Abstract
Development of effective second-line treatment options for patients with BRAFwtNRASwt or BRAFwtNRASmut melanoma resistant to immune checkpoint blockade (ICB) is crucial. While systemic delivery of agonist CD40 (aCD40) plus anti-PD1 (αPD1) showed activity in patients with ICB-resistant melanoma, the objective response rate was modest (15%), in part due to induction of B regulatory cells (Bregs) which suppress CD8+ effector T cell responses. We previously reported that RAS/RAF/PI3K-inhibition elevates CD40 expression in melanoma cells and sensitizes tumors to ICB. Here, we show that combined treatment with a RAS/PI3K/AKT-pathway inhibitor rigosertib (RGS), and/or a MEK1/2 inhibitor trametinib (T), plus aCD40, overcomes the ICB resistance of BRAFwtNRASwt and BRAFwtNRASmut melanoma tumors growing in C57BL/6 mice. In addition, overexpression of CD40 in these melanoma cells effectively reverses ICB-resistance and aCD40 + αPD1 treatment induces tumor regression. Mechanistically, RGS + T suppress aCD40-associated CD11b+PD-L1+ Bregs, promoting CD8+ T-cell mediated killing in melanoma. scRNA-Seq analyses confirm CD40-associated CD11b+ Bregs across cancer types in patients. Our data demonstrate that addition of RAS/PI3K/AKT and MEK inhibitors to aCD40 resolves the issue of aCD40 induction of CD11b+PD-L1+ Bregs and provides alternative therapeutic options for ICB-resistant BRAFwtNRASwt or BRAFwtNRASmut metastatic melanoma.
Similar content being viewed by others
Introduction
Recent advances in immune checkpoint blockade (ICB) therapy for melanoma, such as targeting PD1, PD-L1, CTLA4, and LAG3, illustrate the power of ICB for enhancing anti-melanoma immune responses and improving clinical therapeutic outcomes1,2,3,4,5,6. While the first line of treatment for most patients with metastatic melanoma is currently ICB therapy, when primary or acquired ICB resistance develops, other options are essential. Primary resistance to ICB occurs in 40~55% of patients with melanoma treated with αPD1 and >70% of patients treated with αCTLA47. Acquired resistance also occurs in 25~30% of patients who relapse after exhibiting an initial response to αPD18. While an increasing number of preclinical and clinical studies have combined targeted therapies with ICB9,10, many patients still need a suitable 2nd line therapeutic approach after progressing on ICB.
Aberrant activation of RAS/RAF/PI3K signaling is due to driver mutations in BRAF or NRAS genes in melanoma and other tumor types11,12,13. When coupled with loss of one or more tumor suppressor genes, these mutations are associated with cancer onset, progression, and drug resistance. Therapeutic approaches that successfully target the RAS/RAF/PI3K pathway14,15,16, including BRAFV600, MEK, KRASG12C, and KRASG12D inhibitors, are being used to treat tumors driven by these pathways. Currently, ongoing therapeutic approaches include combining therapies that target the RAS/RAF/MEK/ERK and RAS/PI3K/AKT/mTOR with ICB inhibitors, though the sequence of delivery of these therapies remains controversial. While patients with BRAFmut tumors who are resistant to ICB are often treated with BRAF/MEK inhibitors, there is a crucial need to develop better second-line treatment options for patients with BRAFwt (NRASwt or NRASmut) melanoma who are ICB resistant17.
One potential treatment approach is to combine αPD1 with agonist CD40 (aCD40) antibodies. CD40, a member of the TNF receptor superfamily, is expressed in a variety of cell types and CD40–CD40 ligand (CD40L) interaction gives rise to many immune events, including the licensing of dendritic cells to activate CD8+ effector T cells, as well as the facilitation of B cell activation, proliferation, and differentiation6. Human aCD40 is emerging as an option for cancer treatment, and early-phase clinical trials explored its efficacy as a monotherapy or in combination with radiotherapy, chemotherapy, ICB, and other immunomodulatory approaches6. Systemic delivery of aCD40 plus αPD1 achieved prolonged response in patients with αPD1-resistant melanoma; however, the objective response rate was modest (15%)18. Relevant to these observations, a significant issue is that aCD40 treatment is associated with an increase in immunosuppressive CD11b+CD19+ B regulatory cells (Bregs) in preclinical glioma models19. These CD11b+ Bregs impair anti-tumoral T cell function and αPD1 therapeutic responses. In the murine model of experimental autoimmune hepatitis, IL-10-induced CD11b+ Bregs inhibited T-cell proliferation, interferon (IFN)γ production, and T-cell receptor (TCR) signaling20. In a case report of a melanoma patient whose B cells were depleted entirely by rituximab treatment prior to the diagnosis of metastatic melanoma, treatment with the αPD1 inhibitor nivolumab resulted in sustained tumor remission, suggesting that the depletion of all B cells (including Bregs) may have improved the efficacy of ICB in melanoma21. There is no consensus around canonical surface markers that identify Bregs in cancer, while they are characterized by the expression of various markers that indicate their immunosuppressive function, such as PD-L1, TIM-1, CD1d, CD27, and/or TIGIT22. Characterized primarily by their immunosuppressive functions and secretion of anti-inflammatory cytokines and growth factors, including IL-10, TGF-β, IDO1 and IL-35, Bregs suppress the activity of cytotoxic T cells and natural killer cells and promote regulatory T cell expansion22. On the other hand, tertiary lymphoid structures in melanoma tumors, comprised of the co-association of tumor-associated CD8+ T cells and CD20+ B cells, are associated with improved response to ICB and patient survival23. This divergence underscores the dual nature of B cells in melanoma dynamics and highlights the need for an improved understanding of aCD40-induced CD11b+ Breg-specific signaling pathways and therapies.
We previously demonstrated that oncogenic activation of the RAS/RAF/PI3K pathway by NRAS mutation suppressed the expression of CD40 in melanoma cells24,25. The inhibition of this pathway with rigosertib (RGS), a non-ATP-competitive small molecule RAS/PI3K/AKT-pathway inhibitor, synergized with ICB responses in a manner that was dependent on increased CD40 expression of melanoma cells, CD40-dependent immunogenic melanoma cell death, and CD8+ T effector cell responses24. However, only occasional tumor regression occurred, and the residual disease was associated with insufficient inhibition of MEK/ERK activity.
Here we show that significant tumor regression can be achieved by combining a low dose of agonist CD40 (aCD40) with a full spectrum RAS/PI3K/AKT-pathway inhibition via RGS and the MEK1/2 inhibitor, trametinib (RGS + T). While toxicity and induction of CD11b+ Bregs are two key issues raised by systemic aCD40 therapy, the addition of RGS + T boosts CD40 expression on melanoma cells, selectively suppresses the proliferation of CD11b+ Bregs, and restores IFNγ responses in T effector cells. Altogether, our data demonstrate a combination therapy of RGS, trametinib, and aCD40 that is predicted to be effective for patients with BRAFwt (NRASwt or NRASmut) metastatic melanoma who are resistant to ICB therapy. In addition, based on the transcriptome profile of murine and human melanoma, we further uncover a four-gene signature specific for CD11b+ Bregs, which is significantly associated with worse overall survival of patients across a broad spectrum of human cancer types.
Results
Agonist CD40 induces CD11b+ regulatory B cells in melanoma
While PD-L1+ Bregs were reported to suppress T cell responses in a PD-L1-dependent manner in melanoma patients with bone metastasis26, CD11b+ Bregs have not yet been identified in TME of melanoma patient tumors. Using immunofluorescent and H&E staining, we identified CD11b+CD20+ Bregs in tumors of metastatic melanoma patients resistant to αPD1 therapy (Fig. 1A). The CD11b+CD20+ Bregs were identified in both Melan-A- (Supplementary Fig. 1A) and Melan-A+ (Supplementary Fig. 1B) patient melanoma tissues. The published positivity rates of melanoma-specific marker Melan-A (MART-1) are considerably discordant for various patient tumor entities ranging from 69%–100% in primary and metastatic melanoma27. Analysis of two single-cell (sc)RNA-Seq datasets28,29 revealed that ~12% of tumor-infiltrated B cells (TIB) were CD11b+ Bregs in melanoma patient tumors at baseline (Fig. 1B, C, n = 818 B cells from 26 patients, and n = 1379 B cells from 30 patients, respectively). There was no significant alteration of the CD11b+ Breg frequency in patient tumor samples pre- (n = 10) and post-αPD1 (n = 15) therapy (Fig. 1D). Nevertheless, by comparing paired pre- vs post-αPD1 samples (n = 3), we observed that the two tumors of αPD1-responders exhibited 67% and 82% reduction of percentage of CD11b+ Bregs, in sharp contrast to the non-responder with an 8% reduction of CD11b+ Bregs from baseline, suggesting greater attenuation of CD11b+ Bregs in melanoma tumors may correlate with a better αPD1 response. However, due to the small sample size of paired pre- and post-treatment tumors, we could only identify a trend toward an association between reduced CD11b+ Breg and patient response to αPD1. Additional studies will be needed to validate the role of CD11b+ Breg in αPD1 treatment responses.
A Immunofluorescence staining in tumors of patients with metastatic melanoma resistant to αPD1 therapy. DNA (blue), CD8α (green), CD20 (red) and CD11b (yellow). Scale bars indicate 2 mm (left) and 50 μm (right). These results are representative of immunofluorescence staining performed on melanoma tumors from two patients. B–F Single-cell (sc) RNA-Seq analysis identifies CD20+CD11b+PD-L1+ regulatory B cells in patient melanoma tumors. References: (B) Cell 2018, 175, 984–997. doi: j.cell.2018.09.006. C Cell 2018, 175, 998–1013.e1–e20. doi: j.cell.2018.12.034. Samples with total B-cell size <5 were excluded. D Frequency of CD20+CD11b+PD-L1+ regulatory B cells in patient melanoma tumors at baseline and post αPD1 treatment. Data were derived from (C). E, F Patient melanoma scRNA-Seq analysis reveals a high CD40 mRNA expression in CD20+CD11b+PD-L1+ Bregs. E n = 818 B cells from 26 patients, two-sided t-test; and (F) n = 1379 B cells from 30 patients. One-way analysis of variance (ANOVA) with post hoc test.
To access the possible mechanism for the preferential induction of aCD40 on CD11b+ Breg over CD11b- B cell counterparts, we quantified CD40 expression in human and murine melanoma tumors, as well as murine splenic B cell samples. Our analysis of the two available human melanoma scRNA-Seq datasets28,29 revealed a significantly higher positivity and intensity of CD40 mRNA in CD11b+PD-L1+ Bregs compared to CD11b-PD-L1- B cells in patient tumors (Fig. 1E), while αPD1 did not alter the CD40 mRNA level in TIB cells (Fig. 1F). Consistently, we identified that aCD40 increased CD40 protein expression from 45% to 75% positivity in tumor-infiltrated CD11b+ Bregs in mice, and these CD11b+ Bregs also exhibited a significantly higher CD40 positivity than CD11b- B cells (35%) in aCD40-treated NRASmut 1014 murine melanoma tumors (Fig. 2A). We next questioned whether aCD40 alone independent of other factors in the TME is sufficient for CD40 induction in splenic B cells. Indeed, in vitro experiments showed that aCD40 increased CD40 protein positivity in CD11b+ Bregs from <10% to ~70% while only 40% of CD11b- splenic B cells are CD40+ post aCD40 exposure (Fig. 2B). Agonist CD40 consistently increased the CD40 intensity in CD11b+ Bregs as compared to CD11b- B cells (Fig. 2B). Taken together, the higher endogenous and inducible levels of CD40 expression on CD11b+ Bregs, compared to CD11b- B-cell counterparts, may position them to respond better to the proliferative signal under CD40 agonism19.
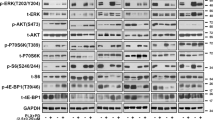
A 1014 cells were injected in C57BL/6 female mice. Treatment of C57BL/6 mice with aCD40 (30 μg/mouse every 3 days, intraperitoneal) started at Day 10 post tumor cell inoculation. Analysis of CD40 levels by flow cytometry of CD45+CD19+ B cells prepared from tumor samples at day 22 post-treatment (n = 5 per group) using a one-sided Mann–Whitney U test. B Splenocytes were isolated from C57BL/6 mice by magnetic separation, serum-free starved for 4hrs, and stimulated with or without 12.5 μg/ml aCD40. After 4 days of culture, splenic B cells were isolated by magnetic separation and stained for flow cytometric analysis (n = 6 per group). C, D Tumor volume of 1014 melanoma in C57BL/6 female mice. Treatment with 200 μg/mouse αPD1 (every 3 days, intraperitoneal), 100 μg/mouse αCTLA4 (every 3 days, intraperitoneal), 30 μg/mouse agonist CD40 (aCD40, every 3 days, intraperitoneal), or IgG control antibody, starts at day 10 post tumor cell inoculation. E Frequency of CD19+CD11b+ regulatory B cells in 1014 melanoma tumors or tumor draining lymph node (TDLN) after 22 days of aCD40 treatment (every 3 days, intraperitoneal) TDLN IgG, n = 5; TDLN aCD40, n = 5; Tumor IgG, n = 4; Tumor aCD40, n = 5. F Flow cytometric analysis of CD45+CD19+ B cells in the tumor microenvironment of 1014 melanoma after 3 weeks treatment of aCD40. Data were replicates from one experiment (n = 5 mice per group). G C57BL/6 mice were intravenously injected with formalin-fixed 2 × 106 metastatic B16F10-Luc2 cells via the tail vein. Splenic CD8+ T and total B cells were isolated by magnetic separation after 2 days of tumor antigen priming. Total B cells were stimulated with 12.5 μg/ml aCD40 for 2 days and CD11b+ B cells were isolated by magnetic separation. CD8+ cytotoxic T cell killing assay were performed via co-culturing CD11b+ B cells, CD8+ T cells and B16F10-Luc2 cells in the conditioned RPMI complete medium containing 0.5 μg/ml of IL-7, 30U/ml of IL-2, and CD3/CD28 Dynabeads at a bead-to-cell ratio 2:1. Dead cells were removed and the luciferase activity of remaining live tumor cells was quantified to calculate % inhibition of CTL killing capacity. (n = 3 independent experiments). Created in BioRender. Yan, C. (2026) https://BioRender.com/83jrya2. Exact p values are provided as a Source Data file. H Splenic B cells were stimulated with 12.5 μg/ml aCD40 for 4 days. The CD11b- and CD11b+ B cells were isolated by magnetic separation and restimulated with 12.5 μg/ml aCD40 for 30 min. Whole-cell extracts were harvested and immunoblotted. β-actin was used as a loading control for densitometry quantification. n = 3 independent experiments. One-way analysis of variance (ANOVA) with post hoc test. Source data are provided as a Source Data file.
We next evaluated the ICB response in a murine allograft syngeneic model of NRASmut 1014 melanoma. When tumors reached a size of 5 mm diameter 10 days after tumor cell inoculation, a 3-week treatment of αPD1 (200 μg/mouse, every 3 days, intraperitoneal), αCTLA4 (100 μg/mouse, every 3 days, intraperitoneal), αPD1 + αCTLA4, or IgG control was initiated. The tumor growth was not altered compared to IgG control-treated tumors, suggesting a complete resistance of NRASmut 1014 melanoma to ICB therapy (Fig. 2C). Importantly, the ICB-resistant treatment-naive 1014 tumors exhibited an 82.9% reduction of tumor growth (adj.p < 0.001) in response to systemic low-dose aCD40 treatment (30 μg/mouse, every 3 days, intraperitoneal) as compared to IgG control (Fig. 2D). FACS analysis of the immune content of tumors post treatment revealed that residual disease after continual treatment response was associated with an induction of CD11b+CD19+ B cells. Compared to IgG control-treated samples, aCD40 treatment resulted in a significant enrichment of these CD11b+CD19+ B cells in the tumor microenvironment (TME), from 60% at baseline to 80% of total TIB, compared to induction from <1% to ~5% in the tumor-draining lymph nodes (TDLN) (Fig. 2E). Phenotypic analysis by flow cytometry of aCD40-induced TIB revealed a B2-like (CD19+CD43-CD5-B220lowCD23lowCD21low) lineage feature30,31, while possessing integrin CD11b (a myeloid cell marker) and immune regulatory B-cell markers (PD-L1highCD1dlow) (Fig. 2F). In addition, aCD40-induced TIB retained MHC-II expression, but exhibited suboptimal co-stimulatory function (CD86-) or activation (CD69-) (Fig. 2F). Consistently, these CD11b+CD19+ B cells exhibit a high expression of PD-L1, a previously identified marker of Bregs24,26, reaching ~87% with a 26.5-fold increase in the cell surface PD-L1 expression level among aCD40-induced TIB in the TME. We further compared the potency of reported stimuli, such as aCD40 and lipopolysaccharide (LPS)19,20, on the retention and expansion of CD11b+CD19+ B cells in a splenocyte in vitro assay (Supplementary Fig. 2A–E). After 48 h of stimulation, splenocytes treated with 12.5 µg/ml aCD40 or 1 µg/ml LPS exhibited an increase in the total B cells from 20% to 40–50%. Among these expanded B cells, 8–12% were CD11b+. Notably, about 40% of the in vitro expanded CD11b+ B cells were PD-L1+, among which 25% also produced the immunosuppressive cytokine IL-10. In sharp contrast, there was no detectable expression of PD-L1 on CD11b- B cells and merely 10% of CD11b- B cells were IL-10+ (Supplementary Fig. 2E), suggesting the Breg population is enriched in the CD11b+ B cell subsets. Furthermore, aCD40-induced CD11b+ B cells exhibited a higher level of MHC-II (Supplementary Fig. 3A) and a distinct cytokine secretion profile compared to CD11b- B cells (Supplementary Fig. 3B). Among 80 screened cytokine proteins, CD11b+ B cells secreted significant higher levels (fold-change > 2) of EGF, IL-12p70, IL-12p40, I-TAC (CXCL11), MDC (CCL22), OPN, Prolactin, P-selectin, SCF, TPO, VCAM-1, VEGF, Fas-L, IL-5, MCP-5 (CCL12), MIP-1α (CCL3), and TNF-RI, compared to CD11b- B cells. Agonist CD40 stimulation specifically induced LIX (CXCL5) secretion by CD11b+ B cells but not CD11b- B cells, and further increased the secretion of IL-12p70, TPO and MCP-5 (CCL12) in both CD11b- and CD11b+ B cells. To functionally evaluate whether aCD40-induced CD11b+CD19+ B cells inhibited CD8+ T-cell cytotoxicity, we isolated CD8+ T cells from mouse spleens two days after ‘tumor priming’ via tail vein injection of formalin-fixed luciferase-expressing B16F10 (B16F10-luc2) cells. Subsequently, the ex vivo cytotoxicity of the CD8+ T cell targeting of B16F10-luc2 cells was evaluated by incubating the CD8+ T cells with live B16F10-luc2 tumor cells (Effector:Target = 50:1) with different ratios of CD11b+ B cells isolated by magnetic separation (Fig. 2G). As expected, aCD40-induced CD11b+ B cells exhibited a direct and potent suppression of tumor-primed CD8+ T-cell killing of B16F10-luc2 cells in a dose-dependent manner. Notably, with a 1:1 ratio, aCD40-induced CD11b+ B cells suppressed ~60% of the killing capacity of CD8+ T cells (Fig. 2G). Thus, we concluded that the aCD40-induced CD11b+ B cells negatively regulate the anti-tumor immune function of CD8+ T cells in melanoma. Importantly, compared to the CD11b- B cell counterparts, aCD40-induced CD11b+ Bregs exhibited higher endogenous levels of RAS-pathway activity (heightened levels of total- and phospho- ERK as well as total AKT), making them vulnerable to RAS pathway-targeted therapies (Fig. 2H), such as the MEK inhibitor trametinib that might target both ERK-addicted tumor cells and CD11b+ Bregs in the TME.
Anti-PD1 resistant 1014 NRAS mut melanoma tumors respond to inhibitors of the RAS/MEK/PI3K pathway
We previously reported that the inhibition of the RAS/PI3K/AKT pathway via rigosertib (RGS), a non-ATP-competitive small molecule, synergized with ICB to inhibit tumor growth in a manner that was dependent on melanoma-cell selective CD40 induction and CD8+ T-cell responses24. However, only occasional tumor regression occurred with this treatment regime, which was associated with insufficient MEK/ERK inhibition in BRAFmut melanoma24. Similarly, we demonstrate here that RGS suppressed 70% of tumor growth of the ICB-resistant 1014 NRASmut melanoma after four weeks of treatment (adj.p < 0.001) (Fig. 3A). The TME of RGS-treated tumors were enriched with 2-fold higher frequency and 10-fold higher activation of CD8+ T cells (p = 0.003 and p = 0.004, respectively), as well as M1 macrophage polarization (p = 0.0135) to facilitate anti-tumor immune responses (Fig. 3B). Consistent with our findings in BRAFmut melanoma24, in 1014 NRASmut melanoma cells treatment with 1 μM RGS resulted in early inhibition (60 min) of AKT activity, but it did not fully inhibit MEK/ERK activity, at either 60 min or 24 h treatment timepoints (Fig. 3C). Therefore, we next combined RGS (1 μM) with the MEK inhibitor trametinib (1 μM), which together effectively inhibited both PI3K/AKT and MEK/ERK pathways in 1014 NRASmut melanoma cells (Fig. 3C). Remarkably, treatment with 0.75 μM RGS, 0.5 μM trametinib, or R + T (0.5 μM + 0.5 μM) converted a significant number of CD40-negative 1014 melanoma cells to CD40-positive cells (1.5-fold adj.p = 0.0249 increase for RGS, and 3.5-fold adj.p < 0.001 increase for trametinib alone or RGS + T, respectively) (Fig. 3D). Notably, there was no significant difference in the percentage of CD40+ 1014 cells in response to treatment with trametinib verses RGS + T, suggesting a dominant role of RAS/RAF/MEK inhibition via trametinib in CD40-induction resulted from the combined therapeutic regimen.
A Tumor volume of 1014 melanoma in C57BL/6 female mice. Treatment with 300 mg/kg rigosertib (RGS, 5 days a week, oral gavage) or vehicle control (H2O) starts at day 10 post tumor cell inoculation. B Tumor samples were collected from each treatment group at 4 weeks of treatment and immune profiled by FACS analysis. All graphs show Mean ± SEM, n = 5 samples per group. C 1014 tumor cells were treated with 1 μM trametinib (T) and/or 1 μM RGS for 30–60 min. Whole-cell extracts were harvested and immunoblotted. β-actin was used as a loading control for densitometry quantification. n = 3 independent experiments. Source data are provided as a Source Data file. D RGS and T induce CD40 expression in 1014 cells. Cells were treated with RGS + /- T for 24 h. CD40 protein expression on the cell surface was detected by anti-CD40-FITC staining and flow cytometric analysis. n = 3 independent experiments. E Tumor volume of 1014 melanoma in C57BL/6 female mice. Treatment with 200 μg/mouse αPD1 (every 3 days, intraperitoneal), 300 mg/kg rigosertib (RGS, 5 days a week, oral gavage), and/or 1 mg/kg trametinib (T, 5 days a week, oral gavage), starts at day 8 post tumor cell inoculation, n = 7 ~ 10 mice per group. Created in BioRender. Yan, C. (2026) https://BioRender.com/83jrya2. Exact p values are provided as a Source Data file. F Definition of time to resistance to drug: (1) the tumor is > 100 mm3 and (2) has a > 30% increase of tumor volume compared to the previous measurement. Survival curves (resistance-free probabilities or survival probabilities) of treatment groups are estimated using the Kaplan–Meier Plotter and compared using the log-rank test, n = 7 ~ 9 mice per group. Exact p values are provided as a Source Data file. G Mouse body weight recorded for 20 days of treatment. Veh, n = 10; T, n = 8; RGS, n = 7, RGS + T, n = 9 mice. H Tumor and (I) Tumor-draining lymph node (TDLN) samples were collected from each treatment group at day 20 of treatment and immune profiled by FACS analysis, Veh, n = 5; T, n = 4; RGS, n = 4, RGS + T, n = 4 tumors. For tumor samples, we collected 200,000 live singlet cells from each tumor and recovered ~6000 CD45+ cells per tumor, regardless of treatment groups. Tc, CD8+CD3+ cytotoxic T cells. Th, CD4+CD3+ T helper cells. Exact p values are provided as a Source Data file.
To access the feasibility and mechanism of broad-spectrum RAS/MEK/PI3K-pathway inhibition in NRASmut melanoma to sensitize ICB therapy, we pre-treated 1014 NRASmut melanoma tumors with two doses of αPD1 (200 μg/mouse, intraperitoneal) to validate the ICB non-responsive phenotype and then continued administration of αPD1 (200 μg/mouse, every 3 days, intraperitoneal) and added either vehicle control, RGS (300 mg/kg, 5 days a week, oral gavage), trametinib (1 mg/kg, 5 days a week, oral gavage), or RGS + T (5 days a week, oral gavage) (Fig. 3E). Combinatorial treatment of RGS + T plus αPD1 resulted in superior inhibition tumor growth of 1014 NRASmut melanoma compared to either αPD1 alone, or αPD1 plus single arm of RAS-pathway inhibition (adj.p < 0.001, compared to RGS, or adj.p = 0.052 compared to trametinib). In addition, 12.5% (1/8) and 0% (0/7) of the tumors completely regressed (CR), after αPD1 plus trametinib or RGS treatment respectively, compared to 0% (0/7) complete tumor regression in the control group. Notably, 44% (4/9) NRASmut melanoma tumors receiving combinatorial treatment of RGS + T plus αPD1 exhibited CR after 20 days of treatment and an extended median αPD1-resistance-free survival (median 27 days vs. 6 days for αPD1 alone, p < 0.0001, Fig. 3F). RGS + T plus αPD1 treatment did not result in significant loss of mouse body weight, suggesting a safe toxicity profile at the doses used here (Fig. 3G).
To our surprise, immune profiling of the TME revealed that the 1014 NRASmut melanoma tumors exhibited the classic feature of so-called “Immune Desert” tumors, which are characterized by a lack of CD8+ T cells, and are associated with poor responsiveness to ICB32. The baseline frequency of T cells in the 1014 tumors may be affected by different vehicles used for targeted therapy, the administration of αPD1, and/or the endpoint of tumor growth (Fig. 3B, H). Notably, tumors in the combinatorial treatment group exhibited increased frequency (from 0% to 20% in total CD3+ T cells) and activation (from 0% to 60%) of CD8+ T cells, along with a reduced frequency in CD206+F4/80+ M2 macrophages (from 60% to 20% of total macrophages) (Fig. 3H). Interestingly, while trametinib promoted T-cell enrichment and the decrease of M2 macrophages in the TME, RGS preferentially increased CD8+ T-cell activation. Consistent with the elevated anti-tumor immune response in the TME, analysis of the TDLN revealed that combinatorial treatment resulted in increased frequencies of CD4+ T helper cells (Th), CD11c+MHCII+ dendritic cells (DC), and MHCII+F4/80+ M1 macrophages (Fig. 3I). Taken together, our data suggest an initial lead-in treatment with αPD1, followed by RGS + T inhibition of the RAS pathway is a potent therapeutic strategy for durable tumor control of ICB-resistant 1014 NRASmut melanoma.
RAS/MEK/PI3K inhibition selectively attenuates aCD40-induced CD11b+ regulatory B cells and promotes CD8+ T effector cell activity
Our group previously demonstrated that while activation of the oncogenic RAS pathway in melanoma tumor cells masks CD40 expression, inhibition of the RAS/MEK/PI3K pathway results in an upregulation of CD40 membrane expression in melanoma cells24,25. Here, we hypothesized that RAS/MEK/PI3K inhibition to restore CD40 expression in RAS-addicted melanoma cells, combined with a low-dose aCD40, may provide effective second line therapy for ICB-resistant tumors. However, one critical issue is that aCD40 induction of immunosuppressive CD11b+ Bregs19,20 could potentially inhibit response to ICB (Fig. 2). Surprisingly, we found that RGS selectively and directly inhibited the proliferation and expansion of aCD40-induced splenic CD11b+ IL-10-producing B cells in vitro (Fig. 4A). Interestingly, RGS did not abrogate LPS induction of splenic B cells (Fig. 4A). As a result of this RGS suppression of aCD40-induced CD11b+ Bregs, there was a significant increase in interferon (IFN)γ-producing T cells and activated CD8+ T cells in the splenic CD11b+ Breg cell co-cultures (Fig. 4B). We next asked whether inhibition of the RAS/MEK/PI3K pathway would suppress aCD40-induction of CD11b+ Bregs and restore T-effector cell responses in vivo. In these experiments we combined low-dose aCD40 (30 μg/mouse, every 3 days, intraperitoneal) with vehicle, RGS (300 mg/kg 5 days a week, oral gavage), trametinib (1 mg/kg, 5 days a week, oral gavage), or RGS + T, in a murine preclinical study of αPD1-primed 1014 NRASmut melanoma (Fig. 4C). Consistent with the deep and durable RAS signaling inhibition of RGS + T (Fig. 3C), combined RGS + T in combination with 30 μg/mouse aCD40 resulted in superior tumor growth inhibition as compared to vehicle (adj.p < 0.001), RGS alone (adj.p < 0.001) or trametinib alone (adj.p = 0.051), all in combination with aCD40. Notably, nearly 95% regression of the 1014 melanoma tumors was achieved with the aCD40 + RGS + T combinatorial therapy after 27 days of treatment in our preclinical trial. We observed a transient 10% decrease of mouse body weight in the first week of aCD40 treatment in all mice (30 μg/mouse, every 3 days, intraperitoneal) (Fig. 4D), although this was quickly reversed in all aCD40-treated mice without intermittent or reduced dosing. At the experimental endpoint of day 27 post-treatment, there was no evidence of toxicity based upon changes in serum levels of alanine aminotransferase (ALT), aspartate aminotransferase (AST), and blood urea nitrogen (BUN) (Fig. 4E).
A, B Splenocytes were isolated from C57BL/6 mice, serum-free starved for 4 h, and stimulated with 12 μl/ml anti-CD3/CD28 Dynabeads plus 12.5 μg/ml agonist CD40 (aCD40). After 5 days of culture, cells were stained for FACS analysis. Exact p values are provided as a Source Data file. C Tumor volume of 1014 melanoma, created in BioRender. Yan, C. (2026) https://BioRender.com/83jrya2, exact p values are provided as a Source Data file; and D Mouse body weight of tumor-bearing C57BL/6 female mice. Treatment with 200 μg/mouse αPD1 (2 lead-in doses, every 3 days, intraperitoneal), 30 μg/mouse aCD40 (every 3 days, intraperitoneal), plus 300 mg/kg rigosertib (RGS, 5 days a week, oral gavage) and/or 1 mg/kg trametinib (T, 5 days a week, oral gavage), starts at day 8 post tumor cell inoculation (n = 10 per group). E Serum level of liver/kidney enzymes alanine aminotransferase (ALT), aspartate aminotransferase (AST) and blood urea nitrogen (BUN) at day 27 post-treatment (n = 5 per group). Live CD45+ leukocytes in (F) Tumor and (G) Tumor-draining lymph node (TDLN) samples were concatenated after downsampling to, 25,000 and 10,000 events respectively, for t-SNE analysis through flow cytometry. All graphs show Mean ± SEM, n = 5 per group. H–K Frequency of B cells and effector cells in 1014 tumors and TDLN samples at day 27 post-treatment (n = 5 per group). Exact p values are provided as a Source Data file. L L308 mouse protein array of 1014 tumor lysate samples at day 27 post-treatment. Data were pooled from replicates of one experiment (n = 5 mice per group). Exact p values are provided as a Source Data file.
To evaluate the breadth of antitumor immune response, we performed FACS of immune cells in the TME (Fig. 4F) and tumor-draining lymph nodes (TDLNs) (Fig. 4G) followed by tSNE analysis. We observed a significant increase of total CD45+ leukocytes specifically in the TME but not in the TDLNs. Consistent with our in vitro findings (Fig. 4A), RGS significantly reduced tumor-infiltrating CD11b+ Breg frequency from 10% in the aCD40+vehicle treated tumors to <2%, with less effect from trametinib (6% CD11b+ Bregs) (Fig. 4H). Importantly, the remaining TIB cells were activated (CD69+ B cells) with a frequency around 4% in the aCD40 + vehicle treated tumors, and this percentage increased to ~12% in the aCD40 + RGS + T treated tumors (adj.p = 0.0019), confirming a reduction in the immunosuppression function of CD11b+ Bregs in response to RAS/PI3K/AKT pathway inhibition by RGS. The unleashed anti-tumor effector response in the aCD40 + RGS + T treated tumors was also evidenced by an elevation of CD8+ T-cell frequency (from 25% to 45% of total T cells, adj.p < 0.001) and their activation (from 20% to 25% of total CD8+ T cells, adj.p = 0.032), as well as an increase in M1 macrophages (from 8% to 16% of total macrophages, adj.p = 0.011) in the TME (Fig. 4I). Consistently, aCD40 + RGS + T treatment resulted in reduction in CD11b+ Bregs in the TDLNs (Fig. 4J), along with enrichment of activated CD8+ T cells and M1 macrophages (Fig. 4K), demonstrating an improved cancer immunity at both induction and effector sites that are associated with therapeutic outcome in the ICB-resistant NRASmut tumors. To gain further insight into molecular mechanisms of anti-tumor activity of aCD40 and RGS + /- T combined treatments, tumor cytokines and growth factors were examined in combined drug-treated versus aCD40+vehicle-treated tumor lysates using a L308 protein array (Fig. 4L). Among the significantly altered proteins with |Log2Fold-change | ≥1.5 compared to aCD40 + Veh, we observed that adding RGS to aCD40 decreased the levels of CCR4, a chemokine receptor for regulatory T cells (Tregs)33, while trametinib decreased IL-11 levels. IL-11 expression by tumor fibroblasts has been reported to be correlated with poor prognosis of solid cancers34. Interestingly, combining either RGS or trametinib with aCD40 significantly downregulated IL-335 and epigen (an EGRF ligand)36, two proteins known to regulate angiogenesis and tumor cell growth. In contrast to the single arm inhibition of the RAS/MEK/PI3K pathway, full spectrum inhibition of RAS via RGS + T resulted in an upregulation of 14 proteins, including proinflammatory toll-like receptor (TLR)1, IL-17 and IL-9R37, IL-17C38 and IL-22 binding protein (IL-22BP)39, etc. Importantly, significant induction of tumor necrosis factor (TNF)α, was identified as a common feature of aCD40 treatment when combined with targeted inhibition of RAS/MEK/PI3K via either RGS alone, trametinib alone, or RGS + T combinatorial therapy (Fig. 4L). To determine whether CD19+ B cells or CD8+ T cells are required for the RGS + T + aCD40 anti-melanoma activity, we conducted depletion experiments of CD19 or CD8 cells. Interestingly, the depletion of CD8 cells (Supplementary Fig. 4A) but not depletion of CD19+ total B cells (Supplementary Fig. 4B), significantly abrogated 1014 melanoma tumor growth inhibition by RGS + T + aCD40 treatment. These results suggest that CD8+ T cells play an important role in the anti-tumor activity triggered by combinatorial therapy. While total B cell depletion did not alter tumor growth, this may be partially explained by the heterogeneity of B cell subpopulations.
We next questioned whether the lead-in αPD1 is required for the therapeutic benefit of aCD40 plus RAS/MEK/PI3K inhibition in ICB-resistant NRASmut tumors. We found that while low-dose 30 μg/mouse aCD40 alone did not significantly suppress 1014 tumor growth in this αPD1-naive setting (adj.p = 0.564), RGS + /- T with or without low-dose aCD40 resulted in a significant inhibition. Notably, either adding aCD40 to RGS + T, or adding RGS + T to aCD40, significantly improved the tumor growth control of 1014 melanoma (adj.p = 0.029 or 0.011, respectively) (Supplementary Fig. 5A). Consistently, we observed a ~ 5% mild and transient reduction of mouse body weight in the first week of aCD40 treatment (Supplementary Fig. 5B). In addition, systemic delivery of aCD40 triggered CD11b+ Breg induction in tumors (adj.p = 0.058) and RGS + T successfully attenuated aCD40 induction of CD11b+ Bregs (adj.p = 0.005) and M2 macrophages (adj.p = 0.0014) and also enriched total (adj.p = 0.0001) and activated CD8+ T cells (adj.p = 0.0108) in the TME (Supplementary Fig. 5C). The combination of RGS + T and aCD40 promoted the percentage and intensity of MHC-II protein expression on CD11b- B cells (Supplementary Fig. 5D). While aCD40 plus RGS + T reduced the viability of CD45- cells in tumors, including tumor cells, it sharply increased the number of CD40 positive CD45- cells (from 5% to 15%) and resulted in an 40% elevation of CD40 intensity of CD45- cells (Supplementary Fig. 5E), suggesting an effective CD40-dependent positive feedback loop of combinatorial drug-induced tumor eradiation in the absent of ICB therapy24.
To test whether RAS-pathway inhibition and aCD40-induced anti-tumor effects are gender-dependent, we administrated RGS + T with or without low-dose aCD40 to male mice bearing 1014 NRASmut melanoma tumors (Supplementary Fig. 6A). When followed after a lead-in of two doses of αPD1 (200 μg/mouse), low-dose 30 μg/mouse aCD40 alone, or RGS + T RAS-targeted therapy alone, resulted in 60% and 75% tumor growth inhibition, respectively, after 3 weeks of treatment. Consistent with our observations in female mice, aCD40 combined with RGS + T in male mice offered a significantly superior ~90% tumor regression, compared to RGS + T (adj.p = 0.04) or aCD40 alone (adj.p = 0.001).
Clinically relevant levels of trametinib are well studied to transiently suppress T cell functions in vitro, but do not result in relevant effects on T cell functions in vivo40. However, the direct effect of RGS on T-cell status and function is unclear. We performed mass spectrometry to determine tumor and plasma levels of RGS in 1014 tumor-bearing mice after three weeks of treatment (Supplementary Fig. 6B). We confirmed that the clinical-grade RGS used in our experiments was 99.96% pure. While effective for tumor growth control (Fig. 3A), the tumor level of RGS was undetectable, and the RGS concentration in plasma was ~0.8 μM in mice. Multiple mechanisms of action of RGS have been reported, including inhibition of microtubules41,42, all reported preclinical studies show it to be a potent inhibitor of tumor growth. Thus, we next compared the effect of 0 ~ 10 μM RGS on mouse splenic T cell status and function with 0 ~ 10 μM vincristine (VCR), an FDA-approved chemotherapy medication used to treat various types of cancer with a defined mechanism of action to inhibit microtubule formation in the mitotic spindle and impede the proper division of cancer cells (Supplementary Fig. 6C, D). We found that 0.1 ~ 10 μM RGS slightly (1.5-fold) promoted T-cell viability compared to untreated (0 μM) T cells, while 0.1 ~ 1 μM of VCR maintained the control level of T-cell viability and 10 μM of VCR reduced T-cell viability by 90% (Supplementary Fig. 6C). Consistently, RGS (0.1 to 10 μM) did not suppress the activation of T-cells (CD69+), while 1 μM and 10 μM of VCR reduced the percentage of activated T-cells by 20% and 95%, respectively (Supplementary Fig. 6D). Lastly, either 1 μM RGS or VCR resulted in a 15% reduction in IFNγ-producing effector T cells, while high-dose 10 μM of RGS and VCR reduced 60% and 90% IFNγ production in effector T cells, respectively. Taken together, the physiological dose of 0.8 μM RGS exhibited a better safety profile compared to VCR, and there was no significant toxicity on T-cell viability, activation, or effector function. These data argue that in these experiments, RGS is acting through a pathway different than VCR, which inhibits microtubule function.
CD11b+ Bregs reduce CD8+ T cells in B16F10 melanoma and promote melanoma lung metastasis and αPD1-resistance
Given the promising therapeutic outcome of combined RAS/MEK/PI3K inhibition with low dose aCD40 against NRASmut melanoma, we next tested whether this regimen was suitable for NRASwtBRAFwt melanoma in preclinical trials (Fig. 5A). We separated the mice into two arms, one with the lead-in of two doses of αPD1 treatment (200 μg/mouse, intraperitoneal) when palpable tumors reached a diameter of 5 mm, and one without αPD1. In each arm, tumor-bearing mice were administrated either vehicle + IgG control (30 μg/mouse, intraperitoneal), RGS (300 mg/kg, 5 days a week, oral gavage) + trametinib (1 mg/kg, 5 days a week, oral gavage) + IgG, vehicle + aCD40 (30 μg/mouse, every 3 days, intraperitoneal), or RGS + T + aCD40. We found that without lead-in αPD1, only the combinatorial treatment of RGS + T + aCD40, but not RGS + T alone or aCD40 alone, resulted in a significant ~80% inhibition on the tumor growth of B16F10 NRASwtBRAFwt melanoma (Fig. 5B). On the other hand, if lead-in αPD1 treatment was used, follow-up treatment of RGS + T, aCD40, or the combinatorial regimens of RGS + T + aCD40, were all capable of inhibiting B16F10 tumor growth at a similar potency of 70 ~ 80% growth inhibition (Fig. 5B). While lead-in αPD1 did not affect the toxicity profile of combinatorial therapy, ~5% transient reduction of mouse body weight occurred in mice treated with the two initial doses of aCD40, which recovered in the second week of treatment (Fig. 5C). These data support the scientific rationale for treating patients with BRAFwt melanoma with first line αPD1 therapy followed by broad spectrum RAS/MEK/PI3K inhibition plus a low-dose CD40 agonist as the second line therapy for αPD1-refractory progressive disease.
A, B Tumor growth, created in BioRender. Yan, C. (2026) https://BioRender.com/83jrya2, exact p values are provided as a Source Data file, and (C) Mouse body weight of B16F10 NRASwtBRAFwt melanoma in C57BL/6 female mice, n = 8 mice per group. Treatment with 200 μg/mouse αPD1 (lead-in 2 doses, every 3 days, intraperitoneal), followed by 30 μg/mouse aCD40 (every 3 days, intraperitoneal), with or without 300 mg/kg rigosertib (RGS, 5 days a week, oral gavage) and 1 mg/kg trametinib (T, 5 days a week, oral gavage), starts at day 8 post tumor cell inoculation. D Tumor samples were collected from each treatment group at day 20 of treatment and immune profiled by FACS analysis. No αPD1-prime groups, Veh + IgG, n = 6, RGS + T + IgG, n = 8, Veh + aCD40, n = 8, RGS + T + aCD40, n = 7 tumors; with αPD1-prime groups, Veh + IgG, n = 5, RGS + T + IgG, n = 5, Veh + aCD40, n = 8, RGS + T + aCD40, n = 8 tumors. Exact p values are provided as a Source Data file. E The baseline frequency of CD11b+ B cells and CD8+ Tc cells in melanoma tumors at 1-month post tumor cell inoculation (1014 tumors, n = 5; B16F10 tumors, n = 3). Exact p values are provided as a Source Data file. F The lung colonization model was established by intravenously injecting 0.5 × 106 metastatic B16F10-Luc2 cells via the tail vein. Splenic B cells were treated with 12.5 μg/ml aCD40 for 2 days then CD11b+ B cells were isolated by magnetic separation for adoptive transfer. Starting at day 2 post tumor cell inoculation, ~2 × 106 CD11b+ B cells were intravenously injected via the tail vein (once a week) in 200 μl HBSS buffer. Treatment of 200 μg/mouse αPD1 (every 3 days, intraperitoneal) starts at day 2 post tumor cell inoculation (n = 5 mice per group). Twenty-one days after tumor implantation, tumor lung was weighed and subtracted from tumor-free lungs in mice. Created in BioRender. Yan, C. (2026) https://BioRender.com/83jrya2. Exact p values are provided as a Source Data file. G Flow cytometry, exact p values are provided as a Source Data file, and (H, I) Immunohistochemistry (IHC) staining of B16F10 melanoma lung metastatic tumors after 21 days of treatment. The percentage of positive red IHC staining cells of Ki67 and CD8 was quantified from 20 fields of each FFPE slide sample per group (n = 5 tumors per group). Exact p values are provided as a Source Data file. Tc, CD8+CD3+ cytotoxic T cells. Th, CD4+CD3+ T helper cells.
Consistent with the immune profile of NRASmut melanoma tumors, we observed that RGS + T + /- aCD40 drug responding B16F10 NRASwtBRAFwt tumors exhibited an increased anti-tumoral response of elevated DCs, macrophage M1/M2 ratio, CD8+ effector T cells, along with a reduction of CD11b+ myeloid cells in the TME (Fig. 5D). Following the lead-in αPD1 therapy, aCD40 monotherapy resulted in a trend toward increased tumor-infiltrating CD11b+ Bregs (adj.p = 0.090), while the addition of RGS + T successfully reduced CD11b+ Bregs to below the control level (adj.p = 0.024). Notably, there was a striking difference in the baseline CD11b+ Breg level between B16F10 and 1014 tumors (Fig. 5E). While CD11b+ Bregs constitute ~6% of all leukocytes in the 1014 tumors, <1% of all leukocytes were CD11b+ Bregs detected in the B16F10 tumors. The low level of baseline CD11b+ Breg in B16F10 tumors was associated with a significantly higher baseline CD8+ T cell content in the B16F10 TME. As a result, the marginal level of endogenous CD11b+ Bregs in B16F10 tumors did not provide enough power to validate the functional impact of CD11b+ Bregs reduction in unleashing T-cell anti-tumor responses in this model. Therefore, we next performed experiments to adoptively transfer aCD40-induced CD11b+ Bregs into C57BL/6 mice bearing B16F10 lung metastases where the B16F10 melanoma cells had been genetically engineered with a luciferase reporter (B16F10-Luc2) (Fig. 5F). Splenic B cells were treated with aCD40 to expand the CD11b+ Breg population and these Bregs were isolated by magnetic separation as described in Methods (Fig. 5F). Subsequently, ~2×106 purified CD11b+ Bregs were injected i.v. into B16F10-Luc2 tumor-bearing mice each week. Twenty-one days after tumor implantation, tumor bearing lungs were removed, weighed, and compared to tumor-free lungs from mice. While αPD1 reduced outgrowth of B16F10-Luc2 melanoma cells in the lung, adoptive cell transfer (ACT) of aCD40-induced CD11b+ Bregs significantly increased the lung tumor mass and partially reversed the therapeutic responses of αPD1 (Fig. 5F). Consistently, flow cytometric analysis of the tumor bearing lungs revealed that ACT of CD11b+ Bregs directly increased CD11b+ Breg content in melanoma lung tumors, while reducing the CD45+ tumor-infiltrating total leukocytes, especially the total and activated CD8+ T cells in the TME (Fig. 5G). Immunohistochemistry (IHC) staining confirmed the reduction of tumor-infiltrating CD8+ T cells (Fig. 5H). Importantly, while αPD1 therapy effectively attenuated tumor cell division based on Ki67+ staining (Fig. 5I), ACT of the CD11b+ Bregs completely blocked the αPD1-triggered enrichment of CD8+ T cells in the melanoma lung metastatic tumors, suggesting a vital role of immunosuppressive CD11b+ Bregs in the development of αPD1-resistance.
CD40 overexpression in melanoma cells reduces CD11b+ Bregs in tumors, improves CD8+ T cell activity, and overcomes αPD1-resistance
To mechanistically explore the role of CD40-induction in melanoma cells and its crosstalk with immune cells in the TME, we developed CD40 overexpression (CD40-OE) clones in two melanoma cell lines, 1014 and B16F10 (Supplementary Fig. 7). The antibiotic combination of blasticidin, hygromycin and puromycin was used for initial clonal selection, proceeded by single colony selection and expansion, and followed by flow-sorting of membrane CD40+ tumor cells, and validation of CD40 protein levels via western blotting and FACS. For the 1014 NRASmut melanoma cell line, we established five CD40-OE clones and nine CD40-Ctrl clones. For B16F10 NRASwtBRAFwt melanoma cell line, 12 CD40-OE clones and 10 CD40-Ctrl clones were selected. Representative clones that had specific induction of CD40 expression and marginal changes in PD-L1 expression were selected for further in vivo tumor model characterization. Comparison of CRISPR/dCas9-based CD40 overexpressing 1014 and B16F10 melanoma cells, to CD40-Ctrl cells with non-specific CRISPR/dCas9 lentiviral particles, resulted in a 46.8-fold and 132-fold overexpression of CD40 protein on the melanoma cell surface (Supplementary Fig. 7A, B). To evaluate the expression of CD40 in these CD40 overexpressing murine melanoma cells with human melanoma cells, we examined CD40 mRNA expression in human melanoma cell lines and saw that human melanoma cell lines exhibited >100-fold higher CD40 mRNA expression compared to murine melanoma cell lines (Supplementary Fig. 7C). However, only 10% of melanoma cells in patient melanoma tumors were CD40 positive based upon scRNA-Seq analysis, indicating a small population of human melanoma cells express high levels of CD4024. Our constructed CD40-OE melanoma cell monoclones, therefore, mimic the physiological expression level of CD40 in 10% of melanoma cells in human lesions.
To further evaluate the effects of continuous expression of higher levels of CD40 on melanoma cells, we inoculated 3x105 cells from selected CD40-Ctrl or CD40-OE 1014 NRASmut melanoma cell clones into female C57BL/6 mice (Fig. 6). For each melanoma cell clone, mice were separated into five treatment groups with five mice per treatment group, including (1) Vehicle (Veh) + IgG; (2) Veh + aCD40 (30 μg); (3) Veh + αPD1 (200 μg); (4) aCD40 + αPD1; (5) RGS (300 mg/kg) + IgG. We observed that compared to CD40-Ctrl tumors, 1014 CD40-OE melanoma tumors exhibited reduced tumor growth in vivo, and melanoma-cell-selective CD40 overexpression reversed αPD1 resistance (adj.p = 0.025). When CD40-OE tumors were treated with aCD40 + αPD1, tumors failed to grow over the 22-day time frame of the assay (adj.p < 0.001) (Fig. 6A). Treatment effect analyses revealed a marked sensitization role of melanoma-cell-selective induction of CD40 to improve response to αPD1 alone (adj.p = 0.07) and aCD40 + αPD1 (adj.p < 0.001) (Fig. 6B). RGS treatment of CD40-OE melanoma tumors resulted in unaltered tumor growth rate over vehicle control, compared to CD40-Ctrl tumors where RGS significantly inhibited tumor growth. These data suggest that a key role of RGS treatment in melanoma tumors is to reverse RAS-mediated suppression of CD40 expression, such that if CD40 expression is already high, RGS treatment is less effective. Detailed immune profiling of the TME in these experiments demonstrated that treatment-responding tumors exhibited a significantly higher immune cell infiltration, including CD8+ T cells, along with a reduction of CD11b+ Bregs (Fig. 6C). Importantly, the intratumoral frequencies of CD11b+ Bregs and CD8+ T cells were significantly and negatively correlated with each other (Fig. 6D), confirming the defined immunosuppressive role of CD11b+ Bregs on inhibiting the CD8+ T effector cells in melanoma tumors (Figs. 4, 5). Comparable results were observed in the B16 F10 NRASwtBRAFwt melanoma model with a partial response to αPD1 +/- aCD40. While aCD40 (adj.p = 0.03), αPD1 (adj.p = 0.025), or aCD40 + αPD1 (adj.p = 0.001) delayed the growth of CD40-Ctrl B16F10 tumors, completely halted tumor growth was only observed in CD40-OE B16F10 tumors receiving aCD40 + αPD1 treatment for 18 days (adj.p < 0.001) (Fig. 6E). Consistent with the results in 1014 melanoma, results from the B16F10 model confirmed a reduced response to RGS therapy (adj.p = 0.002) in CD40-OE tumors compared to the CD40-Ctrl tumors (Fig. 6F), highlighting CD40 induction in melanoma cells is the driving mechanism for RGS therapy. In addition, we consistently observed the increase of total CD45+ leukocyte enrichment in the TME of drug-responding B16F10 tumors, including total and activated CD8+ T cells (Fig. 6G). The percentage of tumor-infiltrating CD8+ T cells was significantly and negatively correlated with the B16F10 tumor burden (Spearman p = 0.0003, r = −0.5399) (Fig. 6H), suggesting that CD8+ T cell responses triggered by RAS-pathway inhibition and CD40 agonism play a key role in melanoma tumor control. However, due to the limited amount of endogenous CD11b+ Bregs in B16F10 tumors (Fig. 5E), this therapeutic model of B16F10 melanoma did not provide us enough power to validate the causal relationship between reduced immunosuppressive CD11b+ Bregs and elevated CD8+ T cell responses, as revealed in the ACT model of B16F10 melanoma (Fig. 5F–I).
Treatment with 200 μg/mouse αPD1 (every 3 days, intraperitoneal), 30 μg/mouse aCD40 (every 3 days, intraperitoneal), or 300 mg/kg rigosertib (RGS, 5 days a week, oral gavage), starts at day 8 post tumor cell inoculation, n = 5 mice per group. A Tumor volume, created in BioRender. Yan, C. (2026) https://BioRender.com/83jrya2, exact p values are provided as a Source Data file, and (B) Treatment effect on immunotherapy-resistant NRASmut 1014 melanoma growth in C57BL/6 female mice. Exact p values are provided as a Source Data file. C, D Flow cytometric analysis of the 1014 tumors at day 22 post-treatment. Exact p values are provided as a Source Data file. E Tumor volume, created in BioRender. Yan, C. (2026) https://BioRender.com/83jrya2, exact p values are provided as a Source Data file, and F Treatment effect on NRASwtBRAFwt B16F10 melanoma growth in C57BL/6 mice, n = 5 mice per group. Exact p values are provided as a Source Data file. G Flow cytometric analysis of the B16F10 tumors at day 18 post-treatment. n = 5 mice per group. Tc, CD8+CD3+ cytotoxic T cells. Exact p values are provided as a Source Data file. H Correlation analysis between CD8+CD3+ Tc cell frequency in total CD45+ cells in B16F10 tumors and endpoint tumor weight at day 18 post-treatment.
To provide a more comprehensive assessment of drug responses in CD40-OE vs CD40-Ctrl melanoma tumors, we performed bulk RNA and T-cell receptor (TCR) sequencing analysis on the 1014 tumors, including tumor cells, stromal cells, and immune cells (Fig. 7A, Supplementary Table 2 and Supplementary Fig. 8). CIBERSORT (LM22) deconvolution analysis suggested the percentage of CD8+ T cells significantly increased in CD40-OE tumors in response to treatment with αPD1 +/- aCD40 (adj.p = 0.0008) (Fig. 7B), confirming our flow cytometric results (Fig. 7C). Gene enrichment analysis of 1014 CD40-OE tumor transcriptome revealed upregulated gene signature hallmarks of IL-2/STAT5 signaling, inflammatory responses, JAK/STAT3 signaling and IFNα/γ responses, while a reduction in gene signature hallmarks of hypoxia, oxidative phosphorylation and Myc targets, compared to the CD40-Ctrl tumors at baseline (Fig. 7C) or CD40-Ctrl tumors treated with aCD40 + αPD1 (Fig. 7D). Sequencing of the TCR-β chain further confirmed an increase in the total count of TCR clonotypes in CD40-OE tumors at baseline and with the treatment of αPD1 +/- aCD40 (Fig. 7E, F). For example, the drug effect of Veh + αPD1 in CD40-OE is 6.38 times greater than the drug effect of Veh + αPD1 in CD40-Ctrl (p < 0.001) (Fig. 7G and Supplementary Table 2). Similarly, the drug effect of aCD40 + αPD1 in CD40-OE is 4.86 times greater than the drug effect of aCD40 + αPD1 in CD40-Ctrl (p = 0.005). Notably, there were no significant differences between the drug effect of RGS + IgG in CD40-OE and CD40-Ctrl, confirming the RGS induction of CD40 in melanoma cells is responsible for CD40-dependent tumor cell killing observed. As expected, distribution analysis of TCR-β clonotype abundances revealed that lower abundant (1 ~ 10 copies) clonotypes exhibited a higher number in quantity in drug-responding tumors (Fig. 7H), suggesting there was a strong polyclonal expansion of T-cell clonotypes. CDR3 amino acid sequences and V genes are shown in Fig. 7I and Supplementary Fig. 8A, where the TRBV26 gene exhibited the highest interaction with treatment response. Importantly, the relative abundance of TCR-β clonotypes with small/medium-expansion exhibited a biased expansion in response to the drug treatment in CD40-OE tumors (adj.p = 0.019), while hyper-expanded TCR clonotypes were not associated with beneficial therapeutic outcomes (Fig. 7J). Indeed, clonotype tracking of the top 20 small/medium-expanded clonotypes in the TRBV26 gene further confirmed the treatment-triggered increase of polyclonal TCR expansion (Fig. 7K).
A Correlation heatmap of top 25% differentially altered genes summarized across all treatment groups (15,903 genes plotted). Heatmap3 was used for cluster analysis and visualization. Significantly differential expressed genes with absolute fold change ≥ 2 and FDR adjusted p value ≤ 0.05 were detected by DESeq2. B CIBERSORT analysis of a leukocyte signature matrix (LM22) to deconvolution of 1014 tumors. Exact p values are provided as a Source Data file. C, D Hallmark Gene Set Enrichment Analysis (GSEA) of the transcriptomic profile of 1014 tumors at baseline or with agonist CD40 (aCD40) plus αPD1 treatment at 22 days using GSEA package. E–K TCR-β chain sequencing results were evaluated using the Archer Immunoverse analyzer. CDR3 sequences and frequency tables were extracted from the manufacturers’ analysis platform and imported for analysis using the Immunarch package (https://immunarch.com) in R. Comparisons were made within specific tumor types and across different tumor types (CD40-Ctrl vs CD40-OE) to evaluate drug effects, with the analysis conducted on a natural logarithmic scale. Number of (E) total and (F) unique TCR clonotypes. Exact p values are provided as a Source Data file. G Treatment effect (ratio of TCR clonotypes among treatment to clonotypes among Veh + IgG control) comparisons between CD40-Ctrl and CD40-OE tumors were performed using multiple comparisons test for generalized linear hypothesis test based on the negative Binomial generalized linear model. All graphs show Mean ± SD. H Estimated distribution of clonotype abundances. I Table summary of FDR (False Discovery Rate) adjustment. J Estimated relative abundance (also known as clonal space homeostasis), which characterizes the proportion of repertoire occupied by clonal groups with specific abundances indicated. K The top 20 most abundant TCRs of small- and medium-expanded clonotypes in TRBV26 are shown. Flow between samples indicates shared TCRs. Boxed CDR3s indicate most clonal TCRs in TRBV26. The Holm correction was used to adjust the p value for multiple comparisons. A–K Pooled values of n = 3 tumors per group.
Rigosertib plus CD40 agonism attenuates the growth of patient-derived organoids and the identification of protumor CD11b+PD-L1+ Bregs in patient melanoma tumors
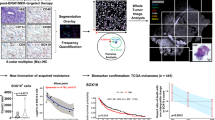
Testing potential therapeutics on patient biopsy samples ex vivo is an attractive approach to making treatment decisions. To determine whether our findings regarding the therapeutic benefit of combined RAS-inhibition and CD40 agonism observed in mouse ICB-resistant melanoma models are recapitulated in human melanoma, we have developed a 3D co-cultured model of patient-derived organoids (PDOs) with immune cells from cancer patients43. The viability of melanoma cells and T cells in the fine-needle aspiration (FNA)-PDOs were assessed via microscopy imaging and flow cytometry following drug treatment for 10 days (Supplementary Fig. 9). We observed that ~90% of CD45- melanoma cells in the FNA-PDO cultures are viable cells. There is no difference in the viability of PDO samples derived from primary verses the lymph node tissues. In 3 of 4 PDOs developed in our lab, RGS + aCD40 + αPD1 resulted in a clear inhibition of αPD1-refractory tumor growth (Fig. 8A). We found that PDO-3529 derived from the primary melanoma tumor, but not one from a melanoma-involved lymph node, was sensitive to RGS + aCD40 + αPD1, suggesting the tissue-specific TME may dictate the action and outcome of the combinatorial treatment. Notably, the only non-responding melanoma organoid, PDO-3101, was derived from a patient with a history of renal transplant and chronic immunosuppression (Fig. 8A). Together, our data support evidence for efficacy RAS-pathway inhibition plus low-dose aCD40 in human melanoma.
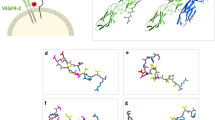
A PDO cultures were developed using previously frozen patient melanoma tissue that was thawed and immediately subjected to fine needle aspiration. IL-2 was added to all cultures to ensure the survival and proliferation of T cells. PDOs were grown in organoid media containing 5% Matrigel. After allowing PDOs to establish for 3–4 days, treatments were initiated as follows: RGS, CD40 agonist (aCD40), αPD1, RGS + aCD40, RGS + αPD1, or RGS + aCD40 + αPD1. Cultures were allowed to grow for 10 days, and the organoid area was assessed and quantified at day 10, PDO-2624, n = 18 per group; PDO-3453A, n = 18 per group; PDO-3529 primary, n = 12 per group; PDO-3529LN, n = 12 per group; PDO-3101, n = 12 per group. All graphs show Min to Max with all points +/− SD. Exact p values are provided as a Source Data file. B, C, F Melanoma scRNA-Seq dataset28. Analysis of malignant and immune cell gene expression of 6173 scRNA-seq profiles from 32 human melanoma tumors. D, E, G Melanoma scRNA-Seq dataset29. Transcriptome analysis of 16,291 individual immune cells from 49 tumor samples of melanoma patients treated with checkpoint inhibitors. C, E Functional enrichment of Geneset and KEGG pathway over-representation analysis were performed on differentially expressed genes of CD20+CD11b+PD-L1+ Bregs in patient melanoma tumors using WebGestaltR package (NULL). F, G Patient melanoma scRNA-Seq analysis reveals a distinct transcriptome profile of CD20+CD11b+PD-L1+ Bregs. All graphs show Median ± SD. H The Breg gene signature (Sig. Bregs) is associated with poorer patient OS in multiple cancer types. Kaplan-Meier plots show OS for patients in the TCGA pan-cancer datasets44. Patients were split into high and low expression of the Breg gene curated signature. Median OS survival is represented on the graph when available. P-values were calculated by log-rank test. GBM, glioblastoma; LGG, low-grade glioma; LUSC, lung squamous cell carcinoma; STAD, stomach adenocarcinoma; MESO, mesothelioma; OV, ovarian carcinoma.
Our data highlight the functional importance of CD11b+PD-L1+ Bregs in driving aPD1 resistance in murine models of melanoma. To further analyze the importance of these Bregs in human melanoma, we analyzed scRNA-Seq data from two publicly available melanoma datasets28,29 (Fig. 8B–E). The human melanoma dataset #1 covers malignant and immune cell gene expression of 6,173 scRNA-seq profiles from 32 human melanoma tumors28. The human melanoma dataset #2 compiles transcriptome analysis of 16,291 individual immune cells, without malignant cell profiling, from 49 tumor samples of melanoma patients treated with checkpoint inhibitors29. While the mean expression of CD40 mRNA in all cells in the TME significantly correlates with the proportion of CD11b+PD-L1+ Bregs in the melanoma tumors (dataset #1, p = 0.002, r = 0.535; dataset #2, p = 0.000, r = 0.557), melanoma-cell-specific CD40 level exhibited a negative correlation with the proportion of CD11b+PD-L1+ Bregs in the TME (dataset #1, p = 0.078, r = −0.486) (Fig. 8B, D). In addition, the level of CD11b+PD-L1+ Bregs is negatively correlated with the mRNA of CD8A (dataset #1, p = 0.158, r = −0.256; dataset #2, P = 0.000, r = −0.555) and IFNG (dataset #1, p = 0.08, r = −0.314) in the tumors, confirming an immune suppressive function of these Breg cells in patient melanoma tumors. Functional enrichment of Geneset and KEGG pathway over-representation analysis was performed on the differentially expressed genes of CD11b+PD-L1+ Bregs in patient melanoma tumors in reference to CD11b-PD-L1- B cells using WebGestaltR package (Fig. 8C, E). Consistently, the transcriptome profile of CD11b+PD-L1+ Bregs in patient melanoma tumors resembles gene signatures of “negative regulation of immune system process” (Geneset FDR ≤ 0.05), “neutrophil mediated immunity” (Geneset FDR ≤ 0.05), “negative regulation of cell activation” (Geneset FDR ≤ 0.05), “primary immunodeficiency” (KEGG pathway p = 0.0009), and “negative regulation of cell activation” (KEGG pathway p = 0.0007). Importantly, we identified a 4-gene signature of CD11b+PD-L1+ Bregs (Sig.Bregs) in human melanoma tumors, including ITGAM (CD11b), CD274 (PD-L1), TNFRSF13B (TNF Receptor Superfamily Member 13B) and AHNAK, which were simultaneously expressed only in these Breg cells, and not in any other cellular populations in the TME (Fig. 8F, G and Supplementary Fig. 10A, B). We next probed this 4-gene Sig.Bregs in the pan-cancer TCGA dataset44 (Fig. 8H). The patients with a high level of baseline Sig.Bregs exhibited significantly worse overall survival (OS) in 6 of 12 screened cancer types, including glioblastoma (GBM, p = 0.013), low-grade glioma (LGG p = 0.00075), lung squamous cell carcinoma (LUSC, p = 0.095), stomach adenocarcinoma (STAD, p = 0.046), mesothelioma (MESO, p = 0.0053) and ovarian carcinoma (OV, p = 0.02). Notably, while a worse prognosis of Sig.Bregs was identified, the frequency of high Sig.Bregs was not altered during the cancer disease progression, suggesting CD11b+PD-L1+ Breg cells may serve as an intrinsic immunosuppressive function for primary resistance to ICB (Supplementary Fig. 10C).
Finally, we quantified the frequency of tumor-infiltrating CD11b+ B cells across cancer types using The Single Cell Portal (singlecell.broadinstitute.org) (Supplementary Fig. 11). The keywords “cancer” or “tumor” identified 101 studies. We further filtered the studies with available gene expression profiles of ITGAM (CD11b) and MS4A1 (CD20) or CD19, which identified 10 studies, including three melanoma cohorts (28,111 cells from 98 patient tumors), 1 thymoma cohort (64,649 cells from four patient tumors), 3 colon cohorts (48,115 cells from normal colon, 371,223 cells from 62 patient tumors, and 26,556 cells from 33 small intestinal biopsies from 11 patients with environmental enteropathy [EE]), one prostate cancer cohort (2,170 cells from 14 patient tumors), one glioma cohort (201,986 cells from 18 patient tumors), and one breast cancer cohort (130,246 cells from 26 patient primary tumors). The baseline frequency of CD11b+ B cells among total TIB cells is low in prostate cancer (1.06%), breast cancer (0.16 ~ 2.71%), normal colon (0.36%) and colon cancer (0.92%), compared to melanoma (5.70 ~ 9.29%) and glioma (7.22%). The scRNA-Seq profile of breast and thymoma datasets also revealed that there is a 10 ~ 15-fold enrichment of CD11b+ B cells in the memory B cells compared to the naive B cell counterparts (Supplementary Fig. 11). Lastly, there is a clear increase of CD11b+ B cells in the colon of EE (8.86%) compared to the normal colon (0.36%), which is associated with elevated pro-inflammatory cytokine expression in the EE cohort. One possible scenario is that our host immune system may utilize CD11b+ B cells to counteract or restrain over-whelming inflammation. Indeed, intestinal CD11b+ B cells were reported to ameliorate colitis in an IgA-dependent manner45.
Altogether, our work demonstrated that broad-spectrum RAS-pathway inhibition, via combining RGS + MEK inhibitor, for improved CD40 induction and immunogenic cell death selectively in melanoma cells, plus a low, non-toxic dose of aCD40, increased CD8+ T cell responses while minimizing aCD40-induced CD11b+ Bregs and thus inhibiting αPD1-resistant melanoma tumor growth. This combination may provide an effective second line of therapy for ICB-resistant BRAFwt (NRASmut and NRASwt) melanoma. Given the identification of CD11b+ Bregs across an extremely broad spectrum of human cancer types and inflammatory disease conditions, it is worth considering the possibility that CD11b+ Bregs may actively restrain immune surveillance and effector function in multiple settings.
Discussion
Combining targeted therapy with immune checkpoint blockade is emerging as a promising approach for patients with ICB resistance, although the most effective regimens have not been well defined. We have demonstrated that treatment with RGS or trametinib induces the expression of CD40 in melanoma cells and RGS inhibition of tumor growth depends on the expression of CD4024. Our prior work also showed that low levels of CD40 expression in patients with RASmut melanoma, are associated with poor survival and a poor response to ICB therapy25, highlighting the potential for delivery of therapies that induce CD40 expression in αPD1-resistant NRASmut melanoma. Our single-cell (sc)RNA-Seq analysis of melanoma patient tumors (n = 1883 melanoma cells from 14 patients) revealed that while 80% of human melanoma tumors express CD40, only 10% of melanoma cells in the tumors are CD40 positive24.
We demonstrated here that the systemic treatment of aCD40 in preclinical mouse models, while effective in suppressing ICB-refractory melanoma tumor growth, simultaneously increases a CD11b+PD-L1hi Breg population in the TME that is associated with residual disease and therapy resistance (Fig. 2A–G). While the in vitro splenocyte killing assay demonstrated the inhibitory role of CD11b+ Bregs on the direct killing activity of CD8+ T cells (Fig. 2G), CD8+ T cells in the TME can also provide indirect supportive roles, such as IFNγ production in stimulating the innate anti-tumor immune response46, and maintain killing of MHC-I-negative tumor cells through the NKG2D–NKG2DL axis47. Regulatory B cell development and function in response to aCD40 may offer valuable insights into cellular-driven immune modulation and response to immunotherapy in cancer. However, Bregs may potentially arise from every stage of B cell differentiation, and definitive markers of the lineage and function of Bregs (e.g., IL-10, IL-35, IL-27, CD1d, PD-L1) remain elusive and content-dependent30,48. Targeting pro-tumoral Bregs could enhance anti-tumor immunity whilst leveraging anti-tumor B cells to promote the formation and functionality of tertiary lymphoid structures23, consequently bolster immune responses against cancer. Our results reveal the heterogeneity of aCD40-induced CD11b+ Breg subsets. In our attempt to compare aCD40-induced CD11b+ Bregs with the conventional Breg markers in cancer, such as IL-1049 and PD-L124,26, we found that aCD40 induces approximately 40% splenic B cells to differentiate into PD-L1+ Bregs, among which ~25% are also IL-10-producing B10 cells in vitro (Fig. 4A and Supplementary Fig. 2). The aCD40-treated melanoma TME enriches CD11b+ Bregs to ~80% of total TIB in vivo, among which about 87% are PD-L1+ (Fig. 2F) and 10% are IL-10+ B10 cells (Supplementary Fig. 2). In patients with melanoma, the baseline frequency and immunosuppressive potency of PD-L1+ Bregs increased significantly during disease progression (e.g., from 2% in stage I to 10% in stage IV)26. These reported results of PD-L1+ Bregs were consistent with our scRNA-Seq data analysis on the frequency of PD-L1+CD11b+ Bregs in patients with melanoma, representing ~12% at baseline in total TIB (Fig. 1B–D). While we identified and characterized the immunosuppressive function of aCD40-induced CD11b+ Bregs in preclinical models of melanoma growth and lung metastasis (Figs. 1–5), the driver genes responsible for the development and function of aCD40-induced CD11b+ Breg in human melanoma remain unresolved, and the roles of CD11b+ Bregs should be explored beyond melanoma.
Compared to the CD11b- B cell counterparts, CD11b+ Bregs exhibited a higher level of intrinsic and inducible CD40 expression (Figs. 1E, F, 2A, B). The CD11b+ Bregs exhibited heightened endogenous levels of RAS-pathway activation (total-ERK and total-AKT) (Fig. 2H), making them vulnerable to RAS-targeted therapies. While CD11b+ Bregs exhibited a prominent level of PD-L1, the treatment of αPD1 alone failed to rescue CD11b+ Breg-suppressed CD8+ T cell response in the lung metastatic tumor model of melanoma (Fig. 5). Importantly, exposing aCD40-induced CD11b+ Bregs to RAS/MEK/PI3K-pathway inhibition directly reduced Breg cell proliferation and restored the activation and IFNγ production in effector T cells (Fig. 4A, B), opening the door for rational combinations of aCD40 plus RAS/MEK/PI3K-targeted inhibition to enhance therapeutic specificity and durability. We demonstrated that induction of CD40 on melanoma cells via (1) therapeutic intervention of RAS/MEK/PI3K-targeted inhibition (Figs. 3–5) or (2) CRISPR-Cas9-based CD40 overexpression (Figs. 6–7) overcomes tumor resistance to αPD1 of BRAFwtNRASwt and BRAFwtNRASmut melanoma. As expected for the driver oncogenic pathway, we found that near-complete and durable inhibition of RAS signaling via combined rigosertib (inhibit RAS/PI3K/AKT) and trametinib (inhibit RAS/RAF/MEK) would maximize selective induction of CD40 in melanoma cells, as well as inhibition of CD11b+ Breg and melanoma tumor growth, in response to aCD40 plus RAS/MEK/PI3K-targeted therapy. Such observations align with previous reports on the trade-off tolerogenic response of PD-L1 induction in cancer cells triggered by pro-inflammatory cytokines, such as IFNγ50, tumor necrosis factor (TNF)α51, and interleukin(IL)-652, which attenuate the activation of effector T cells and anti-tumor macrophages. Similarly, IL-12 triggers compensatory induction of secondary immunosuppressive signals, including NKG2A, on human CD8+ T cells53. Tumors appear to hijack signals induced by immune-stimulatory cytokines and/or proinflammatory mediators and use these signals to enhance immune checkpoint expression, contributing to tumor adaptive resistance pathways.
While promising in principle, CD40 agonists have been associated with different levels of toxicities both in the clinic and animal models [reviewed in ref. 6]. Our results in the 1014 BRAFwtNRASmut and B16F10 BRAFwtNRASwt melanoma models confirmed the feasibility and safety profile to combine a low-dose 30 μg aCD40 (twice a week) plus 300 mg/kg RGS (QD, 5 days/week) +/− 1 mg/kg trametinib (QD, 5 days/week), with a transient loss of 5 ~ 10% mouse body weight in the first week of aCD40 treatment (Figs. 4D and 5C). No elevation of liver/kidney damage was observed in mice post 3-weeks of therapy (Fig. 4E), suggesting reversible and non-cumulative kinetics of the adverse effects of the combinational treatment. While dosages of CD40 agonists selected for clinical trials are most frequently at 0.01 ~ 0.3 mg/kg (vary from 0.01 to 3.6 mg/kg) depending on the CD40 agonist used, preclinical mouse studies using CD40 rat anti-mouse IgG2a mAb (clone FGK45) have commonly used 1.5 ~ 10 mg/kg (30 ~ 200 μg/dose), with 5 mg/kg (100 μg/dose) being the most frequently employed dosage across treatment groups6. The minimum dosage of 1.5 mg/kg in mice is the exact dosage used in our study (30 μg/dose), equivalent to 0.01 mg/kg in humans after body surface area and metabolic normalization54, validating the safety of initial dosages used in current clinical studies. The current approaches to decrease the aCD40 toxicity include intratumoral delivery of Fc-optimized agonistic CD40 antibodies in mice55 and patients56 with metastatic cancer (melanoma, breast cancer, and renal cell carcinoma) and nanoparticle-based hydrogel-dependent delivery of agonistic CD40 antibody in preclinical mouse melanoma models57. Given the quick (within 1 week) recovery of mouse body weight of our low dose 30 μg aCD40 delivery and the safety profile of RAS-pathway inhibition by RGS58,59 and trametinib60 in the clinical studies, we predicted that the side effects of RAS-pathway inhibition plus a low dose of aCD40 might be transient and manageable in the clinic.
Our TCR-Seq results of the TME support the existence of fewer than expected tumor antigens that elicit the response of a restricted set of identifiable tumor antigen-specific tumor-infiltrating lymphocytes (TILs) (Fig. 7E–K). Though the low/medium number of therapy-induced TCR clonotypes might be unexpected, the responses of these TILs to tumor antigens were highly robust. Our finding that only a few tumor antigen-specific TILs and TCR clonotypes mediate tumor rejection corroborates prior studies61,62,63,64 that have demonstrated and clinically validated the number of reactive neoantigens is about <5 per melanoma tumor, despite the tumors harboring hundreds or thousands of somatic mutations. Our work highlights that while only a few tumor-targeting T-cell clones are invigorated in a tumor, targeting them by immunotherapeutic interventions may be sufficient to mount tumor rejection. Our findings and the insight gathered through others strengthen the notion that identifying a few targetable tumor antigens and their corresponding TIL clones may guide personalized cancer immunotherapy.
Based on recent preclinical65,66 and clinical studies10 in melanoma, sequential treatment with αPD1 + /- αCTLA4 followed by targeted therapy, but not the other way around, may maximize the synergistic benefit between targeted inhibition and immunotherapy14. In the circumstance without lead-in αPD1, adding a low dose of aCD40 to RAS/MEK/PI3K-targeted therapy suppressed αPD1-resistant tumor growth of both NRASmut and NRASwt melanoma (Supplementary Fig. 5). While aCD40-induced CD11b+ Bregs attenuate treatment responses of αPD1-resistant melanoma, adding RAS-pathway inhibition reduced CD11b+ Bregs, unleashing the activity of T effector cells to kill tumor cells, which resulted in durable tumor regression without the reinstatement of αPD1 (Supplementary Fig. 5). Therefore, we posit that broad-spectrum inhibition of the RAS pathway plus low-dose CD40 agonism has the potential to serve as an effective second line of therapy for αPD1-resistant BRAFwtNRASmut and BRAFwtNRASwt melanoma. In support of this posit, treatment of melanoma patient-derived organoids with RGS + aCD40 + αPD1 resulted in an additive therapeutic inhibition of the growth compared to the monotherapies (Fig. 8A). Recently, a new RAS-ON inhibitor, RMC-6236, was developed that inhibits all canonical RAS isoforms, resulting in the inhibition of both the RAF-MAPK and PI3K-AKT pathways. Data from preclinical and an initial patient case study supports the use of RAS-pathway inhibitor in a variety of RAS-addicted tumors as well as in combination with ICB67. There is a need for both comprehensive drug de-escalating preclinical and clinical studies to understand the safety profile of the promising therapy fully. Additionally, combining the new RAS-ON inhibitor of both mutant and wild-type RAS, RMC-623667, with low-dose aCD40 in preclinical studies will be important.
Our previous pan-cancer transcriptome analysis revealed that CD40 was downregulated in 10 different solid tumor types out of 23 screened cancer types as compared to corresponding normal tissues, including skin, breast, lung, colon, brain, ovary, esophagus, prostate, uterus and endometrium25. Consistent with these data, pan-cancer analysis using our defined 4-gene signature (Itgam, Cd274, Tnfrsf13b, and Ahnak) of CD11b+ Bregs indicated a significant association of high Breg signature and poor survival of patients with glioblastoma, low-grade glioma, lung squamous cell carcinoma, stomach adenocarcinoma, mesothelioma, and ovarian carcinoma (Fig. 8H). Our scRNA-Seq analysis of breast and thymoma datasets revealed a 10 ~ 15-fold enrichment of CD11b+ B cells in the memory B cells compared to the naive B cell counterparts (Supplementary Fig. 11). Likewise, Tnfrsf13b and Ahnak are expressed in a subset of B cells producing auto-antibodies. One limitation of gene signature analysis in human databases is the possible involvement of other B cells that are not immunoregulatory, which warrants future functional validation. Overall, these data indicate that treating these tumors with a combination of aCD40 and RAS/MEK/PI3K pathway inhibitors could maximize cell death and minimize the CD11b+ Breg-dependent tolerogenic response.
We are currently conducting a Phase IIb trial treating unresectable/metastatic αPD1-refractory melanoma with RGS plus pembrolizumab in (NCT05764395). The study reported here provides preclinical proof-of-concept studies for future clinical trials that offer an improved treatment plan that combines the specificity and immunogenicity of cancer cell-targeted therapy (e.g., RAS/MEK/PI3K-pathway inhibition and immunotherapy, e.g., CD40-based agonism) to orchestrate efficient, tolerable and durable therapeutic options for patients with BRAFwtNRASwt and BRAFwtNRASmut melanoma that exhibit innate or acquired resistance to ICB therapy.
Methods
Mouse melanoma tumor models
Animal studies were approved by the Vanderbilt Institutional Care and Animal Use Committee (IACUC; protocol number V1700194) and were performed in accordance with Vanderbilt IACUC guidelines. All animals were housed under pathogen-free conditions at the Vanderbilt Animal Care Facility. Three mouse melanoma tumor models were used in the study, including BRAFwtNRASmut 1014, BRAFwtNRASwt B16F10 (Cat#: CCL-6475, ATCC, RRID:CVCL_0159), and B16F10-Luc2 cells (Cat#: CRL-6475-Luc2, ATCC, RRID:CVCL_A4CJ). Cells were cultured in DMEM/F12 medium (Gibco, #11330-032) containing 10% FBS (Sigma, PAA, A11-201). Cell cultures were tested monthly for mycoplasma (e-Myco TM Plus, LiliF Diagnostics), and only mycoplasma-free cells were used for experiments outlined in these studies.
Tumor allografts were established in 8-10-week-old female and male C57BL/6 mice that were purchased from Jackson Labs. Mice were maintained under a 12:12 h light–dark cycle, with lights on at 6:00 a.m. and off at 6:00 p.m. The housing environment was controlled at a relative humidity of 50% and an ambient temperature of 20~22 °C. For the in vivo orthotopic melanoma model, mice received 3 × 105 of tumor cells in 100 µl of serum-free DMEM medium by subcutaneous injection in the lower back. Mouse body weight was assessed once a week, and tumor measurements were taken twice a week with micro-calipers. Tumor volume was estimated as 0.5*length*width*width. Treatment began when tumors reached ~100 mm3 volume on average or at day 8 ~ 10 post tumor cell inoculation and continued until tumors in the experiment exceeded 20 mm in diameter if one tumor was inoculated per mouse; if two tumors were inoculated per mouse, treatment continued until the tumors exceeded 15 mm in diameter, or any tumors became perforated. Percent (%) inhibition of tumor growth = (the tumor volume of IgG control-treated tumors ‒ the average tumor volume of aCD40-treated tumors) divided by the average tumor volume of IgG control-treated tumors x 100%. To establish a lung colonization model, the B16F10-Luc2 cells (Cat#: CRL-6475-Luc2, ATCC, RRID:CVCL_A4CJ), derived from C57BL/6 mice, were i.v. injected into 8-10-week-old female C57BL/6 mice. Three weeks after injection, lungs were harvested, and lung weight was determined and normalized to tumor-free lung. For depletion of CD19+ or CD8+ cells, mice were injected intra-peritoneally with 200 μg of anti-mouse CD19 (Purified in vivo GOLD Functional Grade, clone: 1D3, CAT#: C2117), or anti-mouse CD8α (Purified in vivo GOLD Functional Grade, clone: YTS 169.4, CAT#: C2442), respectively. The same amount of rat IgG2b control (Purified in vivo GOLD Functional Grade, clone: 1-2, CAT#: I-1034) or rat IgG2a control (Purified in vivo GOLD Functional Grade, clone: 1-1, CAT#: I-1177) antibodies were used for CD19 or CD8 depletion controls, respectively. Antibodies were administrated daily for three days prior to the tumor cell injection, and every three days thereafter for the entire experiment. All antibodies were purchased from Leinco Technologies.
Flow cytometry staining and analysis
Tumor tissues were minced on a programmable gentleMACS dissociator (Miltenyi Biotec) and digested with an enzyme solution of collagenase I (1500 CDU, Worthington Biochemical Corp.), Dispase II (1 mg/mL, Worthington Biochemical Corp.) and DNase I (0.1 mg/mL, Worthington Biochemical Corp.). Digested tumors were passed through a 70-μm strainer to obtain a single-cell suspension. Minced samples of mouse spleens, lungs, and lymphoid nodes (LN) were pressed through a 70-μm strainer using a syringe plunger to obtain a single-cell suspension. Erythrocytes were lysed by incubation in an ammonium-chloride-potassium (ACK) buffer for 5 min at room temperature. The detail of antibodies used (Supplementary Table 1), staining, and flow cytometry analysis protocols are according to our previously published methodology24,68. Briefly, cells were incubated with Ghost Dye Violet 450/510/780 (Tonbo Biosciences), an amine-reactive viability dye used to discriminate live/dead cells and washed with FACS buffer (PBS containing 1% v/v FBS). After blocking Fc receptors with anti-mouse CD16/CD32 mAb (BD Biosciences Cat:553142, RRID:AB_394657) in FACS buffer for 20 min, cells were incubated with target antibodies. After staining, cells were washed twice in FACS buffer and fixed with 300 μl/tube of fixation buffer (1% formalin in PBS). For intracellular staining after surface staining, cells were fixing/permeabilizing using Transcription Factor Buffer Set (Cat:562674, BD Pharmingen) per manufacture’s protocol. Data were collected using a 5-laser BD LSR Fortessa flow cytometer and analyzed using FlowJo software (RRID:SCR_008520, Version 10.9.0). For t-SNE analysis, live CD45+ leukocytes were concatenated after downsampling to ~25,000 events for subsequent high-dimensional data analysis to normalize the contribution among samples under different treatments. Samples were then analyzed in parallel by t-SNE and manually gated leukocyte populations were overlaid onto the total t-SNE map using FlowJo 10.9.0 (RRID:SCR_008520). FACS sequential gating strategies were included in Supplementary Fig. 12.
For in vitro T cell function assays, 0 ~ 10 μM rigosertib (Onconova Therapeutics) and vincristine (VCR, MedChemExpress, Cat#: HY-N0488) were examined. Splenic T cells were stimulated for 24 h with a Cell Stimulation Cocktail (Cytek Biosciences, Cat#: TNB-4975) composed of the phorbol ester, PMA (Phorbol 12-Myristate 13-Acetate), a calcium ionophore (Ionomycin), and the protein transport inhibitors Brefeldin A and Monensin. T-cell viability (Ghost Dye), activation (CD69) and IFNγ production were determined using intracellular staining by flow cytometry and analyzed using FlowJo 10.9.0 (RRID:SCR_008520).
Therapeutic treatment regimens
Rigosertib (300 mg/kg, Onconova Therapeutics, currently Traws Pharma, Newtown PA 18940) and/or trametinib (1 mg/kg, LC Laboratories, Cat#: T-8123) were administered five days a week by oral gavage in a total volume of 100 μL. Rigosertib was prepared in H2O and trametinib were prepared in 5% DMSO and 95% corn oil. Corn oil with 5% DMSO was used as the vehicle control. Immunotherapies or equivalent amounts of isotype control, including anti-mouse PD-1 (200 μg/dose, Leinco Technologies Cat# P362, RRID:AB_2737557, clone: RMP1-14 and Leinco Technologies Cat# P377, RRID:AB_2737558, clone: 29 F.1A12), Rat IgG2a control (200 μg/dose, Leinco Technologies Cat# I-1177, RRID:AB_2737530, clone: 2A3), anti-mouse CTLA-4 (100 μg/dose, Leinco Technologies Cat# C1614, RRID:AB_2737453, clone: 9H10) and polyclonal Syrian hamster IgG control (100 μg/dose), anti-mouse agonist CD40 (30 μg/dose, Leinco Technologies Cat# C2825, RRID:AB_2829586, clone: FGK45) and Rat IgG2a control (30 μg/dose, Leinco Technologies Cat# I-1177, RRID:AB_2737530, clone: 2A3), were administered intraperitoneally every three days.
Generation of CD40 overexpressing melanoma cell lines
CD40 overexpressing melanoma cell lines were generated with a CRISPR-Cas9 based transcriptional activation system using CD40 lentiviral activation particles (sc-423444-LAC) and control lentiviral activation particles (sc-437282) (Santa Cruz Biotechnology, Inc). For lentiviral infection, 1014 and B16F10 melanoma cells were plated in 35 mm cell culture dishes in 10% FBS DMEM/F12 (1:1) media. When cells grow to ~40% confluency, 5.0 ug/ml Polybrene (Santa Cruz, CAT#: sc-134220) was added into culture media along with the CD40 or control lentiviral activation particles. After overnight 37 °C incubation, cells were changed to complete medium without Polybrene and incubated overnight. Cells were then trypsinized and plated into 150 mm cell culture dishes. To select successful lentiviruses infected cells, 0.2 ug/ml puromycin (Sigma-Aldrich, CAT#: P8833), 25 ug/ml hygromycin (RPI, CAT#: H75020) and 1.5 ug/ml blasticidin (InvivoGen, CAT#: ant-bl) were used for 1014 cells, while 0.75 ug/ml puromycin, 125 ug/ml hygromycin and 5 ug/ml blasticidin were used for B16F10 cells. After 7~12 days of selection, single monoclone colonies grown on 150 mm dishes were picked up for cell expansion. Selected monoclones were characterized on CD40 and PD-L1 expression, flow sorted and validated using western blotting and flow cytometry.
CD8+ T-cell killing assay
Tail vein injection of 2 × 106 B16F10-Luc2 cells that were fixed with 2% Formalin was performed to prime 8~10-week-old female C57BL/6 mice (RRID:IMSR_JAX:000664). After 2 days, splenic T cells were isolated using Pan T-cell Isolation Kit II (Cat#: 130-095-130, Miltenyi Biotec). Isolated T cells were cocultured with B16F10-Luc cells (Cat#: CRL-6475-Luc2, ATCC) for 28 h in the conditional complete RPMI 1640 medium containing 10% FBS, IL-7 (0.5 mg/mL, Cat#: 577802, Biolegend), IL-2 (30 U/mL, Cat#: 575402, Biolegend), and CD3/CD28 Dynabeads (Cat#: 11456D, Gibco) at a bead-to-cell ratio of 2:1 for various times based on the experimental design. Dead cells were removed by washing, and the remaining live cells were lysed using 2x Cell Lysis Buffer (Cat#: AA-Lys16ml, RayBiotech, Inc.) for luciferase activity quantitation to reflect the CD8+ T-cell killing capacity. In the experiments of CD11b+ B cell cocultures, splenic total B cells were isolated using MojoSort Mouse Pan B-cell Isolation Kit (Cat#: 480051, Miltenyi Biotec) after 2 days of in vivo tumor antigen priming as mentioned above. Total B cells were stimulated with 12.5 μg/ml aCD40 for 2 days and CD11b+ B cells were isolated using mouse CD11b Microbeads UltruPure (Cat#: 130-126-725, Miltenyi Biotec). The isolated CD11b+ B cells, CD8+ T cells and B16F10-Luc2 cells were mixed at different ratios as indicated in the specific figure legend and cocultured for 24~48hrs. Dead cells were then removed, and the luciferase activity of remaining live tumor cells was quantified.
Protein array and western blot analysis
Mouse splenocytes were isolated from C57BL/6 mice and were cultured in 10% FBS RPMI 1640 media with 10 µg/ml CD40 agonist antibody (Leinco Technologies, CAT#: C2825) for five days. B cells were isolated with EasySepTM mouse B cell isolation kit (STEMCELL Technologies, CAT#: 19854) per the manufacturer’s protocol. CD11b+ and CD11b- B cells were separated using EasySepTM mouse CD11b B cell positive selection Kit II (STEMCELL Technologies, CAT#: 18970 A) per the manufacturer’s protocol. CD11b+ B cells or CD11b- B cells were plated in 96 well plate at 1 × 106/well in 1% FBS RPMI 1640 media with 10 µg/ml CD40 agonist antibody (Leinco Technologies, CAT#: C2825) or 10 µg/ml Rat IgG2a Isotype Control antibody (Leinco Technologies, CAT#: I-1177) for 24 h. Cultured media were harvested and cleared by spin at 300 g for five minutes. Supernatants from three wells in each group were pooled together, and 100 µl were used for cytokines measurement with G-Series Mouse Cytokine Antibody Array 1000 (RayBiotech, CAT#: GSM-CAA-1000) per the manufacturer’s protocol. Slides were scanned with GenePix® Microarray Scanner (Molecular Device), and images were quantified with software GenePix Pro (Molecular Device). Tumor lysate samples were prepared as previously described31,69 and were examined in protein arrays using Mouse Antibody L308 Array Kit (308 proteins) (Cat#: AAM-BLG-1-4, RayBiotech), per the manufacturer’s protocol. For each mouse tissue, samples were pooled from five mice per group. The glass chip was scanned on the Cy3 channel of a GenoPix 4000B scanner (Genopix 6.1, Molecular Devices, Sunnyvale, CA). For each spot, the net density was determined by subtracting the background. The relative fold difference in protein amount was determined in reference to the amount present in the control samples. Data were analyzed with GraphPad Prism 10.4.2. For western blot analysis, mouse splenocytes were isolated from C57BL/6 mice and cultured in 10% FBS RPMI 1640 media with 10 µg/ml CD40 agonist antibody (Leinco Technologies, Cat#: C2825) for five days. CD11b+ B cells and CD11b- B cells were cultured in FBS free Opti-MEM media overnight and then treated with 10 µg/ml CD40 agonist antibody (Leinco Technologies, Cat#: C2825) or 10 µg/ml rat IgG2a isotype control (Leinco Technologies, Cat#: I-1177) for 30 minutes. The whole cell lysates were prepared with RIPA buffer (Fisher Scientific, Cat#: PI89900) supplemented with protease inhibitor cocktail (Roche, Cat#: 04693124001) and phosphatase inhibitor cocktail (Roche, Cat#: 4906845001). Pierce™ BCA protein assay was performed to quantitate the amount of protein in each sample and 18 µg of total proteins per sample was resolved by SDS-PAGE. For the 1014 melanoma cell line samples, cells were treated with 1 μM trametinib, 1 μM RGS, or 1 μM trametinib combined with 1 μM RGS, for 60 min and 24 h in DMEM/F12 medium supplemented with 10% FBS. Whole cell lysates were prepared with RIPA buffer and 40 µg proteins of each sample were used to run SDS-PAGE gel. Proteins were transferred to nitrocellulose membrane using Trans-Blot Turbo RTA Transfer Kit (Nitrocellulose, BioRad, Cat#: 1704272). Blots were blocked with Intercept Blocking Buffer (TBS) (LI-COR, Cat#: 927-60001) for 1 h. Primary antibodies were diluted in Intercept Antibody Diluent (TBS) (LI-COR, Cat#: 927-65001). Blots were incubated with primary antibodies overnight at 4 °C. The primary antibodies were purchased from Cell Signaling Technology, including pAKT-Thr308 (Cat#: 13038), total AKT (Cat#: 4691), pERK1/2 (Cat#: 4370), total ERK1/2 (Cat#: 4695) Cruz), with the expression level of β-actin (Invitrogen, Cat#: MA5-15739) as a loading control. After washing with TBS-T buffer, blots were incubated with secondary antibodies IRDye® 800CW Goat anti-Rabbit IgG (H + L) (LI-COR, Cat#: 926-32211) and IRDye® 680RD Goat anti-Mouse IgG (LI-COR, Cat#: 926-68070). Blots were scanned with OdysseyR CLx Imaging System (LI-COR). Signals were quantified with Image Studio Lite Ver 5.2 (LI-COR) and visualized with GraphPad Prism 10.4.2.
Mass spectrometry
The tumor samples were treated with rigosertib for 3 weeks, harvested and homogenized in three times their volume of deionized water to provide a sample that could be extracted for LCMS. Both tumor and plasma samples in 50 μl were extracted with 300 μl of acetonitrile, the sample supernatant was collected and dried under nitrogen before being reconstituted in Mobile Phase A (0.2% FA in Water/ACN 9/1). The samples were then run on a Waters Acquity HPLC with a gradient between Mobile Phase A and Mobile Phase B (0.2% FA in Water/ACN 1/9) over 10 min with the starting conditions of 100% A help for the first minute before the gradient begins and ending at 100% B at 7 min, washing the column for 1.5 min and then re-equilibrating the column for the remaining 1.5 min. The Mass Spectrometry experiment was performed using a TSQ Quantum Ultra AM, run in negative mode with an Electrospray source, monitoring a transition of 450.5 to 151, a spray voltage of 2500 sheath gas of 30 and Aux gas of 10. The data was analyzed using LCQUAN™ Quantitative Software (Thermo Scientific, Cat#: LCQUAN25).
Adoptive cell transfer
The lung colonization model was established by intravenously injecting 0.5 × 106 metastatic B16F10-Luc2 cells in 500 μl HBSS buffer (Corning, Cat#: 21-022-CV) via the tail vein of 8–10- week-old female C57BL/6 mice. Splenic B cells from tumor-free mice were cultured in 20% FBS RPMI containing 12.5 μg/ml aCD40 ex vivo for 2 days. The enriched CD11b+ B cells were then isolated using mouse CD11b Microbeads UltruPure (Cat#: 130-126-725, Miltenyi Biotec) for adoptive transfer. Starting at day 2 post tumor cell inoculation, 1.6 ~ 2 × 106 aCD40-stimulated CD11b+ B cells per mouse were intravenously injected (once a week) via the tail vein in 200 μl HBSS buffer (Corning, Cat#: 21-022-CV). Mice were then treated with 200ug/dose αPD1 or isotype IgG control twice a week. After 1 ~ 3 weeks of tumor implantation, 200 µl D-Luciferin (15 mg/ml, Cat#: eLUCNA-250, Gold Biotechnology) was intraperitoneally injected into each mouse and animal imaging was taken using the IVIS 200 Imaging System (Xenogen Imaging Technologies). Mice were sacrificed when there was a failure to thrive, difficulty breathing or after 3 weeks of tumor implantation. The lungs were weighted, fixed for making FFPE blocks, or proceeded with single cell isolation for flow cytometric analysis.
Pathology, immunohistochemistry and immunofluorescence assessment
Serum AST/ALT/BUN analysis and immunohistochemistry (IHC) analysis on 10% buffered formalin-fixed, paraffin-embedded tissue sections were performed by the Vanderbilt Translational Pathology Shared Resource (TPSR)24,70. Slides were placed on the Leica Bond Max IHC stainer. All steps besides dehydration, clearing and cover slipping are performed on the Bond Max. Slides are deparaffinized and heat-induced antigen retrieval was performed on the Bond Max using their Epitope Retrieval 2 solution for 20 min. Slides were incubated with primary antibodies, including anti-Ki67 (Cell Signaling Technology Cat# 12202, RRID:AB_2620142) diluted at 1:1000, anti-MelanA (Cat#: PA0233, Leica) and anti-CD8 (Cat#: HS-361 003, Synaptic Systems) at a 1:1000 dilution. The Bond Red Polymer Refine detection system (DS9800, Leica, Buffalo Grove, IL) was used for visualization and minimizing the background signal due to melanin. Slides were the dehydrated, cleared and cover slipped. Underlying 40x magnification of EVOS microscopy (M7000, Invitrogen), cells with IHC positive red staining vs blue negative staining of total nucleated cells were counted. The percentage of positive red staining cells was quantified and calculated from 20 fields of each slide sample. For immunofluorescence (IF) assessment, 5μm sections of FFPE samples were mounted in the Nanostring Digital Spatial Profiler (DSP) scan area in the center of the positively charged slide, baked, deparaffinized and rehydrated. Slides were placed on the Leica Bond Max stainer. Heat induced antigen retrieval was performed on the Bond Max using their Epitope Retrieval 2 solution for 20 min. Slides were washed in 1x TBS-T (0.1% Tween 20 in 1xTBS) and incubated with Buffer W (Nanostring) for 1 h at room temperature in a closed humidity chamber. The working IF antibody mix in Buffer W (Nanostring) includes CD20-AF594 (clone: IGE/773, Novus, Cat#: NB110-89474AF647, dilution factor = 1:235), CD8a-AF532 (clone: C8/1779 R, Novus, Cat#: NBP2-54595AF532, dilution factor = 1:30), and CD11b-AF647 (clone: D6X1N, Cell Signaling Technology, Cat#: 79750S, dilution factor = 1:100). Tissue samples were covered with 200 μl of the diluted IF antibody mix in the humidity chamber and incubated at 4 °C overnight. The slides were then washed 3 times in 1x TBS-T for 10 min of each wash, covered with 200 μl 10% neutral buffered formalin and incubated for 30 min at room temperature. Slides proceeded with 2 washes of 1x TBS-T for 5 min each. Meanwhile, the SYTO13 nuclear stain stock (NanoString Morphology Marker Kit, Cat#: 121300303) warmed from −20 °C to room temperature. The SYTO13 stain was 1:10 diluted in 1x TBS solution, and 200 μl diluted SYTO13 solution was added to cover the tissue and incubated for 15 min at room temperature. Slides were washed in 1x TBS-T twice and submerged in 2x SSC Buffer (Sodium Chloride Saline-Citrate, Cat#: S6639, Sigma-Aldrich) in a dark staining jar and stored at 4 °C (protected from light) until scanned on the Nanostring DSP platform at the Vanderbilt University Medical Center VANTAGE (Vanderbilt Technologies for Advanced Genomics) core facility.
Patient-derived organoid (PDO)
Institutional IRB approval and written informed consent from all patients were obtained before study initiation. All patient donors signed informed consent before providing tissue samples. Patient samples were collected as part of NCT01205815 clinical trial on a tissue-collection protocol approved by the Vanderbilt University IRB #100178. Fresh patient tissue was prepared as described43,71. Frozen tissue samples were thawed in a 37 °C bead bath and immediately transferred to a sterile 3 cm dish with prewarmed organoid media prior to following the same procedure as used with fresh tissue. Collected material was centrifuged at 350 g for 5 min at RT and resuspended in 1 mL complete organoid media (2:1:1 DMEM, Ham’s F12, MCDB, 10% FBS, 1% L-glutamine, 1% penicillin-streptomycin, 2% B-27, 1X Primocin) containing 2% Matrigel. 500uL of cell suspension was plated into each of two wells in a 24-well low-attachment cell culture plate. Following a 2-day recovery/growth period, these two wells were combined and diluted in complete media and plated across 21 wells of a 24-well low-attachment cell culture plate. With this protocol, deriving organoid cultures from fine needle aspirates (FNA) of tumor results in establishment of an organoid that is mostly devoid of connective tissue and the cells growing in the organoid stain positively for S100 and are extensively pigmented. The patient organoids that grew out from the FNA were ~90% viable. Rigosertib, CD40 agonist, and/or anti-PD1 or appropriate controls were prepared in complete media to reach final concentrations of 1 μM, 0.2 μg/ml, and 10 μg/ml, respectively. Wells were imaged using a EVOS XL Core Imaging System (Invitrogen) on days 5, 7, and 10 post-treatments. The tissue samples used were destroyed after the analysis. Organoid area measurements were obtained using ImageJ (RRID:SCR_003070) and analysis was performed using Graphpad Prism (RRID:SCR_002798).
Bulk RNA/TCR-Seq
RNA was extracted from tumor tissues using the RNeasy Plus Mini Kit (Cat#: 74134, Qiagen) per manufacturer’s protocol. RNA-Seq was performed using an Illumina next-generation sequencing platform at the Vanderbilt University Medical Center VANTAGE (Vanderbilt Technologies for Advanced Genomics) core facility and analyzed by Vanderbilt Technologies for Advanced Genomics Analysis and Research Design (VANGARD) core facility. RNA reads were aligned to the reference genome using STAR72 (RRID:SCR_004463) and quantified by featureCounts73 (RRID:SCR_012919). The RNA-seq quality was further checked after mapping by RNA-SeQC74 (RRID:SCR_005120), including alignment and duplication rates, GC bias, rRNA content, regions of alignment, 3’/5’ bias, and number of detectable transcripts. DESeq275,76 (RRID:SCR_015687) was used to detect differential expression between two conditions, considering technical and biological variability. CIBERSORT(RRID:SCR_016955), based on the LM22 gene signature, was used to estimate immune cells infiltrating the tumor77. Benjamini–Hochberg correction was applied to adjust for multiple-testing. Functional enrichment analysis was performed by WebGestalt78 (RRID:SCR_006786) and Gene Set Enrichment Analysis (GSEA, RRID:SCR_003199)79. TCRs were sequenced using the TCR Immunoverse all chain assay per the manufacturer’s protocol (Invitae/ArcherDX). Sequencing results were evaluated using Archer Immunoverse analyzer. CDR3 sequences and frequency tables were extracted from the manufacturers’ analysis platform and imported into R for analysis using the Immunarch package (RRID:SCR_023089, https://immunarch.com) in R.
Single-cell (sc)RNA-Seq analysis of human melanoma datasets
Two human melanoma datasets28,29 were downloaded and analyzed by Vanderbilt Technologies for Advanced Genomics Analysis and Research Design (VANGARD) core facility. The 10x Genomics Cell Ranger software was used to preprocess single-cell RNA-seq reads and to generate count matrices of each gene in each cell. Cells with low proportion of uniquely mapping reads ( < 1000), low proportion of expressed genes ( < 300) or high proportion of reads mapped to the mitochondrial RNA ( > 20%) were considered as low quality and excluded from downstream analysis. scRNABatchQC, developed by Dr. Liu at VANGARD, was used to detect any systematic biases, batch effects, or outlier samples80. After quality control, scRNA-seq data was normalized using UMI-filtered counts. Technical and biological variance, including capture efficiency, sequencing efficiency, or cell-cycle stage, were estimated, and adjusted by Seurat (RRID:SCR_016341)81,82. Batch effects if any were corrected using reciprocal principal component analysis-based integration81,82. Cell subpopulations were identified by the shared nearest neighbor modularity optimization and visualized by UMAP (RRID:SCR_018217) using Seurat (RRID:SCR_016341) based on the first 50 principal components generated from the top 2,000 highly variable genes. Cell subpopulations were automatically annotated by scMRMA, a multiresolution marker-based approach developed by Dr. Liu at VANGARD83, and then further manually checked by known marker genes. For scRNA-Seq analysis using the Single Cell Portal (singlecell.broadinstitute.org), the keywords “cancer” or “tumor” identified 101 studies. We further filtered the studies with available gene expression profiles of ITGAM (CD11b) and MS4A1 (CD20) or CD19, which identified 10 studies. To avoid biases resulting from artificially pooling unnormalized datasets, all identification and clustering of immune cell subsets, including B cells, were defined by the original method of publication using the default visualization program on the Single Cell Portal.
Pan-cancer survival analysis
Gene expression data for the TCGA Pan-Cancer (PANCAN), and the TCGA Pan-Cancer Clinical Data Resource was downloaded from cSurvival44. We focused on solid tumors only, so DLBC (Lymphoid Neoplasm Diffuse Large B-cell Lymphoma), FPPP (FFPE Pilot Phase II), and LAML (Acute Myeloid Leukemia) were removed from the analysis. For each sample, we computed the expression of a Breg gene signature (Sig. Breg) as the average expression of Itgam, Cd274, Tnfrsf13b, and Ahnak. Then samples were split into high- and low- Sig. Breg groups by using a median-anchored greedy84. Kaplan–Meier (KM) log-rank tests (default) and Cox proportional-hazards (PH) regression models were applied to assess the association of Sig. Breg with prognosis on overall survival, using survival v3.2.1185.
Statistical analysis
Treatment effects in standard two-group experiments were compared two-sided using a two-sample t-test for the independent samples (e.g., Fig. 1E) or using paired t-test for the correlated samples (e.g., Fig. 1D). A one-way analysis of variance (ANOVA) with post hoc two-sided test was used for more than two-group experiments to compare treatments’ differences (e.g., Figs. 5D, G). For in vivo experiments, differences in tumor weights between treatment groups were compared using one-way ANOVA (e.g., Figs. 3E, 4C, 5B, 6A, E, S5A and S6A). The progression of tumor volume (mm3) over time among groups of mice receiving different therapies was analyzed using a linear mixed-effects regression model. A random intercept was included to account for the correlation of the repeated measures within each mouse and an autocorrelation structure of order 1 [AR(1)] was specified for the residual variance-covariance structure. For two-factor experiments, we modeled a two-way interaction between treatment and day where a significant interaction indicates that the treatment effects vary over time. Model-based (least-square) means with standard error (SE) was reported. Differences in average tumor growth rates (tumor volume over time) between treatment groups were assessed using Wald test (Figs. 3A, 4C, S5A and S6A). For three-factor experiments, we modeled a three-way interaction (i) among treatment, day, and pretreatment to evaluate changes in treatment effects over time stratified by pretreatment (Fig. 5B) (ii) among treatment, day, and cell line to evaluate the tumor growth by cell lines (Fig. 6A, E). Model-based means with SE were presented and tumor growth rates between groups were compared. Drug effects, defined as difference in tumor growth rate between treatment groups, were reported with 95% confidence intervals (CIs) and compared between cell lines using multiple comparisons test for generalized linear hypothesis testing based on the linear mixed-effects model (Fig. 6B, F). Definition of time to αPD1-resistance: (1) the tumor is >100mm3 and (2) has a > 30% increase of tumor volume compared to the previous measurement. Survival curves (resistance-free probabilities (Fig. 3F) or survival probabilities (Fig. 8H) of treatment groups are estimated using the Kaplan-Meier method (RRID:SCR_018753) and compared using the log-rank test. Spearman’s rank correlation was used to measure the strength and direction of a monotonic association between two ranked (continuous) variables. For TCR-Seq analysis, the distribution of clonotype abundances by treatment group and cell lines was calculated to overview the structure of repertoire using repExplore function from immunarch R package (Fig. 7H). The proportion of repertoire occupied by clonal groups with specific frequencies relative abundance was measured by clonal space homeostasis using repClonality function from immunarch R package (Fig. 7J). Count data, including the number of clones (Fig. 7E) and number of unique clonotypes (Fig. 7F), were summarized as mean and standard deviation (SD) on a natural log scale by treatment group and cell line. The relationship between outcomes and treatment groups was assessed using a negative binomial generalized linear model (NB GLM) to account for over-dispersion in count data. Interaction effects between cell type and treatment were evaluated using a likelihood ratio test. Comparisons between treatment groups, adjusted for cell line, were conducted using Wald test. Drug effects with 95% CIs were reported and compared between cell lines using generalized linear hypothesis test for multiple comparisons based on NB GLM (Fig. 7G, Fig. S8A). At the gene level, clonotypes data were also analyzed using NB GLM with interaction term between treatment and cell type (Fig. 7I). To meet the model assumption, data for tumor volume, tumor weights and TCR-Seq analyses were analyzed on the natural log scale. Holm correction was used to adjust the p-value for pairwise comparison for survival curves, tumor growth and TCR-Seq analysis. All tests of statistical significance were two-sided. GraphPad’s Prism 10.2.3 software (RRID:SCR_002798) and R language (RRID:SCR_001905, version 4.4.0) were used for the statistical analysis. Unless otherwise specified, data are presented with Mean with SEM. *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ****p ≤ 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA/TCR-Seq datasets supporting the conclusions of this article are available in the Gene Expression Omnibus under accession GSE291447. The TCGA publicly available data used in this study are available in the data portal of Genomic Data Commons (GDC, https://gdc-portal.nci.nih.gov/legacy-archive/). The single cell RNA-Seq publicly available data used in this study are available in the Single Cell Portal (https://singlecell.broadinstitute.org/single_cell). The remaining data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
References
Wolchok, J. How recent advances in immunotherapy are changing the standard of care for patients with metastatic melanoma. Ann. Oncol. 23, viii15–viii21 (2012).
Hodi, F. S. et al. Improved survival with ipilimumab in patients with metastatic melanoma. N. Engl. J. Med. 363, 711–723 (2010).
Larkin, J. et al. Five-year survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl. J. Med. 381, 1535–1546 (2019).
Tawbi, H. A. et al. Relatlimab and nivolumab versus nivolumab in untreated advanced melanoma. N. Engl. J. Med. 386, 24–34 (2022).
Wolchok, J. D. et al. Long-term outcomes with nivolumab plus ipilimumab or nivolumab alone versus ipilimumab in patients with advanced melanoma. J. Clin. Oncol. 40, 127 (2022).
Zhou, Y., Richmond, A. & Yan, C. Harnessing the potential of CD40 agonism in cancer therapy. Cytokine Growth Factor Rev. 75, 40–56 (2024).
Gide, T. N., Wilmott, J. S., Scolyer, R. A. & Long, G. V. Primary and acquired resistance to immune checkpoint inhibitors in metastatic melanoma. Clin. Cancer Res. 24, 1260–1270 (2018).
Ribas, A. et al. Association of pembrolizumab with tumor response and survival among patients with advanced melanoma. JAMA 315, 1600–1609 (2016).
Yu, C. et al. Combination of immunotherapy with targeted therapy: theory and practice in metastatic melanoma. Front Immunol. 10, 990 (2019).
Wang, Y. et al. Anti-PD-1/L1 lead-in before MAPK inhibitor combination maximizes antitumor immunity and efficacy. Cancer Cell 39, 1375–1387.e1376 (2021).
Vendramini, E. et al. KRAS and RAS-MAPK pathway deregulation in mature B cell lymphoproliferative disorders. Cancers 14, 666 (2022).
Bahar, M. E., Kim, H. J., Kim, D. R. Targeting the RAS/RAF/MAPK pathway for cancer therapy: from mechanism to clinical studies. Sig. Transduct. Target Ther. 8, 455 (2023).
Haugh, A. M. et al. Targeted DNA sequencing of cutaneous melanoma identifies prognostic and predictive alterations. Cancers (Basel) 16, 1347 (2024).
Zhang, Z., Richmond, A., Yan, C. Immunomodulatory properties of PI3K/AKT/mTOR and MAPK/MEK/ERK inhibition augment response to immune checkpoint blockade in melanoma and triple-negative breast cancer. Int. J. Mol. Sci. 23, 7353 (2022).
Miyashita, H., Kato, S., Hong, D. S. KRAS G12C inhibitor combination therapies: current evidence and challenge. Front. Oncol. 14, 1380584 (2024).
Zhu, C. et al. Targeting KRAS mutant cancers: from druggable therapy to drug resistance. Mol. Cancer 21, 159 (2022).
Randic, T., Kozar, I., Margue, C., Utikal, J. & Kreis, S. NRAS mutant melanoma: towards better therapies. Cancer Treat. Rev. 99, 102238 (2021).
Weiss, S. A. et al. A phase II trial of the CD40 agonistic antibody sotigalimab (APX005M) in combination with nivolumab in subjects with metastatic melanoma with confirmed disease progression on anti-PD-1 therapy. Clin. Cancer Res. 30, 74–81 (2024).
van Hooren, L. et al. Agonistic CD40 therapy induces tertiary lymphoid structures but impairs responses to checkpoint blockade in glioma. Nat. Commun. 12, 4127 (2021).
Liu, X. et al. B cells expressing CD11b effectively inhibit CD4+ T-cell responses and ameliorate experimental autoimmune hepatitis in mice. Hepatology 62, 1563–1575 (2015).
Wulfken, L. M. et al. Case report: sustained remission due to PD-1-inhibition in a metastatic melanoma patient with depleted B cells. Front Immunol. 12, 733961 (2021).
Ahsan, N. F. et al. The current understanding of the phenotypic and functional properties of human regulatory B cells (Bregs). Oxf. Open Immunol. 5, iqae012 (2024).
Cabrita, R. et al. Tertiary lymphoid structures improve immunotherapy and survival in melanoma (vol 577, pg 561, 2020). Nature 580, E1–E1 (2020).
Yan, C. et al. Novel induction of CD40 expression by tumor cells with RAS/RAF/PI3K pathway inhibition augments response to checkpoint blockade. Mol. Cancer 20, 85 (2021).
Yan, C. & Richmond, A. Hiding in the dark: pan-cancer characterization of expression and clinical relevance of CD40 to immune checkpoint blockade therapy. Mol. Cancer 20, 146 (2021).
Wu, H. et al. PD-L1+ regulatory B cells act as a T cell suppressor in a PD-L1-dependent manner in melanoma patients with bone metastasis. Mol. Immunol. 119, 83–91 (2020).
Boroojerdi, S. et al. Staining pattern of specific and cross-reacting Melan-A antibodies: a comparative study on 15,840 samples from 133 human tumor types. APMIS 132, 479–491 (2024).
Jerby-Arnon, L. et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell 175, 984–997.e924 (2018).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013.e1020 (2018).
Yang, Y. et al. Pan-cancer single-cell dissection reveals phenotypically distinct B cell subtypes. Cell 187, 4790–4811.e22 (2024).
Yang, J. et al. Targeted deletion of CXCR2 in myeloid cells alters the tumor immune environment to improve antitumor immunity. Cancer Immunol. Res. 9, 200–213 (2021).
Zheng, S. W. et al. Tumor battlefield within inflamed, excluded or desert immune phenotypes: the mechanisms and strategies. Exp. Hematol. Oncol. 13, 80 (2024).
Marshall, L. A. et al. Tumors establish resistance to immunotherapy by regulating T(reg) recruitment via CCR4. J. Immunother. Cancer 8, e000764 (2020).
Nishina, T. et al. Interleukin-11-expressing fibroblasts have a unique gene signature correlated with poor prognosis of colorectal cancer. Nat. Commun. 12, 2281 (2021).
Lopatina, T. et al. IL-3 signalling in the tumour microenvironment shapes the immune response tumour endothelial cell-derived extracellular vesicles. Pharm. Res. 179, 106206 (2022).
Wrenn, E. D. et al. Regulation of collective metastasis by nanolumenal signaling. Cell 183, 395 (2020).
Yan, C. & Richmond, A. Th9 and Th17 cells: the controversial twins in cancer immunity. J. Clin. Invest 130, 3409–3411 (2020).
Han, X. Y. et al. Pan-cancer analysis reveals interleukin-17 family members as biomarkers in the prediction for immune checkpoint inhibitor curative effect. Front Immunol. 13, 900273 (2022).
Xu, L. et al. IL-22: A key inflammatory mediator as a biomarker and potential therapeutic target for lung cancer. Heliyon 10, e35901 (2024).
Kuske, M. et al. Immunomodulatory effects of BRAF and MEK inhibitors: implications for melanoma therapy. Pharm. Res. 136, 151–159 (2018).
Jost, M. et al. Combined CRISPRi/a-based chemical genetic screens reveal that rigosertib is a microtubule-destabilizing agent. Mol. Cell 68, 210–223.e216 (2017).
Jost, M. et al. Pharmaceutical-grade rigosertib is a microtubule-destabilizing agent. Mol. Cell 79, 191–198.e193 (2020).
Vilgelm, A. E. et al. Fine-needle aspiration-based patient-derived cancer organoids. iScience 23, 101408 (2020).
Liu, J. F. et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400 (2018).
Fu, Y. et al. Intestinal CD11b+ B cells ameliorate colitis by secreting immunoglobulin A. Front Immunol 12, 697725 (2021).
Benci, J. L. et al. Opposing functions of interferon coordinate adaptive and innate immune responses to cancer immune checkpoint blockade. Cell 178, 933–948.e914 (2019).
Lerner, E. C. et al. CD8(+) T cells maintain killing of MHC-I-negative tumor cells through the NKG2D-NKG2DL axis. Nat. Cancer 4, 1258–1272 (2023).
Bradford, H. F. & Mauri, C. Diversity of regulatory B cells: markers and functions. Eur. J. Immunol. 54, e2350496 (2024).
Matsumoto, M. et al. Interleukin-10-producing plasmablasts exert regulatory function in autoimmune inflammation. Immunity 41, 1040–1051 (2014).
Garcia-Diaz, A. et al. Interferon receptor signaling pathways regulating PD-L1 and PD-L2 expression. Cell Rep. 29, 3766 (2019).
Lim, S. O. et al. Deubiquitination and stabilization of PD-L1 by CSN5. Cancer Cell 30, 925–939 (2016).
Chan, L. C. et al. IL-6/JAK1 pathway drives PD-L1 Y112 phosphorylation to promote cancer immune evasion. J. Clin. Invest 129, 3324–3338 (2019).
Derre, L. et al. Expression of CD94/NKG2-A on human T lymphocytes is induced by IL-12: implications for adoptive immunotherapy. J. Immunol. 168, 4864–4870 (2002).
Nair, A. B. & Jacob, S. A simple practice guide for dose conversion between animals and human. J. Basic Clin. Pharm. 7, 27–31 (2016).
Singh, M. et al. Intratumoral CD40 activation and checkpoint blockade induces T cell-mediated eradication of melanoma in the brain. Nat Commun 8, 1447 (2017).
Osorio, J. C. et al. Intratumoral Fc-optimized agonistic CD40 antibody induces tumor rejection and systemic antitumor immunity in patients with metastatic cancer. Res Sq. 3, rs.3.rs–4244833 (2024).
Correa, S. et al. Injectable Nanoparticle-based hydrogels enable the safe and effective deployment of immunostimulatory CD40 agonist antibodies. Adv. Sci. (Weinh.) 9, e2103677 (2022).
Garcia-Manero, G. et al. Rigosertib versus best supportive care for patients with high-risk myelodysplastic syndromes after failure of hypomethylating drugs (ONTIME): a randomised, controlled, phase 3 trial. Lancet Oncol. 17, 496–508 (2016).
O’Neil, B. H. et al. A phase II/III randomized study to compare the efficacy and safety of rigosertib plus gemcitabine versus gemcitabine alone in patients with previously untreated metastatic pancreatic cancer. Ann. Oncol. 26, 2505 (2015).
Robert, C. et al. Five-year outcomes with dabrafenib plus trametinib in metastatic melanoma. N. Engl. J. Med 381, 626–636 (2019).
Cohen, C. J. et al. Isolation of neoantigen-specific T cells from tumor and peripheral lymphocytes. J. Clin. Invest 125, 3981–3991 (2015).
Gros, A. et al. Prospective identification of neoantigen-specific lymphocytes in the peripheral blood of melanoma patients. Nat. Med 22, 433–438 (2016).
Kalaora, S. et al. Combined analysis of antigen presentation and T-cell recognition reveals restricted immune responses in melanoma. Cancer Discov. 8, 1366–1375 (2018).
Xie, N. et al. Neoantigens: promising targets for cancer therapy. Signal Transduct. Target Ther. 8, 9 (2023).
Phadke, M. S. et al. Targeted therapy given after anti-PD-1 leads to prolonged responses in mouse melanoma models through sustained antitumor immunity. Cancer Immunol. Res. 9, 554–567 (2021).
Haas, L. et al. Acquired resistance to anti-MAPK targeted therapy confers an immune-evasive tumor microenvironment and cross-resistance to immunotherapy in melanoma. Nat. Cancer 2, 693 (2021).
Jiang, J. J. et al. Translational and therapeutic evaluation of RAS-GTP inhibition by RMC-6236 in RAS-driven cancers. Cancer Discov. 14, 994–1017 (2024).
Yan, C. et al. Inhibition of the PI3K/mTOR pathway in breast cancer to enhance response to immune checkpoint inhibitors in breast cancer. Int. J. Mol. Sci. 22, 5207 (2021).
Yang, J. et al. CXCR2 expression during melanoma tumorigenesis controls transcriptional programs that facilitate tumor growth. Mol. Cancer 22, 92 (2023).
Yan, C. et al. Proximity of immune and tumor cells underlies response to BRAF/MEK-targeted therapies in metastatic melanoma patients. NPJ Precis. Oncol. 6, 6 (2022).
Phifer, C. J. et al. Obtaining patient-derived cancer organoid cultures via fine-needle aspiration. Star. Protoc. 2, 100220 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
DeLuca, D. S. et al. RNA-SeQC: RNA-seq metrics for quality control and process optimization. Bioinformatics 28, 1530–1532 (2012).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Zhang, B., Kirov, S. & Snoddy, J. WebGestalt: an integrated system for exploring gene sets in various biological contexts. Nucleic Acids Res. 33, W741–W748 (2005).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet 34, 267–273 (2003).
Liu, Q. et al. scRNABatchQC: multi-samples quality control for single cell RNA-seq data. Bioinformatics 35, 5306–5308 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Li, J., Sheng, Q., Shyr, Y. & Liu, Q. scMRMA: single cell multiresolution marker-based annotation. Nucleic Acids Res. 50, e7 (2022).
Cheng, X. et al. cSurvival: a web resource for biomarker interactions in cancer outcomes and in cell lines. Brief. Bioinform 23, bbac090 (2022).
Therneau, T. M. & Grambsch, P. M. Modeling Survival Data: Extending the Cox Model (Springer, Berlin, 2000).
Acknowledgements
We thank Dr. Justin Balko (Vanderbilt University Medical Center) for his helpful discussions and technical support on the TCR-Seq analysis. We thank Holly Crandall (Vanderbilt University Medical Center) for coordinating clinical samples for the laboratory correlative study. We thank Dr. Tae Kon Kim (Vanderbilt University Medical Center) for the technical support on the immunofluorescent staining analysis. We are indebted to Dr. Christine Lovly (Vanderbilt University Medical Center), Patricia A. Ward (Vanderbilt University) and Yang (Maggie) Zhou (supported by SyBBURE Searle Undergraduate Research Program at Vanderbilt University) for facilitating these studies. This work was supported by grants from the Lloyd Foundation Melanoma Research Grant (C.Y.), NCI R01-CA116021 (A.R., C.Y.), the Department of Veterans Affairs SRCS Award IK6BX005225 (A.R.) and the Department of Veterans Affairs MERIT Award 101BX002301 (A.R. and C.L.). The authors sincerely thank Onconova Therapeutics (currently Traws Pharma, Newtown PA 18940), for kindly supplying rigosertib for this work. Flow Cytometry experiments were performed in the VUMC Flow Cytometry Shared Resource that is supported by the Vanderbilt Ingram Cancer Center (P30CA68485) and the Vanderbilt Digestive Disease Research Center (DK058404). Sequencing support was provided by the VUMC VANTAGE Core Facility, also supported by the NCI/NIH VICC Cancer Center Grant P30CA68485. We acknowledge the Translational Pathology Shared Resource is supported by NCI/NIH Cancer Center Support Grant P30CA068485, the Shared Instrumentation Grant S10 OD023475-01A1 for the Leica Bond RX.
Author information
Authors and Affiliations
Contributions
Project conception experimental design, manuscript preparation and editing (C.Y., A.R.), consultation regarding human relevance of project design, procurement of human patient samples, and critical reading of manuscript (D.B.J.), pathological analysis of tissues and cells, consultation on organoid development, and critical reading of the manuscript (V.L.W.), collection of experimental data and data analysis (W.L., Jinming Yang, K.B., C.Y., Q.H.), bioinformatics analysis (Jing Yang, Q.L., C.Y.), biostatistical analysis of data (S.C.), funding for project (A.R., C.Y.). All of the authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.B.J. has consulted for AstraZeneca, BMS, Jackson Laboratories, Merck, Novartis, Pfizer, and Teiko. The other authors do not have any competing interests to disclose.
Peer review
Peer review information
Nature Communications thanks Chuanlin Ding and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yan, C., Luo, W., Yang, J. et al. RAS/MEK/PI3K pathway inhibition augments response to CD40 agonism by targeting CD11b+ Bregs thereby overcoming melanoma PD1-resistance. Nat Commun 17, 162 (2026). https://doi.org/10.1038/s41467-025-67315-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67315-1