Abstract
Pulmonary fibrosis (PF) is a progressive lung disease characterized by chronic immune dysregulation and excessive extracellular matrix (ECM) remodeling. Neutrophils, traditionally linked to acute inflammation, are increasingly recognized for their role in fibrosis. In this study, we identified a unique subset of neutrophils undergoing reverse transendothelial migration (rTEM) in the lungs of a silicosis mouse model induced by intratracheal SiO₂ instillation. Using single-cell RNA sequencing (scRNA-seq), spatial transcriptomics (ST), and ECM proteomics, we found that rTEM neutrophils were retained in fibrotic regions via ICAM1-mediated interactions with the ECM. Mechanistically, macrophage-derived cathepsin C (CTSC), accumulating in the ECM, cleaved ICAM1 into its soluble form (sICAM1), which promoted fibroblast activation and exacerbated fibrosis. In vivo depletion of neutrophils or macrophages reduced ICAM1 and CTSC levels, alleviating fibrosis and underscoring their pathogenic roles. These findings establish a direct link between ECM, rTEM neutrophils, and fibrosis, highlighting CTSC and ICAM1 as promising therapeutic targets. All animal experiments were performed using male mice.
Similar content being viewed by others
Introduction
Silicosis is a chronic interstitial lung disease causing progressive lung dysfunction1. Once regarded as primarily affecting workers in developing countries, it has now become a global health concern, impacting an estimated 2.2 million workers in the United States, 2 million in the European Union, and over 23 million in China2,3,4,5,6,7. Silicosis manifests in pathological forms such as simple nodular silicosis, progressive massive fibrosis, and diffuse interstitial fibrosis. Moreover, silica exposure can lead to severe complications, including acute silicoproteinosis, pulmonary tuberculosis, chronic obstructive pulmonary disease, and lung cancer6,8,9. Despite advancements in care and support technologies, no effective pharmacotherapies for silicosis are currently available. The disease is characterized by persistent lung inflammation and advanced fibrosis, progressing even after exposure cessation10,11. Most patients are diagnosed at the irreversible stages (phase II/III), where treatment options are lacking and current approaches fail to halt disease progression3,12. This underscores the need for early diagnostic markers and specific therapeutic targets. The pathological mechanisms of silicosis involve multiple lung cell types, particularly inflammatory cells such as neutrophils, which respond to SiO2 deposition by releasing inflammatory mediators, promoting fibroblast proliferation, and contributing to abnormal ECM remodeling. Therefore, a comprehensive understanding of the molecular mechanisms underlying silica-induced silicosis is critical for advancing its pathophysiological knowledge and identifying therapeutic opportunities.
Immune dysregulation plays a pivotal role in chronic inflammation and fibrosis13,14. Neutrophils, the most abundant immune cells in circulation, are essential effectors of the innate immune response. However, their role in pulmonary fibrosis, particularly during the late stages of irreversible fibrosis, remains poorly defined15. Neutrophils originate in the bone marrow and, following maturation, migrate through the endothelial layer into circulation under the guidance of chemokines. They rapidly accumulate at sites of injury to combat pathogens and mobilize bone marrow reserves16,17,18. Notably, circulating neutrophils are retained in the lung microvasculature, forming the “marginated neutrophil pool”. Early in inflammation, many neutrophils recruited to the lung are derived from this pool, which serves as a readily available reservoir for rapid immune responses19. Clinical studies indicate elevated neutrophil counts in the blood and bronchoalveolar lavage fluid (BALF) of patients with idiopathic pulmonary fibrosis (IPF)20,21. Furthermore, neutrophils exhibit substantial phenotypic and functional heterogeneity under various disease states22,23,24. Of particular interest is a subset of neutrophils undergoing reverse transendothelial migration (rTEM), moving from tissues back into circulation25. These rTEM neutrophils differ from traditional neutrophils that undergo apoptosis and are cleared by macrophages26; instead, they acquire a highly activated phenotype and prolonged survival25. While rTEM neutrophils have been implicated in other chronic inflammatory diseases, their role in silicosis remains largely unexplored. We hypothesize that blocking the activation and recruitment of rTEM neutrophils to the lungs could mitigate irreversible fibrosis in silicosis.
The extracellular matrix (ECM) forms a dynamic scaffold around cells, providing structural and biochemical support, mechanical stability, and protection. Composed of fibrillar proteins, glycosaminoglycans (GAGs), proteoglycans, and other components, the ECM constitutes more than a third of body mass27. Dysregulation of the ECM is directly or indirectly implicated in numerous chronic diseases, including lung fibrosis. In conditions of injury or fibrosis, the ECM undergoes remodeling mediated by matrix-degrading enzymes, resulting in significant changes in composition28,29. An abnormal ECM microenvironment can attract rTEM neutrophils, further influencing immunomodulatory processes30,31. Conversely, neutrophils play active roles in regulating and repairing the ECM, particularly in early tissue repair, by incorporating preexisting matrix into new scaffolds32,33. Investigating the complex interplay between ECM and neutrophils offers potential solutions for addressing failed or excessive tissue repair in lung fibrosis.
In this study, we examined the occurrence and functional roles of rTEM neutrophils in silica-induced lung fibrosis, focusing on macrophage-derived cathepsin C (CTSC)-mediated cleavage of ICAM1. Using scRNA-seq and ST, we identified rTEM neutrophils in a silicosis mouse model and analyzed their association with fibrosis using public datasets. Proteomic analyses, along with in vitro and in vivo experiments, revealed that ECM-accumulated CTSC cleaves ICAM1 into soluble sICAM1, which drives lung fibrosis progression.
Results
rTEM neutrophils are involved in fibrosis progression based on multi-omics analysis
To examine the cellular changes associated with silica-induced fibrosis, we established a murine silicosis model via intratracheal administration of SiO₂. CT imaging of the lungs at 56 days post-exposure revealed grid-like shadows consistent with interstitial fibrosis, while H&E staining confirmed thickened alveolar septa, alveolar structural disorganization, and hemorrhagic plaques (Fig. S1a, b). These findings confirmed the successful induction of silicosis.
Single-cell RNA sequencing (scRNA-seq) was performed on lung tissues from four experimental groups (NS-7d, NS-56d, SiO₂−7d, and SiO₂−56d). t-distributed stochastic neighbor embedding (t-SNE) plot revealed 20 distinct cell clusters across all groups (Fig. S1c). Among these, neutrophil populations displayed significant numerical and proportional changes. In SiO₂-treated groups, neutrophil counts decreased at 7 days but showed a marked increase by 56 days, compared to control groups (Figs. 1a and S1d).
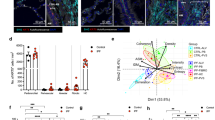
a Pie chart showing the proportion and number of neutrophils in different groups. b Dot plot illustrating marker genes for neutrophil subtypes. c t-SNE plot visualizing the distribution of four neutrophil subtypes, including rTEM neutrophils, in mouse lung tissues across four groups. d Stacked barplot displaying the proportion changes of rTEM and non-rTEM neutrophils. e Pseudotime trajectory analysis using the Monocle2 package to investigate transitions among the four neutrophil subtypes. f Joint analysis of scRNA-seq and spatial transcriptomics data reveals the spatial distribution of rTEM neutrophils within lung tissues. g Gene Ontology (GO) enrichment analysis of marker genes for the four neutrophil subtypes, highlighting top six terms for each subtype and fibroblast activation and proliferation-related terms in rTEM neutrophils. The enrichment was performed with clusterProfiler. For each gene set, over-representation was tested using a hypergeometric test (equivalent to a one-sided Fisher’s exact test) against the specified background gene universe. Analyses were conducted for BP, CC, and MF ontologies. Resulting P values were adjusted for multiple testing using the Benjamini–Hochberg procedure, and significance was determined by FDR (p.adjust) <0.05. Point color encodes the adjusted P value and point size indicates the number of input genes overlapping the term (Count). h Correlation analysis between rTEM neutrophils and fibrosis using transcriptomic data from Pang et al. i Correlation analysis between rTEM neutrophils and fibrosis using scRNA-seq data from Peng et al. (GSE174725). h, i: Module scores between two groups were compared using a two-sided Wilcoxon rank–sum test (Mann–Whitney U). The association between the rTEM module score and the fibrosis module score was evaluated using a two-sided Pearson correlation test (h, i, rightmost panel). Correlation coefficients (R2) and exact P values were calculated using the R function cor.test. Source data are provided as a Source Data file.
The phenotype of rTEM neutrophils is characterized by high expression of intercellular adhesion molecule 1 (ICAM1) and low expression of chemokine receptor 1 (CXCR1, also known as CD181) (ICAM1highCXCR1low)25. Based on this phenotype, 5386 neutrophils were categorized into four subpopulations: reverse transendothelial migration (rTEM) neutrophils (ICAM1highCXCR1low), naïve circulatory (NC) neutrophils (ICAM1⁻CXCR1⁺), tissue-resident (TR) neutrophils (ICAM1lowCXCR1low), and an unknown subtype (ICAM1highCXCR1high) (Fig. 1b, c). Notably, rTEM neutrophil proportions were higher in SiO₂ groups than NS groups (Fig. 1d), and their absolute number increased from day 7 to day 56 post-exposure (Figs. 1c and S1e). Interestingly, although rTEM cells increased in number, their relative percentage declined slightly at the later stage, which was attributed to a more pronounced expansion of TR neutrophils during fibrosis (Fig. S1f–i). This disproportionate growth of the TR subset likely reflects a broader adaptive response to chronic injury. Pseudotime analysis suggested that NC neutrophils transition into rTEM neutrophils during disease progression. However, the eventual fate of rTEM neutrophils remains unclear (Fig. 1e).
Since silicosis is a focal disease, we next investigate the spatial distribution of neutrophil subsets within the lung. An integrated analysis of spatial transcriptomics and scRNA-seq revealed that rTEM neutrophils were primarily localized to inflammatory and fibrotic regions, whereas other neutrophil subsets displayed no distinct spatial patterns (Fig. 1f). Functional analysis indicated that rTEM neutrophils were associated with neutrophil response pathways and uniquely linked to fibroblast proliferation and activation (Fig. 1g), underscoring their potential pro-fibrotic role. In addition, we observed that rTEM neutrophils were enriched in inflammation-associated pathways, such as immune response–activating and immune response–regulating signaling pathways. To further elucidate the functional phenotype of rTEM neutrophils, we defined a pro-fibrotic gene signature by intersecting rTEM-specific marker genes (from scRNA-seq) with published fibrosis-related gene sets. This analysis identified 74 candidate genes potentially mediating fibrotic activity (Fig. S1j). Spatial transcriptomics revealed that these genes are predominantly enriched in regions of inflammation and fibrosis, supporting their relevance in fibrotic microenvironments. Full-resolution plots for all 74 genes are available via Figshare (https://figshare.com/s/fb49da030b4605bb306d). To experimentally validate this signature, we performed qPCR for the top five genes, all of which were significantly upregulated in rTEM neutrophils compared to other subsets (Fig. S1k).To confirm the clinical relevance, rTEM neutrophil and fibrosis scores were assessed in human silicosis lung tissues and bronchoalveolar lavage fluid (BALF) from silicosis patients34,35. Both scores were elevated and significantly correlated, reinforcing their association with fibrosis (Fig. 1h, i).
rTEM neutrophils mediate fibrotic phenotype of fibroblasts
To validate the role of rTEM neutrophils in fibrosis progression, flow cytometry was employed to assess their presence in vivo. The proportion of ICAM1highCXCR1low rTEM neutrophils in SiO₂−56d lung tissues was significantly higher than in NS-56d (Figs. 2a and S2a). A similar increase was also observed in bleomycin-induced pulmonary fibrosis, indicating that rTEM neutrophil accumulation is a shared feature across distinct fibrotic models (Fig. S2b). Immunofluorescence staining revealed elevated ICAM1 levels co-localized with the neutrophil marker Ly6G in SiO₂−56d lungs, further confirming the increased presence of rTEM neutrophils in fibrotic lungs (Fig. 2b). Interestingly, flow cytometry detected approximately 4% rTEM neutrophils, markedly lower than the ~20% estimated by scRNA-seq using our initial definition (ICAM1 counts > 0 and CXCR1 counts = 0). To test whether this difference was driven by threshold settings, we reanalyzed scRNA-seq data using mean-, median-, and MAD-based criteria. All three methods consistently showed rTEM enrichment in fibrotic lungs (Fig. S2c–e), with the MAD method—robust to outliers and zero inflation—yielding the most conservative estimates, down to ~2%. These findings indicate that the absolute percentage differences between scRNA-seq and flow cytometry primarily reflect modality-specific detection principles and threshold definitions rather than biological inconsistency.
a Flow cytometry gating strategy for identifying rTEM neutrophils (ICAM1highCXCR1low) within CD11b⁺Ly6G⁺ lung neutrophils, based on fluorescence-minus-one (FMO) controls. Full gating workflow is shown in Fig. S2A. b Immunofluorescence analysis confirming rTEM neutrophils in lung tissues, with co-localization of Ly6G and ICAM1 signals. Scale bar = 200 μm. c CellChat analysis visualizing cell-cell communication between neutrophil subtypes and fibroblasts. Thicker lines indicate stronger interactions, and darker colors denote higher confidence. d, e Western blots validating the generation of in vitro rTEM neutrophils by showing higher ICAM1 and lower CXCR1 expression in upper chamber cells of the transwell system. f, g Fibroblasts stimulated with upper and lower chamber neutrophil conditioned medium. Western blot analysis shows significant changes in ACTA2 and COL1A2 levels. IL-1β, neutrophils, and mpECs did not show significant effects on fibroblast function in controls. h, i Proliferation and migration of fibroblasts assessed by BrdU incorporation and scratch wound healing assays, respectively. Scale bar = 650 μm. Data are presented as mean ± SEM. n = 5 mice per group for flow cytometry and immunofluorescence, and n = 5 independent experiments for in vitro assays. Biological replicates indicate independent animals or cell cultures. Representative blots/images are shown from n = 5 independent experiments with similar results. Statistical significance was determined using two-sided Student’s t-test with Welch’s correction (a, compared with NS−56d) or one-way ANOVA with Dunnett’s multiple comparisons test (e, g, compared with Con-1). Source data are provided as a Source Data file.
Fibroblasts are the key effector cells in pulmonary fibrosis. Functional analysis revealed that rTEM neutrophils were involved in fibroblast proliferation, activation, and migration (Fig. 1g). CellChat analysis also indicated an increase in interactions between rTEM neutrophils and fibroblasts (Fig. 2c). To further investigate the role of rTEM neutrophils in fibroblast function, we developed an in vitro model utilizing transwell chambers to simulate rTEM, with integrity of the mpECs monolayer assessed via transendothelial electrical resistance (TEER) measurements36 (Fig. S3a, b). Western blot analysis confirmed the rTEM neutrophil phenotype in vitro, characterized by high ICAM1 and low CXCR1 levels in the Exp-upper group (neutrophils in the upper chamber)36 (Fig. 2d, e). Additionally, in the Exp-lower group (neutrophils in the lower chamber), ICAM1 protein levels were also elevated, while CXCR1 protein levels remained unchanged compared to the controls (Fig. 2d, e). According to a study by A. Woodfin et al.36, these ICAM1high neutrophils are characteristic of rTEM neutrophils, supporting the identification of these neutrophils as rTEM in our study. Conditioned medium (CM) from in vitro-generated rTEM neutrophils significantly enhanced COL1A2 and ACTA2 expression in MLg fibroblasts, along with their proliferation and migration (Figs. 2f–i and Fig. S3c, d). These findings suggest that rTEM neutrophils actively contribute to fibroblast activation and pulmonary fibrosis progression.
ICAM1 activates fibroblast and links rTEM neutrophils to ECM interactions
ICAM1, highly expressed on rTEM neutrophils (Fig. 3a), was investigated for its role in pulmonary fibrosis. Recombinant ICAM1 (rmICAM1) significantly enhanced fibroblast activation, as evidenced by increased expression of fibrotic markers COL1A2 and ACTA2 (Fig. 3b, c). Furthermore, rmICAM1 stimulated fibroblast proliferation and migration in MLg cells, as demonstrated by BrdU incorporation and scratch wound healing assays (Fig. 3d–g).
a Violin plot showing elevated ICAM1 expression in rTEM neutrophils. b Recombinant mouse ICAM1 (rmICAM1, 10 ng/ml) significantly enhances fibroblast viability. c Effects of rmICAM1 on fibroblast activation markers COL1A2 and ACTA2, with quantified data shown in the right panel. d–g BrdU incorporation and scratch wound healing assays demonstrating the effects of rmICAM1 on fibroblast proliferation and migration. d, e Scale bar = 200 μm. f, g Scale bar = 650 μm. e and g quantify BrdU-positive cells and wound area, respectively. h CellChat analysis of communication between ECM-producing fibroblasts and rTEM neutrophils, highlighting ligand-receptor contributions in ICAM1 signaling. i Volcano plot of differential ECM proteins in SiO₂-56d vs. NS−56d, with integrin family proteins (e.g., ITGAM, ITGB2) showing the most significant changes. The gray dots represent all the differentially expressed proteins, while the colored dots represent all the integrin family proteins on the ECM. j Barplot showing KEGG terms enriched for differential ECM proteins in SiO₂-56d compared to NS−56d. k–n HDOCK analysis reveals physical binding sites between ICAM1 and integrin family proteins (ITGAM, ITGB2, ITGAL, ITGAX). Representative blots/images are shown from n = 5 independent experiments with similar results. Data are presented as mean ± SEM. n = 5 independent experiments. b, compared with 0 ng/mL group: Dunn’s post hoc test with Benjamini–Hochberg correction for multiple testing. c, f, g, compared with vehicle group: unpaired two-sided t test with Welch’s correction. Source data are provided as a Source Data file.
To elucidate the mechanism by which ICAM1 promotes a fibrotic phenotype, we analyzed cellular interactions between ECM-producing fibroblasts and rTEM neutrophils. ICAM1 signaling was found to mediate this interaction through integrin family proteins, specifically ITGAM, ITGAL, ITGAX, and ITGB2 (Fig. 3h). These integrins, which are closely associated with ECM components, were significantly upregulated in ECM proteomics data from SiO₂-exposed lungs compared to NS controls (Figs. 3i and S4b–d). Additionally, rTEM neutrophils exhibited the highest expression levels of ECM-related genes among all neutrophil subtypes, further supporting their involvement in ECM remodeling (Fig. S4a).
Functional enrichment analysis of differentially expressed ECM proteins revealed significant enrichment in pathways related to ECM-receptor interactions, cell adhesion, and leukocyte migration, highlighting a strong connection between ECM and rTEM neutrophils (Fig. 3j). Molecular docking analysis confirmed direct interactions between ICAM1 and the integrin proteins ITGAM, ITGB2, ITGAL, and ITGAX, suggesting a physical basis for this signaling axis (Fig. 3k–n). These findings underscore the pivotal role of ICAM1 in facilitating fibroblast activation and mediating critical ECM-neutrophil interactions, thereby driving pulmonary fibrosis progression.
ECM-accumulated CTSC cleaves ICAM1 into a soluble form
To further investigate the interaction between rTEM neutrophils and fibrotic ECM, we simulated the rTEM process in vitro (For detailed, see the “Methods”) (Fig. 4a). The results revealed that the mean fluorescence intensity (MFI) of Ly6G was significantly higher in fibrotic ECM (Fib-ECM) compared to normal ECM (Nor-ECM) (Fig. 4b, c), consistent with the high expression of integrin molecules in Fib-ECM. Unexpectedly, the MFI of ICAM1 in Fib-ECM was significantly lower than in Nor-ECM (Fig. 4b, c). To summarize, these results suggest that rTEM neutrophils were trapped in lung ECM during rTEM.
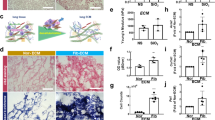
a Workflow showing the seeding of rTEM neutrophils on fibrotic ECM (Fib-ECM from SiO₂-56d lungs) and normal ECM (Nor-ECM from NS−56d lungs). b, c IF analysis of Ly6G and ICAM1 expression on Fib-ECM and Nor-ECM, with (c) quantifying mean fluorescence intensity. Scale bar = 200 μm. d GO enrichment of upregulated ECM proteins in SiO₂-56d compared to NS−56d. e Volcano plot showing differential ECM proteins, with CTSC exhibiting the most significant changes among hydrolytic enzymes. The gray dots represent all the differentially expressed proteins, while the colored dots represent all the hydrolysis-related proteins on the ECM. The p-values shown in the proteomics results were calculated using a two-sided Welch’s t-test, comparing normalized protein intensities between SiO₂-treated samples and control samples. f, g Western blot and immunofluorescence analyses of CTSC expression in lung ECM from NS-56d and SiO₂−56d mice. Scale bar = 200 μm. h HDOCK suggesting CTSC binding to ICAM1, indicating potential cleavage. i Experimental workflow illustrating CTSC-mediated cleavage of ICAM1. j, k Western blots showing ICAM1 cleavage fragments after incubation with recombinant CTSC (j) or lung ECM (k). As purified proteins (200 ng) were used, traditional loading controls (e.g., GAPDH) were not applicable (k). No loading control was applied due to the acellular nature of the ECM (k). Representative images are shown from n = 5 independent experiments with similar results. Data are presented as mean ± SEM. n = 5 independent experiments. c, compared with Nor-ECM group: two-way ANOVA, followed by Šídák’s multiple comparisons test. Source data are provided as a Source Data file. Created in BioRender. Chao, J. (2025) https://BioRender.com/ua4330j.
To determine the mechanism for the reduced ICAM1 in Fib-ECM, we performed GO enrichment analysis for upregulated proteins in Fib-ECM compared with Con-ECM. Results show that these ECM proteins are associated with positive regulation of proteolysis (Fig. 4d). During the disease process, ICAM1 can be cleaved by selective splicing or proteolysis to a variety of membrane-bound and soluble forms37. Notably, among all the hydrolysis-related proteins on the ECM, the increase in CTSC was the most significant and pronounced(Fig. 4e). Both Western blot and immunofluorescence analysis confirmed that CTSC levels were significantly higher and accumulated more in Fib-ECM compared to Nor-ECM (Figs. 4f, g and S5a).
To further investigate the ability of CTSC to cleave ICAM1 into a soluble form, we conducted HDOCK analysis to confirm the interaction between CTSC and ICAM1 (Fig. 4h). We then conducted in vitro cleavage assays using recombinant ICAM1 protein incubated with either recombinant CTSC or decellularized lung ECM, following a modified protocol based on Gocheva et al.38 (Fig. 4i). Western blot analysis revealed a marked reduction in full-length ICAM1 (~ 130 kDa) and the appearance of a single cleaved fragment (~35–40 kDa), a process inhibited by protease inhibitors (Fig. 4j, k). Notably, unlike neutrophil elastase (NE) or cathepsin G (CTSG)39, CTSC did not generate multiple ICAM1 isoforms. Taken together, these findings demonstrate that CTSC, accumulated on the lung ECM, has the ability to cleave ICAM1 to make it soluble form.
CTSC genetic ablation attenuates pulmonary fibrosis and ICAM1 cleavage in vivo
To validate the functional role of CTSC in pulmonary fibrosis, we employed CTSC knockout (CTSC−/−) mice and subjected them to SiO₂-induced lung fibrosis for 56 days (Fig. 5a). CT imaging revealed a marked reduction in high-density areas in the lungs of CTSC-deficient mice compared to wild-type controls, indicating attenuated fibrotic changes (Fig. 5b). Histopathological analyses, including H&E, Sirius Red, and Masson staining, demonstrated substantial improvements in lung architecture, along with reduced collagen deposition in CTSC−/− mice (Fig. 5c). Consistently, hydroxyproline quantification revealed a significant decrease in CTSC−/− mice compared with wild-type controls (Fig. 5d). These results suggest that CTSC deficiency mitigates silica-induced pulmonary fibrosis.
a Schematic of the experimental design: wild-type (CTSC+/+) and CTSC knockout (CTSC−/−) mice were administered either normal saline (NS) or SiO₂ via intratracheal instillation and evaluated after 56 days. b Representative CT images of mouse lungs showing reduced high-density fibrotic regions in CTSC−/−__SiO₂ mice compared to wild-type controls (CTSC+/+__SiO2). c Representative H&E, Sirius Red, and Masson staining images showing improved alveolar architecture and decreased collagen deposition in CTSC−/−__SiO₂ lungs. Scale bar 200 μM. d Hydroxyproline quantification indicating decreased collagen content in CTSC−/−__SiO2 lungs compared with CTSC+/+__SiO2 group. e–g Western blot and densitometric quantification of fibrosis markers COL1A2 and ACTA2 in lung tissues across all four groups. h, i ELISA quantification of soluble ICAM1 levels in (h) lung tissue lysates and (i) bronchoalveolar lavage fluid (BALF), showing reduced ICAM1 cleavage in CTSC-deficient mice compared with CTSC+/+__SiO2 group. Representative images are shown from n = 5 independent experiments with similar results. Data are presented as mean ± SEM. n = 5 mice per group. d, f–i: Statistical significance was determined using two-way ANOVA with Tukey’s post hoc test. Significant comparisons between groups are indicated in the figure panels. Source data are provided as a Source Data file. Created in BioRender. Chao, J. (2025) https://BioRender.com/ua4330j.
Furthermore, Western blot analysis showed significantly lower expression levels of fibrotic markers COL1A2 and ACTA2 in lung tissues from CTSC−/− mice compared to wild-type controls (Fig. 5e–g). To investigate whether CTSC contributes to ICAM1 proteolysis in vivo, we measured soluble ICAM1 (sICAM1) levels using ELISA. CTSC deficiency significantly lowered sICAM1 levels in both lung tissue lysates and bronchoalveolar lavage fluid (BALF), further supporting that CTSC mediates ICAM1 cleavage during fibrosis (Fig. 5h, i).
Macrophage-derived CTSC accumulation in ECM promotes pulmonary fibrosis via ICAM1 hydrolysis
To identify the primary source and role of CTSC accumulation in ECM, a retrospective analysis of CTSC expression was conducted. The Dotplot revealed that macrophages exhibited the highest CTSC expression among cell types (Fig. 6a), which a finding further supported by data from Peng et al.34 (Fig. 6b). To validate this observation, CTSC expression dynamics were assessed in RAW264.7 macrophage cells following SiO₂ treatment. We observed that the expression of CTSC peaked 12 h following SiO2 treatment (Fig. 6c). Additionally, CTSC level in lung tissues was higher in SiO2−56 d and was colocalized with macrophage cell marker F4/80 (Figs. 6d, e and S6a).
a, b Dot plots from GSE183682 and GSE174725 showing highest CTSC expression in macrophages. c Western blot of CTSC levels in RAW264.7 cells treated with SiO₂. d, e Spatial transcriptomics and IF show elevated CTSC expression in SiO₂-56d mouse lungs. Scale bar = 200 μm. f Clodronate liposome workflow for macrophage depletion to investigate effects on ECM-associated CTSC and rTEM neutrophils. g, h IF showing Ly6G and ICAM1 levels in ECM after macrophage depletion, with Nor-ECM as the control group. Scale bar = 200 μm. i Experimental workflow exploring CTSC’s role in ICAM1 cleavage. PI represents a protease inhibitor that inhibits CTSC function, while Ab refers to an anti-CTSC antibody that blocks CTSC function. j ELISA detecting ICAM1 levels in CM-2, confirming CTSC cleaves neutrophil-derived ICAM1. k, l Fibroblasts co-cultured with CM-2 show elevated COL1A2 and ACTA2 levels. m–o BrdU assay and wound healing assessing fibroblast proliferation (m, n) and migration (o, p) after CM-2 treatment. Representative images are shown from n = 5 independent experiments with similar results. Scale bar = 650 μm. Data are presented as mean ± SEM. n = 5 independent experiments. h: two-way ANOVA, followed by Šídák’s multiple comparisons test. j, compared with Con-CM2: one-way ANOVA, followed by Tukey’s test. l, n, p: Unpaired t test with Welch’s correction. Source data are provided as a Source Data file. Created in BioRender. Chao, J. (2025) https://BioRender.com/ua4330j.
To confirm the role of macrophages in CTSC accumulation within ECM, clodronate liposomes (CL) were used to deplete macrophages in silicosis mice (Fig. 6f). Histological analysis revealed that macrophage depletion reduced hemorrhagic plaques, normalized alveolar septa thickness, alleviated ECM disorganization, and decreased collagen deposition in lung tissues (Fig. S6b). Importantly, CTSC levels accumulation in the ECM were significantly reduced in macrophage-depleted mice, further establishing macrophages as the primary source of CTSC in silicosis (Fig. S6c, d).
We next evaluated how macrophage depletion affected rTEM neutrophils. Consistent with the previous results (Fig. 4b, c), Ly6G fluorescence intensity was significantly higher in Fib-ECM compared to Nor-ECM (Fig. 6g, h). However, ICAM1 fluorescence intensity showed no significant differences between these groups (Fig. 6g, h). This observation aligns with the understanding that integrin molecules are derived not only from macrophages but also from other immune cells such as monocytes, dendritic cells, and natural killer cells (Fig. S6e). In summary, macrophages are the main source of CTSC in fibrotic ECM. Macrophage depletion reduced CTSC accumulation and ICAM1 proteolysis but did not affect the retention of rTEM neutrophils in the ECM.
To confirm that macrophage-derived CTSC can cleave ICAM1 and promote pulmonary fibrosis, we simulated the disease microenvironment and generated CTSC in vitro (for details, see the “Methods”) (Fig. 6i). The results demonstrated that SiO₂-stimulated RAW 264.7 macrophages produced CTSC, which interacted with neutrophils to cleave ICAM1 into soluble intercellular adhesion molecule 1 (sICAM1) (Fig. 6j). Notably, Neutralizing CTSC with a specific antibody or protease inhibitor significantly reduced ICAM1 proteolysis (Fig. 6j).
The role of sICAM1 in fibroblast activation was evaluated using conditioned medium (CM-2), which contained elevated levels of sICAM1. CM-2 treatment significantly increased COL1A2 and ACTA2 expression in MLg fibroblasts (Fig. 6k, l), while promoting fibroblast proliferation and migration (Fig. 6m–p). These findings suggest that macrophage-derived CTSC, accumulated on ECM, promotes fibrosis by cleaving ICAM1, releasing sICAM1, and activating fibroblasts.
Neutrophil depletion alleviates pulmonary fibrosis
To validate the role of rTEM neutrophils in pulmonary fibrosis, we depleted neutrophils in vivo using an anti-Ly6G antibody. A dosing regimen involving eight consecutive intraperitoneal injections was employed, with sampling conducted on the final day of treatment (Fig. 7a). Flow cytometry confirmed successful depletion of rTEM neutrophils in lung tissues (Figs. 7b and S7a), as well as in peripheral blood (Fig. S7a, b). CT imaging revealed significantly reduced high-density areas in the lungs of silicosis mice following anti-Ly6G treatment (Fig. 7c). Additionally, HE staining, Sirius Red, Masson staining, and hydroxyproline assays demonstrated improvements in lung architecture and a reduction in collagen deposition and hydroxyproline content following neutrophil depletion (Fig. 7d, e), highlighting a mitigating effect of anti-Ly6G treatment on lung fibrosis. HE staining of other organs, except for the spleen, showed no significant changes following anti-Ly6G antibody administration (Fig. S7c–g).
a Workflow of anti-Ly6G antibody treatment for neutrophil depletion; anti-IgG2a served as the isotype control. b Flow cytometry detecting rTEM neutrophil levels in lungs 10 days after anti-Ly6G treatment. c CT imaging and 3D reconstruction showing reduced high-density lung areas in neutrophil-depleted mice. d Histological analysis (H&E, Masson, Sirius Red) of lung tissues showing improved morphology and reduced collagen deposition. Scale bar = 200 μm. e Hydroxyproline assay quantifying collagen content in lung tissues. f–h Western blot analysis of COL1A2 and ACTA2 levels in lung tissues. i Immunofluorescence showing Ly6G and ICAM1 co-localization in lung tissues. Yellow arrows indicate cells with strong co-localization. Scale bar = 200 μm. j, k ELISA showing reduced sICAM1 levels in lung tissues and BALF after neutrophil depletion. Representative images are shown from n = 5 independent experiments with similar results. Data are presented as mean ± SEM. n = 5 mice. The unit of study was the individual mouse, and samples (lung tissue or protein extracts) were collected from each animal independently. All replicates are biological replicates derived from distinct experimental subjects. b, e, g, h, j, k: two-way ANOVA, followed by Tukey’s test. Groups compared include NS (normal saline control) and SiO₂ (silica-induced fibrosis) treatments, each assessed with two different antibody labels (αIgG2a and αLy6G). Source data are provided as a Source Data file. Created in BioRender. Chao, J. (2025) https://BioRender.com/ua4330j.
Western blot analysis further revealed significant reductions in the expression of fibrosis markers, including COL1A2 and ACTA2, in lung tissues of anti-Ly6G-treated mice (Figs. 7f–h and S7h). To assess the effect of neutrophil depletion on ICAM1 expression, we performed immunofluorescence and ELISA assays. The results showed that anti-Ly6G treatment significantly decreased ICAM1 levels in lung tissues (Figs. 7i and S7i, j) and reduced soluble ICAM1 levels in both lung tissue extracts and BALF (Fig. 7j, k). Taken together, these findings demonstrate that depletion of neutrophils, including rTEM subsets, effectively alleviates silica-induced pulmonary fibrosis by reducing ICAM1 expression and limiting its proteolysis into a soluble form. To further confirm the pro-fibrotic capacity of rTEM neutrophils in a lung tissue context, we treated precision-cut lung slices (PCLS) with conditioned medium derived from rTEM neutrophils. This treatment significantly upregulated the expression of fibrotic markers COL1A2 and ACTA2 (Fig. S8a–c), underscoring the ability of rTEM to promote fibrotic remodeling in lung tissue.
Discussion
In this study, we identified a heterogeneous subtype of reverse transendothelial migrated (rTEM) neutrophils that emerges in the lungs during the progression of silicosis-induced fibrosis. We demonstrated that interactions between rTEM neutrophils and the lung ECM create a sustained cycle of inflammation and fibrosis, ultimately leading to irreversible pulmonary fibrosis. Importantly, the progression of fibrosis observed in this model closely resembled the pathological processes seen in patients with pulmonary fibrosis. The critical role of ICAM1 in rTEM neutrophils was highlighted in this study. ICAM1 expression was found to be essential for the activation and functional changes of lung fibroblasts. Depleting neutrophils in vivo significantly reduced both rTEM neutrophil numbers and ICAM1 levels in lung tissues, contributing to the mitigation of fibrosis. Moreover, we demonstrated that macrophage depletion effectively reduced CTSC production, thereby attenuating ICAM1 proteolysis. However, this intervention did not significantly affect the retention of rTEM neutrophils in the lung ECM, suggesting that macrophage-derived CTSC plays a role in promoting fibrosis through its interaction with ICAM1. These findings provide mechanistic insights into the progression of fibrosis and identify potential therapeutic targets (Fig. 8).
Fibrotic insults induce reverse transendothelial migration (rTEM) of neutrophils①. ICAM1high CXCR1low rTEM neutrophils adhere to and are retained by lung ECM via integrins②. Macrophage‑derived CTSC accumulates in the ECM③ and cleaves neutrophil ICAM1 to generate sICAM1④, which activates fibroblasts and drives collagen deposition⑤. This CTSC–ICAM1 axis links ECM retention of rTEM neutrophils to fibroblast activation in pulmonary fibrosis. Schematic illustration created with Figdraw.com (license ID: SIUUY789f7).
Neutrophils are traditionally associated with acute inflammation due to their short lifespan; however, their role in progressive fibrosis has remained controversial. Recent studies in sepsis-induced lung injury and long COVID-19 have revealed that delayed neutrophil apoptosis and cytoskeletal rearrangement can promote their sequestration in pulmonary capillaries, amplifying inflammatory cascades and potentially contributing to fibrosis40,41,42. Our findings align with this perspective, as neutrophil depletion significantly alleviated lung fibrosis in silicosis mice. This suggests that rTEM neutrophils, in particular, play a pathogenic role in the late stages of fibrosis.
rTEM neutrophils are distinct from classical neutrophils due to their prolonged survival and enhanced inflammatory capacity22,43,44,45. In this study, we found that the number and proportion of rTEM neutrophils in lung tissues increased continuously during silicosis progression, correlating with fibroblast activation and ECM remodeling. These findings consolidate the “double-edged sword” nature of rTEM neutrophils. While they play protective roles in innate immunity, their persistent presence in fibrotic tissues exacerbates disease progression by sustaining inflammation and fibroblast activation46,47. Notably, we also detected elevated rTEM neutrophils in bleomycin-induced fibrosis, suggesting that their accumulation is a common feature of fibrotic lung models likely driven by early excessive inflammation.
Although the proportion of rTEM neutrophils among total neutrophils slightly declined from day 7 to day 56, this reflects a faster expansion of TR neutrophils during late-stage fibrosis. Such expansion may represent an adaptive response to chronic tissue injury, as suggested in other lung inflammation models. We speculate that the accumulation of TR neutrophils may reflect a general adaptive response to chronic tissue injury, as previously suggested in lung inflammation models48,49. Importantly, the absolute number of rTEM neutrophils still rose significantly, supporting enhanced recruitment or retention. Given their potent pro-inflammatory and pro-fibrotic capacity, this accumulation plays a key role in driving fibrotic progression.
A key finding of this study was the interaction between rTEM neutrophils and lung ECM. The ECM, acting as a molecular reservoir and structural scaffold, was shown to retain and “hijack” rTEM neutrophils through ICAM1 and integrin-mediated interactions. This mechanism establishes a feedback loop that perpetuates inflammation and fibrosis. Although managing the excessive infiltration of rTEM neutrophils remains a significant hurdle50, understanding this unique migration and retention event offers opportunities for therapeutic intervention by disrupting the ECM-rTEM neutrophil cycle.
Adhesion molecules are key regulators of cellular function, tissue integrity, and dynamic homeostasis51. They mediate intercellular interactions and trigger intracellular signaling by binding to the cytoskeleton and adapter proteins, ensuring dynamic responses to specific stimuli52. Among adhesion molecules, ICAM1, a member of the immunoglobulin superfamily, has been frequently implicated in inflammation and fibrosis. ICAM1 is a cell surface glycoprotein comprising five extracellular immunoglobulin domains, a transmembrane region, and a short cytoplasmic tail, which interacts with various integrin family members, such as LFA-1 and Mac-1, to regulate leukocyte adhesion and migration53,54. In lung diseases, ICAM1 has been associated with both inflammatory and fibrotic processes. For example, ICAM1 expression is upregulated in idiopathic pulmonary fibrosis and acute lung injury, where it facilitates leukocyte recruitment and the amplification of inflammatory responses55. In our study, ICAM1 was identified as a critical molecule on rTEM neutrophils, serving as an “anchor” that mediates their retention in the lung ECM. Although we did not conduct direct experiments to confirm ICAM1-integrin interactions in this context, previous studies have extensively documented the binding of ICAM1 to integrins and its adhesive functions in inflammatory and fibrotic settings56,57. Based on these well-established mechanisms, we deduce that ICAM1 adheres to ECM components, contributing to the retention of rTEM neutrophils and perpetuating the fibrotic cycle.
Interestingly, ICAM1 not only functions as a cell surface molecule but is also susceptible to cleavage into its soluble form (sICAM1). This dual function of ICAM1, both as an adhesion molecule and a soluble mediator, underscores its complex role in fibrosis. In silicosis, we observed elevated ICAM1 expression on rTEM neutrophils, which likely contributed to their retention in the ECM. Concurrently, macrophage-derived CTSC cleaved ICAM1 into sICAM1, which activated fibroblasts and exacerbated fibrosis. These findings highlight the dual roles of ICAM1 in regulating neutrophil-ECM interactions and promoting fibroblast activation, reinforcing its central position in the fibrotic cascade. Although our study identifies CTSC as a key ECM-associated protease responsible for ICAM1 cleavage and fibrotic progression, we cannot exclude the possibility that other proteases—such as neutrophil elastase, cathepsin G, or matrix metalloproteinase-9—may also contribute to this process under certain conditions39,58. While CTSC exhibited the most prominent upregulation in fibrotic ECM and its genetic deletion significantly reduced sICAM1 levels and fibrosis severity, compensatory activation of other proteases in the absence of CTSC remains a plausible scenario. Future studies involving protease activity profiling, selective inhibitors, or combinatorial knockout models will be necessary to delineate potential redundancy or synergy among protease families in ICAM1 cleavage and fibrotic regulation.
ICAM1 undergoes protein cleavage to produce at least six membrane-bound forms and one soluble form (sICAM1)37. sICAM1 is particularly relevant in the context of lung diseases, where its levels are elevated in the serum and BALF of patients with pulmonary fibrosis, asthma, and chronic obstructive pulmonary disease59,60,61,62,63,64,65. This soluble variant retains the extracellular immunoglobulin domains responsible for binding integrins, allowing it to modulate cell signaling pathways and contribute to disease progression56,66 Our study corroborates these findings, showing a progressive increase in sICAM1 levels in the lung tissues and BALF of silicosis mice during fibrosis progression.
The cleavage of ICAM1 into sICAM1 is mediated by proteases such as matrix metalloproteinase-9 and cathepsin G, as well as CTSC, which was identified in this study as a key ECM protease upregulated during fibrosis39,58 CTSC not only cleaves ICAM1 but also regulates neutrophil extracellular traps and inflammatory responses, linking it to a range of immune-mediated diseases such as rheumatoid arthritis, pneumonia, and viral infections67,68,69 Interestingly, CTSC has also been implicated in cancer metastasis by regulating neutrophil infiltration and ECM remodeling70 Furthermore, the inhibition of elastase-associated neutrophil proteases, which are often upregulated by CTSC, has been shown to alleviate lung injury71 In this study, we demonstrated that macrophage-derived CTSC cleaves ICAM1 on rTEM neutrophils, generating sICAM1, which subsequently activates fibroblasts and promotes fibrosis. The proteolytic activity of CTSC adds another layer of complexity to ICAM1’s role in silicosis, transitioning from an adhesion molecule to a soluble mediator of fibrotic signaling. This finding underscores the importance of understanding the proteolytic regulation of ICAM1, particularly in its transition to sICAM1.
Despite the significant insights gained, our study has limitations. The use of anti-Ly6G antibodies to deplete neutrophils lacks specificity for rTEM neutrophils, potentially affecting other neutrophil subtypes with protective roles in fibrosis. Developing targeted methods to specifically manipulate rTEM neutrophils will be crucial for further understanding their precise contributions to fibrosis. Although our PCLS-based co-culture assays confirmed the pro-fibrotic effects of rTEM neutrophils in vitro, additional in vivo experiments are warranted to validate their pathogenic function under physiological conditions. For example, adoptive transfer of purified rTEM neutrophils into fibrotic mouse lungs would provide stronger causal evidence.
Second, while we demonstrate that sICAM1 generated from rTEM neutrophils promotes fibroblast activation, rTEM cells may also secrete other pro-fibrotic mediators. Thus, a comprehensive profiling of rTEM-derived soluble factors and further functional validation are required to fully understand their contribution to disease progression. Moreover, although the increased trends of rTEM neutrophil were consistent between scRNA-seq and flow cytometry, the absolute proportions differed considerably between platforms—likely reflecting methodological biases inherent to each technique72,73,74,75 rTEM neutrophils are transcriptionally distinct and may be difficult to isolate by conventional flow cytometry due to gradual marker shifts and phenotypic overlap. This underscores the need for complementary validation using single-cell transcriptomics, which allows unbiased clustering and trajectory inference. Another potential limitation of comparing rTEM proportions across platforms is the inherent misalign in definition criteria—flow cytometry relies on protein surface markers (ICAM1highCXCR1low), while scRNA-seq detects transcriptional activity (ICAM1 counts > 0, CXCR1 counts = 0). Although we approximated flow’s protein gating by applying stricter mRNA thresholds, direct translation is constrained by lack of empirical mRNA-to-protein conversion data for neutrophils in our dataset. Nonetheless, concordant enrichment of rTEM neutrophils in fibrotic lungs across both methods validates the biological relevance of this subset, even if absolute proportions differ due to technical and definitional differences.
Finally, while our use of global CTSC knockout mice provides genetic support for the pro-fibrotic role of CTSC in vivo, it does not exclude the involvement of CTSC expressed by other cell types. To more precisely define the role of macrophage-derived CTSC, future studies employing macrophage-specific CTSC knockout models (e.g., LysM-Cre or Cx3cr1-Cre) will be crucial. These models will help isolate the contribution of CTSC within macrophages and clarify whether the observed fibrotic effects are driven by macrophage-intrinsic mechanisms or by systemic or developmental compensatory pathways.
Pulmonary fibrosis is a fatal, progressive disease with limited therapeutic options. In this study, we identified a direct functional link between the lung ECM and neutrophils during the irreversible fibrosis stage of silicosis. Specifically, we demonstrated that a subtype of heterogeneous neutrophils, which underwent rTEM, persisted in the lungs throughout the progression of silicosis. The ECM, acting as both a molecular reservoir and a scaffold for cell survival, facilitated the retention and recruitment of rTEM neutrophils in a spatiotemporal manner. This interaction resulted in the continuous circulation of rTEM neutrophils during the inflammatory and fibrotic stages, ultimately contributing to irreversible fibrosis. Furthermore, our findings suggest that targeting CTSC, a key molecule in the lung ECM, and ICAM1, a core molecule in rTEM neutrophils, represents a potential therapeutic approach for progressive pulmonary fibrosis. We believe that the insights from our mouse models of pulmonary fibrosis will pave the way for breakthroughs in the prevention and treatment of pulmonary fibrosis and other fibrotic diseases.
Methods
Study approval
This work has received approval for research ethics from the Laboratory Animal Care and Use Committee of Southeast University (Approval No. 20190121002).
Animals, strains, and housing
Mus musculus, C57BL/6J mice (male, 6–8 weeks old) were obtained from GemPharmatech Co., Ltd. Ctsc knockout C57BL/6J mice were kindly provided by Prof. Jianmei Ma and Prof. Chi Liu; the line was maintained on a C57BL/6J background in our facility. Mice were housed in the Laboratory Animal Center of Southeast University under SPF conditions (12-h light/dark cycle, 21–25 °C, 40–50% humidity) with food and water ad libitum.
Silicosis model
After anesthetization with pentobarbital, mice received an intratracheal injection of 100 μL silica suspension (50 mg/mL). Control mice were injected with sterile saline (NS) in the same manner. Mice were housed in separate cages and observed for 56 days. Silicon dioxide particles (<5 μm diameter) were dried at high temperature before use. Concentrations used were 5 mg/mL for cellular assays and 50 mg/mL for animal models. All in vivo experiments were performed using male C57BL/6J mice. The rationale for using only male animals was as follows: (i) male mice provide a more stable model, as estrogen cycling in females can influence immune responses and fibrosis progression; (ii) the use of male mice is consistent with the majority of published studies on SiO₂- and bleomycin-induced pulmonary fibrosis, facilitating comparison across studies; and (iii) limiting experiments to one sex reduced the overall number of animals required, in line with the 3 R principle of animal research. Euthanasia was performed by intraperitoneal injection of overdose pentobarbital (150 mg/kg) on the final day of the experiment, in accordance with institutional animal welfare guidelines.
CT scanning
Mouse lung CT imaging was conducted using a Hiscan XM Micro CT scanner (Suzhou Hiscan Information Technology Co., Ltd.) under the following conditions: 60 kV tube voltage, 134 μA tube current, 50 ms single exposure time, and 50 μm resolution. Image reconstruction and analysis were performed using Hiscan Viewer 3.0 (Nanjing Zuangchuang Biotechnology Co., Ltd.).
Cell culture
All cells were maintained at 37 °C in a humidified atmosphere with 5% CO2. MLg(ATCC, Cat#CCL-206) and RAW 264.7 (ATCC, Cat#TIB-71) cells were cultured in DMEM containing 10% FBS, 1% penicillin-streptomycin (P/S), and 1% glutamine. Primary mouse pulmonary microvascular endothelial cells (mpECs) were isolated from male C57BL/6J mice using CD31 MicroBeads (Miltenyi, 130-097-418) and cultured in Endothelial Cell Medium (ScienCell, #1001) with Endothelial Cell Growth Supplement (#1052) and 1% penicillin–streptomycin at 37 °C, 5% CO₂. Neutrophils were cultured in RPMI-1640 medium with 10% FBS, 1% P/S, 1% glutamine, and 0.1% 2-mercaptoethanol.
Generation of rTEM neutrophils
Following Gocheva et al.36, rTEM neutrophils were generated in a Transwell system. Mouse pulmonary endothelial cells (mpECs) were pre-stimulated with IL-1β (2.5 ng/mL) for 4 h, washed with PBS, and co-cultured with bone marrow-derived neutrophils at 37 °C for 1 h. Nonadherent neutrophils were removed, and the culture continued for 24 h. rTEM neutrophils were collected by washing the mpEC monolayer with DMEM. Three control groups were set up: (1) neutrophils incubated alone; (2) neutrophils co-incubated with mpECs; (3) mpECs incubated after IL-1β stimulation.
Phenotypic identification of rTEM neutrophils
For in vivo identification, single-cell suspensions from mouse lung tissue were treated with Red Blood Cell Lysis Buffer to remove erythrocytes. Single-cell suspensions were stained with Fixable Viability Dye eFluor 506 (eBioscience, Cat#65-0866-14, 1:1000), and Fc receptors were blocked using anti-mouse CD16/CD32 (BD Biosciences, Cat#553141, clone 2.4G2, 1:100). The following fluorochrome-conjugated antibodies were used: anti-CD45-PE-Cy7 (BD Biosciences, Cat#561868, clone 30-F11, 1:200), anti-CD11b-PerCP-Cy5.5 (BD Biosciences, Cat#561114, clone M1/70, 1:200), anti-Ly6G-FITC (eBioscience, Cat#11-9668-80, clone 1A8, 1:200), anti-CD54-BV786 (BD Biosciences, Cat#740855, clone YN1/1.7.4, 1:100), and anti-CD181-PE (BD Biosciences, Cat#566383, clone 8E5, 1:100). Samples were acquired on a Cytek Aurora spectral cytometer. Flow cytometry gating was performed in the following order: exclusion of debris (FSC-A vs SSC-A), live cells (viability dye–), singlets (FSC-A vs FSC-H), CD45⁺ leukocytes, and CD11b⁺Ly6G⁺ neutrophils. rTEM neutrophils were identified as ICAM1high CXCR1low cells, with gating thresholds defined based on fluorescence-minus-one (FMO) controls. The complete gating strategy and FMO validation are shown in Fig. S2A.
For the in vitro rTEM neutrophils identification, cells from Transwell inserts were lysed, and ICAM-1 and CXCR1 expression was detected by western blot.
Single-cell transcriptome sequencing (GSE183682)
Lung tissue samples were collected from mouse silicosis model groups: NS-7 d, NS-56 d, SiO2-7 d, and SiO2−56 d (one sample from each group). After perfusion, hearts were rinsed three times with PBS buffer. Lung tissues were sliced into small pieces (approximately 0.5 mm³), and individual cells were isolated using a lung isolation kit. A Gel Bead in Emulsion (GEM) was generated following the manufacturer’s protocol, and quality was assessed using an Agilent 4200 system. Cell suspensions were loaded onto a Chromium Single-Cell Controller (10× Genomics) using a single-cell 5′ Library (v3), Gel Bead Kit, and Chromium Single-Cell G Chip Kit. Libraries were constructed with 150 bp paired-end reads and sequenced on an Illumina NovaSeq6000 platform using NovaSeq Control Software v1.7 by Beijing BIO Biotechnology Co.
scRNA-seq reanalysis of rTEM neutrophils with alternative thresholds
rTEM neutrophils were initially defined in the scRNA-seq dataset as ICAM1 counts > 0 and CXCR1 counts = 0 within the neutrophil cluster. To assess the robustness of rTEM proportion estimates to threshold definitions, we reanalyzed the same dataset using three alternative strategies, all based on log-normalized expression values: (1) Mean-based threshold—ICAM1high defined as expression greater than the global mean of ICAM1 across all neutrophils; CXCR1low defined as expression less than or equal to the global mean of CXCR1. (2) Median-based threshold—ICAM1high defined as expression greater than the global median of ICAM1 across all neutrophils; CXCR1low defined as expression less than or equal to the global median of CXCR1. (3) Median Absolute Deviation (MAD)-based threshold—ICAM1high defined as expression greater than k × MADbaseline (ICAM1) calculated from baseline-control neutrophils; CXCR1low defined as counts = 0 (optional ≤ MADbaseline). This method was chosen for its robustness to extreme values and zero inflation in scRNA-seq data, and k was tuned to yield conservative estimates in fibrotic groups. For each approach, neutrophils were classified into rTEM (ICAM1highCXCR1low), NC (ICAM1lowCXCR1high), TR (ICAM1lowCXCR1low), or unknown (ICAM1highCXCR1high), and rTEM proportions were calculated as a percentage of total neutrophils in each experimental group. The MAD-based criterion was selected as the strictest approximation to the protein “high/low” logic in flow cytometry because it is robust to extreme values and zero inflation in scRNA-seq data.
Spatial transcriptome sequencing (GSE183683)
Lung tissues were trimmed near the hilum, frozen in optimal cutting temperature compound (OCT), and stored at −80 °C. Sections were adhered to slides, fixed, and stained with H&E before imaging. Visium spatial libraries were constructed using the Visium Spatial Library Construction Kit and sequenced on an Illumina NovaSeq6000 by CapitalBio Technology.
Proteome analysis of the extracellular matrix
ECM proteins from NS-56d and SiO2−56d mouse lungs were extracted using the Compartmental Protein Extraction Kit (Millipore, Cat 2145). Extracts were divided for SDS-PAGE and trypsin digestion. Proteomic data were generated on an LC–MS/MS platform (Shanghai Luming Biological Technology Co., Ltd.) using vendor-provided acquisition software (ProteinPilot, v.5.0).
One 6-plex TMT set was acquired comprising six channels (126–131) mapped to sample IDs: 126 = com111, 127 = con116, 128 = con117, 129 = M80, 130 = M101, 131 = M107. For between-group statistics we compared M (M80/M101/M107; n = 3 biological replicates) versus Con (com111/con116/con117; n = 3 biological replicates). Proteins were reduced/alkylated (iodoacetamide to 10 mM, 15 min, dark) and acetone-precipitated (−20 °C ≥ 4 h; 8000 × g, 10 min) before trypsinization (Trypsin-TPCK 1:50, 200 mM TEAB, 37 °C overnight). Peptides were TMT-labeled in 100 mM TEAB: TMT reagent reconstituted in anhydrous acetonitrile (88 µL), 41 µL added per sample, 1 h at room temperature; reaction quenched with 8 µL 5% hydroxylamine for 15 min. Labeled peptides were combined. High-pH reversed-phase fractionation was performed on an Agilent 1100 HPLC with a Zorbax Extend-C18 (2.1 × 150 mm, 5 µm) column; mobile phases were ACN/H₂O adjusted to pH 10 (A: 2/98; B: 90/10), flow 300 µL min−1, with a 75 min gradient (8–60 min collected each minute into 15 tubes). Peptide fractions were loaded onto Evotips and separated on an Evosep One using the 30SPD method (1 µL min−1) with 15 cm analytical column; mobile phases: A = H₂O/0.1% FA, B = ACN/0.1% FA. MS data were acquired on a Bruker timsTOF Pro (Bruker Daltonics; supplier listed in the report) with nano-ESI 1.5 kV, drying gas 180 °C at 3.0 L min−1, MS1 m/z 100–1700, TIMS 1/K₀ range 0.6–1.6, collision energy 20–59 eV. Raw data were processed in MaxQuant v1.6.17.0 using the UniProt reviewed Mus musculus (taxonomy 10090) FASTA. Enzyme: Trypsin/P; fixed modification: Carbamidomethyl (C); variable modifications: Oxidation (M), Protein N-term Acetylation; first search tolerance 20 ppm, main search 10 ppm; max missed cleavages = 2; instrument type: Bruker TIMS. Protein-level FDR was controlled at <1% (4228 protein groups; 25,987 peptides identified). Proteins with ≥1 unique peptide were retained; proteins with expression values present in ≥50% of samples in any group were kept. For proteins with ≤50% missingness, missing values were imputed by the within-group mean. Intensities were median-normalized and log₂-transformed before downstream analyses. Quality control included PCA and sample-to-sample correlation analyses on the set of confidently quantified proteins. Differentially expressed proteins (DEPs) between M and Con were defined using fold-change and two-sided t-tests with the vendor’s thresholds FC ≥ 2.0 and p < 0.05; in this comparison, 270 DEPs were reported (M_vs_Con).
Decellularization of lung matrices
Decellularized ECM was harvested based on a previous report with some modifications76 Freshly harvested lung tissues were embedded in OCT and cryosectioned into 300 μm slices using a Leica cryostat. Tissue sections were placed in 6-well plates and incubated with 5 mL of 1% SDS solution under gentle agitation at room temperature for 1 h, a step repeated once with fresh SDS. Subsequently, sections were incubated overnight in fresh 1% SDS at room temperature with continuous shaking. The next day, the SDS solution was replaced with 5 mL of 1% Triton ×-100, and the sections were agitated at room temperature for 1 h, repeated three times. This was followed by overnight incubation in fresh 1% Triton ×-100 under the same conditions. After detergent removal, sections were rinsed twice with 1× PBS and four times with ddH₂O. To remove residual nuclear material, sections were treated with 5 mL of 1 M NaCl for 1 h at room temperature, then rinsed again with 1× PBS (2 times) and ddH₂O (2 times). DNA digestion was performed by incubating the sections with 5 mL of DNase I solution (20 μg/mL) containing 4.2 mM MgCl₂ for 1.5 h at room temperature under agitation. After enzymatic treatment, tissues were washed twice with ddH₂O. For sterilization, sections were treated with 4.8% acetic acid for 20 min in a biosafety cabinet, followed by three washes with sterile 1× PBS. The decellularized ECM slides were stored at 4 °C until use.
Simulation of neutrophils rTEM in vitro
To simulate dynamic rTEM, rTEM neutrophils were induced in Transwell inserts. After nonadherent neutrophils were removed, lung ECM (Nor/Fib) was placed in the lower chamber to mimic the microenvironment. ECM was fixed with 1× paraformaldehyde after 24 h and analyzed by immunofluorescence staining.
ScRNA-seq and spatial transcriptomics integration analysis
Cell-type-specific marker genes identified from the scRNA-seq dataset were used as gene signatures. Using the AddModuleScore function in Seurat, we calculated module scores to estimate the spatial enrichment of each cell type, with higher scores indicating greater abundance. To validate spatial distribution patterns, we further applied cell2location to integrate the scRNA-seq and spatial transcriptomics data, generating probabilistic maps of cell-type abundance across tissue sections
Enrichment analysis
Unless otherwise specified, Enrichment analysis was performed using the R package “clusterProfiler” for genes meeting criteria of |log2 fold change| > 1 and adjusted p-value < 0.01.
Signature scores and their correlation analysis
The rTEM signature genes were identified from our scRNA-seq dataset (GSE183682), and the fibrosis signature was derived from the Comparative Toxicogenomics Database (CTD), which contains a list of 947 genes associated with pulmonary fibrosis. These two signature scores were computed using Seurat’s AddModuleScore function, followed by an assessment of their correlation.
Macrophage depletion
Clodronate liposomes (100 μL, 5 mg/mL) were injected via tail vein into silicosis model mice every 7 days for 56 days. Control liposomes were used as controls. Mice were grouped based on similar weight and status before treatment.
Neutrophils depletion
To deplete neutrophils, anti-mouse Ly6G (Clone 1A8, Bioxcell) and an IgG2a isotype control (2A3, Bioxcell) were administered via intraperitoneal (i.p.) injection at a dose of 100 µg/20 × g body weight. The treatment protocol included daily injections for the first 6 days, followed by injections every other day for 4 additional days, based on other reports77,78
Mouse primary endothelial cells
To isolate mouse primary endothelial cells, one-week-old mice were anesthetized with pentobarbital, and a thoracic incision was made to expose the heart and lungs for cardiac perfusion. The lungs were excised, minced with surgical scissors, and digested with collagenase I and DNase for 1 h at 37 °C with gentle shaking. The resulting supernatant was centrifuged and filtered through a 70-µm cell strainer to obtain a single-cell suspension. Mouse pulmonary endothelial cells (mpECs) were then isolated using CD31 MicroBeads (130-097-418, Miltenyi Biotec) following the manufacturer’s protocol.
Isolation of mouse bone marrow neutrophils
To isolate mouse bone marrow neutrophils, wild-type (WT) mice, approximately 6–8 weeks old, were anesthetized with pentobarbital. A neck dissection was performed to remove the femurs and tibiae of the hind limbs, and the bone marrow was flushed with phosphate-buffered saline (PBS) to collect a single-cell suspension. Mouse primary bone marrow neutrophils were then isolated using the Neutrophil Isolation Kit (130-097-658, Miltenyi Biotec) following the manufacturer’s protocol.
Resistance across monolayers
Following previously established protocols with modifications79 3 × 105 mpECs per well were cultured on 0.3 cm2 Transwell® plates. The transendothelial electrical resistance (TEER) was monitored during cell growth using a microtiter Milli cell ERS-2 system. The resistance values were calculated using the following formula:
Unit Area Resistance = Resistance (Ω) × Effective Membrane Area (cm2).
Immunofluorescence staining
Immunofluorescence staining was performed as previously described80 Briefly, lung tissues and ECM sections (8 μm) were blocked with 10% fetal sheep serum in 0.3% Triton ×-100 for 2 h and incubated with primary antibodies overnight at 4 °C. Sections were stained with Cy3/Alexa Fluor-conjugated secondary antibodies and counterstained with DAPI. Images were captured using an Olympus IX70 fluorescence microscope with accompanying Olympus software (v3.28). For immunofluorescence staining, samples were incubated with ICAM-1 antibody (Proteintech, Cat#10020-1-AP, 1:200), Ly6G antibody (eBioscience, Cat#11-9668-80, clone 1A8, 1:200), CXCR1 antibody (ABclonal, Cat#A16386, 1:200), α-SMA antibody (Proteintech, Cat#14395-1-AP, 1:200), collagen I antibody (Proteintech, Cat#14695-1-AP, 1:200), and cathepsin C antibody (Santa Cruz, Cat# sc-74590, clone D-6, 1:200). Secondary antibodies conjugated to Alexa Fluor 488, 594, or 647 were used for detection. Nuclei were counterstained with DAPI (ProLong™ Diamond, Cat# P36971).
Hematoxylin and Eosin staining
Hematoxylin and eosin (H&E) staining was performed on mouse lung tissue and lung extracellular matrix (ECM) samples. The tissues were fixed in 1× paraformaldehyde (PFA) and embedded in OCT compound to prepare 8-µm frozen sections. These sections were stained using a Hematoxylin and Eosin Staining Kit (C0105M, Beyotime). The procedure involved washing the sections with distilled water for 2 min, staining them with hematoxylin and eosin for 10 min, and rinsing off the excess stain with tap water for approximately 10 min. The tissue sections were then dehydrated in 75% ethanol, followed by 80% anhydrous ethanol for 10 s, and 90% xylene for 5 min. Finally, the tissue sections were mounted with neutral gum, air-dried, and images were captured using a fluorescence microscope (Olympus IX70, USA).
Hydroxyproline Assay
Mouse lung tissues (~0.1 × g) were digested in extraction solution at 100 °C for 4 h. The pH was adjusted to 6–8, and the solution was centrifuged at 16,000 rpm for 20 min. Hydroxyproline was quantified using the Hydroxyproline Assay Kit (Solarbio).
Western blotting
Western blotting was performed as previously described81 Briefly, protein lysates were prepared in RIPA buffer, quantified, and separated by SDS-PAGE. Proteins were transferred to PVDF membranes, blocked with 5% skim milk, and incubated with primary antibodies overnight at 4 °C. Membranes were incubated with HRP-conjugated secondary antibodies and imaged using a Tanon scanner. The following primary antibodies were used for Western blotting: ICAM-1 polyclonal antibody (Proteintech, Cat#10020-1-AP, 1:1000), CXCR1 rabbit polyclonal antibody (ABclonal, Cat#A16386, 1:1000), collagen type I polyclonal antibody (Proteintech, Cat#14695-1-AP, 1:1000), α-smooth muscle actin polyclonal antibody (Proteintech, Cat#14395-1-AP, 1:1000), cathepsin C monoclonal antibody (Santa Cruz, Cat# sc-74590, clone D-6, 1:1000), β-actin monoclonal antibody (Proteintech, Cat#66009-1-Ig, clone 2D4H5, 1:5000), and GAPDH monoclonal antibody (Proteintech, Cat#60004-1-Ig, clone 1E6D9, 1:5000). HRP-conjugated secondary antibodies (Proteintech, Cat# SA00001-1/2, 1:2000) were used.
Cell counting kit-8 assay
Cell viability was assessed using the CCK-8 Assay Kit (APExBIO). Treated cells were incubated with the reagent at 37 °C for 1–4 h, and absorbance was measured at 450 nm using a microplate reader.
Bromodeoxyuridine labeling
Treated slides were incubated with BrdU-containing medium, followed by fixation in 1× PFA overnight. Cells were denatured in 2 N HCl/0.3% Triton ×-100, neutralized with boric acid, and stained with BrdU primary antibodies. BrdU incorporation was detected using BrdU antibody (Santa Cruz, Cat# sc-32323, clone IIB5, 1:200), followed by Alexa Fluor–conjugated secondary antibodies (1:2000). Nuclei were counterstained with DAPI and imaged using an Olympus IX70 fluorescence microscope.
Cell migration assay
Cell migration was evaluated using a scratch wound healing assay. Confluent cell monolayers were scratched using a pipette tip and photographed at 0, 6, 12, and 24 h. Gap distances were quantified using ImageJ.
Enzyme-linked immunosorbent assay (ELISA)
Soluble ICAM-1 levels in BALF, lung tissue, and culture supernatants were quantified using an ELISA kit (R&D Systems). Absorbance at 450 nm was measured using a microplate reader.
Protein cleavage assay
For recombinant CTSC cleavage of ICAM1, recombinant mouse ICAM1 protein (R&D Systems) was incubated with recombinant CTSC (R&D Systems) in cleavage buffer containing 25 mM MES, 50 mM NaCl, and 5 mM DTT at pH 5.5. The reaction was performed at 37 °C for 2 h. After incubation, the reaction was terminated by adding SDS loading buffer, followed by boiling for 5 min. Samples were then analyzed by Western blotting to detect ICAM1 cleavage products. As the assay involved purified recombinant proteins, traditional loading controls (e.g., β-actin or GAPDH) were not applicable. Instead, equal amounts of ICAM1 (200 ng per lane) were loaded across all conditions to ensure comparability.
For ECM-associated CTSC activity, decellularized lung extracellular matrix (ECM) was first incubated with recombinant cathepsin L (CTSL; R&D Systems) at 37 °C for 20 min in cleavage buffer (25 mM MES, 50 mM NaCl, 5 mM DTT, pH 5.5) to activate endogenous CTSC retained within the ECM. After pre-treatment, recombinant ICAM1 protein was added and incubated with the ECM at 37 °C for 6 h. The reactions were stopped by the addition of SDS loading buffer and boiling for 5 min, and the samples were subsequently analyzed by Western blot to evaluate ICAM1 cleavage. As the ECM is a non-cellular substrate, conventional loading controls were not applicable. To ensure transparency, we explicitly state the amount of recombinant protein (200 ng) and ECM loaded for each condition in the main figure (Fig. 4K).
Preparation and culture of precision-cut lung slices (PCLS)
Mouse precision-cut lung slices (PCLS) were prepared as follows. Briefly, C57BL/6J mice (6–8 weeks old) were euthanized, and the lungs were inflated with 1.5 mL of 2% low-melting-point agarose (Sigma) in pre-warmed (37 °C) Advanced DMEM/F-12 tissue medium. The inflated lungs were immediately placed on ice for agarose solidification, then excised and embedded in 4% agarose for slicing. Using a vibratome (Leica VT1200S), lung tissues were sectioned into 250 µm thick slices. Slices were then washed and kept individually in 24-well plates in DMEM/F-12 supplemented with 0.1% FBS, 1% Penicillin/Streptomycin at 37 °C, 5% CO2, and 100% humidity. Tissue viability was assessed using the LDH Cytotoxicity Detection Kit (Roche) according to the manufacturer’s protocol. PCLS treated with 1% Triton ×-100 in PBS served as the positive control.
Quantification and statistical analysis
All statistical analyses were performed using GraphPad Prism 9.0.0. Two-sided Student’s t-test was used for two-group comparisons, while one-way or two-way ANOVA was applied for multi-group analyses. Data are presented as mean ± SEM, with significance defined as *P < 0.05, **P < 0.01, or ***P < 0.001. Single-cell RNA-seq and spatial transcriptomic data were processed and analyzed in R v4.3.2 using Seurat v4.4.0 (CRAN), Harmony v1.1.0 (CRAN), CellChat v1.6.0 (GitHub), and clusterProfiler v4.10.0 (Bioconductor). Integration of scRNA-seq and spatial data was performed following established Seurat pipelines. GSVA v1.50.0 (Bioconductor) was used for pathway enrichment. ECM proteomic downstream analyses were conducted in R v4.3.2 with standard Bioconductor workflows. Structural modeling and molecular docking were conducted with the HDOCK server (http://hdock.phys.hust.edu.cn/). Image analysis and quantification were performed using ImageJ (Fiji distribution, v1.54, https://fiji.sc/).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this study are available in the article and its Supplementary Information. Source data are provided as Source Data file and may be obtained from the corresponding authors upon request. The sequencing data reported in this paper are deposited in NCBI Gene Expression Omnibus (GEO) database with accession numbers GEO: GSE183683 ((https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE183683), GSE183682). The proteomics data have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository under accession code PXD028194 (https://www.iprox.cn/page/project.html?id=IPX0003445000). No restrictions apply to data access. Source data are provided with this paper.
References
Cavalin, C. et al. Beyond silicosis, is the world failing on silica hazards? Lancet Respir. Med. 7, 649–650 (2019).
Wise, J. Doctors call for ban on cutting artificial stone after reporting first UK cases of silicosis. BMJ 386, q1755 (2024).
Leung, C. C., Yu, I. T. & Chen, W. Silicosis. Lancet 379, 2008–18 (2012).
Steenland, K. & Ward, E. Silica: a lung carcinogen. CA Cancer J. Clin. 64, 63–9 (2014).
Maciejewska, A. Occupational exposure assessment for crystalline silica dust: approach in Poland and worldwide. Int. J. Occup. Med. Environ. Health 21, 1–23 (2008).
Sato, T., Shimosato, T. & Klinman, D. M. Silicosis and lung cancer: current perspectives. Lung Cancer (Auckl.) 9, 91–101 (2018).
Chen, W. et al. Long-term exposure to silica dust and risk of total and cause-specific mortality in Chinese workers: a cohort study. PLoS Med. 9, e1001206 (2012).
Rees, D. & Murray, J. Silica, silicosis and tuberculosis. Int. J. Tuberc. Lung Dis. 11, 474–84 (2007).
Ehrlich, R. I. et al. Lung function loss in relation to silica dust exposure in South African gold miners. Occup. Environ. Med. 68, 96–101 (2011).
Kirby, T. Australia reports on audit of silicosis for stonecutters. Lancet 393, 861 (2019).
Castranova, V. & Vallyathan, V. Silicosis and coal workers’ pneumoconiosis. Environ. Health Perspect. 108, 675–84 (2000).
The Lancet Respiratory Medicine The world is failing on silicosis. Lancet Respir. Med. 7, 283 (2019).
Kamiya, M. et al. Immune mechanisms in fibrotic interstitial lung disease. Cell 187, 3506–3530 (2024).
Taru, V. et al. Inflammasomes in chronic liver disease: hepatic injury, fibrosis progression and systemic inflammation. J. Hepatol. 81, 895–910 (2024).
Shenderov, K. et al. Immune dysregulation as a driver of idiopathic pulmonary fibrosis. J. Clin. Invest. 131, e143226 (2021).
Sokol, C. L. & Luster, A. D. The chemokine system in innate immunity. Cold Spring Harb. Perspect. Biol. 7, a016303 (2015).
Witter, A. R., Okunnu, B. M. & Berg, R. E. The essential role of neutrophils during infection with the intracellular bacterial pathogen Listeria monocytogenes. J. Immunol. 197, 1557–65 (2016).
Liew, P. X. & Kubes, P. The neutrophil’s role during health and disease. Physiol. Rev. 99, 1223–1248 (2019).
Giacalone, V. D. et al. Neutrophil adaptations upon recruitment to the lung: new concepts and implications for homeostasis and disease. Int. J. Mol. Sci. 21, 851 (2020).
Achaiah, A. et al. Neutrophil lymphocyte ratio as an indicator for disease progression in idiopathic pulmonary fibrosis. BMJ Open Respir. Res. 9, e001202 (2022).
Obayashi, Y. et al. The role of neutrophils in the pathogenesis of idiopathic pulmonary fibrosis. Chest 112, 1338–43 (1997).
Silvestre-Roig, C., Hidalgo, A. & Soehnlein, O. Neutrophil heterogeneity: implications for homeostasis and pathogenesis. Blood 127, 2173–81 (2016).
Qi, X. et al. Identification and characterization of neutrophil heterogeneity in sepsis. Crit. Care 25, 50 (2021).
Ng, L. G., Ostuni, R. & Hidalgo, A. Heterogeneity of neutrophils. Nat. Rev. Immunol. 19, 255–265 (2019).
Buckley, C. D. et al. Identification of a phenotypically and functionally distinct population of long-lived neutrophils in a model of reverse endothelial migration. J. Leukoc. Biol. 79, 303–11 (2006).
Soehnlein, O. & Lindbom, L. Phagocyte partnership during the onset and resolution of inflammation. Nat. Rev. Immunol. 10, 427–39 (2010).
Theocharis, A. D. et al. Extracellular matrix structure. Adv. Drug Deliv. Rev. 97, 4–27 (2016).
Bonnans, C., Chou, J. & Werb, Z. Remodelling the extracellular matrix in development and disease. Nat. Rev. Mol. Cell Biol. 15, 786–801 (2014).
Schiller, H. B. et al. Time- and compartment-resolved proteome profiling of the extracellular niche in lung injury and repair. Mol. Syst. Biol. 11, 819 (2015).
Karamanos, N. K. et al. A guide to the composition and functions of the extracellular matrix. FEBS J. 288, 6850–6912 (2021).
Bollyky, P. L. et al. The role of hyaluronan and the extracellular matrix in islet inflammation and immune regulation. Curr. Diab Rep. 12, 471–80 (2012).
Kolahian, S. et al. Immune mechanisms in pulmonary fibrosis. Am. J. Respir. Cell Mol. Biol. 55, 309–22 (2016).
Fischer, A. et al. Neutrophils direct preexisting matrix to initiate repair in damaged tissues. Nat. Immunol. 23, 518–531 (2022).
Peng, Z. et al. Impaired interferon-gamma signaling promotes the development of silicosis. iScience 25, 104647 (2022).
Pang, J. et al. Comparative transcriptome analyses reveal a transcriptional landscape of human silicosis lungs and provide potential strategies for silicosis treatment. Front Genet 12, 652901 (2021).
Woodfin, A. et al. The junctional adhesion molecule JAM-C regulates polarized transendothelial migration of neutrophils in vivo. Nat. Immunol. 12, 761–9 (2011).
Ramos, T. N., Bullard, D. C. & Barnum, S. R. ICAM-1: isoforms and phenotypes. J. Immunol. 192, 4469–74 (2014).
Gocheva, V. et al. Distinct roles for cysteine cathepsin genes in multistage tumorigenesis. Genes Dev. 20, 543–556 (2006).
Robledo, O. et al. ICAM-1 isoforms: specific activity and sensitivity to cleavage by leukocyte elastase and cathepsin G. Eur. J. Immunol. 33, 1351–1360 (2003).
Williams, A. E. & Chambers, R. C. The mercurial nature of neutrophils: still an enigma in ARDS? Am. J. Physiol. Lung Cell Mol. Physiol. 306, L217–L30 (2014).
Lv, Y. et al. YAP controls endothelial activation and vascular inflammation through TRAF6. Circ. Res 123, 43–56 (2018).
Tang, Y. et al. Protein kinase C-delta inhibition protects blood-brain barrier from sepsis-induced vascular damage. J. Neuroinflammation 15, 309 (2018).
Nourshargh, S., Renshaw, S. A. & Imhof, B. A. Reverse migration of neutrophils: where, when, how, and why? Trends Immunol. 37, 273–286 (2016).
de Oliveira, S., Rosowski, E. E. & Huttenlocher, A. Neutrophil migration in infection and wound repair: going forward in reverse. Nat. Rev. Immunol. 16, 378–91 (2016).
Burn, T. & Alvarez, J. I. Reverse transendothelial cell migration in inflammation: to help or to hinder? Cell Mol. Life Sci. 74, 1871–1881 (2017).
Colom, B. et al. Leukotriene B4-neutrophil elastase axis drives neutrophil reverse transendothelial cell migration in vivo. Immunity 42, 1075–86 (2015).
Ji, J. & Fan, J. Neutrophil in reverse migration: role in sepsis. Front Immunol. 12, 656039 (2021).
Herrero-Cervera, A., Soehnlein, O. & Kenne, E. Neutrophils in chronic inflammatory diseases. Cell Mol. Immunol. 19, 177–191 (2022).
Yang, L. et al. Neutrophils in tissue injury and repair: molecular mechanisms and therapeutic targets. MedComm 6, e70184 (2025).
Hirano, Y., Aziz, M. & Wang, P. Role of reverse transendothelial migration of neutrophils in inflammation. Biol. Chem. 397, 497–506 (2016).
Harjunpaa, H. et al. Cell adhesion molecules and their roles and regulation in the immune and tumor microenvironment. Front. Immunol. 10, 1078 (2019).
Cavallaro, U. & Dejana, E. Adhesion molecule signalling: not always a sticky business. Nat. Rev. Mol. Cell Biol. 12, 189–97 (2011).
Lee, I. T. et al. Overexpression of HO-1 protects against TNF-alpha-mediated airway inflammation by down-regulation of TNFR1-dependent oxidative stress. Am. J. Pathol. 175, 519–32 (2009).
Staunton, D. E. et al. Primary structure of ICAM-1 demonstrates interaction between members of the immunoglobulin and integrin supergene families. Cell 52, 925–33 (1988).
Cheng, S. E. et al. Thrombin induces ICAM-1 expression in human lung epithelial cells via c-Src/PDGFR/PI3K/Akt-dependent NF-kappaB/p300 activation. Clin. Sci. 127, 171–83 (2014).
Qian, W. J. et al. Intercellular adhesion molecule-1 (ICAM-1): From molecular functions to clinical applications in cancer investigation. Biochim. Biophys. Acta Rev. Cancer 1879, 189187 (2024).
DiStasi, M. R. & Ley, K. Opening the flood-gates: how neutrophil-endothelial interactions regulate permeability. Trends Immunol. 30, 547–56 (2009).
Fiore, E. et al. Matrix metalloproteinase 9 (MMP-9/gelatinase B) proteolytically cleaves ICAM-1 and participates in tumor cell resistance to natural killer cell-mediated cytotoxicity. Oncogene 21, 5213–5223 (2002).
Tsoutsou, P. G. et al. ICAM-1, ICAM-2 and ICAM-3 in the sera of patients with idiopathic pulmonary fibrosis. Inflammation 28, 359–64 (2004).
Lee, J. S., Shin, J. H. & Choi, B. S. Serum levels of IL-8 and ICAM-1 as biomarkers for progressive massive fibrosis in coal workers’ pneumoconiosis. J. Korean Med. Sci. 30, 140–4 (2015).
Okuda, R. et al. Soluble intercellular adhesion molecule-1 for stable and acute phases of idiopathic pulmonary fibrosis. Springerplus 4, 657 (2015).
Li, X. et al. Associations of serological biomarkers of sICAM-1, IL-1beta, MIF, and su-PAR with 3-month mortality in acute exacerbation of idiopathic pulmonary fibrosis. Med. Inflamm. 2020, 4534272 (2020).
Fernandez, I. E. et al. Systematic phenotyping and correlation of biomarkers with lung function and histology in lung fibrosis. Am. J. Physiol. Lung Cell Mol. Physiol. 310, L919–L27 (2016).
Lopez-Campos, J. L. et al. Increased levels of soluble ICAM-1 in chronic obstructive pulmonary disease and resistant smokers are related to active smoking. Biomark. Med. 6, 805–11 (2012).
Shijubo, N. et al. Soluble intercellular adhesion molecule-1 (ICAM-1) in sera and bronchoalveolar lavage fluid of patients with idiopathic pulmonary fibrosis and pulmonary sarcoidosis. Clin. Exp. Immunol. 95, 156–61 (1994).
Witkowska, A. M. & Borawska, M. H. Soluble intercellular adhesion molecule-1 (sICAM-1): an overview. Eur. Cytokine Netw. 15, 91–8 (2004).
Korkmaz, B. et al. Structure-based design and in vivo anti-arthritic activity evaluation of a potent dipeptidyl cyclopropyl nitrile inhibitor of cathepsin C. Biochem. Pharmacol. 164, 349–367 (2019).
Hamon, Y. et al. Neutrophilic cathepsin C is maturated by a multistep proteolytic process and secreted by activated cells during inflammatory lung diseases. J. Biol. Chem. 291, 8486–8499 (2016).
Korkmaz, B. et al. Neutrophil proteinase 3 and dipeptidyl peptidase I (cathepsin C) as pharmacological targets in granulomatosis with polyangiitis (Wegener granulomatosis). Semin. Immunopathol. 35, 411–421 (2013).
Xiao, Y. et al. Cathepsin C promotes breast cancer lung metastasis by modulating neutrophil infiltration and neutrophil extracellular trap formation. Cancer Cell 39, 423–437.e7 (2021).
Korkmaz, B. et al. Lung protection by cathepsin C Inhibition: a new hope for COVID-19 and ARDS?. J. Med. Chem. 63, 13258–13265 (2020).
Salcher, S. et al. High-resolution single-cell atlas reveals diversity and plasticity of tissue-resident neutrophils in non-small cell lung cancer. Cancer Cell 40, 1503–1520 e8 (2022).
Bailey, J. I. et al. Profibrotic monocyte-derived alveolar macrophages are expanded in patients with persistent respiratory symptoms and radiographic abnormalities after COVID-19. Nat. Immunol. 25, 2097–2109 (2024).
Su, E. Y. et al. Direct comparison of mass cytometry and single-cell RNA sequencing of human peripheral blood mononuclear cells. Sci. Data 11, 559 (2024).
Oetjen, K. A. et al. Human bone marrow assessment by single-cell RNA sequencing, mass cytometry, and flow cytometry. JCI Insight 3, e124928 (2018).
Booth, A. J. et al. Acellular normal and fibrotic human lung matrices as a culture system for in vitro investigation. Am. J. Respir. Crit. Care Med. 186, 866–76 (2012).
Finisguerra, V. et al. MET is required for the recruitment of anti-tumoural neutrophils. Nature 522, 349–53 (2015).
Coffelt, S. B. et al. IL-17-producing γδ T cells and neutrophils conspire to promote breast cancer metastasis. Nature 522, 345–348 (2015).
Srinivasan, B. et al. TEER measurement techniques for in vitro barrier model systems. SLAS Technol. 20, 107–26 (2015).
Yang, S. et al. Single-cell transcriptome sequencing-based analysis: probing the mechanisms of glycoprotein NMB regulation of epithelial cells involved in silicosis. Part Fibre Toxicol. 20, 29 (2023).
Zhou, X. et al. Macrophage-derived MMP12 promotes fibrosis through sustained damage to endothelial cells. J. Hazard Mater. 461, 132733 (2024).
Acknowledgements
This work was supported by grants from the Basic Research Program of Jiangsu Province (No. BK20253013, C.J.), and also supported by Jiangsu Province Science and Technology Plan Project “Provincial Frontier Technology R&D Program” (BF2024054, C.J.) and the National Natural Science Foundation of China (No.82373547, C.J.).This work was also supported by grants from the National Key R&D Program of China (No. 2022YFC2504403, Z.X.X.) and the Young Scientists Fund of the National Natural Science Foundation of China (Grant No.82204004, H.J.). This work was also supported by the Natural Science Foundation of Jiangsu Province, China (No. BK20221184, F.S.C.). We thank LetPub (www.letpub.com) for linguistic assistance and pre-submission expert review. This research work is supported by the Big Data Computing Center of Southeast University. We thank Xin Zheng for her assistance in creating the graphic abstract. We thank Dr. Jianming Zeng and his bioinformatics team for their reference codes. We thank Prof. Chi Liu and Prof. Jianmei Ma for providing the CTSC knockout mice.
Author information
Authors and Affiliations
Contributions
C.J. and F.S.C. conceived and supervised the study. Y.L.L., S.P.P., W.J., X.M.J., and F.S.C. designed the experiments. Y.L.L., S.P.P., W.J., and X.M.J. performed and analyzed the main experiments. Z.S.F., C.H.B., L.W., Z.W., and Z.X.X. contributed to experimental work. X.M.J., Y.K.Q., R.Y., M.R.Q., Z.B.Y., L.C., and M.J.M. provided additional support for experiments and manuscript preparation. Y.L.L., S.P.P., C.J., W.J., and F.S.C. wrote the manuscript, and all authors reviewed and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Georges Helou, Clifford Taggart, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, L., Sun, P., Wang, J. et al. Extracellular matrix anchored neutrophils drive pulmonary fibrosis in mice. Nat Commun 16, 11655 (2025). https://doi.org/10.1038/s41467-025-66633-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66633-8