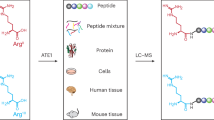
Abstract
N-terminal arginylation (Nt-arginylation) is a multifunctional post-translational modification (PTM) with roles in protein quality control, organelle homeostasis and stress signaling, but its study has been limited by technical challenges. Here, we develop an integrated approach combining N-terminomics with machine learning-based filtering to identify in cellulo Nt-arginylation. Using Arg-starting missed cleavage peptides as proxies for ATE1-mediated arginylation, we train a transfer learning model to predict mass spectra and retention times. By applying the prediction models with an additional statistical filter, we identify 134 Nt-arginylation sites in thapsigargin-treated HeLa cells. Arginylation is enriched in proteins from various organelles, especially at caspase cleavage and signal peptide processing sites. Eight of twelve tested proteins are further validated for their interaction with p62 ZZ domain. Temporal profiling reveals that ATF4 increases early post-stress, followed by arginylation at caspase-3 substrates and ER signal-cleaved proteins. Our approach enables sensitive detection of rare N-terminal modifications, offering potential for biomarker and drug target discovery.
Similar content being viewed by others
Introduction
Protein Nt-arginylation is mediated by ATE1 (Arginyl-tRNA-protein transferase 1)1 for degradation through the ubiquitin-proteasome system as a pivotal component of the Arg/N-degron pathway2. Nt-arginylation not only promotes the degradation of short-lived protein fragments but also triggers autophagic processes in a concentration-dependent manner3,4. Disruption of ATE1 leads to defects in cardiovascular development and angiogenesis5, neurodegenerative disease6,7, and carcinogenesis8. The ability to identify Nt-arginylation is thus a key to understanding the mechanism behind aberrant proteostasis. While affinity-based methods are powerful, direct confirmation relies on mass spectrometry (MS)9,10,11,12,13. Recently, a method to induce Nt-arginylation in vitro and profile the product using liquid chromatography (LC)-MS has been described14.
Positional proteomic techniques such as COFRADIC15 and TAILS16 have been developed to study N-terminal modifications, which would also be suitable for Nt-arginylation15,16,17. However, the identification of Nt-arginylation based solely on mass shift risks generating false positives because of mass ambiguities associated with other unknown N-terminal modifications and amino acid combinations with similar masses18. Recent advancements in machine learning (ML) for predicting MS characteristics, particularly retention time (RT)19 and fragment spectra20, enhance peptide identification rates and enable assessment of the validity of modified peptides. Moreover, the integration of ML models into transformer architectures has improved performance and facilitated transfer learning, reducing the size of required training data significantly through the fine-tuning of pre-existing models21,22,23.
Here, we introduce ML for stringent filtering of N-terminomics MS data generated to profile in cellulo Nt-arginylation. Identifications of Nt-arginylation were assessed by analyzing fragment spectra, RT, and fragment mass errors, comparing them to predicted values obtained using ML-based prediction models. The N-terminomics MS data from ER stress-induced HeLa cells were refined to 134 high-confidence Nt-arginylation sites through false discovery rate (FDR) control and statistical analysis. The Nt-arginylome was further validated using a p62-ZZ domain-derived bait called R-catcher, which has an affinity for arginylated proteins. Temporal dynamics in the Nt-arginylome following ER stress induction were monitored by parallel reaction monitoring MS (PRM-MS). Our approach significantly enhances understanding of Nt-arginylation substrates by reducing common mis-annotations using ML algorithms.
Results
A tandem database search is necessary but insufficient for confident Nt-arginylation calls
To investigate the Nt-arginylome, we designed three treatment conditions: (i) thapsigargin (TG) plus the proteasome inhibitor MG132, to enhance in cellulo Nt-arginylation; (ii) MG132 alone; and (iii) untreated control. HeLa cells under each condition were subjected to N-terminal peptide enrichment using our established iNrich method followed by LC–MS/MS analysis (Fig. 1a)24. To increase proteome coverage, both trypsin and chymotrypsin were used for digestion, resulting in a total of six experimental datasets. The MS data were searched twice sequentially (“tandem database search”) against the human protein database25. The first search was designed to exclude MS spectra matching peptides with N-terminal arginine residues that originate from the protein sequence itself rather than from true post-translational arginylation events (referred to as “Arg-starting peptides” hereafter). Such cases arise when consecutive trypsin cleavage sites (e.g., –XRRX–) are cleaved at the first site but missed at the second, or when a chymotrypsin cleavage site occurs immediately upstream of an arginine. The second search was then performed on the remaining unassigned spectra to identify Nt-arginylation as a genuine post-translational modification (PTM)26.
a Profiling of Nt-arginylome in ER stress-induced HeLa cells. Cells are treated with a proteasome inhibitor (MG132) or MG132 and thapsigargin (MGTG), on which N-terminomics is performed using iNrich method. Mass spectra searched are tried for matching to protein sequences in a two-stage database search method. During the N-terminal peptide enrichment process, peptides with free Nt-amine (α and ε-amine) are labeled with D3-acetyl (D3Ac). b Schematic representation of ML-based filtering of mass spectra for putative Nt-arginylated peptides. Spectra of the Arg-starting peptides derived from missed cleavage of consecutive tryptic sites and their RT were used as training material for the fine-tuning of MS2 prediction model and RT prediction model. Spectra identified as Nt-arginylation in the database search are examined to determine if their fragment spectra or RT align with the predictions made by each model. In the mass error test (MET) filtering step, a null hypothesis that there is no difference in mass error distributions for the two sets of fragment ions, specifically b-ions and y-ions, is examined. The PSMs that have all three ML-based filtering consensuses are further analyzed as high confidence Nt-arginylome.
We identified 392 putative Nt-arginylated peptides corresponding to 1217 PSMs. The majority were derived from trypsin-digested samples, whereas only 8 peptides and 15 PSMs were obtained from chymotrypsin digestion (Supplementary Data 1). The searches found protein N-termini with reproducible numbers across replicates and treatments. By contrast, the putative Nt-arginylated peptides yielded varied outcomes depending on TG treatment, demonstrating that Nt-arginylation is responsive to ER stress conditions and is consistent with its rarity among protein N-termini (Supplementary Fig. 1). Although many Arg-starting peptides were removed in the first round of tandem database search, a considerable fraction remained in the list. Sequence logo analysis of residues surrounding the arginylation site revealed strong enrichment of arginine at P3 to P1 (nth residue in protein sequence before arginylated site) (Supplementary Fig. 2a, b)27. For P1 = R peptides, lysine or arginine was frequently observed at P2, suggesting potential mis-annotation of missed trypsin cleavage products as Nt-arginylated peptides (Supplementary Fig. 2c). This further indicates that FDR filtering alone may be insufficient to completely exclude such PSMs of Arg-starting peptides. Even after excluding 53 known near-isobaric ambiguities (e.g., P1 = R; P2–P1 = GV/VG), arginine remained enriched at P3–P2 (Supplementary Fig. 2d–f)18. Nonetheless, we also observed high levels of aspartate in P4 and P1, reminiscent of the caspase cleavage site DXXD motif, suggesting presence of genuine Nt-arginylation28,29. Our results highlight that defining the Nt-arginylome solely by the mass shift corresponding to Arg can be confounded by mis-annotations arising from uncharacterized modifications, specific amino acid combinations, or combinations thereof18. Manual inspection of individual PSMs remains necessary to identify authentic Nt-arginylated peptides and minimize false positives. However, systematic filtering based on the physicochemical properties of arginylation can substantially improve this process, providing a more reliable framework for large-scale Nt-arginylome studies.
Trainable features of Nt-arginylation: peptides containing protein sequence-derived arginine at the N-terminus
We hypothesized that Arg-starting peptides (Supplementary Fig. 3a) could serve as proxies for understanding the characteristics of Nt-arginylated peptides due to their identical chemistry. In this regard, we evaluated the physicochemical characteristics of the Arg-starting peptides with 17,669 PSMs obtained from a conventional database search (see details in Methods) on the same LC-MS/MS dataset (Supplementary Fig. 3b and Supplementary Data 2). Consistent with previous findings13,18, we observed the following characteristics: (i) high b-ion fragment intensities (average area difference in Arg-starting peptides: 25.7%; non-Arg-starting peptides: 45.8%) (Supplementary Fig. 3c, d)18, ii) a relatively low RT distribution than non-Arg-starting peptides (normalized Δt = 0.186) (Supplementary Fig. 3e). Based on these attributes, a prediction model trained with the mass spectra of Arg-starting peptides is valuable for discovering the bona fide Nt-arginylome. However, constructing such a model would require at least millions of mass spectra, as previously suggested30. To resolve this matter, we leveraged transfer learning strategy with a pre-trained MS2 prediction model, which is part of the recently launched AlphaPeptDeep algorithm, a large language model (LLM) dedicated to proteomics employing transformer layers23. By fine-tuning the LLM via transfer learning using the spectra of Arg-starting peptides, we predicted the MS2 spectra and RT of the putative Nt-arginylated peptides identified from the tandem database search, and compared them to the experimental data. In addition, we employed a statistical method based on the mass errors of MS2 fragment ions to verify the accuracy of identification for Nt-arginylated peptides (Fig. 1b and Supplementary Fig. 3f).
Machine-learning-based MS2 spectrum prediction and filtering
To assess the practicality of improving a pre-trained model via transfer learning with LLM, we fine-tuned the pre-trained tryptic peptide MS2 model of the AlphaPeptDeep Python package. The fine-tuning process utilized all spectra we acquired, both non-acetylated and acetylated (N = 311,547 for a trypsin model and N = 251,969 for a chymotrypsin model), including 17,669 PSMs of Arg-starting peptides. The non-acetylated peptides likely reflect incomplete depletion during the N-terminal enrichment process. We also built additional MS2 models by conventional training from scratch or by transfer learning with the same spectra except for Arg-starting peptides. We then evaluated the performance of each model by calculating Pearson’s correlation coefficient (PCC) between the predicted and the observed fragment intensities (Fig. 2a and Supplementary Data 3) of Arg-starting peptides (N = 9569 in trypsin and N = 8100 in chymotrypsin). From the fine-tuned MS2 prediction model, 87.8% of Arg-starting peptide spectra had a PCC of at least 0.9 (PCC90) while only 56.4% of PCC90 was obtained by the pre-trained MS2 model and 75.9% by “from scratch” model. MS2 prediction performance of the fine-tuned MS2 model was comparable to reported AlphaPeptDeep benchmarks23. The other models revealed a strong predictive accuracy for y-ions, but they did not achieve the same results for b-ions. The significance of including Arg-starting peptide spectra is evident, as the fine-tuned model without these peptides failed to exceed the performance of the pre-trained model and even showed reduced predictability for b-ions. On the other hand, a chymotryptic peptide prediction model using a transfer learning from trypsin MS2 model with chymotryptic peptides showed moderately increased prediction performance as PCC90 increased by 9.1% compared to the model without transfer learning (from scratch model) (Supplementary Fig. 4a, b).
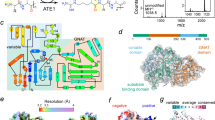
a Assessment of prediction accuracy for MS2 spectra with varying training methods. Plotted are the distribution of Pearson’s correlation coefficient (PCC) of PSMs for Arg-starting peptides according to prediction models (left), percentage of PSMs with PCC ≥ 0.9 (center) and fragment ion species-specific PCC (right). Box plots in this figure show the median (center line), interquartile range (IQR; box limits), and whiskers extending to 1.5 × IQR; outliers beyond this range are shown as individual points. b PCCs obtained by comparing the PSMs searched as Nt-arginylated peptides to the predicted spectra generated using the fine-tuned MS2 prediction model. Bin size: 0.025. c, Boxplots that compare ion species-specific PCC values for b-ion and y-ion. n indicates the number of PSMs for putative Nt-arginylated peptides in each PCC score group of b. d Discriminatory power of similarity measures between true positive and false positive Nt-arginylation PSMs using receiver-operating characteristic (ROC) curves. The PSMs were generated using a decoy database specialized for Nt-arginylation search. PCC Pearson’s correlation coefficient, COS cosine similarity, SPC Spearman’s correlation coefficient, spec_FNR spectral false negative rate, spec_FPR spectral false positive rate, percolator percolator score. e Cumulative false discovery rate by order of percolator score and PCC similarity score. A PSM with a lower index has a higher score. f Distribution of ion species specific PCC values for b-ion and y-ion. n indicates the number of PSMs for putative Nt-arginylated peptides divided into two groups based on the PCC score of (e) corresponding to 1% FDR.
When we compared the measured mass spectra of the well-known Nt-arginylation proteins with the predicted spectra, i.e., CALR|18E, P4HB|18D, and FBLN1|30D (denoted as gene name|arginylation site), the prediction accuracy was significantly high with average PCC of 0.954 ± 0.117 for the CALR|18E (36 PSMs), 0.931 ± 0.081 for the P4HB|18D (37 PSMs), and 0.960 ± 0.038 for the FBLN1|30D (12 PSMs), while the PCC values were 0.891 ± 0.121, 0.915 ± 0.094, and 0.938 ± 0.061, respectively, when predicted with pre-trained model (Supplementary Fig. 4c–e and Supplementary Data 1). The median PCC was 0.878 for all 1217 PSMs including 15 chymotryptic PSMs of putative Nt-arginylated peptides (Supplementary Fig. 4f). We categorized the prediction results into three groups based on the PCC values: high (PCC ≥ 0.9), moderate (0.9 > PCC ≥ 0.6) and low (PCC < 0.6) (Fig. 2b). In the high group, the median PCC of b-ions and y-ions was 0.976 and 0.978, respectively, compared to the moderate group with 0.264 and 0.920 and the low group with 0.051 and 0.510 for b-ions and y-ions, respectively (Fig. 2c). The gathered data illustrate that database search alone relies primarily on y-ions of tryptic peptides, while b-ion signals are required for sufficient confirmation of Nt-arginylation.
Next, we aimed to determine the cut-off for the PCC score for maximizing sensitivity and specificity in identifying authentic Nt-arginylated peptides. In pursuit of this goal, we constructed a decoy database by altering the protein sequences, wherein consecutive arginine residues were consolidated into singular arginine (Supplementary Fig. 5a). The application of the decoy database causes Arg-starting peptides to become Nt-arginylated peptides and thus allows for the distinction of true positives from false positives by determining whether the peptides originated from the altered location of the decoy database or not. With the decoy database, 11,407 PSMs containing Nt-arginylation modification were identified, with 144 of them being deemed false (Supplementary Data 4). There was no significant difference (p-value = 0.94) in the Percolator scores of the database search output between the true and false positives (Supplementary Fig. 5b). In contrast, ROC analysis indicated that PCC values of the MS2 prediction-based rescoring method demonstrated improved performance, achieving an AUROC of 0.624 (Fig. 2d and Supplementary Fig. 5c). Several other metrics reflecting the similarity between predicted and observed spectra also showed higher AUC values than the Percolator score. PCC outperformed the Percolator score in controlling the FDR. The cumulative FDR obtained by sorting the Percolator scores was always higher than the FDR obtained by PCC scores, and the deviation was particularly large in spectra with high scores. (Fig. 2e). We chose a cut-off value of 0.869 for PCC in the trypsin dataset, which achieved a cumulative FDR of 1%. The median PCC for b-ions and y-ions was 0.973 and 0.976, respectively (Fig. 2f). For the chymotrypsin dataset, we set a PCC threshold at 0.9, a decision driven by the limited number of 15 PSMs associated with Nt-arginylated peptides. Using this PCC cut-off, 623 out of 1217 PSMs of Nt-arginylated peptides were accepted.
Retention time modeling using machine learning
Arg-starting peptides exhibited another distinct feature, earlier elution than ordinary tryptic peptides during reversed-phase LC (Supplementary Fig. 3e). For all sequential LC-MS/MS runs performed on fractionated samples originating from a single sample, a total of 12 RT models were generated, one per run23. Comparison of the observed RT and the predicted RT for Arg-starting peptides in the 12 fine-tuned RT models showed high predictive accuracy with R2 values spanning from a low of 0.955 to a high of 0.980 (Fig. 3a and Supplementary Fig. 6). We used these RT models as filters to determine the presence or absence of arginylation modification of the putative Nt-arginylated peptides.
a RT of Arg-starting peptides predicted by RT prediction model versus observed RT. Blue line, fitted linear regression; red dashed line, Δt95% region. b Predicted RT of peptides with Arg replaced by Gly-Val in Arg-starting peptides of (a). Only 41 out of 2567 PSMs remain within the Δt95% RT interval. c Deviation between the predicted and observed RT of Arg-starting peptides in which Arg is replaced by other types of amino acid/dipeptides. The distributions are expressed as cumulative fractions. Dashed line, RT at Δt95%. d Predicted versus observed RT of the putative Nt-arginylated peptides obtained by tandem database search. Red dashed lines indicate the Δt95% region of the RT model. e A histogram of Nt-arginylation peptides as a function of RT deviation between observed and predicted. f Distribution of precursor mass errors that fit (within) and do not fit (outside) the RT model. A two-sample Kolmogorov–Smirnov (K–S) test shows that the two distributions are different. D distance statistic, P P-value. g Average mass errors of b- and y-fragment ions in each PSMs of the putative Nt-arginylated peptides. h A histogram of Nt-arginylation peptides as a function of P-values obtained from Student’s t-test comparing b-ion errors and y-ion errors. The PSMs are divided into two groups based on P-values. Dashed line, median of each group.
We first evaluated the RT models by comparing the RT changes when the Nt-modification of Arg-starting peptides was replaced with GV, a dipeptide that is identical in mass to Nt-arginylation but is not basic18. GV substitution resulted in a rise in RT (Fig. 3b), with most PSMs (98.7%) falling outside Δt95%, a 95% confidence level derived from the linear model (Supplementary Fig. 7). In addition to GV, the RT model reliably distinguished other substitutions, such as SA and DG, except lysine (K) which exhibits basicity like arginine (Fig. 3c). This demonstrates that the RT models developed through transfer learning of Arg-starting peptides can effectively distinguish Nt-arginylation modification from hydrophobic or neutral modifications with similar masses.
Based on the results, we established a filtering criterion at the 95% confidence level (Δt95%) of the linear regression (Supplementary Fig. 8). Nt-arginylated peptides falling within the prediction interval of Arg-starting peptides were classified as positives (Fig. 3d). Among 1217 PSMs, 544 passed the filter. Intriguingly, most PSMs outside the 95% confidence interval exhibited greater-than-predicted increases in RT (Fig. 3e) suggesting that unknown modifications initially mistaken for Nt-arginylation during the database search are less basic than arginine (Supplementary Fig. 3e). Moreover, the PSMs that passed the RT filter had smaller mass deviations than those that did not pass (Fig. 3f), suggesting that the RT prediction module worked appropriately.
Statistical filtering by fragment mass error analysis
We introduced an additional evaluation module that leverages systematic variations in intrinsic mass measurement inaccuracies caused by mis-annotations31. This module is based on the hypothesis that mass spectrometers produce equivalent measurement errors for fragment ions regardless of whether they are b-ions or y-ions. However, when we compared the error distributions between b-ions and y-ions across 1217 putative Nt-arginylated PSMs, the average error for b-ions was −1.37 ± 14.2 mDa, while for y-ions, it was 0.0383 ± 8.93 mDa (Supplementary Fig. 9a, b). This imbalance between b-ion and y-ion errors was consistent at the spectrum level (Fig. 3g). These findings suggest that b-ion mass errors likely stemmed from mis-annotation of the N-terminal modification, which may originate not only from near-isobaric unknown modifications that have not yet been characterized but also from those already cataloged in previous reports (e.g., GV dipeptide), where such unknown modifications were also suggested as potential contributors to these discrepancies18,32. Indeed, it has been proposed that the observed error discrepancy could serve as a confidence metric for assessing the correctness of N-terminal modifications32,33, which we here termed the mass error test (MET).
We used a two-tailed Student’s t-test to compare the b-ion error distribution with the y-ion error distribution and discarded spectra with P-values less than 0.05 (Fig. 3h). MS2 and RT prediction modules could also remove several spectra with heterogeneous mass errors (Supplementary Fig. 9c, d). Nevertheless, we observed significant discrepancies between b- and y-ion mass error distributions in many PSMs having agreement with the MS2 and RT prediction modules (Supplementary Fig. 9e–h). These findings suggest that MET helps eliminate mis-annotations of other modifications with physicochemical properties similar to Nt-arginylation. Using the MET module, we narrowed down the 1217 PSMs to 844.
Performance of integrated machine-learning and statistical filtering
Of 1217 PSMs identified as Nt-arginylated in database search, 394 PSMs were retained after the integrated ML-based filtering (Fig. 4a). The proportion of remaining PSMs that passed filtering varied across the experimental conditions. In DMSO-treated control (MOCK), only 16.1% of PSMs remained (Fig. 4b), while 50.8% for MG132-treated samples (MG132) and 65.3% for MG132- and TG-treated samples (MGTG) were kept. Notably, the absolute number of discarded PSMs was similar across all conditions. These findings indicate that the ML-based filters reflect the intended biological perturbations while reducing false-positive identifications.
a A Venn diagram showing the PSMs of Nt-arginylated peptides that pass each ML-based filtering module. b Number of PSMs for Nt-arginylated peptides that pass all three ML-based filtering modules. c Schematic for annotating the diagnostic ion of Nt-arginylated peptide. d Proportion of the diagnostic ion intensity compared to the total ion intensity. The x-axis represents the PSMs that pass the indicated prediction module. Box plots in this figure show the median (center line), IQR (box limits), and whiskers extending to 1.5 × IQR; outliers beyond this range are shown as individual points. Two-tailed Student’s t-test P-values are shown. e A logo plot showing Nt-arginylation sites that pass (Filtered-in) or fail to pass (Filtered-out) the ML-based filters. The logo for P1’, which is made of only four amino acids, D/E/N/Q, is not displayed. f The pLDDT score distributions determined by AlphaFold for the peptide residue positions of Nt-arginylation sites that pass (left) and fail (right) ML-based filtering. The position value of 0 denotes the residue where Nt-arginylation occurs.
The effectiveness of ML-based filtering was further supported by the frequent detection of fragment ions diagnostic for the modification itself. Previous studies have reported that Nt-acetylation enhances b1 fragmentation34,35. In Arg-starting peptides, the presence of an N-terminal basic residue would increase further the likelihood of b1-ion detections and indeed, the b1-ion has been observed when MS2 spectra of an Arg-starting peptide were compared with those of the same peptides lacking Nt-Arg36. Here, we observed the signature b1-ion of D3-acetylated Arg at 202.138 ± 0.005 m/z (Fig. 4c and Supplementary Fig. 10a–c). Chi-squared test confirmed that the Arg-starting peptides were significantly enriched with b1 ion (1% intensity cutoff, P-value < 2.2 × 10−16) (Supplementary Fig. 10d). The b1-like diagnostic ion is expected at the same m/z in the MS2 spectrum of Nt-arginylated peptide. The 394 PSMs that passed all three filtering modules had higher median relative intensity of diagnostic ion (6.48%) than any other PSM groups that failed at least one of the three modules. The PSMs that failed in all modules had the lowest median, 0% (Fig. 4d). This suggests that all three modules favor PSMs with Arg-starting–like features, and their combination strengthens this enrichment.
Next, we revisited sequence preference of P5–P5’ sites of Nt-arginylation after ML-based filtering. Comparing the logo analysis results before and after filtering, PSMs with arginine at P3 and P2 sites were mostly removed (Fig. 4e). Instead, aspartic acids were observed frequently at P4 and P1 sites which is analogous to the DXXD caspase motif28. In concordance, DAU analysis also showed that arginine is categorized as one of the least preferable residues in P5–P1 sites (Supplementary Fig. 11).
Previous research has demonstrated that the structural context of functionally relevant PTMs is concentrated within the intrinsically disordered region when assessing the predicted protein structure37. Remarkably, structure prediction scores from AlphaFold database on Nt-arginylation sites of 394 PSMs showed low average pLDDT (58.5 ± 23.0) while the sites of PSMs excluded by ML-based filtering exhibited high average pLDDT (75.1 ± 21.0) with significant difference (P = 1.1 × 10−11) (Fig. 4f and Supplementary Data 1). The result is consistent with the notion that PTMs occur in the disordered region of proteins. Notably, the pLDDT scores of the residues in the N-terminal direction from Nt-arginylation sites remained similarly low, which is not observed in the C-terminal direction. Taken together, the comparative analyses show that ML-based filtering is effective for screening mass spectra with Nt-arginylation modification. We note that the inferred consensus is condition-dependent: robust enrichment for protease cleavage motifs is observed in the MG132/MGTG conditions, whereas the MOCK (non-stress) condition yields far fewer Nt-arginylation PSMs and correspondingly weaker motif signal. Thus, the reported consensus/seqlogo primarily reflects stress-enriched arginylation events rather than a universal baseline pattern.
Characterization of ER-stress induced Nt-arginylome
From 394 Nt-arginylation PSMs that passed ML-based filters, 134 Nt-arginylation sites were annotated, showing the highest occurrence in MGTG and the lowest in MOCK (Fig. 5a and Supplementary Data 5). The UPR pathway emerged as the most prominent in the Reactome pathway enrichment analysis using the list of Nt-arginylated proteins (Fig. 5b and Supplementary Data 6). The impact of TG was evident as the gene set enrichment analysis (GSEA) revealed distinct pathways connected to protein metabolism and stress responses in the comparison of Nt-arginylation sites between MGTG and MG132 (Supplementary Fig. 12a). Gene ontology (GO) overrepresentation analysis (ORA) showed that the highest enriched ontologies were GO terms related to cellular component (GOCC), such as focal adhesion, cell-substrate junction, and ER lumen (Supplementary Fig. 12b). A supplementary analysis with GOCC terms reveals that proteins with Nt-arginylation are found in a broad spectrum of subcellular organelles, including ER, nucleus and mitochondria (Supplementary Fig. 12c, d). The protein-protein interaction analysis using STRING database revealed that the arginylome proteins are functionally closely connected with each other even though they are located in various subcellular organelles (Supplementary Fig. 12e). MCL clustering analysis reveals that essential cytoskeletal proteins, such as beta-actin (ACTB), filamin A (FLNA) and moesin (MSN), are at the middle of proteins of ER lumen and cytoplasmic ribonucleoprotein granule (Supplementary Fig. 12f). This suggests a homeostatic role of ATE1, as previously demonstrated by the interaction of Nt-arginylated calreticulin with stress granules38. Of note, the observation of Nt-arginylated proteins in MOCK hints at a mechanism where Nt-arginylation may not induce protein degradation39. These Nt-arginylated proteins also remained stable in MG132 and MGTG (Supplementary Fig. 13).
a A Venn diagram showing the proportion of Nt-arginylation sites that were identified in each experimental condition. b Reactome pathway enrichment analysis of the Nt-arginylated proteins. A one-tailed hypergeometric test was used to obtain P-values. c Number of Nt-arginylation sites with the motif of specified proteases. Cleavage sites were derived from the predictive output of SignalP, TransitP, and Procleave. d Quantitative comparison of Nt-arginylation sites, categorized by the absence or presence of a predicted cleavage site motif, with Δ(Z-score) obtained by comparing MGTG and MG132. Z-scores for each experiment were obtained from the MS1 label-free quantification result of all identified peptides regardless of Nt-arginylation. Box plots show in this figure the median (center line), IQR (box limits), and whiskers extending to 1.5 × IQR; outliers beyond this range are shown as individual points. A two-tailed Student’s t-test P-value are shown. Quantitative distribution of Nt-arginylation sites predicted as cleavage sites for signal peptides (e), caspase-3 substrates (f), and transit peptides (g). Z-scores were obtained as in Fig. 4d. Two-tailed Student’s t-test P-values are shown. h One-carbon metabolism pathway. The proteins marked in red are those where Nt-arginylation sites at the transit peptide cleavage site were discovered. i Spectra mirror plots for the Nt-arginylation sites of Fig. 4h. The observed mass spectrum is shown in the upper plane and the corresponding predicted spectrum is in the bottom plane. Nt-arginylation is indicated by “r” at the beginning of the peptide sequence and the protein name is denoted as (gene name)|(Nt-arginylation site position).
Not only were arginylation sites more abundant under MGTG conditions than under non-stress condition, but they were also markedly enriched in proteolyzed proteins with less ordered structures at the arginylation site4. Sequence logos revealed a pronounced caspase-like DXXD motif under MGTG treatment, whereas MOCK displayed only a modest enrichment of Asp at the P1 position, consistent with basal caspase activity (Supplementary Fig. 14a–c). Using protease-site prediction algorithms40,41,42, we found 56 Nt-arginylation sites that displayed a high likelihood of being protease cleavage sites, including signal peptide, transit peptide, and caspase-3 substrates (Fig. 5c). Logo analysis on these sites indicated that the presence of arginine at the P2 position is strongly associated with signal peptides and transit peptides (Supplementary Fig. 14d, e), largely consistent with a previous report41. By contrast, the result for Nt-arginylation sites lacking known protease motifs revealed no enrichment of arginine at P2 position (Supplementary Fig. 14f, g). In addition to protease-generated N-termini, we observed one candidate isoform-specific Nt-arginylation: SHC1 (P29353) at position 112, which aligns with the annotated start site of the p52SHC (P29353-2) isoform. Because dedicated orthogonal validation for this isoform context has not yet been performed, we report SHC1|112N as a putative isoform-derived Nt-arginylation site pending orthogonal validation. Quantitation of the Nt-arginylome demonstrates increased level of Nt-arginylation at sites cleaved by proteases upon MGTG treatment (Fig. 5d). It was evident that arginylation increased at the N-terminal site newly exposed after the signal peptide was cleaved off or at the N-terminal region newly exposed in the substrate following caspase-3 cleavage (Fig. 5e, f). Mitochondrial proteins containing transit peptides also appeared to increase, although this was not statistically significant (Fig. 5g and Supplementary Fig. 15). We identified six Nt-arginylated mitochondrial proteins, four of which were at the transit peptide cleavage site. Interestingly, two proteins SHMT2|27Q and MTHFD2|36E out of the four are primarily involved in folate pathway, known as one-carbon metabolism (Fig. 5h, i)43. Since these proteins are essential for cancer cell survival and proliferation, the discovery of Nt-arginylation on these proteins may provide a potential means for modulating these therapeutic targets44.
Validation of Nt-arginylome using R-catcher pulldown assay
Nt-arginylation of the identified arginylated proteins was validated using an R-catcher pulldown assay11,26. The R-catcher is derived from the ZZ domain of p62/sequestosome-1, known to have binding affinity toward arginylated protein/peptide. We expected that at least some, if not all, Nt-arginylated proteins would have binding affinity to R-Catcher. Therefore, we arbitrarily selected 12 proteins out of the Nt-arginylome list, cloned and transfected the corresponding genes into HeLa cells. Pull-down assays were performed on the MGTG cells using R-catcher beads (Fig. 6a). As a control, we also used a D129A mutant R-catcher that lacks binding affinity for arginylated peptides. Eight out of the 12 candidate proteins exhibited significant and selective binding to the wild-type R-catcher, similar to positive controls such as HSPA5 and CALR (Fig. 6b, c). We then performed competition assays by pre-charging the R-catcher with an RA dipeptide. All proteins except PDIA3 and CALU lost binding affinity for the R-catcher in the presence of the RA dipeptide but not in the presence of the control AR dipeptide, confirming the specificity of the interaction (Fig. 6d). Nt-arginlyation of these 6 proteins by ATE1 was confirmed by comparing the R-Catcher pulldown assays performed using Ate1 wild-type (Ate1+/+) and knockout (Ate1−/−) mouse embryonic fibroblast (MEF) cells. The interaction was seen only in wild-type MEF cells not in Ate1 KO cells (Fig. 6e). Taken together, our results suggest that arginylated proteins discovered through MS are substrates of ATE1-mediated arginylation. Furthermore, our in vitro R-catcher pull-down assay demonstrates that a significant subset of these proteins can interact with p62, suggesting their potential involvement in autophagy pathways following arginylation.
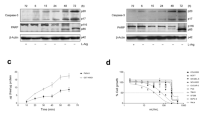
a A schematic representation describing the R-catcher pulldown assay for validation of Nt-arginylation candidate proteins. b R-catcher pulldown assay of Nt-arginylation candidate proteins in HeLa cells treated with MG132 10 μM plus thapsigargin 50 nM. HSPA5 and CALR are positive controls showing that the assay system works properly. P.C. positive control. c List of proteins tested for R-catcher pulldown assay. Position: residue number in the protein sequence where Nt-arginylation occurs; Arginylation site: 10 amino acid sequence surrounding the Nt-arginylation site, spanning 5 residues on each side; Cleavage type: proteases specified by the motif of Nt-arginylation sites. d Dipeptide competition R-catcher pulldown assay to validate the binding is Nt-arginylation-dependent. HeLa cell lysates stimulated with MGTG were incubated with a purified R-catcher in the presence of 25 mM dipeptides, RA or AR. e R-catcher pulldown assay to validate ATE1-dependent arginylation of candidate proteins using Ate1+/+ and Ate1−/− mouse embryonic fibroblasts. P.C. positive control. The bands of immunoblot are representatives of two (e) or three independent experiments (b, d).
Temporal changes of Nt-arginylation in response to ER stress
We next monitored temporal changes of Nt-arginylation in response to ER stress. After treating HeLa cells with MGTG, arginylated proteins and the corresponding unmodified proteins were detected by Western blot and MS at regular intervals (Fig. 7a). We aimed to monitor 21 Nt-arginylation sites with their corresponding unmodified sites primarily associated with UPR-related biological processes, apoptosis, autophagy, and caspase cascade (Supplementary Data 7). In addition, irrespective of arginylation, six peptides derived from six proteins previously established as markers of the UPR and its downstream processes were also included. Although a commercial antibody recognizing the RE and RD motifs is available, the specific proteins targeted in this study have not been validated. To address this limitation, we employed PRM-MS to detect Nt-arginylated proteins, except in cases where antibodies recognizing individual arginylated proteins could be applied. Fourteen synthetic peptides (ISTDs) were also included to modulate RTs.
a A schematic representation of parallel reaction monitoring mass spectrometry (PRM-MS) performed in this study. For each Nt-arginylation target (Nt-arg.), the corresponding unmodified (unmod.) peptides were also monitored. b Observed versus predicted RT of tryptic peptides of HeLa digest. c,d Box plots comparing retention time deviation (c) and PCC similarity value (d) between predicted and observed MS spectra for synthetic ISTD peptides and target peptides. Box plots show in this figure the median (center line), IQR (box limits), and whiskers extending to 1.5 × IQR; outliers beyond this range are shown as individual points. A two-tailed Student’s t-test was used to analyze the results. e,f Log2 fold changes of N-arginylation sites comparing MG132 to MOCK (e) and MGTG to MOCK (f). Two-tailed Student’s t-test P-values are shown. g-i Normalized intensities of target peptides as a function of drug treatment duration. Shown are quantitation results for a tryptic peptide of ATF4 (g), Nt-arginylated peptide starting at the 18th residue of CALR (h), and Nt-arginylated peptide starting at the 18th residue of P4HB (i). Data are presented as mean values ± SD. j Immunoblot analysis of ATF4, activated CASP3, HSPA5, CALR and P4HB during ER stress. R-HSPA5, R-CALR, and R-P4HB represent Nt-arginylated forms of HSPA5, CALR, and P4HB, respectively. k Schematics of UPR leading into caspase-3 activation and cleavage of its substrates. l-n Normalized intensities of substrate peptides at the caspase cleavage sites as a function of drug treatment duration. EIF4B|46D with Nt-arginylation (l), FLNA|1502 N with Nt-arginylation (m), and FLNA|1502N without Nt-arginylation (n). Data are presented as mean values ± SD. o Knockdown of ATE1 in HeLa cells. shC, control shRNA. Knockdown efficiency of ATE1 was confirmed by Western blotting. The bands of immunoblot are representative of biological duplicate. p PRM-MS analysis of mitochondrial transit sites (SSBP1|17E, MTHFD2|36E, and UQCRHL|15D) from subcellularly fractionated samples (cytosol vs. mitochondria). Shown are the PRM intensity ratios of Nt-arginylated peptides to their unmodified counterparts, normalized to the MOCK/Cytosol condition. Two-tailed Student’s t-test P-values are shown for experiments performed in biological triplicate.
In PRM-MS, whole-cell lysate was digested with trypsin and analyzed in a single run of LC-MS acquiring only the MS2 spectra of the target peptides repeatedly in a predefined time duration. Since the N-terminal peptide was not enriched beforehand, the peptide we monitored was not the same as that we found in the profiling experiment: lysine was left unacetylated thus susceptible to tryptic cleavage. For example, if Arg-DEPEWVKTER is the peptide detected in the profiling experiment, Arg-DEPEWVK was attempted to be monitored in the PRM-MS. For this, we had to predict theoretical MS2 spectra and RT for the new peptides with ML. Tryptic digests of HeLa cell lysates were analyzed immediately before each of the six consecutive PRM experiments under identical LC conditions, and the peptide-RT data matrix was fine-tuned by transfer learning. After testing several gradient conditions, we selected a 24% B gradient that allowed even distribution of multiple target peptides across the entire LC running time (Supplementary Fig. 16). The fine-tuned RT prediction model gave R2 of 0.994 and an MAE (mean absolute error) of 0.52 min (Fig. 7b and Supplementary Data 8). The performance of the fine-tuned MS2 model was measured as an average PCC of 0.957 (Supplementary Fig. 17). We further optimized PRM parameters, i.e., acquisition time and isolation width (Supplementary Figs. 18 and 19).
We then performed 42 PRM-MS runs, monitoring the target peptides and ISTDs in duplicated samples of three different conditions, MGTG, MG, and MOCK, collected at seven time points from 0 to 48 h after ER stress activation. Each PRM-MS run was evaluated with the 14 ISTDs included therein. The average ΔRT and MS2 PCC of ISTDs were 0.610 ± 1.537 min and 0.876 ± 0.098, respectively, across all 42 PRM-MS runs (Supplementary Fig. 20a, b). The values for the target peptides were 0.144 ± 1.738 min and 0.854 ± 0.166, respectively (Fig. 7c). No significant difference was observed between ISTDs and the targets in the MS2 similarity score (Fig. 7d). For further analysis, we chose only the MS2 scans with PCC > 0.7 (ref. 45) and utilized the three fragment ions with the highest intensity for quantification (Supplementary Fig. 20c).
Of 21 Nt-arginylation sites and 21 unmodified sites, 15 arginylated and 11 unmodified sites could be detected (Supplementary Fig. 21 and Supplementary Data 9). The PRM-MS result displayed a notable increase of Nt-arginylation in almost all sites upon MG132 or MGTG treatments, with strongest effects observed in MGTG (Fig. 7e, f, Supplementary Figs. 22 and 23). By contrast, the corresponding unmodified sites showed little change under ER stress, remaining near baseline under the treatments compared to MOCK. (Supplementary Fig. 24). In fact, the unmodified counterparts did not show a temporal reduction; conversely, their absolute intensities exhibited a modest increase over time, which trend was observed across all treatment categories, including MOCK (e.g., ERO1A|24E). This pattern suggests that the expression of these proteins may be influenced by factors independent of ER stress, such as starvation during cell culture. Consequently, such baseline changes may obscure the apparent reduction of the unmodified sites by arginylation under ER stress conditions.
Most significantly, the increase in arginylation began later than the increase in the amount of ATF4 protein, a UPR transcription factor46. ATF4 demonstrated its initial rise at 3 h post-treatment (Fig. 7g) while arginylation of CALR|18E and P4HB|18D increased at 12 h post-treatment in MGTG and 18 h in MG132 (Fig. 7h, i). Immunoblot analysis using the antibodies to Nt-arginylated CALR and Nt-arginylated P4HB showed similar temporal changes to those observed by PRM-MS (Fig. 7k and Supplementary Fig. 25). Furthermore, HSPA5, a well-known substrate of ATE1, also showed an increase in Nt-arginylation at a similar time.
Other notable Nt-arginylation sites were those located at predicted caspase cleavage sites. We monitored four sites by PRM-MS, two of which are in EIF4B47 and FLNA48. These two sites (EIF4B|46D and FLNA|1502N) have previously been shown to be cleaved by caspases but to our knowledge, their occurrence following UPR induction have not been reported (Fig. 7l). UPR induction led to a notable rise in Nt-arginylation after 12 h in both MG132 and MGTG treatments relative to MOCK, with levels stabilizing at 12 h in the MGTG condition; this increase preceded the alteration seen at signal or transit peptide cleavage sites (Fig. 7m, n and Supplementary Fig. 24). Immunoblotting using cleaved caspase-3-specific antibody demonstrated that these temporal changes were consistent with caspase-3 activation (Fig. 7k). In contrast, the unmodified form of FNLA|1502N was detected at low levels, making the change unclear. This implies that FLNA is immediately Nt-arginylated once cleaved by caspase-3 (Fig. 7o). The PRM-MS analysis results confirmed that the targets of caspase-3 are indeed subject to modification via Nt-arginylation in the context of the UPR.
Guided by the time-resolved PRM results, we next examined the subcellular localization of Nt-arginylated proteins, with particular attention to mitochondrial proteins. To this end, we combined subcellular fractionation with PRM to quantify Nt-arginylated peptides in each organelle fraction. Specifically, cytosol and mitochondria were fractionated from ATE1 knockdown (shATE1) HeLa cells, and paired peptides (Nt-arginylated and corresponding unmodified) were quantified (Fig. 7p and Supplementary Fig. 26). For SSBP1|17E, MTHFD2|36E, and UQCRHL|15D, the Nt-Arg:unmodified ratio increased upon MGTG treatment and decreased with shATE1, with the strongest effects observed in the mitochondrial fraction (Fig. 7q). These results indicate that the detected transit-site Nt-arginylation events are ATE1-dependent and enriched in mitochondria.
Discussion
Mass-spectrometry workflows can now measure proteomes from individual cells; however, PTM studies remain bottlenecked by low stoichiometry and weak MS signals. Nt-arginylation illustrates these challenges: without a dedicated affinity-based purification method, database searches of MS data frequently miscall Arg-starting peptides from missed trypsin cleavage as bona fide sites. As a result, only ~32% of initial PSMs survive rigorous scrutiny by our ML-based filters, which preserve specificity yet enhance overall sensitivity49. Previous studies addressed such false positives through extensive manual validation, including mass ambiguity tables and filtering of ambiguous PTMs and missed cleavages13,18. While effective, these approaches were highly labor-intensive. In the present study, we aimed to automate these steps by incorporating ML-based RT and MS2 prediction modules, enabling accurate large-scale analysis with reduced effort. Although manual validation will remain necessary until these algorithms are fully benchmarked, our results demonstrate that key diagnostic features—such as prominent b ions, RT shifts, and mass ambiguity checks—can be reliably captured in an automated framework. Thus, our pipeline should be viewed as a complement to expert curation, reducing the burden of large-scale PTM analysis while maintaining confidence in site identification.
Our study focused on the identification of Nt-arginylation sites, which exhibit a significant inclination towards positive charges, resulting in a higher likelihood of ionization and detection during LC-MS analysis. These characteristics, when utilized alongside N-terminal peptide enrichment, have the potential to generate considerable synergistic benefits for constructing ML-based filtering modules. The N-terminomics method we applied blocks primary amines, forcing peptides to fragment into mostly y‑ion series, whereas Nt‑arginylated peptides yield pronounced b‑ions and elute earlier in LC. These contrasting features enhance the performance of RT and MS2 prediction models, supplying orthogonal evidence to cull false positives. Indeed, many peptides that were searched as Nt-arginylated peptides but discordant to the RT model tended to have positive RT bias, suggesting a mis-annotation from more hydrophobic modifications or amino acid combinations.
Our Nt-arginylome defines stress-responsive ATE1 targets, including protease-generated neo-termini such as caspase cleaves. In particular, some hits are mitochondrial enzymes involved in folate metabolism, linking Nt-arginylation to UPR-induced metabolic reprogramming that elevates translation of these enzymes50. Why a largely degradative tag marks folate enzymes remains puzzling. ATE1 also mediates autophagic clearance of cytosolic mitochondrial DNA, implying a wider role in eliminating mislocalized mitochondrial proteins; mis-localized folate enzymes are indeed catalytically aberrant51. Based on shATE1 and mitochondrial fractionation PRM analyses, arginylation of the mitochondrial proteins appears to be ATE1-dependent, and the arginylated forms remain localized within mitochondria. Although the precise mechanism is unclear, our results suggest that arginylation likely occurs after mitochondrial import and removal of the transit peptide. Considering that ATE1 has been detected in mitochondria as well as in the cytosol52, this observation is not unexpected. Importantly, this represents a pathway distinct from the canonical Arg/N-degron pathway, in which mistargeted proteins are typically degraded in cytosol. In the case of SSBP1, for example, arginylation has been shown to regulate mitochondrial respiration rather than promote protein degradation14.
The present arginylome does not include arginylation at canonical N-termini generated by Met-aminopeptidases. Interestingly, however, we detected one Nt-arginylation site at the second residue of an N-terminus–truncated isoform of SHC1. At this stage, it remains uncertain whether this modification originates from Met-aminopeptidase processing of the shorter SHC1 isoform or from proteolytic cleavage of the canonical longer isoform. This observation raises the possibility that Nt-arginylation may contribute to isoform-dependent regulation, providing a potential mechanism for functional diversification of SHC1. Nevertheless, additional analyses will be necessary to delineate the precise origin of this modification and to clarify its biological significance.
We benchmarked our stress‑responsive, in cellulo Nt‑arginylome against Lin et al.’s in vitro ATE1 dataset14. Among 229 sites from Lin and 134 from this study, only 10 overlapped, corresponding to 10 common proteins out of 119 and 161, respectively (Supplementary Fig. 27a–c). Five of these shared sites stem from signal-peptide cleavage and four from mitochondrial-transit cleavage, underscoring methodology and biological context‑dependent disparities (Supplementary Fig. 27d). We additionally detected five caspases‑derived Nt‑arginylation sites absent from Lin’s list; their levels rose in MGTG but were nearly undetectable in MOCK cells, highlighting the tight link between in cellulo arginylation and caspase processing (Supplementary Fig. 27e)53,54. Comparing our Nt-arginylation sites to those reported in other previous studies, only four sites were found in common (Supplementary Data 10)11,55,56. The limited overlap likely reflects differences in experimental contexts of earlier studies, where in vitro or cell-free assays were used, compared with in cellulo experiments. In addition, our experimental design introduces a bias toward degradation-associated Nt-arginylation. Although this bias enabled us to sensitively detect rare substrates, the resulting dataset does not fully capture the functional diversity of the arginylome. Taken together, these results suggest that different experimental paradigms reveal complementary subsets of the arginylome and expand the current knowledge base by incorporating stress-responsive substrates.
The temporal quantifications by PRM-MS clarified the dynamics of relative Nt-arginylation under UPR stress. The upregulation of ATF4 (not arginylation) is expected to be the earliest phenotype while the activation of caspase-3 is the latest under sustained presence of stress that could lead to pro-apoptotic events57. Nt-arginylation has a pivotal function over autophagy-apoptosis balance in UPR stress, as explained in studies of Nt-arginylated HSPA5 (R-HSPA5)4,58. Interestingly, we found that caspase-3 activation and hence appearance of its cleaved substrates underwent much earlier inception than the appearance of R-HSPA5. Integrating previous research on the anti-apoptotic role of Nt-arginylation, these early Nt-arginylation of caspase-3 targets might also be anti-apoptotic flux as a result of adapting to UPR stress.
In this study, we introduced ML-based filtering to profile Nt-arginylation, generating a more comprehensive map during the UPR. Although direct enrichment methods for Nt-arginylation are lacking, our N-terminomics approach with stringent filtering offers an effective interim solution. Our strategy may enhance understanding of the role of Nt-arginylation in stress responses such as autophagy and apoptosis.
Methods
Cell culture
HeLa (ATCC, #CCL-2) was grown in DMEM (Gibco, Rockville, MD, USA) medium supplemented with 10% FBS (Gibco) and 1% penicillin/streptomycin (Gibco). The HeLa cells were sourced from ATCC. Cultures were maintained in an atmosphere of 5% CO2 and 95% air in a humidified incubator at 37 °C. Cells were grown to >90% confluence. For ER stress experiments, HeLa cells were treated with 10 µM MG-132 and/or 0.1 µM TG for 24 h. For TG chase experiments, HeLa cells plated in 6-well plates at about 90% confluency were treated with DMSO as control, MG132, or MG132/TG for defined lengths of time (0, 3, 6, 12, 18, 24, and 48 h). Cells were harvested by trypsinization, washed thrice with ice-cold PBS (phosphate-buffered saline, pH 7.4; Gibco), and resuspended in an appropriate lysis buffer.
ATE1 knockdown and mitochondrial fractionation
Knockdown of ATE1 in HeLa cells was achieved by lentiviral shRNA transduction59. HeLa cells were infected with titered virus particles encoding either a control shRNA (shC) or shATE1#69. Thirty-six hours after transduction, the virus-containing medium was replaced with DMEM medium. In this knockdown setting, ER stress induction and proteasome inhibition were maintained for 12 h. Mitochondria of HeLa cells were then isolated using the Mitochondria Isolation Kit for Cultured Cells (Thermo Fisher Scientific, Cat# 89874), according to the manufacturer’s instructions. The purity of the mitochondrial fraction was verified by western blotting with organelle-specific marker proteins; β-tubulin for the cytosolic fraction and cytochrome c for the mitochondrial fraction. All experiments were performed in biological triplicate.
Cell lysis and protein digestion
For the N-terminome experiment by iNrich, cells were lysed in iNrich lysis buffer (0.2 M EPPS, pH 8.0, 6 M guanidine, 20 mM TCEP, 80 mM 2-chloroacetamide) containing 1 × HALT protease inhibitor cocktail (Thermo Scientific). Lysate was boiled for 10 min at 600 rpm and 95 °C, disrupted with ultrasonication (BranSonic 400B), and cleared for 10 min at 10,000 × g and 4 °C. Protein concentrations were determined with Pierce BCA Protein Assay Kit (Thermo Scientific). Proteins were precipitated by adding 8 × volumes of acetone and 1 × volume of methanol to the lysate and incubating overnight at −80 °C. Precipitates were washed twice with methanol and dried briefly.
For PRM-MS and global proteomics, samples were prepared differently. Cells were lysed in 8 M urea in 50 mM Tris-HCl, pH 8.0 containing 1 × HALT protease inhibitor cocktail (Thermo Scientific), and disrupted with BranSonic 400B sonifier. The lysate was cleared for 10 min at 10,000 × g and 4 °C. Proteins in the lysate were reduced (5 mM DTT, 45 min at 25 °C and 600 rpm), alkylated (20 mM 2-chloroacetamide, 45 min at 25 °C and 600 rpm), and then diluted to bring the urea concentration to <0.8 M using 50 mM Tris-HCl, pH 8.0. Digestion was performed by adding trypsin (Promega, 1:50 enzyme-to-substrate ratio) and incubating overnight at 25 °C and 600 rpm. Digests were acidified to pH <3 by addition of trifluoroacetic acid (TFA) to 0.5% and were desalted using HLB solid-phase extraction (SPE) cartridges (Waters; wash solvent: 0.1% TFA; elution solvent: 0.1% FA in 50 % acetonitrile (ACN)). Eluates were dried by vacuum centrifugation and stored at −20 °C.
Enrichment of N-terminal peptides
The precipitated protein sample (1 mg) was reconstituted to 4 mg/mL in 0.25 mL reaction buffer (6 M guanidine in 0.2 M EPPS, pH 8.0). Enrichment of N-terminal peptides was carried out using the iNrich strategy24, involving D3-acetylation of protein amines, tryptic digestion, and negative selection of internal peptides with NHS-activated agarose resin. Briefly, proteins in the sample were labeled with 200 mM D6-acetic anhydride and 200 mM pyridine for 2 h at 25 °C with end-over-end rotation. Labeled proteins were digested with trypsin or chymotrypsin (Promega, 1:50 enzyme-to-substrate ratio) overnight at 25 °C with end-over-end rotation. The peptides were loaded onto the HLB SPE column (Waters). Depletion of internal peptides was performed by adding 330 mg of NHS-activated agarose dry resin and incubating for 2 h at 25 °C with end-over-end rotation. The unbound N-terminal peptides were transferred to the stationary phase of the SPE column by drawing under a controlled vacuum. SPE-bound peptides were washed for 20 mL of 0.1% TFA and were eluted using 1 mL of 0.1% FA in 50% ACN. N-terminal peptides were dried by vacuum centrifugation and stored at −20 °C.
Peptide fractionation by basic reversed-phase liquid chromatography (bRPLC)
For basic reversed-phase liquid chromatography (bRPLC) of N-terminal peptide samples, 100 μg peptides were reconstituted in bRPLC solvent A (10 mM ammonium formate, pH 10) and loaded onto an XBridge BEH C18 RPLC column, 130 Å, 3.5 μm (4.6 × 250 mm) and coupled to a 1290 UHPLC system (Agilent). Samples were washed using solvent A for 10 min at 0.5 mL/min and subsequently eluted applying a two-step gradient from 0 to 40% bRPLC solvent B (10 mM ammonium formate, pH 10 in 90% ACN) in 38.5 min, to 70% B in 14 min, and holding at 70% B for 10 min. A total of 168 fractions (0.5 min each) were collected, and then every 12th fraction was pooled to create 12 fractions. In total, 144 pooled fractions were prepared, corresponding to three treatment conditions, two protease types, and biological duplicates (12 × 3 × 2 × 2 = 144). The pooled fractions were dried and stored at −20 °C until LC-MS analysis.
LC-MS/MS of N-terminal peptide samples
LC-MS measurements of N-terminal peptide samples were performed with an Ultimate 3000 RSLCnano system coupled to a Q-Exactive mass spectrometer (Thermo Fisher Scientific). bRPLC fractionated peptide samples were reconstituted in 5 μL of 0.1% FA in 2% ACN. 2 µL of samples were injected onto a PepMap 100 trap column (75 µm × 20 mm, Thermo Fisher Scientific), washed with 0.1% FA in 2% ACN for 10 min at a flow rate of 5 µL/min and subsequently transferred to an EASY-Spray PepMap RSLC, 2 µm analytical column (75 µm × 500 mm, Thermo Fisher Scientific). Peptides were separated at 300 nL/min using a 110 min linear gradient from 2.5 to 37.5% LC solvent B (0.1% FA in 80% ACN) in LC solvent A (0.1% FA). MS1 spectra were recorded in the Orbitrap from 400 to 1800 m/z at a resolution of 70,000 and using an automatic gain control (AGC) target value of 1e6 charges and a maximum injection time (maxIT) of 30 ms. Up to 12 of the most abundant precursors (topN) were selected for HCD fragmentation at 27% normalized collision energy (NCE). MS2 spectra were acquired at 17,500 resolutions using an isolation window of 2.0 m/z, an AGC target value of 5e4 charges, and a maxIT of 120 ms. The dynamic exclusion was set to 30 s.
Protein sequence databases
A UniProt human reference protein database (Release 2023_02) with common contaminants was used throughout the study. We also constructed specialized decoy databases from this database for FDR estimation of Nt-arginylation search. In case that trypsin was used for protein digestion, any consecutive arginine residues in the original database were consolidated in a single arginine. In our experimental workflow, lysine was modified before digestion and therefore, the residue was left untouched in the database. In the case of chymotrypsin, any arginine (or arginines) that immediately follows phenylalanine, leucine, methionine, tryptophan, or tyrosine was removed.
Database search of mass spectral data
Proteome Discoverer v2.4 with its built-in search engine SequestHT was used to identify and quantify Nt-arginylated peptides. Fragment mass spectra were searched against the protein database in two stages (“tandem database search”). Mass spectra that failed to pass Percolator validation in the first stage search were collected and used as input data for the second stage search. Search parameters for the first and the second searches were identical except for the types of modifications included. Parameters for both stages were ±10 ppm for precursor tolerance, ±0.05 Da for fragment ion tolerance, up to 2 missed cleavages for the trypsin dataset, and up to 5 missed cleavages for chymotrypsin dataset, minimum peptide length of six amino acids, fixed modification of carbamidomethylation of cysteine and D3-acetylation of lysine, variable modification of methionine oxidation, and cleavage specificity at carboxy-terminal end. Parameters included only in the first stage were N-terminal acetylation and N-terminal D3-acetylation as variable modifications. Parameters for the second stage search were pyro-glutamation at N-terminal glutamate, D3-acetyl-arginylation (+201.1305 Da) at N-terminal aspartate and glutamate, and D3-acetyl-arginylation-deamidation (+202.1145 Da) at N-terminal asparagine and glutamine as variable modification. Minora feature detector module was used for LFQ. Parameters for consensus workflow regarding LFQ: Precursor abundance, area; internal normalization for experimental bias correction, and total peptide amount. For mass spectra of global proteomics, parameters of D3-acetylation modifications were removed. Unless stated otherwise, Proteome Discoverer’s default parameters were applied.
When collecting MS2 spectra of Arg-starting peptides resulting from missed cleavage and using decoy databases to estimate FDR of Nt-arginylation modification, we performed database search in a single step (“conventional database search”). The same parameters as for tandem database search were applied except the following: cleavage specificity to both ends; N-terminal acetylation, N-terminal D3-acetylation, and pyro-glutamylation at N-terminal glutamate as variable modifications.
Construction of MS2 prediction models
AlphaPeptDeep v1.1.5 was used to build MS2 prediction models from the training dataset containing PSMs processed with Proteome Discoverer. Four fragment types (b+, b++, y+ and y++) were used to train and predict. Unless stated otherwise, AlphaPeptDeep default parameters were applied. To build a “from scratch” model, the pDeep model with ModelMS2Transformer was used. The training parameters were: epoch=100 and batch_size = 100. To build a fine-tuned model, a pre-trained model (“generic”) was used with the “train_ms2_model” function. The training parameter was: epoch=50.
Spectral similarity scoring
Similarities between two spectra (for example, observed vs. predicted spectra) were determined by calculating PCC, cosine similarity (COS), Spearman’s correlation coefficient (SPC), spectral FPR, and spectral FNR. These similarity scores were computed using Python (v3.9.18, function calc_ms2_similarity) or R (v4.4.1, function cor). Spectral FPR and spectral FNR for each comparison were calculated as:
where false positive is a fragment ion appearing in the observed spectrum but not in the predicted spectrum, false negative is that appearing in the predicted spectrum but not in the observed spectrum, and true positive is that appearing in both spectra.
FDR estimation of MS2 prediction model for detecting Nt-arginylation
FDR for detecting Nt-arginylation was estimated based on the fact that Arg-starting peptide is searched as a missed cleavage peptide in the original database, but as Nt-arginylated peptide without the first residue Arg in the decoy database. Any PSMs found in this way were treated as true positives. Nt-arginylation PSMs that mapped to sequences other than those immediately following the modified sequence were treated as false positives. By sorting the Nt-arginylated PSMs in descending order of a specific score and applying an arbitrary threshold value, we could calculate the FDR. Conversely, the threshold was set to satisfy an arbitrary FDR value. In most cases, FDR was set to be less than 0.01.
Construction of RT prediction models
AlphaPeptDeep v1.1.5 was used to build RT prediction models from the training dataset containing PSMs processed with Proteome Discoverer. PSMs from each LC-run were used separately for the training, taking into account the degradation of analytical columns with use. The observed RTs were normalized by dividing by the time length of the LC gradient. Fine-tuning of a pre-trained model was performed with the same parameters as for MS2 except using train_rt_model function instead of the train_ms2_model function. We then collected pairs of RT predicted by the fine-tuned RT prediction model versus actual measured RT, performed linear regression, and calculated the RT deviation that included 95% of all data points (Δt95%) by using R (function lm and metrics).
The fine-tuned RT prediction model thus constructed was used in the next step to predict the RT of PSM for Nt-arginylation in the search result. The RT prediction was performed in Python (function predict_rt). If the predicted RT is within Δt95%, the PSM is considered true, otherwise it is considered false.
Each RT prediction model was only used to analyze the PSMs of Nt-arginylation identified in the same dataset used to build that model. In LC-PRM-MS experiments, a mixture of HeLa digest and PRTC (Thermo Fisher Scientific) as internal standards was analyzed by LC-MS/MS in DDA mode before every six PRM experiments. The dataset collected in this DDA mode was used to build an RT prediction model, which was used to analyze the following six PRM data.
Mass error test
For each PSM of Nt-arginylation, m/z errors of fragment ions were calculated by subtracting theoretical m/z values from the measured m/z values. A two-sided Student’s t-test was performed between the m/z errors of b ions and those of y ions in R (function t.test). A PSM with a P-value ≥ 0.05 (that is, an insignificant P-value) was considered true, otherwise, it is considered false.
Structural annotation of Nt-arginylation sites
For structural annotation of the Nt-arginylated sites, we utilized information in the UniProt knowledgebase or used bioinformatic tools such as dagLogo27, DeepLoc, SignalP40, TargetP41, Procleave42 and Alphafold60. The dagLogo R/Bioconductor package v1.28.1 was used to analyze differential amino acid usage (DAU) of Nt-arginylated peptides. A background model was prepared from UniProt protein database (Release 2023_02), with an argument of “fisher” test type. A total of 10 amino acids from P5 to P5’ in the sequence were tested against the background model for statistical significance (function testDAU). Visualization was performed by functions of dagHeatmap for heatmap plot and dagLogo for sequence logo plot. The predicted protein localization was determined using DeepLoc v2.0. The sequences of proteins corresponding to the identified Nt-arginylation sites were extracted from the database and used for DeepLoc analysis. Each protein was assigned a localization that exhibited a probability exceeding 0.5. Proteases predicted to cleave each Nt-arginylation sites were acquired using Procleave algorithm, inputting the sequence from P4 to P4’ into the algorithm. Predictions were made for all 27 accessible proteases. pLDDT scores of AlphaFold were acquired using protti R package (v0.9.1, function fetch_alphafold_prediction) together with AlphaFold database version v4. The pLDDT score was calculated for a total of 21 amino acids, including the N-terminal 10 and the C-terminal 10 amino acids from the Nt-arginylation site.
Functional annotation of Nt-arginylation sites
Reactome61, GO62, and STRING63 were used for functional annotation of Nt-arginylation sites. Reactome, Gene Ontology, and GESA results were obtained using clusterProfiler R package v4.12.6 (function enrichPathway, enrichGO, and gsePathway, repectively)64. The P-values were corrected by Benjamini-Hochberg method65 and the cutoff was set to 0.05. Protein-protein interaction network was obtained from STRING with medium confidence (combined score > 0.4) and visualized using Cytoscape v3.9.166.
Plasmid construction for the arginylated protein candidates
HeLa cells, as well as wild-type (+/+) and Ate1−/− MEFs, were cultured in Dulbecco’s Modified Eagle Medium (DMEM/high glucose; HyClone, Cat# SH30243.01) supplemented with 10% fetal bovine serum (FBS; Gibco, Cat# 16000044) in a 5% CO2 incubator at 37 °C.
Total RNA was isolated using TRI Reagent (Molecular Research Center, Cat# TR 118). Two micrograms of the isolated total RNA was utilized for cDNA synthesis using TOPscript™ RT DryMIX (Enzynomics, Cat# RT220). The constructs for arginylated protein candidates were generated through PCR amplification from a human cDNA library and subsequently subcloned into the pcDNA 3.1 myc/his B plasmid (Invitrogen) using specific restriction sites. The primer information used in these experiments is provided in Supplementary Table 1).
R-catcher pulldown assay
Plasmids encoding the arginylated protein candidates were transiently transfected into HeLa cells using XtremeGene HP DNA transfection reagent (Roche, Cat# C756V59) according to the manufacturer’s protocol. After transfection for 24 h, cells were treated with 10 μM MG132 and 50 nM thapsigargin, followed by incubation for 24 h. Cells were then collected in cold phosphate-buffered saline (PBS) and centrifuged at 500 × g for 5 min. The cell pellets were resuspended in hypotonic buffer containing protease and phosphatase inhibitors. The resuspended cells were lysed by undergoing at least five cycles of freezing and thawing using liquid nitrogen and 37 °C water bath, followed by centrifugation at 15,928 × g for 20 min at 4 °C. We have previously described detailed protocols for the R-catcher pulldown assays11,67. Briefly, 300 μL of purified R-catcher WT and mutant (D129A) proteins were conjugated with 120 μL (50% slurry) of Strep-Tactin Sepharose resin overnight at 4 °C. Subsequently, 400 μg of cell lysates from MG132- and thapsigargin-treated cells were diluted in 940 μL of binding buffer (0.05% Tween 20, 10% glycerol, 0.2 M KCl, and 20 mM HEPES at pH 7.9) and mixed with 60 μL (in packed volume) of R-catcher-conjugated beads. The mixtures were gently rotated at 4 °C for 3 h. The beads were collected by centrifugation at 4600 × g for 1 min, washed five times with 1 mL of binding buffer at 4 °C for 10 min, resuspended in 50 μL SDS sample buffer, and heated at 100 °C for 10 min. The protein samples were then separated by SDS-PAGE and transferred onto a polyvinylidene difluoride (PVDF) membrane (Cytiva, Cat# 10600023) at 35 V overnight at 4 °C. The protein-bound PVDF membrane was subsequently blocked with 5% skim milk in TBS-T buffer for 1 h at room temperature. The membrane was then incubated with the primary Myc antibody overnight at 4 °C, followed by a 1-h incubation with a host-specific HRP-conjugated mouse secondary antibody at room temperature. Protein bands were visualized using an enhanced chemiluminescence (ECL) solution (Thermo Fisher Scientific, Cat# 32106) and X-ray films.
PRM-MS
PRM-MS was performed on the same LC-MS instrument as in the profiling experiments. A 2 µg peptide sample spiked with 125 fmol of PRTC peptide standard (Thermo Scientific) was injected onto a PepMap 100 trap column (75 µm × 20 mm, Thermo Fisher Scientific), washed with 0.1% FA in 2% ACN for 10 min at a flow rate of 5 µL/min and subsequently transferred to an EASY-Spray PepMap RSLC analytical column (75 µm × 150 mm, Thermo Fisher Scientific). Peptides were separated at 300 nL/min using a 55 min linear gradient from 2.0 to 24.0% LC solvent B (0.1% FA in 80% ACN) in LC solvent A (0.1% FA). Similar MS settings as described above were used, but the MS was operated in PRM mode with the following adjustments: PRM MS1 resolution was 70,000 at m/z 200 (Orbitrap), maxIT = 30 ms; targeted MS2 resolution = 35,000. Targeted MS2 spectra were recorded at a resolution of 35,000 and using an AGC target value of 2e5 charges, a maxIT of 200 ms, an isolation window of 1.2 m/z, and an isolation offset of 0.4 m/z. The number of targeted precursors per cycle was set to 20. The first mass was fixed to 200 m/z. Per six PRM LC-MS analyses, 0.5 µg of HeLa digest/PRTC standard (Thermo Fisher Scientific) was injected and analyzed with DDA method and identical LC gradients to PRM-MS. The inclusion list was comprised of 65 targets including 15 PRTC, 6 UPR related proteins, 21 Nt-arginylated peptides and 21 corresponding non-Nt-arginylated peptides. The charge state of each target peptide was determined using the “iep” application of EMBOSS. The PRM acquisition times for the targets were specified as −2 min for the start time and +4 min for the end time, based on the predicted RT. Out of 15 PRTCs, 14 except the most hydrophobic ones were used as internal standards (ISTD).
PRM spectral assignment to each target used a two-step criterion: (i) MS1 precursor agreement within 0.001 m/z, and (ii) MS2 spectral matching to model-predicted fragments with a 5 ppm fragment-ion tolerance. PRM spectra exhibiting PCC values greater than 0.7, which represented the minimum PCC of PRTC standards, were selected, of which the spectra having at least 4 fragments were used for subsequent analysis. The intensities for fragment ions were integrated to get target peptide intensities and then normalized by the PRTC intensities to correct for run-to-run variation. The normalization was done using the crmn R package (v0.0.21). The final LFQ intensities were acquired using MSFragger (v4.0) and IonQuant (v1.10.12) in the platform of FragPipe (v21.1). Match-between-run workflow “LFQ-MBR” was used under the default parameters except for the following: fragment ion tolerance to ±0.05 Da, arginylation (+156.1011 Da) at N-terminal aspartate and glutamate, and arginylation-deamidation (+157.0851 Da) at N-terminal asparagine and glutamine as variable modification.
Immunoblot analysis
Cells were lysed using ice-cold RIPA buffer (Sigma Aldrich, R0278) supplemented with a protease inhibitor cocktail and agitated for 30 min at 4 °C. Lysates were subsequently probe-sonicated and centrifuged at 16,000 × g for 20 min at 4 °C. Protein concentrations were determined using Pierce BCA Protein Assay Kit (Thermo Scientific). For western blot analysis, 1 µg of protein was separated by SDS-PAGE to detect all targets except cleaved caspase-3. For the detection of cleaved caspase-3, the same lysate was concentrated using 3 kDa molecular weight cutoff centrifugal filter (Amicon), followed by protein quantification with BCA assay, and 5 µg of protein was separated by SDS-PAGE. After completion of the electrophoresis, proteins were transferred to a PVDF membrane at 100 V for 1 h. The membrane was blocked with 5% skim milk in TBS-T (20 mM Tris, 150 mM NaCl, and 0.1% Tween 20, pH 7.5) for 1 h at room temperature, followed by overnight incubation at 4 °C with the appropriate primary antibodies, diluted in a phosphate buffered saline (PBS) solution containing 1% bovine serum albumin (BSA) and 0.02% sodium azide. After incubation, the membranes were washed with TBS-T three times and treated with the rabbit IgG-HRP secondary antibodies (1:1,000,000 dilution in 5% skim milk) for 1 h. The membranes were washed with TBS-T and visualized with the ECL chemiluminescent substrate (Thermo Fisher Scientific, A38555). Subsequently, the PVDF membranes were stripped with West Ez Stripping Buffer (GenDEPOT, S2100-050) following the manufacturer’s protocol and re-probed to detect multiple proteins of interest (POIs) and a loading control on the same blot.
The antibodies used are as follow: rabbit polyclonal anti-HSPA5 (Cell Signaling Technology, #3183, lot 11, 1:5000), rabbit polyclonal anti-R-HSPA5 (Sigma–Aldrich, #ABS2103, lot 4117365, 1:5000), rabbit polyclonal anti-CRT (courtesy of Dr. Yong Tae Kwon, 1:2000), rabbit polyclonal anti-R-CRT (courtesy of Dr. Yong Tae Kwon, 1:2000), rabbit polyclonal anti-PDI (courtesy of Dr. Yong Tae Kwon, 1:1000), rabbit polyclonal anti-R-PDI (courtesy of Dr. Yong Tae Kwon, 1:2000), rabbit monoclonal anti-ATF4 (Cell Signaling Technology, 11851, lot 6, 1:2000), rabbit polyclonal anti-caspase-3, cleaved form (Cell Signaling Technology, #9661, lot 47, 1:500), rabbit polyclonal anti-GAPDH (Abcam, #ab9485, lot 1064471-1, 1:10,000), mouse monoclonal anti-alpha-tubulin (Santa Cruz Biotechnology, #sc-5286, lot D2310, clone B-7, 1:1000), rabbit monoclonal anti-cytochrome c (Cell Signaling Technology, #4280, lot 3, clone 136F3, 1:1000), mouse monoclonal anti-ATE1 (Santa Cruz Biotechnology, #sc-271220, lot F2921, clone E-6, 1:2000), mouse monoclonal anti-beta-actin (Sigma-Aldrich, #A1978, lot 0000227449, clone AC-74, 1:30,000), mouse monoclonal anti-Myc (Santa Cruz Technology, #sc-40, lot C2224, clone 9E10, 1:2000), horse anti-mouse IgG-HRP (Cell Signaling Technology, #7076, lot 38, 1:5000) and goat anti-rabbit IgG-HRP (Cell Signaling Technology, #17492, lot 33, 1:100,000).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The MS and processed proteome data generated in this study have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository under accession codes PXD058868 (N-terminome profiling data) and PXD058872 (PRM-MS data). The identical datasets were also deposited in the KPOP (Korea ProteOme rePository, https://kbds.re.kr/KPOP) under accession ID KAP241007. All other data generated in this study are provided in the Supplementary Information/Source Data file. Source data are provided with this paper.
Code availability
The code for the ML-based filtering and data analyses is available on GitHub (https://github.com/syju1984/Nt-arginylationFiltering) and Zenodo (https://doi.org/10.5281/zenodo.17247948) under MIT license.
References
Balzi, E., Choder, M., Chen, W. N., Varshavsky, A. & Goffeau, A. Cloning and functional analysis of the arginyl-tRNA-protein transferase gene ATE1 of Saccharomyces cerevisiae. J. Biol. Chem. 265, 7464–7471 (1990).
Elias, S. & Ciechanover, A. Post-translational addition of an arginine moiety to acidic NH2 termini of proteins is required for their recognition by ubiquitin-protein ligase. J. Biol. Chem. 265, 15511–15517 (1990).
Ji, C. H. et al. The N-degron pathway mediates ER-phagy. Mol. Cell 75, 1058–1072 e1059 (2019).
Yoo, Y. D. et al. N-terminal arginylation generates a bimodal degron that modulates autophagic proteolysis. Proc. Natl. Acad. Sci. USA 115, E2716–E2724 (2018).
Kwon, Y. T. et al. An essential role of N-terminal arginylation in cardiovascular development. Science 297, 96–99 (2002).
Brower, C. S., Piatkov, K. I. & Varshavsky, A. Neurodegeneration-associated protein fragments as short-lived substrates of the N-end rule pathway. Mol. Cell 50, 161–171 (2013).
Ma, S., Attarwala, I. Y. & Xie, X. Q. SQSTM1/p62: a potential target for neurodegenerative disease. ACS Chem. Neurosci. 10, 2094–2114 (2019).
Moorthy, B. T. et al. The evolutionarily conserved arginyltransferase 1 mediates a pVHL-independent oxygen-sensing pathway in mammalian cells. Dev. Cell 57, 654–669 e659 (2022).
Rai, R. et al. Arginyltransferase regulates alpha cardiac actin function, myofibril formation and contractility during heart development. Development 135, 3881–3889 (2008).
Karakozova, M. et al. Arginylation of beta-actin regulates actin cytoskeleton and cell motility. Science 313, 192–196 (2006).
Seo, T. et al. R-catcher, a potent molecular tool to unveil the arginylome. Cell. Mol. Life Sci. 78, 3725–3741 (2021).
MacTaggart, B. et al. Global analysis of post-translational side-chain arginylation using pan-arginylation antibodies. Mol. Cell. Proteom. 22, 100664 (2023).
Wong, C. C. et al. Global analysis of posttranslational protein arginylation. PLoS Biol. 5, e258 (2007).
Lin, Z. et al. An unbiased proteomic platform for ATE1-based arginylation profiling. Nat. Chem. Biol. 21, 1970–1980 (2025).
Gevaert, K. et al. Exploring proteomes and analyzing protein processing by mass spectrometric identification of sorted N-terminal peptides. Nat. Biotechnol. 21, 566–569 (2003).
Kleifeld, O. et al. Isotopic labeling of terminal amines in complex samples identifies protein N-termini and protease cleavage products. Nat. Biotechnol. 28, 281–288 (2010).
Varshavsky, A. N-degron pathways. Proc. Natl. Acad. Sci. USA 121, e2408697121 (2024).
Xu, T., Wong, C. C., Kashina, A. & Yates, J. R. 3rd Identification of N-terminally arginylated proteins and peptides by mass spectrometry. Nat. Protoc. 4, 325–332 (2009).
Bouwmeester, R., Gabriels, R., Hulstaert, N., Martens, L. & Degroeve, S. DeepLC can predict retention times for peptides that carry as-yet unseen modifications. Nat. Methods 18, 1363–1369 (2021).
Zhou, X. X. et al. pDeep: predicting MS/MS spectra of peptides with deep learning. Anal. Chem. 89, 12690–12697 (2017).
Ekvall, M., Truong, P., Gabriel, W., Wilhelm, M. & Kall, L. Prosit transformer: a transformer for prediction of MS2 spectrum intensities. J. Proteome Res. 21, 1359–1364 (2022).
Lou, R. et al. DeepPhospho accelerates DIA phosphoproteome profiling through in silico library generation. Nat. Commun. 12, 6685 (2021).
Zeng, W. F. et al. AlphaPeptDeep: a modular deep learning framework to predict peptide properties for proteomics. Nat. Commun. 13, 7238 (2022).
Ju, S. et al. iNrich, rapid and robust method to enrich N-terminal proteome in a highly multiplexed platform. Anal. Chem. 92, 6462–6469 (2020).
Desai, H. et al. Chemoproteogenomic stratification of the missense variant cysteinome. Nat. Commun. 15, 9284 (2024).
Ju, S. & Lee, C. Identification of protein arginylation by encapsulated n-terminal peptide enrichment method. Methods Mol. Biol. 2620, 229–241 (2023).
Ou, J. et al. dagLogo: an R/Bioconductor package for identifying and visualizing differential amino acid group usage in proteomics data. PLoS ONE 15, e0242030 (2020).
Mahrus, S. et al. Global sequencing of proteolytic cleavage sites in apoptosis by specific labeling of protein N termini. Cell 134, 866–876 (2008).
Umeda, T. et al. Limited proteolysis of filamin is catalyzed by caspase-3 in U937 and Jurkat cells. J. Biochem. 130, 535–542 (2001).
Gessulat, S. et al. Prosit: proteome-wide prediction of peptide tandem mass spectra by deep learning. Nat. Methods 16, 509–518 (2019).
Kim, M. S., Zhong, J. & Pandey, A. Common errors in mass spectrometry-based analysis of post-translational modifications. Proteomics 16, 700–714 (2016).
Hoernstein, S. N. et al. Identification of targets and interaction partners of arginyl-tRNA protein transferase in the moss physcomitrella patens. Mol. Cell. Proteom. 15, 1808–1822 (2016).
Lee, H. et al. Distinguishing N-terminal methylation from near-isobaric modifications by statistical analysis of mass error distributions of fragment ions. J. Proteome Res. 24, 4804–4815 (2025).
Yalcin, T., Khouw, C., Csizmadia, I. G., Peterson, M. R. & Harrison, A. G. Why are B ions stable species in peptide spectra? J. Am. Soc. Mass Spectrom. 6, 1165–1174 (1995).
Medzihradszky, K. F. Peptide sequence analysis. Methods Enzymol. 402, 209–244 (2005).
Hiserodt, R. D., Brown, S. M., Swijter, D. F., Hawkins, N. & Mussinan, C. J. A study of b1+H2O and b1-ions in the product ion spectra of dipeptides containing N-terminal basic amino acid residues. J. Am. Soc. Mass Spectrom. 18, 1414–1422 (2007).
Bludau, I. et al. The structural context of posttranslational modifications at a proteome-wide scale. PLoS Biol. 20, e3001636 (2022).
Goitea, V. E. & Hallak, M. E. Calreticulin and arginylated calreticulin have different susceptibilities to proteasomal degradation. J. Biol. Chem. 290, 16403–16414 (2015).
Mallia-Milanes, B. et al. TAILS proteomics reveals dynamic changes in airway proteolysis controlling protease activity and innate immunity during COPD exacerbations. Am. J. Physiol. Lung Cell. Mol. Physiol. 315, L1003–L1014 (2018).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 40, 1023–1025 (2022).
Almagro Armenteros, J. J. et al. Detecting sequence signals in targeting peptides using deep learning. Life Sci. Alliance 2, e201900429 (2019).
Li, F. et al. Procleave: predicting protease-specific substrate cleavage sites by combining sequence and structural information. Genom. Proteom. Bioinform. 18, 52–64 (2020).
Ducker, G. S. et al. Reversal of cytosolic one-carbon flux compensates for loss of the mitochondrial folate pathway. Cell Metab. 23, 1140–1153 (2016).
Chen, J., Cui, L., Lu, S. & Xu, S. Amino acid metabolism in tumor biology and therapy. Cell Death Dis. 15, 42 (2024).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Fusakio, M. E. et al. Transcription factor ATF4 directs basal and stress-induced gene expression in the unfolded protein response and cholesterol metabolism in the liver. Mol. Biol. Cell 27, 1536–1551 (2016).
Bushell, M. et al. Disruption of the interaction of mammalian protein synthesis eukaryotic initiation factor 4B with the poly(A)-binding protein by caspase- and viral protease-mediated cleavages. J. Biol. Chem. 276, 23922–23928 (2001).
Wejda, M. et al. Degradomics reveals that cleavage specificity profiles of caspase-2 and effector caspases are alike. J. Biol. Chem. 287, 33983–33995 (2012).
Nesvizhskii, A. I. Proteogenomics: concepts, applications and computational strategies. Nat. Methods 11, 1114–1125 (2014).
Reich, S. et al. A multi-omics analysis reveals the unfolded protein response regulon and stress-induced resistance to folate-based antimetabolites. Nat. Commun. 11, 2936 (2020).
Liu, C. et al. Cytoplasmic SHMT2 drives the progression and metastasis of colorectal cancer by inhibiting beta-catenin degradation. Theranostics 11, 2966–2986 (2021).
Jiang, C. et al. Regulation of mitochondrial respiratory chain complex levels, organization, and function by arginyltransferase 1. Front. Cell Dev. Biol. 8, 603688 (2020).
Wang, H. & Julien, O. CaspSites: a database and web application for experimentally observed human caspase substrates using n-terminomics. J. Proteome Res. 22, 454–461 (2023).
Piatkov, K. I., Brower, C. S. & Varshavsky, A. The N-end rule pathway counteracts cell death by destroying proapoptotic protein fragments. Proc. Natl. Acad. Sci. USA 109, E1839–E1847 (2012).
Cha-Molstad, H. et al. Amino-terminal arginylation targets endoplasmic reticulum chaperone BiP for autophagy through p62 binding. Nat. Cell Biol. 17, 917–929 (2015).
Carpio, M. A., Lopez Sambrooks, C., Durand, E. S. & Hallak, M. E. The arginylation-dependent association of calreticulin with stress granules is regulated by calcium. Biochem. J. 429, 63–72 (2010).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Song, X. et al. Crosstalk between apoptosis and autophagy is regulated by the arginylated BiP/Beclin-1/p62 complex. Mol. Cancer Res. 16, 1077–1091 (2018).
Nawale, L. et al. ATE1 promotes breast cancer progression via arginylation-dependent regulation of MAPK-MYC signaling. Cell Commun. Signal. 23, 390 (2025).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Fabregat, A. et al. The reactome pathway knowledgebase. Nucleic Acids Res. 46, D649–D655 (2018).
Thomas, P. D. et al. PANTHER: making genome-scale phylogenetics accessible to all. Protein Sci. 31, 8–22 (2022).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Xu, S. et al. Using clusterProfiler to characterize multiomics data. Nat. Protoc. 19, 3292–3320 (2024).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate - a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 57, 289–300 (1995).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Seo, T., Han, G. & Cha-Molstad, H. N-Terminal arginylation pull-down analysis using the R-Catcher tool. Methods Mol. Biol. 2620, 219–228 (2023).
Acknowledgements
This work was supported by grants from the National Research Foundation of Korea (RS-2023-00279134, RS-2022-NR068428, RS-2024-00444177), a grant from the National Research Council of Science & Technology (GTL24022-000), a KIST intramural program (Grand Challenge), and the KRIBB Research Initiative Program (KGM1062413).
Author information
Authors and Affiliations
Contributions
S.J. and C.L. conceptualized the study. S.J. and C.L. developed the methodology. S.J. wrote the algorithm. L.N., S.L., and J.G.K. performed immunoblot validation. S.L. and H.L. performed targeted proteomics. D.H.K. and H.C.M. provided resources. S.J., L.N., S.L., J.G.K., H.L., N.P., H.C.M., and C.L. curated the data. All authors wrote the paper. C.L. supervised the project and was the project administrator. C.L. acquired funding. These authors contributed equally: S.J., L.N., and S.L. These authors jointly supervised this work: Hyunjoo Cha-Molstad and Cheolju Lee.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Anna Kashina and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ju, S., Nawale, L., Lee, S. et al. Implementing N-terminomics and machine learning to probe Nt-arginylation. Nat Commun 17, 172 (2026). https://doi.org/10.1038/s41467-025-66883-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66883-6