Abstract
The recent global outbreaks of mpox highlight the urgent need for both fundamental research and antiviral development. However, studying the mpox virus (MPXV), with its large and complex genome, remains challenging due to the requirement for high-containment facilities. Here, we describe a strategy for de novo assembly of MPXV clade IIb genomes in bacterial artificial chromosomes using transformation-associated recombination cloning. Leveraging CRISPR-Cas9 and Lambda Red recombination, we engineer replication-defective MPXV particles with dual deletions of OPG96 (M2R) and OPG158 (A32.5 L)—genes essential for virion assembly, that are capable of recapitulating key stages of the viral life cycle. We apply this system to screen a compound library and identify G243-1720, a potent anti-poxvirus inhibitor with broad activity in vitro and in vivo. G243-1720 blocks the formation of extracellular enveloped virions and cell-cell spread. Resistance mutation selection, crystallographic analysis, analytical ultracentrifugation, and mass photometry reveal that, despite its distinct chemical structure, G243-1720 shares a mode of action with tecovirimat, both functioning by affecting dimerization of protein OPG57 (F13). Our findings underscore the potential of G243-1720 as a promising broad-spectrum anti-poxvirus lead compound and demonstrate the utility of replication-defective MPXV particles as a reliable platform for viral biology studies and antiviral development.
Similar content being viewed by others
Introduction
Mpox virus (MPXV), the causative agent of the zoonotic disease mpox, is an enveloped double-stranded DNA virus belonging to the genus Orthopoxvirus of the family Poxviridae, which also includes variola virus (VARV), vaccinia virus (VACV), and cowpox virus (CPXV)1. MPXV strains are classified into two distinct clades: Clade I, found mainly in Central Africa (particularly the Democratic Republic of the Congo, DRC), is associated with severe clinical symptoms and substantial mortality (4–11%); Clade II, prevalent in West Africa, is characterized by milder symptoms and a mortality rate of less than 1%2,3. For over 50 years, MPXV outbreaks in Africa have been driven predominantly by zoonotic spillover and caused isolated, self-limiting outbreaks with poor human-to-human transmission4.
In 2022, MPXV clade IIb caused the first widespread community transmission of mpox outside Africa5,6, prompting the World Health Organization (WHO) to declare mpox a Public Health Emergency of International Concern (PHEIC). Subsequently, due to the rising number of cases caused by clade Ib, the WHO declared a second mpox PHEIC in 20247, following its earlier reprioritization of mpox as a neglected tropical disease with pandemic potential. This underscores the urgent need for effective prevention and treatment strategies8.
Poxviruses enter host cells via fusion with either the plasma membrane or the membrane of an intracellular acidified vesicle9. Following fusion of the viral envelope with the host membrane, the core—containing the ~200 kb viral genome (encoding nearly 200 proteins) and structural components (e.g., RNA polymerase)—is released into the cytoplasm10. The transcription system of the core then drives the expression of early genes, which facilitate immune evasion, genome release, and subsequent DNA replication. Newly synthesized genomes serve as templates for sequential expression of intermediate and late genes, initiating viral particle assembly11.
The earliest hallmark of particle assembly is the formation of rigid, crescent-shaped structures within viral factories12,13,14,15. These crescents require at least nine essential proteins, comprising three structural and six regulatory (e.g., L2 (now renamed orthopoxvirus gene 9616, OPG96), A30.5 (OPG158)) factors, collectively termed viral membrane assembly proteins (VMAPs). Among these, OPG96 and OPG158 are the smallest and critical for regulating crescent formation. The crescent extends with constant curvature and eventually closes to form immature virions (IV), which then mature into the infectious intracellular mature virus (IMV), also called mature virus (MV). IMV particles mediate cell-to-cell transmission but are released primarily upon cell lysis. A subset of IMV acquires a double-layer envelope from the trans-Golgi network or early endosomes, forming an intracellular enveloped virus (IEV), also called a wrapped virus (WV). IEV particles are transported along microtubules to the cell periphery, where they fuse with the plasma membrane to become cell-associated enveloped virus (CEV) on the cell surface. CEV can induce actin tail formation to propel itself towards neighboring cells for direct spread; alternatively, it may be released into the extracellular space as extracellular enveloped virus (EEV), also called enveloped virus (EV), to mediate systemic dissemination12,13,14,15.
The large and complex MPXV genome and the high biosafety containment (e.g., BSL-3) required to handle the live virus hinder studies on its molecular biology, pathogenesis, and antiviral drug development. Consequently, the development of tools such as replication-defective viruses—which enable genome manipulation and recapitulate viral infection under lower biosafety conditions—provides a valuable platform for scientific research.
Despite the global spread of MPXV, few drugs are approved for clinical use. Currently, only two therapeutics, tecovirimat (ST-246, TOPXX) and cidofovir or its prodrug brincidofovir (CMX001), are authorized for mpox treatment17. Tecovirimat dimerizes the orthopoxvirus protein P37 (also called F13) encoded by the F13L (OPG57) gene, to inhibit the formation of EEV particles and cell-cell spread18,19, but its efficacy remains unproven in clinical settings17,20,21. Cidofovir or brincidofovir can inhibit viral DNA polymerase activity22, but is associated with gastrointestinal and hepatic toxicity. Thus, the identification of antivirals against MPXV is important.
To establish a safe and reliable alternative to authentic MPXV for antiviral research, we assembled a circular, full-length MPXV genome within a bacterial artificial chromosome (BAC) by transformation-associated recombination (TAR) cloning in yeast. We deleted MPXV homologs of OPG96 (VACV L2R) and OPG158 (VACV A30.5 L)—both essential for viral envelope formation23,24,25—to generate replication-defective MPXV (rdMPXV) particles that mimic the viral life cycle. Using this platform, we performed a proof-of-concept screen of a small-compound library and identified G243-1720, a promising antiviral lead compound and discovered it has a mechanism of action analogous to tecovirimat.
Results
Assembly of the MPXV genome lacking OPG96 (MPXV gene M2R)
Compared to clade I MPXV strains, MPXV clade IIb was associated with mild disease symptoms during the 2022 outbreak, and so was selected for the construction of rdMPXV particles. The entire viral genome was divided into 23 fragments (F1–F23), with 80–100 bp overlaps between adjacent fragments. The MPXV OPG96 (M2R) lies within fragment F10, and encodes a 92 aa protein that is highly conserved in orthopoxviruses and essential for viral envelope formation and subsequent morphogenesis of IMV (intracellular mature virion) and EEV (extracellular enveloped virion)23,26. Therefore, this gene was selected for deletion from the MPXV genome. As illustrated in Fig. 1A, adjacent fragments (grouped in sets of four), excluding F1, F2, and F11, were assembled by transformation-associated recombination (TAR) cloning in yeast to generate fragments B, C, E, F, and G.
A Schematic of the assembly of the MPXV genome. The full-length genome was divided into 23 fragments (F1–F23), with deletion of OPG96 in F10. Every four adjacent fragments (except F1, F2, and F11) were assembled via TAR cloning in yeast to generate fragments B, C, E, F, and G. The mGreenLantern (mGreen)-P2A-Gaussia luciferase reporter cassette and the yeast/E. coli shuttle plasmid pBAC was inserted into the tk gene (OPG101) within F11, forming fragment D. Fragments B to G were further assembled through TAR cloning to obtain plasmid pBAC-IΔ96. Fragments F1 and F2 in plasmids were digested with SapI, ligated to form fragment A, and assembled with AscI-linearized pBAC-IΔ96 via Gibson assembly, yielding plasmid pBAC-MPXVΔ96, which contains the full-length MPXV genome with OPG96 deleted. B Schematic of the generation of replication-defective MPXV particle (rdMPXVΔ96). CV-1-96 cells were infected with FPV for 2 h, followed by transfection with pBAC-MPXVΔ96. Cytopathic effects were observed at 120 h post-transfection, and rdMPXVΔ96 particles were harvested and used to infect CV-1-96 and CV-1-WT cells (MOI 0.05, 48 h). Representative fluorescence images from one experiment were shown. Scale bar, 400 μm. C, D Growth kinetics and Gaussia luciferase activity of rdMPXVΔ96 in CV-1-WT and CV-1-96 cells (MOI 0.05). Cells and supernatants were collected at the indicated time points for viral titration (C); supernatants alone were used for measuring luciferase activity (D). Error bars represent standard deviations from three independent experiments (n = 3; mean ± s.d.). E Transmission electron microscopy (TEM) of viral particle assembly in rdMPXVΔ96-infected CV-1-WT and CV-1-96 cells (MOI 1, 36 h). Scale bar, 2 μm. Representative images from three independent experiments are shown. Source data are provided as a Source Data file.
The reporter expression cassette, mGreenLantern (mGreen)-P2A-Gaussia luciferase, under the control of the viral late promoter P11, was inserted together with the linearized yeast/E. coli shuttle plasmid pBAC, into the thymidine kinase (tk) gene (OPG101) within fragment F11, yielding fragment D. Fragments B to G were then assembled by TAR cloning to generate intermediate plasmid pBAC-IΔ96, which lacks OPG96 (Fig. 1A). The plasmid pBAC-IΔ96 exhibited stable maintenance in E. coli over 10 passages (Supplementary Fig. 1A), as confirmed by next-generation sequencing (NGS). No mutations or indels (insertions or deletions) were found in the viral genome or reporter region compared to the reference sequence (Supplementary Fig. 1B, C).
The remaining fragments F1 and F2 in plasmids were digested with the Type IIS restriction enzyme SapI and ligated in vitro to form fragment A. Fragment A was then inserted into the AscI-linearized pBAC-IΔ96, yielding the plasmid pBAC-MPXVΔ96, which contains the full-length MPXV genome lacking OPG96 (Fig. 1A).
Generation of replication-defective MPXV particles lacking OPG96
To rescue the replication-defective virus particles, wild-type CV-1 cells (CV-1-WT) were transduced with a lentivirus expressing OPG96 with codon-optimized coding sequences. The resulting stable cells (CV-1-96) were infected with fowlpox virus (FPV) to provide poxvirus transcriptional enzymes and then transfected with pBAC-MPXVΔ96 (Fig. 1B). The green fluorescence from the expression of mGreen reporter and cytopathic effect were monitored daily. On day 5 post-transfection, distinct viral plaques were observed (Fig. 1B), and the replication-defective MPXV particles lacking OPG96 (rdMPXVΔ96) were harvested.
With the deletion of OPG96, which encodes a protein crucial for virion assembly, the rdMPXVΔ96 particles could only enter and express virus proteins in CV-1-WT cells, but were unable to form progeny IMV or EEV to infect neighboring cells. As expected, only single mGreen-positive cells were detected in CV-1-WT when compared to CV-1-96 cells, which expressed OPG96 (Fig. 1B). The growth kinetics and Gaussia luciferase expression of rdMPXVΔ96 in CV-1-WT and CV-1-96 cells were measured. Abortive replication in CV-1-WT cells was observed, while the virus titer reached around 107 FFU in CV-1-96 cells (Fig. 1C). Although moderate increase of luciferase activity was detected in CV-1-WT cells, the significant increase in CV-1-96 cells suggested that rdMPXVΔ96 underwent multiple cycles of replication, expressing abundant luciferase (Fig. 1D). Transmission electron microscopy revealed structurally intact virions in CV-1-96 cells infected with rdMPXVΔ96 (Fig. 1E). For rdMPXVΔ96 infection in CV-1-WT cells, only short “spicule”-coated membrane arcs resembling crescent segments were detected around the large dense aggregates (Fig. 1E), which was consistent with the previous report describing a VACV mutant not expressing OPG96 (VACV gene L2R)23. Thus, these results demonstrated that rdMPXVΔ96 can only complete the entire lifecycle and propagate in OPG96 trans-complementing cells.
The defective replication property of rdMPXVΔ96 upon serial passaging was further confirmed in passage 1 (P1), P5, and P10 viruses, in which no viral amplification was detected in CV-1-WT cells (Supplementary Fig. 2A,B). PCR analysis of viral genomic DNA from P5 and P10 viruses revealed no indels in either the OPG96-deleted region or the reporter gene region (Supplementary Fig. 2C). Depth-of-coverage analysis of NGS reads also revealed no mutations or indels in the deleted OPG96 region and the whole genome compared to the rdMPXVΔ96 reference sequences (Supplementary Fig. 2D,E). Additionally, analysis of NGS reads indicated no introduction of FPV DNA into the P5 rdMPXVΔ96 genome (Supplementary Fig. 2F). The above results indicate that the replication-defective rdMPXVΔ96 particles exhibit stability in their trans-complementing cells.
Generation of replication-defective MPXV particles with dual deletions of OPG96 and OPG158 (MPXV gene A32.5 L)
The VACV protein OPG158 (A30.5) interacts with OPG96 (L2), and the deletion of OPG158 also blocks the formation of the virion envelope, resulting in the failure to produce infectious viral particles24,25. To further strengthen safety, we constructed replication-defective virus particles lacking both OPG96 and OPG158 (MPXV gene A32.5 L). The CRISPR/Cas9 gene editing in combination with the Lambda Red recombination system was employed in E. coli to achieve this goal27.
As illustrated in Fig. 2A, starting from E. coli DH10B containing the plasmid pBAC-IΔ96, the plasmid pEcCas, which constitutively expresses Cas9 and inducibly expresses the Lambda Red system, was transformed into DH10B for genome editing and recombination. The sgRNA-expressing plasmid pEcgRNA (targeting OPG158) and a homologous repair DNA template containing only the flanking sequences of the OPG158 open reading frame (ORF) were then introduced into the cells. Cas9/sgRNA-mediated cleavage of OPG158 within pBAC-IΔ96, combined with Lambda Red-induced homology-directed repair, resulted in deletion of OPG158. After removing both pEcgRNA and pEcCas plasmids in E. coli, the plasmid with dual deletions of OPG96 and OPG158 was obtained, validated by NGS (Supplementary Fig. 3A–C), and designated as pBAC-IΔ96,158. Using the virus rescue method applied for rdMPXVΔ96, replication-defective MPXV with double deletions (rdMPXVΔ96,158) was rescued in CV-1 cells trans-complementing both codon-optimized MPXV OPG96 and OPG158 (CV-1-96,158).
A Flowchart of CRISPR-based editing in combination with the Lambda Red system to delete OPG158 in plasmid pBAC-IΔ96, resulting pBAC-IΔ96,158 (see the “Methods” section for details). B, C Growth kinetics and Gaussia luciferase activity of rdMPXVΔ96,158 in CV-1-WT, CV-1-96, and CV-1-96,158 cells (MOI 0.05). Cells and supernatants were collected for viral titration (B); supernatants were used for measuring luciferase activity (C). D-E. Body weight change and viral replication in the lungs of dormice. Dormice were inoculated intranasally with PBS (mock), 5 × 105 focus-forming unit (FFU) of rdMPXVΔ96,158 particles, or 1 × 105 FFU of wild-type MPXV. Body weight was monitored (D), and viral genomic copies in the lungs were determined by qPCR (E). F, G TEM analysis of rdMPXVΔ96,158-infected cells (F) and virions (G). Representative images from three independent experiments are shown. Scale bar, 2 μm. H. Susceptibility of cells to rdMPXVΔ96,158 (MOI 0.5, 48 h) analyzed by flow cytometry. I Infection efficiency of rdMPXVΔ96,158 in WT and ΔB3GAT3 A549 cells (MOI 0.5, 0.15, 0.05; 48 h). J–L EC50 of 7D11 antibody (J), tecovirimat (K), and cidofovir (L) in CV-1-96,158 cells infected with rdMPXVΔ96,158 (MOI 0.01 or 0.1, 30 h). M Fluorescence microscopy of the inhibitory effect of tecovirimat (0.125 μM) or cidofovir (25 μM) on rdMPXVΔ96,158 (MOI 0.1, 30 h) in CV-1-96,158 cells (n = 3). Representative images from three independent experiments are shown. Scale bar, 400 μm. Error bars represent standard deviations from three independent experiments (n = 3; B, C, H–L; mean ± s.d.). An animal experiment with n = 3–5 dormice was performed once for bodyweight loss (D; mean ± s.d.) and tissue titration (E; median values). Unpaired, two-sided t-test (I); ****P < 0.0001. Source data are provided as a Source Data file.
Due to the dual-gene deletion, the rdMPXVΔ96,158 particles could theoretically only enter and express proteins in CV-1-WT and CV-1-96 cells, but be unable to form progeny virions unless propagated in the trans-complementing CV-1-96,158 cell line. This was confirmed by viral growth kinetics and Gaussia luciferase expression, which demonstrated that rdMPXVΔ96,158 could complete multi-cycle propagation in the CV-1-96,158 cells, but not in the CV-1-WT or CV-1-96 cells (Fig. 2B, C). Additionally, the defective replication of rdMPXVΔ96,158 was evaluated in dormice, a natural host for MPXV infection28,29. Over 14 days, comparable increases in bodyweight were observed in mock- and rdMPXVΔ96,158-inoculated mice, whereas a significant reduction in body weight was noted in mice challenged with wild-type MPXV (Fig. 2D). Furthermore, rdMPXVΔ96,158 did not exhibit productive replication in the lungs of dormice by day 14, in contrast to wild-type MPXV (Fig. 2E).
Transmission electron microscopy was used to examine virion assembly. Structurally intact virions were observed in rdMPXVΔ96,158-infected CV-1-96,158 cells, but not in both CV-1-WT and CV-1-96 cells (Fig. 2F), consistent with previous reports on VACVs lacking either gene23,25. Additionally, intact virions released from rdMPXVΔ96,158-infected CV-1-96,158 cells were detected (Fig. 2G).
The stability of replication-defective rdMPXVΔ96,158 was studied. After 10 passages in CV-1-96,158 cells, rdMPXVΔ96,158 maintained the single-cycle replication property in CV-1-WT and CV-1-96 cells (Supplementary Fig. 3D). PCR analysis of the genomic DNA from the P10 virus confirmed no indels in OPG96, OPG158, and reporter regions (Supplementary Fig. 3E). NGS analysis further revealed no mutations and indels in the deleted OPG96 and OPG158 regions, and the whole genome of the P5 and P10 rdMPXVΔ96,158 compared to the rdMPXVΔ96,158 reference sequences (Supplementary Fig. 3F–H). Thus, these results demonstrate that rdMPXVΔ96,158 exhibits favorable safety and stability. Because of dual-gene deletions, rdMPXVΔ96,158 possesses a higher safety profile compared to rdMPXVΔ96, which has only a single gene deleted.
Application of rdMPXVΔ96,158 particles in cell biology and antiviral assessment
To expand the application of replication-defective rdMPXVΔ96,158 particles, the susceptibility of commonly used cell lines to rdMPXVΔ96,158 infection was evaluated. As shown in Fig. 2H and Supplementary Fig. 4A, HaCaT, CV-1, Vero E6, BS-C-1, and HFF-1 cells were highly susceptible to infection, as indicated by expression of mGreen reporter, while 293 T, Huh7, and SW13 cells exhibited low susceptibility. It is intriguing that the susceptibility of skin and fibroblast cells, such as HaCaT and HFF-1, may correlate with the MPXV tissue tropism. Previous studies have reported that heparan sulfate and chondroitin sulfate are involved in the attachment of VACV virions to the cell membrane30,31,32,33. The B3GAT3 gene, responsible for heparan sulfate and chondroitin sulfate biosynthesis34, was found to be important for MPXV infection, as its deletion reduced the infection efficiency of rdMPXVΔ96,158 in A549 cells (Fig. 2I and Supplementary Fig. 4B).
Next, the capacity of rdMPXVΔ96,158 particles for antiviral development was explored. In VACV, the specific neutralizing antibody 7D11 targeting the L1 viral protein (OPG95) can block infection35. This antibody also inhibited rdMPXVΔ96,158 infection with an EC50 of 0.83 μg/mL (Fig. 2J). Tecovirimat, which targets the orthopoxvirus P37 protein encoded by OPG57 (VACV F13L) and blocks wrapping of IMV, so that the intracellular enveloped virus (IEV, also called wrapped virus, WV) is not formed, and cidofovir, which targets viral DNA polymerase to inhibit DNA synthesis, are FDA-approved drugs for emergency use in MPXV patients17,36. Both drugs effectively inhibited rdMPXVΔ96,158 spread (tecovirimat) or replication (cidofovir), with EC50 values of 0.003 μM (tecovirimat) and 10.450 μM (cidofovir) in CV-1-96,158 cells (Fig. 2K–M). Similar results were observed in human keratinocytes (HaCaT) stably-expressing OPG96 and OPG158 (HaCaT-96,158) (Supplementary Fig. 4C–E). The antiviral activities of tecovirimat and cidofovir assessed using replication-defective rdMPXVΔ96,158 particles were consistent with previous reports using wild-type MPXV in HFF, HFK, Vero E6, or other cell lines (Supplementary Table 1)19,36,37,38,39,40.
Screening of antiviral compounds using the rdMPXVΔ96,158
As a proof-of-concept, we performed a high-throughput screen of a small-compound library using the rdMPXVΔ96,158 particles. Highly susceptible HaCat-96,158 cells were pre-treated with individual compounds at 10 μM for 1 h, followed by infection with rdMPXVΔ96,158 for 60 h in the presence of the compounds. The numbers of mGreen-positive cells and DAPI-stained nuclei were analyzed by high-content imaging to calculate the number of infected cells versus the total number of cells per well (Fig. 3A).
A Schematic of antiviral compound screening (see the “Methods” section for details). B Validation of three candidate compounds in HaCaT-96,158 cells infected with rdMPXVΔ96,158 (MOI 0.01, 60 h). C EC50 and CC50 of two candidate compounds in HaCat-96,158 cells infected with rdMPXVΔ96,158 (MOI 0.01, 60 h). D Chemical structure of G243-1720. E Fluorescence microscopy analysis of the inhibitory effect of G243-1720 (0.412 μM) on rdMPXVΔ96,158 (MOI 0.01, 60 h) in HaCaT-96-158 and CV-1-96,158 cells. Scale bar, 400 μm. F EC50 and CC50 of G243-1720 in CV-1-96,158 cells infected with rdMPXVΔ96,158 (MOI 0.01, 60 h). G EC50 of G243-1720 in wild-type MPXV-infected Vero E6 cells (200 PFU, 72 h). H Inhibitory effect of G243-1720 on wild-type MPXV (MOI 0.1, 72 h) in Vero E6 cells determined by qPCR targeting OPG47 (gene F3L). I EC50 of G243-1720 in VACV-infected HaCaT cells (MOI 0.0002, 36 h). J The inhibitory effect of G243-1720 (10 µM G243-1720 or 1 µM tecovirimat) on other orthopoxviruses in Vero E6 cells by plaque assay (CPXV-BR or CPXV-E, 300 PFU, 72 h; RPXV, 500 PFU, 48 h; CMLV, 500 PFU, 120 h). K EC50 of G243-1720 in HSV-infected HaCaT cells (MOI 0.005, 36 h). L Inhibitory effect of G243-1720 (10 µM) on VSV (MOI 10, 6 h) and SINV (MOI 10, 21 h) in A549 cells. M Antiviral efficacy of G243-1720 in SCID mice inoculated intranasally with 1 × 106 TCID50 of MPXV. One hour later, mice were administered vehicle control (twice daily, intragastric injection [i.g.]), G243-1720 (twice daily, 22.5 mg/kg [intraperitoneal injection, i.p.] or 45 mg/kg [i.g.]), or tecovirimat (once daily, 50 mg/kg [i.g.]). Error bars represent standard deviations from two (n = 2; G) or three (n = 3; B, C, F, H–I, and K, L) independent experiments (mean ± s.d.). An animal experiment with n = 5 mice was performed once (M, median values). One-way ANOVA with Sidak’s test (H, M); unpaired, two-sided t-test (L); *P < 0.05; **P < 0.01; ****P < 0.001; ns, not significant. Representative images from two (J) or three (E) independent experiments are shown. Source data are provided as a Source Data file.
Compounds that demonstrated more than 90% inhibition of viral replication and maintained over 80% relative cell number were selected as candidates for further validation. Three candidate compounds were re-evaluated at concentrations of 10 μM and 2 μM to confirm their antiviral activity (Fig. 3B). Among these, Merimepodib, reported previously to inhibit MPXV replication, was identified41, a finding that underscores the reliability of our screening procedure.
Two compounds—G243-1720 and AG-1478—were selected to determine the CC50 and EC50 in HaCaT-96,158 cells. Results showed that G243-1720 had the lowest EC50 (0.23 μM) and the highest CC50 (> 100 μM) among them (Fig. 3C). The compound G243-1720 consists of a methylchlorobenzene moiety linked via an aminosulfonyl group to a bicyclic structure, which includes an azepane attached to a carbonyl group and a five-membered nitrogen-containing ring with two methyl groups and a methylated nitrogen atom (Fig. 3D). The compound G243-1720 is synthesized and authenticated (Supplementary Data 1). Immunofluorescent images showed that G243-1720 significantly inhibited the rdMPXVΔ96,158 infection in both HaCaT-96,158 and CV-1-96,158 cells (Fig. 3E), with an EC50 of 0.142 μM in the latter case (Fig. 3F). Similarly, plaque assay indicated that G243-1720 has an EC50 of 0.194 μM in Vero E6 cells infected with wild-type MPXV (Fig. 3G), and 0.16 μM of G243-1720 reduced the number of genome copies by 60% (Fig. 3H). Overall, G243-1720 exhibited potent anti-MPXV activity with low cytotoxicity and was selected for further investigation.
To validate the efficacy of G243-1720 against other orthopoxviruses and its broad-spectrum activity in cell lines from multiple species, the inhibition of VACV replication in HaCaT, A549, CV-1, Vero E6, BHK-21, and MEF cells was examined. The results showed that G243-1720 effectively inhibits VACV replication in these cell lines, with EC50 values ranging from 0.15 to 1.5 μM (Fig. 3I and Supplementary Fig. 5A). Additionally, G243-1720 exhibited inhibitory effect on the replication of other orthopoxviruses like CPXV-BR, CPXV-E, CMLV, and RPXV, but not FPV (a member of the distantly related Avipoxvirus genus), HSV-1 (herpes simplex virus type 1), and the RNA viruses VSV (vesicular stomatitis virus) and SINV (Sindbis virus) (Fig. 3J–L and Supplementary Fig. 5B). These findings demonstrate that G243-1720 specifically inhibits the replication of orthopoxviruses.
Pharmacokinetics (PK) of G243-1720 were studied in ICR mice to assess the bioavailability (Supplementary Fig. 5C,D). The compound demonstrated rapid absorption, achieving a maximum plasma concentration (Cmax) of ~30,000 ng/mL within ~12 min (Tmax) after intraperitoneal (i.p., 22.5 mg/kg) or intragastric (i.g., 45 mg/kg) administration, with a half-life (T1/2) of 0.828 h or 1.85 h, respectively. Plasma compound concentrations remained above the antiviral EC50 value (0.194 μM, 84.97 ng/mL) for ~4 h (i.p.) or ~8 h (i.g.). Moreover, no body weight loss was observed in severe combined immunodeficiency (SCID) mice treated with G243-1720 twice daily for up to 7 days (Supplementary Fig. 5E).
To evaluate the antiviral efficacy of G243-1720 in vivo, SCID mice were inoculated intranasally with MPXV and subsequently were treated with the compound. Viral titers in the lungs were determined at day 10 post-infection (Fig. 3M). Twice-daily administration of G243-1720 via the intraperitoneal (i.p., 22.5 mg/kg) or intragastric (i.g., 45 mg/kg) route reduced the median viral titers from 5.48 logs in the vehicle control to 3.28 logs and 3.92 logs, respectively. As a positive control, once-daily dosing of tecovirimat (i.g., 50 mg/kg) reduced the titers to 2.78 logs. Although G243-1720 demonstrated slightly lower potency than the FDA-approved tecovirimat, its effective antiviral activity against MPXV, combined with favorable pharmacokinetic properties, supports its potential as a promising candidate for further development.
G243-1720 inhibits the formation of EEVs and virus spread
To investigate how G243-1720 inhibits poxvirus infection, we first examined which stages of the viral life cycle the compound might act on. Using tecovirimat as a control, we added the compounds during the entry, post-entry, or entire infection period (Fig. 4A). In HaCaT-96,158 cells infected with rdMPXVΔ96,158, G243-1720 significantly inhibited infection at the post-entry or entire infection period but not during entry, a phenotype similar to tecovirimat treatment (Fig. 4B). Similarly, VACV infection in wild-type HaCaT (HaCaT-WT) cells showed comparable results (Fig. 4C). These data suggest that G243-1720 likely targets viral replication, assembly, or release.
A Schematic of the time-of-addition assay (see the “Methods” section for details). B, C Time-of-addition assay for rdMPXVΔ96,158 in HaCat-96,158 cells (MOI 0.3, 24 h) (B) and VACV in HaCaT-WT cells (MOI 0.002, 24 h) (C). Cells were treated with tecovirimat (0.05 μM) or G243-1720 (4 μM). D–F Effect of G243-1720 on viral replication and late gene expression of rdMPXVΔ96,158 (MOI 0.5, 24 h) in HaCaT-WT cells. Percentage of mGreen-positive cells (D), relative mean fluorescence intensity (E), and relative Gaussia luciferase activity (F) in supernatants were analyzed. G–I. TEM analysis of the effect of G243-1720 on IMV morphogenesis of VACV in CV-1-WT. Cells pre-treated with DMSO (G), tecovirimat (0.2 μM) (H), or G243-1720 (10 μM) (I) were infected with VACV (MOI 0.5, 36 h). Representative images from three independent experiments are shown. Scale bar, 2 μm. J, K Titration of IMVs from VACV-infected HaCaT-WT (J) and rdMPXVΔ96,158-infected HaCat-96,158 cells (K). Cells pre-treated with tecovirimat (1 μM) or G243-1720 (10 μM) were infected (VACV: MOI 0.008; rdMPXVΔ96,158: MOI 0.05 or 0.25, 36 h), washed, and collected for titration. L Titration of EEVs released from rdMPXVΔ96,158-infected CV-1-96,158 cells. Cells pre-treated with tecovirimat (1 μM) or G243-1720 (10 μM) were infected (MOI 0.5, 5 h) and washed. Supernatants were collected after 48 h for EEV titration. M–O Effect of G243-1720 on the cell-to-cell spread of VACV and rdMPXVΔ96,158. Plaque size of VACV (MOI 0.0005, 24 h) in HaCaT-WT cells (M) and rdMPXVΔ96,158 (MOI 0.002, 50 h) in HaCat-96,158 cells (N) were analyzed. Representative images of plaque formation from three independent experiments are shown (O). Scale bar, 2000 μm. Error bars represent standard deviations from three independent experiments (n = 3; B–F and J–N; mean ± s.d.). One-way ANOVA with Sidak’s test (B–F and J–N); *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.001; ns, not significant. Source data are provided as a Source Data file.
To further explore the impact of G243-1720 on viral DNA replication and late gene expression, HaCaT-WT cells were treated with varying concentrations of G243-1720, tecovirimat, or cidofovir, followed by infection with rdMPXVΔ96,158. As indicated by the mGreen reporter, the percentages of virus-infected cells treated with these three compounds were comparable to those of the DMSO control (Fig. 4D). However, analysis of the mean fluorescence intensity of mGreen-positive cells revealed that only cidofovir treatment, which inhibits viral replication, showed decreased intensity (Fig. 4E). Similarly, only cidofovir reduced Gaussia luciferase activity (Fig. 4F). These results suggest that G243-1720 does not interfere with the DNA replication or late gene expression.
To determine whether G243-1720 affects viral particle assembly, particularly IMV formation, VACV-infected CV-1-WT cells were analyzed by transmission electron microscopy. Normal IMV morphology was observed in cells treated with DMSO, G243-1720, or tecovirimat, indicating that G243-1720 does not impair IMV assembly (Fig. 4G–I). Moreover, similar titers of infectious IMVs were detected in VACV-infected HaCaT-WT cells treated with G243-1720 or tecovirimat, albeit lower than those in the DMSO-treated control (Fig. 4J). Likewise, comparable titers were observed in HaCaT-96,158 cells infected with rdMPXVΔ96,158 after treatment with G243-1720 or tecovirimat (Fig. 4K). These results demonstrate that G243-1720 does not interfere with IMV assembly or infectivity.
Next, we asked whether G243-1720 impacts IEV formation and EEV release, which are critical for the long-range spread of the orthopoxviruses26. We first examined EEV production in the supernatant of CV-1-96,158 cells infected with rdMPXVΔ96,158 at 48 h post-infection under G243-1720 treatment. Plaque assays revealed that, compared to DMSO-treated control, both G243-1720 and tecovirimat significantly reduced the titer of extracellular virus (Fig. 4L). We also investigated the effect of G243-1720 on viral plaque size. After treatment with G243-1720 or tecovirimat, VACV-infected HaCaT-WT or rdMPXVΔ96,158-infected HaCaT-96,158 cells were overlaid with methylcellulose, and the plaque size was measured. As shown in Fig. 4M–O, both compounds inhibited cell-cell spread and plaque formation. These findings suggest that G243-1720 might act during IEV formation, its transport to the cell surface or virus release, sharing a mechanism similar to that of tecovirimat.
G243-1720 targets protein OPG57 (F13) of MPXV
To elucidate the potential target of G243-1720, we selected resistant viral mutants by serially passaging rdMPXVΔ96,158 under increasing G243-1720 concentrations (Supplementary Fig. 6A). Resistant mutations were enriched through treatment with varying doses of G243-1720 in HaCaT-96,158 and CV-1-96,158 cells. After 5 or 10 passages, MPXV genomes were sequenced and compared with the control virus passaged in DMSO-treated cells. In addition to non-specific mutations (Supplementary Fig. 6B), two G243-1720-specific mutations, I372N and D294V, were identified in the OPG57 protein (VACV F13) at P5 and P10 in HaCaT-96,158 cells, respectively (Fig. 5A and Supplementary Fig. 6B). Similarly, two single mutations, I372N and A290V, in OPG57 were detected in CV-1-96,158 cells at P5 (Fig. 5A and Supplementary Fig. 6C). Viruses carrying these OPG57 mutations exhibited resistance to high concentrations of G243-1720, as evidenced by enhanced cell spread compared to the DMSO-treated control (Fig. 5B and Supplementary Fig. 7A, B).
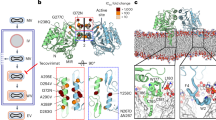
A Specific mutation was identified in the OPG57 gene of rdMPXVΔ96,158. P5 and P10 viruses selected under G243-1720 or DMSO treatment were subjected to next-generation sequencing (NGS) and Sanger sequencing. Three specific mutation sites in the OPG57 gene were identified. B Resistance of the OPG57I372N mutation to G243-1720. HaCat-96,158 cells were pre-treated with G243-1720 (1.5 μM) for 1 h, followed by infection with P5 of rdMPXVΔ96,158 or rdMPXVΔ96,158-OPG57I372N (MOI 0.0005 or 0.005) for 40 h. Cells were fixed and imaged by fluorescence microscopy. Scale bar, 400 μm. C Resistance of OPG57I372N to tecovirimat. HaCat-96,158 cells pre-treated with tecovirimat (0.1 μM) for 1 h were infected with P5 of rdMPXVΔ96,158 or rdMPXVΔ96,158-OPG57I372N (MOI 0.005 or 0.03) for 72 h. Cells were fixed and imaged. Scale bar, 400 μm. D Comparison of resistance mutations in the OPG57 identified for G243-1720, tecovirimat, and IMCBH, another OPG57-targeting small-molecule. Representative images from three independent experiments are shown (B, D).
Tecovirimat targets the OPG57 protein (F13), and specific mutations (e.g., A290V, D294V, and I372N) confer resistance to its inhibitory effects42,43,44,45 (Fig. 5C). Similarly, another OPG57-targeting small-molecule, IMCBH, is rendered ineffective by mutations like D294V43,46 (Fig. 5C). Since the same three mutations emerged under G243-1720 selection, we hypothesized that G243-1720-resistant viruses would also exhibit cross-resistance to molecules like tecovirimat. As expected, tecovirimat failed to inhibit the cell spread of virus carrying the OPG57I372N mutation selected by G243-1720 (Fig. 5D). Although G243-1720 shows a different chemical formula compared to tecovirimat and IMCBH (Supplementary Fig. 7C), the findings of resistance mutations suggest that G243-1720 may target the same viral protein (OPG57) and share similar active sites.
G243-1720 induces the dimerization of the OPG57 protein
Structural data and functional assays have shown that tecovirimat inhibits MPXV egress by inducing OPG57 dimerization18. To test whether G243-1720 uses the same mechanism, we purified a soluble version of OPG57 (sOPG57) and evaluated its oligomerization state in the presence of G243-1720 using analytical ultracentrifugation (AUC), mass photometry, and size-exclusion chromatography coupled with small-angle X-ray scattering (SEC-SAXS) (Fig. 6). Using AUC, we observed that 10 µM G243-1720 induces a shift of sOPG057 from a monomeric form (S = 3.5) to a homodimer (S = 4.6) like that formed by tecovirimat (S = 4.8) (Fig. 6A). Similarly, using mass photometry, in presence of 10 µM G243-1720, we observed 50% of homodimers, versus 80% in presence of 10 µM tecovirimat and 5% in the absence of drugs. The introduction of escape mutants A290V and I372N separately reduced the level of G243-1720-induced dimerization by almost half, while the double mutant D294V/I372N rendered sOPG57 insensitive to G243-1720-induced dimerization (Fig. 6B). In line with these results, SEC-SAXS showed a monodisperse dimer population compatible with the crystallographic dimer (Fig. 6C, D and Supplementary Fig. 8A–C).
A Analytical ultracentrifugation of sOPG057 in the absence (gray line) or presence of 10 μM tecovirimat (yellow line) or 10 μM G243-1720 (green line). Experimentally derived sedimentation coefficient values (Svedberg units (S)) are indicated above each peak. c(s) is the concentration of protein with sedimentation coefficient s. B Mass-photometry assay used to estimate the dimerization activity of G243-1720 in solution using sOPG057WT and G243-1720 escape mutants. The y-axis represents the proportion of sOPG057 dimer, the different columns represent the dimerization degree of sOPG057WT or different OPG057 mutants in the absence or presence of 10 μM G243-1720 or tecovirimat. Data are mean ± s.d. of three independent experiments (n = 3). C Experimental SAXS profile (green dots) and theoretical profiles (dashed lines) for one monomer of sOPG057 (blue dashed line) and the dimer (pink dashed line) are shown. I(q), scattering intensity as a function of the scattering vector q; q(Å-1), scattering vector; Å, Angstrom; X2, chi-squared test. D Orthogonal views of a representative dummy atom model (green) reconstructed from SAXS data. For comparison, we included below a model of the crystallographic sOPG057 dimer with an outline of the dummy model. Data in (B) are from three independent experiments, and each was performed in triplicate. Statistical significance was determined by unpaired, two-tailed Student’s t-test (n = 3; mean ± s.d.; ns, not significant; ****P < 0.0001). Source data are provided as a Source Data file.
To confirm that G243-1720 binds to the same pocket as tecovirimat, cubic crystals of sOPG57 were obtained and then soaked in a solution containing 1 mM of G243-1720 for 18 h. The resulting maps revealed an additional electron density at the dimer interface consistent with the shape of G243-1720; however, the presence of a symmetry axis prevented unambiguous modeling of the molecule (Supplementary Fig. 8D–G). Overall, these findings confirmed that G243–1720 induces OPG57 dimerization by binding to the tecovirimat-binding pocket.
Discussion
In this study, the full-length MPXV genome was assembled in a BAC plasmid with deletions of both OPG96 and OPG158, and used to generate replication-defective rdMPXVΔ96,158 particles that undergo normal replication in trans-complementing cells. The single-cycle replication property of rdMPXVΔ96,158 with dual gene deletions enables its use in lower-biosafety-level facilities (e.g., BSL-2). As a proof-of-concept, we leveraged this platform for a high-throughput screen of a small-compound library, identifying G243-1720. This antiviral compound inhibits MPXV by inducing OPG57 dimerization, a mechanism analogous to the clinically approved drug tecovirimat.
VACV, one of the most extensively studied poxviruses, provides a foundational model for constructing replication-defective viral particles. Genes that are essential for viral replication include: a. genes involved in DNA replication, such as OPG116 (D4)47,48, OPG110 (H5)49, and OPG79 (I3)50; b. genes related to intermediate-stage viral gene transcription, such as OPG134 (A8) and OPG150 (A23)51; and c. genes involved in virion morphogenesis, such as OPG96 (L2)23,25, OPG158 (A30.5)24,25, OPG112 (H7)52, and OPG78 (I2)53. To facilitate the subsequent use of replication-defective particles to investigate the life cycle of MPXV, we selected OPG96 and OPG158 for trans-complementation. These genes are distantly located within the MPXV genome and are both required for virion morphogenesis. To minimize the risk of generating replication-competent virus through recombination, we codon-optimized the deleted genes and expressed them via separate lentiviral vectors in cells for trans-complementation.
Most existing replication-defective VACVs were derived from infectious virus, where the viral genome was edited within complementing cell lines, with replication-defective particles obtained after multiple purification rounds23,25,53. Additionally, rescue of live poxviruses could be achieved in helper virus-infected cells by transfecting either segmented genomic DNA with homologous arms54 or circularized genomes containing BAC elements derived from concatemeric viral DNA55. In contrast, we employed yeast TAR cloning to assemble the full-length MPXV genome de novo. Due to the presence of inverted terminal repeats (ITRs) in fragments F1 and F2, we first assembled the genome without these fragments into a circular intermediate plasmid (pBAC-IΔ96), which was stable upon successive passaging in E. coli. The remaining F1 and F2 fragments were then inserted to construct pBAC-MPXVΔ96, which was used to produce replication-defective particles.
Traditional methods for constructing mutant poxviruses rely on homologous recombination in infected cells using selection markers56, an approach limited by low efficiency, the need for multiple purification steps for generating mutants. Similarly, conventional editing of BAC plasmids bearing VACV genomes depends on homologous recombination with selection markers57, restricting iterative modifications. To overcome these limitations, we developed a platform combining CRISPR-based genome editing with the Lambda Red recombination system in E. coli, enabling rapid, efficient, and multiplexed modifications of the stable intermediate plasmid pBAC-IΔ96. This system significantly facilitates manipulation of the MPXV genome and is applicable to other BAC systems containing large viral genomes, such as herpesviruses and coronaviruses58,59.
Using our rdMPXVΔ96,158 particles, we screened a small-molecule compound library and identified G243-1720 as a promising anti-MPXV candidate. Intriguingly, although G243-1720 inhibited IEV formation and cell-cell spread similar to tecovirimat, it possesses a distinct chemical structure. The OPG57 protein, which is critical for IMV envelopment by trans-Golgi/early endosome-derived membranes, a key step in generating egress-competent virions60,61, is highly conserved among orthopoxviruses and is the target of tecovirimat43. Notably, mutations I372N, A290V and D294V in OPG57 confer tecovirimat resistance in CMLV, VACV, or MPXV42,43,44,45. The identification of these same resistance mutations in MPXV OPG57 following G243-1720 selection, along with observed cross-resistance to tecovirimat, strongly suggests that both compounds share a similar mechanism of action through OPG57 targeting. Crystallographic analysis, mass photometry, AUC and SEC-SAXS confirmed that G243-1720 binds the tecovirimat-binding pocket and induces the dimerization of OPG57 protein, but with slightly lower potency compared to both tecovirimat and IMCBH, another OPG57-targeting compound.
Prior to this work, only two OPG57-targeting inhibitors were known: IMCBH (first reported in 1969)62 and tecovirimat (ST-246, discovered in 2002)63. While IMCBH showed complete cytopathic effect (CPE) inhibition in chicken embryo fibroblasts, its potency was reduced in mammalian cells, and it failed to protect mice and rabbits from VACV infection62. Tecovirimat was approved for treatment of infection with VARV based primarily on safety and pharmacokinetic trials in healthy volunteers and efficacy studies in animal models17,20. However, a recent double-blind, randomized, placebo-controlled trial of endemic mpox caused by MPXV clade I revealed no significant differences between placebo and tecovirimat in lesion resolution time or PCR positivity rates20,64. This outcome is likely explained by the mechanism of action of tecovirimat, which blocks virus spread but not replication. Clinical deployment may therefore occur too late to impact disease progression in symptomatic patients. Additional confounding factors, including suboptimal dosing, comorbidities, inadequate food intake, drug-drug interactions, or genetic variability, may further influence therapeutic outcomes20,64, underscoring the need for additional clinical evaluations.
Given the essential role of OPG57 protein in virulence and highly attenuated phenotype resulting from its deletion, OPG57 remains a compelling antiviral target61. G243-1720, identified here as only the third reported OPG57 inhibitor after IMCBH and tecovirimat, demonstrates broad-spectrum anti-viral activity against multiple orthopoxviruses with consistent efficacy in cellular and mouse models. It represents a lead compound that shares a tecovirimat-like mechanism of action while exhibiting a distinct chemical structure, warranting further investigation of its therapeutic potential.
In summary, our engineered replication-defective MPXV particles establish a robust, convenient, and reliable platform for high-throughput antiviral discovery, offering significant advantages for both fundamental virology studies and therapeutic development against orthopoxviruses.
Methods
Animals and ethics
Male ICR mice (4–6 weeks old), male SCID mice (4–5 weeks old), and male dormice (10–12 weeks old) were housed under controlled environmental conditions (23 ± 2 °C, 30–70% relative humidity, 15 air changes per hour). Animals were group-housed (5 mice per cage) in individually ventilated polycarbonate cages with sterile corn cob bedding and provided ad libitum access to sterile food and water. All animal experiments were conducted in accordance with the National Institutes of Health (NIH) Guide for the Care and Use of Laboratory Animals and were approved by the Animal Care and Use Committees of the Changchun Veterinary Research Institute (IACUC approval no. AMMS-11-2023-041).
Cells and viruses
BS-C-1 (Cell Bank of the Chinese Academy of Sciences #SCSP-5506), CV-1, HaCaT, SW13, Huh7, Vero E6 (Cell Bank of the Chinese Academy of Sciences, #GNO17), HFF-1 (ATCC #SCRC-1041), U-2 OS (ATCC #HTB-96), BHK-21 (ATCC #CCL-10), HEK 293 T (ATCC #CRL-3216), A549 (ATCC #CCL-185), and HeLa (ATCC #CCL-2) were cultured at 37 °C in Dulbecco’s Modified Eagle’s Medium (Hyclone) supplemented with 10% fetal bovine serum (FBS), 10 mM HEPES, 1 mM sodium pyruvate, 1× non-essential amino acids, and 100 U/mL penicillin-streptomycin. All cell lines were routinely tested and confirmed free of mycoplasma contamination. VACV WR strain was propagated in CV-1 cells. Fowlpox virus (FPV) vaccine strain (DCEP25, Boehringer-Ingelheim) was propagated in primary chick embryo fibroblasts (CEF). MPXV IIb strain (GenBank: PP778666.1) was grown in Vero E6 cells. CPXV-Brighton Red strain (CPXV-BR), CPXV isolated from an elephant (CPXV-E), camelpox virus (CMLV) CMS strain, and rabbitpox virus (RPXV) Utrecht strain were propagated in RK13 cells. Herpes simplex virus type 1 (HSV-1) (kindly provided by Prof. Liang, Shanghai Institute of Immunity and Infection), SINV, and VSV were propagated in BHK-21 cells.
Transformation-associated recombination (TAR) cloning of plasmids
The complete genome of MPXV (MPXV-BY-IMB25241, GenBank: ON568298), with a length of 197,378 bp, was divided into 23 fragments (F1–F23). Adjacent fragments were designed with 80–100 bp homologous arms, and OPG96 (M2R) in fragment F10 was deleted. As illustrated in Fig. 1, every four adjacent fragments (except F1, F2, and F11) were assembled in Saccharomyces cerevisiae strain VL6-48 using the yeast/E. coli shuttle plasmid pBAC. Assembled plasmids were purified from VL6-48 and transformed into E. coli DH10B for amplification, resulting in plasmids designated pBAC-B, -C, -E, -F, and -G. For pBAC-D construction, the reporter cassette (mGreenLantern-P2A-Gaussia luciferase) under the control of the late viral promoter P11 was synthesized and inserted into OPG101 (J2R gene, thymidine kinase (tk)) of fragment F11. The modified F11 fragment was cloned into the shuttle plasmid. Fragments B–G (digested from pBAC-B to pBAC-G) were co-transformed with linearized pBAC-D into yeast. Assembled plasmids were purified and transformed into DH10B, yielding the final plasmid pBAC-IΔ96, which lacks F1, F2, and OPG96.
Yeast transformation using the LiAc/SS carrier DNA/PEG
Yeast transformation via lithium acetate was performed as described previously65. Briefly, VL6-48 cells were grown in 2× YPAD medium, washed, and resuspended in sterile water. A 100 μL yeast suspension was transferred to a microcentrifuge tube and mixed with transformation solution containing 36 μL lithium acetate (1.0 M), 240 μL PEG 3350 (50% w/v), 50 μL single-stranded carrier DNA (2.0 mg/mL), and 34 μL DNA (fragments and linearized shuttle vector). The mixture was heat-shocked at 42 °C for 40 min to facilitate DNA uptake. Cells were harvested by centrifugation, resuspended in sterile water, plated on selective medium, and incubated at 30 °C for 3–4 days. Positive clones were validated by PCR and sequencing. Plasmids were extracted and electroporated into E. coli DH10B for amplification.
CRISPR-based editing of the plasmid in E. coli
To knock out OPG158 (A32.5 L) via homologous recombination, a DNA template encompassing only the upstream and downstream sequences of OPG158 was PCR-amplified and joined by overlapping-extension PCR. The spectinomycin resistance gene in the pEcgRNA vector (Addgene #166581) was replaced with an ampicillin resistance gene, and an sgRNA (5ʹ-TTTGGAACGTTTGATAAAAG-3ʹ) targeting OPG158 was cloned to generate pEcgRNA-sgOPG158. The plasmids pBAC-IΔ96 (constructed above) and pEcCas (Addgene #73227) were transformed into DH10B cells, and the Lambda Red recombination system was induced with 10 mM L-arabinose. The sgRNA-expressing plasmid pEcgRNA-sgOPG158 and the DNA recombination repair template (lacking OPG158 coding sequences) were co-electroporated into DH10B cells harboring pBAC-IΔ96 and pEcCas. Positive clones lacking OPG158 were identified by PCR and sequencing. After overnight culture in selection medium, the pEcgRNA-sgOPG158 and pEcCas plasmids were eliminated, resulting in DH10B cells containing only the plasmid pBAC-IΔ96,158 with dual deletions of OPG96 and OPG158.
Generation of stable cell lines
The codon-optimized OPG96 coding sequence was cloned into pLV-EF1α-IRES-puro (Addgene #85132), in which the puromycin resistance gene was replaced with mCherry. Similarly, the codon-optimized OPG158 coding sequence was cloned into the same plasmid, where puromycin resistance was replaced with mIFP. Lentiviruses were packaged by co-transfecting HEK 293 T cells with psPAX2 (Addgene #12260) and pMD2.G (Addgene #12259). Wild-type (WT) CV-1 or HaCaT cells were transduced with lentiviruses and sorted for mCherry+ or mCherry+/mIFP+ cells to establish stable cell lines: CV-1-OPG96 (CV-1-96), CV-1-OPG96-OPG158 (CV-1-96,158), HaCaT-OPG96 (HaCaT-96), and HaCaT-OPG96-OPG158 (HaCaT-96,158).
Assembly of full-length MPXV genome and virus rescue
Fragments F1 and F2 in plasmids were digested with SapI and ligated using T4 DNA ligase to form fragment A. Fragment A and the AscI-linearized pBAC-IΔ96 plasmid were assembled via Gibson assembly to generate the plasmid pBAC-MPXVΔ96, which contains the full-length MPXV genome lacking OPG96. Similarly, the pBAC-MPXVΔ96,158 plasmid lacking OPG96 and OPG158 was constructed in the pBAC-IΔ96,158 backbone. CV-1-96 or CV-1-96,158 stable cell lines were infected with FPV for 2 h. After washing, the pBAC-MPXVΔ96 or pBAC-MPXVΔ96,158 plasmid was transfected into the cells, and the presence of green fluorescence and cytopathic effect (CPE) was monitored daily. Supernatants and cells were harvested and subjected to three freeze-thaw cycles at −80 °C to generate the P0 virus stock. The resulting replication-defective viral particles—with a single deletion of OPG96 (rdMPXVΔ96) or dual deletions of OPG96 and OPG158 (rdMPXVΔ96,158)—were designated accordingly.
Viral growth kinetics and luciferase assay
In 12-well plates, cells were infected with the virus at an MOI of 0.05 for 2 h, followed by a medium change to DMEM containing 2% FBS. At the indicated time points, cells and supernatants were collected together and subjected to three freeze-thaw cycles at −80 °C to release the virus. To determine the titer, monolayers of CV-1-96 or CV-1-96,158 in 96-well plates were inoculated with serially diluted virus for 2 h and then overlaid with methylcellulose for 36 h. Cells were fixed with 4% paraformaldehyde (PFA) in PBS for 1 h and washed with PBS. Green fluorescent foci were counted. To measure Gaussia luciferase activity, only the supernatants were collected. The luciferase activity was determined using the Secrete-Pair Gaussia Luciferase Assay Kit (iGeneBio #LF061) according to the manufacturer’s instructions. Briefly, 10 μL of the supernatant was added to a 96-well plate, followed by 100 μL of assay reagent, which was gently pipetted to mix thoroughly. After incubation at room temperature for 1 min, luminescence was recorded using a FlexStation 3 (Molecular Devices) with an integration time of 1 s per well.
Electron microscopy
CV-1-WT and CV-1-96 cells were infected with rdMPXVΔ96 at an MOI of 1. CV-1-WT, CV-1-96, and CV-1-96,158 cells were infected with rdMPXVΔ96,158 at an MOI of 1. At 36 h post-infection, cells were scraped and fixed with 2.5% glutaraldehyde for 1 h. Samples were further fixed with 1% osmium tetroxide at 20 °C for 2 h, dehydrated through a graded ethanol series, and infiltrated overnight with acetone. After polymerization at 60 °C for 48 h, ultrathin sections were prepared using a Leica UC7 microtome. Sections were stained with uranyl acetate and lead citrate, dried, and imaged using a FEI Tecnai G2 20 Twin transmission electron microscope. Electron microscopy analysis of IMV formation was performed in CV-1-WT cells pre-treated for 1 h with DMSO, tecovirimat (0.2 μM), or G243-1720 (10 μM), followed by infection with VACV at an MOI of 0.5 for 36 h. Cells were subsequently collected, fixed, processed into electron microscopy samples, and visualized as described above.
Assessment of cell susceptibility
The cell lines from different species and organs were infected with rdMPXVΔ96,158 at an MOI of 0.5 for 48 h. Representative images were captured using a fluorescence microscope. To determine infection efficiency, cells were trypsinized, fixed with 2% paraformaldehyde (PFA) for 10 min, and analyzed by flow cytometry. To examine the role of heparan sulfate and chondroitin sulfate in MPXV infection, a B3GAT3-knockout clonal cell line was generated in A549 cells by introducing a ribonucleoprotein (RNP) complex containing Cas9 protein and crRNA. Following limiting dilution and expansion, clonal knockout cells were selected and validated by sequencing. The sgRNA targeting B3GAT3 was designed as: 5ʹ-CCAGAGCCCATACCTGGCAT-3ʹ.
Assessment of viral stability
Trans-complemented stable cells in 12-well plates were infected with replication-defective MPXV particles at an MOI of 0.01 for 2 h. After washing, cells were maintained in DMEM supplemented with 2% FBS to allow viral propagation for 96 h, followed by three freeze-thaw cycles at −80 °C to release virions. Virus particles were then used to infect new cells at an MOI of 0.01 for passaging as described. Genomic DNA from passage 5 (P5) and passage 10 (P10) was extracted. Regions encompassing OPG96, OPG158, or the mGreenLantern-P2A-Gaussia luciferase reporter cassette were amplified for gel electrophoresis and Sanger sequencing. Genomic DNA was also subjected to Next-Generation Sequencing (NGS) to analyze mutations or contamination.
Plaque reduction neutralization (PRNT) assay
The neutralizing antibody 7D11 (Antibody System #RVV13903) targeting OPG95 (VACV L1) was serially diluted in DMEM containing 2% FBS, starting from 80 μg/ml in 5-fold increments. The diluted antibody was mixed in equal volumes with 2% FBS DMEM containing 240 focus-forming units (FFU) of rdMPXVΔ96,158 (final volume: 100 μL) and incubated at 37 °C for 1 h. CV-1-96,158 cells in a 96-well plate were infected with the mixture for 2 h, overlaid with methylcellulose for 30 h, and fixed with 4% PFA. Plaques were quantified using an ImmunoSpot® Analyzer.
Compound screening
HaCat-96,158 cells in 96-well plates were pre-treated with individual small-molecule compounds from a library (3185 compounds; TargetMol, #LC00) at a final concentration of 10 µM for 1 h. Cells were then infected with rdMPXVΔ96,158 at an MOI of 0.001 in the presence of the compounds. After 60 h of infection, cells were fixed and stained with DAPI. Infection efficiency and relative cell number in each well were analyzed using the Operetta CLS High-Content Analysis System. Compounds showing >90% viral inhibition and >80% relative cell number were selected for validation at final concentrations of 10 µM and 2 µM using the same method.
Half-maximal effective concentration (EC50) of compounds
HaCaT-96,158 cells in 96-well plates were pre-treated with serial dilutions of compounds (starting at 100 µM) for 1 h. Without washing, cells were infected with rdMPXVΔ96,158 at an MOI of 0.001 in the presence of the compounds. After 60 h of infection, cells were fixed and stained with DAPI. The infection efficiency in each well was analyzed using the Operetta CLS High-Content Analysis System. Using GraphPad Prism, the compound concentration and viral infection efficiency in treated cells were first log-transformed (base 10), followed by nonlinear regression analysis (curve fitting) to determine the EC50 value. Alternatively, the EC50 of compounds was determined by plaque assay. Briefly, confluent monolayers of Vero E6 cells in 6-well plates were pretreated with serial dilutions of compounds (starting at 60 µM) for 1 h, infected with wild-type MPXV (200 PFU) for 2 h, and overlaid with methylcellulose in the presence of the compounds. After 72 h of infection, cells were fixed, stained with crystal violet, and plaques were counted. To assess the inhibitory effect of compounds on different orthopoxviruses, Vero E6 cells were pretreated with G243-1720 (10 µM) or tecovirimat (1 µM) for 1 h, and infected with viruses for plaque assays (CPXV-BR or CPXV-E, 300 PFU, 72 h; RPXV, 500 PFU, 48 h; CMLV, 500 PFU, 120 h).
Half-maximal cytotoxic concentration (CC50) of compounds
HaCat-96,158 cells in 96-well plates were treated with serial dilutions of compounds (starting at 100 µM). After 60 h of treatment, cell viability was assessed using the CellTiter-Lumi™ Steady Luminescence Cell Viability Assay Kit (Beyotime #C0069L) according to the manufacturer’s instructions. Briefly, CellTiter-Lumi™ reagent was added to each well and mixed by shaking for 2 min. After incubation at room temperature for 10 min, luminescence was recorded using a FlexStation 3 (Molecular Devices) with an integration time of 0.5 s per well. The luminescence intensity in each well was normalized by dividing it by the fluorescence intensity of DMSO-treated control wells. Using GraphPad Prism, the compound concentration and its corresponding relative luminescence intensity were first log-transformed (base 10), followed by nonlinear regression (curve fitting) to determine the CC50 value.
Time of addition experiments
HaCaT-WT cells were seeded in 96-well plates. Full-time group: cells were pre-treated with compounds for 1 h, infected with VACV at an MOI of 0.002 for 2 h in the presence of compounds, washed, and cultured in 2% FBS DMEM with compounds for 24 h. Entry group: cells were pre-treated with compounds for 1 h, infected with VACV (MOI = 0.002) for 2 h with compounds, washed, and cultured in 2% FBS DMEM without compounds for 24 h. Post-entry group: cells were infected with VACV (MOI = 0.002) for 2 h, washed, then cultured in 2% FBS DMEM with compounds for 24 h. After treatment, cells were collected and analyzed for infection efficiency by flow cytometry. Experiments involving rdMPXVΔ96,158 were performed in HaCaT-96,158 cells.
Determination of viral replication and late gene expression
HaCaT-WT cells seeded in 96-well plates were pre-treated with compounds for 1 h. Cells were infected with rdMPXVΔ96,158 at an MOI of 0.5 for 2 h. After removing the viral inoculum by washing, cells were cultured in DMEM supplemented with 2% FBS and compounds. At 24 h post-infection, supernatants were collected, and Gaussia luciferase activity was measured using the Secreted Gaussia Luciferase Assay Kit (iGeneBio #LF061). Cells were harvested to determine the percentage of mGreen-positive cells and mean fluorescence intensity by flow cytometry.
Pharmacokinetics and toxicity of compounds in mice
Male ICR mice (4–6 weeks, 18–22 g) were used to determine the pharmacokinetics of the compounds. Compounds were dissolved in a vehicle solution containing 10% DMSO and 30% w/v HP-β-CD in 0.9% saline. Following intraperitoneal (22.5 mg/kg) or intragastric (45 mg/kg) administration, blood samples were collected into K2EDTA-containing tubes via retro-orbital bleeding at 0.0833, 0.25, 0.5, 1, 2, 4, 6, 8, 12, and 24 h. Plasma was obtained by centrifugation at 3500 × g for 3 min. Compound concentrations were determined using the SCIEX Triple Quad 5500 + LC-MS/MS System.
For toxicity monitoring, severe combined immunodeficiency (SCID) mice aged 4-5 weeks were administered either vehicle control via intragastric administration twice daily, or G243-1720 via intraperitoneal (22.5 mg/kg) or intragastric (45 mg/kg) route twice daily. Body weight was measured on days 0, 3, and 7. G243-1720 was dissolved in a vehicle solution containing 10% DMSO and 30% w/v HP-β-CD in 0.9% saline.
Virus infection and antiviral efficacy in animals
To evaluate the replication of rdMPXVΔ96,158 in vivo, male dormice aged 10-12 weeks were inoculated intranasally with PBS (mock control), 5 × 105 focus-forming unit (FFU) of rdMPXVΔ96,158 particles, or 1 × 105 FFU of wild-type MPXV in a volume of 30 μl. Body weight was monitored over 14 days. Lung tissues were harvested on days 3, 7, and 14 post-infection for DNA extraction. Viral genomic copies were quantified by qPCR targeting OPG47 (F3L) using the following primers and probe: Fwd, 5ʹ-CTCATTGATTTTTCGCGGGATA-3ʹ; Rev, 5ʹ-GACGATACTCCTCCTCGTTGGT-3ʹ; Probe, 5ʹ-CATCAGAATCTGTAGGCCGT-3ʹ.
To assess the antiviral efficacy of G243-1720, male SCID mice aged 4–5 weeks were inoculated intranasally with 1 x 106 TCID50 of MPXV IIb strain (GenBank: PP778666.1) in a volume of 50 μl. Compounds were dissolved in a vehicle solution containing 10% DMSO and 30% w/v HP-β-CD in 0.9% saline. One hour post-infection, mice were given vehicle control via intragastric route twice daily, G243-1720 via intraperitoneal (22.5 mg/kg) or intragastric (45 mg/kg) administration twice daily, or tecovirimat (50 mg/kg) via intragastric injection once daily. At day 10-post infection, mice were euthanized, and lung tissues were collected for viral titer determination. Tissue homogenates were serially diluted (10-fold) and inoculated onto Vero E6 monolayer in 96-well plates in a volume of 100 μl. Viral titers were calculated based on cytopathic effects using the method of Muench and Reed.
Determination of infectious intracellular mature virus (IMV) titer
HaCaT-WT cells seeded in 12-well plates were pre-treated with compounds for 1 h, followed by infection with VACV for 2 h. Unbound virus was washed off, and cells were cultured in DMEM containing 2% FBS and compounds. At specified time points, cells were washed twice with PBS and collected to determine the infectious titer of IMV. To determine the titer, CV-1-WT monolayers in 96-well plates were inoculated with serially diluted virus for 2 h and overlaid with methylcellulose for 30 h. Cells were fixed with 4% paraformaldehyde (PFA) for 1 h, washed with PBS, and the formation of green fluorescent foci was quantified. Experiments involving rdMPXVΔ96,158 were performed in HaCaT-96 or CV-1-96,158 cells.
Determination of infectious extracellular enveloped virus (EEV) titer
CV-1-96,158 cells in T-75 flasks were pre-treated with G243-1720 (10 μM) or tecovirimat (1 μM) in 2% FBS DMEM for 1 h, followed by infection with rdMPXVΔ96,158 at an MOI of 0.5 for 5 h. Cells were washed twice with PBS and cultured in 2% FBS DMEM with compounds for 48 h. Supernatants were collected by centrifugation at 500 × g for 10 min. IMVs in the supernatant were neutralized with VACV IMV neutralizing antibody (mAb 7D11, 50 μg/mL, targeting OPG95, L1), and used to infect CV-1-96,158 cells in a 96-well plate for 2 h. Cells were washed twice with PBS and cultured in 2% FBS DMEM with 1% methylcellulose for 36 h. After fixation with 4% PFA, plaques were quantified using an ImmunoSpot® Analyzer.
Cell–cell spread assay
HaCaT-WT cells were pre-treated with tecovirimat (0.2 μM) or G243-1720 (10 μM) for 1 h, followed by infection with VACV at an MOI of 0.0005 for 2 h. For replication-defective MPXV particles (rdMPXVΔ96,158), HaCaT-96,158 cells were pre-treated with compounds and infected at an MOI of 0.002 for 2 h. After washing to remove unbound virus, cells were overlaid with 1% methylcellulose in 2% FBS DMEM supplemented with the compounds. After 24 h (VACV) or 50 h (rdMPXVΔ96,158), cells were fixed with 4% PFA for 1 h, and images were captured using a fluorescence microscope. The area of mGreen-positive cells was analyzed using ImageJ.
Identification of G243-1720-resistant MPXV mutations
As illustrated in Fig. S5A, HaCaT-96,158 or CV-1-96,158 cells seeded in 12-well plates were infected with rdMPXVΔ96,158 for 2 h. The virus was washed off, and cells were cultured in 2% FBS DMEM containing increasing concentrations of G243-1720. At day 4 post-infection, virus from the highest compound concentration that still supported viral replication was passaged onto new cells in medium containing a higher concentration of G243-1720. This process was repeated for 10 passages. P5 and P10 viruses were sequenced by NGS, and G243-1720-specific mutations were identified by comparing them to control viruses passaged in DMSO-treated cells.
sOPG57 production, purification, and crystallization
Soluble OPG57 (VACV strain Western Reserve, Uniprot code: P04021) was produced in a pET-28a (+) vector (Novagen) with an N-terminal His- and Strep-tag followed by a thrombin cleavage site, as reported elsewhere18. The mutations C53A, A290V, I372N and the double mutation D294V/I372N were inserted into this plasmid. E. coli BL21 (DE3) cells (New England Biolabs) were transformed with these plasmids, and protein expression was induced overnight at 16 °C with 0.25 mM isopropyl β-d-1-thiogalactopyranoside (IPTG). Cells were harvested from 3 L of culture, resuspended in 40 mL cold TNE 8 buffer (Tris-HCl 10 mM, pH 8, NaCl 150 mM, EDTA 1 mM) supplemented with one tablet of complete protease inhibitor (Roche) and then lysed by sonication. After removing the insoluble material by centrifugation at 20,000 x g (30 min, 4 °C), the recombinant protein was purified using Strep-tag-based affinity chromatography in a StrepTrap HP 5 mL column (Cytiva). The protein was treated with 5 mM Tris (2-carboxyethyl) phosphine hydrochloride (TCEP) (Thermo Scientific) for 10 min at room temperature, and aggregates were removed by size-exclusion chromatography in TN 8 buffer (Tris-HCl 10 mM pH 8, NaCl 100 mM).
For crystallization, we removed the purification tags by digestion with 1.5 units of thrombin (Cytiva) per 0.1 mg of protein overnight at 4 °C, treated the protein with 5 mM TCEP for 10 min at room temperature and purified the protein by size exclusion chromatography in TN 8 buffer. The protein was concentrated to 12 mg/mL, crystallized using 1 M Na3 citrate, 0.1 M imidazole, pH 8 and the crystals were soaked for 18 h in the crystallization solution supplemented with 1 mM G243-1720 and 10 % DMSO. Shorter soaking times produced less clear electron densities for G243-1720, probably due to lower occupancy. After soaking, the crystals were cryoprotected with 33% (v/v) glycerol and x-ray diffraction data were collected on beamlines PROXIMA-1 and PROXIMA-2 at the synchrotron SOLEIL (St Aubin, France). The diffraction images were integrated with XDS (version January 10, 2022)66 and the phases were determined using a crystallographic model of OPG57 (PDB ID: 9FHS) as a template using PHASER67. To obtain the final models, the structures were built iteratively and refined using phenix.refine (Phenix version 1.19.2-4158)68 and coot69 using isotropic B factor and TLS groups as refinement strategy. All the models were validated using Molprobity70. The crystallographic statistics are provided in Supplementary Table 2. To facilitate the modeling, the final map was corrected using a bulk-solvent mosaic model available in the PHENIX program (phenix.mosaic). The final map showed an additional electron density at the interface compatible with G243-1720. Because the crystal only diffracted to 3.4 Å, there is a 2-fold symmetry axis crossing the ligand binding pocket, and the occupancy of the ligand is not complete; the density did not show clear features, and we have not succeeded in placing the molecule in the density without ambiguity to study ligand/protein contacts. Coordinates and structure factors have been deposited in the Protein Data Bank under the accession code 9QQV. Figures showing the crystallographic models were generated with PyMol v3.0.3 (Schrödinger, LLC)
Mass photometry
Mass photometry (MP) experiments were done using the TwoMP instrument (Refeyn Ltd, Oxford, UK) using filtered (0.22 μm) TN 8 buffer (10 mM Tris-HCl, pH 8, 100 mM NaCl). Contrast-to-mass calibrations were performed using two references (BSA and urease, both purchased from Sigma Aldrich) diluted in TN 8 buffer, covering a mass range from 66 kDa to 272 kDa. Samples were loaded on clean and dry microscope coverslips assembled into silicone gaskets. Immediately prior to mass photometry measurements, 2 μL of sOPG57 protein was diluted alone or supplemented with the antivirals in 18 μL of TN 8 into the gasket hole and mixed twice. The final concentrations were 25 nM for sOPG57 and 10 μM for the antivirals. Data were collected using AcquireMP v2.3 (Refeyn Ltd), and movies of 2936 frames at a 49 Hz frame rate were recorded. The images were processed and analyzed using DiscoverMP v2.3 (Refeyn Ltd). To extract mole fractions (percentage of each species), all particle contrasts obtained from each movie were converted to mass and calculated mole fractions as the area of each Gaussian curve. The percentage of sOPG57 dimer was plotted using Prism Graphpad v9.0.2.
Analytical ultracentrifugation (AUC)
Sedimentation velocity experiments were carried out at 20 °C in an Optima AUC analytical ultracentrifuge (Beckman Coulter) equipped with double-UV and Rayleigh interference detection. Purified sOPG57 proteins at 0.2 mg ml−1 in the absence or presence of tecovirimat or G243-1720 (10 μM) were centrifuged at 42,000 r.p.m. (23,600 g) using an AN60-Ti rotor and 12 mm thick double sector centrepieces. Absorbance and interference profiles were recorded every 5 min. Buffer viscosity (η = 1.018 cP) and density (ρ = 1.0054 g ml−1) at 20 °C were estimated with SEDNTERP 1.09. Partial specific volumes at 20 °C were estimated based on amino acid sequences using SEDNTERP 1.09 software. Data were analyzed with SEDFIT 16.171 using a continuous size distribution c(S) model. Theoretical sedimentations of the complex were generated using HydroPro 1072.
Size-exclusion chromatography coupled with small-angle X-ray scattering (SEC-SAXS)
Data were collected on the SWING beamline at Synchrotron Soleil (France) using the online HPLC system. The column was equilibrated in the preparation buffer containing 10 mM Tris, pH 8, 100 mM NaCl and 10 μM G243-1720. These experiments have been performed using sOPG57 digested with thrombin. sOPG57 samples at 4.6 mg ml−1 were prepared in a buffer and injected into a size exclusion column (Superdex 75 increase 5/150 mm), cooled at 15 °C, eluting directly into the SAXS flow-through capillary cell at a flow rate of 200 µl min−1. The data were analyzed using FOXTROT and PRIMUS from ATSAS 3.273, from which Guinier was generated. Scattering curves were selected for stable Rg at the apex of the elution profile; the selected curves were averaged, and the buffer signal was subtracted. The scattering curve of the complex was extracted from the main peak using LC and Evolving Factor analysis in RAW 2.3.174. From these corrected scattering curves, the pair distribution function was computed using GNOM (version 5.0)75, and the normalized Kratky plot was generated. Using the structure of sOPG57/G243-1720 (PDB ID: 9QQV), the experimental curve was compared to the theoretical curve using CRYSOL (version 2.8.3)75. Ab initio models were generated with DAMMIN (version 5.3)76, and for each model, sedimentation characteristic was calculated with hydropro (version 10)72. The SAXS statistics are provided (Supplementary Tables 3 and 4).
Statistical analysis
Statistical significance was assigned when P-values were <0.05 using Prism Version 9 (GraphPad). Data analysis was determined by an ANOVA or unpaired t-test, depending on data distribution and the number of comparison groups.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Coordinates and structure factors have been deposited in the Protein Data Bank under the accession code 9QQV. All other relevant data generated in this study are provided in the Supplementary Information/Data and Source Data file. Source data are provided with this paper.
References
Relich, R. F. & Loeffelholz, M. J. Taxonomic changes for human viruses, 2020 to 2022. J. Clin. Microbiol. 61, e0033722 (2023).
Bunge, E. M. et al. The changing epidemiology of human monkeypox-A potential threat? A systematic review. PLoS Negl. Trop. Dis. 16, e0010141 (2022).
Dimitrakoff, J. Monkeypox virus infection across 16 countries-April-June 2022. N. Engl. J. Med. 387, e69 (2022).
Van Dijck, C. et al. Emergence of mpox in the post-smallpox era-a narrative review on mpox epidemiology. Clin. Microbiol. Infect. 29, 1487–1492 (2023).
Isidro, J. et al. Phylogenomic characterization and signs of microevolution in the 2022 multi-country outbreak of monkeypox virus. Nat. Med. 28, 1569–1572 (2022).
Otu, A., Ebenso, B., Walley, J., Barceló, J. M. & Ochu, C. L. Global human monkeypox outbreak: atypical presentation demanding urgent public health action. Lancet Microbe 3, e554–e555 (2022).
Du, M., Liu, M., Niu, B. & Liu, J. The global alarm bell is ringing due to the threat of potential severe cases and deaths caused by clade I of monkeypox virus. Lancet Infect. Dis. 24, 10e616–e61(2024).
Ndembi, N. et al. Evolving epidemiology of Mpox in Africa in 2024. N. Engl. J. Med. 392, 666–676 (2025).
Mercer, J. et al. Vaccinia virus strains use distinct forms of macropinocytosis for host-cell entry. Proc. Natl. Acad. Sci. USA 107, 9346–9351 (2010).
Schmidt, F. I., Bleck, C. K. & Mercer, J. Poxvirus host cell entry. Curr. Opin. Virol. 2, 20–27 (2012).
Harrison, S. C. et al. Discovery of antivirals against smallpox. Proc. Natl. Acad. Sci. USA 101, 11178–11192 (2004).
Condit, R. C., Moussatche, N. & Traktman, P. In a nutshell: structure and assembly of the vaccinia virion. Adv. Virus Res. 66, 31–124 (2006).
Roberts, K. L. & Smith, G. L. Vaccinia virus morphogenesis and dissemination. Trends Microbiol. 16, 472–479 (2008).
Liu, L., Cooper, T., Howley, P. M. & Hayball, J. D. From crescent to mature virion: vaccinia virus assembly and maturation. Viruses 6, 3787–3808 (2014).
Moss, B. Poxvirus membrane biogenesis. Virology 479-480, 619–626 (2015).
Senkevich, T. G., Yutin, N., Wolf, Y. I., Koonin, E. V. & Moss, B. Ancient gene capture and recent gene loss shape the evolution of Orthopoxvirus-Host interaction genes. mBio 12, e0149521 (2021).
Gessain, A., Nakoune, E. & Yazdanpanah, Y. Monkeypox. N. Engl. J. Med. 387, 1783–1793 (2022).
Vernuccio, R. et al. Structural insights into tecovirimat antiviral activity and poxvirus resistance. Nat. Microbiol. 10, 734–748 (2025).
Yang, G. et al. An orally bioavailable antipoxvirus compound (ST-246) inhibits extracellular virus formation and protects mice from lethal orthopoxvirus Challenge. J. Virol. 79, 13139–13149 (2005).
Wilkin, T. Determining effective therapy for Mpox. N. Engl. J. Med. 392, 1547–1548 (2025).
Lenharo, M. Hopes dashed for drug aimed at monkeypox virus spreading in Africa. Nature 632, 965 (2024).
Chan-Tack, K. et al. Benefit-risk assessment for brincidofovir for the treatment of smallpox: U.S. Food and Drug Administration’s Evaluation. Antivir. Res. 195, 105182 (2021).
Maruri-Avidal, L., Weisberg, A. S., Bisht, H. & Moss, B. Analysis of viral membranes formed in cells infected by a vaccinia virus L2-deletion mutant suggests their origin from the endoplasmic reticulum. J. Virol. 87, 1861–1871 (2013).
Maruri-Avidal, L., Weisberg, A. S. & Moss, B. Direct formation of vaccinia virus membranes from the endoplasmic reticulum in the absence of the newly characterized L2-interacting protein A30.5. J. Virol. 87, 12313–12326 (2013).
Carten, J. D., Greseth, M. & Traktman, P. Structure-function analysis of two interacting vaccinia proteins that are critical for viral morphogenesis: L2 and A30.5. J. Virol. 96, e0157721 (2022).
Smith, G. L., Vanderplasschen, A. & Law, M. The formation and function of extracellular enveloped vaccinia virus. J. Gen. Virol. 83, 2915–2931 (2002).
Li, Q. et al. A modified pCas/pTargetF system for CRISPR-Cas9-assisted genome editing in Escherichia coli. Acta Biochim. Biophys. Sin. 53, 620–627 (2021).
Earl, P. L., Americo, J. L., Cotter, C. A. & Moss, B. Comparative live bioluminescence imaging of monkeypox virus dissemination in a wild-derived inbred mouse (Mus musculus castaneus) and outbred African dormouse (Graphiurus kelleni). Virology 475, 150–158 (2015).
Song, G. et al. Establishment of an animal model for monkeypox virus infection in dormice. Sci. Rep. 15, 4044 (2025).
Chung, C. S., Hsiao, J. C., Chang, Y. S. & Chang, W. A27L protein mediates vaccinia virus interaction with cell surface heparan sulfate. J. Virol. 72, 1577–1585 (1998).
Hsiao, J. C., Chung, C. S. & Chang, W. Vaccinia virus envelope D8L protein binds to cell surface chondroitin sulfate and mediates the adsorption of intracellular mature virions to cells. J. Virol. 73, 8750–8761 (1999).
Lin, C. L., Chung, C. S., Heine, H. G. & Chang, W. Vaccinia virus envelope H3L protein binds to cell surface heparan sulfate and is important for intracellular mature virion morphogenesis and virus infection in vitro and in vivo. J. Virol. 74, 3353–3365 (2000).
Pokorny, L. et al. The vaccinia chondroitin sulfate binding protein drives host membrane curvature to facilitate fusion. EMBO Rep. 25, 1310–1325 (2024).
Sammon, D. et al. Molecular mechanism of decision-making in glycosaminoglycan biosynthesis. Nat. Commun. 14, 6425 (2023).
Wolffe, E. J., Vijaya, S. & Moss, B. A myristylated membrane protein encoded by the vaccinia virus L1R open reading frame is the target of potent neutralizing monoclonal antibodies. Virology 211, 53–63 (1995).
Bojkova, D. et al. Drug sensitivity of currently circulating Mpox viruses. N. Engl. J. Med. 388, 279–281 (2023).
Warner, B. M. et al. In vitro and in vivo efficacy of tecovirimat against a recently emerged 2022 monkeypox virus isolate. Sci. Transl. Med. 14, eade7646 (2022).
Smith, S. K. et al. In vitro efficacy of ST246 against smallpox and monkeypox. Antimicrob. Agents Chemother. 53, 1007–1012 (2009).
Smee, D. F., Sidwell, R. W., Kefauver, D., Bray, M. & Huggins, J. W. Characterization of wild-type and cidofovir-resistant strains of camelpox, cowpox, monkeypox, and vaccinia viruses. Antimicrob. Agents Chemother. 46, 1329–1335 (2002).
Yang, X. et al. Identification of neutralizing nanobodies protecting against poxvirus infection. Cell Discov. 11, 31 (2025).
Hishiki, T. et al. Identification of IMP dehydrogenase as a potential target for anti-Mpox virus agents. Microbiol. Spectr. 11, e0056623 (2023).
Smith, T. G. et al. Tecovirimat resistance in Mpox patients, United States, 2022-2023. Emerg. Infect. Dis. 29, 2426–2432 (2023).
Duraffour, S. et al. ST-246 is a key antiviral to inhibit the viral F13L phospholipase, one of the essential proteins for orthopoxvirus wrapping. J. Antimicrob. Chemother. 70, 1367–1380 (2015).
Garrigues, J. M. et al. Identification of tecovirimat resistance-associated mutations in human Monkeypox virus-Los Angeles County. Antimicrob. Agents Chemother. 67, e0056823 (2023).
Alarcon, J. et al. An Mpox-related death in the United States. N. Engl. J. Med. 388, 1246–1247 (2023).
Schmutz, C., Payne, L. G., Gubser, J. & Wittek, R. A mutation in the gene encoding the vaccinia virus 37,000-M(r) protein confers resistance to an inhibitor of virus envelopment and release. J. Virol. 65, 3435–3442 (1991).
Holzer, G. W. & Falkner, F. G. Construction of a vaccinia virus deficient in the essential DNA repair enzyme uracil DNA glycosylase by a complementing cell line. J. Virol. 71, 4997–5002 (1997).
Mayrhofer, J. et al. Nonreplicating vaccinia virus vectors expressing the H5 influenza virus hemagglutinin produced in modified Vero cells induce robust protection. J. Virol. 83, 5192–5203 (2009).
Boyle, K. A., Greseth, M. D. & Traktman, P. Genetic confirmation that the H5 protein is required for vaccinia virus DNA replication. J. Virol. 89, 6312–6327 (2015).
Greseth, M. D., Czarnecki, M. W., Bluma, M. S. & Traktman, P. Isolation and characterization of vΔI3 confirm that vaccinia virus SSB plays an essential role in viral replication. J. Virol. 92, e01719–17 (2018).
Warren, R. D., Cotter, C. A. & Moss, B. Reverse genetics analysis of poxvirus intermediate transcription factors. J. Virol. 86, 9514–9519 (2012).
Meng, X., Wu, X., Yan, B., Deng, J. & Xiang, Y. Analysis of the role of vaccinia virus H7 in virion membrane biogenesis with an H7-deletion mutant. J. Virol. 87, 8247–8253 (2013).
Hyun, S. I., Weisberg, A. & Moss, B. Deletion of the vaccinia virus I2 protein interrupts virion morphogenesis, leading to retention of the scaffold protein and mislocalization of membrane-associated entry proteins. J. Virol. 91, e00558-17 (2017).
Noyce, R. S., Lederman, S. & Evans, D. H. Construction of an infectious horsepox virus vaccine from chemically synthesized DNA fragments. PLoS One 13, e0188453 (2018).
Domi, A. & Moss, B. Cloning the vaccinia virus genome as a bacterial artificial chromosome in Escherichia coli and recovery of infectious virus in mammalian cells. Proc. Natl. Acad. Sci. USA 99, 12415–12420 (2002).
Wyatt, L. S., Earl, P. L. & Moss, B. Generation of recombinant vaccinia viruses. Curr. Protoc. Protein Sci. 89, 5.13.11–15.13.18 (2017).
Domi, A. & Moss, B. Engineering of a vaccinia virus bacterial artificial chromosome in Escherichia coli by bacteriophage lambda-based recombination. Nat. Methods 2, 95–97 (2005).
Tischer, B. K. & Kaufer, B. B. Viral bacterial artificial chromosomes: generation, mutagenesis, and removal of mini-F sequences. J. Biomed. Biotechnol. 2012, 472537 (2012).
Hao, M., Tang J., Ge, S., Li, T. & Xia N. Bacterial-artificial-chromosome-based genome editing methods and the applications in herpesvirus research. Microorganisms 11, 589 (2023).
Blasco, R. & Moss, B. Extracellular vaccinia virus formation and cell-to-cell virus transmission are prevented by deletion of the gene encoding the 37,000-Dalton outer envelope protein. J. Virol. 65, 5910–5920 (1991).
Vliegen, I., Yang, G., Hruby, D., Jordan, R. & Neyts, J. Deletion of the vaccinia virus F13L gene results in a highly attenuated virus that mounts a protective immune response against subsequent vaccinia virus challenge. Antivir. Res. 93, 160–166 (2012).
Kato, N., Eggers, H. J. & Rolly, H. Inhibition of release of vaccinia virus by N1-isonicotinoly-N2-3-methyl-4-chlorobenzoylhydrazine. J. Exp. Med. 129, 795–808 (1969).
Jordan, R., Leeds, J. M., Tyavanagimatt, S. & Hruby, D. E. Development of ST-246® for treatment of poxvirus infections. Viruses 2, 2409–2435 (2010).
Group, P. W. et al. Tecovirimat for Clade I MPXV infection in the democratic Republic of Congo. N. Engl. J. Med. 392, 1484–1496 (2025).
Gietz, R. D. & Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D. Biol. Crystallogr. 66, 133–144 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D. Biol. Crystallogr. 68, 352–367 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Williams, C. J. et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Ortega, A., Amoros, D. & Garcia de la Torre, J. Prediction of hydrodynamic and other solution properties of rigid proteins from atomic- and residue-level models. Biophys. J. 101, 892–898 (2011).
Franke, D. et al. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 50, 1212–1225 (2017).
Hopkins, J. B. BioXTAS RAW 2: new developments for a free open-source program for small-angle scattering data reduction and analysis. J. Appl. Crystallogr. 57, 194–208 (2024).
Svergun, D. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Svergun, D. I. Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing. Biophys. J. 76, 2879–2886 (1999).
Acknowledgements
Grants from the Shanghai Municipal Science and Technology Major Project (ZD2021CY001 to R.Z.), Program of Shanghai Academic Research Leader (22XD1420600 to R.Z.), and Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (2023-PT310-02 to R.Z.) supported this work. We acknowledge support from the Institut Pasteur and the French National Research Agency (ANR: ANR-22-CE11-0003) to P.G.C. We would like to acknowledge Xiaoqing Sun, Yao Wang, and Shen Cai from the Key Laboratory of Medical Molecular Virology (MOE/NHC/CAMS), Shanghai Frontiers Science Center of Pathogenic Microorganisms and Infection, School of Basic Medical Sciences, Fudan University, for their assistance with next-generation sequencing and data analysis. We would like to thank Guangzhou Eighth People’s Hospital of Guangzhou Medical University for providing MPXV samples. We also thank colleagues at the Biosafety Level 3 Laboratory of Changchun Veterinary Research Institute of Chinese Academy of Agricultural Sciences for their help with the experiments. We thank the staff of the Institut Pasteur Crystallography facility for help with crystallization trials and the staff of the PX1, PX2 and SWING beamline at synchrotron SOLEIL (St Aubin, France) for beamline support.
Author information
Authors and Affiliations
Contributions
J.C., L.H., R.V., N.S., J.T., X.C., Z.H., J.L., L.B., B.R., A.H., Yiqi Z., and Y.L. performed the experiments. J.C., L.H., R.V., P.G.C., and R.Z. designed the experiments. Yuyi Z. and S.T. provided technical or material support. J.X., Q.C., Y.L., G.L.S., Y.X., H.L., P.G.C., P.Z., and R.Z. provided administrative, supervision support. J.C., L.H., R.V., N.S., L.B., P.G.C., and R.Z. performed data analysis. J.C. and R.Z. wrote the initial draft of the manuscript, with the other authors contributing to editing into the final form.
Corresponding authors
Ethics declarations
Competing interests
J.C., L.H., Y.X., P.Z., and R.Z. have filed patents on the replication-defective MPXV particles and G243-1720 compound. The remaining authors declare that they have no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, J., Hu, L., Vernuccio, R. et al. Development of a replication-defective mpox virus platform for fundamental and therapeutic research. Nat Commun 17, 798 (2026). https://doi.org/10.1038/s41467-025-67487-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67487-w