Abstract
Characterization of the genetic and molecular architecture underlying egg production traits in chickens is essential for improving the rate of genetic gain through intensive artificial selection. Here, to explore the dynamic landscape of regulatory effects across egg-laying stages in chickens, we generated 1272 RNA-seq samples of four tissues, i.e., the hypothalamic–pituitary–ovarian axis and liver, and paired whole-genome sequence data from 358 hens. We detected 1008 genes with stage-specific regulatory effects in at least one tissue excluding the pituitary. Out of them, 12.60, 52.78 and 32.84% were mediated by alterations in cell type composition, transcriptional factor activity, and gene co-expression networks among laying stages, respectively. Out of 80 significant loci associated with egg production traits that were detected in a large population (n = 12,952), 37 and 5 colocalized by shared and stage-specific regulatory effects, respectively. Furthermore, orthologues of these colocalized genes are enriched for the heritability of reproductive traits in pigs, cattle, and humans. In summary, we provide a resource for understanding reproduction-relevant gene regulation and highlight the importance of context-specific regulatory effects in deciphering complex traits.
Similar content being viewed by others
Introduction
Chicken provides a vital source of animal protein for humans, significantly contributing to the resolution of food shortages caused by the rapid global population growth1. As an essential model organism, chickens also serve as a biological model in research on domestication2, immunology3, and developmental biology4. Moreover, throughout the egg-laying process, hens often develop various diseases—including oophoritis5, osteoporosis6, fatty liver hemorrhagic syndrome7—that similarly affect humans and other animals. Therefore, studying the molecular and genetic basis of egg laying in hens can serve as a model for understanding the mechanisms underlying these fertility-relevant complex traits.
The egg-laying cycle of hens is generally divided into three phases: Pre-laying, Peak-laying, and Late-laying stages. During the Pre-laying stage (typically between 19 and 25 weeks of age), hens gradually reach sexual maturity, accompanied by a rapid increase in egg production. This rate is sustained throughout the Peak-laying stage (typically between 26 and 55 weeks of age) before sharply declining during the Late-laying stage (after 55 weeks of age). Several studies8,9,10 have indicated that egg-laying stages are regulated by various reproductive endocrine factors and hormones, primarily through the hypothalamic-pituitary-gonadal (HPG) axis. For instance, gonadotropin-releasing hormone (GnRH) produced by the hypothalamus stimulates the pituitary gland to release follicle-stimulating hormone (FSH) and luteinizing hormone (LH)11. However, differences in gene expression and regulation within the HPG axis tissues at different egg-laying stages remain largely unknown.
Although 3028 genomic loci associated with reproductive traits have been identified in chickens (Chicken QTLdb12, release 54), most of these loci are located in non-coding genomic regions, posing challenges in elucidating their mechanisms and pinpointing causative mutations and genes. Systematic characterization of regulatory variants (e.g., expression quantitative trait loci, eQTL) has been proposed to be an essential approach to illustrate the genetic and molecular mechanisms underlying complex phenotypes13. For example, the Genotype-Tissue Expression (GTEx) project offers a framework to explore the genetic mechanisms underlying complex traits by identifying eQTL across multiple tissues in adult humans14,15. The FarmGTEx consortium extended this framework to domestic animals, such as cattle and pigs16,17. Recent studies in humans have also proposed that gene regulation is highly dependent on specific biological contexts, such as developmental stages18 and cell type compositions19. Although the pilot phase of ChickenGTEx identified tissue-specific regulatory effects20, public data still have notable limitations. First, the quality of RNA-seq data obtained from public databases is inconsistent, and several confounders are difficult to correct. Moreover, the lack of paired WGS data for most public RNA-seq samples limits the number and coverage of variants available for molQTL mapping. Another critical limitation is the lack of metadata annotations, which has impeded the comprehensive investigation of context-specific regulatory effects in the chicken GTEx pilot phase. The developmental GTEx project for humans is currently ongoing21, and previous studies have identified regulatory mutations specific to developmental stages of the human brain18, underscoring the significance of analyzing specific regulatory effects within stage context. However, studies systematically investigating context-specific regulatory effects remain limited, partly because such designs are difficult to achieve in humans but can be carefully implemented in farm animals22.
In this study, we developed a well-designed egg-laying stage-specific ChickenGTEx resource (Fig. 1A) by collecting 1272 RNA-seq samples from four tissues, including the HPG axis and liver, obtained from 358 hens across the Pre-, Peak-, and Late-laying stages. We also generated paired deep whole-genome sequencing (WGS, an average of 26.42×) data from these animals. We systematically identified regulatory effects across tissues and egg-laying stages, and investigated potential factors (e.g., changes in cell type composition, transcription factor, and gene regulatory networks) that mediate stage-specific regulatory effects. By integrating GWAS results from the same chicken population (n = 12,952), we demonstrated that the importance of stage-specific regulatory effects in illustrating the molecular mechanisms underlying body weight at the age of 49 days (BW49), age at first egg (AFE), the total number of eggs at days 210 (EN210), days 300 (EN300), days 400 (EN400) and days 210–400 (EN210-400). Ultimately, we found that the orthologues of the fine-mapped genes for chicken reproductive traits were significantly enriched in genes associated with reproductive traits in human, pig, and cattle, suggesting that the gene regulation of reproductive traits is conserved to some extent across vertebrates.
A The overall workflow of the chicken laying-stage GTEx. All samples were collected from a single commercial chicken population. In total, 358 hens representing three distinct laying stages—pre-laying (20 weeks of age), peak-laying (30 weeks), and late-laying (58 weeks)—were selected. Four tissues (hypothalamus, pituitary, ovary, and liver) were collected from each individual, generating 1272 RNA-seq datasets. In parallel, genomic DNA was extracted from blood and subjected to whole-genome sequencing (WGS) with an average depth of 26×. Based on these multi-omics data, we performed molecular quantitative trait loci (molQTL) mapping for gene expression, exon expression, enhancer activity, alternative splicing, and tissue-specific expression ratios, and investigated the temporal and tissue-specific regulatory patterns of these molQTL. In addition, low-depth whole-genome sequencing followed by genotype imputation was conducted for 12,952 hens from the same population with recorded egg-production phenotypes. Genome-wide association studies (GWAS) were performed for six complex traits, and the identified QTL were integrated with molQTL to explore their regulatory relationships. Finally, cross-species comparative analyses were carried out to investigate the conservation of reproductive trait regulation among chickens, humans, pigs, and cattle. Illustrative images were created in BioRender. Wang, Y. (2025) https://BioRender.com/fydtay2. B Hierarchical clustering was performed using the expression levels (quantified as transcripts per million, TPM) of the 4000 genes showing the highest variance across all samples, together with the 4000 splicing events exhibiting the greatest variability. This analysis illustrates the overall similarity and tissue-specific clustering patterns among the 1272 RNA-seq datasets.
Results
Data summary and molecular phenotypes
In a commercial chicken population, we genotyped 13,186 hens using low-depth sequencing technology (0.247 ± 0.07×, LDS) and recorded their body weight at 49 days of age (BW49) as well as detailed egg production traits, including age at first egg (AFE), egg number at 210 days (EN210), 300 days (EN300), 400 days (EN400), and between 210 and 400 days (EN210-400) (Figs. 1A, S1A). To study laying stage-specific regulatory effects, we selected 358 hens with considerable genetic variation at three distinct egg-laying stages (i.e., 20 weeks, 30 weeks, and 58 weeks of age) in the same population. To enhance genetic diversity and ensure a more representative sample, individuals were randomly chosen, with close genetic relationships excluded based on pedigree information. We then generated WGS data (average 26.42×) of all these 358 animals and 1309 RNA-seq samples (an average of 37.18 million clean read pairs) from their four tissues, representing the hypothalamic–pituitary–ovarian axis and liver. Over 90% of the animals had RNA-seq data from three or more tissues. (Fig. S1B). After removing potential label error samples (see “Methods”), 12,952 animals with LDS data were retained for GWAS and 1272 RNA-seq data for mapping laying-specific regulatory effects (Fig. S1C, D, Supplementary Datas 1–2). Genotype principle competent analysis (PCA) showed that the RNA-seq samples were representative of the whole population, and there was no population stratification among the samples from the three different egg-laying stages (Fig. S1E).
We quantified four molecular phenotypes from the RNA-seq data, including the expression levels of 25,016 genes, 78,530 exons, and 13,374 enhancers, and the alternative splicing abundance of 15,593 genes across tissues and stages (Supplementary Data 3). PCA and hierarchical clustering analysis based on four molecular phenotypes consistently demonstrated that RNA-seq samples were clustered by tissue types primarily, followed by laying stages (Figs. 1B, S2A,S3). The expression of genes and their corresponding exons exhibited a stronger correlation compared to alternative splicing (Fig. S2B). Additionally, we identified 8,001,349 common variants (minor allele frequency [MAF] > 0.05) from the WGS data for subsequent molecular quantitative trait loci (molQTL) mapping.
The gene expression landscape across tissues and laying stages
We identified 2256, 938, 2658, and 1972 genes with tissue-specific expression in the hypothalamus, pituitary, ovary, and liver, respectively. The functions of these tissue-specific genes corresponded to the known physiological functions of the respective tissues (Fig. 2A). For example, hypothalamus-specific genes were significantly enriched in the nervous system development pathway, while liver-specific genes were enriched in small molecule and lipid transport pathways. The time-course analysis of gene expression revealed six distinct gene clusters with varying expression patterns across egg-laying stages (Figs. 2B, S4). In general, across all the gene clusters, the hypothalamus shared more genes with the pituitary. Specifically, in the gene cluster with high expression at the Peak stage, the HPG axis exhibited a higher degree of gene expression similarity (Fig. 2B).
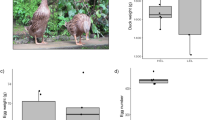
A Heatmap showing the expression patterns of tissue-specific genes (log₂FC > 2) across the four tissues. Each row represents one gene, and each column corresponds to one individual sample. A total of 2256, 938, 2658, and 1972 genes exhibited tissue-specific expression in the hypothalamus, pituitary, ovary, and liver, respectively. Colors represent standardized (z-score normalized) expression levels. The color bar above the heatmap indicates the four tissue types. The boxes display Gene Ontology (GO) biological processes significantly enriched among tissue-specific genes (P-values adjusted using the Benjamini–Hochberg method). For each tissue, only the two most significantly enriched GO biological processes are shown. B The heatmap summarizes the time-course analysis, highlighting the similarity of stage-specific expression patterns across tissues. Genes were grouped into six clusters according to their expression dynamics across the three laying stages. For example, Group 1 includes genes that are highly expressed specifically at the peak laying stage. The odds ratio represents the degree of enrichment of overlapping genes between tissues within the same expression-pattern group relative to that expected by chance. A total of 1272 samples were included in the analysis. Illustrative images of tissues were created in BioRender. Wang, Y. (2025) https://BioRender.com/pnkedie. C The number of differentially expressed genes (DEGs) across laying stages in the ovary and the corresponding Gene Ontology (GO) enrichment are shown. The boxes display Gene Ontology (GO) biological processes significantly enriched among tissue-specific genes (P-values adjusted using the Benjamini–Hochberg method). The heatmap displays the number of DEGs with higher expression in the query stage compared to the reference stage. Row labels indicate the query stages, and column labels indicate the reference stages. For example, the cell at row “Pre” and column “Peak” represents the number of genes upregulated in the pre-laying stage relative to the peak-laying stage. D Violin plots showing the expression changes of CD3D and PTPRC across the three laying stages in the ovary. The plots indicate the median (center dashed line) and the 25th and 75th percentiles. Sample sizes for the three stages are as follows: pre-laying (n = 116), peak-laying (n = 110), and late-laying (n = 115). P-values, derived from unpaired and two-sided t-tests, indicate the significance of differences in gene expression between stages. E Co-expression network modules identified in the ovary and their correlations with egg-laying stages. Numbers in parentheses indicate the number of genes within each module. Asterisks denote the significance of correlations (* P < 0.05, ** P < 0.01, *** P < 0.001), as determined by two-sided Pearson correlation tests. The panel on the right shows the results of Gene Ontology (GO) enrichment analysis for genes in module ME10.
The ovary exhibited the highest number of stage-specific expressed genes, with the majority detected in the Peak- and Late-laying stages (Figs. 2C, S5A). These genes were significantly enriched in responses to external biological invasions and immune functions (Fig. 2C). For example, the CD3D and PTPRC genes were significantly and highly expressed at the Peak and Late stages, and they are typically considered as marker genes of immune cells23 (Fig. 2D). This result suggests that hens become more susceptible to external microbial invasion after the onset of egg production.
To identify co-expression patterns among genes, we employed weighted gene correlation network analysis (WGCNA)24, and identified 20, 29, 21, and 40 modules in the hypothalamus, pituitary, ovary, and liver, respectively. Among these co-expression modules, 81.82% were significantly associated with different stages (Figs. 2E, S5B). For example, in the ovary, ME10 was positively correlated with the Pre (r = 0.815, p = 2.38e–82), but negatively correlated with the Late stage (r = −0.724, p = 2.85e–56), respectively. GO enrichment analysis revealed that genes within this module were significantly enriched in transcription-related biological processes, such as mRNA metabolic processes and RNA splicing (Fig. 2E).
Discovery and fine mapping of molQTL
The average cis-heritability of gene expression, alternative splicing, exon expression and enhancer expression was 0.16, 0.067, 0.078 and 0.076, respectively, and was consistent across stages and tissues (Fig. S6A). By conducting cis-molQTL mapping for each stage-tissue combination, we detected 20,815 (83.21%) of the 25,016 tested genes, 10,397 (66.68%) of the 15,593 genes with alternative splicing events, 40,584 (51.68%) of the 78,530 exons, and 7242 (54.15%) of the 13,374 enhancers that were significantly regulated by at least one genetic variant, referred to as eGene, exGene, sGene, and eEnhancer, respectively (Fig. 3A). The cis-heritability of eGenes is significantly higher than that of non-eGenes, and the heritability of eGenes increases with the number of independent QTL. However, there is no difference in their expression levels (Fig. S6B). Supplementary Data 4 provides a summary of the number of independent molQTL detected for each tissue, developmental stage, and molecular phenotype, along with the average confidence interval size for each QTL. These data offer a comprehensive overview of the QTL identified across different tissues and stages, as well as their estimated boundaries based on LD (r² ≥ 0.8). Combining samples from different laying stages within tissues (i.e., the merge group) led to the detection of a greater proportion of e-molecular phenotypes (Fig. 3A), with correspondingly smaller confidence intervals (Supplementary Data 4). Additionally, the eQTL detected in addition to those overlapping between single and merged stages showed smaller effect sizes (Fig. S6C). The lead variants of molQTL were enriched around the transcription start site (TSS) of genes (Fig. S6D). Additionally, eGenes had higher phastCons scores25 (which quantify the evolutionary conservation of genomic regions across species, with higher scores indicating greater constraint, and obtained from UCSC) compared to non-eGenes, indicating that eGenes are more evolutionarily constrained across species (Fig. S6E). The absolute effect sizes of lead variants of eGenes were significantly and negatively correlated with both minor allele frequency (Spearman rho = −0.23, P = 3.72e–91) and the gene’s kME (a measure of the representativeness of a gene within a module) (Spearman rho = −0.49, P = 6.31e–121) (Fig. 3B, C). This indicates that eQTL with larger effect sizes tend to have lower allele frequencies, and that their corresponding eGenes exhibit lower membership within modules, suggesting that large-effect eQTL may be subject to negative selection.
A Bar plot showing the proportion of molecular phenotypes with at least one molecular QTL, relative to the total number of tested molecular phenotypes, in each tissue–stage combination. In the hypothalamus, pituitary, ovary, and liver, 21,829, 20,783, 21,770, and 18,050 genes; 57,981, 56,137, 53,488, and 47,568 exons; 8921, 8074, 7391, and 6893 enhancers; and 64,969, 59,251, 87,063, and 48,364 alternative splicing events were analyzed, respectively. Illustrative images of tissues were created in BioRender. Wang, Y. (2025) https://BioRender.com/pnkedie. B Spearman correlation between the absolute values of eQTL effect sizes and minor allele frequency, showing only the top associated SNP for each eGene. A density contour map is overlaid to illustrate point density, with red areas indicating regions of higher density. The P-value was derived from a two-sided Spearman’s rank correlation test assessing the significance of the correlation between effect size and minor allele frequency. C Spearman correlation between the absolute values of eQTL effect sizes and module kME (module eigengene-based connectivity), showing only the top associated SNP for each eGene. The P-value was derived from a two-sided Spearman’s rank correlation test assessing the significance of the correlation between eQTL effect size and module connectivity. D Spearman correlation between the number of fine-mapped credible sets (identified using SuSiE) and the number of conditionally independent eQTL (identified using tensorQTL). The size of each dot is scaled according to the number of genes represented. The P-value was derived from a two-sided Spearman’s rank correlation test assessing the significance of the correlation. E Fine-mapping resolution of eQTL identified using SuSiE. The x-axis represents the number of SNPs within each credible set, and the y-axis represents the number of credible sets containing the corresponding number of SNPs. F Distribution of distances between the transcription start site (TSS) of each target gene and the top SNPs of molecular QTL of varying ranks obtained from the tensorQTL independence analysis. Colors represent different molQTL ranks, with molQTL of rank greater than 4 combined into a single group for visualization. G Enrichment odds ratios of molQTL before and after fine-mapping (left panel) and enrichment of credible sets containing only a single variant (right panel) within functional genomic regions. Circles represent all nominally significant variants, triangles represent variants within fine-mapped credible sets, and squares represent SNPs from credible sets containing only a single variant.
To determine whether molecular phenotypes are influenced by multiple independent molQTL, we performed conditionally independent and fine-mapping analyses using the “cis_independent” mode from tensorQTL26 and SuSiE27, respectively. As expected, the number of conditionally independent eQTL was significantly correlated with the number of credible sets (CS) identified through fine mapping (Spearman rho = 0.54, P-value = 7.31e–33; Fig. 3D). The majority of molecular phenotypes are primarily regulated by a single independent cis-molQTL (Fig. S7), and 83.53% of eGenes, 75.34% of eEnhancers, 75.41% of exGenes, and 77.39% of alternative splicing events have at least one detectable CS (Figs. 3D, S8). A total of 1612, 1524, 155, and 1290 CS in eQTL, exQTL, enQTL, and sQTL contained a single variant, respectively (Figs. 3E, S8), indicating that our fine-mapping has a high resolution. Among the molecular phenotypes influenced by multiple independent QTL, lead molQTL were more likely to be enriched around the TSS compared to the others (Fig. 3F).
All molQTL were significantly enriched in 5’UTR, 3’UTR, stop gain, and stop loss sites (Fig. 3G). Additionally, sQTL exhibited the highest level of enrichment in splice-related regions. In terms of chromatin states, all molQTL were enriched in regions associated with promoters and enhancers, with enQTL exhibiting a greater enrichment in enhancer-like regions (Fig. S9). Furthermore, fine-mapped molQTL showed a higher enrichment in functional regions compared to all the molQTL (Figs. 3G, S9). We also performed an enrichment analysis for CS containing only a single variant. The results showed that, compared to all CS, those with only one variant exhibited a further increase in enrichment in functional regions (Figs. 3G, S9).
Compared to the pilot ChickenGTEx20, we identified more eGenes and independent eQTL in the hypothalamus, pituitary, and ovary due to the larger sample size and more genotypes tested (8 M vs. 1.5 M) in this study (Fig. S10). More specifically, a total of 29.53, 34.59, 16.24, and 56.87% of eQTL could be replicated in ChickenGTEx, measured by π1 values, in the hypothalamus, pituitary, ovary, and liver, respectively. Conversely, 61.72, 71.66, 68.52, and 60.28% of eQTL in ChickenGTEx were replicated in this study (Fig. S11). We also found that for the validated eQTL (P-value < 0.05 in validation), the effect sizes of loci identified in our study were significantly correlated with those from the chicken GTEx (Fig. S11).
Tissue-sharing pattern of molQTL
Around 50% of molQTL were shared across all the tissues, whereas around 20% were detected in a single tissue (Fig. 4A). The molecular phenotypes corresponding to tissue-specific molQTL exhibited lower cis-heritability and smaller effect sizes compared to those associated with tissue-shared molQTL (Figs. 4A, S12A). The effect correlation of molecular phenotype QTL was higher between the hypothalamus and pituitary (Figs. 4B, S12B). We identified 697, 452, 1415, and 516 tissue-specific eGenes in the hypothalamus, pituitary, ovary, and liver, respectively (eGenes with an LFSR < 0.05 in only one tissue). Moreover, we found a significant overlap between tissue-specific eGenes and genes with differential expression across tissues (Hypergeometric test, P-value = 2.83e–8, Fig. S12C), with 25.79% to 29.65% of tissue-specific eGenes also exhibiting differential expression across tissues (Fig. 4C). Gene Ontology (GO) annotation revealed that tissue-specific eGenes in the ovary are significantly enriched in cellular and metabolic processes (Fig. S12D).
A The bar plot (top panel) shows the proportion of four types of molecular QTL shared across different numbers of tissues, as determined by an LFSR < 0.05 threshold obtained using MashR. The box plot (bottom panel) displays the cis-heritability of molecular phenotypes, calculated using LDAK, corresponding to molecular QTL shared across varying numbers of tissues. In the box plot, the center line represents the median, the box bounds represent the 25th and 75th percentiles, and the whiskers indicate 1.5 × the interquartile range. A total of 37,309, 27,977, 5566, and 17,454 eQTL, exQTL, enQTL, and sQTL were detected in the hypothalamus; 32,132, 27,473, 5339, and 12,460 in the pituitary; 42,668, 34,464, 5305, and 39,367 in the ovary; and 21,182, 22,260, 4616, and 10,647 in the liver, respectively. B Heatmap showing the Spearman correlation of eQTL effect sizes between tissues, where effect sizes were estimated using MashR. Only the top SNP–eGene pair was included for each eGene in this analysis. C Stacked bar plot showing the proportion of tissue-differentially expressed genes among tissue-specific eGenes across the four tissues. Tissue-differentially expressed genes were identified using the Wilcoxon rank-sum test. Tissue-specific eGenes were defined as eGenes with an LFSR < 0.05 in only one tissue, as determined by MashR. D Illustrative diagram showing the definition of gene expression ratios between tissues. For gene (a) in a tissue1–tissue2 pair, the gene expression ratio is defined as (TPM_a1 + 0.01)/(TPM_a2 + 0.01), where a small constant (0.01) is added to prevent division by zero and to stabilize the ratio for genes with low or zero expression. Only genes expressed in both tissues (TPM > 0.1 in at least 20% of samples) and samples overlapping between tissues were included in the analysis. An inverse normal transformation was applied to the ratios before QTL mapping. Illustrative images of chicken and tissues were created in BioRender. Wang, Y. (2025) https://BioRender.com/pnkedie. E Stacked bar plot showing the identification of expression ratio QTL genes (erGenes) and their overlap with expression QTL genes (eGenes) across all tissue pairs and laying stages. Blue bars represent genes with only eQTL detected, orange bars represent genes with both eQTL and erQTL detected, and red bars represent genes with only erQTL detected. Illustrative images of tissues were created in BioRender. Wang, Y. (2025) https://BioRender.com/pnkedie. F Density plot showing the distribution of PPH.4 values from coloc, representing the posterior probability that the associations of two molecular phenotypes are driven by a shared causal variant. G Associations and colocalization of genetic variants with the gene expression ratio between the hypothalamus and pituitary and with CPXM2 expression in the hypothalamus during the late laying stage. The box plot on the right shows the distributions of CPXM2 expression and expression ratios across the three genotypes of the top erQTL variant (rs733049819). PPH.4 values from Coloc are shown in the figure, representing the posterior probability that the associations for two traits (e.g., molecular QTL and GWAS signal) share a single causal variant. Sample sizes are as follows: hypothalamus (n = 113), pituitary (n = 112), and hypothalamus/pituitary ratio (n = 110). In the box plot, the center line represents the median, the box bounds represent the 25th and 75th percentiles, and the whiskers indicate 1.5 × the interquartile range. Illustrative images of tissues were created in BioRender. Wang, Y. (2025) https://BioRender.com/pnkedie. H Associations and colocalization of genetic variants with the gene expression ratio between the ovary and pituitary and with GSTA3 expression in the ovary during the peak laying stage. The box plot on the right shows the distributions of GSTA3 expression and expression ratios across the three genotypes of the top erQTL variant (rs739157915). PPH.4 values from Coloc are shown in the figure, representing the posterior probability that the associations for two traits (e.g., molecular QTL and GWAS signal) share a single causal variant. Sample sizes are as follows: ovary (n = 110), pituitary (n = 98), and ovary/pituitary ratio (n = 94). In the box plot, the center line represents the median, the box bounds represent the 25th and 75th percentiles, and the whiskers indicate 1.5 × the interquartile range. Illustrative images of tissues were created in BioRender. Wang, Y. (2025) https://BioRender.com/pnkedie.
To further explore the crosstalk of molQTL between tissues, we proposed a molecular phenotype, the between-tissue gene expression ratio, defined as the ratio of gene expression between two tissues within the same individual. We then performed erQTL (expression ratio QTL) mapping across all tissue pairs and developmental stages (Fig. 4D). Overall, across all tissue pairs and developmental stages, 1.39–6.94% of all tested genes were identified as erGenes, which exhibited a high degree of overlap with eGenes in corresponding tissues (88.85–94.86%) (Fig. 4E). However, the colocalization analysis revealed a U-shaped distribution of PPH.4 between the erQTL and their corresponding eQTL. Among all erQTL and eQTL pairs, 35.05% (PPH.4 < 0.3) could not be colocalized, while 35.26% were colocalized with PPH.4 > 0.7 (Fig. 4F). For example, between the hypothalamus and pituitary, the erQTL of CPXM2 colocalized with its eQTL in the hypothalamus (Fig. 4G, PPH.4 = 0.953). The tissue-specific eQTL of CPXM2 in the hypothalamus led to a correlation between gene expression and the expression ratio, ultimately resulting in the colocalization of erQTL and eQTL. Another example is the erQTL of GSTA3 between the ovary and pituitary, where the erQTL and eQTL of GSTA3 are controlled by different causal variants (Fig. 4H, PPH.4 = 0.043). This suggests that the correlation in gene expression levels between tissues may be influenced by genetic factors beyond the effects captured by local eQTL.
Laying stage-specific regulatory effects
By conducting a meta-analysis using MashR, we identified 65.27% to 77.83% of molQTL (77.83% of eQTL, 65.95% of exQTL, 68.30% of enQTL, and 65.27% of sQTL) shared across the three egg-laying stages. (Fig. 5A, S13A). The effects of all molQTL showed a high correlation across different stages within tissues. However, the correlation of eQTL effects between different tissues at the same stage was significantly lower than that of the other three types of molQTL (Figs. 5B, S13B). As expected, we observed that the hypothalamus and pituitary displayed higher effect correlations among all molQTL (Fig. S13B). Furthermore, molQTL shared across more stages exhibited larger effect sizes compared to stage-specific molQTL (Fig. S13C). Notably, the correlation of effect sizes between tissues for stage-specific eQTL was significantly lower than that for stage-shared eQTL (Fig. 5C), and stage-specific eGenes (LFSR < 0.05 in only one stage) also tend to be tissue-specific (Fig. 5D).
A The bar plot shows the proportion of four types of molecular QTL shared across different laying stages in the ovary, as determined by an LFSR < 5% threshold obtained using MashR. B The violin plot illustrates the correlation of four types of molecular QTL effect sizes within tissues between stages and within stages between tissues. The number of tests is as follows: within-tissue between-stage, n = 12; within-stage between-tissue, n = 18. In each violin plot, the middle dashed line represents the median, and the upper and lower dashed lines represent the 75th and 25th percentiles, respectively. C Heatmap showing the pairwise Spearman correlation of cis-eQTL effect sizes between tissues, with the upper triangle representing stage-shared eQTL and the lower triangle representing stage-specific eQTL. Effect sizes were estimated using MashR. D UpSetR plot depicting the number of stage-specific eGenes shared across the four tissues. Stage-specific eGenes were defined as eGenes detected in only one laying stage within the same tissue. E UpSetR plot depicting the number of stage-interaction eGenes (left) and eGenes (right) shared across the four tissues. The results show that stage-interaction eGenes exhibit strong tissue specificity, whereas most eGenes are broadly shared across tissues. F Stacked bar plot showing the proportion of stage-differentially expressed genes among stage-interaction eGenes. G Example of a stage-interaction eGene for PLD5 in the liver, demonstrating differential expression across the three laying stages. The three colors represent the distinct genotypes of the top SNP (rs16253696). In the box plot, the center line represents the median, the box bounds represent the 25th and 75th percentiles, and the whiskers indicate 1.5 × the interquartile range. The three colored lines indicate the fitted changes in gene expression across the laying stages for each genotype. Sample sizes are as follows: pre-laying stage (n = 106), peak-laying stage (n = 81), and late-laying stage (n = 107). H Example of a stage-interaction eGene for DFFA in the ovary, where the stage interaction is unrelated to stage differential expression. The three colors represent the distinct genotypes of the top SNP (rs16179949). In the box plot, the center line represents the median, the box bounds represent the 25th and 75th percentiles, and the whiskers indicate 1.5 × the interquartile range. The three colored lines indicate the fitted changes in gene expression across the laying stages for each genotype. Sample sizes are as follows: pre-laying stage (n = 116), peak-laying stage (n = 110), and late-laying stage (n = 115).
Furthermore, by conducting stage-interaction molQTL mapping, we identified 1289 stage-ieGenes, 988 stage-ieExons, 123 stage-ieEnhancers, and 685 stage-isGenes that were identified in at least one tissue (Figs. 5E, S14). The MAF of the lead interacting variants showed a very high correlation across the three stages (Correlation Coefficient: 0.88–0.93) (Fig. S15), indicating that the identified interaction effects were not caused by MAF differences among stages. Consistent with the results from MashR, molecular phenotypes with stage interaction QTL also tend to be tissue-specific (Figs. 5E, S14). For example, in the hypothalamus, pituitary, ovary, and liver, only 22.97%, 10.79%, 6.64%, and 8.42% of ieGenes, respectively, are shared with other tissues. However, when considering all eGenes, the majority (73.65%) are shared across two or more tissues (Fig. 5E). Similar to tissue-specific, 19.41–51.36% of stage-ieGenes were also differentially expressed between stages (Fig. 5F). For example, the PLD5 in the liver exhibited very low expression levels at Pre-stage, resulting in a smaller eQTL effect size, which gradually increased with rising expression levels at the Peak and Late laying stages (Fig. 5G). In contrast, some stage interactions could not be explained by stage-specific differentially expressed. For instance, in the ovary, the eQTL effect of the DFFA gradually decreased as the laying stages progressed without a significant change in expression level (Fig. 5H).
The biological mediators of stage interaction eQTL
We obtained cell proportions in the hypothalamus, ovary, and liver by deconvolution of bulk RNA-seq data (Fig. S16). After filtering out rare cell types, we identified 11 cell types with significant proportion differences across at least two egg-laying stages (Fig. S17). For example, in the ovary, the proportions of fibroblasts and macrophages were significantly higher during the Peak and Late-laying stages. This is consistent with the significant enrichment of immune-related biological processes in genes highly expressed during the Peak and Late stages (Fig. 2C). We then further analyzed the relationships between stage-specific ieGenes and three biological contexts—cell composition, transcription factors, and co-expression modules—using both causal inference test (CIT)28 and mediation29 methods (Fig. 6A). By comparing the two approaches, we found that the mediation method provided higher power (Fig. S18); therefore, we retained only the results from the mediation analysis in the final report (Fig. 6A). Among the 1008 stage-specific ieGenes detected in tissues other than the pituitary (due to the lack of single-cell RNA-seq data for this tissue), we found that 60.02% were significantly mediated by at least one biological factor (P-value ACME < 0.01, Fig. 6B). The full list of significant mediation effects is provided in Supplementary Data 5.
A Schematic illustrating the mediation model framework used in this study. In this model, X represents the interaction term between stage and genotype (stage × G), Y denotes the target gene expression, and M represents the mediator, which includes three classes of biological factors: cell component proportions, transcription factor (TF) expression, and module eigengenes from co-expression networks. TE (total effect) refers to the overall effect of X on Y; ADE (average direct effect) represents the effect of X on Y that is not mediated by M; and ACME (average causal mediation effect) quantifies the portion of the effect of X on Y that is mediated through M. B UpSetR plot depicting the number of stage-interaction eGenes mediated by the three biological contexts. To objectively reflect the proportion across tissues with available data, the pituitary was excluded from this analysis due to the absence of single-cell data. C Example of a stage-interaction eGene for THBS2 in the liver, mediated by the endothelial cell component. The three colors represent the distinct genotypes of the top SNP (rs731944622). The y-axes of the left and middle panels show THBS2 expression levels (log₂(TPM + 0.25)). The x-axis of the left panel represents the three laying stages, while the x-axis of the middle panel represents the endothelial cell component proportion. In the box plot, the center line represents the median, the box bounds represent the 25th and 75th percentiles, and the whiskers indicate 1.5 × the interquartile range. The right panel shows the effect sizes from the mediation model, including the Total Effect (TE), Average Direct Effect (ADE), and Average Causal Mediation Effect (ACME) and P-values were calculated to assess the statistical significance of each effect. Bars indicate the 95% confidence interval bounds of each effect estimate. Sample sizes are as follows: pre-laying stage (n = 106), peak-laying stage (n = 81), and late-laying stage (n = 107). D Example of a stage-interaction eGene for SRGAP1 in the ovary, mediated by the transcription factor TFEC. The three colors represent the distinct genotypes of the top SNP (rs313000832). The y-axes of the left and middle panels show SRGAP1 expression levels (log₂(TPM + 0.25)). The x-axis of the left panel represents the three laying stages, while the x-axis of the middle panel represents TFEC expression levels. In the box plot, the center line represents the median, the box bounds represent the 25th and 75th percentiles, and the whiskers indicate 1.5 × the interquartile range. The right panel shows the effect sizes from the mediation model, including the Total Effect (TE), Average Direct Effect (ADE), and Average Causal Mediation Effect (ACME) and P-values were calculated to assess the statistical significance of each effect. Bars indicate the 95% confidence interval bounds of each effect estimate. Sample sizes are as follows: pre-laying stage (n = 116), peak-laying stage (n = 110), and late-laying stage (n = 115). E Example of a stage-interaction eGene for ZNF385D in the pituitary, mediated by the eigengene of co-expression module ME21. The three colors represent the distinct genotypes of the top SNP (rs315127948). The y-axes of the left and middle panels show ZNF385D expression levels (log₂(TPM + 0.25)). The x-axis of the left panel represents the three laying stages, while the x-axis of the middle panel represents the eigengene value of the co-expression module ME21. In the box plot, the center line represents the median, the box bounds represent the 25th and 75th percentiles, and the whiskers indicate 1.5 × the interquartile range. The right panel shows the effect sizes from the mediation model, including the Total Effect (TE), Average Direct Effect (ADE), and Average Causal Mediation Effect (ACME) and P-values were calculated to assess the statistical significance of each effect. Bars indicate the 95% confidence interval bounds of each effect estimate. Sample sizes are as follows: pre-laying stage (n = 99), peak-laying stage (n = 98), and late-laying stage (n = 112).
Among them, 12.60% of stage-interacting eQTL were mediated by cell composition (Fig. 6B). For example, in the liver, the stage-interacting eQTL of THBS2, which encodes an extracellular matrix (ECM) glycoprotein that inhibits blood vessel and endothelial cell formation30, was mediated by the proportion of endothelial cells (Fig. 6C). Moreover, a total of 52.78% of stage-specific ieGenes were mediated by transcription factors. For example, in the ovary, the stage-interacting eQTL of SRGAP1 was mediated by the expression of TFEC, and the effect of the eQTL varied according to the expression levels of the transcription factor, consistent with the patterns observed across different egg-laying stages (Fig. 6D). Additionally, 32.84% of stage-specific ieGenes were mediated by co-expression module eigengenes. For example, the stage-interacting eQTL of ZNF385D in the pituitary was significantly mediated by the eigengene of module ME21 (Fig. 6E) and this module was significantly negatively correlated with the Pre stage (cor = −0.664, P-value = 1.1e–40) and positively correlated with the Late stage (cor = 0.928, P-value = 2.74e–133) (Fig. S5B). Furthermore, we found that some ieGenes are mediated by multiple biological contexts. For instance, 49.26% of stage-specific ieGenes were mediated by at least two contexts (Fig. 6B), suggesting an interplay between different biological contexts.
Interpreting genetic regulation behind complex traits
Figure S19 illustrates the distribution and statistical summary (mean, standard deviation) of the six traits, providing a clearer understanding of the variation present in each. Based on a genome-wide association study (GWAS) of these traits in the LDS population, we identified a total of 100 loci with suggestive significance across the genome (P-value < 1e–5; Fig. S20A). Supplementary Data 6 provides more detailed information for each GWAS locus. Molecular QTL explained a higher proportion of phenotypic heritability compared to a MAF-matched random variant set, with eQTL explaining the highest proportion (Fig. S20B). These results highlight the strong explanatory power of molecular QTL in complex traits.
Colocalization analysis revealed that 53% (53 out of 100) of GWAS loci colocalized (PPH.4 > 0.8) with at least one molecular QTL (Fig. 7A, Supplementary Data 7), with eQTL and exQTL explaining the largest proportion of GWAS loci, followed by sQTL and enQTL. Compared to eQTL from matching tissues in the chicken GTEx, this study explained a greater number of GWAS loci (40 versus 17), emphasizing the importance of using populations with the same genetic background for colocalization (Fig. S20C). We also identified several GWAS loci that colocalized with specific molQTL. For instance, PIK3R1 influences EN210-400 through alternative splicing rather than gene expression in ovary (Fig. 7B) and this sQTL also identified as a significant stage interaction sQTL (P-value = 1.11e–07, Fig. S20D). Notably, 15% (6 out of 40) of the GWAS loci explained by all eQTL can only be colocalized with stage-specific or stage-interaction eQTL (Fig. 7A), highlighting the importance of the stage context for revealing regulatory effects and elucidating complex traits. For example, in the ovary, the eQTL of AMH colocalized with the GWAS loci for EN210, specifically in the Pre-laying stage (Fig. 7C). This gene has also been reported to be associated with follicular development for females31.
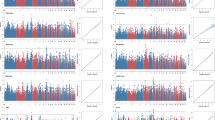
A UpSetR plot depicting the number of GWAS loci colocalized with four classes of molecular QTL (eQTL, exQTL, enQTL, and sQTL). The inset UpSetR plot shows, within the set of all eQTL–GWAS colocalizations, the breakdown across three eQTL categories: stage-interaction eQTL (stage-ieQTL), stage-specific eQTL (stage_specific), and other eQTL. B Interpretation of GWAS loci for egg number from ages 210 to 400 days, where the GWAS loci colocalize with the sQTL of PIK3R1 but not with the corresponding eQTL. The purple star indicates the SNP with the highest colocalization posterior probability (PPH.4), rs16762093. The −log10(P-values) for the GWAS and molQTL associations are shown in the plot, reflecting the statistical significance of each variant. The color of each point represents the linkage disequilibrium (r²) value between surrounding SNPs and rs16762093. C An example of stage-specific colocalization between the GWAS locus on chromosome 28 for egg number at 210 days and the eQTL of AMH in the ovary. The AMH eQTL exhibits stage specificity, being detected only in the pre-laying stage. The purple star indicates the SNP with the highest colocalization posterior probability (PPH.4), rs314098017. The color of each point represents the linkage disequilibrium (r²) value between surrounding SNPs and rs314098017. The −log10(P-values) for the GWAS and molQTL associations are shown in the plot, reflecting the statistical significance of each variant. The bottom panel shows the effect sizes of rs314098017 on AMH expression across the three laying stages, with bars representing the standard errors of the effect estimates. Sample sizes are as follows: GWAS (n = 12,952), pre-laying stage (n = 116), peak-laying stage (n = 110), and late-laying stage (n = 115).
We also estimated the genetic correlations among the six complex traits (Fig. 8A). The results revealed a strong positive genetic correlation between early egg production (EN210) and body weight, as well as a negative correlation with AFE. As a result of the genetic correlation, some molecular QTL can simultaneously explain multiple complex traits. For example, the eQTL of IGF2BP1, a key gene widely associated with growth traits in animals32,33,34,35, colocalizes with GWAS signals for BW49, AFE, and EN210, but not with EN300, EN400, or EN210–400 (Figs. 8B, S21). It is speculated that IGF2BP1 influences early development, thereby affecting early body weight, age at first egg, and early egg production. At the same time, we found that the colocalization exhibited tissue specificity, and no eQTL associated with IGF2BP1 was detected in the ovary (Fig. 8B).
A Genetic correlations among six complex traits, with SNP-based heritability (estimated using GCTA-REML) shown in parentheses. Red indicates positive genetic correlations, whereas blue indicates negative genetic correlations. B Colocalization of multiple complex traits with the IGF2BP1 eQTL on chromosome 27, showing tissue specificity of the eQTL (not detected in the ovary). The purple star indicates the SNP with the highest colocalization posterior probability (PPH.4), rs15242731. The −log10(P-values) for the GWAS and molQTL associations are shown in the plot, reflecting the statistical significance of each variant. The color of each point represents the linkage disequilibrium (r²) value between surrounding SNPs and rs15242731. C LDSC enrichment analysis of five complex human traits within genes colocalized with GWAS loci for age at first egg in chickens. Summary statistics for the five human traits were obtained from the GWAS Catalog (https://www.ebi.ac.uk/gwas/). The y-axis represents enrichment, determined by calculating the proportion of trait heritability attributed to SNPs within the specific annotation relative to the total SNPs in that annotation. Bars indicate the standard errors of the enrichment estimates. D Eleven out of twenty-six genes associated with age at first egg (AFE) in chickens show genome-wide significance for age at menarche in humans. Gene-based P-values were obtained from the GWASATLAS (https://atlas.ctglab.nl/). E Enrichment fold change between chicken reproduction-related genes (co-localized with loci for age at first egg and egg number) and female fertility QTL in pigs and cattle. QTL for reproduction-related traits in pigs and cattle were obtained from the Animal QTL Database (release 55). The enrichment fold change represents the ratio of the observed overlap between chicken reproduction-related genes and pig or cattle fertility QTL genes to the overlap expected by chance. Illustrative images of pigs and cattle were created in BioRender. Wang, Y. (2025) https://BioRender.com/pnkedie.
Common genes regulate reproduction traits in chicken and mammals
We investigated the overlap of genes affecting reproduction traits between chicken and humans based on the 1-to-1 orthologous genes. For the 36 genes colocalized with AFE, 26 are orthologous to human genes. We used LDSC36 to calculate heritability enrichment of genes colocalized with AFE in chickens on human complex traits. Results showed that AFE-related genes were more enriched for age at menarche and age at natural menopause compared to other complex traits, such as BMI, hip circumference, and height (Fig. 8C). For more details, we examined reported human GWAS results and found that 11 out of 26 homologous genes have been associated with age at menarche in humans (Fig. 8D). Additionally, we observed a significant enrichment of chicken reproduction-related candidate genes within the female fertility-associated QTL regions in both pigs and cattle. For instance, the enrichment fold change for the pig trait “Number of litters” was 62.93 (p = 0.0015) and for “Age at puberty” in cattle was 5.78 (p = 7.47e-5) (Fig. 8E). These results suggest a potential conservation of genetic regulation of fertility between chickens and mammals.
Discussion
Investigating the genetic regulation of molecular phenotypes in specific biological contexts and their impact on complex traits is critically important; however, well-designed population-based GTEx studies remain scarce. In this study, we established the laying-stage ChickenGTEx resource to elucidate stage-specific regulation during the egg-laying stages in chickens and its impact on complex traits. Compared to the recently released pilot phase ChickenGTEx20, which was mainly based on publicly available datasets, we detected more molecular QTL in HPG axis. For instance, we detected 10,186 eGenes in the ovary, whereas the ChickenGTEx dataset reported only 821, demonstrating that our study provides a substantial complement to the ChickenGTEx project. More importantly, we investigated stage-specific regulation across three egg-laying stages and its interactions with other biological contexts. By integrating molQTL with GWAS of complex traits, we elucidated the contribution of stage-specific regulation to complex traits and identified 124 candidate genes along with their corresponding tissues, molecular phenotypes, and the specific egg-laying stages in which they are functionally active. Overall, this study offers valuable resources and insights for advancing the understanding of reproductive trait regulation in chickens and other vertebrates.
More specifically, we observed that eQTL with larger effect sizes tend to have lower allele frequencies and lower connectivity (kME) within co-expression modules, suggesting that regulatory variants with large effects may be subject to stronger purifying (negative) selection. This reflects evolutionary constraints that help maintain the stability of core regulatory networks, a pattern consistent with observations from previous studies18,37. Further fine-mapping analysis identified numerous candidate causal mutations, laying the foundation for subsequent functional validation of molQTL. Both shared and tissue-specific molQTL were identified across tissues. Although we found that the likelihood of detecting an eQTL was independent of the expression level of the gene (Fig. S6B), similar conclusions have been reported in other studies17,20. We observed a significant overlap between tissue-specific eGenes and genes that are differentially expressed across tissues (Figs. 4C, S12C). This suggests that some tissue-specific gene expression may be caused by tissue-specific regulatory variants. However, there are still 2240 genes that, despite exhibiting tissue-specific regulatory effects, do not show tissue-specific expression. Additionally, 2646 genes that are differentially expressed across tissues do not detect tissue-specific eQTL (Fig. S12C). This suggests that other potential mechanisms, such as post-transcriptional regulation or environmental influences, may also contribute to these observations. To further explore cross-tissue regulatory effects, we introduced an additional molecular phenotype—the expression ratio between tissues—and found that 35.05% of erQTL are influenced by genetic factors distinct from those regulating the corresponding eQTL, providing insights into the genetic regulation of gene expression across tissues. Focusing on eQTL, we identified a number of stage-specific or -interaction eQTL. Notably, these QTL exhibited high tissue specificity, suggesting that during organismal development, tissue-specific functions and cellular microenvironments may promote the emergence of tissue- and stage-specific regulatory effects to support functional adaptation. However, these hypotheses remain speculative and require further experimental validation. Furthermore, we found that 60.02% of stage-interaction eQTL are mediated by cell proportions, transcription factor expression, or co-expression modules. Among these, transcription factors account for the largest proportion at 52.78%, consistent with previous studies that transcription factors as key regulatory factors during developmental stages38. Other factors, such as hormonal fluctuations, epigenetic modifications, micro environmental changes, and additional signaling pathways, likely also contribute to context-specific regulatory effects but were not captured in our current dataset, underscoring the need for additional data to fully elucidate these mechanisms. As laying stage and chronological age are biologically intertwined and cannot be fully separated, we acknowledge that some of our findings might reflect age-dependent regulatory effects rather than mechanisms specific to egg laying stages. Further investigation with samples from additional developmental stages and finer time points will be necessary to disentangle these effects and validate our observations.
By conducting GWAS on six complex traits in a large chicken population, we identified 100 GWAS loci, 80 of which are associated with chicken reproductive traits. Furthermore, through colocalization with molQTL, we explained 53 of the 100 loci and identified 124 candidate causal genes Similar to previous GTEx studies17, we found that some GWAS loci (15 of 53) could only be colocalized with specific molecular QTL, For example, we found that PIK3R1 influences EN210-400 through a splicing QTL in the ovary. Notably, PIK3R1 is a member of the PI3K/AKT pathway and serves as a key regulator of cellular processes such as proliferation and apoptosis39, highlighting the importance of investigating more detailed molecular phenotypes. We clarified the distinct roles of stage-specific and stage-interaction QTL in interpreting complex trait GWAS loci. For example, the stage-specific eQTL of AMH in the ovary regulates AFE and EN210 during the Pre-laying stage. This is consistent with studies in humans, where AMH levels steadily increase, peaking and plateauing around age 25. Subsequently, serum AMH levels begin to decline until menopause, when AMH production ceases40, emphasizing the importance of investigating stage-specific regulatory effects. Finally, our analysis revealed the potential conservation of regulatory genes associated with reproductive traits in chickens across mammals, including humans, pigs, and cattle, highlighting the value of using chickens as a model to study complex traits in other species.
Although our study provides valuable insights into the mechanisms of regulatory mutations across tissues and laying stages, several limitations and challenges remain. First, the GWAS of complex traits was conducted in a single commercial chicken population. Although the sample size was large (n = 12,952), the strong relatedness among individuals resulted in a small effective population size, which limited the power of GWAS. This issue has also been observed in dairy cattle41,42. In the future, it would be great to consider multi-breed GWAS analysis in chicken or even other farm animals. Second, further investigation into the patterns and differences in regulatory variations across a broader array of tissues requires a more extensive range of tissue types and developmental stages. The limited sample sizes for individual tissues and stages also restrict the exploration of trans-regulation. In addition, the number of tissues being analyzed remains limited. While our study focused on key components of the HPG axis, egg production is also influenced by other physiological systems, such as skeletal and metabolic tissues. Expanding tissue types in future studies will be crucial for achieving a more comprehensive understanding of the regulatory landscape underlying egg production traits. Finally, this study employed a deconvolution-based approach to examine the interaction between eQTL and cell types. However, a more comprehensive investigation of cell-type-specific regulatory mechanisms necessitates further validation with single-cell data. Moreover, the molecular phenotypes analyzed in this study were exclusively derived from transcriptomic data. Future studies integrating multi-omics strategies to characterize more precise molecular phenotypes will be essential for uncovering the genetic regulatory mechanisms underlying complex traits.
Methods
Tissue samples collection and RNA extraction
All tissue samples were obtained from a pure line commercial White Plymouth rock population comprising tens of thousands of chickens. We selected 119, 119, and 120 hens at 20 weeks, 30 weeks, and 58 weeks of age, respectively, corresponding to the Pre-, Peak-, and Late-laying stages of egg production for the population. To ensure representative sampling and increase the genetic diversity, individuals were randomly selected while avoiding close genetic relationships (e.g., half-sibs or closer) based on pedigree records. For each sample, we collected three gonadal axis tissues: the hypothalamus, pituitary, and ovary, along with liver tissue. All tissue samples were immediately frozen in liquid nitrogen and stored at −80 °C.
A total RNA of tissues was extracted using TRIeasy™ LS Total RNA Extraction Reagent (Yeasen, Shanghai, China), Qsep100 Bio-fragment analyzer (Bioptic, Jiangsu, China) was employed to detect the RNA quality, and RQN (RNA Quality Number) value > 7 was considered available. To remove rRNA (ribosome RNA), we adopted a previous study43 and designed 145 probes (Tsingke, Beijing, China) complemented to 5 s, 5.8 s, 12 s, 16 s, 18 s, and 28 s rRNA, these primers were mixed, and each primer was 1 mM in pool. The removal procedures of rRNA consisted of probe hybridization, RNase H enzymic digestion, and DNase I enzymic digestion; these procedures were performed according to the manufacturer guidelines (12258ES08, Yeasen, Shanghai, China). Subsequently, the produced clean RNA was constructed RNA library by the Hieff NGS® Ultima Dual-mode RNA Library Prep Kit following the manufacturer’s instrument (12308ES08, Yeasen, Shanghai, China). Finally, the obtained library employed a Qsep100 Bio-fragment analyzer to detect the library quality, and paired-end sequencing (100 bp) was performed using the DNBSEQ-T7 platform.
Whole genome sequence data analysis
DNA was extracted from the blood of all 358 hens using the DNeasy Blood & Tissue Kit (Qiagen 69506). The extracted DNA was assessed with a NanoDrop spectrophotometer and verified on a 1% agarose gel. All samples were then quantified using a Qubit 2.0 Fluorometer and subsequently diluted to a concentration of 40 ng/mL in 96-well plates. Subsequently, DNA sequencing libraries were constructed using the IGT® Enzyme Plus Library Prep Kit V3 (C11112, iGeneTech, Beijing, China) following standard protocols and sequenced on the DNBSEQ-T20×2RS platform (paired-end 150 bp), with an average sequencing depth of 26.43× (Supplementary Data 2). The raw data generated from sequencing were first subjected to quality control and adapter trimming using fastp44 (version 0.23.4). The processed reads were then aligned to the GRCg7b reference genome using bwa45 mem (version 0.7.17). The resulting BAM files were sorted with samtools46 (version 1.3.1), and duplicate reads were marked using Picard tools (https://broadinstitute.github.io/picard/). Base quality score recalibration (BQSR) was performed using GATK47 4.0.1.2, with known sites obtained from the chicken dbSNP48 (version 106). The processed BAM files were used to generate gVCF files through the GTX49 program, which integrates GATK Best Practices with FPGA-based hardware acceleration. Subsequently, joint SNP calling for all samples was performed using the GTX joint function, resulting in a total of 17,613,842 initial variants. A hard filtering method was applied using parameters “QD < 2.0||MQ < 40||FS > 60.0||SOR > 3.0||MQRankSum < −12.5 || ReadPosRankSum < −8.0” and remove indels, leaving 9,993,387 variant sites for further analysis. Afterward, for molQTL mapping, we removed SNPs with a minor allele frequency (MAF) < 0.05 within each tissue-stage group separately, ensuring that variants with sufficient allele frequency in each specific group were included in the analysis.
Corrections of sample labeling errors for RNA-seq and WGS
To eliminate the potential impact of labeling errors that may have occurred during sample collection or experimentation on subsequent analyses, we employed two strategies to detect and correct labeling errors in RNA-seq and WGS samples: 1) Checking the consistency of genotypes from RNA-seq data across different tissues from the same individual; 2) Comparing the consistency between RNA-seq genotypes and WGS-derived genotypes from the same individual.
First, we used GLIMPSE50 (version 1.1.1) for genotype imputation on the RNA-seq data, using the commercial breed panel (CBP) from the GCRP51,52 as the reference. Post-imputation, only variants with an INFO score greater than 0.4 were retained. We then combined these imputed genotypes with the clean variant set obtained from the WGS data and calculated the PI_HAT (the overall IBD value between two individuals) under both conditions (1 and 2) using Plink53 1.9 with the --genome parameter. Pairs of samples with identical labels but “PI_HAT < 0.8” or with different labels but “PI_HAT ≥ 0.8” were defined as mislabeled sample pairs. Subsequently, an iterative matching process was performed to correct the sample labels using the WGS data labels as the reference. A total of 37 RNA-seq samples that could not be corrected were identified and excluded. The final number of samples remaining in each group is shown in Fig. 1A. All label correction processes were performed using chromosome 6.
Low-depth sequence data analysis and genotype imputation
We collected blood samples from a total of 13,329 hens, each of which had at least one phenotypic record, from the same population as the RNA-seq samples. DNA was extracted from all blood samples using the same method as described in the previous step. Subsequently, genomic libraries were constructed using a Tn5 transposase-based method, following the detailed procedures outlined in54. All samples were sequenced using the MGI-seq 2000 platform with a targeted sequencing depth of 0.5 × (192 samples per sequencing lane). The raw data underwent quality control and trimmed using fastp44 (version 0.23.4) to remove low-quality reads and adapter sequences. Samples with sequencing depths lower than 0.1 × (calculated as clean sequence base number/reference genome base number) were excluded, leaving a set of 13,186 samples. The distribution of sequencing depths is shown in Fig. S1A.
All samples that passed quality control were aligned to the GRCg7b reference genome using the bwa45 (version 0.7.17) mem algorithm. Genotype imputation was then performed using GLIMPSE50, with a reference panel comprising the 358 high-depth GATK clean variant set obtained previously (phased by Beagle55 5.4). The imputed dataset retained only variants with an INFO score greater than 0.4 and a minor allele frequency (MAF) greater than 0.01, resulting in a final set of 8,742,439 variants for subsequent analysis. We corrected potential sample mislabeling using pedigree records by computing the Mendelian error rate between parent-offspring pairs in the pedigree. Samples with labeling errors were removed based on the distribution of Mendelian error rate (Fig. S1C), leaving a total of 12,952 samples for further analysis.
RNA-Seq data analyses and definition of molecular phenotype
For all raw RNA-seq data, we performed quality control and adapter trimming using fastp (version 0.23.4). The clean reads were then aligned to the GRCg7b reference genome using STAR56 (version 2.7.3a) with the following parameters “--chimSegmentMin 10, --twopass1readsN −1, --outFilterMismatchNoverLmax 0.03, --alignIntronMin 20, --alignSJoverhangMin 8 and --sjdbOverhang 99”. High-quality samples were defined as those with a unique mapping rate of > 60% and a clean reads count of > 8,000,000. The results indicated that all samples passed the quality control (Supplementary Data 1). We defined four types of molecular phenotypes: gene expression, exon expression, enhancer expression, and alternative splicing. We obtained annotation information for 29,521 genes and 315,300 exons (only genes or exons located on chromosomes were considered) from the GRCg7b genome annotated in Ensembl v110. For enhancers, we extracted chromatin state annotation data for 23 tissues from the chicken FAANG57 project. Regions annotated as active enhancers (E6) were merged across all tissues, excluding regions overlapping with gene regions and those longer than 2 kb. We then used LiftOver to convert the genomic coordinates from GRCg6a to GRCg7b, resulting in a final set of 22,702 enhancer annotations. We used featureCounts58 from subread (version 2.0.5) to obtain the raw count data for each gene, exon, and enhancer, and subsequently removed batch effects caused by experimental and sample collection variations using ComBat59 from sva60 R package (version 3.50.0). The corrected counts were converted into normalized expression levels (i.e., Transcripts Per Million, TPM). In each group (tissue × stage or merged stage), we retained only genes, exons, and enhancers with a TPM (Transcripts Per Million) greater than 0.1 in at least 20% of the samples for next analysis.
We utilized the LeafCutter61 to identify and quantify alternative splicing events. Starting with the BAM files produced by STAR56 alignment, we generated junction files for each sample using the “bam2junc.sh” script. Subsequently, intron clusters were defined across samples with the leafcutter_cluster.py script, employing the parameters “-m 50 and -l 500000”. We then mapped these intron clusters to their corresponding genes by applying the “map_clusters_to_genes.R” script (available at https://github.com/broadinstitute/gtex-pipeline).
To refine our dataset, introns were filtered out if they did not meet specific criteria: less than 50% of samples contained detectable reads, or the read count was lower than max(10, 0.1n), where n is the number of samples. Additionally, we excluded introns with minimal variability, defined as ∑i(|zi| < 0.25) ≥ n−3 and ∑i(|zi| > 6) ≤ 3, where zi represents the z-score of the i-th cluster’s read fraction across samples. The number of valid molecular phenotypes detected in each group is provided in Supplementary Data 3. We performed the tree clustering of all the RNA-Seq samples using the ggtree62 package and conducted PCA clustering with the prcomp function in R.
Differential gene expression analysis across tissues and egg laying-stages
We conducted a differential gene expression analysis across tissues and within different egg-laying stages using the Wilcoxon rank-sum test method63. Firstly, we normalized the count (after batch effects corrected) to TMM (Trimmed Mean of M-values) by edgeR64 package and further converted to CPM (Counts Per Million). For the calculation of P-values, each gene’s CPM values were input into the wilcox.test function. Multiple testing correction of P-values was performed using the Benjamini & Hochberg method65. Genes with a log2 fold change (log2FC) > 2 and a false discovery rate (FDR) < 0.05 were defined as differentially expressed genes (DEGs) between tissues. Similarly, within the same tissue, genes with FC > 2 and FDR < 0.05 were considered DEGs between laying stages.
Time-course clustering
We utilized the Mfuzz66 package (v.2.32.0) to examine changes in gene expression across three egg-laying stages within each tissue. The genes were clustered based on their expression changes using a c-means clustering method. The number of clusters for each tissue was determined by the elbow method applied to the principal component analysis (PCA). Subsequently, all genes were grouped into six distinct clusters based on their expression patterns (Fig. 2B). To detect differences in gene expression patterns across stages between tissues, we calculated the odds ratios of overlapping genes between different tissues for each specific expression pattern cluster. For example, genes with high expression specifically at the Peak stage across the three egg-laying stages were defined as cluster A. The odds ratio (OR) of overlapping genes in cluster A between tissue 1 and tissue 2 was calculated as “OR = a*d/b*c”, where a is the number of overlapping genes clustered in cluster A for both tissues, b is the number of genes not clustered in cluster A in tissue 2, c is the number of genes not clustered in cluster A in tissue 1, and d is the number of genes not appearing in cluster A in either tissue 1 or tissue 2.
Construct co-expression networks
We performed robust Weighted Gene Correlation Network Analysis (WGCNA) on genes within each tissue to construct co-expression networks using the WGCNA24 R package (version 1.69). Network dendrograms for each TOM were generated using average linkage hierarchical clustering of the dissimilarity TOM (1 – TOM) and modules were subsequently constructed using the cutreeDynamic function. Highly correlated modules were combined with the mergeCloseModules function using a merge cut height of 0.25.
Estimating cis-heritability
For gene, exon, and enhancer expression analysis, we used the edgeR64 package to convert the count values with batch effects corrected into TMM (Trimmed Mean of M-values). The resulting TMM matrix was then subjected to an inverse normal transformation for subsequent molecular QTL (molQTL) mapping analysis. For alternative splicing events, the filtered data were normalized across samples using the “prepare_phenotype_table.py” script. The final normalized PSI (Percent Spliced In) values were stored in a BED format file, which was subsequently used for sQTL (splicing QTL) mapping.
We used LDAK67 version 5.2 to fit the following mixed linear model Eq. (1) to estimate the cis-heritability of molecular phenotypes:
where y is a vector containing the molecular phenotypes normalized across samples, and β is a vector corresponding coefficients of quantitative covariates X, including 10 phenotype principal components (PCs), 5 genotype PCs and stages (for merged group). The term g1 represents the genetic values of SNPs in the cis-region (defined as within ±1 Mb of the transcription start sites (TSS) of a gene or enhancer), with g1 ~ N (0, \({{{\bf{G}}}}_{{{\bf{1}}}}{{{\rm{\sigma }}}}_{{{\rm{g}}}1}^{2}\)). The term g2 represents the genetic values of SNPs outside the cis region, with g2 ~ N (0, \({{{\bf{G}}}}_{{{\bf{1}}}}{{{\rm{\sigma }}}}_{{{\rm{g}}}2}^{2}\)). The term ϵ represents the residuals, with ϵ ~ N(0, \({{{\rm{I}}}{{\rm{\sigma }}}}_{{{\rm{e}}}}^{2}\)). G1 and G2 are the genomic relationship matrices (GRM) constructed from SNPs in the cis and non-cis regions, respectively, and I is the identity matrix. The parameters \({{{\rm{\sigma }}}}_{{{\rm{g}}}1}^{2}\), \({{{\rm{\sigma }}}}_{{{\rm{g}}}2}^{2}\) and \({{{\rm{\sigma }}}}_{{{\rm{e}}}}^{2}\) represent the variances explained by SNPs in the cis region, SNPs in the non-cis region, and random residuals, respectively. Cis-heritability is defined as \({{{\rm{\sigma }}}}_{{{\rm{g}}}1}^{2}/({{{\rm{\sigma }}}}_{{{\rm{g}}}1}^{2}+{{{\rm{\sigma }}}}_{{{\rm{g}}}2}^{2}+{{{\rm{\sigma }}}}_{{{\rm{e}}}}^{2})\).
molQTL mapping
We utilized the GPU-accelerated software tensorQTL26 (version 1.0.4) to perform cis-QTL mapping on standardized molecular phenotypes across all tissues and laying stage (i.e., SNPs located within 1 Mb upstream and downstream of the gene’s TSS). Similar to the approach used for heritability estimation, the first 10 phenotype principal components (PCs), the first 5 genotype PCs and stages (for merged group) were included as covariates. The parameter “--model cis_nominal” was employed to calculate all nominal associations for each variant-molecular phenotype pair. Subsequently, we used the parameter “--model cis” to conduct permutations for computing empirical P-values for each molecular phenotype. The molecular quantitative trait loci (molQTL) are genetic loci that are significantly associated with the variation in molecular phenotypes across individuals. More specifically, molQTL linked to gene expression, exon expression, enhancer expression, and alternative splicing are further referred to as expression QTL (eQTL), exon QTL (exQTL), enhancer QTL (enQTL), and splicing QTL (sQTL), respectively. The genes with at least one molQTL are referred to as molGenes. For example, a gene harboring eQTL or sQTL is referred to as an eGene or sGene, respectively. Additionally, we utilized the “--mode cis_independent” module to determine the number of independent molQTL for each ePhenotype. To estimate confidence intervals for each identified QTL, we employed a linkage disequilibrium (LD)-based approach13. Specifically, for each lead SNP, we defined the confidence interval as the genomic region spanning from the furthest upstream to the furthest downstream SNP that is in high LD (r² ≥ 0.8) with the lead SNP, based on pairwise LD calculated from the corresponding population genotype data.
Fine-mapping of molQTL
To fine-map the molQTL identified in each of four tissues, we employed the Sum of Single Effects Regression (SuSiE) method, a Bayesian variable selection framework that assumes multiple causal variants may be present within a given locus while accounting for linkage disequilibrium (LD) among variants27. SuSiE models the genetic effect as the sum of multiple “single-effect” components, where each component represents a regression on a single causal variant, thereby enabling the joint inference of multiple causal signals without requiring prior specification of the number of causal variants. Specifically, SuSiE iteratively fits a series of sparse regression models using a Bayesian framework, estimating the posterior inclusion probabilities (PIP) for each variant, which reflects the probability of the variant being causal, given the data and the LD structure. Based on these PIPs, SuSiE constructs credible sets (CS)—minimal sets of variants that together contain the causal variant with high probability.
The summary statistics for the cis-region of each ePhenotype were used as input, while the corresponding genotype matrix and linkage disequilibrium (LD) matrix were generated using PLINK53 1.9. Variants with posterior inclusion probabilities (PIPs) summing up to 90% were identified as credible sets (CS).
Functional enrichment of molQTL
We annotated all variants used in the study for sequence ontology and regulatory elements using SnpEff68 and the 15 chromatin states annotated by the chicken FAANG57 project. Subsequently, we calculated the enrichment odds ratio (OR) for each set of molQTL across different annotation categories using the following formula Eq. (2):
Here, a represents the number of variants that are both molecular QTL and overlap with the annotation category; b represents the number of variants that fall within the annotation category but are not molecular QTL; c represents the number of variants that are molecular QTL but do not fall within the annotation category; and d represents the number of variants that are neither molecular QTL nor within the annotation category.
Shared and specific molQTL across tissues and laying stages
We carried out a meta-analysis of molQTL across various tissues and egg-laying stages using MashR69 (version v0.2.79). For this analysis, we focused on the z-scores from tensorQTL26 (derived from the ratio of slope to standard error) for the leading cis-molQTL. The mash function was utilized to estimate the effect sizes (posterior means) and their corresponding significance (local false sign rates, LFSR). A molQTL was considered active in a specific tissue or egg-laying stage if its LFSR was below 0.05. To evaluate the genetic similarity between tissues concerning gene expression regulation, we computed the Spearman correlation coefficients of effect size estimates for cis-molQTL between tissue or stage pairs, concentrating on SNPs with an LFSR below 0.05 in at least one group. We defined eGene as tissue- or stage-specific if it exhibited an LFSR < 0.05 in only one tissue or stage.
Gene expression ratios between tissues and erQTL mapping
To further determine whether the cross-talk of gene expression between tissues is genetically regulated, we defined gene expression ratios between tissues (6 tissue pairs for 4 tissues). Firstly, counts (after correcting batch effects) were normalized to TPM (see above). For gene(a) in the tissue1-tissue2 pair, the gene expression ratio is defined as (TPMa1 + 0.01)/(TPMa2 + 0.01), where a small constant (0.01) is added to prevent division by zero and to stabilize the ratio for genes with low or zero expression. Only genes expressed in both tissues (with TPM > 0.1 in at least 20% of samples) and samples overlapping between tissues were included, and an inverse normal transformation was performed before QTL mapping. Subsequently, we identified expression ratio erQTL using the same method applied to other molecular phenotypes (see above).
Colocalization between erQTL and eQTL
To demonstrate whether eQTL and erQTL shared genetic regulatory mechanisms, we performed colocalization analysis between erQTL and the corresponding gene’s eQTL within individual tissues using the coloc70 R package version 5.2.3. We used PPH.4, defined as the posterior probability for association with both the molecular phenotype and shared signals, to assess the likelihood of colocalization between QTL.
Single-cell data analysis and deconvolution
We obtained single-cell RNAseq data from external populations for three tissues: the hypothalamus, liver, and ovary. The original single-cell sequencing data were processed using the DNBelab C Series scRNA analysis software71 (MGI). Reads were aligned to the GRCg7b reference genome to generate a digital gene expression matrix by STAR56. The quality control parameters, including gene counts per cell, UMI counts per cell, and the percentage of mitochondrial genes, were specified. Genes expressed in fewer than three cells and cells with fewer than 200 detected genes were removed. Additionally, cells with more than 25% mitochondrial gene expression were filtered out. The data from each sample were normalized using the default options in the “NormalizaData” function. Highly variable genes were then identified using the “FindVariableFeatures” function, selecting them based on their average expression and variability. For each sample, “DoubleFinder” was used with default parameters to eliminate potential doublets. The cell cluster was identified using the “FindClusters” function from Seurat72 v5.1.0, applying a standard integration process and setting a threshold of P-value < 0.01 for statistical significance. To annotate cell populations, we utilized the expression patterns of differentially expressed genes in conjunction with known cell markers from the literature. Genes with a |log2FC| > 1 and an adjusted P-value < 0.05 were identified as marker genes. Subsequently, the CIBERSORT73 tool was used for cellular deconvolution analysis for each bulk RNA-seq sample. In tissues, cell types with a mean value < 1% or more than 80% of samples having a value of 0 were removed.
Laying stage interaction molQTL
For molecular phenotypes with at least one significant QTL, we used tensorQTL26 to fit the following model to identify laying stage interaction QTL Eq. (3):
where y is a vector containing the molecular phenotypes normalized across samples, and β is a vector corresponding coefficients of quantitative covariates X, including 10 phenotype principal components (PCs) and 5 genotype PCs. g is the vector of genotype effect of SNPs in the cis-region, i is the stage term, g*i is the interaction term between genotype and laying stage, and ϵ represents the residuals. The P-values for g*i were first corrected using the Bonferroni method based on the number of independent variants tested for each molecular phenotype. Subsequently, the adjusted P-value of the most significant variant in each molecular phenotype was adjusted using the Benjamini-Hochberg (BH) correction. Genes with eQTL that exhibited a significant interaction with developmental stage (FDR < 0.2) were referred to as stage-interaction genes (ieGenes), and the corresponding eQTL were considered as stage-interaction eQTL (ieQTL).
Mediation analysis
In order to more precisely evaluate whether the genotype-by-stage (G × Stage) interaction effect on gene expression is mediated by other biological contexts, we conducted a formal mediation analysis using the mediation29 package (version 4.5.0) and CIT package28 (version 2.3.2) in R. We considered three classes of potential mediators: cell type proportions, transcription factor expression levels (annotation from AnimalTFDB474), and module eigengenes derived from weighted gene co-expression network analysis (WGCNA). For the mediation method, for each stage-interaction eGene (stage-ieGene), we tested all three types of mediators by fitting two models:
Mediator model Eq. (4):
Outcome model Eq. (5):
Where Y represents gene expression, G × M denotes the interaction effect between genotype and the mediator, β1 represents the stage effect, β2 is the genotype effect, β3 represents the interaction effect between genotype and stage, β4 is the mediator effect, βc denotes the covariate effect, and ϵ is residual. We assessed whether the observed genotype-by-stage interaction effects on gene expression could be explained by potential mediators by estimating the Total Effect (TE), Average Causal Mediation Effect (ACME), and Average Direct Effect (ADE). TE refers to the overall effect of the genotype-by-stage interaction on gene expression, without accounting for the mediator. ADE represents the portion of this effect that is not transmitted through the mediator. ACME quantifies the portion of the total effect that is transmitted through the mediator (i.e., the indirect path from genotype-by-stage to the mediator, then to gene expression). Statistical significance of ACME, ADE, TE, and Proportion Mediated was assessed at a 95% confidence level, with confidence intervals computed using a nonparametric bootstrap procedure with 999 Monte Carlo draws. A mediation effect was considered significant when the ACME P-value < 0.01.
For the Causal Inference Test (CIT), we adopted the formal hypothesis testing framework implemented in the R package cit to evaluate whether the regulatory effects of stage-interacting eQTL (ieQTL) on gene expression could be statistically mediated by three classes of biological variables: cell type proportions, transcription factor expression, and co-expression modules. For each triplet (L, G, T), where L represents the interaction term between stage and genotype (stage × G), G is the mediator, and T is the gene expression trait, the CIT framework tests four necessary conditions: (i) L is associated with T, (ii) L is associated with G conditional on T, (iii) G is associated with T conditional on L, and (iv) L is independent of T conditional on G. Each condition was assessed using general linear models, and the largest p-value among these four tests was used as an omnibus test statistic following the intersection-union test (IUT) principle. We applied this framework separately for all candidate mediator-stage-ieGene triplets. To identify significant mediation effects, we used a permutation-based association threshold of p < 0.01 for condition (iii) (i.e., association between T and G conditional on L).
GWAS of complex traits
We collected 12,952 hens with at least one phenotypic record from the same population as the RNAseq samples for genome-wide association studies (GWAS). The phenotypes analyzed included body weight at 49 days of age (BW49), age at first egg (AFE), egg number at 210 days of age (EN210), egg number at 300 days of age (EN300), egg number at 400 days of age (EN400), and egg number between 210 and 400 days (EN210-400). Genotypes were obtained using low-depth sequencing combined with genotype imputation (as described previously). We employed the GCTA75 software to fit mixed linear models for each complex trait, and candidate GWAS loci were identified using a significance threshold of P < 1e-5. GWAS loci were defined as chromosomal regions adjacent pairs of significant variants were less than 1 Mb from each other. We applied the same LD-based strategy as used in the molQTL analysis to define confidence intervals of each GWAS loci. We applied the same LD-based strategy used in the molQTL analysis to define the confidence intervals for each GWAS locus. Specifically, for each lead SNP, the confidence interval was defined as the genomic region spanning from the furthest upstream to the furthest downstream SNP in high LD (r² ≥ 0.8), based on pairwise LD calculated from the genotype data.
Integrate analysis of molQTL and GWAS
We performed integrative analysis of complex trait GWAS and molQTL using colocalization by the coloc70 R package to prioritize molecular features that share an underlying genetic cause with GWAS loci. For each molQTL, we first checked whether any overlap existed with a GWAS locus. For overlapping molQTL-GWAS loci pairs, we extracted the summary statistics and conducted colocalization analysis. The posterior probability of colocalization (PPH.4) was estimated using the coloc_abf function, and molecular features with PPH.4 > 0.8 were defined as colocalized.
Common genes regulate reproduction in chicken and mammals
To investigate whether genes affecting reproduction-related complex traits in chicken exhibit conservation across species, we firstly collected data on five human complex traits from the GWAS Catalog76 (https://www.ebi.ac.uk/gwas/): age at menarche, age at natural menopause, BMI, hip circumference, and standing height. Among these, age at menarche and age at natural menopause were considered analogous to the trait “age at first egg (AFE)” in chickens, while the remaining three traits were used as control traits. We mapped the 36 functional genes colocalized with AFE in chickens to their corresponding human orthologues using one-to-one orthologous gene mapping from the Ensembl database (v102), identifying 26 homologous genes shared between chickens and humans. Subsequently, we conducted linkage disequilibrium (LD) score regression36 analysis for these 26 homologous genes. Heritability enrichment was determined by calculating the proportion of trait heritability attributed to SNPs within the specific annotation relative to the total SNPs in that annotation. Additionally, we queried the GWASATLAS77 (https://atlas.ctglab.nl/) database to examine the association of these 26 genes with human reproduction-related traits.
For non-human mammals, we retrieved QTL annotations related to female fertility for two major agricultural species, pig and cattle, from the AnimalQTL database (release 55)12. The associated genes for the relevant traits (Fig. 8E) were identified using the GTF annotation file. Subsequently, we calculated the enrichment fold change by comparing the chicken fertility candidate genes (AFE or EN related) identified in this study (Supplementary Data 4) with the trait-associated genes in both pig and cattle.
Ethical statement
All animal procedures in this study were conducted in accordance with the guidelines of the Institutional Animal Care and Use Committee (IACUC) of China Agricultural University (permission number: SKLAB-2014-06-07).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw RNA-seq data generated in this study have been deposited in the NCBI SRA database under BioProject ID: PRJNA1188634. Additionally, the raw whole-genome sequencing data have been deposited in the SRA database under BioProject ID: PRJNA1198198. The full summary statistics of molQTL mapping results for all molecular phenotypes are publicly available at https://ngdc.cncb.ac.cn/chickengtex/static/download/molQTL.tar.gz. The fine-mapping results of molecular QTL are publicly available at https://ngdc.cncb.ac.cn/chickengtex/static/download/fine-mapped_molQTL.rar. The GWAS summary statistics for six complex traits are available on Figshare [https://doi.org/10.6084/m9.figshare.28333362.v1]. The raw genotype and phenotype data underlying the six complex traits are available under restricted access for commercial reasons. Access can be requested from the corresponding author, Xiaoxiang Hu (huxx@cau.edu.cn), and will be granted for research purposes. All requests will be reviewed and responded to within two weeks. Source data are provided with this manuscript. Source data are provided with this paper.
Code availability
The code required to reproduce the analyses is available on Zenodo (https://doi.org/10.5281/zenodo.14902956) and http://chicken.farmgtex.org78.
References
Khalid, Z. An updated review on chicken eggs: production, consumption, management aspects and nutritional benefits to human health. Food Nutr. Sci. 06, 1208–1220 (2015).
Wright, D. et al. The genetic architecture of domestication in the chicken: effects of pleiotropy and linkage. Mol. Ecol. 19, 5140–5156 (2010).
Garcia, P., Wang, Y., Viallet, J. & Macek Jilkova, Z. The chicken embryo model: a novel and relevant model for immune-based studies. Front. Immunol. 12, 791081 (2021).
Flores-Santin, J. & Burggren, W. W. Beyond the chicken: alternative avian models for developmental physiological research. Front. Physiol. 12, 712633 (2021).
Mubarak, M., AM, A.-E. G. & Bastawrous, A. Pathological changes of the reproductive tract of laying hens and their causative agents. Assiut Vet. Med. J. 40, 97–120 (1998).
Webster, A. Welfare implications of avian osteoporosis. Poult. Sci. 83, 184–192 (2004).
Wolford, J. & Polin, D. Lipid accumulation and hemorrhage in livers of laying chickens.: a study on Fatty Liver-Hemorrhagic Syndrome (FLHS). Poult. Sci. 51, 1707–1713 (1972).
Padmanabhan, V., Karsch, F. J. & Lee, J. S. Hypothalamic, pituitary and gonadal regulation of FSH. Reprod. Suppl. 59, 67–82 (2002).
Eppig, J. J., Wigglesworth, K. & Pendola, F. L. The mammalian oocyte orchestrates the rate of ovarian follicular development. Proc. Natl. Acad. Sci. USA 99, 2890–2894 (2002).
Ahmed, A. A., Ma, W., Ni, Y., Wang, S. & Zhao, R. Corticosterone in ovo modifies aggressive behaviors and reproductive performances through alterations of the hypothalamic-pituitary-gonadal axis in the chicken. Anim. Reprod. Sci. 146, 193–201 (2014).
Komatsu, K. & Masubuchi, S. Observation of the dynamics of follicular development in the ovary. Reprod. Med. Biol. 16, 21–27 (2017).
Hu, Z.-L., Park, C. A. & Reecy, J. M. Bringing the animal QTLdb and CorrDB into the future: meeting new challenges and providing updated services. Nucleic Acids Res. 50, D956–D961 (2021).
Zhu, X. et al. Mapping the regulatory genetic landscape of complex traits using a chicken advanced intercross line. Nat. Commun. 16, 5841 (2025).
Consortium, G. et al. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Consortium, G. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Liu, S. et al. A multi-tissue atlas of regulatory variants in cattle. Nat. Genet. 54, 1438–1447 (2022).
Teng, J. et al. A compendium of genetic regulatory effects across pig tissues. Nat. Genet. 56, 112–123 (2024).
Wen, C. et al. Cross-ancestry atlas of gene, isoform, and splicing regulation in the developing human brain. Science 384, eadh0829 (2024).
Kim-Hellmuth, S. Cell type-specific genetic regulation of gene expression across human tissues. Science 369, eaaz8528 (2020).
Guan, D. et al. Genetic regulation of gene expression across multiple tissues in chickens. Nat. Genet. 57, 1298–1308 (2025).
Coorens, T. H. H. et al. The human and non-human primate developmental GTEx projects. Nature 637, 557–564 (2025).
Fang, L. et al. The Farm Animal Genotype–Tissue Expression (FarmGTEx) project. Nat. Genet. 57, 786–796 (2025).
Chen, Y. et al. The Prognostic model established by the differential expression genes based on CD8+ T cells to evaluate the prognosis and the response to immunotherapy in osteosarcoma. Med. Inflamm. 2023, 6563609 (2023).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 9, 1–13 (2008).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Taylor-Weiner, A. et al. Scaling computational genomics to millions of individuals with GPUs. Genome Biol. 20, 228 (2019).
Wang, G., Sarkar, A., Carbonetto, P. & Stephens, M. A simple new approach to variable selection in regression, with application to genetic fine mapping. J. R. Stat. Soc. Ser. B Stat. Methodol. 82, 1273–1300 (2020).
Millstein, J., Chen, G. K. & Breton, C. V. cit: hypothesis testing software for mediation analysis in genomic applications. Bioinformatics 32, 2364–2365 (2016).
Tingley, D. et al. Mediation: R package for causal mediation analysis. J. Stat. Softw. 59, 1–38 (2014).
Liao, X., Wang, W., Yu, B. & Tan, S. Thrombospondin-2 acts as a bridge between tumor extracellular matrix and immune infiltration in pancreatic and stomach adenocarcinomas: an integrative pan-cancer analysis. Cancer Cell Int. 22, 213 (2022).
Weenen, C. et al. Anti-Mullerian hormone expression pattern in the human ovary: potential implications for initial and cyclic follicle recruitment. Mol. Hum. Reprod. 10, 77–83 (2004).
Sheng, Z. et al. Genetic dissection of growth traits in a Chinese indigenous× commercial broiler chicken cross. BMC Genomics 14, 1–12 (2013).
Ma, M. et al. Genome-wide association study for carcase traits in spent hens at 72 weeks old. Ital. J. Anim. Sci. 18, 261–266 (2019).
Hansen, T. V. et al. Dwarfism and impaired gut development in insulin-like growth factor II mRNA-binding protein 1-deficient mice. Mol. Cell. Biol. 24, 4448–4464 (2004).
Wang, K. et al. The chicken pan-genome reveals gene content variation and a promoter region deletion in IGF2BP1 affecting body size. Mol. Biol. Evol. 38, 5066–5081 (2021).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Zeng, J. et al. Signatures of negative selection in the genetic architecture of human complex traits. Nat. Genet. 50, 746–753 (2018).
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Vallejo-Díaz, J., Chagoyen, M., Olazabal-Morán, M., González-García, A. & Carrera, A. C. The opposing roles of PIK3R1/p85α and PIK3R2/p85β in cancer. Trends Cancer 5, 233–244 (2019).
Demirdjian, G. et al. Performance characteristics of the Access AMH assay for the quantitative determination of anti-Müllerian hormone (AMH) levels on the Access* family of automated immunoassay systems. Clin. Biochem. 49, 1267–1273 (2016).
Jiang, J. et al. Functional annotation and Bayesian fine-mapping reveal candidate genes for important agronomic traits in Holstein bulls. Commun. Biol. 2, 212 (2019).
Freebern, E. et al. GWAS and fine-mapping of livability and six disease traits in Holstein cattle. BMC Genomics 21, 41 (2020).
Phelps, W. A., Carlson, A. E. & Lee, M. T. Optimized design of antisense oligomers for targeted rRNA depletion. Nucleic Acids Res. 49, e5 (2021).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. Imeta 2, e107 (2023).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, https://doi.org/10.1093/gigascience/giab008 (2021).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Phan, L. et al. The evolution of dbSNP: 25 years of impact in genomic research. Nucleic Acids Res. 53, D925–D931 (2025).
Bu, L. et al. Improving read alignment through the generation of an alternative reference via an iterative strategy. Sci. Rep. 10, 18712 (2020).
Rubinacci, S., Ribeiro, D. M., Hofmeister, R. J. & Delaneau, O. Efficient phasing and imputation of low-coverage sequencing data using large reference panels. Nat. Genet. 53, 120–126 (2021).
Zhu, D. et al. GCRP: integrated global chicken reference panel from 11,951 chicken genomes. Genomics, Proteomics & Bioinformatics, https://doi.org/10.1093/gpbjnl/qzaf032 (2025).
Zhang, Z. et al. The efficient phasing and imputation pipeline of low-coverage whole genome sequencing data using a high-quality and publicly available reference panel in cattle. Anim. Res. One Health 1, 4–16 (2023).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet 81, 559–575 (2007).
Yang, R. et al. Accelerated deciphering of the genetic architecture of agricultural economic traits in pigs using a low-coverage whole-genome sequencing strategy. Gigascience 10, giab048 (2021).
Browning, B. L., Tian, X., Zhou, Y. & Browning, S. R. Fast two-stage phasing of large-scale sequence data. Am. J. Hum. Genet 108, 1880–1890 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Pan, Z. et al. An atlas of regulatory elements in chicken: a resource for chicken genetics and genomics. Sci. Adv. 9, eade1204 (2023).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general-purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Li, Y. I. et al. Annotation-free quantification of RNA splicing using LeafCutter. Nat. Genet. 50, 151–158 (2018).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T. Y. ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Li, Y., Ge, X., Peng, F., Li, W. & Li, J. J. Exaggerated false positives by popular differential expression methods when analyzing human population samples. Genome Biol. 23, 79 (2022).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc.: Ser. B 57, 289–300 (1995).
Kumar, L. & Futschik, M. E. Mfuzz: a software package for soft clustering of microarray data. Bioinformation 2, 5 (2007).
Berrandou, T. E., Balding, D. & Speed, D. LDAK-GBAT: fast and powerful gene-based association testing using summary statistics. Am. J. Hum. Genet 110, 23–29 (2023).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly. (Austin) 6, 80–92 (2012).
Urbut, S. M., Wang, G., Carbonetto, P. & Stephens, M. Flexible statistical methods for estimating and testing effects in genomic studies with multiple conditions. Nat. Genet. 51, 187–195 (2019).
Wallace, C. Eliciting priors and relaxing the single causal variant assumption in colocalisation analyses. PLoS Genet. 16, e1008720 (2020).
Han, L. et al. Cell transcriptomic atlas of the non-human primate Macaca fascicularis. Nature 604, 723–731 (2022).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Shen, W. K. et al. AnimalTFDB 4.0: a comprehensive animal transcription factor database updated with variation and expression annotations. Nucleic Acids Res. 51, D39–D45 (2023).
Yang, J. A., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Cerezo, M. et al. The NHGRI-EBI GWAS Catalog: standards for reusability, sustainability and diversity. Nucleic Acids Res. 53, D998–D1005 (2025).
Watanabe, K. et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 51, 1339–1348 (2019).
Guan, D. Chicken genotype-tissue expression (ChickenGTEx) project. Zenodo https://doi.org/10.5281/zenodo.14902956 (2025).
Acknowledgments
The analysis was performed on the high-performance computing platform of the State Key Laboratory of Agrobiotechnology and the National Research Facility for Phenotypic and Genotypic Analysis of Model Animals. We would like to thank Jiangli Ren, Yini Jia, Linlin Zhang, Ce Wang, Jiatong Yu, and Ran Song for their genomic sequencing and data analyses. We also thank Yang Xi, Zexi Cai, Goutam Sahana and Doug Speed for discussions. This study is funded by the National Key R&D Program of China (Grant No. 2021YFD1300100 to X.X.H.), Biological Breeding-National Science and Technology Major Project (Grant No. 2023ZD0406805 to X.X.H.), the National Natural Science Foundation of China (Grant No. 32272862to X.X.H.), Key-Area Research and Development Program of Guangdong Province (Grant No. 2022B0202100002 to X.X.H.) and the 2115 Talent Development Program of China Agricultural University.
Author information
Authors and Affiliations
Consortia
Contributions
Conceptualization, X.H., L.F., and C.F.; methodology, D.Z., L.F., K.S., and X.H.; formal analysis, D.Z., K.S., C.L., Y.Y., H.L., Z.B., and L.T.; investigation, D.G., Y.Z., Z.J., Z.X., and C.F.; funding acquisition, X.H., Y.W., and C.F.; supervision, L.F. and X.H.; writing – original draft, D.Z., L.F., and K.S.; writing– review & editing, D.Z., L.F., X.H., C.F., K.S., B.F., and Y.W. All authors read, edited and approved the final manuscript. The ChickenGTEx Consortium contributed to data generation and validation.
Corresponding authors
Ethics declarations
Competing interests
Ziqin Jiang and Zhenqiang Xu are current employees of Wen’s Nanfang Poultry Breeding Co., Ltd. All the other authors have declared no competing interests.
Peer review
Peer review information
Nature Communications thanks Dominic Wright and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, D., Shi, K., The ChickenGTEx Consortium. et al. Egg-laying ChickenGTEx resource deciphers context-specific regulatory effects on fertility traits. Nat Commun 17, 553 (2026). https://doi.org/10.1038/s41467-025-67245-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67245-y