Abstract
The combination of DNA damage response inhibitors (DDR inhibitors) has emerged as a promising strategy for anticancer therapy. Herein, we demonstrate that the combined administration of poly (ADP-ribose) polymerase (PARP) and ataxia telangiectasia and Rad3-related (ATR) inhibitors elicits PANoptosis across diverse cellular lineages, including non-cancerous cell populations. The induction of PANoptosis is dependent on mitotic entry and ZBP1-dependent PANoptosome (ZBP1-RIPK1-Caspase8-Caspase6). We identify the Aurora kinase B (AURKB) as the upstream regulator, essential for phosphorylation of ZBP1 which impacts ZBP1-dependent PANoptosome assembly and activation. The Trp53-/- Brca1-/- model of ID8 and two patient-derived xenograft (PDX) models further confirm the occurrence of PANoptosis following combination administration in vivo. The toxicity is mitigated in ZBP1-knockout mice. This study unveils a mechanism that dictates cell fate during DDR inhibitors-induced aberrant mitosis, emphasizing the critical balance between efficacy and safety in optimizing DDR inhibitors combination therapies.
Similar content being viewed by others
Introduction
The DNA damage response (DDR) occurs continuously in cells in response to the various types of endogenous and exogenous stressors1. The DDR includes a diverse repertoire of mechanisms that help maintain the cell viability and genetic integrity by regulating the cell cycle, chromatin remodeling, and programmed cell death (PCD)1,2. For instance, the DDR enables cells to repair DNA damage by activating checkpoints that stall the cell cycle. Alternatively, the DDR can also eliminate cancer cells with unrepaired DNA damaged by inducing the programmed cell death3,4. Any event compromising the DDR can lead to genomic instability, promoting tumor development and progression. However, the compromised DDR also represents a therapeutic vulnerability in the context of cancer2,5. DDR-targeting agents have become an alternative and effective therapy for tumors by inducing cancer cell lethality6. Inhibitors of multiple members involved in DDR-regulated pathways have been rapidly developed and demonstrated antitumor activity in several preclinical and clinical studies4,7.
PARP inhibitors (PARPi) monotherapy has exerted benefits for patients with homologous recombination-deficient (HRD) tumors in the clinic by exploiting the concept of synthetic lethality8. Besides PARPi, other inhibitors regulating cell cycle checkpoints kinase, such as ATRi, CHK1i, and WEE1i, have also been explored in monotherapy strategies against tumors in clinical trials9. To maximize the clinical utility of those inhibitors, the strategy of combining PARPi with cell cycle checkpoints kinase inhibitors is being vigorously examined3. The combination of PARPi with ATRi has been demonstrated to exhibit synergistic effects in suppressing tumor growth through preclinical experiments10,11. Clinical trials further revealed that platinum-resistant patients can benefit from the combination of PARPi with ATRi (NCT03462342)12. The same has been found for PARPi-resistant recurrent high-grade serous ovarian cancer (HGSOC) (NCT03462342, NCT02576444)13,14.
The mechanistic rationale behind PARPi and ATRi combination is that PARPi induces DNA damage that activates the ATR pathway to promote cell survival15. Thus, the concomitant perturbation of signal transducer proteins of the ATR pathway produces synergistic antineoplastic activity3. PARPi induces replication stress (RS) and DNA damage, resulting in G2 cell cycle arrest. In addition to causing the replication fork collapse, ATRi abrogates the G2 arrest16. The administration of DNA-damaging agents such as PARPi and ATRi causes DNA damage and forces cells to enter mitosis in the presence of unrepaired DNA damage, leading to the activation of spindle-assembly checkpoint (SAC), the arrest of mitotic processes, and subsequent enhancement of PCD17,18.
Recently, researchers have discovered PANoptosis, a unique cell death pathway19,20. PANoptosis is a distinct programmed cell death pathway that is driven by caspases and receptor-interacting protein kinases (RIPKs) and regulated by the PANoptosome complex20,21,22,23. Mechanistically, ZBP1-dependent PANoptosome has been characterized as a crucial upstream executor of PANoptosis, whereby the assembly of ZBP1, RIPK1, caspase 8, and caspase 6 plays a pivotal role19,24,25,26,27,28. To date, it remains unknown whether PANoptosis is involved in the cancer cell death caused by DDR inhibitors combination therapy.
In this study, we report that ZBP1-dependent PANoptosome (ZBP1-RIPK1-Caspase 8-Caspase 6) is crucial for PANoptosis induction following PARPi and ATRi combination therapy. Furthermore, we discover that AURKB functions as the upstream regulator that phosphorylates ZBP1, subsequently affecting the assembly and activation of the ZBP1-dependent PANoptosome. Our findings are validated in Trp53−/− Brca1−/− model of ID8 and two patient-derived xenograft (PDX) models in which PARPi and ATRi combination-induced PANoptosis. Finally, the alleviated toxicity induced by combination therapy is observed in ZBP1−/− mice. These results suggest that phosphorylation of ZBP1 by AURKB triggers the activation of ZBP1-dependent PANoptosome after treatment with PARPi and ATRi combination, leading to cell-intrinsic antineoplastic activity and toxicity.
Results
PARP and ATR inhibitors combination therapy induces PANoptosis
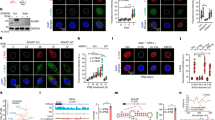
DDR inhibitors have been reported to induce DNA damage and apoptosis16. Interestingly, we observed that increasing the concentration and duration of treatment with cell cycle checkpoint inhibitors (ATRi, WEE1i, and CHK1/2i) led to the emergence of pyroptotic membrane blebbing (Supplementary Fig. 1A–C) and lactate dehydrogenase (LDH) release (Supplementary Fig. 1D–F), whereas the combination of PARPi (talazoparib) and ATRi (ceralasertib) elicited an enhanced membrane vacuolization phenotype compared to monotherapy (Fig. 1A). Using an apoptosis and necrosis detection kit, we found that the combination therapy resulted in a higher number of dead cells compared to monotherapy with either talazoparib or ceralasertib. This was evidenced by increased staining with propidium iodide (PI, indicating apoptosis, pyroptosis, or necroptosis) and oxazole yellow (YP1, indicating apoptosis or necroptosis) (Fig. 1B–D). Furthermore, the combination therapy elevated LDH release and the percentage of Annexin-V+PI+ cells compared to either monotherapy (Fig. 1E–G). Similar effects were observed when PARPi was combined with other cell cycle checkpoint inhibitors (Adavosertib or AZD-7762), which also led to increased LDH release and Annexin-V+PI+ cells (Fig. 1E–G).
A Phase-contrast imaging assay of OVCAR8 or OVCAR3 cells treated with 250 nM Talazoparib or/and 2.5 μM Ceralasertib for 24 h. The red arrowheads indicate the large bubbles emerging from the plasma membrane. Scale bar = 100 μm. B Representative images show that PI+ cells (red) may undergo apoptosis or pyroptosis, and YP1+ cells (green) may undergo apoptosis or necroptosis in OVCAR8 and OVCAR3 cells treated with 250 nM Talazoparib or/and 2.5 μM Ceralasertib for 24 h. Scale bar = 100 μm. C, D Quantification of the YP1+ and PI+ cells from (B). E Comparison of LDH release-based cell death in OVCAR8 cells treated with 250 nM Talazoparib plus cell cycle checkpoints kinase inhibitors for 24 h. ATRi: 2.5 μM Ceralasertib, WEE1i: 250 nM Adavosertib, CHK1/2i: 25 nM AZD-7762. Flow cytometry (F) and quantification analysis (G) with Annexin V/PI staining evaluating the percentages of live cells (Annexin V−/PI−), early apoptotic cells (Annexin V+/PI−), and late apoptotic cells or pyroptotic cells (Annexin V+/PI+) in OVCAR8 cells treated with 250 nM Talazoparib plus cell cycle checkpoints kinase inhibitors for 24 h. ATRi: 2.5 μM Ceralasertib, WEE1i: 250 nM Adavosertib, CHK1/2i: 25 nM AZD-7762. H, I Immunoblot analysis of GSDME, PARP1, pMLKL, tMLKL, and caspase 8 in OVCAR8 and OVCAR3 cells treated with Talazoparib or/and Ceralasertib, Olaparib or/and Ceralasertib for 24 h. Tala: Talazoparib, Cera: Ceralasertib, Ola: Olaparib. GAPDH was used as the internal control. Data were repeated thrice independently in (A, H, I) with similar results. The data in (C, D, E, G) are representative of three independent experiments and presented as the mean ± SEM. Statistical significance was determined using two-tailed one-way ANOVA with Tukey’s multiple comparisons test in (C, D, E, G). Source data are provided as a Source data file.
We further investigated the impact of combining talazoparib with ceralasertib on the molecular pathways regulating cell death in OVCAR8 and OVCAR3 cells. Treatment with either talazoparib or ceralasertib alone resulted in minimal proteolysis of GSDME, generating active fragments capable of forming membrane pores and inducing pyroptosis. However, talazoparib and ceralasertib combination led to the massive cleavage of GSDME, thereby promoting robust pyroptotic cell death (Fig. 1H, I). Similarly, there was an increase in the cleavage of PARP1 and caspase 8 in combination group, indicating the apoptotic pathway was activated (Fig. 1H, I). Cells treated with the talazoparib and ceralasertib combination also led to the phosphorylation of MLKL, a marker of necroptosis (Fig. 1H, I). We used another PARPi, olaparib, in combination with ceralasertib and observed similar results (Fig. 1H, I). Therefore, these results suggest that the combination of PARPi and ATRi triggers PANoptosis, a unique cell death modality, in ovarian cancer cells20.
Additionally, the induction of PANoptosis by ATRi and PARPi was dose-dependent, with low concentrations being sufficient to induce the response, and higher concentrations showing more pronounced effects. It was demonstrated by the increased cleavage of GSDME, PARP1, and caspase 8 (Fig. 1H, I), the enhanced phosphorylation of MLKL-S358 (Fig. 1H, I), the higher percentage of Annexin-V+PI+ cells (Supplementary Fig. 2A–D), and higher LDH release (Supplementary Fig. 2E, F).
Similar dose-dependent PANoptosis features were observed in other ovarian cancer models, including A2780 cells, two HRD primary ovarian cancer cells (POC-1 and POC-2, confirmed by the absence of Rad51 foci upon PARPi treatment using Rad51 as a marker for HR and geminin to identify S/G2 phase cells (Supplementary Fig. 2G, H)), and BRCA1-mutant UWB1.289 cells, where the cleavage of GSDME and PARP, and the MLKL-S358 phosphorylation were increased with higher combination therapy doses (Supplementary Fig. 2I–L).
Further, we assessed PANoptosis induction in HEK293T and two other non-cancerous cells, such as BMDM and IOSE-80. The combination therapy resulted in similar PANoptosis pattern in HEK293T cells. Specifically, the emergence of membrane blebbing (Supplementary Fig. 3A), the percentage of YP1+ and/or PI+ cells (Supplementary Fig. 3B, C), the release of LDH (Supplementary Fig. 3E), the percentage of Annexin-V+PI+ cells (Supplementary Fig. 3G, H), and the cleavage or the phosphorylation of PANoptosis-related proteins (Supplementary Fig. 3J) all increased in HEK293T after treating with talazoparib and ceralasertib combination. The PANoptosis was less pronounced in bone marrow-derived macrophages (BMDM), as the membrane blebbing was not significantly increased (Supplementary Fig. 3A), though the other PANoptosis characteristics were increased in the combination therapy group compared to the control, as demonstrated by the percentage of YP1+ and/or PI+ cells (Supplementary Fig. 3B, D), the release of LDH (Supplementary Fig. 3F), the percentage of PI+ Annexin-V+ cells (Supplementary Fig. 3G, I), and the cleavage of PANoptosis proteins (Supplementary Fig. 3K). Similarly, treatment of normal ovarian epithelial cells (IOSE-80) with talazoparib and ceralasertib also induced PANoptosis in a dose-dependent manner, as demonstrated by increased cleavage of GSDME and PARP1, as well as MLKL-S358 phosphorylation (Supplementary Fig. 3L, M). Together, these findings suggest the co-administration of talazoparib and ceralasertib induced PANoptosis.
Preventing mitosis entry attenuates PANoptosis induction by PARP and ATR inhibitors combination therapy
Treatment with ATRi sensitizes cells to PARPi by prematurely forcing cells into mitosis entry while DNA damage from PARPi is still unresolved15. Flow cytometry analysis revealed an increase in the proportion of mitotic pH3(S10) + cells after talazoparib and ceralasertib treatment (Supplementary Fig. 4A, B). Mechanistically, ATRi can directly repress ATR-CHK1-WEE1 signaling and therefore activate cyclin dependent kinase 1 (CDK1)29 (Fig. 2A). As expected, CDK1 activation revealed by the phosphorylation of T161 site and dephosphorylation of T14/T15 sites was observed in single ceralasertib or the combination treatment groups (Fig. 2B). CDK1 is a key regulator of mitosis entry, and its activity can be specifically inhibited by RO-3306 (Fig. 2A). Flow cytometry results demonstrated that RO-3306 reduced the increase in mitotic pH3(S10) + cells induced by talazoparib and ceralasertib treatment (Supplementary Fig. 4C, D). RO-3306 decreased the proportion of mitotic pH3(S10)+ and DNA-damaged γH2AX+ cells induced by the combination (Fig. 2C, D). Importantly, RO-3306 treatment inhibited the cleavage of GSDME, and PARP1 and the phosphorylation of MLKL induced by talazoparib and ceralasertib combination treatment (Fig. 2E). Similarly, siRNA-mediated knockdown of CDK1 (si-CDK1) also inhibited PANoptosis induction by talazoparib and ceralasertib, as shown by reduced cleavage of GSDME and PARP1, and decreased phosphorylation of MLKL (Supplementary Fig. 4E). Importantly, re-expression of wild-type CDK1 (CDK1-WT) rescued the PANoptosis induction, further confirming that CDK1 activation was required for this process (Supplementary Fig. 4E), suggesting the activation of CDK1 was required for PANoptosis induction. We investigated the role of CDK1 in mitosis by enriching mitotic cells after talazoparib and ceralasertib treatment. RO-3306 treatment inhibited the PANoptosis phenotype in these enriched cells (Supplementary Fig. 4F). Accordingly, the combination treatment inducing large pyroptotic bubbles disappeared when cells were co-treated with RO-3306 (Fig. 2F). Consistently, the percentage of PI+ and/or YP1+ cells (Fig. 2G, H) and the release of LDH (Fig. 2I) decreased when CDK1 activity was blocked. Taken together, these findings suggest that the PANoptosis induced by the talazoparib and ceralasertib combination therapy is mitotic entry dependent.
A Model diagram illustrating the effect of ATR pathways activation on CDK1 and cell cycle arrest when DNA damage occurs in cells. RO-3306 can inhibit the activation of CDK1. ATRi can inhibit the ATR and thereby activate the CDK1. Created in BioRender. Liao, Z. (2025) https://BioRender.com/qbbh4m4. B Immunoblot analysis in OVCAR8 cells treated with 250 nM Talazoparib or/and 2.5 μM Ceralasertib for 24 h. GAPDH was used as the internal control. Data were repeated thrice independently with similar results. Representative images (C) and quantification (D) of the γH2AX+cells (red) and pH3s10+ cells (green) in OVCAR8 cells treated with 250 nM Talazoparib, 2.5 μM Ceralasertib, or/and 9 μM RO-3306 for 24 h. Scale bar = 100 μm. Data were repeated thrice independently with similar results. E Immunoblot analysis in OVCAR8 cells treated with 250 nM Talazoparib, 2.5 μM Ceralasertib, or/and 9 μM RO-3306 for 24 h. GAPDH was used as the internal control. Data were repeated thrice independently with similar results. F Phase-contrast imaging assay of OVCAR8 cells treated with 250 nM Talazoparib, 2.5 μM Ceralasertib, or/and 9 μM RO-3306 for 24 h. The red arrowheads indicate the large bubbles emerging from the plasma membrane. Scale bar = 100 μm. Data were repeated thrice independently with similar results. Representative images (G) and quantification (H) of the PI+ cells (red) and YP1+ cells (green) in OVCAR8 cells treated with 250 nM Talazoparib, 2.5 μM Ceralasertib, or/and 9 μM RO-3306 for 24 h. Scale bar = 100 μm. I Comparison of LDH release-based cell death in OVCAR8 cells treated with 250 nM Talazoparib, 2.5 μM Ceralasertib, or/and 9 μM RO-3306 for 24 h. The data in (D, H, I) are representative of three independent experiments and presented as the mean ± SEM. Statistical significance was determined using two-tailed one-way ANOVA with Tukey’s multiple comparisons test in (D, H, I). Source data are provided as a Source data file.
ZBP1-centered PANoptosome mediates PANoptosis induced by PARP and ATR inhibitor combination therapy
Previous studies have demonstrated that PANoptosis is regulated by a ZBP1-dependent complex20,24. Interestingly, the protein level of ZBP1 increased in the combination treatment compared with the control group (Fig. 3A). Knockdown (KD) of ZBP1 using two independent siRNA attenuated the activation of PANoptosis induced by the combination treatment. This attenuation was demonstrated by decreased cleavage of PANoptosis proteins and abolished phosphorylation of MLKL (Fig. 3A and Supplementary Fig. 5A), disappearance of pyroptotic bubbles (Fig. 3B), the lower percentage of YP1+ and/or PI+ cells (Fig. 3C, D), the lower percentage of PI+ Annexin-V+ cells (Fig. 3E, F), and the decreased release of LDH (Fig. 3G). In addition, the mRNA level of ZBP1 also increased in the combination group in comparison to the control group (Fig. 3H). As ZBP1 was an interferon (IFN)-I-induced sensor, the KD of IFNAR1 attenuated ZBP1 mRNA and protein expression (Fig. 3H, I), suggesting that IFN signaling pathway acted as an upstream regulator of expression of ZBP1 after talazoparib and ceralasertib combination therapy.
A Immunoblot analysis of ZBP1, GSDME, PARP1, pMLKL, tMLKL, γH2AX, and pH3s10 in control and ZBP1-KD OVCAR8 cells treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. GAPDH was used as the internal control. Data were repeated thrice independently with similar results. B Phase-contrast imaging assay of control or ZBP1-KD OVCAR8 cells treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. The red arrowheads indicate the large bubbles emerging from the plasma membrane. Scale bar = 100 μm. Data were repeated thrice independently with similar results. Representative images (C) and quantification (D) of the PI+cells (red) and YP1+ cells (green) in control and ZBP1-KD OVCAR8 cells treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. Scale bar = 100 μm. E Flow cytometry and F quantification analysis with Annexin V/PI staining evaluating the percentages of live cells (Annexin V−/PI−), early apoptotic cells (Annexin V+/PI−), and late apoptotic cells or pyroptotic cells (Annexin V+/PI+) in control and ZBP1-KD OVCAR8 cells treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. G Comparison of LDH release-based cell death in control and ZBP1-KD OVCAR8 cells treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. H, I OVCAR8 cells with IFNAR1 knockdown were treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. H qPCR analysis was performed to evaluate the relative mRNA expression of ZBP1 in both control and IFNAR1-KD cells, with GAPDH serving as the internal control. I Protein levels of ZBP1 and IFNAR1 were assessed via immunoblot analysis, with GAPDH used as the loading control. J Schematic representation of full-length ZBP1 or indicated mutants. K–O After the KD of ZBP1, OVCAR8 cells were transfected with plasmids expressing Myc-tagged ZBP1 and its truncation mutants, and then were subjected to 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. K Immunoblot analysis of Myc tag, GSDME, PARP1, γH2AX, and pH3s10. GAPDH was used as the internal control. L Phase-contrast imaging assay of OVCAR8 cells. The red arrowheads indicate the large bubbles emerging from the plasma membrane. Scale bar = 100 μm. Representative images (M) and quantification (N) of the PI+ cells (red) and YP1+ cells (green). Scale bar = 100 μm. O Comparison of LDH release-based cell death. The data in (D, F, G, H, N, O) are representative of three independent experiments and presented as the mean ± SEM. Statistical significance was determined using two-tailed Student’s t test in (D, F, G) and one-way ANOVA with Tukey’s multiple comparisons test in (H, N, O). Source data are provided as a Source data file.
To elucidate the domains of ZBP1 required for PANoptosis, in ZBP1-knockdown OVCAR8 cells, we introduced siRNA-resistant wild-type ZBP1 (ZBP1-WT) or truncation mutants lacking specific domains, including Zα, RHIM, or C domains (ΔZα, ΔRHIM, ΔC), as depicted in Fig. 3J. The silencing efficiency of ZBP1 is shown in Supplementary Fig. 5B. While the ZBP1-ΔC-Myc mutant displayed similar responses compared to the full-length ZBP1-Myc, the ZBP1-ΔZα-Myc and ZBP1-ΔRHIM-Myc mutants were unable to induce PANoptosis (Fig. 3K). Indeed, compared with the full-length ZBP1 form, the expression of either Zα or RHIM mutant proteins prevented the cleavage of PANoptosis-related proteins (Fig. 3K), and membrane blebbing (Fig. 3L), the increased in YP1+ and/or PI+ cells (Fig. 3M, N), and the release of LDH (Fig. 3O) during combination therapy. These findings reveal that both the Zα and RHIM domains are essential for ZBP1-mediated PANoptosis. It is well known that Z-NA binds to the Za domain of ZBP1, thereby activating ZBP130. Therefore, we examined the levels of Z-NA following combination treatment. The results showed that, along with the upregulation of ZBP1, Z-NA levels also increased after talazoparib and ceralasertib treatment (Supplementary Fig. 5C). Taken together, these findings indicate that the Zα and RHIM domains of ZBP1 play pivotal roles in mediating PANoptosis during PARP and ATR inhibitors combination therapy.
In addition to ZBP1, RIPK1, caspase 8, and caspase 6 are all well-established components of PANoptosome. Consistent with their cooperative roles played in PANoptosome formation, the KD of RIPK1 (Supplementary Fig. 5D, G, J, K, N–P), caspase 8 (Supplementary Fig. 5E, H, J, L, N–O, Q), and caspase 6 (Supplementary Fig. 5F, I, J, M–O, R) all blocked the induction of PANoptosis. We further investigated whether these members of PANoptosome were also required for PANoptosis in HEK293T. Similar to the results obtained in OVCAR8 cancer cells, the KD of ZBP1 (Supplementary Fig. 6A, E, I, M), RIPK1 (Supplementary Fig. 6B, F, J, N), caspase 8 (Supplementary Fig. 6C, G, K, O), and caspase 6 (Supplementary Fig. 6D, H, L, P) also prevented the induction of PANoptosis in HEK293T cells. Collectively, these data suggested talazoparib and ceralasertib combination treatment-induced PANoptosis is regulated by ZBP1-dependent PANoptosome.
Aurora kinase B is essential for ZBP1-mediated PANoptosis in PARP and ATR inhibitors combination therapy
To further investigate the regulators of ZBP1 and PANoptosis induced by the combination of talazoparib and ceralasertib, we sought to identify the interacting proteins of ZBP1 through immunoprecipitation mass spectrometry (IP-MS) analysis. More than 200 associated bound polypeptides were detected, encompassing protein involved in a wide range of cellular functions, including the regulation of cell death pathways and mitotic sister chromatid separation (Fig. 4A, B). Notably, critical regulators of cell death such as AURKB, RIPK1, and caspase 8 were among these identified interacting proteins (Fig. 4A). The identification of AURKB is particularly intriguing due to its well-established involvement in mitosis and cell death processes. To validate the interaction between ZBP1 and AURKB, we performed proximity ligation assay (PLA) experiments in OVCAR8 cells treated with talazoparib and ceralasertib. The results confirmed the interaction and demonstrated co-localization of ZBP1 and AURKB on chromatin in mitotic cells (Fig. 4C, D). In addition, we found that AURKB and ZBP1 predominantly colocalize at centromeres (Supplementary Fig. 7A). To further validate our findings, we established a ZBP1 overexpression system in HEK293T cells to elucidate the molecular interaction mechanism between ZBP1 and AURKB. Co-immunoprecipitation (Co-IP) assays demonstrated that exogenous ZBP1 directly interacted with endogenous AURKB (Fig. 4E). Additionally, we observed that endogenous AURKB formed specific interaction complexes with endogenous ZBP1 (Fig. 4F, G), further substantiating the biological relationship between these two proteins. These results collectively confirm the specific molecular interaction between ZBP1 and AURKB in a cellular context. The interaction between AURKB and ZBP1 was found to be mediated by the RHIM domain of ZBP1 (Fig. 4H). The KD of AURKB using two independent siRNA abolished the ZBP1-dependent PANoptosis induced by talazoparib and ceralasertib combination treatment in OVCAR8 (Fig. 4I and Supplementary Fig. 7B) and HEK293T cells (Fig. 4J). Similar results were observed using AURKB inhibitor (Fig. 4K). Further cell cycle analysis revealed that KD of AURKB and AURKB inhibitors caused cells to arrest in the G2/M phase (Supplementary Fig. 7C–F). These findings collectively demonstrate that AURKB is an essential interacting protein for ZBP1-dependent PANoptosis induced by talazoparib combined with ceralasertib.
A The scatter plot exhibiting the interacted proteins of ZBP1 from LC-MS. The proteins with Log2FC > 2 identified as the interactors of ZBP1. We transfected 293T cells with either the ZBP1-Myc plasmid or the vector-Myc plasmid for 24 h and then treated with talazoparib and ceralasertib for another 24 h, followed by protein immunoprecipitation using Myc magnetic beads. The beads were then sent for mass spectrometry analysis. B Reactome gene sets enriched of the interacted proteins of ZBP1. Representative images (C) and quantification (D) of the PLA to detect the physical association between AURKB and ZBP1. Scale bar = 20 μm. E Co-immunoprecipitation (CO-IP) assay and WB were carried out to assess the interactions between AURKB and ZBP1 in 293T cells transfected with Myc-tagged ZBP1 or not (vector-Myc) and treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h or not. F, G Endogenous CO-IP assay to assess the interactions between AURKB and ZBP1 in HEK293T or OCVAR8 cells treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h or not. H CO-IP assay to assess the interactions between AURKB and ZBP1 or its truncation mutants in HEK293T cells transfected with HA-tagged AURKB, Myc-tagged ZBP1 or its truncation mutants, and treated with 250 nM Talazoparib and 2.5 μM Ceralasertib. Immunoblot analysis of AURKB, GSDME, PARP1, pMLKL, tMLKL in control or AURKB-KD OVCAR8 (I) and in control or AURKB-KD HEK293T (J) cells. K Immunoblot analysis of GSDME, PARP1, pMLKL, and tMLKL in OVCAR8 cells treated with 250 nM Talazoparib and 2.5 μM Ceralasertib and/or 1 μM Barasertib for 24 h. L Immunoblot analysis of pAURKB, AURKB, and ZBP1 in OVCAR8 cells treated with 250 nM Talazoparib or/and 2.5 μM Ceralasertib for 24 h. GAPDH was used as the internal control. M Model diagram illustrating the T232 phosphorylation level of AURKB during cell cycle and the phosphonull AURKB-T232A mutants. Created in BioRender. Liao, Z. (2025) https://BioRender.com/jgpa23z. N–R After the KD of AURKB, OVCAR8 cells were transfected with plasmids expressing HA-tagged AURKB-WT or phosphonull AURKB-T232A mutants and then were subjected to 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. N Phase-contrast imaging assay of OVCAR8 cells. The red arrowheads indicate the large bubbles emerging from the plasma membrane. Scale bar = 100 μm. Representative images (O) and quantification (P) of the PI+cells (red) and YP1+ cells (green). Scale bar = 100 μm. (Q) Comparison of LDH release-based cell death. (R) Immunoblot analysis of AURKB, GSDME, PARP1, pMLKL, tMLKL, γH2AX, and pH3s10. GAPDH was used as the internal control. The data in (D, P, Q) are representative of three independent experiments and presented as the mean ± SEM. Data were repeated thrice independently in (E–L, N) with similar results. Statistical significance was determined using two-tailed Student’s t test in (D) and one-way ANOVA with Tukey’s multiple comparisons test in (P, Q). Source data are provided as a Source data file.
Next, we investigated whether the kinase activity of AURKB affected PANoptosis during talazoparib and ceralasertib combination therapy. The phosphorylation level at T232 site of AURKB, an important site for its kinase activity31, increased both in ceralasertib monotherapy or the combination treatment (Fig. 4L). The expression of ZBP1 was induced following talazoparib combined with ceralasertib (Fig. 4L), which was consistent with our previous result (Fig. 3A), indicating that the combination treatment, rather than monotherapy, could activate ZBP1-dependent PANoptosis. We further transfected OVCAR8 cells with siRNA-resistant AURKB-WT or phosphonull AURKB mutant AURKB-T232A plasmids after the KD of AURKB (Fig. 4M). We verified the expression of the AURKB plasmids (Supplementary Fig. 7G). Reintroduction of AURKB-WT restored PANoptosis induced by talazoparib and ceralasertib, as evidenced by the appearance of pyroptotic bubbles (Fig. 4n), an increased percentage of YP1+ and/or PI+ cells (Fig. 4O, P), elevated LDH release (Fig. 4Q), and restored cleavage of PANoptosis-related proteins with phosphorylation of MLKL (Fig. 4R). In contrast, AURKB-T232A expression failed to rescue these PANoptotic features. Further cell cycle analysis revealed that reintroduction of AURKB-T232A into AURKB-depleted cells remained arrested in G2/M phase (Supplementary Fig. 7H, I). It indicated that the kinase activity of AURKB was essential for the combination treatment-induced PANoptosis.
AURKB phosphorylates ZBP1 to drive PANoptosis
We further investigated whether AURKB, functioning as a kinase, could phosphorylate ZBP1 and consequently impact PANoptosis. Our findings revealed that AURKB-WT, but not AURKB-T232A, increases the phosphorylation level of ZBP1 (Fig. 5A). LC-tandem mass spectrometry (LC-MS/MS) shotgun peptide sequencing was conducted to reveal the phosphorylation sites of ZBP1 that are regulated by AURKB. High-quality higher-energy collisional dissociation (HCD) spectra revealed that AURKB phosphorylated five sites of ZBP1, including S27, S289, S309, T386, and T393 (Fig. 5B). Phosphorylation levels at these sites were reduced in the AURKB-T232A group compared to AURKB-WT (Fig. 5B).
A Immunoprecipitation assay indicates ZBP1 phosphorylation levels in 293T cells transfected with Myc-tagged ZBP1 and HA-tagged AURKB or its phosphonull TA mutants. Immunoprecipitated ZBP1 was probed with anti-phosphor-ser/thr antibody. B A schematic of the human ZBP1 sequence highlighting the phosphorylation sites observed by MS is shown. Fold changes in the abundance of phosphorylated peptides of ZBP1 in 293T cells transfected with AURKB-WT relative to 293T cells transfected with vectors are shown. Dots indicate data from three independent experiments. Additionally, fold changes in the abundance of phosphorylated peptides of ZBP1 in 293T cells transfected with AURKB-TA relative to 293T cells transfected with AURKB-WT are shown. C, D After the KD of ZBP1, OVCAR8 cells were transfected with plasmids expressing Myc-tagged ZBP1 or its phosphonull mutants and then were subjected to 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. Immunoblot analysis of ZBP1, GSDME, PARP1, pMLKL, and tMLKL. GAPDH was used as the internal control. E The S309 phosphorylation site of ZBP1 in 293T cells after treated with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h was analyzed by LC-MS/MS. F–I After the KD of ZBP1, OVCAR8 cells were transfected with plasmids expressing Myc-tagged ZBP1 or its phosphonull S309A mutants and then were subjected to 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h. F Phase-contrast imaging assay of OVCAR8 cells. The red arrowheads indicate the large bubbles emerging from the plasma membrane. Scale bar = 100 μm. Representative images (G) and quantification (H) of the PI+ cells (red) and YP1+ cells (green). Scale bar = 100 μm. I Comparison of LDH release-based cell death. J The validation of ZBP1-S309 antibody using dot-blot assay. The vertical axis represents the concentration of the antibody. K In vitro phosphorylation assay of ZBP1 by purified protein AURKB or CDK1. L Immunoblot analysis in OVCAR8 cells after treated by 250 nM Talazoparib and 2.5 μM Ceralasertib and/or 1 μM Barasertib for 24 h. M After treating with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h, OVCAR8 cells were shaken off to enrich mitotic cells and then treated with 1 μM Barasertib for 6 h. Immunoblot analysis of pAURKB, AURKB, pZBP1, and ZBP1 in OVCAR8 cells. GAPDH was used as the internal control. Representative images (N) and quantification (O) of the PLA to detect the physical association between AURKB and pZBP1-S309. Scale bar = 20 μm. P CO-IP assay to assess the interactions among ZBP1, RIPK1, CASP8, and CASP6 after transfected with Myc-tagged ZBP1 or its phosphonull S309A mutants and treated with 250 nM Talazoparib and 2.5 μM Ceralasertib in 293T cells. CASP8: caspase 8, CASP6: caspase 6. Q Schematic overview of the mechanism of DDR inhibitors-induced PANoptosis during aberrant mitotic progression. Created in BioRender. Liao, Z. (2025) https://BioRender.com/bstbek5. The data in (H, I, O) are representative of three independent experiments and presented as the mean ± SEM. Data were repeated thrice independently in (A, C, D, F, K–M, P) with similar results. Statistical significance was determined using two-tailed Student’s t test in (O) and one-way ANOVA with Tukey’s multiple comparisons test in (H, I). Source data are provided as a Source data file.
To identify the most crucial phosphorylation sites among the five, we generated individual serine-to-alanine mutants for each site. OVCAR8 cells were transfected with either siRNA-resistant ZBP1-WT or one of the single-mutant plasmids following ZBP1 knockdown. Mutations at T386 and T393 sites had minimal impact on PANoptosis, consistent with their location in the less critical C-terminal domain (Fig. 5C). We therefore focused on S27, S289, and S309. Among these, the mutation of S309 (S309A) effectively inhibited PANoptosis, whereas mutations at S27 or S289 did not (Fig. 5D). Moreover, LC-MS/MS shotgun peptide sequencing analysis of cells treated with talazoparib and ceralasertib revealed an increase in S309 phosphorylation levels upon combination therapy (Fig. 5E). The introduction of ZBP1-S309A led to a reduction in pyroptotic bubbles (Fig. 5F), YP1+ and/or PI+ cell percentages (Fig. 5G, H), and LDH release (Fig. 5I), confirming the pivotal role of S309 phosphorylation in PANoptosis.
To further elucidate the role of S309, we developed a site-specific antibody targeting ZBP1-S309 phosphorylation. The antibody’s specificity was validated (Fig. 5J). In vitro kinase assay revealed that AURKB, but not CDK1, mediates phosphorylation of ZBP1 at S309 (Fig. 5K). Inhibition of AURKB reduced the phosphorylation of ZBP1-S309 during combination therapy (Fig. 5L). Further experiments showed that the inhibition of AURKB in mitotic cells after combination treatment could also reduce phosphorylation at ZBP1-S309 (Fig. 5M). Additionally, the PLA assay indicated a colocalized interaction between pZBP1-S309 and AURKB following combination treatment (Fig. 5N, O). Moreover, disruption of S309 phosphorylation impaired ZBP1 interactions with RIPK1, caspase 8, and caspase 6, emphasizing the critical role of S309 phosphorylation in the assembly of the ZBP1-dependent PANoptosome (Fig. 5P).
To determine whether this mechanism is conserved in mice, we further investigated its role in mouse ovarian cancer ID8 cells. Knockdown of Aurkb in ID8 cells reversed combination treatment-induced PANoptosis, indicating that Aurkb plays a critical role in this process in mice as well (Fig. S7J). Notably, knockdown of Ripk1 also reversed PANoptosis (Supplementary Fig. 7K), suggesting its involvement in this pathway, while knockdown of Aurka did not reverse PANoptosis (Supplementary Fig. 7L). To examine ZBP1 phosphorylation in mice, we transfected mouse Zbp1 plasmids into ID8 cells. Despite optimization, Zbp1 plasmids expression remained low, limiting mass spectrometry detection. However, Western blot analysis still showed increased ZBP1 phosphorylation after AURKB overexpression, as well as PARPi and ATRi combination treatment (Supplementary Fig. 7M), confirming ZBP1’s responsiveness. Due to the persistently low Zbp1 protein abundance in ID8 cells, we subsequently employed Chinese Hamster Ovary (CHO) cells as an alternative mammalian expression system to achieve robust mouse ZBP1 protein levels. Phosphoproteomic analysis of CHO cells revealed increased mouse ZBP1 phosphorylation following AURKB overexpression, as well as PARPi and ATRi combination treatment (Supplementary Fig. 7N). The specific phosphorylation sites elevated in mouse ZBP1 included S127, S273, T356, and S384 (Supplementary Fig. 7O).
The findings of our study provide insights into the mechanism underlying PANoptosis induced by PARPi and ATRi combination therapy, wherein AURKB phosphorylates ZBP1 to initiate the assembly of a ZBP1-dependent PANoptosome and drive PANoptosis (Fig. 5Q).
PARP and ATR inhibitors combination therapy induces PANoptosis in vivo
After confirming that the combination of PARPi and ATRi induces PANoptosis in vitro, we next explored whether this effect could be observed in vivo. We generated a cell line-derived xenograft (CDX) in C57BL/6 mice using a Trp53−/− Brca1−/− knockout ID8 cell line to evaluate the therapeutic efficacy of orally administered talazoparib and ceralasertib (Supplementary Fig. 8A). The combination therapy showed strong antitumor effects compared to the control, but the benefit was less than the sum of the single agents. This sub-additive effect may result from a ceiling effect, as the Trp53 and Brca1 double-deficient model is already highly sensitive to PARP inhibition. Thus, the limited added benefit may reflect model-specific constraints rather than true drug antagonism (Supplementary Fig. 8B–D). Furthermore, the combination therapy-induced PANoptosis within the tumor tissues, as demonstrated by increased expression levels of HMGB1, calreticulin, IL-1β, cleaved caspase 3, ZBP1, and AURKB (Supplementary Fig. 8E).
To further validate the critical role of ZBP1 in vivo, we performed CDX models using ZBP1 wild-type (WT) and ZBP1-knockout (KO) OVCAR8 cells in BALB/c nude mice. Mice were treated with a combination of talazoparib and ceralasertib (Supplementary Fig. 8F). In contrast to the ZBP1 WT group, where combined treatment suppressed tumor growth, ZBP1 knockout abrogated this therapeutic effect (Supplementary Fig. 8G–I). Specifically, the loss of ZBP1 prevented the tumor regression induced by combination treatment (Supplementary Fig. 8G–I). Moreover, ZBP1 deficiency rescued tumor cells from undergoing PANoptosis in response to the combination therapy (Supplementary Fig. 8J), highlighting the role of ZBP1 in mediating treatment-induced PANoptotic cell death in vivo.
To further investigate the role of AURKB in this process, we utilized an HGSOC PDX model and examined whether the addition of barasertib, an AURKB inhibitor, could mitigate the effects of talazoparib and ceralasertib combination therapy (Supplementary Fig. 8K). Indeed, barasertib rescued the tumor-suppressive effects of the combination treatment (Supplementary Fig. 8L–N). Barasertib also rescued the PANoptosis induced by the combination treatment (Supplementary Fig. 8O), underscoring the role of AURKB in mediating PANoptosis.
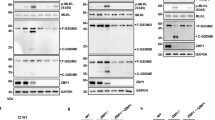
We further validated the antitumor effects of the ceralasertib and another PARPi (olaparib) in the second HGSOC PDX model. Using the HGSOC PDX model, we validated the antitumor effects of the combination therapy with olaparib and ceralasertib (Fig. 6A–C). The combination group showed a slight decrease in mice weight without statistical significance (Fig. 6D). Western blot analysis of tumor protein extracts confirmed that the combination therapy induced the proteolytic cleavage of PANoptosis-associated proteins in the PDX tumor tissues, which are concomitant with elevated expressions of pZBP1-S309 and pAURKB-T232 (Fig. 6E–G). In addition, the hallmarks of immunogenic cell death, defined as HMGB1 secretion and calreticulin exposure, and pro-inflammatory marker IL-1β expression were also elevated in the PDX tumor tissues collected from mice treated with the combination therapy (Fig. 6F, G). These findings suggest that olaparib combined with ceralasertib induces PANoptosis in PDX models, exerting an antitumor effect.
A Model diagram illustrating a PDX models were established on BALB/c nude mice using the tumor tissues from a high-grade serous ovarian cancer patient, and were treated with vehicle or the combination of olaparib (50 mg/kg/day) and ceralasertib (50 mg/kg/day), (n = 5 in each group). Created in BioRender. Liao, Z. (2025) https://BioRender.com/4xb7ilp. B From the beginning of treatment, tumor volumes were measured to plot tumor growth curves. N = 5 mice in each group. C Photograph of tumor tissues from PDX models. Scale bar = 1 cm. D From the beginning of treatment, mouse weight was measured to plot mice growth curves. N = 5 mice in each group. E Immunoblot analysis in tumor tissues of PDX models. GAPDH was used as the internal control. The lane numbers in the figure correspond to different treatment conditions applied to PDX tumor tissue samples. CASP3: caspase 3. Data were repeated thrice independently with similar results. Representative images (F) and quantification (G) of immunohistochemical staining with indicated antibodies in tumor specimens. Scale bar = 100 μm. Dots indicate the individual mouse. N = 5 mice in each group. The data in (B, D, G) are representative of five biological replicates and are presented as the mean ± SEM. Statistical significance was determined using two-tailed Student’s t test in (B, D, G). Source data are provided as a Source data file.
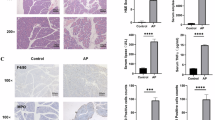
To investigate whether the combination treatment-induced toxicity in normal tissues, we conducted comprehensive histological analyses. The combination treatment resulted in toxic effects on bone marrow, spleen, kidney, liver, and lung; however, no significant toxic alterations were observed in the heart (Supplementary Fig. 9A). Specifically, following olaparib and ceralasertib administration, bone marrow displayed compensatory hyperplasia, splenic corpuscles showed marked reduction in number, glomeruli exhibited atrophy accompanied by renal tubular hydration, liver tissues presented focal punctate necrosis, and lungs demonstrated alveolar wall thickening (Supplementary Fig. 9A). Importantly, immunohistochemical staining (IHC) confirmed that the combination therapy activated PANoptosis pathways in bone marrow and spleen, as evidenced by elevated levels of multiple PANoptosis markers (Supplementary Fig. 9B–D). Collectively, these findings demonstrate that PARPi and ATRi induce PANoptosis in normal tissues.
Deficiency of ZBP1 alleviates PARP and ATR inhibitors combination therapy-induced toxicity in mice
Subsequently, we conducted further investigations to ascertain whether the toxic impact of PARPi combined with ATRi could be mitigated by ZBP1 deficiency. The co-administration of olaparib and ceralasertib decreased the body weight of the mice, while this impact was mitigated in ZBP1−/− mice (Fig. 7A). The combination therapy induces hematotoxicity in mice, leading to a decrease in leukocyte and erythrocyte counts, as well as a reduction in hemoglobin levels (Fig. 7B). The deficiency of ZBP1 alleviated the hematotoxicity induced by olaparib combined with ceralasertib (Fig. 7B). Besides, the H&E staining revealed deficiency of ZBP1 attenuated the toxicity on bone marrow, spleen, kidney, liver, and lung to some extent (Fig. 7C and Supplementary Fig. 10A). The IHC staining of bone marrow and spleen showed that ZBP1 deficiency decreased the expression of ZBP1, HMGB1, IL-1β, calreticulin, and cleaved-caspase 8 induced by the combination treatment with olaparib and ceralasertib (Fig. 7D–F and Supplementary Fig. 10B–D). Additionally, we found that combination treatment induced the upregulation of PANoptosis-related proteins in the spleens of wild-type ZBP1 mice, whereas this effect was not observed in ZBP1-knockout mice (Supplementary Fig. 10E). Taken together, those results indicate that ZBP1 deficiency alleviated olaparib and ceralasertib combination-induced systemic toxicity in mice.
A–F ZBP1+/+ or ZBP1−/− mice were treated with vehicle or the combination of olaparib (50 mg/kg/day) and ceralasertib (50 mg/kg/day) (n = 5 in each group). A From the beginning of treatment, mouse weight was measured to plot mice growth curves. N = 5 mice in each group. B The blood tests showing the comparison of leukocyte, erythrocyte, and hemoglobin. N = 5 mice in each group. C Representative images of HE staining in bone marrow, spleen, and kidney specimens. The black arrowheads in spleen indicate the splenic corpuscles. The black arrowheads in kidney indicate the renal tubules were hydrated, while the red arrowheads indicate the glomeruli were atrophied. Scale bar = 100 μm. Representative images (D, E) and quantification (F) of immunohistochemical staining with indicated antibodies in bone marrow and spleen specimens. Scale bar = 100 μm. Dots indicate the individual mouse. N = 5 mice in each group. The data in (A, B, F) are representative of five biological replicates and are presented as the mean ± SEM. Statistical significance was determined using two-tailed one-way ANOVA with Tukey’s multiple comparisons test in (A, B, F). Source data are provided as a Source data file.
Discussion
In the present study, we found that PARPi combined with ATRi induces PANoptosis, which has been validated in multiple animal models, including Trp53−/− Brca1−/− ID8 and two PDX models. Mechanistically, AURKB phosphorylates ZBP1 at the S309 site to activate the assembly of the ZBP1-dependent PANoptosome to trigger PANoptosis. While PANoptosis is driven in cancer cells, it also operates, albeit to a lesser extent, in non-cancer cells, potentially explaining the observed toxicity of this drug combination, as evidenced by the mitigated treatment-related toxicity in ZBP1 knockout mice. These findings provide a theoretical foundation for understanding their anticancer effects and toxicity, crucial for optimizing the use of DDR inhibitors combination approaches in the clinic.
DDR inhibitors target specific proteins, such as PARP, ATR, CHK1, and WEE1, disrupting DDR pathways or G2/M checkpoints9. Once cells have already committed mitosis in the presence of unresolved DNA damage, the accumulation of DNA damage results in mitotic catastrophe32,33. Mitotic catastrophe causes cells to enter an irreversible state of senescence or directly triggers apoptotic and necrotic cell death pathways33. These processes are orchestrated by protein kinases, including AURKB plays a crucial role in mediating BIM/BID-driven apoptosis34,35. Interestingly, in this study, we found that apoptosis is not the only PCD pathway triggered by the mitotic catastrophe induced by PARPi combined with ATRi; instead, we identified PANoptosis as the predominant cell death mechanism. PANoptosis is a distinct programmed cell death pathway that is driven by caspases and RIPKs and regulated by the PANoptosome complex. Importantly, we identified AURKB, a key mitotic kinase protein, as an upstream activator driving PANoptosis.
AURKB plays a crucial role in determining the fate of mitotic cells by exerting significant effects through phosphorylation of multiple proteins during mitotic progression. Specifically, it is involved in regulating chromosome alignment and spindle-assembly checkpoint (SAC) by phosphorylating kinetochore proteins (including KNL1, MIS12, and NDC80) and SAC proteins (such as HEC1)36,37. AURKB has also been shown to phosphorylate caspase-2 and cGAS, thereby inhibiting their respective activities to avoid stimulating death pathways and ensure proper mitosis38,39. The kinase activity of AURKB is tightly regulated through phosphorylation of the residue T232, which is catalyzed by CDK140,41. By constructing antibodies specific to ZBP1-S309, we demonstrated the crucial role of AURKB in mediating PANoptosis by which phosphorylation of ZBP1 at the S309 site initiates the assembly of PANoptosome and activates PANoptosis. Although Ser309 is outside the RHIM domain, phosphorylation at this site may induce allosteric changes that either facilitate or stabilize ZBP1s interactions with RIPK1, caspase 8, and caspase 6. The introduction of negative charges through phosphorylation may alter the local electrostatic environment, thereby propagating conformational changes toward the RHIM region and optimizing its structural configuration for interaction with RIPK142. It is noteworthy that Ser309 in human ZBP1 is not conserved in mice/rats. We observed that human ZBP1 can be phosphorylated by AURKB at Ser309 and trigger PANoptosis, but the corresponding serine residue is not conserved in mouse ZBP1, suggesting this regulation may be human-specific. However, the precise regulatory sites and potential involvement of other signaling pathways require further investigation in mice.
The multifunctional protein ZBP1 triggers a signaling cascade that can initiate the innate immune responses and regulate cell death pathways upon activation43,44. During mitotic progression, we found that ZBP1 acts as a crucial sensor in initiating the downstream PANoptosome assembly. Our data demonstrated that the Zα and RHIM domain of ZBP1 were required for the activation of the protein and to mediate PANoptosis in response to PARPi and ATRi combination. Additionally, we observed an increase in Z-NA following combination treatment. Given that Z-NA binds to the Za domain of ZBP1 to activate it30, Z-NA may serve as a trigger for ZBP1 activation upon combination treatment. The ZBP1-dependent PANoptosome, including ZBP1, RIPK1, caspase 8, and caspase 6, was found to be required to trigger PANoptosis in response to DDR inhibitors treatment, which was consistent with previous studies24. These findings suggest that tumors with intact ZBP1-driven PANoptotic pathways, especially pyroptotic and necroptotic pathways, might elicit greater benefits from PARPi and ATRi combination therapy.
DDR inhibitors have shown exceptional activity in many preclinical cancer models. Unfortunately, its establishment in the clinic has been challenging due to the toxicity that emerges from those combination therapies45. Our study found that PARPi combined with ATRi induces PANoptosis. Importantly, the toxicity that results from this combination therapy is alleviated in ZBP1-deficient mice. The occurrence of PANoptosis induced by the combination therapy happens most likely in cells with high proliferative activity, as it relies on mitotic entry. Considering that cancer cells exhibit relatively elevated proliferation rates compared to normal tissues, there exists a potential window for the application of PARPi and ATRi. In summary, comprehending the underlying mechanisms of toxicity caused by DDR inhibitors offers the opportunity to improve the administration doses and schedule of the drugs and thus improve their usage.
In this study, the combination of PARPi and ATRi was found to exert antitumor effect and induce toxicity through PANoptosis during aberrant mitotic progression. We identified the mitotic kinase AURKB as an essential regulator of ZBP1, responsible for triggering the assembly of PANoptosome and mediating PANoptosis. Overall, this study unveils a mechanism that dictates cell fate during DDR inhibitors-induced aberrant mitosis, emphasizing the critical balance between efficacy and safety in optimizing DDR inhibitors combination therapies.
Methods
Study approval
All experimental procedures involving mice in this study were approved by the Institutional Animal Care and Use Committee of Tongji Hospital (Approval Number: TJH-202308015) and were performed in strict accordance with the guidelines for the ethical use and care of animals established by the Chinese Council on Animal Care.
The clinical samples of HGSOC used in this study were from patients in the NANT (neoadjuvant niraparib monotherapy) clinical trial (NCT04507841). In accordance with the Helsinki Declaration, this study received formal ethical review and approval from the Institutional Review Board of Tongji Hospital Affiliated to Tongji Medical College, Huazhong University of Science and Technology (Permit Number: S122-5).
Cell lines and cell culture
Human ovarian cancer cell line OVCAR8 was obtained from MDACC’s Characterized Cell Line Core. Human ovarian carcinoma OVCAR3 cells, UWB1.289, human embryonic kidney cell line HEK293T, and U2OS were obtained from the American Type Culture Collection (ATCC). Human ovarian epithelial cell line IOSE-80 cells and CHO cells were purchased from Seven Innovation (Beijing)Biological Technology Co., Ltd. ID8 cells (C57BL/6-derived ovarian carcinoma model) were kindly provided by Professor K. Roby (Department of Anatomy and Cell Biology, University of Kansas, U.S.A.). All cell lines were tested monthly for Mycoplasma using PCR and were not passaged more than 30 times.
OVCAR8 cells and OVCAR3 cells were cultured in RPMI-1640 medium. HEK293T cells, IOSE-80 cells, and ID8 cells were cultured in Dulbecco’s modified Eagle medium (DMEM). U2OS cells were cultured in McCoy’s 5A Modified medium. All the media were supplemented with 10% FBS (fetal bovine serum) and 10,000 U/mL of PS (penicillin-streptomycin). Primary mouse BMDMs were cultivated for 6 days in DMEM supplemented with 10% FBS, 30% L929-conditioned medium, 1% non-essential amino acids, and 1% PS. BMDMs at a density of 1 × 106 cells/well in 12-well plates or 5 × 105 cells/well in 24-well plates were seeded into growth media overnight before use. All cell lines were cultured at 37 °C in a 5% CO2 atmosphere.
Compounds and antibodies
Talazoparib (HY-16106), Ceralasertib (HY-19323), Adavosertib (HY-10993), and RO-3306 (HY-12529) were purchased from MCE (New Jersey, USA). AZD-7762 (S1532) was purchased from Selleck (Texas, USA). The primary antibodies and concentrations were as follows: anti-GAPDH (Abclonal, A19056; 1:1000 dilution), anti- GSDMD (Abcam, ab210070; 1:1000 dilution), anti-GSDME (Abcam, ab215191; 1:1000 dilution), anti-PARP1 (CST, 9532; 1:1000 dilution), anti-cleaved caspase 3 (CST, 9664; 1:1000 dilution), anti-cleaved caspase 8 (CST, 9496; 1:1000 dilution), anti-MLKL (Abcam, ab184718; 1:1000 dilution), anti-pMLKL-S358 (CST, 91689; 1:1000 dilution), anti-pH3-S10 (CST, 53348; 1:1000 dilution), anti-γH2AX (CST, 9718; 1:1000 dilution), anti-RIPK1 (CST, 3493; 1:1000 dilution), anti-ZBP1 (Adipogen, AG-20B-0010; 1:1000 dilution), anti-AURKB (CST, 3094; 1:1000 dilution), anti-AURKB-T232 (CST, 2914; 1:2000 dilution), anti-Myc tag (Proteintech, 60003-2-Ig; 1:2000 dilution), anti-HA tag (Proteintech, 66006-2-Ig; 1:10000 dilution), anti-CDK1 (Abclonal, A11420; 1:1000 dilution), anti-CDK1-T161 (Abclonal, AP0324; 1:400 dilution), anti-CDK1-T14 (Abclonal, AP1465; 1:1000 dilution), anti-CDK1-Y15 (Abclonal, AP0016; 1:500 dilution), anti-Rad51 (GeneTex, GTX100469; 1:500 dilution), anti-Geminin (Zenbio, 222297; 1:200 dilution).
siRNA transfection and plasmid transfection
For small interfering RNA (siRNA) transfection, cells were seeded into six-well plates at 70% confluency, and the introduction of siRNA into cells was carried out with Lipofectamine 3000 (Invitrogen, L3000075) following the manufacturer’s protocol.
The siRNAs directed against AURKB (GCAGAAGAGCTGCACATTT or ACACCCGACAUCUUAACGC) were designed and synthesized by RIBOBIO.
The siRNAs designed and synthesized by Generalbiol are as follows:
The siRNAs directed against ZBP1 (AGGAAGACAUCUACAGGUU or AACCAAGUCCUCUACCGAA);
The siRNAs directed against RIPK1 (GGAUAAUUUUGGAAAUCAUTT);
The siRNAs directed against caspase 8 (GAUACUGUCUGAUCAUCAATT);
The siRNAs directed against caspase 6 (GCAAUCACAUUUAUGCAUATT);
The siRNAs directed against CDK1 (ACAGGUCAAGUGGUAGCCA).
For plasmid transfection, cells were seeded into six-well plates at 70% confluency and transfected with plasmids using Lipofectamine 8000 (Beyotime, C0533). Point mutations in ZBP1 and AURKB were generated following the protocol of ClonExpress II One Step Cloning Kit (Vazyme, C214).
Immunofluorescence staining for γH2AX and pH3
After fixation in 4% formaldehyde in PBS and permeabilization in 0.5% Triton X-100/PBS, cells were blocked in 1% bovine serum albumin in 0.5% Triton/PBS for 1 h. Cells were then stained with specific primary antibodies (anti-pH3S10, Abcam, ab5176, 1:200; anti-γH2AX, CST, 80312, 1:200) at 4 °C for 12 h, followed by incubation with secondary antibodies (anti-Rabbit AlexaFluor 488 antibody, Jackson Immunoresearch, 711-545-152, 1:200; anti-Mouse AlexaFluor 594 antibody, Jackson Immunoresearch, 715-585-151, 1:200) in the dark for 1 h. After counterstained with DAPI (C1005, Beyotime), cells were examined under a fluorescent microscope. For quantification, foci and nuclei were segmented, and then the integrated intensity of all foci normalized to cell area was taken as staining intensity. For each condition, at least 300 cells were counted.
Live/dead cell assay (LDH)
After cells being seeded into 12-well plates and treated for the indicated periods, the culture media were collected. The release of LDH was assessed using Cytotoxicity LDH Assay Kit (Dojindo, CK12) according to the manufacturer’s instructions. LDH levels were assessed by measuring the absorbance at 490 nm.
Flow cytometry for apoptosis
Cells were seeded in triplicate in 12-well plates for 24 h, followed by treatment with test compounds for the indicated times. After trypsinization into a cell suspension, cells were washed with PBS by centrifugation, followed by staining with Annexin V and propidium iodide (PI) using the FITC Annexin V Apoptosis Detection Kit I (BD Biosciences, 556547). The fluorescence intensity of cells with positive staining was measured by a Beckman Coulter CytoFLEX Flow Cytometer. Flow cytometry data were collected using CytExpert (version 2.4). The analysis and visualization of the flow cytometry data were carried out by FlowJo V10 software.
Flow cytometry for cell cycle
Cells with specific treatments were collected, washed with cold PBS, and then fixed with cold 70% ethanol overnight at 4 °C. Afterwards, cells were washed with cold PBS and stained with propidium iodide (PI) staining buffer (50 µg/mL PI, 4 mM sodium citrate buffer, and 10 µg/mL RNase A) and incubated at 37 °C for 15 min protected from light. At least 10,000 cells per tube were counted by flow cytometry. The relative distribution of cells over the different cell cycle phases was analyzed using the FlowJo V10 software.
Immunoblotting
Cell or tissue samples were treated with specific treatment and then lysed with RIPA lysis buffer (Servicebio, G2002-100) supplemented with EDTA-free protease inhibitor cocktail (MCE, HY-K0010) and phosphatase inhibitor cocktail I (HY-K0021) and phosphatase inhibitor cocktail II (MCE, HY-K0022). After incubation at 4 °C for 30 min, the lysates were sonicated for 30 s and clarified by centrifugation at 12,000 RPM and 4 °C for 15 min. Protein concentration was measured using Coomassie dye (Beyotime, ST1119). The proteins were boiled in SDS loading buffer at 100 °C for 10 min and separated by performing 10% SDS-polyacrylamide gel electrophoresis and subsequently transferred onto PVDF membranes. The membranes were then blocked-in protein-free rapid blocking solution (Boster, AR0041) and incubated at 4 °C overnight with specific primary antibodies, followed by incubation with a horseradish peroxidase-conjugated secondary antibody (Antgene, ANT019, ANT020). Blot bands were visualized using a WesternBrightTM ECL kit (Advansta, 190113-13) and enhanced chemiluminescence (BioRad).
Real-time quantitative PCR (qPCR)
For qPCR assay, total RNA (1 μg) was purified from cells with indicated treatment using FastPure Cell/Tissue Total RNA Isolation Kit V2 (Vazyme, RC112) and was reversed to cDNA using HiScript IV RT SuperMix (Vazyme, R423). ChamQ Universal SYBR qPCR Master Mix (Vazyme, Q711) was used for qPCR test. The relative abundance of target gene expression was quantified using the delta-delta-Ct (ddCt) method with GAPDH as the internal control. The primers used were as follows: IFNAR1 (forward, CGCCTGTGATCCAGGATTATCC; reverse, TGGTGTGTGCTCTGGCTTTCAC), GAPDH (forward, GGAGCGAGATCCCTCCAAAAT; reverse, GGCTGTTGTCATACTTCTCATGG).
PLA assay
For the PLA assays, cells were deposited on glass slides, harvested, and processed according to the manufacturer’s instructions (Sigma, DUO92102). Briefly, the cells were pretreated with respect to fixation, retrieval, and permeabilization, and then blocked with Duolink Blocking Solution for 1 h at 37 °C. Blocking solution was then washed, and samples were incubated with the primary antibodies, which were diluted by Duolink Antibody Diluent in the humidity chamber. Afterwards, samples were incubated with PLUS and MINUS PLA probes for 1 h, Ligation buffer for 30 min, and the Amplification buffer for 100 min. Finally, the slides were mounted with an overslip using a minimal volume of Duolink® PLA Mounting Medium with DAPI and analyzed using a fluorescence or confocal microscope. The primary antibodies for PLA were: anti-ZBP1 (Adipogen, AG-20B-0010; 1:1000 dilution), anti-AURKB (CST, 3094; 1:1000 dilution).
Immunoprecipitation
The cells were seeded into six-well plates and transfected with the indicated plasmids. Subsequently, the cells were washed with PBS and harvested, followed by lysis in an ice-cold immunoprecipitation buffer for 15 min. The lysis buffer (50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 1% Triton X-100, 1 mM EDTA, 0.5% sodium deoxycholate) containing EDTA-free protease inhibitor cocktail (MCE, HY-K0010) and phosphatase inhibitor cocktail I (HY-K0021), and phosphatase inhibitor cocktail II (MCE, HY-K0022). After centrifugation at 13,000 × g for 15 min, a 40 μL aliquot of the clarified lysate was pipetted to a clean centrifuge tube as the total cell lysate sample. The remaining lysates were incubated with anti-Myc, anti-Flag, anti-HA, or Protein A/G magnetic beads (MCE) overnight at 4 °C with gentle rotation. Subsequently, bead-bound proteins were washed with ice-cold lysis buffer for 3 times and boiled in SDS loading buffer at 100 °C for 10 min. The immunoprecipitated proteins were separated with SDS-PAGE, transferred to polyvinylidene difluoride membrane, and analyzed by immunoblotting with the indicated antibodies.
Mass spectrometry
We transfected 293T cells with either the ZBP1-Myc plasmid or the vector-Myc plasmid and treated cells with 250 nM Talazoparib and 2.5 μM Ceralasertib for 24 h, followed by protein immunoprecipitation using Myc magnetic beads. The beads were then sent for mass spectrometry analysis. Beads samples were incubated in the reaction buffer (1% SDC/100 mM Tris-HCl, pH 8.5/10 mM TCEP/40 mM CAA) at 95 °C for 10 min for protein denaturation, cysteine reduction, and alkylation. The eluates were diluted with equal volume of H2O and subjected to trypsin digestion overnight by adding trypsin at a ratio of 1:50 (enzyme: protein, w/w) for overnight digestion at 37 °C. The next day, TFA was used to bring the pH down to 6.0 to end the digestion. After centrifugation (12,000 × g, 15 min), the peptide was purified using self-made SDB desalting columns. Afterwards, the samples were fractionated into various fractions in a C18 analytical column (75 μm × 25 cm, C18, 1.9 μm, 100 Å). LC-MS/MS data were acquired using a Q Exactive HF mass spectrometer coupled with UltiMate 3000 RSLCnano system, and analyzed with MaxQuant (V1.6.6) using the Andromeda database search algorithm.
In vitro phosphorylation assay
Flag-tagged recombinant human AURKB protein was purchased from Proteintech (Cat: 81352), GST-tagged recombinant human CDK1 & Cyclin B1 protein was obtained from SinoBiological (Cat: C22B-10G), and GST-tagged recombinant human ZBP1 protein was obtained from CUSABI (Cat: CSB-EP861990). 0.5 μg of the recombinant ZBP1 protein was incubated with AURKB or CDK1 in the kinase buffer (CST, 9802) together with 1 mM cold ATP (MCE, HY-B2176) for 1 h at 30 °C, which was stopped by adding 5× loading buffer and boiling for 5 min at 95 °C. The products were analyzed by SDS-PAGE.
ZBP1-S309 antibody preparation
The ZBP1-S309 antibody was custom-made by Jingjie PTM Biolab (Hangzhou) Co., Inc. Phosphorylated peptides containing the target site and flanking sequences were synthesized, along with corresponding non-phosphorylated peptides. The phosphorylated peptides were emulsified with Freund’s adjuvant and injected into New Zealand White rabbits. Booster immunizations were performed at 2-week intervals. Serum was collected when sufficient antibody titers were detected.
The antibodies were purified by ammonium sulfate precipitation and protein-A affinity chromatography, followed by phospho-peptide affinity purification. Negative selection was performed using non-phosphorylated peptide columns to eliminate cross-reactivity. The specificity of purified antibodies was validated by Dot Blotting using both phosphorylated and non-phosphorylated peptides as coating antigens.
Immunohistochemistry
Tissue samples were fixed in 4% formalin, then processed and embedded in paraffin by standard procedures. Sections were subjected to deparaffinization and antigen retrieval and endogenous peroxidase blockade, followed by blocking with goat serum, and then were incubated with primary antibodies at 4 °C overnight. Subsequently, sections were subjected to HRP-conjugated secondary antibody incubation and DAB staining. Staining signals were quantified by ImageJ software, and for every section, the mean of three readings was calculated and considered as the final result. Sections were also stained with hematoxylin and eosin (H&E) and examined by a pathologist blinded to the experimental groups.
Experimental design in vivo
All procedures involving tumor burden in this therapeutic study were conducted in strict compliance with the guidelines approved by the Institutional Animal Care and Use Committee of Tongji Hospital. The maximum allowable tumor burden, as defined in our approved protocol, was 10% of the animal’s body weight. Furthermore, the humane endpoint was implemented immediately upon any observation of tumor ulceration, infection, or necrosis. Throughout the study, we ensured through diligent monitoring that no tumor burden exceeded this limit by applying the humane endpoint prior to its attainment. C57BL/6 mice, NCG mice, BALB/c nude mice, and ZBP1 mutant mice (4–6 weeks old) were purchased from GemPharmatech (Nanjing, China), bred at the Animal Center at Tongji Hospital, and maintained under SPF (specific pathogenfree) conditions after passing the quarantine inspection. All animals were acclimatized for a period of 1 week prior to the initiation of any experimental procedures. Mice were maintained with a 12 h light/ dark cycle and were fed standard chow, which was sterilized every week.
The PDX model was constructed by subcutaneous xenograft method. The clinical samples of ovarian cancer were from patients recruited in NANT trial. Female patients enrolled in this study underwent cytoreductive or exploratory surgery at Tongji Hospital and were diagnosed as HGSOC. All tumor specimens were histopathologically confirmed as HGSOC, with subsequent HRD genetic profiling performed using next-generation sequencing platforms. All participants provided written informed consent for the use of clinical and molecular data and received systemic staging evaluations and standardized treatment regimens aligned with internationally recognized ovarian cancer management guidelines.
Detailed experimental procedures were as follows: After the obtained patient’s fresh ovarian tumor tissue was transported at low temperature, it was placed in RPMI-1640 medium and washed repeatedly, after which tissue blocks of about 3 × 3 × 3 mm3 was prepared and transplanted into the subcutaneous flank of female BALB/c nude mice or NCG mice to maintain the primary characteristics of the tumor to the greatest extent. Only female mice were used in this study because ovarian cancer is a sex-specific disease that occurs exclusively in females. Sex was therefore a biological variable integral to the experimental design rather than an independent variable to be tested. When the volume of the subcutaneous xenograft of PDX model grew to 100 mm3, the experimental animals were randomly divided. BALB/c nude mice PDX models were randomized into two cohorts: vehicle control (solvent) and experimental group receiving combination therapy (Olaparib 50 mg/kg/day + Ceralasertib 50 mg/kg/day). NCG mice PDX models were stratified into four treatment arms: vehicle control (solvent), experimental group 1: Talazoparib 0.33 mg/kg/day + Ceralasertib 50 mg/kg/day, experimental group 2: Barasertib 5 mg/kg/day monotherapy, experimental group 3: Triplet combination (Talazoparib 0.33 mg/kg/day + Ceralasertib 50 mg/kg/day + Barasertib 5 mg/kg/day). During the experiment, the body weight and tumor volume of mice were measured regularly, and the tumor volume calculation formula was V = length × width2/2. At the end of the experiment, Hematologic sampling via retroorbital venous plexus was performed. Tumor tissues and major organs were harvested for immunohistochemical detection and HE staining analysis.
For the construction of Trp53−/− Brca1−/− ID8 subcutaneous tumor model, 5 × 106 ID8 cells were resuspended in a 1:1 mixture of PBS and Matrigel (Corning, 354234) and injected subcutaneously into female C57BL/6 mice to establish a xenograft model. Female mice were used exclusively in this study as the research focuses on ovarian cancer, which is a female-specific malignancy. Upon reaching 100 mm³ tumor volume in subcutaneous Trp53−/− Brca1−/− ID8 mouse models, animals were randomized into 4 therapeutic cohorts (n = 5 per cohort): 1. Vehicle control; 2. Talazoparib (0.33 mg/kg/day); 3. Ceralasertib (50 mg/kg/day); 4. Combination (Talazoparib 0.33 mg/kg/day + Ceralasertib 50 mg/kg/day). Longitudinal monitoring included biweekly measurements of body mass and tumor dimensions (calculated as length × width2/2). Terminal procedures involved retroorbital blood collection for blood count analysis, followed by 4% paraformaldehyde fixation of tumor and vital organs for histopathological assessment (IHC and H&E).
For the construction of OVCAR8 subcutaneous tumor model, 5 × 106 OVCAR8 sg Control or OVCAR8 sg ZBP1 cells were resuspended in a 1:1 mixture of PBS and Matrigel (Corning, 354234) and injected subcutaneously into female BALB/c nude mice to establish xenograft models. Female mice were used exclusively in this study as the research focuses on ovarian cancer, which is a female-specific malignancy. Upon reaching 100 mm3 tumor volume, animals were randomized into 4 therapeutic cohorts (n = 5 per cohort): 1. sg Control + Vehicle; 2. sg Control + Combination (Talazoparib 0.33 mg/kg/day + Ceralasertib 50 mg/kg/day); 3. sg ZBP1 + Vehicle; 4. sg ZBP1 + Combination (Talazoparib 0.33 mg/kg/day + Ceralasertib 50 mg/kg/day). Longitudinal monitoring included measurements of body mass and tumor dimensions (calculated as length × width2/2).
Zbp1 gene knockout (Zbp1−/−) C57BL/6 mice were purchased from GemPharmatech (Nanjing, China). Zbp1 wild-type (Wt) and knockout (Zbp1−/−) mice were randomly divided respectively, and both set up as control group (solvent) and experimental group (combination treatment of Olaparib 50 mg/kg/day plus Ceralasertib 50 mg/kg/day). Mice were randomly distributed into each group with a balanced representation of both sexes. Body weight was monitored regularly throughout the dosing period. At the end of the experiment, peripheral blood was collected through the retroorbital venous plexus for routine blood testing. Major organs were fixed with 4% formaldehyde for subsequent immunohistochemical detection and HE staining analysis.
Statistical analysis
GraphPad Prism 9.0 software was used for statistical analysis and data plotting. Data are shown as mean ± SEM. Statistical significance was determined by t tests (two-tailed) for two groups, one- or two-way ANOVA (with Dunnett’s or Tukey’s multiple comparisons tests) for three or more groups. P value < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD055407. All remaining data generated in this study are available within the Article and its supplementary files. Source data are provided with this paper.
References
Pilié, P. G., Tang, C., Mills, G. B. & Yap, T. A. State-of-the-art strategies for targeting the DNA damage response in cancer. Nat. Rev. Clin. Oncol. 16, 81–104 (2019).
Pearl, L. H., Schierz, A. C., Ward, S. E., Al-Lazikani, B. & Pearl, F. M. G. Therapeutic opportunities within the DNA damage response. Nat. Rev. Cancer 15, 166–180 (2015).
Xiong, J., Barayan, R., Louie, A. V. & Lok, B. H. Novel therapeutic combinations with PARP inhibitors for small cell lung cancer: a bench-to-bedside review. Semin. Cancer Biol. 86, 521–542 (2022).
Groelly, F. J., Fawkes, M., Dagg, R. A., Blackford, A. N. & Tarsounas, M. Targeting DNA damage response pathways in cancer. Nat. Rev. Cancer 23, 78–94 (2023).
da Costa, A. A. B. A., Chowdhury, D., Shapiro, G. I., D’Andrea, A. D. & Konstantinopoulos, P. A. Targeting replication stress in cancer therapy. Nat. Rev. Drug Discov. 22, 38–58 (2023).
Concannon, K., Morris, B. B., Gay, C. M. & Byers, L. A. Combining targeted DNA repair inhibition and immune-oncology approaches for enhanced tumor control. Mol. Cell 83, 660–680 (2023).
O’Connor, M. J. Targeting the DNA damage response in cancer. Mol. Cell 60, 547–560 (2015).
Luo, Y. et al. Neoadjuvant PARPi or chemotherapy in ovarian cancer informs targeting effector Treg cells for homologous-recombination-deficient tumors. Cell 187, 4905–4925.e24 (2024).
Carrassa, L. & Damia, G. DNA damage response inhibitors: Mechanisms and potential applications in cancer therapy. Cancer Treat. Rev. 60, 139–151 (2017).
Kim, H. et al. Targeting the ATR/CHK1 axis with PARP inhibition results in tumor regression in BRCA-mutant ovarian cancer models. Clin. Cancer Res. 23, 3097–3108 (2017).
Wilson, Z. et al. ATR inhibitor AZD6738 (Ceralasertib) exerts antitumor activity as a monotherapy and in combination with chemotherapy and the PARP inhibitor olaparib. Cancer Res. 82, 1140–1152 (2022).
Shah, P. D. et al. Combination ATR and PARP inhibitor (CAPRI): a phase 2 study of ceralasertib plus olaparib in patients with recurrent, platinum-resistant epithelial ovarian cancer. Gynecol. Oncol. 163, 246–253 (2021).
Mahdi, H. et al. Ceralasertib-mediated ATR inhibition combined with olaparib in advanced cancers harboring DNA damage response and repair alterations (olaparib combinations). JCO Precis. Oncol. 5, 1432–1442 (2021).
Wethington, S. L. et al. Combination ATR (ceralasertib) and PARP (olaparib) inhibitor (CAPRI) trial in acquired PARP inhibitor-resistant homologous recombination-deficient ovarian cancer. Clin. Cancer Res. 29, 2800–2807 (2023).
Zimmermann, M. et al. Guiding ATR and PARP inhibitor combinations with chemogenomic screens. Cell Rep. 40, 111081 (2022).
Kim, H. et al. Combining PARP with ATR inhibition overcomes PARP inhibitor and platinum resistance in ovarian cancer models. Nat. Commun. 11, 3726 (2020).
Bakhoum, S. F., Kabeche, L., Murnane, J. P., Zaki, B. I. & Compton, D. A. DNA-damage response during mitosis induces whole-chromosome missegregation. Cancer Discov. 4, 1281–1289 (2014).
Shi, C. et al. Aurora kinase A inhibition induces synthetic lethality in SMAD4-deficient colorectal cancer cells via spindle assembly checkpoint activation. Oncogene 41, 2734–2748 (2022).
Lee, S. et al. AIM2 forms a complex with pyrin and ZBP1 to drive PANoptosis and host defence. Nature 597, 415–419 (2021).
Wang, Y. & Kanneganti, T.-D. From pyroptosis, apoptosis and necroptosis to PANoptosis: A mechanistic compendium of programmed cell death pathways. Comput. Struct. Biotechnol. J. 19, 4641–4657 (2021).
Karki, R. et al. ZBP1-dependent inflammatory cell death, PANoptosis, and cytokine storm disrupt IFN therapeutic efficacy during coronavirus infection. Sci. Immunol. 7, eabo6294 (2022).
Pandeya, A. & Kanneganti, T. D. Therapeutic potential of PANoptosis: innate sensors, inflammasomes, and RIPKs in PANoptosomes. Trends Mol. Med. 30, 74–88 (2024).
Karki, R. et al. Synergism of TNF-α and IFN-γ triggers inflammatory cell death, tissue damage, and mortality in SARS-CoV-2 infection and cytokine shock syndromes. Cell 184, 149–168.e17 (2021).
Zheng, M., Karki, R., Vogel, P. & Kanneganti, T. D. Caspase-6 is a key regulator of innate immunity, inflammasome activation, and host defense. Cell 181, 674–687.e613 (2020).
Karki, R. & Kanneganti, T. D. PANoptosome signaling and therapeutic implications in infection: central role for ZBP1 to activate the inflammasome and PANoptosis. Curr. Opin. Immunol. 83, 102348 (2023).
Sundaram, B. et al. NLRP12-PANoptosome activates PANoptosis and pathology in response to heme and PAMPs. Cell 186, 2783–2801.e2720 (2023).
Sundaram, B. et al. NLRC5 senses NAD(+) depletion, forming a PANoptosome and driving PANoptosis and inflammation. Cell 187, 4061–4077.e17 (2024).
Kuriakose, T. et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci. Immunol. 1, aag2045 (2016).
Tang, Z. et al. ATR inhibition induces CDK1-SPOP signaling and enhances anti-PD-L1 cytotoxicity in prostate cancer. Clin. Cancer Res. 27, 4898–4909 (2021).
Yuan, F. et al. Z-DNA binding protein 1 promotes heatstroke-induced cell death. Science 376, 609–615 (2022).
Yasui, Y. et al. Autophosphorylation of a newly identified site of Aurora-B is indispensable for cytokinesis. J. Biol. Chem. 279, 12997–13003 (2004).
Bakhoum, S. F., Kabeche, L., Compton, D. A., Powell, S. N. & Bastians, H. Mitotic DNA damage response: at the crossroads of structural and numerical cancer chromosome instabilities. Trends Cancer 3, 225–234 (2017).
Vitale, I., Galluzzi, L., Castedo, M. & Kroemer, G. Mitotic catastrophe: a mechanism for avoiding genomic instability. Nat. Rev. Mol. Cell Biol. 12, 385–392 (2011).
Tanaka, K. et al. Targeting Aurora B kinase prevents and overcomes resistance to EGFR inhibitors in lung cancer by enhancing BIM- and PUMA-mediated apoptosis. Cancer Cell 39, 1245–1261.e1246 (2021).
Bertran-Alamillo, J. et al. BID expression determines the apoptotic fate of cancer cells after abrogation of the spindle assembly checkpoint by AURKB or TTK inhibitors. Mol. Cancer 22, 110 (2023).
Carmena, M., Wheelock, M., Funabiki, H. & Earnshaw, W. C. The chromosomal passenger complex (CPC): from easy rider to the godfather of mitosis. Nat. Rev. Mol. Cell Biol. 13, 789–803 (2012).
McVey, S. L., Cosby, J. K. & Nannas, N. J. Aurora B tension sensing mechanisms in the kinetochore ensure accurate chromosome segregation. Int. J. Mol. Sci. 22, 8818 (2021).
Li, T. et al. Phosphorylation and chromatin tethering prevent cGAS activation during mitosis. Science 371, eabc5386 (2021).
Lim, Y. et al. Phosphorylation by Aurora B kinase regulates caspase-2 activity and function. Cell Death Differ. 28, 349–366 (2021).
Tsukahara, T., Tanno, Y. & Watanabe, Y. Phosphorylation of the CPC by Cdk1 promotes chromosome bi-orientation. Nature 467, 719–723 (2010).
Matthews, H. K., Bertoli, C. & de Bruin, R. A. M. Cell cycle control in cancer. Nat. Rev. Mol. Cell Biol. 23, 74–88 (2022).
Ardito, F., Giuliani, M., Perrone, D., Troiano, G. & Lo Muzio, L. The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy (Review). Int. J. Mol. Med. 40, 271–280 (2017).
Liu, R. et al. ZBP1-mediated apoptosis and inflammation exacerbate steatotic liver ischemia/reperfusion injury. J. Clin. Investig. 134, e180451 (2024).
Karki, R. et al. ADAR1 restricts ZBP1-mediated immune response and PANoptosis to promote tumorigenesis. Cell Rep. 37, 109858 (2021).
Fang, Y. et al. Sequential therapy with PARP and WEE1 inhibitors minimizes toxicity while maintaining efficacy. Cancer Cell 35, 851–867.e857 (2019).
Acknowledgements
For schematic drawing, the authors thank the Biorender website (https://biorender.com/) for abundant image materials. This study was supported by the National Science Foundation of China (82441045 to Y.F., 82272707 to Y.F., 81874109 to Y.F., 81772787 to Q.G., and 82072889 to Q.G.), the National Key Technology Research and Development Program of China (2022YFC2704200 and 2022YFC2704205 to Q.G.), and Innovation Group Project of Hubei Province (2023AFA036 to Y.F.). M.L. is a member of the Fonds de Recherche du Québec-Santé–funded (FRQS-funded) Centre de Recherche du CHUS and is supported by the Institut de Recherche sur le Cancer de l’Université de Sherbrooke, Canada Research Chair on Development of Personalized Therapies for Ovarian Cancer Patients, Stand up to Cancer, and Natural Sciences and Engineering Research Council of Canada. G.B.M. was supported by the Breast Cancer Research Foundation and a kind gift from the Sheldon and Miriam Adelson Medical Research Foundation.
Author information
Authors and Affiliations
Contributions
W. Gong, Z. Liao, G. Zhao, and Y. Fang conceived the study. D. Liu, G. Zhao, Q. Gao, and Y. Fang supervised the investigation. W. Gong, Z. Liao, F. Xiong, and L. Wang carried out and validated the experiments with the help of S. Shi, M. Luo, Y. Zhang, Q. Xu, Y. Wang, Y. Wu, L. Zhu, and D. Kuang. W. Gong, Z. Liao, and F. Xiong wrote the manuscript, and M. Labrie, G.B. Mills, D. Ma, G. Zhao, and Y. Fang revised the manuscript. D. Liu, G. Zhao, Q. Gao, and Y. Fang acquired the funding.
Corresponding authors
Ethics declarations
Competing interests
G.B.M.—SAB/Consultant: AstraZeneca, BlueDot, Chrysallis Biotechnology, Ellipses Pharma, ImmunoMET, Infinity, Ionis, Lilly, Medacorp, Nano-string, PDX Pharmaceuticals, Signalchem Lifesciences, Tarveda, Turbine, Zentalis Pharmaceuticals; Stock/Options/Financial: Catena Pharmaceuticals, ImmunoMet, SignalChem, Tarveda, Turbine; Licensed Technology: HRD assay to Myriad Genetics, DSP patents with Nano-string. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hiroshi Katayama and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gong, W., Liao, Z., Xiong, F. et al. Aurora kinase B phosphorylates ZBP1 to drive PANoptosis following treatment with PARP and ATR inhibitors combination. Nat Commun 16, 11672 (2025). https://doi.org/10.1038/s41467-025-66591-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66591-1