soil input datasets. A.C.R. and A.C. provided climate input datasets. K.K. and E.M.L. provided crop profile background information and data. S.A and C.Z. provided modeling guidance. All coauthors supported the writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Prakash Jha and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, M., Guarin, J.R., Freduah, B.S. et al. Climate-crop models to support opportunity crop adaptation in Africa. Nat Commun 16, 11186 (2025). https://doi.org/10.1038/s41467-025-66180-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66180-2