Abstract
The role of soil microorganisms in supporting multiple ecosystem functions (multifunctionality) remains poorly understood across diverse environmental conditions. Here, we investigate 484 soils from 27 European countries spanning a range of climatic and edaphic contexts. We assess the contribution of climate, soil properties, and soil microbiome traits (i.e., the relative abundance of co-occurring taxa) to explain six key functional proxies related to soil structure, biochemical activity, and productivity. We find the highest multifunctionality values in grasslands, woodlands, loamy and acidic soils, and temperate humid regions, and the lowest in croplands, alkaline soils, and drier regions. Soil properties explain 12–31% of variation in multifunctionality, with microbial biomass and nitrogen content emerging as the strongest predictors. The soil microbiome accounts for 2–14% of unique variance in multifunctionality but explains more than 25% of variation in enzymatic activities and primary productivity in clay-rich soils and soils originating from temperate dry regions. Specific taxa, particularly within Actinobacteria, Acidobacteria, and the fungal genus Mortierella consistently emerge as strong predictors of ecosystem multifunctionality. Our findings highlight that ecosystem multifunctionality is jointly shaped by soil properties and microbial communities. We argue that specific taxa hold potential as context-dependent indicators for multifunctionality monitoring across environmental gradients.
Similar content being viewed by others
Introduction
Soils are the foundation of terrestrial ecosystems and represent an important habitat for organisms across the tree of life1. Soils also contribute to a wide range of ecosystem services including food production, climate regulation, as well as cultural and educational services2. These ecosystem services are directly supported by ecosystem functions, such as nutrient cycling, water holding capacity, and primary production3,4,5. In recent years, research has increasingly focused on ecosystem multifunctionality, recognizing that ecosystem services depend on multiple functions simultaneously, often assessed through measurable functional proxies (e.g., enzymatic activities, soil respiration)6,7,8. Previous research has shown that greater soil biodiversity is associated with higher levels of multiple functions, such as nutrient cycling and primary production9,10,11.
Microorganisms are sensitive, ubiquitous, and react quickly to environmental stress, reflecting cumulative impacts12, and potentially constituting an important indicator of changes in ecosystem multifunctionality13. This is especially important as sequencing is becoming a cheaper and highly standardized technology allowing direct comparison of data across different studies and national surveys14. However, assessments on the capacity of the soil microbiome in explaining ecosystem multifunctionality at larger scale are urgently needed to support the monitoring and conservation of soils.
Unfortunately, there are still important uncertainties about the capacity of the soil microbiome to predict multifunctionality across large spatial scales. First, despite the enormous diversity of soil biota (particularly microbial diversity), the focus is still often placed on a narrow subset of organisms. For instance, mycorrhizal fungi and nitrogen-fixing bacteria are among the few microbial functional groups currently assessed in Europe as potential indicators of ecosystem multifunctionality15,16. Second, despite growing interest in microbial indicators of ecosystem multifunctionality, current studies do not specifically address how multiple environmental contexts, such as land use, soil texture, and climate, influence the relationship between the taxonomic identity of soil organisms and their capacity to predict multifunctionality17. Finally, although overall microbial diversity (i.e., taxa richness) has been shown to positively influence multifunctionality in global and national contexts, similar large-scale analyses linking microbial community structure to multifunctionality are lacking for Europe11,18,19. This gap is especially relevant considering that less than 40% of European soils are currently categorized as healthy, despite the continent’s long history of land management and its significant role in global food production and biodiversity20. These knowledge gaps hinder our ability to evaluate how the soil microbiome relate to specific ecosystem functions across diverse contexts (i.e., soil types, climatic regions, or land uses).
Here, we use a harmonized pan-European observational field survey (named LUCAS Soil) to quantify the contribution of the soil microbiome in explaining ecosystem multifunctionality. We analyze 484 fresh soil samples collected across diverse environmental conditions, using categorical groupings to test the effects of land use, climatic region, soil texture, and soil pH, as these factors have previously been shown to structure soil microbial communities and ecosystem multifunctionality21. Using metabarcoding, we obtain site-level information on the relative abundance of bacteria and fungi. We assess ecosystem multifunctionality by integrating several functional proxies that capture key functions linked to broader ecosystem services. Specifically, we measure soil aggregate stability (flood regulation, habitat for organisms), net primary productivity (food and fiber/timber provision), and basal respiration (climate regulation). In addition, we quantify enzymatic activities including N-acetyl-glucosaminidase (nitrogen cycling), phosphatase (phosphorus cycling), and xylosidase (carbon cycling), all of which are involved in organic matter decomposition. Together, these indicators provide a multidimensional picture of ecosystem functioning, which we summarize as ecosystem multifunctionality.
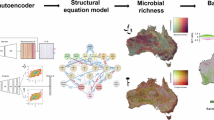
To quantify the contribution of the soil microbiome to explain multifunctionality, we apply Random Forest models to identify the most important predictors of multifunctionality, considering a wide set of abiotic (climate and soil properties) and biotic (microbial community composition) variables. To quantify the relative contributions of these predictor groups, we perform variance partitioning analysis, allowing us to attribute the unique and shared variance in multifunctionality to climate, soil properties, and the soil microbiome. We finally employ structural equation models (SEMs) to test for the direct and indirect effects of climate, soil properties, and the soil microbiome on ecosystem multifunctionality. We hypothesize that soils from less-disturbed environments (e.g., woodlands) would support higher multifunctionality due to more favorable conditions for microbial activity. We also hypothesize that the contribution of microbial taxa to multifunctionality would be context-dependent, varying across land use, climatic region, pH, and soil texture.
Results and discussion
We analyzed 484 soils across Europe to investigate how ecosystem multifunctionality varies with land use, climatic region, soil texture, and pH, and we also investigated the relative contribution of the soil microbiome compared to soil properties and climate as indicators of multifunctionality. For each soil, we measured six functional proxies: soil basal respiration, xylosidase, N-acetylglucosaminidase, phosphatase activities, soil aggregation, and primary productivity (Table S1). Consistent with our first hypothesis, our study found the highest multifunctionality values in less-disturbed grassland and woodland soils (average multifunctionality value = 0.33 ± 0.09, n = 92, and 0.31 ± 0.10, n = 165, respectively), loam soils (0.31 ± 0.09, n = 179), acidic soils (0.30 ± 0.09, n = 290), and soils originating from temperate humid locations (0.31 ± 0.08, n = 249). On the contrary, cropland soils (0.25 ± 0.07, n = 227), alkaline soils (0.25 ± 0.09, n = 131), and soils originating from drier regions (0.25 ± 0.10, n = 122) showed the lowest multifunctionality values (Fig. 1, Table S3). Multifunctionality in cropland soils was 24.2% lower than in grasslands, and 19.4% lower than in woodlands. Our study also found that a combination of soil properties (i.e., total nitrogen, organic carbon, and microbial biomass), together with the relative abundance of a few taxa, significantly associate with multifunctionality (Fig. 2). More importantly, the soil microbiome explained a portion of variation of multifunctionality across soils. Specifically, the community composition of bacterial and fungal modules (i.e., co-occurring OTUs) explained between 2.27% and 14.08% of unique variance (Fig. 3, Table S4). Soil properties accounted for 12.2–31.4% of the unique variance in multifunctionality, whereas climate explained very little (0–1%), suggesting that climate influences multifunctionality mainly through indirect effects. At the level of individual functional proxies, the soil microbiome explained more unique variance than soil properties or climate for primary productivity and enzymatic activities (Figures S4–S7), while soil properties dominated to explain variation in soil aggregation and basal respiration (Figures S8–S9).
Results are shown for land-use types (A), climatic regions (B), soil texture types (C), and soil pH (D). Chi-squared (χ²) statistic, eta-squared (η2), and significance (p value) are indicated following Kruskal-Wallis non-parametric test. Different letters represent significant differences among groups (p value < 0.05). In the boxplots, the central line indicates the median, the box limits correspond to the 25th (Q1) and 75th (Q3) percentiles (interquartile range), and the whiskers extend to 1.5 × IQR. Points beyond the whiskers represent outliers.
Results are shown for different land uses (A1–A3), soil texture classes (B1–B3), pH classes (C1–C3), and climatic regions (D1–D3). Importance is shown for each predictor as the increase in mean-squared error (%). For each model, the ten most important predictors are displayed. Model performance is reported as cross-validated R2 (CV R2) and out-of-bag error (OOB). Asterisks indicate significance after non-parametric permutation test with 1000 permutations: **; p value < 0.010, *; p value < 0.050.
Results are grouped by land use (A), climatic region (B), soil texture type (C), and pH (D). The number of soils included in each group is indicated above each stacked bar plot as “n”. Different colors indicate unique variance explained by climate (blue), soil properties (yellow), soil microbiome (red), and shared variance (gray). Individual values for each category are available in Table S3.
Context-dependency of soil microbiome contributions to multifunctionality
Our findings indicate that the potential of the soil microbiome to act as indicator of ecosystem multifunctionality is strongly context-dependent, varying with soil characteristics and environmental conditions. We observed that in croplands, the unique variance in multifunctionality explained by the soil microbiome (13.94%) was comparable to that explained by soil properties (13.92%), suggesting that microbial composition in these intensively managed systems is an important factor explaining ecosystem performance (Fig. 3). A similar pattern was evident in soils from temperate dry climates, in neutral pH conditions (pH between 6.5 and 7.3), and in clay-rich soils, where the predictive power of the microbiome closely matched that of soil properties (Fig. 3, Table S4). These results suggest that under certain soil and environmental configurations, the microbiome may serve as a reliable indicator for assessing ecosystem functioning. In contrast, in soils from other contexts, such as those originating from woodlands and continental climatic regions, the unique contribution of the microbiome to explaining multifunctionality was noticeably lower compared to that of soil properties (Fig. 3). Our results contrast with previous studies showing that multifunctionality in intensively managed ecosystems (e.g., croplands) depends primarily on external inputs (e.g., fertilizers), as we demonstrate that these systems also rely on their soil microbiome (Gossner et al., 2016). We also suggest that the utility of the soil microbiome as indicator of multifunctionality is not universal but depends on the interaction between microbial communities and their surrounding environment, which is in line with previous research showing that soil microbial diversity depends on the interplay between soil properties and climate21,22.
At the level of individual functions, the soil microbiome explained substantial unique variance (>25%) for xylosidase activity in dry and clay-rich soils, phosphatase activity in neutral and clay-rich soils, and N-acetylglucosaminidase activity in neutral soils (Figures S4–S6). The soil microbiome was also the main predictor of primary productivity, particularly in alkaline and sandy soils (Figure S7). On the other hand, soil properties (e.g., organic carbon) were the main predictors of basal respiration. Future studies should address whether the freeze–thaw of soil samples might have disrupted microbial community signals and make respiration more strongly dependent on substrate availability than on community composition.
We hypothesized that microbial indicator taxa of multifunctionality vary across soil types, and our results confirmed the hypothesis. Random forest analyses revealed that the contribution of microbial groups to ecosystem multifunctionality is context-dependent and varies across land uses and environmental gradients (Fig. 2). In croplands, multiple microbial modules (particularly bacterial modules 3, 5, 7, and 9, and fungal modules 13 and 1) emerged as strong predictors of multifunctionality. Taxonomically, most bacterial modules were dominated by Proteobacteria, representing 30–35% of OTUs in nearly all modules (Figure S10, Supplementary dataset 1). Module 7, however, was distinct in being dominated by Actinobacteria, especially the genus Gaiella, a pattern also observed to a lesser extent in module 3. Both modules 3 and 7 also contained numerous OTUs affiliated with Sphingomonas, Nocardioides, Solirubrobacter, and Streptomyces (Actinobacteria). Module 1, together with modules 2 and 5, included a high proportion of Acidobacteria. Module 5 also contained many OTUs within Bacteroidetes, particularly taxa from the genus Flavobacteria (Figure S10). Fungal modules were largely dominated by Ascomycota, which accounted for 48–61% of OTUs across modules (Figure S11). Modules 1, 3, and 5 contained many OTUs associated with the genus Mortierella, while modules 5 and 13 included members of Glomeromycota (arbuscular mycorrhizal fungi). In woodlands, fungal module 5, enriched in Mortierella and symbiotic fungi such as Archaeorhizomyces and members of Glomeromycota, emerged among the top predictors of multifunctionality for certain soil categories (Fig. 2). Notably, Glomeromycota (arbuscular mycorrhizal fungi) have repeatedly been shown to support plant growth and contribute to ecosystem multifunctionality23,24, although they represented as small proportion of all fungal sequences in our dataset ( ≈ 1.00%). Similarly, previous research at the regional scale have identified Actinobacteria and members of Mortierella as hubs in microbial co-occurrence networks25,26. We finally acknowledge that all taxa identified here as potential indicators of multifunctionality are strict aerobes, which is consistent with sequencing bulk soil from the top 20 cm where oxygen is abundant. Future studies should extend to deeper layers, where community composition may shift and alternative microbial indicators could emerge27.
Taken together, our results open the path towards the development of biomarkers where the abundance of these taxa could be monitored through qPCR assays, integrated into soil health indices, or used as features in predictive models28. Given their sensitivity to land-use and soil conditions, these taxa may serve as early-warning markers for shifts in ecosystem multifunctionality.
To explore how environmental context shapes the relationships between microbial communities, soil properties, climate, and ecosystem multifunctionality, we employed multi-group structural equation models (SEMs). These models tested direct contributions of climate, soil properties, and the microbiome to multifunctionality, as well as indirect effects of climate and soil mediated through the microbiome (Fig. 4, Figure S3, Supplementary dataset 2). For each environmental classification (land use, climatic region, soil texture, and pH class), we compared unconstrained models (allowing regression paths to vary among groups) with constrained models that assumed equal relationships across groups. Unconstrained models consistently outperformed constrained ones based on likelihood ratio tests, supporting the idea that the effects of microbial, soil, and climatic factors on multifunctionality are context dependent (Table S5). Among the classification schemes tested, grouping by land use yielded the best model fit, as indicated by the lowest AIC in chi-squared comparisons (Fig. 4). This, together with previous research, suggests that land use is a primary organizing factor in determining how environmental drivers influence multifunctionality29.
Multi-group SEMs were built for land use categories (A), climatic regions (B), soil types (C), and pH classes (D). For each SEM, colors (red, green, blue) indicate soil grouping categories. Numbers adjacent to arrows indicate standardized path coefficients across all variables in each category. Predictor variables included in each box and a priori model are available in Figure S3. Model fit parameters are indicated as chi-squared (X2), degrees of freedom (d.f.), p value of two-sided test (P), Akaike Information Criterion (AIC), and root mean squared error of approximation (RMSEA). Full details including individual path coefficients are available in Supplementary Dataset 2.
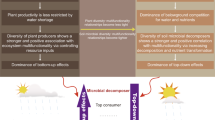
SEMs also confirmed that while the direct effects of climate, soil properties, and the microbiome were relatively balanced in croplands and grasslands, woodlands exhibited a high influence of soil properties on multifunctionality (Fig. 4). This may reflect the greater influence of accumulated organic matter and nutrient pools in woodland soils, together with microclimatic buffering by tree cover30. These factors strengthen the role of soil properties in regulating ecosystem multifunctionality in woodlands, while reducing the relative contribution of climate and microbial community composition.
SEMs showed that the direct effect of climate (air temperature and precipitation) on ecosystem multifunctionality was strongest in soils from dry regions, where water limitation potentially amplifies both direct and microbiome-mediated climate effects (Fig. 4). In soil originating from continental regions, soil properties also played a dominant direct role, exceeding their influence in other regions. Analyses by soil texture showed that in loam soils, the microbiome had a greater direct influence on multifunctionality than soil properties, a pattern not observed in clay or sandy soils, where soil properties were more dominant. Loamy soils occupy the middle of the soil texture spectrum, reflecting a balanced composition of sand, silt, and clay31. Such conditions likely support diverse and active microbial communities, enabling them to directly regulate multiple ecosystem processes. Finally, pH classification revealed that in alkaline (pH > 7.3) and neutral (pH 6.5–7.3) soils, the microbiome and soil properties contributed equally to multifunctionality (Fig. 4). In contrast, in acidic soils (pH <6.5), multifunctionality was driven primarily by soil properties, with the microbiome playing a minor role. We argue that acidic soils impose stronger physiological constraints on microorganisms, enhancing the relative importance of soil chemical properties such as pH and nutrient availability in regulating ecosystem multifunctionality32. Taken together, these findings highlight the highly context-specific nature of soil functioning and underscore the need to consider environmental background when interpreting the role of the soil microbiome. To confirm the mechanisms proposed (regarding the influence of loam texture, woodland organic matter, or acidic soils on the relationship between the soil microbiome and multifunctionality), controlled manipulative experiments are needed. Laboratory or field studies altering soil texture, pH, or organic inputs could directly test microbial contributions to multifunctionality.
European-level assessment of the link between microbiomes and functions
We here assess ecosystem multifunctionality across Europe by simultaneously comparing multiple environmental factors, including land use (croplands, grasslands, and woodlands), climatic regions (continental, temperate dry, and temperate humid), soil texture (clay, loam, and sand), and soil pH classes (acidic, neutral, and alkaline soils). Previous works have modeled multifunctionality in grasslands and croplands, showing that agricultural practices can pose risks to ecosystem functions by creating trade-offs between biodiversity and functions such as productivity and nutrient cycling, with pesticide use and fertilization emerging as key drivers33,34. Similarly, a European-scale study covering 94 soils in 13 countries found that interactions between land use and climatic zones drive ecosystem functions in croplands and grasslands35. More recently, Sünnemann and collaborators (2023) found a Europe-wide decline in ecosystem multifunctionality under rising temperatures and dry conditions, worsened by fertilizer and pesticide application. Our study adds primary productivity as an additional ecosystem function, thereby moving beyond soil-focused analyses toward a broader ecosystem perspective of multifunctionality. We argue that future studies should incorporate direct measurements of actual biomass, such as kilograms of dry weight per hectare, because these are likely to provide more accurate estimates than satellite-derived proxies36. Remote sensing has been used in similar studies and captures vegetation greenness or productivity indices at coarse resolution, while field-based biomass measurements directly reflect the material available for carbon storage, reducing uncertainty and improving the precision of multifunctionality assessments37,38. Finally, ecosystem multifunctionality relies on the number of functions integrated7. In this study, six ecosystem functions were measured across all the 484 sites, including primary productivity, which is a key function. Future studies should also incorporate direct measurements of soil gas emissions (e.g., CO₂, CH₄, N₂O), as these are critical processes linking microbial activity to climate regulation39,40.
Considering that habitat conversion to croplands will likely increase worldwide for food production and to satisfy needs of a growing population, our results stress that the capacity of ecosystems to supply multiple functions simultaneously could be compromised if conversion to croplands is preferred. In line with this, Jeanneret and collaborators (2021) estimated that conversion to arable lands negatively impacts biodiversity of vascular plants and arthropods across Europe. Similarly, conversion to cropland causes homogenization of microbial communities41,42. A similar study found that European croplands were particularly abundant in potentially pathogenic fungi compared to grasslands and woodlands, but the richness of beneficial taxa (e.g., mycorrhizal fungi) decreased43. Our results add to these that the conversion of natural ecosystems to croplands not only has an impact on soil biodiversity but also on multifunctionality. Moreover, we show that the contribution of the soil microbiome to multifunctionality is context-dependent. We acknowledge that comparing multifunctionality across highly disturbed ecosystems (croplands) and natural or semi-natural ecosystems (woodlands or grasslands) is challenging, since these are very different ecosystem types with different disturbance regimes and the dynamics of each ecosystem might differentially impact multifunctionality. For example, woodlands are stable ecosystems dominated by trees where primary productivity is rather constant over time, whereas croplands are subjected to crop rotations, whereas grasslands are often dominated by annual herbs and forbs, with consequences for organic carbon stocks, and productivity44. Taken together, we argue that despite being part of the same landscape, soils from different land uses, textures, climatic regions, and pH classes respond to distinct microbial predictors of multifunctionality. This indicates that monitoring should adopt context-specific microbial indicators rather than a uniform approach.
Our continental analysis provides evidence that ecosystem multifunctionality is shaped by both environmental context and biotic composition, with land use, soil properties, climate, and the soil microbiome each contributing to ecosystem multifunctionality. Multifunctionality was highest in grasslands and woodlands, in loam-textured and acidic soils (pH <6.5), and in temperate humid regions. The best predictors of ecosystem multifunctionality overall were microbial biomass and nitrogen content. Random forest and variance partitioning analyses further revealed that the soil microbiome (i.e., co-occurring microbial OTUs) explains a substantial and context-dependent share of multifunctionality. The soil microbiome was the main factor contributing to variation in enzymatic activities and primary productivity, while soil basal respiration and soil aggregation were mostly explained by soil properties. We also found that modules dominated by Actinobacteria (e.g., Gaiella, Sphingomonas), Acidobacteria, and Mortierella frequently emerged as strong predictors of ecosystem multifunctionality. Our use of multi-group structural equation models further highlights that the strength and nature of relationships among climate, soil properties, the microbiome, and multifunctionality differ across environmental classifications. Among all groupings, land use emerged as the most influential structuring variable. Collectively, these findings underscore the need for context-aware approaches in using microbial indicators for soil monitoring. As land-use change intensifies globally, understanding how local conditions modulate microbiome-function relationships is essential to designing effective, scalable soil health strategies and sustaining ecosystem services under environmental change.
Methods
Field survey and soil sampling
Our study was built upon the EU Statistical Office’s Land Use and Coverage Area Frame Survey (LUCAS) Soil, the largest pan-European scheme for assessing soil characteristics in relation to land cover and land use45,46. The sampling approach used a composite strategy, where the final sample at each location consisted of five combined topsoil (0–20 cm) subsamples. The initial subsample was obtained precisely at the coordinates of the designated LUCAS point, while the four additional subsamples were collected 2 m from the central point, aligning with the cardinal directions (North, East, South, and West). To minimize the impact of seasonality and temporal variation, all samples were collected over the shortest possible timeframe during spring/summer 2018. For this study, a total of 484 soil samples were collected, comprising 165 from woodlands, 92 from grasslands, and 227 from croplands. Based on the Köppen–Geiger climate classification, 113 samples originated from sites in a continental climate, 122 from temperate dry regions, and 249 from temperate humid regions (Figure S1).
Soil edaphic factors and climatic variables
Soil samples were used to determine a range of edaphic factors following standard procedures46. Total phosphorus content (mg kg-1) was measured by the ISO 11263:1994 protocol following phosphorus solubilization in a sodium hydrogen carbonate solution. Total nitrogen content (g kg−1) was measured following the ISO 11261:1995 protocol. Extractable potassium content (mg kg−1) was determined with atomic absorption spectrophotometry47. Soil pH was measured according to the ISO 10390:1994 standard using 0.01 M CaCl₂ as the extractant, while the proportions of silt, clay, and sand were determined using laser diffraction particle size analysis following ISO 13320:2009. Based on these measurements, samples were further grouped by pH and texture. For soil texture classification, we used the USDA system integrated within the soil texture wizard48. We classified 290 soils as acidic (pH < 6.5), 63 as neutral (pH 6.5–7.3), and 131 as alkaline (pH > 7.3). Regarding texture, 129 soils were identified as clay, 179 as loam, and 176 as sand.
Soil organic carbon was measured following the ISO 10694:1995 protocol. Briefly, the total carbon content in each sample was determined using an elemental analyzer following dry combustion and corrected for carbonate content (ISO 10693:1994), resulting in soil organic carbon (g kg−1). Soil bulk density was measured following the ISO 11272:2017 procedure49. Soil bulk density was expressed as the dry weight of soil divided by its volume (g cm−3), which includes the volume of soil particles and the pores among them50. Soil microbial biomass (expressed as μg Cmic g−¹ soil dry weight) was estimated as a proxy using substrate-induced respiration, based on the respiration response to glucose addition measured directly after potential basal respiration on the same soil samples. Climatic variables, including monthly mean air temperature and precipitation (averaged values over the 1970–2000 period) were obtained for each sampling location from the WorldClim database (worldclim.org).
Soil microbial community composition
Soil samples (n = 484) were analyzed for bacterial and fungal biodiversity using DNA metabarcoding. DNA was extracted using the Qiagen DNeasy PowerSoil HTP 96 Kit, with three 0.2 g aliquots per sample pooled post-extraction51. DNA quality and quantity were assessed using the Qubit™ 1X dsDNA HS Assay Kit. PCR amplification was performed in triplicate using 5 × HOT FIREPol® Blend Master Mix (Solis BioDyne, Tartu, Estonia) in 25 μl volume. The PCR conditions for bacterial amplification included 55 °C annealing temperature, 26 cycles, and 1.5 ng of DNA template in 1 µl, following an optimized protocol derived from the Earth Microbiome Project (https://earthmicrobiome.org/protocols-and-standards/16s) to minimize PCR bias while ensuring sufficient yield for sequencing. For fungal amplification, the PCR conditions included an annealing temperature of 55 °C, 30 cycles, and 1.5 ng of DNA template in 1 µl. Amplified DNA was tagged with multiplex identifier tags, pooled, and verified on a TBE 1% agarose gel. Primer sets for barcode amplification of 16S rRNA gene were 515 F (GTGYCAGCMGCCGCGGTAA) and 926 R (GGCCGYCAATTYMTTTRAGTTT), targeting the bacterial V4-V5 hypervariable region52,53 and ITS9mun (GTACACACCGCCCGTCG) and ITS4ngsUni (CGCCTSCSCTTANTDATATGC) for the fungal ITS region54. Sequencing was performed using Illumina MiSeq platform with 2 ×300 paired-end mode for bacterial amplicons and PacBio Sequel II platform for fungal amplicons.
The Illumina and PacBio amplicon data (for bacteria and fungi, respectively) were demultiplexed using LotuS2 and paired-end reads were assembled using FLASH 1.2.1055,56. For bacteria, zero-radius operational taxonomic units (zOTUs) were generated using the UPARSE algorithm (usearch version 10.0.02457). The process involved merging paired-end reads, trimming off 16S primer sequences, quality filtering, and denoising. For fungi, 98%-OTUs were obtained following the VSEARCH algorithm58. Our datasets, consisting of 484 soils, included 79,593 zOTUs (bacteria) and 35,152 98%-OTUs (fungi). The bacterial dataset was rarefied at 40,000 sequences per sample, and the fungal dataset was rarefied at 1000 sequences per sample. Taxonomy was assigned to the bacterial dataset using the Ribosomal Database Project v1659 and to the fungal dataset using MegaBLAST searches against the UNITE 9.1 database60.
To identify groups of co-occurring microbial taxa, we used the WGCNA (Weighted Gene Co-expression Network Analysis) R package61. OTU tables containing relative abundances were first formatted with samples as rows and OTUs as columns. We assessed the data for missingness using the goodSamplesGenes function, followed by hierarchical clustering (hclust) to detect potential sample outliers. A soft-thresholding power was selected based on scale-free topology and mean connectivity criteria (pickSoftThreshold, with blockSize = 1000). An adjacency matrix was then constructed using this threshold and transformed into a Topological Overlap Matrix (TOM) to compute dissimilarity between OTUs. Taxa were clustered using hierarchical clustering, and modules were defined via the cutreeDynamic function, with a minimum module size of 15 for fungi and 100 for bacteria. Module eigengenes were calculated (moduleEigengenes function) and used as indicators of microbial community composition. Finally, OTUs were taxonomically annotated and assigned to their corresponding modules for interpretation. This network-based approach allowed us to reduce community data dimensionality and link modules to environmental variables and multifunctionality.
Ecosystem multifunctionality
We assessed six functional proxies across 484 soils (Table S1): soil basal respiration, three enzymatic activities (N-acetylglucosaminidase, phosphatase, and xylosidase), net primary productivity, and soil aggregate stability. These proxies capture different ecosystem processes and associated services. Soil basal respiration reflects microbial activity and relates to climate regulation. N-acetylglucosaminidase breaks down chitin and complex carbohydrates, informing nitrogen cycling and organic matter decomposition. Phosphatase releases inorganic phosphate from organic compounds, a key process in phosphorus cycling. Xylosidase cleaves xylose from hemicellulose, contributing to plant litter decomposition and carbon cycling. Net primary productivity reflects primary production and food provision, while soil aggregate stability represents soil structure, linked to erosion regulation, flood regulation, and habitat provision. Our selection of functions was based on biotic or abiotic processes that can be measured as a rate or directly contribute to ecosystem services7.
Potential soil basal respiration (μL O2 g−1 soil dry weight h−1) was measured using an O2-microcompensation apparatus: respiration was determined at 20 °C using the pressure difference between a sealed chamber containing the sample and a control chamber at atmospheric pressure. For this, 5–7 g of thawed soil, which had been previously acclimated for five days at 4 °C and sieved at 2 mm, was used. Measurements of potential basal respiration lasted from 22 to 42 hours, depending on the time taken for respiration rates to reach detectable levels. A timeframe of 5–7 consecutive measurement hours, in which respiration was stable, was used to calculate the average potential basal respiration of the sample62,63. The activities of the soil enzymes N-acetylglucosaminidase, xylosidase, and phosphatase were measured based on 4-methylumbelliferone (MUF)-coupled substrates63. Briefly, fresh soil samples (250 mg each) were suspended in 50 μM acetate buffer (pH 5), sonicated to disrupt soil aggregates, and incubated at 25 °C for 60 min in 96-well microplates containing substrates, MUF dilutions for quenching and extinction coefficients, and controls for substrate and soil suspensions. Reactions were stopped with 2 M NaOH, and fluorescence was measured in eight technical replicates using an Infinite 200 PRO instrument (Tecan Group, Männedorf, Switzerland). Enzymatic activities were quantified as substrate turnover rates, expressed as nmol of MUF per gram of dry soil per hour. Net Primary Productivity (NPP) was estimated at each sampling location using MODerate resolution Imaging Spectroradiometers (MODIS) imagery from NASA’s Terra and Aqua satellites, specifically through the MOD17 PSN/NPP algorithm. NPP was calculated as the sum of eight-day net photosynthesis (PSN) products based on the Fraction of Absorbed Photosynthetically Active Radiation (FAPAR). FAPAR, which indicates the solar radiation absorbed by plants, is derived using variables such as the maximum radiation conversion efficiency, ground temperature, and vapor pressure64. The NPP data for 2018, aligning with LUCAS sampling locations, were averaged across ±6 months surrounding the sampling times at a 500 m resolution, and expressed as g C m−2 yr−1. Soil aggregation was determined from the percentage of water-stable aggregates (WSA) following the wet-sieving method63,65. Briefly, 4 g of fresh soil (FM) were placed into 0.25-mm sieves and allowed to rewet by capillarity for 5 min. Samples were then wet-sieved for 3 min and dried overnight at 70 °C, after which dry matter was measured. Coarse Matter (CM) was measured after another night at 70 °C. Finally, the percentage of water-stable aggregates was calculated as follows:
To quantify ecosystem multifunctionality, we calculated four distinct indices based on the six measured functions. The first index was a simple average, where all functions were standardized to a 0–1 scale and then averaged, assuming equal contribution of each function. The second index applied a z-score transformation, standardizing each function using the formula z = (X − µ)/σ, where X is the function value, µ is the mean, and σ is the standard deviation6,66. This approach facilitates direct comparability among functions by expressing values in terms of standard deviations from the mean. The third index was a weighted average, which accounted for the fact that three of the six functions were enzymatic activities representing similar processes. In this index, each enzymatic function was assigned a relative weight of 0.33, while non-redundant functions retained full weight. In addition, we calculated a multiple-threshold index by assessing the number of functions (0–6) performing above predefined thresholds (25%, 50%, and 75% of the maximum observed value). We also analyzed each function individually to provide detailed insights into how predictors influenced both single functions and overall multifunctionality. Because the calculated indices were highly correlated (Figure S2), we selected the weighted index for downstream analyses (see next section). This index best reflects ecosystem multifunctionality, as it corrects for the overrepresentation of enzymatic activities (3 out of the 6 measured proxies).
Statistical analyses
All statistical analyses were conducted in R version 4.2.167. We explored the relative contribution of the soil microbiome, soil properties, and climate in explaining patterns on multifunctionality by means of random forest analyses, variance partitioning, and structural equation models (SEMs). We first investigated how multifunctionality and individual soil functions varied across different environmental groupings. Specifically, we categorized our 484 soil samples based on land use (cropland = 227, grassland = 92, woodland = 165), climatic region (continental = 113, temperate dry = 122, temperate humid = 249), soil texture (clay = 129, loam = 179, sand = 176), and pH class (acidic = 290, neutral = 63, alkaline = 131). We then used Kruskal-Wallis tests followed by pairwise Dunn’s tests (Bonferroni correction) to assess group differences. For each test, we reported the chi-squared statistic, p value, and eta-squared as a measure of effect size. Since overlaps among soil groupings could confound interpretations, we quantified the percentage of overlap between categories. With few exceptions, overlap was limited. However, woodland, continental, and sandy soils largely coincided with acidic soils (Table S2). This indicates that patterns attributed to these groupings may partly reflect their strong association with soil pH.
To identify key predictors of multifunctionality (weighted index), we employed Random Forest (RF) regression using the rfPermute package with 1000 trees68. Models were trained on standardized data and evaluated using out-of-bag (OOB) R² and 10-fold cross-validation (repeated 3 times). Variable importance was assessed based on the increase in mean squared error when each predictor (microbial modules, soil properties, climate) was permuted.
Next, we partitioned the variance in multifunctionality using adjusted R² to determine the unique and shared contributions of three predictor groups: climate (precipitation, temperature), edaphic factors (excluding total nitrogen due to collinearity with SOC), and microbial composition (bacterial and fungal modules). Unique and shared contributions were calculated through a nested linear models.
Finally, to disentangle direct and indirect effects, we used multi-group Structural Equation Models (SEMs) with the lavaan package69. Predictors were grouped into climate, soil properties, and microbial variables (see Figure S3 for a priori model). We fit SEMs and estimated standardized path coefficients. A constrained model was also fit to test whether relationships differed significantly across groups. Direct effects on multifunctionality and indirect effects on the microbiome were summarized by group and category.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that supports the findings of this study are freely available in figshare with the identifier https://doi.org/10.6084/m9.figshare.28645625. Raw DNA sequences can be accessed through the European Soil Data Centre (ESDAC) portal: https://esdac.jrc.ec.europa.eu/content/soil-biodiversity-dna-bacteria-and-fungi. The raw data (DNA sequences) generated in this study have been deposited in the Sequence Read Archive (SRA) database under BioProject ID PRJNA952168. Detailed information on the taxonomic composition of bacterial and fungal modules is available as Supplementary Dataset 1. Detailed information on Structural Equation Models (SEMs) results is availabel as Supplementary Dataset 2.
Code availability
Code used to perform all analyses described in this study is freely available at https://github.com/fromerob/Multifunctionality.git.
References
Anthony, M. A., Bender, S. F. & van der Heijden, M. G. A. Enumerating soil biodiversity. Proc. Natl. Acad. Sci. 120, e2304663120 (2023).
Adhikari, K. & Hartemink, A. E. Linking soils to ecosystem services—a global review. Geoderma 262, 101–111 (2016).
Calzolari, C. et al. A methodological framework to assess the multiple contributions of soils to ecosystem services delivery at regional scale. Geoderma 261, 190–203 (2016).
Greiner, L., Keller, A., Grêt-Regamey, A. & Papritz, A. Soil function assessment: review of methods for quantifying the contributions of soils to ecosystem services. Land Use Policy 69, 224–237 (2017).
Banerjee, S. & van der Heijden, M. G. A. Soil microbiomes and one health. Nat. Rev. Microbiol 21, 6–20 (2023).
Byrnes, J. E. K. et al. Investigating the relationship between biodiversity and ecosystem multifunctionality: challenges and solutions. Methods Ecol. Evol. 5, 111–124 (2014).
Garland, G. et al. A closer look at the functions behind ecosystem multifunctionality: A review. J. Ecol. 109, 600–613 (2021).
Creamer, R. E., Barel, J. M., Bongiorno, G. & Zwetsloot, M. J. The life of soils: Integrating the who and how of multifunctionality. Soil Biol. Biochem 166, 108561 (2022).
Romero, F. et al. Soil microbial biodiversity promotes crop productivity and agro-ecosystem functioning in experimental microcosms. Sci. Total Environ. 885, 163683 (2023).
Romero, F., Jiao, S. & van der Heijden, M. G. A. Impact of microbial diversity and pesticide application on plant growth, litter decomposition and carbon substrate use. Soil Biol. Biochem. 208, 109866 (2025).
Delgado-Baquerizo, M. et al. Microbial diversity drives multifunctionality in terrestrial ecosystems. Nat. Commun. 7, 10541 (2016).
Doran, J. W. & Zeiss, M. R. Soil health and sustainability: managing the biotic component of soil quality. Appl. Soil Ecol. 15, 3–11 (2000).
Knight, C. G. et al. Soil microbiomes show consistent and predictable responses to extreme events. Nature 636, 690–696 (2024).
George, P. B. L. et al. Divergent national-scale trends of microbial and animal biodiversity revealed across diverse temperate soil ecosystems. Nat. Commun. 10, 1107 (2019).
Riggi, L. G. A. et al. Establishing assessment criteria for soil bioindicators: insights from case studies in Europe. Ecol. Indic. 178, 114063 (2025).
Tang, B. et al. Mycorrhization enhances plant growth and stabilizes biomass allocation under drought. Glob. Change Biol. 30, e17438 (2024).
Bardgett, R. D. & van der Putten, W. H. Belowground biodiversity and ecosystem functioning. Nature 515, 505–511 (2014).
Delgado-Baquerizo, M. et al. Microbial richness and composition independently drive soil multifunctionality. Funct. Ecol. 31, 2330–2343 (2017).
Jing, X. et al. The links between ecosystem multifunctionality and above- and belowground biodiversity are mediated by climate. Nat. Commun. 6, 8159 (2015).
van Zanten, H. H. E. et al. Circularity in Europe strengthens the sustainability of the global food system. Nat. Food 4, 320–330 (2023).
Labouyrie, M. et al. Interaction effects of pH and land cover on soil microbial diversity are climate-dependent. Environ. Microbiol. 26, e16572 (2024).
Siles, J. A. et al. Land use interacts with climate to influence microbial diversity-to-biomass ratios across Europe via soil organic carbon and nitrogen. Mol. Ecol. n/a, e17806 (2025).
Romero, F., Argüello, A., de Bruin, S. & van der Heijden, M. G. A. The plant–mycorrhizal fungi collaboration gradient depends on plant functional group. Funct. Ecol. 37, 2386–2398 (2023).
van der Heijden, M. G. A. et al. Mycorrhizal fungal diversity determines plant biodiversity, ecosystem variability and productivity. Nature 396, 69–72 (1998).
Ishimoto, C. K. et al. Microbial co-occurrence network and its key microorganisms in soil with permanent application of composted tannery sludge. Sci. Total Environ. 789, 147945 (2021).
Toju, H., Tanabe, A. S. & Sato, H. Network hubs in root-associated fungal metacommunities. Microbiome 6, 116 (2018).
Xu, Z. et al. Depth-dependent effects of tree species identity on soil microbial community characteristics and multifunctionality. Sci. Total Environ. 878, 162972 (2023).
M, H. S. et al. Bacteria as emerging indicators of soil condition. Appl. Environ. Microbiol 83, e02826–16 (2016).
Allan, E. et al. Land use intensification alters ecosystem multifunctionality via loss of biodiversity and changes to functional composition. Ecol. Lett. 18, 834–843 (2015).
Wardle, D. A. et al. Ecological linkages between aboveground and belowground biota. Science 304, 1629–1633 (2004).
Brady, N. C., Weil, R. R. & Weil, R. R. The Nature and Properties of Soils. Vol. 13 (Prentice Hall Upper Saddle River, 2008).
L, L. C., Micah, H., Rob, K. & Noah, F. Pyrosequencing-based assessment of soil ph as a predictor of soil bacterial community structure at the continental scale. Appl. Environ. Microbiol. 75, 5111–5120 (2009).
Vazquez, C., de Goede, R. G. M., Rutgers, M., de Koeijer, T. J. & Creamer, R. E. Assessing multifunctionality of agricultural soils: Reducing the biodiversity trade-off. Eur. J. Soil Sci. 72, 1624–1639 (2021).
Wittwer, R. A. et al. Organic and conservation agriculture promote ecosystem multifunctionality. Sci. Adv. 7, eabg6995 (2021).
Zwetsloot, M. J. et al. Soil multifunctionality: Synergies and trade-offs across European climatic zones and land uses. Eur. J. Soil Sci. 72, 1640–1654 (2021).
Scurlock, J. M. O., Johnson, K. & Olson, R. J. Estimating net primary productivity from grassland biomass dynamics measurements. Glob. Change Biol. 8, 736–753 (2002).
Running, S. W. et al. A continuous satellite-derived measure of global terrestrial primary production. Bioscience 54, 547–560 (2004).
Rillig, M. C. et al. Increasing the number of stressors reduces soil ecosystem services worldwide. Nat. Clim. Change 13, 478–483 (2023).
Bahram, M. et al. Structure and function of the soil microbiome underlying N2O emissions from global wetlands. Nat. Commun. 13, 1430 (2022).
Jeanneret, P. et al. An increase in food production in Europe could dramatically affect farmland biodiversity. Commun. Earth Environ. 2, 183 (2021).
Peng, Z. et al. Land conversion to agriculture induces taxonomic homogenization of soil microbial communities globally. Nat. Commun. 15, 3624 (2024).
Gossner, M. M. et al. Land-use intensification causes multitrophic homogenization of grassland communities. Nature 540, 266–269 (2016).
Labouyrie, M. et al. Patterns in soil microbial diversity across Europe. Nat. Commun. 14, 3311 (2023).
Henderson, K. A., Bauch, C. T. & Anand, M. Alternative stable states and the sustainability of forests, grasslands, and agriculture. Proc. Natl. Acad. Sci. 113, 14552–14559 (2016).
Orgiazzi, A. et al. LUCAS Soil Biodiversity and LUCAS Soil Pesticides, new tools for research and policy development. Eur. J. Soil Sci. 73, e13299 (2022).
Orgiazzi, A., Ballabio, C., Panagos, P., Jones, A. & Fernández-Ugalde, O. LUCAS Soil, the largest expandable soil dataset for Europe: a review. Eur. J. Soil Sci. 69, 140–153 (2018).
Yeung, V., Miller, D. D. & Rutzke, M. A. Atomic absorption spectroscopy, atomic emission spectroscopy, and inductively coupled plasma-mass spectrometry. Food Anal. 129, 150 (2017).
Moeys, J. The soil texture wizard: R functions for plotting, classifying, transforming and exploring soil texture data. CRAN. R-Project 1–104 (2018).
Fernández-Ugalde, O. et al. LUCAS 2018 Soil Module—Presentation of Dataset and Results. https://doi.org/10.2760/215013 (Publications Office of the European Union, 2022).
Panagos, P. et al. Soil bulk density assessment in Europe. Agric. Ecosyst. Environ. 364, 108907 (2024).
Morita, H. & Akao, S. The effect of soil sample size, for practical DNA extraction, on soil microbial diversity in different taxonomic ranks. PLoS ONE 16, e0260121 (2021).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. 108, 4516–4522 (2011).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Tedersoo, L. & Anslan, S. Towards PacBio-based pan-eukaryote metabarcoding using full-length ITS sequences. Environ. Microbiol Rep. 11, 659–668 (2019).
Özkurt, E. et al. LotuS2: an ultrafast and highly accurate tool for amplicon sequencing analysis. Microbiome 10, 1–14 (2022).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Tedersoo, L. et al. The Global Soil Mycobiome consortium dataset for boosting fungal diversity research. Fungal Divers. 111, 573–588 (2021).
Cole, J. R. et al. Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42, D633–D642 (2014).
Nilsson, R. H. et al. The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 47, D259–D264 (2019).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 1–13 (2008).
Smith, L. C. et al. Large-scale drivers of relationships between soil microbial properties and organic carbon across Europe. Glob. Ecol. Biogeogr. 30, 2070–2083 (2021).
Sünnemann, M. et al. Climate change and cropland management compromise soil integrity and multifunctionality. Commun. Earth Environ. 4, 394 (2023).
Romero, F. et al. Soil health is associated with higher primary productivity across Europe. Nat. Ecol. Evol. 8, 1847–1855 (2024).
Seybold, C. A. & Herrick, J. E. Aggregate stability kit for soil quality assessments. Catena 44, 37–45 (2001).
Maestre, F. T. et al. Plant species richness and ecosystem multifunctionality in global drylands. Science 335, 214–218 (2012).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/.
Archer, E. rfPermute: estimate permutation p-values for random forest importance metrics. R package version 1, (2016).
Rosseel, Y. et al. Package ‘lavaan’. Retrieved June 17, 2017 (2017).
Acknowledgements
F.R. acknowledges funding from the Novo Nordisk Foundation through a Postdoctoral fellowship (grant reference number NNF24OC0094454). The LUCAS Survey is coordinated by Unit E4 of the Statistical Office of the European Union (EUROSTAT). The LUCAS Soil sample collection is supported by the Directorate-General Environment (DG-ENV), Directorate-General Agriculture and Rural Development (DG-AGRI) and Directorate-General Climate Action (DG-CLIMA) of the European Commission. M.D-B. acknowledges support from the Spanish Ministry of Science and Innovation for the I + D + I project PID2020-115813RA-I00 funded by MCIN/AEI/10.13039/501100011033. M.v.d.H and F.R. acknowledge funding from the Swiss National Science Foundation (Switzerland) through grant no. 310030–188799 and from the European Union Horizon 2020 research and innovation program under grant agreement no. 862695 EJP SOIL-MINOTAUR.
Author information
Authors and Affiliations
Contributions
F.R. contributed to conceptualization, methodology, formal analysis, investigation, data curation, writing – original draft, writing – review & editing, and visualization. M.L. contributed to conceptualization, methodology, formal analysis, investigation, writing – review & editing, and data curation. A.O. contributed to conceptualization, methodology, resources, writing – review & editing, and data curation. C.B. contributed to methodology and resources. P.P. contributed to conceptualization, methodology, resources, writing – review & editing, project administration, and data curation. A.J. contributed to conceptualization, methodology, resources, writing – review & editing, project administration, and data curation. L.T. and M.B. contributed to resources, writing – review & editing, and data curation. N.E., M.S., and C.A.G. contributed to resources and writing – review & editing. D.T. contributed to methodology. I.R. and S.J. contributed to writing – review & editing. S.M. contributed to project administration and funding acquisition. M.C.R. and A.L. contributed to resources and writing – review & editing. M.D.B. contributed to writing – review & editing. M.v.d.H. contributed to conceptualization, methodology, resources, writing – original draft, writing – review & editing, supervision, project administration, and funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xiangang Hu, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Romero, F., Labouyrie, M., Orgiazzi, A. et al. The soil microbiome as an indicator of ecosystem multifunctionality in European soils. Nat Commun 17, 705 (2026). https://doi.org/10.1038/s41467-025-67353-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67353-9