Abstract
The recruitment of telomerase to telomeres is a tightly regulated process which is stimulated by replication stress and the DNA damage response regulatory kinase ATR, via an unknown mechanism. Here, we demonstrate that nuclear filamentous actin is important for the stable interaction of telomerase with telomeres in immortal human cells, resulting in productive telomere elongation by telomerase in an actin-dependent manner. This process is regulated by both ATR and mTOR kinases, and employs other regulators of actin structure and function, such as WASP, ARP2/3 and myosin. Nuclear filamentous actin serves as a site for telomerase recruitment, which is mediated by telomere tethering on actin fibers in response to replication stress, allowing telomerase to localize to telomeres containing stalled replication forks. Overall, these data demonstrate that, in human cells which express telomerase, telomeric replication stress triggers the recruitment of telomerase to telomeres via a nuclear actin network, enabling telomere length maintenance.
Similar content being viewed by others
Introduction
Telomeres are nucleoprotein complexes comprised of tandem DNA repeats, TTAGGG in vertebrates, which cap the ends of linear chromosomes to maintain the integrity of the genome and ensure cellular survival1. Given their resemblance to damaged DNA, exposed telomeres activate the DNA damage response (DDR) which can result in chromosome end-to-end fusions. To suppress this response, telomeres are bound by the six-protein complex shelterin, which includes TRF1 and TRF2, which directly bind to duplex telomeric repeats and mediate binding of the remaining subunits: POT1, TPP1, RAP1, and TIN22,3. Shelterin proteins perform various functions to maintain telomere structure and function, including regulating telomere length.
In human stem and germline cells and ~85% of cancers, telomere synthesis is performed by the ribonucleoprotein enzyme telomerase4. Human telomerase associates with a large number of different proteins5; however, the minimal catalytic core is the telomerase reverse transcriptase (hTERT) and telomerase RNA (hTR)5,6,7. Recruitment of telomerase to telomeres is a highly regulated process4,8, predominantly occurring during S phase9,10; this cell cycle control is at least partially regulated by TRF1, which negatively regulates telomerase recruitment outside S phase11,12. TPP1 is also critical for telomerase recruitment, as it binds hTERT via an exposed N-terminal domain termed the TEL patch13. The interactions between TPP1, POT1, and TIN2 are also necessary to facilitate telomerase recruitment and processivity at telomeres14,15,16.
If telomeres become critically short, they can trigger activation of a DDR through either ATR or ATM17, PIKK (phosphatidylinositol-3 kinase-related) family kinases which govern the DDR. Shelterin typically represses the activation of both ATR and ATM at telomeres, specifically via POT1 and TRF2, respectively18,19,20,21. Furthermore, replication fork stalling within the telomere results in ATR activation22. Almost paradoxical, however, is the emerging evidence that the DDR is also crucial for regulation of telomerase presence at telomeres. In yeast, telomerase preferentially extends the shortest telomeres in a manner dependent on Tel1 (the yeast homolog of ATM)23,24,25,26. Furthermore, Tel1 is required for telomerase recruitment in budding yeast27, and both Tel1 and Rad3 (the fission yeast homolog of ATR) are required in fission yeast28,29,30. In human cells, ATR or ATM deficiency or mutation results in telomere shortening or instability31,32. Telomerase regulation by ATM/ATR may rely upon TRF1 displacement, as phosphorylation of TRF1 at an ATM/ATR target site results in its depletion from telomeres and heightened telomerase recruitment11,33. Furthermore, the depletion or inhibition of ATR or ATM results in reduced telomerase presence at telomeres11,34. This axis is supported by the finding that induced replication stress promotes telomerase recruitment11,35 and ultimately causes telomere lengthening11,22. These studies together suggest a tightly regulated process for telomerase recruitment which is coordinated by DNA replication, the DDR and the telomere maintenance machinery. However, the downstream substrate(s) of ATR involved in recruiting human telomerase to telomeres have not yet been identified, and the link between replication stress and telomerase recruitment has not been mechanistically characterized.
An emerging body of research has illustrated that a key regulator of the DDR is nuclear filamentous actin (F-actin). Although classically considered cytoplasmic, nuclear F-actin forms in response to double strand breaks (DSBs) and replication stress, where it facilitates the DDR by re-localization of damaged sites to the nuclear periphery for repair or fork restart36,37,38,39,40,41. This occurs globally in response to DNA damage, but also specifically at telomeres undergoing replication stress38,42. Under replication stress conditions, the polymerization of nuclear F-actin is dependent on ATR, whose activity is required for downstream phosphorylation of another PIKK family kinase, mTOR38, in turn regulating F-actin through the Wiskott-Aldrich syndrome protein (WASP) family43.
The convergence of these data suggests there is controlled actin-mediated interplay between DNA replication, the DDR, and telomere maintenance. Here we demonstrate that nuclear F-actin regulates telomerase recruitment in an ATR- and mTOR-dependent manner in response to replication stress. This recruitment results in productive telomere lengthening by telomerase, which is upregulated by replication stress and dependent on F-actin polymerization. Specifically, nuclear actin filaments appear to act as sites for telomerase recruitment, potentially due to telomere tethering on F-actin under conditions of replication stress. Furthermore, the recruitment of telomerase to telomeres occurs in proximity to stalled replication forks and F-actin. These data reveal a new function for the nuclear F-actin network in telomere maintenance.
Results
F-actin polymerization facilitates telomerase recruitment to telomeres
To better visualize hTR presence at telomeres, we modified an existing fluorescence in situ hybridization (FISH) protocol11,44 by generating SABER (signal amplification by exchange reaction) probes45 targeting both hTR and telomere sequences, greatly amplifying the intensity of the foci (Supplementary Fig. 1a-b), particularly hTR. Cells were also labeled with EdU to identify S phase cells for quantification, as this is predominantly when telomerase recruitment to telomeres occurs within the cell cycle9,10,11. The hTR SABER FISH signal is highly specific, as hTR foci were not detected in the hTR-negative WI-38 VA1346 cell line (Supplementary Fig. 1c). In untreated HEK293T cells, the number of co-localizing hTR and telomere foci ranged from 0 – 5 per S phase cell with a mean of ~1 (Fig. 1a, c, and Supplementary Fig. 2a-b), consistent with previous quantitation using standard FISH9,10,11; this small number of co-localizing foci in fixed cells reflects a “snapshot” of the dynamic interactions between telomerase and telomeres across S phase35,47. To exclude the possibility of these events being stochastic associations, we performed a rotation control48 to simulate random distribution of foci within nuclei. One fluorescence channel (telomeres) was rotated 90°; this resulted in significantly less hTR/telomere co-localizations (Supplementary Fig. 1d-e), demonstrating that while these events are infrequent, they occur more often than would be expected by chance.
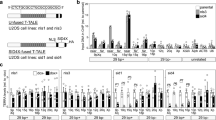
a Representative images of SABER FISH probing for telomeres (pink) and hTR (yellow) in 293T cells stained with DAPI (blue). Cells were treated with actin polymerization inhibitors LatB (0.2 μM), LatA (50 nM) or DMSO (no treatment control, NTC) for 16 h, ± 1.5 μM APH for 30 min. b Phalloidin staining (white) of 293T cells treated with DMSO (NTC), 0.2 μM LatB or 50 nM LatA for 16 h, also stained with DAPI (blue) and imaged by super-resolution Airyscan microscopy. c Average hTR/telomere co-localizations in S phase 293T cells following treatment with DMSO (NTC), LatB or LatA (for 16 h), ± 1.5 μM APH (for 30 min). Significance for the latrunculin - APH samples is expressed relative to NTC - APH; significance for the latrunculin + APH samples is expressed relative to NTC + APH. d Western blot of 293T cells treated with DMSO (NTC), LatB or LatA, ± APH, probed for Chk1 and Chk1 pS345, with vinculin as a control. e, f Percentage of S phase (EdU-positive) 293T cells with nuclear F-actin, visualized by transfection with either (e) nuclear-actin-ChB or (f) mCherry-actin. Cells were treated with DMSO or APH (1.5 μM) for 30 min. g Average hTR/telomere co-localizations in HeLa cells treated with control siRNA or two different ATR siRNAs. h Average hTR/telomere co-localizations in 293T cells treated with control or ATRIP pooled siRNAs ± 1.5 μM APH for 30 min. Significance for siATRIP - APH is expressed relative to siCtrl - APH; significance for siATRIP + APH is expressed relative to siCtrl + APH. i Representative images of SABER FISH for hTR (yellow) and telomeres (pink) with DAPI staining (blue) of nuclei in the indicated cell lines treated with DMSO (NTC) or 1.5 μM APH for 30 min. j Quantitation of telomerase recruitment to telomeres from (i). k SABER FISH for hTR (yellow) and telomeres (pink) with DAPI staining (blue) in the indicated cell lines treated with 0.2 μM LatB for 16 h. l Normalized quantitation of telomerase recruitment from (k). For all microscopy images, scale bar = 2 μm; co-localizations indicated by white arrows. Mean and number of nuclei quantified per replicate are listed in Source Data. All bar graphs displayed as mean ± SEM; n = 3 independent biological replicates; One-way ANOVA with Šídák’s multiple comparisons test (except e, f; paired two-tailed t test).
Given that the recruitment of mammalian telomerase to telomeres is promoted by replication stress11,49, and the emerging understanding that F-actin is involved in resolution of stalled replication forks38,41,50, we explored whether actin polymerization is also involved in telomerase recruitment. Treatment of 293T cells with two inhibitors of actin polymerization, latrunculin A and latrunculin B (LatA and LatB), which both bind to monomeric F-actin to prevent its polymerization51,52, resulted in a significant reduction in telomerase recruitment to telomeres in S phase cells (Fig. 1a-c, Supplementary Fig. 2a-b). Furthermore, while short-term treatment (30 min) with the polymerase inhibitor aphidicolin (APH) is sufficient to induce a DDR (as measured by phosphorylation of the ATR substrate Chk1; Fig. 1d) and promote an increase in telomerase recruitment, this increased recruitment was prevented by inhibiting actin polymerization using either LatA or LatB (Fig. 1c, Supplementary Fig. 2a-b). Treatment with the latrunculins (with or without APH) did not cause substantial perturbation to cell cycle progression (Supplementary Fig. 2c), suggesting that the observed effects on telomerase are due to F-actin facilitating recruitment, and not due to fluctuations in the cell cycle.
While we have previously demonstrated that replication stress promotes nuclear F-actin formation in S phase38, this was after 24 h incubation with low dose (0.4 μM) APH, unlike the short period of higher dose (1.5 μM) APH used above to induce telomerase recruitment without causing DNA DSBs11. To confirm that a short incubation with APH also promotes nuclear F-actin formation, we quantified the number of S phase cells with nuclear F-actin. This was performed by visualizing either endogenous nuclear actin using an actin chromobody53, tagged with both blue fluorescent protein (BFP) and a nuclear localization signal (NLS; nuclear-actin-ChB), or by overexpressing mCherry and NLS-tagged actin36,38. Using either approach, approximately 10% of unperturbed S phase 293T cells possess visible nuclear F-actin, which increased threefold after treatment with APH for 30 min (Fig. 1e, f, Supplementary Fig. 2d-e), confirming that this dose of replication stress is sufficient to induce nuclear F-actin polymerization.
These results support the hypothesis that replication stress promotes telomerase recruitment through activation of the DNA replication stress response and subsequent F-actin polymerization. The requirement of ATR for telomerase recruitment has only been previously examined in 293T cells11, so to validate the generality of this pathway, the requirement of ATR for recruitment was also confirmed in the cervical cancer cell line HeLa. Knockdown of ATR by siRNA resulted in a significant reduction in telomerase recruitment in HeLa cells (Fig. 1g, Supplementary Fig. 2f-g), which was not accompanied by an observable perturbation to cell cycle profiles (Supplementary Fig. 2h). To further confirm that ATR-mediated telomerase recruitment employs the canonical ATR response to replication stress, ATR interacting protein (ATRIP)54,55 was also depleted (Supplementary Fig. 2i). Following ATRIP knockdown in 293T cells, both with and without replication stress, there was a significant reduction in telomerase recruitment without major cell cycle perturbations (Fig. 1h, Supplementary Fig. 2j-l), confirming that ATR functions in this pathway via its canonical DDR signaling role.
Subsequently, the levels of telomerase recruitment in response to replication stress was examined in a panel of telomerase positive human cell lines, including HeLa, lung adenocarcinoma A549, fibrosarcoma HT1080, and colorectal carcinoma HCT116 cells. Following APH treatment to induce replication stress, all cell lines displayed a significant increase in telomerase recruitment with no substantial alteration to their cell cycle profile (Fig. 1i, j, Supplementary Fig. 2m-n). Treatment of this panel of cell lines with LatB to inhibit actin polymerization also recapitulated the reduction in telomerase recruitment observed in 293T cells (Fig. 1k, l, Supplementary Fig. 2o-p). Overall, these results provide evidence that the polymerization of actin in telomerase positive human cell lines is important for telomerase recruitment to telomeres, and that replication stress-induced recruitment requires ATR-mediated F-actin formation.
F-actin polymerization facilitates telomere extension by telomerase
Given that inhibition of nuclear F-actin polymerization affected the recruitment of telomerase to telomeres, we investigated whether this would also impact telomere synthesis by telomerase. To address this, we utilized an in situ telomere extension assay which employs a variant of hTR with a mutated template sequence, TSQ1-hTR (tolerated sequence 1), that results in addition of GTTGCG repeats to telomeres instead of the canonical TTAGGG56. We adapted this assay to detect TSQ1-positive telomeres in 293T cells via SABER FISH45 following overexpression of hTERT, dyskerin and TSQ1-hTR. This system only detects TSQ1-positive telomeres after transfection of the TSQ1-hTR variant, but not wild type (WT) hTR, and does not require endogenous hTR (Supplementary Fig. 3a-d).
Following TSQ1 telomerase overexpression, the number of TSQ1-positive telomeres significantly increased following short-term (30 or 60 min) induction of replication stress in a time-dependent manner (Fig. 2a, b). There was also an increased presence of telomerase at TSQ1-positive telomeres following replication stress (Fig. 2c).
a Representative images of 293T cells stained using SABER FISH probes against canonical telomeres (TTAGGG; pink), TSQ1 telomere sequence (GTTGCG; cyan) and hTR (yellow). Cells were transfected with TSQ1 telomerase (hTR, hTERT and dyskerin) 8 h prior to treatment with DMSO or 0.2 μM LatB for a subsequent 16 h. Cells were treated with DMSO or APH (1.5 μM) for 30 or 60 min immediately before fixation. S phase cells were identified with EdU (not shown). Co-localizations between hTR and TSQ1-positive telomeres are indicated by white arrows. Scale bars, 2 μm. Average number of (b) TSQ1-positive telomeres and (c) co-localizations between hTR and TSQ1-positive telomeres in 293T cells shown in (a). Mean and number of nuclei quantified per replicate are listed in Source Data. d Direct telomerase assay of endogenous telomerase purified from 293T cells treated with DMSO or 0.2 μM LatB for 16 h, followed by treatment with DMSO or 1.5 μM APH for 30 min. Asterisk indicates radiolabelled DNA loading control. e Normalized telomerase activity of telomerase purified from 293T cells shown in (d), relative to NTC. p values compared to NTC: 0.5592 (APH), 0.6169 (LatB), 0.7325 (LatB + APH). All bar graphs displayed as mean ± SEM; n = 3 independent biological replicates; One-way ANOVA with Šídák’s multiple comparisons test.
To investigate the role of F-actin in this process we treated cells with LatB and observed a slight reduction in the number of TSQ1-positive telomeres (with and without telomerase), although this was not statistically significant (Fig. 2a–c). However, when cells were treated with APH after inhibition of F-actin polymerization, there was no increase in TSQ1-positive telomeres, or telomerase at those telomeres. These results were not due to variable levels of telomerase expression, as the number of hTR foci did not change between conditions (Supplementary Fig. 3e).
To exclude the possibility that the changes in TSQ1-positive telomeres were a by-product of altered telomerase assembly or activity, we treated 293T cells with APH and/or LatB and purified the endogenous telomerase to perform a direct telomerase activity assay (Fig. 2d). Regardless of APH or LatB treatment, there was no change in the total activity of telomerase in these cells (Fig. 2d, e), indicating that the results observed in the TSQ1 assay were specifically due to changes to telomerase recruitment. Taken together, these data demonstrate that replication stress results in increased productive telomere extension in situ, which is governed by nuclear F-actin-mediated telomerase recruitment.
Telomerase recruitment is facilitated by known regulators of nuclear F-actin
Nuclear actin polymerization and function is controlled and regulated by a wide array of proteins (Fig. 3a). Monomeric actin is transported into the nucleus by importin 9; this transport is regulated by cofilin 157,58. Branching of both nuclear and cytoplasmic actin fibers is mediated by ARP2/3, which is regulated by WASP through mTOR43,59. Consistent with our previous work38, we observed that the nuclear F-actin network (visualized by either chromobody or exogenous mCherry-NLS-actin expression) includes fibers that branch at angles of ~70° (Supplementary Fig. 4a), supporting that ARP2/3 and its regulation through WASP are required for formation of a branched nuclear actin network which facilitates telomerase recruitment. F-actin depolymerization is necessary for fiber reorganization, and can be performed by cofilin 160,61, which is regulated by LIM kinase 1 (LIMK)62. Furthermore, myosin motor proteins aid F-actin function by transporting cargo and aid remodeling of the F-actin network63.
a Model of replication stress-induced polymerization of nuclear F-actin, highlighting proteins involved in nuclear actin import, regulation, polymerization, and function. Created in BioRender. Harman, A. (2025) https://BioRender.com/1h81xvg. b, c Representative SABER FISH images of 293T cells probed for hTR (yellow) and telomeres (pink) with DAPI (blue) staining. Cells were treated with DMSO (NTC), 0.2 μM INK128 (mTORi), 200 μM CK-666 (ARP2/3i), 5 μM wiskostatin (WASPi), 10 μM LIMKi-3 (LIMKi), 10 mM BDM (MYHi) or 10 nM BTS (MYHi #2) for 16 h. Cells were subsequently treated with DMSO or 1.5 μM APH for 30 min. d, e Average hTR/telomere co-localizations in 293T cells shown in (b) and (c). Significance above the columns for the inhibitor treated samples - APH is expressed relative to NTC - APH; significance for the inhibitor + APH samples is expressed relative to NTC + APH. f Representative SABER FISH images of 293T cells transfected with control, cofilin 1 (CFL1) or importin 9 (IPO9) siRNA, ± 30 min APH (1.5 μM) treatment. Cells were stained with DAPI (blue) and probed for hTR (yellow) and telomeres (pink). g Western blot of 293T cells transfected with control, CFL1 or IPO9 siRNA, probing for CFL1, IPO9 and actin as a control. h Telomerase presence at telomeres in 293T cells transfected with control, cofilin 1 (CFL1) or importin 9 (IPO9) siRNA, ± 30 min 1.5 μM APH treatment. Significance above the columns for each siRNA treated sample - APH is expressed relative to siControl - APH; siRNA + APH sample significance is expressed relative to siControl + APH. i SABER FISH images of HeLa, A549, HT1080 and HCT116 cells treated with DMSO (NTC) or 0.2 μM INK128 (mTORi) for 16 h, probing for hTR (yellow) and telomeres (pink) with DAPI staining (blue). j Normalized telomerase presence at telomeres from (i). For all microscopy images, scale bar = 2 μm. Co-localizations are indicated by white arrows. Mean and number of nuclei quantified per replicate are listed in Source Data. All bar graphs displayed as mean ± SEM; n = 3 independent biological replicates; One-way ANOVA with Šídák’s multiple comparisons test.
Given that these proteins are all established actin regulatory elements which have been demonstrated to function in nuclear F-actin polymerization in response to replication stress, or in the import of actin into the nucleus, we decided to confirm whether actin-mediated telomerase recruitment utilizes these known regulatory elements. 293T cells were treated with small molecule inhibitors of five key actin regulators: the mTOR inhibitor INK12864, the ARP2/3 inhibitor CK-66665, the WASP inhibitor wiskostatin66, the LIMK inhibitor LIMKi 367, and the myosin inhibitors BDM (2,3-butanedione monoxime)68 and BTS (N-benzyl-p-toluene sulfonamide)69. Chemical inhibition of these actin regulators resulted in decreased telomerase presence at telomeres (Fig. 3b–e). The decrease observed after LIMKi treatment was not statistically significant, but this is likely because it regulates cofilin, which has both positive and negative effects on nuclear actin polymerization; indeed, there was no significant difference in the proportion of nuclear F-actin-positive S phase cells after LIMKi treatment alone (Supplementary Fig. 4b). The cells were also treated with APH briefly to induce replicative stress; while inhibition of these regulatory elements did not impair Chk1 phosphorylation or cell cycle progression (Supplementary Fig. 4c-d), all inhibitors prevented the increase in telomerase recruitment induced by replication stress (Fig. 3d, e), and for LIMKi treatment there was a concomitant inhibition of the replication stress-induced increase in S phase nuclear F-actin (Supplementary Fig. 4b).
We next tested the role of two regulators of nuclear actin import by siRNA knockdown in 293T cells, specifically cofilin 1 (encoded by the gene CFL1) and importin 9 (IPO9). Knockdown of these proteins resulted in a decrease in telomerase recruitment, although this was not statistically significant (Fig. 3f–h, Supplementary Fig. 4e-g). For cofilin, this was likely because the reduction in F-actin-positive S-phase cells was also not significant (Supplementary Fig. 4h). However, cells treated with APH following knockdown retained a normal cell cycle profile (Supplementary Fig. 4i) and functional DDR (Supplementary Fig. 4j), but showed no significant increase in telomerase recruitment upon replication stress (Fig. 3h, Supplementary Fig. 4f), which for cofilin was paralleled by no significant increase in F-actin-positive S phase cells (Supplementary Fig. 4h). Overall, these data demonstrate that the increased telomerase recruitment induced by replication stress requires a functional regulatory system to promote nuclear import and polymerization of actin.
To establish the universality of this pathway, the selected panel of telomerase positive cell lines (HeLa, A549, HT1080, and HCT116) was treated with INK128 to assess its effect on telomerase recruitment. Following mTOR inhibition, all cell lines displayed decreased telomerase recruitment without major changes to cell cycle progression (Fig. 3i, j, Supplementary Fig. 4k-l). Overall, the results from these experiments indicate that the known nuclear actin regulatory network is important for facilitating telomerase localization to telomeres, both under endogenous conditions and following induction of exogenous replication stress.
Nuclear F-actin polymerization is required for telomerase recruitment
The role of nuclear F-actin in resolving replication stress38,41, in combination with our observation that knockdown of IPO9 impacted telomerase recruitment (Fig. 3h, Supplementary Fig. 4f), suggested that it is the nuclear pool of F-actin that specifically regulates telomerase recruitment, rather than an indirect effect of perturbations to cytoplasmic actin. To directly test this hypothesis, we utilized expression constructs encoding WT actin or a polymerization-deficient actin mutant (R62D) tagged with an NLS36,38. Both constructs, along with an empty vector as a control, were transfected into 293T cells, where only R62D actin disrupted the polymerization of nuclear F-actin, without impacting cytoplasmic F-actin (Fig. 4a, b, Supplementary Fig. 5a–c). Expression of exogenous WT actin did not impact telomerase recruitment under endogenous or stressed conditions; however, expression of the R62D mutant significantly decreased recruitment in both conditions (Fig. 4c, d). To further validate this result, expression of WT and R62D actin was then repeated with simultaneous siRNA knockdown of IPO9 to reduce the nuclear pool of endogenous actin (Supplementary Fig. 5d–f). IPO9 knockdown reduced telomerase recruitment in cells transfected with empty vector as expected; however, expression of exogenous WT actin was able to rescue this phenotype by replenishing the nuclear actin pool (Fig. 4e, f). Conversely, expression of the mutant R62D mutant significantly reduced telomerase recruitment, regardless of IPO9 knockdown. These data provide evidence that the regulation of telomerase recruitment specifically involves nuclear F-actin filaments.
a Super-resolution Airyscan microscopy of 293T cells transfected with vectors encoding WT or polymerization-deficient mutant (R62D) 3xNLS-actin, using empty vector as a control. Cells were co-transfected with a BFP- and NLS-tagged actin chromobody (nuclear-actin-ChB) to visualize nuclear F-actin (green). b Western blot of 293T cells transfected with empty vector (EV) or vector encoding WT or mutant (R62D) 3xNLS-actin, probing for actin and vinculin as a control. Exogenous actin is tagged with a 3×NLS. Both exogenous (NLS-actin) and endogenous (Endo. actin) are indicated by black arrows. c Representative images of 293T cells probed for hTR (yellow) and telomeres (pink) with DAPI (blue) staining following transfection with vectors encoding WT or mutant (R62D) 3xNLS-actin and treated with DMSO or 1.5 μM APH for 30 min. Empty vector (EV) was used as a control. d Average telomerase presence at telomeres in 293T cells transfected with empty vector (EV) or vector encoding WT or mutant (R62D) 3xNLS-actin, treated with DMSO or APH. Significance above the columns for actin transfected samples - APH is relative to EV - APH; significance for actin transfected + APH samples is relative to EV + APH. e Representative SABER FISH images of 293T cells probed for hTR (yellow) and telomeres (pink) with DAPI (blue) staining. Cells were transfected with empty vector (EV) or vectors encoding WT or mutant (R62D) 3xNLS-actin together with control or IPO9 siRNA. f Average hTR/telomere co-localizations in 293T cells transfected with vectors encoding WT or mutant (R62D) 3xNLS-actin, with or without IPO9 siRNA. Empty vector (EV) and control siRNA were used as controls. Significance above the columns for actin transfected samples - siIPO9 is relative to EV - siIPO9; significance for actin transfected + siIPO9 samples is relative to EV + siIPO9. For all microscopy images, scale bar = 2 μm. Co-localizations are indicated by white arrows. Mean and number of nuclei quantified per replicate are listed in Source Data. All bar graphs displayed as mean ± SEM; n = 3 independent biological replicates; One-way ANOVA with Šídák’s multiple comparisons test.
Telomerase recruitment to telomeres occurs in proximity to F-actin filaments
One of the emerging functions of nuclear F-actin in resolving genomic stress is the re-localization of stalled replication forks or sites of DNA damage to the nuclear periphery37,38,40. It has also been demonstrated that telomeres undergoing replicative stress are repositioned toward the nuclear periphery in an F-actin dependent manner38,42. Given this, one potential hypothesis for replication stress-mediated telomerase recruitment is that it occurs closer to the periphery following induction of replication stress and re-localization of telomeres. To investigate this, the nuclear periphery was first identified by DAPI staining of 293T cells. Individual hTR or telomere foci, as visualized by FISH, were then identified alongside co-localized hTR/telomere foci indicating telomerase recruitment (Fig. 5a). To account for changes in nucleus size, nuclei were divided into six equal segments, with foci scored according to which segment they were located within (Fig. 5b). There was no significant difference in localization between hTR or telomeres; however, telomerase interacting with telomeres was significantly further from the nuclear periphery than telomeres or hTR alone (Fig. 5c). This suggests that while the nuclear periphery serves as a site for DNA repair, it is not where telomerase recruitment predominately occurs under basal conditions. Short-term exposure to APH also failed to induce movement of telomeres towards the nuclear periphery (Supplementary Fig. 6a); instead telomeres moved slightly closer to the nuclear interior. While it remains possible that re-localization occurred but was not detected due to invaginations in the nuclear envelope, we favor the interpretation that telomere re-localization to the nuclear periphery requires prolonged exposure to replication stress38,42.
a Representative images showing telomere (pink), hTR (yellow) and co-localizing hTR/telomere (orange) foci, detected using CellProfiler. 293T nuclei were stained using DAPI (gray) to identify the nuclear periphery (white outline). b Schematic for segment-based separation of foci. Nuclei were identified using DAPI and separated into 6 equal segments; each focus was scored according to the segment it was located within. c Distribution of telomeres, hTR, or hTR/telomere co-localizations within the nuclei of 293T cells. Data are displayed as percentages of foci within each segment. Statistics were performed using χ2 tests. n = 3 independent biological replicates, representing foci from at least 30 nuclei per replicate. d Representative image of HeLa cells expressing nuclear-actin-ChB to visualize nuclear F-actin (green), with telomere (pink) and hTR (yellow) staining by SABER FISH, imaged via super-resolution Airyscan microscopy. e Line scan of telomere (pink) and hTR (yellow) signals to demonstrate co-localization along chromobody-marked nuclear F-actin (green). f Mean minimum distance of telomeres, hTR and hTR/telomere co-localizations from nuclear F-actin in HeLa cells treated ± 1.5 μM APH for 60 min to promote F-actin polymerization. n = 3 or 4 independent biological replicates, representing foci from at least 11 nuclei per replicate. Significance shown immediately above the columns for foci from APH treated cells are expressed relative to the same focus from cells without APH treatment. g Representative image of HeLa cells transfected with mCherry- and NLS-tagged actin (green), stained with DAPI (blue) and probed for telomeres (pink) and hTR (yellow) by FISH. Cells were imaged via super-resolution Airyscan microscopy. h Line scan of telomere (pink) and hTR (yellow) signals to demonstrate co-localization along mCherry-tagged nuclear F-actin (green). i Mean minimum distance of telomeres, hTR, and hTR/telomere co-localizations from mCherry-tagged nuclear F-actin. HeLa cells were treated with 1.5 μM APH for 60 min to promote F-actin polymerization. n = 3 independent biological replicates, representing foci from at least 36 nuclei per replicate. For all microscopy images, scale bar = 2 μm. Mean and number of nuclei/foci quantified per replicate are listed in Source Data. All bar graphs displayed as mean ± SEM; f and i analyzed by One-way ANOVA with Šídák’s multiple comparisons test.
We instead hypothesized that F-actin might serve as a direct site of interaction between telomeres and telomerase. To test this, FISH against hTR and telomeres was performed on HeLa cells expressing nuclear-actin-ChB, with or without brief treatment with APH to promote F-actin formation. Neither nuclear-actin-ChB expression nor APH treatment impacted the cell cycle progression of the BFP-positive cells (Supplementary Fig. 6b-c). Cells were visualized by confocal laser scanning microscopy using an Airyscan super-resolution detector (Carl Zeiss), where co-localized hTR/telomere foci were observed residing on F-actin (Fig. 5d, e). To quantify this, individual and co-localizing hTR/telomere foci were identified along with F-actin, and the minimum distance between each focus and the nearest actin fiber was then calculated (Supplementary Fig. 6d). While the mean distance of hTR and telomere foci from actin fibers were not significantly different, telomerase foci co-localizing with telomeres were on average significantly closer to F-actin than either unbound telomerase or telomeres in cells treated with or without APH (Fig. 5f). These results were due to more foci residing directly on or adjacent to F-actin, as well as fewer foci which were distant (>1 μm) from F-actin (Supplementary Fig. 6d-h). While APH treatment increases the number of F-actin-positive nuclei (Fig. 1e, f), it did not affect the mean distance of hTR and telomere foci from nuclear F-actin compared to untreated cells (Fig. 5f), indicating that exogenous replication stress increases the frequency of F-actin formation and telomerase recruitment events, but not their underlying mechanism.
While it has been demonstrated that the actin chromobody stains canonical, dynamic nuclear F-actin structures that can form and dissociate in the presence of the chromobody37,38,60, and we also see cells with few, short, or thin actin fibers (Supplementary Fig. 6i), we nevertheless validated this result by overexpressing mCherry and NLS-tagged actin36 and repeating this analysis. Overexpression of this NLS-tagged actin (without mCherry) alone does not promote telomerase recruitment (as would be expected following nuclear F-actin stabilization; Fig. 4c, d), and mCherry-tagged actin can form both long and short nuclear actin fibers (Supplementary Fig. 6j), which can also dissociate38. hTR/telomere co-localizations were again observed residing on nuclear F-actin (Fig. 5g, h). Although the minimum distance was larger than we observed with the chromobody, co-localizations were on average significantly closer to nuclear F-actin than foci which did not co-localize (Fig. 5i). The distribution of foci was also comparable (Supplementary Fig. 6k-m). Taken together, these results demonstrate that telomerase recruitment to telomeres occurs in proximity to nuclear F-actin, which likely serves as a direct site for their interaction.
Telomerase recruitment occurs via tethering of telomeres to F-actin
To explore the dynamics of telomerase recruitment further, live cell imaging was performed using CRISPR-modified HeLa cells expressing endogenous hTERT tagged with a FLAG-HaloTag and endogenous TRF2 tagged with the fluorescent protein HA-mEOS3.2, to allow visualization of telomerase and telomeres, respectively47. These cells were transfected with the nuclear-actin-ChB, synchronized by double thymidine block (Supplementary Fig. 7a), and released into S phase for 3–5 h prior to live cell imaging using super-resolution microscopy (Fig. 6a, Supplementary Movies 1 and 2). Synchronization by double thymidine block also causes replication stress and therefore promotes formation of nuclear F-actin filaments of varying lengths (Fig. 6a, Supplementary Fig. 7b). Telomerase is known to rapidly bind and dissociate from telomeres in S phase, while longer interactions with telomeres are rarer35,47. These longer interactions are known to represent successful recruitment of telomerase, leading to engagement of telomerase with the 3′ DNA overhang and productive telomere extension70. Given this, we assessed these long telomerase/telomere interactions to determine whether proximity to F-actin would help mediate this process. Stable interactions (> 30 s) between telomerase and telomeres (Fig. 6b) were classified based on where they occurred relative to F-actin; this included on or adjacent to a fiber, or in proximity to F-actin (~1 μm maximum distance), with all other events classified as distal (Fig. 6c). Stable interactions were also separately classified based on whether the initial interaction between telomerase and the telomere had already occurred prior to the start of the movie (Fig. 6d, “Previously recruited”), or if the recruitment event was observed occurring during the movie (i.e. at least one focus was observed prior to it co-localizing with its partner; Fig. 6d, “Observed recruitment”). Using these classifications, ~45% of observed recruitment events occurred directly on F-actin, with ~35% of the remaining events occurring adjacently (Fig. 6d). By comparison, previous recruitment events were imaged directly on or adjacent to F-actin only ~15% and ~25% of the time, respectively. The distribution of these events was significantly different, which suggests that recruitment may occur in proximity to F-actin, but this is not required for the continued interaction between telomerase and a telomere.
a Still image from live cell imaging experiments. Nuclear-actin-ChB to visualize nuclear F-actin (green) was transfected into CRISPR-modified HeLa cells expressing HA-mEOS3.2-tagged TRF2 (pink) and FLAG-HaloTag-tagged hTERT (visualized using JF-646-HaloTag ligand; yellow). Scale bar, 2 μm. b Time course of live cell imaging experiment showing hTERT (yellow) and TRF2 (pink) foci co-localizing on a nuclear F-actin fiber (green), over the indicated time in minutes. c Classification used for live cell imaging quantification. d Quantification of live cell imaging experiments, displaying distribution of foci with respect to F-actin. ‘Observed recruitment’ includes co-localizations where one or both of the hTERT or TRF2 foci were observed alone prior to recruitment. ‘Previously recruited’ includes co-localizations where both foci were first observed already co-localized. Quantified TRF2 and hTERT foci did not co-localize with the other. Statistics were performed using χ2 tests. n = 52 (observed recruitment), 51 (previously recruited), 247 (telomeres) and 36 (hTERT). Data were from 80 nuclei across 8 independent experiments. e Observed recruitment events (d; excluding distal events) classified according to which constituent focus was closest to nuclear F-actin (n = 49). f Examples of observed recruitment event classification in (e) where TRF2, hTERT or both foci are closest to a nuclear actin fiber. g Observed recruitment events (d; excluding distal events) classified according to which constituent focus arrived at nuclear F-actin first (n = 49). h Time course of live cell imaging experiment over the indicated time in minutes, showing telomere (TRF2; pink) focus mobility in relation to F-actin (green). Top panels display a mobile telomere which is off F-actin, while the bottom panels show a static telomere on F-actin. Scale bar, 2 μm. i Examples of cumulative mean squared displacement (MSD) of two separate TRF2 foci in relation to F-actin. MSD is plotted over time for a focus which is not on F-actin (gray) or one that remains associated with F-actin (green). j, k Mean MSD (μm2/s) of (j) telomere (TRF2) or (k) telomerase (hTERT) foci from live cell imaging experiments. MSD was calculated for each focus on a frame-by-frame basis, and each MSD value was classified as being “Off fiber” or “On fiber” based on the intensity of nuclear-actin-ChB signal within the focus region. hTERT foci were also classified as “Off TRF2” or “On TRF2” based on the intensity of TRF2 signal with the focus region. TRF2 data are displayed as mean MSD from 14 experiments (each experiment mean shown as a dot), representing 136 nuclei. n = 25,071 (Off fiber) and 3938 (On fiber) MSD values. hTERT data are displayed as mean MSD from 8 experiments (each experiment mean shown as a dot), representing 103 nuclei. n = 10,094 (Off TRF2/Off fiber), 1313 (Off TRF2/On fiber), 2025 (On TRF2/Off fiber) and 258 (On TRF2/On fiber) MSD values; analyzed using paired two-tailed t test for (j) or One-way ANOVA with Šídák’s multiple comparisons test for (k). Mean and number of nuclei/foci quantified per replicate are listed in Source Data.
It was also observed by live cell imaging that telomeres displayed a similar profile to observed recruitment events; a majority of telomeres which were never bound by telomerase were on or adjacent to F-actin (Fig. 6d). In contrast, telomerase foci which never interacted with a telomere were primarily located distally from F-actin. When the position of the telomere and telomerase foci relative to actin fibers was examined, in ~50% of observed recruitment events the telomere was located closer to F-actin than telomerase, while telomerase was closer only ~10% of the time (the remaining ~40% were equidistant) (Fig. 6e, f). We also determined which focus arrived at the actin filament first for these events, which revealed that in ~80% of observed recruitment events, the telomere was located at or near F-actin before telomerase (Fig. 6g). These data together suggest that under replication stress conditions, telomeres are the first to interact with actin filaments, which ultimately could facilitate the ability of telomerase to interact with those telomeres.
We observed that telomeres located on F-actin were less mobile than those distant from fibers (Fig. 6h, Supplementary Movie 3); this was quantified by measuring the mean squared displacement (MSD) of each individual telomere on a frame-by-frame basis and classifying each focus as “on” or “off” F-actin based on the intensity of nuclear-actin-ChB signal within the focus (examples of individual telomeres in Fig. 6i, Supplementary Fig. 7c-e). From this analysis, telomeres which were on F-actin were ~40% less mobile than other telomeres (Fig. 6j, Supplementary Fig. 7f). We subsequently performed the same analyses for individual hTERT foci but also determined whether each focus was “on” or “off” TRF2 at any given time point using the same approach as for F-actin. For hTERT foci which did not overlap with TRF2 there was a slight, but significant, increase in hTERT mobility when located on F-actin (Fig. 6k, Supplementary Fig. 7g). Conversely, hTERT foci interacting with TRF2 were less mobile on average, and this did not depend on actin fibers (Fig. 6k, Supplementary Fig. 7g-k), consistent with previous observations35,47 that stable telomerase recruitment to telomeres significantly reduces its movement. Furthermore, hTERT foci were overall more mobile than telomeres (Fig. 6j, k, Supplementary Fig. 7f-g), even when co-localizing with TRF2 foci on F-actin, which is consistent with the scanning behavior of telomerase at telomeres35. Overall, these data suggest that under replicative stress, telomeres are restrained by F-actin and that these telomeres may serve as a site for telomerase recruitment to facilitate telomere elongation.
Stalled replication forks promote telomerase recruitment at F-actin
While it is known that replication stress promotes telomerase recruitment11, the exact mechanism behind this process remains unclear. One possible explanation for this phenomenon is that replication fork stalling within a telomere triggers telomerase recruitment to that same telomere. To examine the hypothesis that telomerase recruitment occurs in proximity to stalled replication forks, PCNA (proliferating cell nuclear antigen) foci were used as a marker of DNA replication71. Immunofluorescence (IF) against PCNA was combined with SABER FISH for hTR and telomeres (Fig. 7a) in 293T cells in the presence or absence of a 30 min incubation with APH. While a basal level of co-localization between all three elements was observed in untreated cells, APH treatment resulted in ~4-fold more hTR-telomere co-localizations that overlapped with PCNA foci (Fig. 7b, Supplementary Fig. 8a-b). As expected, APH treatment also increased telomere-PCNA or hTR-telomere co-localizations ~twofold (Fig. 7c, d), indicative of increased telomeric replication stress and stress-induced telomerase recruitment, respectively. Interestingly, when hTR-telomere co-localizations were compared according to their overlap with PCNA, the proportion of recruitment events which coincided with PCNA approximately doubled following APH treatment (Fig. 7e, Supplementary Fig. 8c), supporting the hypothesis that replication fork stalling may promote telomerase recruitment.
a Representative image of 293T cells stained using SABER FISH probes against telomeres (pink) and hTR (yellow) with immunofluorescence against PCNA (cyan). S phase cells were identified with EdU (blue). Bottom row: zoomed images of the boxed co-localizations from the top row (excluding EdU). Co-localizations between hTR/telomere/PCNA are indicated by white arrows in the merge panel and circles in the zoomed panels. b–d Average co-localizations between (b) hTR/telomere/PCNA, (c) telomere/PCNA or (d) hTR/telomere in 293T cells treated ± 1.5 μM APH for 30 min. Data are displayed as mean ± SEM; n = 3 independent biological replicates; paired two-tailed t tests. e Distribution of hTR/telomere co-localizations (shown in d) which also co-localized with PCNA foci in 293T cells ± 1.5 μM APH. Statistics were performed by χ2 test. f Representative image of HeLa cells transfected with nuclear-actin-ChB to visualize nuclear F-actin (green). Cells were treated with 1.5 μM APH for 60 min to promote nuclear F-actin polymerization. Cells were stained via SABER IF-FISH to visualize telomeres (pink), hTR (yellow) and PCNA (cyan) before imaging using super-resolution Airyscan microscopy. Bottom row: zoomed images of the boxed co-localization from the top row. Co-localization between hTR/telomere/PCNA is indicated by white arrows in the merge panel and a circle in the zoomed panels. g Line scan of telomere (pink), hTR (yellow) and PCNA (cyan) signals to demonstrate co-localization along nuclear F-actin (green). h Mean minimum distance of hTR/telomere co-localizations (left) and telomeres (right) from nuclear F-actin. Foci which overlapped with PCNA were compared to those which did not overlap. Data displayed as mean ± SEM from 5 independent biological replicates; One-way ANOVA with Šídák’s multiple comparisons test. Cells were treated with 1.5 μM APH for 60 min to promote nuclear F-actin polymerization. For all microscopy images, scale bar = 2 μm. Mean and number of nuclei/foci quantified per replicate are listed in Source Data.
Given the convergence between replication, telomerase recruitment and actin polymerization, we also performed SABER IF-FISH against PCNA, hTR and telomeres in HeLa cells expressing the nuclear-actin-ChB to assess the spatial relationship between these factors. PCNA foci which reside along nuclear F-actin have been demonstrated to extensively co-localize with FANCD238, indicating they are undergoing replication stress and may represent stalled replication forks72. Cells were briefly treated with APH to induce F-actin polymerization and imaged by super-resolution microscopy, in which hTR/telomere/PCNA co-localizations were observed residing on F-actin (Fig. 7f, g). Using the same strategy as described above (Fig. 5), the minimum distance from F-actin for each focus alone and co-localizing with other foci was quantified. Both telomeres and hTR/telomere co-localizations which overlapped with PCNA were significantly closer to F-actin than those that did not overlap with PCNA (Fig. 7h, Supplementary Fig. 8d–f), while there was no significant difference in the distance from F-actin between PCNA, telomerase, or telomeres which did not co-localize with each other (Supplementary Fig. 8g-h). Together, these data indicate that F-actin directly facilitates telomerase recruitment to telomeres undergoing replication stress.
Discussion
In this study, we have demonstrated that nuclear F-actin is necessary for the recruitment of telomerase to telomeres in immortal human cell lines. This process is regulated by the kinases ATR and mTOR, and relies upon a network of known actin regulators to facilitate nuclear actin polymerization. Telomeres are tethered on nuclear F-actin, limiting their random movement around the nucleus, and thereby serve as a site for telomerase recruitment and ultimately productive telomere extension. Telomerase interactions with telomeres predominately occur near stalled telomeric replication forks which are tethered to nuclear actin fibers. Overall, this suggests a tightly regulated process which governs the occurrence and timing of DNA replication, the replication stress response, and telomere maintenance. Our data support a model wherein telomeric replication stress, a natural occurrence in the highly repetitive telomere sequence22,73,74, triggers the DDR and nuclear F-actin polymerization (Fig. 8a), ultimately resulting in telomere length maintenance. We hypothesize that this occurs via telomere tethering to nuclear F-actin (Fig. 8b), which subsequently promotes telomerase recruitment (Fig. 8c), ultimately resulting in telomere extension by telomerase (Fig. 8d).
During S phase, DNA replication proceeds through telomeres which can result in fork stalling, activating ATR to facilitate restart or repair. a ATR in turn activates mTOR to facilitate F-actin polymerization. b Stalled replication forks within telomeres are restrained on F-actin to facilitate both replication fork restart and (c) telomerase recruitment. Should replication fail to restart, then telomeres are re-localized to the nuclear periphery for repair. If replication is restarted (and subsequently completed), then (d) telomere-bound telomerase can extend telomeres, which does not need to occur in proximity to nuclear F-actin post-recruitment. However, should telomeric replication fail to restart, this would result in a DSB which would serve as a site for de novo telomere synthesis by telomerase. Created in BioRender. Harman, A. (2025) https://BioRender.com/6xg85cw.
One potential outcome of replication stress is the sudden loss of telomeres75,76,77; a mechanism cancer cells may use to overcome this is targeting telomerase to those telomeres to ensure that telomere length is maintained. This is supported by the finding that, in yeast and human cells, telomerase expression aids cell survival following replication stress78,79,80. Telomerase can interact with telomeres through short ‘probing’ or long static interactions, and while we did not examine ‘probing’ telomerase molecules, other work has shown these molecules possess diffusive searching movement35,47. This could suggest that telomerase does not itself track along actin filaments to locate a critically stressed or shortened telomere, but instead that telomeres are restrained to facilitate encounters with telomerase as it diffuses through the nucleus. This is supported by our findings that telomeres residing on F-actin are less mobile than other telomeres (Fig. 6h–j), while telomerase mobility is actually higher in proximity to F-actin (Fig. 6k). Furthermore, the long static interactions of telomerase with telomeres, which represent productive interactions with telomeric DNA70, predominately occur in proximity to F-actin (Fig. 6d) and are typically preceded by a telomere arriving at nuclear F-actin first (Fig. 6g).
While it was initially surprising that telomerase mobility on nuclear F-actin slightly increased in the absence of telomeres (Fig. 6k), we hypothesize that telomerase is incapable of stably binding to (or being bound by) F-actin, and its diffusive behavior35,47 results in its rapid interaction and dissociation from F-actin in search of telomeres. This is supported by our observation that telomerase mobility is dampened by stable interaction with telomeres, regardless of whether it occurs at nuclear F-actin (Fig. 6k). Furthermore, while we observed recruitment events occurring closer to F-actin, already established telomerase-telomere interactions displayed a distribution closer to that of telomerase foci alone (Fig. 6d). This suggests that while F-actin serves as a site for telomerase recruitment, it is likely not required for telomere elongation following successful recruitment. Also surprising was the fact that telomerase mobility, even when bound to telomeres, is ~2–3-fold higher than telomere mobility (Fig. 6j, k). However, we hypothesize this is due to the diffusion of telomerase along telomeres35 as it interacts with TPP1 at telomeres, likely until it is either recruited to the telomeric end or dissociates from the telomere.
Stalled replication forks move along actin filaments towards the nuclear periphery to facilitate the recovery of DNA replication after stalling38. Under mild long-term replicative stress conditions (24 h) telomeres are also re-localized to the nuclear periphery38,42, unlike the tethering we observed following short-term (30 min) APH treatment (Fig. 6j). Cells treated briefly with APH also show no difference in telomere distribution within the nucleus (Supplementary Fig. 6a). We therefore hypothesize that telomeres tether to F-actin rapidly in response to replication stress (Fig. 8b), which would both facilitate resolution by fork restart38 and help position telomerase at telomeres, ready to extend telomeres whose replication reaches the end. However, if replication stress cannot be resolved, the interaction between telomeres and nuclear F-actin could instead promote telomeres to re-locate to the nuclear periphery for repair38,42, taking telomerase with them. At least in yeast, stalled replication forks are not thought to re-localize to the nuclear periphery unless they collapse81,82; however, stalled forks within DNA capable of forming secondary structures can be re-localized82. Therefore, it is possible that, in human cells, particularly difficult substrates requiring repair, such as collapsed forks or stalled forks in difficult to replicate regions, are repositioned to facilitate this process42.
This re-localization could also result in fork restart and subsequent telomerase-mediated telomere extension. Alternatively, if actin-tethered forks instead collapse and result in DSBs, the presence of telomerase would allow rapid de novo telomere addition; this model is supported by the demonstration in yeast that telomerase is bound to DSBs which re-localize to the nuclear periphery83. This approach would allow the maintenance of telomere length and genome integrity by either resolving the replication stress, freeing telomeres for telomerase activity, or alternatively allowing telomerase to add de novo telomeres as a failsafe in the event that the DSB could not be repaired. This is supported by our observation of telomerase recruitment at nuclear F-actin after both short-term (Fig. 5d–i) and long-term (Fig. 6a–d) replication stress.
Although it is unknown exactly how stalled telomeric replication forks might restart, it potentially involves the CST complex, which is well established to maintain telomeres8 but also aids in global replication fork restart84 through its ability to protect stalled forks from degradation by the nuclease Mre1185. Another possible mechanism for overcoming stalled telomeric forks is the activation of dormant replication origins. While unperturbed telomere replication is thought to largely occur from subtelomeric regions, origins have been observed within telomeres86,87. Furthermore, fork stalling using APH can lead to TRF2-mediated dormant origin firing within telomeres88. This is a potential mechanism by which cells can ensure that telomeres are completely replicated if fork stalling occurs and cannot be restarted, which could in turn allow telomerase recruitment to the replicated telomere.
Our model for replication stress-mediated telomerase recruitment is supported by other studies, as long-term treatment of cells with APH to induce mild replication stress ultimately results in telomere lengthening22. Expression of mutant POT1 (POT1-ΔOB), which cannot bind to the single-stranded telomeric overhang, results in telomere-specific replication stress, which causes telomere re-localization to the nuclear periphery42, telomerase recruitment35 and telomere extension11,89. Furthermore, depletion of mouse POT1b (which regulates single-stranded telomere overhang length) results in initial telomere shortening accompanied by a telomeric DDR which results in ATR-dependent telomerase recruitment49,90. However, long term growth of POT1b-deficient tumors ultimately causes telomere hyper-elongation in a telomerase-dependent manner49.
While chemically-induced replication stress using APH heightens recruitment, the inhibition of ATR11, actin polymerization, or actin regulators (Figs. 1, 3) result in a significant reduction in telomerase presence at telomeres under both endogenous and stressed conditions, which supports the notion that these are not two separate pathways, but one pathway which is overstimulated by exogenous induction of replication stress. While nuclear actin fibers are not detectable in every individual cell in unperturbed conditions, ~10–20% of unperturbed S phase human cancer cells possess observable nuclear actin filaments (Fig. 1e, f and38,41). It is also possible that more transient or thinner nuclear actin filaments, below the limits of detection of the methods of visualization used here, may contribute to telomerase recruitment, which is supported by our observation that cells without visible nuclear F-actin can also have hTR/telomere co-localizations (Supplementary Fig. 9). Elevated replication stress is a hallmark of cancer cells91,92, so it can be concluded that cancer cells likely utilize the same mechanisms to bring telomerase to telomeres under endogenous and exogenous stress-induced conditions. This is further supported by our finding that telomerase foci at telomeres were located approximately the same distance from nuclear F-actin in the presence or absence of exogenous replication stress (Fig. 5f). One outstanding question from this and earlier work42 is the mechanism by which F-actin facilitates telomere movement. Inhibition of myosin decreased telomerase recruitment (Fig. 3e) and reduced movement of stalled replication forks38. This suggests that the motor function of myosins in actin filament movement or transport of cargo along actin fibers plays a direct role in the re-localization of stressed telomeres, although the mechanism of myosin involvement in this process remains unknown.
One aspect we did not explore in this work is the specific mechanism(s) by which the actin regulatory elements (Fig. 3a) function in the context of replication stress-induced nuclear F-actin formation and telomerase recruitment. Both importin and cofilin are involved in nuclear import of actin, and several proteins we examined (mTOR, WASP and LIMK) regulate downstream effectors of actin function. However, the specific role of the actin nucleator ARP2/3 in this process remains to be determined. ARP2/3 is involved in polymerization of branched F-actin93 and has been implicated in nuclear F-actin polymerization in response to DNA damage50. This is likely regulated by Arpin, an ARP2/3 inhibitory regulator, as its depletion promotes homologous DNA repair94. We observe a branched nuclear F-actin network (Supplementary Fig. 4a), which suggests ARP2/3-mediated polymerization of actin branches, although it has been reported that ARP2/3 can also facilitate linear F-actin polymerization95. It is therefore possible that ARP2/3 performs multiple functions in mediating telomerase recruitment via nuclear F-actin regulation, and may work in concert with other regulators or actin nucleators.
The factor(s) that specifically facilitate binding between stalled replication forks at telomeres and F-actin to allow telomere tethering to nuclear F-actin (Fig. 6j) have also not yet been identified. Both ARP2/3 and WASP associate with replication forks and facilitate the accumulation of RPA at stalled forks50, suggesting a more direct role in the DDR outside their canonical role in regulation of nuclear F-actin polymerization. While the latter study did not examine telomeres specifically, it is likely that this process also occurs at stalled telomeric forks to maintain their integrity, which may consequently result in telomere tethering to F-actin. Whether ARP2/3 and/or WASP facilitate telomere tethering, either directly or indirectly, remains to be elucidated.
Overall, our data reveal a mechanism by which telomerase responds to telomeric replication stress in order to maintain telomere integrity and length. We have shown that telomerase recruitment to telomeres requires ATR- and mTOR-mediated nuclear actin filament formation, and that stable, long-lived telomerase interactions with telomeres occur in the vicinity of these filaments, ultimately allowing for productive telomere synthesis. We hypothesize that stressed telomeres are restrained by F-actin, allowing scanning telomerase molecules to find these telomeres and extend their length following complete DNA replication (Fig. 8b–d). Facilitation of telomerase recruitment would also allow telomerase to rescue any stressed telomeres that could not restart and thereby resulted in a DSB. Overall, this work suggests that there is a tight balance between replication stress and telomere maintenance, where the natural stress generated from telomere replication is exploited to facilitate telomerase recruitment, while telomerase can also protect cells from critical replication stress and sudden telomere loss.
Methods
Cell lines
HEK293T (T. Adams, CSIRO), WI-38 VA13/2RA (American Type Culture Collection), A549 (American Type Culture Collection), HT1080 (American Type Culture Collection), HCT116 (G. Chenevix-Trench), HeLa-EM2-11ht (J. Schmidt, Michigan State University) and CRISPR-modified HeLa-EM2-11ht cells expressing FLAG-Halo-tagged hTERT and HA-mEOS3.2-tagged TRF247 were all grown in a humidified 37 °C incubator with 5% CO2. HCT116 cells were cultured in McCoy’s 5A medium, supplemented with 10% fetal bovine serum (FBS) and 2 mM L-glutamine. All other cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FBS. Cell Bank Australia validated all cell line identities by short-tandem-repeat profiling and tested all cells for mycoplasma.
Cell culture
Where necessary, cells were treated with chemical inhibitors of actin-associated proteins for 16 h prior to harvesting or sample preparation. APH (Sigma-Aldrich, A4487) treatment (1.5 μM) was performed for 30–60 min immediately prior to harvesting or sample preparation, while maintaining chemical inhibition of other proteins where appropriate. Dimethylsulfoxide (DMSO; Sigma-Aldrich, D2650) was used as a vehicle control for all non-treated controls. LatB (Cayman Chemical, 10010631), LatA (Calbiochem, 428021), INK128 (Cayman Chemical, 11811), CK-666 (Sigma-Aldrich, SML0006), Wiskostatin (Sigma-Aldrich, W2270), LIMKi 3 (Calbiochem, 435930), BDM (Sigma-Aldrich, B0753) and BTS (Calbiochem, 1576-37-0) were used in cell treatments at concentrations indicated in figure legends.
siRNA transfection
siRNA transfections were performed according to the manufacturer’s instructions. Briefly, for knockdown by individual siRNAs (Qiagen), cells were transfected with 30 pmol siRNA and 5.5 μl Lipofectamine RNAiMAX (Thermo Fisher Scientific) in 200 μl Opti-MEM. For knockdown using Dharmacon pooled siRNAs, cells were transfected with 50 pmol pooled siRNA and 4 μl of DharmaFECT 1 (Dharmacon) each added separately to 200 μl of Opti-MEM, before combining with 1600 μl DMEM + 10% FBS. Cells were grown for 48 h after transfection to allow for knockdown. siRNAs and catalog numbers are listed in Supplementary Table 1.
Plasmid transfection
The hTR-TSQ1 encoding plasmid was generated from pApex-3-dyskerin-U3-hTR96 by replacing the 5′ end of hTR with the TSQ1 variant sequence. Telomerase overexpression was performed by transfecting dyskerin-hTR plasmids (WT or TSQ1) with an hTERT encoding plasmid (pApex-3-hTERT-SV40-puro)96. 2 μg total plasmid (19:1 dyskerin-hTR:hTERT plasmids) was transfected with 3.75 μl Lipofectamine 3000 (Thermo Fisher Scientific) in 250 μl Opti-MEM. Cells were grown for 24–72 h after transfection to allow for telomerase expression.
Nuclear WT and R62D mutant actin expression vectors were generated from pmCherry-C1 actin-3×NLS P2A mCherry and pmCherry-C1 R62D actin-3×NLS P2A mCherry vectors (D. Mullins, University of California)36, respectively, by removing mCherry to enable hTR-telomere SABER FISH. BFP- and NLS-tagged actin chromobody (nuclear-actin-ChB) was generated from the NLS-GFP-actin chromobody vector (Chromotek) by replacing green fluorescent protein (GFP) with BFP. For expression of either exogenous actin (with or without mCherry) or the nuclear-actin-ChB, cells were transfected with 1–2.5 μg plasmid DNA and 3.75 μl Lipofectamine 3000 in 250 μl Opti-MEM. Cells were grown for 48 h after transfection to allow for expression.
Flow cytometry
Following treatment or transfection, cells were harvested with trypsin, pelleted at 500 g for 5 min and washed twice with phosphate buffered saline (PBS). Cells were resuspended in 1 ml PBS, before adding 5 ml of cold 70% ethanol dropwise while vortexing. Cells were left at 4 °C at least overnight to fix. The day before analysis, cells were repelleted and washed twice with PBS. Cells were resuspended in fresh propidium iodide (PI) solution (50 μg/ml PI, 0.5 μg/ml RNase A, 5% Triton X-100 in PBS) and left overnight at 4 °C to stain. Cells were then analyzed using an LSRII flow cytometer and FACSDiva software (Becton Dickinson). Cell cycle profiles were generated using FlowJo analysis software (Version 10.8). An example of the gating strategy is shown in Supplementary Fig. 10.
Fluorescence microscopy
All fixed samples (unless otherwise stated) were imaged by fluorescence microscopy using an AxioImager Z.2 microscope (Carl Zeiss), using a ×63/1.4NA oil-immersion objective, appropriate filters, an Axiocam 506 monochromatic camera (Carl Zeiss) and Zen Blue Pro v.2.3.69.01015 (Carl Zeiss). Samples were imaged with 13 Z-stacks at 0.25 μm intervals using consistent exposure times between all treatments and experiments. Pixel intensity histograms were adjusted equally across figure panels for presentation only.
SABER FISH for hTR and telomeres, with immunofluorescence
FISH against hTR and telomeric DNA, with or without IF, was adapted from existing hTR-telomere FISH11,44 and SABER (signal amplification by exchange reaction) FISH protocols45. SABER probes were extended by primer extension reaction (1X PBS, 10 mM MgSO4, 6 mM dNTPs [dATP, dCTP, dTTP], 8 U/μl Bst LF polymerase [New England Biolabs, M0275S], 2 μM target hairpin, 5 μM target probe) at 37 °C for 2 or 6 h, for telomere or hTR probes, respectively. Reactions were heat inactivated at 80 °C for 20 min, evaluated by agarose gel electrophoresis, and then stored at −20 °C until required. All probe and hairpin sequences are listed in Supplementary Table 2.
Cells were seeded on poly-L-lysine coated coverslips for 24 h prior to transfection, inhibitor treatment, or fixation. Cells were labeled with 20 μM EdU for 15–30 min immediately prior to fixation (with subsequent APH treatment where appropriate). Cells were washed once with PBS before permeabilization with 0.5% IGEPAL (in PBS) for 10 min, then fixed with 2% paraformaldehyde (PFA, in PBS) for 15 min. Cells were washed twice with MilliQ water, before further permeabilization with 1:1 methanol:acetone solution for 10 min. Cells were washed once with PBS, then EdU was labeled with AF488 using the Click-iT® Plus reaction kit (Thermo Fisher Scientific). Cells were then washed once with PBS before gradient ethanol dehydration (70%, 90% and 100% ethanol) for 2 min each and brief air drying. Probe mastermix was prepared by diluting extended hTR and telomere probes 1:100 in RNAse-free water, before denaturing at 90 °C for 2 min, followed by cooling on ice for 3 min. For each coverslip, 1 μl probe mastermix was added to 30 μl FISH hybridization buffer (2X SSC [0.3 M NaCl, 30 μM Tri-sodium citrate, pH 7], 10% dextran sulfate, 2 mM vanadyl-ribonucleoside complex, 0.02% bovine serum albumin (BSA), 1 μg/μl E. coli tRNA, 50% deionized formamide) before pipetting onto slides and covering with coverslips containing cells. Slides were then heated at 80 °C for 5 min before hybridizing overnight at 37 °C in a humidified chamber. Coverslips were then washed twice with FISH wash 1 (2× SSC, 50% formamide, 0.1% sodium dodecyl sulfate [SDS]) for 30 min, three times with FISH wash 2 (4× SSC, 0.1% Tween-20) for 5 min, and once with PBS briefly before allowing to air dry. Coverslips were placed onto slides containing imager probes (1 μM in PBS) and incubated in a humidified chamber at 37 °C for 1–1.5 h. Coverslips were washed with pre-warmed (37 °C) PBS for 15 min, twice for 5 min, and then rinsed twice with MilliQ water before air drying. Coverslips were mounted with Prolong Gold with DAPI and left overnight to cure.
For IF-FISH, the same protocol was followed until the EdU labeling step; EdU was instead labeled with AF350. Following this, the coverslips were blocked with ABDIL buffer (20 mM Tris [pH 7.5], 2% BSA, 0.2% fish skin gelatin, 120 mM NaCl, 0.1% Triton X-100, 0.1% sodium azide) in a humidified chamber at room temperature (RT) for 1 h. Anti-PCNA antibody (Cell Signaling Technology, 2586T) was then diluted in ABDIL (1:1000) and added to coverslips, which were incubated overnight in a humidified chamber at 4 °C. Coverslips were washed with PBS for 5 min three times, before incubating with Anti-mouse Alexa Fluor 488 (Thermo Fisher Scientific, A-11001) secondary antibody diluted in ABDIL (1:2000) for 1 h at RT in a humidified chamber. Coverslips were washed three times with PBS for 5 min, re-fixed with 2% PFA for 15 min, washed twice with MilliQ water, before continuing with gradient ethanol dehydration and the remaining steps outlined above. Finally, coverslips were mounted with Prolong Gold.
TSQ1, telomere and hTR SABER FISH
SABER probe extension reactions were performed as described above, except that TSQ1 probes and SABER branches were extended for 12 h at 37 °C. For TSQ1 SABER FISH, unextended telomere probes were used instead of extended probes, while the hTR probe was unchanged. All probe and hairpin sequences are listed in Supplementary Table 2.
Cells were seeded on poly-L-lysine coated coverslips for 24 h prior to transfection with TSQ1-hTR-DKC1 and hTERT expression plasmids. Samples were left to recover for 8 h before LatB treatment, at which point the cells were left for 16 h. Cells were then incubated with 20 μM EdU for 30 min, following which cells were treated with 1.5 μM APH for 30–60 min. Cells were fixed and permeabilized as above, before EdU was labeled with AF350 and samples were ethanol dehydrated. Probe mastermix was prepared by diluting unextended telomere probes and extended TSQ1 and hTR probes 1:100 in RNAse-free water, before denaturing at 90 °C for 2 min, followed by cooling on ice for 3 min. For each coverslip, 1 μl probe mastermix was added to 30 μl FISH hybridization buffer before pipetting onto slides and covering with coverslips containing cells. Slides were then heated at 80 °C for 5 min before hybridizing overnight at 37 °C in a humidified chamber.
Coverslips were washed four times with 60 °C SSCT (2X SSC, 0.1 % Tween-20) for 5 min, twice with RT SSCT for 2 min, then briefly with PBS before allowing to air dry. Extended secondary SABER branch probe (for TSQ1) was diluted 1:100 in RNAse-free water, before denaturing at 90 °C for 2 min, followed by cooling on ice for 3 min. For each coverslip, 1 μl probe mastermix was added to 30 μl FISH branch hybridization buffer (2X SSC, 10% dextran sulfate, 30% deionized formamide) before pipetting onto slides and covering with coverslips and hybridizing in a humidified chamber at 37 °C for 1.5 h. Coverslips were then washed as per above (SSCT/SSCT/PBS) before drying. The tertiary SABER branch probe was prepared and hybridized as per the secondary, except that hybridization was at 30 °C for 60 min. Coverslips were washed again as above, except that 55 °C SSCT was used instead of 60 °C SSCT, before drying. The quaternary SABER branch probe was prepared and hybridized as per the tertiary probe. Coverslips were washed again as above using 55 °C and RT SSCT, before being held overnight at 4 °C in PBS.
The following day, PBS was removed before air drying the coverslips. Coverslips were then placed onto slides containing imager probes (1 μM in PBS) and incubated in a humidified chamber at 37 °C for 1–1.5 h. Coverslips were washed with pre-warmed (37 °C) PBS for 15 min, twice for 5 min, and then rinsed twice with MilliQ water before air drying. Coverslips were mounted with Prolong Gold and left overnight to cure.
Nuclear F-actin imaging, with FISH/IF-FISH
Cells were seeded on poly-L-lysine coated coverslips for 24 h prior to transfection with nuclear-actin-ChB or mCherry-tagged-NLS-actin. Samples were subsequently transfected with siRNAs or treated with chemical inhibitors where appropriate. For S phase identification, cells were incubated with 20 μM EdU for 30 min before fixation with ice-cold 4% PFA for 15 min, washing with PBS, and permeabilization with 0.5% Triton X-100 (in PBS) for 10 min. EdU was then labeled using either AF594 (for nuclear-actin-ChB samples) or AF488 (for mCherry-actin samples), before ethanol dehydration, drying and then mounting with Prolong Gold (with DAPI for mCherry-actin samples). Samples were imaged by fluorescence microscopy (described above).
For FISH/IF-FISH, cells were treated with 1.5 μM APH for 1 h to promote F-actin formation immediately before fixing with ice-cold 4% PFA for 15 min. Cells were then washed with PBS, permeabilized with 0.5% Triton X-100 (in PBS) for 10 min and washed with PBS before ethanol dehydrating. Cells were then prepared for FISH or IF-FISH as above, excluding EdU steps, and mounted with Prolong Gold before being left to cure overnight. For samples transfected with mCherry-actin, the telomere SABER FISH imager was changed from Cy3 to FAM to allow visualization alongside mCherry. Samples were imaged by super-resolution microscopy (described below).
Phalloidin staining
Cells were seeded on poly-L-lysine coated coverslips for 24 h prior to staining. Cells were washed once with PBS, then fixed with ice-cold 4% PFA for 15 min, washed with PBS, permeabilized with 0.5% Triton X-100 (in PBS) for 10 min and washed with PBS. Cells were stained using phalloidin (fluorescein isothiocyanate labeled), diluted to 2% in PBS, overnight at 4 °C in a humidified chamber. Coverslips were then washed four times with PBS for 5 min each, before gradient ethanol dehydration (70%, 90% and 100% for 2 min each), air drying, and then mounting with Prolong Gold with DAPI. Samples were imaged by super-resolution microscopy (described below).
Immunopurification of telomerase
Cells (1–2 × 107) were harvested with trypsin, pelleted at 500 g for 5 min in a LoBind microcentrifuge tube (Eppendorf) and washed once with PBS before flash-freezing in liquid nitrogen. All subsequent steps were performed at 4 °C unless otherwise stated. Cells were lysed on ice using 1 ml of cold Buffer A (20 mM HEPES-KOH [pH 8], 300 mM KCl, 2 mM MgCl2, 0.1% Triton X-100, 10% glycerol) supplemented with 1 mM phenylmethanesulfonyl fluoride (PMSF) before rotating for 1 h. Cell debris was removed by centrifugation at 16,000 g for 30 min. hTERT antibody97 (20 μg) was added to cell lysate before rotating for 30 min, following which 20 μl of Protein G agarose beads was added and rotated for 1 h. Agarose beads were collected in a micro-spin column (GE Healthcare) by vacuum suction and washed with 5 ml of cold Buffer A. Columns were transferred to a microcentrifuge tube and centrifuged at 2000 × g for 10 s to remove residual buffer. The remaining steps were performed at RT.
The column was plugged before addition of 100 μl of Buffer A, supplemented with 1 mM dithiothreitol (DTT) and 2.5 μl of 1 mM hTERT peptide antigen solution (ARPAEEATSLEGALSGTRH). The solution was rotated for 30 min to allow dissociation of telomerase from the agarose beads; eluate was collected in a LoBind tube by centrifuging at 2000 × g for 10 s.
Direct telomerase activity assay
Telomerase extension reactions were performed in LoBind microcentrifuge tubes; 86 μl of telomerase containing eluate was used for a 100 μl reaction containing 50 mM KCl (300 mM final concentration including KCl from eluate), 1 mM dNTPs (dATP, dGTP, dTTP), 10 mM DTT, 50 μCi α−32P-dGTP, and 1 μM biotinylated telomeric DNA substrate (biotin- CTAGACCTGTCATCA(TTAGGG)3). Reaction was incubated overnight at 37 °C before stopping the reaction with 50 μl of stop mix (20 mM EDTA, 250 cpm 32P-end-labeled loading control [32P-CTAGACCTGTCATCACTAGACCTGTCATCA-biotin]) and briefly vortexing.
Dynabead M-280 Strepavidin suspension (20 μl) was added to a fresh LoBind tube, which was then placed onto a DynaMag-2 magnet to pellet beads before removing supernatant. The tube was removed from the magnet before washing beads with 200 μl of bind/wash buffer (10 mM Tris-Cl [pH 7.5], 1 mM EDTA, 2 M NaCl), brief vortexing, a pulse-spin, before placing the tube on the column again for 30 s before removing the supernatant. Bind/wash buffer (100 μl) was added to the beads, before addition of the telomerase extension reaction and mixing by rotation at RT for 1 h. Beads were washed as above with 200 μl of bind/wash buffer three times, before washing once in 200 μl TE buffer and then resuspending beads in 10 μl elution buffer (81% deionized formamide, 0.9 x TBE, 0.009% bromophenol blue, 0.009% xylene cyanol, 500 μM D-biotin). The beads were heated at 80 °C for 10 min before placing on the magnet and collecting the supernatant. Samples were run on a 10% polyacrylamide/ 8 M urea sequencing gel using 1 x TBE (90 mM tris base, 90 mM boric acid, 1 mM EDTA) for ~75 min. The gel was then transferred to thin Whatman filter paper, covered with plastic wrap, dried at 70 °C for 1 h and then imaged by phosphorimaging using a Typhoon FLA 9500 (Amersham).
Preparation of nuclear and cytoplasmic extracts
Cells were washed with PBS, harvested with trypsin, pelleted at 500 g for 5 min and washed once with PBS. 1 × 106 cells were aliquoted into a fresh microcentrifuge tube for each sample, pelleted and then resuspended in ~20 volumes (200 μl) of hypotonic buffer (10 mM HEPES-KOH [pH 8], 10 mM KCl, 1 mM MgCl2, 1 mM DTT). Cells were placed on ice for 5-10 min to swell, as observed by microscopy, before adding 1/20 volume of 10% Triton X-100 to lyse cell membranes. Tubes were mixed by inversion and kept on ice to observe cell membrane lysis by microscopy. Nuclei were then pelleted at 1000 × g for 1 min at 4 °C. The cytoplasmic fraction was collected in a separate tube, and the nuclei were washed twice with 1 ml hypotonic buffer. Fractionation purity was determined by western blot using vinculin and histone H2A as cytoplasmic and nuclear markers, respectively. 4× cell equivalents of nuclei were loaded relative to cytoplasmic fractions for all blots.
Western blot
Cells were harvested with trypsin and washed with PBS. Cell lysates were prepared at 1 × 104 cells/μl in 4× LDS (106 mM Tris-HCl, 141 mM Tris-base, 2% lithium dodecyl sulfate, 10% glycerol, 0.22 mM Brilliant Blue G250, 0.175 mM Phenol Red) supplemented with 50 mM DTT and 2% Benzonase for 15 min at RT. Lysates were denatured at 68 °C for 10 min, before loading 7 × 104 cell equivalents onto Nu-PAGE 4-12% gradient Bis-Tris gels and electrophoresed in MES SDS running buffer at 150 V for 70 min. Proteins were transferred to Amersham 0.45 μm nitrocellulose (GE Healthcare) in transfer buffer (25 mM Tris-Base, 192 mM glycine, 10% methanol) at 30 V overnight at 4 °C. Membranes were blocked in 5% skim milk powder or BSA in TBST (15 mM Tris-HCl, 4.6 mM Tris-Base, 137 mM NaCl, 0.1% Tween-20) for 1 h before incubation with gentle rotation overnight at 4 °C with primary antibodies diluted in either 5% skim milk powder or BSA in TBST. Antibodies used were anti-Chk1 (1:1000, Cell Signaling Technology, 2360S), anti-Chk1 (p345) (1:1000, Cell Signaling Technology, 2348S), anti-Vinculin (1:10000, Sigma-Aldrich, V9131), anti-Cofilin1 (1:1000, Cell Signaling Technology, 5175S), anti-Importin9 (1:1000, Thermo Fisher Scientific, PA141395), anti-Actin (1:2500, Sigma-Aldrich, A5441), anti-p70 (1:1000, Cell Signaling Technology, 2708S), anti-p70 (T389) (1:1000, Cell Signaling Technology, 9234S), anti-Histone 2 A (1:1000, Cell Signaling Technology, 12349S), anti-ATR (1:1000, Cell Signaling Technology, 2790S) and anti-ATRIP (1:1000, Thermo Fisher Scientific, PA1-519). Membranes were washed three times for 10 min with TBST, before probing for 1 h at RT with secondary antibodies (anti-mouse HRP, Dako, P0447; anti-rabbit HRP, Dako, P0448) diluted 1:2000 in 5% skim milk powder in TBST. Membranes were washed three times for 10 min with TBST, before developing signal using either Amersham ECL or ECL prime detection reagent. Blots were imaged using a BioRad ChemiDoc™ MP imaging system.
Live cell imaging
HeLa cells expressing FLAG-Halo-tagged hTERT and HA-mEOS3.2-tagged TRF247 were seeded on sterile 35 mm FluoroDish plates (Coherent Scientific) with cover glass bottoms. Cells were allowed to settle overnight before transfecting with nuclear-actin-ChB. Cells were left for 8 h and then synchronized by addition of 2 mM thymidine for 15 h, releasing for 9 h, and addition of 2 mM thymidine for 16 h. Cells were released into S phase with fresh DMEM ( + 10% FBS) for 3–5 h before labeling FLAG-HaloTag-hTERT. HaloTag ligand (Promega, P6711) was prepared by coupling with Janelia Fluor 646 (Tocris, 6148)98 following the ligand manufacturer’s instructions, followed by purification using liquid chromatography/mass spectrometry (LC/MS) on a C18 column (3.5 µm, 21.2 mm×150 mm) with a 3-90% acetonitrile/water gradient at 2 mL/min and detection at 656 nm. JF-646-Halo ligand (50 nM) in DMEM was added to cells for 1–5 min, before washing the cells three times with DMEM and returning cells to the incubator in DMEM ( + 10% FBS) for 15 min. Cells were washed 3 times with DMEM, before addition of DMEM without phenol red (+ 10% FBS) prior to imaging. Cells were imaged by super-resolution microscopy (described below).
Super-resolution microscopy
Super-resolution imaging was performed using an LSM 880 AxioObserver confocal laser scanning fluorescent microscope (Carl Zeiss) fitted with a super-resolution Airyscan detector using a Plan-Apochromat ×63/1.4NA M27 oil-immersion objective and ZEN Black 2.3 Pro v14.0.20.201 (Carl Zeiss). All samples/cells were imaged using ‘Resolution versus sensitivity’ mode, appropriate filter sets, unidirectional scanning, 1×1 binning and frame averaging (4 frames for fixed samples, 2 frames for live cell imaging).
Fixed samples were imaged using 405 nm (0.2% excitation power, 790 detector gain), 488 nm (1.16% excitation power, 790 detector gain), 561 nm (0.5% excitation power, 913 detector gain), and 633 nm (2.66% excitation power, 860 detector gain) lasers. Live cells were imaged at 37 °C, 20% O2 and 5% CO2, using 405 nm (0.2% excitation power, 790 detector gain), 561 nm (0.5% excitation power, 913 detector gain) and 633 nm (5% excitation power, 903 detector gain) lasers. Cells were briefly (10–20 s) exposed to the 405 nm laser immediately before imaging to convert HA-mEOS3.2-TRF2 from green to red wavelength. Imaging frame time was ~0.5 s per channel, with all channels imaged consecutively for 250–1000 frames.
Quantification and statistical analysis
Automated FISH/IF-FISH image analysis
Microscopy images were converted into extended projections of Z-stacks using ZEN blue desk software v2.3 (Carl Zeiss) and exported as TIFF files for analysis using CellProfiler v2.2.099 or v4.2.6100. Individual nuclei and foci were identified using intensity-based thresholding strategies. Individual foci were considered overlapping with another focus if at least 25% of the first focus was contained within the second. Triple co-localization events (hTR and telomere with TSQ1 or PCNA) were counted if 25% of an hTR focus was within a telomere focus, and if either also overlapped (any amount) with a TSQ1 or PCNA focus. At least 100 S phase cells were quantified per treatment, with each treatment performed in triplicate.
Rotation control for FISH co-localization
CellProfiler was used to identify nuclei based on DAPI staining; nuclear masks were then exported as binary images for identification with a custom Python script. Each nucleus was identified, and the largest symmetrical circle or square completely contained within the mask was determined. Each circle and square was used to rotate that region of the original telomere channel image 90°. Each image was exported and analyzed, alongside the unmodified telomere channel, using the same CellProfiler strategy as described above.
Measurement of minimum distance to nuclear periphery or F-actin
Measurement of the minimum distance to the nuclear periphery or F-actin was performed using custom scripts for Fiji v1.53q101. Individual nuclei, foci, and co-localizations were identified using CellProfiler v2.2.0 as above, and all identified objects were exported as binary images for import into Fiji. F-actin was detected using the FilamentDetector plugin (v2.0.0) for Fiji. For each focus/co-localization, the distance between the centroid and each point along the nuclear periphery or the nuclear F-actin network was calculated, with the lowest value returned. For measuring the minimum distance to the nuclear periphery, each nucleus was separated into six equal segments based on its radius.
Classification of live cell imaging recruitment events
Prior to analysis, live cell imaging data was processed using a custom Fiji script to improve foci clarity. CZI files were imported into Fiji, and each channel was separated for frame averaging and processing. Images were averaged on a 10-frame basis, before background subtraction (2 pixel rolling ball radius by sliding paraboloid) and Gaussian blur (sigma 1). Channels were merged and exported as TIFF files for analysis in Imaris software v9.9.1. Recruitment events were classified manually where hTERT and TRF2 foci overlapped at least 25% for a minimum of 2 consecutive averaged frames (~30 s). Recruitment events were classified as ‘observed’ only if one of the two foci was observed in at least the two previous consecutive averaged frames prior to recruitment; all other events were classified as ‘previously recruited’. Recruitment events and individual hTERT or TRF2 foci were sub-classified based on the distance to a visible actin fiber. Foci and events were classified as ‘on fiber’ if one (or both) focus was at least ~10% overlapping with F-actin. Anything under ~10%, with no observable gap between focus and F-actin was classified as ‘adjacent’. Foci/co-localizations within ~1.5 μm were classified as ‘proximal’, with everything else classified as ‘distal’.
MSD analysis
Telomere tracking was performed using live cell imaging data without any processing; hTERT tracking was performed on frame-averaged imaging data (as described above). Data were imported into Imaris v9.9.1, where the nuclei were identified using the ‘Surfaces’ function and the nuclear-actin-ChB channel. To correct for cell shifting or migration, we utilized previously written code for MATLAB R2023b to ‘register’ and align nuclei consistently within the frame38. The registration operation from MATLAB was returned to Imaris, which was completed using the ‘Align image without interpolation’ function. Following registration, foci were identified using the ‘Spots’ function in Imaris (0.5 μm focus diameter, >20 quality for TRF2 foci, >10 quality for hTERT foci). Foci were then tracked using ‘Autoregressive Motion’ tracking. Tracks were generated based on ~0.5 μm maximal distance movement between frames, 0 gaps within each track, and a minimum track length of 15–30 s for TRF2 foci, and 60 s for hTERT foci. The MSD was calculated for each timepoint (except T0) within each track and classified based on the intensity of nuclear-actin-ChB signal within the focal mask; hTERT foci were also classified based on the intensity of TRF2 signal within the focal mask. All timepoints from cells within an experiment were pooled together to generate a single average MSD value per classification (“On fiber” or “Off fiber”).
Statistics and reproducibility
Statistical analyses were performed using GraphPad Prism 10.5.0. One-way ANOVA with Šídák’s multiple comparisons test was used for all experiments unless otherwise noted. Experiments with only two sample conditions (Fig. 1e, f, Fig. 6j, Supplementary Fig. 7f, Fig. 7b–d, Supplementary Fig. 8a-b, e-f) were analyzed using two-tailed t tests. Segment or percentage-based data (Fig. 5c, Supplementary Fig. 6a, e, g, m, Fig. 6d, Fig. 7e, Supplementary Fig. 8h) were analyzed using χ2 tests.
The number of independent biological replicates of each experiment are provided in the figure legends and source data. In most cases n = 3 independent biological replicates, with at least 100 cells in each experiment, were used for quantitation. No data were excluded from analysis. These numbers were selected based on literature within the field, including our previously published work. Representative experiments were analyzed using WebPower102 to determine statistical power (Supplementary Table 3). All representative blots and images shown in the figures were obtained from at least three independent experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files. Source data are provided with this paper.
Code availability
Custom scripts are available at GitHub (https://github.com/ash-harman/Harman_2025_Nature_Comms).
References
Nassour, J., Schmidt, T. T. & Karlseder, J. Telomeres and cancer: Resolving the paradox. Annu Rev. Cancer Biol. 5, 59–77 (2021).
de Lange, T. Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev. 19, 2100–2110 (2005).
Smith, E. M., Pendlebury, D. F. & Nandakumar, J. Structural biology of telomeres and telomerase. Cell. Mol. Life Sci. 77, 61–79 (2020).
Roake, C. M. & Artandi, S. E. Regulation of human telomerase in homeostasis and disease. Nat. Rev. Mol. Cell Biol. 21, 384–397 (2020).
Fu, D. & Collins, K. Purification of human telomerase complexes identifies factors involved in telomerase biogenesis and telomere length regulation. Mol. Cell 28, 773–785 (2007).
Lue, N. F. & Autexier, C. Orchestrating nucleic acid-protein interactions at chromosome ends: telomerase mechanisms come into focus. Nat. Struct. Mol. Biol. 30, 878–890 (2023).
Cohen, S. B. et al. Protein composition of catalytically active human telomerase from immortal cells. Science 315, 1850–1853 (2007).
Lim, C. J. & Cech, T. R. Shaping human telomeres: from shelterin and CST complexes to telomeric chromatin organization. Nat. Rev. Mol. Cell Biol. 22, 283–298 (2021).
Tomlinson, R. L., Ziegler, T. D., Supakorndej, T., Terns, R. M. & Terns, M. P. Cell cycle-regulated trafficking of human telomerase to telomeres. Mol. Biol. Cell 17, 955–965 (2006).
Jady, B. E., Richard, P., Bertrand, E. & Kiss, T. Cell cycle-dependent recruitment of telomerase RNA and Cajal bodies to human telomeres. Mol. Biol. Cell 17, 944–954 (2006).
Tong, A. S. et al. ATM and ATR signaling regulate the recruitment of human telomerase to telomeres. Cell Rep. 13, 1633–1646 (2015).
Smogorzewska, A. et al. Control of human telomere length by TRF1 and TRF2. Mol. Cell Biol. 20, 1659–1668 (2000).
Nandakumar, J. et al. The TEL patch of telomere protein TPP1 mediates telomerase recruitment and processivity. Nature 492, 285–289 (2012).
Abreu, E. et al. TIN2-tethered TPP1 recruits human telomerase to telomeres in vivo. Mol. Cell Biol. 30, 2971–2982 (2010).
Latrick, C. M. & Cech, T. R. POT1-TPP1 enhances telomerase processivity by slowing primer dissociation and aiding translocation. EMBO J. 29, 924–933 (2010).
Wang, F. et al. The POT1-TPP1 telomere complex is a telomerase processivity factor. Nature 445, 506–510 (2007).
d’Adda di Fagagna, F. et al. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426, 194–198 (2003).
Celli, G. B. & de Lange, T. DNA processing is not required for ATM-mediated telomere damage response after TRF2 deletion. Nat. Cell Biol. 7, 712–718 (2005).
Denchi, E. L. & de Lange, T. Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 448, 1068–1071 (2007).
Guo, X. et al. Dysfunctional telomeres activate an ATM-ATR-dependent DNA damage response to suppress tumorigenesis. EMBO J. 26, 4709–4719 (2007).
Karlseder, J. et al. The telomeric protein TRF2 binds the ATM kinase and can inhibit the ATM-dependent DNA damage response. PLoS Biol. 2, E240 (2004).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Arneric, M. & Lingner, J. Tel1 kinase and subtelomere-bound Tbf1 mediate preferential elongation of short telomeres by telomerase in yeast. EMBO Rep. 8, 1080–1085 (2007).
Sabourin, M., Tuzon, C. T. & Zakian, V. A. Telomerase and Tel1p preferentially associate with short telomeres in S. cerevisiae. Mol. Cell 27, 550–561 (2007).
Hector, R. E. et al. Tel1p preferentially associates with short telomeres to stimulate their elongation. Mol. Cell 27, 851–858 (2007).
Bianchi, A. & Shore, D. Increased association of telomerase with short telomeres in yeast. Genes Dev. 21, 1726–1730 (2007).
Goudsouzian, L. K., Tuzon, C. T. & Zakian, V. A. S. cerevisiae Tel1p and Mre11p are required for normal levels of Est1p and Est2p telomere association. Mol. Cell 24, 603–610 (2006).
Moser, B. A., Chang, Y. T., Kosti, J. & Nakamura, T. M. Tel1ATM and Rad3ATR kinases promote Ccq1-Est1 interaction to maintain telomeres in fission yeast. Nat. Struct. Mol. Biol. 18, 1408–1413 (2011).
Moser, B. A., Subramanian, L., Khair, L., Chang, Y. T. & Nakamura, T. M. Fission yeast Tel1(ATM) and Rad3(ATR) promote telomere protection and telomerase recruitment. PLoS Genet 5, e1000622 (2009).
Yamazaki, H., Tarumoto, Y. & Ishikawa, F. Tel1(ATM) and Rad3(ATR) phosphorylate the telomere protein Ccq1 to recruit telomerase and elongate telomeres in fission yeast. Genes Dev. 26, 241–246 (2012).
Metcalfe, J. A. et al. Accelerated telomere shortening in ataxia telangiectasia. Nat. Genet 13, 350–353 (1996).
Pennarun, G. et al. ATR contributes to telomere maintenance in human cells. Nucleic acids Res. 38, 2955–2963 (2010).
Wu, Y., Xiao, S. & Zhu, X. D. MRE11-RAD50-NBS1 and ATM function as co-mediators of TRF1 in telomere length control. Nat. Struct. Mol. Biol. 14, 832–840 (2007).
Lee, S. S., Bohrson, C., Pike, A. M., Wheelan, S. J. & Greider, C. W. ATM kinase is required for telomere elongation in mouse and human cells. Cell Rep. 13, 1623–1632 (2015).
Laprade, H. et al. Single-molecule imaging of telomerase RNA reveals a recruitment-retention model for telomere elongation. Mol. Cell 79, 115–126.e116 (2020).
Belin, B. J., Lee, T. & Mullins, R. D. DNA damage induces nuclear actin filament assembly by Formin-2 and Spire-1/2 that promotes efficient DNA repair. Elife 4, e07735 (2015).
Caridi, C. P. et al. Nuclear F-actin and myosins drive relocalization of heterochromatic breaks. Nature 559, 54–60 (2018).
Lamm, N. et al. Nuclear F-actin counteracts nuclear deformation and promotes fork repair during replication stress. Nat. Cell Biol. 22, 1460–1470 (2020).
Ryu, T. et al. Heterochromatic breaks move to the nuclear periphery to continue recombinational repair. Nat. cell Biol. 17, 1401–1411 (2015).
Schrank, B. R. et al. Nuclear ARP2/3 drives DNA break clustering for homology-directed repair. Nature 559, 61–66 (2018).
Palumbieri, M. D. et al. Nuclear actin polymerization rapidly mediates replication fork remodeling upon stress by limiting PrimPol activity. Nat. Commun. 14, 7819 (2023).
Pinzaru, A. M. et al. Replication stress conferred by POT1 dysfunction promotes telomere relocalization to the nuclear pore. Genes Dev. 34, 1619–1636 (2020).
Mok, K.-W., Chen, H., Lee, W. M. & Cheng, C. Y. rpS6 regulates blood-testis barrier dynamics through Arp3-mediated actin microfilament organization in rat Sertoli cells. An in vitro study. Endocrinology 156, 1900–1913 (2015).
Stern, J. L., Zyner, K. G., Pickett, H. A., Cohen, S. B. & Bryan, T. M. Telomerase recruitment requires both TCAB1 and Cajal bodies independently. Mol. Cell Biol. 32, 2384–2395 (2012).
Kishi, J. Y. et al. SABER amplifies FISH: enhanced multiplexed imaging of RNA and DNA in cells and tissues. Nat. Methods 16, 533–544 (2019).
Bryan, T. M., Marusic, L., Bacchetti, S., Namba, M. & Reddel, R. R. The telomere lengthening mechanism in telomerase-negative immortal human cells does not involve the telomerase RNA subunit. Hum. Mol. Genet 6, 921–926 (1997).
Schmidt, J. C., Zaug, A. J. & Cech, T. R. Live cell imaging reveals the dynamics of telomerase recruitment to telomeres. Cell 166, 1188–1197 e1189 (2016).
Dunn, K. W., Kamocka, M. M. & McDonald, J. H. A practical guide to evaluating colocalization in biological microscopy. Am. J. Physiol. -Cell Physiol. 300, C723–C742 (2011).
Takasugi, T., Gu, P., Liang, F., Staco, I. & Chang, S. Pot1b−/− tumors activate G-quadruplex-induced DNA damage to promote telomere hyper-elongation. Nucleic Acids Res. 51, 9227–9247 (2023).
Nieminuszczy, J. et al. Actin nucleators safeguard replication forks by limiting nascent strand degradation. Nucleic Acids Res. 51, 6337–6354 (2023).
Wakatsuki, T., Schwab, B., Thompson, N. C. & Elson, E. L. Effects of cytochalasin D and latrunculin B on mechanical properties of cells. J. Cell Sci. 114, 1025–1036 (2001).
Coué, M., Brenner, S. L., Spector, I. & Korn, E. D. Inhibition of actin polymerization by latrunculin A. FEBS Lett. 213, 316–318 (1987).
Plessner, M., Melak, M., Chinchilla, P., Baarlink, C. & Grosse, R. Nuclear F-actin formation and reorganization upon cell spreading. J. Biol. Chem. 290, 11209–11216 (2015).
Zou, L. & Elledge, S. J. Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science 300, 1542–1548 (2003).
Saldivar, J. C., Cortez, D. & Cimprich, K. A. The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat. Rev. Mol. Cell Biol. 18, 622–636 (2017).
Diolaiti, M. E., Cimini, B. A., Kageyama, R., Charles, F. A. & Stohr, B. A. In situ visualization of telomere elongation patterns in human cells. Nucleic acids Res. 41, e176 (2013).
Dopie, J., Skarp, K. P., Rajakylä, E. K., Tanhuanpää, K. & Vartiainen, M. K. Active maintenance of nuclear actin by importin 9 supports transcription. Proc. Natl. Acad. Sci. USA 109, E544–552 (2012).
Pendleton, A., Pope, B., Weeds, A. & Koffer, A. Latrunculin B or ATP depletion induces cofilin-dependent translocation of actin into nuclei of mast cells. J. Biol. Chem. 278, 14394–14400 (2003).
Firat-Karalar, E. N. & Welch, M. D. New mechanisms and functions of actin nucleation. Curr. Opin. cell Biol. 23, 4–13 (2011).
Baarlink, C. et al. A transient pool of nuclear F-actin at mitotic exit controls chromatin organization. Nat. Cell Biol. 19, 1389–1399 (2017).
Lappalainen, P. & Drubin, D. G. Cofilin promotes rapid actin filament turnover in vivo. Nature 388, 78–82 (1997).
Yang, N. et al. Cofilin phosphorylation by LIM-kinase 1 and its role in Rac-mediated actin reorganization. Nature 393, 809–812 (1998).
Titus, M. A. Myosin-driven intracellular transport. Cold Spring Harb. Perspect. Biol. 10, a021972 (2018).
Maiso, P. et al. Defining the role of TORC1/2 in multiple myeloma. Blood J. Am. Soc. Hematol. 118, 6860–6870 (2011).
Hetrick, B., Han, M. S., Helgeson, L. A. & Nolen, B. J. Small molecules CK-666 and CK-869 inhibit actin-related protein 2/3 complex by blocking an activating conformational change. Chem. Biol. 20, 701–712 (2013).
Peterson, J. R., Lokey, R. S., Mitchison, T. J. & Kirschner, M. W. A chemical inhibitor of N-WASP reveals a new mechanism for targeting protein interactions. Proc. Natl. Acad. Sci. 98, 10624–10629 (2001).
Ross-Macdonald, P. et al. Identification of a nonkinase target mediating cytotoxicity of novel kinase inhibitors. Mol. Cancer Ther. 7, 3490–3498 (2008).
Higuchi, H. & Takemori, S. Butanedione monoxime suppresses contraction and ATPase activity of rabbit skeletal muscle. J. Biochem. 105, 638–643 (1989).
Cheung, A. et al. A small-molecule inhibitor of skeletal muscle myosin II. Nat. cell Biol. 4, 83–88 (2002).
Schmidt, J. C., Zaug, A. J., Kufer, R. & Cech, T. R. Dynamics of human telomerase recruitment depend on template-telomere base pairing. Mol. Biol. Cell 29, 869–880 (2018).
Essers, J. et al. Nuclear dynamics of PCNA in DNA replication and repair. Mol. Cell Biol. 25, 9350–9359 (2005).
Lossaint, G. et al. FANCD2 binds MCM proteins and controls replisome function upon activation of s phase checkpoint signaling. Mol. Cell 51, 678–690 (2013).
Ohki, R. & Ishikawa, F. Telomere-bound TRF1 and TRF2 stall the replication fork at telomeric repeats. Nucleic Acids Res. 32, 1627–1637 (2004).
Fouche, N., Ozgur, S., Roy, D. & Griffith, J. D. Replication fork regression in repetitive DNAs. Nucleic Acids Res. 34, 6044–6050 (2006).
Crabbe, L., Verdun, R. E., Haggblom, C. I. & Karlseder, J. Defective telomere lagging strand synthesis in cells lacking WRN helicase activity. Science 306, 1951–1953 (2004).
Miller, K. M., Rog, O. & Cooper, J. P. Semi-conservative DNA replication through telomeres requires Taz1. Nature 440, 824–828 (2006).
Zhang, T. et al. Looping-out mechanism for resolution of replicative stress at telomeres. EMBO Rep. 18, 1412–1428 (2017).
Xie, Z. et al. Early telomerase inactivation accelerates aging independently of telomere length. Cell 160, 928–939 (2015).
Meena, J. K. et al. Telomerase abrogates aneuploidy-induced telomere replication stress, senescence and cell depletion. EMBO J. 34, 1371–1384 (2015).
Chang, M. et al. Telomerase is essential to alleviate pif1-induced replication stress at telomeres. Genetics 183, 779–791 (2009).
Nagai, S. et al. Functional targeting of DNA damage to a nuclear pore-associated SUMO-dependent ubiquitin ligase. Science 322, 597–602 (2008).
Su, X. A., Dion, V., Gasser, S. M. & Freudenreich, C. H. Regulation of recombination at yeast nuclear pores controls repair and triplet repeat stability. Genes Dev. 29, 1006–1017 (2015).
Oza, P., Jaspersen, S. L., Miele, A., Dekker, J. & Peterson, C. L. Mechanisms that regulate localization of a DNA double-strand break to the nuclear periphery. Genes Dev. 23, 912–927 (2009).
Stewart, J. A. et al. Human CST promotes telomere duplex replication and general replication restart after fork stalling. EMBO J. 31, 3537–3549 (2012).
Lyu, X. et al. Human CST complex protects stalled replication forks by directly blocking MRE11 degradation of nascent-strand DNA. EMBO J. 40, e103654 (2021).
Drosopoulos, W. C., Kosiyatrakul, S. T. & Schildkraut, C. L. BLM helicase facilitates telomere replication during leading strand synthesis of telomeres. J. Cell Biol. 210, 191–208 (2015).
Drosopoulos, W. C., Kosiyatrakul, S. T., Yan, Z., Calderano, S. G. & Schildkraut, C. L. Human telomeres replicate using chromosome-specific, rather than universal, replication programs. J. Cell Biol. 197, 253–266 (2012).
Drosopoulos, W. C. et al. TRF2 Mediates Replication Initiation within Human Telomeres to Prevent Telomere Dysfunction. Cell Rep. 33, 108379 (2020).
Loayza, D. & De Lange, T. POT1 as a terminal transducer of TRF1 telomere length control. Nature 423, 1013–1018 (2003).
Gu, P. et al. Distinct functions of POT1 proteins contribute to the regulation of telomerase recruitment to telomeres. Nat. Commun. 12, 5514 (2021).
Macheret, M. & Halazonetis, T. D. DNA replication stress as a hallmark of cancer. Annu Rev. Pathol. 10, 425–448 (2015).
Sundararajan, V. et al. Nuclear pCHK1 as a potential biomarker of increased sensitivity to ATR inhibition. J. Pathol. 259, 194–204 (2023).
Cao, L. & Way, M. The stabilization of Arp2/3 complex generated actin filaments. Biochem. Soc. Trans. 52, 343–352 (2024).
Simanov, G. et al. The Arp2/3 inhibitory protein Arpin inhibits homology-directed DNA repair. Biol. Cell 116, e2400073 (2024).
Wagner, A. R., Luan, Q., Liu, S. L. & Nolen, B. J. Dip1 defines a class of Arp2/3 complex activators that function without preformed actin filaments. Curr. Biol. 23, 1990–1998 (2013).
Tomlinson, C. G., Holien, J. K., Mathias, J. A., Parker, M. W. & Bryan, T. M. The C-terminal extension of human telomerase reverse transcriptase is necessary for high affinity binding to telomeric DNA. Biochimie 128-129, 114–121 (2016).
Cohen, S. B. & Reddel, R. R. A sensitive direct human telomerase activity assay. Nat. Methods 5, 355–360 (2008).
Grimm, J. B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat. Methods 12, 244–250 (2015). 243 p following 250.
Kamentsky, L. et al. Improved structure, function and compatibility for CellProfiler: Modular high-throughput image analysis software. Bioinformatics 27, 1179–1180 (2011).
Stirling, D. R. et al. CellProfiler 4: Improvements in speed, utility and usability. BMC Bioinforma. 22, 1–11 (2021).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. methods 9, 676–682 (2012).
Cohen, J. Statistical power analysis for the behavioral sciences. (Routledge, New York, 2013).
Acknowledgements
We thank Jens Schmidt and Tom Cech for providing hTERT- and TRF2-labeled HeLa cells used for live cell imaging, Andrew Robinson for purification of the JF646-HaloTag ligand, and Alexander Sobinoff for providing the TSQ1 gene fragment. We are grateful to Josh Stern for initial discussions regarding the possibility of actin regulation of telomerase recruitment. We thank the Children’s Medical Research Institute and the ACRF Telomere Analysis Center, supported by the Australian Cancer Research Foundation, for providing access to microscopy equipment. Cell cycle analysis was performed at the Westmead Scientific Platforms, which are supported by the Westmead Research Hub, the Cancer Institute New South Wales, the National Health and Medical Research Council and the Ian Potter Foundation. This project was supported by grants from the Australian Research Council (Discovery Project grant DP190103572 to T.M.B.), Perpetual Foundation (IPAP2022/0347 to T.M.B. and A.H.), Ernest & Piroska Major Foundation (#IPAP2019-0916 to S.B.C.), Cancer Council NSW (RG 25-12 to S.B.C. and T.M.B.) and the Arcus Foundation (to T.M.B.). A.H. was supported by the Profield Foundation and the Neil and Norma Hill Foundation.
Author information
Authors and Affiliations
Contributions
A.H., M.K., and T.M.B. designed the project and conceived hypotheses; A.H., M.K., N.M.M., D.R.N., S.R., K.W., and S.B.C. performed experiments and analyzed data; A.H. and W.E.H. prepared custom scripts for Fiji and Python; A.H., N.M.M., D.R.N., K.W., S.B.C., A.J.C, N.L., and T.M.B. interpreted results; A.H. and T.M.B. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Harman, A., Kartawinata, M., Maroun, N.M. et al. Nuclear actin and DNA replication stress regulate telomere maintenance by telomerase. Nat Commun 16, 10193 (2025). https://doi.org/10.1038/s41467-025-66506-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66506-0