Abstract
We report long-term clinical and immunologic outcomes from the MM1636 trial (NCT03047928), which evaluated a peptide vaccine targeting IDO and PD-L1 in combination with nivolumab in patients with advanced melanoma. At a five-year follow-up, the combination demonstrated durable clinical efficacy, with a 25.5-month median progression-free survival. Serum proteomic profiling identified vaccine-specific immune signatures, with increase in CCL3, CCL4, and TNFα emerging as biomarkers of long progression-free survival. Increase in these markers were not observed in a matched anti-PD-1 monotherapy cohort, suggesting distinct immune modulation by the vaccine. Functional studies confirmed vaccine-induced targeting of myeloid cells and associated increase in these cytokines. These findings provide evidence for durable benefit from immune modulatory vaccination and nominate CCL3, CCL4, and TNFα as candidate biomarkers for response.
Similar content being viewed by others
Introduction
Immune checkpoint inhibitors (ICIs) have dramatically improved outcomes for patients with metastatic melanoma1. However, approximately 50% of patients exhibit primary resistance to ICIs, underscoring the need for more effective therapeutic strategies. Current efforts to enhance efficacy include the development of novel ICIs, adoptive cell therapy, and cancer vaccines2,3,4,5; either patient-specific vaccines targeting neoantigens or immune-modulatory vaccines designed to reverse the immunosuppressive tumor microenvironment to an immune-permissive environment, facilitating immune-mediated destruction of cancer cells6,7,8.
From 2017 to 2020, we conducted a phase I/II clinical trial evaluating a combination of nivolumab, an anti-PD-1 ICI, with a peptide vaccine targeting PD-L1 and indoleamine 2,3-dioxygenase (IDO) in 30 patients with advanced, anti-PD-1 naïve melanoma in Cohort A (Clinicaltrials.gov ID: NCT03047928)9. The study treatment included a maximum of 15 doses of the vaccine and up to two years of nivolumab. Vaccine administration followed a priming and maintenance schedule, while nivolumab was administered every two to four weeks, depending on treatment duration. Patients were monitored for safety, clinical efficacy, and treatment-induced immune modulation. The trial results were originally published in 20219, and follow-up data on subgroup efficacy in 202310. Here, we present long-term survival outcomes and comprehensive immunologic correlates of durable response based on serum proteomics.
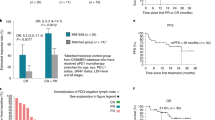
At data cut-off (November 4, 2024), all 30 patients had discontinued treatment, and 15 patients remained alive. Baseline characteristics are presented in Supplementary Table 1. The overall response rate (ORR) was 80%, with 50% of patients achieving complete response (CR) and 30% partial response (PR). The median progression-free survival (mPFS) was 25.5 months, and the median overall survival (mOS) was 60.0 months (Fig. 1A, B). The three- and five-year OS rate was 70% and 50%, respectively. Most responses occurred early, with 22 of 24 responders achieving a response at the first evaluation scan at 12 weeks (Supplementary Fig. 1).
Kaplan–Meier curve of A the progression-free survival, B overall survival, C duration of response for all patient achieving an objective response (partial response (PR) or complete response (CR)), D duration of response for patients receiving either PR or CR, and E a swimmer plot showing the duration of response for the IDO/PD-L1 vaccine and nivolumab and for the next line of treatment.
Durability of response was notable among patients achieving CR, with a median duration of response (DoR) of 53.2 months, compared to 19.3 months in those with PR (Fig. 1C, D). At data cut-off, seven of 15 CRs had ongoing responses (Fig. 1E). All patients with PR eventually experienced disease progression. Among the 23 patients who progressed, 19 received additional systemic therapy, eight of whom received anti-PD-1 reinduction (Supplementary Table 2). Of these, 63% responded again, and three have ongoing responses. These findings suggest both long durability of primary response and retained responsiveness upon re-exposure to ICIs.
Despite the strong clinical efficacy observed, DoR was shorter than in pivotal trials such as CheckMate 067 and RELATIVITY-047, where median DoR exceeded 100 months in some treatment arms1,3. A possible explanation is the shorter median duration of ICI therapy in our study (10.5 months), dictated by national treatment limits for patients with confirmed response. Earlier discontinuation may increase relapse risk in responders, a hypothesis supported by real-world data linking treatment duration to outcome11,12. Also, the optimal number of vaccinations with the IDO/PD-L1 vaccine is not defined and could potentially influence the duration of response.
Comparison of survival curves further underscores the benefit of the combination. The three- and five-year OS rate of 70% and 50%, respectively, are equivalent or superior to that reported in the nivolumab arms of CheckMate 067 (53% and 44%, respectively)1 and RELATIVITY-047 (49% and not-reached, respectively)3. The high early response rate and long mPFS suggest that vaccine-induced immune priming may amplify initial responses and offer more patients prolonged disease control. Results from the IOB-013 phase III trial (ClinicalTrials.gov ID: NCT05155254) are awaited to validate these findings and clarify the duration-response relationship in the context of vaccine therapy.
To identify immunological biomarkers associated with durable clinical response, we conducted a multiplex proteomic analysis of serum using the Olink Immuno-oncology panel. Samples were obtained at baseline and six weeks (6w) after treatment initiation from 22 patients. A comparable group of 25 patients (Supplementary Table 1) receiving first-line anti-PD-1 monotherapy in the noninterventional MM1632 trial13 with matched sample timing served as a reference cohort. Initial assessment of changes in protein levels from baseline to 6w after treatment initiation revealed a shared treatment effect in both cohorts, but the vaccine-treated patients showed distinct shifts in serum protein levels of CCL17, CCL4, IL6, MCP-1, and CD4, which were not observed in monotherapy (Supplementary Fig. 2). Conversely, markers such as CD244, IL33, and ICOSLG changed only in patients receiving monotherapy. These patterns suggest that the IDO/PD-L1 vaccine modifies systemic immune responses beyond those triggered by PD-1 blockade alone.
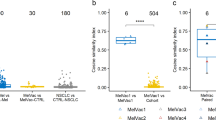
To investigate whether changes in serum protein levels were related to long-term response, we applied elastic-net regression14 to identify proteins associated with long-term outcome. We stratified vaccine-treated patients into long and short PFS groups (above or below median PFS) and utilized logistical regression analysis of elastic-net (Supplementary Fig. 3). The model selected 21 proteins associated with PFS group, including 10 positively and 11 negatively associated to long PFS (Supplementary Fig. 4). The most strongly associated were CCL3, CCL4, and TNFα, showing significant upregulation from baseline to 6w in the long PFS group, but not in the short PFS group (Fig. 2A, Supplementary Fig. 5). None of these markers were significantly upregulated in either PFS groups in the monotherapy cohort (Fig. 2A). Odds ratios ranged from 0.357 (CXCL12) to 1.65 (ICOSLG) (Supplementary Fig. 4), showing varying importance. In addition, univariate logistic regression showed that CCL3 and CCL4 were associated with the PFS group (Fig. 2B). These changes were not significant in the monotherapy cohort, indicating a vaccine-specific immune signature. CCL3, CCL4, and TNFα can be produced by dendritic cells, monocytes, and macrophages, which are all known targets of IDO/PD-L1 vaccination, suggesting that the vaccine-specific immune modulation includes myeloid proinflammatory cytokine signaling, although T cells can also produce them.
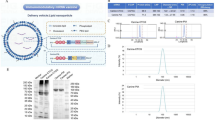
A Change in serum levels of selected proteins from baseline to 6w for patients treated with αPD-1+vaccine with PFS short (n = 10) or PFS long (n = 12), or treated with anti-PD-1 monotherapy (αPD-1-mono) with PFS short (n = 12) or PFS long (n = 13). Two-tailed paired T-test was used to test differences in mean change between time points for each group. B Proteins selected by elastic-net regression as predictive for achieving long PFS. The p-value of univariate logistic regression is reported. C Comparison of changes in protein level from baseline to 6w between patients with short versus long PFS. Two-tailed unpaired T-test was used to test differences in mean change between time points for each group. D Total impact for proteins selected by elastic-net regression for either response group. E CCL3, CCL4, and TNFα quantified with Bio-Plex Multiplex Immunoassay in supernatant of overnight cultures of MonoMac1 cells, CD4+ IDO1-specific T cell clone from patient MM1636.23, or a co-culture of the two at an E:T ratio of 1:10 are shown. Each condition was carried out in triplicate. Box plots show the 25% quantile, median, and 75% quantile for the lower bound, center line, and upper bound of the box, respectively. Box plot whiskers are drawn to the lowest or highest data point within 1.5 * IQR from the lower or upper bound of the box for the lower and upper whiskers, respectively. F CCL3, CCL4, and TNFα quantified with Bio-Plex Multiplex Immunoassay in the supernatant collected from overnight co-cultures of MM1636.05 melanoma cancer cell line, autologous PBMC-derived CD14+ cells with no additional cells (control), or after the addition of autologous PD-L1-specific CD4+ T cell clone, or autologous peripheral blood lymphocytes (PBLs) isolated from PBMCs. Two-tailed unpaired T-test was used to test differences in mean change between groups. Each condition was carried out in triplicate. Box plots show the 25% quantile, median, and 75% quantile for the lower bound, center line, and upper bound of the box, respectively. Box plot whiskers are drawn to the lowest or highest data point within 1.5 * IQR from the lower or upper bound of the box for the lower and upper whiskers, respectively.
To evaluate the clinical relevance of these markers, we examined the magnitude of change and its correlation with outcomes. The largest differential impact between long and short PFS groups was observed for CCL3, CCL4, and TNFα (Fig. 2C). Interestingly, CD5 and ICOSLG also showed higher odds ratios for long PFS, but was not uniquely modulated in the vaccine cohort, suggesting broader roles in T cell activation rather than vaccine-specific effects. A composite score (total impact) incorporating odds ratios and observed fold changes further highlighted CCL3, CCL4 and TNFα as potential biomarkers of vaccine efficacy, as these proteins showed the largest difference in total impact between the long and short PFS groups (Fig. 2D). To underline the coherence of elastic-net results, all proteins showed a more negative total impact for short PFS compared to long PFS. These three proteins were consistently included in elastic-net regression models when using three other patient stratification methods (Supplementary Figs. 6, 7, and 8).
We further performed a comparison of protein changes between the two treatment cohorts using elastic-net regression in the anti-PD-1 monotherapy group (Supplementary Fig. 9). Of the 23 proteins identified in the vaccine combination cohort, only nine overlapped with those associated with outcome in monotherapy, suggesting distinct mechanisms of immune activation. Importantly, changes in CCL3, CCL4, and TNFα were not among the monotherapy-associated proteins, indicating their relevance as vaccine-induced immune correlates. The divergent behavior of CXCL11, which was negatively associated with PFS in the vaccine cohort but positively associated in monotherapy, further emphasizes differential immune dynamics (Supplementary Fig. 10).
To explore the mechanistic basis for the observed changes, we tested in vitro model co-cultures of vaccine-expanded T cells and target-expressing myeloid cells. As previously described, the IDO1-specific T cell clone from the vaccine-treated patient specifically recognizes HLA-matched IDO1+ malignant myeloid MonoMac1 cells9. We observed that co-culture of IDO+ MonoMac1 cells with the CD4⁺ IDO-specific T cell clone led to prominent increases in CCL3, CCL4, and TNFα in the local environment, with minimal to no secretion detected when either cell was cultured separately (Fig. 2E). Similar changes in the cytokine profile were observed when the PD-L1-specific CD4⁺ T cell clone was co-cultured with autologous cancer cells and PBMC-derived CD14⁺ cells mimicking the TME-like conditions in vitro. Increased CCL3, CCL4, and TNFα secretion was detected compared to co-culture lacking the PD-L1-specific T cells (Fig. 2F). These findings were confirmed to be linked to the role of vaccine-expanded T cells targeting tumor-resident myeloid cells (supplementary Fig. 11) and thus driving the specific proinflammatory cytokine response.
One limitation of our study is the absence of a randomized comparator arm, which limits the interpretation of vaccine-specific effects. To address this, we took advantage of matched samples from a patient cohort treated with anti-PD-1 monotherapy. The patient cohorts were similar in baseline characteristics, although three patients in the MM1636 study previously received ipilimumab. The identification of distinct biomarkers in the vaccine cohort suggests that the observed immune changes are not attributable to PD-1 blockade alone. Additionally, the use of elastic-net regression, while powerful, does not provide formal significance values, necessitating complementary validation approaches. Our use of univariable regression and direct groupwise comparisons strengthens confidence in the observed associations, and our mechanistic in vitro studies corroborating the findings further extend the confidence, although they did not include anti-PD-1 treatment. Future efforts will investigate the exact cell dynamics of the altered cytokine levels, as we can only conclude that CCL3, CCL4, and TNFα are increased in the co-culture setting without determining the cellular source of the increase.
In conclusion, this five-year analysis of the MM1636 trial demonstrates durable clinical efficacy and distinct immunologic reprogramming induced by an immune-modulatory peptide vaccine targeting IDO and PD-L1 in combination with nivolumab. The identification of an increase in CCL3, CCL4, and TNFα as vaccine-specific biomarkers of long-term benefit indicates a vaccine-induced immune modulation increasing anti-tumor efficacy, which supports further development of this strategy. Our data provide a compelling rationale for integrating immune-modulatory vaccination into immunotherapy regimens for patients with advanced melanoma and confirm the justification of the ongoing IOB-013 phase III trial.
Methods
Trial design
The trial was an investigator-initiated, single-center, non-randomized phase I/II clinical trial including 30 anti-PD-1 naïve patients with advanced melanoma. The primary endpoints were safety and feasibility, the secondary endpoint was assessment of treatment-induced immunomodulatory changes, and the tertiary endpoint was clinical efficacy evaluated by Response Evaluation Criteria in Solid Tumors (RECIST) version 1.1.
The study treatment comprised a maximum of 15 doses of the IDO/PD-L1 vaccine in the adjuvant Montanide ISA. The first six IDO/PD-L1 vaccines were administered subcutaneously every two weeks; the remaining nine vaccines were administered every four weeks. The anti-PD-1 antibody Nivolumab (3 mg/kg) was administered every two weeks for 24 cycles; thereafter, nivolumab (6 mg/kg) was administered every four weeks for up to a maximum of two years or until progression. If a confirmed objective response (partial or complete response) was obtained, treatment could be terminated after a minimum of six months of therapy.
The trial candidates were included for screening and treatment at the Department of Oncology, Herlev Hospital, Copenhagen, Denmark. The clinical trial was conducted according to Good Clinical Practice (GCP) and the Declaration of Helsinki. The trial was approved by the Capital Region of Denmark Ethical Committee (H-17000988) and the Danish Medical Agency (2017011073), and was monitored by the GCP unit, Copenhagen, and was registered at http://www.clinicaltrials.gov, ID: NCT03047928.
Patient inclusion
Thirty patients were enrolled in the trial from December 2017 to June 2020. Inclusion criteria were diagnosis of metastatic or locally advanced melanoma, no prior exposure to anti-PD-1 therapy, a minimum of one target lesion according to RECIST v.1.1, and ECOG performance status (PS) of 0 or 1. Exclusion criteria were the presence of active autoimmune disease, treatment with systemic steroids or other experimental drugs, and the presence of >4 brain metastases if the lesions measured >1 cm.
Clinical evaluation
Clinical tumor response was evaluated every three months until progression using PET-CT scans. Patients with ongoing response after last treatment administration were evaluated every three to six months (every 3 months the first year, every 4 months the second year, and every six months in later years) until five years after last treatment. The clinical responses were registered as complete response (CR), partial response (PR), stable disease (SD), or progressive disease (PD).
Comparison cohort of patients receiving anti-PD-1 monotherapy
A separate cohort of 25 patients with unresectable melanoma stage IV receiving anti-PD-1 monotherapy in a noninterventional clinical trial (MM1632) with blood samples collected at the same time points was used as a reference group for the analyses of serum proteins. The MM1632 trial was an observational clinical trial prospectively collecting blood samples from patients with advanced melanoma receiving standard treatment with immune checkpoint inhibitors at the Department of Oncology, Herlev Hospital13. Baseline characteristics for the 25 patients receiving anti-PD-1 monotherapy in MM1632 are shown in Supplementary Table 1.
Olink analysis
Serum protein levels were assayed using the Olink Immuno-Oncology panel (v.3111) containing 96 markers for immune activation relevant in cancer. Blood was collected at baseline and six weeks (6w) for 22 patients treated with nivolumab in combination with the IDO-PD-L1 vaccine. Blood samples from the 25 patients from MM1632 receiving anti-PD-1 monotherapy were used as a reference group.
All samples were run on two different plates for the Olink Normalized Protein eXpression (NPX) analysis. All samples within a study were run on the same plate.
QC was carried out by removing samples or proteins where the missing data frequency exceeded 10%, and imputing data for samples where the missing data frequency was below 10%. No samples or proteins were removed due to these criteria. Data was imputed using the k-nearest neighbors method (impute.knn function from the impute package (v.1.58.0)).
For proteins with more than 1 value under the limit of detection (LoD), all values below LoD were adjusted to the LoD value. LoD values were determined and supplied by OLINK.
The following was imputed for MM1636: IL2 for 1 sample, IL33 for 1 sample, IL5 for 1 sample, and IL4 for 3 samples.
The following was imputed for MM1632: IL1α for 3 samples, IL2 for 2 samples, IL5 for 2 samples, and IL4 for 5 samples.
To characterize changes in serum protein levels related to clinical efficacy, patients were stratified into two groups based on progression-free survival (PFS), with long PFS defined as PFS > median and short PFS defined as PFS < median.
R version 4.3.2 was used for the analysis of Olink data. For the analysis and validation of results, the following packages were used: glmnet (version 4.1-8), ggpubr (version 0.6.0), dplyr (version 1.1.4), rms (version 6.7-1), corrplot (version 0.92), lmtest (version 0.9-40), and conflicted (version 1.2.0). Illustrations were created using ggplot2 (version 3.5.0), ggrepel (0.6.0), and sjPlot (2.8.17).
Luminex in vitro protein data
In vitro secretion of protein was done by co-culturing target cells (MonoMac1 or CD14+ cells isolated from frozen PBMCs) with cancer cells derived from tumor biopsies from the patients (MM1636.23 or MM1636.05, respectively) for 48 h (1:1 ratio, cultured in RPMI + 10 % fetal bovine serum). Then, T cells specific for IDO or PD-L1 derived from the corresponding patient (as described previously15) or an appropriate control (CD14-fraction from the CD14+ cell isolation from patient PBMCs) were added (1:10 ratio with target cells). Cells were cultured together for 24 h, supernatant was taken, and protein concentration was determined using the Bio-Plex Pro Human Cytokine Screening Panel, 48-Plex kit (cat. No. #12007283) from Bio-Rad according to the manufacturer’s instructions.
Statistical analyses
Overall survival (OS) and PFS were calculated as the time from the date of first treatment until the date of death/last seen alive and the date of progression/last follow-up without progression, respectively. Melanoma-specific survival (MSS) was calculated as the time from the date of first treatment until the date of death due to melanoma/last seen alive or death by other reason than melanoma. Overall response rate (ORR) was defined as the percentage of patients achieving PR or CR as the best overall response (BOR). Survival curves were generated with the Kaplan-Meier method, and Swimmer’s plots were designed in GraphPad Prism v. 5.01. The median follow-up time was calculated by the reverse Kaplan-Meier method.
Statistical analysis of Olink results was carried out in R using the stat_compare_means function from ggpubr when the T-test was used. Association between individual variables was assessed using univariate logistical regression by applying the glm function from base R. Changes in protein levels between time points were assessed using a paired T-test, and comparisons of the ratio of 6w and baseline between studies were done using a non-paired T-test. The test used is specified in the figure legends.
Identification of proteins associated with clinical outcome was done using elastic-net regression. Elastic-net regression was adopted due to the presence of multiple protein biomarkers that are correlated, and the high number of parameters compared to samples. Additionally, we aim to identify the biomarkers that are most predictive of our outcomes while accounting for the interdependence among other proteins in the analyses and relevant clinical covariates16. Briefly, elastic-net regression is a well-known shrinkage method (sparse selection), which is adopted to produce a more accurate model for complex data. Using elastic-net regression, we strived to obtain an optimally performing model, that is, to minimize both bias and variance. Elastic-net regression analysis was carried out using the glmnet function from the glmnet package. The difference between NPX values at 6w and baseline was calculated (6 W NPX value – baseline NPX value) and used as input together with age (above or below median), gender (male or female), and PD-L1 status (>1% or not). Outcome was PFS > median or PFS < median, and thus a logistic regression model was used for elastic-net regression.
Best α-value was chosen by using 10-fold cross-validation on a range of possible values (between 0 and 1, incrementing with 0.01) using the cv.glmnet function, and selecting the value for α providing the lowest error rate. The lambda.min value inherent to the optimal α-value was used. Seed was set at 42.
Resulting beta coefficients for selected variables were converted to odds ratios by taking the exponential function of the output values. The resulting odds ratio denotes an increase in odds ratio for achieving long PFS per unit of change in protein levels from baseline to 6w.
Proteins selected as predictive by elastic-net regression were used in downstream analysis. Singular predictive power was assessed by performing univariate logistic regression analysis, and changes in protein level by response group were assessed by calculating the change and comparing groups with a two-tailed T-test.
Total impact was calculated by multiplying the resulting odds ratios (per unit increase as described above) with observed change for each individual and taking the mean of the resulting total impact for each group, to incorporate both the effect size (odds ratio values obtained from elastic net) and the change in protein levels (as measured with Olink) into one number.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data generated in this study are available within the article and its supplementary data files. Additional raw data can be Access to raw data for any figure in the manuscript can be requested to the corresponding authors. All requests for data, including raw data and analyzed data, and materials will, within a reasonable time frame, be reviewed by the CCIT-DK office according to institutional policies and applicable laws. Any data and materials that can legally be shared will be released via data and material transfer agreements. Source data are provided with this paper.
Code availability
All code used to analyze data and generate figures is available upon request.
References
Wolchok, J. D. et al. Final, 10-year outcomes with nivolumab plus ipilimumab in advanced melanoma. N. Engl. J. Med. 392, 11–20 (2025).
Rohaan, M. W. et al. Tumor-infiltrating lymphocyte therapy or ipilimumab in advanced melanoma. N. Engl. J. Med. 387, 2113–2125 (2022).
Long, G. V. et al. Relatlimab and nivolumab versus nivolumab in previously untreated metastatic or unresectable melanoma: overall survival and response rates from RELATIVITY-047 (CA224-047. J. Clin. Oncol. 40, 360385 (2022).
Andersen, M. H. Anti-cancer immunotherapy: breakthroughs and future strategies. Semin. Immunopathol. 41, 1–3 (2019).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Andersen, M. H. The specific targeting of immune regulation: T-cell responses against Indoleamine 2,3-dioxygenase. Cancer Immunol. Immunother. 61, 1289–1297 (2012).
Sørensen, R. B. et al. Spontaneous cytotoxic T-Cell reactivity against indoleamine 2,3-dioxygenase-2. Cancer Res. 71, 2038–2044 (2011).
Dey, S. et al. Peptide vaccination directed against IDO1-expressing immune cells elicits CD8+and CD4+T-cell-mediated antitumor immunity and enhanced anti-PD1 responses. J. Immunother. Cancer 8, e000605 (2020).
Kjeldsen, J. W. et al. A phase 1/2 trial of an immune-modulatory vaccine against IDO/PD-L1 in combination with nivolumab in metastatic melanoma. Nat. Med. 27, 2212–2223 (2021).
Lorentzen, C. L., Kjeldsen, J. W., Ehrnrooth, E., Andersen, M. H. & Marie Svane, I. Long-term follow-up of anti-PD-1 naïve patients with metastatic melanoma treated with IDO/PD-L1 targeting peptide vaccine and nivolumab. J. Immunother. Cancer 11, e006755 (2023).
Marron, T. U. et al. Considerations for treatment duration in responders to immune checkpoint inhibitors. J. Immunother. Cancer 9, e001901 (2021).
Maloney, A. K. et al. Nivolumab maintenance improves overall survival of patients with advanced melanoma who experience severe immune-related adverse events on nivolumab plus ipilimumab. J. Immunother. Cancer 12, e009061 (2024).
Kverneland, A. H. et al. Supervised clustering of peripheral immune cells associated with clinical response to checkpoint inhibitor therapy in patients with advanced melanoma. Immunooncol. Technol. 20, 100396 (2023).
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net. J. R. Statist. Soc. B 67, 301–320 (2005).
Mortensen, R. E. J. et al. Pre-existing TGF-β-specific T-cell immunity in patients with pancreatic cancer predicts survival after checkpoint inhibitors combined with radiotherapy. J. Immunother. Cancer 11, e006432 (2023).
FDA-NIH Biomarker Working Group. BEST (Biomarkers, EndpointS, and other Tools) Resource (Food and Drug Administration (US) and National Institutes of Health (US), 2016).
Acknowledgements
The study was funded through a research funding agreement between IO Biotech and the CCIT-DK, Herlev Hospital, and the Oncology Department at Herlev Hospital.
Author information
Authors and Affiliations
Contributions
S.K.P. analyzed clinical data and wrote the manuscript. M.B. analyzed protein data and wrote the manuscript. E.M. carried out experiments. C.L.L. carried out the clinical trial. J.W.K. carried out the clinical trial. S.U.T. advised on data analysis methods. A.K. advised on data analysis methods. M.B.Z. advised on manuscript writing. M.H.A. supervised the project and revised the manuscript. I.M.S. supervised the project and revised the manuscript. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.B.Z. is a stakeholder at IO Biotech. M.H.A. is named as an inventor on various patent applications relating to therapeutic uses of IDO and PD-L1 peptides. These patent applications are assigned to the company IO Biotech, which is developing immune-modulating vaccines. M.H.A. is the founder, shareholder, and advisor for IO Biotech. E.M. is employed at IO Biotech. I.M.S. has received honoraria for consultancies and lectures from TILT Biotherapeutics, IO Biotech, Novartis, BMS, Genmab, MSD, Takeda, Sanofi Aventis, and Janssen Cilag; research grants from Evaxion Biotech, Adaptimmune, IO Biotech, Asgaard Biotech, TILT Biotherapeutics, and Enara Bio; and financial support for attending symposia from MSD. I.M.S. is the founder, shareholder, and advisor for IO Biotech. Other authors declare no conflict of interest.
Peer review
Peer review information
Nature Communications thanks Cong Zhou, Meghan Mooradian, Craig Slingluff, and the other anonymous reviewer for their contribution to the peer review of this work. [A peer review file is available.]
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pedersen, S., Byrdal, M., Martinenaite, E. et al. Five-year clinical outcome and immune biomarkers of durable response from the MM1636 trial on IDO/PD-L1 vaccination and PD-1 blockade in first line metastatic melanoma. Nat Commun 17, 806 (2026). https://doi.org/10.1038/s41467-025-67508-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67508-8