Abstract
Archaeal membrane lipids are markedly distinct from those in bacteria and eukaryotes, serving as biomarkers for unraveling their ecological and biogeochemical roles. Recent advancements in high-resolution mass spectrometry-based lipidomic research facilitate detailed cellular-level characterizations of lipid compounds. However, the lack of a comprehensive and dedicated database severely limits large-scale, high-throughput investigations of archaeal lipids. We present ArchLips, a comprehensive database containing 219,348 in silico molecular structures and tandem mass spectra of 199,248 corresponding archaeal lipid compounds. ArchLips enables the automatic and accurate annotation of archaeal lipid compounds characterized by high-resolution mass spectrometry from both pure cultures and environmental samples, serving as a transformative tool for enhancing our understanding of archaeal diversity and its ecological and evolutionary significance within global ecosystems.
Similar content being viewed by others
Introduction
Archaea are widely recognized as a distinct branch on the tree of life alongside bacteria and eukaryotes and among the earliest microbial forms on Earth1,2,3,4,5. Advances in culture-independent high-throughput sequencing and bioinformatic approaches have catalyzed the discovery of a large number of metabolically versatile archaeal species, driving burgeoning research into the diversity, ecological functions, roles in biogeochemical processes, evolution and biotechnological applications of these organisms. To date, more than 30 archaeal phyla grouped into 4 superphyla (Euryarchaeota, DPANN, TACK, and Asgardarchaeota) have been identified across a wide range of environments, from extreme habitats to diverse mesophilic ecological niches, host-associated microbiomes, and mammalian guts6,7,8,9,10,11,12,13. Notably, the recent discovery of the Asgard archaeal superphylum has revealed a close phylogenomic relationship with Eukarya, reigniting debates on the origins of eukaryotes and underlining the crucial role of archaea in early evolution of life7,14,15,16,17. These milestone findings have prompted intensified research efforts to further elucidate the biological and ecological impacts of these enigmatic microorganisms.
Membrane lipids are the fundamental components in cellular organisms, performing essential functions in maintaining cell morphology and integrity, regulating osmotic permeability for substance transport, and as signaling or energy storage molecules. One of the most intrinsic differences between archaea and bacteria/eukaryotes is the structure of their cell membrane lipids. The substantial differences in the stereochemistry of the glycerol backbone (glycerol-1-phosphate in archaea vs. glycerol-3-phosphate in bacteria/eukaryotes), the hydrocarbon chains (isoprenoids in archaea vs. fatty acyls in bacteria and eukaryotes), and the nature of the ether/ester bonds linking these components have raised “lipid divide” questions regarding the evolutionary origins and divergence of microbial lipid biosynthesis pathways18,19,20.
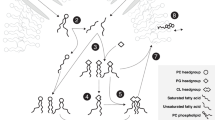
The cell membrane of archaea contains monolayer or bilayer lipids consisting of isoprenoid glycerol dialkyl glycerol tetraethers (e.g., iGDGTs) and/or glycerol dialkyl diethers (e.g., archaeol) as core lipids, which can be modified by methylation, cyclization, cross-linking (H-shaped), hydroxylation and unsaturation to increase the structural diversity (Fig. S1). The second level of structural diversity in archaeal lipids arises from the head groups attached to the core lipid structures. These head groups often contain various sugar moieties (glycolipids) and/or phosphate group (glycerophospholipids), which can be further modified with polar molecules such as choline, ethanolamine, serine, forming amphipathic intact polar membrane lipids in archaeal cells (IPLs, Fig. S1). In addition, archaeal cardiolipin analogues have been identified and function as fundamental membrane components in some lineages of archaea21,22. Other lipids embedded in the membrane like quinone or carotenoid have also been identified and involved in electron transfer in some archaea23.
These distinct lipids not only help archaea survive in diverse environmental conditions but also serve as molecular “fingerprints” in sediments and rocks, offering clues about paleoclimates and ecosystems. Examining archaeal lipids in pure cultures and diverse environmental settings has led to applying multiple lipids-based geochemical or paleoenvironmental proxies such as the TEX86 (TetraEther indeX of tetraethers consisting of 86 carbon atoms) proxy for paleotemperature and the MI (Methane Index) proxy for ancient marine hydrate identification24,25. Increasing evidence also shows that archaeal lipids can be used as biomarkers to reflect paleo ocean redox changes and nutritional status, which are more related to the physiology and biochemistry of archaea26,27,28. The structural and compositional differences of lipid components (including intact polar lipids) among various archaeal species (or phyla) allow them to be widely utilized as chemotaxonomic markers21,29,30,31. Archaeal lipids have also been widely used for tracing archaeal community dynamics and biogeochemical activity across diverse environments32,33,34,35. Thus, great promises lie ahead with more archaeal lipids being used in paleo-climate as well as paleo-ecosystem studies36,37.
The analysis of archaeal lipids has been transformed by direct infusion mass spectrometry (shotgun lipidomics) or high-resolution liquid chromatography-mass spectrometry (LC-MS)38,39, which has established lipidomics as a powerful tool in biogeochemistry. Unlike traditional targeted methodologies that quantify a narrow range of characterized lipids40,41,42,43,44, untargeted MS approaches can simultaneously detect thousands of structurally diverse compounds. This capability generates large-scale lipidomic datasets that require high-throughput processing. A major challenge within this workflow is lipid identification, which remains a significant bottleneck. It often necessitates manual interpretation by matching experimental data, such as accurate mass and tandem mass (MS2) spectra to existing spectral libraries45, a process that limits the speed and scope of research.
Due to the limited availability of authentic lipid standards, especially for archaeal lipids, in silico lipid databases have become widely used and show great promise for compound annotation in untargeted MS-based lipidomics46,47. Algorithms such as rule-based modelling or machine learning approaches have been employed to predict or generate in silico mass spectra48,49. Incorporating these spectral libraries into extant cheminformatic tools enabled the development of automated and effective methods for MS data interpretation, and expanded coverage of lipid species45,50,51,52,53 that would otherwise be impossible for manual annotation.
In this work, we present ArchLips, a comprehensive in silico database for archaeal lipids, designed for high-resolution mass spectrometry-based fingerprinting. It covers a wide structural diversity of archaeal lipids, with a full library of 199,248 lipids including in silico MS2 spectra, and enables high-throughput compound annotation through integration with bioinformatic tools (Fig. S2). This database serves as a resource for lipid identification in pure cultures and environmental samples, supporting the discovery of biomarkers and providing insights into the structural diversity of archaeal lipids.
Results
Construction of the general structure library
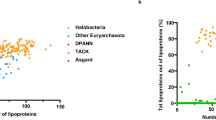
This study began with the compilation of a structure database of archaeal lipids from all available sources (Table 1), including public databases such as LipidMAPS (https://lipidmaps.org/), PubChem (https://pubchem.ncbi.nlm.nih.gov/), and LipidBank (https://lipidbank.jp/). Significant effort was also devoted to collecting publicly available lipid structures through meticulous literature research (e.g., refs. 21,54,55,56,57,58,59,60,61,62,63,64,65,66). We then constructed the structure library that constitutes all currently known archaeal lipids, including their isomers, homologs, and derivatives (Table 2 and Fig. S1). This resulted in a total of 219,348 in silico molecules in distinct structures generated using MarvinSketch, ChemAxon Reactor software, and the RDKit package in Python, which were stored in SDF (MDL molfile) format (Fig. 1B).
A The seven steps involved in the construction of in silico structure library. Steps 1-4 were to construct the structure library using a combinatorial enumeration method based on structures reported in the literature. Steps 5–7 were to construct the in silico MS2 spectra based on the fragmentation pattern and the ion intensity of archaeal lipids. B The four types of in silico libraries constituting the ArchLips database: an structure library, a full spectral library, a filtered high-confidence library and a library containing the M + 1 spectra.
The structure library included basic chemical information such as molecular formula, exact mass, SMILES, Inchi, InchiKey, and logP. The shorthand nomenclature for archaeal diether lipids followed the MS-DIAL lipidomics database (LipidBlast), which adheres to the LipidMAPs convention at the molecular species level67,68. Under this nomenclature system, the isoprenoid chain composition is specified, though the sn-positions are not necessarily resolved, for example, PG-Archaeol (20:0_20:0) and 2Gly-Archaeol (20:0_20:0). For consistency, traditional names were retained for archaeal cardiolipins, but with their isoprenoidal chain lengths explicitly defined, as in TGD-PA (20:0_20:0_20:0_20:0) and S-DGD-PA (20:0_20:0_20:0_20:0). For tetraether lipids, traditional names were preserved, while regioisomer compounds were labelled using suffixes such as “a,” “b,” and “c” (e.g., GDGT-4a, GDGT-4b, and GDGT-4c to denote isomers of GDGTs with four cyclopentane rings in different configurations)56,57.
We defined structural boundaries for archaeal core lipids to avoid a combinatorial explosion of generated structures46. For instance, the length of each isoprenoidal carbon chain in diether lipids (e.g., archaeol) was limited to C20-C30, with a maximum of two additional methyl or hydroxyl groups occurring at the C3 position of the carbon chains. The library also included macrocyclic archaeol containing 0-4 rings and unsaturated archaeol with up to 12 double bonds. However, stereoisomers and regioisomers, such as variations in double bond position or sn2/sn3 positions of the isoprenoid carbon chains, were currently not considered in this database.
Archaeal tetraether lipids (e.g., GDGTs) containing a maximum of 8 cyclopentane moieties served as the primary structures for further generation of tetraether lipids of possible structures constrained by the boundary condition mentioned above. The resulting diverse derivatives were refined to include methylated GDGTs (Me-GDGTs; 1-3 additional methyl groups) and hydroxylated GDGTs (OH-GDGTs; 1-2 hydroxyl groups), GDGTs with cyclohexane rings (S-GDGTs; 1-2 rings), unsaturated GDGTs (1-8 double bonds), and H-shaped configurations (GMGTs). Selected regioisomers of GDGTs, defined under specific chromatographic conditions, were generated to represent variations in the cyclopentyl ring configuration between the etherified chains56,69. Other stereoisomers, such as tetraether lipids with parallel glycerol backbones were excluded from the database as they lack definitive chromatographic conditions to directly separate them70.
The MDL molfile structures for archaeal intact polar lipids (IPLs) were generated by arranging a library of 79 known head groups (Fig. S1, Table S2) to the core lipid structures described above. A consideration for archaeal tetraether lipids is that their structure contains two terminal glycerol units. Consequently, for specific head group combinations54,71,72, we generated IPLs with head groups attached to both ends. An example of such a doubly-attached structure is Gly-(PGly)-GDGT (Glycosyl-GDGT-phosphatidylglycosyl) (Table S2).
Construction of the ArchLips spectral library
We used LipidBlast as a foundational template for developing an in silico spectral library of archaeal lipids because it is easy to operate and formulate structures46,47. Building upon the template we integrated all structural information into a dedicated Excel file to improve data organization, optimize MS2 calculations, and streamline library exportation. Specifically, the MS2 data were organized into multiple Excel sheets, with each sheet corresponding to a lipid class with different core lipids that allow optimized MS2 calculations; the library exportation program was rewritten in Python, employing multiprocessing to efficiently produce spectral library files. The flexible LipidBlast template and streamlined workflow allow the in silico spectral library to be readily updated and expanded to include recently identified lipid classes.
The customized ArchLips spectral library comprised three core components: (1) basic chemical information, e.g., exact mass and a defined formula, (2) diagnostic fragment ion m/z values, and (3) the relative signal intensities of these fragments.
The basic chemical properties in the spectral library were theoretically calculated from the structural input. The m/z values of precursor ions were constructed by incorporating the exact mass and two primary adducts, [M + H]⁺ and [M + NH4]+, which are frequently observed in positive ionization mode (a prevailing methodology for detecting archaeal lipids). The fragment m/z and relative intensity were systematically generated through the following methodology:
Firstly, the m/z of the fragment ion and the relative intensity were intrinsically determined by the stability of chemical bonds for a specific class of lipids and experimentally by the collision energy level used during collision-induced dissociation (CID). We examined the optimal collision energy using 18 archaeal lipids under different levels (Figs. S3–S4). Based on the positive correlation between optimal collision energy and precursor m/z (Fig. S3B), we employed a ramped collision energy in this study: low-mass ions (start at m/z 100) were fragmented at 10–15 eV, while high-mass ions (end at m/z 2000) were fragmented at elevated energies of 55–60 eV (Fig. S3B). The optimized collision energy for the molecular weight-dependent fragmentation characteristics of archaeal lipids enhanced the generation of diagnostic fragment ions and acquisition of high-quality MS2 spectra for spectral library construction.
Secondly, the fragmentation pattern of archaeal lipids was delineated into three distinct regions (Regions I, II, and III) as summarized in ref. 58 (Fig. 2A). Region I was mainly associated with the neutral loss of small molecular moieties occurring for almost all archaeal lipids and the sequential loss of polar headgroups in intact polar lipids. These losses of small molecular moieties typically include the sequential loss of H2O, C3H4O, and C3H6O2, resulting in product ions of e.g., [M + H-18]+, [M + H-56]+, and [M + H-74]+, respectively (Fig. 2A–D).
A MS2 spectrum of Calditiol-GDGT-4a with the fragmentation pattern highlighted with three regions (I, II, and III). Region I refers to the product ions from the neutral loss of small molecules, Region II contains the characteristic ions of archaeal lipids that concern the neutral loss of one diphytanyl chain and subsequent loss of small molecules, and Region III involves the product ions from the further cleavage of the biphytanyl or phytanyl chains. B GTGT-0, an intermediate compound during the biosynthesis of archaeal tetraether lipids. C PG-Archaeol (20:0_25:0), an intact polar lipid with a phosphoglycerol headgroup. D DGD-PA (20:0_20:0_20:0_20:0), an archaeal cardiolipin analogue. The experimental spectrum is shown at the top (black) and the in silico spectrum at the bottom (red).
Region II covered the diagnostic fragments commonly used for the identification and structural elucidation of archaeal lipids. These product ions were mainly derived from cleaving the ether bonds (C-O) of the glycerol tetraether structures. The ether bond cleavage resulted in the neutral loss of diphytanyl or phytanyl chains, along with the sequential loss of H2O, C3H4O, and C3H6O2 as observed in Region I. Moreover, this process usually involves transferring one or two protons from the neutral loss molecules to the charged product ions54.
Region III (m/z < 300) contained fragment ions cleaved from the isoprenoid chains and headgroups, which would provide additional information for structural elucidation of these components (Fig. S5). The relative abundance of ions in this region varied across different classes of archaeal lipids, being prominent in the core lipids of both diether and tetraether types but less frequently observed in lipids such as PS-Archaeol (20:0(OH)_20:0).
After obtaining the fragmentation patterns and relative ion intensities for a representative lipid, these data were used to reconstruct the MS2 spectra for its lipid class. The spectral library was generated based on a modified LipidBlast template for Regions I and II across all lipids, whereas Region III was incorporated for selected lipids to improve identification scores. The relative abundance of the fragment ions was heuristically reconstructed in a modeling effort that integrated known structural features of archaeal lipids, including core structures, side-chain moieties, and cleavage patterns (Fig. 1A).
The above comprehensive workflow culminated in the ArchLips spectral library, which encompassed MS2 spectra for 199,248 archaeal lipids (Table 2), providing a robust resource for systematic identification and structural characterization of these molecules. To improve reliability of the spectral library, we compiled a high-confidence sub-dataset of 29,446 archaeal lipids by curating compounds reported in the literature, their derivatives, and lipids manually verified through rigorous laboratory identification. These entries were supplemented with in silico MS2 spectra to support robust spectral matching (Fig. 1B and Table 2).
Additionally, we addressed challenges posed by automated peak-picking algorithms, which can misassign monoisotopic peaks due to interference from high molecular weight compounds or elevated [M + 1] isotopologue signals (Fig. 3C, D). In doing so, we developed a sub-spectral library comprising 123,686 archaeal lipids, which improved the recognition of compounds with [M + 1]/M ratios exceeding 1 (Fig. S6). The spectral libraries were further strengthened by retention time prediction based on a model built from manually curated dataset for our chromatographic condition (Fig. 1A). The root mean square error (RMSE) for the training dataset was 0.42 min, with a 95% confidence band of ±0.7 min, while the RMSE for the testing dataset was 1.29 min, and a 95% confidence band of ±2.11 min. These complementary strategies collectively refine the accuracy of lipid analysis by resolving peak assignment ambiguities in automated workflows.
A The library search of 148 MS2 spectra of archaeal lipids from the peer-reviewed literature against the high-confidence ArchLips spectral library. B Distribution of archaeal in-source fragmentation (ISF) features among different archaeal lipid classes. ISF-derived features were manually identified from quality control (QC) samples of pure cultures. C Mass spectrum of S-DGD-PA (20:0_20:0_20:0_20:0) showing the dominant M + 1 isotopologue (red peak) arises from the natural abundance of ¹³C in archaeal lipids. D Extracted ion chromatogram showing the distribution of isotopologues in S-DGD-PA (20:0_20:0_20:0_20:0). E Extracted ion chromatogram of 3Gly-GDGT-0 (m/z = 1805.5062, [M + NH4]+) detected in the M. maripaludis mutant strain and QC samples. F Spectral matching of experimental MS2 spectra of 3Gly-GDGT-0 (black, top) against referenced in silico spectra from the ArchLips database.
Library validation using pure culture strains
To validate the accuracy of the ArchLips database, a decoy search of 148 MS2 spectra of archaeal lipids obtained from the peer-reviewed literature was performed against the high-confidence spectral library as described in ref. 46. Over 75.0% of the MS2 spectra were correctly matched, 15.5% were annotated as isomer compounds and 9.5% as false positive or no match (Fig. 3A). The decoy searches of the high-confidence spectral library and full spectral library against the non-archaeal-dominated public lipid spectral databases resulted in a relatively low hit rate for the GNPS (0.8%, 6.6%) and LipidBlast library (0.2%, 0.6%), respectively (Table S3).
Next, the performance of the ArchLips database was evaluated using four pure archaea cultures that have existing lipid profiles22,29,69,73. The example of MS2 spectra match between the experimental spectra and the in silico spectral library was shown in Figures S7–S10 (including MS2 spectra detected in environmental samples). When applying the high-confidence spectral library using retention time (RT) for filtering, 235 lipid features were successfully annotated with an identification score > 70%. With an RT tolerance of 3 min for filtering, 209 mass features were annotated. Additionally, when using the high-confidence spectral library containing M + 1 spectra, both with and without a RT tolerance of 3 min for filtering, identified 259 and 236 lipid features, respectively.
The lipid annotation results obtained using the high-confidence spectral library without incorporating retention time for compound filtering, were further subjected to manual verification and statistical analysis. In total, 292 lipid features were annotated, corresponding to 139 lipid species across four archaeal pure culture strains, including diethers, tetraethers, cardiolipins, GDDs, and quinones (Fig. 4A). Among them, only 3 diether lipids were shared by all four archaeal strains and 45 lipids were shared by subsets of these strains (Fig. 4A), which accounted for 34.5% of total lipid species. The remaining majority (65.5%) of the identified lipids were strain-specific. For example, the lipidome of N. maritimus and S. acidocaldarius were both dominated by tetraether lipids (Fig. 4B), yet they were still distinguishable using the overall lipidomic profiles by PCA analysis (Fig. 4C). Furthermore, some exclusive biomarkers such as methoxy archaeol (MeO-AR, m/z = 667.6962, [M + H]+) for Nitrososphaeria29 were identified in N. maritimus (Fig. 5A). In contrast, the lipidomes of M. maripaludis and H. larsenii were primarily composed of diether lipids, which were also discriminable between these two strains (Fig. 4B, C). For M. maripaludis mutant strain heterologously expressing Tes homologs from M. acetivorans C2A, a trace amount of GDGT-0 (m/z = 1302.3197, [M + H]+)74 was identified along with an intact polar lipid of 3Gly-GDGT-0 (m/z = 1805.5062, [M + NH4]+) (Fig. 3E, F).
A The intersection of 139 lipid compounds identified from different archaeal species is illustrated using the UpSet plot. B The relative abundance of five major lipid categories in four archaeal strains. C The PCA plot is based on the relative abundance of all annotated lipids from the four pure cultures. Note: the relative abundances reported do not reflect the true relative abundances of lipids in the samples due to the ionization efficiencies of different lipid classes could not be determined. Only lipids detected in all replicate samples were retained for the UpSet plot.
A Diether lipids; (B) tetraether lipids lipid features involved in the FBMN. The filled color of the nodes indicated the distribution of lipid features in four archaeal strains. The color of the inner rings represents annotated features (red), and the color of the outer rings refers to the adduct types. The width of the line connecting the nodes represents the level of mass spectral similarities, with thicker lines indicating greater similarity between two nodes. C The bar chart showing the features with MS2 and associated in FBMN and annotated features retained in FBMN.
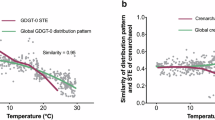
A feature-based molecular network (FBMN) was constructed to characterize the spectral similarities between identified lipids and unknown features75,76. The results revealed the FBMN contained a total of 770 nodes, which formed 97 clusters with at least two connected nodes (Fig. S11). These features in FBMN accounted for 28.9% of features with MS2, and 200 of them were annotated using the ArchLips database. Figure 5A, B showed two clusters of archaeal diether and tetraether lipid-related features, respectively. The cluster of tetraether lipids in Fig. 5B is primarily associated with calditol-GDGTs, which are suggested to be produced by thermoacidophilic archaea in the Crenarchaeota phylum (e.g., S. acidocaldarius) with a putative calditol synthase (cds)73. Besides, several nodes were likely lipid features corresponding to [M + 1] isotopologue peaks that generated chimeric mass spectra and exhibited similar fragmentation patterns to their corresponding monoisotopic species, such as those observed at m/z 831.73 in Fig. 5A and m/z 1457.3 in Fig. 5B.
Archaeal lipidome applied to diverse natural habitats
To identify archaeal lipids in environmental samples using the ArchLips database, 52 environmental samples were collected from diverse geological settings and habitat types such as surface and subsurface marine sediments, cold seep sediments, hot spring sediments, acid mine drainage (AMD), aerated soils and permafrost soils. A range of 415–616 archaeal ether lipid features (assigned to 176–270 lipid compounds) and 46-non-ether lipid features (assigned to 18 to 24 lipid compounds) were annotated using the untargeted spectral libraries of ArchLips database (Figs. S12, S13). These numbers were obviously higher than those obtained from conventional targeted approaches that focused on only a few compounds in environmental samples. Comparison among these samples showed first-order differences in lipid distribution pattern between marine and terrestrial environments (Fig. 6A).
A Relative abundance of archaeal lipid classes, (B) Clustering analysis using Euclidean distance and Ward.D method, (C) PCA analysis and (D) Shannon diversity of archaeal lipids. AMD, acid mine drainage (n = 10): AHTL and FK represent samples collected from Tongling and Fankou AMD, respectively. Hot springs (n = 6): hot spring sediment samples collected from Tengchong (GMQ and DRTY). Marine sediment (n = 20): cold seep sediment collected from the South China Sea (ROV01), a sediment core from Pearl River estuaries (GSD) and surface sediment samples from the East China Sea (ESC). Soils (n = 16): soil samples collected from the western region of Sichuan Province (CX) and a sample from Wuhan (YJS), and samples from Qinghai-Tibetan plateau permafrost (NMCVI and LR). The horizontal black lines indicate medians, the box edges represent the interquartile range (25–75th percentiles), and whiskers show the range within 1.5 times the interquartile range. p-values were calculated using two-sided Wilcoxon rank-sum tests. The “*” indicates the significance level of p < 0.05,“**” indicates the significance level of p < 0.01, and “***” indicates the significance level of p < 0.001. Note: the relative abundances reported do not reflect the true relative abundances of lipids in the samples due to the ionization efficiencies of different lipid classes could not be determined.
Clustering analysis based on the relative abundance of archaeal lipids further depicted the habitat types influencing the marine-terrestrial lipid distribution (Fig. 6B and Fig. S14). Among marine samples, cold seep (ROV01) samples formed a separate branch from surface and subsurface sediments of typical marine environments. In the terrestrial group, samples from extreme environments, such as hot spring sediments and acid mine drainage (AMD), clustered together. Notably, lipid composition varied significantly between acidic and alkaline hot spring samples, with alkaline hot springs clustering with AMD samples. This pattern aligns with differences observed in 16S rRNA-based archaeal community composition between acidic and alkaline hot springs (Fig. S15). We also noticed that the soil samples formed separate branches, further dividing into subgroups of aerated soils (CX and NMCVI) and wet permafrost soils (LR), mirroring soil water content or oxygen availability-driven archaeal adaptations in soils77.
The archaeal lipid composition was also evaluated using principal component analysis (PCA) (Fig. 6C). The first three principal components explained 53.8% of the variance in archaeal lipids, with samples from different biomes primarily separating, consistent with the hierarchical clustering results (Fig. 6B). We further assessed the Shannon-Weiner diversity of archaeal lipidomes (Fig. 6D). The result showed that hot spring samples exhibited the highest lipid diversity, followed by AMD samples. At the same time, soil and marine sediment had the lowest diversity. This pattern does not align with archaeal community diversity inferred from 16S rRNA gene analysis, where AMD samples and marine sediments showed the highest diversity and soils had the lowest (Fig. S16).
Discussion
We present a comprehensive database, ArchLips, that is constructed using a combinatorial and heuristic algorithm, which should fill a critical gap in existing lipidomic resources. It was meticulously developed to enhance archaeal lipid identification in four key aspects: (1) comprehensiveness, (2) accuracy, (3) efficiency, and (4) enhanced biological interpretation. These components are detailed below:
Comprehensiveness
Although previous studies have reported diverse structures of archaeal lipids, this information remains fragmented within the literature36,78. Our database curated and integrated these data into a comprehensive repository that now constitutes 219,348 archaeal lipid structures, which includes a complete range of core lipids and a wide range of intact polar lipids (IPLs) (Fig. 1B and Fig. S1). These structures allow us to generate over 380,000 in silico MS2 spectra (two adducts for each lipid) to improve lipid identification. This resource enables precise lipid searches using accurate mass-based approaches commonly applied in shotgun lipidomics and supports structure-driven lipidomics through untargeted MS2 spectral matching against in silico databases. In addition, the flexible template design allows for seamless updates or incorporation of archaeal lipid classes into the database.
Accuracy
ArchLips demonstrated high accuracy in identifying archaeal lipids by matching 148 curated MS² spectra from peer-reviewed literature against the high-confidence spectral library (Fig. 3A). Among 148 MS2 spectra reported in the literature, 90.5% of them were correctly annotated or annotated as isomer compounds using our libraries. Decoy searches further validated the database’s reliability, while low hit rates in LipidBlast and GNPS comparisons (Table S3) underscored its specificity for archaeal lipid structures. Validation using pure archaeal cultures and environmental samples confirmed its applicability across diverse settings (Figs. S7–S10). Additionally, ArchLips enhances identification confidence through computer-generated reproducible and efficient spectral libraries, minimizing reliance on manual interpretation, which is often influenced by analyst expertise.
Efficiency
Compared to targeted approaches, liquid chromatography separation coupled to untargeted mass spectrometry-based lipidomics enabled a more comprehensive detection of a broad range of lipid species72,79. On the other hand, this approach usually generates expansive datasets that pose significant data processing challenges for manual peak integration and subsequent lipid identification. The efficiency of our database is characterized by being capable of batch identification of hundreds of archaeal lipid species from multiple samples (Fig. 4A and Fig. S14) through cheminformatic tools such as MZmine80 and MS-DIAL software39,81. This allows us to streamline data processing pipelines for profiling archaeal lipids (e.g., Fig. S2) and to calculate lipid-based paleoecological and paleoclimate proxies (Fig. S17). Furthermore, the ArchLips database can be integrated with other public lipidomic databases (e.g., LipidBlast), supporting comprehensive, cross-domain characterization of archaeal, bacterial, and eukaryotic lipids (Fig. S14).
A curated subset of the ArchLips database has been utilized or referenced in previous research to facilitate the annotation and characterization of archaeal lipids in diverse biological samples22,33,82,83. We also applied the ArchLips spectral library for high-throughput lipid identification to a lipidomic dataset of four halobacterial strains analyzed on a Q Exactive Orbitrap MS (available for download from MassIVE under accession codes MSV000089830)84,85. The raw data were processed using MS-DIAL software, leading to the successful annotation of 219 lipid features, which enabled a direct comparison of lipid compositions between different strains. These lipids represented a broad spectrum of archaeal lipid classes that included both previously reported84,85 and recently discovered species (Fig. S18), indicating the ArchLips database can be used for lipid identification across different mass spectrometry platforms.
Enhanced biological information
Increasing studies have demonstrated that the clustering tree based on archaeal lipids closely resembles the topology of the 16S rRNA gene tree22,30,31. Utilizing an untargeted lipidomic approach can significantly enhance the biological interpretation of lipid data by capturing the full structural diversity of lipidomes within complex samples. Furthermore, by integrating global lipid annotation with feature-based molecular networking (FBMN), we were able to comprehensively resolve the structural diversity of archaeal lipids between different strains (Fig. 5C), which supports and reinforces previous findings22,29.
While lipidomic approaches are increasingly used to elucidate microbial degradation86 and microbial adaptations to stressors such as temperature or redox gradient in environmental samples76,87,88,89,90,91, the adaptation of archaeal lipidome to those environmental stressors remains poorly understood. The analysis of 52 environmental samples successfully identified 176-270 archaeal lipids, and the overall distribution indicated that distinct compositional patterns of archaeal lipids across habitat types, such as marine vs. terrestrial, extreme vs. mesophilic environments. These lipidomic signatures largely aligned with the archaeal community structures based on 16S rRNA sequencing (Fig. S15). This indicates that the overall distribution patterns of archaeal lipidome are likely primarily mediated by shifts in archaeal community composition. Moreover, discrepancies between lipidomic and phylogenetic profiles indicate that physiological adaptations of archaeal lipids in certain dominant species may also occur in response to environmental changes. At the same time, confounding factors such as DNA and lipid degradation, as well as source differentiation (e.g., terrestrial inputs vs. water-column-derived lipids deposited in sediments), continue to complicate the interpretation of archaeal lipids in environmental samples and therefore require careful evaluation.
While ArchLips represents a significant advancement for high-throughput identification of archaeal lipids, our analysis revealed several key limitations needing careful consideration using spectral matching approaches. The caution for using automated lipid annotation and identification lies in the presence of structural isomers, as well as instrumental factors such as in-source fragmentation and poor chromatographic separation, which challenge unambiguous lipid annotation of these isomer compounds and require manual validation for accurate identification (Fig. S2).
The ArchLips database was designed to incorporate a wide range of isobaric and isomeric lipids of archaea, which reflected the structural diversity of archaeal membrane lipids that have been identified through a variety of lipid identification approaches in the literature. Such isomers include, for example, structural isomers of GDGTs with the positional combinations of a number of cyclopentane (and sometimes cyclohexane) rings and double bonds along the C40 isoprenoid chains56,57,59,60,64. We meticulously curated the in silico MS2 spectra of these lipids and incorporated their characteristic fragment ions to facilitate differentiation. Nevertheless, the annotation of isomeric lipids based solely on spectral matching remains challenging and is not always reliable. For example, GDGTs containing cyclopentane rings may be assigned as unsaturated GDGTs, and vice versa, because they generate identical product ions and thus rank as top hits during spectral matching. Despite this, such compounds can often be distinguished through manual verification that integrates retention time (elution order) and the relative abundances of diagnostic fragment ions64.
Other structural isomers, such as GDGTs with varying cyclopentyl ring configurations between the etherified chains (e.g., GDGT-4a, GDGT-4b, and GDGT-4c), were also included in the database along with their in silico MS2 spectra. Nevertheless, these isomers could not be reliably resolved using reverse-phase chromatography coupled with MS2 analysis in this study, as they frequently coeluted and produced composite fragmentation spectra that complicate automated matching. Currently, such isomers are more appropriately reported at the level of the total number of cyclopentyl rings (e.g., as GDGT-4). It is worth noting that normal-phase chromatography has been shown to separate these compounds56,57,69, therefore, orthogonal or emerging separation techniques, such as two-dimensional chromatography, ion mobility, or supercritical fluid chromatography (SFC), may provide more comprehensive resolution of archaeal lipid diversity in future studies.
Another limitation came from the inherent nature of the electrospray ionization (ESI) technique, which causes in-source fragmentation (ISF) of certain molecules during the ionization process, remaining prevalent in mass spectrometry-based lipid analysis92. The ISF features are derived from the premature fragmentation of labile lipid species, in which functional groups are lost during ionization in the ion source. The ISF features were then chosen to cleave under CID, which generates fragment ions with MS² spectra identical to authentic lipid molecules and increases the likelihood of false-positive identifications.
In this study, we manually attributed 64 annotated lipid features to ISF and 16 features to uncategorized features from four pure archaeal cultures, accounting for 33.8% of the total annotations. These ISF-derived features predominantly occurred in glycolipids, glyco-phospholipids, and cardiolipins (Fig. 3B), compromising the reliability of lipid identification. To mitigate this effect of ISF on lipid identification, ISF-derived features can also be computationally recognized and annotated using bioinformatic tools. For instance, the ISFrag package92, originally developed for LC-MS-based metabolite analyses, can be adapted to identify ISF in archaeal lipidomic searches.
The retention time (RT) provides an orthogonal parameter to improve confidence in lipid identification, particularly by reducing false-positive assignments arising from ISFs and structural isomers. For the ArchLips database, we constructed RT prediction models based on a dataset of 265 lipids identified from both pure cultures and environmental samples. By applying a 3 min RT tolerance for filtering, the accuracy of lipid identification was substantially improved, resulting in the annotation of 209 mass features using the high-confidence spectral library in pure culture samples.
Overall, our archaeal lipidomic results provide a more comprehensive reflection of lipid diversity of archaea than previous methods and demonstrate promise for establishing high-resolution lipid-based ecological indicators and paleoclimate proxies, which may help open up research directions such as lipo-stratigraphy93 or paleo-microbial ecology into deep geological time when molecular DNA becomes less accessible or non-existing.
Methods
Archaeal cultures and sediments
Two wildtype pure cultures and two mutant strains of archaea were examined in this study, which included a Nitrosopumilus maritimus strain SCM194, a Haloferax larsenii JCM 13917, a Methanococcus maripaludis mutant strain (pMEV4-ma_1486) and a mutant strain of Sulfolobus acidocaldarius (p1561-grsB-grsA). N. maritimus is the first pure strain of marine group I Thaumarchaeota (now the class Nitrososphaeria)94, which grows chemolithoautotrophically and obtains energy by performing aerobic ammonia oxidization, thus called ammonia-oxidizing archaea (AOA). The lipidome of this strain has been extensively examined and covers a suite of archaeal tetraether lipids, including the characteristic biomarkers crenarchaeol and its regio-isomer crenarchaeol for AOA28,66,95. H. larsenii and M. maripaludis are two euryarchaeotes belonging to the halophilic and methanogenic archaeon, respectively. The strains of these two euryarchaeotes do not produce tetraether lipids, but diethers as the major lipid content22. In this study, a M. maripaludis mutant was used, which produces GDGT-0 through the expression of Tes homologs74. S. acidocaldarius is a thermoacidophile belonging to Crenarchaeota, which is capable of producing a high diversity of tetraether lipids, including the Calditol-GDGTs and GDGTs up to 8 cyclopentane rings. The Sulfolobus acidocaldarius mutant strain (p1561-grsB-grsA) was used for examining enhanced biosynthesis of GDGTs with more cyclopentyl rings (e.g., GDGT-5 to GDGT-8).
The four archaeal strains were cultivated according to previously published cultivation protocols. Briefly, the N. maritimus strain SCM1 was aerobically cultured in triplicate at 30 °C with 1 L Synthetic Crenarchaeota Medium (SCM)94. The cells were harvested at the stationary phase by filtering through a 0.22 μm PVDF membrane filter. The M. maripaludis S0001 mutant (pMEV4-ma_1486) strain was grown in duplicate in McF medium under a gas stream of N2/CO2 (80:20) at 37 °C74. The S. acidocaldarius mutant (p1561-grsB-grsA) strain was cultured in triplicate at 75 °C in Brock medium supplemented with 0.1% NZ-Amine and 0.2% sucrose69. H. larsenii (JCM 13917) was grown in triplicate in DSMZ 589 liquid medium at 37 °C and pH 7.4 with 200 rpm shaking22. Biomass of the latter three strains at the stationary phase was collected by centrifugation at 10,000 g for 10 min.
This study also examined a total of 52 sediment and soil samples, including those have been used in previous studies as well as newly reported samples, covering a wide range of environmental gradients, including sediment from hot springs in Tengchong, Yunan Province, sediment from acid mine drainages in Anhui and Guangdong provinces96, permafrost soil from Tibet Plateau97, soil from Western Sichuan Plateau, and a sediment core from the South China Sea98, and sediment from the East China Sea99. These samples were collected from 2015 to 2023 and stored at −20 °C in the laboratory until analyses. Detailed sample information is shown in Table S1. The samples were freeze-dried and the lipids were extracted as described below. In addition, to increase the coverage of environmental samples, this study also reanalyzed total lipid extracts from a cold-seep sediment core33 in the same batch with other environmental samples to perform lipid identification with our ArchLips database.
Lipid analysis
Total lipid extracts (TLEs) of all samples were prepared using the Bligh & Dyer method modified by Sturt et al.100. Briefly, wet biomass and freeze-dried environmental samples were extracted twice using a combination of methanol (MeOH): dichloromethane (DCM): phosphate buffer (PB) (2:1:0.8, v/v/v) and subsequently extracted twice with a solvent mixture of MeOH: DCM: trichloroacetic acid buffer (TCA) (2:1:0.8, v/v/v). During each extraction, samples were sonicated for 10 min and then centrifuged at 1690 g for 5 min. The supernatants were combined, and the solvent ratio was adjusted to 1:1:0.9 (v/v/v) by adding an extra amount of DCM and ddH2O. After phase separation was achieved, the DCM layer was collected, and the remaining aqueous layer was re-extracted twice with DCM. Finally, the DCM phase was combined, dried under a gentle nitrogen gas stream, and stored at −80 °C until further analysis.
An aliquot of the TLEs was dissolved in methanol for injection. 10 μL of each sample analyzed in the same batch was pooled as a QC sample for peak alignment. Lipids were separated using an ACQUITY I-Class Ultra performance liquid chromatography (UPLC) on a C18 EXCEL UPLC column (2.1 × 150 mm, 2 μm; ACE) maintained at 45 °C with a reverse phase gradient. The eluting gradient was modified according to Zhu et al.79: 0–5 min 100% A, 5–10 min 0–24% B, 10–36 min 24–60% B, 36–38 min 60–90% B, 38–45 min 90% B, 45–45.1 min 90–100% A, 45.1–55 min 100% A. Solvent A (Methanol) and solvent B (isopropanol) are both amended with 0.1% NH4OH (25–30% NH3 basis) and 0.04% formic acid (>99.0%). The flow rate was 0.3 ml/min and the total run time was 55 min.
MS data were acquired in a Resolution mode by a Waters SYNAPT G2-Si quadrupole time-of-flight mass spectrometer (qTOF) coupled to an electrospray ionization (ESI) source operated at positive ion mode. The MS source parameters were identical to Chen et al.101: capillary 2.5 KV, source temperature 120 °C, source offset 80 V, sampling cone 45 V, desolvation gas 800 L/hr at 350 °C, cone gas 50 L/hr, nebulizer 6.5 bar. The mass range for MS1 and MS2 was set to 100–2000 and 50–2000, respectively. The MS2 spectrum was acquired by FAST-DDA mode, and the top 5 most abundant ions were selected for MS2 via collision-induced dissociation (CID) with a mass-dependent ramped collision energy that started at 10 eV, and ended at 15 eV for low mass ions (m/z 100); and started at 55 eV, ended at 65 eV for high mass ions (m/z 2000). The qTOF mass spectrometer was calibrated with sodium iodide (m/z 50–2000; Residual mass error <0.5 ppm) and a real-time calibration with leucine enkephalin solution ([M + H]+, m/z 556.2771).
Data processing
The raw data obtained by qTOF were converted into mzML format with the Waters2mzML script (V1.2.0)102 and the mzxml-precursor-corrector script was employed to fix the precursor values103. The processed mzML files were imported into the MS-DIAL software for data processing. Parameters included a minimum peak height of 1000 amplitude and a mass slice width of 0.1 Da. The deconvolution parameters for MS2Dec were set with a sigma window value of 0.5 and MS2 abundance cut-off of 10 amplitude. The peaks were aligned with a retention time tolerance of 0.3 min and an MS1 tolerance of 0.05 Da. Lipids were annotated with mass error within 0.01 Da for MS1 and 0.05 Da for MS2, and total scores over 70% that matched the libraries were considered as annotated compounds. A cutoff at 1% relative abundance of MS2 spectrum was used for compound identification to reduce the false positives caused by [M+Na]+ ions. The performance of spectral libraries in the ArchLips database was evaluated for both pure cultures and environmental samples with or without a 3 min retention time tolerance for compound filtering using the parameters described above. As this study primarily assessed the lipid annotation performance of the ArchLips database across diverse samples, additional manual verification was performed to remove entries with evidently poor spectral matches.
The feature table exported from the MS-DIAL software can be further processed using our python script (Combine_Adducts_For_MS_dial) to find possible adducts of identified features ([M + H]+, [M + NH4]+, and [M+Na]+). Subsequently, the peak area of different adducts was combined into a lipid compound for proxy calculation. Due to the lack of authentic standards for most archaeal lipids, their ionization efficiencies could not be determined. Consequently, the reported relative abundances do not represent true quantitative values, but serve to compare lipid distribution patterns across different samples104.
Statistical and clustering analyses were performed in R (v4.3.1, https://www.r-project.org/). The peak area of all annotated lipids were exported from the MS-DIAL software, and the lipid abundance data (peak area) of environmental samples were preprocessed by replacing zero values with one-fifth of the minimum non-zero value, followed by normalization to the total abundance per sample, log transformation, and z-score standardization by feature. The processed data were then clustered using Euclidean distance and Ward’s linkage, and visualized as heatmaps with the heatmap.2 function (gplots v3.2.0). The Shannon-Weiner diversity indices for archaeal lipids and for archaeal communities that derived from 16S rRNA sequencing at the ASV (Amplicon Sequence Variant) level were calculated as follows:
where pi corresponds to the proportion of lipid i or ASV i (i = 1, 2, …, s), s is the total number of lipids or ASVs.
Methane index of archaeal lipids in cold seeps were calculated with the following equation24:
Feature-based molecular networking construction
A molecular network was created with the Feature-Based Molecular Networking (FBMN) workflow on GNPS75,76,105. The mass spectrometry data processed with MS-DIAL were exported to GNPS for FBMN analysis. The data was filtered by removing all MS2 fragment ions within ±17 Da of the precursor m/z. The MS2 spectra were window-filtered by choosing only the top 6 fragment ions in the ± 50 Da window throughout the spectrum. The minimum fragment ion intensity in the MS2 spectra was set to 100. The precursor ion mass tolerance was set to 0.01 Da, and the MS2 fragment ion tolerance to 0.05 Da. A molecular network was created where edges were filtered to have a cosine score above 0.7 and more than 6 matched peaks. Further, edges between two nodes were kept in the network if and only if each of the nodes appeared in each other’s respective top 10 most similar nodes. The maximum size of a molecular family was set to 100, and the lowest-scoring edges were removed from molecular families until the molecular family size was below this threshold. The molecular networks were visualized using Cytoscape software. Network visualization was performed using Cytoscape software (https://cytoscape.org/).
Curation of the ArchLips database
In silico molecule structures
The core structures of tetraether lipids were manually drawn using the MarvinSketch software (V23.16.0), while the core structures of diether lipids were generated using the ChemAxon Reactor software (V21.3.0). The core structures were stored in EXCEL files to be extended or modified with JChem for EXCEL software. Additional structural information, including molecular formula, exact mass, SMILES, Inchi, InchiKey, and logP, was also calculated using the RDKit package (https://www.rdkit.org) in Python (https://www.python.org/).
The structures of intact polar lipids were generated by connecting the core structure templates with diverse polar headgroups. A total of 79 head groups were obtained from the literature (e.g., refs. 21,54,55,56,57,58,59,60,61,62,63,64,65,66, LipidMAPs (https://www.lipidmaps.org/) and LipidBank (https://lipidbank.jp/) were taken under consideration, including 39 phosphoric groups, 38 glycosyl groups (including a calditol group), one methylated and one acetylated group were also included (Fig. S1 and Table S2). Their structure (MDL molfiles) and structural information were also created and stored in an EXCEL file. The structures of intact polar lipids (IPLs) were created by combining the SMILES code with the RDKit. The generated SMILES were then transformed into the MDL molfile structures, with the structural information (e.g., formula, exact mass, InChI, InChIKey, logP) calculated using the RDKit and stored in SDF files. The MDL molfile structural files of IPLs were further standardized by the RDKit package using core lipids as templates. The structure of cardiolipins and quinones was constructed following a similar protocol (Fig. 1A).
In silico MS2 spectral library
A modified LipidBlast template was used for generating the in silico spectra library47. Two main precursor ions with adducts of [M + H]+ and [M + NH4]+, which are commonly detected under positive ion mode, are incorporated. This template preserved all structural information on a dedicated sheet, with MS2 data organized into multiple EXCEL sheets, each corresponding to a lipid class with similar core lipids. For example, all the PG-archaeol homologs or derivates with different unsaturation and chain lengths were stored in the same sheet. The first 28 columns of each MS2 sheet included the molecule name, structural information, and sidechain properties, followed by columns recording the m/z and intensity of MS2 peaks. The core lipid MS2 sheet served as a template, recording original sidechain properties shared by subsequent sheets, with other MS2 sheets synchronizing data from this template.
Structural information for each MS2 sheet was extracted from the structure sheet using the built-in VLOOKUP function of EXCEL, allowing updates and expansions by editing the structure sheet templates. The MS2 data and structural information were then exported as MSP-formatted library files, divided into 17 files according to lipid type to reduce the computational burden. The library exportation program, rewritten in Python, employed multiprocessing to handle multiple files simultaneously, expediting the process. During exportation, peak intensities for each molecule were standardized by normalizing the highest peak intensity to 1000 and removing peaks with intensities lower than 0.1. The exportation process for approximately 400,000 MS2 spectra took less than 8 min, resulting in an MSP file of 1.6 GB.
The retention time (RT) prediction
The RT prediction was performed by the Retip package (v0.5.4)106 in R (v4.3.1, https://www.r-project.org/). A dataset contained 265 identified lipids from the environment, and pure culture samples were randomly divided into a training set and testing set (4:1). Five different algorithms (Random Forest, BRNN, XGBoost, lightGBM and Keras) were used to build prediction models based on the training set, and their prediction performances were calculated and compared with the testing set. The model with the lowest RMSE (root-mean-square error) value (the BRNN model) was used to predict RT for in silico database of archaeal lipids.
Parameters for library validation
A total of 148 tandem mass spectra (MS2) of archaeal lipids and the characteristic peaks in MS2 reported in previous publications were digitized using the WebPlotDigitizer (v4.6)107. Those spectra were then formatted into an MGF file and annotated with the high-confidence spectral library. The LipiDex (v1.1) was used for library validation108. All MS2 data to be tested in this part (the high confidence spectral library and full spectral library of ArchLips database, open source libraries of LipidBlast and GNPS, and 148 archaeal lipid MS2 from literature) were transformed into brief versions of MSP files or MGF formatted files. The search parameters were set as ± 0.01 Da of MS1 or MS2 tolerance. The peaks with m/z > 61 in MS2 spectra were retained for similarity scoring.
Decoy search
The MSP formatted LipidBlast library (positive, version 68) was downloaded from http://prime.psc.riken.jp/compms/msdial/main.html#MSP, which included 81 classes, 377,313 molecules, and 554,041 spectra. The full GNPS library was downloaded from https://ccms-ucsd.github.io/GNPSDocumentation/gnpslibraries/, which covered 591,512 molecules. Neither libraries contain archaeal lipids, so the annotation can confidently be categorized as a false positive result. Further, the libraries were annotated against themselves to evaluate the replicate spectra.
DNA extraction and sequencing
Total DNA was extracted from about 0.5 g of sediment or soil using FastDNA SPIN for soil kit (MP Biomedicals, LLC). The V4 region of prokaryotic 16S rDNA was amplified by PCR with universal prokaryotic primers (515 F: GTGYCAGCMGCCGCGGTAA and 806 R: GGACTACNVGGGTWTCTAAT)109. The 50 μl PCR mixture contained 25 μl of 2 ×Premix Taq DNA polymerase (TaKaRa), 0.2 mM of each primer, 3 μl of template DNA and ddH2O. Thermocycling process included the following steps: initial denaturation for 30 s at 94 °C, followed by 30 amplification cycles consisting of 30 s denaturation at 94 °C, 30 s annealing at 58 °C and 30 s elongation at 72 °C. The PCR products were pooled and purified by EZNA Gel Extraction Kit (Omega, USA) and sent to Guangdong Magigene Biotechnology Co., Ltd. for sequencing (Illumina Miseq).
The raw 16S sequencing data of most environmental samples have been analyzed and published in refs. 33,96,97,98,99, including acid mine drainages in Anhui and Guangdong provinces96, permafrost soil from Tibet Plateau97, sediment core of cold seeps33 and a sediment core from the South China Sea98, and sediment from the East China Sea99. The raw sequencing reads of 48 environmental samples were collected and reanalyzed using the Quantitative Insights into Microbial Ecology (QIIME2, version 2023.5) software with default settings110. Taxonomy was assigned using the SILVA v138 99% dereplicated reference database (https://www.arb-silva.de/)111.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data necessary to evaluate the conclusions of this study are provided in the main text, figures, Supplementary information, and Source data files. The converted raw data (mzML format) of the archaeal lipidome of pure cultures and environmental samples used in this study are available in MassIVE under the accession codes MSV000097125 and MSV000097126. The processed data and parameter settings for the feature-based molecular network of pure cultures in this study are available at [https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=0de5541052674eff91d31c7c8505e53c]. A previously reported lipidomic dataset from four halobacterial strains, analyzed using a Q Exactive Orbitrap MS and available for download from MassIVE under accession code MSV00008983084,85, was used to evaluate the performance of the ArchLips database for lipid identification across different mass spectrometry platforms. Source data are provided with this paper.
Code availability
All custom scripts to generate the ArchLips spectral library and process the data in this study are available at [https://github.com/magicthree/archaeal_lipid_MS2_library], [https://doi.org/10.5281/zenodo.17580034] and [https://github.com/magicthree/Combine_Adducts_For_MS_dial], [https://doi.org/10.5281/zenodo.17580049]. The ArchLips database are available and download in zenodo [https://doi.org/10.5281/zenodo.16040382].
References
Javaux, E. J. Challenges in evidencing the earliest traces of life. Nature 572, 451–460 (2019.
Woese, C. R. & Fox, G. E. Phylogenetic structure of the prokaryotic domain: The primary kingdoms. Proc. Natl. Acad. Sci. USA 74, 5088–5090 (1977).
Woese, C. R., Kandler, O. & Wheelis, M. L. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc. Natl. Acad. Sci. USA 87, 4576–4579 (1990).
Brueckner, J. & Martin, W. F. Bacterial Genes Outnumber Archaeal Genes in Eukaryotic Genomes. Genome Biol. Evol. 12, 282–292 (2020).
Brunk, C. F. & Martin, W. F. Archaeal Histone Contributions to the Origin of Eukaryotes. Trends Microbiol. 27, 703–714 (2019).
Baker, B. J. et al. Diversity, ecology and evolution of Archaea. Nat. Microbiol. 5, 887–900 (2020).
Liu, Y. et al. Expanded diversity of Asgard archaea and their relationships with eukaryotes. Nature 593, 553–557 (2021).
Rinke, C. et al. A standardized archaeal taxonomy for the Genome Taxonomy Database. Nat. Microbiol. 6, 946–959 (2021).
Tahon, G., Geesink, P. & Ettema, T. J. G. Expanding Archaeal Diversity and Phylogeny: Past, Present, and Future. Annu. Rev. Microbiol. 75, 359–381 (2021).
Lepp, P. W. et al. Methanogenic Archaea and human periodontal disease. Proc. Natl. Acad. Sci. USA 101, 6176–6181 (2004).
Mafra, D. et al. Archaea from the gut microbiota of humans: Could be linked to chronic diseases? Anaerobe 77, 102629 (2022).
DeLong, E. F. Archaea in coastal marine environments. Proc. Natl. Acad. Sci. USA 89, 5685–5689 (1992).
Fuhrman, J. A., McCallum, K. & Davis, A. A. Novel major archaebacterial group from marine plankton. Nature 356, 148–149 (1992).
Orsi, W. D. et al. Metabolic activity analyses demonstrate that Lokiarchaeon exhibits homoacetogenesis in sulfidic marine sediments. Nat. Microbiol. 5, 248–255 (2019).
Imachi, H. et al. Isolation of an archaeon at the prokaryote–eukaryote interface. Nature 577, 519–525 (2020).
Lopez-Garcia, P. & Moreira, D. The Syntrophy hypothesis for the origin of eukaryotes revisited. Nat. Microbiol. 5, 655–667 (2020).
Da Cunha, V., Gaia, M. & Forterre, P. The expanding Asgard archaea and their elusive relationships with Eukarya. mLife 1, 3–12 (2022).
Koga, Y. Early evolution of membrane lipids: how did the lipid divide occur? J. Mol. Evol. 72, 274–282 (2011).
Koga, Y. From promiscuity to the lipid divide: on the evolution of distinct membranes in Archaea and Bacteria. J. Mol. Evol. 78, 234–242 (2014).
Villanueva, L., Schouten, S. & Damste, J. S. Phylogenomic analysis of lipid biosynthetic genes of Archaea shed light on the ‘lipid divide’. Environ. Microbiol. 19, 54–69 (2017).
Bale, N. J. et al. New Insights Into the Polar Lipid Composition of Extremely Halo(alkali)philic Euryarchaea From Hypersaline Lakes. Front. Microbiol. 10, 377 (2019).
Yao, W. et al. Lipidomic chemotaxonomy aligned with phylogeny of Halobacteria. Front. Microbiol. 14, 1297600 (2023).
Elling, F. J. et al. Respiratory quinones in Archaea: phylogenetic distribution and application as biomarkers in the marine environment. Environ. Microbiol. 18, 692–707 (2016).
Zhang, Y. G. et al. Methane Index: A tetraether archaeal lipid biomarker indicator for detecting the instability of marine gas hydrates. Earth Planet. Sci. Lett. 307, 525–534 (2011).
Schouten, S., Hopmans, E. C., Schefuβ, E. & Damste, J. S. S. Distributional variations in marine crenarchaeotal membrane lipids: a new tool for reconstructing ancient sea water temperatures? Earth Planet. Sci. Lett. 204, 265–274 (2002).
Hurley, S. J. et al. Influence of ammonia oxidation rate on thaumarchaeal lipid composition and the TEX86 temperature proxy. Proc. Natl. Acad. Sci. USA 113, 7762–7767 (2016).
Li, Y. et al. Biosynthesis of GMGT lipids by a radical SAM enzyme associated with anaerobic archaea and oxygen-deficient environments. Nat. Commun. 15, 5256 (2024).
Qin, W. et al. Confounding effects of oxygen and temperature on the TEX86 signature of marine Thaumarchaeota. Proc. Natl. Acad. Sci. USA 112, 10979–10984 (2015).
Elling, F. J. et al. Chemotaxonomic characterisation of the thaumarchaeal lipidome. Environ. Microbiol. 19, 2681–2700 (2017).
Koga, Y., Morii, H., Akagawa-Matsushita, M. & Ohga, M. Correlation of Polar Lipid Composition with 16S rRNA Phylogeny in Methanogens. Further Analysis of Lipid Component Parts. Biosci., Biotechnol., Biochem. 62, 230–236 (1998).
Koga, Y. & Nakano, M. A dendrogram of archaea based on lipid component parts composition and its relationship to rRNA phylogeny. Syst. Appl. Microbiol. 31, 169–182 (2008).
Boukhchtaber, D. C. et al. Discovering Hidden Archaeal and Bacterial Lipid Producers in a Euxinic Marine System. Environ. Microbiol. 27, e70054 (2025).
Zhang, T. T. et al. Lipidomic diversity and proxy implications of archaea from cold seep sediments of the South China Sea. Front. Microbiol. 14, 1241958 (2023).
Xu, Y. et al. Intact Ether Lipids in Trench Sediments Related to Archaeal Community and Environmental Conditions in the Deepest Ocean. J. Geophys. Res.: Biogeosci. 125, e2019JG005431 (2020).
Sollai, M., Villanueva, L., Hopmans, E. C., Reichart, G. J. & Sinninghe Damste, J. S. A combined lipidomic and 16S rRNA gene amplicon sequencing approach reveals archaeal sources of intact polar lipids in the stratified Black Sea water column. Geobiology 17, 91–109 (2019).
Pearson, A. & Ingalls, A. E. Assessing the use of archaeal lipids as marine environmental proxies. Annu. Rev. Earth Planet. Sci. 41, 359–384 (2013).
Pearson, A. in Treatise on Geochemistry (Second Edition). (eds. Holland H. D. & Turekian K. K.) 291–336 (Elsevier, Oxford; 2014).
Husen, P. et al. Analysis of lipid experiments (ALEX): a software framework for analysis of high-resolution shotgun lipidomics data. PLoS One 8, e79736 (2013).
Tsugawa, H. et al. A lipidome atlas in MS-DIAL 4. Nat. Biotechnol. 38, 1159–1163 (2020).
Schouten, S., Huguet, C., Hopmans, E. C., Kienhuis, M. V. & Damste, J. S. Analytical methodology for TEX86 paleothermometry by high-performance liquid chromatography/atmospheric pressure chemical ionization-mass spectrometry. Rapid Commun. Mass Spectrom.: RCM 79, 2940–2944 (2007).
Sinninghe Damsté, J. S. et al. The enigmatic structure of the crenarchaeol isomer. Org. Geochem. 124, 22–28 (2018).
Sinninghe Damsté, J. S. Crenarchaeol: the characteristic core glycerol dibiphytanyl glycerol tetraether membrane lipid of cosmopolitan pelagic crenarchaeota. J. Lipid Res. 43, 1641–1651 (2002).
Becker, K. W., Lipp, J. S., Zhu, C., Liu, X.-L. & Hinrichs, K.-U. An improved method for the analysis of archaeal and bacterial ether core lipids. Org. Geochem. 61, 34–44 (2013).
Hopmans, E. C., Schouten, S. & Sinninghe Damsté, J. S. The effect of improved chromatography on GDGT-based palaeoproxies. Org. Geochem. 93, 1–6 (2016).
Law, K. P. & Zhang, C. L. Current progress and future trends in mass spectrometry-based archaeal lipidomics. Org. Geochem. 134, 45–61 (2019).
Kind, T. et al. LipidBlast in silico tandem mass spectrometry database for lipid identification. Nat. Methods 10, 755–758 (2013).
Kind, T., Okazaki, Y., Saito, K. & Fiehn, O. LipidBlast templates as flexible tools for creating new in-silico tandem mass spectral libraries. Anal. Chem. 86, 11024–11027 (2014).
Koelmel, J. P. et al. LipidMatch: an automated workflow for rule-based lipid identification using untargeted high-resolution tandem mass spectrometry data. BMC Bioinforma. 18, 331 (2017).
Wang, F. et al. CFM-ID 4.0 - a web server for accurate MS-based metabolite identification. Nucleic Acids Res. 50, 165–174 (2022).
Ma, Y. et al. An in silico MS/MS library for automatic annotation of novel FAHFA lipids. J. Cheminformatics 7, 53 (2015).
Tsugawa, H. et al. Comprehensive identification of sphingolipid species by in silico retention time and tandem mass spectral library. J. Cheminformatics 9, 19 (2017).
Yuan, B. et al. Discovery of N-Acyl Amino Acids and Novel Related N-, O-Acyl Lipids by Integrating Molecular Networking and an Extended In Silico Spectral Library. Anal. Chem. 95, 8443–8451 (2023).
Zhou, Z. et al. In Silico-Predicted Dynamic Oxlipidomics MS/MS Library: High-Throughput Discovery and Characterization of Unknown Oxidized Lipids. Anal. Chem. 96, 2008–2021 (2024).
Yoshinaga, M. Y. et al. Systematic fragmentation patterns of archaeal intact polar lipids by high-performance liquid chromatography/electrospray ionization ion-trap mass spectrometry. Rapid Commun. Mass Spectrom.: RCM 25, 3563–3574 (2011).
Yoshinaga, M. Y., Kellermann, M. Y., Valentine, D. L. & Valentine, R. C. Phospholipids and glycolipids mediate proton containment and circulation along the surface of energy-transducing membranes. Prog. Lipid Res. 64, 1–15 (2016).
Knappy, C. et al. Mono-, di- and trimethylated homologues of isoprenoid tetraether lipid cores in archaea and environmental samples: mass spectrometric identification and significance. J. Mass Spectrom. 50, 1420–1432 (2015).
Knappy, C. S. et al. Structural complexity in isoprenoid glycerol dialkyl glycerol tetraether lipid cores of and other archaea revealed by liquid chromatography-tandem mass spectrometry. Chem. Phys. Lipids 165, 648–655 (2012).
Knappy, C. S., Chong, J. P. & Keely, B. J. Rapid discrimination of archaeal tetraether lipid cores by liquid chromatography-tandem mass spectrometry. J. Am. Soc. Mass Spectrom. 20, 51–59 (2009).
Liu, X.-L., De Santiago Torio, A., Bosak, T. & Summons, R. E. Novel archaeal tetraether lipids with a cyclohexyl ring identified in Fayetteville Green Lake, NY, and other sulfidic lacustrine settings. Rapid Commun. Mass Spectrom.: RCM 30, 1197–1205 (2016).
Liu, X.-L., Lipp, J. S., Schröder, J. M., Summons, R. E. & Hinrichs, K.-U. Isoprenoid glycerol dialkanol diethers: A series of novel archaeal lipids in marine sediments. Org. Geochem. 43, 50–55 (2012).
Liu, X.-L. et al. Mono- and dihydroxyl glycerol dibiphytanyl glycerol tetraethers in marine sediments: Identification of both core and intact polar lipid forms. Geochim. Cosmochim. Acta 89, 102–115 (2012).
Liu, X. L., Summons, R. E. & Hinrichs, K. U. Extending the known range of glycerol ether lipids in the environment: structural assignments based on tandem mass spectral fragmentation patterns. Rapid Commun. Mass Spectrom.: RCM 26, 2295–2302 (2012).
Zhu, C., Meador, T. B., Dummann, W. & Hinrichs, K. U. Identification of unusual butanetriol dialkyl glycerol tetraether and pentanetriol dialkyl glycerol tetraether lipids in marine sediments. Rapid Commun. Mass Spectrom.: RCM 28, 332–338 (2014).
Zhu, C. et al. Identification and significance of unsaturated archaeal tetraether lipids in marine sediments. Rapid Commun. Mass Spectrom.: RCM 28, 1144–1152 (2014).
Becker, K. W. et al. Unusual Butane- and Pentanetriol-Based Tetraether Lipids in Methanomassiliicoccus luminyensis, a Representative of the Seventh Order of Methanogens. Appl. Environ. Microbiol. 82, 4505–4516 (2016).
Elling, F. J. et al. Effects of growth phase on the membrane lipid composition of the thaumarchaeon Nitrosopumilus maritimus and their implications for archaeal lipid distributions in the marine environment. Geochim. Cosmochim. Acta 141, 579–597 (2014).
Liebisch, G. et al. Update on LIPID MAPS classification, nomenclature, and shorthand notation for MS-derived lipid structures. J. Lipid Res. 61, 1539–1555 (2020).
Kopczynski, D. et al. Goslin 2.0 Implements the Recent Lipid Shorthand Nomenclature for MS-Derived Lipid Structures. Anal. Chem. 94, 6097–6101 (2022).
Zeng, Z. et al. GDGT cyclization proteins identify the dominant archaeal sources of tetraether lipids in the ocean. Proc. Natl. Acad. Sci. USA 116, 22505–22511 (2019).
Liu, X.-L., Lipp, J. S., Birgel, D., Summons, R. E. & Hinrichs, K.-U. Predominance of parallel glycerol arrangement in archaeal tetraethers from marine sediments: Structural features revealed from degradation products. Org. Geochem. 115, 12–23 (2018).
Schouten, S. et al. Intact membrane lipids of “Candidatus Nitrosopumilus maritimus,” a cultivated representative of the cosmopolitan mesophilic group I Crenarchaeota. Appl. Environ. Microbiol. 74, 2433–2440 (2008).
Wörmer, L., Lipp, J. S., Schröder, J. M. & Hinrichs, K.-U. Application of two new LC–ESI–MS methods for improved detection of intact polar lipids (IPLs) in environmental samples. Org. Geochem. 59, 10–21 (2013).
Zeng, Z., Liu, X. L., Wei, J. H., Summons, R. E. & Welander, P. V. Calditol-linked membrane lipids are required for acid tolerance in Sulfolobus acidocaldarius. Proc. Natl. Acad. Sci. USA 115, 12932–12937 (2018).
Zeng, Z. et al. Identification of a protein responsible for the synthesis of archaeal membrane-spanning GDGT lipids. Nat. Commun. 13, 1545 (2022).
Nothias, L.-F. et al. Feature-based molecular networking in the GNPS analysis environment. Nat. Methods 17, 905–908 (2020).
Ding, S. et al. Lipidomics of Environmental Microbial Communities. II: Characterization Using Molecular Networking and Information Theory. Front. Microbiol. 12, 659315 (2021).
Zheng, F. et al. Diverse biological sources of core and intact polar isoprenoid GDGTs in terrace soils from southwest of China: Implications for their use as environmental proxies. Chem. Geol. 522, 108–120 (2019).
Wörmer, L., Lipp, J. S. & Hinrichs, K.-U. in Hydrocarbon and Lipid Microbiology Protocols: Petroleum, Hydrocarbon and Lipid Analysis. (eds. McGenity T. J., Timmis K. N. & Nogales B.) 289–317 (Springer Berlin Heidelberg, Berlin, Heidelberg; 2017).
Zhu, C. et al. Comprehensive glycerol ether lipid fingerprints through a novel reversed phase liquid chromatography–mass spectrometry protocol. Org. Geochem. 65, 53–62 (2013).
Pluskal, T., Castillo, S., Villar-Briones, A. & Oresic, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinforma. 11, 395 (2010).
Tsugawa, H. et al. MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 12, 523–526 (2015).
Chen, H. et al. Engineering archaeal membrane-spanning lipid GDGT biosynthesis in bacteria: Implications for early life membrane transformations. mLife 4, 193–204 (2025).
Zhang, J. et al. Biosynthesis of Hybrid Neutral Lipids with Archaeal and Eukaryotic Characteristics in Engineered Saccharomyces cerevisiae. Angew. Chem. Int. Ed. 62, e202214344 (2023).
Sorokin, D. Y. et al. sp. nov. and gen. nov., sp. nov., the first sulfur-respiring alkaliphilic haloarchaea from hypersaline alkaline lakes. Int. J. Syst. Evolut. Microbiol. 69, 2662–2673 (2019).
Sorokin, D. Y. et al. sp. nov., carbohydrate-utilizing, sulfur-respiring haloarchaea from hypersaline lakes. Syst. Appl. Microbiol. 44, 126249 (2021).
Behrendt, L. et al. Microbial dietary preference and interactions affect the export of lipids to the deep ocean. Science 385, eaab2661 (2024).
Holm, H. C. et al. Global ocean lipidomes show a universal relationship between temperature and lipid unsaturation. Science 376, 1487–1491 (2022).
Liu, W. et al. Unraveling plankton adaptation in global oceans through the untargeted analysis of lipidomes. Sci. Adv. 11, eads4605 (2025).
Ding, S. et al. Characteristics and origin of intact polar lipids in soil organic matter. Soil Biol. Biochem. 151, 108045 (2020).
Becker, K. W. et al. Daily changes in phytoplankton lipidomes reveal mechanisms of energy storage in the open ocean. Nat. Commun. 9, 5179 (2018).
Bale, N. J. et al. Lipidomics of Environmental Microbial Communities. I: Visualization of Component Distributions Using Untargeted Analysis of High-Resolution Mass Spectrometry Data. Front. Microbiol. 12, 659302 (2021).
Guo, J., Shen, S., Xing, S., Yu, H. & Huan, T. ISFrag: De Novo Recognition of In-Source Fragments for Liquid Chromatography–Mass Spectrometry Data. Anal. Chem. 93, 10243–10250 (2021).
Ahangarian, N. et al. Archaeal lipostratigraphy of the Scotian Slope shallow sediments, Atlantic Canada. EGUsphere 2025, 1–29 (2025).
Konneke, M. et al. Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437, 543–546 (2005).
Elling, F. J., Konneke, M., Mussmann, M., Greve, A. & Hinrichs, K. U. Influence of temperature, pH, and salinity on membrane lipid composition and TEX86 of marine planktonic thaumarchaeal isolates. Geochim. Cosmochim. Acta 171, 238–255 (2015).
Hao, Y. Q. et al. Microbial biogeography of acid mine drainage sediments at a regional scale across southern China. FEMS Microbiol. Ecol. 98, fiac002 (2022).
Tang, X. T. et al. Changing microbiome community structure and functional potential during permafrost thawing on the Tibetan Plateau. FEMS Microbiol. Ecol. 99, fiad117 (2023).
Wang, W. X. et al. Contrasting bacterial and archaeal distributions reflecting different geochemical processes in a sediment core from the Pearl River Estuary. AMB Express 10, 16 (2020).
Chen, Y. F. et al. Potential influence of bacterial community structure on the distribution of brGDGTs in surface sediments from Yangtze River Estuary to East China Sea. Chem. Geol. 647, 121934 (2024).
Sturt, H. F., Summons, R. E., Smith, K., Elvert, M. & Hinrichs, K. U. Intact polar membrane lipids in prokaryotes and sediments deciphered by high-performance liquid chromatography/electrospray ionization multistage mass spectrometry-new biomarkers for biogeochemistry and microbial ecology. Rapid Commun. Mass Spectrom.: RCM 18, 617–628 (2004).
Chen, Y. et al. The production of diverse brGDGTs by an Acidobacterium providing a physiological basis for paleoclimate proxies. Geochim. Cosmochim. Acta 337, 155–165 (2022).
Prisching, A. M. Waters2mzML (Version 1.2.0) [Computer software]. https://github.com/AnP311/Waters2mzML/releases/tag/v1.2.0 (2022).
Breaud, C. et al. LC-MS Based Phytochemical Profiling towards the Identification of Antioxidant Markers in Some Endemic Aloe Species from Mascarene Islands. Antioxidants 12, 50 (2023).
Ding, S., Hamm, J. N., Bale, N. J., Sinninghe Damsté, J. S. & Spang, A. Selective lipid recruitment by an archaeal DPANN symbiont from its host. Nat. Commun. 15, 3405 (2024).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Bonini, P., Kind, T., Tsugawa, H., Barupal, D. K. & Fiehn, O. Retip: Retention Time Prediction for Compound Annotation in Untargeted Metabolomics. Anal. Chem. 92, 7515–7522 (2020).
Rohatgi, A. WebPlotDigitizer user manual version 3.4. URL. http://arohatgi.info/WebPlotDigitizer/app, 1–18 (2014).
Hutchins, P. D., Russell, J. D. & Coon, J. J. LipiDex: An Integrated Software Package for High-Confidence Lipid Identification. Cell Syst. 6, 621–625.e625 (2018).
Walters, W. et al. Improved Bacterial 16S rRNA Gene (V4 and V4-5) and Fungal Internal Transcribed Spacer Marker Gene Primers for Microbial Community Surveys. mSystems 1, e00009–e00015 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Acknowledgements
We would like to thank Wenxiu Wang for providing GSD samples, Xiaotong Tang for the QTP permafrost soils and Bu Xu, Dongyu Cui for helping to collect soil samples from the western region of Sichuan Province, Xuemin Zhang for helping with lipid extraction. We gratefully acknowledge Prof. Laura Villanueva for providing valuable comments to improve this manuscript. This research was supported by the National Natural Science Foundation of China (Nos. 32393974 and 42372354 to F.Z.), the Guangdong Basic and Applied Basic Research Foundation (2021B1515120080 to C.Z.); the Shenzhen Key Laboratory of Marine Archaea Geo-Omics, Southern University of Science and Technology (SYSPG20241211173725010 to C.Z.), and F.Z. was supported by the High-Level Special Fund of SUSTech (G03050K001). Computation in this study was supported by the Centre for Computational Science and Engineering at the Southern University of Science and Technology. This work is also contributes to the Ocean Negative Carbon Emissions (ONCE) program and the Advanced Institute for Ocean Research, Southern University of Science and Technology.
Author information
Authors and Affiliations
Contributions
F.Z. and C.Z. conceptualized the project, designed and oversaw the experiments, wrote and revised the manuscript. W.Y. and W.H. performed coding and validation of the scripts, created the database, grew cell cultures, analyzed mass spectral data, and contributed to the writing and revision of the manuscript. W.Z., Y.C., and H.C. performed lipid extraction and lipid structure generation and contributed to the writing and revision of the manuscript. L.H. and Y.Z. helped with environmental sample collection and participated in the manuscript writing. Z.Z., X.L., Y.Z., and S.D. provided comments and advice and helped with the revision of the manuscript. All authors have reviewed and approved the submission of this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, F., Yao, W., He, W. et al. A comprehensive database for high-throughput identification of archaeal lipids using high-resolution mass spectrometry. Nat Commun 17, 588 (2026). https://doi.org/10.1038/s41467-025-67286-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67286-3