Abstract
Filamentous phages are non-lytic phages mutually beneficial to their bacterial hosts. In Neisseria meningitidis, the filamentous phage MDA is associated with invasive diseases thanks to its key role in the formation of biofilm during epithelium colonisation. The infection model for filamentous phages has been defined for phages Ff and CTX. These phages bind to the tips of bacterial pili before being translocated into the periplasm of their hosts. The aim of this study is to investigate the relationship between filamentous phage infection and type IV pili, using the bacterium Neisseria meningitidis and the bacteriophage MDA as model organisms. We show that MDAΦ rather binds to type IV pili along their entire length with preferential binding to positively charged variants of the major fibre-forming pilin, demonstrating a role for antigenic variation in phage infection. Strikingly, bacteria expressing the more positively charged pilin are also the most adhesive, suggesting that MDAΦ primarily target the most adhesive bacteria. Finally, we show that adhesion to human cells is sufficient to amplify the phage-positive meningococcal population. Overall, this study reveals how a filamentous phage can target hyperadhesive bacterial variants and promote their selection, thereby establishing a link between phage infection and bacterial colonisation.
Similar content being viewed by others
Introduction
The contribution of phages to the virulence of their bacterial hosts has been studied mainly for Caudovirales, which represent 96% of the known phages1. In contrast, little is known about inoviruses, a family of bacteriophages carrying a single-stranded circular DNA with unique morphology and infectious cycle characteristics, to which the filamentous phage subfamily belongs. However, filamentous phages are found in several major pathogenic species2. The first filamentous phage of this family was described in Escherichia coli in the 1960s by several groups3,4,5,6. A second well-known filamentous phage, hosted by Vibrio cholerae, was discovered by Matthew K. Waldor and John J. Mekalanos. This phage, called CTXΦ, encodes the cholera toxin genes, which promotes the dissemination of CTX-carrying V. cholerae7. The main feature is the absence of bacteria lysis during the infectious cycle8. The Pf4 phage of Pseudomonas aeruginosa is another example. It remains inactive until the bacteria perceive environmental signals (such as oxidative stress), then Pf4 genes are among the most induced genes in P. aeruginosa biofilms where Pf4 filamentous phages contribute to biofilms as a structural components and as a physical barrier to antibiotics and as an important immune modulator during infection9. Finally, filamentous phages have been described in Neisseria species and in particular in Neisseria meningitidis where the filamentous phage named MDA for Meningococcal Disease Associated phage, also known as Nf1 phage10, plays a key role in biofilm formation and is associated with virulence11,12,13,14. Other filamentous bacteriophages contribute to the virulence of many bacteria in a variety of ways2.
Filamentous phages that infect diderm bacteria must cross both the outer and inner membranes to infect their target. Although the cell surface receptors are unknown for the majority of filamentous phages described so far, a role for retractable bacterial pili has been suspected for their entry, such as Ff and related phages that use the sexual/conjugative pilus F of E. coli2,15. Early observations of f1 phages by electron microscopy suggested that the phages bind to the tip of the pilus15. This was confirmed experimentally for CTXΦ, in which the phage adsorption protein pIII interacted directly with the pilus tip-subunit TcpB16. In the case of N. meningitidis, we have previously shown that MDA phage entry into the host requires retractable type IV pili17, but the bacterial proteins involved in the MDAΦ-pili interaction are not known.
Thanks to pili retraction, the phage is thought to cross the bacterium’s outer membrane through the pilus secretin and then enter the periplasm, where, as previously described, the Ff or CTX phages interact with the TolQRA system for the disassembly and entry of the single-stranded DNA viral genome into the cytosol. In the absence of the TolQR complex, the homologous ExbBD complex can perform phage uptake with TolA18,19. Overall, our knowledge of the molecular mechanisms by which filamentous phages interact with their associated pilus and enter bacteria is incomplete for most filamentous phages and remains to be determined.
N. meningitidis is a commensal bacterium, commonly present asymptomatically in the human nasopharynx. Under certain circumstances, the bacterium can enter the bloodstream and cause sepsis and/or meningitis after crossing the blood-brain barrier20. Survival in the extracellular fluid is dependent on a small number of virulence factors such as the capsular polysaccharide, the iron chelation systems and factor H binding protein21,22. In addition, type IV pili are essential for blood vessel colonisation, a key step in the pathogenesis of N. meningitidis23,24,25,26. Other bacterial attributes, such as Opa proteins, are likely to play an essential role in nasopharyngeal colonisation27. While few virulence factors are associated with meningococcal invasiveness, no virulence factor provides a clear explanation for why certain strains of N. meningitidis are responsible for outbreaks of invasive infection. One exception is the presence of an 8 kb island in the genome of invasive isolates, the meningococcal filamentous MDA prophage10,12,13,14,17. The MDA prophage may be found up to 4 times per genome, with partial heterogeneity in these sequences suggesting that a strain can be re-infected by the same phage or by a phage from another strain28,29. Previous results have suggested that MDA phages enhance epithelial cell colonisation by promoting bacterial-bacterial interactions and increasing bacterial biomass in the nasopharynx11, which in turn should increase the frequency of bacterial dissemination in the bloodstream and/or the spread of bacteria within the human population.
The dissemination of the MDA phage is likely to be associated with the hijacking of the type IV pilus machinery, a feature that appears to be shared with most other inoviruses such as CTX and Pf4. Type IV pili are highly dynamic filaments that generate tensive forces during cycles of elongation and retraction, powered by a dedicated elongation motor called PilF and retraction motor called PilT30. Filament biogenesis relies on a complex multi-protein machinery that, in N. meningitidis, contains 15 proteins essential for biogenesis (PilC1 and C2, D, E, F, G, H, I, J, K, M, N, O, P, Q, W), PilE being the pili-forming pilin and PilQ the secretin through which the pilus crosses the outer membrane. As determined by cryo-electron tomography, the type IV pili biogenesis machinery is a multilayered structure spanning the entire cell envelope31,32. Meningococcal strains express either a class I or a class II pilE gene33. Class I pilE are subject to antigenic variation and type IV pili are therefore considered to be one of the most variable structures on the meningococcal surface. This pilE gene consists of a constant region and a variable domain that can vary by recombination with one of the eight pilS silent cassettes. In addition, in the early 1990s, it was shown that antigenic variation in PilE plays an important role in cell adhesion to the host and escape from the immune response34,35. Conversely, the class II pilins do not undergo antigenic variation and form a class of conserved pilin subunits36. In addition, the PilE protein is highly post-translationally modified (glycosylation and addition of phosphoglycerol)37. Finally, type IV pili have been implicated in numerous functions including bacterial interaction with human cells, DNA uptake, twitching motility and bacterial-bacterial interactions34,38,39,40.
Here, we use N. meningitidis as a model to dissect the molecular mechanism of the interaction between filamentous phages and type IV pili, and how this shapes the bacterial population. Unlike most filamentous phages, we demonstrate that the type IV pili tip adhesin is not involved in infection. Conversely, we observe the binding of MDAΦ to type IV pili along their fibre, an interaction that took advantages of the positive charges carried by certain PilE variants that are also strongly associated with type IV pili-mediated adhesion, suggesting a key role for antigenic variation in MDAΦ infection. Finally, we show that adhesion to human cells is sufficient to allow enrichment of a meningococcal population with phage-positive bacteria, demonstrating the association between adhesion and the presence of the MDAΦ, two features associated with the invasiveness of N. meningitidis.
Results
MDAΦ infection is dependent on retractable type IV pili and specific pilE sequences
Interaction of inoviruses with their host bacteria and subsequent entry into the cytosol is thought to be highly pili-dependent. In the case of MDAΦ, we have previously shown that both the major pilin PilE and the retraction ATPase PilT are required for phage transduction, whereas other minor pilins of the fibre (PilV, ComP, and PilX) are not required17. To gain further insight into the pili-phage interaction, we extended our previous screen by assessing the efficacy of phage transduction in a comprehensive library of individual mutants of the type IV pili machinery, including proteins involved in fibre assembly or in the machinery itself, (PilD, F, G, P, Q, T, T2, U, W, Z and TfpC, TsaP and NMA0415) and proteins that compose the fibre (PilE, PilV, PilX, ComP) or the tip (PilH, I, J, K, C1, and C2) (Fig. 1a–d). Transduction efficiency was quantified in a strain deleted for the MDA prophage (Z∆MDA) and using a purified MDA phage carrying a spectinomycin resistance gene (MDA(orf6-aadA1)Φ) for colony forming unit (CFU) quantification (Fig. 1a). Note that the Z∆MDA strain is derived from a clinical isolate that has not been selected for a specific pilE sequence - as for the NEM8013 2C4.3 strain (clone 12)41 so it corresponds to a mix of pilE sequences. Transduction was compared with transformation efficiency (Fig. 1b) and pili quantity (Fig. 1c). The pili quantity was estimated by immunostaining of the pil mutants introduced into a Z∆MDA pilESB expressing strain34. This strain expresses type IV pili, which can be recognised by the 20D9 antibody42. Transformation and pili quantity are two proxies for the expression of functional type IV pili43. Transduction was impaired in mutants with low pili quantity and low transformation efficiency, as well as in the ∆pilT mutant, confirming that the phage only infects strains expressing fully retractable pili. The contribution of the proteins thought to form the pilus tip (PilH, I, J, K) is difficult to interpret as their respective mutants were very poorly piliated. However, the infection efficiency of the four individual mutants was higher than that of the non-piliated ∆pilE mutant, suggesting that these proteins have a minor role in phage infection. Finally, we studied PilC1 and PilC2, two proteins that are thought to be located at the tip of the type IV pilus based on homology with other type IV pilus models44,45,46. The ∆pilC1/C2 double mutant is barely piliated and does not support efficient phage transduction. Conversely, both ∆pilC1 and ∆pilC2 single mutants were piliated and fully transduced by the phage (Fig. 1a).
Phenotypes of transduction (a), transformation (b) and pili quantity (c) for the ZΔMDA reference strain and its derivative mutants in the pilus machinery genes (green) and pilus fibre genes (orange). Transduction results (a) are expressed as transduction rates (arb. units: arbitrary unit), transformation frequency results (b) as percentages of the reference strain ZΔMDA and pili quantity as raw data (arb. units: arbitrary unit). For transformation and transduction experiments, it should be noted that the background should be due to spontaneous appearance of resistant clones. White-centered dots correspond to CFU below the detection limit of respective experiments (see Methods section). The detection threshold is visualized as a shaded grey area, it corresponds to the frequency of spontaneous resistance to spectinomycin. Exact p-values are provided in the Source Data file. Transduction assays were performed three times (n = 3) in duplicate and statistical analyses were performed using a Brown–Forsythe and Welch ANOVA test (two-sided) with Holm–Sidak’s correction for mutants in the pilus machinery genes and using Ordinary one-way ANOVA test with Dunnett’s correction for mutants in the pilus fiber genes. Transformation frequency assays were performed independently three times (n = 3), and statistical analyses were performed using a One-sample t-test (two-sided) comparing the sample mean with a hypothetical mean of 2. All data were expressed as mean ± SEM. Pili quantifications were performed in triplicate (n = 3) and statistical analyses were performed using a Kruskal–Wallis test with Dunn’s correction. Source data are provided as a Source Data file. d Mutated proteins are indicated on the schematic representation of the type IV pilus machinery. e Transduction rate for the ZΔMDA reference strain and derivatives expressing the PilESA or PilESB variants. Transduction assays were performed three times (n = 3) in duplicate and statistical analyses were performed using an Ordinary one-way ANOVA test with Tukey correction, and data were expressed as mean ± SEM. Source data are provided as a Source Data file.
Finally, we investigated the role of PilE in phage entry and transduction. We took advantage of the antigenic variation of the class I pilE gene and analysed the transduction efficiency of MDA(orf6-aadA1)Φ depending on pilE sequence. We took advantage of previous constructs available in our team in which pilESA and pilESB variant sequences were fused to a kanamycin resistance gene to allow their selection34. We therefore constructed two phage-receptive strains with the two different pilE variant sequences34. The two strains differ in the variable part of PilE and especially in the D-region between the two conserved C-terminal cysteine residues (C120 and C152 for PilESA) (Supplementary Fig. 1a–c). We then evaluated the transduction efficiency in these two strains compared to the parental strain Z∆MDA. The MDA(orf6-aadA1)Φ transduction efficiencies differ among the pilESA, pilESB, and Z∆MDA strains. The pilESA strain exhibits a significantly reduced phage transduction rate compared to Z∆MDA (Fig. 1e, p = 0.0006). The transduction rate of pilESA is also lower than that of pilESB, although this difference is not statistically significant (p = 0.0505). These findings suggest a potential role of the pilE sequence in modulating transduction efficiency. Finally, we investigated the role of PilE post-translational modification in phage transduction (Supplementary Fig. 1d). Mutations in the two genes encoding the glycosyltransferases PglA and PglC or in the gene encoding pilin phosphotransferase B (PptB) have little effect on phage transduction.
PilS loci are a reservoir of pili sequences compatible with MDAΦ infection
Our results above suggested that PilE variants may have different efficiencies for infection by MDAΦ. However, the Fig. 1e raised the question of whether transduced meningococci from the PilESA-expressing strain still express this PilE variant or whether the phage has infected bacteria expressing another PilE variant that is present in the population due to antigenic variation despite the use of the kanamycin resistance gene in frame with pilESA. To answer this question, we first sequenced the pilE gene of 10 independent clones of the Z∆MDA pilESA strains (Fig. 2a, b). Seven clones expressed the PilESA sequence or a closely related sequence (98% identity) and three clones had more than five different mutations compared with the PilESA sequence (Fig. 2b). We then transduced the Z∆MDA pilESA strain with MDA(orf6-aadA1)Φ, selected the transduced population on spectinomycin and sequenced 10 independent clones again (Fig. 2a, c). After transduction, none of the 10 sequences obtained before transduction were recovered (Fig. 2b, c, e). Five of the 10 sequences were identical and were designated PilESD. This sequence carried a D-region completely different from that of PilESA, as well as several other variations between G58 and D113 of PilESA (Fig. 2c, d and Supplementary Fig. 2a, b). Interestingly, the pilESD sequence is closely related to the pilS3 silent locus of the parental strain (Fig. 2d and Supplementary Fig. 2a, b), suggesting that antigenic variation of PilE still occurs despite the addition of the kanamycin resistance gene in frame with pilE. To confirm that antigenic variation in the Z∆MDA pilESA strain helps explain the level of infection by MDA(orf6-aadA1)Φ, we constructed two mutS+ ∆recA mutant Z∆MDA strains expressing either PilESA or PilESD variants, and we infected both strains with MDA(orf6-aadA1)Φ. Deletion of recA is sufficient to abolish the homologous recombination required for antigenic variation47. Meanwhile, the inactivated copy of the mutS gene in the Z∆MDA strain was reverted to WT (mutS+) to limit the spontaneous appearance of mutants29. The strain was subsequently designated Z∆MDA mutS+ΔrecA. Transduction rate was determined as above and compared to the initial Z∆MDA strain (Fig. 2f). It should be noted that phage infection is independent of RecA. We recovered very few colonies with the Z∆MDA mutS+ΔrecA pilESA strain, whereas the Z∆MDA mutS+ΔrecA pilESD strain was transduced as efficiently as the initial Z∆MDA strain. We then confirmed that the efficiency of transformation was in the same range between the pilESA and the pilESD in Z∆MDA mutS+ strains - the recA deletion preventing homologous recombination - (Fig. 2g). While the transformation frequency was decreased in both strains compared to the parental strain, we observed no statistical differences between strains expressing pilESA or pilESD. Overall, these results suggest that: (i) some variants of PilE appear to be more susceptible to infection; (ii) MDAΦ is able to specifically infect a low-abundance clone expressing a specific PilE variant.
a Schematic representation of the methodology used in (b, c, e). The pilE gene of ten clones randomly selected from the population before and after transduction by MDA(orf6-aadA1)Φ and selection on antibiotic was sequenced. b–e The panels depict the amino acid sequence alignments deduced from the sequenced pilE genes of the 10 clones obtained before selection (b) and after (c) selection. The PilESA variant was used as a reference and the regions identical to PilESA were coloured in pink. Gaps were coloured in white and mismatched regions were coloured in black. The percentage of identity relative to the PilESA sequence is indicated to the left of each sequence. d A schematic representation of the pilE gene locus and its eight silent pilS cassettes involved in the antigenic recombination of PilE. The pilS3 cassette, in red, corresponds to the sequence found in the variants framed in red in (c). e A stacked bar plot showing the distribution of the PilE variant relative to the whole population before and after selection. Phenotypes of transduction (f) and transformation (g) for the strain ZΔMDA and its derivatives expressing the PilESA or PilESD variants. Transduction results (f) were expressed as transduction rates (arb. units: arbitrary unit) and transformation results (g) were expressed as percentages of that of ZΔMDA. Experiments were performed three times (n = 3) in duplicate. Statistical analyses were performed using an ordinary ANOVA test with Tukey correction, and data were expressed as mean ± SEM. White centered dots correspond to CFU below the detection limit. The dotted line corresponds to the mean threshold for the appearance of spontaneous nalidixic acid resistant clones. Source data are provided as a Source Data file.
The type IV pili tip is not essential for MDAΦ entry and transduction
The near absence of phage infection in the Z∆MDA mutS+ΔrecA pilESA strain suggests that the pili tip is not the receptor for MDAΦ. To rule out this hypothesis, we decided to further investigate the role of the PilC proteins. The two PilC proteins are bimodular proteins with a similar C-terminal domain that is thought to be anchored at the pilus tip based on the Myxococcus xanthus model45. The N-terminal (N-ter) domain is involved in bacterial adhesion and differs between PilC1 and PilC248,49 (Fig. 3a). Since the ∆pilC1/C2 double mutant strain is apparently barely piliated, we sought to complement a pilC null strain for the expression of recombinant PilC1 mutants of decreasing size (Fig. 3a) and to evaluate transduction and transformation in a strain expressing the PilESD variant (Fig. 3b, c). Deletion of the amino acids from 3 to 555 in the N-terminal part of PilC1 did not affect transduction or transformation frequency, supporting the idea that the N-terminal domain of PilC is not recognised by the MDAΦ and is not required for pili biogenesis. Increasing deletion to 3–843 residues reduced both the transduction and the transformation frequencies of the resulting strain. Interestingly, the truncated protein PilC1Δ3-843 allowed transduction and transformation at a level close to 10% of the control (although not statistically significant for transduction) while truncated PilC1Δ4-977 has lost both and transduction and transformation competency (Fig. 3b, c). This suggests that PilC1Δ3-843 and PilC1Δ4-977 do not complement for piliation and cannot be included in our analysis. This supports the view that PilC does not play a role in MDAΦ entry. In a complementary approach, we investigated a possible interaction between the MDAΦ ORF6 (Supplementary Fig. 3a) and proteins of the pilus tip using the bacterial two-hybrid system. Based on structure and sequence homology with proteins pIII of M13 and CTXΦ pIII of CTXΦ, ORF6 was previously annotated as a phage adsorption protein, and we showed in a prior work that ORF6 is required for MDAΦ transduction17. In the existing model, the adsorption protein should interact with the pili tip to cross the outer membrane. We therefore assessed the interaction of ORF6 with the N-terminal domain of PilC1 or PilC2, with the C-terminal domain of PilC1 and with the globular heads of the pilins PilH, I, J, and K (Supplementary Fig. 3b). No positive result was observed, suggesting that there is no interaction between ORF6 and the N-terminal domain of PilC1, assuming that the proteins are folded properly. Considering that expression of the first 843 amino acids of PilC is not required for phage transduction (see Fig. 3b), our results suggest that entry of the MDAΦ into meningococci relies mostly on a mechanism independent of an interaction between ORF6 and the pilus tip proteins PilC1/2, PilH, PilI, PilJ, and PilK.
a Predicted structures of PilC1 and PilC2 were obtained using Alphafold-3. The deleted parts of PilC1 were shown as a thin line in red. Phenotypes of transduction (b) and transformation (c) for the strain Z5463 expressing the PilESD variant (pilESD) and its derivative double mutant ∆pilC1/C2 complemented or not with the whole pilC1 or pilC2 genes or pilC1 deletion mutants of decreasing size. Transduction results (b) were expressed as transduction rate (arb. units: arbitrary unit) and transformation results (c) were expressed as percentages of that of the strain expressing the PilESD variant. Each experiment was performed three times (n = 3) in duplicate. Statistical analyses were performed using an ordinary ANOVA test with Dunnett correction for transduction rates, a Kruskal–Wallis test with Dunn’s correction for transformation frequencies, and data were expressed as mean ± SEM. White-centered dots correspond to CFU below the detection limit. The dotted line on bar plot c corresponds to the mean threshold for the appearance of spontaneous nalidixic acid resistant clones. Source data are provided as a Source Data file.
The MDAΦ binds to the pilus filament
Considering the role of antigenic variation in MDAΦ transduction, we reasoned that the MDA phage should interact directly with the pilus fibre rather than the tip. The assembly of ORF4 and PilE proteins in filaments makes a two-hybrid assay between them inappropriate. To visualise the possible interaction between a MDAΦ and a pilus, we first performed transmission electron microscopy (TEM) on whole bacteria incubated with phages. To prevent phage internalisation and to facilitate observation of the phage-pili interaction, we used a strain mutated for pili retraction and expressing the PilESD variant. The Z∆MDA∆pilT pilESD strain was thus incubated with purified bacteriophages, washed to remove free bacteriophages and processed for immunogold against the major coat protein ORF4 and TEM (Fig. 4a). Labelled phages were indeed found along the entire length of the pili bundles. Labelling of phage-free bacteria showed no staining (Fig. 4a).
a Representative immunogold labelling of strain ZΔMDA ΔpilT pilESD incubated with MDA(orf6-aadA1) phage (top) and a control without phage (bottom). Phages were labelled with the anti-MDAORF4_N-ter antibody coupled to 10 nm diameter gold particles. We did not observe any labelling of the type IV pili bundle by the antibody. The right images are enlargements of the framed area in the left images. Representative images from the observation of two independent experiments (n = 2). b, c MDA staining on strains expressing different PilE variants was estimated by immunofluorescence as the ratio of phage surface labelling to bacterial surface labelling after phage precipitation (arb. units: arbitrary unit) (b) Statistical analysis was performed using a Brown–Forsythe and Welch ANOVA test (two-sided) with Tamhane T2 correction, and data were expressed as median with 95% CI. Each dot represents one image and images were taken from three different experiments (n = 3), with dots coloured black, grey and white, respectively. Source data are provided as a Source Data file. Representative images used for the quantification are shown in b. Bar = 20 µm. d High-resolution STED microscopy was performed on MDA(orf6-aadA1) phages on a preparation of the strain ZΔMDA ΔpilT pilESE (see methods section). Phages and pili are labelled with anti-MDAORF4_N-ter antibody (anti-phage, green) and anti-PilE 20D9 antibody (anti-pili, red). Bacteria are stained with DAPI. Representative image from several observations of one experiment (n = 1). Bar = 5 µm.
We next assessed MDAΦ binding to pili by quantifying phage precipitation by bacteria expressing either the PilESA or PilESD variant. As above, we used strains mutated for pili retraction. We incubated purified MDA(orf6-aadA1)Φ with the strains Z∆MDA∆pilT pilESA or Z∆MDA∆pilT pilESD and a pili-deficient strain (Z∆MDA ∆pilE) as a negative control (Fig. 4b, c). Bacteria were then washed and immobilised on glass slides before immunostaining with anti-ORF4 antibodies. The aera of ORF4 labelling was analysed using ImageJ and normalised to that of bacteria stained with DAPI (Fig. 4b). As expected, phages did not bind to the non-piliated Z∆MDA ∆pilE strain. The remaining signal for the non-piliated Z∆MDA ∆pilE strain likely corresponds to background and non-specific labelling. However, there is a significant difference in phage precipitation depending on the pilE sequence. MDA phages bound to the PilESA variant at reduced levels, whereas they bound to the strain specifically expressing the PilESD variant (Fig. 4b, c).
To better visualise the phage-pili interaction, we have decided to perform high-resolution STED (Stimulated Emission Depletion) microscopy on precipitated bacteria. However, as the PilESD variant was not recognised by our anti-PilE 20D9 antibody, we had to derive a new strain with phage binding and transduction properties whose pili were labelled by our antibody. To this end, we followed the same strategy as that used to obtain the PilESD strain. We transduced the Z∆MDA strain expressing the PilESB variant (Fig. 1e and Supplementary Fig. 1c) and selected transduced derivatives. We have selected one new pili variant, named PilESE, which was favourable for phage entry and recognised by the 20D9 antibody (Supplementary Fig. 4a, b). We then incubated purified MDA(orf6::aph3’)Φ with the strain Z∆MDA ∆pilT pilESE. The bacteria were washed and immobilised on glass slides before immunostaining with anti-ORF4 and anti-pili 20D9 antibodies. The slides were then processed for high-resolution STED microscopy (Fig. 4d and Supplementary Fig. 4a). Pili staining is shown as dotted lines, which we had previously observed using high-resolution microscopy with the anti-pili 20D9 antibody43,48, and most of the phage staining appears to be associated with pili along their entire length. We also quantified phage retention by ∆pilT strains expressing either the PilESB or PilESE variant and the amount of pili around the strains as in Fig. 4b (Supplementary Fig. 4d, e). The phage retention is the ratio of phage surface labelling to bacterial surface labelling after phage precipitation (see materials and method). As expected, the strain expressing the PilESE variant appeared to retain significantly more phage than the variant expressing PilESB (p = 0.0006). Interestingly, the Z∆MDA ∆pilT pilESB strain produces more pili than the Z∆MDA ∆pilT pilESE strain, but the amount of pili around these two variants is not significantly different (p = 0.5162). Finally, to confirm the absence of a role for ORF6 in the interaction between the phage capsid and pili, we used a phage preparation produced from a strain deleted for orf617 (MDA(orf6::aph3’)Φ). We incubated the purified MDA(orf6::aph3’)Φ preparation with the strain Z∆MDA∆pilT pilESD as above and used the pili-deficient strain (Z∆MDA ∆pilE) as a negative control (Supplementary Fig. 5). We observed no significant difference between the retention of the MDA(orf6-aadA1)Φ and the MDA(orf6::aph3’)Φ on pili. This confirms that ORF6 does not play a role in the primary interaction between the phage and the pili. Taken together, these results suggest that, in contrast to what has been shown for other filamentous phages, MDAΦ preferentially interacts with pili along their fibres rather than at their tips, an association that strongly correlates with the transduction rate.
MDAΦ transduction is associated with positively charged pili
The differences between the PilESA and PilESD variants are essentially differences in amino acids in the D-region, with amino acids having different charges, resulting in a change in the isoelectric point of the pilin (Fig. 5a and Supplementary Fig. 2b). Interestingly, the coat of MDAΦ, consisting of the major coat protein ORF4, has recently been modelled by cryo-EM50. This model showed that the MDAΦ surface was predominantly negatively charged. We therefore reasoned that the interaction between MDAΦ and pili might be facilitated by opposite charges on the negatively charged MDAΦ coat and positively charged pili such as those displaying the PilESD variant. To test this hypothesis, we first selected four clones expressing different PilE variants from the Z∆MDA strain. These variants were selected on the basis of amino acid charge and were named PilEZ1, PilEZ2, PilEZ3, and PilEZ4, with PilEZ1 being the most positively charged and PilEZ4 the most negatively charged (Fig. 5a and Supplementary Fig. 2b). We then modelled in silico the pilus fibre of these strains and the structure of PilESA and PilESD using Alphafold and the cryo-EM reconstruction of the N. meningitidis type IV pilus (5KUA; https://www.rcsb.org/structure/5kua). The Fig. 5a shows the difference in charged amino acids exposed for the six PilE variants studied. PilESD has a majority of positively charged amino acids on its surface, while PilESA has a majority of negatively charged amino acids, with the other four strains falling between these two extremes. We then assessed the frequency of MDA(orf6-aadA1)Φ transduction in these Z∆MDA clones that predominantly express pilEZ1, pilEZ2, pilEZ3, or pilEZ4 (Fig. 5b). We observed a correlation between pili charge and phage transduction: bacteria with positively charged pili were better transduced by phages than those expressing negatively charged pili.
a Reconstruction of type IV pilus fibres for six different PilE variants from the Z∆MDA strain (this strain has a mix of pilE sequences), from the most positively charged surface (on the left) to the most negatively charged surface (on the right). The predicted structures are based on the biological assembly 5KUA and the predicted structure of the PilE subunits obtained using Alphafold-3. Structures were coloured using the coulombic electrostatic potential command of ChimeraX, with the default colour ranging from red for negative potential through white to blue for positive potential. pI (isoelectric point) was determined using the iep tool on Galaxy version 5.0.0.1 with initial parameters. b Transduction rate of Z∆MDA strain and its four derived clones (arb. units: arbitrary unit). The transduction rate of pilESA has been reused from Fig. 1e to facilitate understanding of the figure. Experiments were performed three times in duplicate (n = 3). Statistical analyses were performed using the ordinary ANOVA test with Dunnett’s correction, and data were expressed as mean ± SEM. Source data are provided as a Source Data file. c Amino acid sequence alignment scheme for strains expressing the PilESA and PilESD variants and their derivative mutants in which amino acid 70-71 and/or the D-region (Dreg) were swapped. The PilESA sequence is shown in red and the PilESD sequence is shown in green. d Transduction rate of strains Z∆MDA and its derivatives Z∆MDA mutS+ ∆recA expressing PilESA or PilESD variants and their mutants. White dots correspond to CFU results below the detection threshold. Experiments were performed three times in duplicate (n = 3). Statistical analyses were performed using the Ordinary one-way ANOVA test with Bonferroni correction, and data were expressed as mean ± SEM. Source data are provided as a Source Data file.
We further examined the differences between the amino acid sequences of PilESA and PilESD, focusing on the charged amino acids (Supplementary Fig. 2b). We decided to focus on position 70-71 and the D-region. We therefore constructed a Z∆MDA mutS+ΔrecA pilESA mutant in which we exchanged A70D71 with S70K71 of PilESD and/or the D-region with that of PilESD, and vice versa between PilESD and PilESA (Fig. 5c). The structure of these mutants was then modelled using Alphafold to visualise charged amino acids (Supplementary Fig. 6a, b). We then assessed the MDA(orf6-aadA1)Φ transduction rate in all these mutants (Fig. 5d). The exchange between the D-regions dramatically increased (PilESA to PilESD) or decreased (PilESD to PilESA) the transduction rate, highlighting the role of this region. Modifications at positions 70-71 had little effect on the pilin-exposed charges, and therefore had little effect on the transduction rate. These data support our hypothesis that the interaction between MDAΦ and type IV pili benefits from a positively charged pilus.
Adhesion and transduction both determine the selection of the same pilS cassette
Both MDAΦ and pili are associated with the invasiveness of N. meningitidis. We therefore sought to determine whether adhesion to human cells could select for a PilE variant that favours transduction by MDAΦ or whether strains transduced by MDAΦ could select for a more adhesive PilE variant. We have previously shown that transduction of MDA(orf6-aadA1)Φ in a Z∆MDA pilESA strain, after spectinomycin selection, resulted in a population expressing predominantly the PilESD variant (Fig. 2e), which appeared to correspond to an antigenic variation achieved between the pilE gene and the pilS3 cassette (Fig. 2d and Supplementary Fig. 2a, b). To assess the proportion of pilS3/pilESD variant in an adhesive or transduced population, we designed a qPCR assay with one primer specific to the pilS3 sequence and one primer specific to the constant region of pilE (Supplementary Fig. 2b). A second pair of primers targeting pilT was used as a control. We validated this method on DNA samples from two Z∆MDA ∆recA strains (which avoid antigenic variation) expressing PilESD or PilEZ4 corresponding to an antigenic variation achieved between the pilE gene and the pilS8 cassette (Supplementary Fig. 2a, b). As expected, we obtained a proportion of pilES3 positive bacteria close to 100% for the Z∆MDA ∆recA pilESD strain, and a proportion of pilES3 positive bacteria of 0.01% for the Z∆MDA ∆recA pilEZ4 strain (Supplementary Fig. 7a). We then examined the proportion of bacteria carrying a recombination of the pilE gene with the pilS3 locus in the ZΔMDA pilEZ4 strain. Only 0.1% of the population was pilES3 positive (Supplementary Fig. 7b). This strain was further used to assess the proportion of bacteria expressing the PilES3 variant after adhesion to endothelial or epithelial cells and then the subsequent transduction by MDA(orf6-aadA1)Φ. In a first experiment (Fig. 6a), we collected ZΔMDA pilEZ4 bacteria adherent to FaDu epithelial cells or EA.Hy926 endothelial cells or after incubation in DMEM, and we quantified both the proportion of bacteria expressing PilES3 and the frequency of transduction. The proportion of bacteria expressing PilES3 increased after adhesion to epithelial and endothelial cells (Fig. 6b) from 0.13% of the population to almost 1% and 0.85% respectively. Similarly, adhesion to epithelial and endothelial cells increased the transduction rate by almost fourfold (Fig. 6c, p = 0.0226 and p = 0.0084, respectively). In a second experiment (Fig. 6d), we transduced the ZΔMDA pilEZ4 strain with MDA(orf6-aadA1)Φ and quantified the adhesion of the resulting population. The initial ZΔMDA pilEZ4 strain was incubated in transduction media without phage and kept as a control. The population infected by the phage and selected on spectinomycin became predominantly pilES3 positive (68% of the total population), whereas it remained at 0.18% in the control population (Fig. 6e). After transduction, the spectinomycin-resistant population was 10.6 times more adhesive to epithelial cells and 4.7 times more adhesive to endothelial cells than the initial population (Fig. 6f, g, p < 0.0001 and p < 0.0001 respectively). Therefore, there is a strong link between transduction and adhesion by selection of a specific pilE variant.
a Schematic representation of the methodology used in (b, c). An initial population of the ZΔMDA expressing the PilEZ4 variant was incubated in cell culture media or in the presence of epithelial or endothelial cells. In the former case, bacteria were collected (control). In the latter two cases, only adherent bacteria were collected. b The proportion of bacteria expressing a PilES3 variant was quantified by qPCR in the three populations. c The three resulting populations were transduced with MDA(orf6-aadA1)Φ and the transduction rates were determined (arb. units: arbitrary unit). d Schematic representation of the methodology used in (e–g). An initial population of the ZΔMDA expressing the PilEZ4 variant was incubated in transduction media and collected (control) or transduced with MDA(orf6-aadA1)Φ and selected on spectinomycin (transduced). e Determination of the proportion of bacteria expressing a PilES3 variant in the two populations by quantification by qPCR. Comparison of the adhesion of the two populations on epithelial cells (f) and endothelial cells (g). Data were expressed as a percentage of the control population. b, e qPCR was performed three times independently (n = 3). c, f, g Experiments were performed three times in duplicate independently (n = 3). Dots coloured black, grey and white in b, c and e–g represent the results from each experiment, respectively. Statistical analyses were performed using a Brown–Forsythe and Welch ANOVA test (two-sided) with Dunnett correction for c, an unpaired t-test (two-sided) for f, g, and data were expressed as mean ± SEM. Source data are provided as a Source Data file.
Enrichment of phage-infected bacteria during adhesion
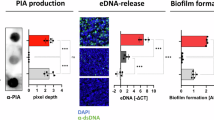
Finally, we asked whether adhesion to human cells would be sufficient to increase the proportion of phage-containing bacteria in a given bacterial population. To answer this question, we brought a Z∆MDA pilEZ4 population into contact with purified MDA(orf6-aadA1)Φ without further antibiotic selection. Bacteria were then incubated on epithelial and endothelial cells or incubated in DMEM as a control. After 3 h, the cells were washed, adhesive bacteria were recovered and the proportion of phage-containing bacteria was assessed by serial dilution on agar plates containing spectinomycin and CFU counting (Fig. 7a). As a control, we confirmed that the growth and adhesion of the two bacterial populations - phage infected or not - were similar (Supplementary Fig. 7c). Bacterial adhesion to epithelial or endothelial cells was sufficient to increase the MDA-carrying population by 2.1 and 2.4 times, respectively, compared to the total population (Fig. 7b, p = 0.0189 and p = 0.0116 respectively). This result highlights the mutual benefit of the MDAΦ-meningococcus interaction in promoting both phage propagation and N. meningitidis colonisation (Fig. 7c).
a Schematic representation of the methodology used in (b). An initial population of the ZΔMDA expressing the PilEZ4 was transduced with MDA(orf6-aadA1)Φ and recovered without selection to have both transduced and non-transduced bacteria in the same population. The population was then incubated in cell culture media and collected or incubated in the presence of epithelial or endothelial cells and only adherent bacteria were collected. The three populations were then plated on agar containing spectinomycin to quantify the percentage of bacteria infected by MDA(orf6-aadA1)Φ. b The proportion of bacteria with MDA(orf6-aadA1) prophage was determined for the three populations. b Experiments were performed three times in duplicate (n = 3). Statistical analyses were performed using a Kruskal–Wallis test with Dunn’s correction for b, and data were expressed as mean ± SEM. Source data are provided as a Source Data file. c Schematic representation of our proposed model of the amplification loop between MDAΦ infection, biofilm formation and enrichment in adhesive PilE variants. The top panel shows the expression of eight possible PilE variants following antigenic variation and coloured according to their PilE charge. Bottom left: a human carrying strains of N. meningitidis with different pili variants and not infected by the MDA phage will spread any PilE expressing bacteria. Bottom right: when an isolate expressing the MDA phage is present in the population, the phage infects bacteria expressing the most adherent PilE variants, which are able to promote themselves in the niche and produce phages. These phage-producing bacteria will form a biofilm at the top of the population11 and will be preferentially disseminated by the host.
Discussion
In this study, we have dissected the molecular interaction between MDAΦ and type IV pili, an interaction that is likely to be intimately involved in shaping the population of PilE variants and thus influencing meningococcal colonisation of the host. We characterised an association between MDAΦ, type IV pili and host colonisation, providing new insights into the role of MDAΦ in the interaction of meningococci with their host and suggesting an unexpected link between the life cycle of this filamentous phage and antigenic variation.
First, we studied phage transduction in type IV pili machinery mutants, and we demonstrated the absolute necessity of retractable pili. Among the piliated mutants of the type IV pili machinery, only the pilT ATPase mutant was not infected by phages. The pilT2 and pilU genes, which encode two other proteins known to be members of the ATPase family of the type II/IV secretion system, were not required for phage infection, which seems consistent with the absence of complementation of a pilT mutant by pilT2 or pilU, as previously shown by Brown et al.30 for transformation with exogenous DNA. Non-piliated mutants, such as those deleted in pilH, I, J, or K or both pilC1 and pilC2, were weakly infected by phages, but more so than a pilE mutant, although it is surprising that pili fibres can be extruded from the PilQ pore without being capped by the tip complex. Another surprising result is the level of transduction and transformation observed in the ∆tfpC mutant. Although TfpC has been described as playing a role in maintaining the extended state of pili, it has not been identified as a factor in pili expression51. Furthermore, Muir and collaborators found that this protein is involved in transformation efficiency52, which was not observed in our study. The ∆tfpC mutant is as transformable and transductible as the wild type, but appears to have slightly fewer pili. It does not appear to be involved in transduction by MDAΦ. This phenotype was observed for insertion and deletion mutant in two different strains. We then investigated in detail the possibility that PilC proteins are involved in MDAΦ binding. The two-hybrid assay failed to demonstrate a possible interaction between the PilC N-terminal domain and ORF6. Besides, our experiments with the PilC deletion mutants and that of the extremely weak phage transduction observed in the PilESA variant strongly suggest that PilE is the only pilin required for phage transduction, and that the pilus tip proteins PilC, H, I, J, K are probably not necessary. Although unlikely, we may not exclude that the phage binds with the very last part of the C-terminal domain of PilC. Similarly, the interaction with PilQ or the involvement of proteins whose absence results in a non-piliated phenotype (such as PilD, PilF, PilP) cannot be excluded. In addition, direct observation by electron microscopy and high-resolution microscopy revealed an interaction between MDAΦ and type IV pili along the pili fibre. To our knowledge, this work is the first demonstration, at the molecular level, of an interaction between the capsid of a filamentous phage and the lateral fibre of type IV pili rather than between the tip of the phage and the tip of the pilus.
We have previously published that the ORF6, located at one end of the phage, should be the adsorption protein required for phage entry across the bacterial membranes17. This conclusion was based on the absence of transduction by MDAΦ mutated for orf6 and on the results obtained for the adsorption proteins of phage Ff and CTX, which recognise pili ends15,53. The Ff and CTX phage adsorption proteins are also involved in the crossing of the inner membrane via an interaction with the TolQRA system. Therefore, although we demonstrated that ORF6 is not involved in the binding to pili, this does not rule out a role for ORF6 in inner membrane crossing. This step needs to be further investigated in the future.
In meningococci, antigenic variation of pilE plays an important role in cell adhesion to the host and that of escape from the immune response34,35. Antigenic variation of pilE with one of the silent pilS loci allows large modifications of the amino acids exposed on the pili surface, leading to changes in charge and hence isoelectric point of pilin in only a subset of the population. Unlike the N. meningitidis laboratory strain 8013, strain Z5463 has not been selected for adhesion to host cells34 and is expected to be able to express a diversity of pilE sequences (due to antigenic variation and the hypermutator phenotype of the strain29). This allows us to observe the preferential infection of phages in N. meningitidis expressing PilE variants associated with high PI, although this variant is expressed by a minority of bacteria. One hypothesis that explains the association of the MDA phages with these PilE variants is electrostatic attraction between the positively charged pilin proteins and the bacteriophages whose capsids are mainly negatively charged, as recently shown by Böhning et al.50, using cryoEM, although we cannot exclude an interaction between the phage capsid and specific amino acids on PilE.
We found a very strong relationship between effective adhesion and effective transduction, with the PilE variant with the highest isoelectric point being the most adhesive and most transducible variant. Therefore, the presence of one MDA phage in a N. meningitidis genome is likely to be associated with the presence of a highly adhesive PilE variant in the population. N. meningitidis genomes contain one to four different MDA prophages, corresponding to multiple infections28 and our works and that of others strongly suggest that infected bacteria are competent for multiple infection by MDA phages10,17.
Finally, the experiment presented in Fig. 7a more closely resembles the natural life cycle of phage and Neisseria, i.e. without direct selection for the presence of the MDAΦ. In this latter experiment, the adhesion to human cells was sufficient to increase the population of MDAΦ-positive bacteria. Considering our results, the infectious cycle of this phage and its role in biofilm formation, we propose a model in which an amplification loop - between MDAΦ infection, biofilm formation and enrichment in adhesive PilE variants - is likely to favour the emergence and maintenance of MDAΦ-positive clones in the most adhesive population, which is also potentially the most virulent (Fig. 7c).
In conclusion, we have described a unique mode of interaction between a filamentous phage and the type IV pili of its host revealing a new type of mutual benefit between a phage and its host.
Methods
Bacterial strains and culture media
The N. meningitidis strains used in this study are listed in Supplementary Data 1 and are derived from Z5463, a naturally transformable serogroup A strain isolated from the throat of a patient with meningitis in the Gambia in 198354. N. meningitidis mutants used in this work are listed in the Supplementary Data 1. It should be noted that the Z∆MDA strain was constructed from the wild-type Z5463 strain. The initial mutant strain was clonal (one colony was selected), but underwent several passages to eliminate the circular cytoplasmic form of MDAΦ. Considering that the frequency of antigenic variation in N. meningitidis is 1 to 2 × 10−3, the resulting strain expresses several different PilE sequences. E. coli strain DH5α was used for DNA cloning and plasmid propagation. E. coli Oxi-Blue55 was used for two-hybrid assay. E. coli Oxi-Blue, derived from the Oxi-BTH strain, facilitates the correct folding of proteins formed by disulphide bonds within its cytoplasm.
The N. meningitidis strains were grown at 37 °C in 5% CO2 on Brain Heart Infusion Agar (BHI Agar, Condalab) plate containing 5% of heat-inactivated horse serum (Gibco), or in gonococcal base (GCB) agar (Difco) plates containing 12 μM FeSO4 and Kellogg’s supplements56 or in BHI liquid medium (Condalab) or in GC-liquid medium [1.5% proteose peptone (Difco); 0.4% K2HPO4, 0.1% KH2PO4, 0.1% NaCl], sometimes with 12 μM FeSO4 and Kellogg’s supplements. E. coli was grown at 30 °C or 37 °C on Lysogeny Broth (LB) agar or in LB liquid medium. For antibiotic selection of E. coli strains, kanamycin (Km) was used at a concentration of 50 µg/ml, erythromycin (Em) at 150 µg/ml, and ticarcillin (Tic) at 100 µg/ml. To select strains derived from N. meningitidis, Km was used at a concentration of 200 μg/ml, spectinomycin (Sp) at 75 μg/ml, Em at 3 μg/ml, chloramphenicol (Cm) at 5 μg/ml, nalidixic acid (Nal) at 1 µg/ml, and tetracyclin (Tet) at 2.5 µg/ml.
Cell lines and culture conditions
The pharynx carcinoma-derived epithelial cell line (FaDu; ATCC #HTB-43) and the human umbilical vein cell line (EA.hy926; ATCC #CRL-2922) were obtained from the American Type Culture Collection (ATCC). The FaDu and EA.hy926 cell lines were grown in Dulbecco’s Modified Eagle Medium (DMEM; Gibco) supplemented with 10% decomplemented foetal calf serum (FCS; Gibco). Cells were grown at 37 °C in a humidified incubator under 5% CO2.
Phage preparation, quantification, and transduction assay
Phage preparation
Phage preparation was performed as previously described using two different strains: ZS2 strain is a derivative of strain Z5463 to which a Sp resistance gene (aadA1) has been added in the MDA prophage (MDA(orf6-aadA1)Φ), just after orf6, without promoter, and Z5463 orf6::aph3’ strain, previously described17, derivative from Z5463 with prophage gene orf6 replaced by a kanamycin resistance cassette without promoter (MDA(orf6::aph3’)Φ). Bacteria were pelleted from 200 ml of an overnight culture in BHI liquid medium containing Sp at 200 µg/ml (for ZS2) or Km at 300 µg/ml (for Z5463 orf6::aph3’). After filtration at 0.45 µm, the supernatant was treated with DNase I (25 µg/ml) for 1 h at 20 °C. Phage was precipitated by the addition of 0.5 M NaCl and 10% polyethylene glycol 6000 and incubated overnight at 4 °C. The phage was then pelleted by centrifugation at 10,000 × g for 30 min and resuspended in PBS 1X (with CaCl2 and MgCl2; Gibco). The phage suspension was centrifuged at 200 × g for 10 min, and the supernatant was collected to remove insoluble particles. It was then filtered through a 0.45 µm filter to prevent contamination. The purified phage preparation was used directly for immunogold labelling, immunofluorescence, or bacterial transduction assays.
qPCR quantification of phage concentration
Phage concentration was quantified by qPCR using two primer pairs: one targeting the phage genome (ORF6-F/R for MDA(orf6-aadA1)Φ or ORF4-F/R for MDA(orf6::aph3’)Φ), and another targeting the Glucose-6-phosphate dehydrogenase (G6PD-F/R) to assess total bacterial chromosome number. qPCR was performed with Luna Universal qPCR Master Mix (NEB) on a QuantStudio 1 (Applied Biosystems), run in triplicates. Cycling conditions were 95 °C for 1 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 30 s. A standard curve was generated using genomic DNA from strain Z5463Δorf1, assumed to contain a single phage copy per genome. The resulting standard equations enabled quantification of circular phage DNA, the relative amount of phage is obtained by NumberORF6/ORF4 – NumberG6PD. After quantification, the average phage quantity is estimated to be 2 × 107 phage/µl.
Transduction assay
Recipient strains were grown overnight on agar plates and resuspended at an OD600 of 1 in transduction liquid medium [containing GC-liquid medium, 12 μM FeSO4, Kellogg’s supplements, 2.5 mM MgCl2, 2.5 mM MgSO4, and 25 µg/ml DNase I]. Two hundred microliters of the bacterial suspension (corresponding to 2 × 108 bacteria) were aliquoted into a 24-well plate and mixed with 30 µl of the MDA(orf6-aadA1) phage preparation. This corresponds approximately to a bacteria:phage ratio of 1:3. The mix was incubated for 15 min at 37 °C with shaking and 15 min without shaking, and 800 µl of transduction liquid medium was added to each well. Plates were incubated for a further 2 h at 37 °C without shaking. If necessary, bacteria were plated on agar plates with or without Sp and stored the next day at −80 °C in BHI/15% glycerol.
To determine the transduction rates, bacteria were diluted and plated on agar plates with or without Sp before counting of CFU the next day. The transduction rate was calculated by dividing the number of CFU obtained on Sp-containing agar plates by that on antibiotic-free agar plates. The efficacy of the phage preparation was verified by transducing 30 μl of the phage preparation into the Z∆MDA strain and was validated if the transduction rate was greater than 10−3. If no bacteria were obtained at the least diluted point on the agar plates, a CFU of 0.9 was arbitrarily assigned and the point was marked as below the detection limit. A detection threshold corresponding to the frequency of spontaneous resistance to spectinomycin was experimentally calculated using the Z∆MDA strain (3.2 × 10−8 ± 4 × 10−8 – SD; n = 3 experiments in duplicate).
Transformation of N. meningitidis
Strains of interest were grown overnight on BHI agar plates and resuspended to OD600 = 1 in liquid transformation medium [containing GC-liquid medium, 12 μM FeSO4, Kellogg’s supplements, MgCl2 at 2.5 mM, and MgSO4 at 2.5 mM]. Five μl of PCR product or 150 ng of plasmid or three µl of genomic DNA were added to 200 µl of this bacterial suspension and incubated for 30 min at 37 °C, 5% CO2 with shaking. Bacteria were then grown for 2 h after the addition of 800 µl of liquid transformation medium. Transformants were selected on BHI agar plates containing corresponding the appropriate selective antibiotic.
To determine the transformation frequency, bacteria were resuspended to OD600 = 0.1 in step 1 transformation media [containing GC-liquid medium, MgSO4 at 5 mM]. One thousand ng of genomic DNA carrying a Nal-resistant cassette was added to 200 µl of this bacterial suspension and incubated for 30 min at 37 °C, 5% CO2 with shaking. Bacteria were then grown without agitation for 2 h after the addition of 800 µl of step 2 transformation liquid medium [containing 12 μM FeSO4 and Kellogg’s supplements except L-Glutamine]. This experiment has been described previously30. After incubation, bacteria were diluted and plated on agar plates with and without Nal before CFU counts the next day. The transformation frequency was calculated by dividing the number of CFU obtained on Nal-containing agar plates by the number of CFU obtained on antibiotic-free agar plates. If no bacteria were obtained at the least diluted point on the agar plates, a CFU of 0.9 was arbitrarily assigned, and the point was marked as below the detection limit.
PCR amplification, plasmid construction and mutant generation
PCR amplification and purification
Sequence amplifications for construction and sequencing were conducted using Q5 High Fidelity DNA polymerase (New England Biolabs) and the oligonucleotides listed in Supplementary Data 2. All oligonucleotides were purchased from Integrated DNA Technologies (eu.idtdna.com). After amplification, PCR products amplified from plasmids were incubated for 1 h at 37 °C with DpnI enzyme (NEB) to digest the methylated plasmid backbones. PCR products were purified using Monarch DNA Gel Extraction kit (New England Biolabs) or Monarch PCR & DNA Cleanup Kit (New England Biolabs). All mutants were verified by PCR amplification using OneTaq DNA Polymerase (New England Biolabs) on bacterial genomic DNA extracted using the Wizard Genomic DNA purification kit (Promega) according to the manufacturer’s instructions.
Plasmid construction and mutant generation
All plasmids used in this work are listed in the Supplementary Data 3.
Z∆MDA pilESA and Z∆MDA pilESB. The pilESA and pilESB alleles fused to the Km resistance gene and the flanking regions (≈250 pb upstream and downstream) were amplified by PCR from the genomic DNA of strains NEM8013pilESA and -pilESB with primers 5′_pilE_Fw and 3′_pilE_Rv34 and used for transformation and allelic exchange in Z∆MDA. Bacteria were then isolated on Km, and pilE sequences were verified by Sanger sequencing. The same methodology was used to construct Z∆MDA pilESD and Z∆MDA pilESE with PCR amplification from the genomic DNA of transduced clonal strains obtained respectively from transduction of Z∆MDA pilESA and Z∆MDA pilESB.
Obtention of mutS+ derivatives. We took advantage of the plasmid containing the Z2491mutS+ allele with a Sp resistance cassette inserted after the gene, leaving a downstream intergenic sequence at the 3′end, allowing the allelic exchange to be performed (as described by Omer et al.29). The Sp resistance gene was swapped with a Cm resistance gene by PCR mutagenesis using appropriate primers (Supplementary Data 2). The ZΔMDA mutS+ strain was obtained by transformation of a vector containing the mutS+ allele with Cm resistance downstream of the gene and allelic exchange. Bacteria were then isolated on Cm and verified by PCR.
Z∆MDA mutS+ and Z∆MDA mutS+ ΔrecA -pilESA and -pilESD and derived mutants. The pilESA and pilESD alleles fused to the Km resistance gene and their flanking regions (≈250 pb upstream and downstream) were cloned into the pMiniT 2.0 vector as above and using primers 5′_pilE_Fw and 3′_pilE_Rv. The resulting plasmids (namely pMiniT_pilESA-KmR and pMiniT_pilESD-KmR) were sequenced and used as templates for site-directed PCR mutagenesis using the appropriate primers (Supplementary Data 2). PCR products corresponding to the pilE loci were obtained from wild-type and mutagenised pMiniT_pilESA-KmR or pMiniT_pilESD-KmR, using primers 5′_pilE_Fw and 3′_pilE_Rv and were introduced into N. meningitidis strain ZΔMDA mutS+ by transformation and allelic exchange. Bacteria were then isolated on Km plates, and pilE sequences were verified by Sanger sequencing. To obtain Z∆MDA mutS+ ΔrecA -pilESA, -pilESD or other derivatives variants of pilE, a genomic DNA from Z∆MDA ΔrecA was introduced into strains Z∆MDA mutS -pilESA, -pilESD or other derivatives variants of pilE by transformation and allelic exchange. Bacteria were isolated on Tet plates, and pilE sequences were verified by Sanger sequencing and recA deletion was verified by PCR.
Z∆MDA pilESB. The pilESB allele was cloned into pMiniT resulting in plasmid pMiniT_pilESB-KmR. The Km resistance cassette fused to pilE gene was swapped with a Sp resistance gene by PCR-based mutagenesis using appropriate primers (Supplementary Data 2). The resulting pilESB variant was introduced into strain Z∆MDA at the locus of pilE by transformation. Cloning and transformation strategies were the same as described for pMiniT_pilESA-KmR.
pNM99Ptet-pilC1 and -pilC2 and derived plasmids. The pilC1 and pilC2 genes from strain Z5463 were cloned into the pMiniT 2.0 vector as described above. To avoid phase variation, the poly(G) sequences present in the two pilC signal peptide sequences were swapped with an amino acids identical alternative sequence using PCR mutagenesis with the primers C1.Z_polyG_Fw/Rv and C2.Z_polyG_Fw/Rv. The resulting plasmids, pMiniT- pilC1fixed(Z5463) and pMiniT-pilC2fixed(Z5463), were sequenced and used as a matrix for cloning of pilC1 and pilC2 into pNM99Ptet57, where gene expression is under the control of a tet promoter (primers pNM99_TET_Fw/Rv, pilC1_Z5463_pNM99_Fw/Rv, pilC2_Z5463_pNM99_Fw/Rv). Deletion mutants of pilC1 were obtained by PCR mutagenesis using the appropriate primers directly on the pNM99Ptet-pilC1 plasmid. Mutations were verified by Sanger sequencing.
Z∆MDA ΔpilC1 ΔpilC2 complemented for PilC1 and PilC2 and derived mutants. Markerless null mutants were constructed as described by Wuckelt et al.57. Briefly, the gene of interest was first swapped by allelic exchange with the APG cassette carrying a Km resistance marker (aphA-3) and two counter-selectable marker genes (pheS* and galK). The markerless null mutant was then obtained by removing the APG cassette by allelic exchange using a PCR product of the 5′ and 3′ regions flanking the gene of interest. Using this strategy, we first obtained a Z∆MDA ΔpilC2 mutant and we inserted the complementation construct from the pNM99Ptet-pilC1 and -pilC2 plasmids (see above) by transformation and allelic exchange. Complemented bacteria were selected on Em plates. We then obtained the ΔpilC1 mutant using the same strategy as for the ΔpilC2 mutant. The PilC expression, required for transformation, was allowed by the addition of anhydrotetracycline (20 ng/ml). The primers used are described in the Supplementary Data 2.
Transposon mutants (ΔpilC1, ΔpilD, ΔpilE, ΔpilF, ΔpilG, ΔpilH, ΔpilI, ΔpilJ, ΔpilK, ΔpilQ, ΔpilT, ΔpilT2, ΔpilU, ΔpilV, ΔpilW, ΔpilX, ΔpilZ, ΔtsaP, ΔpglA, ΔpglC) and other deletion mutants (ΔpilP, ΔpilC2, ΔNMA0415, ΔtfpC, ΔpptB, ΔpilC1/C2, ΔpilT, ΔrecA). We took advantage of mutants originally generated in strain NEM8013 by in vitro transposition58. Regions of interest were amplified by PCR on the genomic DNA and then introduced into Z∆MDA by transformation of purified PCR products and allelic exchange. Bacteria were then isolated on Km plates, and mutants were verified by PCR. Other deletion mutants (except for the markerless mutants ΔpilC1 and ΔpilC2, see above) were obtained by inserting an antibiotic resistance cassette in place of the target gene (from START to STOP codons, except for pilC2 where the last 537 nucleotides were retained) by allelic exchange after transformation into the strain Z∆MDA. The PCR product carrying the cassette flanked by regions upstream and downstream of the target gene was obtained by overlap PCR using primers listed in Supplementary Data 2. All transformants were selected on BHI agar plates containing a selective antibiotic. Mutations in the piliation machinery and pilus fiber genes were also introduced into strain Z∆MDA pilESB by transformation with genomic DNA from the corresponding Z∆MDA mutant, followed by allelic exchange.
Two-hybrid system. The Bacterial Adenylate Cyclase Two-Hybrid System Kit (BACTH System Kit) from Euromedex was used. BACTH plasmids were designed to detect adenylate cyclase-fused protein expression. A 6His-tag was added to pKNT25/pKT25 and a FLAG-tag was added to pUT18/pUT18C. Divergent primers carrying sequences for the tag halves were designed to anneal just after the ATG start codon or just before the stop codon of the free extremity of the adenylate cyclase subunits. The PCR products form the complete tag upon plasmid ligation (same ligation method as for pilE mutagenesis). Then, to add inserts to the BACTH vector backbone, complementary primers were designed to overlap on 15–30 bp, and the two PCR products (insert and vector backbone) were purified and assembled using the NEBuilder HiFi DNA Assembly kit (NEB). The plasmids corresponding to tolA3 in pUT18C, gIII in pKT25, orf6 in pKNT25, pUT18, pKT25, pUT18C, pilC11-475 in pKNT25, pUT18, pilC22-492 in pKNT25, pUT18, pilC1476-1021 in pKT25, pUT18C, pilH30-213 in pKT25, pUT18C, pilI35-189 in pKT25, pUT18C, pilJ40-314 in pKT25, pUT18C, pilK53-197 in pKT25, pUT18C are listed in Supplementary Data 3.
Bacterial adenylate cyclase two-hybrid assay
Competent Oxi-Blue E. coli were co-transformed with 10 ng each of two recombinant plasmids encoding T18 and T25, respectively, Flag and 6His tagged. Transformed bacteria were plates on LB-agar plate supplemented with ampicillin and kanamycin. For each co-transformants, 3 clones were inoculated in duplicate (2×3 clones) into LB wells with Tic, Km, and IPTG 0.5 mM (isopropyl-β-D thiogalactopyranoside). Plates were incubated for 24 h at 37 °C with shaking, then 5 µl of each culture were plated in duplicate on LB agar plates supplemented with Tic, Km, IPTG 0.5 mM and X-gal 40 µg/ml (5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside). Plates were incubated at 30 °C, and the colour of the colonies was assessed after 24 to 48 h.
Adhesion to human cells
The 24-well plates were seeded with EA.Hy926 or FaDu cells and grown to confluence. Bacterial strains grown overnight on BHI agar plates were adjusted to OD600 = 0.05 in DMEM/10% FCS and cultured for 2 h at 37 °C with agitation and 5% CO2. The number of bacteria in the inoculum was estimated and cells were infected with 5 × 106 bacteria (Fig. 6a–g) or 1 × 108 bacteria (Fig. 7a, b). The exact number of CFU was determined by serial dilution and enumeration of CFU. After 1 h (Fig. 6d) or 3 h (Figs. 6a and 7a and Supplementary Fig. 7) of contact, unbound bacteria were removed by six washes with 500 µl of PBS and the cells were detached by mechanical action in 500 µl DMEM/10% FCS. If necessary, bacteria were plated on BHI agar plates and stored the next day at −80 °C in BHI/15% glycerol. To determine the adhesion frequency, adherent bacteria were diluted and spread on agar plates before counting of CFU the next day. The adhesive frequency was obtained by dividing the number of CFU of adherent bacteria by the number of CFU of the inoculum.
Bacterial preparation prior to immunolabeling for pilus quantification
In a 24-well plate, 2 × 10⁷ bacteria were deposited onto coverslips precoated with D-polylysine the day before. The plate was centrifuged at 2500 × g for 1 min and then incubated at 37 °C for 40 min. The medium was removed and the wells were washed once with PBS before adding 500 µl of PBS 1X/4% PFA for 20 min. Finally, the wells were washed with PBS, and the coverslips were immunolabelled.
Phage precipitation
Bacterial strains grown overnight on BHI agar plates were adjusted to OD600 = 0.05 in transduction liquid medium without DNase I and incubated at 37 °C for 2 h. Approximately 2 × 108 bacteria were transferred to a 2 ml tube. One hundred µl of 107 MDA(orf6-aadA1) phage/µl solution or MDA(orf6::aph3’) phage/µl solution was added to the bacteria, and the tubes were filled up to 1.4 ml with transduction liquid medium (without DNase I). The mix was incubated for 30 min at 37 °C, then centrifuged at 400 × g for 4 min, 800 µl of supernatant was removed and centrifuged again for 3 min (to pellet bacteria that have settled to the sides of the tubes). The supernatant was removed and the pellet was resuspended in 800 µl PBS 1X in a 1.5 ml tube. The suspension was centrifuged again at 400 × g for 4 min, 600 µl of supernatant was removed, centrifuged again for 3 min, and finally the pellet was resuspended in 100 µl of PBS 1X/4% PFA (paraformaldehyde; Life Technologies). Twenty µl of bacteria in PBS/4% PFA were spread on a slide and dried for 2 h before immunolabelling.
Phage/pili labelling and quantification
The slides or coverslips were washed with PBS/0.1 M NH4Cl for 2 min and incubated with PBS/4% BSA for 5 min. MDA(orf6-aadA1)Φ and MDA(orf6::aph3’)Φ were stained with the rabbit anti-ORF4 antibody used at 1/100 dilution. For tests with strains expressing PilESB and PilESE, pili were stained with mouse anti-PilE antibody 20D9, used at 1/1000 dilution. Secondary antibodies were goat anti-rabbit antibody conjugated with Alexa Fluor 488 (Molecular Probes) used at 1/400 dilution, and goat anti-mouse antibody conjugated with Alexa Fluor 546. Bacteria were detected by DNA staining using 100 ng/ml DAPI (49,6-Diamidine-29-phenylindole dihydrochloride). Coverslips were mounted in Mowiol. Images were captured using a Zeiss Apotome fluorescence microscope and images were collected and processed using the ZEN (ZEISS Efficient Navigation) software. MDA binding to bacteria and pili quantity were estimated by immunofluorescence as the ratio of phage surface or pili surface labelling to bacterial surface labelling estimated by staining bacterial DNA with DAPI (arb. units: arbitrary unit).
Stimulated-emission-depletion (STED) microscopy
Bacteria with or without bound phages in PBS/4% PFA were spread on 22-mm high-precision cover glasses with thickness of 170 μm, (No. 1.5H, Marienfield) and dried for 2 h. The slides were then washed with PBS/0.1 M NH4Cl for 2 min and blocked with 4% BSA–PBS for 10 min, labelled with monoclonal anti-PilE antibody 20D9 (1:1000 in 4% BSA–PBS) and monoclonal anti-ORF4 antibody (1:100 in 4% BSA–PBS) and then incubated with goat anti-mouse Alexa Fluor 514 secondary antibody fragments (1:1000) and goat anti-rabbit ATTO550 secondary antibody fragments (1:500). Bacteria were detected by DNA staining with 100 ng/ml DAPI. Slides were mounted with ProLong Gold antifade reagent (Invitrogen), refractive index: 1.47.
Images were captured on a Leica TCS SP8 STED microscope, controlled by LASX software, equipped with a 660 nm depletion laser and a 100x NA1.4 objective. Images were acquired using a gated hybrid detector.
The images were then deconvolved using the Huygens software with the ‘Deconvolution Express, standard profile’ parameter.
Electron microscopy: negative staining and immunogold-labelling
Bacteria in PBS/4% PFA with and without co-precipitated MDAΦ were adsorbed onto Carbon/Formvar-coated 200-square-mesh copper grids (Euromedex, Souffelweyersheim, France) for 15 min. The grids were then washed twice for 1 min with PBS and placed sequentially on drops of the following reagents at room temperature: PBS/0.1 M glycine (10 min), PBS/1% BSA (bovine serum albumin)/5% normal goat serum (15 min), and then on a drop of anti-ORF4 N-terminal antibody diluted 1/50 in PBS/1% BSA (60 min). After three washes in PBS, the grids were placed on a drop of Fab2’ goat anti-rabbit conjugated to 10 nm diameter gold (Aurion, Wageningen, The Netherlands) diluted 1/40 in PBS/1% BSA for 60 min. The grids were then washed three times in PBS (1 min), fixed in PBS/1% glutaraldehyde (5 min), washed twice in PBS and twice in distilled water. The grids were then treated with 1% uranyl acetate, air-dried and viewed. Images were acquired using a Jeol 1400 TEM (Jeol, Croissy-sur-Seine, France) operating at 120 keV and equipped with a RIO CMOS camera (Ametek SAS, Elancourt, France).
Quantification of the proportion of pilS3 D-region by qPCR
We estimated the proportion of bacteria with the pilS3 D-region sequence at the pilE locus by qPCR. One pair of primers was designed to anneal to the pilE locus. One primer was designed in the pilS3 D-region and one primer on the conserved upstream part of the pilE gene (pilS3-F/R). The second pair of primers (pilT-F/R) was designed to anneal to the pilT locus, allowing the quantification of the total number of chromosomes. qPCR was performed on genomic DNA using Luna Universal qPCR Master Mix (NEB). All samples were performed in three independent experiments. The cycling conditions were as follows: 95 °C for 1 min, and 40 cycles of 95 °C for 15 s and 60 °C for 40 s on QuantStudio1 (Applied Biosystems). For the standard curve, genomic DNA from Z∆MDA mutS+ ΔrecA pilESD (considered to contain 100% of bacteria expressing pilE D-region S3) was used. Serial dilutions of this DNA were amplified and a standard curve equation was recorded for each primer pair and used to quantify the samples. For each quantification, DNA from pilESD clonal and pilEZ4 clonal strains, consisting of 100% and 0% of bacteria expressing pilE D-region S3, respectively, were used as positive and negative controls.
Bioinformatics
Pilus models were generated by running the amino acid sequences of PilE, PilC1, PilC2 in Alphafold 3 (https://golgi.sandbox.google.com) with general parameters. Several generated PilE models were aligned to subunits of the biological assembly 5KUA using ChimeraX version: 1.7.1. The structures were coloured using the coulombic electrostatic potential command of ChimeraX with the default colouring range. Folding structures of PilC1 and PilC2 were also obtained by running the amino acid sequences in Alphafold 3 with general parameters.
Sequence alignments were performed using Clustal-omega (1.2.4) (https://www.ebi.ac.uk/jdispatcher/msa/clustalo). Isoelectric points of PilE sequences were determined from their PilE amino acid sequences using the tool iep45/5.0.0.1 in Galaxy France.
Statistical analysis
Statistical analyses were performed using GraphPad Prism 8.02. Taking into account the number of replicates, the normality of the distribution for the whole data sets was assessed using QQ plots. When necessary, data were log transformed. Homogeneity of variance was tested using the Brown–Forsythe F-test. If the distribution of the data conformed to the law of normality and the variance of the groups was homogeneous, multiple comparison analyses were assessed using a one-way ANOVA with multiple comparisons test and appropriate correction, and data were expressed as mean ± SEM or median ±95% confidence interval (relevant p-values were reported in the figures). In the case of different variances, a Brown–Forsythe and Welch ANOVA test was performed. A two-way ANOVA with Sidak correction was performed in the Supplementary Fig. 7. The H0 hypothesis was rejected at a significance level of p ≤ 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw data and statistical analysis are listed in the Source Data file. Source data are provided with this paper.
References
Dion, M. B., Oechslin, F. & Moineau, S. Phage diversity, genomics and phylogeny. Nat. Rev. Microbiol. 18, 125–138 (2020).
Hay, I. D. & Lithgow, T. Filamentous phages: masters of a microbial sharing economy. EMBO Rep. 20, e47427 (2019).
Ray, D. S., Bscheider, H. P. & Hofschneider, P. H. Replication of the single-stranded DNA of the male-specific bacteriophage M13. Isolation of intracellular forms of phage-specific DNA. J. Mol. Biol. 21, 473–483 (1966).
Hofschneider, P. H. Untersuchungen über “kleine“ E. coli K 12 Bakteriophagen: 1. und 2. Mitteilung. Z. f.ür. Naturforsch. B 18, 203–210 (1963).
Loeb, T. Isolation of a bacteriophage specific for the F+ and Hfr mating types of Escherichia coli K-12. Science 131, 932–933 (1960).
Marvin, D. A. & Hoffmann-Berling, H. A fibrous DNA phage (fd) and a spherical RNA phage (fr) specific for male strains of E. coli: part II. Physical characteristics. Z. f.ür. Naturforsch. B 18, 884–893 (1963).
Schwartzkopf, C. M. et al. Tripartite Interactions between Filamentous Pf4 Bacteriophage, Pseudomonas aeruginosa, and Bacterivorous Nematodes. https://doi.org/10.1101/2022.10.11.511824 (2022).
Burckhardt, J. C. & Tropini, C. Inoviruses. Curr. Biol. 33, R1272–R1274 (2023).
Secor, P. R. et al. Pf bacteriophage and their impact on Pseudomonas Virulence, mammalian immunity, and chronic infections. Front. Immunol. 11, 244 (2020).
Kawai, M., Uchiyama, I. & Kobayashi, I. Genome comparison in silico in Neisseria suggests integration of filamentous bacteriophages by their own transposase. DNA Res. 12, 389–401 (2005).
Bille, E. et al. A virulence-associated filamentous bacteriophage of Neisseria meningitidis increases host-cell colonisation. PLoS Pathog. 13, e1006495 (2017).
Bille, E. et al. Association of a bacteriophage with meningococcal disease in young adults. PLoS ONE 3, e3885 (2008).
Bille, E. et al. A chromosomally integrated bacteriophage in invasive meningococci. J. Exp. Med. 201, 1905–1913 (2005).
Brynildsrud, O. B. et al. Acquisition of virulence genes by a carrier strain gave rise to the ongoing epidemics of meningococcal disease in West Africa. Proc. Natl. Acad. Sci. USA 115, 5510–5515 (2018).
Jacobson, A. Role of F Pili in the Penetration of Bacteriophage fl. J. Virol. 10, 835–843 (1972).
Gutierrez-Rodarte, M., Kolappan, S., Burrell, B. A. & Craig, L. The Vibrio cholerae minor pilin TcpB mediates uptake of the cholera toxin phage CTX. J. Biol. Chem. 294, 15698–15710 (2019).
Meyer, J. et al. Characterization of MDAΦ, a temperate filamentous bacteriophage of Neisseria meningitidis. Microbiology 162, 268–282 (2016).
Pellegri, C., Moreau, A., Duché, D. & Houot, L. Direct interaction between fd phage pilot protein pIII and the TolQ-TolR proton-dependent motor provides new insights into the import of filamentous phages. J. Biol. Chem. 299, 105048 (2023).
Samire, P. et al. Decoupling filamentous phage uptake and energy of the TolQRA motor in Escherichia coli. J. Bacteriol. 202, e00428-19 (2020).
Stephens, D. S., Greenwood, B. & Brandtzaeg, P. Epidemic meningitis, meningococcaemia, and Neisseria meningitidis. Lancet 369, 2196–2210 (2007).
Perkins-Balding, D., Ratliff-Griffin, M. & Stojiljkovic, I. Iron Transport Systems in Neisseria meningitidis. Microbiol. Mol. Biol. Rev. 68, 154–171 (2004).
Rosenstein, N. E., Perkins, B. A., Stephens, D. S., Popovic, T. & Hughes, J. M. Meningococcal disease. N. Engl. J. Med. 344, 1378–1388 (2001).
Bernard, S. C. et al. Pathogenic Neisseria meningitidis utilizes CD147 for vascular colonization. Nat. Med. 20, 725–731 (2014).
Coureuil, M., Bourdoulous, S., Marullo, S. & Nassif, X. Invasive meningococcal disease: a disease of the endothelial cells. Trends Mol. Med. 20, 571–578 (2014).
Coureuil, M. et al. Meningococcus Hijacks a β2-adrenoceptor/β-Arrestin pathway to cross brain microvasculature endothelium. Cell 143, 1149–1160 (2010).
Join-Lambert, O. et al. Meningococcal interaction to microvasculature triggers the tissular lesions of Purpura Fulminans. J. Infect. Dis. 208, 1590–1597 (2013).
Hill, D. J., Griffiths, N. J., Borodina, E. & Virji, M. Cellular and molecular biology of Neisseria meningitidis colonization and invasive disease. Clin. Sci. 118, 547–564 (2010).
Al Suwayyid, B. A. et al. Meningococcal disease-associated prophage-like elements are present in Neisseria gonorrhoeae and some commensal Neisseria species. Genome Biol. Evol. 12, 3938–3950 (2020).
Omer, H. et al. Genotypic and phenotypic modifications of Neisseria meningitidis after an accidental human passage. PLoS ONE 6, e17145 (2011).
Brown, D. R., Helaine, S., Carbonnelle, E. & Pelicic, V. Systematic functional analysis reveals that a set of seven genes is involved in fine-tuning of the multiple functions mediated by type IV pili in Neisseria meningitidis. Infect. Immun. 78, 3053–3063 (2010).
Chang, Y.-W. et al. Architecture of the type IVa pilus machine. Science 351, aad2001 (2016).
Wang, F. et al. Cryoelectron microscopy reconstructions of the Pseudomonas aeruginosa and Neisseria gonorrhoeae type IV Pili at sub-nanometer resolution. Structure 25, 1423–1435.e4 (2017).
Cehovin, A. et al. Sequence conservation of pilus subunits in Neisseria meningitidis. Vaccine 28, 4817–4826 (2010).
Nassif, X. et al. Antigenic variation of pilin regulates adhesion of Neisseria meningitidis to human epithelial cells. Mol. Microbiol. 8, 719–725 (1993).
Tinsley, C. R. & Heckels, J. E. Variation in the expression of pili and outer membrane protein by Neisseria meningitidis during the course of meningococcal infection. J. Gen. Microbiol. 132, 2483–2490 (1986).
Wörmann, M. E. et al. Sequence, distribution and chromosomal context of class I and class II pilin genes of Neisseria meningitidis identified in whole genome sequences. BMC Genomics 15, 253 (2014).
Jen, F. E. C. et al. Dual Pili post-translational modifications synergize to mediate meningococcal adherence to platelet activating factor receptor on human airway cells. PLoS Pathog. 9, e1003377 (2013).
Berry, J.-L., Cehovin, A., McDowell, M. A., Lea, S. M. & Pelicic, V. Functional analysis of the interdependence between DNA uptake sequence and its cognate ComP receptor during natural transformation in Neisseria species. PLoS Genet. 9, e1004014 (2013).
Merz, A. J., So, M. & Sheetz, M. P. Pilus retraction powers bacterial twitching motility. Nature 407, 98–102 (2000).
Wolfgang, M. et al. PilT mutations lead to simultaneous defects in competence for natural transformation and twitching motility in piliated Neisseria gonorrhoeae. Mol. Microbiol. 29, 321–330 (1998).
Nassif, X. et al. Roles of pilin and PilC in adhesion of Neisseria meningitidis to human epithelial and endothelial cells. Proc. Natl. Acad. Sci. USA 91, 3769–3773 (1994).
Imhaus, A. & Duménil, G. The number of Neisseria meningitidis type IV pili determines host cell interaction. EMBO J. 33, 1767–1783 (2014).
Barnier, J.-P. et al. The minor pilin PilV provides a conserved adhesion site throughout the antigenically variable meningococcal type IV pilus. Proc. Natl. Acad. Sci. USA 118, e2109364118 (2021).
Rudel, T., Scheuerpflug, I. & Meyer, T. F. Neisseria PilC protein identified as type-4 pilus tip-located adhesin. Nature 373, 357–359 (1995).
Treuner-Lange, A. et al. Tight-packing of large pilin subunits provides distinct structural and mechanical properties for the Myxococcus xanthustype IVa pilus. Proc. Natl. Acad. Sci. USA 121, e2321989121 (2024).
Treuner-Lange, A. et al. PilY1 and minor pilins form a complex priming the type IVa pilus in Myxococcus xanthus. Nat. Commun. 11, 5054 (2020).
Koomey, M., Gotschlich, E. C., Robbins, K., Bergström, S. & Swanson, J. Effects of recA mutations on pilus antigenic variation and phase transitions in Neisseria gonorrhoeae. Genetics 117, 391–398 (1987).
Barnier, J.-P. et al. Type IV pilus retraction enables sustained bacteremia and plays a key role in the outcome of meningococcal sepsis in a humanized mouse model. PLoS Pathog. 17, e1009299 (2021).
Morand, P. C., Drab, M., Rajalingam, K., Nassif, X. & Meyer, T. F. Neisseria meningitidis differentially controls host cell motility through PilC1 and PilC2 components of type IV Pili. PLoS ONE 4, e6834 (2009).
Böhning, J. et al. Structure of the virulence-associated Neisseria meningitidis filamentous bacteriophage MDAΦ. Proc. Natl. Acad. Sci. USA 122, e2420157122 (2025).
Hu, L. I. et al. Discovery of a new Neisseria gonorrhoeae type IV pilus assembly factor TfpC. mBio 11, e02528–20 (2020).
Muir, A. et al. Construction of a complete set of Neisseria meningitidismutants and its use for the phenotypic profiling of this human pathogen. Nat. Commun. 11, 5541 (2020).
Ford, C. G. et al. Crystal structures of a CTXphi pIII domain unbound and in complex with a Vibrio cholerae TolA domain reveal novel interaction interfaces. J. Biol. Chem. 287, 36258–36272 (2012).
Achtman, M. et al. Purification and characterization of eight class 5 outer membrane protein variants from a clone of Neisseria meningitidisserogroup A. J. Exp. Med. 168, 507–525 (1988).
Pellegri, C., Bouveret, E. & Houot, L. Protein–protein interactions: oxidative bacterial two hybrid. in Bacterial Secretion Systems (eds Journet, L. & Cascales, E.) Vol. 2715 225–233 (Springer US, New York, NY, 2024).
Kellogg, D. S., Cohen, I. R., Norins, L. C., Schroeter, A. L. & Reising, G. Neisseria gonorrhoeae II. Colonial variation and pathogenicity during 35 months in vitro. J. Bacteriol. 96, 596–605 (1968).
Wuckelt, M. et al. Expanding the genetic toolbox for Neisseria meningitidiswith efficient tools for unmarked gene editing, complementation, and labeling. Appl. Environ. Microbiol. 90, e0088024 (2024).
Rusniok, C. et al. NeMeSys: a biological resource for narrowing the gap between sequence and function in the human pathogen Neisseria meningitidis. Genome Biol. 10, R110 (2009).
Acknowledgements
This work was supported by the research grants ANR-23-CE15-0004-01 from ANR and ANR-18-IDEX-0001-EMERGENCE from Université Paris-Cité (to EB); INSERM (to MC); Université Paris-Cité (PhD scholarship to CM and MW) and Année de recherche from AP-HP and ARS (île de France) to AB. We warmly thank Dr Laetitia Houot and Dr Callypso Pellegri for tools and advice for the two-hybrid assay. We thank Pr Isabelle Martin-Verstraete for fruitful discussions on the two-hybrid assay. We thank Pr Nicolas Biais for his comments and help all along this work.
Author information
Authors and Affiliations
Contributions
C.M., A.B., H.L., M.C., E.B. designed research; C.M., A.B., M.W., M.M., J.M., C.I. performed research; M.M., J.M., B.D., C.I. contributed reagents/analytic tools; C.M., A.B., M.W., M.M., H.L., A.J., C.I., M.C., E.B. analyzed data; M.C. and E.B. validated data; M.C. and E.B. contributed to funding acquisition; M.C. and E.B. coordinated the research; C.M., M.C., E.B. wrote the paper; C.M., H.L., A.J., X.N., M.C., E.B. reviewed and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Daniel Stein, Hank Seifert and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mouville, C., Brizard, A., Wuckelt, M. et al. Neisseria meningitidis filamentous phage MDA promotes colonisation by selecting hyperadhesive pili variants. Nat Commun 17, 744 (2026). https://doi.org/10.1038/s41467-025-67441-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67441-w