Abstract
Mass General Brigham, an integrated healthcare system based in the Greater Boston area of Massachusetts, annually serves 1.5 million patients. We established the Mass General Brigham Biobank (MGBB), currently encompassing 142,238 participants, to unravel the intricate relationships among genomic profiles, environmental context, and disease manifestations within clinical practice. In this manuscript, we described genetic and phenotypic diversity and their interaction in the MGBB by employing population genetics, geospatial assessment, and association analyses of rare and common genetic variants. The population genetic variation captured the demographic histories in the Greater Boston area throughout American history, highlighting communities tied to shared genetic and environmental factors. Furthermore, our phenome-wide association analysis with socioeconomic factors, as well as common and rare genetic variants, prioritized disease causes across a tertiary academic medical center and community-based health system. Our investigation underscores the potency of unbiased, large-scale analyses in a healthcare-affiliated biobank, elucidating the dynamic interplay across genetics, immigration, structural geospatial factors, and health outcomes in one of the earliest American sites of European colonization.
Similar content being viewed by others
Introduction
Determinants of health include a complex interplay of sociodemographic, structural, genetic, and environmental factors that are also contextually dependent on time and geography. Disease risk prediction models and therapeutic paradigms are largely agnostic to many of these important features yet are intended for broad use. Such training datasets often lack the breadth and depth of information and the inherent diversity across features required for equitable applications. The United States populace is highly diverse, marked by complex migration patterns and dynamic social constructs and represents multilevel health contributors. For example, it is widely recognized that the prevalence of diseases is closely linked to individual or neighborhood social deprivation, which further varies across smaller domains and regions1,2. Furthermore, these determinants differentially contribute to health outcomes depending on local factors3,4.
Contemporary healthcare-associated biobanks represent a new opportunity to discover novel determinants of health and augment translation to clinical care. Such endeavors represent a recent collaborative synergy of large-scale population-based5,6,7,8 and local healthcare-biobanks9,10,11,12,13. Understanding how DNA sequence variation tracks with contemporary and historical population demographics can provide insights into differential disease burdens. For example, rs5742904 (c.10580G>A, p.Arg3527Gln) in APOB, a founder pathogenic mutation for familial hypercholesterolemia has significantly higher allele frequency in Old Order Amish people. It substantially explains the increased risk for coronary artery disease in this population14,15. Important insights related to genetic variations and clinical outcomes often require profiling diverse participants. For example, G6PD deficiency16 has long been recognized as a prevalent genetic hemolytic disease in Sub-Saharan Africa. A more recent example is the discovery of the association between disruptive variants in PCSK917,18 and reduced coronary artery disease risk in West African ancestry, where these variants are more prevalent, which has facilitated the development of novel therapeutics. Studying diverse participants across a spectrum of diseases is crucial to assess the penetrance of disease-associated monogenic alleles19 and polygenic models20.
Recent analyses of biobanks in the United States have uncovered the complex genetic structure of Hispanic and/or Latinx groups tracing their origins to the Americas9,12. In these efforts, it has been demonstrated that the fine genetic structure within biobanks can identify varying disease risks by capturing both genetic and social structures, thereby contributing to the advancement of personalized medicine. Separately, recent advances in data size and methodology have enabled us to precisely characterize the complex population dynamics associated with multiple colonization and admixture events21,22,23. However, the interplay across these features, or their interaction with large-scale genetic association studies using whole-genome imputed or sequenced data, remains understudied.
The Greater Boston area represents among the earliest European colonization of the United States with sequential ongoing migration from diverse groups. In this study, we examined genetic variation coupled with sociodemographic, clinical, and environmental/geospatial factors in the Mass General Brigham Biobank (MGBB). By applying a network-based clustering algorithm with recent reference datasets, we established fine genetic clusters with subcontinental resolution within MGBB. These clusters exhibited distinct genetic properties, geographic distributions, and genetic disease risks. In combination with rare and common variant genetic association analyses, we gained further insights into the different disease risks among these clusters. Collectively, this study highlights the power of large-scale, unbiased analyses within a healthcare-based biobank to understand the complex interplay between genotypes and phenotypes, paving the way for increasingly personalized interventions.
Results
Participant recruitment and electronic health record (EHR) based phenotyping
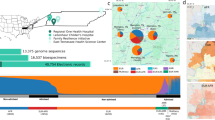
Since 2010, 142,238 individuals within the Mass General Brigham (MGB) network, the largest healthcare system in Massachusetts, have consented to participate in the MGBB as of May 11, 2023 (Fig. 1a, Supplementary Table 1). Among participants, 99.5% (n = 141,519) consented to re-contact. 56.8% of participants were female (n = 80,851, Supplementary Fig. 1). Median age at consent was 51 (interquartile range; IQR 35–63) years for female participants and 58 (43–68) years for males. Self-reported races were 84.4% White, 4.5% Black, and 3.0% Asian. Self-reported ethnicities were 86.6% non-Hispanic and 2.44% Hispanic (Supplementary Fig. 1). The participants are primarily cared for at the two flagship MGB hospitals, both located in Boston, MA, and their associated clinics – Massachusetts General Hospital (MGH) and Brigham and Women’s Hospital (BWH, Fig. 1b). The biobank data is interlinked with EHR encompassing phenotype data across the MGB network, as well as notable specialty centers in Boston, MA, including the Mass Eye and Ear Institute (MEEI) and Dana-Farber Cancer Institute (DFCI). To generate systemically annotated prevalent/incident outcomes, we extracted International Classification of Diseases codes, Ninth (ICD9) and Tenth (ICD10) revisions, from the EHR and mapped them to PheCodes24. We identified 1577 of 1860 possible PheCodes with at least one event (Fig. 1c). The median number of ICD codes per PheCode was 7 (range, 1–134). We identified a median of 34 (IQR 11–74) prevalent events per person. Participants were followed for a median of 4.29 (IQR 2.47–5.9) years after inclusion to MGBB with a median of 10 (3–23) incident events per person. Precise geolocation information is also available in a large fraction of MGBB participants (N = 118,102) and 82.7% (97,729/118,102) were from Massachusetts and 73.4% (71,735/97,729) of those were from the greater Boston area (Fig. 1d).
a The number of the participants in MGBB. The columns represent the cumulative number of individuals who have consented to the MGBB. Colors indicate the vital status of participants as of July 2023. b The number of hospital encounters of participants the MGB Network. The left bar chart indicates the total number of encounters by hospital. The top bar chart indicates the number of individuals presented to the hospitals in the combinations shown in the middle panel. Please note that these encounters include sites where recruitment did not take place. c The number of PheCodes outcomes in MGBB. The columns indicate the number of outcomes in PheCode-category. Colors distinguish between incident and prevalent cases. d The geographical distribution of the participants in MGBB across Greater Boston. The point indicates the geolocation of the participants and colors indicate the number of the participants at each point. MGH Massachusetts General Hospital, BWH Brigham and Women’s Hospital, MEEI Mass Eye and Ear Institute, FH Faulkner Hospital, NWH Newton-Wellesley Hospital, DFCI Dana-Farber Cancer Institute. Basemap: Stamen Terrain via Stadia Maps. © Stadia Maps © Stamen Design © OpenMapTiles © OpenStreetMap contributors.
Fine-scale clustering of genetic similarity in MGBB
Extending beyond traditional low-dimensional projections of genetic similarity from genome-wide data, we utilized high-dimensional principal components (PCs) to achieve greater granularity. Using genome-wide genotyping arrays, we genotyped 53,306 participants in the MGBB. By employing the top 30 genetic PCs and a network-based clustering approach25, we identified 30 data-driven genetically similar clusters (Fig. 2a, Supplementary Figs. 2 and 3, Supplementary Tables 2 and 3). In conventional PC spaces, these clusters are located in overlapping regions, indicating the value of incorporating higher-dimensional information for a better description of genetic clusters and their histories. The largest cluster (cluster 1, ordered by sample size) includes 11,875 (22.3%) MGBB individuals. The smallest (cluster 30) includes only one MGBB participant as well as 27/27 reference Sardinian individuals from Human Genome Diversity Project (HGDP)26, suggesting the origin of this individual. As such, unsupervised clustering with diverse participants from the 1000 Genomes Project Phase 3 (1KG)27 and HGDP reference panels28 allowed us to infer the genetic similarity between these clusters and participants worldwide in an unbiased manner.
a PCA representation of the genetic clusters in MGBB. Each dot represents a participant, with colors indicating distinct genetic clusters identified through graph-based clustering from genetic principal components (Methods). The numbers indicate cluster identification. The detailed cluster information will be found in Supplementary Table 3. b Population differentiation in MGBB revealed by ADMIXTURE analysis. The heatmap displays the proportions of ADMIXTURE components (K = 10) within each genetic cluster. The columns at the top of the heatmap represent the number of MGBB participants in each cluster. c Effective population size estimated using coalescent pattern in MGBB genetic clusters. The horizontal axis shows generations ago from the present. The vertical axis shows the effective population size. Black lines indicate estimates, and gray lines indicate 95% confidence interval. The numbers on the top of charts indicate genetic cluster. UKBB British and Irish are the estimates from self-reported White British and Irish in UKBB. PC principal component, 1KG 1000 genomes project, HGDP Human Genome Diversity Project, MGBB Mass General Brigham Biobank, UKBB UK Biobank.
Cluster 1 (N = 11,875) was genetically similar to the Western European participants in the reference dataset [CEU (Utah residents with Western or Northern European ancestry) and GBR (British from England or Scotland) in 1KG, French and Orcadian in HGDP, Supplementary Fig. 3]. Cluster 2, the second largest cluster (N = 10,802), includes GBR as cluster 1, but is more enriched by Orcadian (northern Scottish origin) suggesting different origin of these large European groups in Boston. In addition to cluster 1 and 2, we identified eight European-like clusters that cluster with Italian (cluster 3, N = 5757), Russian (cluster 4, N = 4142), Spanish (cluster 13, N = 968), Adygei (cluster 17, N = 468), Finnish (cluster 25, N = 35), Basque, and Sardinian reference participants reflecting known patterns of demography in the Greater Boston Area. We also identified two distinct Admixed-American-like clusters: Cluster 6, with a population of 3211, enriched with PUR (Puerto Rican in Puerto Rica); and Cluster 9 comprising 1184 individuals with Colombian, Maya, PEL (Peruvian in Lima, Peru); Pima, CLM (Colombian in Medellín, Colombia), and MXL (Mexican ancestry in Los Angeles, CA). Additionally, four African-like clusters were identified: Cluster 7, with 2151 individuals, enriched with African Caribbean in Barbados and African Ancestry in Southwest USA; Cluster 18, specific to Nigerian Africans (N = 64, Esan in Nigeria, Yoruba in Ibadan, Nigeria); Cluster 23, specific to Kenyan Africans (N = 40, Bantu and Luhya in Webuye, Kenya); and Cluster 19 with other Western Africans (N = 15, Mandinka, Mende people in Sierra Leone, Gambian in Western Divisions in the Gambia). Furthermore, three East-Asian-like clusters were identified: Cluster 20, with 175 individuals, specific to Japanese; Cluster 21, comprising 240 individuals, specific to Uygur; and Cluster 10, including 749 individuals, with other East Asians. We identified a single large cluster (cluster 11, n = 560) enriched in South Asian reference participants.
Even with a diverse reference dataset, eight clusters comprising 9874 (18.8%) MGBB participants did not have enrichments of specific populations from the reference dataset. Among these un-annotated clusters, seven clusters (5, 8, 12, 14, 15, 16, and 22) exhibited genetic similarities to European populations. We calculated pairwise Fixation Index (FST) values among clusters, and then constructed a phylogenetic tree of the clusters (Supplementary Fig. 4a). The observed population differentiation between clusters further corroborates the genetic similarities between reference populations and residents of Greater Boston.
To infer continuous population structures within these genetic clusters, we conducted ADMIXTURE29 analyses, many of which show similar patterns of structure across increasing numbers of ancestral components (Supplementary Fig. 4b). Using cross-validation, ten was the best fit number of components (Fig. 2b and Supplementary Fig. 5). We identified two European components (distinguished by components 1 and 9). The component 1 was most enriched in the Finnish-like cluster (cluster 25), and relatively enriched in northern European-like clusters (1, 2, and 4) more than the southern European-like clusters (3, 13, and 15). In contrast, component 9 was enriched in the southern European-like cluster. We also observed a third component included in the European-like cluster (component 3), which is prominent in the Kalash (Indo-European in northwest Pakistan) and other Pakistani reference populations. While this was enriched in southern European-like clusters, it was more enriched in un-annotated European-like genetic clusters 5, 8, 12, 14, and 22 than other annotated European genetic clusters, possibly consistent with Middle Eastern origins as this group is poorly represented in reference datasets.
Cluster 5—the 5th largest cluster in this study (n = 3514)—is one of such un-annotated European-like clusters. By comparison of allele frequencies between gnomAD30 and our dataset, we found that cluster 5 has allele frequencies most similar to the Ashkenazi Jewish reference population (Supplementary Table 4). We also observed strong enrichment of skin neoplasms and inflammatory bowel diseases which are known to be enriched in established Ashkenazi cohorts (Supplementary Fig. 6a). We also observed significant enrichment of Ashkenazi Jewish founder mutations (e.g., APC I1307K, c.3920 T > A, p.Ile1307Lys, BRCA1 185delAG, c.68_69del, p.Glu23fs, BRCA1 5382insC, c.5266dup, p.Gln1756fs, BRCA2 6174delT, c.5946del, p.Ser1982fs)31,32 in this genetic cluster (Supplementary Fig. 6b). These founder mutations were also enriched in un-annotated European-like clusters 12 and 15 suggesting close genetic relationships between these clusters to the Ashkenazi-Jewish-like cluster 5.
Effective population size of genetic clusters in the Greater Boston area
To characterize the genetic clusters observed in the MGBB, we estimated the historical transition of effective population size of each cluster using genome-wide genetic data. We conducted identity by descent (IBD) based estimation for effective population sizes (Ne, Fig. 2c, and Supplementary Fig. 7, Supplementary Fig. 8). Our results were consistent with some prior results conducted outside of the U.S. For example, we replicated a previously described bottleneck event in the Ashkenazi-Jewish-like participants (cluster 5, Fig. 2c). The lowest Ne was estimated to be 1170 (95% confidence interval; 95%CI 1100–1270) individuals 28 generations ago33. We observed similar bottleneck events in clusters 12 and 15 around the same generation [minimal Ne was 4,510 (95%CI 4220–5130) in cluster 12 and 32,600 (30,000–35,700) in cluster 15] consistent with the aforementioned sharing pattern of Ashkenazi founder mutations with cluster 5. The largest genetic cluster 1 indicates a population bottleneck occurring approximately 12 generations ago. Notably, the cluster 2 which is the second largest in MGBB and genetically similar to cluster 1 does not exhibit such a bottleneck, suggesting different historical trajectories for these genetically similar participants. This timeframe coincides with the initial colonization of the Boston area by British settlers. This event is not evident in the British or Irish participants from the UK Biobank (UKBB) here or in previous studies34,35 (Fig. 2c), suggesting a unique founder event among British Americans likely due to initial U.S. colonization. We also observed a significant bottleneck event in the Admixed-American participants, specifically in clusters 6 and 9, with a pronounced magnitude in the Puerto-Rican-like cluster 6 [minimal Ne was 11,300 (11,100–11,600)]. However, we did not observe such bottlenecks for other clusters.
Genetic clusters, geographic and socioeconomic factors and disease risks
Geospatial information provides valuable insights into the environment surrounding the participants. First, to contextualize genetically similar participants and geolocation information, we used geospatial scan statistics to explore the geographical structure of MGBB genetic clusters. We observed 22 statistically significant regions of geographical enrichment among 13 genetic clusters in smaller than 4-km radius areas in the Boston area (Fig. 3a, Supplementary Fig. 9). We observed concentrations of genetically similar participants, which recapitulate the colonization and segregation histories of the Greater Boston Area. One example of strong enrichment was observed in the southern area (Roslindale/Mattapan/Dorchester and separately Roxbury) by cluster 7 enriched by ACB/ASW (African Caribbean in Barbados and African ancestry in Southwest U.S., expected number 105 and observed number 725, P < 1 × 10−17). Another strong enrichment is observed north of Boston (Charlestown/Chelsea) by cluster 6 enriched by PUR (Puerto Rican in Puerto Rico, expected number 81 and observed number 224, P < 1 × 10−17). This enrichment also extends to Boston’s South End and further to Roxbury, Hyde Park, and Jamaica Plain, potentially reflecting the historical impact of residential segregation in these areas. We also observed enrichment of Ashkenazi-Jewish-like (cluster 5) and East-Asian-like clusters (cluster 10) in areas seeded by early founding communities, such as Back Bay/Brookline (cluster 5) and Allston (cluster 10). In the Cambridge area, we observed significant enrichment of multiple clusters (clusters 1, 5, 10, 11, 12, and 13), potentially related to the high density of academic institutions and the commercial biomedical industry attracting multiple communities and individuals.
a Geographical enrichment of genetically similar groups in the greater Boston area. The circles indicate the areas with significant enrichment after multiple testing correction for the corresponding genetic sub-clusters. The geographic enrichment was scanned in unbiased manner. Only genetic clusters with significant enrichment were shown. b Social Deprivation index (SDI) map across Massachusetts and the Greater Boston area. The color indicates SDI value, with darker red indicating higher deprivation status (higher SDI). c Phenome-wide association analysis for SDI. We tested the logistic linear association between the SDI and 1561 PheCode-based outcomes (prevalence and incidence) in MGBB participants. The model was adjusted for age, sex, and the first ten genetic principal components. An association was considered statistically significant if P-value was less 3.2 × 10−5 (0.05/1561). The color of the bars indicated the direction of the effect of SDI and red color indicates positive associations between SDI and disease risk and blue indicates negative associations (higher SDI suggests higher deprivation). The P-value for the SDI term was computed by multivariable logistic regression model. The exact P-values are listed in Supplementary Table 5. d The disease frequency (prevalence + incidence) by deprivation status and genetic similarities. The color of bars indicated deprivation status (higher or lower than the median SDI). The largest 8 clusters are displayed. Complication of Birth, Other and unspecified complications of birth; puerperium affecting management of mother (PheCode 654). Basemap: Stamen Terrain via Stadia Maps. © Stadia Maps © Stamen Design © OpenMapTiles © OpenStreetMap contributors.
The western European-like clusters cluster 1 and cluster 2 were similar in conventional PC space (Supplementary Fig. 2) and ADMIXTURE analysis (Supplementary Fig. 4b)29, but well differentiated by network-based clustering (Fig. 2a and Supplementary Fig. 2) as well as geospatially. The CEU/GBR-like cluster 1 was enriched in central areas of the Boston (Beacon Hill), representing the earliest sites of British colonization. Cluster 2 (Orcadian-like, tagging northern populations of the British Isles including those hailing from Scotland and Ireland) is enriched in two different geographical locations, including Chelsea and South Boston, similarly potentially reflecting historical residential segregation of Irish immigrants during a secondary wave of immigration. These different geographical enrichments of cluster 1 versus cluster 2 reflect the distinct recent histories of these two genetically close European-like participants aligned with population size estimation.
Socioeconomic status was correlated with geographic distributions1. Using geocoded location information for each participant in our study, we calculated a Social Deprivation Index (higher SDI indicating greater deprivation) for each participant (Fig. 3b). To systemically identify the associations between socioeconomic status and disease risk in MGBB, we associated SDI with phenome-wide outcomes captured by EHR, adjusting for genetic principal components. We found SDI was significantly associated with 402 out of 1561 phenome-wide outcomes (Bonferroni P < 0.05/1561 = 3.2 × 10−5, Supplementary Figs. 10a and 10b, Supplementary Table 5). Among 402 significant associations in the MGBB, 369 associations were tested in the external dataset (UKBB). 98.1% of the associations showed directional concordance and 84.8% were significantly replicated (P < 0.05/369 = 1.3 × 10−4, Supplementary Fig. 10c). Greater SDI was generally associated with increased disease prevalence and incidence (388 out of 402). The strongest SDI-associated PheCodes was with Tobacco use disorder [PheCode 318, odds ratio (OR) per one standard deviation (SD) of SDI was 1.53 (95% CI 1.48–1.59)], followed by Mood disorders [PheCode 296, OR = 1.26 (95% CI 1.23–1.30)], and Depression [PheCode 296.2, 1.25 (1.22–1.29)]. We observed stronger associations for PheCodes categorized under ‘Mental Disorders’, followed by ‘Uncharacterized Symptoms’, ‘Respiratory Systems’, and ‘Circulatory Systems’ in deprived individuals (Fig. 3c). No PheCodes in Congenital Anomalies were associated with SDI. However, several PheCodes in ‘Neoplasms’ or ‘Pregnancy Complications’ were inversely associated with SDI. For example, ‘Prostate Cancer’ [PheCode 185, 0.85 (0.80–0.90)] or ‘Other and unspecified complications of birth puerperium affecting management of mother and postpartum’ [PheCode 654, 0.71 (0.66–0.76)] had significant/nominal inverse associations with SDI (Fig. 3d).
Using coronary artery disease (CAD) as an example of a common complex disease condition, we identified a significant association between SDI and CAD independent from clinical and genetic risk. The association remained significant even after adjustments for clinical risk score (Pooled Cohort Equation, PCE)36, and polygenic risk score37 [PRS, OR1SD-SDI 1.24 (1.16–1.34), OR1SD-PCE 1.69 (1.60–1.78), OR1SD-PRS 1.51 (1.39–1.63), in the multivariate model adjusted by the first ten genetic PCs, Supplementary Fig. 10d].
Exome sequencing in MGBB
Using high-coverage whole-exome sequencing in the same group of individuals, we systemically identified rare coding variants in MGBB. There were significant differences in variant distributions across clusters (alternate allele count, Supplementary Fig. 11a). For instance, the Ashkenazi-Jewish-like cluster 5 had fewer singleton variants [median 21 (IQR 17–25) for cluster 4, and 75 (52–105) for others in all individuals, 139 (125–156) and 467 (387−557) for down sampled cohort. Supplementary Figs. 11b and 11c]. In contrast, there were significantly more singletons in clusters 12 and 15, even though they are closely related to cluster 5 [324 (299–357) and 400 (370–425), respectively].
We identified median 15 [12–18] rare (Minor allele frequency, MAF < 0.01), high-confidence autosomal predicted loss-of-function (pLoF) variants per participant (Supplementary Fig. 11d). Overall, we observed higher number of alternate allele-counts in AFR or AMR participants consistent with previous reports. We observed 2 times higher number of synonymous variants in AFR participants, but this trend is milder in missense or pLoF variants (Supplementary Fig. 11d). The largest number of pLoF variants were observed in African-like clusters [median 23 (IQR 20.25–25.75) in cluster 23, 22 (17–24) in cluster 18, 20 (19–22) in cluster 27, 20 (17.0–23.0) in cluster 7]. The Northern European-like clusters 2 and 1 had the fewest pLoFs [13 (11.0–16.0) in cluster 2 and 14 (11–17) in cluster 1]. We also identified 1,425 individuals (2.8% of total participants) with at least one rare autosomal pLoF homozygous genotype across 760 genes.
We next explored established pathogenic variants (Fig. 4a) in MGBB. 2.6% (1318/50,625) of participants carry a potentially actionable pathogenic/likely pathogenic variant (actionable findings) per the American College of Medical Genetics and Genomics secondary findings guideline (ACMG SF, version 3.1)38,39. These included 6 homozygotes (one TP53, one LDLR, and 4 MUTYH), and 7 potential compound heterozygotes (2 BTD, 3 MUTYH, one ATP7B, and one GAA). Across genetic clusters, we observed substantial differences in the prevalence of these pathogenic variants (Fig. 4b). In general, European participants showed higher rates for known actionable findings. Despite lower numbers of alternate allele counts in the European participants, the genetic diagnostic rate was significantly higher in European participants (Fig. 4c). Namely, pLoF variants on ACMG SF v3.1 genes found in African-like and Admixed-American-like participants have significantly lower likelihood of being annotated with a high-quality (more than equal two-stars) pathogenic/likely pathogenic annotation in comparison to the European-like participants [ORAfrican-like = 0.27 (95%CI 0.18–0.41) and ORAdmixed-American-like = 0.48 (0.35–0.65), tested by Fisher’s exact test, Fig. 4d], at least partly related to the overrepresentation of causative variants recurrently observed in European participants in the ClinVar40,41 database. In addition, we observed variability in the prevalence of actionable findings among European-like sub-clusters. The highest prevalence of actionable findings was observed in clusters 12 (4.7%) and 5 (4.4%). Conversely, clusters 3 (2.4%) and 2 (2.1%) showed a lower prevalence of actionable findings than the mean rate over the MGBB (2.6%).
a Pathogenic variants identified in the MGBB. The heights of bars indicate number of carriers of pathogenic/likely-pathogenic variants for ACMG actionable genes. The colors of the bars indicate the mode of inheritance of the genes. The numbers after the gene symbols also indicate carrier counts. b Carrier frequency of individuals with pathogenic variants in ACMG actionable genes, categorized by genetic similarity. The dotted line represents the average frequency in the MGBB. c Number of pLoF variants in ACMG actionable genes with clinical annotations. The heights of bars indicate the proportion of the annotated variants in the observed pLoF variants in ACMG actionable genes. In (b) and (c), only the genetic clusters with more than 100 individuals are shown. d Annotation rate of pLoF variants in ACMG actional genes. The squares indicate estimated odds ratios in reference to European-like participants. The error bars indicate 95% confidence interval. The odds ratio and 95% confidence interval were estimated by two-sided Fisher’s exact test. e Summary of phenome-wide gene burden testing in the MGBB. We conducted exome-wide phenome-wide association analysis across 1482 PheCodes based outcomes in 15,866 genes. The heights of bars indicate the number of significant associations (P < 2.16 × 10−9 = 5 × 10−2/23,191,897 phenotype-transcript pairs) for designated genes. The color of each column corresponds to the associated PheCode-category. The P-value for the rare deleterious-variant burden was calculated using logistic regression implemented in the REGENIE software. The exact P-values are listed in Supplementary Table 6. MGBB Mass General Brigham Biobank, ACMG American College of Medical Genetics and Genomics, pLoF predicted loss of function, AFR African-like, AMR Admixed-American-like, EAS East-Asian-like, EUR European-like, SAS South-Asian-like.
To understand the clinical consequences of rare coding variants, we performed exome-wide and phenome-wide association study (PheWAS) across 1482 PheCodes and 15,866 genes in 50,622 participants (“Methods”). We identified 71 significant associations (P < 2.16 × 10−9, 0.05/23,191,897 tested phenotype-transcript pairs, Supplementary Table 6) in the burden of rare pLoF and deleterious missense variants with 17 genes, which included 9 ACMG SF v3.1 genes across 60 clinical outcomes (Fig. 4e). In addition to the genes associated with known traits, we found significant associations between PTEN deleterious variants and increased risk for secondary hypothyroidism. This link was not described by previous rare variant targeted analysis42,43 while PTEN deleterious variants have been known to be causal for hamartoma syndrome including thyroid cancers and abnormalities44,45. Nevertheless, we highlight numerous persistent risk signals from known Mendelian mechanisms of disease in MGBB. For these associated trait-gene pairs, we assessed the penetrance and prevalence to further describe the rare-variant burden on the disease in clinical settings (Supplementary Fig. 12). We observed a wide range of penetrance (3.88% to 100%) and generally low prevalence (0.197% to 19.0%).
Genome-wide PheWAS in MGBB
To further explore the relationship between genotype and phenotype in MGBB, we conducted a comprehensive genome-wide PheWAS using ICD code-based PheCodes. We associated over 20 million common variants with 1483 PheCodes in 52,374 participants, which were imputed using the TOPMed imputation reference panel (r2, “Methods”)46. We identified 140 associations that reached genome-wide significance (P < 3.37 × 10−11, 5 × 10−8/1483 tested phenotype, Fig. 5a, Supplementary Fig. 13a, Supplementary Table 7). We refined the prognosis of identified known low-frequency monogenic variants. For instance, we observed that the variant rs6025 (F5, c.1601 G > A, p.Arg534Gln; Factor V Leiden) is strongly associated with Congenital deficiency of other clotting factors, including factor VII [OR = 12.21 (95%CI 9.98–14.94), P = 1.6 × 10−75]. Similarly, rs113993960—a pathogenic variant in CFTR (c.1521_1523del, p.Phe508del)—is associated with Cystic fibrosis [OR = 14.87 (11.98–18.46), P = 4.3 × 10−95].
a Genome-wide and Phenome-wide scan in the MGBB. We conducted association analysis between common genetic variants (Minor allele counts ≥40) and 1483 PheCodes (Case counts ≥60). The columns represent the number of significant associations (P < 3.37 × 10−11) on each chromosome. We annotated chromosomes with the representative locus in the chromosome. The exact P-values are listed in Supplementary Table 7. b Manhattan plot of GWAS for Rhesus isoimmunization during pregnancy in women (n = 29,159). The horizontal axis displays the genomic coordinates from chromosome 1 to chromosome X. The vertical axis represents the strength of association in negative log10 P-value. The significantly associated variants in the RHD locus are highlighted. The P-values for the imputed genotype dosage were calculated using the logistic regression model implemented in the REGENIE software. c Odds ratio for Rhesus isoimmunization during pregnancy by rs55794721 genotypes. The dots and error bars represent the estimated odds ratios and 95% confidence intervals compared to the reference homozygotes ([C/C]). d RHD read counts from Whole Blood RNA sequence data obtained from the GTEx dataset. The horizontal axis displays the number of reads aligned to the RHD gene, categorized by rs55794721 genotypes. e Frequencies of rs55794721 homozygotes across genetic clusters in MGBB. The horizontal axis corresponds to the genetically similar groups in MGBB, while the vertical axis represents the group wise frequency of rs55794721 homozygotes ([G/G]). MGBB Mass General Brigham Biobank, ACMG American College of Medical Genetics and Genomics, AFR African-like, AMR Admixed-American-like, EAS East-Asian-like, EUR European-like, SAS South-Asian-like, GWAS Genome-Wide Association Study, GTEx Genotype-Tissue Expression.
Some of these variants exhibited a pronounced recessive effect on the phenotype. A prime example is the variant rs55794721, which is associated with Rhesus isoimmunization in pregnancy (Fig. 5b). This medical condition exemplifies recessive inheritance resulting from the deletion of the RHD gene. As anticipated, the OR for heterozygotes was not significant [ORHetero = 0.976 (95%CI 0.553–1.72)] compared to the strong effect observed in homozygotes [ORHomo = 18.9 (95%CI 12.4–29.7), Fig. 5c]. Recent large-scale sequencing analysis of structural variants47 identified high linkage disequilibrium (LD) between rs72660908 and a large deletion affecting RHD (R2 > 0.99). Supporting this, we confirmed very low expression level of RHD48 in whole blood transcriptome associated with risk allele at rs72660908 (G, Fig. 5d) and median 0 coverage by exome sequencing in MGBB (Supplementary Fig. 13b). We also observed a significant enrichment of cases among individuals who were homozygous for rs55794721, with 145 out of 202 cases having the A/A genotype at this locus. As previously reported, individuals with the homozygous alternate allele for rs55794721 were dominantly observed in continental European-like participants and we observed very few copies in African-like/East-Asian-like participants. Furthermore, the frequency fluctuates even among European-like participants as illustrated by different frequencies among clusters we denote as European-like. The highest prevalence was observed in cluster 16 (17.7%) followed by cluster 1 (17.1%) and lowest in cluster 5 (9.0%, Fig. 5e), suggesting different genetic risk architecture among European-like sub-clusters.
Another noteworthy example is the association between rs73404549 and sickle cell anemia. This variant is in strong LD with rs334 (HBB c.20 A > T, p.Glu7Val, HbS), a well-established pathogenic variant for sickle cell anemia. Despite high medical relevance, rs334 was not included in the TOPMed reference panel. We re-evaluated the impact of rs334 using exome sequencing data on sickle cell anemia and clinical red blood cell counts. rs334 showed stronger and larger effect size for sickle cell anemia than imputed rs73404549 (ORrs334 = 63.41 ± [39.22–102.53], Prs334 = 1.28 × 10−151, βrs73404549 = 28.96 [19.72–42.52], Prs73404549 = 1.89 × 10−64).
Discussion
In this study, we conducted multidimensional investigations into the structure of a modern healthcare-based biobank based at one of the earliest sites of durable European colonization. We show how expanded immigrant communities in the U.S. often exhibit genetic similarities to contemporary continental populations and reflect common bottlenecks. However, we also observe distinct bottlenecking effects of early colonization and patterns of admixture, and identify genetically similar participants groups not well represented in reference datasets. Using geospatial indices, genetic similarities, and phenome-wide outcome data, we described the architecture of diseases associated with regional socioeconomic factors such as area-level poverty, education level, single-parent households, living in rented housing units or overcrowded housing units, living without care or unemployment49. We further leverage rich genotyping and phenotyping to clarify several clinically relevant genetic associations complementing clinical and environmental features. This work advances an overall goal of comprehensively quantifying heterogeneous health determinants that uniquely vary across diverse communities in the U.S.
Leveraging population genetics, we delineated the complex genetic components present within the Boston area. Most importantly, our data likely reflect the continued segregation and marginalization of specific genetically similar groups, despite decades of efforts to mitigate racial disparities50,51,52,53. Our integrated dataset will provide a valuable resource for studying and devising strategies to address these complex socio-health consequences. In addition, while our findings align well with prior studies on nationwide cohorts22,23,34,35, our research offers further granular insights into the individual-level histories of the participants, including lifestyle, genetic, and social risk factors associated with the diseases. Further efforts are required to address and reduce this stratification in this area. Area-defined SDI improved prediction performance when incorporated into existing clinical54,55 and genetic risk stratification models56,57 for common complex diseases. In this study, by integrating large scale EHR data and geographical information, we systemically assessed the impact of SDI on Phenome-wide scale across genetically diverse population and drew several clinical implications. First, our systemic assessment suggests that although SDI is a significant contributor to a wide range of diseases, the impacts of SDI are significantly varied across disease domains. For example, while mental and cardiopulmonary diseases were more prevalent among individuals experiencing social deprivation, cancers and congenital diseases are observed almost equally, irrespective of deprivation status. Conversely, we observed several inverse associations between cancer and adverse pregnancy outcome diagnoses and deprivation status, which may reflect greater recognition in the context of greater healthcare access. Furthermore, SDI is differentially yet ubiquitously associated with a wide array of health outcomes across various genetic groups. Finally, although the effect of SDI persisted across various genetic clusters, the varying magnitude of association suggests an interaction between social deprivation and genetic factors as previously suggested58,59.
In addition to enabling detailed disease modeling, healthcare biobanks are unique and powerful resources for exploring rare genetic conditions, disease outcomes, and facilitating implementation in clinical medicine60. First, we confirmed a larger number of pLoF variants among individuals genetically similar to Africans, consistent with previous reports61,62. Next, we identified individuals carrying actionable variants, as defined by a curated database. However, these individuals predominantly belong to clusters we denoted as European-like, who have relatively fewer LoF variants overall. This observation implies a bias against non-European individuals in previous literature used for data curation, potentially resulting from disparities in clinical genetic testing19,63. Using an unbiased genomic scan, our study uncovered several significant associations, which may further refine prognosis within healthcare settings. Furthermore, we confirmed a penetrant association between an upstream variant of the RHD gene and Rhesus isoimmunization during pregnancy13,64, while also clarifying varied prevalence across diverse communities. Bringing these findings together, we highlight that healthcare biobanks, compared to general population-based biobanks, are enriched with uncommon outcomes, and associated genetic variations, thereby offering an ideal environment to study clinically pertinent scenarios.
Nevertheless, our study warrants several limitations. First, most of our enrollment occurred in tertiary hospitals. While this enabled us to include patients with rare and more severe conditions, the prevalence may not reflect the general population due to inclusion bias as previously described65. Second, due to the nature of voluntary participation in the Biobank, MGBB may not fully represent Greater Boston area residents. Also, MGBB participants are centralized in the greater Boston area of Massachusetts, with recruitment primarily from the two largest hospitals in Boston. Communal and geospatial characteristics are likely to vary in other New England regions and more broadly across the U.S. While we highlight the non-random geographic distribution of genetic alleles, further investigations of geographically distinct locales are necessary. Moreover, while our study provides detailed insights into European-like participants, the resolution for non-European participants is less robust due to limited sample sizes, reflecting the demography of the included region. Furthermore, given non-random geographic distribution of alleles, various non-genetic factors may confound the relationship between genetic background and socioeconomic status, warranting careful interpretation of the results.
In conclusion, by utilizing population genetics, we discerned specific genetic clusters within the MGBB. These clusters reflect the colonization histories of the Greater Boston area and exhibit existing social structures and better contextualizing risks for genetically similar participants. Individual-level clinical and lifestyle risk factors in combination with community context, structural factors, and genetic variation advance disease modeling toward precision medicine initiatives.
Methods
Patient recruitment in MGBB and study protocols
MGBB, previously known as Partners Biobank, is an integrated research initiative based in Boston, Massachusetts. It collects biological samples and health data from consenting individuals at Massachusetts General Hospital, Brigham and Women’s Hospital, and local healthcare sites within the MGB network66. This repository of samples and data supports researchers aiming to decipher disease mechanisms, enhance personalized medicine, and innovate therapeutic solutions. Since July 1st, 2010, the MGBB has enrolled 142,238 participants, and extracted DNA from 88,665 participants’ samples (62.3%). All participants provided written/electronic informed consent for broad biological and genetic research. The study protocol to analyze MGBB data was approved by the Mass General Brigham Institutional Review Board under protocol number 2018P001236. The study protocols to analyze UKBB data was approved under protocol number 2021P002228 and performed under UKBB application number 7089.
Genotype quality control and imputation
53,306 individuals were genotyped by Illumina Global Screening Array (Illumina, CA) in four batches (13,140 in the 1st batch, 11,649 in the 2nd batch, 5976 in the 3rd batch, and 22,541 in the 4th batch). Genotypes were called using the Z-call software67. After genotype calling, we conducted quality control with the following steps. We re-aligned genotyping probes to the GRCh38 reference genome using the blast software68 and extracted probes with perfect unique match. We removed indels and multi-allelic sites, and removed variants with high missingness (>2%) and low minor allele counts (<2). After genotype quality control, we estimated continental level genetic similarity using the 1KG dataset. We extracted common, high-quality SNPs (missingness <1%, MAF > 1%) across MGBB and the 1KG dataset. After pruning SNPs, we computed SNP weights for genetic principal component using the 1KG dataset. Then, we projected MGBB participants into the same principal component space using 10 PCs. Using genetic PCs in 1KG dataset as a feature matrix, we trained a K-nearest neighbor model for 1KG reference populations (AFR, AMR, EAS, EUR, and SAS) to assign population labels to MGBB participants. With these inferred labels, we calculated Hardy-Weinberg disequilibrium for each population and removed variants with P < 1 × 10−6. Finally, we compared the allele frequency in these populations with gnomAD allele frequency, then removed variants with deviation from gnomAD allele frequency (Chi-square value > 300) in genetically similar group. These quality control procedures were done by genotyping batch. We took the intersection of the variants in these four batches and generated dataset for imputation. Using the same set of variants, we imputed the genotypes by TOPMed imputation server69. We used TOPMed multi-ancestry imputation reference panel (TOPMed r2 panel) including 97,256 reference samples and 308,107,085 variants. Pre-phasing was carried out by Eagle software70, and imputation was conducted by Minimac4 software69. After the imputation, we merged all the four batches by vcftools71 and converted to bgen file by PLINK2 software (June 6, 2022)72 for the downstream analysis.
Exome sequencing and quality control
Exome sequences were performed by on Illumina NovaSeq instruments (Illumina, CA) with a custom exome capture kit (Human Core Exome, Twist Bioscience, CA), with a target of at least 20x coverage at > 85% of target sites. Alignment, processing, and joint calling of variants were performed using the Genome Analysis ToolKit (GATK, version 4.1)73 following GATK best practices. The joint called dataset containing all 53,420 individuals processed by Hail framework74 for further (1) genotype, (2) variant, and (3) sample quality controls. First, we split the multi-allelic site into biallelic by split_multi_hts function. Following this process, we removed low-quality genotypes and genotypes called by unbalanced allele balance. Following genotype quality control, we conducted variant-level quality control. First, we filtered variants in the low complexity region or outside of the target region (broad_custom_exome_v1.Homo_sapiens_assembly38.bed) with 50 bp flanks. We excluded (i) monomorphic variants and, (ii) variants with high missing rate (>10%). Using a quality-controlled variant set, we conducted sample-level quality control. We collected sample QC metric by hail’s sample_qc function. We implemented five hard filters (percent chimeric reads, percent contamination, call rate, mean depth, and mean genotyping quality, Supplementary Fig. 14) and four soft filters (number of singletons, Ts/Tv ratio, Het/Hom variant ratio, and Insertion/Deletion ratio, Supplementary Fig. 15). For soft filters, we obtained residuals of metrics regressing by the first ten genetic PCs and excluded +/–4 SD outliers. Finally, using only unrelated quality-controlled samples, we computed Hardy-Weinberg P-values by continental genetic similarity estimated from genotyping data. Hardy-Weinberg P-values in chromosome X was computed only for Female participants. We excluded variants with Hardy-Weinberg P-values < 1 × 10−6 or monomorphic variant. After quality control steps, 7,895,027 variants in 22 autosomes and chromosome X were found in 50,625 individuals remained.
Relatedness inference
We utilized the pc_relate75 function from hail (version 0.2) to adjust for the presence of an admixed participants within the MGBB, using 91,615 pruned, common (MAF > 1%) variants that are located outside the major histocompatibility complex (chromosome 6 24,000,000–37,000,000 base pair). Among 53,306 individuals, we identified 3147 pairs with a kinship greater than 0.0884.
Derivation of genetic principal components
To obtain insights utilizing reference populations, first we combined array genotypes from unrelated MGBB participants with recently generated whole-genome sequence datasets from genetically diverse populations including 3380 individuals from 1KG and HGDP28. We intersected 495,213 autosomal, non-palindromic variants outside the high LD region with minor allele counts ≥10. After merging two datasets, we pruned variants by PLINK2 software72 with –indep-pairwise option 1000 100 0.2 resulting in 257,754 variants. Using these genotypes, we derived the weight for each variant for PCs excluding related samples. Using derived weights, we calculated 30 PCs for all the individuals from MGBB, 1KG, and HGDP which were used in subsequent analysis.
Fixation index
Pairwise Fixation indices (FST) were computed among in MGBB-, 1KG- and HGDP- populations using PLINK2 software. The phylogenetic tree was constructed neighbor-joining method76 implemented by ape R package77.
ADMIXTURE analysis
Using PCs derived above, we conducted admixed component analysis using ADMIXTURE software (version 1.3.0)29. We optimized the number of admix component K from 1 to 20 and found that K = 10 showed the least cross-validation error (Supplementary Fig. 5).
Genetic similarity clustering
To derive fine-scale genetic clusters in the participants, we conducted Graph-based clustering which is frequently used in single-cell RNA-seq clustering analysis implemented in Seurat software (version 4.1.0)25. Though Seurat is primarily tailored for single-cell RNA seq data analyses, we leveraged its robust clustering capabilities for genetic similarity clustering. Using the first 30 PCs derived above, we constructed a nearest-neighbor graph and classified individuals into distinct clusters using the Louvain algorithm, a default clustering approach in Seurat with resolution parameter of 0.2. As Seurat identified the clusters in an unsupervised mode, we used individuals from the 1KG or HGDP as a “spike in” positive controls (true labels).
Effective population size estimation
To estimate the effective population size using haplotype sharing information, we used IBDNe in combination with the hap-ibd. First, we phased the genotypes of unrelated MGBB participants with SHAPEIT software (version 4.2). Then, using hap-ibd software (version 1.0, 15Jun23.92f)78, we calculated IBD sharing, and this output was fed into IBDNe software (version 23Apr20)79 to determine the effective population size for each genetically similar group. To compare the effective population size trajectories in British/Irish-like participants in UKBB and MGBB, we computed effective population size in down-sampled, unrelated UKBB self-reported White British or Irish participants (UKBB Field ID 21000) to the same sample size as MGBB British/Irish-like participants (N = 11,508 and N = 10,802, respectively), using microarray-based genotypes. To confirm the finding, we also used the newly established HapNe81 software and obtained consistent results (Supplementary Fig. 8).
Variant annotation
We annotated WES data using the VEP software (version 107)80, supplemented with the Loftee30 and dbNSFP81 plugins. The “—pick” option was enabled to prioritize the canonical transcript. Additionally, in silico predictions from dbNSFP (version 4.2) were employed to prioritize missense variants.
Pathogenic variant annotation
We downloaded ClinVar database40,41 on Aug 16, 2022, and annotated all the variants identified by exome sequence using snpEff software (version 5.0e)82. We identified 536,729 variants registered in the ClinVar Database overall. To identify the carriers of pathogenic/likely pathogenic variants in the ACMG SF v3.1 actionable genes38,39, we only used variants with review status “reviewed_by_expert_panel”, “criteria_provided,_multiple_submitters,_no_conflicts”
Disease phenotyping
We obtained patient data from the electronic health record system spanning the MGB network. We extracted all ICD9 and ICD10 codes assigned to each patient. To enhance the interpretability and powered analysis of the disease outcome, we employed the PheWAS R package (version 1.2)83 to map these codes to corresponding PheCodes24. In this study, we use PheCodes as the sole clinical outcomes for analyzing socioeconomic status and conducting genetic analyses. The PheWAS package utilizes a comprehensive catalog of PheCodes (https://phewascatalog.org/phecodes). We curated all occurrences of each ICD code and assigned the corresponding PheCodes, irrespective of the number of times each ICD code was identified. For ICD codes that did not map to valid PheCodes, those entries were marked as not applicable (N/A). We excluded all unmapped or N/A PheCodes from the analysis to ensure only valid, interpretable phenotypes were included. This mapping process facilitated a more standardized and consistent representation of the patient’s conditions for subsequent analyses. To determine the prevalence or incidence of diseases, we considered the date of blood draw for genotyping as the reference date. By aligning with the corresponding date of PheCodes occurrences, we identified the prevalent or incident outcomes related to the date of enrollment in MGBB.
Geocoding
The participants’ current address data was geocoded using the DeGAUSS framework84, a collection of geospatial tools designed for cleaning and formatting geographic data. This process converts the address information into standardized spatial data, specifically latitude and longitude coordinates. Our analysis focused on participants residing in Massachusetts. We excluded (1) Participants whose addresses were located outside of the state of Massachusetts, (2) Participants for whom the geocoding process failed. We successfully geocoded 118,102 participants and 48,369 with genotype data. We defined Greater Boston based on the definition by the Metropolitan Area Planning Council (https://www.mapc.org/get-involved/subregions/#sub) including 22 cities and 79 towns.
Spatial enrichment analysis
We utilized the Bernoulli model in SaTScan85. Under this model, individuals belonging to a specific genetic cluster were treated as “cases,” while all other individuals were treated as “controls.” This model compares the rates of cases in different areas to determine if the rate of cases inside the potential cluster area is significantly different from outside. To avoid detecting overly large and potentially less meaningful clusters, we limited our scan by setting the maximal diameter of the spatial cluster window. Specifically, we restricted this to a maximum radius of 4 km.
Area-based deprivation score index
For individuals whose addresses were successfully geocoded, we proceeded with the following steps: (A) We assigned each individual’s address to a corresponding U.S. Census tract. Census tracts are small, relatively stable geographic areas that are defined by the United States Census Bureau. They are designed to be relatively homogeneous units with respect to population characteristics, economic status, and living conditions. (B) We then merged this Census tract-level data with SDI (2018 SDI, downloaded from https://www.graham-center.org/maps-data-tools/social-deprivation-index.html)49. SDI is a composite measure of area level deprivation based on seven demographic characteristics collected in the American Community Survey (ACS) and used to quantify the socioeconomic variation in health outcomes. The final SDI is a composite measure of seven demographic characteristics collected in the ACS: percent living in poverty, percent with less than 12 years of education, percent single-parent households, the percentage living in rented housing units, the percentage living in the overcrowded housing unit, percent of households without a car, and percentage non-employed adults under 65 years of age. This approach allows for a detailed, regional census tract-level analysis of the social conditions experienced by the study participants.
Phenome-wide association analysis of social deprivation index
We conducted a phenome-wide association analysis for SDI in 38,526 individuals in the MGBB with clinical and geospatial information. Using area-based SDI as an exposure, we tested its association with binary PheCode-based outcomes. To control for known confounding factors such as race/ethnicity and to ensure population-agnostic results, we included the first ten genetic principal components, along with age and sex, in the logistic regression model. The SDI was normalized to have a mean of zero and a standard deviation of one. In the replication analysis, we utilized the Townsend Deprivation Index at recruitment in the UKBB (UKBB Field 22189) in the same manner. We restricted the analysis to PheCodes with more than 10 cases, resulting in 1561 effective tests in the MGBB. The replication analysis was conducted for 369 of the PheCodes with more than 10 cases in the UKBB, out of 402 PheCodes with significant associations in the MGBB. Phenome-wide significance was set at P < 3.2 × 10−5 = 0.05/1561, and significant replication was considered at P < 1.3 × 10−4 = 0.05/369.
Clinical risk, genetic risk, and social risk for CAD
We calculated the 10-year Atherosclerotic Cardiovascular Disease (ASCVD) risk scores based on the PCE using the PooledCohort R package35,36,86. The PCE accounts for sex, race, age, total and HDL cholesterol, systolic blood pressure, antihypertensives prescription, current smoking, and prevalence of diabetes mellitus. For missing values, we performed multiple imputation by chained equations using the mice R package87, using enrollment age, sex, and race as predictors. PRS was computed using GPS-mult model recently published (https://www.pgscatalog.org/score/PGS003725/)37. CAD was defined by the existence of the ICD codes (ICD9, ICD10) in their EHR. We assessed the individual association of 10-year ASCVD risk, CAD-PRS, and SDI with CAD based on logistic regression adjusting the first 10 genetic principal components. Age and sex were not introduced to the model as PCE used these variables.
Phenome-wide genetic association study
We conducted a phenome-wide rare variant association study (RVAS) for exome genotypes and genome-wide association study (GWAS) for imputed genotypes using the Regenie software (version 3.2.5)88. We associated rare variant burden or imputed genotype dosage with 1482 or 1483 PheCodes under additive model, respectively. For sex specific endpoints, only males or females were included in the analysis where appropriate.
The step 1 was fit using pruned common variants derived from microarray-derived genotypes (MAF > 1%, pruned by PLINK2 software72 with option –indep-pairwise 1000 100 0.9).
For RVAS, we generated masks comprised of predicted loss of function (high confidence by the Loftee software30) and damaging missense variants predicted by >90% of 29 in silico prediction programs included in dbNSFP (version 4.2)81 with MAF < 0.001. To account for multiple hypothesis testing, the significance threshold was set at P < 2.16 × 10−9 by dividing 0.05 with 23,191,897-tested trait × transcript pairs.
For GWAS, the significance threshold was set at P < 3.37 × 10−11 by dividing the conventional genome-wide significant threshold 5 × 10−8 with 1483 tested phenotypes. To define distinct associated loci, we added the flanking region (±500,000 base-pairs) for all the variants with genome-wide significance (P < 3.37 × 1011) and merged all overlapping regions.
In phenome-wide RVAS and GWAS, MGBB participants were analyzed as one group based on the sensitivity analysis described in Supplementary Information Text and Supplementary Fig. 16. Briefly, we estimated Lambda GC (observed chi-squared value divided by expected value) at top 1 percentile of the test statistics for rare variant burden PheWAS and common variant PheWAS and compared the distributions of Lambda GCs across various study settings.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Publicly funded genotyping and exome sequencing data from 13,500 MGBB participants generated in this study have been deposited in dbGaP (accession phs002018). Additional genotyping and exome sequencing data are available under restricted access through industry collaborations; access can be requested under the institutional review board protocol for the current study. Summary data for the leading genetic and clinical associations are provided in the Supplementary Table file. Complete summary statistics from the genetic association analyses are available for download on the Association To Function Knowledge Portal (https://a2f.hugeamp.org/).
Code availability
The analysis codes are available at Zenodo (https://doi.org/10.5281/zenodo.12518901). The docker/singularity images used in the analysis are publicly available through docker hub (https://hub.docker.com/u/skoyamamd).
References
Townsend, P., Phillimore, P. & Beattie, A. Health and Deprivation: Inequality and the North (Routledge, 1988).
Li, X., Memarian, E., Sundquist, J., Zöller, B. & Sundquist, K. Neighbourhood deprivation, individual-level familial and socio-demographic factors and diagnosed childhood obesity: a nationwide multilevel study from Sweden. Obes. Facts 7, 253–263 (2014).
Bann, D., Johnson, W., Li, L., Kuh, D. & Hardy, R. Socioeconomic inequalities in childhood and adolescent body-mass index, weight, and height from 1953 to 2015: an analysis of four longitudinal, observational, British birth cohort studies. Lancet Public Health 3, e194–e203 (2018).
Fan, J. X., Wen, M. & Li, K. Associations between obesity and neighborhood socioeconomic status: Variations by gender and family income status. SSM Popul. Health 10, 100529 (2020).
Akiyama, M. et al. Genome-wide association study identifies 112 new loci for body mass index in the Japanese population. Nat. Genet. 49, 1458–1467 (2017).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Zhou, W. et al. Global Biobank Meta-analysis Initiative: powering genetic discovery across human disease. Cell Genom. 2, 100192 (2022).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Belbin, G. M. et al. Toward a fine-scale population health monitoring system. Cell 184, 2068–2083.e2011 (2021).
Verma, A. et al. The Penn Medicine BioBank: towards a genomics-enabled learning healthcare system to accelerate precision medicine in a diverse population. J. Pers. Med. 12, 1974 (2022).
Brumpton, B. M. et al. The HUNT study: A population-based cohort for genetic research. Cell Genom. 2, 100193 (2022).
Johnson, R. et al. The UCLA ATLAS Community Health Initiative: promoting precision health research in a diverse biobank. Cell Genom. 3, 100243 (2023).
Zawistowski, M. et al. The Michigan Genomics Initiative: a biobank linking genotypes and electronic clinical records in Michigan Medicine patients. Cell Genom. 3, 100257 (2023).
Soria, L. F. et al. Association between a specific apolipoprotein B mutation and familial defective apolipoprotein B-100. Proc. Natl Acad. Sci. USA 86, 587–591 (1989).
Shen, H. et al. Familial defective apolipoprotein B-100 and increased low-density lipoprotein cholesterol and coronary artery calcification in the Old Order Amish. Arch. Int. Med. 170, 1850–1855 (2010).
Luzzatto, L., Ally, M. & Notaro, R. Glucose-6-phosphate dehydrogenase deficiency. Blood 136, 1225–1240 (2020).
Cohen, J. C., Boerwinkle, E., Mosley, T. H. & Hobbs, H. H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006).
Cohen, J. et al. Low LDL cholesterol in individuals of African descent resulting from frequent nonsense mutations in PCSK9. Nat. Genet. 37, 161–165 (2005).
Manrai, A. K. et al. Genetic misdiagnoses and the potential for health disparities. N. Engl. J. Med. 375, 655–665 (2016).
Aragam, K. G. et al. Limitations of contemporary guidelines for managing patients at high genetic risk of coronary artery disease. J. Am. Coll. Cardiol. 75, 2769–2780 (2020).
Baharian, S. et al. The great migration and african-american genomic diversity. PLOS Genet. 12, e1006059 (2016).
Han, E. et al. Clustering of 770,000 genomes reveals post-colonial population structure of North America. Nat. Commun. 8, 14238 (2017).
Dai, C. L. et al. Population histories of the United States revealed through fine-scale migration and haplotype analysis. Am. J. Hum. Genet. 106, 371–388 (2020).
Denny, J. C. et al. Systematic comparison of phenome-wide association study of electronic medical record data and genome-wide association study data. Nat. Biotechnol. 31, 1102–1110 (2013).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Bergstrom, A. et al. Insights into human genetic variation and population history from 929 diverse genomes. Science 367, eaay5012 (2020).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Koenig, Z. et al. A harmonized public resource of deeply sequenced diverse human genomes. Genome Res. 34, 796–809 (2024).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Laken, S. J. et al. Familial colorectal cancer in Ashkenazim due to a hypermutable tract in APC. Nat. Genet. 17, 79–83 (1997).
Levy-Lahad, E. et al. Founder BRCA1 and BRCA2 mutations in Ashkenazi Jews in Israel: frequency and differential penetrance in ovarian cancer and in breast-ovarian cancer families. Am. J. Hum. Genet. 60, 1059–1067 (1997).
Carmi, S. et al. Sequencing an Ashkenazi reference panel supports population-targeted personal genomics and illuminates Jewish and European origins. Nat. Commun. 5, 4835 (2014).
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Browning, S. R. et al. Ancestry-specific recent effective population size in the Americas. PLoS Genet. 14, e1007385 (2018).
Goff, D. C. et al. 2013 ACC/AHA guideline on the assessment of cardiovascular risk: a report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines. J. Am. Coll. Cardiol. 63, 2935–2959 (2014).
Patel, A. P. et al. A multi-ancestry polygenic risk score improves risk prediction for coronary artery disease. Nat. Med. 29, 1793–1803 (2023).
Miller, D. T. et al. ACMG SF v3.1 list for reporting of secondary findings in clinical exome and genome sequencing: A policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 24, 1407–1414 (2022).
Miller, D. T. et al. ACMG SF v3.0 list for reporting of secondary findings in clinical exome and genome sequencing: a policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 23, 1381–1390 (2021).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Landrum, M. J. et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 46, D1062–D1067 (2018).
Backman, J. D. et al. Exome sequencing and analysis of 454,787 UK Biobank participants. Nature 599, 628–634 (2021).
Jurgens, S. J. et al. Analysis of rare genetic variation underlying cardiometabolic diseases and traits among 200,000 individuals in the UK Biobank. Nat. Genet. 54, 240–250 (2022).
Pilarski, R. PTEN hamartoma tumor syndrome: a clinical overview. Cancers 11, 844 (2019).
Tischkowitz, M. et al. Cancer Surveillance Guideline for individuals with PTEN hamartoma tumour syndrome. Eur. J. Hum. Genet. 28, 1387–1393 (2020).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Huang, L. et al. TOP-LD: a tool to explore linkage disequilibrium with TOPMed whole-genome sequence data. Am. J. Hum. Genet. 109, 1175–1181 (2022).
Aguet, F. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Butler, D. C., Petterson, S., Phillips, R. L. & Bazemore, A. W. Measures of social deprivation that predict health care access and need within a rational area of primary care service delivery. Health Serv. Res. 48, 539–559 (2013).
Krieger, N. et al. Structural racism, historical redlining, and risk of preterm birth in New York City, 2013–2017. Am. J. Public Health 110, 1046–1053 (2020).
Lynch, E. E. et al. The legacy of structural racism: associations between historic redlining, current mortgage lending, and health. SSM - Popul. Health 14, 100793 (2021).
Lee, E. K. et al. Health outcomes in redlined versus non-redlined neighborhoods: a systematic review and meta-analysis. Soc. Sci. Med. 294, 114696 (2022).
Motairek, I. et al. Historical neighborhood redlining and contemporary cardiometabolic risk. J. Am. Coll. Cardiol. 80, 171–175 (2022).
Kimenai, D. M. et al. Socioeconomic deprivation: an important, largely unrecognized risk factor in primary prevention of cardiovascular disease. Circulation 146, 240–248 (2022).
Schultz, W. M. et al. Socioeconomic status and cardiovascular outcomes: challenges and interventions. Circulation 137, 2166–2178 (2018).
Bann, D., Wright, L., Hardy, R., Williams, D. M. & Davies, N. M. Polygenic and socioeconomic risk for high body mass index: 69 years of follow-up across life. PLoS Genet. 18, e1010233 (2022).
Cromer, S. J. et al. Association and interaction of genetics and area-level socioeconomic factors on the prevalence of type 2 diabetes and obesity. Diabetes Care 46, 944–952 (2023).
Mostafavi, H. et al. Variable prediction accuracy of polygenic scores within an ancestry group. Elife 9, e48376 (2020).
He, Y. et al. Prediction and stratification of longitudinal risk for chronic obstructive pulmonary disease across smoking behaviors. Nat. Commun. 14, 8297 (2023).
Blout Zawatsky, C. L. et al. Returning actionable genomic results in a research biobank: analytic validity, clinical implementation, and resource utilization. Am. J. Hum. Genet. 108, 2224–2237 (2021).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Sun, K. Y. et al. A deep catalogue of protein-coding variation in 983,578 individuals. Nature 631, 583–592 (2024).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Flegel, W. A. The genetics of the Rhesus blood group system. Blood Transfus. 5, 50–57 (2007).
Lee, Y. H. et al. Impact of selection bias on polygenic risk score estimates in healthcare settings. Psychol. Med. 53, 7435–7445 (2023).
Boutin, N. et al. Implementation of Electronic Consent at a Biobank: an opportunity for precision medicine research. J. Pers. Med. 6, 17 (2016).
Goldstein, J. I. et al. zCall: a rare variant caller for array-based genotyping: genetics and population analysis. Bioinformatics 28, 2543–2545 (2012).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Loh, P. R., Palamara, P. F. & Price, A. L. Fast and accurate long-range phasing in a UK Biobank cohort. Nat. Genet. 48, 811–816 (2016).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Hail Team. Hail 0.2. https://github.com/hail-is/hail.
Conomos, M. P., Reiner, A. P., Weir, B. S. & Thornton, T. A. Model-free estimation of recent genetic relatedness. Am. J. Hum. Genet 98, 127–148 (2016).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Zhou, Y., Browning, S. R. & Browning, B. L. A fast and simple method for detecting identity-by-descent segments in large-scale data. Am. J. Hum. Genet. 106, 426–437 (2020).
Browning, S. R. & Browning, B. L. Accurate non-parametric estimation of recent effective population size from segments of identity by descent. Am. J. Hum. Genet. 97, 404–418 (2015).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Liu, X., Li, C., Mou, C., Dong, Y. & Tu, Y. dbNSFP v4: a comprehensive database of transcript-specific functional predictions and annotations for human nonsynonymous and splice-site SNVs. Genome Med. 12, 103 (2020).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Carroll, R. J., Bastarache, L. & Denny, J. C. R. PheWAS: data analysis and plotting tools for phenome-wide association studies in the R environment. Bioinformatics 30, 2375–2376 (2014).
Brokamp, C., Wolfe, C., Lingren, T., Harley, J. & Ryan, P. Decentralized and reproducible geocoding and characterization of community and environmental exposures for multisite studies. J. Am. Med. Inf. Assoc. 25, 309–314 (2018).
Kulldorff, M. A spatial scan statistic. Commun. Stat. - Theory Methods 26, 1481–1496 (1997).
Yadlowsky, S. et al. Clinical implications of revised pooled cohort equations for estimating atherosclerotic cardiovascular disease risk. Ann. Intern. Med. 169, 20–29 (2018).
van Buuren, S. & Groothuis-Oudshoorn, K. mice: multivariate imputation by chained equations in R. J. Stat. Softw. 45, 1–67 (2011).
Mbatchou, J. et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat. Genet. 53, 1097–1103 (2021).
Acknowledgements
We would like to thank our collaborators Drew Hite, Thanh Long Nguyen, Trang Nguyen, Jason Flannick, Noël Burtt, and the Knowledge Portal Network for their support with the data deposit at the Association To Function Knowledge Portal (https://a2f.hugeamp.org/). This work was supported by the following funding sources. Japan Society for the Promotion of Science (202160643), S.K.; National Institutes of Health (NIH, K99HL169733), S.K.; Uehara Memorial Foundation, S.K.; NHGRI (K99/R00MH117229), Y.W. and A.R.M.; MGH Executive Committee for Research Fund for Medical Discovery, K.P.; NIH (K99HL177340), S.J.C.; AHA career Development Award, S.M.U.; Burroughs Wellcome Fund, S.M.U.; TOPMed fellowship, Y.S.; NIH (K08HL161448), A.C.F.; NIH (R01HL164629), A.C.F.; NIH (R01HG009922), R.C.G.; NIH (R01HL143295), R.C.G. and M.S.L.; NIH (OT2OD026553), R.C.G., J.W.S., and E.W.K.; NIH (U01TR003201), R.C.G. and M.S.L.; NIH (R37MH107649), B.M.N.; NIH (R01MH118233), J.W.S.; NIH (U01HG008685), J.W.S., M.S.L., and E.W.K.; NIH (OT2OD002750), M.S.L.; NIH (OT2HL161841), E.W.K.; NIH (P30AR070253), E.W.K.; European Union’s Horizon 2020 research and innovation program (101016775), A.R.M.; NIH (U01HG011719), A.R.M. and P.N.; NIH (R01HL127564), P.N.
Author information
Authors and Affiliations
Contributions
R.C.G., M.J.D., B.M.N., P.T.E., E.W.K., A.R.M., and P.N. conceptualized this project. S.K., K.P., S.J.C., H.K., and E.W.K. curated phenotype data. S.K., Y.S., A.C.F., and M.S.L. curated genotype data. S.K., M.U., and S.M.U. analyzed data. S.K., Y.W., A.C.F., P.T.E., E.W.K., A.R.M., and P.N. interpreted data. S.K., A.R.M., and P.N. prepared the initial draft. Y.W., K.P., M.U., S.J.C., W.E.H., R.C.G., M.J.D., B.M.N., P.T.E., J.W.S., E.W.K., A.R.M., and P.N. provided critical review and edits for the manuscript. S.H., W.E.H., R.C.G., M.J.D., B.M.N., P.T.E., S.A.S., E.W.K., A.R.M., and P.N. supervised the project. S.H., W.E.H., P.T.E., J.W.S., M.S.L., S.A.S., and E.W.K. managed the project administration. P.T.E., J.W.S., M.S.L., E.W.K., and P.N. obtained funding for the project.
Corresponding authors
Ethics declarations
Competing interests
K.P. reports research grants, paid to her institution, from Allelica, Apple, Amgen, AstraZeneca, Boston Scientific, Genentech/Roche, and Ionis. Y.S. reports serving as a consultant for Arboretum Lifesciences. A.C.F. reports being co-founder of Goodpath and Avigena, and serving as scientific advisor to MyOme, HeartFlow, Aditum Bio, and Arboretum Health, all unrelated to the present work. R.C.G. receives compensation for advising the following companies: Allelica, Fabric, Genomic Life and Juniper Genomics; and is co-founder of Genome Medical and Nurture Genomics. M.J.D. is a founder of Maze Therapeutics and is a member of the scientific advisory board for Neumora Therapeutics, Inc. (formerly known as RBNC Therapeutics). B.M.N. is a member of the scientific advisory board at Deep Genomics and Neumora Therapeutics, Inc. J.W.S. is a member of the Scientific Advisory Board of Sensorium Therapeutics (with equity), and has received grant support from Biogen, Inc. J.W.S. is PI of a collaborative study of the genetics of depression and bipolar disorder sponsored by 23andMe for which 23andMe provides analysis time as in-kind support but no payments. P.N. reports research grants from Allelica, Amgen, Apple, Boston Scientific, Cleerly, Genentech/Roche, Ionis, Novartis, and Silence Therapeutics, personal fees from AIRNA, Allelica, Apple, AstraZeneca, Bain Capital, Blackstone Life Sciences, Bristol Myers Squibb, Creative Education Concepts, CRISPR Therapeutics, Eli Lilly & Co, Esperion Therapeutics, Foresite Capital, Foresite Labs, Genentech/Roche, GV, HeartFlow, Incyte, Magnet Biomedicine, Merck, Novartis, Novo Nordisk, TenSixteen Bio, and Tourmaline Bio, equity in Bolt, Candela, Mercury, MyOme, Parameter Health, Preciseli, and TenSixteen Bio, royalties from Recora for intensive cardiac rehabilitation, and spousal employment at Vertex Pharmaceuticals, all unrelated to the present work. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Mashaal Sohail and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Koyama, S., Wang, Y., Paruchuri, K. et al. Genetics and context for precision health in Greater Boston. Nat Commun 16, 11661 (2025). https://doi.org/10.1038/s41467-025-66598-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66598-8