Abstract
Cryptochromes (CRYs) are photolyase-like blue-light/ultraviolet-A (UV-A) receptors that regulate diverse aspects of plant growth and development. Maize (Zea mays), a major crop often grown under high UV-B radiation, harbors four copies of CRY. However, whether CRYs in maize have evolved to improve UV tolerance or acquire novel functions remains unclear. In this study, CRISPR-Cas9-engineered Zmcry mutants are used to investigate the functions of ZmCRYs in maize. The findings reveal that ZmCRYs act redundantly in mediating blue light signaling and inhibiting mesocotyl elongation. Furthermore, we demonstrate that ZmCRYs mediate blue light-enhanced UV-B stress tolerance in maize by upregulating the expression of genes involved in the biosynthesis of UV-B stress tolerance-related metabolites, including phenylpropanoids, flavonoids, and fatty acids. Further analyses show that blue light modulates both the accumulation and composition of epidermal waxes, suggesting that blue light enhances epidermal wax accumulation for UV-B stress tolerance. Notably, we identify that ZmCRY1 directly interacts with GLOSSY2 (GL2), a key acyltransferase in very-long-chain fatty acid metabolism, in a blue light-strengthened manner to mediate blue light-promoted C32 aldehyde accumulation, shedding new light on the enigma of the aldehyde-forming pathway in plants. These results highlight the critical roles of ZmCRY1s in mediating blue light-regulated epidermal wax biosynthesis and UV-B tolerance in Zea mays.
Similar content being viewed by others
Introduction
Cryptochromes (CRYs) are photolyase-like blue-light/UV-A receptors that share high sequence and structural similarity with DNA photolyases1,2,3, but they lack inherent DNA repair activity, distinguishing them functionally from their photolyase ancestors. These evolutionarily related flavoproteins perform diverse biological functions: DNA photolyase reverses far-UV light (200–300 nm)-induced damage (e.g., growth delay, mutagenesis, and cell death) via blue light (350–450 nm)-dependent photoreactivation across organisms from bacteria to plants and animals3,4, while CRYs regulate growth and development in plants and the circadian clock in both plants and animals5.
In Arabidopsis, the two primary CRY photoreceptors (CRY1 and CRY2) exhibit overlapping yet differentiated functions. They are involved redundantly in mediating photomorphogenesis and thermomorphogenesis, including hypocotyl elongation inhibition6,7,8. CRY1 also plays critical roles in stomatal opening, balances photosynthesis and defense by mediating pathogen-triggered stomatal closure9, and regulates plant height and shade avoidance responses—functions conserved across Arabidopsis, soybean, and maize10,11,12,13. In soybean, CRY1 is further essential for blue light-induced root nodule formation, promoting symbiotic nitrogen fixation in natural ecosystems14. CRY2 is well characterized for its role in regulating floral initiation15,16,17. A functionally equivalent dominant mutation, CRY2W374A, enhances species adaptation and rapid evolution, potentially serving as a genetic basis for ruderality in Brassicaceae18. CRY2 also participates in thermomorphogenesis, with its protein level regulated by both blue light and ambient temperature6,7. Notably, CRY2 acts in darkness to inhibit root growth, coordinating developmental processes between aboveground and underground organs19. Despite lacking DNA repair activity, CRYs collectively confer resistance to UV-B damage in plants20.
Maize, a major crop cultivated under high light intensity and elevated UV-B radiation, harbors three CRY1 homologs and one CRY2. High-resolution structures of the active photolyase-related (PHR) domain of ZmCRY1a and ZmCRY1c reveal conserved photoactive conformations similar to AtCRYs21,22, suggesting conserved photomorphogenesis roles that may inform breeding for high-density tolerance10,23. However, whether maize CRY paralogs have evolved novel functions or if maize employs alternative strategies to cope with UV-B stress remains unclear.
A primary line of defense against UV-B in terrestrial plants is the epidermal cuticle, where less than 10% of incident UV-B is transmitted24. The epicuticle contains UV-B-screening compounds—including flavonoids, anthocyanins, and phenolic acids, as well as waxes—that synergistically reflect and absorb UV light25,26,27. Cuticular wax accumulation and composition are modulated by environmental cues such as light wavelength, temperature, and osmotic stress28,29,30,31,32,33. Light is known to upregulate transcription of genes encoding wax biosynthetic enzymes28,32,34. Genetic analyses combined with weighted gene coexpression network analysis indicate that phytochromes mediate light regulation of epicuticle components and their evolution in land plants31. In faba bean, blue light shifts the dominant primary alcohol in cuticular wax32. However, whether CRYs mediate blue light-regulated wax biosynthesis or directly participate in this process remains unknown.
Cuticular waxes are predominantly composed of very long chain (VLC) aliphatics, derived from fatty acid synthesis and elongation. De novo synthesis of C16–C18 fatty acids occurs in plastids35, catalyzed by fatty acid synthases with chain-length-specific condensing enzymes (KAS III (C2–C4), KAS I (C4–C16), and KAS II (C16–C18))36,37,38. These fatty acids are then extended to VLC fatty acids (VLCFAs; ≥ C20) in the endoplasmic reticulum (ER) via multienzyme elongase complexes35,39, with β-ketoacyl-CoA synthases (KCSs) dictating chain-length specificity40,41. Key KCS members—such as KCS5/ECERIFERUM 60 (CER60) and KCS6/CER6—promote elongation beyond C26 (Arabidopsis) or C28, in cooperation with putative BAHD acyltransferases like Arabidopsis CER2, CER2-LIKE, and maize GL2, GL2-LIKE42,43,44,45,46.
VLC Fatty Acyl-CoAs (VLC acyl-CoAs) are processed into wax components via two pathways47: the alcohol-forming pathway (producing even-chain primary alcohols and alkyl esters) and the alkane-forming pathway (producing aldehydes, odd-chain alkanes, secondary alcohols, and ketones)47,48. Alkanes are proposed to derive from VLC aldehydes via decarbonylation, catalyzed by a CER1-CER3 heterodimer complex with cytochrome b5 (CYTB5) as a cofactor49,50,51,52,53. Fatty alcohol oxidase 3 (FAO3) and FAO4b also contribute to aldehyde production by suppressing primary alcohol accumulation54. However, the enzymes responsible for VLC aldehyde formation and the precise mechanism of VLC alkane synthesis remain to be further investigated.
Notably, yeast reconstitution experiments suggest that CER2 and CER2-LIKE enable CER6 to synthesize longer-chain VLCFAs, while subsequent studies suggest the yeast endogenous FAE complex may be associated with this extension46. Based on genetic evidence demonstrating GL2-mediated alterations in VLCFA composition, coupled with yeast reconstitution experiments42,43,44,45,46, it is hypothesized that maize GL2 (and GL2-Like) and Arabidopsis CER2 (and CER2-Like) function in the VLCFA elongation pathway—upstream of aldehyde, alcohol, or hydrocarbon formation—potentially through interaction with the KCS6-composed FAE complex. GLOSSY2 (GL2), a maize putative BAHD acyltransferase, may play a unique role in alkane pathway regulation. Overexpression of GL2 in Arabidopsis cer2-5 mutant or Col-0 induces accumulation of C32 aldehyde, C31 alkane, C31 secondary alcohol, and C31 ketone—components not observed upon overexpression of AtCER2 or maize GL2-LIKE55. Functional divergence between GL2 and GL2-LIKE is further highlighted by their differential dependence on the BAHD catalytic HXXXDX motif: this motif is required for GL2-LIKE activity but not for GL2, indicating evolutionary specialization55. However, the specific biochemical functions of GL2 in fatty acid elongation remain unclear.
This study investigates the roles of ZmCRY1s in mediating blue light-enhanced UV-B tolerance in maize. We show that ZmCRYs regulate transcription of genes involved in the biosynthesis of UV-B-protective metabolites (phenylpropanoids, flavonoids, anthocyanins, and fatty acids56) and modulate epidermal wax accumulation and composition. Furthermore, ZmCRY1 directly interacts with GL2 in a blue light-strengthened manner to promote C32 aldehyde formation, shedding light on VLC aldehyde generation. These findings reveal how blue light organizes UV resistance at the plant surface, potentially enhancing overall UV-B tolerance.
Results
ZmCRYs mediate blue light signaling in maize
To investigate the function of CRYs in maize, we searched maize genome sequence databases and classified the ZmCRY family. Maize CRYs are encoded by a multigene family comprising four members, which are phylogenetically classified as ZmCRY1s (ZmCRY1a, ZmCRY1b, and ZmCRY1c) and ZmCRY2 (Fig. 1a). ZmCRY1a and ZmCRY1b share 92% amino acid sequence identity, indicating a close relationship and likely functional similarity, whereas ZmCRY1c shares 78% and 75% amino acid sequence identity with ZmCRY1a and ZmCRY1b, respectively. Subcellular localization analysis revealed that ZmCRY1a and ZmCRY1b are predominantly nuclear, with minor cytoplasmic presence. In contrast, ZmCRY1c localizes to both compartments, with a significantly higher proportion in the cytoplasm (Supplementary Fig. 1a).
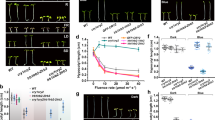
a Phylogenetic tree of CRY homologues among Maize (Zea mays), Rice (Oryza sativa), Soybean (Glycine max)77,78 and Arabidopsis (Arabidopsis thaliana). Distances were estimated using the neighbor-joining algorithm. The numbers at the nodes represent the percentage of 1000 bootstraps. The scale bar indicates the average number of amino acid substitutions per site. b–e Representative photographs of indicated genotypes grown for 7 days under continued blue light (b, 20 μmol m−2 s−1), red light (c, 20 μmol m−2 s−1), white light (d, 40 μmol m−2 s−1) and dark (e). White arrow heads indicate the boundary between mesocotyl and first internode. Scale bars = 5 cm. Mesocotyl lengths of the indicated genotypes under different light conditions are shown on the right. Error bars represent the standard deviation (SD, n = 9). Data are presented as mean values ± SD. Different letters indicate statistically significant differences; no significant differences between genotypes under red light, as determined by one-way ANOVA with Tukey’s multiple comparisons test (P < 0.05).
To elucidate their biological functions, we obtained uniform Mutator (Mu) insertion lines of CRY1s from the Maize Genetics Cooperation Stock Center. These lines carry Mu insertions at 1377 bp (CRY1a: UFMu-08910, designated cry1a-mu), −10 bp (CRY1b: UFMu-09028, designated cry1b-mu), and 426 bp (CRY1c: UFMu-08243, designated cry1c-mu) relative to their respective start codons. PCR, RT-PCR, and Sanger sequencing confirmed the presence of Mu elements and their impact on CRY1s expression, with no detectable corresponding CRY1 transcripts in their Mu insertion lines (Supplementary Fig. 1b–d). Phenotypic analysis showed that cry1a-mu and cry1c-mu single mutants exhibit longer mesocotyls than the wild-type W22 inbred line under blue light and white light. The cry1a cry1c-mu double mutant displays even longer mesocotyls than the single mutants and wild type under these light conditions, whereas no significant differences were observed among the Mu insertion lines under red light or in darkness (Supplementary Fig. 1e–h). These results suggest that these CRY1s may function redundantly in mediating blue light signaling in maize.
To generate multiple cry knockout mutants, we used CRISPR-Cas9 technology to target the four cryptochrome genes in the B73 background. Guide RNA 1 (gRNA1) was designed to target conserved regions of ZmCRY1s, and gRNA2 targeted ZmCRY2. We identified Zmcry quadruple mutants (Zmcry-q) with mutations including base deletions in ZmCRY1a (d45, 45-base deletion), ZmCRY1b (d124), ZmCRY1c (d14), as well as a 15-base deletion followed by a 19-base insertion in ZmCRY2 (d15, i19) (Supplementary Fig. 2a). Zmcry-q was backcrossed to B73 and self-pollinated to remove the Cas9 transgene, enabling the isolation of various single, double, and Zmcry1s triple (Zmcry1abc) mutants. Phenotypic analysis under blue light and white light revealed that, compared to wild-type B73, the single mutants Zmcry1a and Zmcry1c exhibit longer mesocotyls. Mesocotyl length increased as the number of functional ZmCRYs decreased across double (Zmcry1ab, Zmcry1ac, Zmcry1bc, Zmcry1acry2), triple (Zmcry1abc), and quadruple (Zmcry-q) mutants, and no significant differences were observed among these genotypes under red light and in darkness (Fig. 1b–e).
We generated overexpression lines of 35S::YFP-ZmCRY1a (CRY1a-OE) and 35S::YFP-ZmCRY1c (CRY1c-OE) in the maize Hi-II background. Three independent transgenic lines were recurrently backcrossed into Chang7-2 for five generations. Phenotypic analysis showed that all overexpression lines in the Chang7-2 background exhibit shorter mesocotyls than the wild type specifically under blue light and white light, while no significant differences were observed under red light or in darkness (Supplementary Fig. 2b–f). For phenotypic analysis within a uniform genetic background, randomly selected CRY1a-OE and CRY1c-OE lines were recurrently backcrossed into the B73 background for five generations. These lines also showed shorter mesocotyls than B73 (Fig. 1b–e). Collectively, these results indicate that ZmCRYs function redundantly in mediating blue light signaling to regulate maize photomorphogenesis.
To determine whether the blue light receptor function of ZmCRYs is conserved with that of Arabidopsis CRYs (AtCRYs), we overexpressed ZmCRYs in Arabidopsis. YFP-ZmCRY1s mediated blue light inhibition of hypocotyl elongation in transgenic Arabidopsis seedlings, whereas YFP-ZmCRY2 did not (Supplementary Fig. 3a–d). Photoexcited AtCRY2 undergoes polyubiquitination and subsequent degradation via the 26S proteasome57, and AtCRY1 has recently been shown to degrade in response to high-intensity blue light58. To assess ZmCRYs protein stability under blue light, we constructed ZmCRY-Flag overexpression lines in Arabidopsis, using endogenous AtCRY2 protein and an AtCRY1-Flag overexpression line as controls. Transgenic seedlings were pretreated in darkness for 48 h, then exposed to medium blue light (20 μmol m–2 s–1) or high blue light (100 μmol m–2 s–1) for a 1-h time course. Under medium blue light, ZmCRY2 protein levels decreased similarly to AtCRY2, while ZmCRY1s levels remained unchanged (Supplementary Fig. 3e, f). Under high blue light, ZmCRY1a and ZmCRY1b underwent degradation analogous to AtCRY1, whereas ZmCRY1c remained relatively stable (Supplementary Fig. 3g, h). The stability of ZmCRY1c may be attributed to its predominantly cytoplasmic localization, whereas the E3 ubiquitin ligases Light-Response Bric-a-Brack/Tramtrack/Broads (LRBs) responsible for CRY degradation are nuclear-localized7,58,59. These results suggest that ZmCRY1s and ZmCRY2 are functionally conserved with their Arabidopsis orthologs as blue light receptors.
ZmCRYs mediate the blue light-induced transcription of genes involved in UV-B stress tolerance in maize
Current understanding indicates that most CRY-mediated blue light responses in plants are associated with changes in nuclear gene expression. To explore the ZmCRY-dependent blue light-regulated transcriptome, we conducted transcriptome analysis using 7-day-old seedlings of B73 and Zmcry-q grown under long day (LD, 16-h white light/8-h dark) conditions. The seedlings were pretreated in darkness for 48 h, followed by either 1-h blue light (20 μmol m–2 s–1) irradiation or maintenance in darkness. Transcriptome analysis revealed that blue light mediated a total of 5764 differentially expressed genes (DEGs) in the comparison between B73_Blue and B73_Dark (Supplementary Data 1). Venn diagram analysis showed that 45.49% (2,622 genes) of these DEGs observed in wild-type plants under blue light compared to darkness were not differentially expressed in Zmcry-q mutants, indicating that these genes are dependent on the presence of one or more CRY genes (Fig. 2a). A heatmap generated from ZmCRY-mediated DEGs clearly illustrated the blue light-regulated genes mediated by ZmCRYs (Fig. 2a, b).
a Venn diagram showing the overlap between sets of DEGs from blue light treatment or not in B73 (B73_Blue vs. B73_Dark), and in cryq (cryq_Blue vs. cryq_Dark) at 28 °C. b Heatmap generated from ZmCRYs-mediated DEGs. c KEGG pathway enrichment analysis of blue light-upregulated genes. Significantly enriched KEGG pathways were defined as overrepresented terms in the blue light-upregulated gene set, identified using a one-sided (right-tailed) Fisher’s exact test with a liberal statistical significance threshold of P < 0.05. d RT-qPCR validates the already known blue light signaling pathway regulators and the up-regulated fatty acid biosynthesis enzymes detected in the transcriptome. Seven-day LD-grown B73 and Zmcry-q, pretreated with 48-h dark, then moved to blue light (20 μmol m−2 s−1) for a 2-h time course. ZmUBQ served as internal control. Error bars, SD of three biological replicates. Data are presented as mean values ± SD.
Blue light regulates various aspects of plant metabolism, development, and morphology by modulating the expression of numerous genes. In addition to the conserved major blue light responses observed in Arabidopsis, Gene Ontology (GO) enrichment analysis revealed several distinct responses in maize, including biosynthesis and metabolic processes of multiple compounds (Supplementary Fig. 4a, b and Supplementary Data 2). This suggests that blue light may promote fatty acid accumulation by enhancing fatty acid biosynthesis while repressing fatty acid β-oxidation. Furthermore, Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of blue light-upregulated genes in B73 identified several metabolites associated with UV-B stress tolerance, including those involved in phenylpropanoid biosynthesis, flavonoid biosynthesis, and fatty acid biosynthesis (Fig. 2c and Supplementary Data 3). These results imply that fatty acid accumulation may be an important regulatory target of blue light signaling in maize.
To validate the transcriptome data, we performed quantitative reverse transcription PCR (RT-qPCR) using seedlings transferred from darkness to blue light (20 μmol m−2 s−1) over a 2-h time course. For known blue light signaling pathway regulators detected in the transcriptome, such as the HY5 homolog gene BASIC LEUCINE ZIPPER 61 (bZIP61) and bZIP80, the AtCOP1 homolog gene UBIQUITIN-PROTEIN LIGASE 13 (ZmUPL13) and ZmUPL14, SUPPRESSOR OF PHYTOCHROME A 1 (SPA1)-related 3 and SPA1-related 4, REPRESSOR OF UV-B PHOTOMORPHOGENESIS 2 (ZmRUP2) and CRY3, their expression was upregulated in B73 after blue light irradiation, but this induction was abolished in Zmcry-q mutants (Fig. 2d and Supplementary Fig. 4c). We further verified the expression of genes associated with de novo fatty acid synthesis (fatty acyl-acyl carrier protein (ACP) thioesterase B (FatB), β-ketoacyl-ACP synthase I (KAS I), and KAS II36,37), those involved in fatty acid elongation (KCS3, KCS4), and genes related to cuticular wax biosynthesis (GL1, GL1-5, and GL4). These genes were induced by blue light, with partial gene induction dependent on CRYs (Fig. 2d and Supplementary Fig. 4d). These findings suggest that blue light may induce fatty acid biosynthesis and wax accumulation through the regulation of gene expression31,32.
Blue light affects the accumulation and the composition of epidermal wax in maize
Given that blue light induces the expression of genes related to fatty acid biosynthesis and wax accumulation via ZmCRYs, we hypothesized that blue light and CRYs might modulate wax accumulation. To test this, we analyzed the effects of different light wavelengths on wax accumulation in maize. Seven-day-old dark-grown seedlings were exposed to various light treatments (UV-B (1 μmol m−2 s−1), blue (20 μmol m−2 s−1), red (20 μmol m−2 s−1), and far-red light (10 μmol m−2 s−1)) for 48 h, after which their epidermal wax compositions were measured. Compared to darkness, all light treatments significantly promoted epidermal wax accumulation (Fig. 3a). The dominant C32 primary alcohol showed a marked increase across all light treatments. Notably, blue light exposure led to a 1.86-fold increase in total cuticular wax, a 0.35-fold decrease in C28 primary alcohol, a 4.56-fold increase in C32 primary alcohol, and the appearance of C32 aldehyde. This finding indicates that the changes in wax composition under blue light are specifically attributed to increased chain lengths of cuticular wax products. Additionally, blue light and UV-B irradiation exerted consistent effects on wax composition. These results indicate that UV-B, blue, red, and far-red light all enhance epidermal wax accumulation, while exerting distinct effects on wax composition.
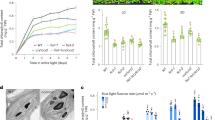
a Epidermal wax composition analysis under different light conditions. The 7-day-old dark-grown seedlings treated with 48 h radiation under indicated lights. Blue (Blue light, 20 μmol m−2 s−1), Red (Red light, 20 μmol m−2 s−1), Far-red (Far-red light, 10 μmol m−2 s−1), NUV (Narrow-band UV-B light, 1 μmol m−2 s−1). Error bars represent SD (n = 3). Data are presented as mean values ± SD. The asterisks indicate a significant difference from dark grown B73 based on a two-sided Student’s t-test (*P < 0.05, **P < 0.01). b Epidermal wax composition affected by CRY mediated blue light were analyzed. Seven-day-old YFP-CRY1c, B73, and Zmcry-q were grown in dark or continued blue light (20 μmol m−2 s−1). Error bars represent SD. (n = 3). Data are presented as mean values ± SD. The asterisks indicate a significant difference based on a two-sided Student’s t-test (*P < 0.05, **P < 0.01). Different letters in total wax indicate statistically significant differences, as determined by one-way ANOVA with Tukey’s multiple comparisons test (P < 0.05). c, d CRY1c enhances maize tolerance to UV-B stress. c Plants of the indicated genotypes were grown in soil under long-day (16/8) conditions in blue light (20 μmol m−2 s−1) for 10 days and irradiated with (+UV) or without (−UV) broadband UV-B (5 μmol m−2 s−1) for 6 h on day 11 and allowed to recover for 2 days in blue light. d Measurement of PSII maximal quantum yield (Fv/Fm) of the first leaves of the indicated genotypes shown in (c). Fv/Fm was measured and quantified with an imaging fluorometer. Error bars represent SD (n = 5 ( − UV), and n = 11 ( + UV)). Data are presented as mean values ± SD. Different letters indicate statistically significant differences, as determined by two-way ANOVA with Šídák’s multiple comparisons test (P < 0.05).
To determine whether ZmCRYs mediate blue light-regulated wax accumulation, we measured epidermal wax compositions in 7-day-old seedlings of CRY1c-OE, B73, and Zmcry-q mutants grown in either darkness or continuous blue light. No significant difference in total wax content was observed among genotypes under both darkness and blue light conditions, suggesting that blue light-promoted epidermal wax accumulation is not ZmCRY-dependent (Fig. 3b) and that other blue light receptor families may redundantly regulate this process. Compared to darkness, blue light exposure consistently increased total cuticular wax across genotypes: in B73, this was accompanied by a 0.27-fold decrease in C32 fatty acid and a 2.82-fold increase in C32 primary alcohol. C32 aldehyde accumulation was undetectable in any genotype under darkness. Under blue light, however, C32 aldehyde levels were significantly higher in CRY1c-OE line (1.45 ± 0.26 μg cm–2) than in B73 (0.83 ± 0.24 μg cm–2) and much lower in Zmcry-q mutant (0.33 ± 0.03 μg cm–2). These results indicate that the blue light-induced C32 aldehyde biosynthesis is primarily ZmCRY-dependent (Fig. 3b). Furthermore, they suggest that ZmCRYs may mediate blue light responses involved in the formation of very-long-chain (VLC) aldehydes, a process that requires further clarification47.
ZmCRYs enhance the UV-B stress tolerance in maize
Maize is cultivated under high light intensity year-round and thus must cope with elevated UV-B radiation. Our transcriptome analyses support the notion that blue light contributes to UV-B tolerance at the transcriptional level, as previously reported20. To evaluate the potential UV-B protective capacity of the blue light-ZmCRY pathway, we measured the maximum quantum yield of Photosystem II (Fv/Fm) as an indicator of UV-B stress tolerance. For this assay, 10-day-old seedlings of CRY1c-OE, B73, and Zmcry-q grown under LD blue light (20 μmol m–2 s–1) were subjected to two broadband UV-B treatments of varying intensities: (1) high intensity (60 μmol·m–2·s–1) for 10 h, followed by 1-h dark adaptation before Fv/Fm measurement; (2) medium intensity (5 μmol·m–2·s–1) for 6 h, followed by a 2-day recovery period before Fv/Fm measurement and an additional week of growth to assess survival rate. Results showed that Zmcry-q was more sensitive to UV-B stress than B73 under both treatments, while CRY1c-OE exhibited greater tolerance. After 1-h dark adaptation following high-intensity UV-B irradiation, the average Fv/Fm values were 0.561 (B73), 0.681 (CRY1c-OE), and 0.435 (Zmcry-q) (Supplementary Fig. 5b, c). After a 2-day recovery following medium-intensity UV-B irradiation, the average Fv/Fm values were 0.516 (B73), 0.718 (CRY1c-OE), and 0.148 (Zmcry-q) (Fig. 3c, d). Additionally, the average survival rate of CRY1c-OE (90.5%) was significantly higher than that of B73 (61.9%), whereas Zmcry-q showed the lowest survival rate (33.3%) after medium-intensity UV-B irradiation (Supplementary Fig. 5d–f). In the Chang7-2 background, both CRY1a-OE and CRY1c-OE lines also displayed enhanced UV-B tolerance compared to wild-type Chang7-2 (Supplementary Fig. 5g, h). Collectively, these findings indicate that ZmCRYs mediate blue light-enhanced UV-B tolerance in maize.
Epidermal wax acts as a primary defense against UV-B radiation33. Blue light modulates both the accumulation and composition of epidermal wax in maize; however, ZmCRYs primarily mediate the blue light-regulated changes in wax composition rather than total wax accumulation. The specific mechanisms by which ZmCRYs influence wax composition, as well as the role of C32 aldehyde or its downstream derivatives in UV-B tolerance, require further investigation.
ZmCRY1 physically interacts with GLOSSY2 in a blue light-enhanced manner
To investigate how ZmCRY influences wax composition and the unique functions of ZmCRY1c due to its distinct cytoplasmic localization, we employed two strategies to identify the ZmCRY1c-interacting proteins under blue light: yeast two-hybrid (Y2H) screening in vitro and in vivo co-immunoprecipitation coupled with mass spectrometry (IP–MS). In the Y2H assay, using ZmCRY1c as bait to screen a maize cDNA library under blue light (20 μmol·m–2·s–1), we sequenced 230 positive clones and identified 87 proteins that interact with CRY1c in yeast (Fig. 4a, b and Supplementary Data 4). Among these, 4 clones encoded various fragments of GLOSSY2 (GL2), a member of the BAHD acyltransferase superfamily involved in the elongation-reductive pathway of very-long-chain fatty acids (VLCFAs). In the IP-MS experiment, we affinity-purified ZmCRY1c complexes from YFP-ZmCRY1c overexpression line (CRY1c-OE) under blue light (20 μmol·m–2·s–1) or in darkness. Analysis of co-purified proteins identified known interactors such as the COP1 homologue ZmUPL13 and the gibberellin receptor GID1L260,61,62, as well as GL2, which was previously identified via the Y2H assay (Supplementary Data 5). GL2 specifically enhances the elongation of C30 and C32 VLC acyl-CoA63 and indirectly promotes the formation of C32 aldehyde, C31 alkane, C31 secondary alcohol, and C31 ketone55. Consequently, we selected GL2 as a candidate and performed further protein–protein interaction assays to determine whether blue light enhances the interaction between ZmCRY1c and GL2.
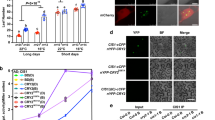
a Histidine auxotrophy assays showing the interaction between ZmCRY1c and GL2 in a blue light-strengthened manner. Yeast cells were grown at –LWHA medium in the dark or under blue light (20 μmol m–2 s–1). P53/T was a positive control pair. b β-gal assays of yeast cells grown on –LW medium at 28 °C in darkness (D) or under blue light (B, 20 μmol m–2 s–1) for the indicated times. Error bars represent the SD of three independent biological replicates. Data are presented as mean values ± SD. ZmCRY1cD381A is a site-specific mutant of ZmCRY1c that cannot be activated by blue light. c Co-localization of ZmCRY1c and GL2 in the cytoplasm in maize protoplasts. BF, brightfield. Merge, overlay of the GFP, CFP, and brightfield images. Scale bar = 10 μm. d Gray-value analysis of fluorescent signals along the dashed diagonal line in the left images. e, f BiLC and BIFC assays showing in vivo protein interactions between ZmCRY1c and GL2. Leaf epidermal cells of Nicotiana benthamiana were co-transformed to express the indicated fusion proteins. In BIFC assay, ZmCRY1cD381A-cCFP and nYFP-KCS4 served as negative controls. Scale bar = 200 μm. g Co-IP assays showing that ZmCRY1c interacts with GL2 in a blue light-strengthened manner in plant cells. Seven-day-old 28 °C LD-grown seedlings were pretreated in darkness for 48 h, then exposed to blue light (B, 20 μmol m–2 s–1) for 2 h. Input: immunoblots showing the abundance of GL2 and YFP-ZmCRY1c in total protein extracts. GL2 immunoprecipitation (IP): IP products precipitated by the anti-GL2 antibody. Total proteins (Input) or IP products of GL2-beads (GL2 IP) were probed in immunoblots with an anti-GL2 or anti-GFP antibody. In a and c–g, three independent experiments were performed with similar results.
Y2H assays revealed that ZmCRY1c and GL2 exhibit a strengthened physical interaction under blue light but a weak interaction in darkness (Fig. 4a). Quantification via β-galactosidase assay showed that the interaction was strengthened only when cells were transferred from darkness to blue light, not when kept in darkness (Fig. 4b). No interaction was detected between GL2 and the flavin-deficient ZmCRY1c variant ZmCRY1cD381A64 (Fig. 4b). ZmCRY1c colocalizes with GL2 in the cytoplasm of Nicotiana benthamiana leaves and maize leaf protoplasts (Fig. 4c, d and Supplementary Fig. 6a, b). We confirmed this interaction in plant cells using bimolecular fluorescence complementation (BiFC) and bimolecular luminescence complementation (BiLC) assays. Strong yellow fluorescent protein (YFP) fluorescence was observed in the cytoplasm of N. benthamiana leaves transiently co-infiltrated with ZmCRY1c-cCFP and nYFP-GL2 constructs, indicating a robust interaction, whereas leaves co-infiltrated with ZmCRY1cD381A-cCFP and nYFP-GL2 showed no interaction (Fig. 4e and Supplementary Fig. 6c). Similarly, strong luminescence was observed in leaves co-infiltrated with ZmCRY1c-nLUC and cLUC-GL2 constructs in BiLC assays (Fig. 4f). Additionally, ZmCRY1a and ZmCRY1b also interacted with GL2 in BiLC assays, while ZmCRY2 showed a weak interaction with GL2 in the BiLC assay (Supplementary Fig. 6d). No interaction was detected between KCS4 and ZmCRY1c in BiFC and BiLC assays (Fig. 4e and Supplementary Fig. 6e). The GL2 homologue, GL2-LIKE, showed a weak interaction with ZmCRY1c, as indicated by faint luminescence in the BiLC assay compared to GL2, but yeast containing ZmCRY1c and GL2-LIKE failed to survive on histidine-deficient (–LWHA) medium in Y2H assays (Supplementary Fig. 6f, g). The in vivo interaction of ZmCRY1c and GL2 was confirmed by co-immunoprecipitation assay. Seedlings pretreated in darkness for 48 h were exposed to blue light for 2 h or kept in darkness before harvesting. Results showed that ZmCRY1c co-precipitated with GL2 in blue light-treated samples, whereas minimal co-precipitation was detected in darkness-maintained samples (Fig. 4g). Collectively, these results reiterate that blue light promotes the interaction between ZmCRY1c and GL2.
The N-terminal PHR domain of ZmCRY1c, which encompasses the chromophore-binding region, was sufficient to mediate the interaction with GL2 in yeast cells (Supplementary Fig. 6h), aligning with the observation that their interaction is blue light-strengthened. Subsequently, Yaqi L. et al. have determined the crystal structure of the constitutively active CRY1c PHR domain (ZmCRY1c-PHRW368A22) in complex with GL2, which provides structural insights into their interaction64.
ZmCRY1 modulates aldehyde accumulation together with GLOSSY2
As described64, the interaction interface between ZmCRY1c-PHRW368A and GL2 primarily involves the α16-α17 loop, α17 of ZmCRY1c (residues 396–419), as well as the α2, β1-β2 loop of GL2 (residues 25–33, 70–76). To validate this, we screened for amino acid site mutations in the Maize EMS-induced Mutant Database (MEMD)65. Among the 14 EMS mutants of GL2, two mutants, gl2V24M and gl2G106D, exhibited a water-beading phenotype similar to that of gl2-ref mutant66 (Fig. 5a, b and Supplementary Fig. 7a). These mutations are located in close proximity to Phe71 of GL2, a residue directly interacting with ZmCRY1c-PHRW368A64(Supplementary Fig. 7b). To assess the impact of these mutations on protein interaction, we cloned the mutated cDNAs and performed interaction assays. In Y2H assays under blue light, yeast co-transformed with constructs harboring GL2G106D failed to grow on –LWHA medium, indicating that this mutation severely disrupted the interaction with ZmCRY1c (Fig. 5c). In contrast, the GL2V24M mutation had a milder effect on the interaction compared to the wild-type GL2 (Supplementary Fig. 7c). BiLC assays further confirmed this: the luminescent signal from ZmCRY1c-nLUC and cLUC-GL2G106D was faint compared to that with cLUC-GL2, reflecting a significant reduction in interaction strength, while cLUC-GL2V24M showed a milder decrease in interaction with ZmCRY1c-nLUC (Fig. 5d). Additionally, pull-down assays with ZmCRY1c-PHRW368A corroborated these findings: the G106D mutation nearly abolished the interaction, while V24M had a weaker effect (Fig. 5e). Consistent with the defects in protein-protein interaction, these gl2 site mutants displayed epidermal wax compositions comparable to gl2-ref, characterized by reduced levels of major components such as C32 primary alcohol and C32 aldehyde, alongside accumulated C22–C30 VLCFAs (Fig. 5f). These results highlight the importance of the interaction interfaces (INT) of GL2 with ZmCRY1c for GL2’s function.
a Schematic illustration of the GL2 protein sequence with all EMS mutant alleles. b Water beading phenotype of the gl2-ref, gl2G106D, and gl2V24M mutants and wild-type B73 after water spraying. c Y2H histidine auxotrophy assays showing that the site mutation G106D largely disrupts the interaction of GL2 with ZmCRY1c. Yeast cells were grown on –LWHA medium under blue light (20 μmol m–2 s–1). Three independent experiments were performed with similar results. d BiLC assay showing the site mutations V24M and G106D of GL2 affect its interaction with ZmCRY1c to different degrees. A representative picture is shown, and the relative luciferase activities are shown on the right. Leaf epidermal cells of Nicotiana benthamiana were co-transformed with the indicated fusion proteins. Error bars represent the SD of three independent biological replicates. Data are presented as mean values ± SD. The asterisks indicate a significant difference based on a two-sided Student’s t-test (*P < 0.05, **P < 0.01). e HIS-pull down assay on the interaction between ZmCRY1c-PHRW368A and GL2 variants. SDS-PAGE gel shows the influence of His-GL2 site mutations on the complex formation. The numbers below the gels denote the ratio calculated by ZmCRY1c-PHRW368A to His-GL2. Three independent experiments were performed with similar results. f Epidermal wax composition analysis of 10-day-old LD white light (50 μmol m−2 s−1)-grown seedlings of the indicated genotypes. Error bars represent SD (n = 3). Data are presented as mean values ± SD. The asterisks indicate a significant difference from B73 based on a two-sided Student’s t-test (*P < 0.05, **P < 0.01).
Previous studies have shown that overexpression of GL2 in Arabidopsis induces the production of C32 aldehyde, C31 alkane, C31 secondary alcohol, and C31 ketone55. To determine if blue light regulates GL2-mediated C32 aldehyde accumulation, we generated GL2 overexpression lines in the B73 background and selected three independent lines for wax composition analysis (Supplementary Fig. 8a). Epidermal waxes were measured in 7-day-old seedlings grown in darkness or under blue light (20 μmol m–2 s–1). Results showed that C32 aldehyde accumulation in GL2 overexpression lines was significantly induced by blue light but undetectable in darkness, whereas C32 primary alcohol induction in GL2 overexpression lines was light-independent (Fig. 6a). This indicates that blue light regulates GL2’s function in promoting C32 aldehyde accumulation.
a Epidermal wax composition analysis of GL2 overexpression lines. D: 7-day-old dark-grown or B: LD blue light-grown seedlings of the indicated genotypes. Error bars represent SD (n = 3). Data are presented as mean values ± SD. The asterisks indicate a significant difference from B73 based on a two-sided Student’s t-test (*P < 0.05, **P < 0.01). b Epidermal wax composition analysis of 7-day-old LD blue light (20 μmol m−2 s−1)-grown seedlings of the indicated genotypes. Error bars represent SD (n = 3). Data are presented as mean values ± SD. The asterisks indicate statistically significant differences from CRY1c-OE, as determined by two-way ANOVA with Dunnett’s multiple comparisons test (*P < 0.01, **P < 0.001). c Analysis of UV-B stress tolerance in maize. Plants of the indicated genotypes were grown in soil under long-day (16/8 h) conditions in white light (50 μmol m−2 s−1) for 10 days and irradiated with (+UV) or without (−UV) broadband UV-B (5 μmol m−2 s−1) for 6 h per day for 2 days, then allowed to recover for 2 days in white light. d Measurement of PSII maximal quantum yield (Fv/Fm) of the first leaves of the indicated genotypes shown in (c). Fv/Fm was measured and quantified with an imaging fluorometer. Error bars represent SD (n = 5 for −UV, and n = 8 for +UV). Data are presented as mean values ± SD. Different letters indicate statistically significant differences, as determined by one-way ANOVA with Tukey’s multiple comparisons test (P < 0.05). e, f BiLC assay showing the competitive interaction of ZmCRY1c and KCS6 with GL2. A representative picture is shown, and the relative luciferase activities are shown below the picture. Leaf epidermal cells of Nicotiana benthamiana were co-transformed with the indicated fusion proteins. Error bars represent the SD of three independent biological replicates. Data are presented as mean values ± SD. The asterisks indicate a significant difference based on a two-sided Student’s t-test (*P < 0.05, **P < 0.01). ns, no statistically significant differences. g A hypothetical model for blue light-regulated wax biosynthesis via the ZmCRY1-GL2 signaling pathway.
To verify whether C32 aldehyde accumulation in CRY1c-OE lines depends on GL2, we analyzed YFP-ZmCRY1c overexpression lines in gl2-ref mutant background (CRY1c-OE/gl2-ref) (Supplementary Fig. 8b). Seven-day-old dark-grown seedlings were irradiated with blue light (20 μmol m–2 s–1) for 48 h, after which epidermal waxes were measured. The results showed that C32 aldehyde accumulation in CRY1c-OE line was abolished in the absence of functional GL2, demonstrating that ZmCRY1c regulates C32 aldehyde accumulation in a GL2-dependent manner (Fig. 6b).
To evaluate the UV-B tolerance of wax-deficient gl2 variants and GL2’s contribution to CRY-mediated UV-B tolerance, 10-day-old seedlings of B73, gl2 variants, CRY1c-OE, and CRY1c-OE/gl2-ref grown under LD blue light (20 μmol m–2 s–1) were treated with high-intensity broadband UV-B light (60 μmol m–2 s–1) for 10 h, followed by 1-h dark adaptation before Fv/Fm measurement (Supplementary Fig. 8c, d). Separately, these genotypes grown under LD white light (50 μmol m–2 s–1) were treated with medium-intensity broadband UV-B light (5 μmol m–2 s–1) for 6 h per day over 2 consecutive days, with a subsequent 2-day recovery period prior to Fv/Fm measurement (Fig. 6c, d). Results revealed that gl2 variants were more sensitive to UV-B stress than B73 under both treatment regimens. Additionally, the absence of functional GL2 attenuated the enhanced UV-B tolerance of CRY1c-OE; however, CRY1c-OE/gl2-ref still exhibited slightly greater resistance than B73 (Fig. 6c, d and Supplementary Fig. 8c, d). These findings underscore the multifaceted regulation of UV-B tolerance by CRY20.
GLOSSY2 plays dual roles in fatty acid elongation and wax maturation through competitive partner interactions
GL2/CER2 proteins function uniquely as cofactors with KCS6 in the biochemical and physiological processes of elongating VLC acyl-CoAs up to C3442,43,44,45. We hypothesized that ZmCRY1c might enhance the interactions between these proteins to facilitate VLC acyl-CoA elongation. To test this, we performed BiLC assays; no luminescence was observed in leaves co-infiltrated with ZmCRY1c-nLUC and cLUC-KCS6 constructs in BiLC assays (Supplementary Fig. 8e). Furthermore, co-expression of ZmCRY1c with KCS6-nLUC and cLUC-GL2 reduced the chemiluminescent signal; a similar reduction was observed when ZmKCS6 was co-expressed with ZmCRY1c-nLUC and cLUC-GL2 (Fig. 6e, f). These results suggest that ZmCRY1c and KCS6 may compete for binding to GL2 rather than forming a ternary complex. Collectively, these findings indicate that GL2 interacts with KCS6 to promote VLCFA elongation in darkness. Under blue light, activated ZmCRY1c can outcompete KCS6 for GL2 interaction, thereby enhancing aldehyde accumulation.
Our working model proposes that blue light-activated ZmCRY1 enhances phenylpropanoid biosynthesis, flavonoid biosynthesis, fatty acid biosynthesis, and cuticle synthesis by promoting the transcription of genes encoding related enzymes, including FatB, KAS I, KAS II, KCS, GL1, GL1-5, and GL4. Additionally, ZmCRY1 directly interacts with GL2 in a blue light-enhanced manner to facilitate C32 aldehyde accumulation (Fig. 6g). In this way, blue light affects the accumulation of phenylpropanoids, flavonoids, and cuticular waxes, collectively enhancing the UV-B tolerance of maize through the coordinated regulation of metabolite biosynthesis and epidermal barrier modification.
Discussion
Cryptochromes (CRYs) are blue-light receptors that regulate diverse aspects of plant growth and development, including photomorphogenesis8, plant height, shade avoidance10,11, floral initiation15,16,18, root length, and nodule formation14,19. Specifically, previous studies have reported that CRY1 mediates blue-light-induced expression and accumulation of RUP1 and RUP2, which regulate the balance between monomeric and homodimeric forms of the UV-B photoreceptor UVR8, thereby modulating UV-B signalling activity. These studies suggest that CRY1, phyB, and UVR8 redundantly regulate UV-B tolerance20. In the current work, we demonstrate that blue light-activated maize CRYs mediate blue light-regulated wax biosynthesis and directly interact with GL2 to facilitate C32 aldehyde accumulation (Fig. 6g), organizing UV stress resistance at the plant’s outermost barrier, potentially enhancing overall UV-B tolerance27.
RNA-seq analysis reveals that blue light regulates compound biosynthesis and metabolic processes in maize, whereas similar studies in Arabidopsis show fewer genes differentially regulated under blue light (Supplementary Fig. 4a, b and Data 2, 3). Yeast two-hybrid screening and IP-MS results further show that ZmCRY1c has the potential to interact with numerous proteins, regulating their functions in the cytoplasm or membrane systems (Supplementary Data 4). These findings underscore the need for further investigation into the roles of photoreceptors in metabolic processes, expanding our understanding of how light influences agricultural traits.
In addition to various light wavelengths affecting cuticular waxes in land plants31,32, we further demonstrate that UV-B, red, far-red, and blue light all significantly promote epidermal wax biosynthesis and accumulation (Fig. 3a). Specifically, blue light and UV-B irradiation induce consistent changes in wax composition. These observations collectively indicate that different light wavelengths exert both synergistic and specific regulatory effects on epidermal waxes, highlighting the complexity of light-mediated responses in plant cuticles.
We identified that photoactivated ZmCRY1 directly interacts with GL2, a putative BAHD family acyltransferase involved in VLCFA elongation, mediating blue light-regulated changes in epidermal wax composition. This uncovers a direct molecular link through which light signaling, via photoreceptors, regulates fatty acid metabolism, filling a critical knowledge gap in our understanding of how plants translate light cues into metabolic adjustments for stress defense. Notably, the ZmCRY1-GL2 interaction is not conserved, at least in Arabidopsis64, suggesting that the blue light-ZmCRY1-GL2 module that regulates epidermal wax composition may be a maize-specific pathway or an evolutionarily diverged mechanism. This uniqueness not only enriches our understanding of the evolutionary plasticity of light-regulated wax biosynthesis across plant lineages but also highlights the existence of lineage-specific adaptive strategies in integrating light signals for cuticle-mediated stress resistance.
A series of wax-deficient mutants have been identified and cloned in Arabidopsis, barley, maize, sorghum, and rice, revealing their potential functions in wax biosynthesis, transport, or regulation. Although the major steps of the wax biosynthetic pathway have been well characterized47,67,68, the fine-tuning regulation remains to be fully elucidated. The products of alcohol- and alkane-forming pathways generate variable wax components in response to environmental factors such as humidity, temperature, or other uncharacterized stimuli. In Arabidopsis, the SOH1–CER3–CER1 module has been identified as a key shunting mechanism. The CER3-CER1 heterodimer catalyzes the decarbonylation of aldehydes to alkanes under water-deficit stress49,50,51,52,53, while the CER3-SOH1 complex converts aldehydes into 1-alcohols in response to high-temperature stimuli29. These findings highlight aldehydes as central intermediates in the flexible shunting between alcohol- and alkane-forming pathways, emphasizing their importance in plant adaptation to environmental changes. Here we report a mechanism by which blue light promotes the interaction between ZmCRYs and GL2, inducing C32 aldehyde production to enhance plant adaptation to diverse environmental stimuli.
GL2 and GL2-LIKE, which cooperate with KCS6, are recognized as major factors involved in the elongation of VLCFA-CoA precursors longer than C28 in both the alkane- and alcohol-forming pathways42,43,44,45. In this study, we show that blue light-activated ZmCRY1s interact with GL2, whereas ZmCRY1c and KCS6 do not form a complex but instead compete for binding to GL2 (Fig. 6e, f; Supplementary Fig. 8e). Furthermore, GL2-LIKE exhibited a weaker interaction with ZmCRY1c compared to GL2 (Supplementary Fig. 6f, g). It has been reported that overexpression of GL2—but not GL2-Like—induces the formation of C32 aldehyde, C31 alkane, C31 secondary alcohol, and C31 ketone55. Additionally, the BAHD catalytic HXXXDX motif is required for GL2-LIKE activity but not for GL255. All these results hint that the CRY1-GL2 complex may indirectly facilitate the VLCFA-CoA biosynthetic flux toward aldehyde synthesis in maize, potentially by recruiting additional binding partners, and this effect appears to be specific to GL2 rather than GL2-LIKE. These findings shed new light on the VLC aldehyde generation, although the precise underlying molecular mechanisms require further elucidation.
Methods
Plant materials and growth conditions
Maize (Zea mays ssp. mays) inbred line B73, Chang7-2, W22, and hybrid line Hi-II were used. To generate maize transformants overexpressing ZmCRYs, the full-length coding sequences of ZmCRYs amplified from B73 leaves were cloned into pB7WGY2 vector69. For GL2 overexpression in the B73 background, the full-length coding sequence of GL2 was subcloned into pEGAD-Myc vector. For CRISPR-Cas9-engineered Zmcry mutants in B73 background, gRNA1 and gRNA2 were cloned into pBUE411-2gR vector. All resulting constructs were delivered into Agrobacterium tumefaciens strain EHA105, and maize transformation was performed by Weimi Biotechnology (Jiangsu) Co., Ltd. UniformMu insertion lines of CRY1s (UFMu-08910, UFMu-09028, and UFMu-08243) were obtained from the Maize Genetics Cooperation Stock Center. glossy2 alleles (mutant IDs: EMS4-1836a9, EMS4-0eb086, EMS4-02f442, EMS4-0eb085, EMS4-0eb084, EMS4-02f441, EMS4-0eb071, EMS4-0eb070, EMS4-02f427, EMS4-02f426, EMS4-02f425, EMS4-02f423, EMS4-0eb06f) were acquired from the Maize EMS-induced Mutant Database (MEMD; http://maizeems.qlnu.edu.cn/). The glossy2-ref mutant was previously described63,66. CRY1c-OE/gl2-ref was generated by crossing CRY1c-OE (in B73 background) with glossy2-ref mutant.
For Arabidopsis experiments, the Columbia (Col-0) ecotype served as the wild type. AtCRY1-Flag overexpression line was previously described9. Full-length coding sequences of ZmCRYs, cloned in pB7WGY2 (carrying YFP tag) and pCambia1300 (carrying Flag tag), were transformed into Col-0 background.
For high intensity UV-B stress analysis in maize (Supplementary Figs. 5b, c and 8c, d), plants were grown at 28°C under LD conditions with blue light (20 μmol m–2 s–1) for 10 days. On day 11, they were treated with (+UV) or without (−UV) broadband UV-B (60 μmol m−2 s−1) for 10 h, then allowed to 1-h dark adaptation before measuring the maximal quantum yield of PSII (Fv/Fm).
For experiments in Fig. 3c, d, plants were grown at 28°C under LD conditions with blue light (20 μmol m–2 s–1) for 10 days. On day 11, they were treated with (+UV) or without (−UV) broadband UV-B (5 μmol m−2 s−1) for 6 h, then allowed to recover for 2 days before measure Fv/Fm; for survival rate analysis (Supplementary Fig. 5d–f), plants were grown for an additional week post-treatment. For each biological replicate, eight seedlings were planted per pot, and the number of surviving plants was recorded (three independent biological replicates were performed).
For experiments in Fig. 6c, d, plants were grown under LD conditions with white light (50 μmol m−2 s−1) for 10 days. They were treated with (+UV) or without (−UV) broadband UV-B (5 μmol m−2 s−1) for 6 h per day over 2 consecutive days, then allowed to recover for 2 days before measuring Fv/Fm.
For different light treatments, blue light (460 ± 40 nm), red light (675 ± 40 nm), far-red light (740 ± 40 nm), and narrowband UV-B (310 ± 10 nm) were generated using LEDs manufactured by Qiding Optoelectronic (Shanghai) Co., Ltd. Broadband UV-B was provided by TL 40 W/12 UV fluorescent tubes (Philips). The intensities of white light, blue light, red light, and far-red light were measured with an ILT1400 Radiometer Photometer, while UV-B light was measured with a LUYOR-340 UV Light Meter.
RNA-seq and transcriptome analysis
For RNA-seq experiments, B73 and zmcry-q seedlings were grown for 7 days under LD (white light, 50 μmol m−2 s−1), pretreated with 48-h darkness, followed by transfer to blue light (20 μmol m−2 s−1) for 1 h. The second leaves were harvested, and total RNA was isolated using RNAiso Plus (Takara). Three biological replicates were independently prepared. RNA purification, reverse transcription, library construction, and sequencing were performed at Shanghai Majorbio Biopharm Biotechnology Co., Ltd. (Shanghai, China) according to the manufacturer’s instructions. RNA-seq transcriptome libraries were prepared following Illumina® Stranded mRNA Prep, Ligation protocol (San Diego, CA) using 1 μg of total RNA. The sequencing libraries were run on the NovaSeq X Plus platform (PE150) using NovaSeq Reagent Kit. The raw paired-end reads were trimmed and quality-controlled by fastp70 with default parameters. Then clean reads were separately aligned to reference genome with orientation mode using HISAT271 software. The mapped reads of each sample were assembled by StringTie72 in a reference-based approach. To identify DEGs (differentially expressed genes) between different samples, the expression level of each transcript was calculated according to the transcripts per million reads (TPM) method. RSEM73 was used to quantify gene abundances. Differential expression analysis was performed using the DESeq274. DEGs with |log2FC | ≥ 1 and FDR < 0.05 were considered to be significantly differentially expressed genes. GO functional enrichments were analyzed at https://geneontology.org. KEGG pathway enrichments were analyzed on the online platform of Majorbio Cloud Platform (https://cloud.majorbio.com/).
Phylogenetic analysis
Amino acid sequences were downloaded from UniProt (https://www.uniprot.org) and aligned using MUltiple Sequence Comparison by Log-Expectation (MUSCLE) in the MEGA7 software with the default settings for protein multiple alignment. Evolutionary distances were computed using Poisson correction analysis. The bootstrap method with 1000 replicates for phylogeny testing was used.
Yeast two-hybrid assay (Y2H)
For the Y2H library screening, the coding sequence of ZmCRY1c was fused in-frame with the GAL4 DNA binding domain (BD) of the bait vector pBridge (Clontech). The resulting constructs were transformed into Y2HGold competent yeast cells. The Zea mays whole transcriptome cDNA library, cloned into the prey vector pGADT7, was constructed by OE Biotech (Shanghai). The cDNA was reverse transcribed from an equal mixture of mRNA extracted from various tissues, including roots, stems, sheaths, leaves (at the VE, V2, and V6 stages), ears, and tassels. The cDNA library contained a total of 1.28 × 10⁷ Colony-Forming Units (CFUs). This library was transformed into Y187 competent yeast cells. The Y2H library screening assays were performed using yeast mating according to the manufacturer’s instructions (Matchmaker User’s Manual, Clontech, California). Screening plates were incubated (colony side facing downward) at 30°C under continuous blue light (20 μmol m−2 s−1) for 3–5 days. Survived yeast clones were sequenced and subjected to a second round of point-to-point validation.
For the point-to-point assays, the sequences encoding ZmCRY1c-PHR (a.a. 1–500), ZmCRY1c-CCE (a.a. 501–688), GL2, GL2G106D, GL2V24M were cloned into pBridge (Clontech). The resulting constructs were transformed into Y2HGold competent yeast cells. The sequences encoding ZmCRY1c-PHR, ZmCRY1c-CCE, GL2, GL2G106D, GL2V24M, GL2-LIKE were cloned into pGADT7. The resulting constructs were transformed into Y187 competent yeast cells. The Y2H assays were performed using yeast mating according to the manufacturer’s instructions (Matchmaker user’s manual, Clontech, California). P53/T was a positive interaction control (provided by Takara Bio).
Immunoprecipitation–mass spectrometry (IP–MS)
Samples (2.5 g) of ZmCRY1c-OE and B73 (7-day-old seedlings grown under LD, pretreated with 48-h darkness followed by 2-h blue light, or maintained in darkness) were collected. A total of 4 samples were analyzed: Sample 1: ZmCRY1c-OE under blue light (biological repeat 1); Sample 2: ZmCRY1c-OE under blue light (biological repeat 2); Sample 3: B73 under blue light; Sample 4: ZmCRY1c-OE in darkness. Sample 3 and 4 served as no-transgene control and no-blue-light control, respectively. The frozen samples were ground into powder in liquid nitrogen and transferred to a mortar with 2 mL of extraction buffer A (50 mM Tris-HCl pH 7.6, 150 mM NaCl, 5 mM MgCl2, 10% v/v glycerol, 0.5% v/v NP40, 5 mM DTT, 1 mM PMSF, 1× protease inhibitor cocktail). After grinding, the homogenate was incubated on a rotary mixer for 30 minutes (4°C, 35 rpm) and centrifuged at 14,000 g for 10 minutes. The supernatant was collected and centrifuged again at 14,000 g for 10 minutes. Then, 3 mL of buffer B (buffer A without NP40) was added and incubated with Anti-GFP Affinity agarose beads (Smart-Lifesciences #SA070005, 25 μL, pre-washed with buffer B twice) on a rotary mixer for 30 minutes (room temperature, 15 rpm). After incubation, the beads were washed 6 times with washing buffer C (50 mM Tris-HCl pH 7.6, 150 mM NaCl, 5 mM MgCl2, 0.2% v/v NP40, 5 mM DTT, 1 mM PMSF, 1× protease inhibitor cocktail) and 6 times with buffer D (buffer C without NP40). The resulting beads were eluted with buffer E (2% SDS, 100 mM Tris pH 8.0, 10 mM TCEP, 50 mM CAA).
The eluted proteins were digested using a modified filter-aided sample preparation (FASP) method75: briefly, eluted proteins were added to 200 μL of UA buffer (8 M urea and 50 mM Tris-HCl, pH 8.0), loaded onto a 10 kDa ultrafiltration centrifuge tube (Sartorius), and centrifuged at 14,000 g for 30 min (repeated twice). Samples were subjected to reductive alkylation by adding 100 μL of UA buffer containing 10 mM TCEP and 20 mM CAA, followed by shaking at 650 rpm for 30 min. Then, 300 μL of UA buffer was added, and the mixture was centrifuged at 14,000 g for 30 min (repeated twice). Next, 300 μL of 50 mM NH4HCO3 solution was added, followed by centrifugation at 14,000 g for 20 min (repeated twice). 2 μL of trypsin solution (1 μg of trypsin in 100 μL of 100 mM NH4HCO3) was added, and the samples were shaken at 650 rpm at 37°C for 12 h. The digests were collected by centrifugation at 14,000×g for 20 min, and the filtrate was desalted using C18 cartridges (Thermo Scientific) after adding 0.1% trifluoroacetic acid (TFA). The peptide concentration was determined using the Pierce Quantitative Colorimetric Peptide Assay kit (Thermo Scientific). Samples were analyzed by timsTOF Pro 2 (Bruker). MS data were acquired as shotgun proteomics (IP-MS) and analyzed using Paser ver2023 software. The raw data were searched against the Maize_UP000007305_4577 Reference proteome database from Uniprot (https://www.uniprot.org/proteomes/UP000007305) and processed as described methods76.
BiFC and subcellular localization assays
In the BiFC assay, constructs expressing ZmCRY1c or GL2 fused to the C-terminus of CFP (a.a. 155–238) or N-terminus of YFP (a.a. 1–172) were transformed into Agrobacterium strain GV3101. For subcellular localization assays, constructs expressing ZmCRY1a, ZmCRY1b, or ZmCRY1c fused to eCFP were transformed into Agrobacterium strain GV3101. For co-localization assays in Nicotiana benthamiana, constructs expressing ZmCRY1c or GL2 fused to CFP or YFP were transformed into Agrobacterium strain GV3101. Overnight agrobacterial cultures were collected by centrifugation, resuspended in MES buffer (pH 5.6) to an OD600 of 0.6, and incubated at room temperature for 2 h before infiltration. Using a 1-mL syringe (without the metal needle), we gently press-infiltrated the Agrobacterium suspensions into the healthy leaves of 3-week-old N. benthamiana. Plants were maintained under LD white light conditions for 2 days after infiltration.
For the co-localization assay in maize protoplasts, constructs expressing ZmCRY1c or GL2 fused to GFP or CFP were transformed into E. coli MC1601. High-concentration constructs (1 to 2 μg/μL) were used for subsequent steps. Protoplasts were transformed with these constructs, cultured at 22°C for 14–16 h, then collected by centrifugation at 100 g for 2 min. Most of the supernatant was removed, and fluorescence signals were observed using a confocal microscope (Leica TSC SP8 STED 3X).
BiLC assays
ZmCRY1a, ZmCRY1b, ZmCRY1c; GL2, GL2V24M, and GL2G106D were fused to sequences encoding either the C- or N-terminus of firefly luciferase. These constructs were transformed into Agrobacterium strain GV3101. After infiltration into N. benthamiana leaves, plants were maintained under LD white light conditions for 2 days. The leaves were infiltrated with luciferin solution (1 mM luciferin and 0.01% Triton X-100), and images were captured using a CCD camera 5 min later.
Pull-down assays
The genes encoding ZmCRY1c-PHRW368A, along with GL2, GL2V24M, or GL2G106D, were cloned into pET-duet vector, with a 6×His tag at the N-terminus of GL2 or GL2 point mutations. E. coli BL21 (DE3) strain was used for protein expression. The bacteria were cultured at 37°C, 200 rpm, then induced with 0.25 mM isopropyl β-D-thiogalactopyranoside (IPTG) for 12 h. Bacterial cells were collected, resuspended in buffer A (20 mM Tris-HCl, pH 8.0, 100 mM NaCl, 1 mM TCEP) supplemented with 1 mM PMSF, lysed using a high-pressure cell disruptor at 700 bar, and centrifuged at 20,000 g for 1 h. The supernatant was incubated with Ni-NTA beads (Qiagen) at 4°C for 1 h. After incubation, the Ni-NTA beads with bound proteins were washed with buffer A supplemented with 25 mM imidazole. The proteins were eluted using buffer A supplemented with 250 mM imidazole, then concentrated and further purified using a Superdex-200 10/300 column.
Immunoblot
Immunoblot was carried out as described previously. For anti-ZmCRY1c-CCE and anti-GL2 polyclonal antibody production, the coding sequence of ZmCRY1c-CCE and full-length GL2 were cloned into pET28a vector (Novagen). Protein expression and purification were carried out as described in pull-down assays. Rabbit immunization with the purified target proteins were performed by Orizymes Biotechnology (Shanghai) Co., Ltd. Briefly, two New Zealand rabbits were used for the preparation of polyclonal antibodies. Freund’s complete adjuvant was used for the first injection, while subsequent immunizations used Freund’s incomplete adjuvant. The injection doses for each rabbit were 250 μg, 500 μg, and 500 μg of protein on days 0, 14, and 35, respectively. The rabbit serum was harvested on day 49. The anti-ZmCRY1c-CCE and anti-GL2 antibodies were used at a dilution of 1:3000. The anti-GFP (Abicode, #M0802-3a) and anti-Flag (Smart-Lifescience, #SLAB01C) monoclonal antibodies were used at 1:3000.
GC-MS analysis of wax composition
Wax extraction and gas chromatography-mass spectrometry (GC-MS) analyses were performed according to the described methods with some modifications76. In different assays, the horizontally expanding parts of the second leaves (2–4 cm from the tip) were taken, and 4 leaves were collected as one replicate and immediately immersed in 10 mL of chloroform with 20 μg of N-tetracosane as an internal standard for 30–60 s. The sample leaves were photographed, and the leaf area of each sample was calculated using Adobe Photoshop 2020. The extracts were then allowed to evaporate to dryness in a fume hood and subsequently transferred to a 2 mL tube. The dried extracts were dissolved in 60 μL of pyridine and derivatized by adding 60 μL of N, N-bis(trimethylsilyl)trifluoroacetamide (BSTFA; Macherey-Nagel) and incubating at 70°C for 1 h. The derivatized samples were filtered into a new sample vial and analyzed using gas chromatography with a flame ionization detector (Agilent GC 7890 Technologies) and GC-MS (Agilent gas chromatograph coupled to an Agilent MSD 5977B quadrupole mass selective detector). GC was performed using an HP-5MS column (Agilent, 30 m × 0.25 mm i.d., 0.25 μm film thickness, 5% phenyl methyl siloxane stationary phase). Helium (purity > 99.999%) was used as the carrier gas at a constant flow rate of 1.0 mL/min. The GC oven temperature was programmed as follows: an initial temperature of 50°C for 2 min, ramped at 5°C/min to 300°C, and held at 300°C for 15 min. The mass range was recorded from m/z 50 to m/z 700. The electron energy was maintained at 70 eV, while the source and quad temperatures were kept at 230°C and 150°C, respectively. Peaks were identified by comparing with standards and using the National Institute of Standards and Technology library (NIST, version 14). The peak areas were determined by cumulative scoring, and the contents of each ingredient were calculated by comparing with the C24 standard. The wax load per unit leaf area was calculated based on the sample leaf area. Three replicates were conducted for each experiment.
Statistical analysis
Phenotype analysis and gene expression analysis were assessed as described in the figure legends. P values were calculated using two-sided Student’s t-tests or one-way ANOVA with Tukey’s multiple comparisons test in GraphPad Prism 9 and are shown in bar graphs or source data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequence data from this work can be found in the Ensembl Gramene or UniProt under the following accession numbers: Zm00001eb244770 (ZmCRY1a), Zm00001eb182820 (ZmCRY1b), Zm00001eb081200 (ZmCRY1c), Zm00001eb382070 (ZmCRY2), Zm00001eb071110 (GL2), Zm00001eb008920 (ZmKCS6), AT4G08920 (AtCRY1), AT1G04400 (AtCRY2), GLYMA_04G101500 (GmCRY1a), GLYMA_06G103200 (GmCRY1b), GLYMA_14G174200 (GmCRY1c), GLYMA_13G089200 (GmCRY1d), GLYMA_10G180600 (GmCRY2a), GLYMA_02G005700 (GmCRY2b), GLYMA_20G209900 (GmCRY2c), Os02g0573200 (OsCRY1a), Os04t0452100 (OsCRY1b), Os02g0625000 (OsCRY2). For RT-qPCR: Zm00001d015743 (bZIP61), Zm00001eb385610 (bZIP80), Zm00001d018207 (ZmUPL13), Zm00001d052138 (ZmUPL14), Zm00001d014990 (ZmRUP2), Zm00001eb361760 (SPA1-related 3), Zm00001eb154420 (SPA1-related 4), Zm00001eb389640 (CRY3), Zm00001eb109290 (KAS I), Zm00001eb373980 (KAS I), Zm00001eb278580 (KAS II), Zm00001eb364580 (FAB2), Zm00001eb277000 (FATB), Zm00001eb377350 (FATB), Zm00001eb167740 (KCS3), Zm00001eb344070 (KCS4), Zm00001eb313510 (GL1), Zm00001eb190120 (GL4), and Zm00001eb233880 (GL1-5). RNA-seq data are available from the National Center for Biotechnology Information Gene Expression Omnibus under the accession number GSE285557. The mass spectrometry proteomics raw data have been deposited in the ProteomeXchange partner repository with the dataset identifier PXD071057. The source data for Figs. 1−6 and Supplementary Figs. 1−6, 8 are provided as a Source Data file. All primers used in this study are listed in Supplementary Data 6. Source data are provided with this paper.
References
Cashmore, A. R. The cryptochrome family of photoreceptors. Plant Cell Environ. 20, 764–767 (1997).
Lin, C. Blue light receptors and signal transduction. Plant Cell 14, S207–S225 (2002).
Sancar, A. Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors. Chem. Rev. 103, 2203–2237 (2003).
Jansen, M. A. K., Gaba, V. & Greenberg, B. M. Higher plants and UV-B radiation: balancing damage, repair and acclimation. Trends Plant Sci. 3, 1360–1385 (1998).
Selby, C. P. & Sancar, A. A cryptochrome/photolyase class of enzymes with single-stranded DNA-specific photolyase activity. Proc. Natl. Acad. Sci. USA 103, 17696–17700 (2006).
Ma, D. et al. Cryptochrome 1 interacts with PIF4 to regulate high temperature-mediated hypocotyl elongation in response to blue light. Proc. Natl. Acad. Sci. USA 113, 224–229 (2016).
Ma, L. et al. Light-response Bric-A-Brack/Tramtrack/Broad proteins mediate cryptochrome 2 degradation in response to low ambient temperature. Plant Cell https://doi.org/10.1093/plcell/koab219 (2021).
Ahmad, M. & Cashmore, A. R. HY4 gene of A. thaliana encodes a protein with characteristics of a blue-light photoreceptor. Nature 366, 162–166 (1993).
Hao, Y. et al. The blue-light receptor CRY1 serves as a switch to balance photosynthesis and plant defense. Cell Host Microbe 33, 137–150.e136 (2025).
Fan, X. et al. Maize cryptochromes 1a1 and 1a2 promote seedling photomorphogenesis and shade resistance in Zea mays and Arabidopsis. Crop J. 11, 1192–1203 (2023).
Li, Z. et al. The mechanism of low blue light-induced leaf senescence mediated by GmCRY1s in soybean. Nat. Commun. 15, 798 (2024).
Keller, M. M. et al. Cryptochrome 1 and phytochrome B control shade-avoidance responses in Arabidopsis via partially independent hormonal cascades. Plant J. 67, 195–207 (2011).
Pedmale, U. V. et al. Cryptochromes interact directly with PIFs to control plant growth in limiting blue light. Cell 164, 233–245 (2016).
Wang, T. et al. Light-induced mobile factors from shoots regulate rhizobium-triggered soybean root nodulation. Science 374, 65–71 (2021).
Guo, H., Yang, H., Mockler, T. C. & Lin, C. Regulation of flowering time by Arabidopsis photoreceptors. Science 279, 1360–1363 (1998).
Liu, H. et al. Photoexcited CRY2 interacts with CIB1 to regulate transcription and floral initiation in Arabidopsis. Science 322, 1535–1539 (2008).
Zhao, Z. et al. CRY2 interacts with CIS1 to regulate thermosensory flowering via FLM alternative splicing. Nat. Commun. 13, 7045 (2022).
Li, L. Z. et al. Common evolutionary trajectory of short life-cycle in Brassicaceae ruderal weeds. Nat. Commun. 14, 290 (2023).
Zeng, D., Lv, J., Li, X. & Liu, H. The Arabidopsis blue-light photoreceptor CRY2 is active in darkness to inhibit root growth. Cell 188, 1–17 (2024).
Tissot, N. & Ulm, R. Cryptochrome-mediated blue-light signalling modulates UVR8 photoreceptor activity and contributes to UV-B tolerance in Arabidopsis. Nat. Commun. 11, 1323 (2020).
Ma, L. et al. Structural insights into the photoactivation of Arabidopsis CRY2. Nat. Plants 6, 1432–1438 (2020).
Shao, K. et al. The oligomeric structures of plant cryptochromes. Nat. Struct. Mol. Biol. 27, 480–488 (2020).
Chen, S. et al. Cryptochrome 1b represses gibberellin signaling to enhance lodging resistance in maize. Plant Physiol. 194, 902–917 (2024).
DeLucia, E. H., Day, T. A. & Vogelman, T. C. Ultraviolet-B and visible light penetration into needles of two species of subalpine conifers during foliar development. Plant Cell Environ. 15, 921–929 (1992).
Gonzalez Moreno, A., de Cozar, A., Prieto, P., Dominguez, E. & Heredia, A. Radiationless mechanism of UV deactivation by cuticle phenolics in plants. Nat. Commun. 13, 1786 (2022).
Agati, G. & Tattini, M. Multiple functional roles of flavonoids in photoprotection. N. Phytol. 186, 786–793 (2010).
Long, L. M. et al. The maize epicuticular wax layer provides UV protection. Funct. Plant Biol. 30, 75–81 (2003).
Shepherd, T. & Wynne Griffiths, D. The effects of stress on plant cuticular waxes. N. Phytol. 171, 469–499 (2006).
Li, S. et al. Deciphering the core shunt mechanism in Arabidopsis cuticular wax biosynthesis and its role in plant environmental adaptation. Nat. Plants https://doi.org/10.1038/s41477-024-01892-9 (2025).
Gordon, D. C., Percy, K. E. & Riding, R. T. Effects of u.v.-B radiation on epicuticular wax production and chemical composition of four Picea species. N. Phytologist 138, 441–449 (1998).
Qiao, P. et al. Transcriptomic network analyses shed light on the regulation of cuticle development in maize leaves. Proc. Natl. Acad. Sci. USA 117, 12464–12471 (2020).
Huang, L. et al. Responses of cuticular waxes of faba bean to light wavelengths and selection of candidate genes for cuticular wax biosynthesis. Plant Genome 13, e20058 (2020).
Gonzalez, R. et al. Responses to ultraviolet-B radiation (280-315 nm) of pea (Pisum sativum) lines differing in leaf surface wax. Physiol. Plant. 98, 852–860 (1996).
Hooker, T. S., Millar, A. A. & Kunst, L. Significance of the expression of the CER6 condensing enzyme for cuticular wax production in Arabidopsis. Plant Physiol. 129, 1568–1580 (2002).
Kunst, L. & Samuels, A. L. Biosynthesis and secretion of plant cuticular wax. Prog. Lipid Res. 42, 51–80 (2003).
Ohlrogge, J. & Browse, J. Lipid biosynthesis. Plant Cell 7, 957–970 (1995).
Ohlrogge, J. B. & Jaworski, J. G. Regulation of fatty acid synthesis. Annu. Rev. Plant Physiol. Plant Mol. Biol. 48, 109–136 (1997).
Shimakata, T. & Stumpf, P. K. Isolation and function of spinach leaf β-ketoacyl-[acyl-carrier-protein] synthases. Proc. Natl. Acad. Sci. USA 79, 5808–5812 (1982).
Joubes, J. et al. The VLCFA elongase gene family in Arabidopsis thaliana: phylogenetic analysis, 3D modelling and expression profiling. Plant Mol. Biol. 67, 547–566 (2008).
Haslam, T. M. & Kunst, L. Extending the story of very-long-chain fatty acid elongation. Plant Sci. 210, 93–107 (2013).
Hegebarth, D. et al. Arabidopsis ketoacyl-CoA synthase 16 (KCS16) forms C(36) /C(38) acyl precursors for leaf trichome and pavement surface wax. Plant Cell Environ. 40, 1761–1776 (2017).
Haslam, T. M., Mañas-Fernández, A., Zhao, L. & Kunst, L. Arabidopsis ECERIFERUM2 is a component of the fatty acid elongation machinery required for fatty acid extension to exceptional lengths. Plant Physiol. 160, 1164–1174 (2012).
Pascal, S. et al. The Arabidopsis cer26 mutant, like the cer2 mutant, is specifically affected in the very long chain fatty acid elongation process. Plant J. 73, 733–746 (2013).
Haslam, T. M. et al. ECERIFERUM2-LIKE proteins have unique biochemical and physiological functions in very-long-chain fatty acid elongation. Plant Physiol. 167, 682–692 (2015).
Wang, X. et al. A beta-Ketoacyl-CoA synthase is involved in rice leaf cuticular wax synthesis and requires a CER2-LIKE protein as a cofactor. Plant Physiol. 173, 944–955 (2017).
Alexander, L. E. et al. The impact of the GLOSSY2 and GLOSSY2-LIKE BAHD-proteins in affecting the product profile of the maize fatty acid elongase. Front. Plant Sci. 15, 1403779 (2024).
Joubès, J. & Domergue, F. Biosynthesis of the plant cuticle. In Hydrocarbons, oils and lipids: diversity, origin, chemistry and fate (Handbook of Hydrocarbon and Lipid Microbiology). (ed. Wilkes, H.) (Springer, Cham, 2020).
Samuels, L., Kunst, L. & Jetter, R. Sealing plant surfaces: cuticular wax formation by epidermal cells. Annu. Rev. Plant Biol. 59, 683–707 (2008).
Kurata, T. et al. The YORE-YORE gene regulates multiple aspects of epidermal cell differentiation in Arabidopsis. Plant J. 36, 55–66 (2003).
Chen, X., Goodwin, S. M., Boroff, V. L., Liu, X. & Jenks, M. A. Cloning and characterization of the WAX2 gene of Arabidopsis involved in cuticle membrane and wax production. Plant Cell 15, 1170–1185 (2003).
Bourdenx, B. et al. Overexpression of Arabidopsis ECERIFERUM1 promotes wax very-long-chain alkane biosynthesis and influences plant response to biotic and abiotic stresses. Plant Physiol. 156, 29–45 (2011).
Bernard, A. et al. Reconstitution of plant alkane biosynthesis in yeast demonstrates that Arabidopsis ECERIFERUM1 and ECERIFERUM3 are core components of a very-long-chain alkane synthesis complex. Plant Cell 24, 3106–3118 (2012).
Pascal, S. et al. Arabidopsis CER1-LIKE1 functions in a cuticular very-long-chain alkane-forming complex. Plant Physiol. 179, 415–432 (2019).
Yang, X. et al. Fatty alcohol oxidase 3 (FAO3) and FAO4b connect the alcohol- and alkane-forming pathways in Arabidopsis stem wax biosynthesis. J. Exp. Bot. 73, 3018–3029 (2022).
Alexander, L. E. et al. Maize Glossy2 and Glossy2-like genes have overlapping and distinct functions in cuticular lipid deposition. Plant Physiol. 183, 840–853 (2020).
Shoaib, N. et al. Potential of UV-B radiation in drought stress resilience: a multidimensional approach to plant adaptation and future implications. Plant Cell Environ. 47, 387–407 (2024).
Yu, X. et al. Arabidopsis cryptochrome 2 completes its posttranslational life cycle in the nucleus. Plant Cell 19, 3146–3156 (2007).
Miao, L. et al. Arabidopsis cryptochrome 1 undergoes COP1 and LRBs-dependent degradation in response to high blue light. N. Phytol. 234, 1347–1362 (2022).
Christians, M. J., Gingerich, D. J., Hua, Z., Lauer, T. D. & Vierstra, R. D. The light-response BTB1 and BTB2 proteins assemble nuclear ubiquitin ligases that modify phytochrome B and D signaling in Arabidopsis. Plant Physiol. 160, 118–134 (2012).
Wang, H., Ma, L. G., Li, J. M., Zhao, H. Y. & Deng, X. W. Direct interaction of Arabidopsis cryptochromes with COP1 in light control development. Science 294, 154–158 (2001).
Yang, H. Q., Tang, R. H. & Cashmore, A. R. The signaling mechanism of Arabidopsis CRY1 involves direct interaction with COP1. Plant Cell 13, 2573–2587 (2001).
Xu, P. et al. Blue light-dependent interactions of CRY1 with GID1 and DELLA proteins regulate gibberellin signaling and photomorphogenesis in Arabidopsis. Plant Cell 33, 2375–2394 (2021).
Tacke, E. et al. Transposon tagging of the maize Glossy2 locus with the transposable element En/Spm. Plant J. 8, 907–917 (1995).
Liu, Y. et al. Structural assembly of maize CRY-GL2 photosignaling complex provides insights into its regulatory role in cuticular wax biosynthesis. Sci. Adv. 11, eadz0136 (2025).
Lu, X. et al. Gene-indexed mutations in maize. Mol. Plant 11, 496–504 (2018).
Zheng, J. et al. Co-expression analysis aids in the identification of genes in the cuticular wax pathway in maize. Plant J. 97, 530–542 (2019).
Koornneef, M., Hanhart, C. J. & Thiel, F. A genetic and phenotypic description of Eceriferum (cer) mutants in Arabidopsis thaliana. J. Heredity 80, 118–122 (1989).
Dietrich, C. R. Molecular and genetic characterization of genes involved in maize cuticular wax biosynthesis. Retrospective Theses and Dissertations 366 https://doi.org/10.31274/rtd-180813-12005 (2002).
Karimi, M., De Meyer, B. & Hilson, P. Modular cloning in plant cells. Trends Plant Sci. 10, 103–105 (2005).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wisniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Tao, Z. et al. ACL1-ROC4/5 complex reveals a common mechanism in rice response to brown planthopper infestation and drought. Nat. Commun. 15, 8107 (2024).
Zhang, Q. et al. GmCRY1a with a latitudinal cline in photoperiodic flowering of soybean. Proc. Natl. Acad. Sci. USA 105, 21028–21033 (2008).
Lyu, X. et al. GmCRY1s modulate gibberellin metabolism to regulate soybean shade avoidance in response to reduced blue light. Mol. Plant 14, 298–314 (2021).
Acknowledgements
The authors thank Dr. Jun Zheng and Dr. Guoying Wang from the Chinese Academy of Agricultural Sciences (CAAS) for the glossy2-ref mutant line, and Dr. Chunyi Zhang from CAAS and Dr. Xiaoduo Lu from QILU Normal University for the glossy2 EMS alleles. Additionally, the authors thank the Instrument Analysis Center of Shenzhen University and are grateful for the support from the Public Service Platform for Instruments and Equipment at the College of Life Sciences and Oceanography, Shenzhen University. This work was supported in part by the National Key R&D Program of China (2024YFA1306700 to H.L.) and National Natural Science Foundation of China (32330006, 31825004, and 32150007 to H.L., 32100197 to F.F., 32300284 to Z.Z., 32170248 to Y.w.L). Z.Z. is supported by the Foundation of the Youth Innovation Promotion Association of CAS.
Author information
Authors and Affiliations
Contributions
Z.Z., F.F., and H.L. conceived the project. Z.Z., F.F., and Y.w.L. performed most of the experiments; Y.q.L. performed the pull-down assay and structural analysis. Y.q.L., Y.w.L., F.W., Y.N., H.f.L., and Y.H. provided materials. W.H., S.W., X.L., J.L., and J.W. provided technical assistance. Z.Z., P.Z., and H.L. analyzed the data. Z.Z. and H.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ronald Pierik and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, Z., Feng, F., Liu, Y. et al. Blue-light-enhanced interaction between ZmCRY1s and GL2 modulates epidermal wax composition in Zea mays. Nat Commun 17, 869 (2026). https://doi.org/10.1038/s41467-025-67587-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67587-7