Abstract
Computational pathology has advanced through foundation models, yet faces challenges in multimodal integration and capturing whole-slide context. Current approaches typically utilize either vision-only or image-caption data, overlooking distinct insights from pathology reports and gene expression profiles. Additionally, most models focus on patch-level analysis, failing to capture comprehensive whole-slide patterns. Here we present mSTAR (Multimodal Self-TAught PRetraining), the pathology foundation model that incorporates three modalities: pathology slides, expert-created reports, and gene expression data, within a unified framework. Our dataset includes 26,169 slide-level modality pairs across 32 cancer types, comprising over 116 million patch images. This approach injects multimodal whole-slide context into patch representations, expanding modeling from single to multiple modalities and from patch-level to slide-level analysis. Across oncological benchmark spanning 97 tasks, mSTAR outperforms previous state-of-the-art models, particularly in molecular prediction and multimodal tasks, revealing that multimodal integration yields greater improvements than simply expanding vision-only datasets.
Similar content being viewed by others
Introduction
The recent advancements in foundation models (FMs)1,2,3,4,5 for computational pathology (CPath) have demonstrated considerable progress in an incredibly broad spectrum of clinical tasks, such as cancer diagnosis, treatment and prognosis. Despite encouraging performance in general-purpose pathology foundation models, there are still several unresolved challenges.
First, massive multimodal data in line with clinical practices is under-utilized for pretraining, such as pathology reports and gene expression profiles. Existing pathology FMs either focus on vision-only2 or image-caption data1,3, in which the information provided by captions is insufficient to provide whole slide context for authentic slide-level oncological tasks although attempting to incorporate different modalities. The power of multimodal data has been repeatedly substantiated not only in the general machine learning community6,7 but also in the field of medical cancer research8,9,10. In the clinical workflow, as shown in Fig. 1a and examples in Supplementary Fig. 1, pathology reports often provide the most clinically relevant information of whole slides in real-world scenarios, while patients’ gene expression profiles offer insights into quantitative molecular dynamics that can complement the qualitative morphological view provided by a slide. The integration of these slide-level multimodal data can establish a broad and holistic perspective, thereby undoubtedly enhancing the capabilities of PFMs for various clinical tasks.
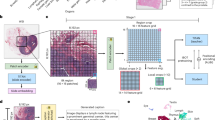
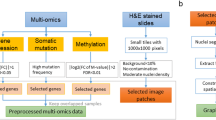
a The workflow in clinical practice for diagnosis, treatment and prognosis of oncology, which primarily involves three common modalities data: WSIs, pathology reports and gene expression profiles. b The overview of mSTAR paradigm. mSTAR consists of two stages: 1) Slide-level Contrastive Learning, and 2) Patch-level Self-Taught Training. c–e statistics of data used in this study, including (c) Venn Graph of cases across various modalities, d the number of cases in pretraining data across different cancer types. e the distribution of word count for pathology reports. f evaluation scheme in this study: including held-out, independent, external and zero-shot. The illustration is presented in Sec. ? g the distribution of datasets across different types of tasks for different evaluation scheme, and the detailed information about every dataset is presented in Supplementary Table 1. h The average performance spanning 15 types of 97 tasks across 7 categories of applications: Pathological Diagnosis, Molecular Prediction, Report Generation, Survival Prediction, Multimodal Fusion, Zero-shot Slide Classification, and Zero-shot Slide Retrieval. Zero-shot tasks, which require a well-aligned vision-language space, are evaluated for vision-language models only, i.e., PLIP, CONCH and mSTAR. Source data are provided as a Source Data file and presented in Supplementary Table 2 as well. This figure was created in BioRender. Zhou, Z. (https://BioRender.com/r035ixv).
Second, existing efforts in pathology FMs are predominantly aimed at the modeling of patch/ROI-level data1,2,3, leading to limited contexts for slide-level oncological applications. Conventional models typically treat individual patch images as independent samples for pretraining a patch extractor, and subsequently employ multiple instance learning (MIL)11,12,13 to perform slide-level modeling based on embedded patch features. Recent concurrent works4,14 have attempted to pretrain the slide-level FM, or incorporate gene data into the lightweight slide aggregator pretraining15. However, pretraining a slide aggregator on top of pre-extracted patch features from a fixed trained patch extractor poses an inherent limitation that the quality of patch features of the slide aggregator inevitably constrains the upper bound of pretraining performance. Since slide-level multimodal self-supervised signals fail to guide patch-level feature extraction, the pretraining objectives’ misalignment of these two independent stages inevitably results in suboptimal performance. Furthermore, the lightweight architecture of pretrained aggregators necessitated by efficiently handling massive patches inherently limits their capacity to absorb multimodal information during pretraining.
In this work, we integrate three complementary modalities for pathology foundation models: pathology slides, specialized pathology reports, and gene expression data. A collection of 26,169 slide-level modality pairs from 10,275 patients across 32 cancer types (Fig. 1c-e) is used to develop a whole-slide pretraining paradigm termed Multimodal Self-TAught PRetraining (mSTAR, Figs. 1b and 2), encompassing over 116 million pathological patch images. This approach leverages slide-level contrastive learning to pretrain a slide aggregator, which is then used to inject whole-slide contextual information into a patch feature extractor. The model is evaluated on a spectrum of 97 clinical tasks across 15 application types (Fig. 1f-g and Supplementary Table 1), including molecular prediction, report generation, and multimodal fusion. Results (Fig. 1h) indicate that incorporating multiple modalities during pretraining enhances performance across tasks related to the respective modalities and can achieve competitive outcomes with less data than vision-only models requiring larger-scale slide datasets.
Results
The overview of mSTAR
The proposed mSTAR aims to provide a novel whole-slide pretraining paradigm that injects multimodal knowledge into the pathology foundation model. Compared with existing pathology foundation models, mSTAR has the following innovative designs to fully unleash its power in a wide spectrum of pathological downstream tasks. First, clinical multimodal data are fully harnessed in pretraining to endow the pathology FM with multimodal knowledge for comprehensive perspectives in clinical tasks. Second, the whole-slide pretraining paradigm provides an alternative way to obtain whole-slide contexts for pathology FMs through self-taught training. To the best of our knowledge, this is the first work to inject multimodal knowledge at the whole-slide context into a pathology FM, broadening the contextual understanding for CPath from patch-level to slide-level and from unimodal to multimodal knowledge. The overview of mSTAR is shown in Fig. 2, consisting of two stages of pretraining.
mSTAR is a whole-slide pretraining paradigm comprising two-stage pretraining. a Stage 1 aims to inject multimodal knowledge into a slide aggregator by slide-level contrastive learning among WSIs, pathology reports and gene expression data. b Stage 2 aims to seamlessly propagate multimodal knowledge learned at the slide level into the patch extractor by Self-Taught training, which leverages the slide aggregator pretrained in Stage 1 as “Teacher” and enforces patch extractor to be “Student''. This figure was created in BioRender. Zhou, Z. (https://BioRender.com/evctgc8).
In the first stage, the objective is to inject multimodal knowledge into the slide aggregator by slide-level contrastive learning among three modalities, i.e., WSIs, pathology reports and RNA-Seq profiles. Note that the pretrained slide aggregator will act as a bridge that propagates multimodal knowledge into the patch extractor in the next stage. To this end, as shown in Fig. 2a, we first utilized a pretrained patch extractor, a state-of-the-art pathology foundation model named UNI2, to encode each patch image of a slide into patch features. Then the resulting patch features are fed into a slide aggregator and integrated into a slide-level representation which is subsequently aligned with other modalities through inter-modality contrastive learning. Furthermore, to mitigate the influence of heterogeneity across different types of cancers, the pretraining of the slide aggregator is also supervised by inter-cancer contrastive learning. This approach brings samples of the same cancer type closer together while concurrently pushing samples of different cancer types apart.
In the second stage, the pretrained slide aggregator acquiring multimodal knowledge, can serve as the teacher model to seamlessly propagate multimodal knowledge at the slide-level context into the patch extractor, called Self-Taught Training (Fig. 2b). Specifically, the patch extractor is pretrained through encouraging the extracted patch features to be as similar as possible to those re-embedded by the pretrained aggregator. At the same time, to avoid catastrophic forgetting, we also enforce a similarity constraint between the extracted features and those embedded by the exponential moving average (EMA) patch extractor.
With these two stages, multimodal knowledge at the whole-slide context can be seamlessly embedded into foundation models. As a result, the model acquires the ability to comprehend both patches and the entire WSI, which facilitates downstream tasks at different levels. In the end, the pathology foundation model can achieve advanced abilities with the extended context from patch-level to slide-level and from unimodal to multimodal knowledge. More details of mSTAR can be found in Section 4.2.
Pathological diagnosis
We start with evaluating the pathological diagnostic capabilities based on pathological morphology, including pathological subtyping, metastasis detection, morphology prediction, pathological grading and pathological staging. These tasks commonly appear in pathology reports, forming a fundamental component of such reports and thus holding significant clinical importance. To evaluate these tasks, we collected 21 datasets from both publicly available and institutional sources consisting for 3 types of evaluation strategies, i.e., 8 independent cohorts on the 7:1:2 split, 3 held-out cohorts that are TCGA data held out from pretraining data and 10 external cohorts for testing only.
Specifically, for pathological subtyping task, we include breast cancer on BRCA-PathSubtype16 as a held-out cohort, brain tumor on GBMLGG_PathSubtype16 as a held-out cohort and EBrains_PathSubtype17 as an external cohort, head and neck cancer on HANCOCK_PathSubtype18 as an independent cohort, gastric cancer NFGC_PathSubtype, YN1_PathSubtype and YN3_PathSubtype as 3 external cohorts, lung cancer on TCGA-NSCLC16 as a held-out cohort and Lauren classification of gastric cancer on NFGC_Lauren and YN3_Lauren as two external cohorts, resulting in 3 held-out, one independent and 6 external cohorts. For metastasis detection task, we perform breast metastasis detection on CAMELYON19,20 as an independent cohort, lung metastasis detection on NF_Metastatic as an independent cohort and QFS_Metastatic as an external cohort, and meanwhile we further predict their primary locations on NF_Metastatic_Fine as an independent cohort and QFS_Metastatic_Fine as an external cohort. For morphology prediction, we assess whether perineural invasion is present on NFGC_Perineural as an independent cohort and YN3_Perineural as an external cohort, while we evaluate whether vascular invasion is present on NFGC_Vascular as an independent cohort and YN3_Vascular as an external cohort. Additionally, we evaluate pathological grading on PANDA21 as an independent cohort and pathological staging on HANCOCK-TStage18 as an independent cohort. The task distribution for pathology diagnosis is demonstrated in Fig. 3e, which covers common types of cancerous sites. The details of every dataset are described in Section 4.3.
a The overall performance on pathological diagnosis. b The performance on 8 independent datasets. c The performance on 10 external datasets. The red lines and the values reported at the top of figures (a, b and c) refer to the averaged performance across datasets. Each point represents a dataset, with the size of the point indicating the standard deviation. d The performance on 3 held-out datasets. The minima and maxima bounds of boxes represent the minimum and maximum performance among corresponding datasets, respectively. e Task distribution of pathological diagnosis across sites for different evaluation. f The overall performance on Pathological Subtyping across 10 datasets. g The performance on 6 external datasets of Pathological Subtyping. Error bars represent standard errors across datasets for all bar plots in (f–g). h, i The visualized validation of attention scores from mSTAR on h) CAMELYON and i) PANDA datasets. P-value for every group of experiments is given through one-sided Wilcoxon signed-rank test between mSTAR and the second-best FM. * represents P < 0.05, ** means P < 0.01 and *** indicates P < 0.001. Detailed Performances of every dataset are presented in Supplementary Fig. 2 and Supplementary Table 7. Source data are provided as a Source Data file.
As baselines, we evaluate the recent pathology foundation models (FMs) including PLIP 1, CONCH3, UNI2, CHIEF14 and GigaPath4 as well as the classical R5022. To perform these tasks, following the standard practice in computational pathology2, we used foundation models to extract features from each patch and adopted attention-based multiple instance learning (ABMIL)11 trained from scratch as the slide-level aggregator to perform slide-level prediction. ABMIL is a simple yet robust MIL approach, which is usually used for evaluation in previous foundation model research2,3. In particular, CHIEF and GigaPath claimed that their patch extractor should be used in conjunction with their pretrained aggregator. Therefore, we finetuned their pretrained aggregator paired with their extracted features on every downstream dataset to ensure the best performance.
All comparisons are based on the metric of Macro-AUC, a commonly used and objective classification metric, which does not rely on the selection of the decision threshold and is insensitive to the sample ratio of various classes. To examine statistical differences between mSTAR and the second-best FMs, the one-sided Wilcoxon signed-rank test was performed on various datasets.
From an overall perspective, we assessed the average performance for mSTAR and compared foundation models across 21 diverse datasets. The overall result demonstrates that mSTAR achieved the best performance with a +1.37% increase overall (P < 0.001) compared to the second-best model, UNI, as shown in Fig. 3a. Compared with slide-level FMs, mSTAR obtained +3.31% (P < 0.001) performance gain over GigaPath, the best slide-level baseline. From the perspective of consistency, mSTAR stood out on 18 out of 21 datasets, ranking at the first place. To evaluate the generalizability, we assessed 8 independent datasets and 10 external datasets. For independent cohorts, mSTAR demonstrates about +1% improvement (P < 0.01) compared to the second-best FM and +2.32% increase (P < 0.001) over GigaPath. It is worth noting that mSTAR exhibits superior generalizable capability on external cohorts with +2.14% improvement (P < 0.001) over the second-best FM, and meanwhile exhibits +4.16% (P < 0.001) increase over GigaPath. For held-out cohorts, mSTAR showcases performance comparable to that of other FMs.
Pathological subtyping is of utmost importance in clinical practice, as it forms the foundation for developing personalized treatment plans and enhancing treatment effectiveness. Therefore, we specifically evaluate mSTAR’s performance on such crucial tasks. The overall performance across 10 datasets demonstrates +1.69% increase (P < 0.01) compared to the second-performing FM. Furthermore, mSTAR shows significant improvement of +2.41% (P < 0.01) on external cohorts, suggesting the strong generalizable capability.
To validate whether the predictions of mSTAR align with clinical understanding, we visualize the predicted attention scores from mSTAR and compared the results with the given human-annotated ground-truth ROI on CAMELYON and PANDA datasets, as shown in Fig. 3h and i.
From the results of Fig. 3h and i, we can see the areas of interest for mSTAR successfully matches with the ground truth. These indicate mSTAR possesses intelligence for pathological diagnosis.
Molecular prediction
Molecular prediction has significant clinical implications in targeted therapy and risk stratification, allowing for tailored treatment plans. For example, HER2-positive breast cancer can be treated with HER2-targeted drugs like trastuzumab23. However, unlike pathological examination, genome sequencing remains largely inaccessible, especially in underdeveloped areas due to its high cost. Fortunately, with the benefit of pretraining on the modality combination of WSIs and gene expression data, mSTAR is more likely to possess a promising capability of molecular prediction only based on easily accessible pathological images. Therefore, in this study, we investigate molecular prediction solely based on pathological images for 3 categories of crucial molecular tasks, including gene mutation prediction, immunohistochemistry (IHC) biomarker prediction and molecular subtyping. To this end, we collected 35 datasets sourced from both public databases and medical institutions for evaluation and followed the same setting as the pathological diagnosis for slide-level prediction. The results are as follows:
Gene mutation prediction
Following the setting of CHIEF14, we predicted gene mutation related to FDA (Food and Drug Administration)-approved targeted therapies presented in OncoKB24 and high-frequency mutations25 in 10 cancer types held out from pretraining data. The positive mutation ratios are presented in Fig. 4g. The overall performance (Fig. 4a) exhibits the superiority of mSTAR in mutation prediction with +2.6% increases (P < 0.001) in Macro-AUC, compared to the second-best FM. On held-out cohorts, mSTAR surpassed the second-best FM by +2.81%. In particular, among the 18 genes Supplementary Fig. 3a and Supplementary Table 8–9, mSTAR excels in the prediction of ARID1A in endometrial carcinoma (UCEC) with +5.23% increase (P < 0.001), KRAS with +5.14% (P < 0.001) in lung adenocarcinoma (LUAD), GATA3 with +3.2% (P < 0.001) improvement and PIK3CA +2.46% (P < 0.001) increase in invasive breast carcinoma (BRCA), KRAS in cutaneous melanoma (SKCM) with +2.73% increase and EGFR with +1.81% (P < 0.001) in LUAD. All of these gene mutations have significant clinical relevance26,27,28,29,30,31,32, indicating the potential of mSTAR in biomedical research.
a Overall Performance of Gene Mutation Prediction on 23 datasets. b Performance of Mutation Prediction on 18 held-out datasets. c Overall Performance of Immunohistochemistry (IHC) Biomarker Prediction on 10 datasets. d Performance of IHC Biomarker Prediction on 4 independent datasets. e Overall Performance of Molecular Subtyping on 7 datasets. f Performance of Molecular Subtyping on 4 held-out datasets. In subfigures b, d and f, the minima and maxima represent the minimum and maximum performance among corresponding datasets, respectively, while the center and the bound of box represent the mean performance, 25% and 75% percentiles, respectively. The red lines and the values reported at the top of figures (a–f) refer to the averaged performance across datasets. Each point represents a dataset, with the size of the point indicating the standard deviation. g Positive and Negative Ratios of gene mutation for every mutation dataset, including genes with high-frequency mutations highlighted in green and genes related to FDA-approved therapies highlighted in red. h–j Internal (In) v.s. External (Ext) Evaluation. h Performance of Mutation Prediction on 5 internal and 5 external datasets. i Performance of IHC Biomarker Prediction on 3 internal and 3 external datasets. j Performance of Molecular Subtyping on 3 internal and 3 external datasets. Error bars represent standard errors across datasets for all bar plots in h-j. P-value for every group of experiments is given through one-sided Wilcoxon signed-rank test between mSTAR and the second-best FM. * represents P < 0.05, ** means P < 0.01 and *** indicates P < 0.001. Detailed performances of every dataset spanning 10 cancer types are presented in Supplementary Fig. 3 and Supplementary Table 8--11. Source data are provided as a Source Data file.
For external validation, mSTAR still obtains about 0.7 Macro-AUC on average and outperformed the second-best FM by about +2% improvements across 5 external cohorts (Fig. 4h). Specifically, mSTAR achieves +2.04% improvement (P < 0.001) in TP53 of BRCA, +2.06% increase (P < 0.001) in EGFR of LUAD and +1.11% increase (P < 0.001) in KRAS of LUAD.
Additionally, mSTAR predicts the mutation status in 14 of the 18 genes with Macro-AUC greater than 0.6 on held-out cohorts, as shown in Supplementary Fig. 3b and Supplementary Table 8. Mutations with excellent performance greater than 0.8 include TP53 in BRCA (0.8366; 95% CI 0.8105-0.8627) and glioblastoma multiforme (GBM) (0.8282; 95% CI 0.7780-0.8784), CIC in low-grade glioma (LGG) (0.9157; 95% CI 0.8952-0.9362), PTEN in UCEC (0.9008; 95% CI 0.8737-0.9279). This showcases mSTAR can provide reliable prediction of these crucial biomarkers33,34,35,36,37 for biomedical research.
IHC biomarker prediction
Immunohistochemistry (IHC) is widely used in clinical pathology, primarily for detecting specific proteins in tissue samples and distinguishing between tumors with similar pathological features, enabling more precise targeted therapy and improving patients’ outcomes. However, IHC examination usually requires extra expensive costs. Therefore, if easily accessible H&E slides can be used to predict IHC biomarkers, it would significantly advance the widespread adoption of precision cancer diagnostics, especially in underdeveloped areas. To assess mSTAR’s performances on IHC biomarker prediction, we collected 10 datasets for evaluation, including 3 held-out datasets and 3 external as well as 4 independent datasets from collaborative medical institutions. Specifically, we involved common biomarkers comprising ER, HER2, PR and CK5 for breast cancer, along with CK7 for lung cancer.
From an overall perspective, mSTAR outperforms the second-performing FM by +1.8% (P < 0.001, Fig. 4c). Across 4 independent datasets, mSTAR performs the best overall (Fig. 4d) by +1.44% (P < 0.001) over UNI, the second-best FM, while consistently surpassing other FMs on all 4 independent datasets (Supplementary Fig. 3b) with significant differences (P < 0.001). Specifically, mSTAR can achieve over 0.8 of Macro-AUC on 3 out of 4 tasks including ER and CK5 for breast cancer and CK7 for lung cancer, as well as almost 0.8 of Macro-AUC on HER2 (0.7951 ± 0.0125). This indicates mSTAR is capable of offering a reliable prediction for these vital and common biomarkers, probably resulting in a great reduction of the cost of IHC examination.
To test the generalization of mSTAR, 3 external datasets spanning ER, PR and HER2 in breast cancer are collected from the collaborative hospital for evaluation. The overall performance is presented in Fig. 4g. mSTAR consistently showcases superiority in both internal and external evaluations with +1.81% (P < 0.001) and +1.02% (P < 0.001) increases over the second-best FMs, UNI and CONCH, respectively. For the examination of ER (Supplementary Table 10), although we observed a decline in performance compared to the internal cohorts, mSTAR still maintains Macro-AUC above 0.85 (0.8526 ± 0.0071), resulting in the promising generalization in the external cohort.
Molecular Subtyping aims to categorize cancers based on their molecular and genetic characteristics, thereby assisting in identifying patients with distinct responses to treatment, and prognostic outcomes. In this study, we investigate molecular subtyping on 4 cancers including Breast Invasive Carcinoma (BRCA), Colon Adenocarcinoma and Rectum Adenocarcinoma (CRC), Glioblastoma Multiforme and Brain Lower Grade Glioma (GBMLGG) and Head and Neck Squamous Cell Carcinoma (HNSC) on 4 held-out datasets (BRCA_MolSubtype, CRC_MolSubtype, GBMLGG_MolSubtype and TCGA_HNSC_HPV) and 3 external datasets (ZJ1_Breast MolSubtype, EBrains_MolSubtype and HANCOCK_HPV).
From the overall perspective, we observed a performance gain of +1.78% (P < 0.001) over UNI, the second-best FM (Fig. 4e). When we delve into different evaluation strategies, +1.24% (P < 0.001) improvement can be seen in held-out cohorts (Fig. 4f). When taking a close on internal and external cohorts of breast, brain and head&neck cancers (Fig. 4j), mSTAR surpassed CHIEF (the second-best FM on internal cohorts) by +1.77%, while exceeding UNI (the second-best FM on external cohorts) by +1.1%. It is worth noting that in the FMs compared, mSTAR is the only one that maintains an AUC above 0.85 for both internal and external datasets, demonstrating strong generalization ability. Furthermore, mSTAR keeps the consistent superiority over 3 external cohorts (Supplementary Table 11).
To sum up, through the joint pretraining of pathological images and gene expression data, mSTAR demonstrates superior performance and strong generalization across mutation prediction, IHC biomarker prediction and molecular subtyping. This capability can provide reliable predictions in clinical applications and biomedical research, making it possible to utilize accurate molecular information in a cost-effective manner.
Vision-language evaluation
Strong language-related capabilities are one of the key features of foundational models, reflecting their potential in open-world scenarios where downstream tasks are conducted without further training, that is, zero-shot learning capability, especially in resource-constrained scenarios where access to sufficient data and computational resources may be limited. Furthermore, pathological report writing is a time-consuming process in pathologists’ clinical workflow. As such, automatic report generation can significantly streamline workload for pathologists, which also heavily relies on the foundation model’s language capabilities. With the benefit of the involvement of pathology reports during pretraining, mSTAR is expected to possess great language capabilities. Therefore, in this study, we assess mSTAR’s language abilities from three aspects, that is, zero-shot slide classification, zero-shot retrieval and report generation.
Zero-shot’s capability always relies on a well-aligned vision-language space. Therefore, we use vision-language foundation models as baselines, i.e., PLIP and CONCH. To produce slide-level predictions for patch extractors, following the setup of CONCH3, MI-Zero38 was adopted through top-K patches voting based on patch similarities to class prototypes or reports embedded by the pretrained text encoders (Fig. 5a).
a The scheme of zero-shot evaluation. For zero-shot classification, we used class prompts as the text input. For zero-shot retrieval, the text input is a pathology report. b Performance of zero-shot slide classification on 6 independent datasets. The ʻOverallʼ refers to the averaged performance across these 6 datasets. Error bars represent 95% CI with 1000 bootstrap replicates for all bar plots. P-value is given through one-sided Wilcoxon signed-rank test between mSTAR and the second-best FM. c Performance of zero-shot retrieval on an external dataset for Image-to-Text and Text-to-Image tasks. The results on held-out TCGA dataset are presented for reference only to be compared with zero-shot’s capability. d Performance of report generation on one held-out TCGA dataset and two external datasets. P-value for every group of experiments is given through one-sided Wilcoxon signed-rank test between mSTAR and the second-best FM. Detailed performances of every dataset are presented in Supplementary Table 15–17. Source data are provided as a Source Data file.
Zero-shot Slide Classification
In this study, we assess every FM on 6 slide classification tasks independent from TCGA data, CAMELYON, PANDA, UBC-OCEAN, BCNB-ER, BCNB-PR and BCNB-HER2.
Across 6 tasks, mSTAR outperforms other FMs on half of the tasks and performs best on the overall result (Fig. 5b and Supplementary Table 15). Specifically, compared to the second-best FM, mSTAR achieves clear enhancement in these tasks by +3.9% on average (P < 0.001) with a significant difference. In particular, on CAMELYON, a remarkable rise of +10.4% (P < 0.001) is observed compared to CONCH, the second-best FM. Furthermore, we see performance enhancement over the second-best FM by +16.2% (P < 0.001) on BCNB-ER and +19.9% (P < 0.001) on BCNB-PR, respectively.
Zero-shot slide retrieval
The capability of zero-shot whole-slide retrieval can assist pathologists in seeking similar cases for reference, effectively enhancing diagnostic precision and consistency as well as reducing the workload for pathologists.
In this study, we explore two settings: Image2Text refers to providing an image for the model to find the corresponding report, while Text2Image does the reverse. Although the source of held-out data is the same as pretraining data, the data itself is totally held out from pretraining data. We presented results on held-out data for reference only to be compared with zero-shot’s results on external cohorts. The results (Fig. 5c and Supplementary Table 16) demonstrate that mSTAR has a clear and significant advantage in this dataset, since mSTAR successfully aligned the vision and language spaces during the pre-training stage. To investigate whether mSTAR can exhibit the advantage on external data, we collected a dataset spanning breast and lung cancers from collaborative hospitals, comprising 500 cases of WSI-Report pairs. Despite performance decreases on the external cohort, results (Fig. 5c and Supplementary Table 16) demonstrate that mSTAR still performs the best among all vision-language FMs, with +9.4% of Recall@50 on Text2Image and +3.6% of Recall@50 on Image2Text.
Report generation
Automated generation of pathology reports has enormous potential in simplifying the report-writing process and reducing the workload burden on pathologists. To assess mSTAR’s capability of report generation, we collected one pan-cancer TCGA dataset with 840 cases held out from pretraining data and two external cohorts including Nanfang of lung cancer from 250 patients and ZJ-First of breast cancer from 250 patients. Since pathology reports generally include numerous contents invisible in whole slide images, such as macro descriptions, we first leverage GPT-4o-mini to filter out these irrelevant descriptions. The prompts used for cleaning reports are presented in Supplementary Table 26. The detailed process regarding the quality control for reports is presented in Section 4.3. In this study, we finetuned HistGen39, a pathology report generation model, based on patch features extracted by different foundation models.
From the quantitative perspective, we evaluated multiple metrics including BLEU, METEOR and ROUGE-L to assess various aspects of the generated text, such as precision of n-grams (contiguous sequences of words), order, alignment, recall, etc. In the held-out TCGA cohort, across different metrics, mSTAR consistently outperformed the second-best approach (Fig. 5d and Supplementary Table 1). In one external cohort, Nanfang, we observe significant improvements in these three metrics compared to the second-best FM: +6.91% of BLEU_1, +1.17% of METEOR and +4.61% of ROUGE_L. This indicates mSTAR has a better generalizable ability for report generation, instead of just memorizing the contents of reports. In another external cohort, ZJ-First, mSTAR demonstrates increases in BLEU metric, indicating mSTAR excels at generating precise long sentences. For METEOR, mSTAR achieves a comparable performance with the second-best FM, while a performance decline of 1.41% is present in ROUGE_L, which indicates the generated texts of mSTAR are less fluent.
We continued to qualitatively evaluate the quality of generated reports. The case studies for every cohort are presented in Supplementary Fig. 7. The texts highlighted in red are matched with the ground-truth report, while the ones highlighted in blue contradict the true report.
First, we investigate how mSTAR and the competitive FMs perform on held-out datasets. For the case (a) and (b) in Supplementary Fig. 7, the texts generated by R50 are almost unrelated to the ground truth and always the same. The reason is probably the pretraining materials are not specific to the pathology domain. The diversity of generated texts from PLIP, CONCH and CHIEF is better than that of R50. For example, PLIP, CONCH and CHIEF can identify the histologic type and simple margin information for case (a), despite missing more details about diagnosis. However, CONCH and CHIEF showcase poor relatedness and the generated texts from PLIP are too short for case (b). UNI and GigaPath are aware of more content types that need to be generated, although the specific predictions are always inaccurate. In other words, the generated texts from UNI and GigaPath contain a lot of hallucinations contradicting to the ground-truth report, such as case (b). mSTAR is able to identify the necessary content and make more accurate predictions cautiously, leading to fewer hallucinations. However, it still fails to count, such as the number of lymph nodes. Then, when examining the external cohorts closely, they demonstrate the same characteristics. This indicates that mSTAR has the generalizable capability of report generation, instead of just memorizing the template reports.
Given that the current generation capabilities are not perfect, mSTAR excels compared to other models overall. This can be attributed to two main factors. First, these foundational models are encoder-based and do not incorporate decoders during pretraining, resulting in a significant distribution gap between encoded features and generated texts. Second, the effectiveness of existing report generation methods that are fine-tuned on foundational features remains restricted.
Survival prediction
Prognostic analysis is an intricate clinical endeavor, which can inform clinical guidelines and practices, helping healthcare providers make evidence-based decisions regarding patient care. It is so complicated that it always necessitates a thorough analysis from a multitude of facets. In this regard, multimodal data has proven instrumental in enabling more comprehensive prognostic assessments8,9,10,40. Therefore, it is crucial to explore the role of multimodal knowledge within the broader whole-slide context in enhancing prognostic estimation. In this study, we assessed 3 prognostic tasks, Overall Survival (OS), Disease-Free Survival (DFS) and Recurrence-Free Survival (RFS) on top of pathological tissue slides. All endpoints in our study were chosen based on their broad usage in clinical oncology and alignment with public datasets. We believe this preserves the clinical utility that makes our findings meaningful for both researchers and practicing oncologists. To this end, we collected 10 held-out cohorts covering 10 cancer types from TCGA, and 4 external and 2 independent cohorts for generalizable validation from public databases and collaborative medical institutions. The distribution relationship between tasks and cohorts can be seen in Fig. 6c. The distribution of samples for every cohort is presented in Supplementary Table 1.
a Comparison of C-Index between mSTAR and compared methods on 9 held-out datasets. b Comparison of C-Index between mSTAR and compared methods on 4 external datasets. The red lines and the values reported at the top of figures (a, b) refer to the averaged performance across datasets. Each point represents a dataset, with the size of the point indicating the standard deviation. c Task distribution of various survival endpoints for different evaluation. d The performance (C-Index and 95% CI) on independent cohorts. `out' refers to the partitions held out from pretraining data. `idpt' means independent datasets with a data source that differs from the pretraining data. `ext' represents external datasets where data originates from a source distinct from the training data used for fine-tuning and is used solely for testing, without any training involved. Error bars represent 95% CI with 1000 bootstrap replicates for all bar plots. P-value for every group of experiments is given through one-sided Wilcoxon signed-rank test between mSTAR and the second-best FM. * represents P < 0.05, ** means P < 0.01 and *** indicates P < 0.001. Detailed performances of every dataset are presented in Supplementary Table 18. Source data are provided as a Source Data file.
First, we investigate the performance of held-out cohorts over 10 datasets, which demonstrates a slight improvement of +0.5% (P < 0.001) overall, compared to UNI, the second-best FM. For consistency of performance increases, mSTAR performed best compared to other foundation models, achieving the top performance on 6 out of 10 datasets. However, UNI, ranking in the second place, performs the best on 2 out of 10 datasets.
Although the improvement in the held-out cohort is not promising, mSTAR demonstrates strong generalization, achieving an average increase of +2% (P < 0.001) on 4 external cohorts, especially on OS_ZJ1 of breast cancer for overall survival with +2.88% increase (P < 0.001). Across 4 external cohorts and 2 independent cohorts, mSTAR also shows consistent superiority, which indicates a great generalizable ability of survival prediction.
To demonstrate patients’ stratification performance, we examined the Kaplan-Meier curves characterized by mSTAR for every task (Supplementary Fig. 5), where 12 out of 14 tasks showcased the statistical difference between low-risk and high-risk groups, via Logrank Test41.
Multimodal fusion
Multimodal data typically provides a more comprehensive understanding of cancer, excelling in various clinical applications, such as treatment response prediction for neoadjuvant chemotherapy42 and prognostic analysis43. However, multimodal data integration often suffers from the heterogeneity of different modalities, leading to limited performance. As a result, whether pathological features from foundation models can be well aligned to other modalities plays a crucial role in multimodal analysis. With the benefit of multimodal pretraining, they can align with each other by contrastive learning, thereby potentially alleviating inter-modal heterogeneity. Therefore, in this study, we examine whether mSTAR facilitates multimodal fusion by assessing multimodal overall survival prediction tasks.
To validate this, we replaced pathological features with ones extracted by various extractors in existing multimodal fusion models for 9 cancer survival prediction tasks held out from pretraining data, to observe the differences that would arise. Specifically, to reduce biases caused by multimodal integration approaches, 4 recent multimodal fusion models were employed in this study to make the multimodal slide-level prediction, including MCAT8, Porpoise43, MOTCat9 and CMTA10.
On the whole, mSTAR has clearly outperformed other SOTA methods by a wide margin. Considering average rank, mSTAR ranked between 1.22 and 1.67 among various fusion models and the overall rank is 1.47, which left the second-best approach UNI far behind (Fig. 7a) ranking at 2.68 on average. For average C-Index (Fig. 7b), mSTAR achieved consistent and notable enhancement in multimodal fusion with a significant difference, with average performance increases of +1.8% (P < 0.001). Among different multimodal fusion models, mSTAR outperformed the second-best FM by +1% (P = 0.02) for MCAT, +1.1% (P < 0.01) for Porpoise, +2.4% (P < 0.005) for MOTCat and +1.4% (P < 0.005) for CMTA.
The patch extractors of all foundation models are evaluated with different multimodal fusion models (MCAT, Porpoise, MOTCat and CMTA), trained from scratch across 9 TCGA held-out datasets. a Performance of Ranking on 9 datasets of each FM on every multimodal fusion models and “Overall” that refers to the average results among these multimodal fusion methods. b The average C-Index on 9 datasets. c Performance (C-Index and 95% CI) on each dataset. The minima and maxima represent the lower and upper bounds of 95%CI, respectively. The center and the bound of box represent the mean value, 25% and 75% percentiles, respectively. P-value is given through one-sided Wilcoxon signed-rank test between mSTAR and the second-best FM. The colors of legends are shared across all sub-figures. * represents P < 0.05, ** means P < 0.01 and *** indicates P < 0.001. Detailed performances of every dataset are presented in Supplementary Table 19–23. Source data are provided as a Source Data file.
Across various datasets, mSTAR surpassed the second-best FM on 8 out of 9 datasets (Fig. 7b and Supplementary Table 19). Among them, +2.22% on BRCA (P < 0.001), +2.55% on CRC (P < 0.001), +2.17% on HNSC (P < 0.001), +1.89% on LUSC (P < 0.001), +1.83% on UCEC (P < 0.001) are achieved with statistically significant differences. Specifically, based on MCAT, mSTAR surpassed other SOTA approaches on 5 out of 9 tasks (Supplementary Table 20), especially on BRCA (+1.9%, P < 0.001). mSTAR with Porpoise demonstrated superior performance in the majority of tasks, topping 6 out of 9 datasets (Supplementary Table 21), which increased the second-best model by up to +3.8% (P < 0.001). In the case of MOTCat, mSTAR excelled in 6 out of 9 tasks (Supplementary Table 22) with performance increases of up to 3.2% (P < 0.001). For CMTA, mSTAR delivered the highest performance in 6 of 9 (Fig. 7f and Supplementary Table 23), advancing the second-best one by up to 2.9% (P < 0.001).
In a nutshell, the remarkable increases across various datasets and diverse multimodal fusion backbone models vividly demonstrate the tremendous contributions of multimodal knowledge embedded by slide-level multimodal contrastive learning in facilitating multimodal fusion.
Ablation study
In this section, we first quantitatively investigate the impacts of different modalities and each component of pretraining objectives Fig. 8a and c), and disentangle the architecture design of Self-Taught Pretraining paradigm by intuitively showcasing the evolution of feature space Fig. 8 b). In the end, the resource efficiency and effectiveness of scaling vision data only or modalities are explored Fig. 8d).
a averaged performance on pathological diagnosis (3 datasets), molecular prediction (12 datasets) and survival prediction (9 datasets), where ʻBeforeʼ refers to before pretraining, and ʻPʼ, ʻTʼ and ʻGʼ indicate pathology slides, pathology reports and gene data, respectively. Error bars represent standard errors across datasets for all bar plots. b visualization of feature space evolution: from before pretraining (initial) to Stage 1 (pretrained aggregator) and Stage 2 (mSTAR), where the areas in red bounding box are multiple tumor regions (1-7) of the case of patient_042_node_3 of CAMELYON17 dataset. Note that different tumor areas correspond to different spatial positions. c averaged performance (9 TCGA OS datasets) for ablating different pretraining objectives (Inter-modal Loss and Inter-cancer Loss) for survival prediction (Supplementary Table 4). d averaged performance (24 datasets) and resources comparisons between scaling slides only (Virchow) v.s. scaling modalities (mSTAR) for pretraining, with UNI as a baseline. Detailed performances of every dataset are presented in Supplementary Fig. 8 and detailed comparisons are showcased in Supplementary Table 5–6. Source data are provided as a Source Data file.
Impact of different modalities
We conducted ablation studies on different combinational modalities and demonstrated the effectiveness of every part based on changes in performance. The purpose of this experiment is to explore how much various modalities contribute to performance gains; hence, it is only involved in the pre-training of the aggregator at Stage 1. Therefore, to demonstrate its contribution, we only need to compare the performance of pretraining on three modalities with ‘before aggregator pre-training’ and ‘after pre-training with two modalities’.
To this end, we have systematically evaluated proportional datasets for each task category (24 datasets). From the results in the Fig. 8a, we observed that the impact of multimodal data on performance gains differs significantly depending on the task type, and the synergy of three modalities resulted in further improvements. The results demonstrate consistent performance improvements across all types of tasks, with a notable average increase of 5.3% specifically observed in the 12 molecular prediction tasks compared to that of only 2 modalities. More interestingly, we found that text modalities outperform genomic data in diagnostic tasks, which intuitively corresponds to the diagnostic-rich nature of pathology reports. For molecular and prognostic predictions, textual and genomic data demonstrated comparable performance, consistent with prior findings44 that both phenotype (text-derived) and genotype features contribute to stratification.
Impact of each component in pretraining objectives
For the ablation of pertaining loss functions, we also estimate their influences by removing them during pretraining, and observe the changes in performance. Results in Fig. 8c showcases that both objective functions make a difference in the pretraining, and their combination elevates performance to a higher level.
Disentangling self-taught pretraining paradigm
Self-Taught Pretraining paradigm consists of two stages. Stage 1 focuses on training the aggregator while keeping the extractor frozen. Its effectiveness is evaluated by ablating the contribution of the third pre-training modality (as previously discussed). Stage 2 performs self-taught training of the extractor with the aggregator frozen. Here, the extractor’s improvement is validated by comparing its performance to the pretrained UNI baseline, with results verified across 97 tasks. Furthermore, from a feature-space perspective, we visualize the evolutionary dynamics of each stage in Fig. 8b. The clusters progressively coalesce, demonstrating clear separation between tumor and non-tumor regions as training advances. Both quantitative performance evaluation and qualitative feature-space visualization consistently verify the effectiveness of each training phase.
Modality scaling versus vision data scaling
Recent studies have claimed that substantially increasing the number of slides for unimodal pretraining can significantly improve model performance, e.g., Virchow45, a vision-only PFM pretrained on 1.5 million slides. To investigate this, we compared the efficiency of scaling multimodal data versus scaling unimodal visual data alone. Our findings reveal that unimodal scaling exhibits limited efficiency and poor generalization, whereas multimodal pretraining achieves superior performance even with modest data volumes. Multimodal pretraining has the potential to circumvent the substantial effort required for large-scale slide collection.
Parameter-wise, compared to Virchow of 631M, mSTAR merely consists of 415M (34% reduction) parameters including the 303M visual encoder (the same as UNI), 2.67M TransMIL module, 0.94M genomic scBERT, and 108M BioBERT text encoder (smaller than CONCH). Data-wise, when benchmarked against UNI’s performance baseline, we compared mSTAR with Virchow as shown in Fig. 8d. mSTAR achieves competitive improvements with only 22K additional slide-level pretraining samples–a far smaller fraction of the 1.39M extra slides required by Virchow for better performance gains. This 53 × reduction in additional data demonstrates that multimodal data brought higher efficiency per sample compared to vision-only brute-force data scaling, significantly reducing pretraining costs in GPU-hours (Supplementary Table 5) while even enhancing clinical-grade accuracy. Our findings offer a remarkable advantage and a practical pathway in scaling pathology foundation models, especially for resource-constrained medical AI development where large-scale data collection is often impractical. Training-wise, as shown in Supplementary Table 5, while UNI requires 4x8 A100 80GB GPUs (4 nodes, each with 8 GPUs) for 32 GPU hours (1024 GPU hours in total), mSTAR merely needs 4 H800 80GB GPUs (1 node, each with 4 GPUs) for 7 days (672 GPU hours in total). While Virchow does not report exact training durations, its substantially larger model size (631M vs. our 414M parameters) and 53 × greater pretraining data (1.39M vs. our 22K slides) inevitably require far greater computational resources. More comparison can be seen in Supplementary Table 5–6.
Discussion
In this study, we delve into how to harness the full potential of three-level multimodal data to advance the performance of the pathology foundation models effectively, by pretraining the model on over 116 million pathological images of 26k modality pairs from 10,275 patients across 32 major cancer types. Additionally, we explored a new whole-slide pretraining paradigm for CPath, which broadened the context of modeling for better performance on slide-level tasks. For systematical evaluation, we established the largest spectrum of oncological benchmark datasets, covering 7 categories of oncological applications comprising 15 types of 97 oncological tasks. With the benefit of the involvement of pathology reports and gene expression data in pretraining, diverse experimental results demonstrated that mSTAR excelled in not only molecular prediction but also pathological tasks frequently presented in pathology reports at the slide level, such as pathological subtyping, mutation prediction and report generation. Furthermore, multimodal pretraining facilitated multimodal fusion tasks due to a well-aligned multimodal space and endowed the model with more generalized zero-shot’s capabilities.
In the realm of prior investigations into pathology foundation models, two prominent categories have emerged: vision-only models2,4,46 and vision-language models1,3. However, these approaches fail to tap into a vast wealth of information inherent in macroscopic-level pathology reports written by experts and molecular-level gene expression profiles. Pathology reports usually provide authentic expert knowledge in line with the clinical practice, while gene expression profiles serve as robust indicators of oncology status for clinical applications in diagnosis47 and prognosis48. As shown in Fig. 8 and Supplementary Table 3, the involvement of pathology reports and gene expression data can bring extra performance gains. The superiority in molecular prediction and report-related oncological validates modality scalability in pathology foundation models, which potentially provides a guiding conclusion: pathology foundation models can benefit from a more diverse range of modalities.
Recently, beyond working on small patches/ROIs, we noticed that some studies4,14 attempted to work on slide-level foundation models, which pretrained the model on patch features. However, the pretrained performance significantly depends on the quality of patch features, leading to under-performing results compared to mSTAR. In other words, their performance would be limited by the patch extractor. We believe that end-to-end pretraining is a promising solution in the future, while its current implementation is hindered by hardware limitations. Therefore, mSTAR bridges this gap through self-taught training to seamlessly transfer the knowledge captured by the slide aggregator into the patch extractor.
Distinct from previous researches, our study provides the uniqueness in three folds. First, our findings showcase the remarkable power of leveraging multimodal data, especially in enhancing multimodal capabilities. This validates the scalability of modalities, providing the guiding principle for building pathology foundation models. We show that multimodal integration yields disproportionately higher returns than unimodal scaling, offering a pathway to develop performant PFMs without requiring massive slide collections. Second, we found a unified way to bridge the gap between slide-level and patch-level pretraining, bringing us closer to achieving end-to-end pretraining on raw slide data. We believe this innovative unified paradigm will revolutionize the workflow of pretraining for CPath. Moreover, this paradigm allows the injection of multimodal knowledge into pathology foundation models in an appropriate manner, which may hold the potential to harness more modalities to construct a stronger foundation model for CPath. Third, we established the widest range of oncological benchmarks spanning 7 categories of 15 types of 97 oncological tasks.
Although preliminary results are encouraging, this study still has several limitations. First, the challenge of collecting paired multimodal data naturally limits the scale of pretraining data, compared to previous works of pathology foundation models. By expanding the scale of multimodal data for pretraining, we can expect to unlock further potential for enhancing various abilities, such as multimodal capabilities. Fortunately, the growing accumulation of slide data enables future validation of scaling laws in multimodal contexts, a previously unexplored frontier in computational pathology. Second, the present investigation was limited to three modalities; subsequent studies incorporating a broader range of modalities, e.g., IHC, specialized stained slides, and spatial transcriptomics, will be essential for robust validation of the modality scaling law. Third, we still potentially have a long way to go before achieving the true end-to-end foundation model. Before that, mSTAR will serve as an alternative solution to seamlessly bridge slide-level and patch-level pretraining. However, there are still several challenges to be further explored, such as the appropriate way to propagate the pretrained knowledge embedded in the slide aggregator and the architectural design of slide aggregator. In mSTAR, due to a large number of patches of a WSI that would lead to extremely high computational costs, we selected TransMIL with linear time complexity as the slide aggregator. However, the increase in training speed comes at the expense of sacrificing a portion of the performance. Fortunately, a multitude of innovative architectures for handling long sequences are emerging, such as Mamba49, LongNet50, etc, which we explore in concurrent work51. We believe that these new architectures will undoubtedly create new avenues in exploring more efficient and powerful pretraining paradigms for CPath. Lastly, due to the inherent challenge of gathering rare cancer data, there is still a need for further assessment of zero-shot’s performance on real rare cancer cases, although zero-shot’s performance can reflect the performance under the situation of limited data to some extent. In the future, we plan to incorporate more multimodal data into pretraining, such as multi-omics data, explore new efficient pretraining architectures, and keep moving forward in the collection of rare cancer data.
Method
This study has been reviewed and approved by the Human and Artefacts Research Ethics Committee (HAREC). The protocol number is HREP-2024-0212.
Pretraining dataset curation
Data used for pretraining in this study were totally obtained from a publicly available source, the Cancer Genome Atlas Program (TCGA)16, in which we collected 9640 cases (11,765 slides) of diagnostics formalin-fixed paraffin-embedded (FFPE) H&E WSIs, 11,108 pathology reports and 10,234 cases of bulk RNA-Seq data across all 32 cancer types of TCGA. After quality control, we curated a dataset with 8440 WSI-Report pairs, 8965 WSI-RNA-Seq pairs and 8764 Report-RNA-Seq pairs, resulting in 26,169 modality pairs, as shown in Fig. 1c. These data involve over 116 million pathological patch images. Given that numerous downstream tasks were evaluated on TCGA data, we held out some validation and test cases. For 9 cancer datasets comprising over 400 cases, we adopted a split ratio of 7:1:2 for train-validation-test folds. For those cases involving multiple slides, we combined their patches or features into a single case for pretraining at the patient level. This ensured slides belonging to one case were included within the same fold, thereby preventing potential data leakage. We also considered label stratification for survival labels at patient-level, since we primarily evaluated the performance of survival prediction on TCGA data. Note that all cases without survival labels were used for pretraining. Details of data splitting for these 9 cancer datasets are provided in Supplementary Table 24. After data partitioning, we curated 22,127 modality pairs for contrastive learning, consisting of 7083 WSI-Report pairs, 7538 WSI-RNA-Seq pairs and 7506 Report-RNA-Seq pairs. Among these, there were 7947 cases with all three modalities for pretraining. For acquisition of high-quality data, we conducted the subsequent pre-processing procedures for each modality.
WSI pre-processing
To conduct slide- (or patient-) level tasks on WSIs, our processing pipeline involved tissue segmentation, patching, and feature extraction (for pretraining aggregators and evaluation). For tissue segmentation, we employed the CLAM library12, which performed binary thresholding on the saturation channel of a downsampled RGB slide, converted to the hue-saturation-value (HSV) color space. The resulting segmentation mask was obtained by filtering the contours based on their area. The hyperparameters of segmentation are released on our codebase. Furthermore, slides that were corrupted and those containing a small proportion of tissue region were excluded from this study. As a result, we acquired 9608 cases of 11,727 slides for pretraining and evaluation.
To adhere to established practices of previous works2,3,12, we partitioned the segmented tissue regions into 256 × 256 pixels patches at 20 × -equivalent magnification without overlaps and then resized all patches to 224 × 224 pixels for feature extraction. Using pretrained patch extractors that were kept frozen, we pre-extracted embeddings for each patch and stored them for subsequent evaluation purposes.
Report pre-processing
For pathology reports, we curated open-source texts from TCGA and converted them from their original PDF format to editable text format via Amazon Web Services (AWS) Optical Character Recognition (OCR) tools, resulting in 9523 Reports. For quality control, we curated these reports via the powerful language tool, GPT-4, with appropriate prompts provided in Supplementary Table 25, and re-checked them manually to ensure the unchanged original intent. The statistical distribution of word counts for reports is demonstrated in Fig. 1e, in which the majority of cases have word counts below 500.
RNA-Seq Pre-processing
We accessed RNA-Seq data of TCGA from cBioportal database, which were preprocessed and normalized using RSEM52. An inherent difficulty in gene expression modeling arises from the variations in absolute magnitudes observed across different sequencing protocols53. Therefore, we further applied a common preprocessing technique log1p transformation54 for gene expression values. Following previous works55, Gene2Vec56 contributed to better representing the gene names by enforcing that words with similar meanings are assigned similar representations. Therefore, we retained genes present in the Gene2Vec vocabulary. In the end, we obtained 9890 cases RNA-Seq data, each consisting of genes with a length of 17,425.
Pretraining framework
To utilize multimodal knowledge at the whole-slide context for enhancing the pathology foundation model, we propose a whole-slide pretraining paradigm consisting of two-stage pretraining, as shown in Fig. 2. In the first stage, we aim to inject multimodal knowledge into the slide aggregator by contrastive learning, including inter-modality contrastive learning (following CLIP57) and inter-cancer contrastive learning. In the second stage, to seamlessly propagate multimodal knowledge at the slide-level context into the patch extractor, we leverage the slide aggregator pretrained in the first stage, serving as a “Teacher” model, to supervise the pretraining of the patch extractor, termed Self-Taught training. In this way, multimodal knowledge of the whole-slide context can be injected into the pathology FM.
Stage 1 - pretrain slide aggregator
In this stage, we aim to pretrain a slide aggregator that learns multimodal knowledge by contrastive learning with other modalities. Note that the pretrained slide aggregator plays a role of “Teacher” that propagates the learned knowledge into the patch extractor at the next stage. These modules to be trained are highlighted in red boxes in the Fig. 2a, in which we pretrain a 2-layer TransMIL13 as the slide aggregator for WSIs, a Bert-like text encoder (following BioBert-Base-v1.258) for pathology reports, and a Performer (following scBERT55) for RNA-Seq data.
Given these transformer-like encoders, we need to tokenize raw data of every modality into token embeddings before feeding them into their respective encoders. For pathology, we obtained non-overlapping 224 × 224 patches as early mentioned, and then for every patch, we used a pretrained patch extractor, UNI2, to extract patch features, where a patch feature was regarded as a token embedding for the slide aggregator. After gathering 4096 patch features for the i-th patient’s WSIs, \({{{{\mathcal{P}}}}}_{i}={\{{{{{\bf{p}}}}}_{i}^{m}\}}_{m=1}^{M}\), we fed them into the slide aggregator to integrate all patch features and got a 512-dimensional pathological [CLS] token embedding Pi as the slide-level representation, where M is the number of patches and it was fixed into 4096. For cases where the number of patches exceeds 4096, a random selection of 4096 patches is made, while for cases with fewer than 4096 patches, padding is applied using the mean value. For those cases where one patient has more than one WSI, we simply concatenated them together. Note that all patch features were transformed into 512-dimensional features by a linear projection before being forwarded into the aggregator.
For pathology reports, we adopted the text encoder for randomly truncated 512 tokens and outputted the report [CLS] token embedding Ti. For cases where the length of the text is less than 512, the special token ’[pad]’ was padded. The RNA-Seq data was organized as a set of 2-tuple (gi, ei) comprising of the gene name gi and its expression variable ei. Following previous works55,59, to assure that genes with potential co-expression get close together, we employed Gene2Vec56 to generate 200-dimensional gene embeddings for each gene name gi. Gene expression can be viewed as the manifestation or presence of each gene, which has been well-documented within a biological system. Therefore, we applied the term-frequency-analysis method used in previous works55,59 to discretize the continuous expression variable ei through binning technique. Subsequently, the discrete variable was transformed into a 200-dimensional embedding, which was then integrated into the final gene token embedding gi by addition. Through forwarding the gene encoder, we can get the gene [CLS] token embedding Gi. It is worth noting that encoder outputs from report and gene modalities were transformed into 512-dimensional features by a linear projection for contrastive learning.
To optimize the model through pretraining, we incorporate two objectives including inter-modality contrastive learning and inter-cancer contrastive learning. In the case of inter-modality contrastive learning, given the [CLS] representation of each modality, every two modalities can be paired together, which finally yielded three combinations: WSI-report (Pi, Ti), WSI-gene (Pi, Gi) and report-gene (Ti, Gi). During pretraining between every modality pairs, a mini-batch consisted of N samples, e.g., \({\{({{{{\bf{P}}}}}_{i},{{{{\bf{T}}}}}_{i})\}}_{i=1}^{N}\) for WSI-report. Contrastive learning imposes a higher similarity in modality pairs from the same sample. Take WSI-report pairs as an example, and the loss function can be formulated as
where τ is a scale factor of the contrastive loss and it was set by default following CLIP57. Similarly, we can get \({{{{\mathcal{L}}}}}_{P-G}\) and \({{{{\mathcal{L}}}}}_{T-G}\) and finally combine them by addition.
To alleviate the heterogeneity of various cancer types, we utilized inherent cancer labels available in TCGA for the inter-cancer pretraining objective. Specifically, [CLS] tokens of available modalities (regardless of whether they involved two or three modalities) would be concatenated into a single anchor representation ai. Furthermore, positive and negative samples were obtained within the mini-batch, and they were from the same cancer and different cancers, respectively. Similarly, they were constructed in the same way by concatenating the [CLS] tokens from available modalities, leading to a+ and a− for positive and negative samples, respectively. Subsequently, we enforced a triplet loss \({{{{\mathcal{L}}}}}_{triplet}\) for them to bring the samples of the same cancer closer than that of the negative sample:
where a+ and a− represent the farthest positive samples and nearest negative samples within a mini-batch, respectively, following the hard sample mining technique60. Here we used l2 distance for function d( ⋅ ) and ϵ is the margin which was set 0.3 based on smoother stability of loss degradation in the training set. Through these two pretraining objectives, as a result, we can get a well-trained slide aggregator that absorbed multimodal knowledge, which would be the ’Teacher’ for the patch extractor at the next stage.
Stage 2 - pretrain patch extractor
Upon finishing the first stage of pretraining, we can obtain a slide aggregator incorporating multimodal knowledge by being pretrained with multimodal data. In this stage, we leverage the pretrained slide aggregator as “Teacher” to seamlessly propagate multimodal knowledge into pathological patch extractor (ViT-L61), as shown in Fig. 2b, which is termed Self-Taught training. Specifically, for each WSI, we gathered their patch features \({{{{\mathcal{P}}}}}_{i}={\{{{{{\bf{p}}}}}_{i}^{m}\}}_{m=1}^{M}\) of the i-th WSI and fed them into the aggregator pretrained in the previous stage, where M refers to the number of patches of this WSI. Following the setting in the previous stage, M was fixed as 4096. In this way, every patch can be re-embedded into new features \({\hat{{{{\mathcal{P}}}}}}_{i}={\{{\hat{{{{\bf{p}}}}}}_{i}^{m}\}}_{m=1}^{M}\) incorporating multimodal knowledge. With these re-embedded features as the objective guidance, we can pretrain a patch extractor by enforcing the extracted patch feature to get as close as possible to the ones re-embedded by the well-trained aggregator. To achieve this, for each patch, we can query its corresponding re-embedded feature \({\hat{{{{\bf{p}}}}}}_{i}^{m}\) encoded by the aggregator and further tuned the extractor with a loss function that minimizes the discrepancy between patch features encoded by the patch extractor and the corresponding re-embedded features incorporating multimodal knowledge:
where f( ⋅ ) is a linear projection for adjusting the dimension of features and it transformed them into 512-dimensional features. Additionally, to avoid the catastrophic forgetting problem, a siamese structure is employed for the patch extractor consisting of two identical branches, where the parameters of one branch are updated using gradient descent, while the parameters of the other branch are updated using an Exponential Moving Average (EMA) of the parameters from the previous branch, without any gradient updates. Afterward, we enforced a similarity constraint between the patch features \({{{{\bf{p}}}}}_{i}^{m}\) extracted by the branch with gradient updates and those \({\overline{{{{\bf{p}}}}}}_{i}^{m}\) embedded by the branch with EMA updates. In the end, we combined two objectives into a loss function for pretraining the patch extractor:
where λ is a balancing coefficient and it was set 0.6 based on smoother stability of loss degradation. By doing this, the patch extractor was enhanced by multimodal knowledge at the whole-slide context.
Downstream tasks
Comparisons and baselines
To investigate the benefit of enhancing the patch extractor by incorporating multimodal knowledge at the slide level, we compared mSTAR against one general baseline and three SOTA pretrained extractors commonly used in the CPath community: (1) ResNet5022 pretrained on ImageNet-1K62, a commonly used baseline in many slide-level tasks9,13. (2) PLIP1, a vision-language (V-L) architecture (CLIP57) pretrained on OpenPath consisting of over 200k pathological patch-caption pairs. (3) CONCH3, a V-L CoCa63 framework with an additional generative loss pretrained on over 1.17 million pathological patch–caption pairs. (4) UNI2, a pure vision patch extractor pretrained on more than 100 million patches from over 100k WSIs, and (5) CHIEF14 as well as (6) GigaPath4, 2 slide-level vision-only pathology foundation models pretrained on 60,530 and 171,189 slides. Through pre-extracted patch features via these encoders, we can get 1024-dimensional (1024-d) embeddings for ResNet50, UNI, and mSTAR, 512-d embeddings for PLIP and CONCH, 768-d embeddings for CHIEF and 1536-d embeddings for GigaPath.
Models for downstream tasks
WSI classification and survival prediction
For slide-level prediction including classification and survival prediction, we follow the conventional two-stage MIL paradigm comprising pre-extraction of patch features as instances and the training of a MIL aggregator that integrates patch features (or instances) into a single slide-level (or bag) feature. The aggregator took all patch features of a WSI as an input and mapped them into a hidden embedding as a single slide-level representation. Subsequently, the slide-level representation was passed through a fully connected classifier head, resulting in logits. Lastly, based on logits, we performed two types of slide-level tasks including classification supervised by cross-entropy loss with slide labels, and survival prediction (an ordinal regression task) supervised by NLL loss64 with survival labels (event time in month), ranging from various diagnosis and prognosis tasks. Unless otherwise specified, we obtained slide-level predictions by training the widely used attention-based multiple-instance learning (ABMIL)11, a MIL aggregator that integrates all patch features of a WSI into the slide-level representation according to attention scores. For CHIEF and GigaPath, we fully follow the design in their original text that their pretrained patch extractors paired with the corresponding pretrained aggregator were employed. In particular, for patient-level tasks, such as survival prediction, we concatenate features of all slides belonging to a single patient as one case for the patient-level prediction.
We used the same hyper-parameters set for mSTAR and the competing FMs, in which the hidden dimensions are 512 and dropout keeps p = 0.25 after each intermediate layer in the network for regularization. We trained each model for 30 epochs on the training split by an Adam optimizer of the learning rate of 2 × 10−4 along with a cosine learning rate scheduler. The full set of hyperparameters is summarized in Supplementary Table 27.
Multimodal fusion
In the experiments of multimodal fusion, we employed 4 existing SOTA multimodal integration models, MCAT8, Porpoise43, CMTA10 and MOTCat9. It is worth highlighting that the training and evaluation of multimodal datasets held out from TCGA followed the same splits as that of vision-only models, and we simply discarded those cases without paired RNA-Seq data. For the aforementioned four existing multimodal integration models, we followed their default hyperparameters for these models, and detailed hyperparameters for each model are presented in Supplementary Table 28–30. For Porpoise, the input length of RNA-Seq varies across different cancer datasets in TCGA and the hidden dimension for RNA-Seq is fixed as 25, while the hidden dimension of pathological features was first transformed into 512 and then 256. Both modality branches adopted the dropout technique with p = 0.1. Lastly, features from two modalities were fused into a 256-dimensional slide-level feature. For MCAT and MOTCat, the hidden dimension of features was 256 for both modalities and dropout was 0.25 for regularization. Subsequently, features from two modalities were concatenated and integrated into a 256-dimensional slide-level representation. Similarly, CMTA followed the same hyperparameters except the hidden dimension of RNA-Seq which first became 1024 and then 256. For RNA-Seq data of MCAT, CMTA and MOTCat, embeddings were defined based on 6 functional categories according to 65 provided in MCAT by default, including 1) Tumor Supression, 2) Oncogenesis, 3) Protein Kinases, 4) Cellular Differentiation, 5) Transcription, and 6) Cytokines and Growth. More training hyperparameters are provided in Supplementary Table 31.
Zero-shot slide classification and retrieval
We considered the pretrained model as a good zero-shot learner, and employed non-parametric MI-Zero38 that does not rely on parametric training for these tasks, a well-established zero-shot approach for pathology slides. Given that the zero-shot’s capability heavily relies on the well-aligned modality spaces, we only compared against those approaches that are equipped with the text encoder by utilizing the pretrained text encoder as a good classification head, including PLIP and CONCH. The ensembling prompt of templates was used as the textual classification, which was utilized to compute the cosine similarity score with every patch feature. In the end, MI-Zero made the slide-level decision for every slide in the test set based on the majority voting of top-K scores.
Pathological report generation
To do this, we finetune the specific model of report generation, our prior work HistGen39. Given patients’ pathology features from WSIs of each FM, HistGen is able to produce a sequence of words. Specifically, given extracted pathological features from the foundation model, the encoder-decoder architecture of HistGen would encode them into the latent features for report decoding. Subsequently, these features are utilized by the text decoder to generate the report. The quality of the generated report is directly influenced by the quality of the pathological features encoded by each FM. For all optimization hyperparameters, refer to Supplementary Table 32.
Evaluation
We need to clarify that pathology reports were only used during pretraining (for multimodal alignment with WSIs), while all downstream tasks were evaluated using H&E diagnostic slides only as the input with no text data. This aligns with standard foundation model paradigms (e.g., CONCH3, where text aids pretraining but isn’t available during inference). Furthermore, the cases used for downstream evaluation were rigorously excluded from pretraining data. To systematically evaluate mSTAR’s capabilities, as shown in Fig. 1f, following the previous work14, we adopted four evaluation strategies as follows:
‘Held-out’ (out) represents the downstream dataset held out from pretraining data to avoid data contamination for evaluation. The training data included in pretraining data was used for training task-specific models, which were then used for inference on validation and test sets (i.e., held-out cohorts) that were held out from the pretraining data.
‘Independent’ (idpt) underscores that the source of dataset is independent from that of the pretraining data. For these datasets, we always either label-stratified these datasets into 7:1:2 train-validation-test folds or employed 5-fold cross-validation independently. Note that the difference between Held-out and Independent lies in whether the data comes from the same source as the pretraining data.
‘External’ (ext) is used for testing only and its data source is different from training data (from either held-out or independent cohorts) that was utilized to train task-specific models.
‘Zero-shot’ means that foundation models (e.g., mSTAR) are directly applied to make slide-level predictions without further training, rather than relying on additional task-specific models.
The details of all evaluation datasets are demonstrated in Supplementary Table 1.
Datasets
We present a description of each dataset used for evaluation, including 7 categories of oncological applications, covering 15 types of 97 practical clinical tasks. More details are summarized in Supplementary Table 1.
BRCA_PathSubtype16 for Pathological Subtyping (2 classes)
The BRCA_PathSubtype (Breast Invasive Carcinoma) dataset are sourced from TCGA including H&E diagnostic histopathology WSIs. This dataset encompassed cases of primary IDC (Invasive Ductal Carcinoma) and ILC (Invasive Lobular Carcinoma). After excluding slides with inadequate proportional tumor, a total of 985 slides were gathered, comprising 787 IDC and 198 ILC slides. Following the splits for pretraining, which approximately yielded 7:1:2 train-validation-test folds (656:95:234 slides), we ensure validation and test sets held out from pretraining sources.
GBMLGG_PathSubtype16 and EBrains_PathSubtype17 for Pathological Subtyping (3 classes)
The GBMLGG_PathSubtype (Glioblastoma and Brain Lower Grade Glioma) dataset comprises 1276 H&E diagnostic histopathology WSIs in total, consisting of three classes subtypes: Glioblastoma (GB) with 895 slides, Anaplastic Astrocytoma (AASTR) with 164 slides and Oligodendroglioma (ODG) with 217 slides. Following the splits for pretraining, which approximately yields 7:1:2 train-validation-test folds (839:200:237 slides), we ensure validation and test sets held out from pretraining materials. To evaluation models’ generalizable ability, we collected samples of the same subtypes as GBMLGG_PathSubtype from EBrains17 database, leading to 732 slides as an external cohort, EBrains_PathSubtype for pathological subtyping. It consists of 559 slides of Glioblastoma (GB), 89 slides of Anaplastic Astrocytoma (AASTR) and 84 slides of Oligodendroglioma (ODG).
HANCOCK_PathSubtype18 for pathological subtyping (3 classes)
HANCOCK_PathSubtype provides a dataset of head&neck tumors for pathological subtyping of 3 categories: SCC_Conventional-Keratinizing with 427 slides, SCC_Basaloid with 144 slides and SCC_Conventional-NonKeratinizing with 101 slides, resulting in 672 slides totally. We label-stratified the dataset into 7:1:2 train-validation-test splits, yielding 470:68:134 slides.
TCGA-NSCLC16 for pathological subtyping (2 classes)
The TCGA-NSCLC (Non-Small Cell Lung Cancer) dataset comprised NSCLC H&E diagnostic slides from TCGA, including cases of primary lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC). After tissue segmentation, a total of 1,053 slides were obtained, consisting of 541 LUAD and 512 LUSC slides. Similarly, we used the same pretraining splits train-validation-test of an approximate ratio 7:1:2 (664:100:289 slides) to avoid data contamination.
NFGC_Lauren and YN3_Lauren for Lauren subtyping of gastric cancer (3 classes)
Lauren subtyping is a common classification system for gastric cancer based on morphology, which typically divides tumors into Diffuse-type, Intestinal-type and Mixed-type that indicate different prognostic outcomes and treatment responses. We utilized the TCGA-STAD dataset as an internal cohort to train a model for Lauren classification. Since the data of TCGA-STAD has been used for pretraining, we collected one external gastric cancer cohort (NFGC_Lauren) of 388 slides from NanFang Hospital (NFH) and another external cohort of 319 slides from the Third Affiliated Hospital of Kunming Medical University in Yunnan (YN3_Lauren) for testing only. NFGC consists of 159 slides of Diffuse-type, 102 slides of Intestinal-type and 127 slides of Mixed-type. For YN3, there are 143 slides of Diffuse-type, 90 slides of Intestinal-type and 86 slides of Mixed-type.
NFGC_PathSubtype, YN1_PathSubtype and YN3_PathSubtype for pathological subtyping (3 classes)
With TCGA-STAD as an internal, we evaluate the ability of pathological subtyping for 3 crucial categories: Tubular Stomach Adenocarcinoma, Signet Ring Cell Carcinoma of the Stomach and Stomach Adenocarcinoma. For external validations, we collected 3 cohorts, NFGC_PathSubtype, YN1_PathSubtype and YN3_PathSubtype, from 3 medical centers including NFH, the First Affiliated Hospital of Kunming Medical University in Yunnan (YN1) and YN3, leading to 385, 254 and 315 slides for testing only. Specifically, NFGC_PathSubtype of NFH includes 166 slides of Tubular Stomach Adenocarcinoma, 163 slides of Signet Ring Cell Carcinoma of the Stomach and 66 slides of Stomach Adenocarcinoma. YN1_PathSubtype consists of 59 slides of Signet Ring Cell Carcinoma of the Stomach and 195 slides of Stomach Adenocarcinoma, while YN3_PathSubtype comprises 82 slides of Signet Ring Cell Carcinoma of the Stomach and 233 slides of Stomach Adenocarcinoma. Note that all data of external cohorts are used for testing only.
CAMELYON19,20 for breast metastasis detection (2 classes)
This dataset comprises 399 slides from the Cancer Metastases in Lymph Nodes Challenge 2016 (CAMELYON16)19 and 500 slides from the CAMELYON17 challenge20, resulting in 899 slides for the breast metastasis detection of two classes ("normal” v.s. “metastasis”). After removing a corrupted slide, we obtained a total of 898 WSIs (557 normal, 341 metastasis). For training and evaluation, we employed the label-stratified 7:1:2 train-validation-test splits (629:90:179 slides).
NF_Metastatic, NF_Metastatic_Fine, QFS_Metastatic and QFS_Metastatic_fine for lung metastasis detection (2 classes and 6 classes)
Lung Metastasis Detection includes two tasks: metastasis detection and its primary site prediction, denoted by ‘Metastatic’ and ‘Metastatic_Fine’. We curated NF_Metastatic dataset (1,198 slides, 705 cases) from NFH, in which ‘Metastatic’ aims to identify if the tumor is metastatic (314 cases) or primary (391 cases). Another dataset NF_Metastatic_Fine (705 cases) is also established from NFH, in which ‘Metastatic_Fine’ is performed to predict the primary site of metastatic cancer. The primary sites include six distinct classes: LUAD (391 cases), breast (55 cases), colon (186 cases), kidney (25 cases), liver (34 cases), and carcinoma of unknown primary (CUP, 14 cases). Both two datasets are label-stratified into 7:1:2 train-validation-test splits (493:70:142 cases). For external cohorts, we incorporated 530 WSIs (430 cases) from Shandong Provincial Qianfoshan Hospital (QFS), leading to QFS_Metastatic and QFS_Metastatic_Fine cohorts for testing only. QFS_Metastatic dataset included 237 primary cases and 193 metastatic cases, while QFS_Metastatic_Fine comprised 237 LUAD cases, 50 breast cases, 96 colon cases, 30 kidney cases, 10 liver cases, and 7 CUP cases.
NFGC_Perineural and YN3_Perineural for perineural invasion detection in gastric cancer
The morphological presence of Perineural Invasion (PNI) often indicates a more aggressive tumor and poorer survival rates. As such, it is crucial for prognostic evaluation, treatment decisions, and assessing recurrence risk to detect PNI. To this end, NFGC_Perineural dataset (396 cases) was collected NFH, consisting of 255 positive and 141 negative cases. As an internal cohort for training and evaluation, the data was divided into training, validation, and test sets in a ratio of 7:1:2 (277:39:80 cases). Furthermore, an additional cohort of 319 cases (112 positive and 207 negative), YN3_Perineural, was obtained from YN3 for external validation.
NFGC_Vascular and YN3_Vascular for vascular invasion detection in gastric cancer
Vascular Invasion in gastric cancer indicates the presence of tumor cells in blood vessels and is linked to poorer prognosis, higher metastasis risk, and increased recurrence rates. To identify it, we used a dataset consisting of 395 cases from NFH, known as the NFGC_Vascular dataset. This dataset comprises 197 positive cases and 198 negative cases. For model training and evaluation, the data was divided into training, validation, and test sets in a 7:1:2 ratio. Furthermore, we included an external validation set of 319 cases from YN3, which contains 122 positive and 197 negative cases.
PANDA21 for prostate ISUP grading (6 classes)
Derived from the PANDA challenge21, the ISUP (International Society of Urological Pathology) grading task includes a collection of 10,616 prostate cancer core needle biopsies for prostate cancer evaluation of 6 grades (also known as “classes”). After tissue segmentation, slides with a low tumor proportion were excluded, which resulted in 10,202 slides. For training and evaluation, we label-stratified PANDA into 7:1:2 train–validation–test folds (7,143:1,019:2,040 slides).
HANCOCK-TStage18 for pathological T-Staging (4 classes)
In clinical practice, pathologists will divide patients into different stages according to the severity, which can guide treatment decisions and assess the likelihood of metastasis. To assess this task, we utilize HANCOCK-TStage dataset consisting of 705 patients and divided it into 7:1:2 train–validation–test folds (496:67:128 cases) for validation. To be specific, the dataset includes 259 T1, 256 T2, 123 T3 and 67 T4 cases.
18 TCGA datasets for mutation prediction
We used the public TCGA data from the studies held out from pretraining materials to evaluate the performance of gene-level mutation prediction. For every study, we involve high-frequent mutated genes and FDA-approved drug-related genes, leading to 17 datasets across 9 held-out studies. The positive rates are presented in Supplementary Table 12. Additionally, the prediction of tumor mutation burden (TMB), a predictive biomarker in solid tumors that is especially important for immunotherapy, was also evaluated in the TCGA-NSCLC study.
5 CPTAC66 datasets for mutation prediction
With internal cohorts from TCGA, we utilized the data from CPTAC database for external validation on the ability of mutation prediction. The datasets with over 100 cases and mutation rate of at least 30%, and overlap with internal datasets are included, resulting in BRCA_PIK3CA (116 cases), BRCA_TP53 (116 cases), BRCA_TTN (120 cases), LUAD_KRAS (175 cases) and LUAD_EGFR (175 cases). The positive rates are presented in Supplementary Table 12.
10 IHC biomarker datasets
Immunohistochemistry (IHC) typically serves as the biomarker to assess tumor types and differentiation, guide the choice of targeted and immunotherapies, and monitor recurrence in clinical practice. We collected three IHC biomarkers tasks from TCGA: estrogen receptor (ER) with 949 cases, progesterone receptor (PR) with 948 cases, and human epidermal growth factor receptor (HER2) with 646 cases. For training and evaluation, these datasets are divided into train-val-test splits following the pretraining splits to avoid data contamination. With these datasets as internal cohorts, we curated the corresponding tasks from The First Affiliated Hospital of Zhejiang University School of Medicine (ZJ1), leading to IHC_ZJ1_ER (1548 cases), IHC_ZJ1_HER2 (1344 cases) and IHC_ZJ1_PR (1556 cases) for testing only. As doctors in clinical settings typically annotate the fine-grained labels for ER and HER2, we further assessed their expression levels, resulting in two datasets: IHC_ZJ1_ER_Level and IHC_ZJ1_HER2_Level with 7:1:2 splits (1083:154:311 cases for ER and 940:134:270 cases for HER2) for training and evaluation. Furthermore, we also evaluate other biomarkers commonly seen in clinical practice: Cytokeratin 5 (CK5, 961 cases) of breast cancer and Cytokeratin 7 (CK7, 419 cases) of lung cancer. For training and evaluation, we divided them with 7:1:2 into train-val-test splits (672:96:193 cases for CK5 and 293:42:84 cases for CK7). The label distribution of these biomarkers can be found in Supplementary Table 13 and 14.
BRCA_MolSubtype16 and ZJ1_Breast_MolSubtype for molecular subtyping (4 classes)
BRCA_MolSubtype is derived from TCGA, consisting of Triple-Negative Breast Cancer (TNBC) (94 cases), HER2 (56 cases), LumA (228 cases) and LumB (127 cases) classes. For training and evaluation, we label-stratified the dataset into train–validation-test cohorts (323:53:129 cases). For external validation, an external cohort was established with 2,045 cases (585 TNBC, 292 HER2, 307 LumA and 861 LumB).
GBMLGG_MolSubtype16 and EBrains_MolSubtype17 for molecular subtyping (2 classes)
GBMLGG_MolSubtype is derived from TCGA for identifying IDH status, consisting of Positive (362 cases) and Negative (190 cases) classes. For training and evaluation, we label-stratified the dataset into train–validation-test cohorts (401:64:87 cases). For external validation, an external cohort was established with 428 cases (361 Positive and 67 Negative).
TCGA_HNSC_HPV16 and HANCOCK_HPV18 for molecular subtyping (2 classes)
HPV-p16 status is a significant prognostic biomarker regrading different outcomes. To predict HPV-p16 status, we leveraged TCGA_HNSC_HPV (405 cases) derived from TCGA as an internal cohort for identifying HPV status, consisting of Positive (41 cases) and Negative (364 cases) classes. For training and evaluation, we label-stratified the dataset into train–validation-test cohorts (284:39:118 cases). For external validation, an external cohort was established with 332 cases (191 Positive and 141 Negative).
CRC_MolSubtype16 for molecular prediction (4 classes)
The dataset (492 cases) used in this study is derived from the TCGA CRC (Colon Adenocarcinoma and Rectum Adenocarcinoma) dataset, which includes the Colon Adenocarcinoma (COAD) and Rectum Adenocarcinoma (READ) datasets. It comprises four consensus molecular subtypes (CMSs): 74 CMS1, 211 CMS2, 68 CMS3 and 139 CMS4. To facilitate training and evaluation, we stratified the dataset based on labels into train-validation-test cohorts with proportions of 325:49:118 cases, respectively.
10 TCGA cohorts, 4 external cohorts and 2 independent cohorts for survival prediction
"In pretraining splits, we employed case- and label-stratified 7:1:2 training-validation-test splits for 9 TCGA cancer datasets of over 400 cases. We evaluated the capability of survival analysis on the same validation and test sets totally excluded from pretraining data. More information about the 9 TCGA cancer datasets were provided in Supplementary Table 32. We first evaluated 9 Overall Survival (OS) tasks of TCGA across 9 cancer types. With OS_HNSC as the internal cohort, we further collected OS_HANCOCK (747 cases) as an external cohort. Given OS_BRCA as the internal cohort, we curated OS_ZJ1 (454 cases) from ZJ1 of breast cancer for external validation. Additionally, we further evaluate the DFS task of breast cancer on TCGA, resulting in DFS_BRCA (878 cases), serving as the internal cohort with the splits (619:84:175 cases). For external validation, DFS_ZJ1 (454 cases) was curated as an external cohort. Although TCGA-STAD was not held out from pretraining data, we utilized it as a training set to train a model and evaluate it on the curated DFS_YN1 (260 cases) sourced from YN1 for external validation. For independent cohorts, we curated OS_NFCRC (294 cases) of colon cancer from NFH for OS prediction using 5-fold cross-validation, and meanwhile we utilized RFS_HANCOCK (747 cases) for Recurrence-Free Survival (RFS) prediction based on 5-fold cross-validation as well.
9 Survival prediction datasets for multimodal fusion
We collected RNA-Seq data from cBioPortal for 9 TCGA held-out studies to evaluate the performance of multimodal fusion. We followed the same splits as those used in unimodal survival prediction tasks, and excluded those without the paired RNA-Seq data. The final splits of the train-val-test are presented in Supplementary Table 1.
UBC-OCEAN67,68 for ovarian cancer subtyping (5 classes)
The UBC-OCEAN (University of British Columbia - Ovarian Cancer subtypE clAssification and outlier detectioN) dataset consists of 538 slides, which aims to classify ovarian cancer subtypes into 5 categories. After performing tissue segmentation, a total of 527 slides were acquired (98 CC, 122 EC, 221 HGSC, 43 LGSC and 43 MC). The class information is presented in Supplementary Table 36.
BCNB datasets69 for ER (2 classes), PR (2 classes) and HER2 prediciton (2 classes) of biopsy slides
The Early Breast Cancer Core-Needle Biopsy (BCNB) WSI dataset, encompasses core-needle biopsy WSIs obtained from patients diagnosed with early breast cancer. We collected 1038 WSIs paired with ER, PR, HER2 status after tissue extraction.
Pancancer TCGA16 and breast&lung datasets for zero-shot slide retrieval
We utilized pan-cancer TCGA datasets (934 cases) held out from pretraining data to evaluate the performance of Zero-shot Slide Retrieval for Slide-to-Report (Image2Text) and Report-to-Slide (Text2Image) retrieval. Furthermore, to evaluate the generalizability of zero-shot slide retrieval, we curated another cohort consisting of breast and lung cancers from ZJ1 and NFH, resulting in Breast&Lung (500 cases) to ensure a sufficiently large search space. Given that original reports of ZJ1 and NFH are in Chinese, we first translated them into English via GPT-4o-mini before performing retrieval.
TCGA dataset, nanfang and ZJ-first for pathological report generation
During pretraining, we employed training-validation-test splits for some cancer datasets of over 400 cases and other data were put into pretraining materials. Following this setting, we considered all pretraining data containing pathology reports as the training set, and the held-out validation-test sets were re-used, resulting in 7073:452:934 cases for train-validation-test splits. Given TCGA dataset as the internal cohort, we additionally collected two external cohorts: Nanfang (250 cases) and ZJ-First (250 cases) from NFH and ZJ1 of lung cancer and breast cancer, respectively. Similarly, considering original reports of ZJ1 and NFH are in Chinese, they are translated into English via GPT-4o-mini before performing report generation.
Evaluation metrics
For classification tasks, Macro-AUC and its 95% confidence interval (CI) are reported considering alleviating the impact of unbalanced data, which doesn’t depend on the selection of the decision threshold and is not affected by the sample ratio of classes. For survival prediction tasks, we report the commonly used Concordance Index (C-Index) and its 95% CI, which is defined as the probability that two randomly selected individuals will have risk predictions correctly ordered. For zero-shot slide retrieval, we reported Recall @5, @10 and @50. In pathological report generation, in line with our prior studies HistGen39, we report various metrics, BLEU@K70, METEOR71 and ROUGE-L72, to assess the accuracy of predicted captions against the ground-truth captions from different perspectives. BLEU@K measures the similarity between machine-generated text and ground truth by comparing the presence and frequency of n-grams. METEOR is a metric that evaluates precision and recall by matching unigrams while also factoring in synonyms and word variations between the original text and the reference. On the other hand, ROUGE-L measures the similarity in n-gram overlap between the generated texts and the ground truth.
Statistical analysis
Unless otherwise specified, we employ non-parametric bootstrapping with 1000 bootstrap73 replicates to estimate 95% confidence intervals (CI) for all experiments. For each evaluation experiment, the model performing best in the validation split was chosen to be evaluated on test sets or external sets. To assess the observed differences in performance between the two models, we utilize a one-sided Wilcoxon signed-rank test74 for statistical significance, following the previous work4.
Computing software and hardware
We conducted all experiments and analyses in this study using Python (v3.11.5) and PyTorch (v2.2.1, CUDA 11.7) (https://pytorch.org) unless stated otherwise, and these can be reproduced with open-source libraries as described below. To pretrain aggregator, the implementation of the text encoder pretrained on PubMed was maintained by the codebase (https://github.com/dmis-lab/biobert) and its pretrained weights can be assessed in the open-source timm library from Hugging Face (https://huggingface.co). For extractor pretraining, we initialize the backbone with the pretrained weights of UNI codebase (https://github.com/mahmoodlab/uni). OpenSlide (v3.4.1) and openslide-python (v1.3.1) were utilized to support the processing of WSIs in conjunction with CLAM. Implementations of other visual pretrained encoders compared in the study can be accessed through the following links: ResNet-50 pretrained on ImageNet-1K (https://github.com/mahmoodlab/CLAM), PLIP, CONCH, CHIEF and GigaPath (https://github.com/prov-gigapath/prov-gigapath). Implementations of zero-shot learning for WSIs were provided in MI-Zero (https://github.com/mahmoodlab/MI-Zero). For training MIL models for downstream tasks, we adapted the code of ABMIL from the CLAM codebase (https://github.com/mahmoodlab/CLAM). For multimodal survival prediction, we used the off-the-shelf multimodal fusion models: MCAT, Porpoise (https://github.com/mahmoodlab/PORPOISE), MOTCat (https://github.com/Innse/MOTCat) and CMTA. For pathological report generation, HistGen (https://github.com/dddavid4real/HistGen) is applied. We used 4 × 80 GB NVIDIA H800 GPUs (graphics processing unit) for pretraining aggregator and a single 80 GB NVIDIA H800 GPU for pretraining extractor. These GPUs were set up for multi-GPU, multi-node training, employing distributed data-parallel (DDP) techniques. All other experiments for downstream tasks were conducted on single 24 GB NVIDIA 3090 GPUs or single 80 GB H800 GPU.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data availability. This study incorporates a total of 97 oncological tasks for downstream evaluation, in which 69 tasks are evaluated on public datasets and 28 tasks are assessed on private cohorts. Pretraining data consisted of whole-slide images (WSIs) and pathology reports from TCGA (https://portal.gdc.cancer.gov/), and RNA-seq expression profiles from cBioPortal (https://www.cbioportal.org/). Downstream evaluations used publicly available datasets including a subset of TCGA (https://portal.gdc.cancer.gov/), BCNB, CAMELYON16, CAMELYON17, HANCOCK, PANDA (https://www.kaggle.com/c/prostate-cancer-grade-assessment/data), UBC-OCEAN. Regarding the data from Nanfang Hospital of Southern Medical University (NFH), Shandong Provincial Qianfoshan Hospital (QFS), The First Affiliated Hospital of Kunming Medical University in Yunnan (YN1), The Third Affiliated Hospital of Kunming Medical University in Yunnan (YN3), and The First Affiliated Hospital of Zhejiang University School of Medicine (ZJ1), these datasets are not publicly available due to patient privacy obligations, institutional review board requirements, and data use agreements. However, researchers interested in accessing deidentified data may submit a request directly to the corresponding author, subject to obtaining the necessary ethical approvals and complying with institutional policies. The details of these datasets are demonstrated in Supplementary Table 40. Source data for each figure or table are provided with this paper. Source data are provided with this paper.
Code availability
The code and weights of mSTAR have been made available on GitHub (https://github.com/Innse/mSTAR)75. Trained weights are available at Hugging Face (https://huggingface.co/Wangyh/mSTAR). For reproducibility, we archived the exact version used in this study on Zenodo with the DOI [10.5281/zenodo.17273573] and tagged release [v1.0.0].
References
Huang, Z., Bianchi, F., Yuksekgonul, M., Montine, T. J. & Zou, J. A visual–language foundation model for pathology image analysis using medical twitter. Nat. Med. 29, 2307–2316 (2023).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. 30, 850–862 (2024).
Lu, M. Y. et al. A visual-language foundation model for computational pathology. Nat. Med. 30, 863–874 (2024).
Xu, H. et al. A whole-slide foundation model for digital pathology from real-world data. Nature 630, 181–188 (2024).
Alfasly, S. et al. When is a foundation model a foundation model. arXiv preprint arXiv:2309.11510 (2023).
Team, G. et al. Gemini: a family of highly capable multimodal models. arXiv preprint arXiv:2312.11805 (2023).
Khattak, M. U., Rasheed, H., Maaz, M., Khan, S. & Shahbaz Khan, F. Maple: Multi-modal prompt learning. Proceedings of the IEEE/CVF conference on computer vision and pattern recognition. 19113–19122 (2023).
Chen, R. J. et al. Multimodal co-attention transformer for survival prediction in gigapixel whole slide images. Proceedings of the IEEE/CVF international conference on computer vision. 4015–4025 (2021).
Xu, Y. & Chen, H. Multimodal optimal transport-based co-attention transformer with global structure consistency for survival prediction. Proceedings of the IEEE/CVF international conference on computer vision. 21241–21251 (2023).
Zhou, F. & Chen, H. Cross-modal translation and alignment for survival analysis. Proceedings of the IEEE/CVF International Conference on Computer Vision. 21485–21494 (2023).
Ilse, M., Tomczak, J. & Welling, M. Attention-based deep multiple instance learning. International conference on machine learning. PMLR. 2127–2136 (2018).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 555–570 (2021).
Shao, Z. et al. Transmil: transformer based correlated multiple instance learning for whole slide image classification. Adv. neural Inf. Process. Syst. 34, 2136–2147 (2021).
Wang, X. et al. A pathology foundation model for cancer diagnosis and prognosis prediction. Nature 634, 970–978 (2024).
Jaume, G. et al. Transcriptomics-guided Slide Representation Learning In Computational Pathology. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 9632–9644 (2024).
Weinstein, J. N. et al. The cancer genome atlas pan-cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Roetzer-Pejrimovsky, T. et al. The digital brain tumour atlas, an open histopathology resource. Sci. Data 9, 55 (2022).
Dörrich, M. et al. A multimodal dataset for precision oncology in head and neck cancer. medRxiv2024-2005 (2024).
Bejnordi, B. E. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. Jama 318, 2199–2210 (2017).
Bandi, P. et al. From detection of individual metastases to classification of lymph node status at the patient level: the camelyon17 challenge. IEEE Trans. Med. imaging 38, 550–560 (2018).
Bulten, W. et al. Artificial intelligence for diagnosis and Gleason grading of prostate cancer: the panda challenge. Nat. Med. 28, 154–163 (2022).
He, K., Zhang, X., Ren, S. & Sun, J. Identity mappings in deep residual networks. In Computer Vision–ECCV 2016: 14th European Conference, Amsterdam, The Netherlands, October 11–14, 2016, Proceedings, Part IV 14, 630–645 (Springer, 2016).
Slamon, D. et al. Adjuvant trastuzumab in HER2-positive breast cancer. N. Engl. J. Med. 365, 1273–1283 (2011).
Chakravarty, D. et al. Oncokb: a precision oncology knowledge base. JCO Precis. Oncol. 1, 1–16 (2017).
Mendiratta, G. et al. Cancer gene mutation frequencies for the us population. Nat. Commun. 12, 5961 (2021).
Wilson, M. R. et al. Arid1a and PI3-kinase pathway mutations in the endometrium drive epithelial transdifferentiation and collective invasion. Nat. Commun. 10, 3554 (2019).
Boiarsky, D. et al. Molecular markers of metastatic disease in Kras-mutant lung adenocarcinoma. Ann. Oncol. 34, 589–604 (2023).
Romero, R. et al. Keap1 loss promotes Kras-driven lung cancer and results in dependence on glutaminolysis. Nat. Med. 23, 1362–1368 (2017).
Zhu, Z. et al. Gata3 mediates doxorubicin resistance by inhibiting cyb5r2-catalyzed iron reduction in breast cancer cells. Drug Resistance Updates 69, 100974 (2023).
Pascual, J. & Turner, N. Targeting the PI3-kinase pathway in triple-negative breast cancer. Ann. Oncol. 30, 1051–1060 (2019).
Lin, J. J. et al. Five-year survival in EGFR-mutant metastatic lung adenocarcinoma treated with EGFR-TKIs. J. Thorac. Oncol. 11, 556–565 (2016).
Marks, J. L. et al. Prognostic and therapeutic implications of EGFR and Kras mutations in resected lung adenocarcinoma. J. Thorac. Oncol. 3, 111–116 (2008).
Patocs, A. et al. Breast-cancer stromal cells with tp53 mutations and nodal metastases. N. Engl. J. Med. 357, 2543–2551 (2007).
Pollack, I. F. et al. Age and TP53 mutation frequency in childhood malignant gliomas: results in a multi-institutional cohort. Cancer Res. 61, 7404–7407 (2001).
Bunda, S. et al. Cic protein instability contributes to tumorigenesis in glioblastoma. Nat. Commun. 10, 661 (2019).
Tashiro, H. et al. Mutations in pten are frequent in endometrial carcinoma but rare in other common gynecological malignancies. Cancer Res. 57, 3935–3940 (1997).
Lacey Jr, J. V. et al. Pten expression in endometrial biopsies as a marker of progression to endometrial carcinoma. Cancer Res. 68, 6014–6020 (2008).
Lu, M. Y. et al. Visual language pretrained multiple instance zero-shot transfer for histopathology images. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 19764-19775 (2023).
Guo, Z. et al. Histgen: Histopathology report generation via local-global feature encoding and cross-modal context interaction. International Conference on Medical Image Computing and Computer-Assisted Intervention. Cham: Springer Nature Switzerland. 189–199 (2024).
Zhang, Y., Xu, Y., Chen, J., Xie, F. & Chen, H. Prototypical information bottlenecking and disentangling for multimodal cancer survival prediction. In The Twelfth International Conference on Learning Representations. 2024.
Mantel, N. et al. Evaluation of survival data and two new rank order statistics arising in its consideration. Cancer Chemother. Rep. 50, 163–170 (1966).
Yu, Y. et al. 2mo multimodal data fusion enhanced precision neoadjuvant chemotherapy in breast cancer with a multi-task transformer-cnn-mixed learning. Ann. Oncol. 34, S1468–S1469 (2023).
Chen, R. J. et al. Pan-cancer integrative histology-genomic analysis via multimodal deep learning. Cancer Cell 40, 865–878 (2022).
Cuny, M. et al. Relating genotype and phenotype in breast cancer: an analysis of the prognostic significance of amplification at eight different genes or loci and of p53 mutations. Cancer Res. 60, 1077–1083 (2000).
Vorontsov, E. et al. A foundation model for clinical-grade computational pathology and rare cancers detection. Nat. Med. 30, 2924–2935 (2024).
Vorontsov, E. et al. A foundation model for clinical-grade computational pathology and rare cancers detection. Nature medicine. 30, 2924–2935 (2024).
Hong, M. et al. Rna sequencing: new technologies and applications in cancer research. J. Hematol. Oncol. 13, 1–16 (2020).
Beer, D. G. et al. Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nat. Med. 8, 816–824 (2002).
Gu, A. & Dao, T. Mamba: Linear-time sequence modeling with selective state spaces. First conference on language modeling. (2024).
Ding, J. et al. Longnet: Scaling transformers to 1,000,000,000 tokens. arXiv preprint arXiv:2307.02486 (2023).
Yang, S., Wang, Y. & Chen, H. MambaMIL: Enhancing Long Sequence Modeling with Sequence Reordering in Computational Pathology. In Medical Image Computing and Computer Assisted Intervention – MICCAI 2024. MICCAI 2024. Lecture Notes in Computer Science, (eds Linguraru, M. G. et al.) vol 15004. (Springer, Cham, 2024).
Li, B. & Dewey, C. N. Rsem: accurate transcript quantification from rna-seq data with or without a reference genome. BMC Bioinforma. 12, 1–16 (2011).
Sarkar, A. & Stephens, M. Separating measurement and expression models clarifies confusion in single-cell RNA sequencing analysis. Nat. Genet. 53, 770–777 (2021).
Haque, A., Engel, J., Teichmann, S. A. & Lönnberg, T. A practical guide to single-cell RNA-sequencing for biomedical research and clinical applications. Genome Med. 9, 1–12 (2017).
Yang, F. et al. scBERT as a large-scale pretrained deep language model for cell type annotation of single-cell RNA-seq data. Nat. Mach. Intell. 4, 852–866 (2022).
Du, J. et al. Gene2vec: distributed representation of genes based on co-expression. BMC Genom. 20, 7–15 (2019).
Radford, A. et al. Learning transferable visual models from natural language supervision. In International Conference On Machine Learning, 8748–8763 (PMLR, 2021).
Lee, J. et al. Biobert: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 36, 1234–1240 (2020).
Cui, H. et al. scgpt: toward building a foundation model for single-cell multi-omics using generative AI. Nat. Methods 21, 1–11 (2024).
Hermans, A., Beyer, L. & Leibe, B. In defense of the triplet loss for person re-identification. arXiv preprint arXiv:1703.07737 (2017).
Dosovitskiy, A. et al. An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. In International Conference on Learning Representations. (2021).
Deng, J. et al. Imagenet: a large-scale hierarchical image database. In 2009 IEEE conference on computer vision and pattern recognition, 248–255 (Ieee, 2009).
Yu, J. et al. CoCa: Contrastive Captioners are Image-Text Foundation Models. Transactions on Machine Learning Research. (2022).
Zadeh, S. G. & Schmid, M. Bias in cross-entropy-based training of deep survival networks. IEEE Trans. Pattern Anal. Mach. Intell. 43, 3126–3137 (2020).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Edwards, N. J. et al. The CPTAC data portal: a resource for cancer proteomics research. J. Proteome Res. 14, 2707–2713 (2015).
Asadi-Aghbolaghi, M. et al. Machine learning-driven histotype diagnosis of ovarian carcinoma: Insights from the Ocean AI Challenge. Preprint at medRxiv https://doi.org/10.1101/2024.04.19.24306099 (2024).
Farahani, H. et al. Deep learning-based histotype diagnosis of ovarian carcinoma whole-slide pathology images. Mod. Pathol. 35, 1983–1990 (2022).
Xu, F. et al. Predicting axillary lymph node metastasis in early breast cancer using deep learning on primary tumor biopsy slides. Front. Oncol. 11, 4133 (2021).
Kishore, P., Roukos, S., Ward, T. & Zhu, W.-J. Bleu: a method for automatic evaluation of machine translation. In Proceedings of the 40th annual meeting of the Association for Computational Linguistics. pp. 311–318 (2002).
Michael, D. & Lavie, A. Meteor 1.3: Automatic metric for reliable optimization and evaluation of machine translation systems. In Proceedings of the sixth workshop on statistical machine translation, pp. 85–91 (2011).
Lin, C.-Y. Rouge: A package for automatic evaluation of summaries. In Text summarization branches out, pp. 74–81 (2004).
Bradley, E. & Tibshirani, R. J. An introduction to the bootstrap. (Chapman and Hall/CRC, 1994).
Wilcoxon, F. Individual comparisons by ranking methods. In Breakthroughs in statistics: Methodology and distribution (pp. 196-202). (Springer, New York, 1992).
Xu, Y. et al. A multimodal knowledge-enhanced whole-slide pathology foundation model. https://doi.org/10.5281/zenodo.17273573 (2025).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 62202403), Innovation and Technology Commission (Project No. PRP/034/22FX and ITCPD/17-9), Research Grants Council of the Hong Kong Special Administrative Region, China (No. R6003-22 and C4024-22GF) and HKUST Frontier Technology Research for Joint Institutes with Industry (No. OKT24EG01).
Author information
Authors and Affiliations
Contributions
Y.X., Y.W. and H.C. conceived the study. Y.X. designed the pretraining approach and experiments, organized the data for downstream evaluation, and prepared the manuscript. Y.W. performed foundation model pretraining and conducted evaluation on downstream tasks, including molecular prediction, survival prediction, zero-shot prediction and report generation. F.Z. conducted an evaluation of pathological diagnosis, molecular prediction and multimodal fusion. J.M., C.J. and H.Z. collected the data and assisted in results analysis and coding. S.Y. participated in coding for pretraining. J.L. provided insightful interpretation and specialized clinical validation. Z.Z., C.Z., Z.L., L.L. and X.Z. provided preprocessed data for downstream tasks. H.L. assisted in the experimental design. X.W., A.H. and R.C.K.C. assisted in the design of the evaluation on clinical tasks and the interpretation of experimental results. J.W. assisted in curating RNA-Seq data and offered suggestions for experimental designs. H.C. supervised the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, Y., Wang, Y., Zhou, F. et al. A multimodal knowledge-enhanced whole-slide pathology foundation model. Nat Commun 16, 11406 (2025). https://doi.org/10.1038/s41467-025-66220-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66220-x
This article is cited by
-
Leveraging multi-modal foundation models for analysing spatial multi-omic and histopathology data
Nature Biomedical Engineering (2026)
-
Multiple Instance Learning Framework with Masked Hard Instance Mining for Gigapixel Histopathology Image Analysis
International Journal of Computer Vision (2026)
-
Cross-staining pathological diagnosis based on spatially enriched multiple instance learning with clinical embedding
Applied Intelligence (2026)
-
Artificial intelligence in digital pathology diagnosis and analysis: technologies, challenges, and future prospects
Military Medical Research (2026)
-
The role of artificial intelligence-based foundation models and “copilots” in cancer pathology: potential and challenges
Journal of Experimental & Clinical Cancer Research (2025)