Abstract
PIK3CA mutation is frequently concurrent with known oncogenic drivers such as EGFR mutation in lung cancer, raising an interesting question about its real function. Cachexia is a systemic disease arising from tumor-organ crosstalk, significantly contributing to cancer-related mortality. Through integrative study of genetically engineered mouse models (GEMMs) and clinical data, we find concurrent PIK3CA mutant preferentially drives cachexia in EGFR-mutant lung cancer, promoting malignant progression instead of cancer initiation. PIK3CA mutant-mediated cachexia can be overcome by osimertinib (Osi) treatment in Osi-sensitive GEMM. In contrast, chemotherapy, routinely used in clinic for those relapsed from Osi therapy, fails to ameliorate cachexia in Osi-resistant GEMM despite notable tumor suppression. PIK3CA mutant-driven cachexia is mediated through NF-κB activation and can be dampened by combined aspirin treatment. This work provides insights into PIK3CA mutant biological function and mechanisms behind its clinical impacts, and proposes a potential strategy for clinical management.
Similar content being viewed by others
Introduction
Lung cancer is one of the most aggressive diseases worldwide, with non-small cell lung cancer (NSCLC) representing the most prevalent histological subtype1. PIK3CA, encoding the p110α catalytic subunit of PI3K, is mutated in approximately 7–16% of NSCLC2,3. PI3K plays a pivotal role in multiple tumorigenic processes in NSCLC, including promoting glucose transport4, facilitating bone metastasis5, mediating cytokine production6, and contributing to an immunosuppressive microenvironment by modulating immune cell infiltration7. PIK3CA mutations are highly enriched in its helical region (E542K, E545K) and kinase domain (H1047R)8. It’s well established that these mutations are oncogenic and frequently cause aberrant activation of downstream AKT signaling, triggering a cascade of responses that drive lung tumorigenesis9,10,11. Animal model study shows that PIK3CA mutant transgene alone is sufficient to drive lung cancer initiation in mice12. Paradoxically, most PIK3CA mutations are found to occur concurrently with those famous oncogenic drivers13,14,15, e.g., EGFR mutations, KRAS mutations, and ALK fusions, which are known to be mutually exclusive15,16,17,18 and individually suffice to drive lung cancer initiation19,20,21.
Multiple studies have reported that PIK3CA mutations are frequently concurrent with oncogenic EGFR mutations in East Asian NSCLC populations13,14,15. Osimertinib (Osi), a third-generation EGFR tyrosine kinase inhibitor (TKI), is currently the preferable option for EGFR-mutant NSCLC patients, owing to its high efficacy and well-tolerated safety profile22. However, similar to that of early-generation EGFR-TKIs, Osi resistance inevitably develops. Additional C797S mutation represents one of the most prevalent mechanisms in Osi resistance23. When occurring in conjunction with a sensitizing mutation and in the absence of T790M mutation, C797S mutation confers resistance to Osi while preserving sensitivity to first- and second-generation agents. The presence of triple mutants consisting of the sensitizing mutation, T790M, and C797S, leads to resistance against all three generations of EGFR TKIs24. For these patients, chemotherapy, e.g., pemetrexed (PEM) in combination with cisplatin (CDDP), is the remaining option in clinic25.
High-throughput sequencing analyses of TKI-resistant lung cancer specimens have identified the emergence of additional PIK3CA mutation26,27, indicative of its potential contribution to drug resistance. This is further supported by the observation of worse overall survival (OS) of those EGFR-mutant patients with concurrent PIK3CA mutation in their tumors28. However, detailed analyses of clinical TKI responses show that the concurrence of PIK3CA mutation does not affect the therapeutic efficacy of TKI treatments28,29, e.g., almost no difference of progression-free survival (PFS) in patients with or without PIK3CA mutations. This raises another interesting question regarding the real mechanism behinds the clinical impacts of PIK3CA mutant.
Cancer cachexia is a complex and debilitating syndrome characterized by body weight loss primarily due to skeletal muscle atrophy and adipose tissue wasting30, responsible for more than 20% of cancer-related deaths31. Cachexia occurs in approximately 50% of lung cancer patients32 and is associated with poor life quality33, elevated treatment-related toxicity34,35, reduced therapeutic responses36, and increased risk of mortality37,38,39,40. Recent study has begun to focus on the link between specific gene alterations and cachexia development37.
In this work, we uncover an unexpected biological function of PIK3CA mutations in lung tumorigenesis, mainly contributing to cachexia instead of driving lung cancer initiation. Moreover, we provide a reasonable explanation about the clinical observation of paradoxical difference between PFS and OS in link to concurrent PIK3CA mutation. We also propose a potential strategy for clinical management of lung cancer patients with concurrent PIK3CA mutation.
Results
PIK3CA mutant contributes to malignant progression but not lung cancer initiation
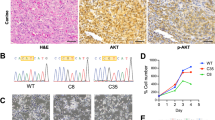
To investigate the real function of PIK3CA mutant in lung tumorigenesis, we generated a genetically engineered mouse model (GEMM) by integrating the LoxP-stop-LoxP-PIK3CA E545K transgene into the Rosa26 locus of C57BL/6 mice (Fig. S1a), in which the PIK3CA mutant can be conditionally expressed via intranasal inhalation of Ad-Cre virus41. Unexpectedly, we found no tumor formation even after 40 weeks of Ad-Cre administration (Fig. S1b, c). This indicates that PIK3CA mutant expression alone seems insufficient to drive lung cancer initiation. Given that PIK3CA mutation and TP53 mutation frequently co-occur in human NSCLC (Fig. S1d), we further crossed PIK3CAE545K to Trp53flox/flox mouse to see if there is any lung tumor formation. Again, we found no tumor formation in this model (Fig. S1e, f). These data collectively point to a dispensable role of PIK3CA mutant in driving lung cancer initiation.
It’s well established that either EGFR mutation, KRAS mutation, or ALK fusion alone suffice in driving lung cancer initiation in mice19,20,21. EGFR mutation is the oncogenic driver most significantly associated with the concurrence of PIK3CA mutation (Fig. S2a). We then crossed PIK3CAE545K mice with EGFRL858R;Trp53flox/flox (EP) mice to generate the EGFRL858R;PIK3CAE545K;Trp53flox/flox (EPP) cohort for further study (Fig. S2b). Both in vitro and in vivo assays demonstrate that PIK3CA mutant can activate the downstream AKT signaling and promotes lung cancer progression (Fig. S2c–k). Further analyses of in situ tumors showed that PIK3CA mutant significantly promoted lung cancer malignant progression, characterized by a greater degree of cellular pleomorphism and nuclear atypia42 (Fig. S2l, m). The tumor-bearing EPP mice exhibited significantly reduced survival compared to the EP mice (Fig. S2n). These data collectively demonstrate that PIK3CA mutant mainly contributes to lung cancer malignant progression instead of driving lung cancer initiation.
PIK3CA mutant drives cachexia in EGFR mutant lung cancer
Interestingly, the tumor-bearing EPP mice displayed a rapid decrease in body weight starting from 6 weeks post Ad-Cre treatment, which was only observed in EP control mice at late stage (Figs. 1a, b and S3a). Further analyses of the body composition revealed that the EPP mice experienced a continuous loss of lean mass along with a decrease in fat mass, aligning with the clinical diagnostic criteria for cachexia as previously established43 (Fig. S3b, c). Moreover, fat imaging visually illustrated a significant decrease of fat tissue in these tumor-bearing EPP mice (Fig. 1c). Compared to WT and EP mice, the EPP mice exhibited significant decrease in muscle mass in gastrocnemius (GAST), tibialis anterior (TA), and quadriceps (QUAD), as well as a decrease in epididymal white adipose tissue (eWAT) in males and the gonadal white adipose tissue (gWAT) in females (Fig. 1d, e). H&E staining analyses revealed the reduction of muscle fiber cross-sectional area (CSA) and adipocyte size in the EPP mice (Fig. S3d–h), histologically confirming these atrophic changes. Moreover, real-time PCR analysis confirmed increased expression of the muscle atrophy-associated marker Atrogin-1 in TA from EPP mice (Fig. 1f).
a Diagram of long-term monitoring strategy of mouse body weight and composition. Created in BioRender. Yue, M. (2026) https://BioRender.com/8v6qob5. b Relative body weight changes of WT, EP, and EPP mice following Ad-Cre treatment for 0–10 weeks (n = 12 mice/group). Data were normalized to baseline and analyzed at 10 weeks. c Fat imaging of WT, EP, and EPP mice post 8 weeks of Ad-Cre treatment. d, e Relative weights of the gastrocnemius (GAST), tibialis anterior (TA), quadriceps (QUAD) (d), and white adipose tissue (WAT) (e) in WT, EP, and EPP mice post 8 weeks of Ad-Cre treatment (n = 8 mice/group). f Real-time PCR detection of mRNA levels in tibialis anterior from WT, EP, and EPP mice post 8 weeks of Ad-Cre treatment (n = 5 mice/group). g–i 24-h activity counts (g), energy expenditure (EE) (h), and respiratory exchange ratio (RER) (i) of WT, EP, and EPP mice post 8 weeks of Ad-Cre treatment (n = 8, 8, 6 mice). j Tumor burden analysis in EP and EPP mice (n = 8 mice/group). k, l Relative weights of the gastrocnemius (GAST), tibialis anterior (TA), quadriceps (QUAD) (k), and white adipose tissue (WAT) (l) in non-tumor bearing WT, and tumor-bearing EP and EPP mice (n = 6, 8, 8 mice). m Bioinformatics analysis schematic based on TRACERx study (Al-Sawaf et al., 2023). Created in BioRender. Yue, M. (2026) https://BioRender.com/dg5w4mq. n Enrichment of cachexia candidate gene list in KEGG pathways analyzed by enrichR (Al-Sawaf et al., 2023). Dot size reflects the number of enriched genes; color indicates the significance of the enrichment. o KEGG pathways enriched in cancer-associated cachexia (CAC) patients vs. non-CAC patients in TRACERx study (Al-Sawaf et al., 2023). Data in (d, e, k, l) were normalized to WT mice of the same sex. Data in (b, d–l) are presented as mean ± SD. Statistical significance was calculated by one-way ANOVA followed by Tukey’s multiple comparisons test (b, d–f, k, l), two-tailed unpaired Student’s t test (j). Source data are provided as a Source Data file.
Metabolic monitoring indicated that EPP mice exhibited a significant decrease in locomotor activity, reduced energy expenditure (EE), and diminished food intake (Figs. 1g, h and S3i), all of which are hallmarks of cachexia in clinical settings32. Moreover, the lower respiratory exchange ratio (RER) in EPP mice indicated a metabolic shift toward the utilization of non-glucose substrates, potentially related to their decreased food intake (Fig. 1i). These characteristics were particularly pronounced at night, coinciding with the activity rhythms of mice (Fig. S3j–l).
To explore the potential impact of tumor burdens upon cachexia, we further analyzed the EP mice and EPP mice with comparable tumor burdens at different time points post Ad-Cre treatment, e.g., the EP mice post 17 weeks of Ad-Cre treatment and the EPP mice post 10 weeks of Ad-Cre treatment (Fig. 1j). Interestingly, a significant difference of cachexia features was still detectable in these two groups, e.g., the EPP mice showed lower levels of mass of skeletal muscles and adipose tissues, and increased Atrogin-1 expression in TA tissues (Figs. 1k, l and S3m). These data suggest that PIK3CA mutant contributes to cachexia through mechanisms other than by promoting tumor burden.
Analyses of cachexia signature proposed by TRACERx study37 highlighted the PI3K-AKT signaling as one of the most significantly enriched pathways (Fig. 1m, n). Compared to patients without cachexia, the PI3K-AKT signature was similarly marginally enriched in those primary tumors from patients with cachexia (Figs. 1o and S4a). Consistently, we observed that PI3K-AKT signaling was similarly enriched in cachectic individuals across two additional datasets of lung tumor RNA sequencing (RNA-seq)44 and serum proteomics45 (Fig. S4b, c). To validate our findings across other cancer types, we further performed Gene Set Enrichment Analysis (GSEA) on a pre-clinical melanoma model46, and observed enrichment of PI3K-AKT signaling in xenografts capable of inducing cachexia (Fig. S4d). These data collectively support the significant role of PI3K-AKT signaling in driving cachexia.
Muscle wasting characteristics in EPP mice recapitulate clinical phenomena
Muscle wasting is the most critical characteristic of cancer cachexia43. We next comparatively analyzed the gene expression profiling of TA derived from WT, EP, and EPP mice (Fig. 2a). Principal component analysis (PCA) revealed separation between TA from EPP mice and those from WT and EP mice (Fig. 2b). Consistently, gene expression pattern of EPP mice differed dramatically from the others (Fig. 2c). We found an enrichment of proteasome and autophagy signaling in TA of EPP mice (Fig. 2d), which has been extensively documented in muscle atrophy47. An enrichment of features associated with muscle disorder-related diseases was also observed, suggesting that muscle atrophy in cancer cachexia may share signaling pathways with other muscle disorders.
a Diagram of RNA sequencing of murine tibialis anterior tissues post 8 weeks of Ad-Cre treatment. Created in BioRender. Yue, M. (2026) https://BioRender.com/tnl1i4u. b Principal component analysis (PCA) plot of tibialis anterior tissues from WT, EP, and EPP mice (n = 4 mice/group). c Heatmap of Pearson correlation coefficient (PCC) among tibialis anterior tissues from WT, EP, and EPP mice (n = 4 mice/group). d Dot plot showing enriched KEGG pathways analyzed by enrichR within the tibialis anterior tissues of EPP vs. EP mice (n = 4 mice/group). The analysis was performed using differentially expressed genes (FDR < 0.05, fold change ≥ 1.5) based on hypergeometric test. Dot size reflects the number of enriched genes; color indicates the significance of the enrichment. e Database information and Venn diagram showing the number of upregulated genes in muscle tissue of cancer cachexia patients vs. non-cancer controls. f Gene expression heatmap of tibialis anterior tissues from WT, EP, and EPP mice (n = 4 mice/group).
We next analyzed two public databases (GSE13056348; GSE13352349) containing expression data from muscle tissues of cachectic patients and their respective controls. We found 42 genes upregulated in the muscles of cachectic patients, with 37 genes shared homologous counterparts in murine tissues with detectable expression (Fig. 2e). We further found that most of these genes were enriched in the muscle tissues of EPP mice (Fig. 2f), suggesting that the muscle wasting features in these mice closely resemble clinical observations.
Osi effectively inhibits lung cancer progression and alleviates cachexia in TKI-sensitive EGFR L858R GEMM with concurrent PIK3CA mutant
Previous studies indicated that PIK3CA mutant drives resistance to TKIs in EGFR-mutant lung cancer26,27. We then assessed the effect of single-agent Osi on mice carrying TKI-sensitive EGFR mutation (EGFRL858R) plus PIK3CA mutation (Fig. 3a). Osi treatment exhibited potent tumor-inhibitory effects, resulting in nearly complete tumor regression supported by dramatic decrease of tumor burdens and tumor numbers (Fig. 3b–d). In parallel, we observed increased weights in the TA, QUAD, GAST skeletal muscles, as well as in WAT tissues, indicative of the notable alleviation of cachexia (Fig. 3e, f). Moreover, Osi treatment mitigated atrophic changes in these tissues, as evidenced by an increase in muscle fiber CSA and adipocyte size (Fig. 3g–k), along with the downregulation of Atrogin-1 in TA tissues (Fig. 3l). These data suggest that, in the TKI-sensitive GEMM, tumor malignancy and cachexia mediated by the concurrent PIK3CA mutant could be effectively suppressed by Osi treatment.
a Diagram of osimertinib treatment strategy. Created in BioRender. Yue, M. (2026) https://BioRender.com/zy8rq63. b–d Representative H&E staining images of lung tissue (b), tumor burden (c), and tumor number (d) in EPP mice following vehicle (Veh) or osimertinib (Osi) gavage (n = 8 mice/group). Scale bar, 2 mm. e, f Relative weights of the gastrocnemius (GAST), tibialis anterior (TA), quadriceps (QUAD) (e), and white adipose tissue (WAT) (f) in non-tumor bearing WT mice without treatment (WT Ctrl) and EPP mice following vehicle (Veh) or osimertinib (Osi) gavage (n = 6, 8, 8 mice). g–i Representative micrographs (g), fiber cross-sectional area (CSA) (h), and fiber CSA distribution (i) of gastrocnemius in non-tumor bearing WT mice without treatment (WT Ctrl) and EPP mice following vehicle (Veh) or osimertinib (Osi) gavage (n = 10 fields/group). Scale bar, 50 μm. j, k Representative micrographs (j) and adipocyte size (k) of white adipose tissue in non-tumor bearing WT mice without treatment (WT Ctrl) and EPP mice following vehicle (Veh) or osimertinib (Osi) gavage (n = 10 fields/group). Scale bar, 50 μm. l Real-time PCR detection of mRNA levels in tibialis anterior from non-tumor bearing WT mice without treatment (WT Ctrl) and EPP mice following vehicle (Veh) or osimertinib (Osi) gavage (n = 6, 5, 5 mice). Data in (e, f) were normalized to WT mice of the same sex. Data in (c–f, h, k, l) are presented as mean ± SD, data in (i) are presented as mean. Statistical significance was calculated by two-tailed unpaired Student’s t test (c, d), one-way ANOVA followed by Tukey’s multiple comparisons test (e, f, h, k, l). Source data are provided as a Source Data file.
Chemotherapy inhibits tumor growth but fails to alleviate cachexia in Osi-resistant EGFR TLCS GEMM with concurrent PIK3CA mutant
Once relapses from Osi therapy, chemotherapy, e.g., PEM and CDDP, is routinely used as the conventional therapeutic strategy for patients25. We next developed EGFRTLCS mice carrying L858R, T790M, and C797S mutations (Fig. S5a), which confer resistance to all three generations of TKIs used in clinical practice24. We generated the EGFRTLCS;Trp53flox/flox (TLCS-P) and EGFRTLCS;PIK3CAE545K;Trp53flox/flox (TLCS-PP) cohorts for further study, and found PI3K-AKT signaling was significantly enriched in mouse lung tumors derived from TLCS-PP mice (Fig. S5b). Compared to the TLCS-P mice, TLCS-PP mice showed an early onset of weight loss (Figs. 4a, b and S5c). Cachexia development in these mice was further characterized by continuous reduction of both lean and fat mass (Fig. 4c, d), and histological wasting of muscle and adipose tissues (Fig. S5d–g).
a Diagram of long-term monitoring strategy of mouse body weight and composition. Created in BioRender. Yue, M. (2026) https://BioRender.com/6tpakmf. b–d Relative changes in body weight (b), lean mass (c), and fat mass (d) of WT, TLCS-P, and TLCS-PP mice over time following Ad-Cre treatment for 0–12 weeks (n = 10 mice/group). Data were normalized to baseline and analyzed at 12 weeks. e Diagram of PEM/CDDP treatment strategy. Created in BioRender. Yue, M. (2026) https://BioRender.com/woyyr6k. f, g Tumor burden (f) and tumor number (g) in TLCS-PP mice following intraperitoneal injection of vehicle or PEM/CDDP (Chemo) (n = 5, 7 mice). h, i Relative weights of the gastrocnemius (GAST), tibialis anterior (TA), quadriceps (QUAD) (h), and white adipose tissue (WAT) (i) in non-tumor bearing WT mice without treatment (WT Ctrl) and TLCS-PP mice following intraperitoneal injection of vehicle or PEM/CDDP (Chemo) (n = 6, 5, 7 mice). Data in (h, i) were normalized to WT mice of the same sex. Data in (b–d, f–i) are presented as mean ± SD. Statistical significance was calculated by one-way ANOVA followed by Tukey’s multiple comparisons test (b–d, h, i), two-tailed unpaired Student’s t test (f, g). Source data are provided as a Source Data file.
We next treated the TLCS-PP mice with the standard second-line chemotherapy regimen of PEM and CDDP (Fig. 4e). We found chemotherapy resulted in a significant reduction of both tumor burdens and numbers, indicative of its effectiveness in suppressing tumor progression (Fig. 4f, g). Consistent with the adverse effects of chemotherapy on cachexia50, the PEM/CDDP combination failed to exert protective effect against cachexia development, as evidenced by the reduced weights of skeletal muscle and adipose tissue and an increased trend of Atrogin-1 expression in TA tissues (Figs. 4h, i and S5h). These data suggest that the PIK3CA mutant-associated cachexia in Osi-resistant cases can’t be alleviated by chemotherapy despite notable tumor regression, emphasizing the need to explore the mechanisms underlying PIK3CA mutant in driving cachexia.
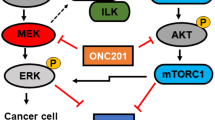
PIK3CA mutant drives cachectic inflammation through NF-κB activation
We then performed RNA-seq on mouse lung tumors derived from both Osi-sensitive and Osi-resistant GEMMs with or without PIK3CA mutant. Enrichment of epithelial-mesenchymal transition and inflammatory response pathways has been documented as a clinical feature of primary lung tumors from cachexic patients37. Interestingly, we observed the enrichment of both pathways in the EPP and TLCS-PP tumors compared to their respective controls (Figs. 5a and S5i, j). Moreover, the EPP and TLCS-PP tumors exhibited the enrichment of NF-κB signaling, which is known as the upstream regulator of multiple cachexia-associated pro-inflammatory factors51 (Figs. 5a and S5k). Upregulation of key factors in these tumors was further confirmed by RNA-seq and real-time PCR analyses (Figs. 5b–d and S5l). Elevated levels of IL-6 were also detected in the serum of EPP and TLCS-PP mice (Fig. S5m, n).
a Top 15 enriched Hallmark pathways analyzed by GSEA in EPP vs. EP tumors (n = 6 tumors/group). Dot size represents normalized enrichment score (NES), color indicates enrichment significance. b, c Gene expression heatmap of cachexia-associated pro-inflammatory factors in EP and EPP tumor (n = 6 tumors/group) (b) and TLCS-P and TLCS-PP tumor (n = 3 tumors/group) (c). d Real-time PCR detection of mRNA levels in indicated tumors (n = 10 tumors/group). e Representative immunofluorescence staining of EP cells expressing vector (Ctrl) or PIK3CA E545K mutant (E545K). p65 in green, nucleus in blue (DAPI). Scale bar, 25 μm. The assays were performed two times with similar results. f Immunoblotting of indicated proteins in EP and EPP tumors. The assays were performed two times with similar results. Numbers below the western blots indicate the relative quantification of band intensities. Dashed line separates lanes from different gels. g Heatmap of indicated gene expression in EP cell line expressing empty vector or PIK3CA E545K mutant (E545K) with or without 10 μM BAY 11-7082 treatment for 6 h. h Indicated gene expression in non-small cell lung cancer samples with high or low levels of PI3K-AKT signaling from The Cancer Genome Atlas (TCGA) database. (Low, n = 336 samples; High, n = 346 samples). i Kaplan–Meier overall survival (OS) of non-small cell lung cancer patients with high or low cachectic inflammation signature from The Cancer Genome Atlas (TCGA) database (Low, n = 191 patients; High, n = 194 patients). Data in d are presented as mean ± SD. Data in (h) are presented as box plot. The center line indicates the median. The lower and upper bounds of each box are the 25th and 75th percentiles (Q1 and Q3). Whiskers extend to the smallest and largest observations within 1.5 × IQR of the box (IQR = Q3–Q1); values beyond this range are plotted individually as outliers. The minima and maxima shown correspond to the ends of the whiskers (the most extreme non-outlying values). Statistical significance was calculated by two-tailed unpaired Student’s t test (d, h), log-rank (Mantel–Cox) test (i). Source data are provided as a Source Data file.
To further clarify the role of NF-κB signaling in this process, we performed immunostaining analyses on the EP cells with ectopic expression of PIK3CA mutant (Fig. S2d). We found that PIK3CA mutant notably increased the nuclear translocation of p65 (Fig. 5e). The enhancement of NF-κB activity was further confirmed by luciferase detection (Fig. S5o). Moreover, the p-p65 levels were elevated in EPP and TLCS-PP tumors (Figs. 5f and S5p). These data indicate that the PIK3CA mutant activates NF-κB pathway. Additionally, cachexia-associated factors were found upregulated following ectopic expression of PIK3CA mutant and attenuated by the treatment with NF-κB inhibitor BAY 11-7082 (Fig. 5g).
We further analyzed NSCLC patient data from The Cancer Genome Atlas (TCGA) database. Increased expression levels of cachexia-associated factors were observed in samples exhibiting elevated PI3K-AKT signaling (Fig. 5h). We next consolidated these factors into a cachectic inflammation signature, and found that a higher cachectic inflammation signature was associated with shorter patient overall survival (Fig. 5i). These data indicate that PIK3CA mutant triggers NF-κB signaling, leading to the upregulation of cachexia-associated pro-inflammatory factors, which are associated with poor prognosis of lung cancer patients.
Combined aspirin treatment attenuates cachexia in EGFR mutant lung cancer with concurrent PIK3CA mutant
Aspirin, a well established non-steroidal anti-inflammatory drug (NSAID), has been reported to inhibit the activity of NF-κB signaling52. To assess the effect of aspirin on inflammation inhibition, we treated EP tumor cells with aspirin in vitro and found that aspirin effectively suppressed NF-κB signaling and the pro-inflammatory factors upregulated by ectopic expression of PIK3CA mutant (Fig. S6a, b). We then treated the TLCS-PP mice with aspirin alone or combined with PEM/CDDP (Fig. 6a). Interestingly, aspirin treatment alone significantly mitigated weight loss without notable tumor regression (Figs. 6b–d and S6c). When combined aspirin with chemotherapy, we observed pronounced tumor regression, and the weight loss became similar to the vehicle group. Notably, we found that aspirin treatment led to increased weights in both skeletal muscle and adipose tissue (Fig. 6e, f), indicating effective suppression of cachexia. Aspirin also resulted in increased muscle fiber CSA and adipocyte size (Fig. S6d–g), histologically confirming the recovery from atrophic changes. Furthermore, aspirin reduced the expression of Atrogin-1 in TA tissues (Fig. 6g). These data suggest that aspirin, despite no tumor inhibition role, effectively mitigates cachexia progression of TKI-resistant lung cancer with concurrent PIK3CA mutant.
a Diagram of combined treatment with PEM/CDDP and aspirin. Created in BioRender. Yue, M. (2026) https://BioRender.com/ci3knzm. b Relative body weight changes during intraperitoneal administration of vehicle, aspirin, PEM/CDDP (Chemo), or combined treatment (Combo) (n = 7, 7, 8, 8 mice). Data were normalized to baseline and analyzed at 14 days. c, d Representative H&E staining images of lung tissue (c) and tumor burden (d) in TLCS-PP mice following vehicle, aspirin, PEM/CDDP (Chemo), or combined treatment (Combo) (n = 7, 7, 8, 8 mice). Scale bar, 2 mm. e, f Relative weights of the gastrocnemius (GAST), tibialis anterior (TA), quadriceps (QUAD) (e), and white adipose tissue (WAT) (f) in non-tumor bearing WT mice without treatment (WT Ctrl) and TLCS-PP mice following vehicle, aspirin, PEM/CDDP (Chemo), or combined treatment (Combo) (n = 6, 7, 7, 8, 8 mice). g Real-time PCR detection of mRNA levels in tibialis anterior from non-tumor bearing WT mice without treatment (WT Ctrl) and TLCS-PP mice following vehicle, aspirin, PEM/CDDP (Chemo), or combined treatment (Combo) (n = 5 mice/group). Data in (e, f) were normalized to WT mice of the same sex. Data in (b, d–g) are presented as mean ± SD. Statistical significance was calculated by one-way ANOVA followed by Tukey’s multiple comparisons test (b, d–g). Source data are provided as a Source Data file.
To further investigate the potential link between aspirin and the PIK3CA mutant, we established a cachexia model based on the transplantation of Lewis lung carcinoma (LLC) cell line with wild-type PIK3CA following a previously described protocol53. We found that LLC transplantation resulted in reduced tissue mass whereas aspirin administration had no significant impact upon tumor growth as well as tissue mass (Fig. S6h–j). Moreover, we observed no improvement of grip strength (Fig. S6k), no changes of Atrogin‑1 level as well as NF-κB signaling in muscle tissue (Fig. S6l, m).
Besides utilizing adjunct anti-inflammatory therapy alongside the current standard chemotherapy regimen, we also evaluated the therapeutic potential of newly approved PI3K inhibitor inavolisib for cachexia management54,55,56. We found that inavolisib effectively suppressed AKT phosphorylation and NF-κB activation in vitro (Fig. S7a). We then treated TLCS‑PP mice with inavolisib and found that inavolisib administration attenuated tumor growth and led to significant increases in both skeletal muscle and adipose tissue weights (Fig. S7b–f). We also observed a reduction of Atrogin-1 expression in TA tissues (Fig. S7g). These findings underscore the important role of PIK3CA mutant in cachexia development and suggest the PI3Kα inhibitor with improved safety might serve as a future therapeutic option for lung cancer patients with PIK3CA mutant.
Discussion
PIK3CA mutation has long been considered an important oncogenic driver in lung tumorigenesis, although it frequently co-occurs with EGFR mutation, KRAS mutation, and ALK fusion13,14. Previous studies have convincingly shown that EGFR mutation, KRAS mutation, or ALK fusion alone is sufficient to drive lung cancer initiation in mice19,20,21, and their co-expression even results in tumor growth inhibition16,18. This raises an interesting question about the real function of PIK3CA mutant in lung tumorigenesis. A previous study demonstrates that doxycycline-induced PIK3CA mutant transgene expression is sufficient to drive lung cancer initiation in mice12. In contrast, we find that Ad-Cre-mediated expression of PIK3CA mutant transgene plays a limited role in mouse lung tumor initiation. Consistently, Trejo CL et al. find no lung tumor formation in mice with endogenous expression of PIK3CA mutant57. No tumor is detectable in this model even after one year of PIK3CA mutant induction58. We reason that this discrepancy might be explained by various mouse models, the strength of PI3K pathway activation, and/or genetic backgrounds57. Nonetheless, clinical studies of multiple cancer types tend to support the incapability of PIK3CA mutant in driving tumor initiation. For example, human endometrial epithelium, which is pathologically normal, is found to harbor frequent PIK3CA mutations59. Similarly, PIK3CA mutations are detectable in pathologically normal human esophagus60. This study also reports no esophageal tumor detectable in PIK3CA-mutant transgenic mice60. These studies collectively demonstrate that PIK3CA mutant does not seem to contribute to cancer initiation as EGFR mutant, KRAS mutant, or ALK fusion does. Instead, it is more likely to play a secondary role in tumorigenesis, especially when concurrent with oncogenic drivers such as EGFR mutant, KRAS mutant, or ALK fusion. In line with this, we find that PIK3CA mutant mainly contributes to lung cancer malignant progression when coexisting with EGFR mutant, e.g., the concurrent PIK3CA mutant drives cachexia via activation of inflammatory signaling and leads to poor survival of mice bearing EGFR-mutant lung cancer. These findings provide a reasonable explanation for the longstanding paradox about the real biological function of concurrent PIK3CA mutant in lung tumorigenesis.
Another paradox exists in the seemingly contradictory impact of the PIK3CA mutant upon PFS and OS of EGFR-mutant lung cancer patients following TKI therapy28,29. Eng J et al. observed shorter survival in 10 patients with concurrent PIK3CA mutation compared with 43 patients harboring EGFR mutation alone; nevertheless, they find no difference of the best objective response, time to best response, time to progression, or TKI duration time between patients with and without PIK3CA mutation28. Similarly, Song Z et al. find that PIK3CA mutations lead to shorter OS, while these mutations do not seem to affect the PFS of EGFR-mutant patients29. Previous studies have observed the acquisition of PIK3CA mutations in relapsed patients after TKI resistance, indicative of its potential contribution to drug resistance, despite occurring at a low rate26,27. However, these mutations do not seem to impact the efficacy of TKIs. For example, Wu SG et al. find that the rate of PIK3CA mutation is comparable in relapsed patients vs. treatment-naïve patients, with no significant impact on therapeutic response and PFS between PIK3CA mutation-positive and -negative cases61. Another study also reports that 22 out of 27 EGFR-mutant patients show partial responses, including those with PIK3CA co-mutations62. Although these studies are based on limited sample sizes, the clinical observations point to a potentially dispensable role of PIK3CA mutant as therapeutic target in EGFR-sensitive patients. Indeed, our mouse model findings support that PIK3CA mutant contributes to lung tumorigenesis by driving cachexia rather than cancer initiation. Since PIK3CA mutant is not important for tumor initiation, tumors are less likely to rely on these mutations for survival. This explains why Osi treatment effectively inhibits the growth of EGFR-mutant lung cancer even with concurrent PIK3CA mutations. We propose that when patients relapse from TKI therapy and the tumors develop TKI resistance, the impact of PIK3CA mutant may become evident in the absence of effective tumor burden control, with cachexia driven by PIK3CA mutant gradually worsening and leading to poor patient prognosis. Our data from both TKI-sensitive and -resistant EGFR-mutant mouse models collectively support this view. Of course, this hypothesis warrants further detailed analyses of clinic data. We find that Osi effectively inhibits both tumor growth and cachexia progression in mice bearing tumors with EGFRL858R (TKI-sensitive) and PIK3CA co-mutations. Moreover, PIK3CA-associated cachexia is unresponsive to second-line chemotherapy, as demonstrated by the reduced tissue weights in mice bearing tumors with EGFRTLCS (TKI-resistance) and PIK3CA co-mutations. This is in line with clinical observations documenting the adverse effects of chemotherapy on cachexia progression50.
Multiple studies suggest a positive regulatory role of PI3K-AKT signaling in NF-κB activation across various pathological contexts63,64,65,66,67, although opposing findings have been reported under specific inflammatory conditions68. We here find that PIK3CA mutant activates NF-κB signaling, leading to increased expression of those cachexia-associated pro-inflammatory factors, such as IL-6 and LIF32, consistent with prior observations in breast cancer69. Systemic inflammation is recognized as the driving force of cachexia development in cancer70. In addition to pro-inflammatory factors secreted by tumor and immune cells, chemotherapy is recognized as a significant source of systemic inflammation in the patients’ macroenvironment71. Recent reports demonstrate that chemotherapy induces the upregulation of pro-inflammatory cytokines and chemokines, contributing to the progression of cachexia72, further underscoring the significance of inflammation in cachexia progression.
Lastly, we find that aspirin, a type of NSAID, effectively inhibits the progression of cachexia. Although inflammation is widely recognized as a hallmark of cachexia development, previous clinical studies indicate that targeting TNF-α, IL-6, or IL-1 individually demonstrates limited effectiveness73. We reason that attributing cachexia progression to a single factor is challenging, as various pro-inflammatory factors interact through complex feedback loops70. Therefore, broader-spectrum anti-inflammatory drugs might offer a more effective intervention. Aspirin has been reported to inhibit IκB kinase-β (IKKβ), thereby blocking NF-κB activation and downstream inflammatory signaling74. We here find that aspirin treatment suppresses cachexia-associated cytokines driven by PIK3CA mutant and results in cachexia attenuation. Notably, besides the inhibition of NF-κB activation in tumor cells, aspirin might have the potential to exert systemic effects in repressing cachexia, which warrants further in-depth investigation. Interestingly, previous studies show that colorectal cancer patients with PIK3CA mutations receiving regular aspirin treatment exhibit improved prognosis75,76. A multicenter, multinational, prospective randomized trial recently provides the first evidence of a positive effect of adjuvant aspirin in colon cancer patients with PIK3CA mutations77, further emphasizing the significance of adjuvant anti-inflammatory treatment in enhancing the prognosis of these patients. A previous study also reveals an association between PIK3CA mutant and weight loss in pancreatic ductal adenocarcinoma patients78. Interestingly, we also found that PI3K-AKT signaling was enriched in melanoma xenografts capable of inducing cachexia, further supporting the role of PIK3CA mutant in driving cachexia across cancer types. Given the significant contribution of PIK3CA mutant to poor prognosis by driving cachexia, it will be important to further investigate its broader implications in lung cancer and beyond.
Methods
Mouse models
The Trp53flox/flox mice were originally provided by Dr. Tyler Jacks (Cambridge, MA). The transgenic mouse models, including PIK3CAE545K, EGFRL858R, and EGFRTLCS mice were generated using CRISPR/Cas9 technology by commercial companies. All mice used were on a C57BL/6J background. The brief process is as follows: homologous recombination vector (donor vector) is composed of a 5′ homology arm, the indicated coding sequence, and a 3′ homology arm. Cas9 mRNA, gRNA, and the donor vector were microinjected into the fertilized eggs of C57BL/6J mice to generate the F0 generation mice.
Mice were maintained in specific pathogen-free facilities and housed in single-sex cages at 20 ± 2 °C, 40–60% humidity under a 12-h light/12-h dark photoperiod with the lights on at 7:00. Mice were fed a standard chow diet ad libitum. At 6–8 weeks of age, mice were treated with Ad-Cre virus (2 × 10⁶ p.f.u.) via nasal inhalation. Approximately equal numbers of male and female mice were used. The time points post-Ad-Cre treatment are indicated for each experiment.
Osimertinib (MCE, HY-15772) was prepared in a solution containing 5% DMSO, 40% PEG300, 5% Tween-80, and 50% saline. PEM (MCE, HY-10820) or aspirin (MCE, HY-14654) was prepared in a solution containing 10% DMSO, 40% PEG300, and 50% saline. Cisplatin (MCE, HY-17394) was formulated in saline. Inavolisib (Selleck, S8668) was prepared in a solution containing 0.5% (w/v) methylcellulose and 0.2% (w/v) Tween-80. For osimertinib treatment, EGFRL858R;PIK3CAE545K;Trp53flox/flox mice were gavage daily with osimertinib (5 mg/kg/day) for 28 days, starting 6 weeks after Ad-Cre infection. Control mice were administered the vehicle solution (5% DMSO: 40% PEG300: 5% Tween-80: 50% saline). For chemotherapy and combination therapy, EGFRTLCS;PIK3CAE545K;Trp53flox/flox mice were administered PEM (50 mg/kg/day) combined with cisplatin (4 mg/kg/day) or aspirin (50 mg/kg/day) or both via intraperitoneal injection for 14 days, starting 10 weeks after Ad-Cre infection. PEM and cisplatin were administered weekly, and aspirin was given daily. Control mice were administered the vehicle solution (10% DMSO: 40% PEG300: 50% saline). For inavolisib therapy, EGFRTLCS;PIK3CAE545K;Trp53flox/flox mice were gavage daily with inavolisib (25 mg/kg/day) for 28 days, starting 8 weeks after Ad-Cre infection. Control mice were administered the vehicle solution consisting of 0.5% (w/v) methylcellulose and 0.2% (w/v) Tween-80. All mice were sacrificed for gross inspection and histopathological examination.
Tumor number, tumor burden, and tumor size were analyzed using ImageJ software. To calculate tumor burden, paraffin-embedded mouse lung tissue sections were processed with H&E staining and scanned using ZEISS Axioscan 7 to obtain lung lobe images. Tumor and lung areas for each lobe were delineated and quantified using ImageJ software. The tumor burden was calculated as the ratio of tumor areas to lung area.
Cachexia model based on LLC transplantation
Male C57BL/6 mice (8–10 weeks old) were received 5 × 10⁶ LLC cells in 100 μL PBS. Aspirin (50 mg/kg/day) was administered via intraperitoneal injection daily starting 7 days post-tumor transplantation. Control mice received the vehicle solution (10% DMSO, 40% PEG300, 50% saline).
Assessment of disease progression and humane endpoints
For the subcutaneous tumor model, disease progression in mice was monitored regularly by measuring tumor size with calipers and recording body weight. For in situ tumor model, disease progression was monitored by recording body weight and observing clinical signs, including activity level, posture, grooming, and general behavior. Experimental endpoints were generally defined based on pre-determined study duration or attainment of humane endpoints. Humane endpoints were defined as a 20% reduction in body weight from tumor induction, cessation of eating and drinking, severe lethargy or moribund state, a maximal tumor diameter of 2 cm, tumor ulceration exceeding 1 cm, severe trauma leading to secondary infection, or significant organ dysfunction. Body weight changes in Figs. 1b and 4b were shown relative to tumor induction, and body weight changes in Fig. 6b were relative to treatment initiation.
Plasmid
The pCDH-PIK3CA E545K plasmid was generated by standard PCR and homologous recombination-based cloning. The psPAX2 plasmid (#12260) and pMD2.G plasmid (#12259) were purchased from Addgene. The NF-κB-Luc plasmid and Renilla plasmid were laboratory-preserved plasmids.
Cell culture and lentivirus infection
293 T and EP cells were cultured in DMEM (HyClone) supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin (P/S). Cells were used for experiments within 10 to 20 passages from thawing. All cell lines were routinely tested for mycoplasma. For the establishment of stable overexpression cell line, 293 T cells were transfected with a 4:3:2 ratio of pCDH-PIK3CA E545K plasmid, psPAX2 plasmid, and pMD2.G plasmid. Lentiviral particles generated were then transduced into EP cells, followed by puromycin selection (3 μg/mL; Sigma-Aldrich) initiated 48 h post-transfection and continued for an additional 2 days. 293 T cell line was purchased from ATCC (CRL-3216), EP cell line was manually established from lung tumors of female EGFRL858R;PIK3CAE545K;Trp53flox/flox mice following Ad-Cre treatment.
Immunoblotting
Whole-cell lysates of cell lines or murine tumors were generated using lysis buffer (10% SDS, 1 mM DTT, and glycerin) and incubated at 100 °C for 10 min. Equal amounts of total protein were separated by SDS-PAGE and transferred onto PVDF membranes. Protein samples were probed with specific antibodies against p110α (CST, 4249, 1:1000), AKT (CST, 9272, 1:1000), p-AKT (CST, 4070, 1:1000), S6 (CST, 2217, 1:1000), p-S6 (CST, 2215, 1:1000), p65 (CST, 8242, 1:1000), p-p65 (CST, 3033, 1:1000), and β-actin (Abclonal, AC026, 1:200000). Protein expression was assessed by Pierce ECL Western Blotting Substrate (Thermo Fisher Scientific) and detected on SAGECREATION (Sage Creation Science Co.).
For quantification, band intensities were quantified using ImageJ. Background was first subtracted, and rectangular regions were then drawn around each band to measure integrated densities. The signal of each target protein was normalized to its corresponding loading control (actin). Ratios were further normalized to the value of the first lane. For Figs. 5f and S5p–S7a actin was measured on the same blot as the target proteins. For Figs. S2d and S2h, actin was measured on the same samples but on different blots. All samples within each subpanel originated from the same experiment, and the corresponding gels/blots were processed in parallel.
RNA Isolation, reverse transcription, and real-time PCR
Total RNA from cultured cells or tissue samples was extracted using TRIzol (Invitrogen), and complementary DNA was synthesized from 1 μg of RNA using the PrimeScript RT Reagent Kit (TaKaRa). Real-time PCR was performed on a LightCycler® 96 System (Roche) using SYBR Green I Master (Roche). β-actin was utilized as the internal control. The following primers were used: Il1b F: 5′-GCAACTGTTCCTGAACTCAAC-3′, R: 5′-ATCTTTTGGGGTCCGTCAACT-3′. Il6 F: 5′-TCTGCAAGAGACTTCCATCCAGTTGC-3′, R: 5′-AGCCTCCGACTTGTGAAGTGGT-3′. Il11 F: 5′-CTGACGGAGATCACAGTCTGGA-3′, R: 5′-GGACATCAAGTCTACTCGAAGCC-3′. Lif F: 5′-AAAAGCTATGTGCGCCTAACA-3′, R: 5′-GTATGCGACCATCCGATACAG-3′. Osm F: 5′-ACGGTCCACTACAACACCAG-3′, R: 5′-CCATCGTCCCATTCCCTGAAG-3′. Ptgs2 F: 5′-TTCAACACACTCTATCACTGGC-3′, R: 5′-AGAAGCGTTTGCGGTACTCAT-3′. Tnf F: 5′-CTGAACTTCGGGGTGATCGG-3′, R: 5′-GGCTTGTCACTCGAATTTTGAGA-3′. Atrogin-1 F: 5′-CTTCTCGACTGCCATCCTGGAT-3′, R: 5′-TCTTTTGGGCGATGCCACTCAG-3′. Nfkb1 F: 5′-CTGAACAATGCCTTCCGGCT-3′, R: 5′-TGGTACCCCCAGAGACCTCAT-3′. Rela F: 5′-GATCGCCACCGGATTGAAGA-3′, R: 5′-GGGGTTCAGTTGGTCCATTG-3′. Actb F: 5′-GGCTGTATTCCCCTCCATCG-3′, R: 5′-CCAGTTGGTAACAATGCCATGT-3′. Oligonucleotides were obtained by chemical synthesis.
Soft agar colony formation assay
For soft agar assay, a bottom layer of 1% agar with complete medium was first solidified. An upper layer was then added, containing 500 cells suspended in a 0.4% agar medium mixture in 6-well plates. After 2–3 weeks of incubation, cells were stained with 0.005% crystal violet, and the number of colonies was counted.
Immunofluorescence analyses
Cells grown on glass coverslips were washed with cold PBS and fixed with 4% paraformaldehyde in PBS for 15 min at room temperature. Fixed cells were permeabilized with 0.2% Triton ×-100 in PBS for 15 min. After blocking with 4% bovine serum albumin in PBS for 1 h, cells were probed with p65 (CST, 8242, 1:500) overnight at 4 °C. After washing three times with PBST, secondary antibodies were added and incubated for 1 h at room temperature. After washed with PBST, the coverslips were mounted onto glass slides using fluorescent mounting medium. Confocal images were captured using a Leica TCS SP8 system with a HC PL APO CS2 63×/1.40 oil objective.
Luciferase reporter gene assay
293 T cells were seeded in 12-well plate at 3 × 105 cells/well. 24 h after seeding, cells were transiently transfected with 1 μg of either PIK3CA E545K plasmid or the control vector, along with 1 μg NF-κB-Luc plasmid and 500 ng Renilla plasmid. 48 h post-transfection, cells were harvested, and the Dual-Luciferase Reporter Assay System (Promega) was used for detection.
Tissue collection
Mice were euthanized, and their body weight was recorded. The lungs with tumors were excised and fixed in 4% formalin; a portion of the tumors was snap-frozen for further analysis. GAST, TA, QUAD, eWAT, and gWAT were dissected and weighed. The white adipose tissues were fixed in 4% formalin for histological examination. One part of the skeletal muscle was snap-frozen for RNA-seq, while the remaining portion was embedded in optimal cutting temperature compound for rapid freezing and subsequent histological examination.
Body composition analysis and fat imaging
Mice were weighed and body composition was measured using Bruker’s minispec LF50 Body Composition Analyzer. The instrument provides measurements of three related components: fat, free body fluids, and lean tissue mass. The imaging of mouse adipose tissue was performed using the NM42-060H-I (Niumag) small animal magnetic resonance imaging scanner.
Metabolic cage
Mice were individually housed in CLAMS metabolic cages (Columbus Instruments) for a duration of 3 days. Oxygen consumption (VO2) and carbon dioxide expiration (VCO2) were measured for 1 min with 14 min intervals at a flow rate set at 0.72 liter per minute. Respiratory exchange ratio (RER) was calculated as the ratio of VCO2 to VO2. Simultaneously, locomotor activity and energy expenditure were recorded using the built-in detection system, while food intake was manually measured at the same designated time each day.
Grip strength measurement
Each mouse was allowed to grasp a bar attached to a force transducer and was steadily pulled horizontally by the tail away from the bar using a KW-ZL-2 grip strength meter. The maximum force exerted before releasing the bar was recorded. Each mouse was tested five times, and the average of the remaining three measurements after exclusion of the highest and lowest values was calculated as the grip strength.
RNA-seq analyses
Raw fastq data from RNA-seq were processed with Trimmomatic79 (v0.39) for adapter trimming and low-quality read filtering. The processed data were then aligned to the mm10 reference genome using STAR80 (v2.5.2b). Genes with zero expression in more than 70% of the samples were filtered out. FPKM normalization and log2 transformation were applied to the raw count data, followed by differential expression analysis using limma81 to identify differentially expressed genes (DEGs) between conditions. Pathway enrichment analysis was conducted using the Enrichr82 method based on the resulting DEGs. In addition, the GSEA analysis was performed using the R package clusterProfiler83 with 1000 permutations and gene_set permutation type (default setting). Genes were ranked based on their fold-change values.
Public datasets analyses
Public datasets were obtained from multiple sources. Gene expression profiles were downloaded from the Gene Expression Omnibus database, including GSE13056348 and GSE13352349, which contain expression data from muscle tissues of cachectic patients and their respective controls. Differential expression analysis was performed by R package limma81. Additional transcriptomic data and gene lists—including the cachexia signature proposed by the TRACEx study37, tumor RNA-seq37,44 and serum proteomics45 data from lung cancer patients, as well as tumor RNA-seq data from melanoma xenografts46—were extracted from supplementary tables of published studies. Pathway enrichment analysis was performed using the Enrichr82 web tool based on downloaded gene lists. GSEA was conducted using the R package clusterProfiler83 with 1000 permutations and gene_set permutation type (default setting). Genes were ranked based on their fold-change values. Normalized enrichment scores (NES) and false discovery rate (FDR) q-values were calculated.
Statistics and reproducibility
GraphPad Prism 9 was used for statistical analyses. Data were presented as mean ± standard deviation (SD) unless otherwise stated. Statistical significance for two experimental samples was determined using two-tailed unpaired Student’s t test. For comparisons among multiple groups, one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test was applied. Survival analysis was performed using the log-rank (Mantel–Cox) test. P value < 0.05 was considered statistically significant. The number of biological replicates (n) is specified in each figure legend and p-values are indicated in the individual figures.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Ethical regulations
The research performed in the present study complies with all ethical regulations. All mice were kept in the specific pathogen-free environment at Shanghai Institute of Biochemistry and Cell Biology and treated in strict accordance with protocols (SIBCB-2101008) approved by the Institutional Animal Care and Use Committee of the Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences.
Data availability
The RNA-seq data generated in this study have been deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2021) at National Genomics Data Center (Nucleic Acids Res 2022), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession code CRA021522. Source data are provided with this paper.
References
Thai, A. A., Solomon, B. J., Sequist, L. V., Gainor, J. F. & Heist, R. S. Lung cancer. Lancet 398, 535–554 (2021).
Cancer Genome Atlas Research, N Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–50 (2014).
Cancer Genome Atlas Research, N Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–25 (2012).
Ghezzi, C. et al. A high-throughput screen identifies that CDK7 activates glucose consumption in lung cancer cells. Nat. Commun. 10, 5444 (2019).
Liu, L. et al. Therapeutic efficacy of TMTP1-modified EVs in overcoming bone metastasis and immune resistance in PIK3CA mutant NSCLC. Cell Death Dis. 16, 367 (2025).
Zhang, J. et al. Tumoral NOX4 recruits M2 tumor-associated macrophages via ROS/PI3K signaling-dependent various cytokine production to promote NSCLC growth. Redox Biol. 22, 101116 (2019).
Collins, N. B. et al. PI3K activation allows immune evasion by promoting an inhibitory myeloid tumor microenvironment. J. Immunother Cancer 10, e003402 (2022).
Arafeh, R. & Samuels, Y. PIK3CA in cancer: the past 30 years. Semin Cancer Biol. 59, 36–49 (2019).
Janku, F., Yap, T. A. & Meric-Bernstam, F. Targeting the PI3K pathway in cancer: are we making headway?. Nat. Rev. Clin. Oncol. 15, 273–291 (2018).
Wang, Y., Rozen, V., Zhao, Y. & Wang, Z. Oncogenic activation of PI K3 CA in cancers: emerging targeted therapies in precision oncology. Genes Dis. 12, 101430 (2025).
Hoxhaj, G. & Manning, B. D. The PI3K-AKT network at the interface of oncogenic signalling and cancer metabolism. Nat. Rev. Cancer 20, 74–88 (2020).
Engelman, J. A. et al. Effective use of PI3K and MEK inhibitors to treat mutant Kras G12D and PIK3CA H1047R murine lung cancers. Nat. Med. 14, 1351–6 (2008).
Wang, L. et al. PIK3CA mutations frequently coexist with EGFR/KRAS mutations in non-small cell lung cancer and suggest poor prognosis in EGFR/KRAS wildtype subgroup. PLoS ONE 9, e88291 (2014).
Chaft, J. E. et al. Coexistence of PIK3CA and other oncogene mutations in lung adenocarcinoma-rationale for comprehensive mutation profiling. Mol. Cancer Ther. 11, 485–91 (2012).
Skoulidis, F. & Heymach, J. V. Co-occurring genomic alterations in non-small-cell lung cancer biology and therapy. Nat. Rev. Cancer 19, 495–509 (2019).
Unni, A. M., Lockwood, W. W., Zejnullahu, K., Lee-Lin, S. Q. & Varmus, H. Evidence that synthetic lethality underlies the mutual exclusivity of oncogenic KRAS and EGFR mutations in lung adenocarcinoma. Elife 4, e06907 (2015).
Tang, S. et al. Counteracting lineage-specific transcription factor network finely tunes lung adeno-to-squamous transdifferentiation through remodeling tumor immune microenvironment. Natl. Sci. Rev. 10, nwad028 (2023).
Ambrogio, C., Barbacid, M. & Santamaria, D. In vivo oncogenic conflict triggered by co-existing KRAS and EGFR activating mutations in lung adenocarcinoma. Oncogene 36, 2309–2318 (2017).
Foggetti, G. et al. Genetic determinants of EGFR-driven lung cancer growth and therapeutic response in vivo. Cancer Discov. 11, 1736–1753 (2021).
Ji, H. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 448, 807–10 (2007).
Qin, Z. et al. EML4-ALK fusions drive lung adeno-to-squamous transition through JAK-STAT activation. J. Exp. Med. 221, e20232028 (2024).
Fu, K., Xie, F., Wang, F. & Fu, L. Therapeutic strategies for EGFR-mutated non-small cell lung cancer patients with osimertinib resistance. J. Hematol. Oncol. 15, 173 (2022).
Thress, K. S. et al. Acquired EGFR C797S mutation mediates resistance to AZD9291 in non-small cell lung cancer harboring EGFR T790M. Nat. Med. 21, 560–2 (2015).
Herbst, R. S., Morgensztern, D. & Boshoff, C. The biology and management of non-small cell lung cancer. Nature 553, 446–454 (2018).
Tan, A. C. & Tan, D. S. W. Targeted therapies for lung cancer patients with oncogenic driver molecular alterations. J. Clin. Oncol. 40, 611–625 (2022).
Le, X. et al. Landscape of EGFR-dependent and -independent resistance mechanisms to osimertinib and continuation therapy beyond progression in EGFR-mutant NSCLC. Clin. Cancer Res. 24, 6195–6203 (2018).
Chmielecki, J. et al. Analysis of acquired resistance mechanisms to osimertinib in patients with EGFR-mutated advanced non-small cell lung cancer from the AURA3 trial. Nat. Commun. 14, 1071 (2023).
Eng, J. et al. Impact of concurrent PIK3CA mutations on response to EGFR tyrosine kinase inhibition in EGFR-mutant lung cancers and on prognosis in oncogene-driven lung adenocarcinomas. J. Thorac. Oncol. 10, 1713–9 (2015).
Song, Z., Yu, X. & Zhang, Y. Mutation and prognostic analyses of PIK3CA in patients with completely resected lung adenocarcinoma. Cancer Med. 5, 2694–2700 (2016).
Ferrer, M. et al. Cachexia: a systemic consequence of progressive, unresolved disease. Cell 186, 1824–1845 (2023).
Argiles, J. M., Lopez-Soriano, F. J., Stemmler, B. & Busquets, S. Cancer-associated cachexia - understanding the tumour macroenvironment and microenvironment to improve management. Nat. Rev. Clin. Oncol. 20, 250–264 (2023).
Baracos, V. E., Martin, L., Korc, M., Guttridge, D. C. & Fearon, K. C. H. Cancer-associated cachexia. Nat. Rev. Dis. Prim. 4, 17105 (2018).
Bye, A. et al. Muscle mass and association to quality of life in non-small cell lung cancer patients. J. Cachexia Sarcopenia Muscle 8, 759–767 (2017).
Ross, P. J. et al. Do patients with weight loss have a worse outcome when undergoing chemotherapy for lung cancers?. Br. J. Cancer 90, 1905–11 (2004).
de Jong, C. et al. The association between skeletal muscle measures and chemotherapy-induced toxicity in non-small cell lung cancer patients. J. Cachexia Sarcopenia Muscle 13, 1554–1564 (2022).
Jin, J., Visina, J., Burns, T. F., Diergaarde, B. & Stabile, L. P. Male sex and pretreatment weight loss are associated with poor outcome in patients with advanced non-small cell lung cancer treated with immunotherapy: a retrospective study. Sci. Rep. 13, 17047 (2023).
Al-Sawaf, O. et al. Body composition and lung cancer-associated cachexia in TRACERx. Nat. Med 29, 846–858 (2023).
Oswalt, C. et al. Associations between body mass index, weight loss and overall survival in patients with advanced lung cancer. J. Cachexia Sarcopenia Muscle 13, 2650–2660 (2022).
Yang, M., Shen, Y., Tan, L. & Li, W. Prognostic value of sarcopenia in lung cancer: a systematic review and meta-analysis. Chest 156, 101–111 (2019).
Mytelka, D. S., Li, L. & Benoit, K. Post-diagnosis weight loss as a prognostic factor in non-small cell lung cancer. J. Cachexia Sarcopenia Muscle 9, 86–92 (2018).
Frese, K. K. & Tuveson, D. A. Maximizing mouse cancer models. Nat. Rev. Cancer 7, 645–58 (2007).
DuPage, M., Dooley, A. L. & Jacks, T. Conditional mouse lung cancer models using adenoviral or lentiviral delivery of Cre recombinase. Nat. Protoc. 4, 1064–72 (2009).
Fearon, K. et al. Definition and classification of cancer cachexia: an international consensus. Lancet Oncol. 12, 489–95 (2011).
Cury, S. S. et al. Low muscle mass in lung cancer is associated with an inflammatory and immunosuppressive tumor microenvironment. J. Transl. Med. 21, 116 (2023).
Wang, D. et al. LCN2 secreted by tissue-infiltrating neutrophils induces the ferroptosis and wasting of adipose and muscle tissues in lung cancer cachexia. J. Hematol. Oncol. 16, 30 (2023).
Graca, F. A. et al. Progressive development of melanoma-induced cachexia differentially impacts organ systems in mice. Cell Rep. 42, 111934 (2023).
Fearon, K. C., Glass, D. J. & Guttridge, D. C. Cancer cachexia: mediators, signaling, and metabolic pathways. Cell Metab. 16, 153–66 (2012).
Judge, S. M. et al. Skeletal muscle fibrosis in pancreatic cancer patients with respect to survival. JNCI Cancer Spectr. 2, pky043 (2018).
Talbert, E. E. et al. Modeling human cancer-induced cachexia. Cell Rep. 28, 1612–1622.e4 (2019).
Kimura, M. et al. Prognostic impact of cancer cachexia in patients with advanced non-small cell lung cancer. Support Care Cancer 23, 1699–708 (2015).
Pahl, H. L. Activators and target genes of Rel/NF-κB transcription factors. Oncogene 18, 6853–6866 (1999).
Kopp, E. & Ghosh, S. Inhibition of NF-kappa B by sodium salicylate and aspirin. Science 265, 956–9 (1994).
Bilgic, S. N. et al. EDA2R-NIK signalling promotes muscle atrophy linked to cancer cachexia. Nature 617, 827–834 (2023).
Turner, N. C. et al. Inavolisib-based therapy in PIK3CA-mutated advanced breast cancer. N. Engl. J. Med. 391, 1584–1596 (2024).
Gough, S. M. et al. Oral estrogen receptor PROTAC vepdegestrant (ARV-471) is highly efficacious as monotherapy and in combination with CDK4/6 or PI3K/mTOR pathway inhibitors in preclinical ER+ breast cancer models. Clin. Cancer Res. 30, 3549–3563 (2024).
Song, K. W. et al. RTK-dependent inducible degradation of mutant PI3Kalpha drives GDC-0077 (Inavolisib) efficacy. Cancer Discov. 12, 204–219 (2022).
Trejo, C. L. et al. Mutationally activated PIK3CA(H1047R) cooperates with BRAF(V600E) to promote lung cancer progression. Cancer Res. 73, 6448–61 (2013).
Green, S., Trejo, C. L. & McMahon, M. PIK3CA(H1047R) accelerates and enhances KRAS(G12D)-driven lung tumorigenesis. Cancer Res. 75, 5378–91 (2015).
Moore, L. et al. The mutational landscape of normal human endometrial epithelium. Nature 580, 640–646 (2020).
Herms, A. et al. Organismal metabolism regulates the expansion of oncogenic PIK3CA mutant clones in normal esophagus. Nat. Genet. 56, 2144–2157 (2024).
Wu, S. G., Chang, Y. L., Yu, C. J., Yang, P. C. & Shih, J. Y. The role of PIK3CA mutations among lung adenocarcinoma patients with primary and acquired resistance to EGFR tyrosine kinase inhibition. Sci. Rep. 6, 35249 (2016).
Endoh, H., Yatabe, Y., Kosaka, T., Kuwano, H. & Mitsudomi, T. PTEN and PIK3CA expression is associated with prolonged survival after gefitinib treatment in EGFR-mutated lung cancer patients. J. Thorac. Oncol. 1, 629–34 (2006).
Chalabi-Dchar, M. et al. Loss of somatostatin receptor subtype 2 promotes growth of KRAS-induced pancreatic tumors in mice by activating PI3K signaling and overexpression of CXCL16. Gastroenterology 148, 1452–65 (2015).
Paul, J. et al. Simultaneous inhibition of PI3Kdelta and PI3Kalpha induces ABC-DLBCL regression by blocking BCR-dependent and -independent activation of NF-kappaB and AKT. Cancer Cell 31, 64–78 (2017).
Dan, H. C. et al. Akt-dependent regulation of NF-kappaB is controlled by mTOR and Raptor in association with IKK. Genes Dev. 22, 1490–500 (2008).
Li, R. et al. Interleukin-7 induces recruitment of monocytes/macrophages to endothelium. Eur. Heart J. 33, 3114–23 (2012).
Ying, H. et al. PTEN is a major tumor suppressor in pancreatic ductal adenocarcinoma and regulates an NF-kappaB-cytokine network. Cancer Discov. 1, 158–69 (2011).
Zhao, L., Lee, J. Y. & Hwang, D. H. The phosphatidylinositol 3-kinase/Akt pathway negatively regulates Nod2-mediated NF-kappaB pathway. Biochem. Pharm. 75, 1515–25 (2008).
Hutti, J. E. et al. Oncogenic PI3K mutations lead to NF-kappaB-dependent cytokine expression following growth factor deprivation. Cancer Res. 72, 3260–9 (2012).
Baazim, H., Antonio-Herrera, L. & Bergthaler, A. The interplay of immunology and cachexia in infection and cancer. Nat. Rev. Immunol. 22, 309–321 (2022).
Swanton, C. et al. Embracing cancer complexity: hallmarks of systemic disease. Cell 187, 1589–1616 (2024).
Englund, D. A. et al. Senotherapeutic drug treatment ameliorates chemotherapy-induced cachexia. JCI Insight 9, e169512 (2024).
Yue, M., Qin, Z., Hu, L. & Ji, H. Understanding cachexia and its impact on lung cancer and beyond. Chin. Med. J. Pulm. Crit. Care Med. 2, 95–105 (2024).
Yin, M. J., Yamamoto, Y. & Gaynor, R. B. The anti-inflammatory agents aspirin and salicylate inhibit the activity of I(kappa)B kinase-beta. Nature 396, 77–80 (1998).
Domingo, E. et al. Evaluation of PIK3CA mutation as a predictor of benefit from nonsteroidal anti-inflammatory drug therapy in colorectal cancer. J. Clin. Oncol. 31, 4297–305 (2013).
Liao, X. et al. Aspirin use, tumor PIK3CA mutation, and colorectal-cancer survival. N. Engl. J. Med. 367, 1596–606 (2012).
Güller, U. et al. 512O Adjuvant aspirin treatment in PIK3CA mutated colon cancer patients: the phase III, prospective-randomized placebo-controlled multicenter SAKK 41/13 trial. Ann. Oncol. 35, S432 (2024).
Narasimhan, A. et al. Identification of potential serum protein biomarkers and pathways for pancreatic cancer cachexia using an aptamer-based discovery platform. Cancers (Basel) 12, 3787 (2020).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–20 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinforma. 14, 128 (2013).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Acknowledgements
We thank Dr. Tyler Jacks and Dr. Liang Chen for providing the mice. This work was supported by the National Key Research and Development Program of China (grants 2022YFA1103900, 2020YFA0803300 to H.J.; 2025YFC3409300, 2025YFF1207900, 2022YFA1004800 to L.C.); the National Natural Science Foundation of China (grants 82303039 to Z.Q.; 82303575 to S.T.; 82341002, 32293192, 82030083 to H.J.; T2341007, T2350003, 12131020, 42450084, 42450135, 12326614, 12426310, 92374001 to L.C.); the Shanghai Sailing Program(23YF1452900 to Z.Q.); the Basic Frontier Scientific Research Program of Chinese Academy of Science (ZDBS-LY-SM006 to H.J.); the Innovative research team of high-level local universities in Shanghai (SSMU-ZLCX20180500 to H.J.); Science and Technology Commission of Shanghai Municipality (23JS1401300 to L.C.); Zhejiang Province Vanguard Goose-Leading Initiative (2025C01114 to L.C.); and JST Moonshot R&D (JPMJMS2021 to L.C.).
Author information
Authors and Affiliations
Contributions
H.J. conceived the idea and designed the experiments. M.Y. and Z.Q. performed all experiments and analyzed the data. S.T. and X.C. performed the bioinformatics analyses. Y.Z. established the primary cell line. Lu. C., C.Y., and Li. C. provided technical assistance and helpful comments. H.J. and M.Y. wrote the manuscript. All authors approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Serkan Kir, who co-reviewed with Samet Agca, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yue, M., Qin, Z., Tang, S. et al. Concurrent PIK3CA mutant promotes cachexia through inflammatory signaling in EGFR mutant lung cancer. Nat Commun 17, 704 (2026). https://doi.org/10.1038/s41467-025-67345-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67345-9