Abstract
GTP-binding protein 1 (GTPBP1) is a widespread translational GTPase closely related to elongation factor eEF1A. The loss of GTPBP1 leads to neurodevelopmental and neurodegenerative disorders in animals. Although linked to translation and quality control mechanisms, GTPBP1 molecular functions remain largely obscure. Similarly to eEF1A, GTPBP1 delivers aminoacyl-tRNA to the ribosome, but the ensuing GTPBP1-mediated elongation is slow. Here, using cryo-EM of mammalian 80S ribosomal complexes bound to GTPBP1 and aa-tRNA with GTP or the non-hydrolysable analog GDPCP, we show that the distinct GTPBP1 architecture and interactions with tRNA underlie slow GTPBP1 dissociation after GTP hydrolysis, resulting in delayed tRNA accommodation. Slow dissociation correlates with an extended proofreading stage and higher accuracy of GTPBP1-mediated decoding, potentially allowing GTPBP1 to elicit its putative quality control functions. GTPBP1 visualization provides the foundation for mapping and elucidating GTPBP1 mutations associated with human diseases.
Similar content being viewed by others
Introduction
GTP-binding protein 1 (GTPBP1) belongs to the widespread GTPBP1/GTPBP2 group of translational GTPases that are closely related to elongation factor eEF1A (and its prokaryotic homolog EF-Tu) delivering aminoacyl-tRNAs to the ribosome, eukaryotic release factor eRF3, and surveillance factor Hbs11. GTPBP1/GTPBP2 are encoded by all major groups of eukaryotes except for yeasts and multicellular plants. In many eukaryotes, both GTPBP1 and GTPBP2 representatives are found, which in humans share ~68% similarity1,2,3,4,5. Like all other translational GTPases, GTPBP1/GTPBP2 contain three conserved domains—a GTPase (G) domain and two β-barrel domains—but they also have long N-terminal and C-terminal extensions that are the most divergent elements in these proteins1,2,3,4. GTPBP1/GTPBP2 are expressed in a wide range of tissues and organs in mice and in humans, and are localized to the cytoplasm6,7.
The loss of either GTPBP1 or GTPBP2 causes neurodegeneration and/or neurodevelopmental disorders in humans8,9,10,11,12 and in mice that are deficient in the main central nervous system (CNS) Arg-tRNA isodecoder13,14. The loss of GTPBP1 also disrupts vascular patterning in zebrafish15. In Drosophila, reduced expression of the GTPBP1/GTPBP2 orthologs Dgp-1 and CG2017 leads to locomotor function defects12,16,17 and neuron-specific repression of Dgp-1 expression results in olfactory learning defects18.
The molecular function of GTPBP1 is associated with translation and surveillance/quality control mechanisms14,19. The studies of neurodegeneration caused by GTPBP1 deletion and Arg-tRNA disruption in mice suggest that GTPBP1 resolves the ribosomes paused during elongation14. GTPBP1 was also proposed to recruit the exosome to promote mRNA degradation in the stalled ribosomal complexes19,20. In Drosophila, Dgp-1 is upregulated in response to stresses, including oxidative stress5, thermal stress21, and hyperoxia22. Furthermore, Dgp-1 is upregulated in response to over-expression of the endoplasmic reticulum (ER) stress-induced dPERK kinase23, and loss-of-function alleles of Drosophila Dgp-1 reduce the viability of Parkin null mutants deficient in Parkin E3 ubiquitin ligase that targets proteins for degradation24. These observations indicate that GTPBP1 may be protective against stresses affecting translation regulation and proteostasis, in keeping with the protein’s roles in translation and/or quality control.
Biochemical experiments revealed that GTPBP1 possesses eEF1A-like elongation activity and can deliver cognate aminoacyl-tRNA to the ribosomal A site in a GTP-dependent manner19. However, in contrast to eEF1A, GTP hydrolysis by GTPBP1 was not followed by rapid peptide bond formation, and GTPBP1-mediated elongation was very slow. Additionally, unlike eEF1A, GTPBP1 could also promote ribosomal delivery of deacylated tRNA. Although GTPBP2 was also shown to bind tRNAs in a GTP-dependent manner, it could not promote translation elongation by delivering aminoacyl-tRNA to the ribosome19.
The structural basis that determines the unusual kinetics of GTPBP1-mediated elongation is unknown. In this work, to elucidate the mechanism of GTPBP1 function, we report cryogenic electron microscopy (cryo-EM) structures of human GTPBP1•aa-tRNA on the 80S ribosome. The intermediates obtained with GTPBP1 and either the non-hydrolysable analog GDPCP or GTP visualize ribosome complexes prior to or after GTP hydrolysis. They bring insights into the ribosome dynamics, highlighting a role for 40S shoulder domain movements in GTPase activation upon mRNA decoding by GTPBP1•aa-tRNA. These structures reveal how the distinct protein architecture of GTPBP1, including an additional N-terminal domain and the shoulder-interacting H-loop in place of the α2 helix of canonical GTPases, is responsible for establishing GTPBP1-specific interactions with tRNA and the ribosome. These interactions differ from those of eEF1A and account for the prolonged association of GTPBP1•aa-tRNA with the ribosome after GTP hydrolysis and delayed accommodation of tRNA in the A site of the ribosome. Our structures and biochemical data suggest the increased decoding accuracy of GTPBP1.
Results
GTPBP1•aa-tRNA•ribosome complexes in the presence of GDPCP
During canonical mRNA decoding, elongation factors (EF: EF-Tu in bacteria and a/eEF1A in archaea or eukaryotes) bound with GTP and aminoacyl-tRNA sample the ribosomal A site. The EF•GTP•aa-tRNA complex initially binds to the small subunit, placed away from the GTPase-activating sarcin-ricin loop (SRL) of the large subunit25,26,27. This placement prevents spontaneous GTP hydrolysis, required for the dissociation of EF from tRNA. If the tRNA anticodon matches the mRNA codon, rearrangements of the small subunit bring the EF closer to the SRL, triggering GTP hydrolysis and EF•GDP dissociation, which allows the accommodation of the tRNA into the A site and formation of a new peptide bond. To visualize the dynamics of GTPBP1 on the ribosome prior to GTP hydrolysis, we took advantage of the non-hydrolysable GTP analog GDPCP (5′-guanosyl-β,γ-methylene-triphosphate), which had allowed visualization of other translational GTPases on the ribosome28,29,30. We incubated GTPBP1•Phe-tRNAGAA•GDPCP with the 80S ribosomes purified from rabbit reticulocyte lysates and programmed with a derivative of β-globin mRNA and containing initiator Met-tRNAiMet bound to the AUG start codon in the P site and the UUC Phe codon in the A-site19. The sample was subjected to cryo-EM. To elucidate the role of GTP hydrolysis in GTPBP1, we assembled an 80S ribosome complex with GTPBP1•Phe-tRNAPhe and GTP, similarly to the complex described above, placed the mixture on ice, and collected two cryo-EM datasets corresponding to 1- and 5-min time points of the reaction.
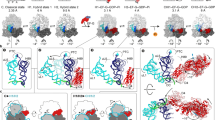
Classification of cryo-EM data yielded ten maps at average resolutions of 2.9 to 3.1 Å, bringing insights into the stages of mRNA-decoding by GTPBP1: ribosomes with a vacant A site (Structures Ia through Ic featuring slightly different conformations of the 40S subunit, as discussed); ribosomes bound with GTPBP1•tRNA•GDPCP, featuring GTPBP1 away from the GTPase-activating SRL (Structures IIa-d) and GTPBP1 docked at the SRL (Structure III); ribosome bound to GTPBP1•tRNA•GDP (Structure IV); and ribosome with tRNAPhe accommodated in the A-site (Structure V) (Fig. 1A, Supplementary Table 1, Supplementary Figs. 1 and 2).
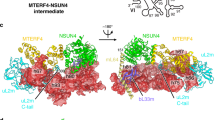
A The ensemble of models resulted from the analysis of the Cryo-EM data of GTPBP1•GDPCP•aa-tRNA and GTPBP1•GDP•aa-tRNA datasets: Structures Ia-c, featuring a vacant A site; Structures IIa-d (undocked GTPBP1) and III (GTPBP1 docked at SRL), featuring ribosomes with GTPBP1•GDPCP and aa-tRNA; Structure IV (GTPBP1•GDP and aa-tRNA), and Structure V with accommodated A-site aa-tRNA. B Cryo-EM density of ribosome with GTPBP1•GDPCP•aa-tRNA (Structure III, 60S subunit is shown in cyan, 40S subunit yellow, aa-tRNA green, GTPBP1 magenta). C Comparison of domain organizations of human GTPBP1 (in Structure III) and eEF1A (PDB: 5LZS).
We first describe the overall structure of GTPBP1, its interactions with tRNA, GTP, and the ribosome, and then discuss the structural rearrangements of the ribosome during mRNA decoding. GTPBP1•Phe-tRNAGAA•GDPCP binds at the ribosomal A site (Fig. 1B). The architecture of GTPBP1 (most well resolved in Structure III, Fig. 2) is similar to that of eEF1A, with the exception of a novel N-terminal domain (aa 62-151), which features two α-helices packed onto a β-sheet, residing on top of the GTPase domain (Fig. 1C). Despite the lack of sequence similarity, this domain structurally resembles archaeal/eukaryotic initiation factors e/aIF1 and the C-terminal domain of bacterial initiation factor IF331,32, according to the DALI fold comparisons33. The N- and C-termini of GTPBP1 (aa 1-61 and 575-669, respectively), which are predicted to form unfolded tails, are not resolved in our maps, consistent with high conformational flexibility (Fig. 1B, C). The two β-barrel domains of GTPBP1 (domains 2 and 3: aa 394-486 and 490-574, respectively) are bound at the shoulder domain of the 40S subunit (Fig. 1B, C). Domain 3 binds the acceptor arm of the A/T tRNA (A-site/Ternary complex), while the 3′ CCA end of tRNA is held in the cleft between the G-domain and domain 2, similar to that in eEF1A complexes (Fig. 1C)28,34.
A Close-up view of GTPBP1 (magenta) and aa-tRNA (green) on the ribosome. B The nucleotide-binding pocket of GTPBP1. C Comparison of the catalytic GTPase pockets of GTPBP1, eEF1A (PDB: 8G6J), and eRF3 (PDB: 5LZT). D Top panel: Distinct interactions of GTPBP1 and eEF1A (PDB: 5LZS) with 40S and the uL14-mediated bridge between the large and small subunits; the complexes were aligned by 28S rRNA. Zoom-in: Potassium-binding site coordinated by the GTPBP1-specific H-helix. E Coloring GTPBP1 according to sequence conservation113 (sequence alignment is shown in Supplementary Fig. 3a). F Interactions of GTPBP1 and aa-tRNA, the colored surface represents the electrostatic potential surface of GTPBP1 (red, negative; blue positive). G Interactions of the tip of aa-tRNA with GTPBP1 (left) and eEF1A (PDB: 8G6J, right).
In the presence of GDPCP mimicking pre-GTP-hydrolysis states, GTPBP1 adopts different positions in the groups of Structures II and Structure III, resolved at ~3.1 and 2.9 Å average resolutions, respectively (Figs. 2 and 3). Structures IIa through IId differ by slight shifts (~1 Å) in GTPBP1 and aa-tRNA positions (Fig. 1A). In this group of structures, the GTPase domain of GTPBP1 (aa 153–385) is placed ~7 Å from the SRL (at the catalytic His256 described below) (Fig. 3A), resembling a pre-activation state of eEF1A and EF-Tu observed during codon sampling29,34,35,36. The elbow of tRNA (at nucleotides 19 and 56) is stabilized by the 28S rRNA of the L11 stalk (helices H43/44) at nucleotides 1981 and 2009 (Fig. 3C). Unlike the pre-activation state in eEF1A structures34, where the codon-anticodon interactions are poorly resolved, the anticodon of tRNA is fully paired with the mRNA codon in Structures IIa through IId (Fig. 3G).
A Comparison of GTPBP1 positions in Structures IIa-d and III aligned by 28S rRNA. B Positions of catalytic His256 relative to the A4607 of the SRL in Structures IIa-d (pale colors) and III (bright colors). C Comparison of tRNA elbow positions in Structures IIa-d (pale colors) and III (bright colors). D Pie chart representing proportions of ribosome particles corresponding to Structures IIa-d and Structure III. Source data are provided as a Source data file. E Worm diagram representing maximal RMSD in the ensemble of GTPBP1•aa-tRNA Structures IIa-d and III aligned via GTPBP1. F Rearrangements of the small subunit of the ribosomes with the vacant A site (Structures Ia-c, left panel) upon binding of GTPBP1•aa-tRNA•GDPCP (Structure IIa, middle panel) and docking of GTPBP1 into the SRL (Structure III, right panel). Cylinders’ lengths and colors are proportional to maximal root-mean-square distances within the ensemble of structures specified on the top of each panel. G Close-up view of the decoding center in Structures IIa-d and III. H Comparison of Phe-tRNAPhe positions in Structures IIa-d through Structure III to Structure V (accommodated tRNA), aligned by 28S RNA.
In Structure III, GTPBP1 is shifted toward the A site (Figs. 2A and 3A), resembling the GTPase activation states of eEF1A and EF-Tu28,34. The GTP substrate (mimicked by GDPCP in our structures) is held in place by conserved regions of GTPBP1, including the switch loops (Fig. 2B and Supplementary Fig. 4a). In translational GTPases, the switch loops must rearrange upon GTP hydrolysis to allow GTPase dissociation from the ribosome36,37. The base of GDPCP interacts with the guanine recognition motif [NT]KxD (where x is any amino acid), which in GTPBP1 is represented by 308TKID311, in contrast to most other translational GTPases featuring NKxD38. Here, the guanine base is stacked on Lys309 and its Watson-Crick edge interacts with Asp311 (Fig. 2B and Supplementary Fig. 4a). The triphosphate of GDPCP interacts with the backbone and sidechains of the P-loop (aa 167-174) and Arg193 of switch 1 (aa 182-212). The γ- and β-phosphates are bound to a Mg2+ ion coordinated by the conserved Ser174 of the P-loop and Thr208 of switch 1 (Fig. 2B and Supplementary Fig. 4a). Recently, a T208A mutation was shown to cause neurodegeneration in humans, highlighting the key importance of this residue for GTPBP1 function12. The γ-phosphate hydrogen bonds with the backbone of the conserved Gly255 in switch 2 (aa 252–269), which is well-resolved in the map next to the catalytic His256 (Fig. 2B and Supplementary Fig. 4a).
The catalytic His256, conserved in translational GTPases (His95 in eEF1A and His300 in eRF3) and required for GTP hydrolysis28,34, is placed at the backbone phosphate of A4607 (A2662 in E. coli) of the SRL (Fig. 2A, C and Supplementary Fig. 4b). The side chain of His256 is rotated ~6 Å from the γ-phosphate (i.e., “outward” conformation), consistent with a catalytically inactive conformation. To allow His256 to move closer, to ~4 Å from the γ-phosphate (i.e., “inward” conformation), a neighboring residue, Arg207, must shift away. In Structure III, Arg207 appears to adopt two alternative conformations, in which the guanidine group interacts either with the backbone phosphate of A4607 or with both A4607 and G4608 (Fig. 2C and Supplementary Fig. 4b). The latter rotamer of Arg207 is similar to the position of Lys276 in eRF3, whose catalytic histidine is placed near γ-phosphate (Fig. 2C)28. Thus, the dynamics of GTPBP1 residues His256 and Arg207 appear critical for GTP hydrolysis near the SRL, consistent with the dependence of the GTPase activity on the ribosome19.
Despite similarity to eEF1A, significant differences in GTPBP1 interactions with its cofactors and the ribosome suggest differences in the dynamics and/or kinetics of GTPBP1-guided tRNA delivery. Perhaps most notable is the absence of helix α2, which in eEF1A and its paralogs eRF3 and Hbs1l is a key part of the switch 1 region, bridging the 40S and 60S subunits during GTPase activation28,34. In eEF1A, Arg37 of α2 stacks on A464 at the shoulder of 18S rRNA, while the other end of the helix reaches toward protein uL14 of the 60S subunit (Fig. 2D)28,34. GTPBP1, however, features a long loop stemming from the GTPase β-sheet that is not part of a switch region (Fig. 2D and Supplementary Fig. 4c, d). On the ribosome, the loop is organized into a short β-hairpin (aa 219–236) and a short α-helix (aa 237–243), which resemble a handle and hence are termed the H-hairpin and H-helix (Fig. 2D and Supplementary Fig. 4c, d). His231 at the tip of the β-hairpin stacks on A464 of 18S rRNA, likely stabilizing GTPBP1 between the small and large subunits (Fig. 2D). Unlike in eEF1A-bound complexes, the N-terminus of uL14 makes no contact with GTPBP1 and is therefore disordered (Fig. 2D). Nevertheless, Glu204 and Lys200 of GTPBP1 are positioned to form salt bridges with Lys109 and Glu111 of uL14, resembling the bridging of Glu68 and Lys64 of eEF1A (Supplementary Fig. 4e). Thus, in the absence of the rigid α-helix of other GTPases, potentially labile interactions of the ionic groups and dynamic H-loop (see below and Fig. 3) of GTPBP1 with the ribosome may confer distinct ribosome association and dissociation pathways.
The interface between H-helix and the N-domain linker features strong density within a pocket formed by backbone carbonyl oxygens (Supplementary Fig. 4c, f). Given the characteristic coordination distances of ~2.8 Å and an octahedral coordination pattern39,40,41, the density likely corresponds to a potassium ion (Fig. 2D and Supplementary Fig. 4c, f)40. The presence of a K+-binding pocket in GTPBP1 further suggests important functional roles for the distinctive H-handle/loop and the N-domain. For example, K+ exchange may play a role in GTPBP1 dissociation from the tRNA or ribosome, which may require a rearrangement of the H-handle and/or the N-terminal domain.
Distinct dynamics of GTPBP1 in comparison to eEF1A or EF-Tu are further emphasized by a distinct particle distribution among cryo-EM classes. Unlike eEF1A and EF-Tu, shown by cryo-EM and FRET studies to predominantly adopt a GTPase-activated state with cognate tRNA29,34,36, GTPBP1 is predominantly detached from the SRL, with Structures IIa-d accounting for ~66% of GTPBP1-bound ribosomes (Fig. 3D). Helix α2 of eEF1A was hypothesized to stabilize GTPase-activated state(s) by forming a bridge with both ribosomal subunits28. The absence of α2 in GTPBP1 therefore may result in GTPBP1 detachment from the large subunit. By contrast, the flexible glycine rich H-loop preserves its contacts with the small subunit in Structures IIa-d and III, helping to accommodate different positions of GTPBP1 (Fig. 3E). These differences are likely critical for GTPBP1-mediated accuracy of mRNA decoding as discussed below.
The N-domain, which does not interact with the ribosome or tRNA, contacts the G-domain through hydrophobic and salt-bridge interactions (Fig. 2E and Supplementary Fig. 4g). Besides several residues at the interface, the N-domain has a low sequence conservation (Fig. 2E and Supplementary Fig. 3a) accompanied by high conservation of the length and spatial organization. This supports an important role of this domain as a structural element, as discussed below.
GTPBP1 interactions with Phe-tRNAPhe reflect the protein’s distinct tRNA substrate preferences and/or tRNA association/dissociation dynamics relative to those of eEF1A. For example, GTPBP1•GTP has high affinities to either amino-acylated tRNAs (Kd of ~3 nM) and deacylated tRNAs (~30 nM;19). By contrast, eEF1A•GTP strongly prefers aa-tRNA (~1–5 nM) over deacylated tRNA (up to 10 µM)42. Whereas the overall pattern of tRNA binding at positively charged interfaces of domain G and β-barrel domains in GTPBP1 is similar to that of eEF1A (Fig. 2F), the phenylalanine’s carbonyl attached at the 3′ CCA end of Phe-tRNAPhe (Fig. 2G and Supplementary Fig. 4h) does not hydrogen-bond with GTPBP1 (4–5 Å), unlike that in the eEF1A complex (2.5 Å; Fig. 2G). This likely underlies the less discriminatory binding of GTPBP1 to aa-tRNA and deacyl-tRNAs.
By contrast, the 3′-CCA76 end appears to be held more strongly by GTPBP1 than eEF1A. The terminal base of the tRNA is buttressed by a hydrophobic/aromatic stacking system in GTPBP1. Here, the adenine of A76 is sandwiched between the conserved Val405, Val408, and Val411 on one side and His445, in turn stacked on Pro451, on the other side (Fig. 2G and Supplementary Fig. 4h). This contrasts with eEF1A providing the acidic Glu293 in the place of histidine. The A76 phosphate interacts with Arg449 of GTPBP1, which is packed onto conserved Trp237 of the H-helix of the G-domain (Fig. 2G). In eEF1A, His296 is placed similarly to Arg449 to interact with the phosphate group, but the histidine does not appear to contact the G-domain (Fig. 2G). Furthermore, the acceptor stem of tRNA at pairs 1:72 and 2:71 is held by a short loop-helix structure (aa 199–205) of the switch 1 region, which must rearrange upon GTP hydrolysis to release a translational GTPase from the ribosome43. In GTPBP1, the conserved His199 and His201 stack within the tRNA minor groove, while the loop-helix of eEF1A with no aromatic residues appears to have a looser interaction in this region (Supplementary Fig. 4i). Overall, the buried surface area between GTPBP1 and aa-tRNA (1655.9 Å2) is 15% higher than that between eEF1A and aa-tRNA (1437.6–Å2) on the ribosome, suggesting a tighter interaction for GTPBP1. These structural differences may contribute to the distinct dynamics of the tRNA release from GTPBP1 and eEF1A upon GTP hydrolysis, as discussed below.
Structural insights into GTPase activation
The movement of GTPBP1 relative to the SRL (from Structures IIa-d to III) is coupled with changes in tRNA and the 40S subunit (Fig. 3), bringing insights into the coordination of codon recognition with GTPase activation. Previous studies showed that in EF-Tu and eEF1A ternary complexes, codon recognition in the ribosomal decoding center of the 40S subunit leads to a closure of the shoulder domain toward the 40S body28,29,30,44. We compared Structures IIa-d and III to Structures Ia-c, which correspond to “pre-delivery” ribosomes, whose A site is vacant and P site is bound with the initiator tRNA (Fig. 3F). To distinguish small stochastic movements of the 40S shoulder in three “pre-delivery” maps, we developed “classification of transferred heterogeneity” (CTH; “Methods”). We find that in “pre-delivery” ribosomes, the shoulder of the 40S subunit undergoes a “rocking” movement within a ~2-Å range at its periphery (Fig. 3F, G and Supplementary Movie 1). In Structures IIa-d to III, the shoulder progressively shifts by ~2.5 Å closer to the body than in the ribosomes with a vacant A site (Fig. 3F, G). Furthermore, the SRL docking of GTPBP1 (Structure IIa-d to III) is accompanied by a subtle small ribosomal subunit (SSU) rotation (0.5–0.7°), resembling the conformational changes in eEF1A-bound complexes28,34 (Fig. 3F, G and Supplementary Movie 1).
Interactions in the decoding center are similar to those with cognate eEF1A complexes28,34. As the anticodon loop of Phe-tRNAGAA base-pairs with the UUC codon (Fig. 3G), nucleotides A1824 and A1825 (A1492 and A1493 in E. coli) flip out of helix 44 of 18S rRNA body to monitor the tRNA-mRNA helix (Fig. 3G). They are joined by nucleotide G626 (G530 in bacteria)—at the shoulder domain—which shifts from its position in the “pre-delivery” ribosomes by ~2.5 Å and, similarly to its bacterial counterpart, rearranges from the syn to anti conformation (Fig. 3G and Supplementary Fig. 4j)29. This range of movement is also observed in bacterial ribosomes with EF-Tu, albeit the overall shoulder movement in the 80S ribosome is less pronounced29,45. In addition, conserved His76 of eukaryotic protein eS30 becomes resolved to interact with the phosphate of nucleotide A36 of the tRNA anticodon, highlighting a potential contribution of eukaryote-specific proteins in codon recognition28 (Fig. 3G). As the 40S shoulder and GTPBP1 shift toward the SRL, the tRNA elbow slides ~8 Å along the H43/44 stalk toward the ribosomal A site, coinciding with the tRNA accommodation trajectory (Fig. 3C, H). The lower resolution of H43/H44 indicates destabilization of its contact with the tRNA (observed in Structures IIa-d), reminiscent of that during aa-tRNA delivery by eEF1A34.
In summary, our maps demonstrate that, similarly to EF-Tu and eEF1A ribosome complexes, the 40S decoding-center rearrangements upon codon-anticodon helix formation are correlated with the 40S shoulder movement toward the body. The movement of the shoulder-bound GTPBP1 brings the GTPase domain toward the SRL for the activation of GTP hydrolysis.
Ribosome interaction with GTPBP1•aa-tRNA in the presence of GTP
In our previous work, time-resolved cryo-EM allowed us to visualize more than a dozen structural intermediates of mRNA decoding by EF-Tu-delivered tRNA upon GTP hydrolysis36. GTP hydrolysis leads to the gradual rotation of the G-domain relative to the β-barrel domains (Fig. 4E), and similar rearrangements were implicated from the comparison of GTP- and GDP-bound isolated structures of eEF1A36,43,46,47 and eRF348. The gradual interdomain rearrangement results in the gradual loss of contacts between aa-tRNA and EF-Tu, allowing EF-Tu dissociation and aa-tRNA accommodation into the peptidyl-transferase center36. Antibiotics kirromycin (for EF-Tu44), and didemnin B, plitidepsin and ternatin-4 (for eEF1A28,34,49) trap these GTPases on the ribosome in the GDP-bound state by binding at the interface between the G domain and second β-barrel, blocking the protein interdomain rearrangements28,44.
A Comparison of GTPBP1 position in Structure III and IV aligned by 28S rRNA. B Cryo-EM maps of the nucleotide-binding pockets in complexes Structure III and IV. To improve interpretability of the Structure IV map, the data were refined with a composite mask101. Black circles indicate location of γ-phosphate. C Close up view of the decoding center in Structure III and IV. D Distribution of particles in datasets obtained in the study. Source data are provided as a Source data file. E Structural rearrangements in EF-Tu after GTP-hydrolysis. The models identified on the figure36 were aligned by domains 2 and 3. F The N-domain can clash with the linker between domains G and 2 after GTP-hydrolysis. The G domain and domains 2 + 3 of GTPBP1 were separately aligned to the corresponding domains of EF-Tu (PDB: 6WD5) from36. G The N-domain linker and H-helix can clash with the domain 2 after GTP-hydrolysis. The G domain and domains 2 + 3 of GTPBP1 were separately aligned to the corresponding domains of EF-Tu (PDB: 6WD6) from36. H The N-domain can interfere with the post-hydrolysis state of GTPBP1 on ribosome. The G-domain of GTPBP1 is aligned against the G-domain of “extended” GDP-bound eEF1A (PDB: 8B6Z). The rest of ribosome is PDB: 5LZS fitted in the map corresponding to PDB: 8B6Z (EMD-15893).
Thorough classification of GTP-datasets at 1- and 5-min time points yielded several classes that closely resemble Structures IIa-d, with the G-domain of GTPBP1 posed away from the SRL and a codon-anticodon interaction in the decoding center (Fig. 4A, C). We term the most well-resolved state Structure IV. We could not identify a sizeable class that resembled the GTPase activation Structure III, with GTPase next to the SRL. This indicates that the GTPase-activated state is only transiently populated by the GTP-bound GTPase, similar to the low abundance “activation” state for EF-Tu•GTP36.
In contrast to EF-Tu in the previous work, and to GTPBP1 with GDPCP described above, the GTPase center of Structure IV contains GDP (Fig. 4B). Gly255 in the switch 2 region, which in GDPCP-bound complexes coordinates the γ-phosphate (Figs. 2 and 3), is disordered in the presence of GDP. This coincides with the lack of density for the γ-phosphate (Fig. 4B), indicating that this region of switch 2 becomes dynamic upon phosphate release, as in other GTPases bound with GDP49. Despite GTP hydrolysis, GTPBP1 domains retain the conformation observed with GDPCP in Structures IIa-d. The short loop-helix structure of switch 1 (aa 199-205) remains well-resolved in contact with the aa-tRNA, contrasting the rearranged switch 1 initiating EF-Tu•GDP release from the ribosome36. The GTPBP1’s GTPase domain resembles that in eEF1A•GDP•aa-tRNA complexed with didemnin B, which blocks EF release from the ribosome28,34,49. The inhibition of GTPBP1•GDP rearrangement into an open conformation to release aa-tRNA is likely caused by the extended N-terminal domain, as discussed below. The abundance of ribosome particles with GTPBP1•GDP•aa-tRNA does not change significantly between the two time points (~5%) (Fig. 4D). By contrast, time-resolved cryo-EM showed rapid depletion of ribosome-bound EF-Tu or EF-G over time36,37. These results indicate a delayed tRNA accommodation after GTP hydrolysis on GTPBP1, which correlates with the previously observed slow peptide bond formation following rapid hydrolysis of GTP19. Accordingly, the abundance of ribosomes with accommodated tRNA (collectively in the A/A and post-peptidyl-transfer A/P states) slowly increases from ~11% in the GDPCP-inhibited complex to 38% and 44% at 1 min and 5 min, respectively, in the GTP complexes (Fig. 4D).
What causes the slow dissociation of GTPBP1? Biochemical studies suggested the NTD of GTPBP1 might be important: removing the N-domain (residues 1-152) (Supplementary Fig. 5) was shown to increase the rate of aa-tRNA delivery by GTPBP1, resembling that by eEF1A19. Furthermore, viral GTPBP1-like proteins without the NTD and H-helix-loop drive elongation at rates similar to that of eEF1A50 (Supplementary Fig. 5). Our structural analysis suggests how the N-terminal domain residing on top of the GTPase domain may help delay GTPBP1 dissociation from the ribosome. In EF-Tu, the GTPase domain “rotates” away from the tRNA acceptor arm to initiate protein dissociation (Fig. 4E)36. A similar rearrangement in GTPBP1 is likely inhibited by the NTD clash/interaction with the linker connecting the GTPase domain and β-barrel domains (Fig. 4F) and/or clash/interaction of the NTD-linker and H-helix-loop with domain 2 (Fig. 4G). Similarly, GTPBP1 is not compatible with the recently observed extended conformation of eEF1A bound to the fully accommodated aa-tRNA35, closely resembling a GDP-bound eEF1A43. Here, GTPBP1’s NTD would severely clash with the SSU (Fig. 4H). These comparisons indicate that the NTD changes the conformational dynamics of GTPBP1, resulting in slower dissociation. The role of the N-domain as a steric block is further supported by its architecture conservation despite high sequence heterogeneity (Fig. 2G).
GTPBP1 promotes high fidelity of mRNA decoding
The high fidelity of mRNA decoding is ensured by kinetic proofreading, a key multi-step discriminatory process of aa-tRNA delivery51,52,53. In kinetic proofreading, two selection steps are separated by an irreversible step that strongly increases the specificity of selection compared to that achieved by just the thermodynamic (i.e., free energy) differences between cognate and near-cognate substrates54. During elongation, the initial thermodynamic selection of the cognate EF•aa-tRNA•GTP complex on the ribosome is followed by the irreversible step of GTP hydrolysis on the EF in complexes with cognate and, less effectively, with near-cognate tRNAs (i.e., with a mismatched anticodon nucleotide)34,55. After GTP hydrolysis, the near-cognate tRNA has another chance to dissociate (i.e., be rejected) during accommodation of tRNA into the peptidyl-transferase center (also see next section and Figs. 5 and 6). Indeed, near-cognate tRNA was shown to dissociate either alone during accommodation36,55,56 or together with EF•GDP55,57,58. Accordingly, slowing down GTP hydrolysis or EF•GDP dissociation from aa-tRNA would increase the off-rate differences between cognate and near-cognate tRNAs, resulting in increased selection accuracy51. Indeed, the slow hydrolysis of the GTP analog guanosine 5′-O-[gamma-thio]triphosphate (GTPγS) improves the fidelity of EF-Tu•aa-tRNA decoding by several orders of magnitude59. Likewise, slowing the dissociation of EF-Tu•GDP from aa-tRNA linearly slows the accommodation of tRNA and peptide bond formation60, resulting in increased proofreading58. By contrast, elevated Mg2+ concentrations, which stabilize ribosome-tRNA interactions, decrease the fidelity of mRNA decoding58,61.
Top panels: Schematic representation of MS(UCU)HL-STOP and MS(UCC)HL-STOP mRNAs. Lower panels: Toeprinting analysis of the activities of GTPBP1 and eEF1H in single-cycle elongation on (A) MS(UCU)HL-STOP and (B) MS(UCC)HL-STOP mRNAs at the indicated free [Mg2+] with in vitro transcribed Ser-tRNASer-AGA, which is cognate to the Ser codon in the first and near-cognate to the Ser codon in the second mRNA, respectively. 80S initiation complexes (ICs) were assembled using individual translational components, as indicated. After assembly of 80S ICs, free [Mg2+] was adjusted to the indicated values before addition of eEF2, eEF1H, GTPBP1, and Ser-tRNASer-AGA. In all cases, subsequent reverse transcription was done at 20 mM free [Mg2+]. The division between sequence and other lanes indicates that these two sets of lanes were derived from the same gel, exposed for different lengths of time. The experiment has been independently performed 3 times with similar results.
GTPBP1 achieves higher accuracy than eEF1A-catalyzed decoding due to distinct protein architectures and interactions with the ribosome (small subunit cyan; large subunit yellow) and aa-tRNA (green) during initial selection and proofreading.
GTPBP1 demonstrates a different behavior on the ribosome than eEF1A, bringing insights into the accuracy of mRNA decoding by GTPBP1. First, GTPBP1 prefers codon-sampling-like conformations away from the SRL rather than GTPase-activated state(s) at the SRL, as the ratio of ribosome particle numbers in Structures IIa-d to those in Structure III is ~3:1 (Fig. 3A). This sharply contrasts the equilibrium of codon recognition and GTPase activation states observed for EF-Tu (~1:6, also formed with GDPCP29) and for eEF1A (~1:62; formed with GTPγS34). The equilibrium shift away from the GTPase-activated state suggests that GTPBP1 allows more time for initial selection or proofreading51. Second, detection of the GDP-bound GTPBP1 in Structure IV prior to GTPBP1 dissociation contrasts the fast dissociation of canonical EFs after GTP hydrolysis36, implying slower tRNA accommodation. Indeed, biochemical data report that peptide bond formation following GTP hydrolysis by GTPBP1 is slower than that for eEF1A19. Since the slower tRNA accommodation correlates with increased decoding accuracy, we asked whether aa-tRNA delivery by GTPBP1 can be more accurate than by eEF1A.
To this end, we compared the activities of GTPBP1 and eEF1A on cognate and near-cognate codons. We assembled 80S initiation complexes from individual components of the translation system19 using mRNAs encoding the MSHL-STOP sequence. The second codon (Ser) was varied to contain UCU (cognate) or UCC (near-cognate), and the ability to promote elongation with Ser-tRNASer-AGA was measured for GTPBP1 or eEF1H (Fig. 5A and Supplementary Fig. 6; eEF1H complex includes eEF1A and its GDP/GTP exchange factor, which is not directly involved in decoding62). Both GTPBP1 and eEF1H induced efficient single-cycle elongation on the cognate Ser codon UCU (Fig. 5A) in the presence of translocase eEF2. By contrast, the activity of GTPBP1 on the near-cognate UCC codon was very low, whereas eEF1H promoted efficient elongation on this codon (Fig. 5B, compare lanes 3 and 9), indicating higher decoding accuracy for GTPBP1 than eEF1H. Elevation of [Mg2+] from 2.5 to 7 mM progressively increased GTPBP1-mediated elongation (Fig. 5B, lanes 3–5), in keeping with the decreased decoding accuracy. Further elevation of [Mg2+] to 10–20 mM nearly abrogated elongation in both cases (Fig. 5B, lanes 6,7,11, and 12), likely due to the inhibition of eEF2-dependent translocation63. In summary, these data indicate that GTPBP1-mediated elongation can be more accurate than eEF1A-mediated elongation, as GTPBP1 allows only low levels of decoding on a near-cognate codon at near-physiological Mg2+ concentrations.
Discussion
We describe structures of mammalian 80S•GTPBP1•aa-tRNA complexes formed in the presence of GDPCP or GTP. Although GTPBP1 only shares 14.9% sequence similarity with the canonical elongation factor eEF1A, its structural organization is very similar to that of eEF1A (RMSD of 2.3 Å). Nevertheless, our structures uncover important differences between GTPBP1 and other translational GTPases, yielding mechanistic insights into functional aspects of GTPBP1.
Our findings suggest the following mechanism of tRNA delivery by GTPBP1•GTP (Fig. 6). The ternary complex binds ribosome with GTPBP1 placed on the 40S shoulder away from the GTPase activating SRL. This position allows the anticodon of the tRNA to interact with the mRNA codon in the A site. Distinct structural features of GTPBP1—the N-terminal extension and the shoulder-interacting H-loop in place of canonical α2 helix—underlie interactions of GTPBP1 with the tRNA and the ribosome that are different from those of eEF1A, appearing to favor conformations with the G-domain posed away from the SRL (as in Structures IIa-d). If the anticodon matches the codon, the decoding center nucleotides G626, A1824 and A1825 stabilize the codon-anticodon helix, inducing a “closure” of the 40S subunit shoulder domain (as in Structures IIa-d). The 40S shoulder movement is less extensive than that found in bacterial ribosomes29,36 and is similar to that observed in eEF1A-bound complexes28,34. GTPase activation (Structure III) appears to require the additional slight (<1°) forward rotation of the small ribosomal subunit, which is substantially smaller than the ~8° rotation observed during tRNA-mRNA translocation64. Nevertheless, the shoulder movement and the subunit rotation appear sufficient to dock GTPBP1 at the SRL to initiate GTP hydrolysis (as in Structure III). The elbow of tRNA, which initially docks at the L11 stalk (a.k.a. the H43/H44 stalk), detaches from the latter as the GTPBP1•aa-tRNA complex moves toward the SRL. Upon GTP hydrolysis and γ-phosphate release, GTPBP1 undocks from the SRL but preserves its overall conformation and remains bound to the 40S subunit and to the tRNA, which reestablishes its contact with the L11 stalk (Structure IV).
Cryo-EM maps reveal that GTP hydrolysis does not quickly lead to the rearrangement of switch 1 of GTPBP1, which is required for the release of aa-tRNA from the protein and aa-tRNA accommodation into the peptidyl-transferase center. Moreover, interdomain rearrangements of GTPBP1 that would be expected to follow GTP hydrolysis are inhibited at least in part by the N-terminal domain (Fig. 4F, G). These findings provide mechanistic explanations for the previously reported slow peptide bond formation following GTP hydrolysis by GTPBP1 and accelerated peptide bond formation when the N-terminal domain of GTPBP1 is deleted19. Indeed, viral GTPBP1-like proteins that lack an NTD and H-helix-loop exhibit high elongation rates similar to those of eEF1A50. In addition, our structures revealed that GTPBP1 interactions with the body of tRNA are tighter than those of eEF1A, perhaps also contributing to tRNA retention and accounting for the ability of GTPBP1 to deliver deacylated tRNA to the A site19.
GTPBP1•GDP must eventually rearrange to release aa-tRNA for accommodation and dissociate from the ribosome (Fig. 6). Indeed, the fraction of accommodated tRNA is higher when GTPBP1 complexes were assembled with GTP than with GDPCP (Fig. 4D). The NTD is connected to the G-domain through a linker coordinating K+-binding pocket (Fig. 2D). This suggests a possibility of the regulation of the NTD association through cation binding65. It remains to be seen how the inhibitory N-terminal domain is housed on the rearranged GTBPB1•GDP. Our alignments suggest that due to steric clashes with the N-terminal domain, the relative positions of the GTPase domain and β-barrel domains likely differ from those in eEF1A.
Because the accuracy of aa-tRNA delivery to the ribosome depends on GTP hydrolysis and aa-tRNA release from a dissociating elongation factor, as discussed in the previous section and in refs. 36,58,59,60,66, the delayed EF dissociation appears to underlie the more precise aa-tRNA delivery by GTPBP1 than by eEF1A (Figs. 5 and 6). GTPBP1 interactions with tRNA are not nucleotide-sequence specific (Fig. 2F, G), in keeping with the delayed delivery of different tRNA species in this work (Fig. 5) and in ref. 19, suggesting that accurate decoding is widespread among codons. GTPBP1 might be required for accurate protein expression at certain stages of cell development or in different cell types, consistent with the differential expression of GTPBP1 in mammalian tissues2,7,14. GTPBP1 is unlikely to compete with the large excess of eEF1A (~100x according to PaxDb67,68) under normal growth conditions, but its elongation function may become more significant under stress or other conditions that impair eEF1A activity, such as eEF1A phosphorylation at Ser300, which inhibits binding of eEF1A to aa-tRNA69, or phosphorylation of eEF1Bδ at Ser133, which slows nucleotide exchange on eEF1A70.
It remains to be determined how the molecular mechanism of GTPBP1-mediated translation defines its cellular roles critical for neurodevelopment and stress responses12,14,15. Although GTPBP1 was implicated in the recruitment of the exosome to promote mRNA degradation that could help resolve stalled ribosomes19,20,71, a strong correlation between mRNA abundance and ribosome pausing in the murine Gtpbp1−/− cerebellum was not established14. Alternatively, GTPBP1 could temporarily bind elongating ribosomes with an unclaimed (e.g., “rare”) codon to prevent the formation of ribosomes with a vacant A site that may induce ribosome disassembly72,73 and/or the integrated stress response14. GTPBP1-bound ribosomes might also correspond to a temporary stalling state, e.g., to enable the dendritic transport of polysomes, in keeping with their stalled or slowed translation74,75. The prolonged association of GTPBP1 with ribosome may attract additional downstream factors. Future studies will elucidate the possible interactions of GTPBP1 with other cellular complexes.
Our structures also bring insights into GTPBP2, a related protein, which does not bind 80S ribosome under conditions sufficient for GTPBP1 binding19. Sequence alignment (Supplementary Fig. 3b) and AlphaFold prediction of GTPBP2’s structure indicate that GTPBP2 features a substantially shorter H-loop than that of GTPBP1 (Fig. 7A, B). Our modeling of GTPBP2 suggests that the protein would be unable to form H-loop-mediated interactions with the 40S subunit. Furthermore, in place of GTPBP1, positively charged and neutral residues that interact with the small ribosomal subunit, GTPBP2 features numerous negatively charged residues (e.g., at positions 421, 481, 483) that disfavor interactions with the rRNA backbone (Fig. 7C, D). Curiously, GTPBP2 retains the N-domain resembling that of GTPBP1, suggesting that the delayed release of its substrate (e.g., tRNA) after GTP-hydrolysis may play a role in GTPBP2 cellular function. Two mutations in this domain of human GTPBP2 (Lys125Arg and Leu93Pro) cause neurodevelopmental impairment and severe intellectual disability12, in keeping with its functional importance. Deficiencies in GTPBP1 and GTPBP2 similarly affect the CNS of human and tRNA-deficient mice12,13,14. However, unlike GTPBP1, GTPBP2 does not seem to interact with the exosome19,20; GTPBP2 and GTPBP1 do not compensate for each other14; and there is no additive phenotype in the brains of mice lacking both GTPBP1 and GTPBP214. Thus, GTPBP1 and GTPBP2 are thought to function at different steps in the same pathway14, which would be in line with GTPBP2 performing an enzymatic activity that is distinct from that of GTPBP1.
A Overlay of a predicted AlphaFold model of GTPBP2 (Q9BX10) on GTPBP1(Structure III); the long unstructured tails of both proteins are hidden. B H-loop of GTPBP2 is shorter than that of GTPBP1. C, D GTPBP2 bears negatively charged substitutions at points where GTPBP1 contacts the ribosome.
In summary, our results provide structural insights into GTPBP1-driven delivery of tRNA to the ribosome, highlighting the similarities with and differences from canonical elongation factors. Similar to studies of other GTPases76,77,78, structures of GTPBP1 may help to develop potential therapies for diseases caused by dysregulation of GTPBP1 function, such as in cancers with altered GTPBP1 expression79. Structure-based development of GTPBP1 GTPase agonists80 may help to improve the neurodegenerative conditions caused by GTPBP1 deficiency12.
Methods
Plasmids
Vectors for bacterial expression of His6-tagged eIF1, eIF1A81, eIF4A, eIF4B82, eIF4G736-1600 and eIF4G736-111583, eIF584, Escherichia coli methionyl-tRNA synthetase85 and GTPBP119 have been described. Transcription vectors for MF-STOP and MS(UCU)HL-STOP mRNAs containing a 5′UTR derived from the native β-globin mRNA, a short open reading frame (ORF), and a 3′UTR composed of nt 16-121 of β-globin ORF have been described19. The transcription vector for MS(UCC)HL-STOP mRNA had the same composition and was made by GeneWiz, South Plainfield, NJ. The mRNA sequences are following; MF-STOP mRNA (shown as DNA sequence): GGCAACAACAACAACACTTGCTTTTGACACAACTGTGTTTACTTGCAATCCCCCAAAACAGACAGAATGTTCTAATGTGTGGAGAAGTCTGCGGTCACTGCCCTGTGGGGAAGGTGAATGTGGAAGAAGTTGGTGGTGAGGACCTGTCCTCTGCAAATGCTGTTATGAACAATCCTAAGGAAGCT; MS(UCU)HL-STOP mRNA (shown as DNA sequence):GGCAACAACAACAACACTTGCTTTTGACACAACTGTGTTTACTTGCAATCCCCCAAAACAGACAGAATGTCTCACCTTTAATGTGTGGAGAAGTCTGCGGTCACTGCCCTGTGGGGAAGGTGAATGTGGAAGAAGTTGGTGGTGAGGACCTGTCCTCTGCAAATGCTGTTATGAACAATCCTAAGGAAGCT; MS(UCC)HL-STOP mRNA (shown as DNA): GGCAACAACAACAACACTTGCTTTTGACACAACTGTGTTTACTTGCAATCCCCCAAAACAGACAGAATGTCCCACCTTTAATGTGTGGAGAAGTCTGCGGTCACTGCCCTGTGGGGAAGGTGAATGTGGAAGAAGTTGGTGGTGAGGACCTGTCCTCTGCAAATGCTGTTATGAACAATCCTAAGGAAGCT. Transcription vectors for tRNAPhe-GAA19, tRNASer-AGA86, and tRNAiMet87 have been described.
mRNA and tRNA preparation
Plasmids for transcription of mRNAs were linearized using HindIII (New England Biolabs, Ipswich, MA), and plasmids for transcription of tRNAiMet, tRNAPhe-GAA, and tRNASer-AGA were linearized using BstNI (New England Biolabs). All RNAs were transcribed using T7 RNA polymerase (Thermo Scientific) and purified by FPLC using a Superdex 75 Increase 10/300 GL column. tRNAPhe-GAA and tRNASer-AGA were aminoacylated using purified native aminoacyl-tRNA synthetases88, while tRNAiMet was aminoacylated using recombinant E. coli methionyl-tRNA synthetase85.
Purification of translation factors, ribosomal subunits, and aminoacyl-tRNA synthetases
Mammalian native 40S and 60S ribosomal subunits, Σ aminoacyl-tRNA synthetases, eIF2, eIF3, eIF5B, eEF1H, and eEF2 were purified from rabbit reticulocyte lysate (RRL) (Green Hectares, Oregon, WI) as described in refs. 88,89. Recombinant His6-tagged eIF1, eIF1A, eIF4A, eIF4B, eIF4G736-1115, eIF4G736-1600, eIF5, and E. coli methionyl tRNA synthetase were expressed in E. coli BL21 (DE3) and purified as described in ref. 88.
Preparation of 80S initiation complexes
80S initiation complexes (ICs) were assembled in vitro using individual purified translation components88. Twelve 200 μl reaction mixtures containing 100 nM 40S subunits, 200 nM MF-STOP mRNA, 270 nM eIF2, 150 nM eIF3, 300 nM eIF1, 300 nM eIF1A, 300 nM eIF4A, 300 nM eIF4B, 150 nM eIF4G736-1600 and 200 nM Met-tRNAiMet were incubated in buffer A (20 mM Tris, 100 mM KCl, 2.5 mM MgCl2, 2 mM DTT, 0.25 mM spermidine, 0.2 mM GTP, 1 mM ATP) for 10 minutes at 37 °C. Then reaction mixtures were supplemented with 50 pmol 60S subunits, 100 pmol eIF5, and 100 pmol eIF5B, and incubation continued for another 10 min at 37 °C. After incubation, reaction mixtures were combined and loaded onto four 10–30% linear sucrose density gradients prepared in buffer B (20 mM Tris, pH 7.5, 100 mM KCl, 2.5 mM MgCl2, 2 mM DTT, and 0.25 mM spermidine) and centrifuged for 1 h 30 minutes at 53,000 rpm at 4 °C in a Beckman SW 55 Ti rotor. Fractions containing 80S ICs were combined, concentrated to 1.3 pmol/μl on Amicon Ultra Ultracel-100K centrifugal filters, transferred to buffer C (20 mM Tris-HCl pH 7.5, 50 mM KCl, 2.5 mM MgCl2, 2 mM DTT, 250 mM sucrose), and stored at −80 °C.
Elongation activity of GTPBP1
To compare the abilities of GTPBP1 and eEF1H to promote elongation on cognate and near-cognate codons, elongation with Ser-tRNASer-AGA was assayed using 80S initiation complexes (ICs) assembled on MS(UCU)HL-STOP or MS(UCC)HL-STOP mRNAs (i.e., containing a cognate or near-cognate codon in the A site, respectively). Elongation reactions were assembled essentially as described in ref. 19. 48S complexes were formed by incubating 25 nM MS(UCU)HL-STOP or MS(UCC)HL-STOP mRNA with 60 nM 40S subunits, 350 nM eIF1, 350 nM eIF1A, 90 nM eIF2, 60 nM eIF3, 300 nM eIF4A, 60 nM eIF4B, 250 nM eIF4G736-1115 and 100 nM Met-tRNAiMet in buffer D (20 mM Tris-HCl pH 7.5, 3.8 mM MgCl2, 100 mM KCl, 0.25 mM spermidine, 2 mM DTT) supplemented with 1 mM ATP, 0.3 mM GTP and 1 U/μl RiboLock RNase inhibitor for 105 minutes at 37 oC. To obtain 80S ICs, reaction mixtures were supplemented with 90 nM of 60S subunits, 200 nM eIF5, and 60 nM eIF5B, and incubation continued for an additional 10 min. After that, MgCl2 was added to achieve indicated concentrations of free [Mg2+]. Elongation was carried out by mixing 80S IC reaction mixtures with 60 nM eEF2, 150 nM eEF1H or GTPBP1, and 300 nM Ser-tRNASer-AGA, after which incubation continued at 37 °C for an additional 20 min. The resulting ribosomal complexes were analyzed by primer extension inhibition88 using AMV reverse transcriptase (Promega, Madison, WI) and a [32P]-labeled primer complementary to nt 149–166 of MS(UCU)HL-STOP and MS(UCC)HL-STOP mRNAs. In all cases, primer extension was done at 20 mM free [Mg2+] to abrogate elongation. cDNA products were resolved in 6% polyacrylamide sequencing gels followed by autoradiography.
Cryo-EM sample preparation
Reaction mixtures containing (final concentrations) 0.56 μM GTPBP1, 1.5 μM in vitro transcribed aminoacylated yeast Phe-tRNAPhe, 1.6 U/μl Ribolock RNAse inhibitor (Thermo Scientific), 0.2 mM GDPCP or GTP, and 1 mM ATP in buffer A were incubated for 5 min at 37 °C to allow GTPBP1•GDPCP/GTP•Phe-tRNAPhe complexes to form. Sucrose-density-gradient-purified 80S ICs and Mg(OAc)2 were then added to the mixture to 0.13 μM and 5 mM, respectively.
For the GDPCP dataset, the sample was incubated for additional 10 min at 37 °C and then 4 μl of it was applied to Quantifoil R2/1 holey-carbon grids (EMSDiasum), which were preliminary glow discharged with 20 mA current with negative polarity for 30 s in a PELCO easiGlow glow discharge unit. A Vitrobot Mark IV (ThermoFisher Scientific) was used to plunge-freeze the grids in liquid-nitrogen-cooled liquid ethane. For the GTP datasets, the reaction mixture was placed on ice, and 4 μl samples were plunge-frozen at 1- and 5-min time points, as described above.
Electron microscopy
Cryo-EM data for 80S•GTPBP1•Phe-tRNAPhe complexes with GDPCP or GTP were collected on a Titan Krios electron microscope (ThermoFisher Scientific) at the UMass Chan Cryo-EM Center, operating at 300 kV and equipped with a Gatan Image Filter (slit width 20 eV) (Gatan Inc.) and a K3 Summit direct electron detector (Gatan Inc.) targeting 0.5–1.5-μm underfocus. SerialEM90 was used to automatically collect 6703 movies (18 frames, 1.6553 e-/Å2 per frame, total dose 29.7955 e-/Å2) for the GDPCP dataset, 14046 movies (20 frames, 1.4877 e-/Å2 per frame, total dose 29.7531 e-/Å2) for the GTP-1-min dataset, and 8277 movies (20 frames, 1.5008 e-/Å2 per frame, total dose 30.0165 e-/Å2) for the GTP-5-min dataset.
Data processing
The movies were aligned during data collection (“on the fly”) using IMOD91 to decompress frames, apply the gain reference, and to correct for image drift and particle damage and bin the super-resolution pixel by 2, yielding 0.83 Å/pixel. The corrected.mrc images were imported into CisTEM92, and images were selected based on the quality of the CTF fits and image appearances (i.e., the absence of grid damage and crystalline ice deposits), yielding 4455, 13721, and 8097 images for the GDPCP, GTP-1-min, and GTP-5-min datasets, respectively. The raw.tif movies corresponding to the selected micrographs were imported into RELION 4.093,94 and were motion-corrected using RELION’s implementation of MOTIONCOR295. The aligned images were used for particle picking in crYOLO96, pretrained on several manually picked images, which yielded 318,381 particles, 1,029,824 particles, and 519,347 particles for the GDPCP, GTP-1-min, and GTP-5-min datasets, respectively. The particles were extracted in RELION 4.0 at 3.32 Å/pixel in the 140 pixel box and 3D refined using the map of the rabbit unrotated ribosome (EMD-923797), low-pass filtered to 30 Å, as the initial reference. The resulting map for the GDPCP dataset contained density for GTPBP1, which allowed for initial placing of GTPBP1 model and creating 3D masks around it. The refined stacks were subjected to maximum-likelihood 3D classifications without additional alignments (into 8 classes, Supplementary Fig. 7). Classes representing damaged particles, free 60S subunits, and rotated ribosomes with hybrid A/P tRNA were discarded. The purified stacks were reextracted at 1.162 Å/pixel, 400-pixel box, and re-refined, followed by cycles of CTF-correction, Bayesian polishing, and 3D-refinement. The shiny stacks were exported to Frealign 9.1198,99 and subjected to 3D classifications without alignments with a 3D mask around GTPBP1. The classes containing GTPBP1 were then exported to cryoSPARC v4.3.0100 and subjected to 3D classifications without alignments with a 3D mask around GTPBP1. The classes representing different conformations of GTPBP1 relative to the ribosome were imported back into RELION 4.0, re-refined, and the resulting maps were used for structure modelling and refinements. Local resolutions of the maps were estimated with RELION 4.0.
To improve local density at the K+-binding pocket and other regions, image stacks were subjected to composite mask refinement101 in RELION 4.0 using a “protein” mask around GTPBP1 and a “micelle” mask around ribosome and GTPBP1, followed by B-factor sharpening (−50 Å2) using bfactor.exe, part of the Frealign v9.11 distribution98,99.
In parallel, the initial stacks refined at 3.32 Å/pixel were exported to Frealign 9.1198,99 and subjected to 3D classification into 8 classes (without alignments) with a generous focus mask around the ribosomal A-site for the quantification of the ribosomal states (e.g., A-site occupancies) in the samples.
The CTH (classification of transferred heterogeneity) approach
We sought to characterize the stochastic movements of the 40S shoulder in 80S ribosomes, but separation of the slightly mobile 40S regions in vacant 80S ribosomes was problematic using standard 3D classifications in RELION 4.0 or Frealign 9.11. To discern small conformational changes, we developed the approach which we term “classification of transferred heterogeneity”, or CTH. It relies on the fact that, unlike a 3D classification, particle refinement with a 3D mask covering the shoulder yielded more resolved maps for this region (Supplementary Fig. 8). Since the alignment of signal within the masked region must “transfer” the local heterogeneity to the rest of ribosome, the heterogeneity at the distant regions must be amplified. Indeed, subsequent classification of the locally refined “pre-delivery” 80S stack, now using a 60S mask, yielded resolved classes with distinct shoulder positions differing by up to 2 Å (Fig. 3F and Supplementary Movie 1). This approach also identified a small fraction of the A-site accommodated tRNA complexes with the larger shoulder shift due to the domain closure, which was not identified by canonical shoulder-focused classification due to the small number of particles.
Model building and refinement
The structure of the rabbit 80S complex with eEF1A•GDP•aa-tRNA28 (PDB: 5LZS) was used as a starting model for the 80S•GTPBP1 complexes. The sequences of mRNA, P-site tRNA, and A/T tRNA were changed in Coot102 to match our mRNA, human Met-tRNAiMet (AGCAGAGUGGCGCAGCGGAAGCGUGCUGGGCCCAUAACCCAGAGGUCGAUGGAUCGAAACCAUCCUCUGCUACCA) and yeast Phe-tRNAPhe (GCGGAUUUAGCUCAGUUGGGAGAGCGCCAGACUGAACAUCUGGAGGUCCUGUGUUCGAUCCACAGAAUUCGCACCA) sequences, respectively. AlphaFold103 model of the human GTPBP1 protein (AF-O00178-F1-model_v4104) was used as a starting model for GTPBP1. The structures were assembled in ChimeraX105, modelled and refined in Isolde106, followed by B-factor and global minimization refinements in Phenix107.
GTPBP1 sequence alignments
For GTPBP1 sequence alignments, we used a set of sequences identified previously to include a broad range of organisms1. The obsolete records were updated using BlastP108, and the Ascomycota sequences were excluded as a distant branch of fungi GTPBP1 featuring a different H-loop organization according to AlphaFold104. Given the absence of the extra N-domain in some GTPBP1-like proteins from giant viruses50 (Supplementary Fig. 5), we checked the remaining sequences using the ChimeraX AlphaFold tool and retained those with the extended N-domain in the predicted models. The resulting 31 unique sequences span from the unicellular amoeboid protist Capsaspora owczarzaki to humans. GTPBP2 sequences were retrieved from Uniprot109 using “[the species names from the GTPBP1 sequences list] AND GTPBP2” as a query, which yielded 15 GTPBP2 sequences. GTPBP1 and GTPBP1/2 sequence sets were aligned using MUSCLE110 at the EMBL-EBI server111, and the alignments were visualized in Jalview 2.11.3.3112. The model of GTPBP1 was colored by sequence conservation in ChimeraX using GTPBP1 alignment113.
Miscellaneous
The structures were visualized, and figure panels were prepared using ChimeraX105. Maximal RMSDs in the ensemble of structures were calculated using a custom Bash script (see Code availability), and ChimeraX was used to visualize the RMSDs as worm or cylinder diagrams. DALI server33 was used to find structural homologs of the GTPBP1’s N-domain. Figures were assembled in Adobe Illustrator. The video was prepared using ChimeraX and Adobe After Effects. Graphs were prepared in Microsoft Excel.
Data availability
The structural models/Cryo-EM maps obtained in this study have been deposited in the RCSB Protein Data Bank / Electron Microscopy Data Bank under the following accession codes: 9YPZ/ EMD-73315 (Structure 1a), 9YQ0/ EMD-73316 (Structure 1b), 9YQ1/ EMD-73317 (Structure 1c), 9YPO/ EMD-73302 (Structure IIa), 9YPS/ EMD-73307 (Structure IIb), 9YPT/ EMD-73308 (Structure IIc), 9YPV/ EMD-73310 (Structure IId), 9YPG/ EMD-73297 (Structure III, including map refined with composite mask around GTPBP1), 9YPW/ EMD-73311 (Structure IV, including map refined with composite mask around GTPBP1 and IV-like map at 5 min time point), 9YPY/ EMD-73314 (Structure V). The coordinate files used in this study are available at the PDB: 5LZS, and at AlphaFold Protein Structure Database: https://alphafold.ebi.ac.uk/entry/O00178. Source data are provided with this paper.
Code availability
Computer code is available from GitHub under https://github.com/DenisSusorov/.
References
Atkinson, C. C. The evolutionary and functional diversity of classical and lesser-known cytoplasmic and organellar translational GTPases across the tree of life. BMC Genom. 16, 78 (2015).
Senju, S. & Nishimura, Y. Identification of human and mouse GP-1, a putative member of a novel G-protein family. Biochem. Biophys. Res. Commun. 231, 360–364 (1997).
Kudo, H., Senju, S., Mitsuya, H. & Nishimura, Y. Mouse and human GTPBP2, newly identified members of the GP-1 family of GTPase. Biochem. Biophys. Res. Commun. 272, 456–465 (2000).
Watanabe, M. et al. Cloning, expression analysis, and chromosomal mapping of GTPBP2, a novel member of the G protein family. Gene 256, 51–58 (2000).
Girardot, F., Monnier, V. & Tricoire, H. Genome wide analysis of common and specific stress responses in adult Drosophila melanogaster. BMC Genom. 5, 74 (2004).
Senju, S., Iyama, K., Kudo, H., Aizawa, S. & Nishimura, Y. Immunocytochemical analyses and targeted gene disruption of GTPBP1. Mol. Cell. Biol. 20, 6195–6200 (2000).
Anisimova, A. S. et al. Human tissues exhibit diverse composition of translation machinery. Int. J. Mol. Sci. 24, 8361 (2023).
Jaberi, E. et al. Identification of mutation in GTPBP2 in patients of a family with neurodegeneration accompanied by iron deposition in the brain. Neurobiol. Aging 38, 216.e11–216.e18 (2016).
Bertoli-Avella, A. M. et al. Biallelic inactivating variants in the GTPBP2 gene cause a neurodevelopmental disorder with severe intellectual disability. Eur. J. Hum. Genet. 26, 592–598 (2018).
Carter, M. T. et al. Clinical delineation of GTPBP2-associated neuro-ectodermal syndrome: report of two new families and review of the literature. Clin. Genet. 95, 601–606 (2019).
Manoochehri, J. et al. Jaberi-Elahi syndrome: exploring a novel GTPBP2 mutation and a literature review. Eur. J. Med. Genet. 70, 104953 (2024).
Salpietro, V. et al. Bi-allelic genetic variants in the translational GTPases GTPBP1 and GTPBP2 cause a distinct identical neurodevelopmental syndrome. Am. J. Hum. Genet. 111, 200–210 (2024).
Ishimura, R. et al. Ribosome stalling induced by mutation of a CNS-specific tRNA causes neurodegeneration. Science 345, 455–459 (2014).
Terrey, M. et al. GTPBP1 resolves paused ribosomes to maintain neuronal homeostasis. Elife 9, 1–22 (2020).
Lo, Y. H. et al. GTP-binding protein 1-Like (GTPBP1l) regulates vascular patterning during zebrafish development. Biomedicines 10, 3208 (2022).
Fedotov, S. A. et al. Overexpression of isoform B of Dgp-1 gene enhances locomotor activity in senescent Drosophila males and under heat stress. J. Comp. Physiol. A Neuroethol. Sens. Neural Behav. Physiol. 205, 897–910 (2019).
Fedotov, S. A. et al. The effect of neurospecific knockdown of candidate genes for locomotor behavior and sound production in Drosophila melanogaster. Fly 8, 176–187 (2014).
Walkinshaw, E. et al. Identification of genes that promote or inhibit olfactory memory formation in Drosophila. Genetics 199, 1173–1182 (2015).
Zinoviev, A. et al. Functions of unconventional mammalian translational GTPases GTPBP1 and GTPBP2. Genes Dev. 32, 1226–1241 (2018).
Woo, K. et al. Modulation of exosome-mediated mRNA turnover by interaction of GTP-binding protein 1 (GTPBP1) with its target mRNAs. FASEB J. 25, 2757–2769 (2011).
Telonis-Scott, M., van Heerwaarden, B., Johnson, T. K., Hoffmann, A. A. & Sgrò, C. M. New levels of transcriptome complexity at upper thermal limits in wild Drosophila revealed by exon expression analysis. Genetics 195, 809–830 (2013).
Gruenewald, C., Botella, J. A., Bayersdorfer, F., Navarro, J. A. & Schneuwly, S. Hyperoxia-induced neurodegeneration as a tool to identify neuroprotective genes in Drosophila melanogaster. Free Radic. Biol. Med. 46, 1668–1676 (2009).
Popovic, R. et al. Combined transcriptomic and proteomic analysis of perk toxicity pathways. Int. J. Mol. Sci. 22, 4598 (2021).
Greene, J. C., Whitworth, A. J., Andrews, L. A., Parker, T. J. & Pallanck, L. J. Genetic and genomic studies of Drosophila parkin mutants implicate oxidative stress and innate immune responses in pathogenesis. Hum. Mol. Genet. 14, 799–811 (2005).
Korostelev, A. A. The structural dynamics of translation. Annu. Rev. Biochem. 91, 245–267 (2022).
Rodnina, M. V. Decoding and Recoding of mRNA Sequences by the Ribosome. Annu. Rev. Biophys. 52, 161–182 (2023).
Prabhakar, A., Puglisi, E. V. & Puglisi, J. D. Single-molecule fluorescence applied to translation. Cold Spring Harb. Perspect. Biol. 11, a032714 (2019).
Shao, S. et al. Decoding mammalian ribosome-mRNA states by translational GTPase complexes. Cell 167, 1229–1240.e15 (2016).
Loveland, A. B., Demo, G., Grigorieff, N. & Korostelev, A. A. Ensemble cryo-EM elucidates the mechanism of translation fidelity. Nature 546, 113–117 (2017).
Voorhees, R. M., Schmeing, T. M., Kelley, A. C. & Ramakrishnan, V. The mechanism for activation of GTP hydrolysis on the ribosome. Science 330, 835–838 (2010).
Biou, V., Shu, F. & Ramakrishnan, V. X-ray crystallography shows that translational initiation factor IF3 consists of two compact α/β domains linked by an α-helix. EMBO J. 14, 4056–4064 (1995).
Fletcher, C. M., Pestova, T. V., Hellen, C. U. T. & Wagner, G. Structure and interactions of the translation initiation factor eIF1. EMBO J. 18, 2631–2637 (1999).
Holm, L. DALI and the persistence of protein shape. Protein Sci. 29, 128–140 (2020).
Holm, M. et al. mRNA decoding in human is kinetically and structurally distinct from bacteria. Nature 617, 200–207 (2023).
Gemmer, M. et al. Visualization of translation and protein biogenesis at the ER membrane. Nature 614, 160–167 (2023).
Loveland, A. B., Demo, G. & Korostelev, A. A. Cryo-EM of elongating ribosome with EF-Tu•GTP elucidates tRNA proofreading. Nature 584, 640–645 (2020).
Carbone, C. E. et al. Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP. Nat. Commun. 12, 7236 (2021).
Leipe, D. D., Wolf, Y. I., Koonin, E. V. & Aravind, L. Classification and evolution of P-loop GTPases and related ATPases. J. Mol. Biol. 317, 41–72 (2002).
Zheng, H. et al. CheckMyMetal: a macromolecular metal-binding validation tool. Acta Crystallogr. Sect. D. Struct. Biol. 73, 223–233 (2017).
Rozov, A. et al. Importance of potassium ions for ribosome structure and function revealed by long-wavelength X-ray diffraction. Nat. Commun. 10, 2519 (2019).
Auffinger, P., Ennifar, E. & D’Ascenzo, L. Deflating the RNA Mg2+ bubble: stereochemistry to the rescue!. RNA 27, 243–252 (2021).
Dreher, T. W., Uhlenbeck, O. C. & Browning, K. S. Quantitative assessment of EF-1α · GTP binding to aminoacyl-tRNAs, aminoacyl-viral RNA, and tRNA shows close correspondence to the RNA binding properties of EF-Tu. J. Biol. Chem. 274, 666–672 (1999).
Crepin, T. et al. Mammalian translation elongation factor eEF1A2: X-ray structure and new features of GDP/GTP exchange mechanism in higher eukaryotes. Nucleic Acids Res. 42, 12939–12948 (2014).
Schmeing, T. M. et al. The crystal structure of the ribosome bound to EF-Tu and aminoacyl-tRNA. Science 326, 688–694 (2009).
Fislage, M. et al. Cryo-EM shows stages of initial codon selection on the ribosome by aa-tRNA in ternary complex with GTP and the GTPase-deficient EF-TuH84A. Nucleic Acids Res. 46, 5861–5874 (2018).
Maruyama, K. et al. Switch of the interactions between the ribosomal stalk and EF1A in the GTP- and GDP-bound conformations. Sci. Rep. 9, 14761 (2019).
Ito, K. et al. Molecular insights into the interaction of the ribosomal stalk protein with elongation factor 1. Nucleic Acids Res. 42, 14042–14052 (2014).
Kong, C. et al. Crystal structure and functional analysis of the eukaryotic class II release factor eRF3 from S. pombe. Mol. Cell 14, 233–245 (2004).
Juette, M. F. et al. Didemnin B and ternatin-4 differentially inhibit conformational changes in eEF1A required for aminoacyl-tRNA accommodation into mammalian ribosomes. Elife 11, e81608 (2022).
Zinoviev, A., Kuroha, K., Pestova, T. V. & Hellen, C. U. T. Two classes of EF1-family translational GTPases encoded by giant viruses. Nucleic Acids Res. 47, 5761–5776 (2019).
Zaher, H. S. & Green, R. Fidelity at the molecular level: lessons from protein synthesis. Cell 136, 746–762 (2009).
Ninio, J. Kinetic amplification of enzyme discrimination. Biochimie 57, 587–595 (1975).
Hopefield, J. J. Kinetic proofreading: a new mechanism for reducing errors in biosynthetic processes requiring high specificity. Proc. Natl. Acad. Sci. USA 71, 4135–4139 (1974).
Boeger, H. Kinetic proofreading. Annu. Rev. Biochem. 91, 423–447 (2022).
Gromadski, K. B. & Rodnina, M. V. Kinetic determinants of high-fidelity tRNA discrimination on the ribosome. Mol. Cell 13, 191–200 (2004).
Gavrilova, L. P., Perminova, I. N. & Spirin, A. S. Elongation factor Tu can reduce translation errors in poly(U)-directed cell-free systems. J. Mol. Biol. 149, 69–78 (1981).
Zhang, J., Ieong, K. W., Johansson, M. & Ehrenberg, M. Accuracy of initial codon selection by aminoacyl-tRNAs on the mRNA-programmed bacterial ribosome. Proc. Natl. Acad. Sci. USA 112, 9602–9607 (2015).
Ieong, K. W., Uzun, Ü, Selmer, M. & Ehrenberg, M. Two proofreading steps amplify the accuracy of genetic code translation. Proc. Natl. Acad. Sci. USA 113, 13744–13749 (2016).
Thompson, R. C. & Karim, A. M. The accuracy of protein biosynthesis is limited by its speed: high fidelity selection by ribosomes of aminoacyl-tRNA ternary complexes containing GTP[γS]. Proc. Natl. Acad. Sci. USA 79, 4922–4926 (1982).
Schrader, J. M., Chapman, S. J. & Uhlenbeck, O. C. Tuning the affinity of aminoacyl-tRNA to elongation factor Tu for optimal decoding. Proc. Natl. Acad. Sci. USA.108, 5215–5220 (2011).
Pape, T., Wintermeyer, W. & Rodnina, M. Induced fit in initial selection and proofreading of aminoacyl-tRNA on the ribosome. EMBO J. 18, 3800–3807 (1999).
Negrutskii, B. S. et al. The eEF1 family of mammalian translation elongation factors. BBA Adv. 3, 100067 (2023).
Borg, A. & Ehrenberg, M. Determinants of the rate of mRNA translocation in bacterial protein synthesis. J. Mol. Biol. 427, 1835–1847 (2015).
Flis, J. et al. tRNA translocation by the eukaryotic 80S ribosome and the impact of GTP hydrolysis. Cell Rep. 25, 2676–2688.e7 (2018).
Page, M. J. & Di Cera, E. Role of Na+ and K+ in enzyme function. Physiol. Rev. 86, 1049–1092 (2006).
Morse, J. C. et al. Elongation factor-Tu can repetitively engage aminoacyl-tRNA within the ribosome during the proofreading stage of tRNA selection. Proc. Natl. Acad. Sci. USA 117, 3610–3620 (2020).
Huang, Q., Szklarczyk, D., Wang, M., Simonovic, M. & Mering, C. von. PaxDb 5.0: curated protein quantification data suggests adaptive proteome changes in yeasts. Mol. Cell. Proteom. 22, 100640 (2023).
Wang, M., Herrmann, C. J., Simonovic, M., Szklarczyk, D. & von Mering, C. Version 4.0 of PaxDb: protein abundance data, integrated across model organisms, tissues, and cell-lines. Proteomics 15, 3163–3168 (2015).
Lin, K. W., Yakymovych, I., Jia, M., Yakymovych, M. & Souchelnytskyi, S. Phosphorylation of eEF1A1 at Ser300 by TβR-I results in inhibition of mRNA translation. Curr. Biol. 20, 1615–1625 (2010).
Sivan, G., Aviner, R. & Elroy-Stein, O. Mitotic modulation of translation elongation factor 1 leads to hindered tRNA delivery to ribosomes. J. Biol. Chem. 286, 27927–27935 (2011).
Zinoviev, A., Hellen, C. U. T. & Pestova, T. V. In vitro characterization of the activity of the mammalian RNA exosome on mRNAs in ribosomal translation complexes. Methods Mol. Biol. 2062, 327–354 (2020).
Filbeck, S., Cerullo, F., Pfeffer, S. & Joazeiro, C. A. P. Ribosome-associated quality-control mechanisms from bacteria to humans. Mol. Cell 82, 1451–1466 (2022).
Inada, T. & Beckmann, R. Mechanisms of translation-coupled quality control. J. Mol. Biol. 436, 168496 (2024).
Dalla Costa, I. et al. The functional organization of axonal mRNA transport and translation. Nat. Rev. Neurosci. 22, 77–91 (2021).
Langille, J. J., Ginzberg, K. & Sossin, W. S. Polysomes identified by live imaging of nascent peptides are stalled in hippocampal and cortical neurites. Learn. Mem. 26, 351–362 (2019).
Lu, S., Jang, H., Gu, S., Zhang, J. & Nussinov, R. Drugging Ras GTPase: a comprehensive mechanistic and signaling structural view. Chem. Soc. Rev. 45, 4929–4952 (2016).
Zhang, H., Cai, J., Yu, S., Sun, B. & Zhang, W. Anticancer small-molecule agents targeting eukaryotic elongation factor 1A: state of the art. Int. J. Mol. Sci. 24, 5184 (2023).
Gao, Y., Dickerson, J. B., Guo, F., Zheng, J. & Zheng, Y. Rational design and characterization of a Rac GTPase-specific small molecule inhibitor. Proc. Natl. Acad. Sci. USA 101, 7618–7623 (2004).
Hu, Y. et al. Pan-cancer analysis revealed the significance of the GTPBP family in cancer. Aging 14, 2558–2573 (2022).
Palsuledesai, C. C. et al. Activation of Rho Family GTPases by small molecules. ACS Chem. Biol. 13, 1514–1524 (2018).
Pestova, T. V., Borukhov, S. I. & Hellen, C. U. T. Eukaryotic ribosomes require initiation factors 1 and 1A to locate initiation codons. Nature 394, 854–859 (1998).
Pestova, T. V., Hellen, C. U. T. & Shatsky, I. N. Canonical eukaryotic initiation factors determine initiation of translation by internal ribosomal entry. Mol. Cell. Biol. 16, 6859–6869 (1996).
Lomakin, I. B., Hellen, C. U. T. & Pestova, T. V. Physical association of eukaryotic initiation factor 4G (eIF4G) with eIF4A strongly enhances binding of eIF4G to the internal ribosomal entry site of encephalomyocarditis virus and is required for internal initiation of translation. Mol. Cell. Biol. 20, 6019–6029 (2000).
Pestova, T. V. et al. The joining of ribosomal subunits in eukaryotes requires eIF5B. Nature 403, 332–335 (2000).
Lomakin, I. B., Shirokikh, N. E., Yusupov, M. M., Hellen, C. U. T. & Pestova, T. V. The fidelity of translation initiation: reciprocal activities of elF1, IF3 and YciH. EMBO J. 25, 196–210 (2006).
Zinoviev, A., Hellen, C. U. T. & Pestova, T. V. Multiple mechanisms of reinitiation on bicistronic calicivirus mRNAs. Mol. Cell 57, 1059–1073 (2015).
Pestova, T. & Hellen, C. U. T. Preparation and activity of synthetic unmodified mammalian tRNAiMet in initiation of translation in vitro. RNA 7, 1496–1505 (2001).
Pisarev, A. V., Unbehaun, A., Hellen, C. U. T. & Pestova, T. V. Assembly and analysis of eukaryotic translation initiation complexes. Methods Enzymol. 430, 147–177 (2007).
Pestova, T. V. & Hellen, C. U. T. Translation elongation after assembly of ribosomes on the Cricket paralysis virus internal ribosomal entry site without initiation factors or initiator tRNA. Genes Dev. 17, 181–186 (2003).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Grant, T., Rohou, A. & Grigorieff, N. CisTEM, user-friendly software for single-particle image processing. Elife 7, e35383 (2018).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Brown, A., Baird, M. R., Yip, M. C. J., Murray, J. & Shao, S. Structures of translationally inactive mammalian ribosomes. Elife 7, e40486 (2018).
Grigorieff, N. Frealign: an exploratory tool for single-particle Cryo-EM. Methods Enzymol. 579, 191–226 (2016).
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: Algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Dou, T., Lian, T., Shu, S., He, Y. & Jiang, J. The substrate and inhibitor binding mechanism of polyspecific transporter OAT1 revealed by high-resolution cryo-EM. Nat. Struct. Mol. Biol. 30, 1794–1805 (2023).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D. Biol. Crystallogr. 66, 486–501 (2010).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database in 2024: providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 52, D368–D375 (2024).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. Sect. D. Struct. Biol. 74, 519–530 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: Recent developments in Phenix. Acta Crystallogr. Sect. D. Struct. Biol. 75, 861–877 (2019).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Bateman, A. et al. UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res. 51, D523–D531 (2023).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Madeira, F. et al. The EMBL-EBI Job Dispatcher sequence analysis tools framework in 2024. Nucleic Acids Res. 52, W521–W525 (2024).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview version 2-A multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Pei, J. & Grishin, N. V. A. L. 2CO: calculation of positional conservation in a protein sequence alignment. Bioinformatics 17, 700–712 (2001).
Acknowledgements
We thank Chen Xu, Kangkang Song, and Christna Ouch for the help with data collection at the cryo-EM facility at UMass Chan Medical School; Darryl Conte Jr., and members of the Korostelev laboratory for helpful comments on the manuscript. This study was supported by the US National Institutes of Health R35GM122602 to T.V.P. and R35GM127094 to A.A.K.
Author information
Authors and Affiliations
Contributions
D.S. conceptualized the study, prepared cryo-EM samples, collected and analyzed cryo-EM data, prepared illustrations and video, and wrote the manuscript draft; A.M. and A.Z. purified proteins, prepared tRNA and mRNA, assembled 80S complexes, performed biochemical experiments, and prepared illustrations; D.G. and A.B.L. provided guidance on cryo-EM analyses; T.V.P. and A.A.K. conceptualized the study, supervised data analyses, wrote the manuscript, and secured funding. All authors contributed to data interpretation and provided feedback on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Susorov, D., Miścicka, A., Golovenko, D. et al. Structural mechanism of mRNA decoding by mammalian GTPase GTPBP1. Nat Commun 17, 148 (2026). https://doi.org/10.1038/s41467-025-66833-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66833-2