Abstract
Soil ecosystems are crucial for supporting life, yet little is known about their biodiversity and its distribution in extreme environments. The Atacama Desert, the driest non-polar desert on Earth, has scarce water, high salinity, and metal-rich water bodies, creating challenging conditions for most organisms. Above-ground life has been partially documented, but its soils remain poorly studied. Here we show that soil nematodes, an abundant and diverse group of invertebrates, display distinct biodiversity patterns across the Atacama at multiple biological scales, including genetic, taxonomic, community, and life-cycle levels. Surveys across dune systems, high-altitude mountains, saline lakes, river valleys, and fog oases reveal unique assemblages in each habitat. We find that asexual taxa are more common at higher altitudes, consistent with patterns of geographical parthenogenesis. Genus richness follows a latitudinal gradient and increases with precipitation. These results demonstrate that even in one of the most extreme terrestrial environments, stable soil communities can persist. However, evidence of simplified soil food webs suggests vulnerability to further environmental change. Our findings provide new insights into the mechanisms shaping biodiversity in arid ecosystems and can inform predictions about soil resilience under global climate-driven aridification.
Similar content being viewed by others
Introduction
Understanding which factors shape the diversity and distribution of species is a major focus of ecological research. Studies examining the interaction of organisms with their immediate environment focus on the influence of ecological, geological, and geographical parameters in diverse systems such as aquatic and soil habitats1,2,3,4,5. From nutrient cycling, mineralization, and pest control6 to carbon sequestration and storage, soils and their processes play a key role in food security, water supply and quality, and in shaping biodiversity7. Globally, at least 58 % of species inhabit soils8, including microorganisms and different size classes of fauna9. Accordingly, soils receive extensive research attention, particularly in arable areas. However, geographical patterns of soil biota in relation to environmental parameters remain poorly understood, largely due to limited data on the true diversity and distribution of many soil organisms8,10.
Even less information exists for soil systems in extreme environments such as deserts. Abiotic factors such as strong temperature fluctuations, minimal annual precipitation, and high evaporation rates drive desert aridity and pose significant challenges for life11. Despite these constraints, deserts and their soils contain spatially distributed microhabitats and niches, including limestone areas, dune systems, and saline patches, that support diverse life forms11. With the global trend of aridification12,13,14, studying biodiversity in desert soils is essential for predicting organismal distributions under global change.
The Atacama Desert, the driest non-polar desert on Earth, receives less than 200 mm of rainfall annually. Despite the long-term stability of its geological system, regional environmental patterns are apparent15,16. Long-term aridity shapes the distribution of sulfates, chlorides, and nitrates in Atacama soils, which in turn affect community composition across taxa17,18,19,20. Diverse microhabitats in the Atacama include alkaline soils, dune systems, fog oases, and soils with high salinity and arsenic content. These support specialized taxa such as microbial life forms, plants, and a limited number of vertebrates21,21,22,23,24,25,26,27,28,29,30,31. Invertebrate biodiversity remains understudied in the Atacama, with research focusing mainly on above-ground fauna32,33,34, leaving soil invertebrates poorly sampled and their distribution patterns unclear8,29,35,36,37. Nematodes, one of the most diverse and abundant metazoan groups, contribute to soil nutrient turnover, carbon sequestration, and regulation of bacterial populations, making them important indicators of soil condition36,38,39,40. Their use as functional indicators is based on life cycle characteristics such as feeding type and reproductive strategy38,41. Members of Nematoda adapt to extreme conditions, including low oxygen in deep-sea environments42, daily freezing in Antarctica43,44, and kilometer-deep mines45. Certain families are prevalent and diverse in deserts, surviving low water availability in systems such as the Mojave46, Chihuahuan47, Monte48, and Namib deserts49. Many nematode genera also evolve parthenogenetic lineages, hypothesized to have a founder advantage in extreme environments (geographical parthenogenesis)50,51. In the Atacama, parthenogenesis may therefore be more frequent than sexual reproduction.
Published records of nematodes from the Atacama are usually based on very few or single individuals35. Consequently, it remains unclear whether soil invertebrate biodiversity patterns align with regional and local ecology, adaptive strategies such as feeding type or reproductive mode, or broader global trends such as environmental gradients in species richness3. In this study, we analyze the factors shaping invertebrate biodiversity in Atacama soils using nematodes as a model system. We sample six localities across the desert, including sand dunes, saline lake shores, and high-altitude mountain regions, to characterize biogeographic patterns in genetic diversity, life history, and community composition. Despite extreme and persistent aridity, we find strong local patterning across niches such as sand dunes, saline lakes, fog oases, and river valleys. Using genus richness and reproductive mode modeling approaches, we identify annual precipitation and temperature heterogeneity as the main factors associated to genus richness, and found elevation to be the strongest predictor of reproductive mode. We show that even under extreme environmental conditions, soil niches can sustain stable and diverse invertebrate communities, although some areas show signs of simplified food webs.
Results
In this study, we aimed to analyze different regions of the Atacama and thus established localities and sampling sites in major areas of the desert: (1) the Altiplano area (Tarapacá region) is more humid and known for supporting more vegetation than the hyperarid core of the Atacama52; (2) the area around the Quebrada de Aroma (Tarapacá region) has moderately alkaline soils with increasing gypsum as altitude decreases and harbors several microorganism taxa such as Proteobacteria and Actinobacteria21, (3) the San Francisco area - further on referred to as Eagle Point (Antofagasta region)—is home to several bacterial species with predominant Firmicutes taxa24, (4) the Salar de Huasco (Tarapacá region) experiences high UV radiation, salinity and arsenic concentrations supporting several microbial taxa25,53 as well as three flamingo species, a nearly threatened puma species and several other vertebrates23, (5) the Paposo area (Antofagasta region) receives water in the form of fog (called camanchaca), it harbors plant species-rich fog oases (known as loma formations)54,55 and is an area of high endemism25,56,57 and the (6) Totoral area (Atacama region) is composed of sand dunes where endemic plants of the Apocynaceae family22, as well as reptiles and birds can be found. In order to analyze patterns of biodiversity in the hyper-arid Atacama Desert using nematodes as a model for soil organisms, we sampled, identified, sequenced, and studied diversity at different levels. We analyzed diversity at the haplotype, genus, family, and community level, including life-history characteristics such as feeding types and reproductive modes. We used linear models as well as a random forest approach to assess the factors that contribute the most to the genus richness and reproductive mode patterns across the desert.
A rich diversity of roundworms persists in the Atacama
We collected 393 nematode morphotypes from 112 samples collected along six localities throughout the Atacama Desert (Fig. 1B, supplementary Fig. S1). Collected nematodes were then sequenced (”Methods”) with a percentage of high quality base pairs above at least 20 (HQ 20 %), 386 sequences from different morphotypes were kept for further analysis. More specifically, we analyzed a total of 39 nematodes from the Altiplano locality, 37 from the Aroma locality, 163 from the Eagle Point locality, 44 from the Paposo locality, 26 from the Salars locality (Salars), and 77 from Totoral dunes locality. Our sampling also included sites where no nematodes could be found (Fig. 1B). These sites are used as a reference for environmental conditions under which nematodes do not seem to persist. We classified the nematodes into 21 families and 36 genera (list of lineages per sample can be found on 10.5281/zenodo.15517342—FamilyGeneraperSample.csv), illustrating a rich diversity of roundworms flourishing in the Atacama. Seven families were found in the Altiplano as well as the Aroma locality, nine families were found in Eagle Point, 12 in the Salars, eight in Totoral Dunes, and ten in Paposo (Fig. 2B). We observed that some generalist families were ubiquitously found, whereas others were geographically restricted. Cephalobidae were present in all localities, whereas representatives of families Panagrolaimidae and Qudsianematidae were found in five of the localities (all except for the Salars). The family Anguinidae was uniquely found in the Eagle Point locality. The families Trichistomatidae and Mermithidae were unique to the Salars locality. Tylencholaimidae was uniquely found in the Paposo locality, and Acuariidae in the Altiplano.
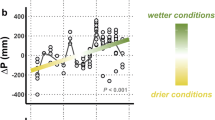
Environmental variables used to model genus richness across the Atacama Desert, with sampling locations shown as points on each map. These maps provide context for the environmental conditions across the study area, including A geographical location of the Atacama Desert, B mean annual precipitation, C temperature range, D elevation, E rock type (classified following138, see Supplementary Table S19), and F soil thickness. All layers represent spatial data, not model predictions. The maps highlight the environmental gradients and heterogeneity present in the Atacama, which underlie the variation in nematode genus richness observed across the sampled sites.
A Jaccard similarity index of families found in six localities, intensity of green depicts similarity between locations (darker green indicates higher similarity). B Number of families found per locality represented by the size of the circles, the proportion of families found with the different feeding types is represented in the pie chart by the different colors.
Genetic diversity and haplotype uniqueness differ between nematode taxa
Given the diversity of retrieved nematode genera and the disparity between ubiquitous and regionally restricted taxa, we wanted to analyze patterns of biogeography on the genetic level. We assessed haplotype clustering and genetic diversity (π, θ) for seven taxonomic groups using 18S rRNA (Table 1). We studied genetic diversity in two ways: per taxonomic group, including all habitats, and per habitat, analyzing one taxonomic group at a time.
There were between 17 to 166 segregating sites for the different soil nematodes in our networks (supplementary Table S8), and we identified differences in haplotype structure and clustering at the genera and family levels (supplementary Figs. S3–S7). For the genera Acrobeloides, Acrobeles, and Panagrolaimus we found 24, 22, and 20 different haplotypes, respectively (Table 1). For the families included in the order Dorylamida and the superfamily Aphelenchoidea, 16 haplotypes were identified, respectively. The family Plectidae was composed of ten different haplotypes. For Alaimidae, only one haplotype was found (two including the reference sequence) and is shared in three habitats (Table 1). Overall, we were not able to find any distinct geographical clustering of haplotypes based on the networks, i.e., individual haplotypes were present in geographically distant localities (a complete list of all identified haplotypes can be found on 10.5281/zenodo.15517342). We found genetic diversity at the genus level to be the highest for Panagrolaimus (π = 0.05788, θ = 0.04221, Fig. 3A) and the lowest for Acrobeloides (π = 0.01278, θ = 0.02192). At the family level, the highest genetic diversity was found for Aphelenchoidea (π= 0.11199, θ = 0.08095, Fig. 3B), and the lowest was found for Plectidae (π = 0.00633, θ = 0.00993).
A Haplotype network of Panagrolaimus based on 18S rRNA sequences. B Haplotype network of Aphelenchoidea based on 18S rRNA sequences. Color refers to sampling locations. Reference sequences were retrieved from the curated 18S-NemaBase120. Accession number and species name are written next to the respective node of haplotype (abbreviated to ''Hap''). Vertical hatchmarks indicate the number of differences from one node to the next.
To better understand the possible effect of geography on genetic distance, we calculated the genetic distance as Euclidean distance and compared it to the geographic distance of the different localities using Mantel statistics based on Spearman’s rank correlation (ρ). For Panagrolaimus, a significant (p < 0.05) isolation-by-distance pattern could be found (r = 0.6, p = 0.041667, 119 permutations); no pattern was found for other taxa analysed. With this approach, it appears that geographically distant haplotypes of Panagrolaimus are also genetically more distant in their 18S rRNA region. Specifically for this genus, we see the highest diversity in the Eagle Point (π=0.06548) and the lowest in Totoral Dunes, where only one haplotype was found for several individuals (supplementary Table S9). These results showcase the high variability of genetic diversity within soil nematode taxa in the Atacama Desert and relation to their habitat.
The nematode community composition varies between habitats
We assessed soil nematode community composition variation in the six localities employing the Jaccard similarity index at the family level. We found the Aroma and Eagle Point localities to share the highest proportion of families (J = 0.78), followed by Altiplano and Aroma (J = 0.75). Conversely, we observed the least amount of shared families between Eagle Point and the Totoral Dunes (J = 0.21) and the Salars locality when compared to Totoral Dunes (J = 0.25)(Fig. 2A). As expected under the isolation-by-distance model, community composition appears to become more dissimilar with increased geographical distance between the different localities (Mantel statistic based on Spearman’s rank correlation ρ, r 0.5153, p = 0.048611, 719 permutations). We were interested to know if a co-occurrence analysis of families would identify a specific geographical pattern, but using 82 family pairs (108 pairs were removed due to unique presence in specific localities) could not retrieve a signal.
Soils in the Atacama are dominated by different categories of nematodes
We wanted to use nematodes as indicators of soil condition throughout different habitats in the desert by assessing their different functional groups. We categorized nematodes into different feeding types, as well as into the colonizer-persister series based on common life-history characteristics reported in literature and assigned using the NINJA application41. Nematodes that are r-strategists (c-p1 colonizer) are considered indicators of resource availability and unstable environments, whereas those that are k-strategists (c-p5 persister) are considered indicators of food web complexity and more stable habitats58. We found that the different habitats show a different structure regarding nematode functional groups. Despite the presence of nematodes belonging to all colonizer-persister categories (c-p) across the desert, r-strategists (categories c-p1 and c-p2) are predominant in the Altiplano locality (57.2 % of families found in the locality) as well as in the Aroma and Totoral Dunes localities. Whereas k-strategists (categories c-p4 and c-p5) were the predominant type of families in the Paposo and Salars localities (supplementary Table S11). By assessing the different feeding types of the nematodes analysed, we found that most families in the different habitats are bacterivores and omnivores (41.2% and 20.1% respectively). The remaining nematode families were categorized as predators, fungivores, and herbivores (18.7%, 14.4%, and 5.6% respectively)(Fig 2B). Families with unicellular eukaryotic feeding type (UEF) were not found in this study (supplementary Table S12). A structuring of the different soil habitats regarding feeding types is apparent, as the Salars only harbor bacterivore, predatory and omnivore families whereas other areas as Totoral Dunes have a more diverse feeding type assemblage with herbivores, fungivores, bacterivores, predators and omnivores present (Fig. 2B).
Climate and latitude determine soil biodiversity in the Atacama
To understand the driving forces and constraints of nematode taxonomic biodiversity across the Atacama Desert more globally, we used a modeling approach incorporating environmental data. We first aimed to predict genus richness and found that the best predictive model using a probabilistic approach incorporated soil thickness and range of temperature. The model showed the highest Akaike weight (0.4709556) and a statistically significant slope for both environmental variables (supplementary Table S16), which indicates that nematode genera richness in the Atacama Desert could be related to reduced soil thickness and more climate heterogeneity, here represented as the range of temperature. To account for variation across localities in our predictions, we further assessed genus richness using linear mixed-effect models specifying the sampling locality as a random effect. Our analysis confirmed that sampling localities had a significant influence on the modeling process, suggesting that spatial variation plays a crucial role in shaping the observed patterns, potentially due to underlying environmental heterogeneity between the different locations. Incorporating locality effects into the models was therefore essential to capture the ecological drivers of genus richness given our dataset. Moreover, we noted that precipitation patterns are not correlated to the distance between localities (Supplementary note 1).
With this approach, we then found that the best predictive model incorporated mean annual precipitation and range of temperature, representing the seasonal thermal amplitude an organism experiences within a given area (temperature heterogeneity). The preferred model showed the highest Akaike weight (0.5263228) and a statistically significant slope for the intercept and both climatic variables (equation 1)(Table 2).
To further validate the results of the tested linear models, we implemented a random forest approach. With this approach, we found that the most important variables for genera richness prediction in our analysis were elevation, mean annual precipitation, latitude, and range of temperature (supplementary Table S17). This corroborates that mean annual precipitation and range of temperature, in congruence with the linear models, have a strong, consistent influence on genus richness prediction. With the three approaches, we found that fewer genera are predicted to occur in the hyper-arid core of the desert (e.g zero for the region of Pampa del Tamarugal), and more genera are predicted to occur, for instance, in the Altiplano region (Fig. 4A). Analyzing the raw data relationship between environmental variables and genus richness, we see that the linear associations were generally weak or non-significant (supplementary Fig. S11), the random forest model captures more complex, potentially nonlinear effects, as shown in the partial dependence plots (supplementary Fig. S12). This highlights that genus richness responds to environmental gradients in a way not fully captured by only linear models. Due to the differing assumptions underlying each modeling approach, a single definitive distribution of genus richness cannot be determined. However, temperature heterogeneity consistently emerged as a strong predictor across all models. Additionally, mean annual precipitation was identified as a significant factor in both linear mixed-effects and random forest models, underscoring the important role of climate in shaping genus richness.
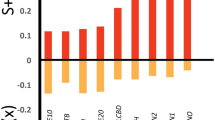
The upper panel (A–C) displays predicted genus richness based on different modeling approaches and environmental variables, dark blue indicates lower genera richness and yellow indicates higher genera richness: A uses a generalized linear model with soil thickness and temperature range as predictors; B applies a linear mixed-effects model based on mean annual precipitation; and C employs a random forest model. In these maps, yellow indicates areas with higher predicted genus richness, while blue indicates areas with lower richness. The lower panel (D–F) presents the predicted probability of asexual reproduction, dark purple indicates higher likelihood of being sexual and light brown indicates higher likelihood of being asexual: D and E use elevation as a predictor in a generalized linear model and a generalized mixed-effects model, respectively, and F shows predictions from a random forest model. In these maps, dark purple indicates areas where sexual reproduction is more likely, while beige shows areas with a higher likelihood of asexual reproduction. ''Soil” is used as an abbreviation for soil thickness, ''RangeT” as an abbreviation for range of temperature, ''MAP” is used as an abbreviation for mean annual precipitation and ''Rock” as an abbreviation for rock type.
Reproductive mode association to environmental variables
The theory of geographical parthenogenesis predicts an excess of asexual taxa in marginal habitats. Given our findings of taxonomically structured nematode communities in different regions of the Atacama Desert, we wanted to test for the specific life-history trait of parthenogenesis (asexual reproduction) using our modeling approach. We established single-worm cultures for a representative subset of our samples. Ten out of the eighteen cultures showed nematodes with parthenogenetic reproduction, i.e., no presence of males and successful propagation from a single unfertilized female. The remaining seven cultures showed sexual reproduction, i.e. presence of males (supplementary Fig. S2G) and no propagation from single females. Sexual strains showed a propagation rate of 0% from single unfertilized female cultures, whereas the asexual strains’ propagation rate was 83.3% ( ± 10.48%). We initially tested variables of elevation and latitude as predictors of reproductive mode along with mean annual precipitation and range of temperature, the former two to test for common variables associated with geographical parthenogenesis, the latter two given their effect on genera richness prediction from our linear mixed-effects and random forest models. We tested generalized linear models as well as generalized mixed-effects models to account for the effect of sampling locality (supplementary Tables S20 and S21). In addition, we tested a model including vegetation (supplementary Fig. S14, supplementary Table S20). The best predictive model for reproductive mode incorporated only elevation, showing the highest Akaike weight and a statistically significant slope (supplementary Table S16), while, for example, plant association did not show any effect on the prediction of reproductive mode. We also tested the effect of sampling locality on the prediction of reproductive mode and found, in congruence with the previous approach, that the model incorporating only elevation was preferred (Table 3). Our validation of these linear models with a random forest approach supports that increasing elevation is the main driver for asexual reproductive mode (represented as 1) in soil nematodes in the Atacama (supplementary Table S18). Our models show that the likelihood of nematode taxa being asexual is greater at higher elevations than in lower ones (e.g. coastal area)(Fig. 4). To account for variation across localities in our predictions, we further assessed genus richness using generalized linear mixed effect models specifying the sampling locality as a random effect. In this case, we found that the sampling localities have no significant effect on the likelihood of a reproductive mode.
Discussion
Altered soil characteristics, along with increasing temperatures, will have an impact on community dynamics and species abundance, especially in times of accelerated global change59,60,61. Dry lands cover around 40% of the Earth’s land surface62. These areas are projected to increase in aridity as a consequence of climate change63. Desertification processes, as an example, soil erosion, acidification, and salinization, have affected over 12% of dry lands already14, partly due to changing precipitation patterns64. Understanding soil biodiversity and species distribution in present-day desert environments could provide a model for further studies on the effect of desertification and the resilience of soil ecosystems. Using nematodes as a model system, we analyzed which factors shape soil biodiversity in an extreme environment, the Atacama Desert in northern Chile. While aridity generally characterizes the Atacama Desert, we found that nematode diversity shows patterning throughout different niches (e.g., sand dunes, saline lake-shore sediments, fog oases), as well as in relation to ecological and geographical parameters. We identified mean annual precipitation and latitude to be the major predictors of genus richness in the desert and elevation was identified as the primary predictor of reproductive mode. Additionally, the characterization of nematodes as indicators of soil condition, based on their life cycle characteristics, revealed that some niches are stable (taxa that are sensitive to pollutants and disturbance) while others exhibit signs of a poor soil food web with high tolerance to pollutants and disturbance. Here, we describe the biogeographic patterns of nematodes, focusing on their diversity, life-history, and community composition in the Atacama Desert, where studies on soil organisms, such as roundworms, are still scarce29,35.
Atacama soils harbor a rich nematode diversity
We analyzed diversity from the haplotype level (genetic diversity) to community level patterns (genera richness and Jaccard similarity at the family level). We report that the Atacama Desert is home to at least 36 different genera of nematodes grouped in 21 families, spread across various habitats. We identified cosmopolitan nematode families, including members of Cephalobidae and Panagrolaimidae, which have been documented in other desert environments such as the Namib Desert65,66, the Judean Desert67, the Mojave Desert68,69, and the Monte Desert70. These families have also been reported in the arid region of Arica in the Atacama Desert37. We found both families in most analyzed areas and aimed to assess the genetic differences among individuals from different localities and their relationship to ecological, geological, and geographical factors. Particularly for the family Panagrolaimidae, all sequenced individuals belonged to only one genus (Panagrolaimus), which displayed an isolation-by-distance pattern (IBD). We were able to detect this pattern in Panagrolaimus specifically, as we found a high enough proportion of sexually reproducing individuals present in our samples (asexually reproducing species do not have gene flow and thus by default show separation). A possible explanation for IBD is limited dispersal, which has been reported to be lower in nematodes of this genus when compared to other species in a laboratory setting71. Generally, dispersal is lower for below-ground organisms when compared to above-ground taxa72. Dispersal can be mediated by wind in small invertebrates like nematodes73, but the effect of wind dispersal has not been studied to date in the Atacama.
Soil biodiversity in the Atacama is highly structured and in parts sensitive to disturbance
By examining communities at both the haplotype and family levels across different desert niches, we identified distinct local patterns. For example, the Jaccard similarity analysis revealed distinct communities in salars compared to other regions. The influx of freshwater from complex underground aquifers into high-altitude wetlands, along with elements like lithium and arsenic, high salt concentrations, and seasonal fluctuations, contributes to the formation of these unique ecosystems53,74,75. Microbial life, proposed as the dominant life form in this area, displays density and structure changes through dry and wet seasons75, which in consequence can have an effect on the nematode community in the area76, given that most families found in the wetlands were bacterivores. Differences in the taxa found in the different areas can also provide information on the complexity of the soil food web; both the salars and Paposo were dominated by persisters that are described to be more sensitive to environmental disturbance and often have a narrow ecological amplitude. Even if these indicators are developed in communities of agricultural soils77, which are expected to differ significantly from the oligotrohpic desert soils, the relative difference in the proportion of persisters among the here investigated localities ( ≈ 10% more than colonizers), highlights the sensitivity of food webs in the salars and Paposo. This aligns with observed and reported threats to these ecological niches that are mining, plant extraction, pasture beyond the area’s capacity, truck transit with dangerous materials, uncontrolled tourism, and habitat loss due to vehicle transit off roads78.
In contrast, in the Altiplano, the Totoral Dunes, and the Aroma area the dominant groups were colonizers that are described to be more resistant to adverse environmental conditions in agricultural soils, indicating poor soil food web function in the absence of some feeding types79,80. Even if the here investigated soils cannot compare to eutrophic agricultural conditions, the relative dominance in colonizers in comparison to the soils from the Salar and Paposo localities indicate altered food web function in Apltiplano, Totoral Dunes, and Aroma. Sand dune systems, such as Totoral Dunes, have already been reported to be dominated by coloniser taxa due to their broad ecological range, with high abundances of the family Cephalobidae throughout coastal sand dunes81,82.
Soil community diversity patterns across the Atacama Desert are primarily shaped by climate
To further understand if broader scale patterns could be found for the soil communities, we implemented a genera richness modeling approach. In isolated areas such as the Atacama Desert, comprehensive field studies that encompass the entire area are challenging due to the desert’s complex topography and large terrain extension. Modeling approaches provide a simplified representation of real-world phenomena83specifically for multifaceted scenarios, as we observed in the Atacama. These models are thus an excellent tool to disentangle the driving factors of species and community distributions by incorporating the complex interplay of ecological, geographical and edaphic factors84.
Our results highlight the key role of climate, particularly mean annual precipitation and temperature heterogeneity, in shaping nematode genera richness. Precipitation, as a proxy for water availability, is a well-established driver of soil biodiversity across various taxa, including protists, bacteria, fungi, arachnids, rotifers, and nematodes85. In desert environments, higher precipitation can enhance biodiversity by improving nutrient availability and soil moisture86. Specifically, in the Atacama Desert, it has been linked to species distribution and habitat suitability in beetles87 and increased microbial diversity at higher elevations21.
Soil nematodes, like other invertebrates88, rely on the thin water film between soil particles for movement, making water availability crucial89. While some can survive dry conditions through anhydrobiosis90,91,92, active nematodes depend on precipitation to sustain mobility and biodiversity. Increased precipitation has also been linked to shifts in nematode communities, particularly an increase in bacterial-feeding nematodes, likely due to a rise in bacterial biomass93.
Beyond precipitation, we found that temperature heterogeneity (seasonal temperature variation) also influences genera richness in the Atacama Desert. While temperature fluctuations are known to shape community dynamics, their precise role in food web interactions remains incompletely understood94. Temperature heterogeneity has been identified as a key factor in population structure95, a driver of increased protist densities96, and a contributor to significant community dissimilarities97. Variations in climate across localities may create distinct micro-niches, fostering the establishment of diverse biological communities.
Although not identified as the primary drivers of nematode diversity in our study, latitude, elevation, and edaphic factors (such as soil thickness and rock type) may still influence community diversity in the extreme environment of the Atacama Desert. These edaphic factors have been proposed as key regulators of soil biodiversity by shaping various environmental conditions, including weathering processes, soil pH, clay formation, water infiltration, and retention capacity98,99. Plant coverage and type can also play an important role in supporting nematode diversity, as plants have been found to be biodiversity hotspots in desert ecosystems for both microorganisms (that serve as food source for nematodes) and invertebrates49,100. However, fine scale analysis of soil and plant properties is not included this study and should be consider in future work.
Latitude, widely recognized as a major determinant of taxonomic diversity, is central to the concept of the latitudinal diversity gradient (LDG)101,102. While the LDG has been empirically observed across numerous taxa103,104,105, its underlying mechanisms remain unclear. Latitude often interacts with other ecological and climatic factors, making it difficult to disentangle its direct effects from confounding influences3,106,107,108.
While climate appears to be the primary driver of nematode biodiversity in the Atacama Desert, it is important to acknowledge that this conclusion is based on the current dataset and represents an initial assessment of potential taxa distribution across this complex landscape. A more comprehensive understanding of biodiversity in the region will require systematic sampling with an increased sample size, particularly capturing diversity in high-altitude and southern areas of the desert that remain difficult to access.
Moreover, the broad patterns identified through different modeling approaches do not account for fine-scale habitat heterogeneity (e.g., water bodies, specific plant associations, or areas with persistent shade), which likely contribute to the availability of distinct ecological niches for nematodes. Despite these limitations, our analysis provides a valuable framework for understanding the potential distribution of nematode taxa in this extreme environment. By offering an initial biogeographic perspective, this study serves as a foundation for future research and targeted field experiments, ultimately enhancing our knowledge of nematode biodiversity in the Atacama Desert.
Asexual nematodes are more prevalent in higher altitudes in the Atacama
The hypothesis of geographical parthenogenesis posits that at high latitudes, high altitudes and in extreme environments, asexually reproducing taxa are more prevalent when compared to sexually reproducing ones109,110,111,112,113. Testing for this hypothesis allowed us to investigate how the relationship between taxa richness and complex biological traits (reproduction) evolved within some nematodes under globally extreme environmental conditions, finding that with increasing elevation, soil nematodes in the Atacama are more likely to reproduce asexually than sexually is supported by the Red Queen Hypothesis (RQH). The RQH suggests that in environments with high biotic interactions, sexual taxa are at advantage, while in harsh conditions with lower interactions, such as high altitudes, asexually reproducing taxa are more prevalent114,115. While our findings appear to indicate that even in an environment with generally harsh (or extreme) conditions, as the Atacama Desert, parthenogenetic taxa are most successful at the margins, we note that larger sampling sizes across multiple taxa will be needed for our understanding of geographical parthenogenesis as a phenomenon in nature. At the very least our data on the reproductive mode of soil nematodes demonstrates that the evolution of biological traits is influenced by ecological, edaphic and geographic conditions in very stable environments, like the Atacama. On a large time-scale the Atacama shows a remarkable climatic stability (aridity for over 150 Ma), showing no significant latitudinal movement since the late Jurassic15. However, on shorter time-scales climatic variation is present, our results reveal nematode diversity across different levels of biological organization but do not provide insights into abundance, that eventually can be highly variable across seasons (e.g., during the Altiplano rainfall season). Nevertheless, we expect that taxonomic diversity remains relatively stable throughout the year for the localities and sampling points analyzed, given that several taxa are capable of undergoing anhydrobiosis.
In summary, soils of the Atacama Desert are home to a diverse fauna of nematodes. While ecological strategies follow regional-local patterns (e.g colonizer-persister distribution), reproductive strategies and genera richness follow a global trend. Even in a system that is seen as an extreme on the environmental scale, diverse communities are distributed along the desert with more genera rich communities in areas with higher precipitation and lower latitudes. Remarkably, asexual taxa appear to be more prominent in the marginal ranges of high altitudes. Future research should prioritize systematic sampling to evaluate the robustness of the model-derived predictions we present in this study.
Our study highlights the necessity of biodiversity and ecosystem health assessments to understand the territory’s biological uniqueness. The Atacama has been historically explored for its notable mineral richness; consequently, mining has been a common activity throughout the desert, threatening the stability of unique ecosystems and the communities that protect and manage these territories116. Focusing on biodiversity research in the Atacama Desert would not only guide our understanding of biological evolution but also our understanding of the limits to life under global change.
Methods
Sampling, extraction and morphological identification
All research conducted in this study complies with relevant ethical regulations. Fieldwork and sampling were carried out under the required permits and permissions, including authorization from the Corporación Nacional Forestal (CONAF; Permit 09/2023) for work in Salar de Huasco. The study was conducted in full cooperation with local communities, including the Comunidad Lickan Antay de Toconao and the Comunidad Indígena Aymara Laguna del Huasco, who granted access to their territories. Local researchers were actively involved in the research process. A field school was conducted in Collaboration with Chilean researchers, university and school students where training on sampling methods, molecular work and data analysis was conducted. Additionally, reports on the results obtained were provided to the Corporación Nacional Forestal in Spanish as soon as the results were obtained. The research was designed to be locally relevant, addressing environmental and conservation questions of importance to the communities. Local and regional literature and prior studies conducted in the area were carefully considered and cited to ensure contextual relevance.
Samples were taken in six different localities across the desert. Here we define a locality as an environmentally distinct area, identified based on visible ecological and landscape features. We determined them based on direct field observation and previously defined priority sites in ref. 117, considering factors such as dominant vegetation, presence of water bodies and landforms while considering habitat suitability for nematodes. We named the localities as Altiplano (ALT), Aroma (ARO), Eagle Point (EPT), Salar de Huasco and Laguna Grande (Salars), Paposo (PAP) and Totoral Dunes (TDT). Sampling in the protected Salar de Huasco was conducted under the authorization (09/2023) of the Corporación Nacional Forestal (CONAF). In this study, 7 samples from Altiplano, 11 samples from Salars, 15 samples from Paposo, 20 samples from Totoral Dunes, 23 samples from Aroma and 21 samples from the Eagle point were analyzed (supplementary Tables S1–S6). Other areas sampled where no nematodes were found and are not part of the defined localities are later on included in the modeling process (supplementary Table S7).
Soil samples of approximately 500g were taken in the different locations using a shovel, from the upper 0–10 cm and up to 30 cm of each site and stored in zip-lock bags. Samples taken near to plants corresponded to sediment ~10 cm away from the plant stem, or from plant litter in the case of Paposo. They were initially checked for the presence/absence of nematodes on-site by extracting a portion from the soil samples using seeding trays with one layer Kimberly-Clark Kimtech science precision wipes 7551. Trays were flooded from the bottom and samples were allowed to re-hydrate overnight. The water was then filtered through sieves of 80 and 120 μm. Nematodes were collected on petri dishes and examined using a Zeiss Stemi 2000 microscope; this procedure was also followed in a laboratory setting for further morphological and molecular work.
Adult nematodes were morphologically identified using a Zeiss Axioplan 2 light microscope and further examined at 10X, 20X and 40X following multiple diagnostic features as shown in supplementary Fig. S2. Examined individuals were categorized as morphotypes to account for morphological variation within nematode families. Morphotype representatives for each family found on the samples were stored in 10μl of nuclease-free water and stored at -20 degrees Centigrade for further processing.
DNA extraction, barcoding and sequence processing
DNA extraction was performed on whole single individual nematodes using a modified version of the HotSHOT DNA extraction method118. PCR was performed using the Cytiva PuReTaq Ready-To-Go™ PCR Beads or REDTaq® ReadyMix™ PCR-mix. The genetic marker 18S rRNA was amplified using the forward (CGCGAATRGCTCATTACAACAGC) and reverse (GGGCGGTATCTGATCGCC) primers described in ref. 119, targeting the V2, V3 and V4 regions. The thermocycling conditions used were: denaturation at 94 °C for 5 min, 35 cycles of amplification (94 °C for 30 s; 54 °C for 30 s; 72 °C for 70 s), followed by a final extension at 72 °C for 5 min. Gel electrophoresis was performed (1.5% gel) to assess the amplicons. PCR products were purified using the Exo-CIP™ Rapid PCR Cleanup Kit (NEB - E1050L). DNA samples were submitted for sequencing to Eurofins Genomics and Genewiz.
Sequences were analyzed using Geneious Prime® (v. 2024.0.5). Only sequences with at least 20% high-quality bases (HQ%) were kept for further processing. Sequences were trimmed at the ends (regions with more than 1% chance of an error per base) and sequence similarity was analyzed using BLAST+ (v. 2.12.0) implementing the curated 18S-NemaBase120 using the blastn function incorporated in Geneious Prime. Best hits were assigned according to the lowest E-value and a high percentage of identity (list with accession numbers and sample information can be found on 10.5281/zenodo.15517342).
Genetic diversity and haplotype uniqueness
Nematode families represented by five or more 18s rRNA sequences from different samples and present in more than one locality were used to generate haplotype networks. Sequences were grouped based on their sequence similarity to reference sequences of the curated 18S-NemaBase database120, identified to family and if possible genus level. Due to lack of resolution in the sequences, some haplotype networks were created based on family level instead of genus level. The grouped sequences were aligned (multiple alignments) using Clustal Omega (v. 1.2.3). Short sequences (<300 bp) or with lack of overlap were removed. Alignments were manually trimmed in Geneious prime according to sequence overlap. Trimmed and curated alignments were used as input for DnaSP (v. 6.12.03)121 to calculate nucleotide diversity (π) and the Watterson’s estimator (θ). Haplotype data files were created in DnaSP and according to this trait matrices defining the occurrences of the different haplotypes were generated and then used as input for PopART (v. 1.7)122. Haplotype networks were constructed using the TCS inference method123. Networks were finalized using Inkscape (v. 1.3.2). Sequence sets were defined in DnaSP to estimate π for each location where enough sequences were present (alignments, haplotype matrix and final output can be found on https://doi.org/10.5281/zenodo.15517342).
Community composition throughout the Atacama
We estimated the Jaccard disimilarity index (β diversity) using presence/absence data of families found in 6 localities (Altiplano, Aroma, Eagle Point, Salars, Paposo and Totoral Dunes), here we analyzed aggregates of communities per locality and do not discriminate by soil sample (bag). We used the vegan package (v 2.6-4)124 and visualized the results inverted (Jaccard similarity) as a heatmap using pheatmap (v. 1.0.12)125 in R. We further on analyzed family co-occurrence using the cooccur function of the cooccur package (v. 1.3)126. To test for isolation-by-distance patterns, we estimated distance between all combinations of sampling localities on centroids using the sf (v. 1.0-16)127,128 and geodist (v. 0.1.0)129 packages. Following, a mantel test was performed comparing Jaccard dissimilarity to geographical distance using the Mantel function of the vegan package with the Spearman’s rank correlation ρ. The nucleotide diversity for each location of the different groups was also tested for isolation-by-distance patterns using the same approach. Nucleotide diversity was transformed to genetic distance by estimating the Euclidean distance using the base R package stats130.
Nematodes as soil indicators
Presence-absence information at the Family level for each locality was used as input for the application NINJA: Nematode INdicator Joint Analysis41. We assessed the coloniser-persister structure assemblage of soil nematode families found in the Atacama Desert as well as the feeding type composition in each of the sampling localities.
Modeling ecological constraints of nematode biodiversity
We evaluated climatic (environmental, e.g. precipitation) and geomorphological (edaphic and geographic, e,g. topographic complexity and latitude respectively) variables as predictors for genera richness in the Atacama Desert. We tested 57 probabilistic models using generalized least squares linear regression from the package nlme (v. 3.1)131,132 on R and 57 linear mixed effect models using lme from the package nlme, using the sampling localities as a random effect. Data on genera presence in the Atacama Desert was obtained as described through the morphological and genetic identification of the nematodes. The geomorphological variables evaluated were elevation, topographic complexity, curvature, soil thickness, rock type and latitude (supplementary Table S13). We obtained elevation data from a digital elevation model at 15 arc sec resolution133 and calculated a) topographic complexity as the standard deviation of elevation in a coarser 0.01 degree resolution grid134 and b) curvature as the second derivative of elevation with respect to the aspect in a 3 × 3 grid135 as implemented in the RichDEM package (v. 0.0.3)136 in Python. Soil thickness and rock-type data were obtained from publicly available datasets compiled respectively by Pelletier et al.,137 and Hartmann et al.,138. The evaluated climatic variables were mean annual temperature, mean annual precipitation, temperature range, and precipitation range. Here, ’range’ is defined as the difference between seasonal (rather than daily) minimum and maximum values, serving as a proxy for climatic heterogeneity. Temperature and precipitation data were obtained from the openly accessible Chelsea version 2 climate dataset139,140. A correlation matrix plot was obtained using the cor and corrplot functions from the corrplot package (v. 0.92)141 to assess variable correlations ( > ∣0.65∣) (Supplementary Fig. S9), only a representative variable from highly correlated pairs was included in the analysis to avoid redundancy. We furhter conducted a principle component analysis using the prcomp function of the stats package (v. 4.3.2)130, the PCA was visualized using the factoextra package (v. 1.0.7)142 in R. We log-transformed soil thickness (supplementary Fig. S8) to fulfill model assumptions. Bivariate relationships between genus richness and each numeric environmental predictor were explored with faceted scatter-plots to obtain the coefficient of determination (R2) using broom (v. 1.0.7)143. Bioinformatic pipelines were optimized using ChatGPT.
We evaluated a total of 114 models, including null models (Supplementary Tables S14 and S15), which incorporated various combinations of selected predictive variables. Half of the tested models were least squares models (GLS), while the other half accounted for the same variables but included localities as a random effect to control for influence of the localities (linear mixed effect models–LME). The variables were selected based on two criteria: 1) variables should not display strong correlation with each other, and 2) the variable contribution to explain the variability along the first and second principal components must be high and among the top variables (supplementary Fig. S10 and S13). Furthermore, the residuals of each model fit were visually evaluated and a variance structure adjustment was performed where necessary following144. The preferred predictive models per approach (GLS and LME) were then selected based on the Bayesian information criterion (BIC) and further estimation of Akaike weights (wm)145, as the latter provides the probability that a model is the best model according to the set of models tested and the experimental data146. The preferred predictive models were then used to obtain a map representing the possible nematode diversity in terms of genera richness. We further on tested variable importance of the previously used variable set using the machine learning algorithm Random Forest using the package randomForest (v. 4.7-1.1)147 to validate the results obtained in the linear models. Training and testing set were created using the rsample (v. 1.2.1). The model was then evaluated using caret (v. 6.0-94). Partial dependence plots (PDPs) were created to visualize the modeled effects of key environmental variables on predicted genus richness using the fitted random forest model using the pdp package (v. 0.8.2)148.
Reproductive mode determination and association to environmental variables
Laboratory cultures were established from nematodes extracted from the soil samples. Worms were propagated on nematode growth medium (NGM) plates containing E. coli strain OP50 as a food source. Single-strain established cultures were analyzed morphologically to identify males and females of a strain. Cultures with only females were categorized as possible parthenogens, and cultures with both males and females present were categorized as possible gonochoristic. In addition, single juvenile stage nematode cultures were propagated (to ensure no mating) to determine the reproductive mode of the strains as described in ref. 149. Between 5 to 30 single juvenile cultures were set up for reproductive mode corroboration per strain. We consider asexual cultures where larvae grew into adults and laid eggs without fertilization from a male.
Models testing elevation, precipitation and mean annual precipitation as predictors for reproductive mode (sexual or asexual). Environmental data was collected as in the previous section. Two model types were employed: (a) a logistic regression analysis using the glm function from the stats package in R (v. 4.3.2), with a binomial family distribution (family = binomial), and (b) a generalized linear mixed-effects model, implemented with glmer from the lme4 package (v. 1.1-36), incorporating localities as a random effect to account for spatial variability in the samples. Following, additional models were tested including vegetation coverage and percentage of phanerophytes estimated for the samples where reproductive mode could be assessed. Plant identification was done through morphological identification. We further checked which variables were most important by using a Random Forest model with the randomForest package (v. 4.7-1.1)147, to confirm the findings from the logistic regression and linear mixed-effect model. The data was split into training and testing sets with rsample (v. 1.2.1), and the model’s performance was then tested using caret (v. 6.0-94).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The sequencing data generated in this study are deposited on GenBank under accession numbers PQ587582 [https://www.ncbi.nlm.nih.gov/nuccore/PQ587582]–PQ587967 [https://www.ncbi.nlm.nih.gov/nuccore/PQ587967]. Accession numbers per sample can also be found in the Zenodo repository [https://doi.org/10.5281/zenodo.13880073] within the file List_of_Families_Genera_Haplotypes_with_accession_20250225.csv.
Code availability
The code implemented in this study along with the data required to run the code can be found on GitHub https://github.com/lauraivillegasr/BiogeographyDesertand is also available as a reproducible run on CodeOcean under the https://doi.org/10.24433/CO.6395549.v3.
References
Nisa, R. U. et al. Influence of ecological and edaphic factors on biodiversity of soil nematodes. Saudi J. Biol. Sci. 28, 3049–3059 (2021).
Döge, Jd. S., de Oliveira, H. V. & Tidon, R. Rapid response to abiotic and biotic factors controls population growth of two invasive drosophilids (Diptera) in the Brazilian Savanna. Biol. Invasions 17, 2461–2474 (2015).
Gaston, K. J. Global patterns in biodiversity. Nature 405, 220–227 (2000).
Alvial, I. E., Orth, K., Durán, B. C., Álvarez, E. & Squeo, F. A. Importance of geochemical factors in determining distribution patterns of aquatic invertebrates in mountain streams south of the Atacama Desert, Chile. Hydrobiologia 709, 11–25 (2012).
Marshall, D. J., Krug, P. J., Kupriyanova, E. K., Byrne, M. & Emlet, R. B. The Biogeography of Marine Invertebrate Life Histories. Annu. Rev. Ecol. Evolut. Syst. 43, 97–114 (2012).
Zhang, X., Ferris, H., Mitchell, J. & Liang, W. Ecosystem services of the soil food web after long-term application of agricultural management practices. Soil Biol. Biochem. 111, 36–43 (2017).
Lal, R., Negassa, W. & Lorenz, K. Carbon sequestration in soil. Curr. Opin. Environ. Sustain. 15, 79–86 (2015).
Anthony, M. A., Bender, S. F. & van der Heijden, M. G. A. Enumerating soil biodiversity. Proc. Natl. Acad. Sci. USA 120 https://doi.org/10.1073/pnas.2304663120 (2023).
Wall, D. H. et al. Soil Ecology and Ecosystem Services (Oxford University Press, United Kingdom, 2012).
Bardgett, R. D. & van der Putten, W. H. Belowground biodiversity and ecosystem functioning. Nature 515, 505–511 (2014).
Ward, D.The Biology of Deserts (Oxford University Press, 2016). https://doi.org/10.1093/acprof:oso/9780198732754.001.0001.
Ma, J. et al. Most high mountainous areas around the world present elevation-dependent aridification after the 1970s. Earths Future 12 (2024).
Chai, R. et al. Human-caused long-term changes in global aridity. Npj Clim. Atmos. Sci. 4 (2021).
Burrell, A. L., Evans, J. P. & De Kauwe, M. G. Anthropogenic climate change has driven over 5 million km2 of drylands towards desertification. Nat. Commun. 11 https://doi.org/10.1038/s41467-020-17710-7 (2020).
Hartley, A. J., Chong, G., Houston, J. & Mather, A. E. 150 million years of climatic stability: evidence from the Atacama Desert, northern Chile. J. Geol. Soc. 162, 421–424 (2005).
Jordan, T. E., Kirk-Lawlor, N. E., Blanco, N., Rech, J. A. & Cosentino, N. J. Landscape modification in response to repeated onset of hyperarid paleoclimate in the Atacama Desert, Chile. Geol. Soc. Am. Bull. 126, 1016–1046 (2014).
Voigt, C., Klipsch, S., Herwartz, D., Chong, G. & Staubwasser, M. The spatial distribution of soluble salts in the surface soil of the Atacama Desert and their relationship to hyperaridity. Glob. Planet. Change 184, 103077 (2020).
Zhang, K. et al. Salinity Is a Key Determinant for Soil Microbial Communities in a Desert Ecosystem. mSystems 4 https://doi.org/10.1128/msystems.00225-18 (2019).
Xi, H. et al. Effects of water and salinity on plant species composition and community succession in Ejina Desert Oasis, northwest China. Environ. Earth Sci. 75 https://doi.org/10.1007/s12665-015-4823-7 (2016).
Pereira, C. S., Lopes, I., Sousa, J. P. & Chelinho, S. Effects of NaCl and seawater induced salinity on survival and reproduction of three soil invertebrate species. Chemosphere 135, 116–122 (2015).
Knief, C. et al. Tracing elevational changes in microbial life and organic carbon sources in soils of the Atacama Desert. Glob. Planet. Change 184, 103078 (2020).
Morales, J. F. “Sinopsis de las Apocynaceae S. Str.: (Apocynoideae y Rauvolfioideae) de Chile”. Darwiniana, nueva serie 1 http://www.scielo.org.ar/scielo.php?script=sci_arttext&pid=S0011-67932013000100004&lng=es&nrm=iso (2013).
de Bienes Nacionales, M. Crea parque nacional “Salar del Huasco", en la Región de Tarapacá. Deja sin efecto los DS N° 24 de 2019 y N° 5 de 2022, no tramitados. Ley 19300 https://www.bcn.cl/leychile/navegar?idNorma=1189761 (2022).
Maza, F. et al. Soil Bacterial Communities From the Chilean Andean Highlands: Taxonomic Composition and Culturability. Front. Bioeng. Biotechnol. 7 https://doi.org/10.3389/fbioe.2019.00010 (2019).
Arndt, H. et al. Mirroring the effect of geological evolution: protist divergence in the Atacama Desert. Glob. Planet. Change 190, 103193 (2020).
Gómez-Silva, B. & Batista-García, R. A. The Atacama Desert: a biodiversity hotspot and not just a mineral-rich region. Front. Microbiol. 13 https://doi.org/10.3389/fmicb.2022.812842 (2022).
Eshel, G. et al. Plant ecological genomics at the limits of life in the Atacama Desert. Proc. Natl. Acad. Sci. USA 118 https://doi.org/10.1073/pnas.2101177118 (2021).
Eduardo Palma, R., Marquet, P. A. & Boric-Bargetto, D. Inter- and intraspecific phylogeography of small mammals in the Atacama Desert and adjacent areas of northern Chile. J. Biogeogr. 32, 1931–1941 (2005).
Marín, C., Rubio, J. & Godoy, R. Chilean blind spots in soil biodiversity and ecosystem function research. Austral Ecol. https://doi.org/10.1101/2021.06.24.449754 (2021).
Acosta, E., Fincke, V., Nitsche, F. & Arndt, H. Novel cercozoan and heterolobosean protists from the rhizosphere and phyllosphere of two endemic cacti from the Atacama Desert. Eur. J. Protistol. 91, 126034 (2023).
Hakobyan, A. et al. Tillandsia landbeckii phyllosphere and laimosphere as refugia for bacterial life in a hyperarid desert environment. Microbiome 11 https://doi.org/10.1186/s40168-023-01684-x (2023).
Zúñiga-Reinoso, A. & Predel, R. Past climatic changes and their effects on the phylogenetic pattern of the Gondwanan relict Maindronia (Insecta: Zygentoma) in the Chilean Atacama Desert. Glob. Planet. Change 182, 103007 (2019).
Pizarro-Araya, J., Alfaro, F. M., Ojanguren-Affilastro, A. A. & Moreira-Muñoz, A. A Fine-Scale Hotspot at the Edge: Epigean Arthropods from the Atacama Coast (Paposo-Taltal, Antofagasta Region, Chile). Insects 12, 916 (2021).
Nitsche, F. et al. Gregarines from darkling beetles of the Atacama Desert, Atacamagregarina paposa gen. et sp. nov. from Scotobius and Xiphocephalus ovatus sp. nov. from Psectrascelis (Coleoptera, Tenebrionidae). Eur. J. Protistol. 90, 126008 (2023).
Gadea, E. Nota sobre Nematodos muscícolas de Atacama (Chile). Miscelanea Zoologica 1-5 https://other.museucienciesjournals.cat/en/other/mz/mz-volum-01-5-1963/nota-sobre-nematodos-muscicolas-de-atacama-chile (1963).
Edgington, S. et al. Heterorhabditis atacamensis n. sp. (Nematoda: Heterorhabditidae), a new entomopathogenic nematode from the Atacama Desert, Chile. J. Helminthol. 85, 381–394 (2010).
Jimenez Roco, M. Contribución al conocimiento de los Nematodos del Departamento de Arica. (Segunda Parte). IDESIA 2, 53–58 (1972).
Yeates, G. W. Nematodes as soil indicators: functional and biodiversity aspects. Biol. Fertil. Soils 37, 199–210 (2003).
Satjarak, A. et al. Microscopic and Metagenomic Evidence for Eukaryotic Microorganisms Associated with Atacama Desert Populations of Giant Equisetum. Am. Fern J. 111 https://doi.org/10.1640/0002-8444-111.2.86 (2021).
Wood, J. R. et al. Ancient parasite DNA from late Quaternary Atacama Desert rodent middens. Quat. Sci. Rev. 226, 106031 (2019).
Sieriebriennikov, B., Ferris, H. & de Goede, R. G. NINJA: An automated calculation system for nematode-based biological monitoring. Eur. J. Soil Biol. 61, 90–93 (2014).
Zeppilli, D. et al. Characteristics of meiofauna in extreme marine ecosystems: a review. Mar. Biodivers. 48, 35–71 (2017).
Yeates, G. W. Two Terrestrial Nematodes from the McMurdo Sound Region, Antarctica, with a Note on Anaplectus arenicola Killick, 1964. J. Helminthol. 44, 27–34 (1970).
Maslen, N. & Convey, P. Nematode diversity and distribution in the southern maritime Antarctic—clues to history? Soil Biol. Biochem. 38, 3141–3151 (2006).
Borgonie, G. et al. Nematoda from the terrestrial deep subsurface of South Africa. Nature 474, 79–82 (2011).
De Ley, P. Past and present highlights of nematode research in the Mojave Desert. Mojave Natl Preserv. Sci. Newsl. 4, 5–10 (2014).
Whitford, W. G. & Steinberger, Y. Chihuahuan Desert Soil Biota. Open J. Ecol. 11, 581–595 (2021).
Boström, S. & Holovachov, O. Description of one new species of Heterocephalobellus Rashid, Geraert and Sharma, 1985 (Rhabditida: Cephalobidae) from Kelso Dunes, Mojave National Preserve, California, USA and Usno, Argentina. J. Nematode Morphol. Syst. 16, 161–166 (2013).
Treonis, A. M., Marais, E. & Maggs-Kölling, G. Nematode communities indicate diverse soil functioning across a fog gradient in the Namib Desert gravel plains. Ecol. Evolut. 12 https://doi.org/10.1002/ece3.9013 (2022).
Schiffer, P. H. et al. Signatures of the Evolution of Parthenogenesis and Cryptobiosis in the Genomes of Panagrolaimid Nematodes. iScience 21, 587–602 (2019).
Hörandl, E. The complex causality of geographical parthenogenesis. N. Phytol. 171, 525–538 (2006).
Bull, A. T., Asenjo, J. A., Goodfellow, M. & Gómez-Silva, B. The Atacama Desert: Technical Resources and the Growing Importance of Novel Microbial Diversity. Annu. Rev. Microbiol. 70, 215–234 (2016).
Castro-Severyn, J. et al. Living to the High Extreme: Unraveling the Composition, Structure, and Functional Insights of Bacterial Communities Thriving in the Arsenic-Rich Salar de Huasco Altiplanic Ecosystem. Microbiol. Spectrum 9 https://doi.org/10.1128/spectrum.00444-21 (2021).
Moat, J. et al. Seeing through the clouds—mapping desert fog oasis ecosystems using 20 years of MODIS imagery over Peru and Chile. Int. J. Appl. Earth Observ. Geoinf. 103, 102468 (2021).
Contreras, M. J. et al. Commonalities between the Atacama Desert and Antarctica rhizosphere microbial communities. Front. Microbiol. 14 https://doi.org/10.3389/fmicb.2023.1197399 (2023).
Jerez, V. Diversidad y patrones de distribución geográfica de insectos coleópteros en ecosistemas desérticos de la región de Antofagasta, Chile. Revista chilena de historia natural 73 https://doi.org/10.4067/S0716-078X2000000100009 (2000).
Pizarro-Araya, J. & Jerez, V. Distribución geográfica del género Gyriosomus Guérin-Méneville, 1834 (Coleoptera: Tenebrionidae): una aproximación biogeográfica. Revista chilena de historia natural 77 https://doi.org/10.4067/S0716-078X2004000300008 (2004).
Bongers, T. & Ferris, H. Nematode community structure as a bioindicator in environmental monitoring. Trends Ecol. Evolut. 14, 224–228 (1999).
Jones, L. M. et al. Climate change is predicted to alter the current pest status of Globodera pallida and G. rostochiensis in the United Kingdom. Glob. Change Biol. 23, 4497–4507 (2017).
Zablocki, O., Adriaenssens, E. M. & Cowan, D. Diversity and ecology of viruses in Hyperarid Desert soils. Appl. Environ. Microbiol. 82, 770–777 (2016).
Prather, C. M. et al. Invertebrates, ecosystem services and climate change. Biol. Rev. 24, 840–856 (2012).
Cartereau, M. et al. Global bioregionalization of warm drylands based on tree assemblages mined from occurrence big data. Front. Biogeograp. 14 https://doi.org/10.21425/F5FBG56435 (2022).
on Climate Change (IPCC), I. P.Climate Change 2021 – The Physical Science Basis: Working Group I Contribution to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change (Cambridge University Press, 2023). https://doi.org/10.1017/9781009157896.
Várallyay, G. The impact of climate change on soils and on their water management. Agronomy Research—Special Issue II 385–396 (2010).
Marais, E. et al. Profiling soil free-living nematodes in the Namib Desert, Namibia. J. Arid Land 12, 130–143 (2019).
Treonis, A. M., Marais, E. & Maggs-Kölling, G. Soil nematode communities vary among populations of the iconic desert plant, Welwitschia mirabilis. Pedobiologia 103, 150943 (2024).
Steinberger, Y., Liang, W., Savkina, E., Meshi, T. & Barness, G. Nematode community composition and diversity associated with a topoclimatic transect in a rain shadow desert. Eur. J. Soil Biol. 37, 315–320 (2001).
De Ley, I., De Ley, P., Baldwin, J., Mundo-Ocampo, M. & Nadler, S. Three new species of Nothacrobeles (Nemata: Cephalobidae) from the Mojave Desert, California. J. Nematol. 4, 482–97 (1999).
De Ley, I. et al. Acromoldavicus mojavicus n. sp. (Nematoda: Cephaloboidea) from the Mojave Desert, California. Nematology 3, 343–353 (2001).
Shahina, F. & De Ley, P. Two new species of Cephalobidae from Valle de la Luna, Argentina, and observations on the genera Acrobeles and Nothacrobeles (Nematoda: Rhabditida). Fundamental Appl. Nematol. 20, 329–347 (1997).
Kreuzinger-Janik, B., Gansfort, B., Traunspurger, W. & Ptatscheck, C. It’s all about food: Environmental factors cause species-specific dispersal. Ecosphere 13 https://doi.org/10.1002/ecs2.4251 (2022).
Berg, M. P. et al. Adapt or disperse: understanding species persistence in a changing world. Glob. Change Biol. 16, 587–598 (2010).
Ptatscheck, C., Gansfort, B. & Traunspurger, W. The extent of wind-mediated dispersal of small metazoans, focusing nematodes. Sci. Rep. 8 https://doi.org/10.1038/s41598-018-24747-8 (2018).
Al-Jawad, J., Ford, J., Petavratzi, E. & Hughes, A. Understanding the spatial variation in lithium concentration of high Andean Salars using diagnostic factors. Sci. Total Environ. 906, 167647 (2024).
Paquis, P. et al. Short-term characterisation of climatic-environmental variables and microbial community diversity in a high-altitude Andean wetland (Salar de Huasco, Chile). Sci. Total Environ. 859, 160291 (2023).
Muschiol, D., Giere, O. & Traunspurger, W. Population dynamics of a cavernicolous nematode community in a chemoautotrophic groundwater system. Limnol. Oceanogr. 60, 127–135 (2015).
Bongers, T. The maturity index: an ecological measure of environmental disturbance based on nematode species composition. Oecologia 83, 14–19 (1990).
Carrasco-Lagos, P., Moreno, R., Figueroa, A., Espoz, C. & de la Maza, C.Sitios Ramsar de Chile (Secretaría Regional Ministerial (SEREMI) del Medio Ambiente de la Región Metropolitana, Facultad de Ciencias, Centro de Investigación e Innovación para el Cambio Climático y Centro Bahía Lomas de la Universidad Santo Tomás, Facultad de Ciencias Forestales y Conservación de la Naturaleza de la Universidad de Chile, Corporación Nacional Forestal (CONAF), 2015). https://portalgeo.sernageomin.cl/Salares/SALAR_DE_HUASCO/FICHA_TECNICA_COMPILADA_SALAR_DE_HUASCO.pdf.
Bongers, T. & Bongers, M. Functional diversity of nematodes. Appl. Soil Ecol. 10, 239–251 (1998).
Du Preez, G. et al. Nematode-based indices in soil ecology: application, utility, and future directions. Soil Biol. Biochem. 169, 108640 (2022).
Ruiz-Cuenca, A. & Abolafia, J. Comparative study of four known species of the genus Acrobeles von Linstow, 1877 (Nematoda, Cephalobidae) with ‘single’ and ‘double’ cuticle from coastal dunes in Spain. Journal of Helminthology 95 https://doi.org/10.1017/S0022149X21000316 (2021).
Ruiz-Cuenca, A. N. & Abolafia, J. Prevalence and distribution of nematodes from coastal Sand Dunes in the Iberian Peninsula. Coasts 3, 263–279 (2023).
Weisberg, M. Forty Years of ‘The Strategy’: Levins on Model Building and Idealization. Biol. Philos. 21, 623–645 (2007).
Guisan, A. & Zimmermann, N. E. Predictive habitat distribution models in ecology. Ecol. Model. 135, 147–186 (2000).
Zhang, J. et al. Water availability creates global thresholds in multidimensional soil biodiversity and functions. Nat. Ecol. Evolut. 7, 1002–1011 (2023).
Hu, Y. et al. Species diversity is a strong predictor of ecosystem multifunctionality under altered precipitation in desert steppes. Ecol. Indic. 137, 108762 (2022).
Zúñiga-Reinoso, A., Ritter, B. & Predel, R. The colonization of the Puna and Atacama Biogeographic Province by sister clades of Psectrascelis (Coleoptera: Tenebrionidae): synchronous expansion without spatial overlap. J. Biogeogr. 48, 1930–1940 (2021).
Schratzberger, M., Holterman, M., van Oevelen, D. & Helder, J. A Worm’s World: Ecological Flexibility Pays Off for Free-Living Nematodes in Sediments and Soils. BioScience 69, 867–876 (2019).
Yeates, G. W. Ecological and behavioural adaptations. 1–24 (CABI Publishing, 2004). https://doi.org/10.1079/9780851998183.0001.
Ricci, C. Anhydrobiotic capabilities of bdelloid rotifers. Hydrobiologia 387/387, 321–326 (1998).
Leeuwenhoek, V. On certain animalcules found in the sediment in gutters of the roofs of houses. J. Zool. 2, 207 (1702).
Rebecchi, L., Boschetti, C. & Nelson, D. R. Extreme-tolerance mechanisms in meiofaunal organisms: a case study with tardigrades, rotifers and nematodes. Hydrobiologia 847, 2779–2799 (2019).
Chen, D. et al. Regional-scale patterns of soil microbes and nematodes across grasslands on the Mongolian plateau: relationships with climate, soil, and plants. Ecography 38, 622–631 (2014).
Zander, A., Bersier, L. & Gray, S. M. Effects of temperature variability on community structure in a natural microbial food web. Glob. Change Biol. 23, 56–67 (2016).
Stager, M. et al. Temperature heterogeneity correlates with intraspecific variation in physiological flexibility in a small endotherm. Nat. Commun. 12 https://doi.org/10.1038/s41467-021-24588-6 (2021).
Hammill, E. & Dart, R. Contributions of mean temperature and temperature variation to population stability and community diversity. Ecol. Evolut. 12 https://doi.org/10.1002/ece3.8665 (2022).
Robinson, S. I., McLaughlin, rB., Marteinsdóttir, B. & O’Gorman, E. J. Soil temperature effects on the structure and diversity of plant and invertebrate communities in a natural warming experiment. J. Anim. Ecol. 87, 634–646 (2018).
Ott, R. F. How lithology impacts global topography, vegetation, and animal biodiversity: a global-scale analysis of mountainous regions. Geophys. Res. Lett. 47 https://doi.org/10.1029/2020GL088649 (2020).
Geroy, I. et al. Aspect influences on soil water retention and storage. Hydrol. Process. 25, 3836–3842 (2011).
Jones, D. L. et al. Life at the extreme: Plant-driven hotspots of soil nutrient cycling in the hyper-arid core of the Atacama Desert. Soil Biol. Biochem. 184, 109128 (2023).
Fischer, A. G. Latitudinal variations in organic diversity. Evolution 14, 64 (1960).
Wallace, A. R. Tropical nature, and other essays (Macmillan & Co., London & New York, 1878).
Qian, H., Dai, Z. & Wang, J. Strong evidence for latitudinal diversity gradient in mosses across the world. Plant Diversity 46, 537–541 (2024).
Economo, E. P., Narula, N., Friedman, N. R., Weiser, M. D. & Guénard, B. Macroecology and macroevolution of the latitudinal diversity gradient in ants. Nat. Commun. 9 https://doi.org/10.1038/s41467-018-04218-4 (2018).
Hillebrand, H. On the generality of the latitudinal diversity gradient. Am. Naturalist 163, 192–211 (2004).
Saupe, E. E. Explanations for latitudinal diversity gradients must invoke rate variation. Proc. Natl. Acad. Sci. USA 120 https://doi.org/10.1073/pnas.2306220120 (2023).
Rapoport, E.Areografía, estrategias geográficas de las especies (Fondo de Cultura Económica, 1975). https://scholar.google.com/citations?view_op=view_citation&hl=es&user=nIGYq10AAAAJ&citation_for_view=nIGYq10AAAAJ:u5HHmVD_uO8C.
Rosenzweig, M. L. Species diversity gradients: we know more and less than we thought. J. Mammal. 73, 715–730 (1992).
Vandel, A. La Parthénogénèse géographique: contribution à l’étude biologique et cytologique de la Parthénogénèse naturelle. Bull. Biologique de. la Fr. et. de. la Belgique 62, 164–281 (1928).
Alonso-Marcos, H. et al. Difference in reproductive mode rather than ploidy explains niche differentiation in sympatric sexual and apomictic populations ofPotentilla puberula. Ecol. Evolut. 9, 3588–3598 (2019).
Kearney, M. Hybridization, glaciation and geographical parthenogenesis. Trends Ecol. Evolut. 20, 495–502 (2005).
Horne, D. J. & Martens, K. Geographical parthenogenesis in European non-marine ostracods: post-glacial invasion or Holocene stability? Hydrobiologia 391, 1–7 (1998).
Law, J. H. & Crespi, B. J. The evolution of geographic parthenogenesis in Timema walking-sticks. Mol. Ecol. 11, 1471–1489 (2002).
Van Valen, L. A new evolutionary theory. Evolutionary Theory https://scholar.google.com/scholar_lookup?title=A%20new%20evolutionary%20law&journal=Evol%20Theory&volume=1&pages=1-30&publication_year=1973&author=Valen%2CL (1973).
Hartmann, M. et al. The Red Queen hypothesis and geographical parthenogenesis in the alpine hawkweed Hieracium alpinum (Asteraceae). Biol. J. Linn. Soc. 122, 681–696 (2017).
Zanetta-Colombo, N. C. et al. Impact of mining on the metal content of dust in indigenous villages of northern Chile. Environ. Int. 169, 107490 (2022).
Dunai, T. J., Melles, M., Quandt, D., Knief, C. & Amelung, W. Whitepaper: earth—evolution at the dry limit. Glob. Planet. Change 193, 103275 (2020).
Montero-Pau, J., Gómez, A. & Muñoz, J. Application of an inexpensive and high-throughput genomic DNA extraction method for the molecular ecology of zooplanktonic diapausing eggs. Limnol. Oceanogr. Methods 6, 218–222 (2008).
Floyd, R., Rogers, A. D., Lambshead, P. J. D. & Smith, C. R. Nematode-specific PCR primers for the 18S small subunit rRNA gene. Mol. Ecol. Notes 5, 611–612 (2005).
Gattoni, K. et al. 18S-NemaBase: Curated 18S rRNA Database of Nematode Sequences. J. Nematol. 55 https://doi.org/10.2478/jofnem-2023-0006 (2023).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evolut. 34, 3299–3302 (2017).
Leigh, J. W. & Bryant, D. PopArt: full-feature software for haplotype network construction. Methods Ecol. Evolut. 6, 1110–1116 (2015).
Clement, M., Snell, Q., Walke, P., Posada, D. & Crandall, K. TCS: estimating gene genealogies. In Proceedings 16th International Parallel and Distributed Processing Symposium (IEEE, 2002). https://doi.org/10.1109/IPDPS.2002.1016585.
Oksanen, J. et al. vegan: Community Ecology Package https://CRAN.R-project.org/package=vegan (2024). R package version 2.6-6.
Kolde, R.pheatmap: Pretty Heatmaps https://CRAN.R-project.org/package=pheatmap (2019). R package version 1.0.12.
Griffith, D. M., Veech, J. A. & Marsh, C. J. cooccur: Probabilistic Species Co-Occurrence Analysis in R. J. Stat. Softw. 69 https://doi.org/10.18637/jss.v069.c02 (2016).
Pebesma, E. Simple Features for R: Standardized Support for Spatial Vector Data. R. J. 10, 439–446 (2018).
Pebesma, E. & Bivand, R.Spatial Data Science: With applications in R (Chapman and Hall/CRC, 2023). https://r-spatial.org/book/.
Padgham, M. & Sumner, M. D.geodist: Fast, Dependency-Free Geodesic Distance Calculationshttps://CRAN.R-project.org/package=geodist (2024). R package version 0.1.0.
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria https://www.R-project.org/ (2023).
Pinheiro, J. C. & Bates, D. M.Mixed-Effects Models in S and S-PLUS (Springer, New York, 2000).
Pinheiro, J., Bates, D. & R Core Team. nlme: Linear and Nonlinear Mixed Effects Models https://CRAN.R-project.org/package=nlme (2024). R package version 3.1-165.
Tozer, B. et al. Global bathymetry and topography at 15 Arc Sec: SRTM15+. Earth Space Sci. 6, 1847–1864 (2019).
Roell, Y. E., Phillips, J. G. & Parent, C. E. Effect of topographic complexity on species richness in the Galápagos Islands. J. Biogeogr. 48, 2645–2655 (2021).
Evans, I. S. Land surface derivatives: history, calculation and further development. Proceedings of geomorphometry 16–20 (2013).
Barnes, R.RichDEM: Terrain Analysis Software http://github.com/r-barnes/richdem (2016).
Pelletier, J. D. et al. A gridded global data set of soil, intact regolith, and sedimentary deposit thicknesses for regional and global land surface modeling. J. Adv. Modeling Earth Syst. 8, 41–65 (2016).
Hartmann, J. & Moosdorf, N. The new global lithological map database GLiM: A representation of rock properties at the Earth surface. Geochem. Geophys. Geosyst. 13 https://doi.org/10.1029/2012GC004370 (2012).
Karger, D. N. et al. Climatologies at high resolution for the Earth’s land surface areas. Sci. Data 4 https://doi.org/10.1038/sdata.2017.122 (2017).
Karger, D. N. et al. CHELSA V2.1 (current) https://www.envidat.ch/#/metadata/chelsa-climatologies (2021).
Wei, T. & Simko, V.R package ’corrplot’: Visualization of a Correlation Matrix https://github.com/taiyun/corrplot (2021). (Version 0.92).
Kassambara, A. & Mundt, F.factoextra: Extract and Visualize the Results of Multivariate Data Analyses https://CRAN.R-project.org/package=factoextra (2020). R package version 1.0.7.
Robinson, D. Broom: An R package for converting statistical analysis objects into tidy data frames https://arxiv.org/abs/1412.3565 (2014).
Zuur, A. F., Ieno, E. N., Walker, N., Saveliev, A. A. & Smith, G. M.Mixed effects models and extensions in ecology with R (Springer New York, 2009). https://doi.org/10.1007/978-0-387-87458-6
Anderson, D. R., Burnham, K. P. & Thompson, W. L. Null hypothesis testing: problems, prevalence, and an alternative. J. Wildl. Manag. 64, 912 (2000).
Portet, S. A primer on model selection using the Akaike Information Criterion. Infect. Dis. Model. 5, 111–128 (2020).
Liaw, A. & Wiener, M. Classification and regression by randomForest. R. N. 2, 18–22 (2002).
Greenwell, B. M. pdp: An R package for constructing partial dependence plots. R. J. 9, 421–436 (2017).
Lewis, S. C. et al. Molecular evolution in Panagrolaimus nematodes: origins of parthenogenesis, hermaphroditism and the Antarctic species P. davidi. BMC Evolut. Biol. 9 https://doi.org/10.1186/1471-2148-9-15 (2009).
Acknowledgements
We would like to thank Julian Pfeiffer, Siwen Ding, Julia Caro-Valenzuela, Thalia Sotiropoulos for their support. We would like to thank the Comunidad Lickan Antay de Toconao and the Comunidad Indígena Aymara Laguna del Huasco for allowing us in their territory, as well as the Corporación Nacional Forestal (CONAF) for providing the authorization (09/2023) for Salar de Huasco. We would like to highlight the importance of collaborating with local Indigenous communities. This allows researchers to learn from their ancestral knowledge on the territories that have been under their management and protection for generations. This ancestral knowledge enriches scientific understanding, and in turn, the data produced can inform and enhance the management of their territories. Such an approach ensures high-quality scientific output that can reach beyond the academic community and have a direct impact on society. This research is part of the DFG funded collaborative research centre CRC1211 (Earth - Evolution at the Dry Limit) and was conducted within suproject B08 [grant number 268236062] awarded to A-M.W. and P.H.S., and the grant ANID/Milenio/ICN2021_044 awarded to M.L.A. The DFG ENP grant (434028868) funded P.H.S.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
O.H., A-M.W. and P.H.S. designed the research. M.L.A. facilitated sampling and provided resources. L.V.,L.P., N.W. and A.Su. performed the research. L.V., L.P., A.St. and E.A-T. analyzed the data. L.V., L.P., O.H, A-M.W. and P.H.S. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Steven Edgington, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Villegas, L., Pettrich, L.C., Acevedo-Trejos, E. et al. Geographic distribution of nematodes in the Atacama is associated with elevation, climate gradients and parthenogenesis. Nat Commun 17, 424 (2026). https://doi.org/10.1038/s41467-025-67117-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67117-5