Abstract
Despite systemic viral suppression, people living with HIV (PLHIV) on antiretroviral therapy (ART) remain highly susceptible to pneumococcal colonisation and disease. Here, we show that long-term ART does not restore nasal mucosal immunity. Using flow cytometry, single-cell transcriptomics, and neutrophil functional assays, we identify a persistent mucosal immune signature in PLHIV-ART > 1 yr marked by epithelial-driven neutrophilic inflammation, T cell exhaustion, and cellular senescence. Neutrophils exhibit mitochondrial stress, senescence-associated secretory phenotype (SASP) gene expression, and impaired oxidative burst, particularly in individuals with pneumococcal carriage. Epithelial cells express elevated neutrophil-recruiting ligand genes, while nasal T cells display pro-apoptotic and exhaustion gene profiles. Neutrophilic inflammation is strongly associated with pneumococcal carriage density, implicating a feedforward loop between inflammation and microbial persistence. Our findings reveal tissue-specific immune dysregulation despite ART and suggest that targeting epithelial-immune signalling or neutrophil senescence may offer novel therapeutic avenues to reduce respiratory pathogen burden in PLHIV.
Similar content being viewed by others
Introduction
People living with HIV (PLHIV) remain at an elevated risk of respiratory infections, including Streptococcus pneumoniae (S. pneumoniae), despite long-term antiretroviral therapy (ART) and viral suppression1,2,3,4. S. pneumoniae continues to be a leading cause of morbidity and mortality in PLHIV on suppressive ART, with studies showing a 30–100-fold increased risk of invasive pneumococcal disease, alongside a 5-fold increase in mortality in PLHIV on suppressive ART than HIV-uninfected5,6,7,8. In addition, nasopharyngeal carriage is more frequent and persistent in PLHIV on suppressive ART than in HIV-uninfected individuals3. Pneumococcal carriage is a prerequisite for pneumococcal disease9. These patterns suggest that immune reconstitution under ART may be incomplete, particularly at mucosal surfaces where pathogen colonisation initiates.
While ART is known to restore peripheral CD4⁺ T-cell counts10,11 and reduce systemic immune activation12, its effects on local immunity at the upper respiratory tract mucosa remain poorly understood. The nasal mucosa, the primary site of pneumococcal colonisation, plays a critical role in host-pathogen interactions13,14,15. However, a limited number of studies have characterised immune cell composition, epithelial-immune interactions, or functional mucosal responses in this tissue in the context of HIV.
Evidence from viral infection studies and controlled human infection models have shown that mucosal inflammation can enhance pneumococcal acquisition and persistence16,17,18. Neutrophils as among the predominant innate immune cells in the nasal mucosa19,20, are key to controlling colonisation21,22. However their recruitment and function are tightly regulated by epithelial cues23,24,25 and their dysregulation could impair mucosal defence. Whether HIV infection alters this epithelial-neutrophil axis and whether such changes persist despite ART remain open questions.
To address this knowledge gap we investigated the upper respiratory mucosal immune landscape in HIV-uninfected adults and PLHIV at different stages of ART. We applied multiparameter flow cytometry single-cell transcriptomics and neutrophil functional profiling to evaluate immune cell composition, epithelial-immune communication and their relationship with pneumococcal carriage. This study aims to define how HIV and ART shape nasal immunity and to identify mechanisms that may underlie persistent pneumococcal colonisation susceptibility in PLHIV on long-term ART.
Results
Cohort characteristics and sampling
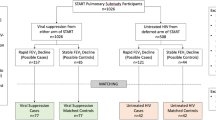
We recruited 132 adults in Blantyre, Malawi. The cohort comprised 37 PLHIV within 3 months of ART initiation (PLHIV-ART < 3 months) and 49 PLHIV on ART for over 1 year (PLHIV-ART > 1 year), both enroled from Gateway Voluntary Counselling and Testing Centre and Lighthouse ART Clinic. In addition, we recruited 46 HIV-uninfected adults (HIV-adults) from Gateway Voluntary Counselling and Testing Centre and surrounding communities in Blantyre through our community engagement work (Fig. 1a). Nasal mucosal cells, nasal lining fluid and peripheral blood were collected from all participants.
Nasal mucosal cells were collected from the inferior turbinate and stained with fluorochrome-conjugated antibodies to identify major immune populations. a Study overview and sampling strategy for HIV-uninfected adults (HIV-adults, n = 46), PLHIV on ART for <3 months (PLHIV-ART < 3 months, n = 37) and PLHIV on ART for >1 year (PLHIV-ART > 1 year, n = 49). b Box plots illustrating the abundance of neutrophils, monocytes and T cells in HIV-adults (n = 15). c, d Comparison of nasal neutrophil and monocyte abundance among HIV- (n = 32), ART < 3 m (n = 25) and ART > 1 year (n = 31) individuals. e, f Frequencies of nasal CD4⁺ and CD8⁺ T cells in HIV-adults (n = 16), PLHIV-ART < 3 months (n = 14) and PLHIV-ART > 1 year (n = 20). g–i Abundance of CD3⁺CD161⁺⁺TCRVα7.2⁺ MAIT cells, CD3⁺TCRγδ⁺ T cells and CD3⁺CD56⁺ natural killer T cells in HIV-adults (n = 18), PLHIV-ART < 3 months (n = 14) and PLHIV-ART > 1 year (n = 20). Immune cell abundance was normalised to epithelial cell count (Epcam⁺) to account for variation in sample yield. Box plots show the interquartile range (25th–75th percentiles). The bold horizontal line inside each boxplot indicated the median. Whiskers extend to the smallest and largest values within 1.5× the interquartile range (IQR) from the lower and upper quartiles, respectively. Individual points beyond the whiskers denote outliers. Statistical significance was assessed using two-sided Wilcoxon rank-sum tests; P < 0.05 was considered significant.
Baseline characteristics, including median age, sex distribution and number of under-five children per household, were comparable across study groups (Table 1). Socioeconomic status (SES) was higher among HIV-adults compared to PLHIV groups, although no significant SES difference was noted between PLHIV-ART < 3 months and PLHIV-ART > 1 year (Table 1). Among PLHIV, the median duration of ART was 1 month (IQR; 0, 1) for PLHIV-ART < 3 months and 5.7 (IQR; 3.1, 9.25) years for PLHIV-ART > 1 year. Median peripheral CD4⁺ T-cell counts were 475 cells/μl (IQR; 285 cells/μl, 674 cells/μl) and 512 cells/μl (IQR; 366 cells/μl, 671 cells/μl) for PLHIV-ART < 3 months and PLHIV-ART > 1 year, respectively. Detectable plasma viral load was observed in 24% (9/37) of PLHIV-ART < 3 months and 10% (5/49) of PLHIV-ART > 1 year (Table 1).
HIV alters nasal immune cell composition despite long-term ART
To investigate the impact of HIV on upper respiratory tract mucosal immunity, we performed flow cytometry for profiling of nasal cells collected using nasal curettes from HIV-adults, PLHIV-ART < 3 months and PLHIV-ART > 1 year (Fig. 1a). In HIV-adults, the nasal mucosa was predominantly populated by CD66b⁺ neutrophils and CD3⁺ T cells, with a smaller proportion of CD14⁺ monocytes (Fig. 1b and Supplementary Fig. S1a). Neutrophil abundance was significantly higher in PLHIV-ART > 1 year compared to HIV-adults, but only marginally higher compared to PLHIV-ART < 3 months, however, this was not statistically significant (Fig. 1c). Monocyte levels were comparable across all cohorts (Fig. 1d). Subset analysis of lymphocyte populations revealed a lower proportion of CD4⁺ T cells in PLHIV-ART < 3 months relative to HIV-adults and PLHIV-ART > 1 year participants, with the latter two groups showing similar levels (Fig. 1e and Supplementary Fig. S1b). There were no significant differences observed in the proportions of CD8⁺ T cells, mucosal-associated invariant T (MAIT) cells (CD3⁺CD161⁺⁺TCRVα7.2⁺), or γδ T cells (CD3⁺TCRγδ⁺) across study groups (Fig. 1f–h). On the other hand, natural killer T (NKT) cells were higher in PLHIV-ART > 1 year compared to PLHIV-ART < 3 months, but not significantly different from HIV-adults (Fig. 1i). These findings indicate persistent perturbations in nasal immune cell composition among PLHIV on long-term ART, particularly involving neutrophil accumulation.
Activated nasal neutrophils accumulate in PLHIV despite long-term ART
Given the enrichment of nasal CD66b⁺ neutrophils in PLHIV-ART > 1 year, we next assessed their activation (CD11b, CD62L)26, maturation (CD10)27 and granule content, tissue residency and maturity (CD63)28 profiles using multiparameter flow cytometry. Uniform Manifold Approximation and Projection (UMAP) of immune cells from matched peripheral blood and nasal samples revealed eight clusters, four of which were CD66b⁺neutrophil subsets (Fig. 2a). Clusters 4 (CD11b⁺CD10⁻CD63+CD62L−CD14−) and 6 (CD11b⁺CD10⁺CD63+CD62L+CD14−) were predominantly observed in blood, while cluster 7 (CD11b⁺CD10⁺CD63−CD62L−CD14−), representing activated mature neutrophils, was enriched in the nasal mucosa. Cluster 7 abundance was elevated in the nasal mucosa of both PLHIV-ART < 3 months and PLHIV-ART > 1 year compared to HIV-adults (Fig. 2a). This was corroborated by conventional gating on an expanded cohort, which showed significantly higher frequencies of CD66b⁺CD11b⁺⁺ neutrophils in PLHIV-ART > 1 year than in HIV-adults (Fig. 2b). These cells strongly correlated with the UMAP-identified cluster 7 phenotype (Supplementary Fig. S2a), confirming population overlap.
Nasal mucosal cells and paired peripheral blood samples were collected from HIV-adults (n = 10), PLHIV-ART < 3 months (n = 10) and PLHIV-ART > 1 year (n = 10). Cells were stained with fluorochrome-conjugated antibodies targeting CD45, CD66b, CD14, CD10, CD11b, CD62L and CD63. a UMAP visualisation of 60,000 concatenated CD45⁺ cells from nasal mucosa and peripheral blood, clustered using FlowSOM. Heatmap indicates average expression of surface markers across clusters. Tables display the relative frequencies of each cluster in the blood and nasal compartments across the three study groups. b Proportion of nasal CD66b⁺CD11b⁺⁺ neutrophils across study groups. c Concentration of myeloperoxidase (MPO) in nasal lining fluid measured by ELISA. Box plots show the interquartile range (25th–75th percentiles). The bold horizontal line inside each boxplot indicates the median. Whiskers extend to the smallest and largest values within 1.5× the interquartile range (IQR) from the lower and upper quartiles, respectively. Individual points beyond the whiskers denote outliers. d Scatter plot showing correlation between nasal neutrophil and monocyte abundance. e Scatter plot showing correlation between nasal neutrophil abundance and MPO levels and nasal monocyte abundance and MPO. Error bars show the 95% confidence interval around the fitted mean. f–h Correlation dot plots between nasal cytokines and myeloid cell subsets in HIV-adults (n = 30), PLHIV-ART < 3 months (n = 29) and PLHIV-ART > 1 year (n = 34) groups. Cytokine levels were measured using a multiplex MSD assay. Dot size reflects statistical significance (P < 0.05); colour denotes Spearman’s correlation coefficient (R). Statistical comparisons were assessed using two-sided Wilcoxon rank-sum test (b, c) and two-sided Spearman correlation (d–h); P < 0.05 was considered significant.
To assess neutrophil-associated inflammation we collected nasal lining fluid using nasosorption filters and measured nasal myeloperoxidase (MPO) levels, which were significantly elevated in PLHIV-ART > 1 year relative to HIV-adults (Fig. 2c). MPO levels correlated with nasal CD66b⁺ neutrophil abundance (Fig. 2d) but not with nasal monocyte abundance (Fig. 2e), implicating neutrophils as the primary MPO source. However, despite elevated neutrophil abundance, nasal concentrations of proinflammatory cytokines and chemokines, including TNF-α, IL-6, IL-8, IL-1β, IFN-γ, IL-2, IL-12p70, IL-13, IL-4 and IL-10, did not differ significantly across study groups (Supplementary Fig. S2b–k). Correlation matrices further revealed that in HIV-adults, neutrophil chemoattractants (IL-8, IL-1β, TNF-α and IL-6) were positively associated with nasal neutrophil abundance (Fig. 2f), whereas these associations were lost in the PLHIV groups (Fig. 2g, h), suggesting dysregulation of neutrophil recruitment pathways. Altogether, these findings indicate neutrophilic inflammation and activation in the nasal mucosa of PLHIV-ART > 1 year, characterised by high neutrophil abundance and high levels of MPO.
Airway epithelial-neutrophil crosstalk is amplified in PLHIV on long-term ART
To investigate the molecular pathways contributing to nasal neutrophilia in PLHIV-ART > 1 year, we performed single-cell RNA sequencing (scRNA-seq) on nasal cells from 3 HIV-adults, 3 PLHIV-ART < 3 months and 5 PLHIV-ART > 1 year from the same cohort (Fig. 3a and Supplementary Table S1). All individuals were asymptomatic and tested negative for common respiratory viruses. Plasma and nasal lining fluid HIV viral loads were undetectable in all PLHIV participants, except in the plasma of one PLHIV-ART < 3 months. Using the 10x Genomics 5′ platform, we generated scRNA-seq libraries from freshly isolated viable nasal cells, yielding 22,119 cells that passed quality control. Dimensionality reduction using UMAP identified 11 major transcriptional clusters (Fig. 3c), which were annotated by canonical gene expression and validated through projection onto the Human Cell Atlas using SingleR and Azimuth29,30,31. The epithelial compartment included goblet cells (CYP2F1, SERPINB3, MUC5AC, SPDEFlow and VMO1), secretory cells (SERPINB3, VMO1 and MUC5AClow), basal cells (CYP2F1 and SERPINB3high), ciliated cells (CAPS, CFAP157, DNAF1 and FOXJ1), squamous cells (SPRR3, SPRR2A and KRT4), Ionocytes, a newly identified epithelial celltype responsible for regulating ion transport and osmotic regulation (CFTR, SCNN1B and RARRES2) and deuterosomal cells, a transient, intermediate airway epithelial cells that serve as precursors to multiciliated cells (CDC20B, CCNO and MSMB). The immune clusters comprised neutrophils (SRGN, CSF3R, GOS2 and CXCL8), T cells (CD3E, CD3D, CD8A and PTPRC), B cells (PTPRC, CD79A, MS4A1 and CD19), monocytes/macrophages (TYROBP, HLA-DPA1 and HLA-DPB1) and dendritic cells (CD83, CD86, LYZ and FCER1G). Further sub-clustering of the 4099 immune cells revealed nine subsets: CD8⁺ T cells, NKT cells, B cells, plasma cells, neutrophils, monocytes, macrophages, dendritic cells and mast cells (Fig. 3d). Consistent with flow cytometry findings (Figs. 1c and 2d), we observed a higher abundance of neutrophils in the nasal mucosa of PLHIV-ART > 1 year compared to HIV-adults (Fig. 3e, f).
Nasal cells from HIV-uninfected adults (HIV-adults, n = 3), PLHIV on ART for <3 months (PLHIV-ART < 3 months, n = 3) and PLHIV on ART for >1 year (PLHIV-ART > 1 year, n = 5) were processed for single-cell RNA sequencing using the 10x Genomics 5′ platform. After quality control and normalisation, 22,119 cells were analysed using the Seurat pipeline with Harmony integration. a UMAP visualisation of all nasal cells coloured by major cell types. b Distribution of major cell clusters across study groups. c Violin plots showing expression of canonical marker genes used for broad cell-type identification. d UMAP visualisation of 4099 immune cells coloured by fine-grained immune cell annotations. e Bar plot showing the distribution of immune cell subsets across study groups. f Box plots comparing neutrophil abundance among HIV–, PLHIV-ART < 3 months and PLHIV-ART > 1 year groups. Box plots show the interquartile range (25th–75th percentiles). The bold horizontal line inside each boxplot indicates the median. Whiskers extend to the smallest and largest values within 1.5× the interquartile range (IQR) from the lower and upper quartiles, respectively. Individual points beyond the whiskers denote outliers. Cell type: NK natural killer cells; Mono monocytes; Mac macrophages; DCs dendritic cells. Statistical significance was determined using the two-sided Wilcoxon rank-sum test; P < 0.05 was considered significant.
Epithelial cues are known to regulate neutrophil recruitment23,24,25. To determine which epithelial-immune interactions may be driving this neutrophilic response, we performed cell-to-cell communication inference using Multinichenet32. In PLHIV-ART > 1 year, epithelial-neutrophil interactions were significantly expanded relative to HIV-adults and PLHIV-ART < 3 months (Fig. 4a). Notably, PLHIV-ART > 1 year showed strong ligand-receptor signalling pathways associated with neutrophil recruitment, including CXCL1-CXCR1, MIF-CD44 and ADGRE5-CD55, originating from basal, secretory, goblet, squamous and ciliated epithelial subsets (Fig. 4a). Gene set scores for these prioritised interactions were elevated in all epithelial subsets of PLHIV-ART > 1 year compared to HIV-adults and were also higher than in PLHIV-ART < 3 months across all epithelial subsets except squamous cells (Fig. 4b–f). In PLHIV-ART < 3 months, distinct ligand-receptor pairs predominated, including ICAM1-ITGAX/ITGB2, ICAM1-MSN, ICAM1-IL2RG and BST2-LILRA5, which also involved major epithelial subsets and neutrophils (Fig. 4a). In contrast, epithelial-neutrophil interactions were sparse in HIV-adults and primarily mediated by MFAP2-NOTCH1 signalling between basal and secretory cells (Fig. 4a).
Epithelial-to-neutrophil ligand-receptor interactions were inferred using the Multinichenet framework to model intercellular communication. a Circos plots illustrating significant epithelial cell-to-neutrophil communication networks in HIV-uninfected adults (HIV-adults), PLHIV on ART for <3 months (PLHIV-ART < 3 months) and PLHIV on ART for >1 year (PLHIV-ART > 1 year). Shown are top-ranked ligand–receptor pairs with adjusted P < 0.05 and expression in >10% of cells. Gene module scores representing average expression of neutrophil recruitment-associated ligand–receptor pairs in epithelial subsets: b basal cells; c ciliated cells; d goblet cells; e squamous cells; f secretory cells. Scores were calculated using the AddModuleScore function in Seurat v5, based on expression of curated neutrophil-recruitment gene sets versus randomly selected control genes. Statistical differences were assessed using Holm-adjusted two-sided Wilcoxon rank-sum tests; adjusted P < 0.05 was considered significant.
Additional analysis of immune cell-neutrophil interactions in PLHIV-ART > 1 year revealed enriched signalling pathways including CXCL1-CXCR1, IL-1β-IL-1R2 and IL-18-IL18RAP, predominantly involving neutrophils, macrophages and dendritic cells (Supplementary Fig. S3). In contrast, PLHIV-ART < 3 months exhibited distinct ligand-receptor interactions, including ICAM1-IL2RG and TNF-TNFRSF1A, primarily associated with neutrophils and T cells, while in HIV-adults, these interactions involved IL-17A-IL17RA, DLK2-NOTCH and MFNG-NOTCH1, involving NKT cells and dendritic cells (Supplementary Fig. S3). Collectively, these findings suggest that airway epithelial cells likely contribute to neutrophil recruitment and activation in the nasal mucosa of PLHIV-ART > 1 year.
Nasal neutrophils in PLHIV-ART > 1 year exhibit transcriptional signatures of stress and senescence
To dissect the transcriptional changes underlying neutrophilic inflammation, we performed differential gene expression analysis on nasal neutrophils using the Model-based analysis of single-cell transcriptomics (MAST) pipeline33. In PLHIV-ART > 1 year, neutrophils showed upregulation of mitochondrial stress-associated genes, including MT-ND4, MT-CO1, LYPD2, PRDX1, MSMB and WFDC2, compared to HIV-adults (Fig. 5a and Supplementary Data 1). In contrast, neutrophils from PLHIV-ART < 3 months exhibited elevated expression of antiviral response genes, including IFITM1, IFITM3, IRF7 and XAF1, relative to HIV-adults (Fig. 5b and Supplementary Data 1). When comparing PLHIV-ART > 1 year to PLHIV-ART < 3 months, mitochondrial stress genes such as MT-CO1, MT-ND6, MT-ND5 and PRDX1 remained upregulated in the former, while antiviral response genes, including IFITM1, IFIT2, ISG15, IRF7 and IL1B, were elevated in the latter (Fig. 5c and Supplementary Data 1).
Differential gene expression analysis was performed on neutrophils using the MAST algorithm via the FindMarkers function in Seurat. Volcano plots showing differentially expressed genes in neutrophils comparing: a PLHIV-ART > 1 year vs. HIV-uninfected adults (HIV-adults); b PLHIV-ART < 3 months vs. HIV−; c PLHIV-ART > 1 year vs. PLHIV-ART < 3 months. d Dot plot showing gene co-expression modules enriched in neutrophils from HIV−, ART < 3 months and ART > 1 year groups, calculated using CEMiTool. Dot colour reflects normalised enrichment score (NES), while size corresponds to −log₁₀(adjusted P value) and intersection size with Reactome pathways. Bar graphs of the top 10 significantly enriched pathways from overrepresentation analysis in e Module 1 (M1), overexpressed in PLHIV-ART > 1 year; and f Module 2 (M2), overexpressed in PLHIV-ART < 3 months. g–j Gene module scores for immune-related KEGG pathways enriched in neutrophils, stratified by study group. k–i Functional assessment of neutrophils using: k phagocytic index (HIV-, n = 25; PLHIV-ART < 3 months, n = 19; PLHIV-ART > 1 year, n = 22); l oxidative index (HIV-, n = 24; PLHIV-ART < 3 months, n = 19; PLHIV-ART > 1 year, n = 20). Indices were calculated by multiplying the frequency of neutrophils that internalised or oxidised beads with their corresponding median fluorescence intensity (MFI). Box plots show the interquartile range (25th–75th percentiles). The bold horizontal line inside each boxplot indicates the median. Whiskers extend to the smallest and largest values within 1.5× the interquartile range (IQR) from the lower and upper quartiles, respectively. Individual points beyond the whiskers denote outliers. Statistical significance was determined using Holm-adjusted two-sided Wilcoxon rank-sum tests; P < 0.05 was considered significant. SASP senescence-associated secretory phenotype; NES normalised enrichment score; −log10 Adj.p-value, −log10 of Holm adjusted p-value; IFN signalling, interferon signalling; MHC1 APC major histocompatibility complex class-1 antigen-presenting cells.
To further interrogate gene expression patterns, we employed CEMiTool34 to identify co-expressed gene modules enriched in neutrophils across the study groups. Two modules were uniquely enriched, module M1 in PLHIV-ART > 1 year and module M2 in PLHIV-ART < 3 months (Fig. 5d). Over-representation analysis of M1 revealed enrichment in pathways linked to homoeostatic regulation and the Senescence-Associated Secretory Phenotype (SASP) (Fig. 5e; Supplementary Fig. S4a and Supplementary Data 2), while M2 was enriched for pathways involved in cytokine signalling, interferon response, antigen presentation and immunoregulatory interactions (Fig. 5f; Supplementary Fig. S4b and Supplementary Data 2). Moreover, SASP pathway gene module scores were significantly higher in neutrophils from both PLHIV-ART > 1 year and PLHIV-ART < 3 months compared to HIV-adults (Fig. 5g and Supplementary Fig. S4c), indicating persistent inflammatory signalling. In contrast, module scores for genes involved in interferon signalling, Nef-mediated HIV replication inhibition and MHC class I antigen presentation were markedly higher in PLHIV-ART < 3months compared to both HIV-adults and PLHIV-ART > 1 year (Fig. 5h–j and Supplementary Fig. S4c), consistent with active antiviral responses during early ART.
Furthermore, based on the premise that the neutrophils in the nasal mucosa were likely recruited from the blood, we evaluated the functional competence of blood neutrophils, assessing phagocytosis and oxidative burst in the three study groups using our in-house published bead-based assay35,36,37. The phagocytic capacity of blood neutrophils was comparable across the study groups (Fig. 5k). In contrast, neutrophil oxidative burst capacity was significantly lower in both PLHIV-ART > 1 year and PLHIV-ART < 3 months relative to HIV-adults (Fig. 5i), suggesting potential impairment in killing capacity. Together, these data indicate that nasal neutrophils from PLHIV-ART > 1 year adopt a transcriptional profile marked by cellular stress and senescence, indicating inflammaging despite virological suppression.
Nasal T cells in PLHIV on long-term ART exhibit transcriptional signatures of exhaustion, senescence and apoptosis
Given that chronic inflammation can drive T cell exhaustion and senescence38, we hypothesised that neutrophilic inflammation in PLHIV-ART > 1 year is associated with altered T cell function. Differential gene expression analysis revealed that nasal T cells from PLHIV-ART > 1 year exhibited upregulation of immune activation genes (HLA-B, HLA-DQA1 and TRAV6), chemokine genes (CXCR4, CCL4 and CCL3) and ageing-associated genes (FKBP5, HMGB2 and BTG1) compared to HIV-adults (Fig. 6a and Supplementary Data 1). In PLHIV-ART < 3 months, nasal T cells showed elevated expression of cytotoxicity-associated genes (GZMH, GNLY, GZMB and NKG7) and immune activation genes (LAG3, HLA-DQA1, HLA-A, IRF1 and STAT1) relative to HIV-adults (Fig. 6b and Supplementary Data 1). Notably, TRAV6, CXCR4 and FKBP5 remained significantly upregulated in PLHIV-ART > 1 year compared to PLHIV-ART < 3 months (Fig. 6c and Supplementary Data 1), suggesting persistent transcriptional changes with long-term ART.
Differential gene expression analysis was performed on nasal T cells using the MAST algorithm via Seurat’s FindMarkers function. Volcano plots showing differentially expressed genes in T cells comparing: a PLHIV-ART > 1 year vs. HIV-uninfected adults (HIV-adults); b PLHIV-ART < 3 months vs. HIV−; c PLHIV-ART > 1 year vs. PLHIV-ART < 3 months. d Dot plot showing co-expression gene modules enriched in T cells across study groups, identified using CEMiTool. Dot size represents −log₁₀(adjusted P value) and colour denotes normalised enrichment score (NES). Bar plots of the top 10 enriched pathways in e Module 1 (M1), overexpressed in PLHIV-ART > 1 year and f Module 2 (M2), overexpressed in PLHIV-ART < 3 months, based on gene set overrepresentation analysis (ORA) using Reactome pathways. Gene module scores for key immunological pathways in nasal T cells, grouped by HIV status: g exhaustion and senescence-associated genes; h activation-related genes; i intrinsic apoptosis genes; and j extrinsic apoptosis genes. Statistical comparisons were performed using a Holm-adjusted two-sided Wilcoxon rank-sum tests; P < 0.05 was considered significant. NES normalised enrichment score; −log10 Adj. p-value, −log10 of Holm-adjusted p-value.
Gene co-expression module analysis using CEMiTool identified distinct modules enriched in each PLHIV group. The M1 module, enriched in PLHIV-ART > 1 year, was associated with pathways related to apoptosis and cell-to-cell communication (Fig. 6d, e; Supplementary Fig. S5a and Supplementary Data 2), while the M2 module, enriched in PLHIV-ART < 3 months, highlighted pathways involved in antigen presentation and immunoregulation (Fig. 6f; Supplementary Fig. S5b and Supplementary Data 2). Gene set scoring further revealed that exhaustion and senescence-associated genes (S100A10, PDCL3, CTLA4, PDCD7, PDCD5, LAG3, KLRG1, TIGIT, CDKN1A and NR4A1) were more highly expressed in T cells from both PLHIV-ART > 1 year and PLHIV-ART < 3 months compared to HIV-adults (Fig. 6g). Similarly, activation-related genes (STAT1, IRF4, HLA-DRA, HLA-DRB1, JUN, GZMB and FOS) were elevated in both PLHIV groups (Fig. 6h, i), indicating ongoing T cell activation.
Considering that chronic T cell activation results in activation-induced cell death39, we explored T cell death pathways by examining intrinsic and extrinsic apoptosis-associated gene sets. The intrinsic apoptosis gene set (BCL2, BAK, CASP3, CASP8, TNFRSF1A, TP53, BCL2L11, APAF1 and NFKB1) showed higher module scores in PLHIV-ART > 1 year compared to PLHIV-ART < 3 months, while no significant differences were observed for the extrinsic apoptosis gene set (FAS, TNFRSF1A, TNFRSF10A and TNFRSF10B) (Fig. 6j). Taken together, these findings suggest that nasal T cells from PLHIV-ART > 1 year are transcriptionally programmed toward heightened immune activation, exhaustion and intrinsic apoptotic susceptibility.
Neutrophilic inflammation is associated with pneumococcal carriage in PLHIV on long-term ART
Next, we investigated whether persistent neutrophilic inflammation in PLHIV-ART > 1 year is linked to increased susceptibility to pneumococcal carriage. Among PLHIV-ART > 1 year, neutrophil abundance was significantly higher in individuals with pneumococcal carriage compared to those without (Fig. 7a). Monocyte abundance followed a similar pattern, with elevated levels in carriage-positive individuals within the same group (Fig. 7b). In contrast, no significant differences in neutrophil or monocyte abundance were observed between carriage-positive and carriage-negative individuals among HIV-adults or PLHIV-ART < 3 months (Fig. 7a, b).
Nasal cells were collected using a Rhinoprobe and stained with fluorochrome-conjugated antibodies to identify neutrophils (CD66b⁺), monocytes (CD14⁺) and epithelial cells (EpCAM⁺). Neutrophil and monocyte abundance was calculated as the ratio of immune cells to epithelial cells. Pneumococcal carriage and density were determined from nasopharyngeal swabs using standard culture methods and Miles and Misra plating. Functional neutrophil activity was assessed by measuring peripheral blood phagocytic and oxidative capacity following ex vivo stimulation with lipopolysaccharide (LPS) and incubation with IgG-coated fluorescent silica beads. Phagocytosis and oxidation indices were computed as the product of median fluorescence intensity (MFI) and the frequency of bead-positive neutrophils. a, b Violin plots comparing nasal neutrophil and monocyte abundance in pneumococcal carriage-negative (HIV-, n = 22; PLHIV-ART < 3 months, n = 15; PLHIV-ART > 1 year, n = 21) and carriage-positive (HIV-, n = 5; PLHIV-ART < 3 months, n = 9; PLHIV-ART > 1 year, n = 12) individuals stratified by HIV status and ART duration. Scatter plots showing correlation between nasal pneumococcal carriage density and c CD11b⁺⁺ neutrophil frequency (HIV-, n = 25; PLHIV-ART < 3 months, n = 18; PLHIV-ART > 1 year, n = 34); d monocyte abundance (HIV-, n = 38; PLHIV-ART < 3 months, n = 31; PLHIV-ART > 1 year, n = 45). Error bars show the 95% confidence interval around the fitted mean. Violin plots comparing peripheral blood neutrophil e phagocytosis index and f oxidation index in carriage-negative (HIV-, n = 16; PLHIV-ART < 3 months, n = 12; PLHIV-ART > 1 year, n = 7) and carriage-positive (HIV-, n = 9; PLHIV-ART < 3 months, n = 7; PLHIV-ART > 1 year, n = 15) individuals. Statistical significance was assessed using a Holm-adjusted two-sided Wilcoxon rank-sum tests; P < 0.05 was considered significant.
Further, we found that the proportion of activated nasal CD11b⁺⁺ neutrophils negatively correlated with pneumococcal carriage density in PLHIV-ART > 1 year, suggesting a potential contribution to carriage density control without clearance (Fig. 7c). No such correlations were observed in the HIV-adults or PLHIV-ART < 3 months groups. Monocyte abundance did not correlate with carriage density in any group (Fig. 7d). Using blood neutrophils as a surrogate of infiltrating nasal neutrophil functional capacity, we observed that the phagocytic capacity was similar regardless of carriage status across all groups (Fig. 7e), but the neutrophil oxidative burst capacity was significantly lower in carriage-positive PLHIV-ART > 1 year compared to their carriage-negative counterparts (Fig. 7f). This impaired neutrophil oxidative burst capacity was not observed in HIV-adults or PLHIV-ART < 3 months. Collectively, these data support an association between persistent nasal neutrophilic inflammation, functional impairment and potential increased susceptibility to pneumococcal carriage in PLHIV-ART > 1 year.
Discussion
Despite long-term viral suppression with ART PLHIV remain disproportionately vulnerable to respiratory infections, including S. pneumoniae1,2. Our findings show that ART does not fully reconstitute upper respiratory mucosal immunity. In PLHIV on ART for longer than 1 year, we observed a distinct nasal immune profile characterised by sustained neutrophilic inflammation, reprogrammed epithelial-immune signalling and transcriptional signatures of T cell exhaustion and cellular senescence. These immune perturbations were associated with increased pneumococcal carriage, offering potential mechanistic insight into the persistent risk of pneumococcal carriage3,4 and potentially invasive pneumococcal disease1,2 in this population.
A central finding was the expansion of nasal neutrophils in PLHIV-ART > 1 year, enriched for mitochondrial stress and SASP-related gene programmes. Notably, SASP-related inflammation can perpetuate neutrophil recruitment through paracrine mechanisms40, suggesting the presence of a chronic, self-reinforcing inflammatory circuit in the nasal mucosa of PLHIV. Our cell-to-cell communication analyses identified epithelial cells as key orchestrators of this altered mucosal state. Moreover, neutrophils exhibited diminished oxidative burst in peripheral blood, especially in pneumococcal carriers. Considering that neutrophils are generally short-lived and that nasal neutrophils likely traffic from peripheral blood, it is plausible to assume that the defective neutrophilic oxidative function extends to nasal neutrophils. This is consistent with prior reports linking neutrophil dysfunction in HIV to increased susceptibility to secondary infections41 and those linking peripheral neutrophil dysfunction to pneumococcal nasopharyngeal carriage susceptibility17. In PLHIV-ART > 1 year, epithelial subsets expressed higher levels of neutrophil-activating ligands genes42,43,44,45,46, such as CXCL1, MIF and ADGRE5, potentially sustaining recruitment of functionally compromised neutrophils. These ligand-receptor interactions were attenuated or absent in HIV-uninfected adults, suggesting that HIV-associated epithelial reprogramming may be a persistent consequence of chronic immune activation or microbial exposure. Given the known impact of HIV on mucosal barrier integrity and the influence of cotrimoxazole prophylaxis in PLHIV on microbiota composition47, epithelial inflammation may reflect both direct and indirect consequences of HIV infection. However, while we used peripheral neutrophils as a surrogate, direct assessment of mucosal neutrophil function remains an important future direction.
Surprisingly, we did not detect corresponding increases in canonical proinflammatory cytokines and chemokines across study groups. It is plausible that chemokine activity is spatially restricted or temporally dynamic, thus, not fully captured by bulk single-timepoint nasal lining fluid measurements. However, neutrophil chemo-attractants correlated positively with neutrophil abundance in HIV-uninfected adults, consistent with physiological recruitment patterns48,49,50,51,52. By contrast, this correlation was absent in ART-treated PLHIV, potentially indicating dysregulated chemotactic signalling or compensatory epithelial-derived pathways. Further spatially resolved and longitudinal studies are needed to ascertain the recruitment pathways sustaining mucosal neutrophilia in ART-treated PLHIV.
We also observed transcriptional evidence of T cell exhaustion, senescence and apoptosis in the nasal mucosa of PLHIV, particularly in those on long-term ART. This immune profile mirrors observations from gastrointestinal53,54 and pulmonary tissues53,54 in HIV and likely reflects incomplete mucosal immune restoration. It is also consistent with observations in peripheral blood, where a more proinflammatory transcriptome state in T cells remained during ART55. Moreover, the upregulation of apoptotic and senescence pathways in nasal T cells from PLHIV also highlights the broad immune dysregulation influenced by SASP40,56,57,58. T cells play an important role in the host response to HIV infection and S. pneumoniae59,60,61,62. Therefore, these altered T cell states may further contribute to suboptimal control of local bacterial burden, though functional validation of these cells remains necessary.
Importantly, our study provides evidence of a strong association between mucosal inflammation, specifically neutrophilic abundance and dysfunction and pneumococcal carriage. Among PLHIV-ART > 1 year, carriage prevalence and density were significantly higher in individuals with increased nasal neutrophil abundance and reduced oxidative function. This association was not observed in HIV-uninfected adults or recently treated PLHIV, suggesting that long-term ART may promote a chronic inflammatory niche conducive to pneumococcal colonisation. Inflammatory mucosal environments are known to alter epithelial glycosylation patterns and increase the availability of host-derived nutrients such as sialic acids, promoting S. pneumoniae adherence, colonisation and outgrowth63,64,65,66. Moreover, pneumococcus can amplify inflammation through epithelial activation67, suggesting a bidirectional relationship. The inability of neutrophils in PLHIV-ART > 1 year to resolve carriage despite increased abundance suggests functional alteration, possibly driven by chronic SASP signalling or microbial persistence.
While our cross-sectional study cannot infer causality, the consistent link between inflammation and pneumococcal carriage, combined with reduced neutrophil function, supports a model in which chronic mucosal inflammation undermines host pneumococcal clearance. These findings echo experimental human carriage studies, where inflammation similarly increased susceptibility to colonisation17,68. Our data suggest that such vulnerability extends to PLHIV, despite immune reconstitution at the systemic level. This study is strengthened by its integration of flow cytometry, single-cell transcriptomics and neutrophil function assays in a well-characterised cohort stratified by ART duration. However, several limitations must be acknowledged. Single-cell analyses were conducted on a limited sample size, potentially underpowering the detection of rare cell subsets. Functional studies were restricted to blood-derived neutrophils and the sample size was smaller, likely underpowering subgroup analysis. Direct assessment of mucosal neutrophils remains an important future direction. We also lacked assessment of microbial community composition, which may shape mucosal immunity and carriage risk.
In conclusion, our findings uncover a persistent mucosal immune signature in PLHIV on long-term ART, marked by epithelial-driven neutrophilic inflammation, neutrophil senescence and T cell dysfunction, that associates with increased susceptibility to pneumococcal carriage. These findings highlight the need for immune interventions beyond systemic viral control, including strategies to reprogramme epithelial-immune interactions, enhance neutrophil function, or interrupt chronic SASP signalling using senotherapeutics56. Addressing mucosal immune dysfunction may be critical for reducing respiratory pathogen burden in this population.
Methods
Study design
The study was a cross-sectional observational cohort design, conducted between September 2021 and December 2023 and aimed to assess upper airway mucosal immune responses in PLHIV at different stages of ART. Healthy adults aged 18–45 years were recruited from Queen Elizabeth Central Hospital Lighthouse HIV Clinic and Blantyre District Office Gateway Clinic in Blantyre, Malawi, during routine HIV or ART clinics and from surrounding communities following community engagement activities. Participants were stratified into three groups: (1) HIV-uninfected adults, (2) PLHIV on ART for less than 3 months (PLHIV-ART < 3 months) and (3) PLHIV on ART for more than 1 year (PLHIV-ART > 1 year). HIV status for PLHIV on ART was self-reported and confirmed in the health passport book (a personal health record), for the HIV-uninfected individuals, this was confirmed using a standard HIV rapid test protocol (2 independent rapid tests). ART duration and regimen were confirmed using official clinic records and ART adherence was confirmed via official clinic pharmacy refill records. In addition, viral control was confirmed using plasma viral load suppression. Exclusion criteria included any signs or symptoms of acute illness, chronic medical conditions, active SARS-CoV-2 infection using polymerase chain reaction (PCR), or self-reported SARS-CoV-2 infection within the preceding 14 days. Participants were screened in accordance with local clinical protocols.
Nasal sample collection
Nasal cells were collected by gently scraping the nasal inferior turbinate using curettes (Rhino-Pro, Arlington Scientific). Two curettes per nostril were transferred into cold RPMI 1640 medium supplemented with 10% Fetal Bovine Serum (Sigma-Aldrich), 1% HEPES (Fisher Scientific) and 1% Penicillin and Streptomycin (Fisher Scientific), transported on ice and processed within 30 min. All samples with visible blood or mucus contamination were excluded from the analysis. For flow cytometry analysis, cells were dislodged by pipetting liquid up and down the tip of the curette, centrifuged at 500 × g for 10 min at 4 °C. The supernatant was completely removed and cells were resuspended in 500 µl of PBS (Fisher Scientific). Nasal lining fluid was collected using a synthetic absorptive matrix (Hunt Development (UK) ltd) and eluted with 200 μL PBS (Fisher Scientific), then centrifuged at 1500 × g for 10 min. The eluate was stored at −80 °C until the day of the analysis. The full list of reagents has been provided in Supplementary Table S2.
Flow cytometry and immunophenotyping
Dislodged nasal cells were centrifuged (500 × g, 5 min), resuspended in PBS and stained with Far Red viability dye (1:500) (Thermo Fisher Scientific) and a panel of fluorochrome-conjugated antibodies. Antibodies included markers for lymphocytes (CD3, CD4, CD8, CD56, CD161, TCRvα7.2 and TCRγδ), neutrophils (CD66b, CD63, CD10, CD11b and CD62L), monocytes (CD14) and epithelial cells (EpCAM). Cells and compensation tubes were acquired using a BD LSRFORTESSA™ cytometer (BD Biosciences). A minimum of 50,000 total events was acquired per sample and FlowJo v10.10.0 (BD Biosciences) was used for analysis. The full list of reagents has been provided in Supplementary Table S2.
Ex vivo neutrophil bead-based phagocytosis and oxidation assay
We adapted a previously described bead-based assay to assess the phagocytosis and oxidative capacity of peripheral blood neutrophils 35,37. The assay uses silica beads derivatised with a calibration fluorochrome (Alexa 405-SE) and the fluorogenic reporter substrate Oxyburst Green, succinimidyl ester (OGSE) (Molecular Probes, Eugene, OR) to measure superoxide burst. When the beads are internalised by neutrophils, they gain fluorescence intensity proportional to the degree of oxidative burst activity in the phagosome. In brief, 100 μL of peripheral blood was stimulated with 10 ng/mL LPS and co-incubated with 120 μL of IgG-coated silica beads. Samples were incubated in the dark at 37 °C for 45 min on a shaker at 250 rpm. The samples were then washed with 500 μL pf PBS to remove the unphagocytosed beads. The supernatant was carefully discarded and samples were stained with live/dead amine-reactive dye for 15 min in the dark, followed by staining with a cocktail of fluorochrome-conjugated antibodies (CD45 PerCP/Cy5.5, CD66b PE and CD16 PE/Cy7; all from Biolegend). Red blood cells were lysed with 2 mL of 1X BD FACS Lysing Solution and samples were washed twice with 500 μL of PBS. Samples were resuspended in FACS buffer and analysed using a BD LSRFortessa™ flow cytometer equipped with FACSDiva software version 8 (BD Biosciences). Data were processed using FlowJo version 10.10.0 (BD Biosciences). The Phagocytic Index was calculated by multiplying the proportion of Alexa 405+ cells with a median fluorescence intensity of Alexa 405, while the Oxidative Index was calculated by multiplying the proportion of OGSE+ cells with a median fluorescence intensity of OGSE.
Streptococcus pneumoniae culture and quantitation
Nasopharyngeal swabs were tested for the presence of S. pneumoniae using World Health Organisation standardised microbiological culture3,69. Briefly 20 μL of the sample was diluted in a 1:10 dilution using sterile PBS and plated on sheep blood agar (SBG) with gentamicin. The plates were incubated overnight at 37 °C in 5% CO2. Pneumococcal isolates were identified by their colony morphology (alpha-haemolytic on blood agar) and optochin disc susceptibility (Oxoid). Isolates with no or intermediate resistance (zone diameter ≥14 mm) to optochin were further confirmed using the bile solubility test. Individuals positive for S. pneumoniae were designated as carriage positive, while those without were designated as carriage negative. Pneumococcal density was determined by serial dilution3,69,70 culture on SBG and reported as colony-forming units per millilitre.
Multiplex cytokine assay
Proinflammatory cytokines (IFN-γ, IL-1β, IL-2, IL-4, IL-6, IL-8, IL-10, IL-12p70, IL-13 and TNF-α) were quantified in nasal lining fluid using a Mesoscale Discovery 10-plex immunoassay. Following blocking and sample incubation, plates were washed and developed with SULFO-TAG detection antibodies. Signals were read using MESO® QuickPlex SQ 120 MM. Assay details followed the manufacturer’s instructions. Samples below the assay detection limit were given the minimum value for that specific target.
RT-qPCR for detection of respiratory viruses
For respiratory virus detection, nucleic acid was extracted from 50 µL of nasal lining fluid samples using the QIAamp Viral RNA Mini Kit (Qiagen, Germany), following a column-based protocol. The process included lysis with Buffer AVL, ethanol precipitation, binding to silica columns, washing with AW1 and AW2 buffers and elution with 50 µL of Buffer. Respiratory viruses were detected using reverse transcription qPCR (RT-qPCR) targeting the following: respiratory syncytial virus, influenza A virus, HRV, influenza B virus, parainfluenza virus types 1–4 (HPIV), human metapneumovirus, adenovirus, enterovirus and human bocavirus. Reactions were conducted using Luna Universal Probe one-step RT-qPCR Kit (New England Biolabs, E3006S) in a 20 µL volume, with 5 µL of extracted viral RNA/DNA. Cycling conditions were 52 °C for 10 min, 95 °C for 2 min, followed by 45 cycles of 95 °C for 10 s and 55 °C for 30 s. Viral positivity was defined as Ct <38. Positive and negative controls were included in each run.
Myeloperoxidase (MPO) enzyme-linked immunosorbent assay (ELISA)
MPO levels in nasal lining fluid were quantified using a commercial ELISA kit (Invitrogen, UK; BMS2038INST) according to the manufacturer’s instructions. Samples stored at −80 °C were thawed, diluted 1:50 and added in duplicate to pre-equilibrated wells. After 3 h of incubation at room temperature with agitation, plates were washed, developed with TMB substrate and the reaction was stopped after 10 min. Absorbance was read at 450 nm. Duplicate readings with >20% variation were excluded. MPO concentrations were interpolated from standard curves using MyAssays.com.
Single-cell library preparation
Participants for scRNA-seq were randomly recruited using the same recruitment criteria and process as the other participants. Only samples with an initial cell count greater than 200,000 cells, no visible mucus and no blood contamination were included and further processed for single-cell library preparation. Single-cell RNA libraries were generated from freshly isolated nasal curette samples, collected from both nostrils, using the Chromium Next GEM Single Cell 5′ Reagent Kit v2 and Chromium Controller (10x Genomics) according to the manufacturer’s protocol. Nasal cells were dislodged from the curette, washed in FACS buffer (PBS with 3% FBS, 5 mM EDTA and 32 U/mL DNase I) by centrifugation at 400 × g for 5 min and the supernatant was discarded. Cells were resuspended in 1 mL of PBS with 3% FBS, filtered through a 40-μm FlowME filter and 10,000–16,000 cells with ≥80% viability were loaded into microfluidic chips to generate gel beads-in-emulsion.
Purified cDNA was quantified using Qubit 4.0 (Thermo Fisher Scientific) and its size distribution assessed using Agilent High Sensitivity D5000 ScreenTape on a TapeStation 4200 system. Libraries were constructed by fragmentation, end-repair, A-tailing, adaptor ligation and index PCR. Final libraries were quantified and quality-checked, then sequenced on an Illumina NextSeq2000 platform (150 bp paired-end reads) to a depth of ≥50,000 reads per cell. Library sequencing was performed by Glasgow Polyomics (University of Glasgow, UK).
Single-cell RNA-seq data processing and analysis
Fastq files were processed using Cell Ranger (v8.0.1, 10× Genomics) and mapped to the human reference genome (GRCh38), including intronic reads to capture unspliced transcripts. The cell ranger count pipeline performed alignment, barcode demultiplexing and UMI counting, generating gene expression matrices for each sample. The quality metrics for each sample have been summarised in a single cell metrics table (Supplementary Data 3) and the raw data files, including fastq files and processed data, have been deposited in the NCBI’s Gene Expression Omnibus and are accessible through the GEO accession number GSE296893. Downstream analysis was conducted in R (v4.4.1) using Seurat (v5.0.1). Cells with fewer than 200 detected genes or >30% mitochondrial gene expression were excluded. Genes expressed in fewer than 3 cells were removed. Samples were merged and normalised using the SCTransform() function, selecting the top 2000 variable genes for downstream clustering and batch effects across samples were corrected using Harmony.
Principal component analysis was performed for dimensionality reduction, followed by clustering using a shared nearest neighbour modularity optimisation algorithm and visualised via UMAP. Cell types were annotated using canonical marker genes and validated with automated label transfer from the Human Lung Cell Atlas via SingleR and Azimuth. Differential expression analysis was performed using the MAST or DESeq2 function in Seurat. Cell-to-cell communication networks were inferred using Multinichenet(), prioritising significant receptor-ligand interactions across epithelial and immune subsets.
Gene module enrichment and overrepresentation analyses were performed using CEMiTool (Bioconductor), integrating transcriptomic data with curated interactomes. Gene set enrichment analysis identified statistically enriched pathways associated with group-specific modules. All visualisations were generated in R using the ggplot2 and ComplexHeatmap packages.
Ethics statement
This study was conducted in accordance with the Declaration of Helsinki and adhered to all relevant ethical regulations. Ethical approval was granted by the National Health Science Research Committee (protocol #21/24/2680, Malawi) and the Liverpool School of Tropical Medicine Research Ethics Committee (protocol #21-035, UK). All participants provided written informed consent prior to enrolment and were compensated for their time according to established ethical guidelines.
Statistical analysis
All statistical analyses were conducted in R version 4.4.1. Unless otherwise specified, comparisons between groups were performed using non-parametric tests (the Wilcoxon test or Kruskal–Wallis test). Categorical data were analysed using Fisher’s exact test. Correlation analyses were performed using Spearman’s rank correlation coefficient. For single-cell transcriptomic data, differential expression was assessed using MAST or DESeq2 as applied in Seurat FindMarkers() function. Multiplex cytokine data were log-transformed before analysis to approximate normality. Where appropriate, p-values were adjusted for multiple testing using the Benjamini–Hochberg false discovery rate method. A two-sided p < 0.05 was considered statistically significant. Statistical approaches and sample sizes for each analysis are detailed in the relevant figure legends and Supplementary Information.
Data availability
The source data generated in this study have been deposited in the GitHub repository and are accessible through https://doi.org/10.5281/zenodo.17580059. The raw single-cell sequencing fastq files and single-cell processed data have been deposited in the NCBI’s Gene Expression Omnibus and are accessible through the GEO accession number GSE296893.
Code availability
Manuscript code and R script that were used to analyse the datasets are available in the GitHub repository https://github.com/japhiri/HIV-PAPER/tree/main.
References
Alexandrova, Y., Costiniuk, C. T. & Jenabian, M.-A. Pulmonary immune dysregulation and viral persistence during HIV infection. Front. Immunol. 12, 35058937 (2021).
Brown, J. et al. Respiratory symptoms in people living with HIV and the effect of antiretroviral therapy: a systematic review and meta-analysis. Thorax 72, 355–366 (2017).
Phiri, J. et al. Estimating pneumococcal carriage dynamics in adults living with HIV in a mature infant pneumococcal conjugate vaccine programme in Malawi, a modelling study. BMC Med. 22, 419 (2024).
Heinsbroek, E. et al. Persisting high prevalence of pneumococcal carriage among HIV-infected adults receiving antiretroviral therapy in Malawi: a cohort study. Aids 29, 1837–1844 (2015).
Garcia Garrido, H. M. et al. Incidence and risk factors for invasive pneumococcal disease and community-acquired pneumonia in human immunodeficiency virus-infected individuals in a high-income setting. Clin. Infect. Dis. 71, 41–50 (2020).
Sadlier, C., O’Connell, S., Kelleher, M. & Bergin, C. Incidence and risk factors for invasive pneumococcal disease in HIV-positive individuals in the era of highly active antiretroviral therapy. Int. J. STD AIDS 30, 472–478 (2019).
Thornhill, J., Sivaramakrishnan, A. & Orkin, C. Pneumococcal vaccination in people living with HIV. Vaccine 33, 3159–3160 (2015).
Owachi, D. et al. Mortality and associated factors among people living with HIV admitted at a tertiary-care hospital in Uganda: a cross-sectional study. BMC Infect. Dis. 24, 239 (2024).
Simell, B. et al. The fundamental link between pneumococcal carriage and disease. Expert Rev. Vaccines 11, 841–855 (2012).
Bishop, J. D., DeShields, S., Cunningham, T. & Troy, S. B. CD4 count recovery after initiation of antiretroviral therapy in patients infected with human immunodeficiency virus. Am. J. Med. Sci. 352, 239–244 (2016).
O’Connor, J. et al. Effect of immediate initiation of antiretroviral therapy on risk of severe bacterial infections in HIV-positive people with CD4 cell counts of more than 500 cells per μL: secondary outcome results from a randomised controlled trial. Lancet HIV 4, e105–e112 (2017).
Hileman, C. O. & Funderburg, N. T. Inflammation, immune activation, and antiretroviral therapy in HIV. Curr. HIV/AIDS Rep. 14, 93–100 (2017).
Zhou, X. et al. Mucosal immune response in biology, disease prevention and treatment. Signal Transduct. Target. Ther. 10, 7 (2025).
Gallo, O., Locatello, L. G., Mazzoni, A., Novelli, L. & Annunziato, F. The central role of the nasal microenvironment in the transmission, modulation, and clinical progression of SARS-CoV-2 infection. Mucosal Immunol. 14, 305–316 (2021).
Tacchi, L. et al. Nasal immunity is an ancient arm of the mucosal immune system of vertebrates. Nat. Commun. 5, 5205 (2014).
Nyazika, T. K. et al. Influenza-like illness is associated with high pneumococcal carriage density in Malawian children. J. Infect. 81, 549–556 (2020).
Jochems, S. P. et al. Inflammation induced by influenza virus impairs human innate immune control of pneumococcus. Nat. Immunol. 19, 1299–1308 (2018).
Mitsi, E. et al. RSV and rhinovirus increase pneumococcal carriage acquisition and density, whereas nasal inflammation is associated with bacterial shedding. Cell Host Microbe 32, 1608–20.e4 (2024).
Gonzalez, R. J. et al. Constitutive immune surveillance of the nasal mucosa by neutrophils. J. Immunol. 206, 111 (2021). 04-.04.
Hurst, J. H. et al. Age-associated differences in mucosal and systemic host responses to SARS-CoV-2 infection. Nat. Commun. 16, 2383 (2025).
Lu, Y. J. et al. Interleukin-17A mediates acquired immunity to pneumococcal colonization. PLoS Pathog. 4, e1000159 (2008).
Nikolaou, E. et al. Experimental human challenge defines distinct pneumococcal kinetic profiles and mucosal responses between colonized and non-colonized adults. mBio 12, https://doi.org/10.1128/mbio.02020-20 (2021).
Deng, Y., Herbert, J. A., Smith, C. M. & Smyth, R. L. An in vitro transepithelial migration assay to evaluate the role of neutrophils in respiratory syncytial virus (RSV) induced epithelial damage. Sci. Rep. 8, 6777 (2018).
Deng, Y. et al. Neutrophil-airway epithelial interactions result in increased epithelial damage and viral clearance during respiratory syncytial virus infection. J. Virol. 94, e02161 (2020).
Hewitt, R. J. & Lloyd, C. M. Regulation of immune responses by the airway epithelial cell landscape. Nat. Rev. Immunol. 21, 347–362 (2021).
Cagle, L. A. et al. Early mechanisms of neutrophil activation and transmigration in acute lung injury. Front. Physiol. 13, 1059686 (2022).
Kwok, A. J. et al. Neutrophils and emergency granulopoiesis drive immune suppression and an extreme response endotype during sepsis. Nat. Immunol. 24, 767–779 (2023).
Pak, V., Budikhina, A., Pashenkov, M. & Pinegin, B. Neutrophil activity in chronic granulomatous disease. In Proc. Immune-Mediated Diseases: From Theory to Therapy 69–74 (Springer, 2007).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–87.e29 (2021).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Nyirenda, J. et al. Spatially resolved single-cell atlas unveils a distinct cellular signature of fatal lung COVID-19 in a Malawian population. Nat. Med. 30, 3765–3777 (2024).
Robin, B. et al. MultiNicheNet: a flexible framework for differential cell-cell communication analysis from multi-sample multi-condition single-cell transcriptomics data. Preprint at bioRxiv https://doi.org/10.1101/2023.06.13.544751 (2023).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Russo, P. S. T. et al. CEMiTool: a bioconductor package for performing comprehensive modular co-expression analyses. BMC Bioinform. 19, 56 (2018).
Afran, L. et al. Defective monocyte enzymatic function and an inhibitory immune phenotype in human immunodeficiency virus-exposed uninfected African infants in the era of antiretroviral therapy. J. Infect. Dis. 226, 1243–1255 (2022).
Jambo, K. C. et al. Asymptomatic HIV-infected individuals on antiretroviral therapy exhibit impaired lung CD4(+) T-cell responses to mycobacteria. Am. J. Respir. Crit. Care Med. 190, 938–947 (2014).
Jambo, K. C. et al. Small alveolar macrophages are infected preferentially by HIV and exhibit impaired phagocytic function. Mucosal Immunol. 7, 1116–1126 (2014).
Mu, W., Patankar, V., Kitchen, S. & Zhen, A. Examining chronic inflammation, immune metabolism, and T cell dysfunction in HIV infection. Viruses 16, 219 (2024).
Sikora, E. Activation-induced and damage-induced cell death in aging human T cells. Mech. Ageing Dev. 151, 85–92 (2015).
Lagnado, A. et al. Neutrophils induce paracrine telomere dysfunction and senescence in ROS-dependent manner. EMBO J. 40, e106048 (2021).
Hensley-McBain, T. & Klatt, N. R. The dual role of neutrophils in HIV infection. Curr. HIV/AIDS Rep. 15, 1–10 (2018).
Raghuwanshi, S. K. et al. The chemokine receptors CXCR1 and CXCR2 couple to distinct G protein-coupled receptor kinases to mediate and regulate leukocyte functions. J. Immunol. 189, 2824–2832 (2012).
Margraf, A., Lowell, C. A. & Zarbock, A. Neutrophils in acute inflammation: current concepts and translational implications. Blood 139, 2130–2144 (2022).
Pellowe, A. S. et al. Endothelial cell-secreted MIF reduces pericyte contractility and enhances neutrophil extravasation. FASEB J. 33, 2171 (2018).
Leemans, J. C. et al. The epidermal growth factor-seven transmembrane (EGF-TM7) receptor CD97 is required for neutrophil migration and host defense. J. Immunol. 172, 1125–1131 (2004).
Lawrence, D. W. et al. Antiadhesive role of apical decay-accelerating factor (CD55) in human neutrophil transmigration across mucosal epithelia. J. Exp. Med. 198, 999–1010 (2003).
Bourke, C. D. et al. Cotrimoxazole reduces systemic inflammation in HIV infection by altering the gut microbiome and immune activation. Sci. Transl. Med. 11, eaav0537 (2019).
Teijeira, A. et al. Differential interleukin-8 thresholds for chemotaxis and netosis in human neutrophils. Eur. J. Immunol. 51, 2274–2280 (2021).
Arokiasamy, S. et al. Endogenous TNFα orchestrates the trafficking of neutrophils into and within lymphatic vessels during acute inflammation. Sci. Rep. 7, 44189 (2017).
Hashizume, M., Higuchi, Y., Uchiyama, Y. & Mihara, M. IL-6 plays an essential role in neutrophilia under inflammation. Cytokine 54, 92–99 (2011).
Oliveira, S. H., Canetti, C., Ribeiro, R. A. & Cunha, F. Q. Neutrophil migration induced by IL-1beta depends upon LTB4 released by macrophages and upon TNF-alpha and IL-1beta released by mast cells. Inflammation 31, 36–46 (2008).
Heeb, L. E. M., Egholm, C. & Boyman, O. Evolution and function of interleukin-4 receptor signaling in adaptive immunity and neutrophils. Genes Immun. 21, 143–149 (2020).
Meziane, O. et al. HIV infection and persistence in pulmonary mucosal double negative T cells in vivo. J. Virol. 94, 10–1128 (2020).
Costiniuk, C. T. et al. HIV persistence in mucosal CD4+ T cells within the lungs of adults receiving long-term suppressive antiretroviral therapy. Aids 32, 2279–2289 (2018).
Pollara, J. et al. Single-cell analysis of immune cell transcriptome during HIV-1 infection and therapy. BMC Immunol. 23, 48 (2022).
Li, X. et al. Inflammation and aging: signaling pathways and intervention therapies. Signal Transduct. Target. Ther. 8, 239 (2023).
Van Avondt, K. et al. Neutrophils in aging and aging-related pathologies. Immunol. Rev. 314, 357–375 (2023).
Yue, Z. et al. Senescence-associated secretory phenotype and its impact on oral immune homeostasis. Front. Immunol. 13, 1019313 (2022).
Peretz, Y. et al. CD160 and PD-1 co-expression on HIV-specific CD8 T cells defines a subset with advanced dysfunction. PLoS Pathogens 8, e1002840 (2012).
Reuter, M. A. et al. HIV-specific CD8+ T cells exhibit reduced and differentially regulated cytolytic activity in lymphoid tissue. Cell Rep. 21, 3458–3470 (2017).
Jambo, K. C. et al. B cell, CD8+ T cell and gamma delta T cell infiltration alters alveolar immune cell homeostasis in HIV-infected Malawian adults. Wellcome Open Res. 2, 105 (2018).
Peno, C. et al. Alveolar T-helper 17 responses to Streptococcus pneumoniae are preserved in ART-untreated and treated HIV-infected Malawian adults. J. Infect. 76, 168–176 (2018).
Siegel, S. J., Roche, A. M. & Weiser, J. N. Influenza promotes pneumococcal growth during coinfection by providing host sialylated substrates as a nutrient source. Cell Host Microbe 16, 55–67 (2014).
Siegel, S. J. & Weiser, J. N. Mechanisms of bacterial colonization of the respiratory tract. Annu. Rev. Microbiol. 69, 425–444 (2015).
Trappetti, C. et al. Sialic acid: a preventable signal for pneumococcal biofilm formation, colonization, and invasion of the host. J. Infect. Dis. 199, 1497–1505 (2009).
Marion, C., Burnaugh, A. M., Woodiga, S. A. & King, S. J. Sialic acid transport contributes to pneumococcal colonization. Infect. Immun. 79, 1262–1269 (2011).
Weight, C. M. et al. Microinvasion by Streptococcus pneumoniae induces epithelial innate immunity during colonisation at the human mucosal surface. Nat. Commun. 10, 3060 (2019).
Minhas, V. et al. In vivo dual RNA-seq reveals that neutrophil recruitment underlies differential tissue tropism of Streptococcus pneumoniae. Commun. Biol. 3, 293 (2020).
Satzke, C. et al. Standard method for detecting upper respiratory carriage of Streptococcus pneumoniae: updated recommendations from the World Health Organization pneumococcal carriage working group. Vaccine 32, 165–179 (2013).
Fadlyana, E. et al. Risk factors associated with nasopharyngeal carriage and density of Streptococcus Moraxella catarrhalis, and Staphylococcus aureus in young children living in Indonesia. Pneumonia 10, 14 (2018).
Acknowledgements
This work was supported by an African Research Leader (ARL) award (MR/T008822/1) to KCJ. This ARL award is jointly funded by the UK Medical Research Council (MRC) and the UK Foreign, Commonwealth and Development Office (FCDO) under the MRC/FCDO Concordat agreement and is also part of the EDCTP2 programme supported by the European Union. We are grateful for the support of the funders and the Glasgow Polyomics sequencing platform for sequencing and demultiplexing the single-cell data. Special thanks to Olympia Hardy and David McGuiness.
Author information
Authors and Affiliations
Contributions
Conceptualisation: J.A.P. and K.C.J.; data curation: J.A.P.; data analysis: J.A.P., K.C.J.; funding acquisition: K.C.J; investigation: N.M., A.K.; methodology: J.A.P., A.K., E.L., R.K.N., J.Z., T.K., L.M., J.T.N. and C.N.; project administration: N.M., A.K.; resources: C.M., H.C.M.; supervision: K.C.J., B.K. and D.F.; validation: J.A.P.; writing–original draft: J.A.P., K.C.J.; writing–review and editing; J.A.P., L.L.S., G.K., L.M., J.T.N., N.M., A.K., C.N., P.C., A.P.C., K.D., T.K., R.N., J.Z., E.L., M.N.M., D.V.M., L.M., S.B.G., B.K., C.M., D.F., H.C.M. and K.C.J.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Andrew Harman and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Phiri, J.A., Sibale, L.L., Kapira, G. et al. Persistent pneumococcal colonisation in antiretroviral-treated HIV infection is associated with nasal inflammation. Nat Commun 17, 565 (2026). https://doi.org/10.1038/s41467-025-67258-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67258-7