Abstract
Intestinal stem cells (ISCs) reside in regionally variable niches that provide diverse microenvironmental cues such as tissue oxygen status, and morphogen signaling. Integration of these cues with ISC metabolism and fate remains poorly understood. Here, we show that cellular redox balance orchestrates niche factors with metabolic state to govern cell fate decisions. We demonstrate that hypoxia and Wnt signaling synergistically restrict the reactive oxygen species generating enzyme NADPH oxidase 1 (NOX1) regionally to the crypt base in the distal colon. NOX1 enables maintenance of an oxidative cell state that licenses cell cycle entry, altering the balance of asymmetric ISC self-renewal and lineage commitment. Mechanistically, cell redox state directs a self-reinforcing circuit that connects hypoxia inducible factor 1α-dependent signaling with post-translational regulation of the metabolic enzyme isocitrate dehydrogenase 1. Our studies show redox balance acts as a cellular rheostat that is central and causative for metabolic control of the ISC cell-cycle.
Similar content being viewed by others
Introduction
The maintenance, renewal and response to stress of post-natal tissues relies on the activity of compartment-specific multi-potent stem cells1,2. The intestinal epithelium undergoes rapid constant renewal that is dynamically regulated by stem cells that reside in the base of crypt glands. In both the small and large intestine, cycling stem cells, marked by high expression of the leucine-rich repeat-containing G-protein coupled receptor 5 (Lgr5), control epithelial cell density and the generation of various differentiated epithelial cell lineages3. Disruption of the balance between self-renewal and differentiation results in a wide range of diseases from developmental disorders to cancer.
Stem cell replication for either self-renewal or progenitor cell population generation is dependent on efficient progression through the cell cycle and requires environment-specific metabolic adaptations for energy and biomass production. Metabolic changes in intestinal epithelial stem-cells (ISCs) are well appreciated to have a critical role in regulating proliferation and differentiation4. The niche environment surrounding all tissue resident stem cells, including ISCs, provides key variables that shape metabolic state, including nutrient provision, oxygen availability and developmental signals such as Wnt5. Most studies to-date have focused on ISC metabolism in the context of maintenance and regeneration in the small intestine6,7,8,9,10. The intestine however exhibits considerable differences in niche environment along its cephalo-caudal axis. The mammalian distal colon is a relatively unique mucosal environment with major differences in blood supply, oxygen tension and nutrient use from the small intestine as well as from the proximal colon11,12. In humans the distal colon also represents a relatively fragile environment as suggested by the observation that major disease states such as inflammatory bowel disease or cancer are concentrated or in some case originate at this tissue site.
Several lines of evidence suggest that ISC metabolism is characterized by relative reliance on aerobic glycolysis and a lower fraction of oxidative phosphorylation with important contributions from neighboring secretory epithelial cells7,13,14. An important factor in maintenance and switching of metabolic state in proliferating cells in other tissues is cellular redox balance, with oxidative metabolism reliant on the generation of both reductive and oxidative equivalents in the form of NADH or NAD+ respectively. Redox signaling and balance in relation to stem cell function has been linked to metabolic homeostasis in other tissues including pluripotent stem cells15,16. However, how cytosolic redox balance in the context of regional niche-specific cues such as oxygen availability, affects intestinal stem cell metabolic status and functions in homeostatic intestinal epithelial renewal and regeneration remains poorly understood.
Here we investigated the role of a major colonic epithelial reactive oxygen species (ROS) producing enzyme, NADPH oxidase 1 (NOX1) in tissue maintenance and regeneration. We show that the unique spatial restriction of NOX1 expression to the distal versus proximal colon crypt base results in a compartment specific role in regulating stem cell renewal, and the balance between proliferation and differentiation in both homeostasis and active regeneration. Using cell-specific redox and stem cell reporter mice, along with single-cell transcriptomics, cysteine proteomics, and cell metabolic studies we show that NOX1 maintains cytosolic redox balance to enable glycolytic metabolism, efficient cell cycle progression and favor distal colonic ISC self-renewal versus differentiation.
Results
NOX1 controls cytosolic redox status in crypt base stem cells in the distal colon
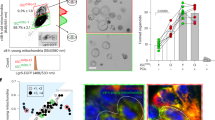
Several NADPH oxidase enzymes have been shown to be functionally important in intestinal epithelial cells along the gastrointestinal tract including NOX1, NOX4 and DUOX217,18. Previous studies have shown increasing Nox1 expression in colonic versus small intestine mucosal tissue19 and have suggested increased expression within colonic stem cells20. We therefore sought to precisely spatially characterize Nox1 expression in WT mice. We found that Nox1 is very specifically expressed in distal versus proximal colonic crypt base epithelial cells (Fig. 1a–c and Supplementary Fig. 1a). Moreover, Nox1 is primarily located within the crypt base stem cell zone, along with Lgr5+ cells, in contrast to DUOX2 which is largely expressed in colonic surface epithelial cells (Fig. 1d and Supplementary Fig. 1b). We also found that Nox1 is most highly expressed in Lgr5high intestinal epithelial cells (IECs) (Supplementary Fig. 1c). To quantitatively assess the intracellular redox status in IECs, we constructed a cell-specific redox reporter mouse (Rosa26-roGFP2-Orp1flox/flox) (Supplementary Fig. 1d, e), where the fluorescence ratio (405 nm and 488 nm) measures relative cytosolic hydrogen peroxide (H2O2) concentration21 and crossed this with Villin-Cre to generate an IEC-specific hydrogen peroxide reporter mouse (Supplementary Fig. 1f). To validate the probe dynamic range and assess levels of oxidized versus reduced probe, colon tissues were treated with either the oxidant diamide or reducing agent dithiothreitol (DTT) (Supplementary Fig. 1g, h). Consistent with the spatial restriction of Nox1 the crypt base in the distal colon is relatively more oxidized than the upper crypt as well as the crypt tip (Fig. 1e, f). Comparison of WT;roGFP2-Orp1Vil to NOX1KO;roGFP2-Orp1Vil showed significantly lower cytosolic H2O2 and relative less cellular oxidation in distal colonic crypt base cells when NOX1 is deficient (Fig. 1g, h). Colonoids generated from the distal colon of NOX1KO redox-reporter mice similarly showed significantly lower steady-state relative cellular H2O2 (Fig. 1i–l). These data along with the observation that the expression of other NOX enzymes were unchanged in NOX1KO mice (Supplementary Fig. 1i) suggest that NOX1 controls cytosolic redox status in crypt base stem cells.
a qRT-PCR for Noxs expression in proximal colons and distal colons. Values were normalized to Rpl13a. Each dot indicates one mouse (n = 6 mice). b, c RNAScope images (b) and quantification of relative Nox1 mRNA intensity (c) of vertical sections of freshly frozen proximal colons and distal colons, probed for Nox1 mRNA in red, counterstained with Hoechst in blue. Each dot indicates one ROI (n = 3 mice, 3 or 4 ROI/mouse). d RNAScope images of vertical sections of freshly frozen proximal colons and distal colons, probed for Nox1 mRNA (green) and Lgr5 mRNA (red) (n = 3 mice). e, f Representative 405/488-nm ratio image (e) and quantification of 405/488-nm ratios (f) of vertical sections from freshly frozen distal colons from R26-RoGFP2-Orp1Villin mice. Each dot indicates one crypt (n = 3 mice). g, h Schematic of the generation of NOX1KO; R26-RoGFP2-Orp1Villin mice (left) and Representative 405/488-nm ratio images (right) and quantification of 405/488-nm ratios (h) of distal colon explant from (g) mice. Each dot indicates one ROI (n = 3 mice, 7 or 8 ROI/mouse). i, j Flow cytometry plot (i) and quantification of 405/488-nm ratios (j) from distal colon-derived colonoids of (g) mice. Each dot indicates one cell (n = 2 mice). k, l Representative 405/488-nm ratio images (k) and quantification of 405/488-nm ratios (l) from distal colon-derived colonoids of (g) mice. Each dot indicates one colonoid (n = 3 mice). Experiments were performed no less than 2 times. Data are presented as mean ± SEM. Statistical significance was determined using two-tailed unpaired Student’s t tests in (c), (f), (h), (j), (l), and Two-way ANOVA in (a); ns, not significant; ****p < 0.0001.
NOX1-derived ROS regulates the balance of self-renewal and differentiation in the distal colon in homeostasis and during regeneration
Given the very specific expression of Nox1 in crypt base epithelial cells along with Lgr5, we hypothesized that NOX1 may play a role in regulating epithelial cellular homeostasis in the distal colon. Assessment of colonic tissues in NOX1KO mice indicated shorter total colon length, distal crypt depth and reduced epithelial cellularity per crypt compared to cage-matched littermate WT mice (Supplementary Fig. 2a and Fig. 2a–c), suggesting that loss of NOX1 is associated with altered epithelial homeostasis at baseline. Measurement of actively cycling epithelial cells via BrdU incorporation experiment showed that NOX1KO mice have reduced proliferation in the distal colon (Fig. 2d, e), but not in the proximal colon (Supplementary Fig. 2b, c), indicating a region-specific effect of NOX1. Consistent with previous reports22, loss of NOX1 leads to increased proportions of MUC2+ goblet cells. In addition, we also found increases in other secretory lineage cells including REG4+ deep crypt secretory cells and CHGA+ enteroendocrine cells in NOX1KO mice (Supplementary Fig. 2d–g). To assess the role of NOX1 in active regeneration of the epithelial compartment, we utilized a fasting-refeeding model (Supplementary Fig. 2h)23 to avoid confounding tissue damage and overt inflammation seen in mucosal injury-based models24. In WT mice we observed that the proportions of Ki67+ cycling IECs were strongly decreased upon fasting and subsequently increased above upon refeeding (Supplementary Fig. 2i, j) indicating active regeneration23. Interestingly, Nox1 expression and cellular H2O2 levels (Fig. 2f, g and Supplementary Fig. 2k) in crypt cells followed a very similar and parallel pattern of changes to Ki67 implying a role for NOX1 function in IEC regeneration upon refeeding. Further evidence for this was provided by NOX1 deficient mice which showed comparable body weight changes (Supplementary Fig. 2l) but impaired regeneration upon refeeding (Fig. 2g, h).
a–c H&E staining images (a), quantification of crypt depths (b) and cell number per crypt (c) of distal colons from WT and NOX1KO littermates under baseline. Each dot indicates one ROI in (b) (n = 3 mice, 4 or 5 ROI/mouse) and one crypt in (c) (n = 3 mice, 14 or 15 crypts/mouse). d, e Flow cytometry plot (d) and quantification (e) of BrdU incorporation in intestinal epithelial cells (Live/dead−CD45−EpCam+) from distal colons of WT and NOX1KO littermates under baseline. Each dot indicates one mouse (n = 6 mice). f–h Representative fluorescence images (g) and quantifications (f) (n = 4 mice, 2 or 3 ROI/mouse) of Nox1 mRNA intensity, quantifications (h) of Ki67+ cells per crypt of distal colons from WT and NOX1KO littermates under fasting and refeeding. Each dot indicates one ROI (n = 7 mice in WT group, n = 5 in NOX1KO group, 8 ROI/mouse). i–k Bright field images (i), colonoids formation numbers (j) and surface area (k) of colonoids generated from isolated crypts in distal colons of WT and NOX1KO littermates, treated with or without NAC or diamide separately. Each dot indicates one replicate in (j) and one organoid in (k) (n = 3 mice per group). l Percentage stem colonoids (spheroid) in WT and NOX1KO three days after plating and in WT following incubation with catalase (n = 4 mice). m Percentage stem colonoids (spheroid) in WT and NOX1KO three days after plating and in NOXKO after addition of glucose oxidase and glucose (n = 4 mice). Experiments were performed no fewer than 2 times. Data are presented as mean ± SEM. Statistical significance was determined using two-tailed unpaired Student’s t tests in (b), (c) and (e), one-way ANOVA in (f), (l) and (m), Two-way ANOVA in (h), (j) and (k); ns, not significant; *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
To understand whether NOX1 mediated changes were related to cellular redox balance we turned to stem-cell derived colonoids, which allow targeted manipulation of the cellular environment and when grown in high-Wnt medium are at baseline an in vitro regeneration model25. Generation of colonoids from the distal colon resulted in decreased secondary organoid formation capacity in NOX1KO compared to WT, as well as reduced surface area (Fig. 2i–k). The NOX1 deficient cellular phenotype was recapitulated in WT colonoids by reducing cellular ROS and increasing relative cellular reductive capacity with n-acetylcysteine (NAC) treatment. Conversely increasing cellular oxidation capacity using diamide partially rescued NOX1KO colonoids (Supplementary Fig. 2m, and Fig. 2i–k), suggesting that NOX1-derived ROS modulates cycling IEC self-renewal. Morphologically, NOX1KO colonoids budded earlier and more extensively than WT, which was recapitulated by treatment with the ROS scavenger catalase in WT colonoids (Supplementary Fig. 2n and Fig. 2l). Increased steady state environmental H2O2 generated by addition of exogenous glucose oxidase in the media partially reversed increased differentiation seen in NOX1KO colonoids as measured by the proportion of cyst-like stem colonoids (Supplementary Fig. 2n and Fig. 2m). Analysis of specific intestinal epithelial cell markers in colonoids showed increased secretory and absorptive lineage markers, but reduced Ki67 expression (Supplementary Fig. 3a). Immunofluorescence to identify mature cell types showed higher proportions of committed IECs with reduced cycling cells in NOX1KO colonoids (Supplementary Fig. 3b-g), consistent with the findings observed in distal colonic tissues. This increase in committed IEC lineage markers induced by NOX1 deficiency was reversed by elevating cellular H2O2 through treatment with the oxidant diamide (Supplementary Fig. 3h, i). Taken together, these data suggest that NOX1-mediated ROS production and consequent alteration of cellular redox balance modulates the balance of epithelial self-renewal and differentiation in the distal colon.
The intestinal stem cell-intrinsic function of NOX1 regulates self-renewal and cell fate in the distal colon
We next investigated whether NOX1-dependent regulation of the balance of IEC proliferation and differentiation were due to changes in either ISC proportion and/or ISC function. We found that Lgr5 expression levels were similar between WT and NOX1KO mice (Supplementary Fig. 4a and Fig. 3a). To further evaluate the question of a functional or proportional role in ISCs, we generated NOX1KO Lgr5 reporter mice (NOX1KO; Lgr5creERT2/+, Supplementary Fig. 4b) to compare with WT; Lgr5creERT2/+ mice. Proportions of Lgr5-EGFP+ ISCs were similar between WT and NOX1KO mice in both the distal and proximal colons (Fig. 3b and Supplementary Fig. 4c). However, we found that NOX1 deficient distal colonic ISCs exhibited reduced BrdU incorporation in steady state conditions with impaired upregulation of incorporation during the active regeneration phase that occurs after re-feeding (Fig. 3c, d). In contrast, and providing further evidence of a region-specific role, loss of NOX1 had no impact on the proportion of actively cycling ISCs in the proximal colon in both the steady state or during regeneration (Supplementary Fig. 4d, e). To further investigate the cell-intrinsic function of NOX1, ISCs were isolated from WT; Lgr5creERT2/+ and NOX1KO; Lgr5creERT2/+ mice separately and plated to generate colonoids. Distal colonic NOX1 deficient ISCs showed impaired primary organoid formation capacity compared to WT ISCs (Fig. 3e, f). Consistent with our tissue data, ISCs isolated from the proximal colon displayed similar primary organoid formation capacity in both WT and NOX1KO mice (Supplementary Fig. 4f, g).
a Quantification of Lgr5 mRNA intensity in Supplementary Fig. 4a from freshly frozen distal colons. Each dot indicates one ROI (n = 3 mice, 2 or 3 ROI/mouse). b Proportions of Lgr5-EGFP+ cells (gated from Live/dead−CD45−EpCam+) from distal colons of WT; Lgr5creERT2+ and NOX1KO; Lgr5creERT2+ naïve mice. Each dot indicates one mouse (n = 5 mice). c, d Flow cytometry plot (c) and proportions (d) of BrdU+ Lgr5-EGFP+ cells (gated from Live/dead−CD45−EpCam+Lgr5-EGFP+) from distal colons of WT; Lgr5creERT2+ and NOX1KO; Lgr5creERT2+ mice under fasting and refeeding. Each dot indicates one mouse (n = 5 mice). e, f Bright field images (e) and quantifications (f) of primary organoid formation per 1000 sorted Lgr5-EGFP+ cells of distal colons from either WT; Lgr5creERT2+ or NOX1KO; Lgr5creERT2+ mice. Each dot indicates one technical replicate (n = 2 mice). g Generation of lineage tracing mice WT; Lgr5tdTomato and NOX1KO; Lgr5tdTomato mice (left) and representative fluorescence images of freshly frozen distal colon sections (right), immunolabelled for tdTomato and EGFP (n = 3 mice). h, i Flow cytometry plot (h) and quantification (i) of total tdTomato+ cells (gated from Live/dead−CD45−EpCam+) from distal colons of (g) mice. Each dot indicates one mouse (n = 6 mice). j, k Flow cytometry plot (j) and quantification (k) of tdTomato+Lgr5-EGFP+ cells (gated from Live/dead−CD45−EpCam+ Lgr5-EGFP+) from distal colons of (g) mice. Each dot indicates one mouse (n = 6 mice). Experiments were performed no less than 2 times. Data are presented as mean ± SEM. Statistical significance was determined using two-tailed unpaired Student’s t tests in (a), (b), (f), (i) and (k), Two-way ANOVA in (d); ns, not significant; *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
ISC cell fate is thought to follow a neutral drift model, whereby ISCs undergo stochastic asymmetric division with either self-renewal to maintain stemness, or generation of committed cell progenitors26. To explore whether loss of NOX1 function acts to alter this cell fate decision process, we employed in vivo genetic lineage tracing to track ISC cell progeny. WT;Lgr5tdTomato and NOX1KO;Lgr5tdTomato mice were generated to enable ISC cell-tracing following tamoxifen induction (Fig. 3g). After induction, committed IECs from ISCs are positive for tdtomato while daughter ISCs are positive for both EGFP and tdtomato. In the distal colon, loss of NOX1 resulted in reduced accumulation of ISC-derived tdtomato+ cells (Fig. 3h, i) and a significantly smaller proportion of EGFP+ tdtomato+ daughter ISCs (Fig. 3j, k) indicating impaired self-renewal capacity. In contrast, loss of NOX1 had no effect on ISC fate in the proximal colon (Supplementary Fig. 4h–k). These results point to a highly location specific role for NOX1 dependent ROS in regulating ISC fate decisions.
Functional NOX1 regulates cell cycle entry in distal colonic ISCs and transient amplifying cells
To understand at what stage NOX1 regulates the process of ISC fate progression and capture the transcriptional signatures involved, we performed single-cell RNA sequencing (scRNA-seq) of EpCAM+CD45− IECs from both proximal and distal colons in NOX1KO and littermate WT control mice. Louvain-clustering of the resulting single-cell expression profiles revealed 17 cell-type /state clusters in distal colon (Fig. 4a, and Supplementary Fig. 5a, b). This included populations of Lgr5+ ISCs and neighboring actively cycling transient amplifying cells (TAs). Consistent with our RNAscope analysis, Nox1 was predominantly expressed in ISCs, TAs and progenitors (Supplementary Fig. 5c). Broad clustering by genotype in distal colonic IECs showed a small shift in cluster distribution between WT and NOX1KO IECs, in contrast to the proximal colon which showed overlapping distributions (Fig. 4b and Supplementary Fig. 5d–f). This was reflected in a shift in the overall cell type/state distribution providing additional evidence for a distal colon specific role (Supplementary Fig. 5e, f and Fig. 4c). Consistent with our immunofluorescence data, NOX1KO distal IECs showed increased relative abundance of secretory progenitors, secretory lineages and several sub-types of committed IECs (Supplementary Fig. 5f). To trace at what stage NOX1-dependent ROS impacts the progression of ISCs, we carried out trajectory analysis of our single cell data. Figure 4d shows lineage trajectories using Lgr5+ ISCs as an origin across increasing pseudotime. Focusing on the early transition from ISC to TA cells we found two distinct cell populations (Fig. 4e–g) characterized by their relative pseudotime (early and late). These populations include cells that are either Lgr5highKi67+ (early) or Lgr5lowKi67+ (late) (Fig. 4g–i) and therefore represent cells at the point of transition between ISC self-renewal and progenitor lineage commitment. A critical factor in this transition is efficient progression through cell cycle allowing stem cells to maintain the balance between self-renewal and differentiation27. To identify specific transcriptional signatures during this transitional state we interrogated the relative expression of key transcription factors involved in regulating cell cycle progression (Supplementary Fig. 5g) and compared these between WT and NOX1KO cells in both early and late TA populations. We found a distinct pattern of expression in NOX1KO with reduced expression of the cell cycle transcription factor E2F1, one of several key factors for cell cycle progression from G1 phase to S phase. In conjunction we found enhanced expression of E2F5, which regulates G0 phase arrest (Fig. 4j and Supplementary Fig. 5g)28. To investigate potentially compromised cell cycle progression in NOX1KO cycling cells, we carried out cell cycle analysis in colonoids. Here we found that NOX1 deficiency increases the number of cells in G0/G1 arrest with fewer cells in S phase versus WT (Fig. 4k, l). Taken together, this suggests that, in the distal colon, NOX1-derived ROS licenses enhanced progression through the cell cycle to regulate the balance of self-renewal versus differentiation in cycling IECs.
a Uniform Manifold Approximation and Projection (UMAP) of distal epithelial cell clusters. b Uniform Manifold Approximation and Projection (UMAP) for distal colonic epithelial cells colored by genotype-WT (grey) and NOX1KO (blue). c Median percentage change in cell-type abundance between WT and NOX1KO in distal and proximal colon. Each dot indicates one cell cluster (n = 2 mice). d Pseudotime trajectory analysis (Monocle3) with Uniform Manifold Approximation and Projection (UMAP) for distal colonic epithelial cells with pseudotime trajectory across epithelial clusters. e Cell proportions across pseudotime for stem cell and transit amplifying cell clusters in WT and NOX1KO. f Total cell proportions for TA cell cluster binned by pseudotime showing two populations (early and late). g Individual cells within TA cell cluster labelled by pseudotime population. h Relative Lgr5 expression in TA cells. i Relative mki67 expression in TA cells. j Heatmap of cell-cycle associated transcription factors in WT and NOX1KO TA cells in both early and late pseudotime populations. k, l Representative flow plots (k) and quantification (l) of cell cycle analysis in WT and NOX1KO colonoids. Each dot indicates one replicate (n = 3 independent experiments). Experiments were performed no less than 3 times in (k) and (l). Data (c) is presented as median ± SD, and (l) is presented as mean ± SEM. Statistical significance was determined using two-tailed unpaired Student’s t tests in (c), Two-way ANOVA in (l); ns, not significant; *p < 0.05 and ****p < 0.0001.
Loss of functional NOX1 leads to rewiring of ISC metabolism through regulation of IDH1 enzyme activity
To explore the underlying mechanisms of how NOX1-dependent ROS alters ISC function we carried out an exploratory study of redox proteomics using the differential cysteine alkylation method29 to assess NOX1-dependent oxidative post-translational modifications on protein cysteine residues (Supplementary Data 1)30. Using stem-like colonoids we analyzed the relative oxidation of protein cysteines in WT versus NOX1KO, identifying 63 putative proteins with differentially oxidized cysteine residues (Supplementary Fig. 6a). Gene set enrichment analysis of this data using gene ontology (GO) indicated that the most enriched pathway term in NOX1KO cells was inhibition of glycolysis (Supplementary Fig. 6b). Several studies have shown that cell metabolism and switching of energy production pathways can dictate intestinal stem cell fate decision in variety of contexts6,8,9,10,31,32,33. ISC self-renewal has been linked with reliance on glycolysis while lineage commitment and differentiation with oxidative phosphorylation (OXPHOS)34. To initially assess metabolic alterations in the setting of NOX1 deficiency, we looked at expression of key glycolytic genes. We found significantly reduced expression of genes associated with glycolytic metabolism in NOX1 deficient cycling IECs (Fig. 5a). To directly investigate cellular metabolism we carried out intracellular metabolic status analysis35. We found that NOX1KO IECs had lower extracellular acidification rate (ECAR) compared to WT, indicating reduced utilization of glycolytic metabolism (Fig. 5b–d). Conversely, NOX1KO IECs displayed enhanced oxidative phosphorylation, as measured by a significantly increased oxygen consumption rate (OCR) (Supplementary Fig. 7a–d), higher mitochondrial DNA copy (Supplementary Fig. 7e) and enhanced mitochondrial membrane potential (Supplementary Fig. 7f, g).
a qRT-PCR for glycolytic gene expression in colonoids from WT and NOX1KO littermates cultured in Wnt-rich WERN medium. Values were normalized to Rpl13a. Each dot indicates one replicate (n = 3 mice). b–d Kinetic line graph (b) of extracellular acidification rate (ECAR) in 2D epithelial monolayers from WT and NOX1KO littermates measured by a Seahorse Analyzer (n = 3 mice). (c) is basal ECAR and (d) is MGC bar graphs. Each dot indicates one replicate (n = 3 mice). e IDH1 enzyme activity assay of colonoids from distal colons of WT and NOX1KO littermates. Each dot indicates one replicate (n = 3 mice). f α-Ketoglutarate levels assay of colonoids (left) from distal colons as well as distal colon tissues (right) of WT and NOX1KO littermates. Each dot indicates one replicate (n = 3 mice) in (left) and one mouse in (right). g, h Flow cytometry plot (h) and quantification (g) of BrdU incorporation in colonoids from distal colons of WT and NOX1KO littermates, treated with or without α-Ketoglutarate or IDH1 inhibitor (compound 13). Each dot indicates one technical replicate (n = 3 mice). Experiments were performed no less than 2 times. Data are presented as mean ± SEM. Statistical significance was determined using two-tailed unpaired Student’s t tests in (c–f), Two-way ANOVA in (a) and (g); ns, not significant; *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
Network analysis of our redox proteomic data also suggested that altered carbon metabolism and glycolysis was one of the main features of potential differentially oxidized proteins (Supplementary Fig. 6c). One of the most differentially altered proteins and a central node in the NOX1-dependent glycolysis network was the cytoplasmic enzyme isocitrate dehydrogenase 1 (IDH1). IDH1 is a key cellular NADP+-dependent metabolic enzyme which reversibly catalyzes the oxidative decarboxylation of isocitrate to non-mitochondrial alpha-ketoglutarate (αKG), an important metabolite regulator of cell metabolism and proliferation36. Measurement of IDH1 enzyme activity in colonoids revealed higher IDH1 enzyme activity in NOX1 deficiency (Fig. 5e). Consistent with this, cellular αKG levels from either NOX1KO colonoids or distal colonic tissues were higher than in WT equivalents (Fig. 5f). Given previous reports that supplementation of αKG can promote differentiation and suppresses proliferation in patient-derived colon tumor organoids37,38, we hypothesized that manipulation of cellular αKG levels could either rescue or phenocopy the proliferation status of NOX1KO or WT cycling IECs respectively. We found cell-permeable αKG treatment could suppress cell cycling and proliferation, recapitulating the phenotype induced by NOX1 deficiency (Fig. 5g, h). Conversely, administration of a wild type IDH1 inhibitor (compound 13)39 was able to restore the reduced cycling and proliferation observed in NOX1KO colonoids (Fig. 5g, h). Taken together, these results suggest that NOX1 alters the metabolic status of distal colonic cycling cells and acts via regulation of IDH1 enzyme activity to modulate ISCs function. Since only two replicates were included in each group for redox proteomics, further work is needed to confirm changes indicated in Supplementary Fig. 6a–c.
The oxygen environment of the distal colonic basal crypt niche regulates NOX1 expression and function via HIF1α signaling
ISC self-renewal and differentiation is critically dependent on input by surrounding environmental niche factors2,40. Our data showing that both the expression and functional effects of NOX1 are restricted to the distal colon led us to explore regionally-specific cues in the crypt base niche environment. The colon harbors the greatest microbial load within the intestine41, and therefore as with other ROS-generating proteins such as Duox219, we speculated whether microbial cues may regulate the highly specific localization of Nox1 expression. We therefore investigated Nox1 expression in the colon of GF mice. Here we found the expression pattern of Nox1 was identical to that seen in SPF mice, with Nox1 highly expressed in the distal colonic crypt base, compared to proximal colon (Supplementary Fig. 8a–c) and suggesting against a major role for intrinsic regulation by the microbial environment.
A major niche factor that also varies along the longitudinal axis of the intestine is tissue oxygen tension, with studies suggesting that the distal colonic mucosa is a relatively more hypoxic environment relative to more proximal intestinal regions42,43. Cellular responses to oxygen tension are primarily mediated by hypoxia-inducible factors (HIFs) such as HIF1α42,44. To investigate this we analyzed the spatial distribution of HIF1α protein in the proximal and distal colon. In addition to increased expression within the surface epithelium, stabilized HIF1α was also detected in crypt bases (Fig. 6a, c)45. We found that both Hif1α mRNA expression levels were higher (Supplementary Fig. 8d), and that there was significantly more stabilized HIF1α protein in crypt base cells in the distal versus proximal colon (Fig. 6a, b). Analysis of NOX1KO versus WT distal colon showed lower levels of stable HIF1α in NOX1KO tissue by both IHC and immunoblot analysis (Fig. 6d–f) suggesting NOX1 function is intrinsically linked to the hypoxic niche environment. In keeping with the data on Nox1 expression, GF mice also showed identical HIF1α expression pattern along the colon as SPF mice (Supplementary Fig. 8e–h), providing further evidence for an intrinsic role for hypoxia/HIF1α but not microbes.
a–c DAB analysis (a) and quantifications (b, c) of HIF1α accumulation in proximal and distal colon FFPE samples. Each spot indicates one ROI (n = 3 mice, 5 ROI/mouse). d, e DAB analysis (d) and quantification (e) of HIF1α accumulation in distal colon FFPE samples from WT and NOX1KO littermates. Each dot indicates one ROI (n = 3 mice, 5 ROI/mouse). f Western blot of HIF1α protein in distal colon tissues from WT and NOX1KO littermates. β-actin as the loading control (n = 3 mice). g qRT-PCR for Nox1 expression in colonoids from distal colons of R26-RoGFP2-Orp1Villin mice cultured either in Wnt-rich WERN medium or non-Wnt ERN medium under normoxia (21% O2), or in Wnt-rich WERN medium under hypoxia (1% O2). Values were normalized to Rpl13a. Each dot indicates one replicate (n = 3 independent experiments for Wnt3a, n = 2 independent experiments for O2). h Oxidation of roGFP in response to treatment indicated in (g). Maximum reduction (purple) and oxidation (yellow) of roGFP following additions of Dithiothreitol (DTT) or diamide respectively are shown as a reference. i Schematic (upper) of transcription factors HIF1α and TCF4 binding sites of Nox1 gene. ChIP-qPCR was carried out with DNA fragments immunoprecipitated with anti-HIF1α (right under) or anti-TCF4 (left under) antibody separately. Values are relative to input signals. Each dot indicates one replicate (n = 2 independent experiments). j, k Flow cytometry fluorescence measurements (j) and quantification (k) of 405/488-nm ratios in distal colon-derived colonoids of R26-RoGFP2-Orp1Villin mice under normoxia (21% O2), or hypoxia (1% O2), treated with or without HIF1α inhibitor LW6. Each dot indicates one cell (n = 3 mice). Experiments were performed no less than 2 times. Data are presented as mean ± SEM. Statistical significance was determined using two-tailed unpaired Student’s t tests in (b), (e), (g) and (i), One-way ANOVA in (c) and (k); ns, not significant; *p < 0.05, **p < 0.01 and ****p < 0.0001.
We next employed colonoids to screen the impact of a variety of niche factors on regulation of NOX1 expression and function. In addition to hypoxia, the crypt base also exhibits high concentrations of the signaling factor Wnt3a, which supports the self-renewal of ISCs46,47. We exposed colonoids to niche factors present at the crypt base, including Wnt3a and reduced oxygen tension, versus microbial cues such as LPS and butyrate which are normally restricted to the crypt tip and surface epithelium in health48,49,50. We found both high Wnt3a and lower oxygen efficiently induced Nox1 mRNA expression (Fig. 6g). Consistent with other reports20,49, butyrate lowered, while LPS increased Nox1 expression (Supplementary Fig. 8i). To assess function, we conducted experiments using colonoids generated from our roGFP2-Orp1 redox reporter mice. Here we found that the presence of hypoxia, in the presence of high Wnt3a, resulted in increased H2O2 concentrations producing a relatively oxidizing cellular environment versus either normoxia alone or normoxia along with high Wnt3a (Fig. 6h), suggesting hypoxia and Wnt3a synergistically regulate NOX1 function. Consistent with the expression data, butyrate lowered the relative cellular oxidation level, and LPS stimulated NOX1-dependent ROS (Supplementary Fig. 8j).
Given the synergistic activation of NOX1 by hypoxia and Wnt3a, we investigated whether Nox1 gene expression is directly regulated by these factors. HIF1α protein is stabilized by hypoxia and binds to hypoxia responsive elements (HREs) to drive a set of responsive genes to help cells adapt to the hypoxic environment44. We therefore interrogated the promotor region of Nox1 and located a putative HRE51. Based on a database of human transcription factor targets, we also found that TCF4, a Wnt3a responsive transcription factor, binds around the TSS region of Nox152,53. To directly test whether Nox1 is a hypoxia responsive gene, we then performed ChIP-qPCR and found that both HIF1α as well as TCF4 bind to regulatory regions in Nox1, in a hypoxia dependent manner (Fig. 6i). To further validate a key role of HIF1α in regulating NOX1 function we treated our hydrogen peroxide reporter colonoids with a HIF1α specific inhibitor (LW6) and assessed cellular redox balance. We found that loss of HIF1α activity under hypoxic conditions led to a relatively reduced cytosolic cellular environment confirming that HIF1α stability is required for enhanced NOX1 function under hypoxia (Fig. 6j, k). In addition, inhibition of HIF1α under hypoxic conditions phenocopied both the decreased BrdU incorporation and reduced daughter cell generation observed in NOX1 deficiency (Supplementary Fig. 8k, l). Taken together, these data suggest that the unique hypoxic niche of the distal colonic crypt base, through the synergistic action of HIF1α and Wnt3a, allows ISCs to maintain a relatively oxidized cytosolic environment.
Cellular redox balance orchestrates metabolic status and niche factors to control ISC cell fate
Our data showing that niche factors within the crypt base cooperate to maintain NOX1 function suggest that cellular redox may be the orchestrating factor in regulating ISC cell fate in the distal colon. To explore this further, we carried out bulk RNA-seq analysis in colonoids from the distal colon of NOX1KO mice compared to WT littermates identifying >600 differentially expressed genes (Supplementary Data 2, Fig. 7a). Gene set enrichment and network analysis (Supplementary Fig. 9a and Fig. 7b) pointed to a central role for glutathione metabolism in the transcriptional signature following loss of NOX1. Redox balance in the cell is in part set by the amount of cytosolic reducing equivalents, with glutathione (GSH) levels, the primary quantitative determinant of reductant capacity. Assessment of the GSH/GSSG ratio in colonoids revealed significantly higher ratios in NOX1KO colonoids (Fig. 7c) both in normoxic and hypoxic conditions. Given our data showing the importance of IDH1 activity, αKG levels and HIF1α in ISC proliferation, we investigated the effect of shifting redox balance on IDH1-dependent metabolism. Manipulation of cellular redox balance towards more reduced with NAC, or more oxidized with diamide treatment, induced alterations in IDH1 enzyme activity status and αKG levels that mirrored the changes observed between WT and NOXKO cells (Fig. 7d and Supplementary Fig. 9b). Correspondingly, under hypoxic conditions, stability of HIF1α in WT colonoids was lower in reducing cellular conditions (NAC), and similar to the administration of additional αKG. αKG functions as the substrate of prolyl hydroxylase domain (PHD) proteins and leads to the protein hydroxylation of HIF1α54. We found αKG treatment in WT recapitulated the levels of destabilized HIF1α seen in NOX1KO colonoids (Fig. 7e, f) and was accompanied by lower cellular H2O2 under hypoxia (Fig. 7g). Both NOX1 deficiency (Supplementary Fig. 6c and Fig. 5e) or treatment with NAC (Fig. 7d) result in increased IDH1 activity and destabilization of HIF1α. Conversely, inhibition of IDH1 in NOX1KO cells either by specific enzyme inhibitor, or by increasing cellular redox with diamide, increased HIF1α stability (Fig. 7e–g). Importantly, we found that hypoxia alone, and therefore increased HIF1α, in a reducing environment (Fig. 7c, h and Supplementary Fig. 9c) was insufficient to rescue the impaired proliferation observed following loss of NOX1. This suggests that redox balance, and maintenance of sufficient oxidative equivalents through the activity of NOX1 is central to ISC function in the distal colon. To provide further evidence for this we assessed cell cycle progression following cellular redox manipulation. Shifting the cellular environment towards reduction in WT colonoids (Supplementary Fig. 9d, e) resulted in impaired transition to S phase, phenocopying loss of NOX1. Correspondingly, a shift to increased oxidation under hypoxia in NOX1KO was able to rescue cell cycle progression (Fig. 7i, j).
a Volcano plot of bulk RNA-seq highlights 614 differentially expressed genes (DEGs, in red; log2 fold change >0.5, adjusted p < 0.05) in NOX1KO colonoids versus WT control colonoids. b String network analysis of (a) with enriched terms highlighted. c GSH/GSSG ratio detection assay of colonoids from distal colons of WT and NOX1KO littermates under either normoxia (21% O2) or hypoxia (1% O2). Each dot indicates one replicate (n = 3 mice). d IDH1 enzyme activity assay of colonoids treated with or without NAC or diamide. Each dot indicates one replicate (n = 2 independent experiments). e, f Representative western blot images (e) and quantification (f) of HIF1α protein in colonoids from WT and NOX1KO littermates under either normoxia (21% O2) or hypoxia (1% O2), WT colonoids were treated with or without NAC, α-Ketoglutarate separately as indicated, NOX1KO colonoids were treated with or without diamide, IDH1 inhibitor (compound 13). β-actin as the loading control (n = 2 independent experiments). g 405/488-nm ratios of colonoids under hypoxia in (e) analyzed by flow cytometry. Each dot indicates one cell (n = 3 independent experiments). h Flow cytometry measurements of BrdU incorporation in distal colon-derived colonoids of WT and NOX1KO littermates under normoxia (21% O2), or hypoxia (1% O2) in Supplementary Fig. 9c. Each dot indicates one replicate (n = 3 independent experiments). i, j Representative flow plots (i) and quantification (j) of cell cycle analysis in NOX1KO colonoids under normoxia (21% O2), or hypoxia (1% O2), treated with or without diamide. Each dot indicates one replicate (n = 2 independent experiments). k–p DAB analysis (k) and quantification (n) of HIF1α intensity, fluorescence images (l) and quantifications (o) of PCNA+ cells, AB/PAS staining (m) and quantification of its relative intensity (p) in healthy patient and Nox1 loss-of-function patient sigmoid FFPE samples (n = 2 independent experiments). q Model illustrating that rheostat-like control of cellular redox regulates IDH1 enzyme activity, αKG production to enhance hypoxia/HIF1α signaling, which in turn transcriptionally reinforces NOX1-dependent ROS and glycolytic metabolism to promote efficient cell cycle progression. r Waddington model depicting the epithelial cell-state landscape in the distal colon as controlled by cellular redox. Under hypoxia, high-Wnt and a relatively oxidizing redox state, the trajectory of ISCs favors self-renewal versus lineage commitment. In the absence of NOX1-dependent ROS, the relatively reductive cellular state decreases the probability of ISC self-renewal and skews the lineage hierarchy, altering proportions of absorptive and secretory IECs. Created in BioRender. Thiagarajah, J. (2025) https://BioRender.com/xtui0xc. Experiments were performed no less than 2 times in (c–p). Data are presented as mean ± SEM, Statistical significance was determined using two-tailed unpaired Student’s t tests in (n–p). One-way ANOVA in (d), Two-way ANOVA in (c), (f–h) and (j); ns, not significant; *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
In the human distal colon, loss of function variants in NOX1 have been shown to be clinically important as risk allele in very early-onset inflammatory bowel disease (VEOIBD). We had previously characterized a Nox1 missense mutation (p.N122H) that abrogated cellular ROS production, and resulted in ulcerative colitis-like VEOIBD in a patient55. Patient biopsy sections showed reductions in crypt basal HIF1α stability (Fig. 7k, n), reduced PCNA+ cycling cells (Fig. 7l, o) as well as increased goblet cell proportions (Fig. 7m, p) mirroring the changes in ISC function observed in mouse NOX1KO tissues and colonoids.
Discussion
The regulation and fate of multipotent stem cells relies on a complex interplay between environment, metabolic status and cellular regulatory factors. Our studies here show that the distal colon crypt base represents a significantly different stem cell environment from more proximal intestinal regions. In the distal colon we show that the niche environment maintains NOX1-derived ROS and synergizes to alter the balance of self-renewal and differentiation in cycling stem cells in both homeostasis and active regeneration states (Fig. 7q, r). We propose that relative redox balance acts as a cellular rheostat whereby a shift to either a more oxidative or reductive state is central to metabolic control of the ISC cell-cycle. The hypoxic and high-Wnt niche environment at the distal colonic crypt base transcriptionally determines the unique spatial restriction of Nox1 expression in cycling cells, contributing to a relatively more oxidized cell state. As a result, lower IDH1-dependent decarboxylation of isocitrate to αKG leads to increased HIF1α stability, enhancing both NOX1 expression and glycolysis in ISCs to metabolically maintain self-renewal13.
Loss of NOX1 function shifts cellular redox balance towards relative reduction or ‘reductive stress’ that lowers efficiency of cell cycle entry, stem and TA cell population progression, and ultimately predisposing towards differentiation, including both secretory and absorptive lineages (Fig. 7r). Our single-cell transcriptomic data uncovered alterations in the E2 transcription factor family in NOX1KO cycling transit amplifying cells with lower expression of the ‘activating’ E2F1, important in promoting S-phase initiation, and elevated expression of the ‘repressor’ E2F528,56. These findings, along with data showing that manipulation of cellular ROS and IDH1 activity can mirror the changes in cell cycle seen between WT and NOX1KO, links together cellular redox balance and metabolism with cell-cycle efficiency in colonic ISCs. Despite increased cellular glutathione in NOX1KO cells, our data does also point to the presence of compensatory mechanisms with concomitant inhibition of NRF2-regulated pathways, potentially to limit reductive stress and allow some ISC proliferation.
ROS is thought to play an important role in various colonic diseases including Crohn’s disease, ulcerative colitis, pouchitis and irritable bowel syndrome17,18. Several studies have established NOX1 loss of function variants in the pathogenesis of VEOIBD55,57,58. Examination of the role of NOX1 in colitis showed that TNF-α stimulates NOX1-dependent ROS and leads to enhanced basal lymphoplasmacytosis and alteration in M-cell signatures59. A few previous studies have also explored the role of NOX1 in intestinal epithelium homeostasis. Denis et al. showed that NOX1 deficiency decreased proliferating cell populations while increased goblet cell numbers by attenuating Wnt as well as Notch signaling pathway in colon, in keeping with our observations of a relative increase in secretory cells22. In another study, NOX1-derived ROS was found to be important for IEC proliferation in distal colon through EGFR activity as suggested by in vitro chemical inhibition studies20. In colonic cancer stem cells, NOX1-dependent mTORC1 activation via S100A9 oxidation is important for proliferation and colon cancer progression60. In many of these studies direct function of NOX1 on cellular ROS were either not assessed or limited by non-quantitative redox measurements. Here, we used a genetically encoded, quantitative ratiometric cell-specific reporter in vivo to precisely delineate the cellular and spatial NOX1-dependent regulation of ROS and redox balance. In conjunction with genetic lineage tracing and scRNAseq analysis, this allowed careful dissection of the complex self-reinforcing mechanism that connects HIF1α-dependent hypoxic signaling to NOX1-dependent cellular ROS regulation of isocitrate dehydrogenase 1 (IDH1) in the cytosol, and maintenance of ISC function.
IDH1 is a key cytosolic metabolic enzyme that links cellular metabolism to epigenetic regulation and redox states via control of αKG levels61. αKG supplementation in ApcMin/+ organoids leads to DNA hypomethylation of genes related to differentiation, attenuating Wnt signaling and driving differentiation in colorectal cancer37. Studies have revealed that wild-type IDH1 is overexpressed in several types of cancers61, although we did not observe any changes in Idh1 mRNA levels in NOX1KO cycling stem cells. αKG supplementation can mimic the phenotype of NOX1-loss-of-function, decreasing redox status under hypoxia and restricting cell cycle progression. IDH1 is one of the most mutated metabolic enzymes across human cancers, with mutant IDH1 enzyme exhibiting neomorphic enzymatic activity54. Mutant IDH1 generates the oncometabolite 2-hydroxyglutarate (2-HG), which inhibits αKG-dependent enzymes, including histone, DNA, and RNA demethylases, metabolic enzymes, and DNA repair factors. Our data connecting NOX1-dependent ROS to IDH1 activity may provide a link between redox state and epigenetic modification in cancer progression. In addition to links to colon cancer, the role of redox balance in setting epigenetic regulation in homeostasis or regeneration after injury in colonic ISCs and the effects on the regulatory pathways outlined by our studies need to be explored further.
An interesting finding in our study was that NOX1 and HIF1α mediated effects were intrinsic to the niche environment and independent of luminal microbial input. Previous studies have outlined the importance of HIF1α in epithelial cells during inflammation and for maintaining colonic regeneration during injury62,63,64. Our studies show significantly increased HIF1α protein in crypt base ISCs in the distal versus proximal colon and suggestive of a different oxygen tension within the distal niche environment. In this setting, HIF1α and NOX1 form a self-reinforcing circuit to maintain cellular oxidative balance and hypoxia-induced transcriptional responses including promotion of glycolysis. Several studies have shown that changes in redox-altering proteins including NOX1, DUOX2 or the combined loss of NOX isoforms, induce microbial changes that affect epithelial function and in the case of Duox2 are directly regulated by commensal microbes19,65,66,67. Nox1 is localized at the distal colonic crypt base which is a site largely shielded from microbes in homeostasis. However, during inflammation or barrier breakdown, as shown for the microbial metabolite butyrate and for other microbial products such pathogen associated molecular patterns (PAMPs), microbial or cytokine-related signaling responses are likely modulated by NOX1-dependent ROS in crypt base stem cells.
Redox balance and signaling are thought to play a role in stem cell function in a variety of tissue compartments, but most studies have focused on the deleterious effects of excessive cellular ROS or ‘oxidative stress’. In embryonic pluripotent stem cells (PSCs) and hematopoietic stem cells (HSCs), stem cells maintain very low cytosolic ROS levels in quiescence and elevated ROS levels are associated with activation and progression through differentiation, but also decreased self-renewal and increased stem cell exhaustion68,69. In contrast, in neural stem cells (NSCs), increasing ROS correlates with stem cell proliferation in vivo, and boosts the capacity of NSCs for neurosphere formation70. These and other studies suggest that the role of redox balance in stem cells depends on context, and is likely driven by the metabolic and regenerative requirements of the specific tissue environment. Adult stem cells in mucosal epithelial tissues such as the distal intestine are required to maintain constant self-renewal and proliferation to maintain the rapidly renewing epithelial barrier and are therefore likely to require a balanced redox state to enable efficient metabolism for biomass production. Our data would suggest that in the setting of physiological hypoxia, as occurs in the distal colon, the set-point for this redox balance favors a more oxidative cellular state to enable efficient cell-cycle progression. The crypt base is a niche environment which at homeostasis is geared towards the maintenance and self-renewal of a multipotent stem cell cellular state but during injury can exhibit a range of adaptations including de-differentiation of progenitor cell populations to enable regeneration. How crypt base redox-balance functions to alter these injury-induced changes will be an interesting topic for further investigation.
In summary, our data suggests that, in the distal colon, a self-reinforcing metabolic-transcriptional circuit centered around maintenance of NOX1-dependent cellular redox balance, regulates ISC function, self-renewal and subsequent cell fate decisions. Our studies show how redox-dependent stem cell metabolic state switching occurs in hypoxic tissue environments, and provide a basis for understanding regeneration dynamics, barrier function and disease propensity in the distal large intestine. Further studies are needed to understand how the dynamic control of stem cell function by hypoxia and redox influence disease-driven processes that disproportionately affect the distal colon such as ulcer formation, inflammation and cancer.
Methods
Animals
C57BL/6 (JAX Stock# 000664), B6.129×1-Nox1tm1Kkr/J (NOX1KO, JAX Stock# 018787) and B6.129P2-Lgr5tm1(cre/ERT2) Cle/J (Lgr5creERT2/+, JAX Stock# 008875) mice were purchased from The Jackson Laboratory. B6.Cg-Gt(ROSA)26Sortm14(CAG−tdTomato)Hze/J (Rosa26tdtomato, JAX Stock# 007914) strain and germ free mice were gifted from Snapper Scott Lab. GF mice were bred and maintained in vinyl isolators in the Animal Resources at Children’s Hospital (ARCH). Stool samples from GF were routinely collected and tested for sterility and exclusion of other bacterial contamination using aerobic and anaerobic conditions culture. Tg(Vil1-cre) 1000Gumstrain (JAX Stock# 021504) was gifted from Meenakshi Rao Lab. To explore the role of NOX1 in intestinal stem cells, we crossed NOX1KO mouse strain and Lgr5creERT2/+ to generate NOX1KO; Lgr5creERT2/+ mouse strain. To generate genetic lineage tracing strain, we bred Lgr5creERT2/+ and Rosa26tdtomato to generate Lgr5creERT2/+; Rosa26tdtomato/+mouse strain. To explore the role of NOX1 in cell fate decision, we bred NOX1KO mouse strain with Lgr5creERT2/+; Rosa26tdtomato/+ to generate NOX1KO; Lgr5 tdtomato mice.
To generate intracellular hydrogen peroxide reporter mice R26-RoGFP2-Orp1flox/flox using the CRISPR-Cas9 method, we used the cloning-free CRISPR/Cas system71. pR26roGFP2Orp1 targeting vector was generated by assembling Orp1-roGFP2 to the AsiSI/MluI restriction sites in Rosa26 targeting vectors pR26 CAG AsiSI/MluI72. 0.61 pmol each of crRNA and tracrRNA was conjugated and incubated with Cas9 protein (30 ng/μl) to prepare ribonucleoprotein complex (RNP) and donor DNA (10 ng/μl) was mixed with RNP for microinjection cocktail2. Microinjection cocktail was injected into 0.5 dpc embryos harvested after mating C57Bl6-Hsd (Envigo) females. Post-injection embryos were reimplanted into CD1 (Envigo) pseudo-pregnant foster females and allowed to term. Tail snip biopsies were collected from pups at P7 for genotyping. Primers for genotyping are listed in supporting information. To generate intestinal epithelial cell specific intracellular hydrogen peroxide reporter mice, we crossed R26-RoGFP2-Orp1flox/flox mice with Tg(Vil1-cre) 1000Gummice to get R26-RoGFP2-Orp1Villin mice.
Mice were bred and housed in a specific pathogen-free animal facility of Animal Resources at Children’s Hospital (ARCH). Age-matched 6- to 12-week-old littermate male and female mice were used for experiments. All animal experiments were approved by the Children’s Institutional Animal Care and Use Committee. Mice were maintained in a 12 hours light/dark cycle, with ad libitum access to food and water, 21.7 °C + / 1.7 °C, 35–70% +/− 5% humidity.
Human intestinal tissue samples
Healthy human sigmoid biopsy tissue was obtained from a 5-year-old male patient during diagnostic colonoscopy as part of routine clinical care under approved Boston Children’s Hospital Institutional Review Board Protocol #P00046566. Nox1 mutant p.N122H human colonic biopsy tissue was obtained from a 8-year-old male patient as a part of a study approved by the institutional review board of the Ludwig-Maximilians-Universität München, and samples were collected with informed consent from the parents/legal guardians of the tissue donors. Biopsy samples were fixed in 10% Neutral Buffered Formalin, embedded in paraffin, and cut into 5–7 μm sections for staining.
3D organoids primary cell culture and treatment
Colonic organoids were generated from isolated crypts collected from distal colon of mice listed above as previously described73. Murine colonic organoids were maintained at 37 °C in a humidified atmosphere containing 5% CO2. Approximately 500 crypts were plated in a 50 μl drop of Matrigel (Corning, 356231) in 24-well plates and overlaid with 500 μl of WENR medium, containing basal crypt media (Advanced DMEM/F12 (Gibco, 12634-028), 100 U/ml penicillin/streptomycin, 10 mM HEPES (Life Technology, 15630-080), 2 mM Glutamax (Life Technology, 35050-061)), supplemented with 1× B27 (Gibco, 17504-044), 1× N2 (Gibco,17502-048), 50 ng/ml rmEGF (Peprotech, 315-09), 50% L-WRN-CM (conditioned medium) (v/v), 10 μM Rock inhibitor Y-27632 (Sigma, Y0503), 250 nM CHIR99021 (Sigma, SML1046) with a final FBS concentration of 10%. Media was changed every 2 days and colonoids were split every 4 days. Cultures were passaged three times prior to experiments. For the measurement of spheroids surface area and the calculation of organoids formation number, we seeded 1000 cells/small cell clumps per well, 24 hrs after seeding, colonoids were treated with or without 1 mM N-Acetyl-L-cysteine (NAC, Sigma, D8199) or 10 μM diamide (Sigma, D3648) and then we took images with a BioTek Cytation5 automated microscope at ×2.5 object on day 3 or 5 after plating. The percentage of stem-like spheroids versus budded colonoids, were manually counted based on brightfield images (BioTek Cytation5 at ×2.5) after 3 days following plating in and treated with or without 200 U catalase (Sigma, C9322) or 1 U glucose oxidase/1 mM glucose (Sigma, G2133). To induce colonoids differentiation, 24 hrs after seeding, primary epithelial cells were exposed to ENR medium containing basal crypt media (Advanced DMEM/F12 (Gibco, 12634-028), 100 U/ml penicillin/streptomycin, 10 mM HEPES (Life Technology, 15630-080), 2 mM Glutamax (Life Technology, 35050-061)), supplemented with 1× B27 (Gibco, 17504-044), 1× N2 (Gibco,17502-048), 50 ng/ml rmEGF (Peprotech, 315-09), 100 ng/ml Noggin (Peprotech, 250-38), 10% R-spondin1-CM (conditioned medium) (v/v) with a final FBS concentration of 10%. For the manipulation the levels of α-Ketoglutarate, 24 hrs after seeding, primary epithelial cells were exposed to Dimethyl α-Ketoglutarate (3 mM, Sigma, 349631), or IDH1 inhibitor 2(compound 13, 2.5 mM, Medchemexpress, HY128661). For the stimulation with different niche factors, 24 hrs after seeding, primary epithelial cells were exposed to either ENR medium, or in WENR medium with or without butyrate (1 mM, Sigma, B5887), LPS (10 μg/ml), antioxidant DTT (2 mM, Sigma, D9779) or oxidant diamide (10 μM, Sigma, D3648) treatment or were cultured either under 21% oxygen (normoxia) or 1% oxygen (hypoxia) conditions in WENR or ENR medium. For linking redox, hypoxia and metabolism rewiring, 96 hrs after seeding, primary epithelial cells were cultured either under 21% oxygen (normoxia) or 1% oxygen (hypoxia) conditions in WENR, with or without Dimethyl α-Ketoglutarate (3 mM), IDH1 inhibitor 2 (compound 13, 2.5 mM), diamide (300 μM) or NAC (5 mM) for 4 hrs.
Human colonoids H514 were generated from a non-diseased pediatric colonic biopsy. Human colonoids were maintained at 37 °C in a humidified atmosphere containing 5% CO2. Colonoids were plated in a 50 μl drop of Matrigel and overlaid with 500 μl of WENR medium, containing basal crypt media (Advanced DMEM/F12 (Gibco, 12634-028), 100 U/ml penicillin/streptomycin, 10 mM HEPES (Life Technology, 15630-080), 2 mM Glutamax (Life Technology, 35050-061)), supplemented with 1× B27 (Gibco, 17504-044), 1× N2 (Gibco,17502-048), 50 ng/ml rhEGF (Peprotech, AF-100-15), 50% L-WRN-CM (conditioned medium) (v/v), 10 mM Nicotinamide (Sigma, N0636), 500 nM A83-01 (Sigma, SML0788), 500 nM SB202190 (Sigma, S7067), 500 μM N-Acetyl-Cysteine (Sigma, A8199), 50 nM Gastrin (Sigma, G9145), 100 nM Prostaglandin E2 (Sigma, P5640), 10 μM Rock inhibitor Y-27632 (Sigma, Y0503), 250 nM CHIR99021 (Sigma, SML1046) with a final FBS concentration of 10%.
RNA isolation and real-time QPCR analysis
Lyse and homogenize snap frozen tissues or colonoids in Matrigel in TRIzol™ Reagent. Then isolate RNA according to the manufacturer’s instructions. Equivalent amounts of purified RNA were converted to cDNA using iScript cDNA synthesis kit (Bio-rad, 1708891) and analyzed for gene expression on a CFX384 real time cycler (Bio-rad) using iTaq Universal SYBR Green Supermix (Bio-rad, 1725271) with Qpcr primers from Primer Bank for the following genes: Nox1, Nox4, Duox2, Ki67, Muc2, Slc26a3, Chga, Reg4, Glut1, Hk1, Ldhα, Pkm, Hif1α, which are listed in the supporting information (Supplementary Table 1). Data were normalized to Rpl13a, and results were shown as fold induction relative to expression levels in WT groups or control groups.
RNAScope
For mouse Nox1 and Lgr5 RNAscope staining, proximal colons and distal colons were collected, embedded in Tissue-Tek OCT compound and snap-frozen in dry ice before freezing at −80 °C. Embedded tissues were warmed to −20 °C, cryosectioned into 14 μm slices and then were fixed with 4% PFA at 4 °C for 15 min and stained for Nox1 and Lgr5 using the RNAscope Multiplex Fluorescent Assay v2 (Advanced Cell Diagnostics, 323100) following the manufacturer’s instructions. Entire sections were imaged using a confocal microscopy at ×20 and images were analyzed in FIJI.
Tissues preparation and images acquisition from intracellular hydrogen peroxide reporter mice (Colon tissues/colonoids)
For freshly fixed frozen samples, proximal colons and distal colons were embedded in Tissue-Tek OCT compound and snap-frozen in dry ice before storage at −80 °C. Embedded tissues were warmed to −20 °C, cryosectioned into 14 μm slices with a cryotome (Leica CM3050) and mounted onto Superfrost Plus slides (Fisher Scientific GmbH). For sections further processed with chemicals, the chemicals were applied immediately after mounting. To induce maximal probe oxidation or reduction, sections were incubated for 10 min on ice with 1 mM diamide (Sigma, D3648) or 20 mM DTT (Sigma, D9779), respectively, before N-ethylmaleimide (Sigma, E3876) treatment. To preserve the endogenous redox state, sections were immediately incubated with 50 mM NEM (Sigma-Aldrich, E3876) for 10 min on ice followed by fixation in 4% PFA in PBS containing 1 mM TO-PRO-3 (using 0.1% dimethyl sulfoxide as a co-solvent) (Invitrogen, T3605) for 15 min at ambient temperature. PFA was washed out twice with ice-cold PBS for 5 min. Sections were then mounted in ProLong Dimond Antifade Mountant (Thermo Fisher Scientific, P36961) and kept at 4 °C until use21.
Guidelines for the use of roGFP2-based redox probes, including microscopy settings and image analysis, have been provided previously21,74,75. Adapted from that, images were acquired using Zeiss LSM 880 using 40 × 1.3 NA objective. For each animal, at least three images were taken from one or more sections. Three channels were imaged (2048 × 2048 pixels at 103 nm resolution). Nuclei stained with TO-PRO-3 was imaged in the far-red channel with the 633 nm excitation laser. For measuring the redox changes, two excitation lasers (405 nm and 488 nm) were used to sequentially image an emission channel (500–540 nm). Images were analyzed using custom written MATLAB code to generate the redox ratio metric images (I405/I488). For the fasted/refed experiment, each channel was first eroded using a structural element of 5 pixels and an intensity threshold of 7 was used to denoise the image and the ratio of the channels was calculated (Supplementary Note 1). For the validation of probe dynamic range experiment, the two channels (405 and 488) are binarized using local adaptive thresholding and morphologically closed with a structuring element with a radius size of 1 to generate the individual channel masks (Supplementary Note 2). The ratio of the two channels is calculated. In both experiments, any ratio greater than 20 was set to 0 to prevent low signal regions. For each image, the median of the non-zero ratio values is calculated. The images were then manually analyzed in ImageJ to specifically measure the oxidation ratio of the crypt base. For live imaging, harvested tissues or colonoids in Matrigel were mounted in a live imaging chamber in Krebs-Hepes buffer (pH7.2) and imaged on a Zeiss LSM 880 microscope using a 20 × 0.8NA objective.
Colon tissue Immunofluorescence microscopy
For FFPE samples, tissues were fixed with 4% paraformaldehyde diluted in PBS at 4 °C overnight. Tissues were processed, paraffin-embedded, and cut into 5 μm sections at BIDMC histology core. 5 μm tissue cross-sections were heated at 60 °C for 10 min prior to deparaffination in xylene for 10 min twice and then rehydrated with decreasing concentrations of ethanol (100, 95, 70, 50 and 0%) for 10 min each at room temperature. Antigen retrieval was performed by pressure cooker in antigen retrieval solution (10 mM citric buffer, pH 6.0) for 15 min on high. Samples were washed in PBS and blocked with 10% donkey serum in PBS for 1 h at 4 °C in a humidified chamber. Then samples were incubated with the indicated primary antibodies, rabbit polyclonal anti-DUOX2 antibody (1:200, Novus Biologicals, NB110-61576), mouse monoclonal anti-E-cadherin antibody (1:200, Bdbiosciences, 610182), mouse monoclonal anti-Chr-A antibody (1:100, Santa Cruz Biotechnology, sc-393941), rabbit polyclonal anti-REG4 antibody (1:200, Sinobiological, 80249-T24), rabbit polyclonal anti-MUC2 antibody (Santa Cruz Biotechnology, no longer available), rat monoclonal anti-human/mouse Ki67 antibody (1:200, Biolegend, 151202), mouse AF488 anti-PCNA (PC10) antibody (1:100, CST, 8580S), rabbit AF555 anti-human Epcam antibody (1:100, Abcam, ab275122), diluted in blocking buffer at 4 °C overnight. Sections were washed twice and then stained with fluorescent secondary antibodies, donkey polyclonal AF488 anti-mouse IgG antibody (1:400, Jackson ImmunoResearch Laboratories, 715-545-150), donkey polyclonal AF568 anti-rat IgG antibody (1:400, Jackson ImmunoResearch Laboratories, 712-575-150), donkey polyclonal AF647 anti-Rabbit IgG antibody (1:400, Jackson ImmunoResearch Laboratories, 711-605-152), donkey polyclonal AF568 anti-Rabbit IgG antibody (1:400, Jackson ImmunoResearch Laboratories, 711-575-152), diluted in blocking buffer for 2 hrs at room temperature. Finally, the sections were washed three times and incubated with PBS containing Hoechst (Invitrogen, H1399) at a 1:1,000 dilution and then mounted with ProLong Dimond Antifade Mountant (Thermo Fisher Scientific, P36961).
For cryosection samples, colons were fixed with 4% paraformaldehyde diluted in PBS for 1–2 hours at 4 °C, washed with PBS three times, incubated with 30% sucrose at 4 °C overnight, and then embedded in OCT (Tissue Tek), snap-frozen in dry ice before storage at −80 °C. Embedded tissues were warmed to −20 °C, cryosectioned into 14 μm slices with a cryotome (Leica CM3050) and mounted onto Superfrost Plus slides (Fisher Scientific GmbH). Samples were then blocked with 10% donkey serum in PBS for 1 h at 4 °C in a humidified chamber and were then incubated with the indicated primary antibodies, chicken polyclonal anti-GFP antibody (1:100, Novus Biologicals, NB100-1614), goat anti-mCherry polyclonal antibody (1:100, OriGene Technologies, AB0040-200), diluted in blocking buffer at 4 °C overnight. Sections were washed twice and then stained with fluorescent secondary antibodies, donkey polyclonal AF488 anti-Chicken IgG antibody (1:400, Jackson ImmunoResearch Laboratories, 703-545-155), donkey polyclonal AF568 anti-Goat IgG antibody (1:400, Jackson ImmunoResearch Laboratories, 705-575-147), diluted in blocking buffer for 2 hrs at room temperature. Finally, the sections were washed three times and incubated with PBS containing Hoechst (Invitrogen, H1399) at a 1:1000 dilution and then mounted with ProLong Dimond Antifade Mountant (Thermo Fisher Scientific, P36961). For each animal, at least two-three images were taken from one or more sections with a ×20 objective on a Zeiss LSM 880 inverted confocal microscope, some images were under automated tile scanning with 10% overlap between individual images.
Organoids tissue Immunofluorescence microscopy
After recovered from Matrigel, colonoids were incubated in 4% paraformaldehyde diluted in PBS with gentle rocking for 45 min at 4 °C. Let the colonoids settle by gravity and wash the colonoids once in immunofluorescence (IF) buffer, PBS containing 0.1% w/v BSA, 0.2% v/v TritonX-100, (0.05% v/v) TWEEN® 20. We then resuspended the colonoids in 1 ml of PBS and transfer them to 1.5 ml Eppendorf tubes. Aspirate PBS and add 1 ml of citrate buffer (pH 6.0) to the colonoids and incubate the tubes in the heating block of 98 °C for 20 minutes. Then aspirate the citrate buffer and add 1 ml of permeabilization/blocking solution, PBS containing 5% v/v serum, 1% v/v TritonX-100, and incubate at room temperature with agitation for 2 hours. Aspirate the permeabilization/blocking solution and wash 3 times in IF Buffer. Add 0.5 ml of the primary antibodies, mouse monoclonal anti-E-cadherin antibody (1:200, Bdbiosciences, 610182), mouse monoclonal anti-Chr-A antibody (1:100, Santa Cruz Biotechnology, sc-393941), rabbit polyclonal anti-REG4 antibody (1:200, Sinobiological, 80249-T24), rabbit polyclonal anti-MUC2 antibody (Santa Cruz Biotechnology, no longer available), rat monoclonal anti-human/mouse Ki67 antibody (1:200, Biolegend, 151202), rabbit polyclonal anti-NHE3 antibody (1:200, Novus Biologicals, NBP1-82574), diluted in IF Buffer and incubate at room temperature overnight with gentle agitation. Aspirate the primary antibody solution and wash 3 times in IF Buffer. Add 0.5–1 ml of the secondary antibodies diluted in IF Buffer supplemented with 10% donkey serum and incubate at room temperature for 2 hours with gentle agitation. Add Hoechst (Invitrogen, H1399) directly to the secondary solution/colonoids to a final concentration of 2–4 μg/ml and incubate at room temperature with gentle agitation for 15–20 min in the dark. Aspirate the staining solution and wash the colonoids once in water and once in PBS and then resuspend the organoid pellet in 50 μl ProLong™ Dimond Antifade Mountant (Thermo Fisher Scientific, P36961). Dispense the organoid/ mountant suspension onto the center of a glass microscope slide within image spacers and cover with a coverslip. Images were taken with a ×20 objective on a Zeiss LSM 880 inverted confocal microscope. This was adjusted from the protocol of Stem Cell Technologies.
Flow cytometry
To determine the intracellular hydrogen peroxide levels in colonoids, we digested colonoids with TripLE (Gibco, 12605010) to get single-cell suspensions. After stained with live/dead dye, cells were immediately treated with 50 mM NEM to clamp the intracellular redox status. roGFP 405/488 ratio analysis by FACS was performed in a Fortessa (BD Biosciences) equipped with 405 nm and 488 nm laser lines. The cells were excited with 405 nm, and 488 nm lasers and fluorescence emission were collected at 525/50 nm or 530/30 nm at each laser line in ratio mode. The oxidation degree of roGFP was calculated adapted from Avia et al and Schwarzländer et al.76,77. Flow data was analyzed using custom written R code to calculate median normalized oxidation ratio. The 405 to 488 ratio was calculated for each cell and the outliers removed using interquartile range as a measure. All the ratios were normalized to their respective median values and scaled using the median value of the DTT treatment (normalized as 0%) and the median value of the DA treatment (normalized as 100%).
To explore the role of NOX1 in ISCs regeneration, we did BrdU incorporation in both mice and in vitro colonoids. For mice experiments, we injected mice IP with 100 μl (1 mg) of BrdU solution (BD Biosciences, 552598). 2 hours later, we sacrificed the mice and isolated colonic crypts with 5 mM EDTA, then digested them with dispase II (Sigma Aldrich, D4693) and DNaseI (Millipore Sigma, D4263) to get single cell suspensions. For organoids culture, we incubated cells with BrdU at a final concentration of 10 μM in cell culture medium for 1 h. After recovered from Matrigel, colonoids were digested with TripLE into single cell suspensions. LIVE/DEAD™ Fixable Near-IR Dead Cell Stain Kit (1:500, Invitrogen™, L10119), Rat Pacific Blue™ anti-mouse CD45 antibody (1:500, Biolegend, 103125), Rat BV605™ anti-mouse EpCam antibody (1:500, Biolegend, 118227), anti-BrdU APC antibody (1:50, BD Biosciences, 552598) was used. Then cells were stained following the manufacturer’s instructions. For cell cycle analysis, cells were stained with 7AAD that binds to total DNA after anti-BrdU staining. Data were acquired on a BD LSRFortessa or a BD Symphony A5 SE spectral flow cytometer and analyzed in Flowjo v10.
For genetic lineage tracing in vivo, mice were intraperitoneally injected with tamoxifen 20 mg/kg and sacrificed 48–72 hrs after induction. As to HIF1α inhibition, mice were injected with or without LW6 20 mg/kg daily 3 days prior to tamoxifen injection. We then isolated colonic crypts with 5 mM EDTA, digested them with dispase II and DNaseI to get single cell suspension. Then dissociated cells were stained with LIVE/DEAD™ Fixable Near-IR Dead Cell Stain Kit (1:500, Invitrogen™, L10119), Rat Pacific Blue™ anti-mouse CD45 antibody (1:500, Biolegend, 103125), Rat BV605™ anti-mouse EpCam antibody (1:500, Biolegend, 118227) and analyzed by BD LSRFortessa analyzer.
For intestinal stem cell isolation, colonic crypts were isolated with 5 mM EDTA from proximal and distal colons of WT; Lgr5creERT2+ and NOX1KO; Lgr5creERT2+ mice. Then crypt pellets were digested with dispase II and DNase I to get single cell suspension. Then dissociated cells were stained with LIVE/DEAD™ Fixable Lime (506) Viability Kit (1:500, Invitrogen™, L34989), Rat Pacific Blue™ anti-mouse CD45 antibody (1:500, Biolegend, 103125), Rat PercP-Cy5.5 anti-mouse EpCam antibody (1:200, Biolegend, 118219) and sorted with BD Melody sorter. Cells were sorted depending on EGFP intensity. Primary Organoids formation analysis was performed with sorted intestinal stem cells.
For intestinal epithelial cells (IECs) isolation for single cell sequencing, we isolated colonic crypts from distal colons and proximal colons of WT and NOX1KO littermates with 5 mM EDTA, digested them with dispase II and DNaseI to get single cell suspension. Then the dissociated cells were stained with LIVE/DEAD™ Fixable Lime (506) Viability Kit (1:500, Invitrogen™, L34989), Rat Pacific Blue™ anti-mouse CD45 antibody (1:500, Biolegend, 103125), Rat PercP-Cy5.5 anti-mouse EpCam antibody (1:200, Biolegend, 118219), and IECs were sorted with BD Melody sorter.
Fasting-refeeding models
During the fasting period, 8–10-week-old mice were placed on a stainless mesh floor with no food to avoid coprophagia, and with ad libitum drinking water. Mice were fasted for 36 hrs and then refed with normal chow for 24 hrs. Mice were monitored every 12 hrs for body weight change23. Colons were harvested for flow cytometry analysis or tissue H&E staining, other immunohistochemical staining.
10X scRNA-seq of mouse colon
Colonic epithelial cells isolated from WT and NOX1KO littermates by BD Melody sorter from proximal colon or distal colon regions were pooled separately across 2 mice at equal ratios and were resuspended in 1% BSA in PBS and counted with trypan blue staining for cell viability and number. Approximately 30,000 cells per sample were loaded onto separate lanes of the Chromium X Controller. Gene expression libraries were prepared according to manufacturer instructions using the 10X Genomics Single Cell 3’ Reagent Kit (v3.1 Chemistry Dual Index). Sequencing was performed on a NovaSeq 6000 through the Broad Clinical Labs using a NovaSeq S2 100 cycle kit to target 20,000 read pairs per cell.
10X scRNA-seq data processing and downstream analysis
Sequencing files were processed using CellRanger (v7.2.0). Analysis was conducted using Seurat (v5.1.0)78 keeping cells containing at least 100 unique genes and features expressed in at least 3 cells. A coarse quality filter was applied to remove cells with greater than 8000 unique genes and 50% of reads mapping to mitochondrial genes. The expression and metadata were converted into Seurat objects, one for proximal and other for distal colon. Gene counts were scaled and normalized using log normalization and the 3000 most variable features selected after variance stabilization were then used in dimensionality reduction by PCA. 30 PCs were used for UMAP dimensional reduction and a granularity of 0.2 and 0.5. The top 10 markers for each cluster were determined and the clusters manually annotated.
Trajectory analysis was performed using Monocle 3.0 with a cluster resolution of 5e-4 and the cluster corresponding to the Stem Cell population was used as the root cell. The low-quality, enteroendocrine and tuft cell clusters were removed from this analysis. 3.5 was used as a cutoff to delineate high and low Transit Amplifying cells. A manually curated list of cell cycle genes was used to generate relative expression under different conditions for the TA cell79. Data can be accessed at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE289397.
Seahorse cellular metabolic analysis
One day prior to assay, transform 3D organoids to 2D monolayer. Colonoids were extracted from Matrigel with an organoid harvesting solution (R&D system, 3700-100-01), and then digested with undiluted TripLE (Gibco, 1265010). Colonoids fragments were suspended in WENR medium and plated on collagen-coated (Sigma, C5533) Seahorse 96-well microplate plate. Meanwhile, hydrate cartridge and warm Seahorse XF Calibrant (Agilent, 100840-000). After 24 hours, cells were washed twice and incubated in the Seahorse Assay Medium (Agilent, 102353-100) supplemented with 12 mM glucose and 2 mM glutamine at 37 °C for 45 min. Oxygen consumption rate (OCR) and extracellular acidification rate (ECAR) were measured with a Seahorse XFe96 Extracellular Flux Analyzer. The OCR and ECAR were measured under basal conditions and after injection of Oligomycin (2 μM, Sigma, 75351), FCCP (2 μM, Sigma, C2920) plus pyruvate (5 mM, Sigma, S8636), rotenone (1 μM, Sigma, R8875) plus antimycin A (1 μM, Sigma, A8674) (Rot.+AA), and 2-DG (50 mM, Sigma, D8375). Metabolic parameters were calculated as follows: Basal OCR = OCRbefore OM–OCRafter Rot+AA, ATP-linked respiration = Basal OCR–OCRafter OM, Maximal Respiratory Capacity (MRC) =OCRafter FCCP+Pyruvate–OCRafter Rot+AA, Basal ECAR = ECARbefore OM–ECARafter 2-DG, Maximal Glycolytic Capacity (MGC) = ECARafterOM - ECARafter 2-DG.
Mitochondrial membrane potential measurement by flow cytometry
Colonoids were treated with TMRM (20 nM, Invitrogen™, M20036) and MitoTracker Green (20 nM, Invitrogen™, M7514) for 30 min at 37 °C. After recovered from Matrigel, colonoids were digested with TripLE into single cell suspensions. Then dissociated cells were stained with live/dead dye and then analyzed by BD LSRFortessa flow cytometer with 561-nm excitation.
Bulk RNA-seq of colonoids
Lyse colonoids recovered from Matrigel in TRIzol™ Reagent. Then isolate RNA according to the manufacturer’s instructions. The quality of RNA samples was determined using an Agilent 5400 Bioanalyzer, and all samples for sequencing had RNA integrity (RIN) numbers >8. cDNA library construction using the NEBNext® Ultra™ II RNA Library Prep Kit for Illumina® was performed by the Novogene Corporation Inc. (Sacramento, CA), and cDNA libraries were sequenced on an Illumina HiSeq 2000 or Illumina Novaseq 6000 instruments.
Bulk RNA-seq data analysis
Quality control for mRNA-seq data (FASTQ files) was conducted using FastQC (v0.11.9) followed by alignment to the mouse genome (GRCm38.p6) using the STAR aligner (v2.7.8a) in 2-pass mode with default parameters and version 25 of the mouse GENCODE primary assembly annotation (GENCODEvM25). Aligned BAM files were further analyzed using RSeQC (v4.0.0). The alignments and quality control steps were run using the Snakemake workflow management system (v6.6.1). Reads belonging to individual genes annotated by GENCODEvM25 were counted using htseq-count (v0.13.5). Differential expression analysis was run using the default workflow provided by the DESeq2 R package (v1.34.0). Volcano plot was charted using the Enhanced Volcano program80. A set of 128 downregulated genes that were on average at least 50% higher in control relative to KO was used to determine pathways altered as a consequence of knockdown. Data can be accessed at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE289209.
Differential cysteine alkylation sample preparation and LC-MS/MS analysis
Differential cysteine alkylation sample preparations were adapted from Bechtel, T. et al.81 and Topf, U. et al.82. 100 μg of cell lysates of colonoids recovered from Matrigel was precipitated in 100% TCA/DPBS at −80 °C for 1 hr or overnight. Samples were pelleted at 14,000 × rpm for 10 min at 4 °C and resuspended in 500 µL of ice-cold acetone by vortexing. Proteins were pelleted at 5000 × rpm for 10 min at 4 °C. Acetone was removed, and the pellet was left air dry to remove traces of acetone. The pellet was resuspended in 100 μL of DAB buffer (6 M urea, 20 mM Tris-HCl, 1 mM EDTA, 0.5% SDS, pH 8.5) and incubated with 10 mM N-ethylmaleimide (NEM) for 2 hrs at 37 °C with agitation. Light NEM is removed by diluting the reaction 3-fold with H2O and then 5-fold with ice-cold acetone. Proteins were precipitated at −20 °C for 2 hrs or overnight and pelleted at 4500 × g for 30 min at 4 °C. The protein pellet was washed again with 500 µL of ice-cold acetone and centrifuged at 4500 × g for 10 min. Proteins were re-solubilized using 80 μL DAB buffer and incubated with 2.5 mM TCEP for 5 min at 37 °C. The reaction was further diluted with 120 μL DAB buffer and incubated with 10 mM D5-NEM (Cambridge Isotope Laboratories) for 2 hrs at 37 °C. After the final alkylation, samples were diluted 3-fold with H2O and then 5-fold with ice-cold acetone for protein precipitation at −20 °C for 2 hrs or overnight. Proteins were pelleted at 4500 × g for 30 min at 4 °C and washed with 500 μL ice-cold acetone. Protein pellets were re-suspended in 200 μL 2 M urea with 1 mM CaCl2 and 2 mg trypsin for overnight digestion at 37 °C. 10 μL formic acid was added to the digested peptides. Samples were de-salted using Sep-Pak C18 columns and eluted with 1.5 mL of 80% LC/MS-grade acetonitrile, 20% H2O, and 0.1% formic acid. The de-salted samples were dried on SpeedVac for further storage at −20 °C.
Tryptic peptide samples were resuspended in 500 μL of high-pH buffer A (95% water, 5% acetonitrile, and 10 mM ammonium bicarbonate). The resuspended peptides were loaded onto a manual injection loop connected to an Agilent 1100 Series HPLC to be further separated on a 25-cm Agilent Extend C18 column using a 60-min gradient ranging from 20 to 35% high-pH buffer B (10% water, 90% acetonitrile, and 10 mM ammonium bicarbonate) in high-pH buffer A. Fractions were collected into a 96-deep-well plate using a Gilson FC203B fraction collector, and samples in every six wells were combined resulting in six pooled fractions to be dried on SpeedVac.
Trypsin digested peptide samples were resuspended in 100 μL buffer A (100% water, 0.1% formic acid), peptide concentration was determined using PierceTM quantitative colorimetric peptide assay. Peptide analysis was performed using an Orbitrap Exploris 240 mass spectrometer coupled to a Dionex Ultimate 3000 RSLCnano system. Peptides were eluted onto an Acclaim PepMap RSLC column (150 mm length, 0.1 mm diameter, 5 μm particle size) and separated with a 160-min gradient ranging of 5–25% MA buffer B (20% water, 80% acetonitrile, and 0.1% formic acid) in MS buffer A at a flow rate of 0.3 mL/min. The spray voltage was set to 2.1 kV. One full MS scan (350-1,800 m/z, 120,000 resolution, RF lens 65%, automatic gain control (AGC) target 300%, automatic maximum injection mtime, profile mode) was obtained every 2 s with dynamic exclusion (repeat count 2, duration 10 s), isotopic exclusion (assigned), and apex detection (30% desired apex window) enabled. A variable number of MS2 scans (15,000 resolution, AGC 75%, maximum injection time 100 ms, centroid mode was obtained between each MS1 scan based on the highest precursor masses, filtered for monoisotopic peak determination, theoretical precursor isotopic envelope fit, intensity (5E4), and charge state (2-6). MS2 analysis consisted of the isolation of precursor ions (isolation window 2 m/z) followed by higher energy collision dissociation (HCD) (collision energy 30%).
Analysis of MS data collected on Orbitrap Exploris 240 was performed using Thermo Proteome Discoverer software. Protein identification was achieved using the SequestHT and Percolator algorithms83 against Mus Musculus proteome UniprotKB database84. The protease enzyme was set as trypsin with a maximum of 2 missed cleavages allowed. The peptide precursor mass tolerance was set to 10 ppm with a fragment mass tolerance of 0.02 Da. Dynamic modifications included methionine oxidation (+15.995), N-terminal acetylation (+42.011), methionine loss (−131.040), and cysteine alkylation by either light-NEM (+125.048) or heavy-NEM (+130.079). The FDR for highly confident peptide identification was set at 1%. The resulting H:L ratios were converted to percent oxidation values by the following equation: \(\frac{H:L}{H:L+1}\). Average percent oxidation values were calculated for each peptide and peptides with standard deviation >50% were removed from the analyses.
Analysis of differential cysteine alkylation redox proteomics data
To identify cysteines that were significantly altered in the oxidative proteomics screen, we calculated two metrics from the average percent oxidation in both WT and NOX1KO colonoids from one biological and two technical replicates. Change in relative ranks and a change in deviation of oxidation was calculated. For the first metric, ranks of all the oxidized cysteines were calculated for both WT and NOX1KO and the absolute value of the difference between the two was determined. A change of rank of at least 40 was used to determine the hits. For the second metric, the measure of the deviation from a linear fit with slope 1 and intercept (0,0) was determined. The magnitude of the deviation gives the magnitude of the effect while the direction determines whether it is less or more oxidized in the NOX1KO. These two parameters were used for the Volcano plot.
Protein networks that were significantly altered in the different analyses were determined using String-DB85. For determination of up and downregulated canonical pathways, Ingenuity Platform Analysis was used. A cutoff of -log(P-value) >=2 and a Z-score of +/−2 was used for identifying the pathways. A positive z-score implies activation while negative z-score implies inhibition of the pathway. Manually curated subsets of pathways and network nodes are shown in the Supplementary Fig 6b and 6c.
IDH1 enzyme activity assay
After recovered from Matrigel, colonoids were washed once with ice-cold PBS and homogenized in 200 μl per well of ice-cold IDH Assay Buffer. Then we centrifuged the lysates at 13,000 × g for 15 min at 4 °C to get supernatant. The concentrations of supernatant were uniformed to 500 μg/ml by BCA analysis (Thermo Scientific™, 23227). Then we analyzed IDH enzyme activity following the manufacturer’s instructions (Sigma, MAK062).
αKG levels measurement
After recovered from Matrigel, colonoids were washed once with ice-cold PBS and resuspended in 100 μl per well of ice-cold Assay Buffer. Colon tissues were homogenized in 200 μl of ice-cold Assay Buffer. After incubating on ice for 20 min, we centrifuged the lysates at 13,000 × g for 15 min at 4 °C to get supernatant. The concentrations of supernatant were uniformed to 1000–2000 μg/ml by BCA analysis (Thermo Scientific™, 23227). Then we analyzed αKG levels following the manufacturer’s instructions (Cayman Chemical, 701350).
GSH/GSSG ratio detection assay
After recovered from Matrigel, colonoids were washed once with ice-cold PBS and homogenized in 100 μl per well of ice-cold PBS/0.5% NP-40. Then we centrifuged the lysates at 13,000 × g for 15 min at 4 °C to get supernatant. The concentrations of supernatant were uniformed to 500 μg/ml by BCA analysis (Thermo Scientific™, 23227). After deproteinization (Abcam, ab204708), we then ran GSH and Total Glutathione assay following the manufacturer’s instructions (Abcam, ab205811).
Gel electrophoresis and immunoblot analysis
After recovered from Matrigel, colonoids were washed once with cold 1× PBS and were lysed in chilled 1× RIPA buffer (10× stock, EMD Millipore) diluted in PBS containing 1 tablet of cOmplete EDTA free protease inhibitor (Millipore Sigma, 11836145001) for 30 minutes on ice. Protein was quantified using a Pierce BCA protein quantification kit. 20 μg of total protein lysates was loaded and separated on Novex™ Tris-Glycine Mini Protein Gels, (4–20%, 1.0 mm, WedgeWell™ format, Thermo Scientific). Proteins were transferred to nitrocellulose membranes, blocked with EveryBlot blocking buffer (Bio-Rad), and incubated with the indicated primary antibodies diluted in blocking buffer at 4 °C overnight. Membranes were washed three times in TBS-T and incubated with HRP-coupled secondary antibodies for 1 h at room temperature. After washed for three times, proteins on the membranes were detected by chemiluminescence using ECL (Thermo Scientific) in a Bio-Rad ChemiDoc Imager. The following primary antibodies and dilutions were used: anti-HIF1α (Novus Biological, NB100-134, 1:1000), anti-HIF1α (CST, 36169S, 1:1000) and anti-β-actin (A5441, Sigma, 1:1000). Western blot images were processed using FIJI and Affinity Designer.
H&E analysis, AB/PAS analysis and DAB analysis of HIF1α
Mice were euthanized and colon tissues were collected and fixed with 4% paraformaldehyde diluted in PBS at 4 °C overnight. Tissues were processed, paraffin-embedded, and cut into 5 μm sections and under H&E (hematoxylin and eosin), AB/PAS (Alcian blue/Periodic acid-Schiff) staining at BIDMC histology core. Colon FFPE samples were deparaffinized, rehydrated and performed with antigen retrieval and we then followed the manufacturer’s instructions for the DAB analysis with VECTASTAIN® Elite® ABC-HRP Kit (Vector Laboratories, PK-6101) and ImmPACT® DAB Substrate Kit (Vector Laboratories, SK-4105). The diluted primary antibody was used, anti-HIF1α (1:1000, Novus Biological, NB100-134). Whole cross-sections were tile-scanned and imaged on a widefield BioTek Cytation5 automated microscope at ×20, and images were analyzed with FIJI.
ChIP-qPCR
ChIP-qPCR was performed as described previously with some modifications from Shen et al.86. Briefly, 2D monolayer cells of human colonoids H514 were crosslinked with 1% formaldehyde for 10 min at room temperature and quenched by 2.5 M glycine for 5 min. Cells were lysed with a nuclei lysis buffer and lysates were sonicated to an average chromatin fragment of 300–400 bp. Take 10% out as input. The following antibodies were used for each ChIP experiment based on instructions: 1:50 dilution anti-HIF1α (Novus Biologicals, NB100-134), 1:50 dilution anti-TCF4 (CST, 2569 T). 40 μl of Protein G Dynabeads (Invitrogen, 10003D) were used for each ChIP experiment. The purified elution was used for following qPCR with primers listed in Key sources table.
Quantification and data analysis
For all quantifications, the exact value of n is described in detail in the figure legends, where n indicates biological replicates, either number of mice, number of wells containing cells or number of areas containing cells. For all microscopy analysis, images were blinded prior to scoring and quantification. All statistical analyses were performed using Microsoft Excel and GraphPad Prism software. Data were represented as mean ± SEM throughout the figures unless indicated, and the level of significance was indicated by asterisks for the following corresponding p-values: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, ∗∗∗∗p < 0.0001. The specific statistical test used to for each experiment is described in detail in the figure legends. For comparisons between two groups, we performed student’s two-sided t tests. For comparisons that have more than two groups or two conditions, one-way ANOVA and Two-way ANOVA were utilized. Sample sizes for all experiments were chosen according to standard practice in the field.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The bulk RNA sequencing data have been deposited in the GEO database under accession code Series GSE289209. The scRNA sequencing data have been deposited in the GEO database under the accession code Series GSE289397. The raw proteomic data have been deposited in the PRIDE database (accession number PXD065916). The cysteine proteomics data and the counts of bulk transcriptomics data are available under the Supplementary. Source data are provided with this paper.
Code availability
Custom Matlab codes written for image analysis are provided as Supplementary Notes 1 and 2 in the Supplementary Information file.
References
Barker, N. Adult intestinal stem cells: critical drivers of epithelial homeostasis and regeneration. Nat. Rev. Mol. Cell Biol. 15, 19–33 (2014).
Brunet, A., Goodell, M. A. & Rando, T. A. Ageing and rejuvenation of tissue stem cells and their niches. Nat. Rev. Mol. Cell Biol. 24, 45–62 (2023).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Jackson, B. T. & Finley, L. W. S. Metabolic regulation of the hallmarks of stem cell biology. Cell Stem Cell 31, 161–180 (2024).
Van der Flier, L. G. & Clevers, H. Stem cells, self-renewal, and differentiation in the intestinal epithelium. Annu. Rev. Physiol. 71, 241–260 (2009).
Cheng, C.-W. et al. Ketone body signaling mediates intestinal stem cell homeostasis and adaptation to diet. Cell 178, 1115–1131.e15 (2019).
Rodríguez-Colman, M. J. et al. Interplay between metabolic identities in the intestinal crypt supports stem cell function. Nature 543, 424–427 (2017).
Berger, E. et al. Mitochondrial function controls intestinal epithelial stemness and proliferation. Nat. Commun. 7, 13171 (2016).
Chen, L. et al. HNF4 Regulates Fatty Acid Oxidation And Is Required For Renewal Of Intestinal Stem Cells In Mice. Gastroenterology 158, 985–999.e9 (2020).
Gao, Y. et al. LKB1 represses ATOH1 via PDK4 and energy metabolism and regulates intestinal stem cell fate. Gastroenterology 158, 1389–1401.e10 (2020).
Jensen, B. A. H. et al. Small intestine vs. colon ecology and physiology: Why it matters in probiotic administration. Cell Rep. Med. 4, 101190 (2023).
Araki, K. et al. Comparison of mucosal microvasculature between the proximal and distal human colon. J. Electron. Microsc. 45, 202–206 (1996).
Li, C. et al. Glycolytic regulation of intestinal stem cell self-renewal and differentiation. Cell Mol. Gastroenterol. Hepatol. 15, 931–947 (2023).
Wei, P., Dove, K. K., Bensard, C., Schell, J. C. & Rutter, J. The force is strong with this one: metabolism (over)powers stem cell fate. Trends Cell Biol. 28, 551–559 (2018).
Ito, K. & Suda, T. Metabolic requirements for the maintenance of self-renewing stem cells. Nat. Rev. Mol. Cell Biol. 15, 243–256 (2014).
Tan, D. Q. & Suda, T. Reactive oxygen species and mitochondrial homeostasis as regulators of stem cell fate and function. Antioxid. Redox Signal 29, 149–168 (2018).
Aviello, G. & Knaus, U. G. NADPH oxidases and ROS signaling in the gastrointestinal tract. Mucosal Immunol. 11, 1011–1023 (2018).
ROS in gastrointestinal inflammation: rescue or sabotage? Br. J. Pharmacol. 174, 1704–1718 (2017).
Grasberger, H. et al. Increased expression of DUOX2 is an epithelial response to mucosal dysbiosis required for immune homeostasis in mouse intestine. Gastroenterology 149, 1849–1859 (2015).
van der Post, S., Birchenough, G. M. H. & Held, J. M. NOX1-dependent redox signaling potentiates colonic stem cell proliferation to adapt to the intestinal microbiota by linking EGFR and TLR activation. Cell Rep. 35, 108949 (2021).
Yuuta F. Mouse redox histology using genetically encoded probes. Sci. Signal. 9, 419 (2016).
Coant, N. et al. NADPH oxidase 1 modulates WNT and NOTCH1 signaling to control the fate of proliferative progenitor cells in the colon. Mol. Cell. Biol. 30, 2636–2650 (2010).
Okada, T. et al. Microbiota-derived lactate accelerates colon epithelial cell turnover in starvation-refed mice. Nat. Commun. 4, 1654 (2013).
Nagai, M. et al. Fasting-refeeding impacts immune cell dynamics and mucosal immune responses. Cell 178, 1072–1087.e14 (2019).
Clevers, H. Modeling development and disease with organoids. Cell 165, 1586–1597 (2016).
Sei, Y., Feng, J., Chow, C. C. & Wank, S. A. Asymmetric cell division-dominant neutral drift model for normal intestinal stem cell homeostasis. Am. J. Physiol.-Gastrointest. Liver Physiol. 316, G64–G74 (2019).
Viatour, P. Bridges between cell cycle regulation and self-renewal maintenance. Genes Cancer 3, 670–677 (2012).
Chen, H.-Z., Tsai, S.-Y. & Leone, G. Emerging roles of E2Fs in cancer: an exit from cell cycle control. Nat. Rev. Cancer 9, 785–797 (2009).
Kisty, E. A., Falco, J. A. & Weerapana, E. Redox proteomics combined with proximity labeling enables monitoring of localized cysteine oxidation in cells. Cell Chem. Biol. 30, 321–336.e6 (2023).
Lennicke, C. & Cochemé, H. M. Redox metabolism: ROS as specific molecular regulators of cell signaling and function. Mol. Cell 81, 3691–3707 (2021).
Stojanović, O., Miguel-Aliaga, I. & Trajkovski, M. Intestinal plasticity and metabolism as regulators of organismal energy homeostasis. Nat. Metab. 4, 1444–1458 (2022).
Mihaylova, M. M. et al. Fasting activates fatty acid oxidation to enhance intestinal stem cell function during homeostasis and aging. Cell Stem Cell 22, 769–778.e4 (2018).
Stine, R. R. et al. PRDM16 maintains homeostasis of the intestinal epithelium by controlling region-specific metabolism. Cell Stem Cell 25, 830–845.e8 (2019).
Wang, D., Odle, J. & Liu, Y. Metabolic regulation of intestinal stem cell homeostasis. Trends Cell Biol. 31, 325–327 (2021).
Di Gioia, M. et al. Endogenous oxidized phospholipids reprogram cellular metabolism and boost hyperinflammation. Nat. Immunol. 21, 42–53 (2020).
Reitman, Z. J. & Yan, H. Isocitrate dehydrogenase 1 and 2 mutations in cancer: alterations at a crossroads of cellular metabolism. J. Natl. Cancer Inst. 102, 932–941 (2010).
Tran, T. Q. et al. α-Ketoglutarate attenuates Wnt signaling and drives differentiation in colorectal cancer. Nat. Cancer 1, 345–358 (2020).
Alpha-ketoglutarate (AKG) inhibits proliferation of colon adenocarcinoma cells in normoxic conditions. Scand. J. Gastroenterol. 47, 5 (2012).
Jakob, C. G. et al. Novel modes of inhibition of wild-type isocitrate dehydrogenase 1 (IDH1): direct covalent modification of His315. J. Med. Chem. 61, 6647–6657 (2018).
Clevers, H., Loh, K. M. & Nusse, R. An integral program for tissue renewal and regeneration: Wnt signaling and stem cell control. Science 346, 1248012 (2014).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016).
Rashi S. Oxygen battle in the gut: hypoxia and hypoxia-inducible factors in metabolic and inflammatory responses in the intestine. J. Biol. Chem. 295, 10493–10505 (2020).
Špela K. Regulation of oxygen homeostasis at the intestinal epithelial barrier site. Int. J. Mol. Sci. 22, 9170 (2021).
Keith, B. & Simon, M. C. Hypoxia-inducible factors, stem cells, and cancer. Cell 129, 465–472 (2007).
Pate, K. T. et al. Wnt signaling directs a metabolic program of glycolysis and angiogenesis in colon cancer. EMBO J. 33, 1454–1473 (2014).
Helmuth G. Tales from the crypt: new insights into intestinal stem cells. Nat. Rev. Gastroenterol. Hepatol. 16, 19–34 (2019).
Beumer, J. & Clevers, H. Cell fate specification and differentiation in the adult mammalian intestine. Nat. Rev. Mol. Cell Biol. 22, 39–53 (2021).
Cheng, H. & Leblond, C. P. Origin, differentiation and renewal of the four main epithelial cell types in the mouse small intestine V. Unitarian theory of the origin of the four epithelial cell types. Am. J. Anat. 141, 537–561 (1974).
Kaiko, G. E. et al. The colonic crypt protects stem cells from microbiota-derived metabolites. Cell 165, 1708–1720 (2016).
Rakoff-Nahoum, S., Paglino, J., Eslami-Varzaneh, F., Edberg, S. & Medzhitov, R. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 118, 229–241 (2004).
Stuart, J. A., Aibueku, O., Bagshaw, O. & Moradi, F. Hypoxia inducible factors as mediators of reactive oxygen/nitrogen species homeostasis in physiological normoxia. Med. Hypotheses 129, 109249 (2019).
Zhang, Q. et al. hTFtarget: a comprehensive database for regulations of human transcription factors and their targets. Genom. Proteom. Bioinform. 18, 120–128 (2020).
Meyer, M. B., Goetsch, P. D. & Pike, J. W. VDR/RXR and TCF4/β-catenin cistromes in colonic cells of colorectal tumor origin: impact on c-FOS and c-MYC gene expression. Mol. Endocrinol. 26, 37–51 (2012).
Shimin, Z. Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1α. Science 324, 261–265 (2009).
Schwerd, T. et al. NOX1 loss-of-function genetic variants in patients with inflammatory bowel disease. Mucosal Immunol. 11, 562–574 (2018).
Trimarchi, J. M. & Lees, J. A. Sibling rivalry in the E2F family. Nat. Rev. Mol. Cell Biol. 3, 11–20 (2002).
Hayes, P. et al. Defects in NADPH oxidase genes NOX1 and DUOX2 in very early onset inflammatory bowel disease. Cell Mol. Gastroenterol. Hepatol. 1, 489–502 (2015).
Khoshnevisan, R. et al. NOX1 regulates collective and planktonic cell migration: insights from patients with pediatric-onset IBD and NOX1 deficiency. Inflamm. Bowel Dis. 26, 1166–1176 (2020).
Hsu, N.-Y. et al. NOX1 is essential for TNFα-induced intestinal epithelial ROS secretion and inhibits M cell signatures. Gut 72, 654–662 (2023).
Ohata, H. et al. NOX1-dependent mTORC1 activation via S100A9 oxidation in cancer stem-like cells leads to colon cancer progression. Cell Rep. 28, 1282–1295.e8 (2019).
Garrett, M. et al. Emerging roles of wild-type and mutant IDH1 in growth, metabolism and therapeutics of glioma. Gliomas (ed. Debinski, W.) (Exon Publications, Brisbane (AU), (2021).
Karhausen, J. et al. Epithelial hypoxia-inducible factor-1 is protective in murine experimental colitis. J. Clin. Invest 114, 1098–1106 (2004).
Colgan, S. P., Campbell, E. L. & Kominsky, D. J. Hypoxia and mucosal inflammation. Annu. Rev. Pathol. 11, 77–100 (2016).
Wang, Y. et al. Long-term culture captures injury-repair cycles of colonic stem cells. Cell 179, 1144–1159.e15 (2019).
Pircalabioru, G. et al. Defensive mutualism rescues NADPH oxidase inactivation in gut infection. Cell Host Microbe 19, 651–663 (2016).
Saldova, R. et al. Characterization of intestinal O-glycome in reactive oxygen species deficiency. PLoS One 19, e0297292 (2024).
Birchenough, G. M. H., Nyström, E. E. L., Johansson, M. E. V. & Hansson, G. C. A sentinel goblet cell guards the colonic crypt by triggering Nlrp6-dependent Muc2 secretion. Science 352, 1535–1542 (2016).
Yanes, O. et al. Metabolic oxidation regulates embryonic stem cell differentiation. Nat. Chem. Biol. 6, 411–417 (2010).
Jang, Y.-Y. & Sharkis, S. J. A low level of reactive oxygen species selects for primitive hematopoietic stem cells that may reside in the low-oxygenic niche. Blood 110, 3056–3063 (2007).
Belle, J. E. L. et al. Proliferative neural stem cells have high endogenous ROS levels that regulate self-renewal and neurogenesis in a PI3K/Akt-dependant manner. Cell Stem Cell 8, 59–71 (2011).
Aida, T. et al. Cloning-free CRISPR/Cas system facilitates functional cassette knock-in in mice. Genome Biol. 16, 87 (2015).
Chu, V. T. et al. Efficient generation of Rosa26 knock-in mice using CRISPR/Cas9 in C57BL/6 zygotes. BMC Biotechnol. 16, 4 (2016).
Yang, D. et al. Nociceptor neurons direct goblet cells via a CGRP-RAMP1 axis to drive mucus production and gut barrier protection. Cell 185, 4190–4205.e25 (2022).
Morgan, B., Sobotta, M. C. & Dick, T. P. Measuring EGSH and H2O2 with roGFP2-based redox probes. Free Radic. Biol. Med. 51, 1943–1951 (2011).
Barata, A. G. & Dick, T. P. In vivo imaging of H2O2 production in drosophila. Methods Enzymol. 526, 61–82 (2013).
Mizrachi, A., Graff Van Creveld, S., Shapiro, O. H., Rosenwasser, S. & Vardi, A. Light-dependent single-cell heterogeneity in the chloroplast redox state regulates cell fate in a marine diatom. eLife 8, e47732 (2019).
Schwarzländer, M. et al. Confocal imaging of glutathione redox potential in living plant cells. J. Microsc. 231, 299–316 (2008).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Riba, A. et al. Cell cycle gene regulation dynamics revealed by RNA velocity and deep-learning. Nat. Commun. 13, 2865 (2022).
EnhancedVolcano: publication-ready volcano plots with enhanced colouring and labeling. https://bioconductor.org/packages/devel/bioc/vignettes/EnhancedVolcano/inst/doc/EnhancedVolcano.html.
Bechtel, T. J., Li, C., Kisty, E. A., Maurais, A. J. & Weerapana, E. Profiling cysteine reactivity and oxidation in the endoplasmic reticulum. ACS Chem. Biol. 15, 543–553 (2020).
Topf, U. et al. Quantitative proteomics identifies redox switches for global translation modulation by mitochondrially produced reactive oxygen species. Nat. Commun. 9, 324 (2018).
Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 4, 923–925 (2007).
UniProt Consortium. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2021).
Szklarczyk, D. et al. The STRING database in 2023: protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Shen, X. et al. EZH1 mediates methylation on histone H3 Lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol. Cell 32, 491–502 (2008).
Acknowledgements
The authors thank the Harvard Digestive Disease Center, funded by P30DK034854, and the IDDRC Gene Manipulation Core, funded by NIH P50 HD105351. J.R.T. is supported by NIH grants R03DK125630, RC2DK122532, R35GM142683 and by the Leona M. and Harry B. Helmsley Charitable Trust. X.Z is supported by R35GM151000 and the Kenneth Rainin Foundation. J.O.M. is a New York Stem Cell Foundation—Robertson Investigator. J.O.M. was supported by the Leona M. and Harry B. Helmsley Charitable Trust, The Pew Charitable Trusts Biomedical Scholars, The Mathers Foundation, The New York Stem Cell Foundation, and The Cell Discovery Network, a collaborative funded by The Manton Foundation and The Warren Alpert Foundation at Boston Children’s Hospital. E.W. is supported by NIH R35GM134964. D.Y. is supported by the National Natural Science Foundation of China (9247410063, 82494520023). We would like to thank Dr. Amy Shepherd for assistance with tissue cryo-sectioning.
Author information
Authors and Affiliations
Contributions
Conceptualization, X.C., J.R.T.; methodology, X.C., K.R., B.B., E.N., A.K., Z.W., M.B., D.Y., U.J., and E.W.; investigation, X.C., K.R., B.B., E.N., A.K., Z.W., A.M., Q.P., and J.R.T.; writing – original draft, X.C., K.R., and J.R.T.; writing—review and editing, X.C., D.Y., E.W., S.S., J.O.M., X.Z., and J.R.T.; resources, E.W., M.R., D.K., and J.R.T.; supervision, M.R., S.S., X.Z., J.O.M., U.J, and J.R.T.; funding: J.R.T.
Corresponding author
Ethics declarations
Competing interests
J.O.M. reports compensation for consulting services with Tessel Biosciences and Radera Biotherapeutics. Michael Rutlin is now a full-time employee of Novo Nordisk. The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Toshio Takahashi, Qingding Wang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, X., Raghunathan, K., Bao, B. et al. A niche-dependent redox rheostat regulates epithelial stem cell fate. Nat Commun 16, 11713 (2025). https://doi.org/10.1038/s41467-025-66636-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66636-5