Abstract
Plant viruses are causal agents of devastating diseases in crops and pose a threat to food security. Viruses transmitted by phloem-feeding insects are frequently restricted to the phloem; the mechanism determining this tissue tropism and its consequences for the viral cycle have remained elusive. Here we show that phloem restriction of tobacco curly shoot virus (TbCSV) depends on the allele of the C4 gene it carries. The Y35 allele produces a plasma membrane-associated C4 protein; TbCSV(Y35) is phloem-limited. The Y41 allele, however, produces two C4 variants, targeted to the plasma membrane and chloroplasts, respectively. Chloroplast-localized C4 interferes with salicylic acid-mediated defenses, which causes a destabilization of PENETRATION3 (PEN3), a subsequent decrease in callose deposition, and the escape of TbCSV(Y41) to the surrounding parenchyma cells. Interestingly, Y41 expands the host range of TbCSV, but Y35 is prevalent in nature. We determine that phloem restriction favors acquisition and transmission by the insect vector, conferring a competitive advantage to TbCSV(Y35). Importantly, PEN3 also determines phloem restriction of an RNA virus. In summary, we demonstrate that PEN3 activity and likely callose deposition confine viruses to the phloem, which favors viral spread by facilitating acquisition by the insect vector.
Similar content being viewed by others
Introduction
Plant viruses cause devastating diseases in crops worldwide, posing a serious threat to global food security. Most plant viruses are transmitted by phloem-feeding insects; following transmission by the vector, many viruses remain restricted to the phloem1,2,3,4. Despite the prevalence of this tissue tropism, the underlying molecular mechanisms as well as its significance for viral performance have so far remained elusive.
Insect-transmitted viruses from the Geminiviridae family are causal agents of economically relevant diseases affecting cash and staple crops in tropical and temperate regions of the world. Most geminiviruses are transmitted by the phloem-feeding whitefly Bemisia tabaci, and most of these are phloem-limited. Many species within this family encode a protein named C4, which is versatile and essential for full infectivity, suppressing plant anti-viral defenses and mediating symptom development5,6,7. C4 proteins from different geminiviruses possess different combinations of targeting signals and show specific subcellular localizations6; the C4 protein encoded by tomato yellow leaf curl virus (TYLCV), for example, displays a dual localization at the plasma membrane and in chloroplasts, moving from the former to the latter during infection8. At the plasma membrane, C4 from TYLCV suppresses the cell-to-cell spread of silencing, while in chloroplasts it interferes with the activation of anti-viral salicylic acid (SA)-dependent defenses8,9. This subcellular compartmentalization that enables multi-functionality of C4 seems to be relevant for the infection, as geminiviruses have evolved independent strategies to accomplish it. Recently, it has been demonstrated that some geminiviruses encode two variants of the C4 protein in their C4 gene, differing in the two N-terminal residues only, resulting from translation initiation from two alternative in-frame start codons; one of these variants localizes to the plasma membrane, while the other is translocated to chloroplasts10. This is the case of tomato golden mosaic virus (TGMV), whose plasma membrane-localized C4 protein inhibits receptor kinases-mediated anti-viral defenses and triggers leaf curling symptoms, while the chloroplast-localized variant promotes degradation of this organelle and hence yellowing symptoms10.
An interesting case is that of tobacco curly shoot virus (TbCSV), of which different isolates coexisting in the field differ in their capacity to encode either one or two C4 variants10, offering a unique opportunity to evaluate the biological relevance of this diverging property. We show that phloem restriction of TbCSV depends on the allele of the C4 gene it carries. The Y35 allele (TbCSV(Y35)) produced a plasma membrane-associated C4 protein that is phloem-limited. The Y41 allele TbCSV(Y41) produced two C4 variants, targeting the plasma membrane and chloroplasts, respectively, that compromise phloem restriction. The chloroplast-localized C4 causes a destabilization of PEN3 (an ATP-binding cassette transporter), a subsequent decrease in callose deposition, and the escape of TbCSV(Y41) to the surrounding parenchyma cells. Importantly, we demonstrate that PEN3 activity, likely through callose deposition, confines viruses to the phloem, which favors viral spread by facilitating acquisition by the insect vector.
Results
TbCSV (Y41) isolates have a lower epidemic rate
Most TbCSV isolates encoding only one C4 variant (localized to the plasma membrane) infect plant species from the Solanaceae family exclusively, while those isolates encoding two variants can additionally infect members of the Amaranthaceae, Cucurbitacea, and Compositae families (Supplementary Fig. 1, Supplementary Table 1). Elegant allele-swap experiments have demonstrated that the presence of the second, chloroplast-localized C4 variant is sufficient to expand the host range of the virus10. Two TbCSV isolates found co-existing in the same geographical area (Yunnan, China) were selected for further comparative characterization: TbCSV(Y35), which carries the Y35 C4 allele, producing only one, plasma membrane-associated, C4 protein, and TbCSV(Y41), which carries the Y41 C4 allele, producing two C4 proteins differing in the two N-terminal amino acids and targeted to the chloroplast and the plasma membrane, respectively.
Intriguingly, despite the increase in potential hosts that the Y41 allele enables, isolates carrying the Y35 allele are prevalent in China (Fig. 1a, Supplementary Table 2), pointing to a potential trade-off imposed by the Y41 allele. To dissect the impact of having two C4 variants for viral performance and identify potential negative effects, a TbCSV(Y35) virus carrying the Y41 C4 allele (TbCSV(Y35-2C4); Fig. 1b) was generated. This recombinant virus triggered the typical yellowing symptoms associated with the chloroplast-localized C4 protein in infected tomato plants (Fig. 1c). All three viruses accumulated to similar levels, indicating that viral performance in the plant is not differentially affected by these C4 alleles (Fig. 1d).
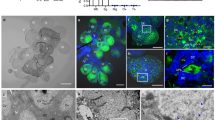
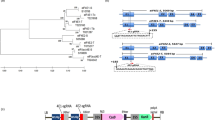
a TbCSV isolates containing one C4 variant only is prevalent in Yunnan, China. The proportion of TbCSV isolates containing two C4 variants is indicated in blue; the proportion of TbCSV isolates harboring only one C4 variant is indicated in red. TbCSV isolates shown in Fig. 1a were collected from tomato plants in Yunnan, China, between 2010 and 2017. b Nucleotide sequences complementary to the virion-sense at the start of the C4 gene of TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4). The encoded amino acid sequences of C4 and C1, encoded in the overlapping gene, are shown. The ATG of the C4 gene is indicated in red letters. c Symptoms caused by TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4) in tomato plants at 28 days post-inoculation (dpi). d Southern blot analysis of TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4) accumulation in tomato plants at 28 dpi. Total DNA was hybridized with a TbCSV CP probe. Total nucleic acids (50 μg DNA) were extracted from tomato plants inoculated with TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) infectious clones at 28 dpi. Total genomic DNA visualized by ethidium bromide staining is shown as a loading control. e Immunofluorescence detection of TbCSV in midguts of Zhejiang II whiteflies after feeding on TbCSV(Y35)-, TbCSV(Y41)-, or TbCSV(Y35-2C4)-infected tomato plants. TbCSV was detected using a monoclonal antibody against the TbCSV coat protein (CP), followed by a commercial 549-conjugated secondary antibody (red); nuclei are stained with DAPI (blue). At least 10 midguts were examined for each treatment. Representative images are shown. Scale bar = 100 μm. f Immunofluorescence detection of TbCSV in primary salivary glands of Zhejiang II whiteflies after feeding on TbCSV(Y35)-, TbCSV(Y41)-, or TbCSV(Y35-2C4)-infected tomato plants. As in (e). At least 10 primary salivary glands were examined for each treatment. Representative images are shown. Scale bar = 20 μm. g, h Immunoblot analysis of virus accumulation in whiteflies after feeding on TbCSV(Y35)-, TbCSV(Y41)-, or TbCSV(Y35-2C4)-infected tomato plants. g Accumulation of TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4) CP in virus-infected tomato plants. Actin was used as a loading control. h TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4) CP accumulation in whiteflies after feeding on virus-infected tomato plants. Histone H3 was used as a loading control. i Immunofluorescence detection of TbCSV in primary salivary glands of Zhejiang II whiteflies after feeding on artificial feed containing TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) virions. At least 10 primary salivary glands were examined for each treatment. Representative images are shown. Scale bar = 20 μm. j Transmission efficiency of TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) by Zhejiang II whiteflies. The upper, middle, and lower panels indicate different numbers of viruliferous whitefly adults (1, 5, or 15, respectively) used per plant. Tomato plants were given an inoculation access period of 72 h with 1, 5, or 15 viruliferous whiteflies using clip cages in three independent biological replicates. Non-viruliferous whiteflies were used as a mock treatment. Photographs were taken 14 days after the inoculation access period. Experiments in (d–h, i) were repeated three times with similar results.
In the absence of changes in the infection per se, effects on insect transmission can account for the differential prevalence of the isolates in nature. To test whether transmissibility was affected by the presence of the two C4 variants, whiteflies were put to feed on tomato plants infected with each of these three viruses, and the presence of the pathogen was detected in the midgut and salivary glands of the insects (Fig. 1e–h). These experiments clearly showed that TbCSV isolates expressing two C4 variants (TbCSV (Y41), TbCSV (Y35-2C4)) accumulated to a lower extent in the insect vector; of note, this is not due to differences in the viral capsid that might affect acquisition, since accumulation of virions added to the insect feed in the whitefly organs was similar (Fig. 1i, Supplementary Fig. 2). As expected from the decreased presence of the virus in the insect vector, transmission efficiency of the isolates with two C4 variants was diminished significantly (Fig. 1j, Supplementary Fig. 3).
The Y41 allele compromises phloem restriction
Chloroplast-localized C4 has been linked to yellowing symptoms in TGMV10. Intriguingly, the presence of the Y41 or the Y35-2C4 alleles in TbCSV resulted in yellowing symptoms in the leaf lamina, not only in tomato (Fig. 1c), but also in the experimental host Nicotiana benthamiana (Fig. 2a). This phenotype prompted the idea that the chloroplast-localized C4 might be acting outside the phloem, the tissue TbCSV is considered restricted to. Differences in symptoms do not rely on changes in viral accumulation in N. benthamiana either (Fig. 2b). Strikingly, immunolocalization of the viral capsid protein (CP) in cross-sections of leaves (Fig. 2c) or leaf veins (Fig. 2d, e) from infected N. benthamiana plants demonstrates that viruses encoding a chloroplast-localized C4 protein (TbCSV(Y41), TbCSV(Y35-2C4)) are not only detected in the phloem of both sides of the bicollateral vascular bundles but also in parenchyma cells outside of the phloem in over 70% of the images, as opposed to TbCSV(Y35) detected only in the phloem, which carries the Y35 allele. In particular, the comparison between TbCSV(Y35) and TbCSV(Y35-2C4) allows the conclusion that the chloroplast-localized C4 is sufficient to compromise phloem restriction, enabling the virus to reach parenchyma cells.
a Viral symptoms induced by TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4) in Nicotiana benthamiana plants at 14 days post-inoculation (dpi). b Southern blot analysis of TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4) accumulation in N. benthamiana plants at 14 dpi. Total DNA was hybridized with a TbCSV CP probe. Total nucleic acids (50 μg DNA) were extracted from N. benthamiana plants inoculated with TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) infectious clones at 14 dpi. Total genomic DNA visualized by ethidium bromide staining is shown as a loading control. c Immunofluorescence detection of TbCSV in leaves of N. benthamiana plants infected by TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4). The distribution of TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) was visualized in a cross-section using an antibody against the TbCSV coat protein (CP) (green). d TbCSV strains expressing two C4 variants (Y41 and Y35-2C4) overcome phloem restriction and reach parenchyma tissue in N. benthamiana plants. The distribution of TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) was visualized in a cross-section using an antibody against TbCSV CP (red). Autofluorescence of highly lignified tissues is shown in blue. PC, parenchyma; P, phloem; X, xylem. White arrowheads indicate TbCSV in parenchyma cells. Scale bar = 50 μm. Statistical analysis showed that TbCSV(Y35) is strictly limited to the phloem in N. benthamiana plants, but TbCSV(Y41) and TbCSV(Y35-2C4) can be observed overcoming phloem restriction and reaching parenchyma cells in over 70% of the images. At least eight individual samples of TbCSV(Y35)-infected N. benthamiana plants, five individual samples of TbCSV(Y41)-infected N. benthamiana plants and six individual samples of TbCSV(Y35-2C4)-infected N. benthamiana plants from three independent biological replicates were used for immunochemistry and statistical analysis. e Immunofluorescence detection of TbCSV in leaf vein sections of AtSWEET11::GFP transgenic N. benthamiana plants. The distribution of TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) and phloem cells was visualized using antibodies against TbCSV CP (red) or GFP (green). Autofluorescence of highly lignified tissues is shown in blue. PC, parenchyma; P, phloem; X, xylem. White arrowheads indicate TbCSV in parenchyma cells. Scale bar = 50 μm. Experiments in (b–d, e) were repeated at least three times with similar results.
Phloem restriction likely requires PEN3-dependent callose deposition
The yellowing of the leaf lamina observed in plants infected with TbCSV carrying the Y41 allele is reminiscent of that displayed by pen3-1 mutant Arabidopsis thaliana plants upon inoculation with the powdery mildew Erysiphe cichoracearum11. PEN3 encodes a plasma membrane-localized ATP-binding cassette transporter required for pathogen-induced callose deposition, which impairs pathogen penetration in a salicylic acid (SA)-dependent manner11,12. The PEN3 orthologue in N. benthamiana, NbPEN3, also localizes to the plasma membrane (Fig. 3a). Since callose deposition at plasmodesmata is known to restrict movement of viruses4,13,14,15,16,17, we sought to investigate whether PEN3 is involved in the phloem limitation of TbCSV(Y35). For this purpose, two independent NbPEN3 knock-out N. benthamiana lines were obtained through CRISPR-Cas9-mediated genome editing (Fig. 3b, Supplementary Fig. 4); these mutant lines showed largely decreased levels of callose (Supplementary Fig. 4b, c). Remarkably, nbpen3 mutants developed yellowing in the leaf lamina during infection with all three TbCSV viruses, regardless of their capacity to encode a chloroplast-localized C4 protein; viral accumulation, nevertheless, remained unchanged (Fig. 3c, d). In agreement with the correlation between phloem escape and leaf yellowing previously established, all three viruses could exit the phloem and reach parenchyma cells in nbpen3 mutant lines (Fig. 3e). Phloem restriction of TbCSV(Y35) in wild-type plants was associated with strong virus-induced PEN3-dependent callose deposition surrounding the vasculature and higher virus accumulation in the phloem (Fig. 3f, Supplementary Fig. 5), which was not observable in wild-type plants infected with TbCSV(Y41) nor in nbpen3 mutants in any case. Our results, therefore, indicate that PEN3-mediated callose deposition is triggered by the TbCSV(Y35) infection and can effectively restrict this viral strain and increase viral titer in the phloem, while TbCSV(Y41) can likely avoid or suppress this response through the action of its chloroplast-localized C4 protein. To test whether callose deposition limits virus to the phloem, we treated TbCSV(Y35)-infected N. benthamiana plants with 2-deoxy-D-glucose (2-DDG), an inhibitor of callose synthesis. Immunoblot and confocal results showed that TbCSV(Y35) escapes the phloem to the surrounding parenchyma cells in 2-DDG-treated leaves, which correlates with lower callose accumulation (Supplementary Fig. 6). These results indicate a direct link between phloem limitation and callose deposition and suggest that the latter plays a critical role in phloem restriction of viruses.
a Cellular distribution of NbPEN3-GFP in epidermal cells of H2B-RFP transgenic N. benthamiana plants. Scale bar = 50 μm. b Immunofluorescence detection of NbPEN3 in leaf vein sections of wild-type (WT) and nbpen3 knock-out N. benthamiana plants. The distribution of NbPEN3 was visualized in a cross-section using an antibody against PEN3 (green). Autofluorescence of highly lignified tissues is shown in blue. PC, parenchyma; P, phloem; X, xylem. Scale bar = 50 μm. c Viral symptoms induced by TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4) in WT and nbpen3 knock-out N. benthamiana plants at 14 days post-inoculation (dpi). d Immunoblot analysis of TbCSV(Y35), TbCSV(Y41), and TbCSV(Y35-2C4) coat protein (CP) accumulation in WT and nbpen3 knock-out N. benthamiana plants at 14 dpi. TbCSV CP was detected using a monoclonal antibody. Actin was used as a loading control. e Immunofluorescence detection of TbCSV in leaf vein sections of WT and nbpen3 knock-out N. benthamiana plants. The distribution of TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) was visualized in a cross-section using an antibody against the TbCSV CP (red). Autofluorescence of highly lignified tissues is shown in blue. White arrowheads indicate TbCSV in parenchyma cells. Scale bar = 50 μm. In TbCSV(Y35)-infected pen3-ko N. benthamiana plants, over 50% of images show virus signal in parenchyma cells. At least eleven individual samples of TbCSV(Y35)-infected pen3-ko N. benthamiana plants, ten individual samples of TbCSV(Y41)-infected pen3-ko N. benthamiana plants and twelve individual samples of TbCSV(Y35-2C4)-infected pen3-ko N. benthamiana plants from three independent biological replicates were used for immunochemistry and statistical analysis. f Callose deposition in leaf vein sections of WT and nbpen3 knock-out N. benthamiana plants infected by TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) at 14 dpi. Callose was stained by using aniline blue. White arrowheads indicate virus-induced callose deposition around the vasculature. Scale bar = 50 μm. Experiments in (a, c–f) were repeated at least three times with similar results.
A Y41-encoded C4 protein variant enables phloem escape
With the aim to elucidate how TbCSV(Y41) overcomes the PEN3-dependent phloem restriction, we first measured NbPEN3 transcript and protein accumulation in the presence of TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4). While no changes were observed in the levels of NbPEN3 mRNA (Fig. 4a), the NbPEN3 protein accumulation was largely decreased in the presence of TbCSV encoding the chloroplast-localized C4 (Fig. 4b). This effect could also be observed at the tissue level: while TbCSV(Y35) and PEN3 co-localized, cells infected with TbCSV(Y41) or TbCSV(Y35-2C4) were devoid of detectable PEN3 (Fig. 4c). Chloroplast-localized C4 has no effects on the NbPEN3 transcription, but was sufficient to trigger the reduction in NbPEN3 content and callose deposition, which could also be observed in transgenic N. benthamiana plants expressing the C4 proteins encoded by the Y41 or the Y35-2C4 alleles, but not the Y35 allele (Fig. 4d, e, Supplementary Fig. 7).
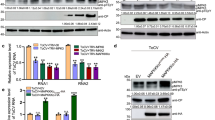
a Relative transcript of NbPEN3 in uninfected, TbCSV(Y35)-, TbCSV(Y41)- or TbCSV(Y35-2C4)-infected N. benthamiana plants. Relative accumulation of NbPEN3 transcripts is normalized against that of Actin. Statistical differences were analyzed by a two-sided, unpaired Student’s t-test (ns: not significant). Data are present as mean values ± SD of three biological replicates. Individual P-values are denoted above the comparison lines. b Western blot of NbPEN3 in uninfected, TbCSV(Y35)-, TbCSV(Y41)-, or TbCSV(Y35-2C4)-infected N. benthamiana plants. NbPEN3 was detected using a commercial antibody. Accumulation of Actin is shown as a control. c Immunofluorescence detection of TbCSV and NbPEN3 in sections N. benthamiana plants infected by TbCSV(Y35), TbCSV(Y41) or TbCSV(Y35-2C4), or mock-inoculated. The distribution of TbCSV(Y35), TbCSV(Y41) or TbCSV(Y35-2C4) was visualized in a cross-section of leaf vein tissues using an antibody against the TbCSV coat protein (CP) (red). The distribution of NbPEN3 was visualized in a cross-section using an antibody against PEN3 (green). Autofluorescence of highly lignified tissues is shown in blue. White arrowheads indicate TbCSV CP in parenchyma cells. Scale bar = 50 μm. d Phenotype of 35S::TbCSV(Y35) C4-Flag, 35S::TbCSV(Y41) C4-Flag, and 35S::TbCSV(Y35-2C4) C4-Flag transgenic N. benthamiana plants. e Western blot of NbPEN3 in wild-type (WT), 35S::TbCSV(Y35) C4-Flag, 35S::TbCSV(Y41) C4-Flag, and 35S::TbCSV(Y35-2C4) C4-Flag transgenic N. benthamiana plants. As in (b). f Relative transcript accumulation of NbPR1 in WT, 35S::TbCSV(Y35) C4-Flag, 35S::TbCSV(Y41) C4-Flag, and 35S::TbCSV(Y35-2C4) C4-Flag transgenic N. benthamiana plants. Relative accumulation of NbPR1 transcripts is normalized to that of the Actin transcript. Statistical differences were analyzed by two-sided, unpaired Student’s t-test (ns: not significant, **P < 0.01, ***P < 0.001). Data are present as mean values ± SD of three biological replicates. Individual P values are denoted above the comparison lines. g, h Identification of the interaction between NbCAS and the chloroplast-localized C4 in vivo and in yeast. g BiFC analysis of the interaction between the chloroplast-localized C4 and NbCAS in epidermal cells of H2B-RFP transgenic N. benthamiana plants. The combination of NbCAS-cYFP/TbCSV(Y35) C4-nYFP serves as a negative control. Scale bar = 20 μm. h Identification of the interaction between NbCAS and TbCSV(Y41) C4 in yeast. The yeast strain Gold co-transformed with the indicated plasmids was subjected to 10-fold serial dilutions and grown on a SD/-Leu/-Trp/-His medium. i Subcellular localization of GFP-NbPEN3 in epidermal cells of WT or NahG transgenic N. benthamiana plants. Scale bar = 20 μm. j Western blot analysis of NbPEN3 in WT and NahG transgenic N. benthamiana plants. k Viral symptoms induced by TbCSV(Y35) in WT and NahG transgenic N. benthamiana plants at 14 dpi. l Immunofluorescence detection of TbCSV(Y35) in sections of WT and NahG transgenic N. benthamiana plants. The distribution of TbCSV(Y35) was visualized in a cross-section of leaf vein tissues using an antibody against the TbCSV coat protein (CP) (red). Autofluorescence of highly lignified tissues is shown in blue. PC, parenchyma; P, phloem; X, xylem. White arrowheads indicate TbCSV in parenchyma cells. Scale bar = 50 μm. In TbCSV(Y35)-infected NahG transgenic N. benthamiana plants, over 60% of images show virus signal in parenchyma cells. At least five individual samples of TbCSV(Y35)-infected NahG transgenic N. benthamiana plants from three independent biological replicates were used for immunochemistry and statistical analysis. Experiments in (c, i, j, l) were repeated at least three times with similar results.
Chloroplast-dependent SA-mediated defenses have been shown to play an anti-geminiviral role, and to be suppressed by C4 orthologues from other geminivirus species8,18. In order to test whether the TbCSV-encoded chloroplast-localized C4 interferes with the activation of SA signaling, expression of the SA-responsive gene NbPR1 was evaluated in the transgenic plants expressing the different C4 proteins. As shown in Fig. 4f, the presence of a chloroplast-localized C4 version is sufficient to hamper the accumulation of the NbPR1 transcript, hence suggesting that this viral protein can suppress SA defenses. To elucidate the underlying molecular mechanism, we tested the interaction between the C4 protein encoded by the Y41 allele and NbCAS, a proven target of other geminivirus-encoded chloroplast-localized C4 proteins8. As shown in Fig. 4g, h, Y41 C4 interacts with NbCAS. In addition, other C4 proteins have been shown to compromise chloroplast integrity10, we observed the chloroplast ultrastructure of wild-type and 35S::TbCSV(Y35) C4 transgenic N. benthamiana plants is not different from that of transgenic N. benthamiana plants expressing the chloroplast-localized C4, but fewer osmiophilic granules can be detected following osmic acid staining in 35S::TbCSV(Y41) C4 and 35S::TbCSV(Y35-2C4) C4 transgenic plants (Supplementary Fig. 8). These results suggest that the chloroplast-localized C4 might interfere with SA signaling through the physical interaction with NbCAS and/or by disrupting the redox balance in chloroplasts. Importantly, the stability of NbPEN3 is regulated in a SA-dependent manner, since NbPEN3 was destabilized in the absence of SA in transgenic NahG plants, encoding a bacterial salicylate hydroxylase that degrades this plant hormone (Fig. 4i, j, Supplementary Fig. 9a). Consistent with these results, SA treatment compromised the degradation of NbPEN3, which confirms the critical role of SA in the regulation of NbPEN3 stability (Supplementary Fig. 9b). As previously observed for the nbpen3 mutant lines, NahG plants also displayed yellowing of the leaf lamina when infected with TbCSV(Y35), which correlated with the escape of this virus outside of the phloem and the absence of callose deposition in infected tissues (Fig. 4k, I, Supplementary Fig. 10). Taken together, these results indicate that the chloroplast-localized C4 protein encoded by the Y41 allele reduces the PEN3 content in infected cells, most likely through the suppression of SA signaling, since SA depletion is sufficient to abrogate PEN3 accumulation and complement the lack of a chloroplast-localized C4 in overcoming phloem restriction.
PEN3-facilitated phloem restriction favors acquisition of the virus by the insect vector
The Y41 C4 allele has a negative impact on insect transmission of the virus (Fig. 1e–j); since the chloroplast-localized C4 protein variant encoded by the Y41 allele suppresses SA signaling, leads to decreased PEN3 accumulation, and in turn enables viral phloem escape, and considering that the insect vector, B. tabaci, is a phloem feeder, it could be hypothesized that invasion of parenchyma would lead to viral dilution, which may ultimately diminish transmission efficiency. To test this idea, we first knocked out PEN3 in tomato (SlPEN3) (Supplementary Fig. 11) and evaluated the effect of this mutation on the viral infection. As shown in Fig. 5a, slpen3 plants displayed yellowing of the leaf lamina upon TbCSV infection irrespective of the C4 allele carried by the virus, although total viral accumulation was not affected, as observed for the nbpen3 mutants (Supplementary Fig. 12). Also in this case, the knock-out of SlPEN3 enabled TbCSV(Y35) phloem escape (Fig. 5b). Importantly, the expansion of tissue tropism facilitated by SlPEN3 knock-out had a negative impact on insect acquisition, as demonstrated by the lower viral accumulation in the midgut of whiteflies fed on these plants (Fig. 5c, d, Supplementary Fig. 11). We next performed the transmission assays with wild-type and SlPEN3 knock-out plants, consistent with the decreased accumulation of the virus detected in the insect vector fed on slpen3 tomato plants, transmission efficiency from slpen3 tomato plants was diminished for all isolates (Fig. 5e–g). These results suggest that PEN3-mediated phloem restriction promotes viral spread by favoring the acquisition of the virus by the insect vector.
a Viral symptom induced by TbCSV(Y35), TbCSV(Y41) and TbCSV(Y35-2C4) in wild-type (WT) and slpen3 knock-out tomato plants at 30 days post-inoculation (dpi). Two independent slpen3 knock-out tomato plants and yellowing symptoms in the leaf lamina of WT and slpen3 tomato plants are shown. Photographs were taken at 30 dpi. b Immunofluorescence detection of TbCSV in sections of WT and slpen3 knock-out tomato plants infected by TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4). The distribution of TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4) was visualized in a cross-section of leaf vein tissues using an antibody against the TbCSV coat protein (CP) (red). Autofluorescence of highly lignified tissues is shown in blue. PC, parenchyma; P, phloem; X, xylem. White arrowheads indicate TbCSV in parenchyma cells. Scale bar = 50 μm. c Immunofluorescence detection of TbCSV in midguts of Zhejiang II whiteflies after feeding on WT or slpen3 knock-out tomato plants infected by TbCSV(Y35), TbCSV(Y41) or TbCSV(Y35-2C4). TbCSV was detected using a monoclonal antibody against the TbCSV coat protein (CP), followed by a commercial 549-conjugated secondary antibody (red), and nuclei were stained with DAPI (blue). At least 10 midguts were examined for each treatment. Scale bar = 100 μm. Representative images are shown. d Immunoblot analysis of virus accumulation in whiteflies after feeding on TbCSV(Y35)-, TbCSV(Y41)-, or TbCSV(Y35-2C4)-infected WT and slpen3 knock-out tomato plants. Total protein of 150 whiteflies was extracted for immunoblot per treatment. Histone H3 was used as a loading control. e–g Immunoblot analysis of TbCSV(Y35) (e), TbCSV(Y41) (f), and TbCSV(Y35-2C4) (g) accumulation in WT tomato plants inoculated with 1, 5, 15, or 45 viruliferous Zhejiang II whiteflies acquiring virus from TbCSV(Y35)-, TbCSV(Y41)-, or TbCSV(Y35-2C4)-infected WT or slpen3 knock-out tomato plants. TbCSV was detected using a monoclonal antibody against the TbCSV coat protein (CP). Actin was used as a loading control. Tomato plants were given an inoculation access period of 48 h with 1, 5, 15, or 45 viruliferous whiteflies using clip cages in three independent biological replicates. Total protein was extracted from WT tomato plants at 14 days after the inoculation access period. Experiments in (b–d) were repeated at least three times with similar results.
PEN3 is a general determinant of viral phloem restriction
To test whether PEN3 mediates phloem restriction in other geminiviruses, we inoculated WT and nbpen3 N. benthamiana plants with tomato yellow leaf curl virus (TYLCV). Nbpen3 plants displayed slightly enhanced symptoms (Fig. 6a), but no difference in viral CP accumulation was detected when compared to the WT plants (Fig. 6b). In spite of similar viral accumulation, TYLCV was found outside of the phloem in NbPEN3 knock-out N. benthamiana plants (Fig. 6c), indicating that phloem restriction of geminiviruses is likely controlled by PEN3.
a Symptoms of TYLCV-infected WT and nbpen3 N. benthamiana plants at 14 days post-inoculation (dpi). Upper panel shows the phenotype of leaves in WT and nbpen3 N. benthamiana plants. Lower panel shows the downward leaf curling induced by TYLCV. b Immunoblot analysis of TYLCV accumulation in WT and nbpen3 N. benthamiana plants at 14 dpi. Actin was used as loading control. c Immunofluorescence detection of TYLCV in cross-sections of leaf veins from infected WT and nbpen3 N. benthamiana plants. The distribution of TYLCV was visualized using antibodies against TYLCV CP (red). Autofluorescence of highly lignified tissues is shown in blue. PC, parenchyma; P, phloem; X, xylem. White arrowheads indicate TYLCV in parenchyma cells. Scale bar = 50 μm. d Symptoms induced by PLRV in WT and nbpen3 N. benthamiana plants at 14 days post-inoculation (dpi). Lower panel shows the downward leaf curling induced by PLRV. e Immunoblot analysis of PLRV accumulation in WT and nbpen3 N. benthamiana plants at 14 dpi. Actin was used as loading control. f Immunofluorescence detection of PLRV in cross-sections of leaf veins from infected WT and nbpen3 N. benthamiana plants. The distribution of PLRV was visualized using antibodies against PLRV CP (red). Autofluorescence of highly lignified tissues is shown in blue. PC, parenchyma; P, phloem; X, xylem. White arrowheads indicate PLRV in parenchyma cells. Scale bar = 50 μm. Experiments in (b, c, e, f) were repeated three times with similar results.
To further assess if PEN3 mediates phloem restriction to viruses in general, potato leaf roll virus (PLRV), an aphid-transmitted phloem-limited RNA virus, was used to inoculate WT and nbpen3 mutant plants. Nbpen3 plants were more susceptible to PLRV infection than the WT, displaying severe downward leaf curling and higher viral accumulation at 14 dpi (Fig. 6d, e). Strikingly, immunolocalization of the viral CP in cross-sections of leaf veins demonstrates that PLRV exits the phloem and reaches parenchyma cells in nbpen3 mutant plants only (Fig. 6f). These results allow the conclusion that PEN3 also determines phloem limitation of an RNA virus.
Discussion
Many plant viruses, such as poleroviruses, criniviruses and most begomoviruses, are phloem-restricted in natural infections, which correlates with their transmission by phloem-feeding insect vectors19,20,21. However, how viruses are restricted to the phloem and what the biological significance of this tissue tropism is, are long-standing questions in plant virology. The different tissue tropisms between TbCSV isolates provide a precious opportunity to investigate these questions. TbCSV (Y35) is an example of such tissue limitation, and its invasion can be recognized by the plant immune system, leading to strong callose deposition surrounding the infected vasculature (Fig. 3f, Supplementary Fig. 5). This response presumably leads to a physical constriction of plasmodesmata associated with decreased trafficking through these channels, and therefore the constraint of lateral viral movement. TbCSV(Y41), however, is capable of compromising callose accumulation during infection through the action of an additional chloroplast-localized C4 effector, cC4. cC4 targets chloroplasts and interferes with SA signaling, promoting PEN3 degradation and compromising callose-mediated phloem restriction. Surprisingly, the difference in tissue tropism between TbCSV(Y35) and TbCSV(Y41) is not reflected in increased virus accumulation (Fig. 1c). This phenomenon might be explained by the fact that geminivirus promoters can display phloem-specific activity, which confers higher replication efficiency in the phloem22; therefore, escaping this tissue would not result in a significant increase in viral load. Determining TbCSV replication efficiency in phloem companion vs parenchyma cells will shed light on this observation. Importantly, the finding that phloem restriction of the unrelated, aphid-transmitted RNA virus PLRV also depends on PEN3 suggests a general role of this protein in meditating phloem limitation of viruses.
Chloroplast immunity is emerging as a cornerstone of plant defence23,24,25,26,27,28,29,30,31. In this study, we show that the chloroplast-localized C4 encoded by the Y41 allele of TbCSV interferes with SA signaling, likely through its direct interaction with NbCAS1 and/or by disrupting the redox balance in chloroplasts. Impaired SA signaling destabilizes PEN3, which in turn compromises virus-induced callose deposition and ultimately virus phloem restriction. This model is supported by several pieces of evidence: (1) The expression of the SA-responsive gene PR1 is repressed in both 35S::TbCSV(Y41) cC4 and 35S::TbCSV(Y35-2C4) cC4 transgenic N. benthamiana plants, indicating a diminished SA response, but not in 35S:TbCSV(Y35) (Fig. 4f); (2) PEN3 stability is regulated in a SA-dependent manner (Fig. 4j, Supplementary Fig. 9); (3) TbCSV(Y35) escapes the phloem and reaches parenchyma cells in the nbpen3 mutant, which correlates with the lack of observable callose deposition (Fig. 3e, f, Supplementary Fig. 5). These results suggest a scenario in which TbCSV(Y41) compromises the PEN3-mediated viral phloem restriction through the action of a chloroplast-localized C4.
C4 is the most divergent canonical geminiviral protein, and the C4 proteins encoded by different geminiviruses have different functions and properties. Even though C4 from TYCLV could shuttle from the plasma membrane to chloroplasts during infection to dampen SA signaling8, its efficiency might not suffice to ensure phloem escape. Further efforts to conduct a comparative study between the strengths of these viral proteins as SA suppressors and their ability to mediate phloem escape will shed light on this apparent conundrum.
The question remains of what the tell-tale of the viral infection is that triggers activation of defence and, ultimately, PEN3-mediated callose deposition. Viruses are devoid of canonical pathogen-associated molecular patterns (PAMPs), inducers of plant immunity; however, in geminiviruses, the replication-associated protein (Rep) or its activity has been shown to activate defense responses8,32, and the genomic DNA has also been proposed to be perceived33,34,35. Viral RNA is believed to act as PAMP36,37. Nevertheless, the nature and mechanism of viral perception in the phloem await clarification.
Phloem restriction has been generally regarded as the result of the deployment of plant defence responses that the invading virus is unable to suppress, although the exact underlying molecular mechanisms have so far remained elusive. Our results show that phloem restriction of the geminivirus TbCSV and other viruses likely relies on PEN3-dependent virus-induced callose deposition, which probably results from recognition of the viral invasion. The chloroplast-localized viral C4 protein encoded by the TbCSV Y41 C4 allele, however, can suppress SA signaling, leading to a decreased accumulation of PEN3, impaired callose deposition, and phloem escape of the virus. This enlarged tissue tropism, nevertheless, has a toll in insect transmission, since a lower proportion of the viral population is accessible to the phloem-feeding insect vector (Fig. 7). The decreased transmission efficiency imposed by phloem escape may underpin the observation that TbCSV strains carrying the Y41 allele are not dominant in nature, despite exhibiting an expansion in host range, and, more generally, that numerous viruses transmitted by phloem-feeding insects remain phloem-restricted through evolution.
Virus-induced PEN3-mediated callose deposition is critical for phloem limitation of geminiviruses, which favors acquisition and subsequent transmission by the phloem-feeding insect vector, the whitefly B. tabaci. In TbCSV(Y41), the virus-encoded chloroplast-localized C4 protein interferes with SA signaling and inhibits PEN3 accumulation, compromising phloem restriction; this expansion in tissue tropism, however, has a toll on the acquisition by the insect vector, since a lower proportion of the viral population is accessible, ultimately decreasing the virus’ epidemic rate.
Methods
Plasmids construction
The NbPEN3 coding sequence (CDS) was obtained from cDNA derived from N. benthamiana plants and cloned into the pGD-GFP vector for the observation of NbPEN3 subcellular localization; TbCSV(Y35) C4, TbCSV(Y41) C4, and TbCSV(Y35-2C4) C4 were cloned into the pCambia-Flag vector individually for the generation of transgenic N. benthamiana plants. For yeast two-hybrid assays (Y2H) and bimolecular fluorescence complementation assays (BiFC), CDS of NbCAS1, TbCSV(Y35) C4, TbCSV(Y41) C4, TbCSV(Y35-2C4) C4 were recombined into pGADT7, pGBKT7, p2YN, and p2YC, respectively, generating AD-NbCAS1, BD-TbCSV(Y41) C4, TbCSV(Y35) C4–nYFP, TbCSV(Y41) C4–nYFP, TbCSV(Y35-2C4) C4–nYFP, and NbCAS1-cYFP. All cDNAs were PCR-amplified using the KOD-Plus-Neo High-Fidelity DNA polymerase (TOYOBO). The resulting PCR fragments were cloned individually into the destination vectors by using the One-Step Cloning Kit (Vazyme). All primers used for plasmid construction are listed in Supplementary Table 3.
Plant materials, growth conditions, and generation of transgenic plants
Wild-type (WT) and transgenic NahG or histone 2B (H2B)-RFP N. benthamiana plants were cultivated in a greenhouse at 26 °C under a 16 h light/8 h dark photoperiod, and seedlings at an approximately 5-leaf stage were used for the experiments. The tomato (Solanum lycopersicum) cultivar ‘Ailsa Craig’ was used in this study. Seeds were sown in trays filled with a blend of peat and vermiculite in a 2:1 ratio (v:v). Upon full development of the second true leaf, seedlings were transplanted into pots containing the same peat and vermiculite mixture. Seedlings were supplied with 1/2 Hoagland’s nutrient solution every two days and were cultivated in a greenhouse equipped with LED lighting, maintaining daytime temperatures around 23–28 °C and nighttime temperatures around 18–23 °C, and seedlings at an approximately 3-leaf stage were used for the experiments. CRISPR/Cas9-based knockout of NbPEN3 and SlPEN3 in N. benthamiana and tomato plants was generated by transformation with the binary vector pBGK01 (Biogle, Changzhou, China) in fusion with a single-guide RNA designed to target the ORFs. The synthesized sgRNA oligo was annealed and then ligated to pBGK01 with One-Step Cloning Kit (Vazyme, Nanjing, China). T2 homozygous transgenic plants were used for subsequent experiments. For overexpression of TbCSV(Y35) C4, TbCSV(Y41) C4 and TbCSV(Y35-2C4) C4, the plasmids carrying 35S::TbCSV(Y35) C4-Flag, 35S::TbCSV(Y41) C4-Flag and 35S::TbCSV(Y35-2C4) C4-Flag were used for transformation in N. benthamiana plants. Immunoblot was conducted to verify the gene expression.
Immunocytochemistry for protein localization in sections of plant tissues
Immunocytochemichemistry for protein localization of plant tissues was performed essentially as described previously38. Leaf tissues (5 mm × 10 mm) of N. benthamiana or S. lycopersicum plants infected by different TbCSV isolates were fixed in 4% paraformaldehyde at 4 °C overnight. The fixative solution was removed, and the materials were washed three times for 5-10 min each with PBST at room temperature. Then the samples were dehydrated sequentially in 25%, 50%, 75%, and 96% ethanol for 60 min each at room temperature. After dehydration, the samples were incubated with wax/ethanol (1:2), wax/ethanol (1:1), wax/ethanol (2:1) and pure wax for 60 min each at 37 °C. Materials were then transferred into a Petri dish, placed evenly at the bottom of the dish, arranged in a final position, and pure wax was poured onto them. The material was placed at 4 °C for 20 min, to let the wax solidify. The material was then sectioned with a thickness of 6–8 μm by using a razor blade and transferred onto a microscope slide for 2 h. The slides were incubated sequentially in 99%, 90%, and 50% ethanol and PBS for 15 min each at room temperature for dewaxing and rehydration. Then the sections were blocked with 2% BSA for 2 h at room temperature, followed by incubation with anti-TbCSV CP monoclonal antibody (1:400) or Anti-PEN3 polyclonal antibody (PHYTOAB, Catalog: PHY0177A) (1:200) in PBST containing 1% BSA overnight at 4 °C and four washes with PBST for 10 min each at room temperature. Next, sections were incubated with 1% BSA solution with goat anti-mouse (1:500) 549-conjugated secondary antibody (Earthox, Catalog: E032410-01) or goat anti-rabbit (1:500) 488-conjugated secondary antibody (Earthox, Catalog: E032220-01) for 2 h at room temperature. After secondary antibody incubation, the material was washed with PBST three times for 10 min each, then covered with a cover glass and observed using a Zeiss LSM780 confocal microscope. The immunofluorescence intensity of TbCSV signal in the phloem of leaf vein sections (fluorescence intensity/selected area) in different treatments was measured using ImageJ software; at least 25 areas from three independent biological replicates were selected in each condition.
Immunofluorescence assay
Immunofluorescence detection of viruses in Zhejiang II whitefly midguts and primary salivary glands was performed essentially as described previously39,40. Intact midguts of whiteflies and intact primary salivary glands of whiteflies at required developmental stages were dissected after a 48 h-access acquisition period (AAP) on virus-infected tomato plants, and the specimens were incubated and fixed in 4% paraformaldehyde for 1 h at room temperature. Samples were then washed in PBST containing 0.5% Triton X-100 three times for 30 min each. The specimens were blocked with 1% BSA for 2 h at room temperature, followed by incubation with anti-PLRV CP polyclonal antibody (1:100) or anti-TbCSV CP monoclonal antibody (1:400) in PBST containing 1% BSA overnight at 4 °C and washed with PBST containing 0.5% Triton X-100 three times for 30 min each. Samples were incubated with 1% BSA solution with goat anti-mouse (1:500) 549-conjugated secondary antibody (Earthox, Catalog: E032410-01) for 2 h at room temperature. All specimens were washed with PBST three times for 30 min each, and DNA was stained with 100 mM 4, 6-diamidno-2-phenylindole (DAPI) for 30 min at room temperature. For each experiment, 10 midguts or primary salivary glands were examined by using a Zeiss LSM780 confocal microscope.
Electron microscopy
Plant tissues (1 mm × 4 mm) were excised from leaves of WT, 35S::TbCSV(Y35) C4-Flag, 35S::TbCSV(Y41) C4-Flag or TbCSV(Y35-2C4) C4-Flag transgenic N. benthamiana plants. The sampled tissues were fixed in 2.5% glutaraldehyde and 1% osmium tetroxide (both in 100 mM phosphate buffer, pH 7.0). The fixed tissues were embedded in Epon 812 resin as instructed by the manufacturer (SPI-EM, Division of Structure Probe, Inc., West Chester, USA). Ultrathin sections (70 nm) were cut from embedded tissues using the Ultracut E Ultramicrotome (Reichart-Jung, Vienna, Austria) and mounted on formvar-coated grids. Thin sections were then stained with uranyl acetate for 10 min followed by lead citrate for 10 min. The stained sections were examined under an electron microscope (HT7820; Hitachi, Japan).
Detection of virus transmission efficiency
Three-leaf stage tomato plants were inoculated with the infectious clones of TbCSV(Y35), TbCSV(Y41), or TbCSV(Y35-2C4). The whitefly adults were placed on TbCSV(Y35)-, TbCSV(Y41)-, or TbCSV(Y35-2C4)-infected tomato plants to feed for a period of 48 h to acquire the virus. These adults were then collected individually, and a given number of adults (1, 5, or 15) were placed on new individual test tomato plants at 21 days post-inoculation (dpi) to feed for another period of 72 h for virus transmission by using clip cages in three independent biological replicates. Non-viruliferous whiteflies were used as a mock treatment. Immediately after the 72 h-virus transmission period, the whitefly adults were removed, and the test plants were sprayed with imidacloprid (20 mg/L) to kill all the whitefly eggs produced during the 72 h inoculation period. The test plants were further cultured for 21 days, and the virus infection status of the test plants was then determined by Immunoblotting.
Callose detection and quantification
Callose staining was performed essentially as described previously41. Callose of wild-type (WT) and nbpen3-ko transgenic N. benthamiana plants was stained with 0.05 mg/mL aniline blue for 20 min at room temperature. The sections were washed with water three times, then the callose was detected at 405 nm wavelength by using a Zeiss LSM780 laser scanning microscope. The relative intensity of the fluorescent signal in the phloem (fluorescence intensity/selected area) in different treatments was measured by using ImageJ software; at least 10 areas from three independent biological replicates were selected in each treatment. Callose quantification was performed by measuring the callose deposition through dot-blot and antigen coating plate (ACP) ELIZA kit (Bioroyee, Catalog: RE5372). For dot-blot assays, 0.2 g of leaf tissues of WT and transgenic N. benthamiana plants were homogenized in 200 μl of isolation buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCI, 5 mM DTT, 1 mM MgCl2). The homogenate was filtered through a 47 μm nylon mesh, the flow-through was centrifuged at 8000 g for 10 min, and the supernatant was spotted onto nitrocellulose membranes and allowed to dry. The membrane harboring proteins was incubated with 3% BSA in TBST buffer for 1 h at room temperature. The membrane was then incubated with TBST containing HRP-conjugated anti-callose antibody or anti-actin antibody at 4 °C overnight. Next, the membrane was washed with TBST three times, and callose bound on the membrane was detected with the ECL western blotting kit (4 A Biotech). For ACP ELIZA, 0.2 g of leaf tissues of WT and transgenic N. benthamiana plants were homogenized 200 μl of isolation buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCI, 5 mM DTT, 1 mM MgCl2). The homogenate was filtered through a 47 μm nylon mesh, the flow-through was centrifuged at 8000 g for 10 min, and the resulting supernatant was incubated in a 96-well microtiter plate at 4 °C overnight, respectively. The plate coated with callose-contained supernatant was washed with PBST buffer three times. Then, PBST containing HRP-conjugated anti-callose antibody was added to the wells coated with callose at 37 °C for 2 h. After washing four times with PBST, an ACP ELIZA was performed to detect the callose bound on the plate.
Quantitative RT-PCR
Total RNA was extracted from N. benthamiana leaf tissues using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. First-strand synthesis was performed using ReverTra Ace qPCR RT Master Mix with gDNA Remover (TOYOBO). Real-time PCR was conducted using SYBR Green I Master (Vazyme) according to the manufacturer’s instructions. Primers were used for the amplification of the targets, and the efficiencies of all primers were verified by normal RT-PCR, gel electrophoresis, and melting curve analysis. Primer sequences are listed in Supplementary Table 3. Expression of the Actin gene was used as an internal control. The value of each bar indicates the average value of three independent measurements.
Semi-in vivo protein degradation assay
Semi-in vivo protein degradation assays were performed essentially as described previously42,43. For semi-in vivo protein degradation analysis, a GFP-NbPEN3 sample was harvested from the leaves infiltrated with Agrobacterium cultures expressing GFP-NbPEN3 in wild-type or NahG transgenic N. benthamiana plants treated with ddH2O or 50 μM SA for 12 h. The samples were extracted gently with native extraction buffer (50 mM Tris-HCl, pH 8.0, 0.5 M sucrose, 1 mM MgCl2, 10 mM EDTA, 5 mM dithionthreitol). For analysis of the GFP-NbPEN3 degradation, the plant extract harboring GFP-NbPEN3 was mixed with cycloheximide (CHX; 100 mM), then ATP was added (to a final concentration of 20 mM). These samples were agitated in an Eppendorf Thermomixer at 25 °C, and equal volumes of sthe ample were removed from the tube at different time points.
Bimolecular fluorescence complementation (BiFC) assay
BiFC assays were performed on 5-week-old N. benthamiana leaves infiltrated with the combination of Agrobacterium tumefaciens EHA105 harboring p2YN-TbCSV(Y35) C4, producing TbCSV(Y35) C4–nYFP, or p2YN-TbCSV(Y41) C4 producing TbCSV(Y41) C4–nYFP, or p2YN-TbCSV(Y35-2C4) C4 producing TbCSV(Y35-2C4) C4–nYFP and A. tumefaciens EHA105 carrying p2YC-NbCAS1 producing NbCAS1-cYFP. The combination of NbCAS-cYFP/TbCSV(Y35) C4-nYFP serves as a negative control. Emission of the YFP interaction signal was detected using Zeiss LSM780 laser scanning microscope (Carl-Zeiss) at 48 h post-inoculation (hpi).
Statistical analysis
All statistical analyses were done with the GraphPad Prism 8.0 software. A two-sided, unpaired Student’s t-test was performed when appropriate. Details about the statistical approach used can be found in the figure legends. Data are represented as mean ± SD as indicated.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
PEN3 sequences can be found in the Sol Genomics Network under accession numbers Niben101Scf04046g00002.1 (NbPEN3) and Solyc03g120980.3.1 (SlPEN3). The authors declare that the data supporting the findings of this study are available within the paper and its supplementary files. A reporting summary for this paper is available as a supplementary file. Source data are provided with this paper.
References
Taliansky, M., Mayo, M. A. & Barker, H. Potato leafroll virus: a classic pathogen shows some new tricks. Mol. Plant Pathol. 4, 81–89 (2003).
Moreno, P., Ambros, S., Albiach-Marti, M. R., Guerri, J. & Pena, L. Citrus tristeza virus: a pathogen that changed the course of the citrus industry. Mol. Plant Pathol. 9, 251–268 (2008).
Bendix, C. & Lewis, J. D. The enemy within: phloem-limited pathogens. Mol. Plant Pathol. 19, 238–254 (2018).
Reagan, C. B. & Burch-Smith, T. M. Viruses reveal the secrets of plasmodesmal cell biology. Mol. Plant Microbe Interact. 33, 26–39 (2020).
Mei, Y., Yang, X., Huang, C., Zhang, X. & Zhou, X. Tomato leaf curl Yunnan virus-encoded C4 induces cell division through enhancing stability of Cyclin D1;1 via impairing NbSKη-mediated phosphorylation in Nicotiana benthamiana. PLoS Pathog 14, e1006789 (2018).
Medina-Puche, L., Orilio, A. F., Zerbini, F. M. & Lozano-Duran, R. Small but mighty: functional landscape of the versatile geminivirus-encoded C4 protein. PLoS Pathog 17, e1009915 (2021).
Kumar, R. & Dasgupta, I. Geminiviral C4/AC4 proteins: an emerging component of the viral arsenal against plant defence. Virology 579, 156–168 (2023).
Medina-Puche, L. et al. A defense pathway linking plasma membrane and chloroplasts and co-opted by pathogens. Cell 182, 1109–1124 (2020).
Rosas-Diaz, T. et al. A virus-targeted plant receptor-like kinase promotes cell-to-cell spread of RNAi. Proc. Natl. Acad. Sci. USA 115, 1388–1393 (2018).
Mei, Y. et al. Two viral proteins translated from one open reading frame target different layers of plant defence. Plant. Commun. 5, 100788 (2024).
Stein, M. et al. Arabidopsis PEN3/PDR8, an ATP-binding cassette transporter, contributes to nonhost resistance to inappropriate pathogens that enter by direct penetration. Plant Cell 18, 731–746 (2006).
Matern, A. et al. A substrate of the ABC transporter PEN3 stimulates bacterial flagellin (flg22)-induced callose deposition in Arabidopsis thaliana. J. Biol. Chem. 294, 6857–6870 (2019).
Zavaliev, R., Ueki, S., Epel, B. L. & Citovsky, V. Biology of callose (β-1,3-glucan) turnover at plasmodesmata. Protoplasma 248, 117–130 (2011).
Burch-Smith, T. M., Brunkard, J. O., Choi, Y. G. & Zambryski, P. C. Organelle-nucleus cross-talk regulates plant intercellular communication via plasmadeamata. Proc. Natl. Acad. Sci. USA 108, E1451–E1460 (2011).
Burch-Smith, T. M. & Zambryski, P. C. Plasmodeamata paradigm shift: regulation from without versus within. Annu. Rev. Plant Biol. 63, 239–260 (2012).
De Storme, N. & Geelen, D. Callose homeostasis at plasmodesmata: molecular regulators and developmental relevance. Front. Plant Sci. 5, 138 (2014).
Amsbury, S., Kirk, P. & Benitez-Alfonso, Y. Emerging models on the regulation of intercellular transport by plasmodesmata-associated callose. J. Exp. Bot 69, 105–115 (2017).
Li, T., Huang, Y., Xu, Z. S., Wang, F. & Xiong, A. S. Salicylic acid-induced differential resistance to the Tomato yellow leaf curl virus among resistant and susceptible tomato cultivars. BMC Plant. Biol. 19, 173 (2019).
Barker, H. Invasion of non-phloem tissue in Nicotiana clevelandii by potato leafroll virus is enhanced in plants also infected with potato Y potyvirus. J. Gen. Virol. 68, 1223–1227 (1987).
Wisler, G. C., Li, R. H., Liu, H. Y., Lowry, D. S. & Duffus, J. E. Tomato chlorosis virus: a new whitefly-transmitted, phloem-limited, bipartite closterovirus of tomato. Phytopathology 88, 402–409 (1998).
Ramos, P. L. et al. Identification of the minimal sequence required for vascular-specific activity of tomato mottle Taino virus replication-associated protein promoter in transgenic plants. Virus Res. 102, 125–132 (2004).
Guan, C. & Zhou, X. Phloem-specific promoter from a satellite associated with a DNA virus. Virus Res. 115, 150–157 (2006).
Caplan, J. L. et al. Chloroplast stromules function during innate immunity. Dev. Cell 34, 45–57 (2015).
de Torres Zabala, M. et al. Chloroplasts play a central role in plant defence and are targeted by pathogen effectors. Nat. Plants 1, 15074 (2015).
Lu, Y. & Yao, J. Chloroplasts at the crossroad of photosynthesis, pathogen infection and plant defense. Int. J. Mol. Sci. 19, 3900 (2018).
Sowden, R. G., Watson, S. J. & Jarvis, P. The role of chloroplasts in plant pathology. Essays Biochem 62, 21–39 (2018).
Fernandez, J. C. & Burch-Smith, T. M. Chloroplasts as mediators of plant biotic interactions over short and long distances. Curr. Opin. Plant Biol. 50, 148–155 (2019).
Kachroo, P., Burch-Smith, T. M. & Grant, M. An emerging role for chloroplasts in disease and defense. Annu. Rev. Phytopathol. 59, 423–445 (2021).
Littlejohn, G. R., Breen, S., Smirnoff, N. & Grant, M. Chloroplast immunity illuminated. New Phytol. 229, 3088–3107 (2021).
Irieda, H. Emerging roles of motile epidermal chloroplasts in plant immunity. Int. J. Mol. Sci. 23, 4043 (2022).
Liu, J. et al. Chloroplast immunity: a cornerstone of plant defense. Mol. Plant 17, 686–688 (2024).
Ding, X., Jimenez-Gongora, T., Krenz, B. & Lozano-Duran, R. Chloroplast clustering around the nucleus is a general response to pathogen perception in Nicotiana benthamiana. Mol. Plant Pathol. 20, 1298–1306 (2019).
Santos, A. A., Carvalho, C. M., Florentino, L. H., Ramos, H. J. O. & Fontes, E. P. B. Conserved threonine residues within the A-loop of the receptor NIK differentially regulate the kinase function required for antiviral signaling. PLoS ONE 4, e5781 (2009).
Teixeira, R. M. et al. Virus perception at the cell surface: revisiting the roles of receptor-like kinases as viral pattern recognition receptors. Mol. Plant Pathol. 20, 1196–1202 (2019).
Teixeira, R. M., Ferreira, M. A., Raimundo, G. A. S. & Fontes, E. P. B. Geminivirus triggers and suppressors of plant antiviral immunity. Microorganisms 9, 775 (2021).
Niehl, A. & Heinlein, M. Perception of double-stranded RNA in plant antiviral immunity. Mol. Plant Pathol. 20, 1203–1210 (2019).
Huang, C. et al. dsRNA-induced immunity targets plasmodesmata and is suppressed by viral movement proteins. Plant Cell 35, 3845–3869 (2023).
Paciorek, T., Sauer, M., Balla, J., Wisniewska, J. & Friml, J. Immunocytochemical technique for protein localization in sections of plant tissues. Nat. Protoc. 1, 104–107 (2006).
Wei, J. et al. Vector development and vitellogenin determine the transovarial transmission of begomoviruses. Proc. Natl. Acad. Sci. USA 114, 6746–6751 (2017).
Pan, L. et al. Differential efficiency of a begomovirus to cross the midgut of different species of whiteflies results in variation of virus transmission by the vectors. Sci. China Life Sci 10, 1254–1265 (2018).
Gui, J., Liu, C., Shen, J. & Li, L. Grain setting defect 1, encoding a remorin protein, affects the grain setting in rice through regulating plasmodesmatal conductance. Plant Physiol. 166, 1463–1478 (2014).
Shen, Q. et al. Tobacco RING E3 Ligase NtRFP1 mediates ubiquitination and proteasomal degradation of a geminivirus-encoded βC1. Mol. Plant 9, 911–925 (2016).
Mei, Y., Ma, Z., Wang, Y. & Zhou, X. Geminivirus C4 antagonizes the HIR1-mediated hypersensitive response by inhibiting the HIR1 self-interaction and promoting degradation of the protein. New Phytol. 225, 1311–1326 (2020).
Acknowledgments
This work was supported by the National Natural Science Foundation of China (W2411024) and the National Key Research and Development Program of China (2022YFD1400804) to Xueping Zhou, and the Basic Research Center of Innovation Program of Chinese Academy of Agricultural Sciences (CAAS-BRC-CB-2025-02) to Fangfang Li. The authors give thanks to Dr. Li Xie in the Analysis Center of Agrobiology and Environment Sciences, Zhejiang University, for the help with transmission electron microscopy.
Author information
Authors and Affiliations
Contributions
X.Z. and Y.M. conceived the project and designed the experiments. Y.M., Y.W., and F.L. performed experiments and analyzed data; X.Z. supervised work; Y.M. and R.L. prepared the figures; Y.M., R.L., and X.Z. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Robert Turgeon and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mei, Y., Wang, Y., Li, F. et al. Defence-mediated phloem restriction of a plant virus facilitates insect transmission. Nat Commun 17, 152 (2026). https://doi.org/10.1038/s41467-025-67827-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67827-w