Abstract
A major concern of COVID-19 is immune escape. While T cells are implicated in protection against severe disease, direct evaluation of their capacity to specifically target SARS-CoV-2 variants remains limited. We aim to narrow this gap by profiling the ability of CD8+ T cells to recognize SARS-CoV-2 mutations after vaccination and observe a high degree of phenotype and repertoire diversity. To better understand the breadth of antigen repertoire coverage, we interrogate the ability of SARS-CoV-2 specific TCRs to bind all point mutations of the immunodominant HLA-A2:S269-277 epitope. While mutation space coverage is surprisingly vast, specific substitutions are not recognized by the vaccine-elicited repertoire. Structural analyses reveal a TCR sequence-based antigen recognition limitation intrinsic to vaccination. Further investigation reveals that this repertoire ‘hole’ is not present in the naive repertoire, suggesting future immune escape through these potential escape mutations could be relieved by modifications to vaccines.
Similar content being viewed by others
Introduction
Immunodominance is a common immunological phenomenon in infection, vaccination, and cancer, where a small set of many possible epitopes is targeted by the immune system1,2. Although metabolically favorable, immunodominance is inherently susceptible to immune escape. It is becoming clearer that the ability of COVID-19 vaccines to elicit neutralizing immunity toward emerging SARS-CoV-2 variants of concern (VOCs) is diminishing3. Furthermore, booster doses with bivalent formulations that contain spike glycoprotein-encoding mRNA from the omicron lineages do not appear to improve antibody neutralization4,5. Taken together, these findings point toward a growing importance of other arms of the immune system as the virus continues to evolve.
While T cells are not generally thought to contribute to sterile immunity, they have been shown to play a key role in reducing COVID-19 severity, particularly in immunocompromised and at-risk groups6. One study evaluating a small group of healthcare workers implicated a strong and diverse T cell response in resisting ‘breakthrough’ infections7. Furthermore, unlike the observed impairment of humoral immunity toward VOCs, several groups have detected no difference between the vaccine-elicited T cell response toward ancestral versus omicron lineages8,9,10. The coverage of potential antigen mutation space by T cells, however, remains unclear.
The ability of T cell receptors (TCRs) to cross-react to multiple epitopes further convolutes the role of T cell memory against VOC lineages. SARS-CoV-2 responding TCRs have been repeatedly reported to cross-react with seasonal coronaviruses11. It is thus reasonable to speculate that TCRs responding to ancestral lineages could respond to VOC mutations as well. Nonetheless, mapping of the mutational coverage space has revealed reproducible ‘blind spots’ where vaccine-elicited repertoires fail to recognize specific substitutions, highlighting both the strength and the limits of TCR degeneracy9.
Evaluating antigen-specific T cells also poses a great challenge. The most common approaches involve stimulating T cells with pools of peptides and assaying the upregulation of activation-induced markers (AIM)8,12,13. While AIM assays have laid much of the groundwork for our understanding of T cell immunity to SARS-CoV-2, their sensitivity can be limited. Other studies have used alternative approaches, such as DNA-barcoded peptide-major histocompatibility complex (pMHC) multimers14,15,16. These studies, however, did not consider VOC-derived epitopes. Deeper knowledge of the capacity of T cells to respond to VOC mutations could improve our understanding of how to better mobilize an immune compartment that may be capable of resisting antigenic drift.
In this work, we aim to investigate a natural T cell response to a common vaccine to better understand TCR-antigen recognition at the amino acid level. To do so, we investigated the ability of CD8+ T cells induced by SARS-CoV-2 challenge to target immunodominant epitopes and their corresponding VOC-derived mutations using a library of DNA-barcoded pMHC tetramers containing a panel of curated ancestral, VOC, and common coronavirus (hCOV) epitopes based on their high prevalence in literature14,15,16,17. After single-cell sequencing, we recalled each cell’s TCR sequence and cognate specificity, finding striking immunodominant to a few unmutated epitopes. To our surprise, despite low diversity in their antigenic targets, many donors had highly diverse TCR repertoires. We hypothesized that integration of these diverse repertoires may enable an individual to maintain durability toward drifting antigens through distinct TCR recognition patterns. Understanding the specific TCR recognition patterns capable of targeting these substitutions could better arm vaccine design in the case of future mutagenesis. To test this hypothesis, we generated transgenic antigen-specific TCR cell lines to evaluate each TCR’s ability to bind to altered peptide ligand (APL) libraries containing all possible single amino acid point mutations of their original cognate epitopes. We observed a high degree of TCR promiscuity that was previously underappreciated. Structural analysis of the antigen-binding pocket pointed to specific TCR features that led to degeneracy at several amino acid positions. Finally, we show that a ‘hole’ in the vaccine elicited repertoire could be filled by TCRs within the vaccine-naïve repertoire through the usage of distinct TCRs with distinct sets of antigen-binding features. Such ‘completeness’ observed within the TCR repertoire suggests that modifications to the vaccine formulation could rescue these escape mutations if they arise in future viral lineages.
Viral evolution poses a persistent challenge to human health. Although vaccines are routinely updated to match emerging viral strains, it is equally important to consider whether our immune system remains sufficiently equipped to counter these evolving threats. Our study reveals that leveraging the complementary breadth of vaccine-induced and naive TCR repertoires could inform the design of next-generation vaccines to anticipate viral escape and maintain population-level immunity against future SARS-CoV-2 variants. This framework can be extended to study T cell responses to other viruses, providing a blueprint for understanding and optimizing vaccine-elicited immunity.
Results
SARS-CoV-2 lineage analysis reveals a low mutation burden on immunodominant CD8+ T cell targets
To delineate SARS-CoV-2 CD8+ T cell targets, we surveyed the literature14,15,16,17 and the immune epitope database (IEDB)18 for commonly cited epitopes (Supplementary Table 1). There was an astoundingly low mutation burden on these epitopes, with only two mutations in Omicron lineages, both restricted to HLA-A*02:01. Based on this information and the cohort available (Supplementary Table 2), we focused on HLA-A*02:01- and HLA-A*03:01-restricted epitopes.
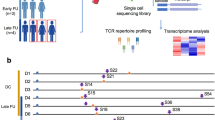
In addition to commonly cited epitopes, we also sought to evaluate the CD8+ T cell response to VOC mutations that may not be well-studied. We thus aligned VOC spike protein sequences from the NCBI Virus Database to the ancestral strain to determine stable mutations within each lineage. We then included mutant epitopes and their ancestral counterparts that were predicted to be strong binders (% Rank < 0.5) by NetMHCpan 4.119 (Fig. 1a). After curating our panel, we evaluated SARS-CoV-2-specific CD8+ T cells before and after COVID-19 vaccination using TetTCR-SeqHD20, which uses DNA-barcoded pMHC tetramers and single cell sequencing to link antigen specificity and TCR sequences to single cell phenotypes (Fig. 1b, c).
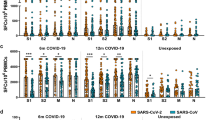
a Mutation analysis of the ancestral spike amino acid sequence to several variants (top). Lineage (left) and percent sequence conservation from the ancestral strain (right) are depicted. Pile-up histogram of epitopes analyzed in this work and their relative positions are shown. Specific mutations analyzed in this work are shown at the bottom. b Overview of TetTCR-SeqHD workflow and experimental design. c Overview of TCR fingerprinting workflow. d Representative gating for sorting antigen-specific T cells before and after vaccination (left). Summary of pre- (n = 12) and post-vaccine (n = 22) spike tetramer-positive T cell frequencies by flow cytometry (right). Each point represents a unique donor. Mean and standard error of the mean summarizing each group are depicted by the middle line and whiskers, respectively. Pre and Post groups were compared by Student’s t-test (two-sided). **P < 0.01. e Mean TCR frequency (determined by Eq. (3)) of CD8+ T cells targeting spike epitopes before and after vaccination from each donor (CONSERVED, Pre n = 10, Post n = 19; WT, Pre n = 7, Post n = 16; VARIANT, Pre n = 8, Post n = 18; Cross-Reactive, Pre n = 4, Post=4; hCOV, Pre n = 7, Post n = 15). Mean and standard error of the mean summarizing each group are depicted by the middle line and whiskers, respectively. b, c were created in BioRender. Malone, M. (2025) https://BioRender.com/ k17aooq.
HLA-A2-restricted CD8+ T cells predominantly target conserved spike epitopes through de novo immunity
We used a panel of 62 spike and 21 control pMHC tetramers (Supplementary Tables 3, 4) to sort antigen-specific CD8+ T cells from the blood of pre- and/or post-SARS-CoV-2 vaccine recipients (Supplementary Table 2). In total, 22 donors were evaluated, 12 of which had pre- and post-vaccine-matched blood draws. As expected, the frequencies of spike+ cells by flow cytometry were higher after vaccination (Fig. 1d, Supplementary Fig. 1a).
After sorting tetramer+ cells, we determined cellular phenotypes by quantifying gene expression and surface protein markers. In total, we retained 15,371 high-quality cells that spanned several distinct naïve, memory, and effector clusters (Supplementary Figs. 5b, c, 6a, Supplementary Table 5). Of note, 7621 of these cells could be assigned putative specificities based on their DNA barcode counts (determined by Eq. (1), also see Supplementary Fig. 3b). To understand the impact of viral mutagenesis on the antigen-specific T cell response, we grouped spike-specific T cells by the lineages of their cognate epitopes. Looking broadly at the clonal frequency (determined by Eq. (3)) of spike specific TCRs, it was clear that T cells targeting conserved epitopes were dominant (Fig. 1e). In contradiction with other publications12,15,21, we observed negligible recruitment of pre-existing immunity from SARS-CoV-2 or hCOV specific T cells (Supplementary Fig. 1c). In fact, of 272 expanded, spike specific clonotypes, we only detected one clone spanning pre- and post-vaccination time points. Additionally, this clone was lower in frequency after vaccination, indicating its lack of invigoration by the vaccine. Interestingly, it was specific to a BA.1-derived epitope, despite the donor’s pre-vaccine blood draw being collected in April of 2021, long before the first cases of Omicron were identified in November of 2021. The donor also reported no SARS-CoV-2 infections prior to the conclusion of the study. It is thus likely that the presence of this omicron-specific clone was a product of unknown, cross-reactive pre-existing immunity.
The CD8+ T cell response to HLA-A2-restricted epitopes is public, immunodominant, and diverse
We next sought to understand the role of individual epitopes in response to COVID-19 vaccination. While we detected T cells specific to many unique SARS-CoV-2 spike epitopes (47 of 51 targets), we focused on epitopes exclusively targeted by the clonal T cell response (n > 1) after vaccination. We found that a surprisingly low number of spike epitopes were targeted by clonal TCRs (Fig. 2a). Of the seven spike epitopes targeted, one HLA-A*02:01-restricted, conserved epitope (S269–277: YLQPRTFLL, defined as COV-S-082 here) was immunodominant and public. Except for one donor, all other 15 HLA-A*02:01-restricted donors had a clonal response to this epitope (Fig. 2b). Additionally, of the donors that had a clonal response, COV-S-082 was the dominant target in all but two donors. Interestingly, this stark, public dominance was also paired with a high TCR polyclonality.
a Frequencies, determined by Eq. (3), of expanded TCR clonotypes (clone cell count > 1) specific to spike epitopes in post-vaccine samples (COV-S-044, Pre n = 0, Post n = 4; COS-S-044.2, Pre n = 1, Post n = 1; COV-S-082, Pre n = 0, Post n = 119; COV-S-085.2, Pre n = 0, Post n = 1; COV-S-086 Pre n = 0, Post n = 1; COV-S-094 Pre n = 0, Post n = 73; hCOV-S-004 Pre n = 4, Post n = 1). b Distribution of spike epitope-specific TCR clonotypes in post-vaccination samples. Lines within the bars are a unique clonotype, where their size is relative to their frequency in the repertoire. Numbers above bars are the total number of cells represented by the bars. c Network analysis of antigen-specific TCR clonotypes by TCR distance (calculated with tcrdist322). Nodes represent unique clonotypes, colored by their cognate specificities. Connections represent a TCR distance less than a pre-determined cutoff. Select communities determined by k-means clustering are circles and labeled. d TCR characteristics of select communities from (c). e Comparison of the NSE (determined by Eq. (2)) between TCRα and TCRβ in their CDR3 sequences and V gene usage in cells specific to COV-S-082. Each point represents a unique donor (CDR3a n = 17; CDR3b n = 17; TRAV n = 15; TRBV n = 17). f Transitional index (ITRAN, determined by Eq. (4)) of expanded (n > 1) COV-S-082 specific T cell clonotypes compared with other SARS-CoV-2 spike epitopes represented in (a) (COV-S-044 n = 5; COV-S-044.2 n = 2; COV-S-082 n = 125; COV-S-085.2 n = 1; COV-S-086 n = 1; COV-S-094 n = 80). g Comparison of memory COV-S-082 specific cells in each broad phenotype for donors with a response. For all boxplots, the center is drawn through the median of the measurement, and the lower and upper bounds of the box correspond to the first and third quartiles (EFF n = 13; MEM n = 16; NAIVE n = 16, MAIT n = 1. Data in (e) were compared by Student’s t-test (two-sided). Data in (f, g) were compared by ANOVA. Pairwise comparisons between specific groups were made via the Tukey test. P values are shown; ****P < 0.0001.
Similar polyclonality toward immunodominant SARS-CoV-2 epitopes has been characterized, but primarily in an HLA-A*01:01-restricted epitope14. We took a similar approach to analyze the antigen-specific TCRs in our data by calculating their pairwise TCR distances using the package tcrdist322, grouping clonotypes into a network, then clustering them into TCR communities (Fig. 2c). This approach revealed many distinct TCR communities within the COV-S-082-specific repertoire. Of note, communities were generally composed of non-overlapping CDR3, V, and J genes from multiple donors (Fig. 2d, Supplementary Figs. 4, 5a–c). We next inferred the diversity of CDR3 and V-gene usage by calculating their respective normalized Shannon’s entropies (NSE23) (determined by Eq. (2)). Interestingly, the TRBV gene usage was significantly more diverse than TRAV usage (Fig. 2e, Supplementary Fig. 6a), suggesting a potential importance of TCRα, specifically its prevalent usage of TRAV12 (Supplementary Fig. 5b), in COV-S-082 recognition. Importantly, TCRs specific to other SARS-CoV-2 and non-SARS-CoV-2 epitopes did not necessarily mirror this TRAV-dominant behavior (Supplementary Fig. 6a).
In addition to the high degree of clonality and TCR diversity within the COV-S-082 specific repertoire, we also observed a high degree of phenotypic diversity across donors (Supplementary Fig. 6b, c) and TCR communities (Supplementary Fig. 6d). To quantitate phenotypic plasticity, we calculated the transitional capacity (determined by Eqs. (4, 5)) of TCR clonotypes as previously described24, finding that COV-S-082 specific clones were significantly more likely to exist in many cell states compared with other specificities (Fig. 2f). Focusing on more general phenotypes, COV-S-082 specific T cells were most represented in the memory compartment (Fig. 2g). Given that immunological memory is also thought to be a critical reservoir for secondary immunity, phenotypically diverse COV-S-082 specific memory T cells could play a meaningful role in combating breakthrough infections7, illuminating the potential impact of the repertoire evaluated in this work.
TCR fingerprinting reveals high mutation coverage by the HLA-A2-restricted TCR repertoire
To understand if the diverse TCR repertoires targeting this public and immunodominant epitope after SARS-CoV-2 challenge were generalizable to other types of antigens and diseases, we compared the repertoire diversities of antigen-specific T cells from several different antigenic sources in our data and several other publications14,15,20. While SARS-CoV-2-specific repertoires exhibited higher diversity than repertoires targeting human CMV epitopes (a persistent virus), they were of comparable diversity to IAV and EBV-specific repertoires (Fig. 3a). It is thus possible that diverse repertoires targeting immunodominant epitopes are commonplace in the T cell response. We hypothesized that the immune system could benefit from this diversity, mounting a strong response to a few epitopes, while limiting commitment to a specific pMHC-TCR binding modality. Such a paradigm could theoretically provide protection against drifting viruses, with each TCR representing a non-redundant ‘safety net’ capable of responding to unique sets of mutations. If true, an understanding of the factors that contribute to these properties could be utilized toward developing better therapeutics targeting diseases beyond the scope of COVID-19 and infectious diseases.
a NSE (determined by Eq. (2)) of TCR repertoires across various antigenic sources from this and other studies compared with COV-S-082 specific repertoires. Each point represents a unique donor (CMV n = 25; EBV n = 43; IAV n = 29; MART1 n = 9; SARS-2 Non-Spike n = 21; SARS-2 Spike n = 54). b Individual TCR fingerprints, each heatmap plot represents one experiment with 172 tetramers stained individually. The YLQPRTFLL wildtype loaded tetramer is assayed once for each TCR, and the result is reused in Y1, L2, Q3, P4, R5, T6, F7, L8, and L9 positions. c Integrated TCR fingerprint (n = 20). Wild-type epitope is boxed in black. Intra-infection mutants detected in a previous study28 are boxed in green. Gray-filled boxes indicate filtered pMHC species based on the T2 stabilization assay (Supplementary Fig. 3f, g). d Number of TCRs capable of binding indicated fraction of pMHC species with a Log2 fold-change > 1 (top). A total of 80% of the possible mutations are covered by TCRs, as indicated by the bracket based on the binding capacities of the 20 TCRs analyzed. Fraction of mutations bound by individual TCRs (bottom). For boxplots in a, the center is drawn through the median of the measurement, and the lower and upper bounds of the box correspond to the first and third quartiles, groups were compared by the Wilcox rank-sum (two-sided). P-values are shown.
To test our hypothesis, we transduced COV-S-082-specific TCRs from several TCR communities into cell lines (Supplementary Table 6). We then characterized the binding patterns by probing each TCR with an APL library containing pMHC tetramers with every possible single amino acid point mutation (172 in total; Fig. 1c). To understand the each pMHC-TCR interaction, we plotted the binding signal of each mutant peptide with respect to the position of its mutation, effectively visualizing each TCR’s unique ‘fingerprint’ (Fig. 3b). Based on the aggregated distribution of top binding TCRs for each pMHC species (Supplementary Fig. 7a, b), we classified pMHCs as binders if they had a fold-change at least 2x above background. To our surprise, TCR promiscuity was much greater than previously reported25, likely reflecting the improved sensitivity of our assay compared with multiplexed recall of DNA barcoded pMHC multimers after next-generation sequencing (NGS), as has been done previously25. Specifically, we observed a threshold phenomenon when using a multiplexed NGS-based approach that was not evident in our flow cytometric approach (Supplementary Fig. 7c, d). We suspect competitive binding between pMHC species during multiplexing limits the ability of weaker species to confer an adequate signal for detection.
As previously reported25, we observed that substitutions on the HLA-A2 anchor residues at positions two and nine did not appear to affect TCR binding (Fig. 3b, Supplementary Fig. 7e). Such a phenomenon may be an artifact of UV-mediated peptide exchange. To clarify discrepancies between biologically relevant pMHC binding and reagent artifact, we performed a T2 MHC stabilization assay26 on our APL library. As expected, substitutions to peptide position two and nine generally reduced MHC stabilization, mostly following canonically and semi-canonically tolerated amino acids27 (Fig. 3b). Additionally, while MHC stabilization correlated well with NetMHC binding predictions, there were a few outliers (Supplementary Fig. 7f, g), advocating for experimental validation of stable pMHC formation whenever possible. Based on the distribution of MHC stabilization, we removed pMHC species that did not stabilize MHC at least 1.5 x greater than background from downstream analyses.
To understand how the integration of diverse TCRs targeting a single epitope could cooperate to resist antigenic drift at the population level, we combined all twenty TCR fingerprints in silico (Fig. 3c). In total, the integrated TCR fingerprint covered 80% of the total mutation space (Fig. 3d), including several low frequency intra-infection escape mutations that have been previously reported during late-stage COVID-1928. TCR promiscuity also correlated well with binding strength to the wild type epitope (R = 0.82, P = 5.02e–05), but was not necessarily restricted by community assignment (Supplementary Fig. 8a). In line with this observation, there were several cases of weaker binding TCRs that covered the mutations missed by stronger binding TCRs (Fig. 3b, Supplementary Fig. 8b), advocating for maximizing mutation coverage through collectivity, rather than individual strength or promiscuity.
Structural analysis of vaccine-elicited TCRs in complex with pMHC reveals a critical amino acid residue restricting mutation coverage
To directly compare the binding patterns of individual TCRs, we hierarchically clustered each TCR fingerprint and visualized them in a single heatmap (Fig. 4a). As expected, TCRs from the same community tended to cluster together, reflecting their similar binding patterns through similar TCR features. Further supporting this paradigm, the binding patterns of TCRs from the same communities were generally well correlated (Fig. 4b). In addition to community-intrinsic patterns, several global TCR patterns also emerged: 1) most TCRs were able to bind to substitutions at positions one, two, and nine, 2) mutation coverage at position five was sparse, and 3) a high degree of variability was detected at positions four, six, seven and eight.
a Heatmap of hierarchically clustered individual TCR fingerprints (23 TCRs plotted). TCR binding strength is normalized by row. TCR communities are annotated on the left. b Pairwise Pearson correlations and hierarchical clustering of each TCR by their binding patterns. Size of the dot represents the inverse P-value. Color intensity represents the correlation coefficient. Select TCR communities are indicated. TCR clones are indicated in the vertical and diagonal axis of the plot. c Interaction of R5 (top) and T6 (bottom) residues of COV-S-082 with SVAR-14 TCRα (orange) and TCRβ (green) during MD simulations. d Same as (c), showing F7 (top) and L8 (bottom) residues. e Total contact between COV-S-082 residues and TCRα/β for 7RTR29 (top) and SVAR-14 (bottom) TCRs during simulation.
To explain some of the global binding patterns, we performed molecular dynamics (MD) simulations of SVAR-14 in complex with HLA-A*02:01:COV-S-082 based on a previously solved crystal structure of a similar TCR that only differed by a single amino acid in the CDR3β29 (Supplementary Fig. 9a, b). Initial examination of the TCR binding pocket corroborated much of our TCR binding data. For example, the general tolerance of each TCR to mutations at Y1, L2, and L9 could be explained by primary contact of these residues with MHC, rather than TCR, through the hydrophobic effect (Supplementary Fig. 9c–e). We also observed a high degree of heterologous TCRα/β contact mediated by multiple H-bonds with R5 and T6 (Fig. 4c, Supplementary Fig. 9e). Particularly toward the R5 residue, such a stable conformation indicates its importance in antigen recognition. This, in combination with the low mutation coverage at this position (Fig. 4a), led us to speculate that R5 may act as an antigenic keystone for the vaccine-elicited repertoire. Based on the presence of a conserved glutamine residue within the CDR1α (Supplementary Fig. 9e), it is possible that contact between R5 and the overrepresented TRAV12 allele may be a key driving force behind recruitment of the vaccine-elicited repertoire. Evaluation of more TCR structures, however, will be imperative to fully understand the importance of the R5 residue to antigen recognition. Finally, F7 and L8 were proximal to TCRβ, but did not durably interact (Fig. 4d, e, Supplementary Fig. 9e). Taken together with the high degree of diversity in TCRβ (Fig. 2e, Supplementary Fig. 5c), it is possible that structural flexibility at F7 and L8 confers elevated diversity and consequent mutation coverage within the COV-S-082-specific repertoire. In line with this speculation, residues proximal to TCRβ were the most variable in their binding patterns (Supplementary Fig. 9f), but this variability could be reduced by drilling down on TCRs with similar TCRβ sequences (Supplementary Fig. 9g, h).
A gap in the vaccine-elicited TCR repertoire is filled by the naïve repertoire through distinct TCR features
We were particularly intrigued by the lack of mutation coverage at position five (Fig. 4a). Given the significant protrusion of R5 into the TCR binding pocket (Fig. 4c), the high degree of stable contact with both TCRα and TCRβ (Fig. 4e), as well as the extreme preferential bias of TRAV12 usage in vaccine recipients (Supplementary Fig. 5b), we wondered if substitutions to R5 corresponded to a ‘hole’ in the T cell repertoire. To address this concern, we first asked if R5-mutant peptides could be presented on common non-HLA-A2 alleles, which would suggest their capacity to be immunogenic in a sizeable fraction of the population. We predicted their binding affinities using NetMHC, revealing minimal binding on non-HLA-A2 backgrounds (Supplementary Fig. 10a). Despite low alternative HLA presentation, we hypothesized the TCR repertoire may be able to cover the apparent hole through the HLA-A2-restricted naïve T cell repertoire. To evaluate the capacity of the naïve T cell repertoire in recognizing the R5-mutant peptides, we next expanded T cells in vitro from pre-2020 (vaccine-naïve) and post-2020 donors on the nineteen R5 APLs and evaluated their ability to bind to pMHC tetramers. We detected binding events from three donors, suggesting that this apparent ‘hole’ in the vaccinated T cell repertoire had the potential to be filled by the naïve T cell repertoire, albeit not likely through current vaccine strategies (Supplementary Fig. 10b).
To better understand the specific TCR features required to respond to R5 mutations, we sequenced pMHC tetramer+ TCRs from the three donors described above. As expected, the TCR sequences were distinct from vaccine-elicited COV-S-082-specific TCRs, including their lack of TRAV12 usage (Fig. 5a). Given their distinct TCR features, we hypothesized that their mutation coverage would be distinct as well. We thus generated cell lines expressing these R5 mutant specific TCRs and fingerprinted them (Fig. 5a). Confirming our hypothesis, the binding patterns of the R5 mutant specific TCRs were distinct from vaccine elicited repertoires (Fig. 4a). Together, they were able to fill much of the mutation space missed by the vaccine stimulated T cell repertoire, binding to 12 unique mutations across Q3, R5, and F7 (Fig. 5b). Furthermore, the extra mutation coverage offered by these three TCRs far exceeded the additional mutation coverage gained by any other TCR community (Fig. 5c), which was reflected in their poor correlation with other TCR binding patterns (|R| < 0.1 in all cases; Supplementary Fig. 7e), even relative to comparisons across distinct TCR communities (Supplementary Fig. 10c).
a TCR fingerprints (top) and characteristics (bottom) for R5-mutant specific TCRs, the wild type epitope is boxed in black. Each heatmap plot represents one experiment with 172 tetramers stained individually. b Boolean fingerprint after integration of R5-mutant specific TCRs and their unique mutation coverage. Boolean binding is represented by a Log2 fold-change > 1. c Fraction of unique mutation coverage (binding) by R5 mutant-specific TCRs and each TCR community. d Dose-response curves and calculated EC50 values for TCR:pMHC species above 50% of the maximum in pre-screen experiment (Supplementary Fig. 5j), each curve represents one experiment. e Spearman’s rank correlation of Log2 fold-change pMHC tetramer binding with EC50 values from (c). f pMHC tetramer binding of all 23 TCRs toward the most dominant COV-S-082 mutations currently in circulation. Mutation is indicated in the vertical axis, TCR clone is indicated in the horizontal axis.
We next asked how an R5 mutant specific TCR could differ structurally from wild-type specific TCRs, given the importance of the R5 residue in the TCR:pMHC binding pocket. MD simulation of SVAR-92 co-complexed with the R5G APL revealed a largely TCRβ-dependent mechanism, driven by H-bonds between T6 and two CDR3β residues (Supplementary Fig. 11a–e). This system was quite distinct from SVAR-14 co-complexed with wild type COV-S-082, where interactions between R5 and TCRα contributed substantially to binding (Fig. 4c–e, Supplementary Fig. 9e). Taken together, these analyses further advocate for the importance of TRAV12 in the vaccine-elicited repertoire, as well as a potential rationale to drive the repertoire away from TRAV12 usage in cases where R5 binding is undesirable.
Finally, given that a TCR’s ability to bind pMHC tetramer does not directly reflect biological function, we next evaluated the cellular function of several TCRs by transducing them into a Nur77 reporter line30 that we engineered to express human CD8. We first pre-screened SVAR-1 and the three R5 mutant-specific TCRs against the wild type, all 19 R5 APLs, and several control epitopes (Supplementary Fig. 12a, c). To more robustly evaluate functional sensitivity, we then titrated peptide species that were above 50% of the maximum signal at 10 µM and calculated EC50 values for TCR:pMHC pairs when appropriate (Fig. 5d, Supplementary Fig. 12b). The EC50 values for the peptides evaluated were well correlated with the Log2 fold-change pMHC tetramer binding values described throughout this work, suggesting our analyses on tetramer binding should be largely generalizable to cellular function (Fig. 5e). Furthermore, many of the R5 APL species were able to elicit a downregulation of CD3 in their cognate TCRs, suggesting relatively strong TCR activation by these pairs (Supplementary Fig. 12b, d).
The most abundant COV-S-082 mutations currently in circulation are covered by TCRs from either the vaccine-elicited or naïve repertoire
To address ongoing SARS-CoV-2 mutagenesis, we evaluated nearly 4 million spike glycoprotein sequences from the NCBI virus database31 and mapped them to the COV-S-082 wild-type sequence (Supplementary Fig. 12e). Four mutations (P4L, R5S, T5I, and P4S) emerged at higher frequencies than others. Interestingly, the P4L mutation was previously reported as an immune escape variant of concern that cannot be recognized by the large CD8 response seen in convalescent and vaccinated individuals32. The recurrence of this known escape mutation confirms that our analysis captures functionally meaningful immune-evasive substitutions. Upon mapping the binding strengths of the 23 TCRs evaluated in this study, as well as two other previously characterized mutations28, we observed coverage by at least one TCR for each mutation (Fig. 5f). While these mutations are not currently dominant in circulation, their cooperation with future mutations in the viral genome remains a possibility. The data we provide here may thus be of critical value to future vaccine efforts when evaluating the potential of novel vaccine candidates.
Discussion
Although the virulence of SARS-CoV-2 has decreased since the peak of the omicron wave, vaccine-resistant lineages continue to emerge33,34. New vaccine strategies will thus continue to be necessary to combat immune evasion. Furthermore, with the prevalence of month-long, chronic COVID-19 cases during post-acute sequelae of COVID-19, diversification of T cell epitopes remains a concern through preferential mutagenesis, enhancing cell-to-cell over individual-to-individual transmission35. Similar biases have been observed in other chronic infections, including HIV36,37, HCV38 and even congenital CMV infections39, where stable and transmissible mutation burdens on T cell epitopes, that are not typically mirrored in acute infections, can commonly persist. Thus, the emergence of T cell-evading mutations on immunodominant epitopes remains a possibility while COVID-19 cases remain elevated. Preemptive knowledge of how the TCR repertoire can resist these mutations will thus be of critical need in future vaccine design efforts.
We have used a high-throughput single-cell sequencing approach to evaluate antigen-specific CD8+ T cells in response to COVID-19 vaccination to address these concerns. Corroborating previous works, we found a strong immunodominant response toward a limited number of spike epitopes in roughly two-thirds of vaccine recipients, up to eight months after vaccination13,14. In contrast to others15,21, we observed primarily de novo immunity, with no pre-existing T cells expanding after vaccination. We suspect this discrepancy is largely due to differences in HLA backgrounds, as has been described40. We also observed minimal cross-reactivity toward SARS-CoV-2 VOC mutant epitopes, with most T cells targeting unmutated epitopes. These T cells, however, were diverse in both their phenotypes and TCR sequences, suggesting a potential functional and physical flexibility against SARS-CoV-2. We evaluated this potential by analyzing specific TCR binding modalities through APL pMHC tetramer libraries, revealing a high degree of TCR promiscuity to potential SARS-CoV-2 mutagenesis. Integration of multiple TCR binding modalities further expanded this paradigm. Application of these principles to both infection, vaccine design, and cell therapies could be beneficial for future therapeutic efforts.
Of particular interest was the minimal binding of the vaccine-elicited repertoire to substitutions on the R5 residue of the COV-S-082 epitope. Based on structural analyses, we hypothesize that this arginine was critical to its protrusion into the TCR:pMHC binding pocket, requiring a precise TCRα conformation for proper recognition. Such a restriction could be mediated through the incredibly prominent usage of TRAV12-related alleles we, and others14,15,40, have observed almost exclusively within vaccine-elicited COV-S-082-specific repertoires. Expanding on this paradigm, TCRs isolated from vaccine-naïve donors recognizing substitutions to R5 had distinct CDR3α sequences and TRAV usage. These data implicate the lack of antigen recognition toward R5 mutations as a vaccine, rather than a repertoire, intrinsic deficit. It is thus possible that modifications to the vaccine could provide protection against future escape mutations, like the ones described in this work, through the naïve repertoire.
A limitation of this study is its lack of infection history from several donors, which limited our ability to discern the true origin of some donor’s immune responses. Given that previous studies have pointed to specific differences in immunity elicited by vaccination versus vaccination and infection14,41,42, it would have been useful if we had detected discrete diversity or community differences between repertoires with unique immune challenge histories. Another lingering question is whether antigen-specific repertoires can maintain the phenotype and repertoire diversities we observed through both time and multiple challenges. Follow-up studies aimed at tracking TCR diversity longitudinally after both infection and vaccination will be prudent to delineating such a phenomenon.
It would also have been of great value to uncover relationships between the immunodominant repertoire diversity observed here and COVID-19 severity and/or resistance to breakthrough infections. Direct evaluation of antigen-specific T cells, however, poses a tremendous challenge during severe COVID-19. One study using large-scale single-cell transcriptomics did find lower TCR repertoire diversity in several naïve and non-naïve CD8+ T cell clusters in severe versus non-severe COVID-19 patients43. Although this study was completed before the use of COVID-19 vaccines, it suggests that diverse repertoires targeting SARS-CoV-2 may contribute to protective immunity. Further advocating this point, another more recent study found improved protection from Ovalbumin-expressing Listeria in mice engineered with diverse, rather than high-avidity mono or oligoclonal, repertoires targeting the SIINFEKL epitope44. They also directly showed enhanced protection from escape mutations during secondary immune responses through recruitment of distinct, low-avidity clones that were non-dominant in the primary response. Based on our data, it is possible that a similar phenomenon could occur during SARS-CoV-2 vaccination. In fact, HLA-B7-restricted nucleocapsid-specific CD8+ T cells have been shown to have a high degree of TCR diversity40, harbored in their promiscuous α/β pairing, and were independently revealed to be associated with less severe COVID-1945. More direct work evaluating the contribution of antigen-specific repertoire diversity to disease and vaccine outcomes, specifically in humans, will be important for applying and understanding this work’s direct practical and therapeutic potential. Knowledge of how different TCRs interact with both wild-type and mutant pMHC could better inform how TCR antigen recognition is mediated, a phenomenon that has long remained elusive in the field. A deeper understanding of such a fundamental principle that controls the behavior of the adaptive immune response will likely surface many future discoveries.
Methods
Sample collection, cohort, and study design
Six healthy adults with no self-reported chronic health conditions were recruited and consented in early 2021 for a longitudinal vaccine study conducted by Laura Su’s group at UPenn with approval from the University of Pennsylvania Institutional Review Board (IRB protocol no. 820884); all donors provided written informed consent for use of their samples in research. Participants were confirmed to have no prior COVID-19 exposure by serologic test and to carry the appropriate HLA-A alleles. All subjects received either Pfizer (BNT162b2) or Moderna (mRNA1273) mRNA vaccines. Leukapheresis samples were collected before vaccination and 176 to 256 days after the second vaccine dose. All samples used in this study were de-identified when transferred to the Jiang lab.
De-identified PBMC samples were also collected from thirteen donors with informed consent via leukoreduction system chambers obtained from We Are Blood (https://weareblood.org/), a local blood bank in Austin, Texas, which provides de-identified blood products and research samples. Because de-identified samples were used, this study does not meet the criteria of human subject research by UPenn IRB regulation, and thus, no IRB protocol is required. Donors were confirmed to have received a COVID-19 vaccine by survey, but their COVID-19 infection history was unknown. Three de-identified PBMC samples were also collected with informed consent at the Human Immunology Core at the University of Pennsylvania without COVID-19 vaccination or infection status obtained under an IRB protocol 705906 that was approved by the University of Pennsylvania Institutional Review Board.
A summary of donor information and metadata pertaining to this study can be found in Supplementary Table 2. No statistical method was used to predetermine sample size.
Cell lines
HEK293T cells (ATCC CRL-3216) were used to produce virus for TCR transductions. Cells were maintained in Dulbecco’s Modified Eagle Medium (DMEM) (10-013-CM, Corning) supplemented with 10% fetal bovine serum (FBS) (97068-091, Avantor) and 1% penicillin/streptomycin (15140122, Gibco) at 37 °C (5% CO2) and passaged every 2–3 days.
A Jurkat 76 (J76)46 cell line with a stable human CD8 knock-in (generously gifted by Dr. Beatrize Carreno) was used as the background to produce transgenic TCR cell lines in this study. Cells were maintained in RPMI 1640 (MT10041CM, Corning) supplemented with 10% FBS and 1% penicillin/streptomycin at 37 °C (5% CO2). Media was replenished every 2–3 days and cells were kept at a concentration between 5e5 and 2e6 cells/mL.
A TAP-deficient T2 (ATCC 174 x CEM.T2) cell line was used to evaluate pMHC stability. Cells were maintained in Iscove’s Modified Dulbecco’s Medium (IMDM, ATCC: 30-2005) supplemented with 20% FBS and 1% penicillin/streptomycin at 37 °C (5% CO2). Media was replenished every 2–3 days and cells were kept at a concentration between 5e5 and 2e6 cells/mL.
Mutation analysis of spike glycoprotein sequences and epitope selection
Complete spike glycoprotein sequences were downloaded from NCBI Virus for AY.103, AY.44, B.1.1.519, B.1.1.7, B.1.351, B.1.617.2, BA.1, BA.2, and P.1 VOC and VOI lineages. After multiple sequence alignment, a consensus sequence was built, and mutations were denoted as mismatches to the ancestral reference (refseq: YP_009724390.1).
Immunodominant CD8+ T cell epitopes were curated from previously published works14,15,16,17 and the Immune Epitope Database (IEDB)18. VOC and VOI lineage-derived antigens were also specifically included for mutations that were predicted by NetMHC-pan 4.119 to be strong binders (rank ≤ 0.5%). In these cases, epitopes from all lineages and the wildtype at the mutation position were included.
Production of DNA-barcoded pMHC tetramers
Crude peptides were synthesized from either Genscript or JPT. Peptide-UV exchange was performed as previously described20. DNA-barcoded tetramers were generated as previously described20 with slight modifications. To reduce non-specific cell binding, 1 molar equivalent of S-HyNic (S-1002, TriLink Biotechnologies) was used during the modification of PE- or APC-labeled streptavidin (405245 or 405243, BioLegend). The barcode construct used was:
5’- ATGGACGACGACGACAAG [10 N] [20NT_BARCODE] TAACGAAGCACCTCGCT [25 A] -3’.
Final pMHC tetramers were generated by combining exchanged pMHC with DNA-barcoded streptavidin at 4 °C for ≥12 h with a minimum of 8X pMHC:streptavidin molar excess and used within one week of UV exchange.
To limit hybrid pMHC tetramer complexes, pMHC tetramers were combined at equimolar concentrations in the presence of excess biotin and always used within ≤ 1 h of pooling. The pool was then centrifuged at 10,000 x g for 2 min to precipitate out aggregates. The supernatant was then filtered, washed, and concentrated in a 100 kDa MWCO Amicon Ultra-centrifugal filter (UFC510096, MilliporeSigma).
Production of DNA-barcoded antibodies
Custom DNA-barcoded antibodies were covalently linked to α-CD57 antibodies using Light-Activated Site-Specific Conjugation (LASIC). Specifically, pure α-CD57 (clone REA769, 13-124-525, Miltenyi, undiluted) was conjugated to the sequence 5’-GACGTGTGCTCTTCCGATCT VNNVNNNNVN GTTAGGTTGTCGAGTGTATTTAGGTT AAAAAAAAAAAAAAAAAAAAAA*A*A -3’ using oYo-Link chemistry (oYo-Link Oligo Custom, AlphaThera) following the manufacturer’s instructions. 1 µg of conjugated antibody was used to stain cells.
Preparation and sorting cells for Rhapsody
PBMCs were collected from leukocyte reduction system (LRS) chambers and isolated using Lymphoprep density-gradient centrifugation (NC0424924, Stemcell). PBMC samples were then stored at −150 °C until use. Upon thaw, CD8+ T cells were purified using EasySep Human CD8+ T cell isolation kits (19053, Stemcell Technologies).
CD8+ T cells were prepared for FACS as previously described20 and stained with the AbSeq and fluorescent antibody panels described in Supplementary Table 7.
Live, Tetramer positive, CD8+CD4−CD19−CD14− cells from each sample were sorted on an AriaFusion (BD Biosciences) as previously described20. Tetramer-negative, CD8+CD4−CD19−CD14− cells were also sorted from each sample into a single tube as a negative control. Sorted cells were then loaded onto a Rhapsody Cartridge (BD) following the manufacturer’s instructions.
Preparation of TetTCR-SeqHD sequencing libraries
Single-cell sequencing libraries were prepared according to the manufacturer’s instructions with some modifications. To amplify pMHC tetramer DNA-barcodes, an additional reverse primer was added during PCR1 and PCR2 as described previously20 (Supplementary Table 8).
Amplification of TCR libraries was modified to improve single-cell capture. Briefly, PCR2 was carried out using custom TCR primers (Supplementary Table 8) and the following thermocycler program: 95 °C denature for 3 min, then 15 cycles of 95 °C for 30 s, 60 °C for 30 s, 72 °C for 1 min, then a final extension at 72 °C for 5 min. The PCR2 product was then purified using Ampure XP beads (A63881, Agencourt). Index PCR was then carried out using 10.5 ng of the purified PCR2 product and the following thermocycler program: 95 °C denature for 3 min, then 7 cycles of 95 °C for 30 s, 60 °C for 30 s, 72 °C for 1 min, then a final extension at 72 °C for 5 min.
A custom-targeted mRNA panel designed by BD was also used to capture mRNA expression.
All final libraries were quantified on a TapeStation 2100 system (Agilent) before pooling. AbSeq, SampleTag, Tetramer, and mRNA libraries were sequenced either on a HiSeq X with a 150 × 150 configuration or a Novaseq SP with a 75 × 225 configuration. TCR libraries were sequenced on either a Miseq V2 with a 75 × 225 configuration or a Novaseq SP with a 75 × 225 configuration. All sequencing runs used 15–20% PhiX.
TetTCR-SeqHD reads pre-processing
Sequencing reads from AbSeq, SampleTag, mRNA, and TCR libraries were pre-processed using BD Rhapsody Targeted Analysis Pipeline (V 1.11.1) on the Seven Bridges platform following the manufacturer’s instructions. Tetramer sequencing libraries were pre-processed as previously described20.
Dimensional reduction and clustering of single cells
Cells from each experiment were filtered based on the distribution of detected mRNA molecules within each cell for that experiment (Supplementary Table 9). Cells identified as multiplets or undetermined by SampleTag calls were also removed. Protein and mRNA expression from each experiment were then combined and used to perform joint probabilistic modeling with totalVI47 to infer protein expression of missing markers in some experiments and implement batch correction (Supplementary Table 7). Combined matrices of background-corrected mRNA and protein expression were then used to perform dimensionality reduction (UMAP algorithm) in Seurat V448. Cells were clustered using the Louvain algorithm in Seurat.
Differential expression analysis
Differential features from Supplementary Fig. 1 were computed using the R package Seurat48. Significance for mRNA features was computed using a negative binomial test on the UMI count data. Significance for AbSeq features was computed using a Wilcoxon Rank Sum test on the totalVI background-corrected data. For Supplementary Fig. 1b, the top 5 most positive and negative significant features of each cluster were plotted. The R package ‘pheatmap’ was used for visualization.
Assigning putative epitope specificities for cells and TCR clonotypes
After tabulating a pMHC tetramer barcode molecule count matrix for each cell, tetramer calls for each cell were established based on the principles of cell hashing described in Stoeckius et al.49. Briefly, after normalization, cells were clustered based on their tetramer UMI counts using k-medoid clustering. The background signal for each tetramer was then modeled using a negative binomial distribution, where cells with counts above the 99th quantile were considered positive for that species. Cells identified as having 2–3 species above background were marked to be possibly cross-reactive, requiring manual inspection to verify. Cells with > 3 species were labeled as ‘sticky’. A second iterative step was then implemented to rescue ‘sticky’ cells if they had a high signal to noise ratio (SNR)50.
Based on a distribution, the cutoff for this ratio was set at SNR > 3. Unless specified otherwise, these higher signal cells were then considered specific to only their top specificity. Finally, a ‘clone specificity’ was determined within groups of cells from each sample that shared CDR3 amino acid sequences for both TCRα and TCRβ. For that, clonotypes were given a specificity if > 50% of cells shared a single specificity call.
TCR and V-gene diversity
TCR, antigen, and V-region diversity were calculated using normalized Shannon entropy (NSE). For TCR diversity, a unique species was a group of cells from the same sample with the same amino acid sequence for both TCRα and TCRβ. For antigen diversity, a unique species was a group of cells from the same sample with the same specificity call. For V-region diversity, a unique species was a group of cells with the same V-gene usage.
Select data used to calculate NSE in Fig. 3a was procured from this work and others14,15,20.
TCR frequency calculation
To correct for differences in Rhapsody cell input for each sample, the TCR frequency for spike-specific cells was calculated as follows:
Where \(\frac{{{{\rm{f}}}}_{{{\rm{clone}}}}}{{{{\rm{f}}}}_{{{\rm{paired}}}}}\) is the ratio of cells with a given TCR clonotype to the total number of cells in the sample with paired TCR chains, fflow is the frequency of spike tetramer+ flow cytometry events to total CD8+ events, and fsort is the ratio of spike tetramer+ cells sorted to the total cells sorted for a given sample.
Transitional index (ITRAN) of SARS-CoV-2-specific cells
The transitional capacity (ITRAN) of SARS-CoV-2-specific cells was determined as described previously by single T cell analysis by RNA sequencing and TCR tracking (STARTRAC)24. Briefly,
in which \({p}_{{cls}}^{t}\) is the ratio of the number of cells with clonotype t in cluster cls to the total number of cells in cluster cls, and
Here, \({p}_{k}^{t}\) is the ratio of the number of cells with TCR clonotype t in cluster k to the total number of clusters with TCR clonotype t. Each TCR clonotype pertains to a group of cells with the same amino acid sequence for both TCRα and TCRβ. ITRAN for single specificities (as in Figure 18E) was calculated similarly, except that clonotypes were further grouped by specificity.
TCR neighborhood and community analysis
TCR neighbors were defined by the pairwise sum of TCRα and TCRβ distances using the package tcrdist322. Likewise, only cells with putative clone specificities (see above) were used for these analyses. The connectivity (neighborhood) of a given TCR clonotype was defined by having a TCR distance < 60. For clearer visualization, clonotypes with connectivity in the top 99th percentile were also removed. Clusters (communities) were determined using k-means clustering. Plotting was done using the R package igraph51. Borders around select communities were manually drawn for visualization. Circos plots of V and J-gene usage were generated using the R package ‘circlize’52.
Unique TCR clonotypes within the communities were used to generate logo plots with the R package ‘ggseqlogo’53 for both TCRα and TCRβ CDR3 regions.
Production of transgenic TCR cell lines
TCR clonotypes (n > 1) with distinct properties were selected based on their TCR community assignment. TCRα and TCRβ consensus sequences were generated from single-cell sequencing data and synthesized into eBlocks (IDT). eBlocks (IDT) were introduced into a vector as previously described20 using HiFi assembly (NEB). Plasmids containing the correct TCR sequence were then transfected into HEK 293 T cells using ViraPower Lentiviral Packaging Kit (K497500, Thermo) following manufacturer’s instructions. Virus was harvested after 48 h and concentrated using an Amicon 30 kDa MWCO Ultra-Centrifugal Filter Unit (UFC803024, MiliporeSigma), treated with DNase I and used directly for transduction.
TCRs were transduced with lentivirus and 6 µg/ml polybrene (H9268-5G, Millipore Sigma) into a CD8-expressing J76 cell line54 (generously gifted by Dr. Beatrize Carreno) or a Nur77-GFP expressing J76 cell line30 (generously gifted by Dr. Paul Thomas). After a 4-day incubation, the virus was removed. After 7 days, cells were tested for murine TCR expression and antigen specificity via flow cytometry using an α-murine TCRβ antibody (109212, clone H57-597, BioLegend, diluted to 0.5 ug/ml) and their cognate pMHC tetramer, respectively. To note, SVAR-91 and SVAR-92 TCR cell lines were generated using human TCR variable region gene fragments synthesized by TwistBio (South San Francisco, CA). These fragments were assembled with human TCR constant gene fragments (TwistBio, CA) into a lentiviral transfer vector with an EF1a promoter55 (courtesy of Dr. Carl June). The resulting viruses were then packaged and transduced into J76 or Nur77-GFP cells, following the same protocol used for other TCRs.
TCR fingerprinting by sequencing and analysis
An APL pMHC tetramer library containing every possible single amino acid point mutation of the HLA-A*02:01-restricted immunodominant peptide described in this study, COV-S-082 (YLQPRTFLL), was synthesized and DNA barcoded and produced as described in “Methods”. The DNA barcode construct for these libraries was: 5’- ATGGACGACGACGACAAG [12 N] [20NT_BARCODE] [12 N] TAACGAAGCACCTCGCT -3’
Twenty-four DNA barcoded tetramers (PE) were randomly selected, pooled and concentrated as described in “Methods”. Transgenic TCR J76-CD8 cell lines were treated with dasatinib (11498–10, Thomas Scientific), then stained with the tetramer pool and AF488 α-CD8 (301006, clone RPA-T8, BioLegend, diluted to 0.5 ug/ml) on ice. In addition to COV-S-082 specific cells, a negative control cell line that does not bind to COV-S-082 was also included. APC α-murine TCR antibody (109211, clone H57-597, BioLegend, diluted to 0.5 ug/ml) was also added. Cells were then washed and stained with 7-AAD (420404, BioLegend, diluted to 0.5 ug/ml) for viability. Live, CD8+Murine TCR+ cells were sorted using an Aria Fusion (BD Biosciences).
Sorted cells were lysed in DNA/RNA Shield (R1200-25, Zymo Research) and treated with proteinase K (D3001-2-5, Zymo Research). DNA was then purified using a Quick-RNA Whole Blood kit (R1201, Zymo Research) without DNase I treatment.
First PCR was carried out on 50% of the purified eluate using Kapa HiFi (KK2601, Roche) and the following thermocycler program: 98 °C for 3 min, then 15 cycles of 98 °C for 30 s, 57 °C for 30 s, and 72 °C for 30 s, then a final extension at 72 °C for 5 min. Index PCR was then carried out on 2 ng of PCR1 product using Kapa HiFi and the following thermocycler program: 98 °C for 3 min, then 6 cycles of 98 °C for 30 s, 57 °C for 30 s, and 72 °C for 30 s, then a final extension at 72 °C for 5 min. See Supplementary Table 10 for list of primers. Index PCR products were then gel purified using a Zymoclean Gel DNA Recovery kit (D4007, Zymo Research) and quantified on a D1000 DNA screen tape (50675582, Agilent Technologies) and TapeStation 2100 System (Agilent Technologies). Final, purified libraries were sequenced at 1000–10,000 reads per cell on a Novaseq S4 lane (150 × 150 configuration).
Sequencing reads were trimmed for length and quality using cutadapt56. UMIs were then extracted, and reads were annotated using UMI Tools57. Annotated reads were then aligned using bowtie258. Erroneous UMIs were then filtered in parallel based on their reads per UMI distributions.
T2 MHC stabilization assay
A TAP-deficient HLA-A2-expressing lymphoblast cell lines (T2; 174 x CEM.T2) was purchased from ATCC. Cells were plated in 96-well plates in 50 μl of serum-free RPMI-1640 media and cultured for 18 h at 37 °C in the presence of 100 uM peptide. Negative controls were also included for each plate with an equivalent volume of DMSO. After incubation, cells were washed, then stained with α-HLA-A2 antibody (343306, clone BB7.2, BioLegend, diluted to 0.5 ug/ml). Antibody was then washed, and cells were analyzed by flow cytometry on an Accuri C-Sampler Plus (BD Biosciences).
MHC stabilization for each peptide was then calculated as the log2 fold-change (Log2_FC) from background (DMSO control) and represented as the mean of two replicates. Based on the Log2_FC distribution, a stabilization signal of 1.5X over background (an approximate Log2_FC of 0.58) was used to classify MHC binders from non-binders.
TCR fingerprinting by flow cytometry and analysis
An altered ligand pMHC tetramer library containing every possible single amino acid point mutation of the HLA-A*02:01-restricted immunodominant peptide described in this study, COV-S-082 (YLQPRTFLL), was synthesized and produced as in TetTCR-SeqHD experiments, except that streptavidin was not modified and DNA-barcoded. TCR-expressing J76 cell lines were then stained individually with PE pMHC and APC α-murine TCR antibody (109211, clone H57-597, BioLegend, diluted to 0.5 ug/ml). Cells were then washed and run on an Accuri C-Sampler (BD Biosciences). To control batch variability, a unique background stain experiment on the background J76-CD8 cell line was run for each pMHC tetramer batch produced.
Binding to each pMHC mutant was quantified by drilling down on murine TCR+ cells, then calculating the log2 fold-change from background staining. Heatmaps were produced using the R package ‘pheatmap’. Other plotting was done using the R package ‘ggplot2’59.
Molecular dynamics simulations
The complexes of SG3 and SVAR-14 TCRs and HLA-A*02:01 loaded with COV-S-082 (YLQPRTFLL) epitope were adopted to perform molecular dynamics (MD) simulations. The structure of SG3:pMHC (PDB: 7RTR29) was used as the starting point, which is referred to as SG3 for short. The SVAR-14:pMHC complex was generated based on SG3 with the MUTATE plugin in Visual Molecular Dynamics (VMD)60. All models were processed with the VMD PSFGEN plugin to add missing atoms. The resulting systems were solvated in rectangular water boxes with the TIP3P water model. Na+ and Cl- ions were then added to these solvated systems to neutralize the systems and maintain salt concentration at ~150 mM. The final systems contained ~0.13 million atoms in total.
All systems were first equilibrated with four steps: (1) 10,000 step energy minimization with heavy atoms fixed, followed by 2 ns equilibration simulations under a 1 fs timestep constrained by 5.0 kcal/mol/Å2 spring; (2) 10,000 step energy minimization with heavy atoms fixed, followed by 2 ns equilibration simulations under 1 fs timestep constrained by 1.0 kcal/mol/Å2 spring; (3) 4 ns equilibration simulation under 1 fs timestep with heavy atoms constrained by 0.2 kcal/mol/Å2 spring; (4) 10 ns equilibration simulation under 1 fs timestep without constraints. Subsequently, more than ~600 ns production simulations were carried out with 2 fs time steps under rigid bond algorithms. Snapshots were saved every 40 ps for further analysis. The last 200 ns MD trajectories were used to calculate the geometries of TCR:pMHC. During the simulations, the temperature of each system was maintained at 310 K with Langevin dynamics, and the pressure was controlled at 1 atm with the Nosé-Hoover Langevin piston method. Particle Ewald Mesh summation was used for electrostatic calculation, and a 12 Å cutoff with 10 to 12 Å smooth switching was used for short-range non-bounded interactions. All simulations were performed with NAMD60 software using CHARMM36m force field with the CMAP correction. The system preparations, trajectory analyses, and illustration renderings were conducted with VMD. Residue contact (Fig. 4e, Supplementary Fig. 5g) was defined as the distance between the heavy atoms of the residue sidechain being less than 0.4 nm.
In vitro expansion of R5 mutant-specific CD8+ T cells
Primary CD8+ T cells from COVID-19 naïve donors were isolated from PBMCs through negative enrichment (19053, EasySepHuman CD8+ T cell Isolation Kit, Stemcell Technologies). From the same sample, primary B cells were also enriched (19054, EasySep Human B cell Enrichment Kit, Stemcell Technologies). One million B cells were then incubated with each of the 19-position five mutant peptides (1 μM each) for one hour in the presence of 0.5 μg/mL multimeric CD40L (AG-40B-0010-C010, AdipoGen Life Sciences), 40 ng/mL IL4 (130-093-917), and 50 ng/mL IL21 (130-095-767). The B cells and one million CD8+ T cells were then combined and co-cultured with an additional 200 U/mL IL2 (5-14-19, Miltenyi) for 3 days, after which a high dosage of IL2 (6000 U/mL) was added to promote T cell growth every other day. After 7 days, the cultures were re-stimulated with peptide, multimeric CD40L, IL4, and IL21 and the process was repeated for another 7 days. At day 14, the cultures were stained with position five mutant pMHC tetramers and live CD8+CD19- tetramer positive cells were single cell sorted directly into 4 μl of lysis buffer (1X PCR buffer II (Thermo Fisher Scientific), 1.35 mM MgCl2, 0.45% NP-40, 4.5 mM DTT, 0.72 U/μl RNase OUT (Invitrogen), and 0.045 mM dNTPs) (PCR kit) on an AriaFusion (BD Biosciences). Lysates were then snap frozen on dry ice and stored at −80 °C until use.
Single-cell TCR sequencing and analysis
TCR sequences were obtained from single-cell lysates as previously described61. Briefly, single T lysates were thawed at 4 °C. Primers binding the constant regions of both TCRα and TCRβ were then added for reverse transcription. PCR1 was then carried out using the same constant region primers, along with TRBV and TRAV primers (Supplementary Tables 11, 12). During PCR2, TCRα and TCRβ chains were amplified separately using nested V and C-region primers. Additionally, C-region primers in this step contained 8-nucleotide cell barcode sequences to increase multiplexing of cells during sequencing. Illumina adapters were then appended to the final libraries during Index PCR. Cells were pooled into TCRa and TCRb pools, run on 2% E-gels (G401002, ThermoFisher Scientific), then gel extracted using a Zymoclean DNA gel extraction kit (D4007, Zymo Research). Final libraries were pooled and sequenced at 1000 reads per cell on a HiSeq X 2 × 150 configuration.
After sequencing, cell barcodes were extracted from sequencing libraries using UMI Tools57. Sequences from each cell were then aligned and annotated using the MixCR package62.
Functional reporter assays
50,000 T2 cells were cultured in serum-free RPMI-1640 and pulsed with peptides at various concentrations for three hours. Cells were washed once, then co-cultured at 1:1 with transgenic TCR Nur77-GFP J76 reporter cell lines for four hours. Cells were then washed and stained with αCD3-PE (317308, Biolegend, diluted to 0.5 ug/ml), αCD19-BV785 (302240, Biolegend, diluted to 0.5 ug/ml), and zombie violet dye (423113, Biolegend, 1:1000 dilution). Cells were washed again, then evaluated flow cytometrically on an Attune NxT Flow Cytometer (ThermoFisher Scientific).
Data visualization
Data were visualized through various R packages, as reported in their respective sections and ggplot2. Final figures were compiled using BioRender.com.
Quantification and statistical analyses
Statistical analyses were performed using the various R packages as indicated in their respective sections. Statistical tests were used as indicated in the figure legends. Sample sizes were determined based on sample availability. ANOVA and Student’s T-test were used to compare multiple groups and between two groups, respectively.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
De-identified human single-cell RNA-seq data have been deposited in the Gene Expression Omnibus (GEO) under the accession GSE246491. In addition, processed datasets derived from these data have also been deposited under the same accession. Flow cytometry data have been deposited in the ImmPort database under the accession SDY3323. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request. All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. Source data are provided with this paper.
Code availability
All original code has been deposited and made publicly available at https://github.com/JiangLabSysImmune/TetTCR-SeqHD_V2. Source data are provided as a Source Data file.
References
Tscharke, D. C., Croft, N. P., Doherty, P. C. & La Gruta, N. L. Sizing up the key determinants of the CD8(+) T cell response. Nat. Rev. Immunol. 15, 705–716 (2015).
Yewdell, J. W. Confronting complexity: real-world immunodominance in antiviral CD8+ T cell responses. Immunity 25, 533–543 (2006).
Planas, D. et al. Considerable escape of SARS-CoV-2 Omicron to antibody neutralization. Nature 602, 671–675 (2022).
Kurhade, C. et al. Low neutralization of SARS-CoV-2 Omicron BA.2.75.2, BQ.1.1 and XBB.1 by parental mRNA vaccine or a BA.5 bivalent booster. Nat. Med 29, 344–347 (2023).
Wang, Q. et al. Antibody response to Omicron BA.4-BA.5 bivalent booster. N. Engl. J. Med 388, 567–569 (2023).
Bange, E. M. et al. CD8(+) T cells contribute to survival in patients with COVID-19 and hematologic cancer. Nat. Med 27, 1280–1289 (2021).
Brasu, N. et al. Memory CD8(+) T cell diversity and B cell responses correlate with protection against SARS-CoV-2 following mRNA vaccination. Nat. Immunol. 23, 1445–1456 (2022).
GeurtsvanKessel, C. H. et al. Divergent SARS-CoV-2 Omicron-reactive T and B cell responses in COVID-19 vaccine recipients. Sci. Immunol. 7, eabo2202 (2022).
Gao, Y. et al. Ancestral SARS-CoV-2-specific T cells cross-recognize the Omicron variant. Nat. Med 28, 472–476 (2022).
Tarke, A. et al. SARS-CoV-2 vaccination induces immunological T cell memory able to cross-recognize variants from alpha to Omicron. Cell 185, 847–859 e811 (2022).
Mateus, J. et al. Selective and cross-reactive SARS-CoV-2 T cell epitopes in unexposed humans. Science 370, 89–94 (2020).
Grifoni, A. et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell 181, 1489–1501 e1415 (2020).
Goel, R. R. et al. mRNA vaccines induce durable immune memory to SARS-CoV-2 and variants of concern. Science 374, abm0829 (2021).
Minervina, A. A. et al. SARS-CoV-2 antigen exposure history shapes phenotypes and specificity of memory CD8(+) T cells. Nat. Immunol. 23, 781–790 (2022).
Francis, J. M. et al. Allelic variation in class I HLA determines CD8(+) T cell repertoire shape and cross-reactive memory responses to SARS-CoV-2. Sci. Immunol. 7, eabk3070 (2022).
Saini, S. K. et al. SARS-CoV-2 genome-wide T cell epitope mapping reveals immunodominance and substantial CD8(+) T cell activation in COVID-19 patients. Sci. Immunol. 6, https://doi.org/10.1126/sciimmunol.abf7550 (2021).
Ferretti, A. P. et al. Unbiased screens show CD8(+) T cells of COVID-19 patients recognize shared epitopes in SARS-CoV-2 that largely reside outside the spike protein. Immunity 53, 1095–1107 e1093 (2020).
Vita, R. et al. The Immune Epitope Database (IEDB): 2018 update. Nucleic Acids Res 47, D339–D343 (2019).
Reynisson, B., Alvarez, B., Paul, S., Peters, B. & Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 48, W449–W454 (2020).
Ma, K. Y. et al. High-throughput and high-dimensional single-cell analysis of antigen-specific CD8(+) T cells. Nat. Immunol. 22, 1590–1598 (2021).
Schulien, I. et al. Characterization of pre-existing and induced SARS-CoV-2-specific CD8(+) T cells. Nat. Med. 27, 78–85 (2021).
Mayer-Blackwell, K. et al. TCR meta-clonotypes for biomarker discovery with tcrdist3 enabled identification of public, HLA-restricted clusters of SARS-CoV-2 TCRs. Elife 10, https://doi.org/10.7554/eLife.68605 (2021).
Wendel, B. S. et al. The receptor repertoire and functional profile of follicular T cells in HIV-infected lymph nodes. Sci. Immunol. 3, https://doi.org/10.1126/sciimmunol.aan8884 (2018).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Bentzen, A. K. et al. T cell receptor fingerprinting enables in-depth characterization of the interactions governing recognition of peptide-MHC complexes. Nat. Biotechnol. https://doi.org/10.1038/nbt.4303 (2018).
Ross, P., Holmes, J. C., Gojanovich, G. S. & Hess, P. R. A cell-based MHC stabilization assay for the detection of peptide binding to the canine classical class I molecule, DLA-88. Vet. Immunol. Immunopathol. 150, 206–212 (2012).
Hobohm, U. & Meyerhans, A. A pattern search method for putative anchor residues in T cell epitopes. Eur. J. Immunol. 23, 1271–1276 (1993).
Agerer, B. et al. SARS-CoV-2 mutations in MHC-I-restricted epitopes evade CD8(+) T cell responses. Sci. Immunol. 6, https://doi.org/10.1126/sciimmunol.abg6461 (2021).
Szeto, C. et al. Molecular basis of a dominant SARS-CoV-2 spike-derived epitope presented by HLA-A*02:01 recognised by a public TCR. Cells 10, https://doi.org/10.3390/cells10102646 (2021).
Guo, X. Z. et al. Rapid cloning, expression, and functional characterization of paired alphabeta and gammadelta T-cell receptor chains from single-cell analysis. Mol. Ther. Methods Clin. Dev. 3, 15054 (2016).
NCBI Virus, Bethesda (MD): National Library of Medicine (US), National Center for Biotechnology Information; 2004-[cited 2024]. Available from https://www.ncbi.nlm.nih.gov/labs/virus/vssi/#/.
Dolton, G. et al. Emergence of immune escape at the dominant SARS-CoV-2 killer T cell epitope. Cell 185, 2936–2951.e2919 (2022).
Kaku, Y. et al. Antiviral efficacy of the SARS-CoV-2 XBB breakthrough infection sera against Omicron subvariants, including EG.5. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(23)00553-4 (2023).
Rasmussen, M. et al. First cases of SARS-CoV-2 BA.2.86 in Denmark, 2023. Euro. Surveill. 28, https://doi.org/10.2807/1560-7917.ES.2023.28.36.2300460 (2023).
Wilkinson, S. A. J. et al. Recurrent SARS-CoV-2 mutations in immunodeficient patients. Virus Evol. 8, veac050 (2022).
Goulder, P. J. et al. Evolution and transmission of stable CTL escape mutations in HIV infection. Nature 412, 334–338 (2001).
Goepfert, P. A. et al. Transmission of HIV-1 Gag immune escape mutations is associated with reduced viral load in linked recipients. J. Exp. Med. 205, 1009–1017 (2008).
Neumann-Haefelin, C. & Thimme, R. Adaptive immune responses in hepatitis C virus infection. Curr. Top. Microbiol. Immunol. 369, 243–262 (2013).
Materne, E. C. et al. Cytomegalovirus-specific T cell epitope recognition in congenital cytomegalovirus mother-infant pairs. Front. Immunol. 11, 568217 (2020).
Nguyen, T. H. O. et al. CD8(+) T cells specific for an immunodominant SARS-CoV-2 nucleocapsid epitope display high naive precursor frequency and TCR promiscuity. Immunity 54, 1066–1082 e1065 (2021).
Reynolds, C. J. et al. Immune boosting by B.1.1.529 (Omicron) depends on previous SARS-CoV-2 exposure. Science 377, eabq1841 (2022).
Lin, D. Y. et al. Association of primary and booster vaccination and prior infection with SARS-CoV-2 infection and severe COVID-19 outcomes. JAMA 328, 1415–1426 (2022).
Ren, X. et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 1895–1913.e1819 (2021).
Straub, A. et al. Recruitment of epitope-specific T cell clones with a low-avidity threshold supports efficacy against mutational escape upon re-infection. Immunity 56, 1269–1284.e1266 (2023).
Peng, Y. et al. An immunodominant NP(105-113)-B*07:02 cytotoxic T cell response controls viral replication and is associated with less severe COVID-19 disease. Nat. Immunol. 23, 50–61 (2022).
Heemskerk, M. H. et al. Redirection of antileukemic reactivity of peripheral T lymphocytes using gene transfer of minor histocompatibility antigen HA-2-specific T-cell receptor complexes expressing a conserved alpha joining region. Blood 102, 3530–3540 (2003).
Gayoso, A. et al. Joint probabilistic modeling of single-cell multi-omic data with totalVI. Nat. Methods 18, 272–282 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Stoeckius, M. et al. Cell Hashing with barcoded antibodies enables multiplexing and doublet detection for single-cell genomics. Genome Biol. 19, 224 (2018).
Ma, K.-Y. et al. High-throughput and high-dimensional single-cell analysis of antigen-specific CD8+ T cells. Nat. Immunol. 22, 1590–1598 (2021).
Nepusz, G. C. a. T. The igraph software package for complex network research. InterJournal Complex Systems, 1695 (2006).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize Implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Wagih, O. ggseqlogo: a versatile R package for drawing sequence logos. Bioinformatics 33, 3645–3647 (2017).
Bear, A. S. et al. Natural TCRs targeting KRASG12V display fine specificity and sensitivity to human solid tumors. J. Clin. Invest. 134, https://doi.org/10.1172/JCI175790 (2024).
Posey, A. D. Jr. et al. Engineered CAR T cells targeting the cancer-associated tn-glycoform of the membrane mucin MUC1 control adenocarcinoma. Immunity 44, 1444–1454 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in Unique Molecular Identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Wickam, H. ggplot2: elegant graphics for data analysis. Springer-Verlag (2016).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph 14, 27–38 (1996). 33−38.
Zhang, S. Q. et al. Direct measurement of T cell receptor affinity and sequence from naive antiviral T cells. Sci. Transl. Med. 8, 341ra377 (2016).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Acknowledgments
We thank Dr. John Wherry and Dr. David Weiner for input and discussion; Dr. Beatrize Carreno for sharing the CD8-expressing J76 cell line; Dr. Paul Thomas for sharing the J76-Nur77 reporter line; the NIH Tetramer core for providing UV-cleavable MHC; the anonymous blood donors and staff at We Are Blood in Austin Texas; Lynn Chen, Max Eldabbas, and Emileigh Maddox of the Human Immunology Core (HIC) at the Perelman School of Medicine at the University of Pennsylvania for providing primary immune cells. This work was supported by NIH U19AI128949 (N.J.), R33CA256086 (N.J.), NIH R01AI66358 (L.F.S), and NFSC T2394512 (Y.Z). The HIC is supported in part by NIH P30 AI045008 and P30 CA016520. HIC RRID: SCR_022380.
Author information
Authors and Affiliations
Contributions
M.J.M. and N.J. conceived and designed the study and interpreted the data. M.J.M. designed and performed the majority of the experiments, analyzed the data, plotted figures, and wrote the manuscript. C.H. discovered and analyzed R5-mutant specific TCRs, established and performed the T2 and Nur77 reporter cell co-culture assays, analyzed mutations of YLQ epitope in circulating viruses, finalized manuscript revision and submission. Y.Z. performed MD simulation, performed related data analysis, and generated related figures. Y.Q. helped with some of the TCR transductions and some preliminary experiments. L.W. processed blood samples and helped with subject recruitment. L.F.S. led the IRB protocol and shared samples. J.L. supervised the MD simulation and related data interpretation. N.J. supervised the study. All authors contributed to manuscript revisions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Katherine Kedzierska and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Malone, M.J., Huang, C., Zhang, Y. et al. Resistance potential of the HLA-A2-restricted immunodominant SARS-CoV-2-specific CD8+ T cell receptor repertoire to antigenic drift. Nat Commun 17, 686 (2026). https://doi.org/10.1038/s41467-025-67336-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67336-w