Abstract
Progressive supranuclear palsy (PSP) is a neurodegenerative disease characterized by abnormal accumulation of the protein tau in the brain, leading to motor and cognitive symptoms that vary between individuals. The reasons for this clinical heterogeneity are unknown. Here we show that distinct molecular forms of tau, particularly high molecular weight (HMW) assemblies, differ in abundance and biological activity across PSP brains. By combining biochemical examination, seed amplification assays, proteomic profiling, and spatial transcriptomics, we identify that HMW tau species drive the strongest aggregation activity in the primary motor cortex. Cases with high tau seeding activity display molecular signatures of altered immune and metabolic pathways. These findings reveal that tau seeding activity reflects underlying molecular heterogeneity in PSP and suggest that measuring 4R-tau seeding capacity could help stratify patients and guide the development of targeted therapeutic approaches.
Similar content being viewed by others
Introduction
Progressive supranuclear palsy (PSP) is a four-repeat (4 R) tauopathy that belongs to the group of frontotemporal lobar degeneration disorders1. Neuropathologically, PSP is defined by neuronal loss, neurofibrillary tangles, and neuropil threads in select subcortical nuclei, accompanied by tufted astrocytes2,3,4, oligodendroglial coiled bodies, and diffuse neuronal cytoplasmic tau immunoreactivity5. Clinically, the most commonly recognized form—Richardson syndrome (PSP-RS)—presents with vertical gaze palsy, axial rigidity, and early falls, although variant clinical phenotypes are increasingly recognized6,7.
Tau is a microtubule-associated protein that stabilizes axonal microtubules and regulates axonal transport8. Under pathological conditions, tau can misfold into β-sheet-rich conformations that assemble into fibrils8. Evidence shows that misfolded tau acts as a seed capable of recruiting native tau and propagating its abnormal conformation in a prion-like manner8,9. This tau seeding mechanism has been demonstrated in vitro and in vivo, where disease-specific tau conformers or “strains” generate distinct pathological and clinical phenotypes10,11,12. Furthermore, over the past decade, studies in transgenic and non-transgenic mice have shown that intracerebral inoculation with PSP brain-derived tau induces the three hallmark PSP cytopathologies11,13,14,15,16. Although tau pathology spreads in these models, none have demonstrated associated behavioral changes or neurodegeneration17. In contrast, inoculation of non-human primates with PSP brain-derived tau produced widespread neuronal tau pathology and both locomotor and executive deficits18. Furthermore, cryo-electron microscopy has also revealed that PSP filaments adopt a distinct tau fold, structurally divergent from those of Alzheimer’s disease (AD) or corticobasal degeneration (CBD)19.
A study has demonstrated that the initial site of neuronal degeneration and tau pathology in PSP (the “pallido-nigro-Luysian” axis and striatum) is consistent across the different clinical subtypes of the disease20. However, it is the varying involvement of other brain regions, with differences in the amount of tau pathology and cellular vulnerability patterns (neuronal versus glial), that correlates with the distinct clinical phenotypes20,21,22,23,24,25.
The reason why the aggregation of the same protein is linked to such variability in clinical presentations remains unclear. Moreover, differences in disease duration, clinical manifestations, and the presence of diverse concomitant pathologies within the same clinical phenotype suggest potential molecular heterogeneity in this disease26,27.
Recent work has begun to link such heterogeneity to the molecular properties of tau assemblies themselves. In AD, high-molecular-weight (HMW) tau species have emerged as key mediators of propagation and toxicity. Takeda and colleagues first identified phosphorylated, PBS-soluble HMW tau species that are efficiently taken up by neurons and transmitted across synapses28 and later detected this seed-competent form in the interstitial and cerebrospinal fluid (CSF) of AD models and patients29. Using size-exclusion chromatography and oligomer-specific ELISAs, Fukumoto et al.30 characterized ≈2000 kDa HMW oligomers that increase with Braak stage and associate with the synaptic protein Bassoon, a mediator of tau spread. Furthermore, soluble tau assemblies with differing seeding efficiencies coexist in AD brains including high-seeding forms driving aggregation, underscoring tau molecular diversity in clinical heterogeneity31. Comparisons of AD brain fractions revealed that these soluble HMW assemblies rival or exceed sarkosyl-insoluble fibrils in seeding potency and glial activation32. Functionally, AD-derived HMW tau impairs hippocampal burst firing and Ca²⁺ channel homeostasis at nanomolar concentrations33, while distinct HMW tau conformers enriched in rapidly progressive AD exhibit accelerated replication kinetics, linking conformer diversity to clinical progression34.
Collectively, these studies establish that HMW tau constitutes a bioactive, transmissible pool of tau assemblies capable of neuronal uptake, seeding, and spread in AD. Although analogous evidence in 4R-tauopathies such as PSP is still limited, the demonstrated transmissibility of patient-derived PSP tau and the potent bioactivity of HMW tau in AD together suggest that comparable assemblies may operate across tauopathies, potentially shaping region-specific vulnerability and clinical heterogeneity.
To explore this hypothesis, we employed seed amplification assays (SAAs), which exploit the self-propagating behavior of misfolded proteins to sensitively quantify pathogenic seeds in real time via thioflavin T fluorescence35. Our group and others have validated this approach to detect α-synuclein and tau seeds across synucleinopathies and tauopathies36,37,38. Applying this framework, we mapped HMW-tau distribution across 20 brain regions from 25 PSP patients, evaluated seeding capacity using 4R-tau SAAs and biosensor cells, and focused on the primary motor cortex—a region rich in HMW tau—where interpatient seeding heterogeneity was observed. By isolating HMW-enriched fractions through size-exclusion chromatography (SEC) and integrating seeding assays with toxicity, neuropathological, proteomic, and spatial transcriptomic analyses, we sought to delineate how distinct tau assemblies contribute to the molecular and clinical diversity of PSP.
Results
Mapping and biochemical characterization of HMW tau in PSP brains
To test the hypothesis that HMW tau could play a role in PSP clinical heterogeneity, we mapped the presence of these species in 20 brain regions from 25 subjects (12 females and 13 males) with a neuropathological diagnosis of PSP (Supplementary Table 1).
To detect these HMW tau species, we immunoblotted the proteins present in the PBS-solubilized fraction from all the regions and patients using an antibody that recognizes the central region of tau (tau HT7, Fig. 1a and Supplementary Fig. 1). For quantification purposes, we defined HMW tau as species with a molecular weight higher than 70 kDa31. The values from the quantification of the HMW tau (normalized to the expression of β-actin) were obtained from all 25 subjects included and were clustered by brain region (Fig. 1b). Values were ranked in ascending order and divided into tertiles, with each tertile assigned a score of 1, 2, or 3, corresponding to low, intermediate, and high levels of HMW tau. These scores were averaged across cases, and a heatmap was generated to illustrate the regional diversity of HMW tau species (Fig. 1c).
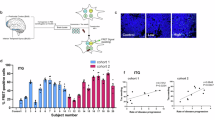
a Representative immunoblot of total tau (clone HT7) illustrating HMW (>70 kDa) and low molecular weight (LMW; <55 kDa) tau species in PBS-solubilized fractions from multiple brain regions of a PSP patient. This immunoblot was performed once on this representative case. A less-exposed film, including molecular weight markers and the full β-actin loading control, is provided in the Source Data file. b Quantification of HMW tau levels across all subjects (n = 25 independent PSP cases) grouped by brain region. Each dot represents an individual biological sample. Individual values are provided in the Source Data file. The data are descriptive and were not subjected to formal statistical testing. c HMW tau regional levels were converted into a semi-quantitative score (0–3) to generate a heatmap, with yellow indicating low levels, orange medium levels, and red high levels. Gray areas represent regions not evaluated. The heatmap illustrates regional trends and does not reflect interpatient variability. Arb units arbitrary units, HMW high molecular weight, LMW low molecular weight, PBS phosphate-buffered saline, PSP progressive supranuclear palsy, SEM standard error of the mean, HT7, tau monoclonal antibody clone HT7. GM gray matter, WH white matter, STN subtalamic nucleus, AC anterior cingulate, kDa kilodalton.
After the analysis, we found that the temporal cortex and the hippocampus were the two regions exhibiting the highest quantity of HMW tau species, followed by the gray matter of the primary motor (precentral) and middle frontal cortices and the amygdala (Fig. 1a–c). Whereas the four regions, which contained the lowest amount of HMW tau were the subthalamic nucleus (STN), the cerebellar dentate nucleus, the gray matter of the occipital cortex and the SN. In all five cortical regions examined, HMW-tau species were always more prominent in the gray matter than in the white matter, with the sole exception of the occipital cortex, where the white matter had more HMW tau than the gray matter (Fig. 1a–c). We correlated the amount of HMW-tau found in each brain region to the neuropathological PSP stage and found a significant but negative correlation in the cerebellum white matter (p = 0.0382, r = −0.444), in the middle frontal gyrus gray (p = 0.0095, r = −0.5929) and white (p < 0.0001, r = −0.7979) matters and in the putamen (p = 0.0056, r = −0.6248).
In addition to the differences found within the amount of HMW tau amongst the different brain regions examined, we also observed large disparities in HMW tau levels among the PSP subjects within the same region (Fig. 1b). Thus, we performed a complete biochemical characterization of the HMW tau present in the primary motor cortex. Although the temporal cortex and hippocampus showed higher levels of HMW tau, the primary motor cortex was chosen because it is heavily involved in the pathophysiology of PSP20 and is less affected by other tau-related comorbidities, such as low or intermediate AD neuropathologic change and aging-related tau astrogliopathy (Supplementary Table 1).
4R-tau seeding capacity shows heterogeneity among PSP patients
To evaluate whether the heterogeneity found in the HMW tau species impacts tau seeding, we evaluated the seeding capacity from the primary motor cortex of the 21 PSP patients, for whom frozen material was available, using two complementary approaches: the 4R-tau SAA and the tau biosensor cells. 4R-tau SAAs are cell-free assays that enable the detection of seeding activity from minute amounts of 4R-tau seeds37. As previously described36,38, we quantified five kinetic parameters from each reaction: lag time, growth phase, T50 (which corresponds to the time needed to reach 50% of maximum aggregation), ThT max, and area under the curve (AUC) of the fluorescence response. Each kinetic parameter was calculated as the mean of the values obtained from each quadruplicate. We defined the AUC as the seeding parameter of interest as it incorporates all the kinetic features of each aggregation reaction, including the speed and extent of aggregation. The AUC values obtained from the 21 subjects included were clustered (Fig. 2a and Supplementary Table 2) and correlated to the amount of HMW tau. There was a robust correlation (r = 0.84, p < 0.0001) between the presence of greater amounts of HMW tau and a higher seeding capacity. However, no significant correlation was found between 4R-tau seeding capacity and age at death (p = 0.4768), biological sex (p = 0.4765), neuropathological PSP stage (p = 0.5915), H1/H2 MAPT haplotype (p = 0.6358) or APOE genotype (p = 0.3579).
a AUC from the 4R-tau SAA of primary motor cortex samples illustrates inter-individual variability in tau seeding activity. Each dot represents the mean AUC of an individual biological sample measured in quadruplicate (n = 21—individual values in Supplementary Table 2). Green diamonds denote the PSP cases selected as high seeders (n = 5) and red asterisks denote the cases selected as low seeders (n = 5) for downstream analyses. The blue asterisk marks PSP#18, which was not included in the low-seeder group due to its low PSP neuropathological stage. Representative aggregation curves of 4R-tau in the presence of primary motor cortex homogenates from two high (green curves)- and two low-seeder (red curves) are shown. The solid line depicts the mean ThT fluorescence of quadruplicate wells, and the shaded band indicates the upper standard error of the mean (SEM). b Representative confocal micrographs of tau biosensor cells incubated with equivalent total protein from primary motor cortex homogenates of high- and low-seeder cases show differential numbers of tau inclusions (green inclusions visualized by FRET fluorescence). Biosensor assays were performed in duplicate per sample; imaging and analysis were performed blinded to seeding category (individual values in Supplementary Table 2). AUC area under the curve, SAA seed amplification assay, PSP progressive supranuclear palsy, ThT thioflavin T, 4R-tau four-repeat tau, FRET Förster resonance energy transfer.
We subtyped the cases into three groups according to their AUC: a top quartile ( ≥ 75th percentile), a middle quartile (25–75th percentile), and a bottom quartile ( ≤ 25th percentile) (Fig. 2a). To assess whether these distinctions could be due to differences in seed concentration or seed characteristics, we performed end-point dilution of the primary motor cortex PBS-solubilized fraction from two PSP cases, one classified as a high seeder and the other a low seeder (Supplementary Fig. 2). Regardless of the amount of brain homogenate used to seed the 4R-tau SAA reaction, the seeding activity of the misfolded tau from the high-seeder PSP case was more potent than that from the low-seeder PSP case. Although we did not find any significant differences between high and low seeders with regard to the PSP neuropathological stage, PSP clinical phenotype, concomitant co-pathologies, sex, MAPT haplotype or APOE genotype, the age at death and disease durations were different. The high seeders, on average, were younger than the low seeders (71.6 versus 76.8 years), and also had a shorter duration of disease (4.4 years) compared to the low seeders (9.2 years, Supplementary Table 1). To illustrate the different kinetic profiles of 4R-tau SAA of high and low seeders, we plotted SAA data from the primary motor cortex of two representative low and two high seeder cases (Fig. 2a).
To validate our 4R-tau SAA results, we measured tau seeding bioactivity using a widely used Förster resonance energy transfer (FRET) based biosensor cell line31,39. We exposed the cells to 6 μg of total protein derived from the PBS-solubilized fraction of the primary motor cortex of the same 21 PSP cases for 48 hours. After the incubation period, we quantified the percentage of FRET-positive inclusions in these cells using an image-based analysis (Supplementary Fig. 3 and Supplementary Table 2). The number of positive FRET inclusions significantly correlated with the 4R-tau SAA AUC values (r = 0.485, p = 0.0243), confirming that there is heterogeneity of tau seeding between PSP patients, especially between those categorized as high and low seeders (Fig. 2b).
Next, to confirm the enrichment of tau seeding activity in the HMW fractions, we performed SEC followed by 4R-tau SAA on the PBS-solubilized fraction of the primary motor cortex from two high seeders and two low seeders. We loaded 450 μg of total protein per patient, and subsequently we immunoblotted 18 SEC fractions from each patient using three antibodies targeting different regions of the tau protein: the N-terminus (tau 6–18), the central domain (tau HT7), and the C-terminus (tau 46).
HMW tau species showed highest immunoreactivity in blots probed with the N-terminus tau antibody (tau 6–18, Supplementary Fig. 4a). In contrast, the central domain antibody (HT7, Supplementary Fig. 4b) detected significantly fewer HMW-tau species, and the C-terminus antibody (tau 46, Supplementary Fig. 4c) minimally detected HMW-tau but effectively recognized physiological tau and LMW tau fragments. Comparing all four patients, high seeders displayed increased HMW-tau levels (fractions 3–6, Fig. 3a, b) compared to low seeders, with no significant differences observed in physiological tau or LMW tau fragments.
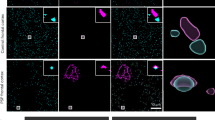
a Immunoblot of SEC fractions (1–18) from the PBS-solubilized primary motor cortex of a high-seeder PSP case, probed with N-terminal tau antibody tau 6–18. This blot was performed once on the indicated sample. A less-exposed film is provided in the Source Data file. b Immunoblot of SEC fractions from a low-seeder PSP case probed with tau 6–18. This blot was performed once on the indicated sample. A less-exposed film, including molecular weight markers is provided in the Source Data file. c Quantification of total tau across SEC fractions by ELISA. Most total tau localized to fractions 12–15, whereas fractions 3–6 contained lower tau levels; in low-seeder samples, tau was not detectable in fractions 3–6. Data are shown as individual values per fraction (see Source Data file). d 4R-tau SAA of SEC fractions. All fractions exhibited measurable 4R-tau seeding; fractions 3–6 showed the highest seeding, fractions 12–15 the lowest, and fractions 16–18 (containing LMW tau) moderate seeding activity. High-seeder cases consistently showed greater seeding across most fractions than low-seeders. SAA measurements for each fraction were acquired in quadruplicate technical replicates; data are presented descriptively without formal statistical testing. Individual values are provided in the Source Data file. PSP progressive supranuclear palsy, SEC size-exclusion chromatography, PBS phosphate-buffered saline, ELISA enzyme-linked immunosorbent assay, SAA seed amplification assay, 4R-tau four-repeat tau, HMW high molecular weight, LMW low molecular weight. RFU relative fluorescence units, AUC area under the curve.
We next quantified total tau in the SEC fractions using ELISA. As expected, most total tau was found in fractions 12–15, while HMW fractions contained lower tau levels. However, caution is needed when interpreting absolute quantitation in these fractions, as steric hindrance may reduce epitope sensitivity31. No differences in total tau levels were observed between high and low seeders, except in fractions 3–6, where tau was undetectable in low seeders (Fig. 3c).
Finally, leveraging the sensitivity of the 4R-tau SAA, we evaluated seeding capacity across the 18 SEC fractions from all patients. Using AUC as the seeding capacity metric, we loaded equal amounts of tau per SAA reaction and conducted four replicates per sample. Data analysis revealed seeding activity in all SEC fractions, with the highest seeding capacity in fractions 3–6, despite their low total tau levels. Fractions 12–15, which contained the largest amount of total tau, exhibited minimal seeding capacity. Fractions 16–18, containing LMW tau, showed greater seeding capacity than fractions 12–15 but less than fractions 3–6. Across most fractions, high seeders consistently exhibited greater seeding capacity than low seeders (Fig. 3d). These results further validate our western blot analysis, as the SEC fractions confirmed the presence of HMW tau species in PSP, albeit at lower abundance than in AD31. Detection in PSP required overexposure of <70 kDa tau bands to visualize these species, and less-exposed versions of all immunoblots are provided in the Source Data file.
Next, we assessed whether primary motor cortex homogenates from high and low seeders exhibit different cytotoxic profiles that might explain the variation in disease duration. We utilized commercially available human astrocytes derived from induced pluripotent stem cells (iAstrocytes). This cell line was chosen due to the prominent astrocytic tau pathology observed in the primary motor cortex of PSP patients20. To ensure consistency with the amount of protein used in the tau biosensor assay, the iAstrocytes were exposed to 6 μg of total protein from the PBS-solubilized fraction of the primary motor cortex for four days (Supplementary Fig. 5a). To evaluate the cytotoxic effect of primary motor cortex-derived homogenates, we used two different approaches36. First, on day in vitro (DIV) 8 and DIV10, we measured the concentration of lactate dehydrogenase (LDH), as damage of the plasma membrane results in a release of this cytosolic enzyme into the surrounding cell culture medium. When we measured extracellular LDH, we found that at both DIV8 and DIV10 LDH was higher in the cells treated with primary motor cortex homogenates derived from either high or low seeder patients compared to untreated cells (p < 0.0001, Supplementary Fig. 5b). However, when we compared the LDH levels between PSP patients, we found significantly higher (p < 0.0001) extracellular LDH expression from the cells exposed to the primary motor cortex homogenates derived from the high-seeder patients compared to low seeder patients (Supplementary Fig. 5b). To corroborate these findings, we performed a cell viability assay (CellTiter-Blue®) at DIV10, where we found a significant decrease in cell viability in iAstrocyte cells exposed to primary motor cortex homogenates derived from high seeders, compared to those exposed to primary motor cortex homogenates derived from low seeders (p < 0.0001, Supplementary Fig. 5c). At DIV10, iAstrocytes were fixed and immunostained using the astrocytic marker glial fibrillary acidic protein (GFAP, Supplementary Fig. 5d). We observed qualitatively that iAstrocytes exposed to primary motor cortex homogenates from high-seeder patients exhibited greater cell loss compared to those exposed to homogenates from low-seeder patients, with both conditions showing reduced cell numbers relative to untreated astrocytes (Supplementary Fig. 5d).
We next characterized tau in the PBS-solubilized homogenates from the primary motor cortex of ten patients (five classified as high and five as low seeders) using two protease-sensitivity digestion assays that had been previously used to define conformations of amyloidogenic proteins31,40. Primary motor-cortex homogenates were incubated with proteinase K (PK, 20 μg/ml) and thermolysin (TL, 25 μg/ml). These banding patterns markedly differed between the high and the low seeders after treatment with PK and TL (Fig. 4). The most divergent PK and TL pattern was observed with the tau antibody against the N-terminus domain (tau 6–18, Fig. 4a). Three out of the five high-seeder cases displayed numerous PK and TL resistant bands that were not evident in the low seeder cases. Furthermore, a more prominent 27 kDa band was observed in the high versus the low seeders. This latter finding was also consistent when the PK and TL digested samples were probed against two other tau antibodies (tau HT7 and tau 46) (Fig. 4b, c). These results indicate inter-individual differences in protease sensitivity between subjects with PSP, and that the tau derived from the high-seeder patients is more resistant to PK and TL digestion.
Representative immunoblots of primary motor cortex samples from high and low seeders, digested with PK or TL, reveal distinct resistance patterns across tau epitopes: a N-terminus (tau 6–18), b central domain (HT7), and c C-terminus (tau 46). All three detections were performed sequentially on the same membrane, with stripping between antibodies over successive days. These blots were performed once on the indicated samples. PSP progressive supranuclear palsy, PK proteinase K, TL thermolysin, HT7 human tau monoclonal antibody clone HT7, tau 6–18 / tau 46 monoclonal antibodies recognizing N-terminal/C-terminal tau epitopes.
Proteomic remodeling between PSP cases with different 4R-tau seeding capacity
To elucidate whether the diversity in HMW tau species and 4R-tau seeding capacity could have an impact on the molecular and cellular mechanisms that might explain the differences in the duration of disease between PSP patients, we performed data-independent acquisition (DIA) analysis, a mass spectrometry technique used in proteomics to comprehensively analyze complex protein mixtures. We analyzed the primary motor cortex proteome of nine PSP patients (four high seeders and five low seeders), which were used for the in vitro experiments (due to tissue unavailability, HS#2 was excluded from the proteomic analysis). We identified 3767 proteins in the primary motor cortex of these patients, of which 484 were differentially expressed in high versus low seeders (Fig. 5a and Source Data File); 247 were highly expressed, and 237 had lower expression in the high versus low seeders. Plotting the differentially expressed proteins (DEPs) demonstrated one cluster involving the four high-seeder cases, another very separate cluster involving three of the low-seeder cases (LS 1, 4 and 5), and a third different proteomic signature in the other two low-seeder cases (LS 2 and 3) (Fig. 5a and Source Data File). Despite the differences in the proteomic signature between the five low-seeder cases, we decided to perform the subsequent bioinformatic analysis with the four high-seeder cases and the five low-seeder cases.
a Heatmap showing unsupervised clustering of DEPs between high and low tau seeders in the primary motor cortex; color scale reflects relative protein abundance. Individual protein values are provided in the Source Data file. b Pathway enrichment analysis of DEPs highlighting common and subset-specific Reactome pathways. Two-sided Fisher’s exact test with Benjamini–Hochberg (BH) correction was used to control the FDR; adjusted P values are reported (Source Data file). c Subcellular compartment analysis of DEPs using Gene Ontology (GO)—Cellular Component terms. Two-sided Fisher’s exact test with BH correction was used; FDR-adjusted P values are reported (Source Data file). d Venn diagram illustrating the overlap between DEPs, known phospho-tau interactors, and proteins previously identified in neurofibrillary tangles. PSP progressive supranuclear palsy, DEPs differentially expressed proteins, Reactome pathway knowledgebase, GO Gene Ontology, BH Benjamini–Hochberg, FDR false discovery rate.
We next examined the differential proteome distributions across specific pathways/biofunctions (Fig. 5b, Supplementary Fig. 6 and Source Data File). Up- and downregulated proteins were independently analyzed. Proteins related to the adaptive immune system, as well as to the metabolism of fatty acids, axon guidance and the cellular response to chemical stress, were both up- and downregulated when comparing high and low seeder patients. However, proteins linked to hemostasis, programmed cell death, regulation of insulin secretion and transport of small molecules were significantly downregulated in high versuslow seeders. In contrast, proteins related to the transcriptional activation of mitochondrial biogenesis, protein localization, or the metabolism of carbohydrates and amino acids were significantly upregulated in the high seeders (Fig. 5b).
We also performed a subcellular mapping of the DEPs. As for the pathway analysis, up- and downregulated proteins were independently analyzed. We found that up- and downregulated proteins had an impact on different cellular structures, and whereas in lytic vacuoles, axons and actin cytoskeleton, both up- and downregulated proteins were found, upregulated proteins were only found in four subcellular compartments: the mitochondrial matrix, the lysosomal lumen, the iron-sulfur cluster assembly complex and the spliceosome complex. However, downregulated proteins were found in many subcellular compartments, including dendrites, endocytic vesicles, neuronal cell body, the nuclear envelope and in the main and distal axons (Fig. 5c and Source Data File).
Finally, a network analysis to evaluate whether any of the DEPs had been previously associated with tau was performed. Our analysis revealed that of the 484 dysregulated proteins, eight proteins had been reported to functionally interact with tau and 27 had been found in the phosphorylated tau interactome of AD neurofibrillary tangles41 (Fig. 5d).
Cases with different 4R-tau seeding capacity can be neuropathologically clustered
Based on the distinct proteomic profiles and the significant differences observed in 4R-tau seeding capacity, we sought to investigate whether these findings correlate with distinctive neuropathological features. To address this, we analyzed the same four cases used in the SEC experiment: two high-seeder cases (HS3 and HS5) and two low-seeder cases (LS4 and LS5). These four cases were not different in the stage of PSP (stage four/five), clinical phenotype (PSP-RS) or in the presence of co-pathologies (Supplementary Table 1). The neuropathological examination was performed by two neuropathologists (S.L.F. and G.G.K.) blinded to the seeding category of the cases. Representative neuropathological features characteristic of PSP are shown in Fig. 6a–e. In the primary motor cortex, gray matter contains neuronal neurofibrillary tangles, tufted astrocytes, and oligodendroglial coiled bodies (Fig. 6a), while the white matter displays thread pathology and additional coiled bodies (Fig. 6b). Higher magnification images illustrate a neurofibrillary tangle (Fig. 6c), a tufted astrocyte (Fig. 6d), and oligodendroglial coiled bodies (Fig. 6e). A thorough evaluation of neuronal, astroglial and oligodendroglial tau-immunopositive inclusions throughout the brain of the four cases revealed a similar distribution and density of total AT8-tau-immunopositive pathology (Fig. 6f). However, the density of each cytopathology differed between regions and cases. In all cases, the density of neurofibrillary tangles was similar, with the highest density observed in the brainstem and globus pallidus (Fig. 6f). In contrast, heat mapping showed variation in the distribution and density of tufted astrocytes and oligodendroglial coiled bodies (Fig. 6g), both between individual cases and between high versus low seeders. Overall, the highest density of tufted astrocytes was observed in the primary motor cortex, basal ganglia and thalamus, and the highest density of oligodendroglial coiled bodies was observed in the primary motor cortex, basal ganglia, thalamus, brainstem and cerebellar white matter. A high density of tufted astrocytes was also found in the parietal cortex of high seeders. Although a clear differentiation between low and high seeders could not be determined when visualizing the neuropathological features, cluster analysis based on the semi-quantitative scores for all brain regions and all cytopathologies (neuronal, astroglial, oligodendroglial) revealed that the two low seeders clustered together and had a distinct pathological profile compared to the high seeders (Fig. 6h).
a–e Representative images of phosphorylated tau (AT8) immunostaining in the precentral (primary motor) cortex of PSP patients. a Gray matter showing neuronal neurofibrillary tangles, tufted astrocytes, and oligodendroglial coiled bodies. b White matter displaying thread pathology and coiled bodies. c Higher magnification of a neurofibrillary tangle. d Higher magnification of a tufted astrocyte. e Higher magnification of oligodendroglial coiled bodies in the precentral cortex. f Comparative images of low seeder (LS#5, left) and high seeder (HS#5, right) pathology, highlighting neurofibrillary tangles, tufted astrocytes, and coiled bodies in the primary motor cortex and putamen, neurofibrillary tangles in the locus coeruleus, and oligodendroglial coiled bodies in the cerebellar white matter. g Heatmap of total tau, neuronal, astroglial, and oligodendroglial tau pathology for two low seeders (LS#4 and LS#5) and two high seeders (HS#3 and HS#5). Horizontal rows represent individual cases, while vertical columns represent tau pathology types. Color coding indicates density: red = high, orange = moderate, yellow = low, white = absence, and gray = areas not examined. h Cluster analysis based on tau-immunopositive cytopathology across brain regions reveals that low seeders cluster together and exhibit a distinct pathological profile compared to high seeders. Scale bars: 100 μm in (a, b), 20 μm in (c, d, f). Each representative micrograph is drawn from one biologically independent PSP case and is representative of the staining pattern observed across the indicated group. Cluster analysis was performed once on the compiled neuropathological scoring dataset. PSP progressive supranuclear palsy, AT8 monoclonal antibody to phosphorylated tau (Ser202/Thr205); NFT neurofibrillary tangle, LS low seeder, HS high seeder.
Spatial transcriptomics confirms molecular differences between PSP subgroups
After confirming that in addition to the differences found in 4R-tau seeding and in the proteomic signature, the high and low seeders could be clustered-based on the constellation of neuropathological features, we performed spatial transcriptomic analysis in the same four cases where a thorough neuropathological examination had been performed (Fig. 7). After confirming that the four cases had good tissue integrity and RNA quality, transcriptomic data was integrated and clustered using standard pipelines, revealing 14 distinct region-types within the primary motor cortex (Supplementary Table 3). Each transcriptomic data spot was also deconvolved using a recent single-cell atlas of primary motor cortex42 (Source Data File). By combining the frequency of layer-specific excitatory neurons, identified clusters, and expert annotation of histology sections, we were able to define the boundaries of each cortical layer across our spatial transcriptomic slices (Fig. 7c). These annotations were confirmed using expression of layer-specific markers (Fig. 7b). Layer 2 highly expressed CUX2 and RASGRF2. Layer 3 highly expressed RORB and SCN4B. Layer 5 highly expressed VAT1L, PCP4, and FEZF2. Whereas Layer 1, 4 and 6 were difficult to distinguish transcriptionally and relied more on anatomical structures for their annotation (Supplementary Fig. 7).
a H&E sections of the four slices of primary motor cortex assayed with Visium spatial transcriptomic. b Marker genes confirm manually annotated cortical layers. c Annotated cortical layers in each slide, white white matter, other colors are the same as in panel d. “Edge” indicates a region of stressed tissue found on the edge of the tissue sample. d Uniform Manifold Approximation and Projection (UMAP) shows annotated layers are transcriptionally distinct. e UMAP confirms successful integration of spatial transcriptomic profiles across slices. f Differentially expressed genes were identified using a negative binomial mixed-effect model, significant genes were identified (<5% FDR) and used for pathway analysis with GSEA. The top 25 significantly (after Benjamini–Hochberg multiple testing correction) enriched Reactome pathways, which overlap with pathways detected through proteomic analysis. Spatial transcriptomic assays were performed once on each of the four independent tissue slices, with internal technical quality control and integrated analysis across slices as described in the Methods. PSP progressive supranuclear palsy, H&E hematoxylin and eosin, Visium 10x Genomics Visium platform, UMAP uniform manifold approximation and projection, DEG differentially expressed gene, FDR false discovery rate, BH Benjamini–Hochberg, GSEA gene set enrichment analysis. LS low seeder, HS high seeder.
The composition of each slide varied in the amount of each layer (Fig. 7c–e and Supplementary Table 3). However, using our cell-type deconvolution results, we observed an increased frequency of oligodendrocytes in both gray (20–40% versus 10–20%) and white matter regions (75% versus 33%) in high seeders versus low seeders (Source Data File). In parallel, we observed a decrease in neuronal signatures in these regions. In addition, we also observed a decrease in astrocytes in Layer 1 and Layer 6, from 44% to 17% in Layer 1 and 29% to 12% in Layer 6 in high seeders. However, due to the small sample size and uneven distribution of layers across samples, the significance of these changes is difficult to assess.
We then performed differential expression analysis between high and low seeders, controlling for the proportions of different cortical layers (two samples per group, Source Data File). Pathway enrichment analysis across all genes revealed many enriched pathways downregulated in high versus low seeders at the transcriptomic level that are consistent with those observed at the protein level, including insulin production, phagocytosis, response to cellular stress and neutrophil degranulation (Fig. 7f). Transcriptomic analysis further revealed downregulation of neurotransmitter release, oxidative phosphorylation, and NF-kappa-B signaling in the high-seeder PSP cases (Source Data File).
Misfolded tau accumulation produces transcriptomic changes
Finally, and taking advantage of the spatial resolution that Visium offers, we examined whether these transcriptomic changes were associated with tau deposition. AT8-tau pathology burden was measured in an adjacent 4 μm tissue section, quantified using Halo (Indica Labs), and aligned with the transcriptomic data (Supplementary Fig. 8a). AT8-tau pathology was quantified as the percentage of each Visium region of interest (ROI) occupied by AT8-immunoreactive signal, and only signal within the physical boundaries of each spot was included in the analysis; surrounding signal was excluded. Unfortunately, from the four cases, only one high-seeder and one low-seeder sample, which exhibited fine alignment with the immunohistochemically stained slide, could be included in the analysis. Using a general linear model, we identified gene expression patterns associated with varying levels of AT8-tau pathology across 1250 Visium regions of interest (ROIs). This identified 69 genes significantly associated with AT8-tau pathology burden, where 58 were upregulated in proximity to AT8-tau pathology, and 11 were downregulated (Supplementary Fig. 8b and Source Data File). We found that these genes were mainly involved in negative regulation of the cell-cycle, negative regulation of transcription factor binding, and extracellular matrix organization (Supplementary Fig. 8c). While these pathways are consistent with our previous results, this analysis was performed on only a portion of two Visium samples and was confounded by the cortical layer. Moreover, we acknowledge that AT8 immunostaining was performed on an adjacent section rather than on the capture slide, and that variability in fixation, antibody penetration, and tissue handling are inherent limitations of studies using postmortem human brain material. As such, these results should be considered preliminary.
Discussion
Our study highlights the differential seeding capacity of 4R-tau in a cohort of clinically and neuropathologically well-characterized PSP cases. While characteristic neuropathological features4 and the tau filament structure19 are thought to be uniform, we reveal previously unrecognized heterogeneity in the distribution of HMW-tau species across 20 distinct brain regions, underscoring the complexity of PSP pathology and the need to consider tau’s biochemical diversity in neuropathological assessments. Second, we demonstrate that HMW-tau species in the primary motor cortex exhibit significantly enhanced seeding capacity, as determined by SEC followed by 4R-tau SAA. Third, we link interpatient variability in these species to differences in 4R-tau seeding capacity, noting that individuals with elevated seeding activity tended to have a shorter disease duration. Fourth, cases stratified by 4R-tau seeding capacity align with clusters of PSP cases grouped by neuropathological features across brain regions, independent of disease stage. Finally, our proteomic and spatial transcriptomic analyses uncover significant dysregulation of the adaptive immune system in high versus low seeder patients.
PSP is a clinically heterogeneous tauopathy, where the regional distribution of 4R-tau pathology correlates with clinical features20,21,22,23,24,25. However, the burden of tau inclusions alone does not explain variability in clinical presentation and progression27. Our findings suggest that tau’s biochemical properties, particularly the seeding capacity of specific HMW fractions, might contribute to disease heterogeneity. These regional and interpatient differences in HMW tau distribution raise critical questions about the underlying mechanisms driving this diversity. Future studies should explore the role of the different tau species in PSP pathology in detergent-soluble and insoluble fractions, as well as the role of LMW tau species.
Our findings also underscore the potential of 4R-tau SAA for stratifying PSP cases based on tau seeding capacity. However, current stratification is limited to analyses using brain homogenates run on the same plate. To extend this approach to living patients, quantitative SAAs must be developed. Additionally, larger cohorts are needed to validate the observed relationship between shorter disease duration and higher tau seeding capacity.
The correlation between high seeding capacity and reduced astrocyte viability in vitro highlights the functional impact of tau heterogeneity. While high seeder-derived tau species appear more cytotoxic, contributions from other proteins present in PSP brain homogenates cannot be excluded. Consistent with prior reports43, our study shows that HMW tau species, despite low total tau levels, exhibit the highest seeding activity. This finding indicates that tau seeding capacity is not necessarily correlated with total tau abundance but is instead influenced by the conformational and biochemical properties of specific tau species. Furthermore, protease resistance differences between high and low seeders suggest that high-seeder tau conformers may adopt aggregation-prone or protease-resistant structures. These findings align with observations in other neurodegenerative diseases31,36,38, where protein conformational diversity underpins phenotypic heterogeneity44.
While our findings provide insights into the molecular heterogeneity linked to tau seeding capacity, we acknowledge that our results are derived from a limited number of cases. Nevertheless, the robustness of our approach is reinforced by its alignment with findings from large-scale transcriptomic analyses, such as those recently conducted by Farrell et al.45, Rexach et al.46, Wang et al.47, and Whitney et al.48, which have significantly advanced our understanding of PSP pathogenesis. Our study builds upon this existing knowledge by specifically exploring the differences between high and low seeders, focusing on the functional heterogeneity linked to seeding capacity.
Our proteomic analysis reveals differentially expressed proteins across subcellular compartments, including dendrites, endocytic vesicles, neuronal cell bodies, the nuclear envelope, and axonal structures. Disruptions in these compartments are likely to impair intracellular trafficking, synaptic integrity, and nuclear-cytoplasmic transport—processes critically affected in tauopathies49,50. Dysregulated endocytic vesicles may interfere with protein recycling and clearance pathways, contributing to tau aggregation51, while alterations in the nuclear envelope exacerbate cellular dysfunction52,53. These findings suggest that subcellular vulnerabilities play a pivotal role in PSP pathogenesis and warrant further investigations, leveraging cell-type-specific approaches, which could provide a more nuanced understanding of how proteomic and transcriptomic changes correlate with specific neuropathological features.
Dysregulated adaptive immunity further distinguishes high and low seeders, with our analyses showing downregulation of cell-cycle regulation and transcription factor binding pathways in regions with misfolded tau. These findings suggest that tau aggregation directly impacts cellular processes critical for neuronal and glial function54,55,56,57,58,59,60. Prior studies measuring microglial activation via PET imaging60 and our transcriptomic and proteomic data support the prominent role of neuroinflammation in PSP. Our earlier study on LBD36 revealed distinct pathways in high seeders, emphasizing mitochondrial dysfunction and lipid metabolism, while PSP is dominated by immune dysregulation and cellular stress. These findings are consistent with recent experimental data59 and human neuropathological evidence61,62,63, underscoring the critical role of T lymphocytes in the pathogenesis of PSP. However, whether tau drives these inflammatory changes or whether inflammation influences tau seeding capacity remains unclear. Future studies using cell-based and animal models will be essential to disentangle this relationship.
Finally, we investigated whether the same cases selected for spatial transcriptomics could be neuropathologically clustered, despite having similar disease stages, clinical phenotypes, and co-pathologies. After a detailed neuropathological examination, we found that high and low seeders exhibited distinct neuropathological profiles, and while total tau-immunopositive pathology was similarly distributed, the density of tufted astrocytes and oligodendroglial coiled bodies varied significantly.
High seeders showed increased tufted astrocytes in the primary motor cortex, basal ganglia, thalamus, and parietal cortex, while oligodendroglial coiled bodies were more prominent in the primary motor cortex, basal ganglia, thalamus, brainstem, and cerebellar white matter. Cluster analysis confirmed distinct neuropathological profiles, with low seeders clustering separately from high seeders. This aligns with the molecular heterogeneity observed in proteomic and transcriptomic profiles, suggesting that tau seeding capacity influences both molecular and neuropathological diversity in PSP.
Our multidisciplinary approach integrates neuropathological, biochemical, proteomic, and transcriptomic data to highlight PSP’s molecular heterogeneity. Stratifying patients based on tau conformational and functional diversity could inform personalized therapies, echoing advances in cancer64,65, where molecular subclassification has transformed outcomes. However, identifying specific seeding-competent tau species driving PSP progression remains a critical unmet need. Pinpointing these species, particularly their interactions with immune pathways, could yield promising therapeutic targets.
Achieving this, however, requires the development of robust, non-invasive biomarkers and assays capable of monitoring tau seeding capacity in vivo37,66,67. Such tools will be essential not only for monitoring early interventions but also for stratifying patients at the outset of clinical trials in order to enhance therapeutic efficacy.
Despite the significant progress achieved in this study, some limitations must be acknowledged: β-actin was used as a normalization factor in western blot analyses due to its widespread use, but alternative normalization strategies, such as total tau, may provide additional context. The small sample size, particularly in proteomic and spatial transcriptomic analyses, limits the generalizability of our findings. Additionally, tissue alignment challenges restricted spatial transcriptomic findings to two cases, highlighting the importance of optimized methodologies, as demonstrated in the AD field68. Finally, as our classification strategy has already enabled the detection of significant differences in OMICs despite a small sample size, validating these findings in larger cohorts will be essential to support the notion that distinct pathogenic pathways may be associated with neuropathological features characteristic of PSP-type pathology
In conclusion, our study reveals that 4R-tau seeding activity can help to delineate molecular subtypes in PSP. These findings provide a foundation for advancing molecular subclassification. By integrating structural, functional, and immune-related aspects, we aim to set the stage for precision medicine approaches tailored to the molecular underpinnings of this devastating disease.
Methods
Human tissue samples
A total of 25 consecutive PSP brain samples (12 females, 13 males) were selected from the University Health Network-Neurodegenerative Brain Collection (UHN-NBC, Toronto, Canada) based on a confirmed neuropathological diagnosis. Autopsy tissues were collected with informed consent from patients or their relatives, in compliance with the approval of the local institutional review board. Consent to publish information that could include indirect identifiers such as age, sex, and diagnosis was obtained. Participants were not compensated. This study was conducted under the University Health Network Research Ethics Board (Approval No. 20-5258). Sex was determined from medical records and confirmed in autopsy reports; gender identity was not collected. Details on age at death, biological sex, and findings from comprehensive neuropathological examinations are provided in Supplementary Table 1. A systematic neuropathological examination was conducted prior to inclusion, adhering to established diagnostic criteria for neurodegenerative conditions and co-pathologies69.
For each brain, one hemisphere was formalin-fixed, dissected into regions, and prepared as formalin-fixed paraffin-embedded (FFPE) tissue sections for neuropathological characterization. The contralateral hemisphere was sliced into ~2 cm thick coronal segments at autopsy, flash-frozen, and stored at −80 °C. Microdissection was performed using a 4-mm tissue punch to collect samples from the gray and white matter of the anterior cingulate, middle frontal, primary motor, middle temporal, and occipital cortices, as well as the amygdala, hippocampus, caudate, putamen, globus pallidus, pons base, substantia nigra, subthalamic nucleus, cerebellar white matter, and dentate nucleus36,38. All tissue punches were stored in low protein-binding tubes (Eppendorf), flash-frozen, and preserved at −80 °C.
Clinical data were obtained by reviewing patient records. Movement disorder specialists independently reviewed and extracted clinical information, including age at diagnosis and death, sex, disease duration (in years), motor and non-motor symptoms, neurological examination findings, and medications. The temporal progression of symptoms was documented, including the order of symptom presentation (i.e., first, second, and third symptoms). Where available, data on the presence, absence, and estimated onset dates of motor and non-motor symptoms were recorded. Additionally, the age of onset for postural instability, oculomotor deficits, Parkinsonism, cognitive impairment, and other movement abnormalities was determined through serial neurological examinations. Baseline scores on the Progressive supranuclear palsy rating scale (PSP-RS) were documented. Predominant PSP phenotypes were classified using Movement Disorder Society diagnostic criteria6.
Genetic analyses
The genotypes for apolipoprotein E (APOE) and microtubule-associated protein tau (MAPT) single-nucleotide polymorphisms were examined in all cases (Supplementary Table 2), using the Illumina GSA array27.
Protein extraction
For the PBS-solubilized fraction, 40–50 mg of frozen micro-dissected tissue was thawed on wet ice and then immediately homogenized in 500 μl of PBS spiked with protease (Roche) and phosphatase inhibitors (Thermo Scientific) in a gentle-MACS Octo Dissociator (Miltenyi BioTec). The homogenate was transferred to a 1.5 ml low protein-binding tube (Eppendorf) and centrifuged at 10,000 × g for 10 min at 4 °C31,36,38. The supernatant was collected and aliquoted in 0.5-ml low protein-binding tubes (Eppendorf) to avoid excessive freeze–thaw cycles. A bicinchoninic acid protein (BCA) assay (Thermo Fisher Scientific) was performed to determine the total protein concentration of all samples.
SDS-PAGE and immunoblotting
Gel electrophoresis was performed using 4–12% NuPAGE Bis-Tris gels (Thermo Fisher Scientific). Proteins were transferred to 0.45-μm nitrocellulose membranes for 60 min at 35 V. The membranes were blocked for 60 min at room temperature in blocking buffer (5% [w/v] skim milk in 1× TBST (TBS and 0.05% [v/v] Tween-20)) or in bovine serum albumin (BSA, 5% [w/v] BSA in 1× TBST) and then incubated overnight at 4 °C with primary antibodies directed against amino acids 6–18 (tau 12, 1:1000 dilution, ref: 806501, Biolegend), 159–163 (tau HT7, 1:1000 dilution, ref: MN1000, Thermo Fisher Scientific) and 404-441 (tau 46, 1:1000 dilution, ref: 13-6400, Thermo Fisher Scientific) of the tau protein. All antibodies were diluted in blocking buffer. The membranes were washed three times with TBST and then incubated for 60 min at room temperature with horseradish peroxidase-conjugated secondary antibodies (1:3000 dilution, ref: 172-1011 and 1:5000 dilution, ref: 170-6515, Bio-Rad) in blocking buffer. β-actin was used as the housekeeping protein for normalization of Western blot analyses. Membranes were incubated with a primary antibody (1:5000 dilution, ref: MAB8929, R&D Systems) for 1 hour at room temperature. Following primary antibody incubation, membranes were washed and incubated with a goat anti-mouse IgG (H + L) secondary antibody conjugated to horseradish peroxidase (1:10,000 dilution, ref: 172-1011, Bio-Rad) for 1 hour at room temperature. Following another three washes with TBST, immunoblots were developed using ECL Western Blotting Detection Reagents (ref: RPN2106, Cytiva) and imaged using X-ray films. For quantification, grayscale images were scanned and imported into Fiji/ImageJ.
4R-tau seed amplification assay (4R-tau SAA)
SAA reactions were performed in 384-well plates with a clear-bottom (Nunc)22. Recombinant 4R-K18 cysteine-free construct with cysteine to serine mutations at residues 291 and 32270,71, and a 3R-K19 cysteine-free construct with a cysteine to serine mutation at residue 32270,72,73 (rPeptide) were thawed from −80 °C storage, reconstituted in HPLC-grade water (Sigma) and filtered through a 100-kDa spin filter (Thermo Fisher Scientific) in 500 µl increments. All reagents used in the reaction buffers were purchased from Sigma. 10 μl of biological sample (1.5 μg of total protein from the PBS-solubilized fraction and 5 pg of total tau from the SEC fractions) were added to the wells containing 20 μl of reaction buffer (0.1 M PB, pH 8, 0.875 M Na3Citrate, 45 μM poly-L-glutamic acid sodium salt), 10 μl of 50 μM ThT and 10 μl of a mixture of 0.5 mg/ml of monomeric 4R-K18 and 0.25 mg/ml of monomeric 3R-K19. The plate was sealed and incubated at 42 °C in a BMG FLUOstar Omega plate reader with cycles of 1 min shaking (500 rpm double orbital) and 1 min rest. The fluorescence measurements (450 ± 10 nm excitation and 480 ± 10 nm emission, bottom read) were taken every 15 min for a period of 72 h. Each sample was tested in quadruplicate, and the same positive and negative controls were added to each plate.
Tau biosensor cells
Tau RD P301S FRET Biosensor (ATCC CRL-3275) cells were cultured at 37 °C, 5% CO2 in DMEM, 10% vol/vol FBS, 0.5% vol/vol penicillin–streptomycin31,39. Cells were plated on Ibidi clear-bottom 96-well plates at a density of 25,000 cells per well. Brain extracts (6 μg of total protein from the PBS-solubilized fraction) were then incubated with Lipofectamine 2000 (Invitrogen, final concentration 1% vol/vol) in Opti-MEM for 10 min at room temperature before being added to the cells. Each sample was tested in duplicate. After 48 h, cells were fixed with 4% PFA for 10 minutes, and then Nucblue (Invitrogen) was added for 10 minutes at room temperature. Cells were imaged in 3 × 3 fields at ×20 magnification using a Nikon ECLIPSE Ti2 confocal microscope. Using the object colocalization IF module on the HALO software (version 3.5, Indica Labs), the total number of cells (DAPI) in the monolayer and tau aggregates (FITC) was quantified blindly to case identity to calculate the number of aggregates per cell (FITC/DAPI).
Size-exclusion chromatography (SEC)
Differently sized tau species were separated via size-exclusion chromatography (SEC) on a Superdex 200 10/300 GL column (GE Healthcare Life Sciences) in sterile-filtered PBS, pH 7.4 (Gibco). PBS-solubilized brain extracts from four PSP cases (450 µg of total protein) were loaded onto the column and fractionated at a flow rate of 0.5 mL/min using an ÄKTA purifier 10 (GE Healthcare Life Sciences). A total of 18 fractions (each 0.5 mL) were collected for each sample, starting 6 mL after injection. Fractions were aliquoted, protein concentration was quantified using micro-BCA (Thermo Fisher Scientific) and stored at −80 °C until further use.
Tau ELISA
Human tau total ELISA kit (ref: KHB0041, Thermo Fisher Scientific) was used to measure the tau present in all SEC fractions according to the manufacturer’s protocol. Absorbance was measured at 450 nm using the SpectraMax i3 (Molecular Devices). Each biological sample was performed in triplicate.
Cell culture and treatment
iCell Astrocytes (iAstro, Cellular Dynamic International) were thawed following company guidelines (iCell Astrocytes User’s Guide, Cellular Dynamic International). Cells were plated in complete maintenance medium (Cellular Dynamic International) on a poly-L-ornithine (ref: A004C, Sigma) and laminin (ref: L2020, Sigma) coated clear-bottom 96-well plate (Ibidi) at a density of 45,000 cells per well. iAstro cells were maintained and then treated with 6 μg of PBS-solubilized brain homogenates on day in vitro (DIV) 6 and 8. Untreated cells were used as a control. On DIV10, cells were prepared for immunocytochemistry or for cell viability assays.
Toxicity assays
Cytotoxicity was determined by the presence of lactate dehydrogenase (LDH). Supernatant was collected at DIV8 and 10, spun at 300 × g for 5 minutes and stored at −80 °C. LDH was measured using the Cytotoxicity Detection Kit according to the manufacturer’s instructions (ref: 11644793001, Roche) and absorbance (ABS) was measured at 490 nm and 600 nm using the SpectraMax i3 (Molecular Devices). LDH values were calculated using the equation (ABS 490 nm–ABS 600 nm) sample - (ABS 490 nm–ABS 600 nm) average media blank (background). Untreated samples were not set to 0 but were just close to the background levels of the media blanks.
Cell viability was determined at DIV10 using the CellTiter-Blue Cell Viability Assay according to the manufacturer’s instructions (ref: 8080, Promega). In brief, live cells were cultured with 20 µl of reagent and 100 µl of medium and incubated for 1 hour in standard cell culture conditions. Fluorescence was measured at 560/590 nm (Ex/Em) using the SpectraMax i3 (Molecular Devices).
Immunofluorescence
Immunofluorescence was performed by first washing the cells in PBS and then fixing with 4% PFA for 10 minutes at room temperature. Cells were washed 3 times with PBS and incubated in 2% BSA in PBS for 1 hour and then in 0.1% TritonX100 + 1% BSA in PBS for 20 minutes, at room temperature. Samples were incubated with anti-GFAP (1:250 dilution, ref: ab7260, Abcam) in 1% BSA in PBS at 4 °C overnight. The following day, cells were washed with PBS and incubated with AlexaFluor-488 donkey-anti-rabbit for 75 minutes at room temperature (1:500 dilution, Thermo Fisher Scientific). Finally, cells were washed 4 times in PBS and incubated at room temperature with Nucblue (Invitrogen) for 10 minutes. Images were obtained on a Nikon ECLIPSE Ti2 confocal microscope (Nikon).
Thermolysin and proteinase K digestions
A concentration of 25 μg/ml of thermolysin and of 20 μg/ml of proteinase K was added to 30 μg of total protein from the PBS-solubilized brain homogenates38,40. Samples were incubated at 37 °C with continuous shaking (600 rpm) for 60 min. Thermolysin digestions were halted with the addition of EDTA to a final concentration of 2.5 mM. Proteinase K digestions were halted with a final concentration of 4 mM PMSF. Samples were resuspended in 1× LDS buffer (Life Technologies) and analyzed by SDS-PAGE followed by immunoblotting, as described above.
Mass spectrometry
Using a 4-mm tissue punch, a microdissection of the primary motor cortex from 9 PSP patients was performed. Samples were homogenized in modified RIPA buffer (2% SDS, 150 mM NaCl, 50 mM Tris pH 8). Protein extraction was performed by mechanical disruption using 1.6 mm stainless steel beads in a NextAdvance Bullet Blender. Samples were incubated at 60 °C for 30 minutes and clarified by centrifugation. Tissue extracts were subjected to TCA precipitation74. Washed protein pellets were solubilized in 900 μL of urea buffer (8 M urea, 150 mM NaCl, 50 mM Tris pH 8, 1× Roche cOmplete protease inhibitor). Protein quantitation was performed using Qubit fluorometry (Invitrogen). 50 μg of each lysate reduced with 15 mM dithiothreitol at 25 °C for 30 minutes followed by alkylation with 15 mM iodoacetamide at 25 °C for 45 minutes in the dark. Digested with 2.5 μg sequencing grade trypsin (Promega) at 37 °C overnight. The final digest volume was 0.5 mL adjusted with 25 mM ammonium bicarbonate. The digest was cooled to 25 °C, acidified with formic acid and desalted using a Waters Oasis HLB solid phase extraction plate. Eluted samples were frozen and lyophilized. A pooled sample was made by mixing equal amounts of digested material from each sample. This pooled sample was used to generate a gas-phase fractionation library.
DIA chromatogram library generation
1 μg of the pooled sample was analyzed by nano LC-MS/MS with a Waters M-class HPLC system interfaced to a ThermoFisher Exploris 480. Peptides were loaded on a trapping column and eluted over a 75μm analytical column at 350nL/min; both columns were packed with XSelect CSH C18 resin (Waters); the trapping column contained a 5μm particle, the analytical column contained a 2.4μm particle. The column was heated to 55 °C using a column heater (Sonation). The sample was analyzed using 6 × 1.5 hours gradients. Six gas-phase fractions (GPF) injections were acquired for 6 ranges: 396–502, 496–602, 596–702, 696–802, 796–902, and 896–1002. Sequentially, full scan MS data (60,000 FWHM resolution) was followed by 26 × 4 m/z precursor isolation windows, another full scan and 26 × 4 m/z windows staggered by 2 m/z; products were acquired at 30,000 FWHM resolution. The automatic gain control (AGC) target was set to 1e6 for both full MS and production data. The maximum ion inject time (IIT) was set to 50 ms for full MS and dynamic mode for products with 9 data points required across the peak; the NCE was set to 30.
Sample analysis
Samples were randomized for acquisition. 1 μg per sample was analyzed by nano LC/MS with a Waters M-class HPLC system interfaced to a ThermoFisher Exploris 480. Peptides were loaded on a trapping column and eluted over a 75 μm analytical column at 350 nL/min; both columns were packed with XSelect CSH C18 resin (Waters); the trapping column contained a 5μm particle, the analytical column contained a 2.4 μm particle. The column was heated to 55 °C using a column heater (Sonation). Samples were analyzed using a 1.5-hour gradient. The mass spectrometer was operated in data-independent mode. Sequentially, full scan MS data (60,000 FWHM resolution) from m/z 385–1015 was followed by 61 × 10 m/z precursor isolation windows, another full scan from m/z 385–1015 was followed by 61 × 10 m/z windows staggered by 5 m/z; products were acquired at 15,000 FWHM resolution. The maximum ion inject time (IIT) was set to 50 ms for full MS and dynamic mode for products with 9 data points required across the peak; the NCE was set to 30.
Data processing
DIA data were analyzed using Scaffold DIA 3.2.1 (Proteome Software), which served several functions:
-
1.
Conversion of RAW files to mzML (ProteoWizard,) including deconvolution of staggered windows. Conversion to DIA format.
-
2.
Alignment based on retention times.
-
3.
Searching for data using the Prosit library (DLIB) and the chromatogram/reference library to create custom ELIB.
-
4.
Filtering of database search results using Percolator at 1% peptide false discovery rate (FDR).
-
5.
Calculation of peak areas for detected peptides using Encyclopedia (0.9.6). For each peptide, the 5 highest quality fragment ions were selected for quantitation.
Data were searched with the following parameters:
Enzyme: Trypsin
Database: UniProt Human
Fixed modification: Carbamidomethyl (C)
Precursor Mass Tolerance: 10 ppm
Fragment Mass Tolerance: 10 ppm
Library Fragment Tolerance: 10 ppm
Peptide FDR: 0.01
Protein FDR: 0.01
Peptide Length: 6–30AA
Max Missed Cleavages: 1
Min. Peptides: 2
Peptide Charge: 2–3
The sample table was exported from Scaffold DIA and further analyzed in Perseus software (version 1.6.14.0)75. Unpaired student’s t test was used for direct comparisons between low and high seeders. Statistical significance was set at P value lower than 0.05. Proteins with a cut-off threshold value higher than 1.33 or lower than 0.77 were considered as differentially expressed proteins. All mass spectrometry data files have been deposited to the ProteomeXchange Consortium76 via the PRIDE partner repository. Data are available with identifier PXD045736. The identification of significantly dysregulated regulatory/metabolic pathways in the primary motor cortex proteomic dataset was made through Metascape platform77 using default settings (minimum overlap: 3, minimum enrichment: 1.5, P < 0.01). The functional protein association network analysis was performed using NetworkAnalyst78. NFT proteome was obtained from Drummond E et al.41.
Histological analysis of tau pathologies
4 µm thick FFPE tissue sections of neocortical regions (frontal middle gyrus, primary motor cortex, parietal inferior and superior gyrus, temporal inferior, middle, and superior gyrus, occipital cortex), basal ganglia (anterior and posterior portion), thalamus and subthalamic nucleus, hippocampus, amygdala, midbrain, pons, medulla oblongata, and cerebellum with dentate nucleus, were examined. In addition to Hematoxylin and Eosin-Luxol Fast Blue, a mouse monoclonal antibody against phosphorylated tau at Serine 202 and Threonine 205 (AT8, 1:1000 dilution, ref: MN1020, Thermo Fisher Scientific) was used for immunohistochemistry. The EnVision detection kit, Peroxidase/DAB, Mouse (Dako) was used to visualize antibody immunoreactivity. For semi-quantitative analyses of observed tau cytopathologies (neuronal neurofibrillary tangles, oligodendroglial coiled bodies, tufted astrocytes), we used a 4-point scale: 0, absent; 1, mild (1–2/20× field of view); 2, moderate (3–5/20× field of view); and 3, severe ( > 5/20× field of view) density27,38. For the figure, images were acquired using a Nikon Eclipse Ci microscope, equipped with a DS-Fi3 microscope camera and NIS-Elements imaging software (Version 1.10.00; Nikon Instruments)
Brain heatmaps
Heatmaps were created to visualize HMW tau levels analyzed via western blot and cell-type-specific tau levels assessed through tau immunostaining. For each brain region, HMW tau levels were converted into a semi-quantitative score. Tau immunoreactivity was also evaluated and assigned a semi-quantitative score by a neuropathologist (G.G.K.).
In both types of heatmaps, yellow represents low levels, orange represents intermediate levels, and red indicates high levels of HMW tau or tau immunoreactivity. Gray was used for regions that were either unavailable or not evaluated, while white represents negative levels.
Each heatmap was generated using a template, with a corresponding color scale created to represent the semi-quantitative scores. Regions within the heatmaps were colored accordingly using Photoshop (2021, Adobe).
VISIUM spatial transcriptomics
Sample collection and sequencing
Primary motor cortex FFPE tissue blocks from four PSP patients with DV200 > 50% were selected for sectioning. Appropriately sized sections were placed within the frames of capture areas on the Visium Spatial Gene Expression Slide (PN-1000188, 10x Genomics) with one section in each capture area (6.5 × 6.5 mm). Tissues were deparaffinized, stained and decross-linked, followed by probe hybridization, ligation, release, and extension. Visium spatial gene expression FFPE libraries were constructed with a Visium Human Transcriptome Probe kit (PN-1000363, 10x Genomics) and Visium FFPE Reagent kit (PN-1000361, 10x Genomics) following the manufacturer’s guidance and sequenced on the Illumina NovaSeq 6000 platforms to achieve a depth of at least 25,000 mean read pairs.
Data quality control and integration
Tissue quality was assessed by calculating the proportion of mitochondrial RNA for each spot, total detected gene per spot and visually inspection of tissue images (Supplementary Fig. 9, Supplementary Table 4). Spots with fewer than 100 detected transcripts were excluded, and spots at the edges of two samples were excluded due to poor quality ( < 200 detected transcripts) detailed below:
-
HS3: Spots with an x-coordinate ≥121 were excluded
-
LS4: Spots with an x-coordinate ≥124 or a y-coordinate ≥75 were excluded
All four slices were merged, and normalized using sctransform (v0.3.3)79 to remove effect of library size and the proportion of mitochondrial reads. The top 2000 highly variable genes were identified using Seurat (v4.1.1)80, and batch effects were removed using canonical correlation analysis (cca). The integrated data was clustered using Louvain clustering with a resolution parameter of 0.8. Clusters were annotated using cell-type-specific markers from publicly available human AD data81 and manual inspection.
Deconvolution
Normal human primary motor cortex scRNAseq was downloaded from: GSE17433242, including cell-type annotations. Marker genes across cell-types were obtained based on any of the following criteria:
-
Known classical markers from the literature (Supplementary Table 5)
-
Classified as highly variable genes using Seurat (v4.4.0) in at least 6/17 different individuals
-
Being among the top 10 most upregulated marker genes for a particular cell-type (based on differential detection rate of a cell-type versus all other cell-types) in at least 6/17 different individuals
This resulted in 2276 unique marker genes for 22 cell-types being kept as the snRNAseq reference data for deconvolution. Raw snRNAseq counts were log2-normalized using a standard depth of 10,000 UMIs/nucleus with a pseudocount of 1. The average normalized expression per nucleus was calculated across all 17 normal samples to generate the expression level reference data.
This snRNAseq reference dataset was used to perform spot-level deconvolution using the depth-adjusted least-squares algorithm82 due to its higher sensitivity for rare, interspersed cell-types such as microglia and T-cells using default parameters. Resulting estimated proportions of cell-types were rounded to 4 decimal places.
Differential expression
Differential expression between high- and low-seeder samples was calculated using a negative binomial mixed-effect model with seeder-type and the cortical layers (Louvain clusters) as fixed effects and VISIUM slice as a random effect, as implemented in nebula (v1.4.2)83. Total UMI counts per spot were included as a fixed effect to reduce potential biases. Significance was assessed using 5% FDR to control for multiple testing. Genes expressed in fewer than 10% of tissue spots were excluded due to lack of power. When comparing across all four samples, the cortical layer was included as a fixed effect.
Quantification of tau load in Visium-analyzed regions of interest
For correlation of Visium gene expression data with AT8 tau pathology, the immediately adjacent sections (4-µm thickness) were stained using a mouse monoclonal antibody against phosphorylated tau at Serine 202 and Threonine 205 (AT8, 1:1000 dilution, ref: MN1020, Thermo Fisher Scientific). Stained sections were scanned using the Huron TissueScope LE120 (Huron) whole slide scanner. Scanned images were then collated with those of the Visium assay (10x Genomics), obtained from the Loupe Browser (10X Genomics). In the HALO software (version 3.5, Indica Labs), a 25 × 25 ROI-replica of the Visium assay was created using the annotation tool (each ROI at 55 µm in diameter and 100 µm distance between adjacent centers). Each ROI was annotated into individual layers (Layer 1–625) to obtain distinct quantification values. Using the collated image described above as a reference, the annotated template was then appropriately placed inside the gray matter at the central region of the primary motor cortex on the AT8-stained serial sections, placing the ROIs on top of the Visium ROIs. AT8 tau pathology in each ROI was quantified using the area quantification module, with results reported as the percentage of positively stained tissue area.
Differential expression in relation to tau protein expression, as estimated from the matched immunohistochemical image, was calculated on only the portions of two slices where AT8 tau pathology was reliably estimated. Quantitative AT8 tau pathology was the fixed effect, and tissue slice was included as a random effect. In addition, we binned AT8 tau pathology into four classes, which were recoded as an ordinal predictor: No AT8 tau pathology = 0, Low AT8 tau pathology = 1, Medium AT8 tau pathology = 2, High AT8 tau pathology = 3, and substituted for the quantitative estimate of AT8 tau pathology in the model. Due to the low power (two sections only) cortical layer was not included in these comparisons.
Pathway enrichments
Reactome pathways were downloaded and used for gene expression enrichment analysis (GSEA) as implemented in the fgsea (v1.20.0) package84. Pathways containing fewer than 10 genes, or more than 500 genes, were excluded to improve the power and specificity of enrichments. Genes were ranked by their estimated log2 fold-change and enrichment scores, and p values were calculated for each pathway using 50,000 permutations to assess significance. Multiple testing was controlled using a 5% FDR.
Statistics and reproducibility
No statistical method was used to predetermine sample size. Cohort size was limited by the availability of frozen and well-characterized postmortem PSP tissue in our brain collection. No analyses were stratified by sex due to the low sample size. No data were excluded from the analyses except when samples failed predefined technical quality criteria (e.g., failed tissue alignment in spatial transcriptomics), which are specified in the relevant figure legends or Methods subsections. The experiments were not randomized. Investigators were not blinded to allocation during experiments or outcome assessment, except for (i) tau biosensor cell assays, (ii) iAstrocyte cytotoxicity experiments, and (iii) neuropathological evaluation of the two high- and two low-seeder cases used for post-proteomic clustering, which were performed blinded to seeding category.
All 4R-tau SAAs were performed in quadruplicate per brain region. Proteinase K and thermolysin digestion assays were performed on the same PSP brains. Tau biosensor cell assays were run in duplicate for each sample, and iAstrocyte cytotoxicity experiments were run in triplicate. Replicate numbers and sample sizes are reported in the figure legends.
Statistical analyses were performed using GraphPad Prism (v9) with a significance threshold of P = 0.05. 4R-tau SAA fluorescence responses were analyzed and plotted in GraphPad Prism. For cytotoxicity assays, differences among untreated astrocytes, low-seeder homogenates and high-seeder homogenates were assessed by one-way ANOVA followed by Tukey’s post hoc test. Correlations were evaluated using Spearman’s rank correlation (r), with exact P values reported in the text or legends. Unsupervised hierarchical cluster analyses of PSP cases were performed using SPSS Statistics (v23).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available as follows. Source data underlying all main-text figures and key graphs are provided in the accompanying Source Data file, and full supporting datasets are provided in the Supplementary Information and the Source Data file. Raw and processed spatial transcriptomic sequencing data have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE302160. Raw proteomic mass spectrometry data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository under accession code PXD045736. All other data needed to interpret, verify and extend the conclusions of this work are included in the paper and its Supplementary Information. Additional information is available from the corresponding author upon request. Source data are provided with this paper.
Code availability
Code for spatial transcriptomic analysis is available on github: https://github.com/tallulandrews/PSP_Spatial.
References
Roesler, T. W. et al. Four-repeat tauopathies. Prog. Neurobiol. 180, 101644 (2019).
Dickson, D. W. Neuropathologic differentiation of progressive supranuclear palsy and corticobasal degeneration. J. Neurol. 246, II6–II15 (1999).
Hauw, J.-J. et al. Preliminary NINDS neuropathologic criteria for Steele-Richardson-Olszewski syndrome (progressive supranuclear palsy). Neurology 44, 2015–2015 (1994).
Roemer, S. F. et al. Rainwater Charitable Foundation criteria for the neuropathologic diagnosis of progressive supranuclear palsy. Acta Neuropathol. 144, 603–614 (2022).
Kovacs, G. G. Invited review: neuropathology of tauopathies: principles and practice. Neuropathol. Appl. Neurobiol. 41, 3–23 (2015).
Höglinger, G. U. et al. Clinical diagnosis of progressive supranuclear palsy: the movement disorder society criteria. Mov. Disord. 32, 853–864 (2017).
Respondek, G. et al. The phenotypic spectrum of progressive supranuclear palsy: a retrospective multicenter study of 100 definite cases. Mov. Disord. 29, 1758–1766 (2014).
Goedert, M., Eisenberg, D. S. & Crowther, R. A. Propagation of tau aggregates and neurodegeneration. Annu. Rev. Neurosci. 40, 189–210 (2017).
Hyman, B. All the Tau we cannot see. Annu. Rev. Med. 74, 503–514 (2023).
Sanders, D. W. et al. Distinct tau prion strains propagate in cells and mice and define different tauopathies. Neuron 82, 1271–1288 (2014).
Clavaguera, F. et al. Brain homogenates from human tauopathies induce tau inclusions in mouse brain. Proc. Natl. Acad. Sci. USA 110, 9535–9540 (2013).
Mirbaha, H. et al. Inert and seed-competent tau monomers suggest structural origins of aggregation. Elife 7, e36584 (2018).
Narasimhan, S. et al. Human tau pathology transmits glial tau aggregates in the absence of neuronal tau. J. Exp. Med. 217, e20190783 (2020).
Ferrer, I. et al. Involvement of Oligodendrocytes in Tau Seeding and Spreading in Tauopathies. Front. Aging Neurosci. 11, 112 (2019).
He, Z. et al. Transmission of tauopathy strains is independent of their isoform composition. Nat. Commun. 11, 7 (2020).
Xu, H. et al. In vitro amplification of pathogenic tau conserves disease-specific bioactive characteristics. Acta Neuropathol. 141, 193–215 (2021).
Priyanka, Qamar, S. H. & Visanji, N. P. Toward an animal model of Progressive Supranuclear Palsy. Front. Neurosci. 18, 1433465 (2024).
Darricau, M. et al. Tau seeds from patients induce progressive supranuclear palsy pathology and symptoms in primates. Brain 146, 2524–2534 (2023).
Shi, Y. et al. Structure-based classification of tauopathies. Nature 598, 359–363 (2021).
Kovacs, G. G. et al. Distribution patterns of tau pathology in progressive supranuclear palsy. Acta Neuropathol. 140, 99–119 (2020).
Sakae, N. et al. Neuropathologic basis of frontotemporal dementia in progressive supranuclear palsy. Mov. Disord. 34, 1655–1662 (2019).
Williams, D. R. et al. Characteristics of two distinct clinical phenotypes in pathologically proven progressive supranuclear palsy: Richardson’s syndrome and PSP-parkinsonism. Brain 128, 1247–1258 (2005).
Williams, D. R. et al. Pathological tau burden and distribution distinguishes progressive supranuclear palsy-parkinsonism from Richardson’s syndrome. Brain 130, 1566–1576 (2007).
Tsuboi, Y. et al. Increased tau burden in the cortices of progressive supranuclear palsy presenting with corticobasal syndrome. Mov. Disord. 20, 982–988 (2005).
Ahmed, Z., Josephs, K. A., Gonzalez, J., DelleDonne, A. & Dickson, D. W. Clinical and neuropathologic features of progressive supranuclear palsy with severe pallido-nigro-luysial degeneration and axonal dystrophy. Brain 131, 460–472 (2008).
Jecmenica Lukic, M. et al. Copathology in progressive supranuclear palsy: does it matter? Mov. Disord. 35, 984–993 (2020).
Couto, B. et al. Protracted course progressive supranuclear palsy. Eur. J. Neurol. 29, 2220–2231 (2022).
Takeda, S. et al. Neuronal uptake and propagation of a rare phosphorylated high-molecular-weight tau derived from Alzheimer’s disease brain. Nat. Commun. 6, 8490 (2015).
Takeda, S. et al. Seed-competent high-molecular-weight tau species accumulates in the cerebrospinal fluid of Alzheimer’s disease mouse model and human patients. Ann. Neurol. 80, 355–367 (2016).
Fukumoto, H., Kao, T. H., Tai, C. Y., Jang, M. K. & Miyamoto, M. High-molecular-weight oligomer tau (HMWoTau) species are dramatically increased in Braak-stage dependent manner in the frontal lobe of human brains, demonstrated by a novel oligomer Tau ELISA with a mouse monoclonal antibody (APNmAb005). FASEB J. 38, e70160 (2024).
Dujardin, S. et al. Tau molecular diversity contributes to clinical heterogeneity in Alzheimer’s disease. Nat. Med. 26, 1256–1263 (2020).
Mate De Gerando, A. et al. Tau seeding and spreading in vivo is supported by both AD-derived fibrillar and oligomeric tau. Acta Neuropathol. 146, 191–210 (2023).
Harris, S. S. et al. Alzheimer’s disease patient-derived high-molecular-weight tau impairs bursting in hippocampal neurons. Cell 188, 3775–3788.e3721 (2025).
Kim, C. et al. Distinct populations of highly potent TAU seed conformers in rapidly progressing Alzheimer’s disease. Sci. Transl. Med. 14, eabg0253 (2022).
Bongianni, M. et al. α-Synuclein RT-QuIC assay in cerebrospinal fluid of patients with dementia with Lewy bodies. Ann. Clin. Transl. Neurol. 6, 2120–2126 (2019).
Martinez-Valbuena, I. et al. α-Synuclein molecular behavior and nigral proteomic profiling distinguish subtypes of Lewy body disorders. Acta Neuropathol. 144, 167–185 (2022).
Martinez-Valbuena, I., Tartaglia, M. C., Fox, S. H., Lang, A. E. & Kovacs, G. G. Four-repeat Tau seeding in the skin of patients with progressive supranuclear palsy. JAMA Neurol. 81, 1228–1230 (2024).
Martinez-Valbuena, I. et al. Alpha-synuclein seeding shows a wide heterogeneity in multiple system atrophy. Transl. Neurodegener. 11, 7 (2022).
Holmes, B. B. et al. Proteopathic tau seeding predicts tauopathy in vivo. Proc. Natl. Acad. Sci. 111, E4376–E4385 (2014).
Lau, A. et al. α-Synuclein strains target distinct brain regions and cell types. Nat. Neurosci. 23, 21–31 (2020).
Drummond, E. et al. Phosphorylated tau interactome in the human Alzheimer’s disease brain. Brain 143, 2803–2817 (2020).
Pineda, S. S. et al. Single-cell dissection of the human motor and prefrontal cortices in ALS and FTLD. Cell 187, 1971–1989.e1916 (2024).
Martinez, P. et al. Bassoon contributes to tau-seed propagation and neurotoxicity. Nat. Neurosci. 25, 1597–1607 (2022).
Martinez-Valbuena I. Phenotype parallels protein seeding capacity in neurodegenerative diseases. Brain Pathol. 34, e13238 (2024).
Farrell, K. et al. Genetic, transcriptomic, histological, and biochemical analysis of progressive supranuclear palsy implicates glial activation and novel risk genes. Nat. Commun. 15, 7880 (2024).
Rexach, J. E. et al. Cross-disorder and disease-specific pathways in dementia revealed by single-cell genomics. Cell 187, 5753–5774.e5728 (2024).
Wang, X. et al. Alzheimer’s disease and progressive supranuclear palsy share similar transcriptomic changes in distinct brain regions. J. Clin. Invest. 132, e149904 (2022).
Whitney, K. et al. Single-cell transcriptomic and neuropathologic analysis reveals dysregulation of the integrated stress response in progressive supranuclear palsy. Acta Neuropathol. 148, 80 (2024).
DeVos, S. L. et al. Antisense reduction of tau in adult mice protects against seizures. J. Neurosci. 33, 12887–12897 (2013).
Wu, J. W. et al. Neuronal activity enhances tau propagation and tau pathology in vivo. Nat. Neurosci. 19, 1085–1092 (2016).
Fecto, F., Esengul, Y. T. & Siddique, T. Protein recycling pathways in neurodegenerative diseases. Alzheimers Res. Ther. 6, 13 (2014).
Eftekharzadeh, B. et al. Tau protein disrupts nucleocytoplasmic transport in Alzheimer’s disease. Neuron 99, 925–940.e927 (2018).
Wolozin, B. & Apicco, D. RNA binding proteins and the genesis of neurodegenerative diseases. Adv. Exp. Med Biol. 822, 11–15 (2015).
Nandakumar, S., Rozich, E. & Buttitta, L. Cell cycle re-entry in the nervous system: from polyploidy to neurodegeneration. Front. Cell Dev. Biol. 9, 698661 (2021).
Herrup, K., Neve, R., Ackerman, S. L. & Copani, A. Divide and die: cell cycle events as triggers of nerve cell death. J. Neurosci. 24, 9232–9239 (2004).
Guzman-Martinez, L. et al. Neuroinflammation as a common feature of neurodegenerative disorders. Front. Pharmacol. 10, 1008 (2019).
Kwon, H. S. & Koh, S.-H. Neuroinflammation in neurodegenerative disorders: the roles of microglia and astrocytes. Transl. Neurodegener. 9, 1–12 (2020).
Langworth-Green, C. et al. Chronic effects of inflammation on tauopathies. Lancet Neurol. 22, 430–442 (2023).
Chen, X. et al. Microglia-mediated T cell infiltration drives neurodegeneration in tauopathy. Nature 615, 668–677 (2023).
Malpetti, M. et al. Neuroinflammation predicts disease progression in progressive supranuclear palsy. J. Neurol. Neurosurg. Psychiatry 92, 769–775 (2021).
Couto, B., et al. Midbrain cytotoxic T cells as a distinct neuropathological feature of progressive supranuclear palsy. Brain 148, 2650–2657 (2025).
Hartnell, I. J. et al. Glial reactivity and T cell infiltration in frontotemporal lobar degeneration with tau pathology. Brain 147, 590–606 (2024).
Backman, E. A. et al. Nigral neuroinflammation and dopaminergic neurons in Parkinson’s disease and atypical parkinsonisms. Ann. Neurol. 97, 1096–1109 (2025).
Tsimberidou, A. M., Fountzilas, E., Nikanjam, M. & Kurzrock, R. Review of precision cancer medicine: evolution of the treatment paradigm. Cancer Treat. Rev. 86, 102019 (2020).
Elson, N. C., Lewis, J. D., Shaughnessy, E. A. & Reyna, C. Lessons from other fields of medicine, Part 1: breast cancer. Handb. Clin. Neurol. 192, 101–118 (2023).
Saijo, E. et al. 4-Repeat tau seeds and templating subtypes as brain and CSF biomarkers of frontotemporal lobar degeneration. Acta Neuropathol. 139, 63–77 (2020).
Wang, Z. et al. Seeding activity of skin misfolded tau as a biomarker for tauopathies. Mol. Neurodegener. 19, 92 (2024).
Miyoshi, E. et al. Spatial and single-nucleus transcriptomic analysis of genetic and sporadic forms of Alzheimer’s disease. Nat. Genet 56, 2704–2717 (2024).
Kovacs, G. G. Molecular pathology of neurodegenerative diseases: principles and practice. J. Clin. Pathol. 72, 725–735 (2019).
Dinkel, P. D., Siddiqua, A., Huynh, H., Shah, M. & Margittai, M. Variations in filament conformation dictate seeding barrier between three-and four-repeat tau. Biochemistry 50, 4330–4336 (2011).
Von Bergen, M. et al. Mutations of tau protein in frontotemporal dementia promote aggregation of paired helical filaments by enhancing local β-structure. J. Biol. Chem. 276, 48165–48174 (2001).
Friedhoff, P., Schneider, A., Mandelkow, E.-M. & Mandelkow, E. Rapid assembly of Alzheimer-like paired helical filaments from microtubule-associated protein tau monitored by fluorescence in solution. Biochemistry 37, 10223–10230 (1998).
Friedhoff, P., Von Bergen, M., Mandelkow, E.-M., Davies, P. & Mandelkow, E. A nucleated assembly mechanism of Alzheimer paired helical filaments. Proc. Natl. Acad. Sci. 95, 15712–15717 (1998).
Jessie, K., Hashim, O. H. & Rahim, Z. Protein precipitation method for salivary proteins and rehydration buffer for two-dimensional electrophoresis. Biotechnology 7, 686–693 (2008).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote) omics data. Nat. Methods 13, 731–740 (2016).
Vizcaíno, J. A. et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 32, 223–226 (2014).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Zhou, G. et al. NetworkAnalyst 3.0: a visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res. 47, W234–W241 (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587. e3529 (2021).
Brase, L. et al. Single-nucleus RNA-sequencing of autosomal dominant Alzheimer disease and risk variant carriers. Nat. Commun. 14, 2314 (2023).
Tsoucas, D. et al. Accurate estimation of cell-type composition from gene expression data. Nat. Commun. 10, 2975 (2019).
He, L. et al. NEBULA is a fast negative binomial mixed model for differential or co-expression analysis of large-scale multi-subject single-cell data. Commun. Biol. 4, 629 (2021).
Korotkevich, G. et al. Fast gene set enrichment analysis. BioRxiv, https://www.biorxiv.org/content/10.1101/060012v3 (2016).
Acknowledgements
The authors would like to acknowledge the patients and their families for their donation. We would also like to acknowledge Dr. Michael Ford and MS Bioworks for their support with the mass spectrometry, and Farzaneh Aboualizadeh from the Princess Margaret Genomic Center and Rami Michael from 10x Genomics for their support with the spatial transcriptomics. This study was supported by the Rossy Family Foundation, the Edmond J. Safra Philanthropic Foundation, the Krembil Foundation, the Maybank Foundation (to G.G.K. and A.E.L.), the Blidner Family Foundation (to I.M.V. and N.P.V.), Ajay Virmani (to I.M.V.), the Canadian Foundation for Innovation (CFI) John R. Evans Leaders Fund (40480) (to G.G.K), the National Institute on Aging of the National Institutes of Health under Award Number R01AG080001 (to I.M.V., S.L.F., and G.G.K.) and the Ontario Research Fund for Small Infrastructure Funds (to G.G.K.). E.S. and J.F.-I. were supported by a grant from the Spanish Ministry of Science, Innovation, and Universities (Ref. PID2019-110356RB-I00/AEI/10.13039/501100011033). E.R. was supported by the G. Harry Sheppard Memorial Research Fund. M.C.T. holds the Marion and Gerald Soloway Chair in Brain Injury and Concussion Research. M.I. was supported by the Krembil Family, and G.G.K. holds the Rossy Chair in PSP research at UHN. The funding bodies did not take part in the design of the study, in the collection, analysis, or interpretation of data, or in writing the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: I.M.V., A.E.L., G.G.K. Methodology: I.M.V. (4R-Tau SAA); S.L. (biosensor cells, spatial transcriptomics); S.Z., M.I. (size-exclusion chromatography); J.L. (protein extraction, western blotting); S.H.Q. (Western blot quantification); A.M.K. (biosensor cell experiments, immunohistochemistry); S.L.F., G.G.K. H.T. (neuropathology); K.S., E.R. (genetics). Formal analysis: I.M.V.; S.L. (biosensor & spatial transcriptomics); E.S., J.F.I. (proteomics); S.L.F., A.K., G.G.K. (neuropathological assessment/clustering); S.H.Q. (Western blotting quantification); K.S., E.R. (genetics); T.A. (spatial transcriptomics). Data curation: E.S., J.F.I. (proteomics); B.C., N.G.R. (clinical data). Resources (participants/brain donations): S.H.F., M.C.T., N.P.V., A.E.L., G.G.K. Writing—original draft: I.M.V. Writing—review & editing: All authors. Supervision: A.E.L., G.G.K. Project administration: A.E.L., G.G.K. Funding acquisition: A.E.L., G.G.K.
Corresponding author
Ethics declarations
Competing interests
I.M.V., G.G.K., and A.E.L. are inventors on a pending patent application related to diagnostic assays for movement disorders. The patent applicant is the inventors (I.M.V., G.G.K., and A.E.L.) and the University Health Network. The application is titled Diagnostic assays for movement disorders and has the application number 18/537,455. The patent is currently pending. The application includes the SAA used in this study; however, in this manuscript, the assay is used solely as a research tool to classify brain tissue samples and not for diagnostic purposes. G.G.K. has a shared patent for 5G4 Synuclein antibody. All other authors declare no competing interests. Outside the submitted work, I.M.V. receives funding from the Michael J. Fox Foundation for Parkinson's Research and consultancy fees from Ferrer. SLF receives funding from the National Health and Medical Research Council, Australia. M.I. is a paid consultant to BioArctic AB. M.C.T receives funding from NIH, Weston Brain Foundation, Tanenbaum Institute for Research in Science of Sport, Canadian Institutes of Health Research, and in-kind funding from Roche; she has served as an advisor to Eisai, Lilly and Novo Nordisk and she conducts clinical trials for Biogen, Anavex, Janssen, Novo Nordisk, BMS, Aribio, Green Valley, UCB, Passage Bio. S.H.F. receives research Funding from the Michael J Fox Foundation for Parkinson's Research, NIH (Dystonia Coalition), Parkinson Canada, and the Weston Foundation. Honoraria from the International Parkinson and Movement Disorder Society. Consultancy/Speaker fees from AbbVie, Lundbeck, Sunovion and Royalties from Oxford University Press. A.E.L. has served as an advisor for AbbVie, Amylyx, Aprinoia, Biogen, BioAdvance, Biohaven, BioVie, BlueRock, BMS, Denali, EG427, Ferrer, Janssen, Lilly, Northera, Pharma 2B, Sun Pharma, UCB; and Ventyx Bio; received honoraria from Sun Pharma and AbbVie; received grants from Brain Canada, Canadian Institutes of Health Research, Edmond J Safra Philanthropic Foundation, Krembil Brain Institute, Michael J. Fox Foundation, Parkinson Foundation, Parkinson Canada, Weston Foundation; is serving as an expert witness in litigation related to paraquat and Parkinson’s disease, received publishing royalties from Elsevier, Saunders, Wiley-Blackwell, Johns Hopkins Press, and Cambridge University Press. G.G.K. reports personal fees from Parexel, other funding from Rossy Family Foundation, from Edmond Safra Foundation, grants from Krembil Foundation, MSA Coalition, Michael J. Fox Foundation, Parkinson Canada, NIH, Canada Foundation for Innovation, and Ontario Research Fund; in addition, G.G.K. received royalties from Wiley, Cambridge, and Elsevier publishers. None of these had an influence on this study. S.L.; E.S.; J.F-I.; S.L.; J.L.; H.T.; B.C.; N.G.R; S.H.Q.; A.M.K.; A.K.; K.S.; E.R.; N.P.V. and T.A. declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Todd Golde and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Martinez-Valbuena, I., Lee, S., Santamaria, E. et al. 4R-tau seeding activity reveals molecular subtypes in progressive supranuclear palsy. Nat Commun 17, 1006 (2026). https://doi.org/10.1038/s41467-025-67744-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67744-y