Abstract
Real or simulated microgravity induces a senescence-like modification of carotid artery in both human and animal observations, with the mechanisms not fully elucidated. Here, we aim to elucidate the role of sterol regulatory element-binding protein 1 (SREBP1, encoded by Srebf1) mediated lipogenesis in the process. Pharmacological activation of SREBP1 directly triggers senescence-like transformation in vascular smooth muscle cells (VSMC), while silencing Srebf1 exerts an opposite effect. Mechanistically, SREBP1-mediated lipogenesis upregulates acetyl-CoA pool to increase histone acetylation, modifying the chromatin accessibility which limiting recruitment of SRF/myocardin complexes to CArG boxes of contractile genes and opening the chromatin accessibility of aging genes. Srebf1 knockdown and local delivery of lentivirus or AAV-mediated VSMC specific expressing sh-Srebf1 significantly attenuates the senescence-like transformation of VSMC both in vitro and in vivo. Our findings reveal a previously unrecognized feature of SREBP1-mediated lipogenesis in vascular biology and SM-induced carotid artery remodeling.
Similar content being viewed by others
Introduction
Space exploration represents a frontier to understand how the human body adapts to extreme environments1,2. Among the various specialties, microgravity exerts a persistent and substantial influence on multiple organs and systems, especially in long-duration space missions3. Astronauts develop several geriatric-like issues, such as bone and muscle mass loss, immune dysfunction, and vascular remodeling, which are also demonstrated in human and animal models designed to simulate microgravity4,5. The carotid artery shows a high sensitivity to various unfavorable impacts6. In both real or simulated microgravity, the carotid artery exhibited a pronounced senescence-like remodeling, including decreased elasticity and increased carotid intima-media thickness (cIMT)2,5,7,8,9,10. Vascular smooth muscle cells (VSMC) are the primary cells that form the tunica media11. Differentiated VSMC exhibit a quiescent, contractile phenotype and express genes involved in contraction, such as smooth muscle actin (Acta2, encoding α-SMA), smooth muscle protein 22-α (Tagln, encoding SM22α), smooth muscle myosin heavy chain (Myh11, encoding MYH11), and calponin (Cnn1, encoding CNN1)12,13. With a notable plasticity, VSMC turn to be reversibly dedifferentiated and transformed to suit the vascular growth, remodeling, and repair upon environmental stimulates12,13,14,15. Dysregulated transformation of VSMC comes along with vascular pathologies in aging, atherosclerosis and aortic aneurysm16. For now, the characteristics and molecular mechanisms behind the phenotypic transformation of VSMC in simulated microgravity are still not fully clarified.
Lipids are key macromolecules engaged in biological activities from cell membrane construction, energy storage, to signaling transduction17. Recent studies indicate significant modification of lipid metabolism in senescence or age-related diseases18,19. Sterol regulatory element binding protein (SREBP, encoded by Srebf) acts as a key transcription factor (TF) controlling the expression of genes in charge of lipid production, degradation and conversion20,21. It has been well known that SREBP1 mainly contributes to fatty acid (FA) synthesis, while SREBP2 to the synthesis and uptake of cholesterol (CL)22. Recent studies unveiled the negative effects of SREBP cleavage-activating protein (SCAP, encoded by Scap) in atherosclerosis23,24. Some other genes, such as Fasn (encoding fatty acid synthase) and Hmgcr (encoding 3-hydroxy-3-methylglutaryl coenzyme A reductase), played a key role in VSMC related diseases25,26.
Lipid metabolites work through multiple downstream biological processes. Among them, epigenetic modifications of chromatin have provided novel insights to etiological exploration of diseases, especially those cannot be fully explained by genetic mechanisms. Recent studies revealed the connection between acetyl-CoA (Ac-CoA), a key metabolite coordinating energy metabolism of different pathways involving lipid homeostasis and chromatin adaptation27,28. For doing this, histone acetylation and deacetylation are modified, which regulate chromatin accessibility by facilitating and blocking TFs access, respectively. This would change the electrostatic interactions between DNA and the histones that assemble the nucleosome29. Accumulating evidence links histone acetylation to aging-related vascular disorders30. The increase in histone-H3 lysine-27 (H3K27) acetylation contributes to vascular senescence by opening up stiffness gene promoters31. Histone deacetylase inhibitors promote histone hyper-acetylation accelerating vascular calcification in preclinical studies32,33. Given the findings depicted above, it was hypothesized that the Ac-CoA pool harmonized the lipid homeostasis and gene expression in carotid artery, which regulated the VSMC phenotypic transformation and senescence-like remodeling in response to simulated microgravity.
In this work, we discover an accumulation of Ac-CoA and the following adjustment of SREBP1-mediated lipogenesis. Furthermore, an altered chromatin accessibility is demonstrated, playing a crucial role in the vascular aging process.
Results
Simulated microgravity leads to senescence-like transformation of VSMC in carotid artery
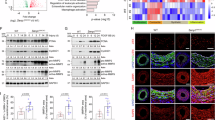
To efficiently simulate the effects of microgravity in vivo, the hindlimb unloaded tail suspended (HU) rat model was established as previously described (Supplementary Fig. S1A)34. Compared with control (CON) rats, 4 weeks of HU rats evinced classical manifestations of microgravity, including atrophy of soleus muscle fibers and natural body weight gain with reduced ratio of soleus to body weight, verifying the efficacy of the model (Supplementary Fig. S1B–E). Pathological staining and vascular ultrasound test revealed a senescence-like structural remodeling in carotid artery of HU rats, which involved a deranged media, broken elastin, and thickened cIMT, but normal lumen diameter (Fig. 1B–D and Supplementary Fig. S1F, G). We further confirmed the downregulated expression of VSMC’s contractile genes (Acta2, Tagln, Myh11, Cnn1), the upregulated expression of aging genes (Cdkn1a, Cdkn2a, Tp53) or senescence-associated secretory phenotype (SASP) genes (Il6, Ccl2) by RT-qPCR, western blotting, immunofluorescence, immunohistochemistry, and higher activity of β-Galactosidase in the carotid artery of HU rats (Fig. 1E–H and Supplementary Fig. S1H–L). Notably, the increased P21 (encoded by Cdkn1a) signals was mainly distributed in VSMC (marked by α-SMA) (Supplementary Fig. S1H, 1H).
A Schematic representation of the experimental protocol in rats. B Representative Masson trichrome staining (Masson’s) and Verhoeff’s Van Gieson staining (EVG) in the carotid artery. Lipid droplets are indicated by black arrows in Masson’s staining. Broken elastic fibers are indicated by red arrows in EVG staining. “L” indicates the side of the vascular lumen. Scale bar = 20 μm. C Representative ultrasound images in the right carotid artery of CON and HU rats. D Quantifications of carotid intima-media thickness (cIMT) of the right carotid artery from CON or HU rats. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t test (n = 6 rats per group). E Relative expression levels of VSMC contractile genes, aging genes, and SASP genes in the carotid artery from CON and HU rats. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t test (n = 3 rats per group). F Representative staining of β-Gal in the carotid artery from CON and HU rats. Scale bar = 2 mm. G Representative immunohistochemical staining of P21 in the carotid artery from CON and HU rats. “L” indicates the side of the vascular lumen. Scale bar = 20 μm. H Representative immunofluorescence images of P21 (red), CD31 (cyan), and α-SMA (green) in the carotid artery from CON and HU rats. Intensity traces (offset white lines) are plotted at right. “L” indicates the side of the vascular lumen. Scale bar = 200 μm or 100 μm (zoom in panel). I Relative expression levels of VSMC contractile genes, aging genes, and SASP genes in MOVAS after control (CTRL) or stimulated microgravity (SM) treatments. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t-test (n = 6 biological replicates). J Immunoblot analyses of α-SMA, SM22α, and P21, upon CTRL or SM treatment on MOVAS. Band quantification is normalized to the first lane. K Representative images of SPiDER-βGal staining (upper panel) or senescence β-Galactosidase staining (lower panel) of MOVAS after CTRL or SM treatment. Scale bar = 50 μm. L Representative FACS images of MOVAS after CTRL or SM treatment by SPiDER-βGal staining. M Representative images of the cell contraction assay of MOVAS after CTRL or SM treatment. Quantifications of cell contraction diameters are shown in the right panel. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t test (n = 5 biological replicates). N GSEA plots showing normalized enrichment scores (NES) and normalized P-value (calculated with an empirical phenotype-based permutation test) for multiple VSMC contractile fate pathways.
This observation prompted us to examine whether there was a similar change in simulated microgravity cell models. To simulate microgravity in vitro, we generated a 2-dimensional rotation (SM model) of mouse aortic smooth muscle cells (MOVAS) as previously described (Supplementary Fig. S2A, B)35. Firstly, the SM cell models exhibited a senescence-like transformation consistent with the HU model, including the downregulation of contractile genes and upregulation of aging or SASP genes (Fig. 1I, J). Moreover, as shown in Fig. 1K, L and Supplementary Fig. S2C, SM treated MOVAS displayed a higher intensity of SPiDER-βGal fluorescence, higher activity of β-Galactosidase, and bigger cell size. In addition, the contractility of MOVAS in the SM group was lower than in CTRL (Fig. 1M). Similarly, SM treated human aortic smooth muscle cells (HASMC) also exhibited senescence-like transformation (Supplementary Fig. S2D–F). However, human umbilical vein endothelial cells (HUVEC), a classic endothelial cell model, had no significant senescent signals following SM treatment (Supplementary Fig. S2G–I).
Moreover, the transcriptomic features of the carotid artery from the HU model (CON as control) and the SM cell model (CTRL as control) were analyzed by bulk RNA-seq. Among 5328 different expression genes (DEGs) of HU rats, 3916 were upregulated, while 1412 were downregulated (Supplementary Fig. S3A–C). Among 961 DEGs of SM cells, 530 were upregulated, while 431 were downregulated (Supplementary Fig. S3D–F). Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis revealed that the DEGs were enriched in lipid metabolism, cardiovascular disease, gene replication and repair, as well as cell growth and death in both HU or SM models (Supplementary Fig. S3G, H). Gene Ontology (GO) based Gene Set Enrichment Analysis (GSEA) demonstrated that the processes related to VSMC biology, such as differentiation, proliferation, and contraction, were significantly inhibited in HU or SM model (Fig. 1N). Moreover, compared with CTRL, SM treated MOVAS exhibited sparser cell density, larger cell size (Supplementary Fig. S2J), lower Ki-67 expression (a marker of cell proliferation) (Supplementary Fig. S2K), and higher apoptosis signaling (Supplementary Fig. S2L). Notably, cells that were shed following SM treatment exhibited stronger apoptotic signaling and higher expression of senescent marker genes (Supplementary Fig. S2M, N).
Together, these findings indicate that the SM leads to a senescence-like transformation of VSMC in the carotid artery.
Simulated microgravity induces lipid accumulation in VSMC
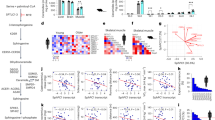
To further investigate the potential mechanisms of senescence-like transformation of VSMC caused by simulated microgravity, we performed combining analysis for the transcriptomes of HU and SM models. Based on KEGG-associated GSEA analysis, multi-pathways involved in lipid metabolism were enriched in the 47 co-activated pathways (NES > 1.0) both in HU and SM models, such as fatty acid metabolism, fatty acid biosynthesis, or steroid biosynthesis (Supplementary Fig. S4A and Fig. 2A). Multi-pathways involved in cytoskeleton and cell contraction were enriched in the 16 co-inhibited pathways (NES < − 1.0), such as TGF-beta signaling pathway, ECM-receptor, and vascular smooth muscle contraction (Supplementary Fig. S4B, C). In addition, heatmap results showed that both HU and SM models led to the activation of cholesterol and fatty acid synthesis pathways, which represented the key pathways for the intracellular lipid homeostasis (Fig. 2B). Further GO analysis indicated that DEGs were also enriched in multi-lipid metabolism-related pathways, both in HU and SM models (Supplementary Fig. S4D, E).
A KEGG based GSEA enrichment analyses for co-activated pathways of CON vs. HU and CTRL vs. SM. Normalized enrichment scores (NES) and normalized P-value (calculated with an empirical phenotype-based permutation test) are shown. B The key genes, regulatory processes (black arrows), and heatmap analysis (middle panel) involved in cholesterol synthesis (left panel) and fatty acid synthesis (right panel) pathways. The Log2-transformed fold change (FC) values are presented in parentheses. C Representative transmission electron microscopy (TEM) images of MOVAS after CTRL or SM treatment. Lipid droplets are indicated by red arrows. Scale bar = 2 μm. D Representative confocal microscopy images of MOVAS after CTRL or SM treatment by Nile-red staining. Scale bar = 50 μm. E Lipid level in MOVAS after CTRL or SM treatment by BODIPY staining. F–K Lipids (mg/g tissue, F), fatty acids (FA) (mg/g tissue, G), cholesteryl ester (ChE) (mg/g tissue, H), monoglycerides (MG) (μg/g tissue, I), diglyceride (DG) (mg/g tissue, J), triglycerides (TG) (mg/g tissue, K) levels in the carotid artery from CON or HU rats detected by lipidomics. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t test (n = 3-4 rats per group).
This observation prompted us to examine whether there was a dyshomeostasis of lipids in SM cells and HU rat models. Firstly, transmission electron microscopy (TEM) results disclosed an accumulated lipid droplets in SM MOVAS (Fig. 2C). This was then confirmed by the staining of Nile Red and BODIPY, two lipophilic fluorescent probes (Fig. 2D, E). Furthermore, rat primary carotid artery smooth muscle cells (RCA-SMCs) were isolated to elucidate whether lipid accumulation remained in SM treated primary cells (Supplementary Fig. S5A, B). Consistent with MOVAS, SM treatment significantly increased the content of fatty acids (FA) and triglyceride (TG), and leading to lipid accumulation in RCA-SMCs (Supplementary Fig. S5C–E). SM-treated HASMC also exhibited lipid accumulation (Supplementary Fig. S5F). In addition, lipidomics was performed to profile the lipid state of the carotid artery (Supplementary Fig. S6 and Supplementary Data 1). We found an abundant distribution of lipids in the carotid artery of rats (Supplementary Fig. S6A). Among the 21 classes (1097 species), TG, FA, and phosphatidylcholine (PC) were the top three in the carotid artery (> 1 mg/g) (Figure S6B). As compared with CON, HU rats showed a higher lipid content of multi-lipid species in carotid artery (Fig. 2F and Supplementary Fig. S6C, D), including FA, cholesterol ester (ChE), monoglycerides (MG), diglycerides (DG), and TG (Fig. 2G–K).
Taken together, these data demonstrate that the SM induces lipid accumulation in VSMC of the carotid artery.
Simulated microgravity activates SREBPs mediated lipogenesis
As a TF, SREBPs activates the expression of numerous genes essential for lipid homeostasis20. To explore its role in lipid accumulation triggered by SM, we performed a GSEA based activated TF enrichment analysis (NES > 1, normalized P < 0.05) for the transcriptomics of HU and SM models. As shown in Fig. 3A, 75 and 102 TFs were significantly enriched in HU and SM models, respectively. There were 21 of these TFs co-enriched in both HU and SM models. Intriguingly, SREBP1 (a crucial TF in lipid biogenesis) was listed among these 21 co-enriched TFs (Fig. 3A). GSEA results indicated the activation of SREBP1 in both HU and SM models (Fig. 3B). Because of the absence of SREBP2 in candidate TFs, it was not listed among the co-enriched 21 TFs. However, as SREBP2 shares a similar DNA-binding motif with SREBP1, it might also be activated during SM (Supplementary Fig. S7A, B).
A GSEA based transcription factors (TFs) enrichment analysis for the activated TFs of two RNA-seq datasets (CON vs. HU and CTRL vs. SM) is shown above. Details of the co-activated TFs (21 in total) are listed below. SREBP1 is highlighted. B GSEA plots showing normalized enrichment scores (NES) and normalized P-value (calculated with an empirical phenotype-based permutation test) for SREBP1 target genes from RNA-seq datasets (CON vs. HU and CTRL vs. SM). C Representative immunohistochemical staining of SREBP1 or SREBP2 in the carotid artery from CON and HU rats. “L” indicates the side of the vascular lumen. Scale bar = 20 μm. D Representative immunofluorescence images of SREBP1 (red), CD31 (cyan), and α-SMA (green) in the carotid artery from CON and HU rats. Intensity traces (offset white lines) are plotted at right. “L” indicates the side of the vascular lumen. Scale bar = 200 μm or 100 μm (zoom in panel). E Immunoblot analyses of precursor-SREBP1/2 (p-SREBP1/2), and nucleic-SREBP1/2 (n-SREBP1/2), from the carotid artery of rats treated with CON or HU. Band quantification is normalized to the first lane. The non-specific bands are marked by “*”. F Relative expression levels of genes involved in SREBP1/2 activation (Srebf1, Srebf2, Scap) and target genes (Acaca and Fasn for SREBP1; Hmgcr and Ldlr for SREBP2) in the carotid artery from CON and HU rats. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t test (n = 3 rats per group). G Immunoblot analyses of precursor-SREBP1/2 (p-SREBP1/2), and nucleic-SREBP1/2 (n-SREBP1/2) of MOVAS treated with CTRL or SM. Band quantification is normalized to the first lane. H Representative confocal microscopy images of SREBP1 (green) and SREBP2 (red) of MOVAS treated with CTRL or SM. Scale bar = 20 μm. I Relative expression levels of genes involved in SREBP1/2 activation (Srebf1, Srebf2, Scap) and target genes (Acaca and Fasn for SREBP1; Hmgcr and Ldlr for SREBP2) of MOVAS treated with CTRL or SM. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t test (n = 6 biological replicates).
Then, we confirmed the upregulated expression of SREBP1 and SREBP2 by immunohistochemistry and immunofluorescence in the carotid artery of HU rats (Fig. 3C and Supplementary Fig. S7C). Notably, the increased SREBP1 signals was mainly distributed in VSMC (marked by α-SMA) (Supplementary Fig. S1H and Fig. 3D). Western blot further demonstrated the increased protein expression of precursor-SREBPs (p-SREBPs) and nucleic-SREBPs (n-SREBPs) in HU rats compared with CON (Fig. 3E). Using RT-qPCR assays, the upregulation of Srebfs (SREBPs encoding genes), Scap (activator of SREBPs), and representative targets of SREBPs was discovered, strongly indicating the activation of SREBPs in HU rats (Fig. 3F). In SM-treated VSMC models, protein levels of p-SREBPs and n-SREBPs were also increased (Fig. 3G and Supplementary Fig. S7D). In addition, the increasing levels of nuclear-distributed SREBPs and transcripts of SREBPs target genes also demonstrated the activation of SREBPs in SM treated MOVAS or HASMC (Fig. 3H, I and Supplementary Fig. S7E, F). However, HUVEC showed no significant changes of SREBPs after SM treatment (Supplementary Fig. S7G). Given to the well-established crucial role of mTOR signaling in activating SREBP1, mechanotransduction, and cell senescence20,36,37,38,39,40, we further confirmed the activation of the mTOR pathway, which was demonstrated by the upregulation of p-mTOR/mTOR and p-AKT/AKT (key kinases bridging mechanical signaling to mTOR activation).
All these results indicate that the activation of SREBPs contributes to the lipid accumulation of the carotid artery and VSMC induced by SM.
SREBP1 directly promotes the senescence-like transformation of VSMC by activating lipogenesis
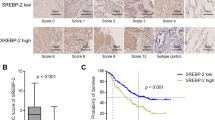
Given the nonidentical roles of SREBP1 and SREBP2 in lipid homeostatic regulation, we tried to examine the potential differences in their contribution to VSMC biology. We firstly analyzed the expression correlation between the SREBPs target genes and contractile (or aging) genes in the transcriptome dataset of human carotid artery from atherosclerotic and healthy individuals (GSE100927)41. As shown in Fig. 4A, SREBPs-targets were positively correlated with aging genes and negatively correlated with VSMC contractile genes. Intriguingly, these correlations were mainly attributed to SREBP1-targets but less to SREBP2-targets (Supplementary Fig. S8A–C). To further elucidate whether such correlations exist in VSMC at the single-cell level, we analyzed the average expression of VSMC contractile, aging, and SREBP1-related genes across different subgroups of VSMC in the carotid arteries from atherosclerosis patients (GSE253903) (Supplementary Fig. S8D–G). Intriguingly, the VSMC subgroup with higher expression of contractile genes exhibited lower expression of SREBP1-related and aging genes. In contrast, the VSMC subgroups with lower expression of contractile marker genes showed higher expression of SREBP1-related and aging genes (Fig. 4B). In addition, H2O2, a canonical senescence-inducer, could directly upregulate the expression of SREBP1 and its target genes, while also induce intracellular lipid accumulation in VSMC (Supplementary Fig. S9A–F), which was consistent with previous studies42.
A Correlation between the expressions of SREBPs target genes and VSMC contractile genes (upper left panel), SREBPs target genes and aging genes (lower left panel), SREBP1 target genes and VSMC contractile genes (upper right panel), and SREBP1 target genes and aging genes (lower right panel) in the carotid artery from healthy and atherosclerosis patients (GSE100927). Correlation and P value (two-tailed) are based on Pearson’s correlation. B Heatmap depicting the average expression level of VSMC contractile genes, SREBP1 related genes, aging genes, and fibroblast genes of each VSMC or fibroblast subgroups in the carotid artery from atherosclerosis patients (single-cell transcriptomic, GSE253903). C Relative mRNA expression levels of genes involved in the maintain of VSMC contractile phenotype of MOVAS transfected with siCtrl, si-Srebf1#3, or si-Srebf2#3. Data are presented as mean ± SD. Statistical significance is calculated with a one-way ANOVA followed by Dunnett’s multiple comparisons test (n = 6 biological replicates). D Representative images of cell contraction assay of MOVAS transfected with siCtrl, si-Srebf1#3, or si-Srebf2#3. E Schematic diagram of the agonistic target of LXR-623. F Immunoblot analyses of precursor-SREBP1/2 (p-SREBP1/2), and nucleic-SREBP1/2 (n-SREBP1/2) of MOVAS after DMSO or LXR-623 treatment. Band quantification is normalized to the first lane. G GSEA based transcription factors (TFs) enrichment analysis for the activated TFs of three RNA-seq datasets (CON vs. HU, CTRL vs. SM, and DMSO vs. LXR-623) is shown above. Details of the co-activated TFs (3 in total) are listed below. SREBP1 is highlighted. H GSEA plots showing normalized enrichment scores (NES) and normalized P-value (calculated with an empirical phenotype-based permutation test) for SREBP1 target genes from RNA-Seq (DMSO vs. LXR-623). I The key genes, regulatory processes (black arrows), and heatmap analysis (middle panel) involved in cholesterol synthesis (left panel) and fatty acid synthesis (right panel) pathways. The Log2-transformed fold change (FC) values are presented in parentheses. J Lipid level in MOVAS after DMSO or LXR-623 treatment by BODIPY staining. K Representative transmission electron microscopy images of MOVAS treated with DMSO or LXR-623. Lipid droplets are indicated by red arrows. Scale bar = 2 μm (left panel), 1 μm (zoom-in panel). L Representative images of the cell contraction assay of MOVAS treated with DMSO or LXR-623. Quantifications of cell contraction diameters are shown in the lower panel. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t test (n = 5 biological replicates). M Immunoblot analyses of MYOCD, α-SMA, and SM22α of MOVAS treated with DMSO or LXR-623. Band quantification is normalized to the first lane. The non-specific bands are marked by “*”. N GSEA plots showing normalized enrichment scores (NES) and normalized P-value (calculated with an empirical phenotype-based permutation test) for VSMC contraction from RNA-Seq (DMSO vs. LXR-623). O Representative FACS images of MOVAS treated with DMSO or LXR-623 by SPiDER-βGal staining. P Representative images of β-Gal staining of MOVAS treated with H2O2 (positive control), DMSO or LXR-623. Scale bar = 200 μm.
Then, we generated Srebf1 (or Srebf2) knockdown MOVAS by siRNA transient transfection (Supplementary Fig. S10A–D). Surprisingly, compared with siCtrl, si-Srebf1 exhibited greater contractile function, which was reflected in higher expression of contractile genes and stronger contractility in MOVAS (Fig. 4C, D and Supplementary Fig. S10E, F). In contrast, si-Srebf2 exhibited lesser contractile function, which was reflected in lower expression of contractile genes and weaker contractility in MOVAS (Fig. 4C, D and Supplementary Fig. S10E, F). To determine whether SREBP1 activation was directly responsible for the senescence-like transformation of VSMC, we performed a targeted agonist (LXR-623) treatment to activate SREBP1 in MOVAS or RCA-SMCs (Fig. 4E). Consistent with previous reports, we found that LXR-623 activated SREBP1 but not SREBP2, demonstrated by increased p-SREBP1, n-SREBP1, as well as its target genes’ expression both in MOVAS, RCA-SMCs, and HASMC (Fig. 4F and Supplementary Fig. S11A–G). We further performed RNA-seq and found 1267 genes were upregulated, while 2278 genes were downregulated in LXR-623 relative to DMSO treated MOVAS (Supplementary Fig. S12A–C). KEGG analysis indicated that the DEGs were enriched in lipid metabolism, cardiovascular disease, gene replication and repair, as well as cell growth and death, which was consistent with HU or SM models (Supplementary Figs. S12D, S3G, H). Notably, as shown in Fig. 4G, H, SREBP1 was significantly enriched as one of the co-activated TFs in all three transcriptomes (CON vs. HU; CTRL vs. SM; DMSO vs. LXR-623). GO analysis further verified the enrichment of DEGs in pathways related to lipid biosynthetic and metabolic processes (Supplementary Fig. S12E). Meanwhile, the heatmap revealed that LXR-623 treatment directly activated FA synthesis but not CL metabolism, suggesting the SREBP1 but not SREBP2 being the target of LXR-623 (Fig. 4I). In addition, MOVAS exhibited lipid accumulation after LXR-623 treatment, demonstrated by the staining of BODIPY (Fig. 4J). These results were further supported by the observation of extensive lipid droplets in LXR-623 treated MOVAS by TEM (Fig. 4K). The LXR-623 induced lipid accumulation was also confirmed in HASMC (Supplementary Fig. S11H).
We next determined the pathological role of LXR-623 for VSMC biology. Compared with DMSO, LXR-623 treated VSMC showed weaker contractility and lower expression of contractile genes (Fig. 4L, M and Supplementary Fig.S11I–L). GO based GSEA results demonstrated that vascular associated smooth muscle cell contraction and differentiation were depressed by LXR-623 (Fig. 4N and Supplementary Fig. S12F). Moreover, LXR-623 directly triggered cell senescence, which was indicated by a higher intensity of SPiDER-βGal fluorescence, higher activity of β-galactosidase, and upregulation of aging and SASP genes (Fig. 4O, P and Supplementary Fig. S11M–O). Furthermore, mTOR signaling was also activated by LXR-623 treatment, which was consistent with SM treatment (Supplementary Figs. S11P, Q, S7G, H).
Taken together, these data reveal that the targeted activation of SREBP1 directly promotes VSMC’ senescence-like transformation.
Upregulated Ac-CoA pool contributes to SREBP1-activation triggered senescence-like transformation of VSMC
It has been well established that the Ac-CoA pool takes a central role in energy production, lipid metabolism, and gene expression (Fig. 5A)27. Therefore, we investigated whether SREBP1 activation disturbed the balance of the Ac-CoA pool in VSMC. We observed that the Ac-CoA levels were significantly enhanced in MOVAS after both LXR-623 and SM treatments, with the citrate, a key substate of endogenous Ac-CoA synthesis, being significantly decreased (Fig. 5B and Supplementary Fig. S13A). Notably, the increased Ac-CoA levels were predominantly localized in the cytoplasm and nucleus, while no significant changes were observed in the mitochondrial Ac-CoA levels (Supplementary Fig. S13B, C).
A Schematic diagram of Ac-CoA metabolism in mammalian cells. B Relative levels of Ac-CoA and citrate in MOVAS treated with DMSO or LXR-623. Data are presented as mean ± SD. Statistical significance is calculated with a two-tailed unpaired t test (n = 6 biological replicates). C Immunoblot analyses of pan-acetylation (Pan-Ac) in MOVAS treated with DMSO or LXR-623. D Representative confocal microscopy images of Pan-Ac (red) in MOVAS treated with DMSO or LXR-623. Scale bar = 50 μm. E The Venn diagram of acetylated proteins (identified by IP-MS) in MOVAS treated with DMSO or LXR-623. F Heatmap of acetylated proteins (identified by IP-MS) in MOVAS treated with DMSO or LXR-623. G Gene ontology analysis of the acetylated proteins which have higher signals in the LXR-623 group compared with the DMSO group. Statistical significance is calculated with a hypergeometric test. H Immunoblot analyses of H3K9Ac, H3K18Ac, and H3K27Ac in MOVAS treated with DMSO or LXR-623. Band quantification is normalized to the first lane. I Schematic diagram of the regulation of sodium acetate (NaAc), Ac-CoA pool, protein acetylation, and deacetylase inhibitor cocktail (DaIC). J Relative mRNA expression levels of Acta2 and Cdkn1a in MOVAS treated with DMSO + Vehicle, DMSO + NaAc (120 mM), LXR-623 + Vehicle, LXR-623 + NaAc (120 mM) (48 h). Data are presented as mean ± SD. Statistical significance is calculated with a two-way ANOVA followed by Tukey’s multiple comparisons test (n = 6 biological replicates). K Immunoblot analyses of α-SMA, SM22α, p21, H3K9Ac, and H3K27Ac in MOVAS treated with DMSO + Vehicle, DMSO + NaAc (120 mM), LXR-623 + Vehicle, LXR-623 + NaAc (120 mM) (48 h). Band quantification is normalized to the first lane. L Relative mRNA expression levels of Acta2 and Cdkn1a in MOVAS treated with DMSO + Vehicle, DMSO + DaIC (1:100), LXR-623 + Vehicle, LXR-623 + DaIC (1:100) (48 h). Data are presented as mean ± SD. Statistical significance is calculated with a two-way ANOVA followed by Tukey’s multiple comparisons test (n = 6 biological replicates). M Immunoblot analyses of α-SMA, p21, H3K9Ac, and H3K27Ac in MOVAS treated with DMSO + Vehicle, DMSO + DaIC (1:100), LXR-623 + Vehicle, LXR-623 + DaIC (1:100) (48 h). Band quantification is normalized to the first lane.
Cellular and nuclear Ac-CoA pool has been reported manipulating the regulation of gene expression by protein’s acetylation, particularly for histones27. In the present study, cellular protein acetylation was amplified substantially by LXR-623 or SM treatment (Fig. 5C, D and Supplementary Fig. S13D). We then performed IP-MS to characterize the acetylated protein profile (anti-Pan-acetylation) in LXR-623 or SM treated MOVAS (Fig. 5E, F and Supplementary Fig. S13E, F). Among the 101 (122 in SM) identified proteins, the 60 proteins in the LXR-623 group (105 in SM group) exhibited hyperacetylation. And these hyperacetylated proteins were significantly enriched in the regulation of gene expression terms, such as nucleosome, chromatin DNA binding, and nucleosome assembly (Fig. 5G, Supplementary Fig. S13G and Supplementary Data 2, 3). These pathways are intimately linked to histone H3 hyperacetylation, which was further confirmed by the increased level of H3K9Ac, H3K18Ac, and H3K27Ac (representative acetylated histone sites) in LXR-623 or SM treated group, compared with DMSO or CTRL (Fig. 5H, Supplementary Fig. S13H–J). Similarly, the increased levels of H3K27Ac were also confirmed in VSMC of HU rat model (Supplementary Fig. S13K).
To determine whether the increased Ac-CoA pool directly plays a role in senescence-like transformation of VSMC, we employed sodium acetate (NaAc, an exogenous Ac-CoA supplements) and Deacetylase Inhibitor Cocktail (DaIC, indirectly increasing endogenous protein acetylation) to the MOVAS (Fig. 5I). As expected, both the Ac-CoA levels and acetylated protein abundance were elevated by these two supplements (Supplementary Fig. S14A–D). In addition, the Acta2 expression was downregulated, and the Cdkn1a was upregulated significantly. The senescence-like transformation of VSMC induced by LXR-623 was further exacerbated upon treatment with NaAc or DaIC, characterized by increased acetylation of H3 and upregulation of P21, while the contractile phenotype was suppressed (Fig. 5J–M, Supplementary Fig. S14F, G). In addition, supplementation with NaAc partially offset the effects of Srebf1 knockdown, including the mitigation of intracellular lipids, H3 acetylation, and the expression of Acta2 and Cdkn1a (Supplementary Fig. S14H–J).
This observation prompted us to examine whether Acly (encoding ATP-citrate lyase) and Acss2 (encoding Ac-CoA synthetase 2) were upregulated, which are two targets of SREBP1 and key enzymes catalyzing the Ac-CoA biosynthesis from endogenous citrate (ACLY) and from exogenous acetate (ACSS2). Intriguingly, Acly and Acss2 were both upregulated in HU rats, SM or LXR-623 treated VSMC (Supplementary Fig. S15A–C). However, the expression of Acss1, which catalyzes the synthesis of mitochondrial Ac-CoA, exhibited no significant changes, in line with the stable levels of mitochondrial Ac-CoA (Supplementary Figs. S15D, E, S13B, C). We further generated Acly or Acss2 knockdown MOVAS by siRNA transient transfection to determine whether ACLY or ACSS2 were involved in the senescence-like transformation mediated by SREBP1 activation induced by SM (Supplementary Fig. S15F–I). Notably, both Acly or Acss2 knockdown significantly alleviated SM-induced lipid accumulation and cellular senescence in VSMC, which indicated by lower signals of BODIPY, SPiDER-βGal, and P21 expression (Supplementary Fig. S15J–L).
Together, these results demonstrate that the upregulated Ac-CoA pool contributes to the SREBP1-activation triggered senescence-like transformation of VSMC.
Upregulated Ac-CoA pool accelerates the phenotypic transformation of VSMC by reshaping chromatin accessibility
Since SREBP1 activation upregulated the intracellular Ac-CoA pool and resulted in an increase of histone acetylation, we next examined whether chromatin accessibility was involved in senescence-like transformation of MOVAS. We performed ATAC-Seq by Tn5 transposase in MOVAS treated with DMSO, LXR-623, DMSO+DaIC, or LXR-623+DaIC (Fig. 6A and Supplementary Fig. S16A). Totally, 69795 peaks were identified which lying across promoter (32.5%), exon (2.8%), intron (32.67%), 5’UTR (0.08%), 3’UTR (1.2%), downstream (0.15%), and distal intergenic (30.6%) (Supplementary Fig. S16B).
A The enrichment of ATAC-seq reads around the Transcription Start Site (TSS). B. Heatmap of ATAC-seq marker peaks across all clusters identified with bias-matched differential testing (n = 3 per group). KEGG analysis of marker peaks across clusters are shown in right panel. The box represents the 25th to 75th percentiles, the whiskers extend to 1.5 times the interquartile range, and the center line indicates the median. C, D GO analysis of marker peaks across Cluster-5 (C) and Cluster-6 (D). Statistical significance is calculated with a hypergeometric test. E Heatmap depicting the average signal intensity of VSMC contractile genes, SREBP1 related genes, and aging genes across different groups in ATAC-seq analysis. F Genome browser of the Acta2 gene (upper panel), Cdkn1a gene (lower panel) in different groups. G Venn plot of TFs enrichment analysis for the cluster-5 genes of ATAC-seq and downregulated DEGs of RNA-seq (DMSO vs. LXR-623). Details of the same transcription factors (3 in total) are listed on the right. H GSEA plots showing normalized enrichment scores (NES) and normalized P-value (calculated with an empirical phenotype-based permutation test) for SRF target genes from RNA-Seq (DMSO vs. LXR-623). I GSEA plots showing normalized enrichment scores (NES) and normalized P-value (calculated with an empirical phenotype-based permutation test) for SRF target genes from RNA-Seq (CON vs. HU and CTRL vs. SM). J Immunoblot analyses of MYOCD and SRF of cytoplasmic (Cyto) and nuclear (nuc) lysates. K Transcriptomic based network Venn plot of SRF target genes’ expression in MOVAS treated with DMSO or LXR-623. Upregulated different expression genes (Up DEGs) are highlighted by red circles. Downregulated different expression genes (Down DEGs) are highlighted by bule circles.
We further divided the genes into eight clusters (cluster 1-8) according to the signals of promoter peaks (22710 peaks of 17703 genes) in each group (Fig. 6B and Supplementary Fig. S16C). Gene sets of cluster-5, −6, and −8 attracted our attention. Cluster-5 represented genes with condensed chromatin accessibility after LXR-623 treatment, which was more condensed after DaIC supplement. Based on KEGG analysis, cluster-5 genes were enriched in pathways of cytoskeleton, including MAPK signaling pathway, cytoskeleton in muscle cells, and regulation of actin cytoskeleton (Fig. 6B). Further GO analysis indicated that cluster-5 were involved in vascular remodeling (angiogenesis; artery development; etc), VSMC dedifferentiation (muscle cell differentiation; positive regulation of cell motility; and migration; etc), and cytoskeleton (regulation of Rho protein signal transduction; etc) (Fig. 6C). Conversely, cluster-6 and −8 represented genes with opened chromatin accessibility after LXR-623 treatment which was more opened after DaIC supplement (Fig. 6B). Based on KEGG and GO analysis, these two clusters were significantly enriched in pathways involving epigenetic remodeling (ATP-dependent chromatin remodeling, chromatin remodeling, etc), aging (cellular senescence, cell cycle, apoptosis, etc), and lipid metabolism (lipid and atherosclerosis, lipid biosynthetic process, acyltransferase activity, etc) (Fig. 6B, D and Supplementary Fig. S16D). Intriguingly, addition of the LXR-623 or DaIC downregulated the chromatin accessibility of key contractile genes, while upregulating the chromatin accessibility of SREBP1-related genes and aging genes (Fig. 6E). For Acta2, the chromatin accessibility was partially blocked by LXR-623 or DaIC. A synergistic effect was uncovered if supplying DaIC after LXR-623 treatment (Fig. 6F). In contrary, the chromatin accessibility of Cdkn1a was opened after treating with LXR-623 or DaIC. A similar synergistic effect was also present when the two supplements were employed (Fig. 6F).
To gain insight into the underlying mechanisms of VSMC’ dedifferentiation triggered by the upregulated SREBP1 activation, we performed TFs enrichment analysis and compared it with that of RNA-seq (Fig. 6G). TFs enrichment analysis suggested that serum response factor (SRF), as one of the three hub TFs, might be involved in the contractile fate inhibition of VSMC (Supplementary Fig. S16E). Based on GSEA analysis, SRF was also the hub TFs contributing to the genes’ downregulation of three transcriptomics performed in this study (CON vs. HU; CTRL vs. SM; DMSO vs. LXR-623) (Supplementary Fig. S16E and Fig. 6H, I). Further results indicated that LXR-623 or SM treatments inhibited the nuclear translocation of SRF and MYOCD (a transcriptional coordinator of SRF), which might contribute to the functional inhibition of SRF (Fig. 6J and Supplementary Fig. S16F, G). In addition, the downregulated SRF targets were mainly enriched in actin cytoskeleton organization, muscle structure development, and response to mechanical stimulus, which contributed to the inhibition of VSMC’ differentiated fate caused by SREBP1 activation and simulated microgravity (Fig. 6K and Supplementary Fig. S16H).
Taken together, these data reveal that upregulated Ac-CoA pool triggers VSMC’ senescence-like transformation by reshaping chromatin accessibility.
Knockdown of Srebf1 alleviates senescence-like transformation of VSMC induced by simulated microgravity
As the SM upregulated the Ac-CoA pool and reshaped chromatin accessibility by activating SREBP1, we investigated whether downregulating Srebf1 using lentivirus (Lv) exerted protection on VSMC senescence and vascular remodeling (Fig. 7A, G). First, we assessed the effects of Lv-mediated Srebf1 knockdown (Lv-sh-Srebf1) on RCA-SMCs. Lv-sh-Srebf1 treatment significantly downregulated the expression of Srebf1, Acly, Acss2, and aging genes, as well as the protein levels of ACLY, ACSS2, P21, and H3 acetylation (Supplementary Fig. S17A–D). Knockdown of Srebf1 alleviated the upregulation of Ac-CoA levels, Srebf1, Acly, and Acss2 genes triggered by SM (Fig. 7B, C). In addition, Srebf1 knockdown partially alleviated the senescence-like transformation of RCA-SMCs due to SM treatment, which was manifested in the restoration of the cell contractility and expression of H3 acetylation, P21, and α-SMA (Fig. 7D–F).
A Schematic diagram of the rescue study of the SM cell model. B Relative level of Ac-CoA in RCA-SMCs treated with Lv-shCtrl + CTRL, Lv-shCtrl + SM, Lv-sh-Srebf1 + CTRL, and Lv-sh-Srebf1 + SM. Data are presented as mean ± SD. Statistical significance is calculated with a two-way ANOVA followed by the Tukey’s multiple comparisons test (n = 6 biological replicates). C Relative mRNA expression levels of Srebf1, Acly, and Acss2 in RCA-SMCs treated with Lv-shCtrl+CTRL, Lv-shCtrl + SM, Lv-sh-Srebf1 + CTRL, and Lv-sh-Srebf1 + SM. Data are presented as mean ± SD. Statistical significance is calculated with a two-way ANOVA followed by the Tukey’s multiple comparisons test (n = 6 biological replicates). D Representative images of cell contraction assay of RCA-SMCs treated with Lv-shCtrl + CTRL, Lv-shCtrl + SM, Lv-sh-Srebf1 + CTRL, and Lv-sh-Srebf1 + SM. E Quantifications of the contraction distance (cm) in D Data are presented as mean ± SD. Statistical significance is calculated with a two-way ANOVA followed by the Tukey’s multiple comparisons test (n = 4 biological replicates). F Immunoblot analyses of α-SMA, P21, H3K9Ac, H3K18Ac, H3K27Ac of RCA-SMCs treated with Lv-shCtrl + CTRL, Lv-shCtrl + SM, Lv-sh-Srebf1 + CTRL, and Lv-sh-Srebf1 + SM. Band quantification is normalized to the first lane. G Schematic diagram of the rescue study of the HU rat model. H The gelation of the laponite hydrogels in vitro. I Representative ultrasound images in the carotid artery of rats treated with Lv-shCtrl + CON, Lv-shCtrl + HU, Lv-sh-Srebf1 + CON, and Lv-sh-Srebf1 + HU. (J). Quantifications of carotid intima-media thickness (cIMT) of right carotid artery of rats treated with Lv-shCtrl + CON (n = 6 rats), Lv-shCtrl + HU (n = 5 rats), Lv-sh-Srebf1 + CON (n = 4 rats), and Lv-sh-Srebf1 + HU (n = 4 rats). Data are presented as mean ± SD. Statistical significance is calculated with a two-way ANOVA followed by the Tukey’s multiple comparisons test (). K Representative Verhoeff’s Van Gieson staining (EVG) in the carotid artery of rats treated with Lv-shCtrl + CON, Lv-shCtrl + HU, Lv-sh-Srebf1 + CON, and Lv-sh-Srebf1 + HU. Broken elastic fibers are indicated by red arrows in EVG staining. “L” indicates the side of vascular lumen. Scale bar = 20 μm. L Relative level of Ac-CoA in carotid artery of rats treated with Lv-shCtrl + CON (n = 6 rats), Lv-shCtrl + HU (n = 5 rats), Lv-sh-Srebf1 + CON (n = 4 rats), and Lv-sh-Srebf1 + HU (n = 4 rats). Data are presented as mean ± SD. Statistical significance is calculated with a two-way ANOVA followed by the Tukey’s multiple comparisons test. M Relative mRNA expression levels of Srebf1, Acta2, Tagln, Cdkn1a, and Tp53 in carotid artery of rats treated with Lv-shCtrl+CON, Lv-shCtrl+HU, Lv-sh-Srebf1 + CON, and Lv-sh-Srebf1 + HU. Data are presented as mean ± SD. Statistical significance is calculated with a two-way ANOVA followed by the Tukey’s multiple comparisons test (n = 4 rats per group). N Representative immunofluorescence images of Pan-Ac (red), and P21 (green) in carotid artery of rats treated with Lv-shCtrl + CON, Lv-shCtrl + HU, Lv-sh-Srebf1 + CON, and Lv-sh-Srebf1 + HU. Scale bar = 200 μm or 100 μm (zoom in panel).
To achieve local intervention of SREBP1 in carotid artery, we used Lv-sh-Srebf1 loaded by Laponite hydrogels (Fig. 7G, H)43. As shown in Supplementary Fig. S18A–D, Laponite hydrogels mixed with Lv effectively introduced the target sequences into VSMC of the carotid artery. Further ultrasound analysis revealed that Lv-sh-Srebf1 mitigated the HU-induced cIMT thickening but not affecting the diameter of the lumen (Fig. 7I, J and Supplementary Fig. S18E, F). Srebf1 knockdown alleviated HU-induced senescence-like media derangement and elastin break (Fig. 7K). Notably, the Ac-CoA levels were significantly increased in the carotid artery of HU rats compared with CON, which was partially reduced by Lv-sh-Srebf1 transfection, which was consistent with RCA-SMCs (Fig. 7L). In addition, in line with attenuated structural remodeling, Lv-sh-Srebf1 treatment partially restored the disordered transcription of Srebf1, Acta2, Tagln, Cdkn1a, and Tp53 as well as the expression of P21 and Pan-Ac in the carotid artery treated with HU (Fig. 7M, N). To achieve local VSMC-specific intervention of SREBP1 in mice CA, we used AAV-Tagln-sh-Srebf1 loaded by Laponite hydrogels (Supplementary Fig. S19A, B). As shown in Supplementary Fig. S19C, Laponite hydrogels mixed with AAV effectively introduced the target sequences into VSMC of the mice carotid artery. VSMC-specific Srebf1 knockdown alleviated HU-induced senescence-like derangement and elastin break of media (Supplementary Fig. S19D). Further ultrasound analysis revealed that AAV-Tagln-sh-Srebf1 mitigated the HU-induced cIMT thickening but not affecting the diameter of lumen (Supplementary Fig. S19E–H). Notably, VSMC-specific Srebf1 knockdown also partially recovered the disordered expression of SREBP1 related genes, aging genes, and contractile genes (Supplementary Fig. S19I–L) in HU mice carotid artery, which was consistent with Lv-sh-Srebf1-treated rats.
Overall, these findings suggest that Srebf1 knockdown imposed functional and morphological benefits on SM induced senescence-like remodeling of carotid artery, highlighting the pathological essence and potential therapeutic value of SREBP1.
Discussion
SREBP1-mediated lipogenesis contributes to multiple aging-related diseases. However, it remains unknown whether it plays a role in VSMC senescence and microgravity-caused vascular remodeling. Our study revealed that SREBP1 activation induced lipogenesis, triggered VSMC senescence, and promoted carotid artery adaptation to SM. These observations were based on studies using HU rats and two-dimensional rotation VSMC models. We also demonstrated that LXR-623, as a well-established agonist of SREBP1, actuated senescence-like transformation in VSMC, while silencing SREBP1 exerts the opposite effects. Mechanistically, we found that SREBP1 upregulates intracellular Ac-CoA pool and histone acetylation, thus reshaping chromatin accessibility of contractile and aging-related genes. Srebf1 knockdown significantly attenuated the senescence-like transformation of VSMC both in vitro and in vivo. As a consequence, our findings bring insights into the pathophysiological action of lipid homeostasis in VSMC biology, providing therapeutic options for lipid-associated vascular diseases (summarized in Fig. 8).
Schematic diagram showing the hypothesis of how the microgravity stress promotes dedifferentiation and senescence of vascular smooth muscle cells.
Phenotypic transformation of VSMC has been identified as the leading feature of age-related vascular diseases44. The most acknowledged switching was from contractile to synthetic phenotype, which acts as a main pathological adjustment of VSMC in atherosclerosis45. Nevertheless, recent lineage tracing and single-cell sequencing studies uncovered macrophage-like, foam cell, mesenchymal stem-like, myofibroblast-like, and osteochondral-like phenotypes of VSMC16. In the present study, we performed transcriptomics and lipidomics in vivo and in vitro confirming a senescence-like transformation of VSMC in SM-induced carotid artery remodeling. Just recently, Marina et al. from NASA conducted an RNA-Seq study on human aortic smooth muscle cells cultured for 3 days in microgravity aboard the International Space Station46. Consistent with our findings, they found the markers of contractile, synthetic, and osteogenic phenotypes were downregulated. All these revelations suggest that phenotypic transformation is one of the common effects of real or simulated microgravity on VSMC biology. It would be a novel target for intervention of microgravity-related vascular remodeling in future.
It has been long known that extracellular physical cues (flow/shear stress, compression, substrate stiffness, and mechanical stress) elicit cell phenotypic plasticity11,47,48,49,50. The underlying mechanisms relate to various pathways, such as integrins, YAP/TAZ, FAK/Src, PI3K/Akt, RhoA/ROCK, mTOR, and Piezo receptor signaling48,51,52. Recently, the role of SREBP-mediated lipid metabolism in mechanical sensing has been gradually recognized. Patrizia et al. demonstrated that SREBP-mediated lipid synthesis responded to mechanical signals through cell–matrix adhesions. Extracellular physical stimulation actuated the mechanical properties of the Golgi apparatus, leading to the accumulation of SCAP/SREBP, which in turn drove SREBP activation53. In addition, Rebecca et al. identified the acto-myosin and extracellular matrix mediating the SREBP1 motivation by mechanical stress54. In addition, geranylgeranyl pyrophosphate activated SREBP1 and promoted fat storage in Drosophila, influencing stem cell fate in mouse upon physical stimulations54. In the current study, we demonstrated that SREBP1-mediated lipogenesis contributes to the phenotypic plasticity of VSMC induced by SM. As for the upstream causes, Golgi apparatus as well as some classical mechanosensitive pathways, such as mTOR, Piezo, YAP/TAZ may be involved as they have been shown playing a role in mechanotransduction or microgravity-induced osteoporosis and skeletal muscle atrophy20,39,55,56,57, which needs further investigation.
Cellular metabolism changes largely along with senescence and age-related diseases, restore of which would be a powerful therapeutic strategy58,59,60,61,62,63. Many lipid subclasses (fatty acids, TG, ether-linked phosphatidylethanolamines) in human plasma are modified during aging, which differs between sexes and is accelerated in the presence of insulin resistance64. Accumulation of glycerol-3-phosphate and phosphoethanolamine alters the senescence gene expression in human fibroblasts65. Cholesterol biosynthetic pathways elicit cellular senescence through ERRα in human hepatocytes66. Recently, Georges et al. verified the central role of lipids in multi-organs aging, and identified bis(monoacylglycero)phosphate accumulation as a conserved signature in both mouse and human67. Likewise, in the present study, SREBP1 activation triggered the VSMC dedifferentiation and senescence by enriching the Ac-CoA pool.
Histone deacetylases (HDACs) regulate transcription by removing histone acetylation68. Notably, the functional interpretation of HDAC inhibitors has been complicated by the prevalent use of pan-inhibitors that simultaneously target multiple HDAC isoforms, often yielding contradictory results even in identical experimental models. A striking example is trichostatin A (TSA), a pan-HDAC inhibitor, which was reported to suppress phenotypic switching in rat carotid artery balloon injury models69, while another study using the same model demonstrated that TSA promoted this process70. However, in our study, DaIC (containing TSA) enhanced global histone acetylation, leading to VSMC differentiation and senescence through chromatin remodeling, specifically by increasing accessibility of aging and lipogenesis related genes while reducing accessibility of contractile gene loci. In addition, in recent years, emerging evidence reported that ACLY and ACSS2 mediated elevation of the acetyl-CoA pool and consequent histone hyperacetylation are pivotal drivers of cellular senescence, VSMC phenotypic switching, and vascular pathogenesis71,72,73,74. Our results further demonstrate that microgravity-induced senescence-like transformation in VSMCs is mechanistically linked to Acly/Acss2 upregulation, acetyl-CoA pool expansion, and subsequent histone hyperacetylation, thereby firstly extending the established ACLY/ACSS2/Ac-CoA axis paradigm to vascular adaptation under simulated microgravity conditions. Moreover, it should be noted that acetyl-CoA homeostasis and histone acetylation are intricately regulated through multiple interconnected pathways, including lipid metabolism, TCA cycle, mitochondrial function, and the sophisticated post-translational protein acetylation75,76. Future studies should focus on elucidating the specific downstream triggered by SREBP1 activation in this context.
Our study identifies SREBF1 as a central regulator in microgravity-induced VSMC senescence. Notably, this finding may bear profound implications for the pathological mechanisms underlying common vascular diseases. Of particular significance, compelling evidence has established that SREBF1, its homolog SREBF2, and its activator SCAP play pivotal roles in the progression of atherosclerosis23,24,77. For instance, SREBF1/MicroRNA-33b exhibits potent effect on unstable atherosclerotic plaque formation via impairing macrophage cholesterol efflux and exacerbating inflammation77. In addition, genetic ablation of SCAP attenuated the development of atherosclerotic lesions, whereas its overexpression exacerbated plaque progression23,24. These independent observations align remarkably with our results, collectively suggesting that SREBF1-mediated lipogenic reprogramming and epigenetic regulation may constitute a convergent hub pathway connecting diverse stressors—including aberrant mechanical cues and metabolic insults—to VSMC dysfunction. Consequently, the SREBF1 mechanism uncovered here under microgravity conditions not only elucidates potential drivers of vascular remodeling during spaceflight but may also offer fresh perspectives on the pathogenesis of ground-based vascular aging and atherosclerotic diseases. Therapeutic targeting of SREBF1 could thus emerge as a broad-spectrum strategy to mitigate vascular pathologies triggered by diverse etiologies.
This study had some limitations. First, although we demonstrated that SREBP1-induced VSMC senescence is involved in vascular remodeling caused by simulated microgravity, additional investigations are needed to explore the potential targets bridging microgravity and SREBP1-activation, and the role of other non-VSMC cells (including fibroblasts, pericytes, and immune cells) in microgravity-related vascular remodeling. Second, our results demonstrate that the Ac-CoA pool and histone acetylation upregulated by SREBP1 activation contributes to VSMC senescence by reshaping chromatin accessibility. However, the other lipid-mediated pathways or acetylated proteins may contribute to VSMC senescence, and future studies will dissect their interplay with epigenetics. Third, although the hindlimb unloading model employed in this study is a well-established standard for investigating microgravity physiology, we acknowledge its inherent limitations34,78. This model primarily replicates certain physiological adaptations to spaceflight by simulating a state of mechanical unloading in the cardiovascular system, rather than creating true microgravity conditions. Consequently, the carotid artery senescence-like alterations and SREBF1 pathway activation observed herein may not solely originate from the absence of gravitational signals, but are likely intricately linked to broader mechanical environment perturbations induced by the model. These include, but are not limited to, localized hemodynamic alterations, dysregulated vascular wall stress, and associated neurohumoral adaptive responses. Future investigations, such as validation during actual spaceflight missions or cross-verification using alternative ground-based models, will be crucial to delineate the specific contributions of these mechanical factors and more precisely elucidate the direct role of gravitational signaling in vascular homeostasis. In addition, our results show that Srebf1 knockdown significantly mitigates VSMC senescence induced by microgravity. Nonetheless, we cannot exclude the possibility that the upstream signals of SREBP1-mediated lipogenesis represent a potent strategy for treating vascular remodeling induced by microgravity or aging. Furthermore, the studies using alternative models to validate the effectiveness of targeting SREBP1-mediated lipogenesis in the treatment of age-related vascular disease should be conducted in the future.
This work reveals a previously unrecognized role of SREBP1-mediated lipogenesis in vascular biology. Our findings not only reinforce the biological significance of lipid homeostasis in VSMC’ plasticity, but also provide insights into the vascular protection during spaceflight and for the therapy of major vascular diseases.
Methods
Animal experiments
Male Sprague-Dawley male rats (6 ~ 7 weeks old, body weight: 200 ± 10 g) and C57BL/6 (6 ~ 8 weeks old) male mice were obtained from laboratory animal center of Fourth Military Medical University, and individually caged within a 12:12 h light/dark cycle at 23 ± 1 °C with standard lab chow and water available free to access. All animal experiments in this study were conducted using male rats/mice exclusively to eliminate potential confounding effects of sex hormones on experimental outcomes, consistent with standard practices in this field of research. Special attention was paid to animal welfare and the minimization of suffering. All animal experiments were followed the rules of the American Association for the Accreditation of Laboratory Animal Care International. All procedures were approved by the Animal Care Committee of the Fourth Military Medical University accredited by AAALAC International (No. 20210429). We have complied with all relevant ethical regulations for animal use.
Hindlimb unloading (HU) models
After 1 week (W) of accommodation, the rats or mice were randomly assigned to control (CON) and HU group. HU was achieved by maintaining the animals in a 30° head-down tilt position using a tail harness, with the fully extended hindlimbs 0.5 cm above the floor79,80. HU duration was 4 W. The CON groups were synchronized timely to HU group. The ratio of left soleus wet weight to body weight (SW/BW, mg/g) was calculated to verify the effectiveness of HU model.
Ultrasound imaging
A Vevo 3100 LAZR ultrasound system (FUJIFILM Visual Sonics, Canada) was used for vascular ultrasound imaging. Rats or mice were anaesthetized by 1.5–2% isoflurane and placed on the platform. Luminal diameter and vessel wall thickness were measured by M-mode ultrasonography at carotid artery.
Morphological analysis
H&E, Masson’s trichrome, and Verhoeff’s Van Gieson staining (EVG) staining were employed for further morphological analysis. After euthanasia, rats or mice were transcardially perfused with PBS followed by 4% PFA. The tissues were isolated and embedded in paraffin. Successive 8 μm sections were cut for further experiments. These sections were stained with H&E, Masson’s trichrome, and EVG to evaluate morphology (Servicebio). All the graphs were taken by Olympus VS200.
In situ transfection of the carotid arteries by lentivirus or AAV loaded in Laponite hydrogels
Lentivirus encoding mCherry protein or sh-Srebf1 was constructed by General Biol Co.Ltd. (Anhui, China) or Tsingke Biotechnology Co., Ltd. (Beijing, China). 50 μL lentivirus (titer: 1.5 × 108 TU/mL) were mixed with 50 μL Laponite hydrogels, and 100 μL of the mixture was applied locally to the adventitia of the carotid arteries of the rats. In Supplementary Fig. S18B, 14 days after the surgery, the rats were euthanized, and the carotid artery were separated from surrounding tissues, harvested, and observed with the IVScope 8200 In Vivo Imaging System (Clinx, Shanghai, China). Similarly, AAV2 encoding mCherry protein or Tagln-mir30-sh-Srebf1 was constructed by General Biol Co.Ltd. (Anhui, China). 25 μL lentivirus (titer: 3 × 1012 Vg/mL) were mixed with 25 μL Laponite hydrogels, and 50 μL of the mixture was applied locally to the adventitia of the carotid arteries of the mice.
Cell culture and transfection
MOVAS, HASMC, and HUVEC cells are obtained originally from ATCC. MOVAS were maintained at 37 °C with 95% air and 5% CO2 in DMEM (Sigma) supplemented with 10% fetal bovine serum (FBS, Gibco), streptomycin (100 μg/ml) and penicillin (100 IU/ml) (Gibco). HASMC were maintained at 37 °C with 95% air and 5% CO2 in Smooth Muscle Cell Medium (ScienCell, Cat. #1101). HUVEC were maintained at 37 °C with 95% air and 5% CO2 in Endothelial Cell Medium (ScienCell, Cat. #1001). Before treatment, cells were seeded in T25 plates, 6-well plates or 20 mm confocal dishes and cultured for 12 h.
Rat primary carotid artery smooth muscle cells (RCA-SMCs) were isolated from the carotid artery of adult male Sprague Dawley rats. Anaesthetize the rats by 2% isoflurane, dissect the neck and excise their carotid artery. After removing the adventitia, the carotid artery tissues were cut open and scraped off the intima gently. Then, the carotid artery were washed with cold DPBS (Sigma), minced into ~ 1 mm3 pieces, and digested using 1 mg/ml collagenaseⅠ(BioFROXX) at 37 °C in a water bath for ~ 30 min. After digestion, the supernatant was collected and mixed with DMEM (Gibco). Cells were collected by centrifuging at 500 × g for 5 min and further resuspended in DMEM supplemented with 10% FBS (Gibco), streptomycin (100 μg/ml) and penicillin (100 IU/ml) (Gibco). The purity of RCA-SMCs was tested by immunofluorescence staining of α-SMA (Proteintech, Cat. #14395-1-AP, 1:200). Isolated RCA-SMCs were maintained in low-glucose DMEM (Gibco) containing 10% FBS (Gibco), and cells at passages 4 to 8 were used for further experiments.
For gene knockdown, MOVAS were transfected with siRNA by TSnanofect V2 transfection Reagent (Cat. #TSV405, Tsingke Biotechnology Co., Ltd. Beijing, China) according to the manufacturer’s protocol. The corresponding sequences are listed in Supplementary Table 2.
For Srebf1 knockdown, RCA-SMCs were infected with lentivirus (Lv) carrying negative control (shCtrl) or knockdown of rat Srebf1 gene (sh-Srebf1) at an MOI of 100 from Tsingke Biotechnology Co., Ltd. (Beijing, China). The corresponding sequences are listed in Supplementary Table 2. After puromycin screening, RCA-SMCs was used for further assays.
All cells were maintained in a humidified 5% CO2 incubator at 37 °C.
Two-dimensional rotation-simulated microgravity cell models (SM)
To simulate microgravity, we used a two-dimensional (2D) clinostat, which was designed and provided by the China Astronaut Research and Training Center (Beijing, China)35,81. Rotation causes a gravity vector that is not recognizable by cells. Therefore, the device prevents the cells from feeling gravity. In the present study, cells were incubated in T25 culture flasks or plated on confocal dishes for following experiments. After cell adhesion, flasks or flasks mounted with coverslips were filled up with culture medium to prevent the presence of air bubbles. The dishes were fixed carefully to the rotating panel of the clinostat system, and rotated at a constant speed for 48 h to simulate microgravity (0.01 g). Cells without rotation (1 g) were cultured in the same manner as the control group.
Senescence β-Galactosidase staining
Senescence β-Galactosidase was stained by using a SA-β-gal detection kit (Beyotime Biotechnology, Shanghai, China). The cells or tissues were immersed in the staining solution and incubated at 37 °C for 12 h. Blue staining were suggested to be SA-β-gal-positive signals.
Cell contraction assay
A Cell Contraction Assay Kit (CBA-021) was purchased from Cell Biolabs, Inc. (San Diego, CA, USA). The contractile ability of the cells was evaluated according to the manufacturer’s instructions. Cells were harvested and resuspended in culture medium. A collagen lattice was prepared by mixing the cell suspension and ice-cold collagen gel solution in a volume ratio of 1:4. Then, 0.1 mL of the cell-collagen mixture (containing 104 cells) was added to a 96-well plate and incubated for 1 hour at 37 °C. After collagen polymerization, 0.1 mL of culture medium was added on top of each collagen gel lattice. 24 h later, the plates were scanned. For data analysis, the area of the gel in each well was analyzed by Fiji Image J (NIH, Bethesda, MD, United States).
Cell apoptosis assay
Annexin V-FITC Apoptosis Detection Kit (Beyotime, Cat. #C1062S) was used to stain apoptotic cells as per the manufacturer’s instructions. Briefly, cells were seeded in T25 plates and treated with SM model for 48 h. Cells were trypsinized, harvested and stained with Annexin-V and PI for 15 min at RT in assay buffer. Annexin-V stained cells were centrifuged and washed with assay buffer. Next, cells were measured by CytoFLEX (Beckman Colter Life Sciences). Data was analyzed with FlowJo v.10 software (BD Life Sciences).
Transmission electron microscopy (TEM)
After washing with PBS, cell samples were fixed in 2.5 % glutaraldehyde and 0.1 M sodium cacodylate buffer at 4 °C overnight. All the samples were embedded and subjected to ultramicrotomy. Transmission electron microscope images were captured by JEM-1400 (Japan Electron Optics Laboratory) and iTEM 5.2 (EMSIS).
Confocal microscopy
After specific treatment, cells in confocal dishes were washed by pre-warmed DPBS twice and followed by incubation with different dyes according to the manufacturer’s protocol. Then, supernatants were discarded, and cells were washed twice using pre-warmed DPBS. Finally, the images of stained cells were taken on Nikon A1 PLUS. The dyes information was listed as following: BODIPY (lipid detection, 3 μM, MCE, Cat. #HY-W090090); Nile Red (lipid detection, 0.5 μM, MCE, Cat. #HY-D0718); SPiDER-βGal (senescence detection, Dojindo, Cat. #SG02).
Immunofluorescence
For cells, MOVAS and RCA-SMCs were seeded on confocal dishes at an appropriate density. After treatments, the cells were washed with DPBS twice and then fixed with 4% formaldehyde buffer (PFA) at 37 °C for 20 min. After washing 2-3 times with PBS, the cells were permeabilized and blocked with 5% BSA and 15% goat serum in 1 × PBS for 1 h at 25 °C. Then, the primary antibodies were incubated at 4 °C overnight, and Alexa Fluor–conjugated secondary antibodies were incubated at 25 °C for 2 h. DAPI was used to stain nuclei. All the graphs were viewed and taken by Nikon A1 PLUS. All the used antibodies were summarized as following: SREBP1 (Proteintech, Cat. #66875-1-Ig, 1:200); SREBP2 (Proteintech, Cat. #28212-1-AP, 1:250); α-SMA (Proteintech, Cat. #14395-1-AP, 1:1000); Pan-Acetylation (Proteintech, Cat. #66289-1-Ig, 1:100); CD31 (Servicebio, Cat. # GB113151-100, 1:5000); Ki-67 (abcam, Cat. #ab16667, 1:2000); P21 (Servicebio, Cat. #GB115313-100, 1:200); SRF (Proteintech, Cat. #16821-1-AP, 1:200); MYOCD (Affinity Biosciences, Cat. #DF2434, 1:200); H3K9Ac (ABclonal, Cat. #A7255, 1:200); H3K27Ac (Beyotime, Cat. #AF5620, 1:200); IgG-mouse (Proteintech, Cat. #B900620, 1:500); IgG-rabbit (Proteintech, Cat. #30000-0-AP, 1:500).
Flow cytometry analysis
After specific treatment, 2 × 105 cells were collected and stained with different dyes according to the manufacturer’s protocol, followed by three washes with specific buffer and centrifugation at 500 × g for 5 min at 25 °C. Then, supernatants were discarded, and cells were resuspended in 300 ~ 500 μL buffer. Finally, the labeled cells were analyzed using a CytoFLEX (Beckman Colter Life Sciences) and FlowJo v.10 software (BD Life Sciences). The dyes information was listed as following: BODIPY (lipid detection, 3 μM, MCE, Cat. #HY-W090090); SPiDER-βGal (senescence detection, Dojindo, Cat. #SG02). Gating strategies were summarized in Supplementary Information file.
Enzyme linked immunosorbent assay (ELISA)
The cells were collected into a 1.5 mL centrifuge tube. 5 × 105 cells were dissolved in 200 μL extraction solution. After sonication for 3 min on ice, the extraction solution was centrifuged at 13,000 × g for 10 min to collect supernatant. The supernatant was placed on ice to be tested. Acetyl-CoA, fatty acid (FA), and triglyceride (TG) levels were measured using an Acetyl-CoA ELISA Kit (Sangon Biotech, Cat. #D751001), FA ELISA Kit (Meimian Industrial Co., Ltd., Cat. #MM-71779R2), and TG ELISA Kit (Meimian Industrial Co., Ltd., Cat. #MM-0610R2) following the manufacturer’s protocol, respectively. The concentration was normalized with the protein content of each sample (BCA Protein Assay Kit, BioVision).
Measurement of citrate levels
Citrate levels in cells and tissue were measured by Citrate Content Assay Kit (Sorlarbio, Cat. #BC2155) following by the recommendations of the manufacturer. The citrate concentration and standard curve were established by an enzyme reader (TECAN, Cat. #30086376, Switzerland). The citrate level was normalized with the protein content of each sample (BCA Protein Assay Kit, BioVision).
Protein isolation and western blot analysis
Nuclear and cytoplasmic proteins were extracted with a nuclear and cytoplasmic protein extraction kit (Yeasen, Cat. #20126ES50) according to the manufacturer’s instructions. Cells, tissue and isolated nucleus samples were washed by ice-cold PBS and lysed with RIPA lysis buffer (Beyotime, Cat. #P0013B) supplemented with phosphatase and protease inhibitors (Roche). Lysates were separated on 10–12.5% SDS-PAGE gels (Beyotime, Cat. #P0012AC), and then transferred to 0.22–0.45 μm polyvinylidene difluoride membranes (Merck Millipore) by Trans-Blot System (Bio-Rad Laboratories). Membranes were then blocked with 5% skimmed milk diluted in TBST for 1 h at 25 °C, and then were incubated in diluted primary antibodies overnight at 4 °C. Subsequently, the membranes were incubated with secondary antibodies conjugated with horseradish peroxidase (HRP) for 1 h at 25 °C. Western HRP substrate (Millipore) and imaging system were employed to blot detection. The antibodies used in this study are as followed: β-Tubulin (Abways, Cat. #AB0057, 1:2000); β-Actin (Sigma, Cat. #A5316, 1:10000); α-SMA (Proteintech, Cat. #14395-1-AP, 1:1000); SM22α (Proteintech, Cat. #, 1:1000); P21 (Servicebio, Cat. #GB115313-100, 1:500); SREBP1 (Proteintech, Cat. #66875-1-Ig, 1:1000); SREBP2 (Proteintech, Cat. #28212-1-AP, 1:1000); MYOCD (Affinity Biosciences, Cat. #DF2434, 1:1000); Pan-Acetylation (Proteintech, Cat. #66289-1-Ig, 1:500); ACLY (Proteintech, Cat. #67166-1-Ig, 1:1000); ACSS2 (Proteintech, Cat. #16087-1-AP, 1:1000); H3K27Ac (Beyotime, Cat. #AF5620, 1:1000); H3K9Ac (ABclonal, Cat. #A7255, 1:1000); H3K18Ac (PTM BIO, Cat. #PTM-114RM, 1:1000); H3 (Proteintech, Cat. #17168-1-AP, 1:1000); SRF (Proteintech, Cat. #16821-1-AP, 1:1000); LaminA/C (Proteintech, Cat. #10298-1-AP, 1:1000); p-AKT (abcam, Cat. #ab38449, 1:1000); AKT (abcam, Cat. #ab8805, 1:1000); mTOR (abcam, Cat. #ab134903, 1:1000); p-mTOR (abcam, Cat. #ab109268, 1:1000).
RNA extraction and real-time quantitative polymerized chain reaction (RT-qPCR)
Total RNA was extracted from carotid artery tissues and cells according to the protocols of Axypre™ Multisource Total RNA Miniprep kit (Axygen). The isolated RNA was then reverse-transcribed by using PrimeScript™ RT Mix with gDNA eraser (Takara). SYBR®Premix Ex Taq™ Ⅱ (Takara) was used for qRT-PCR by using 7500 system (Applied Biosystems). All the primers are listed in Supplementary Table 1. The relative mRNA expression was analysis using the comparative threshold cycle method (2−ΔΔCt) using Actb as the endogenous control gene.
Immunohistochemistry
After deparaffinization, serial sections (4 μm) were soaked in citric acid (pH = 6.0) antigen retrieval buffer (Servicebio, Cat. #G1203) and heated twice for 10 minutes for antigen retrieval in a microwave oven. Then, the sections were placed in 3% hydrogen peroxide solution and incubated at 25 °C in darkness for 25 min to block endogenous peroxidase activity. The sections were then incubated with primary antibodies. HRP-labeled antibodies (Servicebio, Cat. #GB23303, 1:200) were used as secondary antibodies. Nuclei were were counterstained using hematoxylin (Servicebio), and diaminobenzidine (Servicebio, Cat. #G1211) was served as the chromogen. The primary antibodies used in this study are as followed: α-SMA (Proteintech, Cat. #14395-1-AP, 1:100); P21 (Servicebio, Cat. #GB115313-100, 1:50); SREBP1 (Proteintech, Cat. #66875-1-Ig, 1:100); SREBP2 (Proteintech, Cat. #28212-1-AP, 1:100); IgG-mouse (Proteintech, Cat. #B900620, 1:500); IgG-rabbit (Proteintech, Cat. #30000-0-AP, 1:500).
RNA-seq analysis
Total RNA was extracted using Axypre™ Multisource Total RNA Miniprep kit (Axygen) according to the manufacturer’s protocol82. RNA quality was evaluated by Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, CA, USA) and verified using RNase-free agarose gel electrophoresis. After treatment with Oligo(dT) beads, the enriched mRNA was then fragmented and reverse transcribed into cDNA using NEBNext Ultra RNA Library Prep Kit for Illumina (NEB #7530, New England Biolabs, Ipswich, MA, USA). The double-stranded cDNA products were then purified by QiaQuick PCR extraction kit (Qiagen, Venlo, The Netherlands), end repaired, added poly(A), and ligated with Illumina sequencing adapters. The adapter-ligated products were size selected by agarose gel electrophoresis. Finally, PCR amplification and sequencing were completed using Illumina Novaseq6000 by Gene Denovo Biotechnology Co. (Guangzhou, China). Raw reads were further filtered by fastp (V.18.0) to get high-quality clean reads. Short reads alignment tool Bowtie2 (V.2.2.8) was used for mapping reads to ribosome RNA (rRNA) database. The rRNA mapped reads then will be removed. The remaining clean reads were further used in assembly and gene abundance calculation. An index of the reference genome was built, and paired-end clean reads were mapped to the reference genome (Ensembl_release110 for MOVAS; Ensembl release109 for rats) using HISAT V.2.2.4 and other parameters set as a default. The mapped reads of each sample were assembled by using StringTie V.1.3.1 in areference-based approach. For each transcription region, an FPKM (fragment per kilobase of transcript per million mapped reads) value was calculated to quantify its expression abundance and variations, using RSEM software. RNAs differential expression analysis was performed by DESeq2 software between two different groups. The genes with a false-discovery rate (FDR) < 0.05 and fold-change ≥ 1.5 or ≤ 1/1.5 were considered differentially expressed genes. KEGG, GO, and GSEA enrichment analysis was performed using the KEGG, Gene Ontology database (http://www.geneontology.org/), and GSEA database (https://www.gsea-msigdb.org/gsea/index.jsp). Regulatory target gene sets from the GSEA database were used for transcription factors enrichment analysis (https://www.gsea-msigdb.org/gsea/msigdb/index.jsp) as previously described83. All the bioinformatic analysis was performed by Gene Denovo Biotechnology Co., Ltd (Guangzhou, China).
Lipid extraction and lipidomics analysis
Lipidomics analysis was performed by Shanghai Applied Protein Technology Co., Ltd. (Shanghai, China)84. Briefly, samples were first mixed with 200 μL methanol, and then 10 μL internal lipid standards (SPLASH™ LIPIDOMIX™ Mass Spec Standard, AVANTI, 330707-1EA) and 800 μL methyl tert-butyl ether were added. The quality control (QC) samples are a composite prepared by pooling equal volumes of all the samples to be tested. The mixture was adequately vortexed, sonicated for 20 min at 4 °C and then kept for 30 min at room temperature. After that, 200 μL of MS-grade water was added, and the mixture was vortexed and centrifuged at 20000 × g for 15 min at 4 °C. The upper organic solvent layer was obtained and dried under nitrogen. For LC-MS analysis, the samples were re-dissolved in 200 μL of IPA/CAN (9:1, v/v) solvent and centrifuged at 20000 g at 4 °C for 15 min, then the supernatant was injected. The analysis was performed on a UHPLC system (LC-30A, Shimadzu) coupled with QTRAP MS (6500 + , Sciex). The analytes were separated on HILIC (Phenomenex, Luna NH2, 2.0 mm × 100 mm, 3 µm) and C18 column (Phenomenex, Kinetex C18, 2.1 × 100 mm, 2.6 μm). For C18 separation, the column temperature was set at 45 °C. Mobile phase A: 70% acetonitrile + 30% H2O + 5 mM ammonium acetate, mobile phase B: IPA solution. A gradient (20% B at 0 min, 60% B at 5 min, 100% B at 13 min, 20% B at 13.1–17 min) was then initiated at a flow rate of 0.35 mL/min. The sample was placed at 10 °C during the whole analysis process. For Amino separation, the column temperature was set at 40 °C. Mobile phase A: 2 mM ammonium acetate + 50% methanol + 50% acetonitrile, mobile phase B: 2 mM ammonium acetate + 50% acetonitrile + 50% H2O. A gradient (3% B at 0 ~ 3 min, from 3% to 100% B at 3 ~ 13 min, 100% B at 13 ~ 17 min, 3% B at 17.1 ~ 22 min) was then initiated at a flow rate of 400 μL/min. 6500 + QTRAP (AB SCIEX) was performed in positive and negative switch mode. The ESI positive source conditions were as follows: Source temperature: 400 °C; Ion Source Gas1 (GS1): 50 Ion Source Gas2 (GS2): 55; Curtain Gas (CUR): 35; IonSpray Voltage (IS): + 3000 V; The ESI negative source conditions were as follows: Source temperature: 400 °C; Ion Source Gas1 (GS1): 50; Ion Source Gas2 (GS2): 55; Curtain gas (CUR): 35; IonSpray Voltage (IS): − 2500 V. Multiple Reaction Monitoring method was used for mass spectrometry quantitative data acquisition, and Sciex OS was used for quantitative data processing by Shanghai Applied Protein Technology Co., Ltd. (Shanghai, China). The concentrations of lipids (ng/g, normalized to sample weight) were summarized in Supplementary Data 1. Following lipids were quantified: glycerophospholipids (including phosphatidic acid PA, phosphatidylcholine PC, phosphatidylethanolamine PE, phosphatidylglycerol PG, phosphatidylinositol PI, phosphatidylserine PS, lysophosphatidic acid LPA, lysophosphatidylethanolamine LPE, lysophosphatidylserine LPS), Sphingolipids (including ceramide Cer, dihydroceramide DHCer, hexoceramide Hex1Cer, dihexose ceramide Hex2Cer, sphingomyelin SM), Glycerolipids (including monoglycerides MG, diglyceride DG, triglycerides TG), cholesteryl ester (ChE), and fatty acids (FA).
Immunoprecipitation and mass spectrometry (IP-MS)
For protein IP-MS, the cell lysates treated by Cell Lysis Buffer (CST, Cat. #9803S) were incubated with Pan-Acetylation (Proteintech, Cat. #66289-1-Ig) or IgG (Proteintech, Cat. #B900620) primary antibody overnight at 4 °C. Immunoprecipitates were collected and separated by SDS–PAGE. The gel lysis and protein digestion were performed by using the commercially available iST Sample Preparation kit (PreOmics, Germany). The peptides were analyzed by nano-HPLC–MS/MS, and the resulting data were processed by PEAKS Studio version 10.6 (Bioinformatics Solutions Inc., Waterloo, Canada) by Gene Denovo Biotechnology Co. (Guangzhou, China). The results were summarized in Supplementary Data 2 and 3.
ATAC-seq library preparation and sequencing
Briefly, cells were transferred into 5 μL of lysis buffer (10 mM Tris-HCl (pH 7.4), 10 mM NaCl, 3 mM MgCl2, and 0.005% digitonin) in ice for 10 min. The ATAC reaction was performed by adding 4 μL of ddH2O, 4 μL of TTBL (5 ×), and 5 μL TTE mix V5 (Vazyme Biotech, TD711) at 37 °C for 30 min and then stopped by adding 5 μL of TS (5 ×) stop buffer at 25 °C for 5 min. DNA was extracted using ATAC DNA Extract Beads. Then, DNA was dissolved in 20 µl ddH2O and PCR-amplified with 10 μL of index primers (Vazyme Biotech, TD202), 30 μL of CAM (2 ×) (Vazyme Biotech, TD711) with the program of 72 °C for 3 min; 95 °C for 3 min; 98 °C for 10 s, 60 °C for 5 s for 12 cycles; and 72 °C for 1 min. The amplified DNA was size-selected using ATAC DNA Clean Beads (Vazyme Biotech, TD711). The libraries were multiplexed and then sequenced using an Illumina novaseq 6000 with PE 150 method by IGENEBOOK Biotechnology Co., Ltd (Wuhan, China, http://www.igenebook.com). The bioinformatic analysis was performed by Vazyme Biotech Co., Ltd (Nanjing, China).
Statistics and reproducibility
Data were expressed as mean ± SD. Statistical analysis and graphical representation were performed in the GraphPad Prism software (v.9.0.0) or Rstudio (version 4.1). Normality of the data was evaluated using the Shapiro-Wilk test. Two-tailed unpaired Student t tests were used to assess the differences between two groups, when normal distribution was satisfied; otherwise, the Mann–Whitney test was used. The differences between multiple groups were compared using one-way ANOVA or two-way ANOVA, followed by Dunnett’s (one-way) or Tukey’s (two-way) multiple comparisons test, when normal distribution was satisfied; otherwise, the nonparametric Kruskal–Wallis test was performed for further analysis. Two-way mixed-effects ANOVA followed by Sidak’s multiple comparisons test was used to analyze differences in body weight at multiple time-points in order to account for the correlation of repeated measurements of the same rats (Supplementary Fig. S1B; S13H). Statistical significance was set at P < 0.05 (*); two asterisks (**) for P < 0.01; three asterisks (***) for P < 0.001; and four asterisks (****) for P < 0.0001. The representative data shown as pathological staining, immunohistochemistry, immunofluorescence, WB, TEM images were obtained from at least three independent experiments (for Figs. 1B, G, H, J, K; 2C, D; 3C–E, G, H; 4F, K, M, P; 5C, D, H, K, M; 6J; 7F, N).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The bulk RNA sequencing data have been deposited into the Sequence ReadArchive (SRA) with accession numbers PRJNA1140849, PRJNA1140851, PRJNA1140855. The remaining data are available within the Article, Supplementary Information, or Source Data file. Source data and unedited blot images are provided with this paper. Source data are provided in this paper.
References
da Silveira, W. A. et al. Comprehensive multi-omics analysis reveals mitochondrial stress as a central biological hub for spaceflight impact. Cell 183, 1185–1201 (2020).
Garrett-Bakelman, F. E. et al. The NASA Twins Study: A multidimensional analysis of a year-long human spaceflight. Science 364, https://doi.org/10.1126/science.aau8650 (2019).
Jirak, P. et al. How spaceflight challenges human cardiovascular health. Eur. J. Prevent. Cardiol. 29, 1399–1411 (2022).
Strollo, F., Gentile, S., Strollo, G., Mambro, A. & Vernikos, J. Recent progress in space physiology and aging. Front. Physiol. 9, 1551 (2018).
Scott, J. M., Stoudemire, J., Dolan, L. & Downs, M. Leveraging spaceflight to advance cardiovascular research on earth. Circul. Res. 130, 942–957 (2022).
Willeit, P. et al. Carotid intima-media thickness progression as surrogate marker for cardiovascular risk: meta-analysis of 119 clinical trials involving 100 667 patients. Circulation 142, 621–642 (2020).
Hughson, R. L., Helm, A. & Durante, M. Heart in space: effect of the extraterrestrial environment on the cardiovascular system. Nat. Rev. Cardiol. 15, 167–180 (2018).
Zhang, L. F. & Hargens, A. R. Spaceflight-induced intracranial hypertension and visual impairment: pathophysiology and countermeasures. Physiol. Rev. 98, 59–87 (2018).
Zhang, L. F. Region-specific vascular remodeling and its prevention by artificial gravity in weightless environment. Eur. J. Appl. Physiol. 113, 2873–2895 (2013).
Arbeille, P., Provost, R. & Zuj, K. Carotid and femoral artery intima-media thickness during 6 months of spaceflight. Aerosp. Med. Hum. Perform. 87, 449–453 (2016).
Lacolley, P., Regnault, V., Segers, P. & Laurent, S. Vascular smooth muscle cells and arterial stiffening: relevance in development, aging, and disease. Physiol. Rev. 97, 1555–1617 (2017).
Owens, G. K., Kumar, M. S. & Wamhoff, B. R. Molecular regulation of vascular smooth muscle cell differentiation in development and disease. Physiol. Rev. 84, 767–801 (2004).
Chakraborty, R. et al. Histone acetyltransferases p300 and CBP coordinate distinct chromatin remodeling programs in vascular smooth muscle plasticity. Circulation 145, 1720–1737 (2022).
Elmarasi, M. et al. Phenotypic switching of vascular smooth muscle cells in atherosclerosis, hypertension, and aortic dissection. J. Cell. Physiol. 239, e31200 (2024).
Miano, J. M., Fisher, E. A. & Majesky, M. W. Fate and state of vascular smooth muscle cells in atherosclerosis. Circulation 143, 2110–2116 (2021).
Cao, G. et al. How vascular smooth muscle cell phenotype switching contributes to vascular disease. Cell Commun. Signal. 20, 180 (2022).
Zeng, Q. et al. Lipids and lipid metabolism in cellular senescence: Emerging targets for age-related diseases. Ageing Res. Rev. 97, 102294 (2024).
Jeon, Y. G., Kim, Y. Y., Lee, G. & Kim, J. B. Physiological and pathological roles of lipogenesis. Nat. Metab. 5, 735–759 (2023).
Borén, J., Taskinen, M. R., Björnson, E. & Packard, C. J. Metabolism of triglyceride-rich lipoproteins in health and dyslipidaemia. Nat. Rev. Cardiol. 19, 577–592 (2022).
Shimano, H. & Sato, R. SREBP-regulated lipid metabolism: convergent physiology - divergent pathophysiology. Nat. Rev. Endocrinol. 13, 710–730 (2017).
Shao, W. & Espenshade, P. J. Expanding roles for SREBP in metabolism. Cell Metab. 16, 414–419 (2012).
Fernández-Hernando, C. & Moore, K. J. MicroRNA modulation of cholesterol homeostasis. Arterioscler. Thromb. Vasc. Biol. 31, 2378–2382 (2011).
Li, D. et al. Sterol-resistant SCAP overexpression in vascular smooth muscle cells accelerates atherosclerosis by increasing local vascular inflammation through activation of the NLRP3 inflammasome in mice. Aging Dis. 12, 747–763 (2021).
Li, D. et al. SCAP knockdown in vascular smooth muscle cells alleviates atherosclerosis plaque formation via up-regulating autophagy in ApoE(-/-) mice. FASEB J. 33, 3437–3450 (2019).
Cao, K. et al. Glycolysis and de novo fatty acid synthesis cooperatively regulate pathological vascular smooth muscle cell phenotypic switching and neointimal hyperplasia. J. Pathol. 259, 388–401 (2023).
Neggazi, S. et al. Hypothyroidism increases angiotensinogen gene expression associated with vascular smooth muscle cells cholesterol metabolism dysfunction and aorta remodeling in Psammomys obesus. Sci. Rep. 13, 19681 (2023).
Sabari, B. R., Zhang, D., Allis, C. D. & Zhao, Y. Metabolic regulation of gene expression through histone acylations. Nat. Rev. Mol. Cell Biol. 18, 90–101 (2017).
Peleg, S., Feller, C., Ladurner, A. G. & Imhof, A. The metabolic impact on histone acetylation and transcription in ageing. Trends Biochem. Sci. 41, 700–711 (2016).
Nitsch, S., Zorro Shahidian, L. & Schneider, R. Histone acylations and chromatin dynamics: concepts, challenges, and links to metabolism. EMBO Rep. 22, e52774 (2021).
Sen, P., Shah, P. P., Nativio, R. & Berger, S. L. Epigenetic mechanisms of longevity and aging. Cell 166, 822–839 (2016).
Ibarrola, J. et al. Smooth muscle mineralocorticoid receptor as an epigenetic regulator of vascular ageing. Cardiovas. Res. 118, 3386–3400 (2023).
Azechi, T. et al. Trichostatin A, an HDAC class I/II inhibitor, promotes Pi-induced vascular calcification via up-regulation of the expression of alkaline phosphatase. J. Atheroscler. Thromb. 20, 538–547 (2013).
Li, P., Ge, J. & Li, H. Lysine acetyltransferases and lysine deacetylases as targets for cardiovascular disease. Nat. Rev. Cardiol. 17, 96–115 (2020).
Su, Y. T. et al. Acid sphingomyelinase/ceramide mediates structural remodeling of cerebral artery and small mesenteric artery in simulated weightless rats. Life Sci. 243, 117253 (2020).
Liu, C. et al. The mechanosensitive lncRNA Neat1 promotes osteoblast function through paraspeckle-dependent Smurf1 mRNA retention. Bone Res. 10, 18 (2022).
Simcox, J. & Lamming, D. W. The central moTOR of metabolism. Dev. Cell 57, 691–706 (2022).
Bakan, I. & Laplante, M. Connecting mTORC1 signaling to SREBP-1 activation. Curr. Opin. lipidol. 23, 226–234 (2012).
Ortega-Molina, A. et al. A mild increase in nutrient signaling to mTORC1 in mice leads to parenchymal damage, myeloid inflammation and shortened lifespan. Nat. Aging 4, 1102–1120 (2024).
Evers, T. M. J., Holt, L. J., Alberti, S. & Mashaghi, A. Reciprocal regulation of cellular mechanics and metabolism. Nat. Metab. 3, 456–468 (2021).
Nacarelli, T., Azar, A. & Sell, C. Mitochondrial stress induces cellular senescence in an mTORC1-dependent manner. Free Radic. Biol. Med. 95, 133–154 (2016).
Qu, Y. Q. et al. 2-Aminoethoxydiphenylborane sensitizes anti-tumor effect of bortezomib via suppression of calcium-mediated autophagy. Cell Death Dis. 9, 361 (2018).
Kim, Y. M. et al. Sterol regulatory element-binding protein (SREBP)−1-mediated lipogenesis is involved in cell senescence. J. Biol. Chem. 285, 29069–29077 (2010).
Zheng, S. S. et al. Inhibition of neointimal hyperplasia in balloon-induced vascular injuries in a rat model by miR-22 loading Laponite hydrogels. Biomater. Adv. 142, 213140 (2022).
Zhang, F., Guo, X., Xia, Y. & Mao, L. An update on the phenotypic switching of vascular smooth muscle cells in the pathogenesis of atherosclerosis. Cell. Mol. life Sci. 79, 6 (2021).
Gomez, D. & Owens, G. K. Smooth muscle cell phenotypic switching in atherosclerosis. Cardiovas. Res. 95, 156–164 (2012).
Scotti, M. M. et al. Spaceflight effects on human vascular smooth muscle cell phenotype and function. NPJ Microgravity 10, 41 (2024).
Jain, P. et al. Dynamical hallmarks of cancer: Phenotypic switching in melanoma and epithelial-mesenchymal plasticity. Semin. Cancer Biol. 96, 48–63 (2023).
Totaro, A., Panciera, T. & Piccolo, S. YAP/TAZ upstream signals and downstream responses. Nat. Cell Biol. 20, 888–899 (2018).
Song, Y., Soto, J. & Li, S. Mechanical regulation of histone modifications and cell plasticity. Curr. Opin. Solid State Mater. Sci. 24, https://doi.org/10.1016/j.cossms.2020.100872 (2020).
Wang, J., Lü, D., Mao, D. & Long, M. Mechanomics: an emerging field between biology and biomechanics. Protein Cell 5, 518–531 (2014).
Young, K. M. & Reinhart-King, C. A. Cellular mechanosignaling for sensing and transducing matrix rigidity. Curr. Opin. Cell Biol. 83, 102208 (2023).
Li, N. et al. Multiscale biomechanics and mechanotransduction from liver fibrosis to cancer. Adv. Drug Deliv. Rev. 188, 114448 (2022).
Romani, P. et al. Extracellular matrix mechanical cues regulate lipid metabolism through Lipin-1 and SREBP. Nat. Cell Biol. 21, 338–347 (2019).
Bertolio, R. et al. Sterol regulatory element binding protein 1 couples mechanical cues and lipid metabolism. Nat. Commun. 10, 1326 (2019).
Hu, Y. et al. The critical role of the piezo1/β-catenin/ATF4 axis on the stemness of Gli1(+) BMSCs during simulated microgravity-induced bone loss. Adv. Sci. 10, e2303375 (2023).
Wubshet, N. H. et al. Cellular mechanotransduction of human osteoblasts in microgravity. NPJ Microgravity 10, 35 (2024).
Usuki, F. et al. Local Vibration Stimuli Induce Mechanical Stress-Induced Factors and Facilitate Recovery From Immobilization-Induced Oxidative Myofiber Atrophy in Rats. Front. Physiol. 10, 759 (2019).
Zwerschke, W. et al. Metabolic analysis of senescent human fibroblasts reveals a role for AMP in cellular senescence. Biochem. J. 376, 403–411 (2003).
Johmura, Y. et al. Senolysis by glutaminolysis inhibition ameliorates various age-associated disorders. Science 371, 265–270 (2021).
Ogrodnik, M. et al. Obesity-induced cellular senescence drives anxiety and impairs neurogenesis. Cell Metab. 29, 1061–1077 (2019).
Flor, A. C., Wolfgeher, D., Wu, D. & Kron, S. J. A signature of enhanced lipid metabolism, lipid peroxidation and aldehyde stress in therapy-induced senescence. Cell Death Discov. 3, 17075 (2017).
Wiley, C. D. & Campisi, J. The metabolic roots of senescence: mechanisms and opportunities for intervention. Nat. Metab. 3, 1290–1301 (2021).
Mutlu, A. S., Duffy, J. & Wang, M. C. Lipid metabolism and lipid signals in aging and longevity. Dev. Cell 56, 1394–1407 (2021).
Hornburg, D. et al. Dynamic lipidome alterations associated with human health, disease and ageing. Nat. Metab. 5, 1578–1594 (2023).
Tighanimine, K. et al. A homoeostatic switch causing glycerol-3-phosphate and phosphoethanolamine accumulation triggers senescence by rewiring lipid metabolism. Nat. Metab. 6, 323–342 (2024).
Ziegler, D. V. et al. Cholesterol biosynthetic pathway induces cellular senescence through ERRα. NPJ Aging 10, 5 (2024).
Janssens, G. E. et al. A conserved complex lipid signature marks human muscle aging and responds to short-term exercise. Nat. Aging 4, 681–693 (2024).
Zhang, M. et al. HDAC6 Regulates the MRTF-A/SRF axis and vascular smooth muscle cell plasticity. JACC. Basic Transl. Sci. 3, 782–795 (2018).
Kee, H. J. et al. Trichostatin A prevents neointimal hyperplasia via activation of Krüppel like factor 4. Vas. Pharmacol. 55, 127–134 (2011).
Song, S., Kang, S. W. & Choi, C. Trichostatin A enhances proliferation and migration of vascular smooth muscle cells by downregulating thioredoxin 1. Cardiovas. Res. 85, 241–249 (2010).
Bradshaw, P. C. Acetyl-CoA Metabolism and Histone Acetylation in the Regulation of Aging and Lifespan. Antioxidants 10, https://doi.org/10.3390/antiox10040572 (2021).
Chen, W. et al. The SESAME complex regulates cell senescence through the generation of acetyl-CoA. Nat. Metab. 3, 983–1000 (2021).
Grobs, Y. et al. ATP citrate lyase drives vascular remodeling in systemic and pulmonary vascular diseases through metabolic and epigenetic changes. Sci. Transl. Med. 16, eado7824 (2024).
Wu, H. et al. ANK Deficiency-Mediated Cytosolic Citrate Accumulation Promotes Aortic Aneurysm. Circ. Res. 135, 1175–1192 (2024).
Guertin, D. A. & Wellen, K. E. Acetyl-CoA metabolism in cancer. Nat. Rev. Cancer 23, 156–172 (2023).
Pietrocola, F., Galluzzi, L., Bravo-San Pedro, J. M., Madeo, F. & Kroemer, G. Acetyl coenzyme A: a central metabolite and second messenger. Cell Metab. 21, 805–821 (2015).
Nishino, T. et al. SREBF1/MicroRNA-33b axis exhibits potent effect on unstable atherosclerotic plaque formation In Vivo. Arterioscler. Thromb. Vas. Biol. 38, 2460–2473 (2018).
Metzger, C. E., Anand Narayanan, S., Phan, P. H. & Bloomfield, S. A. Hindlimb unloading causes regional loading-dependent changes in osteocyte inflammatory cytokines that are modulated by exogenous irisin treatment. NPJ Microgravity 6, 28 (2020).
Morey-Holton, E. R. & Globus, R. K. Hindlimb unloading rodent model: technical aspects. J. Appl. Physiol. 92, 1367–1377 (2002).
Wei, L. X., Zhou, J. N., Roberts, A. I. & Shi, Y. F. Lymphocyte reduction induced by hindlimb unloading: distinct mechanisms in the spleen and thymus. Cell Res. 13, 465–471 (2003).
Sun, W. et al. The mechanosensitive Piezo1 channel is required for bone formation. ELife 8, https://doi.org/10.7554/eLife.47454 (2019).
Li, B. et al. FARS2 Deficiency causes cardiomyopathy by disrupting mitochondrial homeostasis and the mitochondrial quality control system. Circulation 149, 1268–1284 (2024).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Huang, C. et al. Sensory ASIC3 channel exacerbates psoriatic inflammation via a neurogenic pathway in female mice. Nat. Commun. 15, 5288 (2024).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (grants 82171866-J.B., 82271893-Y.W., 82302090-K.C); the Qinchuangyuan “Scientist + Engineer” Team Development Program of the Shaanxi Provincial Department of Science and Technology (grants 2022KXJ-123-Y.W.); the Natural Science Foundation of Shaanxi Province (grants 2023-JC-YB-819-F.L.); “Clinical Medicine + X” research program of Xijing hospital (grants LHJJ24KQ03-Y.L.).
Author information
Authors and Affiliations
Contributions
J.B., Y.C., Y.W., and K.C. supervised the project. B.L. and K.Z. were major contributors in writing, reviewing, and editing the manuscript. B.L., X.W., Y.L., and Y.J.C. established the models. F.L., H.Z., Q.L., and Z.M. performed experiments. B.L., K.Z., and A.Y. analyzed the data. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Wei Kong, Sergey Tumanov, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, B., Zhou, K., Yin, A. et al. SREBP1-mediated lipogenesis promotes dedifferentiation and senescence of vascular smooth muscle cells through epigenetic remodeling. Nat Commun 17, 835 (2026). https://doi.org/10.1038/s41467-025-67534-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67534-6