Abstract
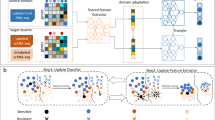
Drug resistance remains a major challenge in cancer treatment. While single-cell profiling offers unprecedented resolution for uncovercovering resistance mechanisms, the potential of emerging foundation models for drug response prediction at the single-cell level is still largely unknown. Here, we introduce scDrugMap, a unified framework featuring both Python toolkits and an interactive web server for benchmarking and predicting drug responses with state-of-the-art foundation models. scDrugMap evaluates eight single-cell foundation models and two large language models across 495,000 cells from 60 datasets, spanning diverse tissues, drugs, cancer types, and treatment conditions. In pooled-data evaluation, scFoundation delivered the strongest performance, particularly in tumor tissue. In cross-data analysis, UCE performed best after fine-tuning, while in zero-shot settings, scGPT achieved the highest accuracy. Together, scDrugMap provides the first systematic benchmark of foundation models for single-cell drug response prediction and offers a powerful, user-friendly platform to accelerate drug discovery and translational precision oncology.
Similar content being viewed by others
Introduction
Drug resistance continues to be a significant challenge in disease treatment, particularly in cancer therapy1. An analysis of 85 cancer drugs approved by the U.S. Food and Drug Administration (FDA) revealed a median response rate (RR) of 41%, with nearly half exhibiting an RR below 40%, and two-thirds showing a complete RR of less than 10%2. Even in biomarker-based personalized cancer treatments, response rates remain modest—30.6% compared to just 4.9% for non-biomarker-based therapies3. Moreover, drug resistance is closely associated with patient survival rates. For example, resistance to KRAS inhibitors, a standard treatment for non-small cell lung cancer (NSCLC), is associated with a median survival of only 6.3 months4. Similarly, in glioblastoma, resistance to chemotherapy and radiotherapy limits median survival post-diagnosis to just 12–15 months, with less than 10% of patients surviving longer than five years5. These findings underline the pressing need for further investigation and advanced approaches to better elucidate drug resistance mechanisms and improve treatment efficacy.
Over the past decade, single-cell RNA sequencing (scRNA-seq) technology has revolutionized our understanding of cellular and molecular heterogeneity, enabling high-resolution analysis of drug responses across diverse cell types and subtypes6,7,8,9,10,11,12,13. For instance, scRNA-seq analysis of NSCLC tumor tissue identified elevated expression of estrogen metabolism enzymes and serum estradiol in cancer cells exhibiting poor pathological responses to PD-1 blockade immunotherapy. Additionally, aged CCL3+ neutrophils interacting with tumor-associated macrophages were identified as potential biomarkers of poor therapy response13. In basal cell carcinoma (BCC), scRNA-seq revealed that novel T cell recruitment, rather than the reinvigoration of pre-existing tumor-infiltrating lymphocytes, was pivotal for effective PD-1 blockade therapy11. Similarly, in inflammatory breast cancer (IBC), single-cell profiling uncovered a shift in cell state from luminal to basal/mesenchymal, implicating the JAK2/STAT3 signaling axis as a key mediator of chemotherapy resistance9. These findings underscore the power of scRNA-seq in uncovering distinct cell states and gene expression profiles driving drug response heterogeneity, enabling the discovery of novel biomarkers and informing personalized therapeutic strategies.
While scRNA-seq offers invaluable insights at single-cell resolution, its high dimensionality, noise, variability, and data sparsity pose significant challenges for interpretation and the extraction of meaningful biological signals. Additionally, differences in sample preparation, library construction, and sequencing platforms across laboratories introduce substantial batch effects and technical variability. To address these issues, recent advancements have introduced large-scale foundation models tailored for single-cell data analysis14,15,16. Prominent examples include scFoundation15, scBERT14, scGPT16, and Geneformer17. These models were pre-trained on large-scale scRNA-seq datasets without being specifically designed for downstream tasks18. Through transfer learning and fine-tuning, single-cell foundation models have demonstrated strong performance across diverse applications, including cell type annotation and batch effect correction18. Notably, certain models offer specialized capabilities—scFoundation for drug response prediction15, scGPT for multi-omics integration (e.g., gene expression and chromatin accessibility)16, and Geneformer for predicting gene dosage sensitivity and chromatin dynamics17. On the other hand, general domain large language models have also been adapted to analyze sequencing data19,20. Despite these advancements, no comprehensive benchmarking studies have rigorously evaluated these foundation models for drug response prediction across diverse single-cell datasets. This gap motivates our study, which aims to develop a unified platform and systematically evaluate existing large models for drug response prediction across a wide range of cancer types, tissues, and therapeutic regimens.
To achieve this, we developed scDrugMap, a unified framework for drug response prediction, and conducted the first comprehensive benchmarking of ten foundation models, comprising eight domain-specific models for single-cell data and two general-purpose natural language models. The curated dataset of scDrugMap includes a primary collection of 326,751 cells derived from 36 single-cell datasets across 23 studies, along with an external validation dataset, which includes 168,486 cells from 24 single-cell datasets across six drug-related studies. By covering a total of 495,237 single cells, we performed two evaluation strategies: pooled-data evaluation and cross-data evaluation. Our results indicate that different foundation models exhibit varying capabilities in drug response prediction, with pooled-data evaluation demonstrating better performance than cross-data evaluation. To facilitate broader application, scDrugMap is developed with a command-line tool and an interactive web server (https://scdrugmap.com/) for seamless drug response prediction using a variety of foundation models. Collectively, our study makes several key contributions: (1) offering model selection guidance based on dataset-specific characteristics and evaluation scenarios; (2) providing an integrated, practical, and interactive tool for drug response prediction, biomarker discovery, and resistance mechanism analysis; (3) establishing a benchmark framework to inform and accelerate future research in drug discovery.
Results
Overview of scDrugMap
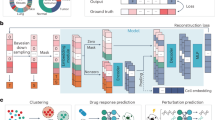
As an integrated framework for drug response prediction using single-cell data (Fig. 1a), scDrugMap consists of three core components: a comprehensive computational pipeline, an interactive web server, and a large-scale curated drug-related dataset—all designed to enable streamlined user access and efficient model evaluation. At the core of scDrugMap are foundation models (FMs), including eight single-cell-specific foundation models (tGPT21, scBERT14, Geneformer17, CellLM22, scFoundation15, scGPT16, CellPLM23, and UCE24) and two general natural language models (LLaMa3-8B25 and GPT4o-mini26). To adapt these models for drug response prediction, scDrugMap supports both zero-shot inference and fine-tuning using Low-Rank Adaptation (LoRA). Users can interact with scDrugMap through either a command-line interface or an intuitive web-based platform, allowing flexible model experimentation and analysis. For benchmarking and validation, we manually curated two distinct single-cell datasets: a primary collection and a validation collection (Data Collection and Preprocessing in Methods). By integrating data curation, model adaptation, and predictive analysis, scDrugMap serves as a comprehensive and accessible resource for advancing single-cell-based drug response research.
a Schematic of the scDrugMap framework, which integrates a benchmarking platform, computational pipeline, interactive web server, and curated drug response datasets. Users can select from a range of foundation models (FMs), including single-cell-specific foundation models (scFMs) and general-purpose language models, and apply different training strategies (layer-freezing or fine-tuning) to predict drug response outcomes (sensitive or resistant). b Categories used for benchmarking model performance, including tissue types, drug types, and cancer types. c Summary of the primary dataset collection, showing the number of datasets (left) and number of cells (right) across tissue types, drug types, and cancer types. Dark shades represent the number of sensitive cells and light shades represent the number of resistant cells. d Summary of the validation dataset collection, with the number of datasets (left) and number of cells (right) across tissue types, drug types, and cancer types. Dark shades represent the number of sensitive cells and light shades represent the number of resistant cells. Source data are provided as a Source Data file. FT fine-tuning, PBMC Peripheral blood mononuclear cells, MM Multiple myeloma, ALL Acute lymphoblastic leukemia, CLL Chronic lymphocytic leukemia, AML Acute myeloid leukemia, NSCLC Non-small cell lung cancer, HCC Hepatocellular cancer. Created in BioRender. Song, Q. (2025) https://BioRender.com/vbfzsbb.
As shown in Fig. 1b, the curated datasets in scDrugMap span 14 cancer types, 3 therapy types, 5 tissue types, and 21 treatment regimens. The primary data collection includes 36 scRNA-seq datasets manually curated from 23 published studies (Fig. 1c), covering 11 major cancer types such as lung cancer, multiple myeloma, and melanoma, along with three therapy categories: targeted therapy, chemotherapy, and immunotherapy. The dataset encompasses four major tissue types: cell line, bone marrow aspirates, tumor tissue, and peripheral blood mononuclear cells (PBMCs) (see Supplementary Data 1). In addition, six distinct scRNA-seq technologies are included in our primary collection: 10x Genomics (n = 27), BD Rhapsody (n = 3), MARS-seq v2.0 (n = 2), DropSeq (n = 2), inDrop v3 (n = 1), and SMART-seq v4 (n = 1). Following quality control, the primary dataset comprised 326,751 single tumor cells annotated with drug response information. Among these, cell lines, targeted therapy, and lung cancer represented the largest cell counts within their respective categories. Importantly, most subgroups (e.g., those for targeted therapy, bone marrow aspirates, and multiple myeloma) exhibited balanced distributions between drug-sensitive and drug-resistant cells (Supplementary Data 2).
For the validation data collection, we manually collected 17 scRNA-seq datasets from six other drug-related studies on solid tumors available in the GEO database7,8,10,11,12,27 (Fig. 1d). This validation dataset includes five cancer types: ovarian cancer, NSCLC, pancreatic cancer, colon cancer, and basal cell cancer, along with three therapy types: targeted therapy, chemotherapy, and immunotherapy. The represented tissue types include cell lines, tumor tissue, and organoids (Supplementary Data 1). After quality control, the validation set comprised 18,856 single-cell transcriptomes. Similar to the primary dataset, the largest proportions were again observed in cell line (tissue), targeted therapy (treatment), and NSCLC (cancer type) categories. Most validation subgroups also maintained a balanced representation of drug-sensitive and drug-resistant cells (Supplementary Data 2). Using these curated data, we implemented two distinct evaluations within scDrugMap: (1) pooled-data evaluation, where models were trained and tested on aggregated data from multiple studies; (2) cross-data evaluation, where models were tested independently on datasets from individual studies.
Pooled-data evaluation in primary data collection
First, we evaluated different FMs in scDrugMap using frozen layer training for predicting drug responses in the primary data collection. The results across various tissue types, cancer types, drug classes, and regimen are shown in Fig. 2a and Supplementary Fig. 1a. Specifically, for the cell line data, which contained the highest number of cells, scFoundation demonstrated the best performance (mean F1 score: 0.971) while scBERT performed the worst (mean F1 score: 0.630). In prostate and pancreatic cancer, LLaMa3 exhibited comparable performance with scFoundation (mean F1 scores: scFoundation-0.990, LLaMa3-0.913 for prostate; scFoundation–1.000, LLaMa3–0.963 for pancreatic cancer). Notably, this high performance may be due to the relatively small sample sizes (10,982 cells in prostate cancer and 6136 in pancreatic cancer) and pronounced imbalance (prostate cancer: 20.85% sensitive cells; pancreatic cancer: 11.38% sensitive cells) (Supplementary Data 2). Additionally, LLaMa3 was the best method for predicting drug response in the CAR-T regimen (mean F1 score: 0.875), and scGPT was the best for the carboplatin regimen (mean F1 score: 0.882). Other metrics-including AUROC, accuracy, precision, recall (Supplementary Fig. 1a), as well as F1 score comparison by Student’s t-test (Supplementary Note 1) further demonstrate that scFoundation achieves superior drug response predictions in most of the datasets among the primary data collection. In contrast, some models demonstrated poor performance in specific contexts. For instance, CellLM and scBERT underperformed in on PBMCs data (mean F1 score: 0.461 and 0.483, respectively) while Geneformer presented poor predictions on immunotherapy datasets (mean F1 score: 0.442). Additionaly, the optimal performances of scFoundation, scGPT, and CellPLM remained stable or showed minor improvement before and after batch effect correction (Supplementary Note 2). Furthermore, to evaluate whether the scFMs outperform traditional methods in drug response prediction, we compared their performances with that of the multilayer perceptron (MLP), scDEAL—a deep-learning model for cancer drug response prediction at single-cell level28, and DREEP, a statistical method for drug response estimation from single-cell transcriptional profiles of tumors29. Our results showed that in liver cancer data, scDEAL, DREEP, and MLP achieved approximately equivalent performances to that of scGPT (0.924), CellLM (0.924), LLaMa3 (0.924), Geneformer (0.924), and scBERT (0.924) (Supplementary Note 3).
F1 scores across tissue, drug, and cancer types using a layer-freezing and b fine-tuning training method. Error bars on the bar plots/dots on the vertical line charts represent standard deviation of the mean F1 score of each method in each category. Bars represent the mean ± s.e.m. of 10 independent folds from cross-validation. Each dot overlaid on the bars indicates the F1 score from an individual fold (n = 10). The folds represent independent validation splits from the same dataset and are considered technical replicates for model evaluation, not independent biological replicates. Source data are provided as a Source Data file. PBMC Peripheral blood mononuclear cells, MM Multiple myeloma, ALL Acute lymphoblastic leukemia, CLL Chronic lymphocytic leukemia, AML Acute myeloid leukemia.
We next evaluated different models using fine-tuning training for predicting drug responses (Fig. 2b; Supplementary Fig. 1b). The results from fine-tuning were overall consistent with those from embedding-based approaches, with scFoundation remaining the best and scBERT remaining the worst across most of the datasets. An exception was observed with melanoma cancer data, where scGPT outperformed scFoundation (mean F1 scores: 0.993 vs 0.978). In liver cancer, CellPLM performed comparably to scFoundation (mean F1 scores, CellPLM: 0.992; scFoundation: 0.997), both outperforming other models. Across drug regimens, scGPT performed similarly to scFoundation for ceritinib (mean F1 scores,scGPT: 0.985; scFoundation: 0.997) and outperformed scFoundation on vemurafenib (mean F1 scores, scGPT: 1.000; scFoundation: 0.990). Similarly, CellPLM demonstrated comparable performance to scFoundation for ceritinib (mean F1 scores, CellPLM: 0.990; scFoundation: 0.997), sorafenib (mean F1 scores, CellPLM: 0.996; scFoundation: 0.999), and vemurafenib (mean F1 scores, CellPLM: 0.977; scFoundation: 0.990). Other metrics, including AUROC, accuracy, precision, recall (Supplementary Fig. 1b), F1 score comparison by Student’s t-test (Supplementary Note 1), as well as correlation between groundtruth and predicted labels (Supplementary Note 4) showed that scFoundation achieves accurate response predictions in diverse datasets.
Unlike the results under the layer-freezing strategy, all models in the fine-tuning setting outperformed the baseline models across various tissue types. Notably, scFoundation achieved the highest mean F1 scores across all tissue categories: 0.940 for peripheral blood mononuclear cells, 0.990 for tumor tissue, 0.962 for bone marrow aspirates, and 0.947 for cell lines (Fig. 2b). Additionally, UCE showed strong performance specifically in tumor tissue analysis, achieving a mean F1 score of 0.869. When evaluating performance across different cancer types, scFoundation remained the top-performing model in all cases except for melanoma, where scGPT outperformed it with a mean F1 score of 0.992, compared to 0.978 for scFoundation. In terms of drug type categories, scFoundation consistently demonstrated the best performance, with mean F1 scores of 0.953 for targeted therapy, 0.996 for chemotherapy, and 0.832 for immunotherapy. Batch effects removal showed minial impact on the model performances (Supplementary Note 2).
Taken together, fine-tuning the FMs generally outperformed layer-freezing based prediction. Among the evaluated models in scDrugMap, scFoundation, when fine-tuned, consistently demonstrated superior performance in drug response prediction across diverse datasets, including various tissues, drugs, cancer types, and regimens. scGPT, also fine-tuned, excelled in predicting responses for melanoma cancer and the vemurafenib regimen. In contrast, CellLM particularly underperformed in immunotherapy and neuroblastoma, under both training strategies.
Cross-data evaluation in primary data collection
To assess the generalizability of FMs in scDrugMap, we performed cross-data evaluation, in which models were trained on certain studies and tested on independent, unseen datasets. This evaluation simulates de novo drug response prediction scenarios in real-world settings.
We first examined model performance using layer-freezing training. In general, most models performed less effectively across various tissue types, drug classes, cancer types, and regimens, compared to the pooled-data evaluation results (Fig. 3a; Supplementary Fig. 2). Nevertheless, a few models showed promising results. Specifically, regarding tissue types, scGPT achieved a mean F1 score of 0.858 in tumor tissue and the performance was statistically significantly higher than that of tGPT (mean F1 score: 0.603, Supplementary Note 1). UCE scored 0.723 in tumor tissue, and CellPLM reached 0.676 in bone marrow aspirate. For drug type, chemotherapy showed the best performance, with UCE and CellLM achieving mean F1 scores of 0.577 and 0.617, respectively. Regarding regimens, the top three performances were observed for UCE and scBERT with ibrutinib (mean F1 scores, UCE: 0.645; scBERT: 0.570), and scGPT with paclitaxel (mean F1 score: 0.566). Across cancer types, LLaMa3 generally showed the lowest variance in F1 scores. scFoundation achieved the highest performance in multiple myeloma, with a mean F1 score of 0.734. CellLM performed best specifically in acute lymphoblastic leukemia (mean F1 score: 0.635) and prostate cancer (mean F1 score: 0.636). We also observed that batch effects removal did not notably alter the performance patterns compared to the uncorrected data (Supplementary Note 2). Given that cross-data evaluations better reflect real-world clinical applications, foundation models typically show lower absolute F1 scores (generally below 0.8) under these more stringent conditions, highlighting the inherent challenge of generalizing across heterogeneous datasets. Nevertheless, they consistently outperform traditional architectures such as scDEAL, demonstrating greater robustness and transferability (Supplementary Note 3). This indicates that cross-data evaluations provide a more realistic measure of clinical utility: while foundation models retain a clear advantage over traditional methods, they still face important limitations, emphasizing the need for future work to further enhance their generalizability and performance.
F1 scores across tissue, drug, and cancer types using A layer-freezing and B fine-tuning training method. Radar plots represent mean F1 scores for different tissue, drug types, and regimens (the radial axis is scaled from 0–1) based on 10-fold cross-validation (n = 10). Violin plots represent the kernel density distribution and the box plots inside represent the median (center line), upper and lower quartiles and 1.5× the interquartile range (whiskers) for all the cancer types. In the circular bar charts, each color segment showed the mean F1 score of the corresponding category across tissue, drug type, and regimen category. In the boxplots, the middle line is the median, the lower and upper hinges correspond to the first and third quartiles, the upper whisker extends from the hinge to the largest value no further than 1.5× the inter-quartile range (IQR) from the hinge, and the lower whisker extends from the hinge to the smallest value at most 1.5× IQR of the hinge. Replicates are technical replicates (10-fold cross-validation) used to estimate model stability. Source data are provided as a Source Data file. MM Multiple myeloma, ALL Acute lymphoblastic leukemia, CLL Chronic lymphocytic leukemia.
Next, we evaluated model performance under the fine-tuning strategy, where UCE demonstrated relatively stronger performance compared to other models across various categories, despite the overall modest performance observed across all models (Fig. 3b; Supplementary Fig. 3). For example, UCE was the best model across tissue types (highest mean F1 score of 0.774 in tumor tissue), drug types (highest mean F1 score of 0.549 in targeted therapy), and regimens (highest mean F1 score of 0.677 in paclitaxel). In contrast, scFoundation, which had performed best under pooled-data evaluation, showed reduced generalizability in the cross-data setting. It was notably the least effective model for the ibrutinib regimen. Interestingly, CellPLM ranked second across drug types (mean F1 score of 0.681 in immunotherapy). Across cancer types, UCE showed top performance in acute lymphoblastic leukemia (ALL, mean F1 score of 0.752), while scFoundation maintained strong results in multiple myeloma (mean F1 score of 0.743). Additional relatively-strong performers included tGPT and CellLM for ALL (mean F1 scores: tGPT-0.728; CellLM-0.718). We also found that batch effects removal did not notably alter the performance patterns compared to the uncorrected data (Supplementary Note 2). Moreover, we conducted a case study on a multiple myeloma dataset using both groundtruth and predicted drug response labels from fine-tuned scGPT (F1 score: 0.2645). Downstream bioinformatics analyses revealed that key biological insights that can be missed when relying on less accurate predicted labels. For example, top up-regulated genes COL1A2 and SEZ6L2 in resistant tumor cells and enriched biological processes such as cell differentiation, cell–cell adhesion, and the p53 pathway in the up-regulated genes were only observed in the ground truth–based analysis (Supplementary Note 5).
To further interpret model behavior, we visualized model-specific embeddings using Uniform Manifold Approximation and Projection (UMAP). These visualizations revealed that scFoundation and scGPT effectively distinguished drug-resistant and drug-sensitive cells (Fig. 4), while other models showed limited separation (Supplementary Fig. 18). In addition, when stratified by scRNA-seq protocols, we observed consistent patterns where scFoundation achieved the best overall performance, while certain models stood out for specific sequencing technologies—for instance, the fine-tuned Geneformer performed notably well on data from the BD Rhapsody system (F1 score of 0.981) (Supplementary Note 6). Collectively, under the cross-data evaluation scenario, most models yielded comparable performance by layer-freezing and fine-tuning, although both showed generally poorer performance than the pooled-data evaluation scenario. When using layer-freezing, scGPT outperformed others in tumor tissue data, while under fine-tuning, UCE consistently delivered the best performance across various tissue types, drug categories, cancer types, and regimens. A model recommendation guide (Supplementary Data 5) and a detailed analysis of model characteristics (Supplementary Data 6) are provided to support model selection for user applications.
UMAP embeddings using layer freezing training method are shown for a scFoundation b scGPT and c UCE, with cells colored cancer type and cell response. Source data are provided as a Source Data file. MM Multiple myeloma, ALL Acute lymphoblastic leukemia, CLL Chronic lymphocytic leukemia, AML Acute myeloid leukemia.
Few-shot learning for drug response prediction
While single-cell FMs such as scFoundation15, UCE24, and scGPT16 demonstrated strong performance in predicting drug responses from single-cell transcriptomic data, we also included a general-purpose foundation model, GPT4o-mini, in the scDrugMap and investigated its applicability in this context. Unlike scFMs specifically pre-trained on single-cell or biomedical data, GPT4o-mini was developed for general natural language tasks. We aimed to assess its zero-shot reasoning capabilities when prompted with single-cell gene expression features for binary drug sensitivity classification (Fig. 5a).
A Prompt display used for GPT4o-mini. For the prompt word template, we first use a technique like the thought chain to prompt the model how it should think about the output, then tell the model the data source and sequence information, and finally we repeat telling the model and give an output template to ensure the consistent of the output format. A complete input example with prompt is also showed. b Radar plots showing the mean F1 scores of GPT4o-mini in predicting drug response across different tissue, drug types, and cancer types (the radial axis is scaled from 0–1). Source data are provided as a Source Data file.
Overall, GPT4o-mini exhibited limited predictive performance, performing near or below baseline levels in most evaluation settings. The model’s highest performance was observed in liver cancer, achieving a mean F1 score of 0.690, which suggests some potential context-specific utility. However, across most other categories, its predictive accuracy was substantially lower. In terms of tissue types, GPT4o-mini achieved mean F1 scores of 0.583 in peripheral blood mononuclear cells, 0.507 in tumor tissue, 0.479 in cell lines, and 0.459 in bone marrow aspirates, highlighting a clear performance gap compared to specialized scFMs. Across drug classes, the model performed best on targeted therapy (mean F1: 0.538), followed by chemotherapy (0.517) and immunotherapy, where performance was notably poor (0.391). When stratified by cancer type, GPT4o-mini’s performance varied widely. Besides liver cancer, the model showed modest performance for chronic lymphocytic leukemia (0.622) and lung cancer (0.594), but struggled on others such as pancreatic cancer (0.190), melanoma (0.320), and breast cancer (0.387) (Fig. 5b).
Additionally, we compared the results using the names of the top 10 highly expressed genes in each cell along with the data source (e.g., GEO accession number) with another two prompting strategies: (1) using the top 100 genes and data source and (2) a chain of thoughts (CoT) prompt, which builds on the original task by adopting a CoT approach, clearly defining the nested data structure (the outer list contains 10 cells, and the inner list contains the 10 highly expressed genes for each cell). This enables the model to follow a predetermined order when assessing drug sensitivity for each cell, helping to reduce ambiguity, improve the clarity and accuracy of reasoning, enhance the consistency and interpretability of results, and provide flexibility for scaling up the task in the future. Across various tissue, drug, and cancer-type datasets, our results showed a consistent pattern: both the top 10-gene and top 100-gene prompts yielded comparable performance based on F1 scores, while the COT prompt achieved relatively higher accuracy in multiple datasets (Supplementary Note 7). These results suggest that, in the absence of domain-specific pretraining or adaptation, general-purpose language models may have limited precision and robustness for drug response prediction in single-cell contexts, highlighting the need for further optimization and domain adaptation.
Model evaluation in the validation data collection
To further assess model generalizability, we evaluated the performance of foundation models in scDrugMap using a separate validation dataset collection (Fig. 6; Supplementary Fig. 4). For this analysis, we applied layer-freezing training method, where each dataset used 10-fold for model training and testing. Among all models, scFoundation demonstrated the best performance across all categories, including tissue types (mean F1 scores, cell line: 0.901; tumor tissue: 0.932; organoid: 0.973), drug types (mean F1 scores, targeted therapy: 0.946; chemotherapy: 0.899; immunotherapy: 0.915), and cancer types (mean F1 scores, NSCLC: 0.997; ovarian cancer: 0.920; basal cell cancer: 0.918; pancreatic cancer: 0.951; colon cancer: 0.847; hepatocellular cancer: 0.980; melanoma: 0.867; breast cancer: 0.985) (Fig. 6a). When categorized by data source, including cell line, fresh biopsy, organoid, and PDX, scFoundation remains achieved the best performance (Supplementary Note 8). Additionally, the strong performance was further supported by AUROC metrics, where scFoundation again led across tissue types, drug types, and cancer types (Fig. 6b). Other metrics including accuracy, precision, and recall also showed that scFoundation provides accurate predictions in this validation data collection (Supplementary Fig. 4).
Radar plots illustrate mean a F1 scores and b AUROC of each model in predicting single-cell drug response using layer-freezing training method across different tissue, drug types, and cancer types. Each radial axis represents the mean of the corresponding scores (0–1, where 1= best performance). The outermost vertices indicate the maximum value for the metric in the given data type. The lines are colored by categories in each data type.
In addition to scFoundation, both scGPT and tGPT demonstrated competitive performance in selected categories. For example, across tissue types, they performed well in tumor tissue (mean F1 scores, scGPT: 0.800; tGPT: 0.863) and organoid (mean F1 scores, scGPT: 0.928; tGPT: 0.946) but not in cell line (mean F1 scores, scGPT: 0.559; tGPT: 0.740) (Fig. 6a). Across drug types, the two models performed similarly, with mean F1 scores ranging from 0.714 of scGPT for targeted therapy to 0.874 of tGPT for chemotherapy. In terms of cancer types, CellPLM, scGPT, and tGPT displayed similar performance, performing well across most cancers (mean F1 scores ranged from 0.755 of CellPLM for basal cell cancer to 0.986 of tGPT for NSCLC). However, in ovarian cancer and melanoma, scGPT showed lower performance compared to the other two models. Specifically, in ovarian cancer, scGPT achieved a mean F1 score of 0.578, while CellPLM and tGPT reached 0.700 and 0.753, respectively. For melanoma, scGPT’s score was 0.589, compared to 0.774 for CellPLM and 0.766 for tGPT.
Computational scalability
In addition to prediction accuracy, scDrugMap also assesses the computational scalability of each foundation model by comparing their architectures, parameter counts, output dimensions, and runtime efficiency (Fig. 7). These factors are critical for selecting models that balance performance with practical deployment needs in large-scale single-cell analysis.
Rows correspond to algorithms ordered chronologically by year and months of publication. The first three columns display model characteristics: whether it uses an encoder-decoder architecture, the type of input embeddings, and whether it is a single-cell foundation model. The next two columns present parameters and output dimensions for each model. The next set of columns show the training and inference time and speed. For each model, the color in each cell is proportional to the corresponding value (scaled between corresponding minimum and maximum values, ignoring values of the two natural language models, shown as dashes). Source data are provided as a Source Data file.
For example, scFoundation, using an encoder-decoder architecture, stood out with its large parameter count (121.2 million), high output dimension of 3072, and efficient training and inference speeds (23.26 it/s and 69.98 it/s, respectively), making it one of the most efficient and powerful models in our benchmark. In comparison, scGPT, which also adopts a dual-architecture approach, had fewer parameters (52.5 million) and a smaller output dimension (512), with lower efficiency (1.43 it/s for training; 1.44 it/s for inference) than scFoundation. Another encoder-decoder model, UCE, offers competitive inference and training speeds, though it has a smaller output dimension of 1280. On the other hand, models like tGPT (decoder-only), scBERT (encoder-only), and Geneformer (encoder-only) used a single architecture and are relatively less efficient, with lower output dimensions (1024 for tGPT, 200 for scBERT, 256 for Geneformer) and slower training and inference speeds. Taken together, these results highlight important trade-offs between architectural complexity, model size, and computational performance. By incorporating these metrics, scDrugMap enables users to make informed decisions about model selection based not only on accuracy but also on runtime efficiency and resource constraints, tailoring deployment to the specific demands of different single-cell datasets and drug response tasks.
Discussion
In this study, we introduce scDrugMap, a comprehensive framework for benchmarking large-scale foundation models in single-cell drug response prediction. We evaluated eight single-cell foundation models and two general-purpose language models using a primary dataset of 326,751 cells and a validation set of 18,856 cells, spanning five tissue types, three drug classes, fourteen cancer types, and twenty-one treatment regimens. While all these datasets were collected through the GEO repository, they were originally generated from independent clinical studies conducted across different laboratories and institutions. Each dataset represents a distinct patient or sample cohort with unique clinical and experimental contexts, ensuring biological and clinical independence rather than repeated sampling from a single source. In addition, we included fresh biopsy and PDX datasets, which provide higher translational relevance and further diversify the data sources beyond conventional in vitro models. Together, these datasets form a highly diversified and representative collection encompassing a broad spectrum of cancer types, data generation platforms, and clinical origins, effectively mitigating concerns about data dependency. Moreover, they constitute a comprehensive and diverse set of publicly available clinical resources for evaluating the drug response prediction capabilities of current foundation models.
Prior to the rise of foundation models, statistical and deep learning (DL) frameworks were widely used for single-cell drug response prediction30. For example, DREEP is a tool that relies on well-established publicly available pharmacogenomic profiles and functional enrichment analysis to predict an individual cell’s vulnerability to hundreds of drugs29. Chen et al. introduces scDEAL, a deep transfer learning model that integrates scRNA-seq data with drug-response annotated bulk RNA-seq to transfer the bulk-level knowledge to the single-cell level28. Another study, DeepDR, provides a DL library designed for drug response prediction31. It supports 135 types of DL models by incorporating a wide range of drug and cell features, along with multiple encoders and fusion modules31. These tools are often specialized for specific biological tasks and were trained on domain-specific data, such as the Genomics of Drug Sensitivity in Cancer (GDSC)32, Cancer Therapeutics Response Portal (CTRP) (http://www.broadinstitute.org/ctrp/), Pharmacogenomics of Responses in Cancer Cell Lines (PRISM) (https://depmap.org), and the Cancer Cell Line Encyclopedia (CCLE)33. Although these frameworks can offer great performance for targeted tasks, they require extensive domain-specific training to improve stability and robustness and may face challenges with scalability and generalization.
In contrast, large-scale, pre-trained foundation models are built on broader knowledge and can transfer learnings across different downstream tasks. Single-cell foundation models, such as scFoundation, scGPT, and scBERT are trained on millions of scRNA-seq data with a variety of cell types from different sources. These models demonstrate the ability to generalize across multiple datasets and tasks without requiring extensive retraining on, for example, specific drug-response data. Foundation models thus represent a promising shift towards more flexible, scalable, and data-efficient approaches in single-cell drug response prediction. However, the advantage of foundation models in this domain is not without limitations. Despite their high computational efficiency and generalizability, our evaluation in scDrugMap demonstrated that fine-tuning consistently yielded better performance than layer-freezing based prediction, indicating that pretrained models still benefit from exposure to task-specific data, particularly in capturing the complex relationships between cellular states and drug responses.
Additionally, our study highlighted that models in pooled-data evaluation outperformed those in cross-data evaluation. This indicates that single-cell FMs benefit from exposure to a broader range of data within a given category. When datasets from multiple studies within the same category (e.g., tumor tissue) are pooled together, the broader diversity may allow the model to learn from heterogeneous sources for more accurate prediction. In contrast, when training is performed on each dataset separately, the model often struggles to generalize, as it is less able to apply its pre-trained knowledge to a narrower and more specific context. Importantly, in cross-data evaluations, which more closely reflect real-world clinical conditions, most models achieved F1 scores below 0.8, indicating that while single-cell FMs show promise, their practical utility remains limited when applied to fully independent datasets. This underscores the need for continued methodological innovation to enhance robustness and cross-study generalization. Furthermore, class imbalance in cross-data scenarios can further deteriorate performance, as measured by area under the precision-recall curve (AUPRC). As shown in Supplementary Fig. 22, the AUPRC values for most FMs remain below 0.7 under both pooled- and cross-data settings, underscoring the difficulty of reliable prediction in imbalanced datasets. This finding highlights an important limitation of current FMs and suggests that future work should explore strategies such as re-weighting, resampling, or tailored fine-tuning to mitigate the effects of class imbalance. Together, these results indicate that while large-scale pooled data can help FMs leverage their broad pre-trained knowledge more effectively, careful consideration of cross-data generalizability and data imbalance is essential for their translation into clinical practice.
Looking ahead, several future directions for single cell foundation models in drug response prediction can be envisioned. First, there remains a need for the incorporation of domain-specific knowledge into model training. Second, integrating multi-modal data sources, such as transcriptomic, proteomic, and genomic data, to enhance the models’ ability to predict drug responses more accurately across diverse patient populations. Third, further exploration of hybrid models combining the strengths of single-cell foundation models and natural language models may lead to more powerful and generalized prediction tools that can effectively handle the complexities of single-cell data while benefiting from broader biological knowledge. Finally, incorporation of explainable AI techniques to enhance model interpretability and biological relevance of predictions made by foundation models, providing clearer insights into the biological mechanisms behind drug responses.
Methods
Overview of scDrugMap framework for drug response prediction
In the scDrugMap framework, we identified recently developed single-cell-related foundation models (scFMs) for drug response prediction. In total, we selected and evaluated ten FMs, comprising eight scFMs and two general-purpose natural language models. The scFMs evaluated in this study include transcriptome-GPT34 (tGPT), single-cell Bidirectional Encoder Representations from Transformers14 (scBERT), Geneformer17, Single-Cell Language Model22 (CellLM), single-cell Foundation Model15 (scFoundation), single-cell Generative Pre-Trained Transformer16 (scGPT), Cell Pre-training Language Model23 (CellPLM), and Universal Cell Embeddings24 (UCE). The two natural language models are Llama3-8B25 and GPT4o-mini26.
Model evaluation settings
In scDrugMap, we evaluated the FMs in two scenarios: the pooled-data scenario and the cross-data scenario. In the pooled-data scenario, datasets from multiple studies within the same category (e.g., tissue type, drug, regimen, or cancer type) were combined into a single dataset. This pooled dataset was then randomly split into training and test sets, allowing the model to learn and be evaluated on data drawn from the same mixed source. In contrast, the cross-data scenario tested the model’s ability to generalize across studies. The model was trained on data from one or more studies and evaluated on data from a completely separate study. In this setting, training and testing were performed using data from different studies, while still remaining within the same category (e.g., tissue type, drug, regimen, or cancer type). This setting can be considered a zero-shot capability evaluation of the model, which better reflects real-world conditions where models are often applied to new, unseen datasets.
Model training strategies
For both evaluation scenarios, we applied two training strategies: layer-freezing and LoRA35 (Low-Rank Adaptation). These strategies were used to train the foundation models (FMs) on the training sets and evaluate their performance on corresponding test sets. For LLaMa3-8B, only the layer-freezing strategy was applied. For GPT4o-mini, we employed a prompt engineering approach similar to that used in GPTCelltype19, without additional fine-tuning.
In the layer-freezing strategy, pretrained scFMs treat each gene (by ENSEMBL ID or gene symbol) as an individual token within a fixed vocabulary (dictionary). To extract feature embeddings for downstream classification, we averaged all token embeddings generated by the scFM. These aggregated embeddings were then passed to a multi-layer perceptron (MLP) for classification. The MLP structure used for classification was defined as follows:
where \({{{\boldsymbol{x}}}}\in {{\mathbb{R}}}^{d}\) is the embedding representation produced by the scFM, \(d\) is the model-specific output dimension, and \(\widehat{{{{\boldsymbol{y}}}}}\) is the predicted probability of drug sensitivity. \({ReLU}\) and \({Sigmoid}\) are the activation functions, and \({{{{\boldsymbol{W}}}}}_{i}\), \({{{{\boldsymbol{b}}}}}_{i}\) represent weights and biases of fully connected layers. We used 10-fold cross-validation, randomly partitioning the data within each biological category (e.g., tissue type, drug class, cancer type, or treatment regimen). The same data partitions were used consistently across models. Only the MLP parameters were trained, while the foundation model weights remained frozen.
In the fine-tuning strategy, we fine-tuned a subset of the model parameters using LoRA, implemented via the peft36 package, while keeping the rest of the model frozen. LoRA enables efficient adaptation of large models by injecting low-rank matrices into trainable layers. We set the LoRA rank to 8, alpha (scaling) to 8, dropout to 0.05, and defined the task type as “SEQ_CLS” (sequence classification). We also conducted a ablation study to evaluate of different LoRA ranks (4, 8, and 16) on the fine-tuning performance of scLLMs for drug response prediction (Supplementary Note 10). The output embeddings from the partially fine-tuned model were again passed through the MLP classifier, and both the MLP parameters and LoRA-injected parameters were trained using backpropagation. Let a pretrained single cell foundation model be denoted as \({scFM}\left(D,W\right)\), where \({{{\boldsymbol{W}}}}=\left\{\widehat{{{{\boldsymbol{W}}}}},{{{{\boldsymbol{W}}}}}_{{ft}}\right\}\), with \(\widehat{{{{\boldsymbol{W}}}}}\) representing frozen weights and \({{{{\boldsymbol{W}}}}}_{{ft}}\) representing the trainable layers. Given a fine-tuning dataset \({D}_{{ft}}={\left\{\left({x}_{i},{y}_{i}\right)\right\}}_{i=1}^{k}\), the objective is to minimize the binary cross-entropy loss:
where \({\Phi }_{{ft}}\) are the parameters of the MLP classifier, and \(L\) is the binary cross-entropy loss.
In LoRA, weight updates are expressed as a low-rank approximation:
where \({{{\boldsymbol{A}}}}\in {{\mathbb{R}}}^{d\times r}\) and \({{{\boldsymbol{B}}}}\in {{\mathbb{R}}}^{r\times k}\) are trainable low-rank matrices, and \(r\) is the rank (set to 8). Here \({{{{\boldsymbol{W}}}}}_{{ft}}\) remains frozen and \(\Delta {{{\boldsymbol{W}}}}\) is learned during fine-tuning.
tGPT
tGPT takes gene expression rankings for generative pretraining on 22.3 million single-cell transcriptomic data. Gene sequence are sorted in descending order of expression, enclosed with special tokens <start> and <end>, and padded with <pad>. We followed instructions from the official tGPT GitHub repository (https://github.com/deeplearningplus/tGPT). For benchmarking, we used the pretrained model ‘transcriptome-gpt-1024-8-16-64’, which is stored in https://huggingface.co/lixiangchun/transcriptome-gpt-1024-8-16-64. The input sequence length of tGPT is 64 and the output embedding dimension is 1024. For layer-freezing training, we average the output of the last layer in the sequence dimension as the input of the downstream classifier. The target module \({W}_{{ft}}\) for LoRA fine-tuning is the attention layer of Transformer block called ‘c_attn’.
scBERT
scBERT uses gene2vec technology to embed genes into a predefined vector space, reflecting the similarity between genes and simplifying model training14. To effectively utilize the transcription levels of each gene, the model discretizes gene expression and converts it into a 200-dimensional vector, similar to the word frequency analysis method in NLP, which serves as the token input for the model. Moreover, scBERT adopts the Performer architecture to mitigate limitation of input length and model efficiency. Though scBERT can handle very long input sequences, we chose to use the first 8000 tokens to maintain uniformity in batch operations. For drug response prediction, we used the pre-trained model provided at https://github.com/TencentAILabHealthcare/scBERT. For the choice of layer-freezing embedding, we keep it consistent with the original operation of scBERT and average the output of the last layer in the sequence dimension. LoRA fine-tuning \({W}_{{ft}}\) includes the key (‘to_k’), value (‘to_v’) and query (‘to_q’) matrices of the model.
Geneformer
Geneformer consists of six standard Transformer encoder blocks. The model features an input size of 2048, an embedding dimension of 256, four attention heads per layer, and a feedforward layer of size 512. Geneformer is pre-trained on 30 million cells and released two versions of the pre-trained model. For this drug response prediction task, we used the pre-trained model named as ‘geneformer-6L-30M_CellClassifier_cardiomyopathies_220224’, which is available and can be obtained from https://huggingface.co/ctheodoris/Geneformer/tree/main. For the input, we follow the same approach as Geneformer by first sorting gene expression values in descending order and then retaining the top 2048 genes. For drug response prediction using fixed-layer embeddings, we utilized the representations extracted from the ‘hidden_states’ output of the final Transformer encoder layer. For LoRA fine-tuning, the target module \({W}_{{ft}}\) included the key (‘key’), value (‘value”), and query (‘query’) matries.
CellLM
Traditional methods rely on the BERT architecture, which leads to anisotropy in the embedding space and inefficient semantic representation. CellLM addresses the computational limitations caused by large batches of data through a new divide-and-conquer contrastive learning method, thereby improving the quality of representation. Trained on 2 million scRNA-seq data, CellLM is the first large language model that learns from both normal cells and cancer cells. To ensure batch consistency, we also limit the max input length to 8000. We choose to take the average of the hidden states output by the encoder in the sequence dimension as the layer-freezing embedding. CellLM utilizes a Performer structure similar to scBERT, so the components involved in \({W}_{{ft}}\) are consistent with scBERT, specifically the key (‘to_k’), value (‘to_v’) and query (‘to_q’) matrices of the model. The pre-trained model for CellLM can be accessed at https://github.com/PharMolix/OpenBioMed/tree/main.
scFoundation
scFoundation has more than 100 million parameters and is trained on over 50 million human single-cell transcriptome data, covering the complex molecular features of all known cell types15. In addition to gene expression values, the input of scFoundation includes two additional indicators: ‘S’ (source) and ‘T’ (target), which represent the input value and the total gene expression, respectively. scFoundation adopts an encoder-decoder design, using only non-zero genes during the encoding stage, while both expressed and non-expressed genes are considered during the decoding stage. We used the pretrained model available at https://github.com/biomap-research/scFoundation. For fixed-layer drug response prediction, the cell embedding was constructed by concatenating four components, as described in the original paper: the representation of ‘S’ and ‘T’, along with the maximum and average embeddings of all genes. LoRA fine-tuning was applied to the model’s output layer.
scGPT
scGPT is pretrained on over 33 million single-cell sequencing data and is further optimized through transfer learning16. For its input, scGPT not only uses information about genes and expression values, but also introduces the special Condition Tokens. The Condition Tokens encompass diverse meta information associated with 502 individual genes, such as modality, batch, and perturbation conditions. Additionally, scGPT can handle unknown genes to address the Out of Vocabulary problem. The model can be accessed at https://github.com/bowang-lab/scGPT. The cell embedding used for layer-freezing based drug response prediction is the <CLS> token (the token defined before pre-training, its representation will be trained during the training phase, and is usually added as the first token in the sequence), which can represent the entire sentence. For LoRA fine-tuning, it targeted the output layer (‘out_proj’).
CellPLM
CellPLM addresses the differences between single-cell data and natural language data that are often overlooked by existing models. Unlike traditional methods that treat genes as tokens and cells as sentences, CellPLM treats cells as tokens and tissues as sentences, using spatial transcriptome data during pre-training to better learn the relationships between cells. They consider the embedding at l-th layer of Transformer as a set of N tokens, where N is the total number of cells in a tissue sample. Additionally, CellPLM introduces a Gaussian mixture prior distribution to mitigate issues of insufficient data and noise. The pre-trained model we used is ‘20230926_85M’ from https://github.com/OmicsML/CellPLM. The cell embedding used for layer-freezing drug response prediction is the output of the last layer in CellPLM. LoRA fine-tuning includes the query, key, and value (‘query_projection’, ‘key_projection’, and ‘value_projection’) matrices in the CellPLM model.
UCE
UCE provides a unified biological latent space through fully self-supervised training with single-cell transcriptome data24. The UCE is capable of capturing significant biological changes and can map any new cell to this embedding space without additional data annotation, model training, or fine-tuning. UCE employs an encoder-decoder structure and performs specific processing on the input, such as adding special tokens (e.g., the CLS token is designed to capture cell-level embeddings when training the model). We used the UCE model provided at https://github.com/snap-stanford/UCE. For layer-freezing drug response prediction, the embeddings was taken from the <CLS> token (the first token in the sequence), as provided by UCE. For LoRA fine-tuning, the output layer out_proj was designated as the target module.
LLaMa3-8B
In addition to the above-mentioned scFMs, we also tested the performance of general FMs in this task, specifically using the open-source LLM released by Meta, Llama-3-8B. Llama-3-8B has nearly 8 billion parameters and is designed to provide efficient natural language understanding and generation capabilities. It is an upgraded version of the Llama series, capable of handling more complex language tasks while optimizing inference speed and resource efficiency. For input processing, to align with the methods used by most scFMs, we first arranged the genes in the sequence from high to low based on expression levels. We then limited the input length to 1024 genes, treating these gene symbols as natural language (separated by space) and used Llama-3-8B’s tokenizer for tokenization. Finally, the embedding output by Llama-3-8B was used for layer-freezing drug response prediction. The model can be obtained by applying for authorization from Meta and downloading it from https://huggingface.co/meta-llama/Meta-Llama-3-8B.
GPT4o-mini
Inspired by GPTCelltype19, we also tested the latest generation of FMs, GPT4o-mini, recently released by OpenAI. It contains knowledge up to July 2024. Like GPTCelltype, we use prompt words to guide the FM’s responses. The specific settings for the model input are as follows:
The input consists of 4 prompt modules, source, and content. The description of each prompt module is shown in Fig. 5a. The \({source}\) is the data source of the current single-cell sequence. We hope to help obtain accurate results by prompting the FM with data source. The \(c{ontent}\) is the name of 10 genes with the highest expression in the single cell sequence. To avoid the problem of hallucination caused by too long input in the large language model, we use the mini batch technique to only let the FM answer 10 sequences in each context [questions on Fig. 5a]. We also evaluated another two prompts: top 100 genes in each cell and data source, and (3) a chain of thoughts (CoT) prompt which builds on the original task by adopting a CoT approach, clearly defining the nested data structure (the outer list contains 10 cells, and the inner list contains the 10 highly expressed genes for each cell). This enables the model to follow a predetermined order when assessing drug sensitivity for each cell, helping to reduce ambiguity, improve the clarity and accuracy of reasoning, enhance the consistency and interpretability of results, and provide flexibility for scaling up the task in the future (Supplementary Note 7). Our experiments were conducted by calling the GPT4o-mini’s API provided by OpenAI (https://openai.com/api).
Data collection and preprocessing
To systematically collect scRNA-seq data, we conducted a literature search in PubMed for studies published up until 2024, using the keywords ‘(drug resistance) AND ((single cell) OR (scRNA)’. Only samples from Homo sapiens with annotated drug response information were included. We also included available datasets collected on drug resistance mechanisms37. This yielded a total of 36 scRNA-seq datasets as the primary data collection, encompassing 11 cancer types, three treatment types, and four tissue types (Supplementary Data 1). To construct an external validation collection, we followed the same curation methodology, focusing on studies published since January 2024. This resulted in 24 scRNA-seq datasets for drug response studies on solid tumors covering five cancer types, three treatment types, and three tissue types (Supplementary Data 1). Both the primary and validation collections included experimental annotations of binary drug response status (sensitive/resistant) at cell-level (Supplementary Data 2). The validation collection was incorporated to further evaluate model generalizability. In addition to the standard train–test split approach (Fig. 6), we also assessed the models in a zero-shot learning setting, directly predicting drug response labels without any retraining (Supplementary Note 11), which suggest that the models can generalize to entirely independent datasets. General natural language models such as GPT5, GPTo4-mini-deep-research and GPT4o-mini were assessed for their zero-shot reasoning capabilities when prompted with single-cell gene expression features for binary drug response prediction (Supplementary Note 12).
Specifically, cells of the nonresponsive samples collected from pre- or post-treatment conditions are labeled as resistant cells, while cells of the responsive samples are labeled as sensitive cells. Quality control and preprocessing were conducted using Seurat v4.3.038 with uniform filtering thresholds (nFeature_RNA > 500 and percent.mt <10%) across all datasets. Although mitochondrial content may vary across tissue types and sequencing technologies, standardized preprocessing was applied to ensure fair and consistent evaluation of model performance. The top 30 principal components (PCs) were then used to cluster the cells at a resolution of 0.5. For datasets exhibiting batch effects, we used Harmony package to correct for them39.
Evaluation metrics
We used four evaluation metrics (F1, AUROC, precision, and recall) for assessing model performance in predicting cell-level drug response. Accuracy is the proportion of correctly predicted samples out of the total number of samples and is most appropriate when the class distribution is balanced. F1 score is the harmonic mean of Precision and Recall, making it suitable for situations where class distribution is imbalanced. F1 can be expressed as: \({\mbox{Precision}}=TP/({TP}+{FP})\), \({\mbox{Recall}}=TP/({TP}+{FN})\), \({{\mbox{F}}}1{{\mbox{Score}}}=(2\times {{\mbox{Precision}}}\times {{\mbox{Recall}}})/({{\mbox{Precision}}}+{{\mbox{Recall}}})\), where TP (True Positive) represents the number of samples correctly predicted as positive, while TN (True Negative) denotes the number of samples correctly predicted as negative. FP (False Positive) refers to the number of samples incorrectly predicted as positive, indicating a false alarm, and FN (False Negative) is the number of samples incorrectly predicted as negative, signifying a missed detection. AUROC represents the area under the Receiver Operating Characteristic (ROC) curve, which is used to evaluate the discriminatory ability of a classification model. The ROC curve plots the True Positive Rate (TPR) against the False Positive Rate (FPR), where \({TPR}=TP/({TP}+{FN})\), \({FPR}=FP/({FP}+{TN})\), \({AUROC}={\int }_{0}^{1}{TPR}({FPR}){dFPR}\). TPR (True Positive Rate) is also known as Sensitivity or Recall. FPR is False Positive Rate. The closer the AUROC value is to 1, the better the model’s ability in predicting accurate drug response.
Web server implementation
We developed scDrugMap as a web-based system for drug response prediction and benchmarking, built on a robust three-tier architecture consisting of the client, server, and database layers. The backend database was implemented using MySQL version 3.23 with the MyISAM storage engine, selected for its simplicity and fast read performance, which suits the platform’s analytical needs. The server layer was constructed using Perl and its DBI (Database Interface) module, enabling efficient interaction with the MySQL database for dynamic content retrieval and user-driven queries. The client layer provides an intuitive, user-friendly web interface designed to facilitate access to curated datasets, model results, and visualization tools. Through this architecture, scDrugMap allows users to seamlessly explore drug response predictions, compare the performance of foundation models, and interact with large-scale single-cell datasets, supporting translational research in drug discovery and precision oncology.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data and results can be downloaded from the scDrugMap web server (https://scdrugmap.com). Details of metadata can be accessed from Supplementary Data 1 and Supplementary Data 2. Source data are provided with this paper.
Code availability
The code used to develop the model, perform the analyses and generate results in this study is publicly available and has been deposited in github at https://github.com/QSong-github/scDrugMap40, under MIT license. All reused libraries retain their original licenses and copyright statements as required. The specific version of the code associated with this publication is archived in Zenodo and is accessible via https://zenodo.org/records/17576321. The web server scDrugMap is available at https://scdrugmap.com and enables users to predict scRNA-seq drug response using different models.
References
Li,Y.,Wang,Z.,Ajani,J. A.&Song,S.Drug resistance and Cancer stem cells.Cell Commun. Signal19,19 (2021).
Chen, E. Y., Raghunathan, V. & Prasad, V. An overview of cancer drugs approved by the US food and drug administration based on the surrogate end point of response rate. JAMA Intern Med. 179, 915–921 (2019).
Schwaederle, M. et al. Association of biomarker-based treatment strategies with response rates and progression-free survival in refractory malignant neoplasms: a meta-analysis. JAMA Oncol. 2, 1452–1459 (2016).
Su, C. Emerging insights to lung cancer drug resistance. Cancer Drug Resist 5, 534–540 (2022).
Oliver, L. et al. Drug resistance in glioblastoma: are persisters the key to therapy?. Cancer Drug Resist 3, 287–301 (2020).
Heo, H. et al. DNA methylome and single-cell transcriptome analyses reveal CDA as a potential druggable target for ALK inhibitor-resistant lung cancer therapy. Exp. Mol. Med. 54, 1236–1249 (2022).
Kwok, H. H. et al. Single-cell transcriptomic analysis uncovers intratumoral heterogeneity and drug-tolerant persister in ALK-rearranged lung adenocarcinoma. Cancer Commun. (Lond.) 43, 951–955 (2023).
Sacchetti, A. et al. Phenotypic plasticity underlies local invasion and distant metastasis in colon cancer. Elife 10, e61461 (2021).
Stevens, L. E. et al. JAK-STAT signaling in inflammatory breast cancer enables chemotherapy-resistant cell states. Cancer Res 83, 264–284 (2023).
Takeuchi, K. et al. Incorporation of human iPSC-derived stromal cells creates a pancreatic cancer organoid with heterogeneous cancer-associated fibroblasts. Cell Rep. 42, 113420 (2023).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med 25, 1251–1259 (2019).
Mukhopadhyay, S. et al. Genome-wide CRISPR screens identify multiple synthetic lethal targets that enhance KRASG12C inhibitor efficacy. Cancer Res 83, 4095–4111 (2023).
Hu, J. et al. Tumor microenvironment remodeling after neoadjuvant immunotherapy in non-small cell lung cancer revealed by single-cell RNA sequencing. Genome Med 15, 14 (2023).
Yang, F. et al. scBERT as a large-scale pretrained deep language model for cell type annotation of single-cell RNA-seq data. Nat. Mach. Intell. 4, 852–866 (2022).
Hao, M. et al. Large-scale foundation model on single-cell transcriptomics. Nat. Methods 21, 1481–1491 (2024).
Cui, H. et al. scGPT: toward building a foundation model for single-cell multi-omics using generative AI. Nat. Methods 21, 1470–1480 (2024).
Theodoris, C. V. et al. Transfer learning enables predictions in network biology. Nature 618, 616–624 (2023).
Bian, H. et al. General-purpose pre-trained large cellular models for single-cell transcriptomics. Natl. Sci. Rev. 11, nwae340 (2024).
Hou, W. & Ji, Z. Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis. Nat. Methods 21, 1462–1465 (2024).
Hu, M. et al. Evaluation of large language models for discovery of gene set function. Nat. Methods 22, 82–91 (2025).
Shen, H. et al. Generative pretraining from large-scale transcriptomes for single-cell deciphering. iScience 26, 106536 (2023).
Zhao, S., Zhang, J. & Nie, Z. Large-scale cell representation learning via divide-and-conquer contrastive learning. arXiv preprint arXiv:2306.04371 (2023).
Wen, H. et al. CellPLM: pre-training of cell language model beyond single cells. bioRxiv, 2023.2010. 2003.560734 (2023).
Rosen, Y. et al. Universal cell embeddings: A foundation model for cell biology. bioRxiv, 2023.2011. 2028.568918 (2023).
AI@Meta Llama 3 Model Card. https://huggingface.co/meta-llama/Meta-Llama-3-8B (2024).
OpenAI GPT-4o mini. https://openai.com/index/gpt-4o-mini-advancing-cost-efficient-intelligence/ (2024).
Franca, G. S. et al. Cellular adaptation to cancer therapy along a resistance continuum. Nature 631, 876–883 (2024).
Chen, J. et al. Deep transfer learning of cancer drug responses by integrating bulk and single-cell RNA-seq data. Nat. Commun. 13, 6494 (2022).
Pellecchia, S., Viscido, G., Franchini, M. & Gambardella, G. Predicting drug response from single-cell expression profiles of tumours. BMC Med 21, 476 (2023).
Erfanian, N. et al. Deep learning applications in single-cell genomics and transcriptomics data analysis. Biomed. Pharmacother. 165, 115077 (2023).
Jiang, Z. & Li, P. DeepDR: a deep learning library for drug response prediction. Bioinformatics 40, btae688 (2024).
Yang, W. et al. Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res 41, D955–D961 (2013).
Barretina, J. et al. Addendum: the cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 565, E5–E6 (2019).
Shen, H. et al. Generative pretraining from large-scale transcriptomes: Implications for single-cell deciphering and clinical translation. bioRxiv, 2022.2001. 2031.478596 (2022).
Hu, E. J. et al. LoRA: Low-Rank Adaptation of Large Language Models. In Proceedings of International Conference on Learning Representations (2022).
Mangrulkar, S. et al. PEFT: state-of-the-art parameter-efficient fine-tuning methods (2022). https://github.com/huggingface/peft (2023).
Liu, X. et al. DRMref: comprehensive reference map of drug resistance mechanisms in human cancer. Nucleic Acids Res. 52, D1253–D1264 (2024).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Korsunsky, I.et al.Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods16, 1289–1296 (2019).
Wang, Q. et al. scDrugMap: Benchmarking Large Foundation Models for Drug Response Prediction. QSong-github/scDrugMap, https://doi.org/10.5281/zenodo.17576320 (2025).
Hancock, D. Y. et al. in Practice and Experience in Advanced Research Computing 2021: Evolution Across All Dimensions Article 11 (Association for Computing Machinery, Boston, MA, USA, 2021).
Boerner, T. J., Deems, S., Furlani, T. R., Knuth, S. L. & Towns, J. in Practice and Experience in Advanced Research Computing 2023: Computing for the Common Good 173–176 (Association for Computing Machinery, Portland, OR, USA, 2023).
Acknowledgements
Q.S. is supported by the National Institute of General Medical Sciences of the National Institutes of Health (R35GM151089). G.W. is supported by the National Institute of General Medical Sciences of the National Institutes of Health (1R35GM150460). This work partially used Jetstream241 through allocation CIS230237 from the Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support (ACCESS)42 program, which is supported by National Science Foundation grants #2138259, #2138286, #2138307, #2137603, and #2138296.
Author information
Authors and Affiliations
Contributions
Q.S. supervised the overall project. Q.S., Y.P. and Q.W. drafted the paper and led the revision process. Q.S., Y.P. and Q.W. were responsible for data collection. Q.W. implement the modeling and expriments. M.Z. developed and maintained the web server. Y.W., Z.T. and G.W. contributed to project discussions and assisted in refining the paper and visualizations. All authors reviewed and approved the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Pengyong Li, Jianzhong Su and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Q., Pan, Y., Zhou, M. et al. scDrugMap: benchmarking large foundation models for drug response prediction. Nat Commun 17, 730 (2026). https://doi.org/10.1038/s41467-025-67481-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67481-2