Abstract
Cancer stem cells (CSCs) play a critical role in tumor initiation, progression, and recurrence. How liver CSCs initiate their self-renewal remains elusive. Here we identify a conserved small nucleolar RNA (snoRNA), SNORA49, which is lowly expressed in liver CSCs, as a negative regulator of CSC self-renewal. SNORA49 knockout enhances the self-renewal capacity of liver CSCs and accelerates hepatocellular carcinoma (HCC) tumorigenesis, whereas overexpression of SNORA49 suppresses tumor formation. Mechanistically, in non-CSCs, SNORA49 is specifically localized in the nucleoplasm to associate with HNRNPU, blocking its interaction with ZC3H18, resulting in inhibition of SOX9 transcription. In liver CSCs, lowly expressed SNORA49 releases HNRNPU to engage with ZC3H18 and enrich on the promoter of SOX9, leading to its transcription. Of note, lipid nanoparticle (LNP)-mediated delivery of SNORA49 RNAs and antisense oligonucleotides (ASOs) targeting SOX9 exerts potent synergistic anti-tumor effect on HCC tumors. Our findings define SNORA49 as a tumor suppressor in liver CSCs, and restoring SNORA49 levels and silencing SOX9 with LNP-delivered system may provide therapeutic strategy for clinical intervention to HCC patients.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is the sixth most common and third most deadly cancer worldwide1. Although various clinical treatments are available for HCC, including surgical resection, radiotherapy, chemotherapy, and target therapy, their overall efficacy remains limited2. The major challenge lies in the heterogeneity and high recurrence of HCC. Cancer stem cells (CSCs), a subpopulation of tumor cells that possess the ability to self-renew and differentiate, have been considered as the root cause of tumor initiation, progression, and metastasis, leading to drug resistance, treatment failure, and tumor recurrence2. Over the past two decades, extensive research has been dedicated to identifying and characterizing liver CSCs using surface markers such as CD133, CD13, CD47, EpCAM, and Lgr53,4,5. Based on these findings, specific therapies targeting liver CSCs have shown significant potential in animal models and preclinical trials, with even greater efficacy when combined with chemotherapy and immunotherapy. For instance, blocking CD47 with an anti-CD47 antibody sensitize HCC cells to sorafenib, promoting cell phagocytosis and inhibiting tumor progression6. In addition, treatment with ANXA3-neutralizing antibodies reduces the expression of CD133, CD24, and EpCAM in liver CSCs, enhancing the effectiveness of sorafenib and regorafenib in HCC7,8. These studies suggested that therapeutic strategies targeting and eliminating CSCs hold great promise for the future of cancer treatment. However, to find effective biomarkers, develop more effective CSC-based therapies, and understand the biological characteristics of CSCs are urgently needed.
Similar to embryonic stem cells, CSCs rely on various stemness signaling pathways and stemness factors to maintain their self-renewal capacity9. Among these factors, the Sex determining region Y-box 9 (SOX9) plays a critical role in carcinogenesis of various malignancies, including HCC, breast cancer, gastric cancer, and lung cancer10,11. For example, in breast cancer, expression of SOX9 endows tumor cells with stem cell-like properties and enhances their tumor-initiating and metastasis-seeding abilities12. In gastric cancer, elevated SOX9 levels induce intestinal metaplasia, a pre-malignant stage in gastric carcinogenesis13, or increase β-catenin levels to accelerate tumor cell proliferation14. In HCC, SOX9 promotes the expansion of liver CSCs via tilting asymmetric division toward symmetric division by inhibiting the Notch antagonist Numb15. In addition, SOX9 functions downstream of YAP and exerts a decisive influence on HCC tumor plasticity in the context of YAP activation16. These findings indicate that SOX9 plays a pivotal role in maintaining liver CSC self-renewal, which may interact with other stemness-related pathways, such as Notch and Hippo signaling, to exert its functions. Nevertheless, the precise mechanism that regulates SOX9 expression in liver CSCs still remains elusive.
Small nucleolar RNAs (snoRNAs) are a type of non-coding RNAs with a length of 60-300 nt, which are originally discovered in the nucleolus17. They are categorized into two major classes: H/ACA box snoRNAs (snoRA) and C/D box snoRNAs (snoRD). Canonically, snoRNAs are known to guide 2’-O-methylation or pseudouridylation of ribosomal RNAs (rRNAs) through complementary base pairing. Confined by their nucleolus-only localization and limited functions, snoRNAs have not initially attracted much attention in cancer research. However, with the advent of whole-genome deep sequencing technologies, emerging studies have identified a variety of novel snoRNAs that are widely expressed in different cancer types. They have localizations beyond the nucleolus and functions beyond rRNA modifications. For example, SNORD50A and SNORD50B are specifically localized in the cytoplasm, the loss of which results in hyperactivation of Ras-ERK signaling in melanoma, ovarian cancer, and lung cancer18. In addition, SNORD27 binds to E2F7 pre-mRNAs in the nucleus, regulating the alternative splicing of E2F7, a transcriptional repressor of cell-cycle-regulated genes, which is often dysregulated in many malignancies19. In HCC, upregulation of SNORD17 promotes cell growth and tumorigenesis by inhibiting p53-mediated cell cycle arrest and apoptosis20. We recently showed that SNORD88B is highly expressed in liver CSCs. Elevated SNORD88B anchors WRN in the nucleolus to inhibit STK4 transcription via recruitment of XRCC5, leading to enhancement of self-renewal of liver CSCs21. These studies indicate that snoRNAs play a critical role in the regulation of tumor initiation and progression. However, the function and mechanisms of snoRNAs in liver CSCs still remain poorly understood. In this study, we identify a conserved snoRNA SNORA49 (originated from EP400 gene transcripts, NCBI ID: 677829) that is lowly expressed in liver CSCs and negatively regulates their self-renewal capacity. Downregulation of SNORA49 promotes enrichment of HNRNPU with ZC3H18 on SOX9 promoter to initiate SOX9 transcription, which induces the self-renewal of liver CSCs and HCC tumorigenesis. Of note, LNP-mediated delivery of SNORA49 RNAs with SOX9 ASOs exerts a potent synergistic anti-tumor effect on HCC tumors.
Results
SNORA49 is lowly expressed in liver CSCs
To explore the role of snoRNAs in liver CSCs, we sorted liver CSCs (CD13+CD133+) and non-CSCs (CD13−CD133−) from human primary HCC tumor tissues by fluorescence-activated cell sorting (FACS) as previously described22 and performed snoRNA transcriptome sequencing (Fig. 1A). We validated the top five downregulated snoRNAs in CSCs using qRT-PCR (Supplementary Fig. S1A). To determine their functions, we depleted these snoRNAs in primary HCC cells using short hairpin RNAs (shRNAs) (Supplementary Fig. S1B) and conducted sphere formation assays to evaluate the self-renewal capacity of tumor cells. Among the five snoRNAs, knockdown of SNORA49 most significantly enhanced sphere formation ability (Fig. 1B). Therefore, we selected SNORA49 to examine its function in the regulation of liver CSCs.
A Volcano plot showing differentially expressed snoRNAs between liver CSCs (CD13+CD133+) and non-CSCs (CD13−CD133−) sorted from HCC samples. Differential expression was assessed using an unpaired two-sided Student’s t test, and P-values were adjusted for multiple testing using the Benjamini-Hochberg method. snoRNAs with adjusted P < 0.05 and Log2FC > 1.2 were considered significant. The top five downregulated snoRNAs were highlighted in blue. B Sphere formation assays with snoRNA silencing. HCC#1 and HCC#2 represent two HCC patient samples. sh#1 and sh#2 represent two short hairpin RNAs designed for each snoRNA. Results are presented as means ± SD. n = 3 independent experiments. Exact P-values from upper to lower: 0.0003, 0.0004, 0.0002, 0.0007. C Schematic annotation of genomic localization of SNORA49. E1, Exon1. The black arrowhead indicates the transcription direction of EP400 and SNORA49. D, E Expression levels of SNORA49 in HCC tumors and peri-tumors (D), and in CSCs and non-CSCs (E) were detected by qRT-PCR (upper panel) and Northern blotting (lower panel). 18S rRNA was used as a loading control. Results are presented as means ± SD. n = 3 independent experiments. Exact P-values from left to right: (D) 0.0009, 0.0019, 0.0003, 0.0001, 0.0103; (E) 0.0002, 6.12E-06, 3.53E-05, 0.0021. F Fractionation of HCC cells followed by qRT-PCR. SNORA49 was localized in the nucleoplasm (upper panel). U1 was used as a positive control for nuclear localization. Immunoblotting showed separation efficiency (lower panel). EEA1, H3 and Nucleolin served as markers for cytoplasm, nucleoplasm, and nucleolus, respectively. Data are presented as means ± SD. n = 3 independent experiments. G Representative FISH images of SNORA49. Scale bar, 10 μm. H Kaplan-Meier survival analysis of liver cancer patients from the TCGA database. Patients were grouped by SNORA49 expression levels. Solid lines indicate the Kaplan-Meier estimate of survival probability, and shaded areas represent 95% confidence intervals. P = 0.0001 by Log-Rank test. * P < 0.05; ** P < 0.01; *** P < 0.001 by two-tailed Student’s t test. Data are representative of at least three independent experiments.
SNORA49 is located in the intronic region of its host gene EP400, between exons 28 and 29, with a length of 137 nt (Fig. 1C). qRT-PCR and Northern blotting revealed that SNORA49 expression was dramatically lower in HCC tumor tissues compared with peri-tumor tissues (Fig. 1D), lower in CSCs compared with non-CSCs (Fig. 1E), and lower in spheres compared with non-spheres (Supplementary Fig. S1C). When normalized to peri-tumor expression, there was also a stepwise decline from peri-tumor tissues to non-CSC tumors and further to liver CSCs (Supplementary Fig. S1D). Moreover, lower expression of SNORA49 in patient tumor tissues was further confirmed via fluorescence in situ hybridization (FISH) (Supplementary Fig. S1E). While SNORA49 was downregulated in liver CSCs compared with non-CSCs, the expression levels of its host gene EP400 remained unchanged (Supplementary Fig. S1F), suggesting that the downregulation of SNORA49 in liver CSCs was independent of its host gene.
Next, we investigated the subcellular localization of SNORA49 in liver CSCs by fractionation and qRT-PCR. We found that SNORA49 was primarily localized in the nucleoplasm, with a small amount in the cytoplasm and nucleolus (Fig. 1F). Consistently, RNA FISH also showed a nucleoplasm-specific localization of SNORA49 in both liver CSCs and non-CSCs (Fig. 1G). We also quantified copy numbers of SNORA49 in liver CSCs and non-CSCs using absolute quantification assays. We validated that SNORA49 was much lower in liver CSCs compared to non-CSCs (Supplementary Fig. S1G). Of note, Kaplan–Meier survival analysis of liver cancer patients, based on data from The Cancer Genome Atlas (TCGA), indicated that higher levels of SNORA49 were associated with better patient prognosis (Fig. 1H), suggesting that SNORA49 may serve as a tumor suppressor gene in HCC. Taken together, SNORA49 is lowly expressed in liver CSCs and correlated with improved survival of HCC patients.
SNORA49 depletion enhances self-renewal capacity of liver CSCs and promotes HCC progression
To explore the function of SNORA49 in liver CSCs, we constructed SNORA49-depleted Huh7 and primary HCC cells via lentivirus-mediated short hairpin RNA interference. Efficient depletion of SNORA49 was confirmed by qRT-PCR (Supplementary Fig. S2A) and Northern blotting (Supplementary Fig. S2B), without affecting the expression of its host gene EP400 (Supplementary Fig. S2C). After SNORA49 depletion, sphere formation of liver CSCs sorted from both Huh7 and primary HCC cells was dramatically enhanced, which could be inhibited by overexpression of SNORA49 (Fig. 2A). Serial passaging assays also demonstrated similar results (Fig. 2B), suggesting that SNORA49 depletion enhanced the self-renewal capacity of liver CSCs. Consistently, we observed an increased proportion of CSCs (CD13+CD133+) in SNORA49-depleted cells (Supplementary Fig. S2D). Furthermore, SNORA49 knockdown enhanced the proliferation capacity of tumor cells by colony formation and cell counting kit-8 (CCK-8) assays (Supplementary Fig. S2E, F). These results indicate that SNORA49 negatively regulates the stemness of CSCs in vitro.
A Sphere formation assay of SNORA49-depleted and rescued CSCs sorted from Huh7 cells and HCC primary cells. Scale bar, 200 μm. B Statistics of serial passaging and sphere formation assays. 1st, 2nd, and 3rd indicate the first, second, and third generations of spheres, respectively. Data are presented as means ± SD. n = 3 biologically independent experiments. Exact P-values from left to right: 0.0011, 0.0011, 0.0027, 0.0002, 3.67E-05, 0.0004, 0.0002, 0.0001, 0.0009, 0.0007, 0.0003, 0.0004, 0.0008, 0.0004, 0.0005, 0.0005, 0.0001, 2.55E-05. C Limiting dilution assay of SNORA49-depleted and control cells. Gradient numbers of cells were subcutaneously injected into BALB/c nude mice and allowed to grow for up to 3 months. Ratios of tumor-free mice for each group were calculated (left panel), and representative images of subcutaneous tumors were shown (right panel). n = 5 for each group. Scale bar, 1 cm. Exact P-values by paired one-sided t test from left to right: 0.0288, 0.0288. D 1 × 106 SNORA49-depleted or control cells were subcutaneously injected into BALB/c nude mice, and tumor growth was measured every 4 days. n = 4 for each group. Representative images of tumors were shown. Data are presented as means ± SD. Scale bar, 1 cm. Exact P-values from left to right: 0.0324, 0.0046, 0.0019, 0.0013. E 5 × 105 luciferase-labeled cells were orthotopically injected into the liver of BALB/c nude mice and visualized by in vivo imaging system (IVIS) one week later. Representative images (left panel) and statistics of bioluminescence signals (right panel) were shown. n = 6 for each group. Data are presented as means ± SD. Exact P-values from upper to lower: 2.81E-07, 2.31E-08. F Representative images of HCC tumors induced by hydrodynamic tail vein injection (HTVI) in Snora49+/+ and Snora49−/− mice. Results from two timepoints (3 weeks and 4 weeks) were shown. Scale bar, 5 mm. G H&E staining of tumors in (F). Scale bar, 0.5 mm. H, I Statistics of tumor numbers (H) and liver-to-body weight ratios (I) of Snora49+/+ and Snora49−/− mice. n = 6 for each group. Data are presented as means ± SD. Exact P-values from left to right: (H) 0.0025, 2.72E-06; (I) 0.0005, 0.0001. * P < 0.05; ** P < 0.01; *** P < 0.001 by two-tailed Student’s t test. Data are representative of at least three independent experiments.
To assess whether SNORA49 exerts inhibitory effects on liver CSCs in vivo, we performed a limiting dilution assay, a well-established approach to evaluate the stemness of tumor cells in vivo. We subcutaneously injected gradient numbers of SNORA49-depleted or control cells into BALB/c nude mice and observed that SNORA49 knockdown remarkably increased tumor incidence (Fig. 2C and Supplementary Fig. S2G). In addition, we subcutaneously injected SNORA49-depleted cells into BALB/c nude mice and monitored tumor growth for approximately one month. We observed that SNORA49-depleted tumors grew much larger than those of control cells (Fig. 2D). Moreover, SNORA49 knockdown significantly increased ratios of liver CSCs (CD13+CD133+) in subcutaneously formed tumors (Supplementary Fig. S2H). To further confirm the impact of SNORA49 depletion on tumor growth, we generated luciferase-labeled SNORA49-depleted cells and injected them into the livers of BALB/c nude mice for in situ tumor monitoring. Bioluminescence imaging revealed that SNORA49 knockdown remarkably promoted orthotopic tumor growth (Fig. 2E). Collectively, SNORA49 inhibits the stemness of human liver CSCs and suppresses HCC tumor growth.
Although we validated the role of SNORA49 in liver CSCs via both in vitro and in vivo assays, these models could not fully recapitulate the complexities of HCC tumorigenesis in vivo. To address this limitation, we generated Snora49 knockout (KO) mice using CRISPR/Cas9 technology (Supplementary Fig. S2I) and subjected them to HCC induction via hydrodynamic tail vein injection (HTVI), a widely used model that closely mimics human HCC tumorigenesis23,24,25,26. In brief, a saline solution containing Sleeping Beauty transposon plasmids (HrasG12V plus shp53) in a volume of 10% of the mouse body weight was rapidly injected into the tail vein. Huge hydrodynamic pressure could force fluid into livers, cause liver expansion, and push plasmid DNAs into hepatocytes, inducing tumor formation. Given that SNORA49 was highly conserved between mouse and human, sharing similar length, intact snoRNA motifs, and a comparable genomic localization (Supplementary Fig. S2J), mouse models provide more pathogenic rationale for liver tumorigenesis. We confirmed the genotype of Snora49 KO mice by PCR (Supplementary Fig. S2K) and verified the complete absence of Snora49 in mouse hepatocytes by Northern blotting (Supplementary Fig. S2L), without effect on the expression of its host gene Ep400, at both RNA and protein levels (Supplementary Fig. S2M, N). Following tail vein injection, we euthanized mice at two time points (3 and 4 weeks) to examine the role of Snora49 in both HCC initiation and progression. We noticed that Snora49 knockout mice exhibited an increased incidence of liver tumors and accelerated tumor growth compared with their wild-type (WT) littermates (Fig. 2F, G). These mice developed more and larger tumor lesions and displayed higher liver-to-body weight ratios, indicating a higher tumor burden (Fig. 2H, I). Taken together, SNORA49 plays a tumor-suppressive role in both HCC initiation and progression.
SNORA49 overexpression impairs stemness of CSCs and suppresses tumor formation in mouse models
We next wanted to examine whether the self-renewal capacity of liver CSCs could be suppressed by upregulating SNORA49 expression. We overexpressed SNORA49 in Huh7 and primary HCC cells using a lentivirus-mediated approach, which was confirmed by qRT-PCR and Northern blotting (Supplementary Fig. S3A). We found that overexpression of SNORA49 remarkably reduced sphere formation (Fig. 3A, B), CSC proportion (Supplementary Fig. S3B), and cell proliferation (Fig. 3C and Supplementary Fig. S3C). Furthermore, SNORA49 overexpression obviously inhibited tumor formation in limiting dilution assays (Fig. 3D and Supplementary Fig. S3D) and inhibited tumor growth both subcutaneously (Fig. 3E) and orthotopically (Fig. 3F and Supplementary Fig. S3E).
A Oncosphere formation assay of SNORA49-overexpressed and control CSCs. Scale bar, 200 μm. B Statistics of serial passaging and sphere formation assay. 1st, 2nd, and 3rd indicate the first, second, and third generations of spheres, respectively. n = 3 biologically independent experiments. Data are presented as means ± SD. Exact P-values from left to right: 8.85E-05, 5.08E-05, 1.57E-05, 0.0014, 0.0004, 0.0005. C Colony formation assay of SNORA49-overexpressed and control CSCs. Scale bar, 5 mm. D Limiting dilution assay of SNORA49-overexpressed and control cells. Ratios of tumor-free mice for each group were calculated (left panel), and representative images of subcutaneous tumors were shown (right panel). n = 5 for each group. P = 0.0069 by paired one-sided t test. Scale bar, 1 cm. E 1 × 106 SNORA49-overexpressed or control cells were subcutaneously injected into BALB/c nude mice. Tumor volumes were measured every 4 days. Scale bar, 1 cm. Data are presented as means ± SD. n = 4 for each group. Exact P-values from left to right: 0.0286, 0.0350, 0.0015, 0.0003. F 5 × 105 luciferase-labeled cells were orthotopically injected into the liver of BALB/c nude mice and visualized by IVIS one week later. Representative images were shown. n = 6 for each group. G Schematic diagram of AAV-mediated Snora49 overexpression in mouse liver, followed by HTVI induction. H Representative images of HCC tumors induced by HTVI in AAV-Scramble and AAV-Snora49 mice. Results from two timepoints were shown. Scale bar, 5 mm. I H&E staining of tumors in (H). Scale bar, 0.5 mm. J, K Statistics of tumor numbers (J) and liver-to-body weight ratios (K) of mice corresponding to (H). n = 6 for each group. Data are presented as means ± SD. Exact P-values from left to right: (J) 0.0003, 0.0002; (K) 0.0161, 0.0012. * P < 0.05; ** P < 0.01; *** P < 0.001 by two-tailed Student’s t test. Data are representative of at least three independent experiments.
To achieve liver-specific overexpression of Snora49 in vivo, we cloned full-length sequences of mouse Snora49 into an adeno-associated virus (AAV) plasmid and used a liver-specific TBG promoter to drive its expression (Fig. 3G). Packaged AAV was then injected into the tail veins of mice, and AAV-mediated overexpression of Snora49 in mouse hepatocytes was confirmed (Supplementary Fig. S3F). Two weeks later, when AAV-mediated Snora49 expression peaked, HCC tumors were induced in these mice using HTVI (Fig. 3G). As expected, overexpression of Snora49 dramatically suppressed tumor induction in these mice (Fig. 3H, I), with fewer and smaller tumor lesions and lower liver-to-body weight ratios compared to those of controls (Fig. 3J, K). Collectively, overexpression of SNORA49 effectively impairs the self-renewal capacity of liver CSCs and inhibits tumorigenesis.
SNORA49 binds to HNRNPU protein to abrogate its interaction with ZC3H18
We then sought to explore the mechanism by which SNORA49 regulated the self-renewal of liver CSCs. Canonically, snoRNAs containing H/ACA boxes mediate the pseudouridylation of target rRNAs via complementary base pairing in the nucleolus17. However, to date, there has been no validated rRNA site targeted by SNORA49, which was consistent with our observations that SNORA49 was localized in the nucleoplasm (Fig. 1G) rather than the nucleolus, where snoRNAs usually bind to rRNAs for modification. This led us to determine its non-canonical functions in the regulation of liver CSCs.
Recent studies have shown that snoRNAs exert non-canonical functions through interactions with various proteins18,20,21. We next performed RNA pulldown assays to identify potential binding proteins of SNORA49 in liver CSCs using biotin-labeled full-length SNORA49 probes (Fig. 4A). After silver staining and mass spectrometry (MS) analysis, we identified an RNA-binding protein, heterogeneous nuclear ribonucleoprotein U (HNRNPU) (Supplementary Fig. S4A) and confirmed their interaction by immunoblotting (Fig. 4B). To further assess whether the binding of SNORA49 and HNRNPU was specific, we conducted an RNA immunoprecipitation assay using a polyclonal HNRNPU antibody in sphere lysates from Huh7 and primary HCC cells. We found that HNRNPU specifically bound to SNORA49 but not to other snoRNAs, such as SNORD17 or SNORA74A (Fig. 4C). After incubating purified human HNRNPU protein with biotin-labeled SNORA49 probes, we observed a shift band in the migration pattern of SNORA49 on gel, indicating an interaction between HNRNPU and SNORA49 (Fig. 4D). Addition of unlabeled probes could compete with the labeled SNORA49 probes for binding to HNRNPU. However, neither SNORA74A or scramble probes with the same size and GC content as SNORA49 could compete with SNORA49 for HNRNPU binding (Supplementary Fig. S4B), further supporting the specificity of SNORA49-HNRNPU interaction. Consistently, both SNORA49 and HNRNPU exhibited nucleoplasm-specific localization by immunofluorescence staining (Fig. 4E). To test which part of SNORA49 interacted with HNRNPU, we performed a truncated RNA probe mapping assay. We found that the 31-60 nt region of SNORA49 was required for the interaction (Fig. 4F). We also generated Flag-tagged HNRNPU constructs lacking specific domains and subjected them to RNA pulldown by SNORA49. Deletion of the SPRY domain abolished the binding between HNRNPU and SNORA49 (Fig. 4G). Of note, the RGG motifs in the C terminal region of HNRNPU, previously shown to be responsible for RNA binding27, were not required for SNORA49 binding.
A Biotin-labeled RNA pulldown assay. Full-length sense and antisense sequence of SNORA49 were transcribed, biotin-labeled and incubated with Huh7 sphere lysates, followed by protein electrophoresis, silver staining and MS analysis. The arrow indicates the band of HNRNPU. B Immunoblotting of HNRNPU pulled down by SNORA49. C RIP assay performed in HCC and Huh7 sphere lysates with an anti-HNRNPU polyclonal antibody, followed by qRT-PCR. SNORD17 and SNORA74A served as controls. n = 3 biologically independent experiments. Data are presented as means ± SD. Exact P-values from left to right: 4.73E-05, 0.1097, 0.0955, 0.0001, 0.1771, 0.5754. D Biotin-labeled SNORA49 was incubated with increasing concentrations of HNRNPU and then subjected to EMSA. E Representative images showing colocalization of SNORA49 and HNRNPU in the nucleoplasm. Scale bar, 10 μm. F Truncated SNORA49 probes were subjected to RNA pulldown. ∆1-30nt, lacking the 1-30 nt while retaining other parts. G Flag-tagged HNRNPU was differently truncated (left panel) and then subjected to SNORA49 pulldown (right panel). ∆SAP, lacking the SAP domain while retaining other domains. FL, full length. H Co-IP assay was conducted with GST-tagged HNRNPU and anti-GST beads in Huh7 sphere lysates, followed by silver staining and MS analysis. Corresponding bands were indicated by arrows. I Immunoblotting validation of binding between HNRNPU and ZC3H18 by reciprocal co-IP with respective polyclonal antibodies in Huh7 sphere lysates. Endogenous RNAs were removed by RNase A before incubation with antibodies. J Co-IP assay was performed in control and SNORA49-depleted Huh7 sphere lysates using an anti-ZC3H18 polyclonal antibody. β-actin was used as a loading control. K, L GST-tagged HNRNPU and Flag-tagged ZC3H18 were purified and subjected to in vitro binding assay. GST-HNRNPU was pre-incubated with increasing concentrations of SNORA49 (K) or SNORA74A (L) and then incubated with Flag-tagged ZC3H18. GST-HNRNPU and the interacting protein Flag-ZC3H18 were pulled down by anti-GST beads. * P < 0.05; ** P < 0.01; *** P < 0.001 by two-tailed Student’s t test. Data are representative of at least three independent experiments.
Next, we sought to determine how SNORA49 exerted its functions through its association with the HNRNPU protein. First, we ruled out the possibility that SNORA49 affects the expression of HNRNPU, as its depletion did not alter the RNA or protein levels of HNRNPU (Supplementary Fig. S4C). Since HNRNPU is involved in RNA splicing28,29, we hypothesized that HNRNPU might engage in the splicing of the EP400 transcript, the host gene of SNORA49, thereby modulating SNORA49 biogenesis. However, depletion of HNRNPU had no effect on SNORA49 expression, as confirmed by qRT-PCR and Northern blotting (Supplementary Fig. S4D, E). These results suggest that the interaction of SNORA49 with HNRNPU may influence the function of HNRNPU instead of its expression.
HNRNPU is implicated in promoting gene transcription in various diseases. It has been shown to bind DDX5 to activate the transcription of LMO4, thereby supporting tumor growth in triple-negative breast cancer30. HNRNPU can also drive chromatin remodeling and facilitate enhancer activation to promote target gene expression in antiviral immunity31. However, since HNRNPU itself is not a transcription factor, it may need to cooperate with other transcription factors to exert its transcriptional promotion effect. To identify the interacting proteins of HNRNPU in liver CSCs, we purified GST-tagged HNRNPU protein and performed co-immunoprecipitation (co-IP) with anti-GST beads using Huh7 sphere lysates (Fig. 4H). Through this approach, we identified zinc finger CCCH-type containing 18 (ZC3H18) as a binding partner of HNRNPU (Supplementary Fig. S4F), and verified their interaction by reciprocal co-IP using their respective polyclonal antibodies (Fig. 4I). We also performed domain-mapping to identify the binding site on HNRNPU that interacted with ZC3H18. Interestingly, HNRNPU lacking the SPRY domain lost the ability to bind to ZC3H18 (Supplementary Fig. S4G), as was the same case for SNORA49 binding to HNRNPU. We also modeled the interaction between HNRNPU and ZC3H18 by Alphafold232. As expected, we found that the second domain of HNRNPU mediated their binding, highlighted by the interactions at the Y350-D353 sites (Supplementary Fig. S4H). This overlap of binding regions prompted us to investigate whether SNORA49 binding could affect the interaction between HNRNPU and ZC3H18.
To address this issue, we conducted co-IP experiments using an anti-ZC3H18 antibody in lysates from SNORA49-depleted and rescued Huh7 spheres. We found that the interaction between HNRNPU and ZC3H18 was remarkably enhanced in SNORA49-depleted cells, while rescued SNORA49 attenuated this enhancement (Fig. 4J). Similar results were observed using an anti-HNRNPU antibody (Supplementary Fig. S4I). Conversely, overexpressed SNORA49 strongly suppressed the binding of HNRNPU to ZC3H18 (Supplementary Fig. S4J, K). In order to eliminate the interference from endogenous cellular factors, we purified GST-tagged HNRNPU and Flag-tagged ZC3H18 to test their interaction in vitro. We incubated GST-HNRNPU with increasing concentrations of SNORA49 and then added Flag-ZC3H18 for co-IP using anti-GST beads. As expected, pre-incubation with SNORA49 hindered the binding of HNRNPU to ZC3H18 in a dose-dependent manner, with high concentrations of SNORA49 completely blocking this interaction (Fig. 4K), whereas another unrelated RNA, SNORA74A, had no such effect (Fig. 4L). Taken together, these data indicate that SNORA49 specifically binds to HNRNPU and impairs its interaction with ZC3H18.
ZC3H18 associates with HNRNPU to initiate SOX9 transcription
To further explore the mechanism by which SNORA49 modulated CSC self-renewal, we performed RNA sequencing of SNORA49-depleted and control Huh7 cells. We found that SNORA49 depletion obviously upregulated several transcription factors (Fig. 5A). We confirmed these changes in SNORA49-depleted Huh7 cells (Supplementary Fig. S5A). In contrast, SNORA49-overexpressed cells showed reversed downregulation of these transcription factors (Supplementary Fig. S5B). We then depleted these transcription factors in Huh7 cells and assessed their impact on sphere formation. Among these candidates, knockdown of SOX9 most significantly impaired the stemness of Huh7 cells (Fig. 5B). We further confirmed upregulation of SOX9 in SNORA49-depleted Huh7 and primary HCC cells by qRT-PCR and immunoblotting (Fig. 5C and Supplementary Fig. S5C). Conversely, in SNORA49-overexpressed cells, both mRNA and protein levels of SOX9 were remarkably reduced (Supplementary Fig. S5D).
A Volcano plot showing differentially expressed genes (FDR < 0.05, Log2FC > 0.8) in SNORA49-depleted Huh7 cells compared with control cells. B Sphere formation assays after silencing top 12 upregulated genes in (A). n = 3 independent experiments. Results are shown as means ± SD. Exact P-value: 2.22E-06. C Expression levels of SOX9 in SNORA49-depleted HCC cells were detected by immunoblotting. β-actin was used as a loading control. D Diagram of SOX9 promoter (left panel) and ChIP assay showing enrichment of ZC3H18 in the -600 ~ -400 fragment of SOX9 promoter (right panel). n = 3 independent experiments. Data are presented as means ± SD. Exact P-value: 6.07E-05. E ZC3H18 was specifically pulled down by biotin-labeled DNA probes of the -600 ~ -400 region of the SOX9 promoter. NSP, non-specific probes. F, G Chromatin accessibility assays of SOX9 promoter (-600 ~ -400 region) in HNRNPU and ZC3H18 depleted (F) or SNORA49-overexpressed Huh7 cells (G). n = 3 independent experiments. Results are shown as means ± SD. Exact P-values from left to right: (F) 0.0019, 0.0017; (G) 0.0021. H Enrichment of ZC3H18 in the -600 ~ -400 region of the SOX9 promoter was reduced in SNORA49-overexpressed Huh7 cells. n = 3 independent experiments. Data are presented as means ± SD. Exact P-value: 4.16E-05. I Dual-luciferase reporter assay of SOX9 promoter (-600 ~ -400 region) in ZC3H18-depleted Huh7 cells. Data are presented as means ± SD. n = 3 independent experiments. Exact P-value: 3.27E-05. J mRNA and protein levels of SOX9 after depletion and rescue of HNRNPU and ZC3H18 in Huh7 cells. Data are presented as means ± SD. n = 3 independent experiments. Exact P-values from upper to lower: 3.20E-04, 0.0001, 0.0003, 3.01E-06. K Representative immunofluorescence images showing consequent downregulation of SOX9 protein in CSCs following ZC3H18 depletion. Scale bar, 10 μm. L FACS analysis of CSC (CD13+CD133+) frequencies in ZC3H18-depleted or rescued Huh7 cells. Data are presented as means ± SD. n = 3 independent experiments. Exact P-values from left to right: 8.17E-05, 8.14E-05. * P < 0.05; ** P < 0.01; *** P < 0.001 by two-tailed Student’s t test. Data are representative of at least three independent experiments.
We then examine how SNORA49 regulated the expression of SOX9. Analysis of the TCGA database revealed a positive correlation between SOX9 expression and transcript levels of both HNRNPU and ZC3H18, with Pearson correlation coefficients (R) of 0.52 and 0.38, respectively (Supplementary Fig. S5E). Since SNORA49 could inhibit the interaction between HNRNPU and ZC3H18, we hypothesized that SNORA49 might impact target genes by interfering with this interaction. ZC3H18 is a well-characterized RNA-binding protein involved in nuclear RNA degradation33,34. Therefore, we speculated that ZC3H18 might bind to the pre-mRNA or mRNA of SOX9, thereby regulating SOX9 expression post-transcriptionally. We thus performed RNA immunoprecipitation experiments using the anti-ZC3H18 antibody but did not detect ZC3H18 binding to the pre-mRNA or mRNA of SOX9 (Supplementary Fig. S5F). They do not interact indirectly via HNRNPU, since there was no binding between HNRNPU and the SOX9 RNAs (Supplementary Fig. S5F). ZC3H18 contains a zinc finger domain, which is associated with DNA-binding activity, and interacts with HNRNPU, a known transcriptional regulator. This prompted us to explore its potential role in transcriptional regulation. It has been reported to bind to the BRCA1 promoter and activate its transcription, promoting ovarian cancer progression35. We next wanted to determine whether ZC3H18 could bind to the SOX9 promoter and cooperated with HNRNPU to regulate SOX9 transcription in liver CSCs. We then performed chromatin immunoprecipitation (ChIP) with a polyclonal antibody against ZC3H18. The 4 kb region upstream of the SOX9 transcription start site was divided into 10 fragments, and we found that ZC3H18 was enriched in the -600 ~ -400 region of the SOX9 promoter (Fig. 5D). We also used biotin-labeled DNA probes of this region to enrich ZC3H18 (Fig. 5E). Moreover, depletion of either HNRNPU or ZC3H18 remarkably reduced chromatin accessibility in the SOX9 promoter (Fig. 5F). Of note, overexpression of SNORA49, blocking the interaction between HNRNPU and ZC3H18, also caused reduction in chromatin accessibility in the SOX9 promoter (Fig. 5G) and impaired the binding of ZC3H18 to this region (Fig. 5H).
To further determine the transcriptional regulation of SOX9, we performed dual-luciferase reporter assays in ZC3H18-depleted (Supplementary Fig. S5G) and SNORA49-overexpressed cells. We cloned the -600 ~ -400 region of the SOX9 promoter into a pGL3 vector to drive luciferase expression. Luciferase activity was obviously reduced in ZC3H18-depleted cells (Fig. 5I), similar to the effect of SNORA49 overexpression (Supplementary Fig. S5H). In addition, SNORA49 overexpression also decreased levels of H3K4 trimethylation (Supplementary Fig. S5I) and H3K27 acetylation (Supplementary Fig. S5J), two histone modifications associated with transcriptional activation, at the SOX9 promoter. Consistent with changes in transcriptional activity, both mRNA and protein levels of SOX9 were remarkably decreased following knockdown of HNRNPU or ZC3H18 (Fig. 5J). Immunofluorescence staining also revealed a concomitant decrease in SOX9 expression following depletion of either HNRNPU or ZC3H18 in liver CSCs (Fig. 5K and Supplementary Fig. S5K). Consequently, proportions of liver CSCs were also remarkably reduced after HNRNPU or ZC3H18 depletion (Fig. 5L and Supplementary Fig. S5L). To explore the possibility that ZC3H18 indirectly regulated SOX9 expression via upstream regulators such as β-catenin, we performed ChIP experiments and found that there was no enrichment of ZC3H18 at the promoter of CTNNB1 (the gene encoding β-catenin) (Supplementary Fig. S5M). Collectively, these data indicate that the association and cooperation of HNRNPU with ZC3H18 are essential and sufficient for SOX9 transcription, which can be disrupted by SNORA49, leading to transcriptional suppression of SOX9 in liver CSCs.
SOX9 is highly expressed in liver CSCs and promotes self-renewal of liver CSCs
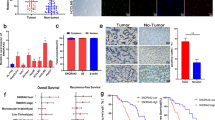
To determine the role of SOX9 in liver CSCs, we examined its expression by qRT-PCR and immunoblotting and observed that SOX9 was highly expressed in HCC tumor tissues compared with normal tissues (Fig. 6A), highly expressed in CSCs compared with non-CSCs (Fig. 6B), and highly expressed in spheres compared with non-spheres (Fig. 6C). Similar observations were validated in HCC patient samples by immunohistochemistry (Fig. 6D). High expression of SOX9 in HCC patient samples was further confirmed from the TCGA database (Fig. 6E), which was negatively correlated with survival outcomes for HCC patients (Fig. 6F). Of note, patients with higher SNORA49 expression and lower SOX9 levels had significantly better survival (Supplementary Fig. S6A). These results indicate that SOX9 plays a critical role in HCC tumorigenesis.
A–C Expression levels of SOX9 in HCC tumors and peri-tumor tissues (A), in CSCs and non-CSCs (B) and in oncospheres and non-spheres (C) were detected by qRT-PCR (upper panels) and immunoblotting (lower panels). β-actin was used as a loading control. Results are presented as means ± SD. n = 3 independent experiments. Exact P-values from left to right: (A) 0.0006, 4.33E-05, 0.0025, 6.57E-06, 6.89E-06; (B) 0.0016, 9.14E-06, 7.53E-06, 7.37E-06; (C) 1.38E-06, 5.67E-05, 2.19E-06, 4.44E-05. D Representative immunohistochemistry images of SOX9 expression in HCC patient samples. SOX9-positive staining is shown in brown. Scale bar, 50 μm. E Expression levels of SOX9 in tumors and peri-tumor tissues from TCGA liver cancer patients. n = 50 for Normal and n = 373 for Tumor. Data are presented as means ± SD. Exact P-value: 1.20E-26. F Kaplan-Meier survival analysis of TCGA liver cancer patients. Patients were grouped by SOX9 expression levels. A 95% confidence interval was indicated by dotted lines. G Sphere formation assay of SOX9-depleted or rescued CSCs sorted from Huh7 and HCC primary cells. Scale bar, 200 μm. H Colony formation assay of SOX9-depleted or rescued CSCs sorted from Huh7 and HCC primary cells. Scale bar, 5 mm. I 1 x 106 SOX9-depleted or control cells were subcutaneously injected into BALB/c nude mice. Tumor volumes were measured every 4 days. n = 4 for each group. Data are presented as means ± SD. Exact P-values from left to right: 0.0107, 0.0135, 0.0016, 0.0003. J SOX9 was depleted in SNORA49-silenced cells (upper panel) or overexpressed in SNORA49-overexpressed cells (lower panel). Representative images of sphere formation and statistical results were shown. Scale bar, 200 μm. n = 3 independent experiments. Data are presented as means ± SD. Exact P-values from upper to lower: 1.79E-05, 5.86E-05, 1.71E-05, 2.34E-05. * P < 0.05; ** P < 0.01; *** P < 0.001 by two-tailed Student’s t test. Data are representative of at least three independent experiments.
Next, we knocked down SOX9 in Huh7 and primary HCC cells (Supplementary Fig. S6B, C). SOX9 depletion dramatically reduced sphere formation (Fig. 6G and Supplementary Fig. S6D), cell proliferation (Fig. 6H), and CSC proportion (Supplementary Fig. S6E) in HCC cells, whereas overexpression of SOX9 could restore these oncogenic features. Moreover, SOX9 knockdown suppressed subcutaneous tumor growth in BALB/c nude mice (Fig. 6I). To further test the link between SNORA49 and SOX9, we silenced SOX9 in SNORA49-depleted cells. We found that enhanced sphere formation caused by SNORA49 knockdown was reversed by SOX9 depletion (Fig. 6J). By contrast, overexpression of SOX9 in SNORA49-overexpressed tumor cells augmented the self-renewal capacity of liver CSCs. Taken together, these results indicate that SOX9 is involved in the enhancement of self-renewal of liver CSCs as a downstream stemness factor of SNORA49.
Lipid nanoparticle delivery of SNORA49 and ASOs against SOX9 exerts synergistic antitumor effect on HCC tumors
In order to test the in vivo role of SOX9 in HCC tumor development, we generated Alb-Cre;LSL-Cas9 mice that constitutively expressed Cas9 in the liver and deleted Sox9 upon delivery of specific sgRNAs. We injected packaged AAV-TBG-sgSox9 and/or AAV-TBG-Snora49 plasmids via mouse tail veins and detected efficient reduction of Sox9 expression (Supplementary Fig. S6F). We then established HCC tumors via HTVI and collected tumors three weeks later (Fig. 7A). We observed that deletion of Sox9 dramatically inhibited tumor formation, with more pronounced effects combining with Snora49 overexpression (Fig. 7B, C). To further evaluate their effects on CSC self-renewal, we sorted tumor cells from HTVI-induced and AAV-treated mouse HCC tumors and injected them subcutaneously into BALB/c nude mice to generate secondary tumors. Both Snora49 overexpression and Sox9 deletion dampened secondary tumor formation, and their combination exerted a synergistic anti-tumor effect (Fig. 7D).
A Schematic diagram of AAV-mediated Sox9 deletion in mouse livers, followed by HTVI induction. B Representative images of HCC tumors induced by HTVI in AAV-sgSox9 and/or AAV-Snora49 mice. Scale bar, 5 mm. C Statistics of tumor numbers in mice corresponding to (B). n = 6 for each group. Data are presented as means ± SD. Exact P-values from upper to lower: 3.29E-06, 0.0001, 2.74E-05. D Secondary tumor formation of HTVI-induced and AAV-treated tumors in (B). 2 x 106 tumor cells were sorted by FACS and then subcutaneously injected into BALB/c nude mice. Tumor volumes were measured every 4 days. n = 4 for each group. Data are presented as means ± SD. Scale bar, 1 cm. Exact P-values from left to right: 0.0049, 0.0012, 0.0026. E Schematic diagram of HTVI induction and LNP administration in mice. F Representative images of HCC tumors in mice treated with LNPs containing Snora49 RNAs and/or Sox9 ASOs. Scale bar, 5 mm. G H&E staining of tumors in (F). Scale bar, 0.5 mm. H Statistics of tumor numbers in mice corresponding to (F). n = 6 for each group. Data are presented as means ± SD. Exact P-values from upper to lower: 4.79E-06, 0.0045, 0.0003. I Survival analysis of tumor-bearing mice treated with LNPs containing Snora49 RNAs and/or Sox9 ASOs. n = 6 for each group. Exact P-values from left to right by Log-Rank test: 0.0161 0.0450, 0.0005. J Schematic diagram of LNP administration in orthotopic PDC models. K 5 × 105 luciferase-labeled primary HCC cells were orthotopically injected into livers of BALB/c nude mice, followed by intravenous injection of LNPs containing SNORA49 RNAs and/or SOX9 ASOs. Representative images and statistical results of IVIS were shown. n = 6 for each group. Data are presented as means ± SD. Exact P-values from upper to lower: 8.22E-10, 4.49E-08, 7.33E-09. * P < 0.05; ** P < 0.01; *** P < 0.001 by two-tailed Student’s t test.
Despite its high efficacy, AAV-mediated gene manipulation has not yet been suitable for clinical applications. Lipid nanoparticles (LNPs) have been used to deliver siRNAs and mRNAs in clinical treatment36,37. To examine the potential of LNP-mediated RNA delivery for HCC treatment, we first tested in vitro transfection and silencing efficiency of delivering human SNORA49 RNAs and SOX9 ASOs into Huh7 cells via LNPs. Full-length human SNORA49 was in vitro transcribed using T7 polymerase, yielding a high quantity of capped RNAs. SOX9-targeting ASOs were chemically modified with 2’-O-methoxyethyl groups at both ends and phosphorothioate modifications in the backbone, enhancing their stability, binding affinity, and pharmacological properties38. LNP delivery achieved obvious SNORA49 overexpression and SOX9 depletion (Supplementary Fig. S7A), which consequently dampened the stemness of liver CSCs as validated by oncosphere formation assay (Supplementary Fig. S7B). To evaluate the therapeutic potential of SNORA49 and SOX9 in mouse HCC models, we encapsulated mouse Snora49 RNAs and ASOs against Sox9 with LNPs and administered them to tumor-bearing mice (Fig. 7E). Two weeks after HTVI, mice were treated with LNPs containing Snora49 RNAs and/or Sox9 ASOs at a dose of 1 mg/kg, administered intravenously once a week (Fig. 7E). We observed efficient RNA delivery in mouse livers via LNPs, reaching ~ 40-fold increase in Snora49 levels and 50% reduction in Sox9 expression, which was even more prominent when Snora49 and ASOs were used together (Supplementary Fig. S7C). LNPs containing Snora49 RNAs or Sox9 ASOs remarkably suppressed tumor growth (Fig. 7F–H). A Combination of Snora49 RNAs with Sox9 ASOs led to more reduction in tumor sizes and numbers, improving survival rates of tumor-bearing mice (Fig. 7I). We also assessed potential side effects of LNP-mediated RNA delivery in mice. Compared with control groups, those treated with either empty LNP vectors or LNP-RNAs displayed normal liver, kidney, and spleen morphology, comparable body weight and organ size, and only a slight but non-significant increase in hepatic expression of cytokines IFN-γ and IL-2 (Supplementary Fig. S7D, E). These results indicated that LNP-mediated RNA delivery had non-detective toxicity, minimal immunogenicity, and favorable biosafety. To further assess the translational potential of this therapy in humanized mouse models, we orthotopically injected luciferase-labeled primary HCC cells into livers of BALB/c nude mice, followed by treatment with empty LNPs or RNA-loaded LNPs (Fig. 7J). In this model, LNPs also achieved efficient RNA delivery and SOX9 silencing (Supplementary Fig. S7F). We found that combination treatment of human SNORA49 RNAs and SOX9 ASOs markedly suppressed orthotopic tumor growth, with greater effects than either agent alone (Fig. 7K). Taken together, LNP-mediated delivery of SNORA49 RNAs and SOX9 ASOs exerts potent synergistic anti-tumor effects on HCC tumors, providing strong preclinical evidence for this approach as a potential therapeutic strategy.
Discussion
The self-renewal of CSCs is tightly regulated by many factors, among which non-coding RNAs have gained a lot of attention in recent years. We and others have previously identified several non-coding RNAs that regulate liver CSCs22,39,40, while most of them are highly expressed in CSCs and exert tumor-promoting functions. In this study, we identified a conserved snoRNA SNORA49, which is downregulated in liver CSCs and exerts a suppressive effect on CSC self-renewal. SNORA49 is localized in the nucleoplasm, where it binds to HNRNPU protein to disrupt the interaction of HNRNPU with ZC3H18, leading to transcriptional repression of the SOX9 gene. Our in vitro experiments and mouse models further confirmed the tumor suppressor role of SNORA49. Of note, co-administration of SNORA49 and ASOs against SOX9 has synergistic anti-tumor effects, which provides new insights into the development of CSC-targeted therapies for HCC patients.
snoRNAs are originally identified in the nucleolus, where they mediate 2’-O-methylation or pseudouridylation of rRNAs41. However, recent studies have shown that some snoRNAs exert their functions in a non-canonical manner42,43. For instance, SNORA73 binds mRNA and 7SL RNA in the cytoplasm to strengthen the association of the SNORA73-targeted mRNA with the signal recognition particle (SPR), thereby facilitating ER translocation and protein secretion43. We recently revealed that elevated SNORD88B anchors WRN in the nucleolus, which promotes XRCC5 interaction with STK4 promoter to suppress its transcription, leading to augment of self-renewal of liver CSCs21. In this study, we found that SNORA49 exhibits a nucleoplasm-specific localization. This localization may be related to the absence of its target rRNAs in the nucleolus and its association with the nucleoplasm-localized protein HNRNPU. Our findings strongly support that SNORA49 does not possess canonical functions, with guiding rRNA modifications in the nucleolus, thereby theoretically excluding the possibility that it regulates liver CSC self-renewal through canonical mechanisms.
HNRNPU is a pluripotent protein involved in various biological processes, including chromatin organization, gene transcription, and RNA processing44,45,46. Here, we identified both SNORA49 and ZC3H18 bind to the same domain of HNRNPU, and SNORA49 binding blocks the interaction between HNRNPU and ZC3H18 to suppress SOX9 transcription. However, lower expressed SNORA49 releases HNRNPU, which then engages with ZC3H18 to enrich on the SOX9 promoter, resulting in SOX9 transcription for the induction of self-renewal of liver CSCs. However, how SNORA49 is downregulated in liver CSCs still needs further investigation.
SOX9 is recognized as a key stemness factor in a wide range of cancers. Previous studies have shown that SOX9 is highly expressed in HCC tumor tissues and liver CSCs15,47. However, how SOX9 is upregulated in HCC tumors is still unclear. In this study, we showed that ZC3H18 recruits HNRNPU to enrich on the promoter of SOX9, leading to its transcription initiation. In normal liver tissues, elevated SNORA49 and ZC3H18 competitively bind HNRNPU to inhibit the interaction of HNRNPU with ZC3H18, leading to inactivation of SOX9 transcription. In contrast, in liver CSCs, lower SNORA49 unleashes ZC3H18 to engage with HNRNPU to enrich on the promoter of SOX9, resulting in its transcription. We also showed that deletion of Sox9 inhibits the development of HCC tumors. Therefore, lowly expressed SNORA49 induces the association of HNRNPU with ZC3H18 to trigger SOX9 transcription, leading to the initiation of self-renewal of liver CSCs.
Treatment of HCC remains a huge challenge worldwide. Despite the use of various drugs and combined therapies, only a small subset of patients with advanced HCC achieves objective responses and survival benefits2. Targeting CSCs offers potential advantages over traditional regimens, given the biological complexity and heterogeneity of HCC. However, the identification of CSC-specific markers or factors that can reliably distinguish liver CSCs or tumor cells from normal tissues remains one of the major challenges in the development of such therapies. Here, we defined SNORA49 is lowly expressed, whereas SOX9 is highly expressed in liver CSCs. Moreover, SNORA49 progressively declines from peri-tumor tissues to tumor tissues and further to liver CSCs, whereas SOX9 exhibits the opposite trend, being markedly upregulated in tumors and liver CSCs but minimally expressed in normal hepatocytes. This inverse and causally linked expression pattern presents a valuable therapeutic window, enabling specific targeting of liver CSCs and tumor cells while minimizing potential toxicity to normal tissues. In addition, in the combined survival analysis of the TCGA liver cancer cohort, patients with a SNORA49HighSOX9Low profile had significantly better overall survival compared to those with a SNORA49LowSOX9High profile. This finding underscores the prognostic value and therapeutic potential of this inverse regulatory relationship. Lipid nanoparticles display low toxicity, biocompatibility and non-immunogenicity48. Especially, LNPs possess a natural propensity to accumulate in the liver after intravenous administration37, making them an ideal vehicle for targeted liver delivery. Here, we demonstrated that LNP-mediated delivery of SNORA49 RNAs and SOX9 ASOs converged to reduce SOX9 expression and exhibited potent synergistic anti-tumor effect on HCC tumors. Efficient delivery of RNAs and ASOs into liver tumors with LNPs makes RNA-based drugs possible to treat cancer patients. We believed that RNA-based drugs will be irreplaceable for tumor therapy in the near future.
In summary, our findings reveal that lowly expressed SNORA49 releases the interaction of HNRNPU with ZC3H18 in liver CSCs to induce SOX9 transcription, leading to the enhancement of liver CSC self-renewal. LNP-mediated delivery of SNORA49 RNAs and SOX9 ASOs exerts a potent synergistic anti-tumor effect on HCC tumors, highlighting the therapeutic potential of this approach for clinical intervention to HCC patients.
Methods
All experiments in this study were conducted in compliance with relevant ethical regulations and approved by the Institutional Committee of the Institute of Biophysics, Chinese Academy of Sciences. Both male and female mice (6–8 weeks of age) were housed under specific pathogen-free conditions with a 12 h light/dark cycle, temperature of 20 ± 2 °C, and 50% relative humidity. Mice were randomly assigned to experimental groups, and no sex-dependent differences were observed in any of the analyses. All animal protocols were reviewed and approved by the Institutional Animal Care and Use Committee of the Institute of Biophysics, Chinese Academy of Sciences. Mice were euthanized before tumors exceeded 15 mm in diameter or developed skin ulceration. No biases were introduced with respect to patient sex or gender.
Cell lines and HCC samples
Human HCC cell lines Huh7 and PLC were provided by Dr. Zeguang Han (Shanghai Jiaotong University, School of Medicine, Shanghai, China), and HEK293T cells were obtained from ATCC (American Type Culture Collection, Manassas, VA, USA). HCC cell lines and HEK293T cells were cultured in DMEM medium supplemented with 10% FBS, 100 μg/ml penicillin G and 100 U/ml streptomycin. Fresh tumor tissues were minced into 1 mm3 pieces and then digested with digestion buffer (0.1% IV collagenase, 0.01% DNase, 0.05% protease dissolved in serum-free DMEM) at 37 °C for 45 min. The supernatant was then filtered through a 70 μm filter, and the filtrate was centrifuged at 50 x g for 1 min to further remove clumps. The supernatant was collected and centrifuged at 150 x g for 8 min. Cells were enriched in the precipitate, and primary HCC cells were obtained after lysing the erythrocytes. All human HCC specimens were obtained from a partial hepatectomy series at Wuxi People’s Hospital (Wuxi, China) and the Department of Hepatobiliary Surgery, PLA General Hospital (Beijing, China). All tissue samples were obtained from consenting patients and approved by the Institutional Review Board of the Institute of Biophysics, Chinese Academy of Sciences, by the Ethics Committee of the Affiliated Wuxi People’s Hospital of Nanjing Medical University, and by the Ethics Committee of the PLA General Hospital. Clinical information of HCC patients in this study is listed in Supplementary Table S1. Materials used in this study are listed in Supplementary Data 1.
Sphere formation assay
2000 Huh7 or primary HCC cells were counted, seeded in low-attachment 6-well plates (Corning) and cultured in serum-free DMEM/F12 medium containing 20 ng/ml epithelial growth factor, 20 ng/ml fibroblast growth factor-2, N2 and B27 for 1-2 weeks. The number of spheres was counted under a stereomicroscope, and the percentage of sphere-initiating cells was calculated as (number of spheres formed) / (number of cells seeded per well) × 100%.
Colony formation assay
2000 Huh7 or HCC cells were seeded into 6-well plates and cultured in DMEM medium containing 10% FBS, 100 μg/ml penicillin G and 100 μ/ml streptomycin for approximately two weeks. After washing with PBS, cells were fixed in 4% PFA for 10 min and then stained with 0.1% crystal violet for 30 min. The plates were gently rinsed with running water and dried naturally.
Cell counting kit-8 (CCK-8) assay
10,000 Huh7 cells were seeded into 96-well plates and cultured in DMEM medium containing 10% FBS, 100 μg/ml penicillin G and 100 μ/ml streptomycin for 12 h to allow cell attachment. Then, 10 μl CCK-8 reagent (ZomanBio) was added to each well, followed by incubation at 37 °C for 2 h. The absorbance at 450 nm (OD450) was measured using a microplate reader. Cell proliferation rates were calculated based on the OD450 values.
Short hairpin RNA (shRNA) interference
Three shRNAs for each gene were cloned into the pSicoR-Puro lentiviral vector (Catalog No. 12084, Addgene). shRNA sequences used in this study are listed in Supplementary Table S2. The vectors were transfected into HEK293T cells with packaging plasmids pMD2.G (Catalog No. 12259, Addgene) and psPAX2 (Catalog No. 12260, Addgene). 48 h after transfection, the supernatant containing lentivirus was collected and filtered through a 0.45 μm filter, mixed with an equal volume of fresh DMEM medium and added to Huh7 or primary HCC cells for 24 h, followed by puromycin selection.
Gene overexpression
Full-length sequence of SNORA49 and coding sequence of HNRNPU, ZC3H18 and SOX9 were cloned into the pLVX-IRES-Puro or pLVX-IRES-GFP plasmid and transfected with packaging plasmids pMD2.G and psPAX2 into HEK293T cells for 48 h. Lentivirus was collected, mixed with an equal volume of DMEM medium, and transfected into target Huh7 or primary HCC cells for 24 h, followed by puromycin selection or GFP sorting.
Quantitative RT-PCR
Total RNAs were extracted from the samples using TRIzol and reverse transcribed into cDNAs by All-In-One 5X RT Master Mix (Abm). Quantitative real-time PCR analysis was conducted using the SYBR Green reaction system on the CFX Connect™ Real-Time PCR Detector (Bio-Rad, USA). All experiments were repeated three times independently and were analyzed using the 2-ΔΔCt method. U6, ACTB and 18S rRNA were used as internal references. Primers used in this study are listed in Supplementary Data 2.
Absolute quantification assay
The sequence of SNORA49 was cloned into the pcDNA3 plasmid. Plasmids were serially diluted and used as templates for qRT-PCR to generate a standard curve of CT values and copy numbers of SNORA49. Total RNAs of 2 × 106 CSCs or non-CSCs were extracted and reverse transcribed to cDNAs, followed by qRT-PCR with SNORA49-specific primers. The obtained CT values were then projected onto the standard curve to inversely calculate the copy number of SNORA49 in CSCs and non-CSCs.
Cellular fractionation
2 × 107 HCC cells were collected and lysed on ice for 10 min with 200 μl lysis buffer (1% NP-40, 0.25% sodium deoxycholate, 10 mM Tris pH 7.4, 140 mM NaCl, 1.5 mM MgCl2, 1 x protease inhibitor cocktail). The lysates were centrifuged at 300 x g for 5 min, and the supernatant contained the cytoplasmic fraction. For further separation of the nuclear and nucleolar fractions, the pellet was resuspended in 200 μl sucrose solution (340 mM, 5 mM MgCl2) and sonicated to break up the floccules in the suspension. Then 200 μl sucrose solution (880 mM, 5 mM MgCl2) was added, and the nucleolar fraction was pelleted by centrifugation at 500 x g for 30 min, leaving the nuclear fraction in the supernatant. Finally, the nucleolar pellet was suspended in 200 μl sucrose buffer (340 mM, 5 mM MgCl2).
RNA fluorescence in situ hybridization and immunofluorescence staining
HCC cells were seeded on slides, fixed in 4% paraformaldehyde (PFA) for 15 min, and permeabilized with 1% Triton X-100 for 15 min. Biotin-labeled SNORA49 probes were diluted to a concentration of 10 ng/μl in hybridization buffer(50% formamide, 5 × SSC, 500 ng/μl yeast tRNA, 1 × Dehardt’s solution, 500 ng/μl sperm DNA, 50 ng/μl Heparin, 2.5 mM EDTA, 0.1% Tween-20) and added to the slides and incubated at 45 °C for 2 h. After washing three times with 2 x SSC, TSA fluorescein (Akoya) was used to amplify the biotin signals according to the manufacturer’s instructions. For immunofluorescence staining, 10% donkey serum was used for blocking for 30 min. The primary antibodies were incubated at room temperature for 2 h or at 4 °C overnight. After washing with PBS for three times, the slides were incubated with secondary antibodies at room temperature for 30 min. The slides were mounted with DAPI and observed using Nikon A1R + confocal microscope.
Flow cytometry
Primary HCC tissues were dissociated into single cells and then were stained with CD45 antibodies to gate CD45-negative tumor cells and stained with FITC-conjugated CD13 and PE-conjugated CD133 antibodies to gate CSCs. Cells were stained with fluorescent antibodies on ice for 30 min and then sorted by flow cytometry. Data were analyzed by FlowJo V10. Flow cytometry gating strategies are listed in Supplementary Data 3.
Limiting dilution assay (LDA) and xenograft growth in nude mice
For LDA, HCC cells were serially diluted, mixed with 100 μl Matrigel and subcutaneously injected into BALB/c nude mice. The percentage of tumor-free mice was calculated 3 months later. For subcutaneous tumor growth, 1 × 106 HCC cells were injected into BALB/c nude mice. Tumor volume was measured every 4 days, starting from day 8 until day 28 and calculated as 1/2*(length)*(width)*(width). All BALB/c nude mice were 6–8 weeks old and provided by Beijing Vital River Laboratory Animal Technology. Mice were sacrificed before the tumor diameter exceeded 15 mm or skin ulcers appeared. Mouse experiments were approved by the Animal Care and Use Committees of the Institute of Biophysics, Chinese Academy of Sciences.
In vivo imaging system (IVIS)
5 x 105 luciferase-labeled Huh7 or primary HCC cells were orthotopically injected into the livers of BALB/c nude mice. 1-2 weeks post-injection, mice were intraperitoneally injected with 3 mg D-luciferin potassium salt and then anesthetized with isoflurane. Luciferase signals were detected by IVIS Lumina3 (PerkinElmer). Data were analyzed by Living Image V4.
Construction of knockout mice by CRISPR/Cas9 technology
Snora49 KO mice in C57BL6/J background were generated using CRISPR/Cas9 technology. Two sgRNAs (listed in Supplementary Table S3) targeting intronic sequences flanking the Snora49 locus were synthesized and injected into zygotes, which were transplanted into the uterus of pseudo-pregnant females to obtain F0 mice. To generate liver-sgSox9 mice, the U6 promoter in the AAV-sgRNA plasmid (Catalog No. 60231, Addgene) was replaced with the liver-specific TBG promoter (AAV-TBG). sgRNAs targeting the coding sequence of Sox9 were cloned into the AAV-TBG plasmid and co-transfected with pHelper and pAnc into HEK293T cells for 48 h for virus packing. Then the culture medium was collected and ultracentrifuged at 25,000 x g for 2 h. The pellet was resuspended with 0.9% NaCl solution and injected intravenously into Alb-Cre;LSL-Cas9 mice in a volume of 200 μl per mouse. The efficiency of AAV-mediated Sox9 knockout was assessed by qRT-PCR and immunoblotting. All mice were euthanized before the tumor diameter exceeded 15 mm or skin ulcers appeared. Alb-Cre mice were obtained from GemPharmatech Co., Ltd, and LSL-Cas9 mice were from Jackson Laboratory.
Hydrodynamic tail vein injection (HTVI)
8-week-old mice with C57BL/6 background were injected with 0.9% NaCl solution containing 20 μg of pBABE-c-mycT58A+HRasG12V plasmid (Catalog No. 11130, Addgene), 20 μg of pT2-shP53 plasmid (Catalog No. 124261, Addgene) and 20 μg of pT2/C-Luc//PGK-SB13 plasmid (Catalog No. 20207, Addgene). The saline was in a volume equivalent to 10% of the mouse body weight and was rapidly injected into the mouse tail vein within 7 s.
Generation of Snora49 overexpression mouse model
To achieve liver-specific overexpression of Snora49, the sequence of Snora49 was cloned into the AAV-TBG plasmid, packaged into AAV, and administered to mice via tail vein injection. Two weeks post-injection, overexpression of Snora49 was detected by qRT-PCR and Northern blotting.
RNA pulldown and mass spectrometry assay
Biotin-labeled sense and antisense probes of SNORA49 were in vitro transcribed using the T7 RNA polymerase (Roche) and biotin labeling agent (Roche). Sphere lysates of Huh7 cells were incubated with 3 mg probes at 4 °C overnight. Then streptavidin magnetic beads were added to pull down RNA probes and their binding proteins. After washing with lysis buffer (RIPA), the proteins were subjected to SDS-PAGE and silver staining. Differential bands were collected for liquid chromatography-tandem mass spectrometry analysis.
For LC-MS/MS analysis, silver-stained gels were destained, reduced with DTT, alkylated with iodoacetamide, and subjected to overnight digestion with trypsin. Peptides were extracted in multiple steps using acetonitrile at different concentrations and subsequently analyzed on a nanoLC system coupled to a Q Exactive mass spectrometer (Thermo Scientific) using a reversed-phase C18 analytical column (75 μm × 20 cm, 3 μm particle size). Peptides were separated at a flow rate of 300 nL/min with mobile phase A (0.1% formic acid in H2O) and mobile phase B (0.1% formic acid in acetonitrile). The gradient was programmed as follows: 4% B at 0 min, 8% B at 8 min, 22% B at 58 min, 32% B at 70 min, 90% B at 73–80 min. The Q Exactive was operated in data-dependent acquisition mode (top 20), consisting of one full MS scan followed by MS/MS of the 20 most intense precursor ions. Full MS scans were acquired over m/z 300–1600. The spray voltage was set to 2.0 kV, and the capillary temperature was 320 °C. Dynamic exclusion was set to 40 s.
Database searching was performed with the SEQUEST HT engine in Thermo Proteome Discoverer (version 1.4.0.288) against the UniProt human 2022 database. Searches were conducted with trypsin specificity, allowing up to two missed cleavages. The precursor ion mass tolerance was set to 10 ppm, and the fragment ion mass tolerance to 20 mDa. Carbamidomethylation of cysteine was specified as a fixed modification, whereas oxidation of methionine and lactylation of lysine were set as variable modifications. Spectral matches were filtered with Percolator, applying a Delta Cn threshold of < 0.1 and a false discovery rate (FDR) cutoff of 1%. Peptides were further filtered by requiring High peptide confidence. Proteins identified by mass spectrometry are listed in Supplementary Data 4.
Domain mapping
HNRNPU was divided into 4 fragments according to the distribution of functional domains. The sequence of full-length HNRNPU or HNRNPU lacking each domain was cloned into the p3xFlag-CMV10 plasmid and then transfected into HEK293T cells. 48 h later, cells were lysed with RIPA and subjected to immunoblotting to confirm the production of truncated HNRNPU proteins. Correctly truncated HNRNPU proteins were then incubated with biotin-labeled SNORA49 probes for RNA pulldown or with ZC3H18 for Co-IP assay.
Electrophoretic mobility shift assay (EMSA)
Human HNRNPU was cloned to the pGEX-6p-hDcp2 plasmid (Catalog No. 72216, Addgene) and purified by GST-tagged protein purification. Biotin-labeled SNORA49 probes were incubated with HNRNPU, with or without unlabeled probes, and then subjected to mobility shift assay using a Chemiluminescent RNA EMSA Kit (Beyotime) according to the manufacturer’s protocol. For competitive EMSA with SNORA74A and scramble probes, full-lengthen SNORA74A sequence (200 nt) was used, and the synthetic scramble RNA has the same lengthen (137 nt) and GC content (48%) as SNORA49.
Co-immunoprecipitation (Co-IP) assay
Sphere lysates of Huh7 cells were treated with RNase A at 37 °C for 30 min to remove endogenous RNAs and pre-cleared with Protein A/G agarose beads for 1 h and then incubated with anti-HNRNPU or anti-ZC3H18 antibodies at 4 °C for 2 h. Protein A/G agarose beads were added again to precipitate the target protein and its potential binding proteins, followed by immunoblotting.
In vitro competitive binding
GST-tagged HNRNPU and Flag-tagged ZC3H18 were purified through affinity chromatography. SNORA49 or SNORA74A was transcribed in large quantities using the T7 high yield RNA transcription kit (Vazyme). GST-HNRNPU was pre-incubated with increasing concentrations of SNORA49 or SNORA74A at 4 °C for 2 h, followed by incubation with Flag-ZC3H18 at 4 °C for an additional 2 h. The complexes were then captured with GST-beads. The interaction between HNRNPU and ZC3H18 was semi-quantified by immunoblotting.
Chromatin immunoprecipitation (ChIP) assay
Huh7 sphere cells were cross-linked with 1% formaldehyde at 37 °C for 10 min, washed twice with PBS and lysed in SDS lysis buffer (1% SDS, 10 mM EDTA, 50 mM Tris). DNA was fragmented into 200 ~ 500-bp fragments by sonication at 300 W for 1 min 4 times. The lysate was incubated with 4 μg anti-ZC3H18 polyclonal antibodies at 4 °C overnight. After incubation with Protein A/G agarose beads at 4 °C for 2 h, DNA was eluted and analyzed by qPCR with specific primers listed in Supplementary Table S4.
Chromatin accessibility assay
The nuclei of Huh7 sphere cells were isolated using the NE-PER nuclear and cytoplasmic extraction kits (Thermo), suspended with 200 μl of DNase Digestion Buffer and digested with 2 μ of DNase I at 37 °C for 5 min. Digestion was stopped with Digestion Stop Buffer. DNAs were extracted and analyzed by qPCR.
Dual-luciferase reporter assay
The sequence of the SOX9 promoter was cloned into the pGL3 vector and co-transfected with the pRL-TK plasmid into Huh7 cells for 36 h. After the cells were lysed, the luminescence signals were detected using a Dual Luciferase Reporter Gene Assay Kit (Beyotime) according to the manufacturer’s instructions.
Transcriptome sequencing
Total RNAs of 2 x 107 SNORA49-depleted and control cells were extracted by TRIzol, followed by DNBSEQ sequencing (BGI). After quality control, clean reads were aligned to the human genome (GCF_000001405.39_GRCh38.p13). Data analysis was performed by R 4.2.
Immunohistochemical staining
The tumor samples were fixed with 4% PFA for 48 h, embedded with paraffin, and sectioned into 10 µm thick slides. The slides were deparaffinized by sequential washes in xylene, 100% ethanol, 95% ethanol, and 85% ethanol, each for 10 min, and then rehydrated in PBS. Antigen retrieval was performed using an antigen retrieval buffer (Tris/EDTA pH 9.0) by heating the slides at 95 °C for 15 min. Endogenous peroxidase was quenched by incubating the slides with 3% hydrogen peroxide (H2O2) for 10 min. The slides were then blocked with 10% donkey serum for 30 min, incubated with primary antibodies for 2 h, and incubated with HPR-conjugated secondary antibodies for 1 h, all at room temperature. The immunoreactivity was visualized using DAB staining, and the slides were counterstained with hematoxylin. Finally, the slides were mounted with neutral balsam for microscopic examination.
Lipid nanoparticle packaging and administration
For large-scale transcription of human and mouse SNORA49, we adopted a co-transcriptional capping reaction system, in which the standard capping reagent m7G(5’)ppp(5’)G was added together with DNA templates, NTPs, and T7 RNA polymerase, following the manufacturer’s instructions (Vazyme). ASOs targeting SOX9 were synthesized and chemically modified (Tsingke). ASOs targeting human and mouse SOX9 have identical sequences, which are listed in Supplementary Table S5. The LNP formulation was based on commercially available ionizable lipids SM-102 (Moderna) and consisted of the following components: SM-102, cholesterol, 1,2-distearoyl-sn-glycero-3-phosphocholine (DSPC), and PEG2000-DMG, with a molar ratio of 50:10:38.5:1.5. The average particle size of LNPs was approximately 100 nm, as determined by dynamic light scattering (DLS). RNAs were packaged into LNPs with a concentration of 0.35 μg/μl. For in vitro LNP administration, Huh7 cells were treated with 100 ng/ml LNP-packaged RNAs for 24 h, followed by collection for assessment of transfection and gene silencing efficiency. For in vivo LNP treatment in mouse models, 1 mg/kg RNAs (corresponding to 57 μl LNPs) were injected into the tail vein of tumor-bearing mice (20 g) once a week. For biosafety assessment, WT mice were treated with empty LNP vectors or RNA-loaded LNPs at the same dose used in HTVI-LNP model. Body and major organ weights were recorded, liver, kidney, and spleen morphology were evaluated, and expression levels of IFN-γ and IL-2 in the liver were measured by qRT-PCR 48 h after injection.
Statistics and reproducibility
Statistical analysis was performed using GraphPad Prism 9.0 and Excel 2019. Unless otherwise stated, statistical significance was assessed by unpaired two-tailed Student’s t tests. A significance threshold of P < 0.05 was applied, with results denoted as * P < 0.05, ** P < 0.01, *** P < 0.001, and ns (not significant). All assays were performed in at least three independent experiments, and representative results are presented.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The snoRNA sequencing dataset analyzed in this study was previously generated and reported in our earlier publication21, which has been deposited to the Gene Expression Omnibus under accession code GSE233521. SNORA49-depletion sequencing data are available from the National Genomics Data Center under accession code HRA010557. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium with the identifier PXD061379. The remaining data are available within the Article, Supplementary Information, Supplementary Data, or Source Data file. Source data are provided in this paper.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74, 229–263 (2024).
Lee, T. K., Guan, X. Y. & Ma, S. Cancer stem cells in hepatocellular carcinoma - from origin to clinical implications. Nat. Rev. Gastroenterol. Hepatol. 19, 26–44 (2022).
Ma, S. et al. Identification and characterization of tumorigenic liver cancer stem/progenitor cells. Gastroenterology 132, 2542–2556 (2007).
Matsumoto, T. et al. Proliferating EpCAM-positive ductal cells in the inflamed liver give rise to hepatocellular carcinoma. Cancer Res. 77, 6131–6143 (2017).
Ang, C. H. et al. Lgr5(+) pericentral hepatocytes are self-maintained in normal liver regeneration and susceptible to hepatocarcinogenesis. Proc. Natl. Acad. Sci. USA 116, 19530–19540 (2019).
Lee, T. K. et al. Blockade of CD47-mediated cathepsin S/protease-activated receptor 2 signaling provides a therapeutic target for hepatocellular carcinoma. Hepatology 60, 179–191 (2014).
Tong, M. et al. ANXA3/JNK signaling promotes self-renewal and tumor growth, and its blockade provides a therapeutic target for hepatocellular carcinoma. Stem Cell Rep. 5, 45–59 (2015).
Tong, M. et al. Efficacy of annexin A3 blockade in sensitizing hepatocellular carcinoma to sorafenib and regorafenib. J. Hepatol. 69, 826–839 (2018).
Yang, L. et al. Targeting cancer stem cell pathways for cancer therapy. Signal Transduct. Target. Ther. 5, 8 (2020).
Laughney, A. M. et al. Regenerative lineages and immune-mediated pruning in lung cancer metastasis. Nat. Med. 26, 259–269 (2020).
Vasaikar, S. et al. Proteogenomic analysis of human colon cancer reveals new therapeutic opportunities. Cell 177, 1035–1049 (2019).
Guo, W. et al. Slug and Sox9 cooperatively determine the mammary stem cell state. Cell 148, 1015–1028 (2012).
Huang, K. K. et al. Spatiotemporal genomic profiling of intestinal metaplasia reveals clonal dynamics of gastric cancer progression. Cancer Cell 41, 2019–2037 (2023).
Santos, J. C. et al. SOX9 Elevation acts with canonical WNT signaling to drive gastric cancer progression. Cancer Res. 76, 6735–6746 (2016).
Liu, C. et al. Sox9 regulates self-renewal and tumorigenicity by promoting symmetrical cell division of cancer stem cells in hepatocellular carcinoma. Hepatology 64, 117–129 (2016).
Liu, Y. et al. Yap-Sox9 signaling determines hepatocyte plasticity and lineage-specific hepatocarcinogenesis. J. Hepatol. 76, 652–664 (2022).
Kiss, T. Small nucleolar RNAs: an abundant group of noncoding RNAs with diverse cellular functions. Cell 109, 145–148 (2002).
Siprashvili, Z. et al. The noncoding RNAs SNORD50A and SNORD50B bind K-Ras and are recurrently deleted in human cancer. Nat. Genet. 48, 53–58 (2016).
Falaleeva, M. et al. Dual function of C/D box small nucleolar RNAs in rRNA modification and alternative pre-mRNA splicing. Proc. Natl. Acad. Sci. USA 113, E1625–E1634 (2016).
Liang, J. et al. Non-coding small nucleolar RNA SNORD17 promotes the progression of hepatocellular carcinoma through a positive feedback loop upon p53 inactivation. Cell Death Differ. 29, 988–1003 (2022).
Gu, Y. et al. SNORD88B-mediated WRN nucleolar trafficking drives self-renewal in liver cancer initiating cells and hepatocarcinogenesis. Nat. Commun. 15, 6730 (2024).
Wang, Y. et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell 16, 413–425 (2015).
Calvisi, D. F. et al. Increased Lipogenesis, Induced by AKT-mTORC1-RPS6 Signaling, Promotes Development of Human Hepatocellular Carcinoma. Gastroenterology 140, 1071–1083 (2011).
Tao, J. et al. Modeling a human hepatocellular carcinoma subset in mice through coexpression of met and point-mutant β-catenin. Hepatology 64, 1587–1605 (2016).
Filliol, A. et al. Opposing roles of hepatic stellate cell subpopulations in hepatocarcinogenesis. Nature 610, 356–365 (2022).
Fan, W. et al. Matrix viscoelasticity promotes liver cancer progression in the pre-cirrhotic liver. Nature 626, 635–642 (2024).
Nozawa, R.-S. et al. SAF-A Regulates interphase chromosome structure through oligomerization with chromatin-associated RNAs. Cell 169, 1214–1227 (2017).
Ye, J. et al. hnRNP U protein is required for normal pre-mRNA splicing and postnatal heart development and function. Proc. Natl. Acad. Sci. USA 112, 3020–3029 (2015).
Isaac, R. et al. TM7SF3 controls TEAD1 splicing to prevent MASH-induced liver fibrosis. Cell Metab. 36, 1030–1043 (2024).
Han, B. et al. HNRNPU promotes the progression of triple-negative breast cancer via RNA transcription and alternative splicing mechanisms. Cell Death Dis. 13, 940 (2022).
Cao, L. et al. The nuclear matrix protein SAFA surveils viral RNA and facilitates immunity by activating antiviral enhancers and super-enhancers. Cell Host Microbe 26, 369–384 (2019).
Bryant, P. et al. Improved prediction of protein-protein interactions using AlphaFold2. Nat. Commun. 13, 1265 (2022).
Chi, B. et al. A sub-element in PRE enhances nuclear export of intronless mRNAs by recruiting the TREX complex via ZC3H18. Nucleic Acids Res. 42, 7305–7318 (2014).
Winczura, K. et al. Characterizing ZC3H18, a multi-domain protein at the interface of RNA production and destruction decisions. Cell Rep. 22, 44–58 (2018).
Kanakkanthara, A. et al. ZC3H18 specifically binds and activates the BRCA1 promoter to facilitate homologous recombination in ovarian cancer. Nat. Commun. 10, 4632 (2019).
Polack, F. P. et al. Safety and efficacy of the BNT162b2 mRNA covid-19 vaccine. N. Engl. J. Med. 383, 2603–2615 (2020).
Hou, X. et al. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 6, 1078–1094 (2021).
Crooke, S. T. et al. Antisense technology: an overview and prospectus. Nat. Rev. Drug Discov. 20, 427–453 (2021).
Wang, Y. et al. Long noncoding RNA lncHand2 promotes liver repopulation via c-Met signaling. J. Hepatol. 69, 861–872 (2018).
Chen, Z. et al. Circular RNA cia-MAF drives self-renewal and metastasis of liver tumor-initiating cells via transcription factor MAFF. J. Clin. Invest. 131, e148020 (2021).
Reichow, S. L. et al. The structure and function of small nucleolar ribonucleoproteins. Nucleic Acids Res. 35, 1452–1464 (2007).
Cheng, Y. et al. A non-canonical role for a small nucleolar RNA in ribosome biogenesis and senescence. Cell 187, 4770–4789 (2024).
Liu, B. et al. snoRNA-facilitated protein secretion revealed by transcriptome-wide snoRNA target identification. Cell 188, 465–483 (2024).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Song, H. et al. HNF4A-AS1/hnRNPU/CTCF axis as a therapeutic target for aerobic glycolysis and neuroblastoma progression. J. Hematol. Oncol. 13, 24 (2020).
Hosen, M. R. et al. Circulating microRNA-122-5p is associated with a lack of improvement in left ventricular function after transcatheter aortic valve replacement and regulates viability of cardiomyocytes through extracellular vesicles. Circulation 146, 1836–1854 (2022).
Ruzinova, M. B. et al. SOX9 expression is superior to other stem cell markers K19 and EpCAM in predicting prognosis in hepatocellular carcinoma. Am. J. Surg. Pathol. 47, 1–11 (2023).
Tenchov, R. et al. Lipid Nanoparticles─from liposomes to mRNA vaccine delivery, a landscape of research diversity and advancement. ACS Nano 15, 16982–17015 (2021).
Acknowledgements
We thank Drs. Xiang Ding, Yihui Xu, and Junying Jia (Institute of Biophysics) for technical support. We also thank Jing Li (Cnkingbio Company Ltd, Beijing, China) for technical support. This work was supported by the National Natural Science Foundation of China (82530038, 82130088, 82271785), National Key R&D Program of China (2020YFA0803501), Strategic Priority Research Programs of the Chinese Academy of Sciences (XDB0570301); and Natural Science Foundation of Beijing (Z231100007223013).
Author information
Authors and Affiliations
Contributions
Z.Y. designed experiments, interpreted results, and wrote the paper; Z.Z., Y.G., and Z.X. performed some experiments and analyzed data; Y.X. designed and constructed mouse models; Z.X., J.W., H.G., and Y.D. provided technical support; Y.C. provided HCC samples and analyzed data; Z.F. initiated, organized, designed, and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Søren Lykke-Andersen and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yi, Z., Zhou, Z., Gu, Y. et al. SNORA49 negatively regulates self-renewal of liver cancer stem cells and hepatocarcinogenesis via suppressing SOX9 transcription. Nat Commun 17, 766 (2026). https://doi.org/10.1038/s41467-025-66486-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66486-1