Abstract
Synthetic biology aims to engineer or re-engineer living systems. To achieve increasingly complex functionalities, it is beneficial to use higher-level building blocks. In this study, we focus on oscillators as such building blocks, propose oscillator-based circuit designs and model the interactions of intracellularly coupled oscillators. We classify these oscillators on the basis of coupling strength: independent, weakly or strongly, and deeply coupled. We predict a wide range of dynamic behaviors to arise in these systems, such as the beat phenomenon, amplitude and frequency modulation, period doubling, higher-period oscillations, chaos, resonance, and synchronization, with the aim of guiding future experimental work in bacterial synthetic biology. Finally, we outline potential applications, including oscillator-based computing that integrates processing and memory functions, offering multistate and nonlinear processing capabilities.
Similar content being viewed by others
Introduction
In synthetic biology (SynBio), the goal is to engineer living organisms to perform specific, useful tasks by designing and constructing biological systems1,2,3,4. SynBio is inherently multidisciplinary, integrating principles from fields such as molecular biology, genetics, electrical engineering, and computer science. Similar to electrical engineering, where simple components like resistors, transistors, and capacitors are assembled to create functional circuits, SynBio uses biological building blocks, such as genes, proteins, and regulatory elements, to construct biological “circuits" that control cellular behavior in predictable ways. As electrical circuits become more complex, higher-level components such as microchips are often employed to streamline the design process. Similarly, in SynBio, achieving more advanced and intricate functionalities could benefit from using higher-level biological building blocks, such as multiple oscillators coupled to one another, which can produce complex, diverse, and useful behaviors.
Insights from physics, chemistry, biology, and engineering show that coupled oscillators exhibit remarkable dynamic behaviors. These phenomena have revolutionized industries over the past centuries, with applications in telecommunications5, micro-electromechanical systems (MEMS)6, healthcare (e.g., cardiac pacemakers)7, and quantum computing8, among others. Beyond practical applications, the study of coupled oscillators enhances our understanding of natural processes such as synchronization in circadian rhythms9, neuronal activity10, and even animal and human behaviors11,12,13.
In biological systems, natural genetic oscillators are coupled at both intracellular and intercellular levels. For instance, cell division and the circadian clock are intracellularly coupled, enabling them to achieve robust synchronization14,15. Similarly, metabolic oscillations have been shown to synchronize with the early and late stages of the cell cycle, as demonstrated in a study in budding yeast16. Intercellular coupling also plays a pivotal role in processes like presomitic mesoderm segmentation during vertebrate embryonic development. Precise spatiotemporal oscillations are essential for the proper formation of vertebrae17,18,19. Another example of coupling can be observed in plant growth, whereby the growth rate and shoot curvature oscillate in anti-phase of one another, effectively dampening curvature and enabling straight growth20,21. These examples highlight the critical role of coupled oscillations in maintaining coordination and functionality across biological systems.
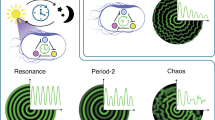
In this study, we explore the potential phenomena that may arise when multiple synthetic oscillatory circuits are introduced within a single living cell, e.g., intracellular coupling. Our goal is to contribute to fundamental advancements in SynBio, just as coupled oscillators have done in many other fields. Figure 1 gives an overview of the oscillators we use in this study as building blocks for constructing coupled oscillatory systems. The first oscillator, the Goodwin oscillator22,23,24,25 consists of a single self-repressing node and exhibits fast, low-amplitude oscillations. As the time period increases, the amplitude likewise grows until protein levels approach saturation. The dual-feedback oscillator, introduced by Stricker et al.25, is the second fastest oscillator in this lineup. The repressilator, designed by Elowitz et al. in 200026, was the first synthetic oscillator in synthetic biology. Since then, numerous variants and alternative circuit topologies have been developed, spanning a wide range of time periods25,27,28,29,30,31,32. While these designs predominantly rely on transcription factor-mediated regulation, the CRISPRlator, our slowest oscillator, relies on CRISPR interference-based repression30. These oscillators span a broad range of amplitudes and time periods, enabling the study of their rich dynamic properties and paving the way for innovative applications in the future.
The time period increases from left to right. While representative time periods are shown, these can vary significantly depending on the parameters used in simulations, biological implementation, or experimental conditions. The blue nodes in the CRISPRlator symbolize that this circuit relies on CRISPR-based repression rather than transcription factor interactions.
One possibility to study coupled oscillators is to use non-autonomous systems and investigate the interactions between an oscillator inside the cell and an external oscillatory input32,33,34. For example, we previously examined how oscillatory light inputs interact with an optogenetic repressilator, referred to as the “optoscillator”32. On the other hand, in autonomous systems, each oscillator operates based on its own internal dynamics, and interactions occur between oscillators, without external time-dependent forcing. Several studies have explored synthetic intercellularly coupled oscillators mediated by small-molecule-based cell-cell communication27,28,35,36,37,38. These efforts focused on populations in which every cell harbors an identical oscillator. Such systems have proven to be a powerful tool for constructing complex systems and investigating intricate dynamic behaviors27,28,35,36,37.
However, intercellular coupling introduces inherent limitations: signaling molecules are required to traverse cell membranes, diffuse through the extracellular space and re-enter neighboring cells in order to influence other oscillators38. This process can significantly attenuate and delay signal transmission. Moreover, studying interactions between distinct oscillators in such setups requires the use of different cell types and the establishment of stable co-cultures, posing additional experimental challenges39. In contrast, intracellularly coupled oscillators enable alternative coupling strategies that bypass these delays and signal degradation, offering a more direct and efficient framework for exploring oscillator interactions.
Notably, intracellular coupling between oscillators can be achieved through direct regulatory interactions, in which components of one circuit modulate the dynamics of another. In addition to these explicit connections, it is well established that competition for shared, limited cellular resources, such as RNA polymerases, ribosomes, proteases, dCas9 and nutrients, can influence the behavior of otherwise orthogonal genetic circuits40,41,42,43,44,45,46,47. While such competitions are typically considered undesirable, they can be deliberately harnessed to couple synthetic circuits. For instance, Prindle and colleagues demonstrated that protease competition can be harnessed to couple oscillators43. A similar mechanism can be envisioned for CRISPRi-based systems, where competition for dCas9 could serve as a coupling strategy. Metabolic networks offer an additional layer of interaction between genetic circuits. Oscillations in metabolic activity can naturally synchronize distinct circuits via fluctuations in shared metabolites48. For example, changes in substrate or product availability can modulate oscillator period and amplitude49. A notable case is the metabolator, a synthetic metabolic oscillator driven by dynamic flux between interconverting metabolite pools50. Thus, intracellular oscillator coupling presents a range of intriguing possibilities, which we explore in this study.
While coupled oscillators have been extensively studied in physics and engineering, their systematic design within single living cells remains largely unexplored. Previous efforts in synthetic biology have focused mainly on externally driven or intercellularly coupled systems25,27,28,32,33,34,36,38. Here, we introduce a framework for autonomous intracellular coupling between synthetic oscillators, enabling complex dynamical behaviors, such as synchronization, resonance, and chaos, to emerge solely from internal regulatory interactions. By treating oscillators as modular building blocks, this work establishes a design paradigm that bridges dynamical systems theory with the practical challenges of scalable genetic circuit engineering.
Figure 2 provides an overview of the classification of coupled oscillators relevant to this study. Coupled oscillators can be divided into two main categories: non-autonomous and autonomous systems. Autonomous systems are self-governing and time-invariant, described by differential equations in which the independent variable (time) does not explicitly appear:
where x denotes the state vector of the system and f(x) defines the intrinsic dynamics. In contrast, non-autonomous systems depend explicitly on time, either through external driving forces or time-varying parameters:
where the additional argument t captures the influence of time-dependent inputs or modulations. This distinction is fundamental in classifying oscillatory dynamics, particularly in biological contexts where intrinsic rhythms (autonomous) can be perturbed or entrained by external cues such as light–dark cycles or chemical stimuli (non-autonomous).
In non-autonomous systems, oscillators within cells are influenced by an external force (dashed orange and turquoise lines). In autonomous systems, however, the oscillators interact directly with each other within the cells (solid orange and turquoise lines). Variations in the type and strength of these interactions give rise to distinct oscillator categories. Independent oscillators occur when no interactions are present (light beige background), while deeply coupled oscillators (dark blue background) arise under infinitely strong interactions. Between these extremes, oscillators can be classified as weakly or strongly coupled (gradient of blue intensity background). Further distinctions can be made regarding the relative frequency of the oscillators (length of orange/turquoise lines) or the direction of coupling (orange or turquoise arrows).
This study focuses on autonomous coupled oscillators, where multiple oscillators interact without external influences. However, as we will show, autonomous unidirectionally coupled systems share notable similarities with their non-autonomous counterparts. We further classify autonomous oscillators on the basis of the strength of their interactions. When the interaction is negligible, we refer to them as independent oscillators, whereas with infinitely strong interaction, we observe deeply coupled oscillators. Between these extremes are weakly and strongly coupled oscillators. The direction of coupling interaction can also be taken into account when categorizing coupling oscillators. When one oscillator influences the other without being affected in return (or the effect is negligible), this interaction is referred to as unidirectional or master-slave coupling. In contrast, when both oscillators mutually influence each other, it is referred to as bidirectional or peer-to-peer coupling51,52,53. Another critical aspect influencing the behavior of coupled oscillators is their relative frequencies.
Since we are utilizing higher-level building blocks (i.e., oscillators) to create these coupled oscillators, establishing simulations at a higher level of abstraction is straightforward. For this purpose, we used GRN_modeler54, our previously developed tool to simulate, and analyze gene regulatory networks (GRNs). It features a user-friendly graphical user interface (GUI) that allows us to construct and model the circuits node by node, significantly accelerating the design and modeling of GRNs.
For our simulations, we employed three distinct models. The first model, developed by Elowitz et al.26, represents the repressilator. In other cases, such as when describing the Goodwin oscillator22, a more detailed model is required to account for the role of enzymatic protein degradation. For these situations, we employed the model created by Tomazou et al.55. For the CRISPRlator, we used a model described by Holló et al.54. We provided a comprehensive summary of these models in our previous work54, and for the reader’s convenience, we summarize them again in the Supplementary Note 1. Throughout this study, we will refer to these models as the Elowitz, Tomazou and CRISPR models, respectively.
One promising application of coupled oscillators is their use in oscillator-based computing (OBC), enabling information processing in response to inputs. Most current biocomputing implementations in synthetic biology rely on the logic-gate metaphor from electronics, producing binary outputs56,57. However, this framework is inherently limiting for complex computations and may not represent the most suitable paradigm for information processing in living cells58,59,60. In OBCs, the information is encoded in the frequency or phase of the oscillators, rather than in the signal’s strength, mimicking the natural neuronal oscillations of the brain61. In the brain, oscillations couple with one another in phase and/or frequency to coordinate complex calculations/actions such as visual processing, memory, cognition, and voluntary movement generation10,62,63,64. OBCs aim to reproduce this behavior, with individual processing units ("neurons”) performing basic calculations in response to multiple signals65. The interactions between oscillators then allow complex, nonlinear calculation functions as the output from the collection of neurons (see Csaba & Porod (2020)65 for a comprehensive review about OBCs). While researchers in other fields have attempted to emulate the functioning of the brain’s action in biological systems66,67,68, to the best of our knowledge, OBCs have not yet been designed in SynBio.
Here, we computationally investigate how the topology and coupling strength of intracellular oscillators influence their behavior, offering insights into the dynamics of intracellular systems. This theoretical framework lays a foundation for experimental efforts to engineer oscillatory circuits. After exploring different scenarios that can arise with intracellularly coupled oscillators, we study their implementation in OBCs and their potential application in SynBio. By presenting this foundational concept, we aim to inspire further research and innovation in the field.
Results
Independent oscillators
We began by examining the simplest case of autonomous coupled oscillators, two systems operating without direct interactions. This scenario involves two independent oscillators controlling the state of a common node through logic gates. Although the assumption of complete independence is highly idealized, as in real cells, oscillators often share molecular components or are interconnected through metabolic activities, it remains valuable to study these systems as a limiting case. The basic principles underlying independent coupled oscillators can be illustrated analytically using simple harmonic functions (Supplementary Note 2).
Using the Tomazou model55, we simulated two independently coupled repressilators with similar frequencies (Fig. 3I). We kept the parameters of the second oscillator constant while gradually varying the concentration of the PROT1 protease, which degrades the proteins of the first repressilator. Although the protease concentration was held constant in the simulations, its activity varied dynamically due to oscillations in the concentrations of its substrates (Supplementary Fig. 6). The adjustment of the protease concentration allowed us to alter the degradation rate of the transcription factors in the first repressilator, thereby modifying its oscillation period, while the period of the second remains constant. Both oscillators control the output node through a NOR gate, resulting in an output signal only when both oscillators are in-phase, i.e., when N1 and N4 are deactivated simultaneously. Due to the slight frequency difference between the oscillators, the phase difference is changing continuously between them, leading to periodic activation and deactivation of the output node. This interference pattern, known from acoustics as the beat phenomenon, can be calculated with the following formula69:
where f4 and f1 are the frequencies of the independent repressilators, and f7 is the beat frequency at the output node (N7). According to this formula, when the frequency difference between the two oscillators is small, the resulting beat frequency is also low, leading to a prominent peak at a long beat period. As shown in Fig. 3I, d and determined by Eq. (3), the time period of the output node, calculated from trajectories with varying protease concentrations, corresponds closely to the period of the beat phenomenon. This ideal case allows us to easily calculate the necessary protease concentration for a given output period.
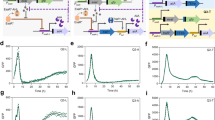
Row I: The Beat Phenomenon. a Circuit topology of two repressilators coupled via a NOR gate at node N7. b Protein concentration trajectories of nodes N1 (red) and N4 (blue), respectively producing proteins P1 and P4. Two segments of the trajectories are presented: one up to 5000 min, illustrating distinct peak separation, and another from 6800 min onward, where the peaks overlap. c Oscillations in the output node N7. Peaks are observed when the oscillations in (b) are in-phase (second half of the graph). Conversely, when the oscillations in (b) are out-of-phase (first half), N7 remains repressed. Gray rectangles indicate the time ranges corresponding to panel (b). d Comparison of theoretical (blue) and simulated (red) beat time period as a function of N1's oscillation period. Row II: Amplitude Modulation. a Circuit topology of a dual-feedback oscillator coupled to a CRISPRlator via N6. b Protein concentration dynamics of nodes N2 (red) and N3 (blue). c Output node N6 oscillations. d Fourier transform (FFT) of N6 (blue), with expected modulation frequency peaks (black dashed lines). f2 and f3 denote the frequencies of N2 (dual-feedback oscillator) and N3 (CRISPRlator), respectively. The detailed descriptions of the models are available in Supplementary Data 1 and in Supplementary Data 2, respectively. An extended figure caption with additional details is available in Supplementary Note 2.2. The figure can be reproduced using the independent3.m and independent_oscillators.m scripts.
For practical applications, this method provides a way to achieve significantly different frequencies without redesigning the oscillatory circuits. Typically, circuit adjustments to alter its frequency (e.g., by modifying promoter strength or protease concentration) only tune the frequency within a limited range. Significant changes often require a full circuit redesign. Leveraging the beat phenomenon, however, allows substantial adjustments to the output node’s period without altering the existing circuit designs, making it a valuable technique for tuning oscillatory systems.
Next, we examined a scenario involving independently coupled autonomous oscillators with significantly different time periods. We coupled a CRISPRlator30,54,70 and a dual-feedback oscillator25 using again a NOR gate (Fig. 3IIa). In our model, the time period of the CRISPRlator is 604.5 minutes, while the time period of the dual-feedback oscillator with the modified Elowitz model is 25.2 minutes (Fig. 3II,b) (for details of the model see the extended figure caption in Supplementary Note 2.2). We observed a phenomenon known as amplitude modulation (AM), in which the amplitude of a high-frequency carrier signal is modulated by a lower-frequency signal. Specifically, the carrier signal from the fast dual-feedback oscillator (P2, red) is modulated by the slow CRISPRlator (P3, blue), resulting in an AM signal at the output node (P6, yellow) (Fig. 3IIb, c). In this simulation, the repression strength from N3 to N6 is low (0.01%), ensuring a continuous carrier signal. For comparison, the scenario with complete repression is shown in Supplementary Fig. 5. Figure 3IId illustrates that under this ideal conditions with completely independent oscillators, the frequency of the output node is indeed determined by the modulation of the dual-feedback oscillator’s frequency, occurring at integer multiples of the CRISPRlator’s frequency. Furthermore, we confirm the independence of both frequency and phase between the uncoupled oscillators: The time-resolved frequency spectrum of the output signal is characterized using wavelet transformation (Supplementary Fig. 8) and the Hilbert transform (Supplementary Fig. 9), providing a detailed view of the modulation dynamics. Finally, we show that AM can also be achieved by coupling a repressilator and a CRISPRlator (Supplementary Fig. 7).
Deeply coupled oscillators
In contrast to independent oscillators, where the coupling strength is zero, in deeply coupled oscillators, the coupling is infinitely strong. Although these circuits could be viewed as a single unified system rather than as coupled oscillators, treating them as coupled systems provides a more intuitive framework for understanding their behavior.
To study the behavior of deeply coupled oscillators, we connected two repressilators through a common edge (a repression) (Fig. 4a), creating a scenario of deep coupling in which the oscillators cannot oscillate at different frequencies or phases. The “first repressilator" comprises the nodes N1, N2, and N3, while the “second repressilator" consists of N1, N4, and N3. We gradually adjusted the promoter strength of the fourth node (Fig. 4b, c) – a node not connected to the common edge. As the promoter strength decreases, the dynamic behavior and time period of the circuit increasingly resemble those of a single repressilator. Initially, due to the coupling, the system exhibits asymmetry: nodes display varying amplitudes, and the overall period is longer than that of the standard repressilator (Fig. 4b, first row). However, as the promoter strength of N4 decreases and the influence of the second oscillator weakens, the system becomes symmetrical (Fig. 4b, third row), with its period converging to that of the three-node repressilator. We would like to emphasize that, in most cases, we highlight these phenomena in coupled oscillators using a single model and a specific example (e.g., in this case, by modifying the promoter strength of N4). However, the system’s oscillation frequency can be fine-tuned through multiple approaches, all of which produce qualitatively similar results to the one presented. For instance, frequency adjustments can be achieved by altering the promoter strength (Fig. 4), modifying the protease concentration (Supplementary Fig. 11), or varying repression strength with a small chemical (Supplementary Fig. 12). Therefore, while this study outlines potential scenarios, experimental implementation will require selecting the most appropriate approach and modeling the corresponding conditions.
a Topology of two deeply coupled three-node repressilators. b Protein trajectories of corresponding nodes, with varying promoter strengths for the N4 node: 30, 15 and 0 molecules/minute, respectively. c Time period of the coupled system as a function of the promoter strength of the fourth node. Scenarios from panel (b) are marked with red dots. d Topology of deeply coupled repressilators with three and five nodes. e Time period of the system as a function of the number of nodes in the coupled oscillators. f Time period of a single repressilator and two deeply coupled repressilators sharing an edge with both rings having an equal number of nodes ("double repressilator"), plotted as a function of the number of nodes in a single ring. The simulations ran for 104 min, with the first 103 min regarded as transient time. The detailed descriptions of the model for the circuit in (a) is available in Supplementary Data 16. The figure can be reproduced using the deeply_coupled_tirv_Elowitz.m and double_repressilator_T.m scripts.
Next, we investigated the behavior of two deeply coupled oscillators when their frequencies vary significantly. We previously demonstrated that the time period of the repressilator family increases linearly with the number of nodes54. In Figures 4d, e, we couple two repressilators with odd numbers of nodes. In the matrix of Fig. 4e, the two axes represent the number of nodes in the first and second circuits. A node number of 1 indicates the absence of the circuit. Therefore, the first column and the last row show the time period of the repressilator without coupling to a second oscillator, while the point (1, 1) represents a system without oscillators. The matrix is symmetric because the system itself is symmetric: the first and second oscillators are interchangeable. As expected, in the uncoupled case, the time period increases linearly with the number of nodes (Supplementary Fig. 10). Interestingly, a similar linear relationship is observed when the two oscillators share an edge and have the same number of nodes, although the overall period is slightly longer due to the shared edge (Fig. 4f). However, when a large circuit (such as a five- or fifteen-node system) is coupled via a shared edge to a smaller (and faster) oscillator, the overall system accelerates to the frequency of the faster oscillator. This occurs because along the faster route, the first common node is repressed earlier, forcing the frequencies and phases of the coupled oscillators to synchronize. As a potential application, this coupling strategy could be used to accelerate larger or slower oscillators by linking them to smaller and faster oscillators. The repressilator can only oscillate with an odd number of nodes; therefore, both of the coupled circuits in this case were designed with an odd number of nodes. If either circuit contains an even number of nodes, the system becomes bistable, and oscillations cease. However, a similar approach can be applied to circuits like the reptolator or actolator, which resemble the repressilator yet are capable of oscillating with an even number of nodes54.
The deeply coupled Goodwin oscillator
Oscillatory circuits are often constructed from delayed negative feedback and self-activation motifs22,71. The Goodwin oscillator represents one of the simplest examples, relying on a single self-repressing node that generates high-frequency, low-amplitude oscillations. We found that this characteristic, “Goodwin-type” behavior, is not restricted to a single node: it can also emerge in multi-node systems when the nodes become synchronized. For instance, the classical toggle switch72,73, typically known for bistable switching, can exhibit fast, low-amplitude oscillations when both nodes oscillate in phase. In this regime, the circuit behaves effectively as a single self-repressing unit (Supplementary Fig. 13).
Interestingly, even the repressilator, a canonical model of sequential (out-of-phase) oscillations, can transition to a Goodwin-type, in-phase oscillatory state under specific coupling conditions. The coexistence and interaction of these two oscillatory modes leads to rich dynamics, including bistability, period-doubling, and even chaotic behavior at strong coupling (Supplementary Fig. 13). These findings highlight that deeply coupled circuits can display unexpectedly diverse dynamical regimes, extending the concept of the Goodwin oscillator far beyond its traditional single-node form. A detailed analysis and supporting simulations are provided in Supplementary Note 3.2, and Supplementary Figs. 13, 14, 15, 16.
Weakly and strongly coupled oscillators
So far, we discussed the independent case, where oscillators do not interact, and the case of deep coupling, where the system behaves like a single oscillator. Next, we explored the cases that fall between these two extremes and that we further categorized into weakly and strongly coupled oscillators. Typically, most studies about coupled oscillators analyze these intermediate categories. Although there is no strict boundary or universally agreed-upon definition to distinguish weak from strong coupling, a common criterion for strong coupling is when the coupling strength (or energy exchange rate) exceeds the system’s loss rates74,75.
Independent oscillators exhibit no synchronization, while deeply coupled oscillators are fully synchronized. In contrast, weakly and strongly coupled oscillators display more nuanced synchronization dynamics, such as partial or limited synchronization. We first studied the effect of synchronization using a model of two coupled repressilators, based on the Tomazou model55 (Fig. 5a–d). Each oscillator is degraded by its own protease, but they also share a common protease, PROT3, allowing us to fine-tune the coupling strength by adjusting the concentration of this shared enzyme (Fig. 5a). In Fig. 5b, both oscillators have identical parameter sets, resulting in the same oscillation frequency. However, a small difference in the initial conditions leads to a phase offset between the oscillations. As we increase the coupling strength, represented by the number of protease molecules PROT3 (from 0 to 16), we observe the transition from independent oscillators (at zero coupling) where the phase difference remains constant, to synchronized oscillators where the phase difference diminishes. As expected, the stronger the coupling strength, the faster the oscillators synchronize.
a–d Synchronization in coupled repressilators. a Topology of two oscillators coupled through PROT3. b Phase difference and protein oscillations for varying PROT3 concentrations. c Protein trajectories in N1 for increasing PROT3 levels (0, 40, and 100 molecules). d Average period (mean ± SD) of nodes 1 and 4 plotted as a function of PROT3. Period estimates were obtained from approximately 80 oscillatory peaks during the simulation. e–h Higher period oscillations and chaos. e Topology of two unidirectionally coupled repressilators. f Bifurcation diagram for the system in (e), chaotic regions are depicted in red. g Topology of a Goodwin oscillator coupled to a repressilator. h Bifurcation diagram for the system in (g), chaotic regions are depicted in red. i–k Resonance. i Topology of two unidirectionally coupled repressilators. j Time period changes in the second oscillator with promoter strength variation in node N6. k Oscillation amplitude at N1 with changes in the promoter strength of N6. Simulations for resonance (i–k) used the Elowitz model; others (a–h) used the Tomazou model. The details of the simulations can be found in Supplementary Data 8, Supplementary Data 9, 10 and in Supplementary Data 11, respectively. An extended figure caption with additional details is available in Supplementary Note 4.1. The figure can be reproduced using the coupled_synchronisation.m, repressilator_repressilator.m, repressilator_goodwin.m and resonance.m scripts.
We then varied the protease concentrations in the coupled repressilators (PROT1 and PROT2), leading to differences in their natural frequencies (Fig. 5c, d). With zero PROT3 concentration, representing the independent case, the oscillators exhibit regular oscillations (Fig. 5c, top row), and their distinct time periods are evident at the start of the curves (Fig. 5d). As the coupling strength increases, we observe higher-period oscillations with alternating large and small peaks (Fig. 5c, middle row). This pattern arises from constructive and destructive interference, similar to the beat phenomenon seen in independent oscillators. However, as the coupling strength further increases, the oscillators’ time periods gradually converge, making the dynamics more complex compared to the uncoupled case. The error bars in Fig. 5d indicate that the time period is not constant during the oscillation. When a critical coupling strength is reached, a bifurcation occurs, causing the individual frequencies of the oscillators to merge, resulting in a single, common time period. Beyond this critical point, the system achieves regular oscillations with constant amplitude and a stable time period (Fig. 5c, bottom row). High phase-locking values (Supplementary Fig. 17I,c) indicate sustained synchronization between the oscillators, while the frequency perturbations observed in the continuous wavelet transform reflect dynamic adjustments in frequency during the synchronization process (Supplementary Fig. 17I). The instantaneous phase difference, calculated using the Hilbert transform (Supplementary Note 2), varies during the synchronization process and stabilizes once the oscillators achieve full synchronization (Supplementary Fig. 17II).
For synchronization to occur, bidirectional coupling is typically required, whereas unidirectional coupling behaves similarly to non-autonomous systems. Previously, we experimentally demonstrated that non-autonomous systems can exhibit complex dynamics such as resonance, period-\({{\mathcal{N}}}\) oscillations or chaos32. A unidirectionally coupled system operates similarly, but with the periodic driving force coming from within the system rather than from an external source, suggesting that similar dynamic phenomena can be observed in these autonomous systems as well.
We already discussed amplitude modulation (AM) with independent coupled oscillators, where the oscillation frequency remains constant, and only the amplitude changes (Fig. 3II). By unidirectionally coupling a slow oscillator to a faster oscillator, both FM (frequency modulation) and AM can be observed (Supplementary Fig. 18). As altering the promoter strength affects both amplitude and frequency, this generally results in imperfect FM and AM. However, using more complex topologies and building on previous studies55,76, it is possible to generate pure AM and FM signals with unidirectionally coupled oscillators (Supplementary Fig. 19).
We next explored the capability of unidirectionally coupled oscillators to produce chaotic oscillations, using two examples. In the first examples, a repressilator is driven by another repressilator (Fig. 5e, f). and the frequency of the driving system is varied by adjusting the protease concentration, PROT2. For the coupled repressilators, we observe wide regions of higher-period oscillations and chaos (Fig. 5e, f, and Supplementary Fig. 20).
In the second example, a Goodwin oscillator drives the repressilator (Fig. 5g). Also here, the frequency of the Goodwin oscillator can be varied with the protease concentration. The observed chaotic region is notably wide (Fig. 5h and Supplementary Fig. 20). This coupled oscillator presents a promising candidate for a synthetic chaotic autonomous oscillator. The large Lyapunov exponents facilitate the observation of chaotic dynamics experimentally, and the wide chaotic region reduces the need for precise experimental fine-tuning. Moreover, the system is relatively simple, consisting of only four nodes, which minimizes the potential burden on cells.
Apart from chaotic oscillations, resonance is another phenomenon of interest in coupled oscillators. In our previous work32, we demonstrated both experimentally and computationally that the non-autonomous optoscillator can exhibit resonance (alongside higher-period oscillations and chaos). Our simulations were based on the Elowitz model26. Here, we predict that resonance can also occur in autonomous coupled oscillators by using two unidirectionally coupled repressilators (Fig. 5i). In our simulations, we varied the frequency of the driving oscillator by changing the promoter strength (Fig. 5j) while keeping the frequency of the other oscillator constant. By adjusting the promoter strength of the N6 node in the master oscillator, we found that the amplitude of protein oscillations achieves its maximum when this parameter is set around 30 molecules per minute (Fig. 5k), which coincides with the point at which the two oscillators have equal frequencies.
An application: oscillator-based computing
Finally, we explored an application of coupled oscillators in oscillator-based computing (OBC): the creation of information processing units with memory functions. To showcase this application, we studied two coupled CRISPRlators (Fig. 6). The coupling is achieved through a shared dCas pool rather than a protease (as was the case in Fig.5). If the dCas concentration is high, there is no competition between the oscillators for the shared species, allowing each oscillator to function independently (Supplementary Fig. 23). In contrast, at low dCas concentrations, CRISPRlators cannot oscillate at different frequencies because a synchronized peak alignment would excessively deplete the dCas pool. This constraint enforces synchronization with a fixed phase difference between oscillators. It is important to note that these simulations only account for the effects related to the dCas pool. In a real system, higher synchronization could arise from factors such as proteases or cellular metabolic activity, which would further strengthen phase locking.
a Schematic of intracellularly coupled oscillators, where small chemical inputs drive binary outputs based on phase differences. b Correlation matrix showing in-phase and anti-phase signals, with an example AND gate operation. c, e, g, i Circuit topologies (top) and protein oscillations (bottom), demonstrating phase locking with low dCas concentration (200 molecules). d, f, h, j Correlation matrices computed from protein time series. k Schematic representation of an alternative (explicit) coupling strategy between oscillators. l The effect of the repression strength between the oscillators (β) to the phase-locking. In panels (l) and (n), the simulations were run for 105 min, with the first 15% treated as transient. Parameters were a1,N2 = 1.1, a1,N6 = 1.2, and in (n) [dCas] = 200 molecules. m Schematic illustration of implicit coupling between oscillators mediated by the time-dependent burden. n Phase-locking value versus dCas concentration for various half-saturation constants K; lower K indicates stronger burden-mediated coupling. The details of the simulations are available in Supplementary Data 15. An extended figure caption with additional details is available in Supplementary Note 5.1. The figure can be reproduced using the computer_CRISPRlator.m, computer_CRISPRlator_sctoch_burden.m and computer_CRISPRlator_sctoch_repress.m scripts.
During the initial oscillations, synchronization occurs, representing the phase in which the system performs its computation. The output of a computation with oscillators is not a single low or high signal intensity; instead, the information is encoded in the frequency or phase of the output. Here, we focus on the phase difference between a node in the first oscillator (N1) and nodes in the second oscillator (N4, N5, N6), while our input is the promoter strength of N3 and N4 (Fig. 6). Once synchronization occurs, the phase-locking effect enables the system to function as a memory unit. Thus, the system initially operates as a processor, performing computations during the start of oscillations, and transitions into acting as a memory unit thereafter.
We first demonstrate that the system can operate in a traditional binary mode based on Boolean algebra, functioning as an AND gate (Fig. 6a, b). The behavior depends on whether N1 is synchronized with N4. We computed the correlation matrix from the oscillatory patterns of protein concentrations at each node (Fig. 6d, f, h, j and Supplementary Fig. 22). However, a big proportion of these matrices are redundant: by identifying the closest node in phase with N1, we can qualitatively infer the remaining correlations. Nodes of the same CRISPRlator always exhibit different phases and non-positive correlations. In other words, the first column of the matrix contains all the qualitative information needed to distinguish the three possible outcomes of the system. In panels (d) and (f) of Fig. 6, N1 is synchronized with N5 (and consequently N2 with N6, and N3 with N4). In Figure 6h, N1 aligns with N6, and in panel (j), it locks with N4. A close phase relationship results in a positive value in the correlation matrix (close to one), which is interpreted as a logical one (true), while a non-positive (close to zero or negative) correlation corresponds to a logical zero (false). In this example, the inputs could be small chemical molecules or light that modulate the promoter strengths of N1 and N4. When both N1 and N4 are activated, corresponding to high concentrations of the input chemicals, they synchronize, resulting in a positive correlation. In all other cases, the correlation is non-positive, enabling the system to function as an AND gate, where the output is true only if both inputs are true.
We can also illustrate the behavior of these coupled oscillators in phase space on the PN1-PN2 plane (Supplementary Fig. 24). This representation resembles the output of a flow cytometry measurement. Interestingly, since we are plotting two periodic signals as functions of each other, we observe Lissajous curves. When the correlation is perfect (i.e., one) and the phase difference is zero, the result is a straight line (Supplementary Fig. 24h). Conversely, with a phase difference of π, the line has a slope of -1. For a phase difference around π/2 (Supplementary Fig. 24a), we observe a closed curve with a large enclosed area. As the phase difference approaches 0 or π, this curve transitions into the straight lines described earlier (Supplementary Fig. 24b–d). If synchronization does not occur, the output of such a flow cytometry experiment would spread across and fill the phase space (Supplementary Fig. 24f, g).
Interestingly, this OBC system based on two coupled CRISPRlators, is not only capable of operating in a binary mode, it can actually produce three distinct outputs dependent on node synchronization, thus behaving as a ternary system (Supplementary Fig. 21a). Specifically, the phase of node N1 aligns with the phase of node N4, N5, or N6 from the other oscillator, producing three possible outcomes. This behavior is evident in the correlation matrices of the oscillatory signals (Figure 6f, h, j), which display three distinct patterns. However, we can observe this ternary behavior by analyzing the protein oscillations of just two nodes and calculating their corresponding correlation values (Supplementary Fig. 21a). The correlation can take three distinct values: − 1, 0, and 1; effectively forming a ternary system. The advantage of a ternary system over a binary one lies in its ability to store more information per unit. Each ternary digit represents three states, thus reducing the number of digits required to encode the same amount of information compared to binary systems. By coupling oscillators with more nodes, the number of possible outputs can be significantly increased. Furthermore, the correlation values can vary according to the promoter strength, resulting in a nonlinear output (Supplementary Fig. 21b), further enriching its dynamic behavior. The output of the system, which represents the correlation between N1 and N2, follows a sigmoid function around a promoter strength of 1.2 min−1. This type of function is commonly used in neural networks. If these nonlinear functions could be computed with OBC in a single step, rather than through the layered architecture of current neural networks, we could unlock a substantial performance boost in tasks like image processing, mimicking the efficiency of biological neurons (Supplementary Fig. 21c).
A critical consideration in the biological implementation of OBC within cells is the presence of noise, an intrinsic feature of all biological systems. To assess its influence, we conducted stochastic simulations (Supplementary Note 5.1) while systematically varying the noise strength, σ∞ and analyzed the resulting phase-locking value (PLV). The PLV quantitatively captures how consistently the phase difference between two oscillators remains stable over time, with values approaching 1 indicating strong phase locking and values near 0 signifying desynchronization (Supplementary Note 2 and Supplementary Equation 17). Our results show that increasing noise levels progressively reduce the phase-locking value (PLV) (Fig. 6l, blue curve), indicating that sufficiently strong noise can disrupt oscillator coupling. Furthermore, while phase locking typically results in the phase difference fluctuating around a stable value, high noise amplitudes lead to pronounced temporal variations in phase difference (Supplementary Fig. 25).
Successfully implementing such circuits experimentally will require coordinated theoretical and experimental efforts to account for these effects. One strategy to mitigate the impact of noise is to increase the coupling strength between oscillators. For example, additional explicit coupling interactions can be introduced to enhance synchronization between oscillators (Fig. 6k), such as repression from node N1 to N4. Ideally, the strength of these interactions could be modulated using chemical inducers, allowing fine-tuned control over oscillator coupling. In our stochastic simulations (Fig. 6k,l), we implemented such repression, with the parameter β scaling the baseline repression strength between nodes. As β increases, phase locking becomes progressively more robust, persisting even under substantial noise. Remarkably, even a modest repression level (10% of the standard repression strength) is sufficient to synchronize the oscillators despite intense noise. This behavior is further illustrated by the instantaneous phase difference between oscillators (Supplementary Fig. 26): under high noise and weak coupling, no phase locking is observed; with moderate coupling, phase locking occurs intermittently; and with sufficiently strong coupling, the phase difference fluctuates narrowly around a constant value, indicating stable and sustained synchronization. In addition to increasing the coupling strength, the overall coherence of the system can also be improved by reducing intrinsic noise. When intracellular oscillators are coupled through diffusible signaling molecules, intercellular synchronization can emerge. In this synchronized state, the population of cells effectively acts as a single oscillator with a larger effective volume, thereby reducing stochastic fluctuations and further enhancing the relative influence of intracellular coupling43.
Moreover, there may be couplings beyond those we design explicitly. We illustrate this effect through the burden and resource limitation imposed by the oscillators on the host cells (Fig. 6m). Because gene expression and mRNA production oscillate in time, the availability of intracellular resources, such as RNA polymerases, ribosomes and overall cellular activity, also vary, potentially introducing an implicit coupling between oscillators40,41,42. We examined this effect in a simple simulation (Fig. 6n), assuming that increased heterologous protein concentration leads to higher burden and consequently decreases overall cellular activity, including mRNA production40,77,78. This was modeled as a simple repression using a Hill function, where the half-saturation constant K was varied. The results show that as the dCas concentration increases, the PLV decreases and the oscillators lose synchrony. However, reducing K strengthens coupling through the burden, and the oscillators remain phase-locked even at high dCas concentrations (blue curve). It is important to emphasize that this burden is only a demonstration of implicit, indirect coupling: in reality, other mechanisms could have similar effects and modify the effective coupling strength between oscillators.
In summary, with OBC, we present a framework for biocomputing in synthetic biology that holds promise for establishing a paradigm in the field, though its practical realization will require addressing technical challenges.
Discussion
Here, we computationally explored intracellularly coupled autonomous oscillators. In other words, we computationally introduced different oscillators within a single cell and aimed to map the range of behaviors that can arise in such systems. To establish a clearer framework, we categorized the oscillators into three distinct groups based on the interaction strength between them: independent, weakly/strongly coupled, and deeply coupled oscillators, with the coupling strength progressing from zero to infinity.
This work establishes a biologically grounded framework for autonomous intracellular coupling between oscillators, creating a basis for synthetic circuits that access more complex yet experimentally attainable dynamical behaviors and expand the reachable architectural design space. In the following discussion, we summarize the phenomena investigated and suggest potential future applications, many of which are inspired by existing implementations of oscillators in other fields.
Using the example of independent oscillators, we exemplify the beat phenomenon (Fig. 3I), which has numerous practical applications. In acoustics, the beat phenomenon is commonly applied to tuning musical instruments, enabling precise adjustments by comparing interference patterns79. Similarly, in optical frequency metrology, it plays a crucial role in comparing and calibrating laser frequencies, facilitating advancements in precision measurement technologies80. In a biological context, this phenomenon could be applied to measure frequency differences between oscillators, providing a basis for highly sensitive biosensors. For instance, one oscillator could be sensitive to a specific chemical compound or physical parameter, while the other remains unaffected, allowing us to measure frequency shifts through the beat phenomenon. The output frequency of the system is highly sensitive to the frequency difference between the oscillators (Fig. 3Id), enhancing the precision and sensitivity of the method (Fig. 7a).
a Beat phenomenon and its use in sensors and the design of new oscillators. b AM and FM in cell-cell communication or as sensing mechanisms. c Synchronization and oscillator-based computing, illustrated using neural network examples and a possibility to fine-tune oscillators. d Resonance effects for selectively activating damped oscillators, enabling new cellular functionalities. e Higher-period oscillations and chaos-generated patterns. Detailed descriptions of these applications are provided in the text.
We proposed several circuits for generating AM and FM signals. In our first example, we coupled independent oscillators with different frequencies to a common output (Fig. 3II). In the second example, we used coupled oscillators (Supplementary Fig. 19) to generate pure AM and FM signals55,76. Recent findings show that living organisms leverage this technique for cell-to-cell communication, benefiting from its enhanced signal-to-noise ratio (Fig. 7b) and enabling them to accurately respond to gradual changes in external stimuli81,82,83,84. For example, bacteria use both AM and FM to broadcast signals that determine whether the population will commit to biofilm formation or cell motility85. In SynBio, the use of AM/FM has recently been demonstrated in mammalian cells by changing the diffusivity and reactivity of oscillatory components of a genetic circuit (based on bacterial MinDE), that can ultimately modulate the frequency and amplitude of oscillations within cells86.
AM is commonly used in signal transmission, such as in telecommunications5, various types of advanced sensors, including Quartz Crystal Microbalance (QCM) sensors87, atomic force microscopy88, electroencephalogram (EEG)89, dielectric characterization of solids and microfluidics90, and biomedical sensors in cochlear implants and in cardiac pacemakers7. AM enhances these sensors’ ability to detect subtle changes, increasing sensitivity and accuracy. Inspired by this, a potential application of AM in SynBio would be its use in whole cell biosensors91, for applications as diverse as detecting polychlorinated biphenyls in wastewater, metallic trace elements in contaminated soil, or hallmark molecules of infectious diseases in the medical field.
Synchronization was first observed and described by Huygens in the 17th century, using the example of pendulum clocks that naturally synchronized when placed on the same surface92. In our simulations, we reproduced that oscillators with the same frequency can shift their phase relative to each other, while oscillators with different frequencies can adjust their time periods until they synchronize, depending on the strength of the coupling. In a cell, oscillators are never completely independent, which opens up the possibility of fine-tuning natural oscillators without directly modifying them. By introducing a synthetic oscillator with a frequency similar to that of a natural oscillator, we could potentially adjust the frequency and phase of a natural oscillator (cell cycle, circadian clock, metabolic oscillation) through synchronization with a synthetic oscillator.
In our numerical example of OBCs, we demonstrated how coupled CRISPRlators can be used to construct logic gates and memory units. However, the potential of OBCs extends far beyond these applications. By moving beyond binary thresholds and Boolean logic and instead utilizing partial synchronization between oscillators, it becomes possible to generate nonlinear functions as output. This approach could prove valuable in tasks such as neural network computations, image analysis (Supplementary Fig. 21c)65,93,94 or in oscillator-based optimization95. Moreover, coupling more than just two oscillators significantly increases the number of possible states, potentially enabling solutions to computationally hard problems in a manner analogous to entangled systems in quantum information theory8,65. However, scaling up the number of coupled oscillators will require intercellular coupling and the development of stable co-cultures38,96,97,98, which represents an important next step in this field. In our example (Figure 6), we observed that information can be stored in the phase of oscillators through phase locking. Alternatively, the information could also be encoded in the frequency of the oscillator65. Nature already employs this principle of frequency encoding. A striking example is found in snakes, whose pit organs can detect temperature differences as small as a few millikelvins99. In this system, sensory information is carried by a stochastic oscillator: minute temperature shifts, particularly when the system operates near a bifurcation point, can induce large changes in firing frequency. This phenomenon illustrates how stochasticity does not merely disrupt OBC but can instead be exploited to achieve exceptional sensitivity. By tuning the system close to a critical transition, weak inputs can be amplified into robust frequency shifts.
Previous studies demonstrated that in a non-autonomous system, when the frequency of external forcing matches the natural frequency of the synthetic oscillator, high-amplitude oscillations, i.e., resonance, can be observed32,33. In this work, we show that a unidirectional coupling between autonomous oscillators produces an analogous behavior, resulting in resonance as well (Fig. 5). Resonance is widely utilized in electrical engineering, such as in radio receivers or band-pass filters, where only a specific frequency range is targeted. In certain methods, resonance is advantageous, such as in Nuclear Magnetic Resonance (NMR) or Electron Paramagnetic Resonance (EPR)100. However, in structures such as tall buildings, resonance-induced vibrations are mitigated by using tuned mass dampers101. In SynBio, oscillatory circuits often exhibit damped oscillatory behavior rather than sustained oscillations. As a potential application, introducing another robust oscillator into the system could drive the damped oscillator at its resonant frequency, forcing it to oscillate. By incorporating multiple damped oscillators with varying natural frequencies and properties, we could harness their resonance and band-pass filter characteristics to selectively activate specific oscillators by tuning the frequency of a master oscillator. This approach ensures that only the activated damped oscillator oscillates, reducing the cellular burden while enabling the desired functionality associated with the selected oscillator (Fig. 7d). Furthermore, the system can be easily fine-tuned by adjusting the master oscillator’s frequency with a single inducer. In another application, resonance could be harnessed to filter and amplify AM and FM signals in cell-to-cell communication.
In SynBio, oscillators have been used to generate spatial patterns27,28,29,32. Bacterial colonies harboring a synthetic oscillator and growing on solid surfaces produce ring patterns. These patterns are formed because only the cells at the colony edge are growing and oscillating, while cells in the inner region are arrested in different phases of the oscillations. We previously demonstrated that using higher-period oscillations, we can produce rings with alternating intensities, and that these patterns are disrupted in chaotic regions32. However, in that case, an externally modulated light signal was required to achieve the effect. With the suggested autonomous chaotic coupled oscillators, it could be possible to generate these patterns without the need for an external signal (Fig. 7e).
Surprisingly, we found that the most well-known repressilator and toggle switch circuits should be able to exhibit Goodwin-type oscillations, caused by enzymatic protein degradation (Supplementary Fig. 13 and Supplementary Note 3.2). In this oscillatory state, nodes synchronize, leading to fast, low-amplitude oscillations. Due to the interactions between different oscillatory states, we were even able to predict chaotic behavior in a small parameter region. A noteworthy experimental implication is that if these fast, low-amplitude oscillations were observed without prior knowledge of this phenomenon, it might be mistaken for malfunctioning behavior, prompting unnecessary circuit modifications. This underscores the importance of theoretical mapping to complement experiments, as even well-characterized circuits can exhibit unexpected behaviors. Learning these possibilities beforehand allows for a better understanding of experimental outcomes and avoids misinterpretation.
The main objective of this study was to explore the potential of intracellularly coupled oscillators. While this work serves as a foundation, we hope it will encourage further computational and experimental studies. For example, studying intracellularly coupled oscillators can deepen our understanding of the mechanisms and interactions that govern ubiquitous natural oscillators. In addition, this research may yield critical insights into how synthetic oscillators interact with their natural counterparts. Such understanding could provide a powerful tool for engineering living systems, enabling the manipulation of biological processes through the coupling of synthetic and natural oscillators.
Further research is also required to experimentally implement synthetic intracellularly coupled oscillators. For instance, for deeply coupled oscillators, we showed that the same phenomena can be achieved through various methods, such as altering promoter strengths, protease concentrations, or repression strengths. Experimental realization will require identifying the optimal method to construct these circuits, as this study focused solely on fundamental phenomena.
However, we are optimistic that implementing coupled oscillators in E. coli is feasible. Prindle and colleagues, for instance, have already provided a proof of concept by successfully coupling two oscillators to achieve frequency modulation43. Regarding noise, it is worth emphasizing that individual oscillators serving as building blocks have been highly optimized to exhibit remarkably robust dynamics with minimal noise and phase drift29,30,31,102,103. For example, certain repressilator variants require up to 455 generations to drift by half a phase102. Furthermore, oscillator frequency can be externally modulated using small molecules or light32,33,34, enabling systematic parameter exploration and fine-tuning. Multiple orthogonal oscillators without shared parts have also been successfully implemented102.
A potential concern is the metabolic burden imposed by multiple oscillators on the host cell. Yet, a five-node repressilator has already been demonstrated, and the addition of one or two nodes represents only a modest increase in network size. More strikingly, a synthetic circuit comprising 35 genes (31 kb of DNA) has recently been implemented in a single E. coli cell, suggesting that accommodating two relatively small oscillators is well within reach. Moreover, CRISPRi-based circuits offer a promising alternative to transcription factor-based oscillators, as orthogonal sgRNAs can be readily designed, and their transcription generally imposes a low cellular burden104. We have previously shown that E. coli can host two functional three-node networks simultaneously30. While high levels of dCas9 can be toxic29,30,31,102,103, this limitation can be mitigated using engineered dCas9 variants or by employing dCas12a, which has been reported to be less toxic105,106.
A central challenge in experiments with coupled oscillators will be the need to record a sufficient number of oscillatory cycles to reliably characterize system dynamics. Since most synthetic oscillators depend on actively growing cells, experimental setups that allow to maintain long-term cultures will be required. Employing faster oscillators, such as the Goodwin oscillator, could be advantageous in this context. At these extended timescales, optimizing the evolutionary stability of the circuits may also become necessary.
When considering real-world applications of those circuits, environmental factors, notably temperature, must also be taken into account. The frequency of synthetic oscillators is known to be temperature-sensitive, which can affect their performance and dynamic behavior25,107. One way to address this limitation is through the design of temperature-compensated circuits, in which oscillatory periods remain stable despite fluctuations in temperature. The circadian clock provides a natural example of such robustness108,109,110, and Hussain and colleagues have shown that temperature compensation is also achievable in a synthetic oscillator111.
Another critical factor in these systems is the coupling strength. For independent or deeply coupled oscillators, determining the coupling mechanism is relatively straightforward, but for weakly or strongly coupled oscillators, careful consideration of coupling strength is essential. We proposed two potential mechanisms: protease-mediated coupling (Fig. 5) and coupling via the dCas system (Fig. 6). However, coupling could also emerge from other limited cellular resources40,41,42,43,44,45,46,47,48,49 (Fig. 6). In cases where the origin of synchronization is unclear, the well-established Kuramoto model38,112,113 could serve as a useful framework for studying coupled oscillators. Finally, in OBC systems, computation occurs during the initial phase of oscillation, yet our understanding of this transient phase remains limited. This gap highlights the need for additional experiments and, potentially, the refinement or expansion of the current modeling approaches.
Biological noise also modulates the coupling strength necessary for stable synchronization. As shown by our stochastic simulations, higher noise levels lead to a progressive decline in phase-locking (Fig. 6l), indicating that strong noise can disrupt coupling. However, coupling can be restored either by incorporating additional explicit interactions or through coupling mediated by limited shared resources (Fig. 6k–n). Thus, our findings suggest that OBC remains feasible and robust even under biologically noisy conditions.
The proposed systems operate on timescales of minutes to hours, which raises the question of whether such dynamics are sufficiently fast for real-time applications. However, several existing synthetic biology platforms already perform computation on comparable or even slower timescales57,114, and yet interest in this field is expanding rapidly because of its transformative potential. OBCs may help overcome this limitation by leveraging coupling and synchronization to enable inherently parallel computation, thereby offering the prospect of significant acceleration. Theoretical analyses65 also indicate that OBCs could solve computationally complex problems more efficiently than conventional digital systems. In such cases, slower timescales may be an acceptable trade-off for enhanced computational capability. Although these prospects remain speculative, we anticipate that ongoing conceptual and technological advances will continue to stimulate progress in this emerging area.
In conclusion, we believe that intracellularly coupled oscillators hold considerable promise for synthetic biology, both as a fundamental design principle and as a platform for diverse applications, making them a compelling direction for future research.
Methods
We used the GRN_modeler54 to design and model the circuits. Simulations were conducted in MATLAB (version 2023b) using the SimBiology toolbox with the Sundials solver. The simulations were performed with relative tolerances set to 10−8 and absolute tolerances set to 10−10, unless stated otherwise in the figure captions.
We determined the time period (T) of the oscillators and, for the beat phenomenon in Fig. 3, calculated the standard deviation of T using MATLAB’s findpeaks function.
Linear stability analysis
During the linear stability analysis (Supplementary Note 3.2) of the Goodwin oscillator, the fixed point was determined using MATLAB’s fsolve function, where the right-hand side of the ODE system was set to zero. The Jacobian matrix was then evaluated at the fixed point. If the real part of all eigenvalues of the Jacobian was negative, the fixed point was considered stable. For cases where the fixed point was unstable and had complex eigenvalues, stable periodic orbits were identified by starting simulations from the unstable fixed point. A 2 ⋅ 103 min simulation was run, and the highest and lowest values of the protein oscillations were calculated from the second half of the simulation.
In the case of the toggle switch, a similar procedure was performed; however, we had to check the stability of multiple fixed points. The fixed points were found first analytically with the solve function, then it was evaluated with the vpa function at the given protease concentrations. The stability of the fixed point was determined based on the eigenvalues of the Jacobian matrix.
Calculating the Lyapunov exponents
In the bifurcation diagrams for the visualization of the SPOs and the chaotic regions, each point represents the outcome of a simulation lasting 2 ⋅ 105 minutes. The first half of the simulation was discarded as transient behavior, while the second half was used to construct the bifurcation diagram. Following this, the simulation was extended to compute the Lyapunov exponents. These exponents were calculated using COPASI (version 4.38)115 via Python3.10 and the BasiCo interface (version 0.74)116 in MATLAB. Specifically, after a preliminary 103 min simulation, the Lyapunov exponents were determined from a subsequent 104 min simulation, with orthogonalization performed every 102 min. For these calculations, a relative tolerance of 10−4 and an absolute tolerance of 10−6 were applied. To accurately identify positive Lyapunov exponents and differentiate them from near-zero values, we computed the standard deviation (σ) of the exponents closest to zero. Exponents were considered positive if they exceeded 3σ, ensuring that the regions in the bifurcation diagrams marked with red (indicating positive Lyapunov exponents) were not artifacts of small numerical errors, particularly when theoretical values should be zero.
The methods used for stochastic simulations are presented in Supplementary Note 5.2.3, while the linear stability analysis is detailed in Supplementary Note 3.2. Stochastic simulations were implemented in C++ (g++ 11).
Statistics & reproducibility
All results were generated from numerical simulations using the code provided in the accompanying repository. Every figure and dataset can be reproduced exactly using the supplied scripts with default parameters and random seeds.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data used in this study are generated computationally and can be fully reproduced using the source code provided. As no external datasets were used, all numerical results and figures can be regenerated directly from the code. Please see the Code Availability section for details. The detailed descriptions of the models are available in Supplementary Data 1–16, and are cited in the relevant figure legends.
Code availability
This code is based on the GRN_modeler platform54, which provides tools for constructing, simulating, and analyzing gene regulatory networks. The code used to generate the results presented in this study is available in the following GitHub repository: https://github.com/SchaerliLab/GRN_modeler/tree/main/analysing_examples/coupled_oscillators. An archived version of the code used in this study is also available via Zenodo117.
References
Endy, D. Foundations for engineering biology. Nature 438, 449–453 (2005).
Cameron, D. E., Bashor, C. J. & Collins, J. J. A brief history of synthetic biology. Nat. Rev. Microbiol. 12, 381–390 (2014).
Purnick, P. E. M. & Weiss, R. The second wave of synthetic biology: from modules to systems. Nat. Rev. Mol. Cell Biol. 10, 410–422 (2009).
Khalil, A. S. & Collins, J. J. Synthetic biology: applications come of age. Nat. Rev. Genet. 11, 367–379 (2010).
Pursley, M. B. Reference Data for Engineers (Ninth Edition), (Newnes, Woburn, 2002).
Agrawal, D. K., Woodhouse, J. & Seshia, A. A. Synchronization in a coupled architecture of microelectromechanical oscillators. J. Appl. Phys. 115, 164904 (2014).
Hannan, M. A., Abbas, S. M., Samad, S. A. & Hussain, A. Modulation techniques for biomedical implanted devices and their challenges. Sensors 12, 297–319 (2012).
Babbush, R., Berry, D. W., Kothari, R., Somma, R. D. & Wiebe, N. Exponential quantum speedup in simulating coupled classical oscillators. Phys. Rev. X 13, 041041 (2023).
Gonze, D., Bernard, S., Waltermann, C., Kramer, A. & Herzel, H. Spontaneous synchronization of coupled circadian oscillators. Biophys. J. 89, 120–129 (2005).
Ramayya, A. G. et al. Theta Synchrony Is Increased near Neural Populations That Are Active When Initiating Instructed Movement. eneuro 8, ENEURO.0252–20.2020 (2021).
Strogatz, S. H. & Stewart, I. Coupled oscillators and biological synchronization. Sci. Am. 269, 102–109 (1993).
Henry, M. J. et al. An ecological approach to measuring synchronization abilities across the animal kingdom. Philos. Trans. R. Soc. B Biol. Sci. 376, 20200336 (2021).
Duranton, C., Bedossa, T. & Gaunet, F. Interspecific behavioural synchronization: dogs exhibit locomotor synchrony with humans. Sci. Rep. 7, 12384 (2017).
Bieler, J. et al. Robust synchronization of coupled circadian and cell cycle oscillators in single mammalian cells. Mol. Syst. Biol. 10, 739 (2014).
Yan, J. & Goldbeter, A. Robust synchronization of the cell cycle and the circadian clock through bidirectional coupling. J. R. Soc. Interface 16, 20190376 (2019).
Papagiannakis, A., Niebel, B., Wit, E. C. & Heinemann, M. Autonomous metabolic oscillations robustly gate the early and late cell cycle. Mol. Cell 65, 285–295 (2017).
Herrgen, L. et al. Intercellular coupling regulates the period of the segmentation clock. Curr. Biol. 20, 1244–1253 (2010).
Meijer, W. H. & Sonnen, K. F. From signalling oscillations to somite formation. Curr. Opin. Syst. Biol.39, 100520 (2024).
Tsiairis, C. D. & Aulehla, A. Self-organization of embryonic genetic oscillators into spatiotemporal wave patterns. Cell 164, 656–667 (2016).
Bastien, R., Guayasamin, O., Douady, S. & Moulia, B. Coupled ultradian growth and curvature oscillations during gravitropic movement in disturbed wheat coleoptiles. PloS ONE 13, e0194893 (2018).
Moulia, B., Douady, S. & Hamant, O. Fluctuations shape plants through proprioception. Science 372, eabc6868 (2021).
Goodwin, B. C. Temporal Organization in Cells; A Dynamic Theory of Cellular Control Processes (London, Academic Press, 1963).
Purcell, O., Savery, N. J., Grierson, C. S. & di Bernardo, M. A comparative analysis of synthetic genetic oscillators. J. R. Soc. Interface 7, 1503–1524 (2010).
Borg, Y., Alsford, S., Pavlika, V., Zaikin, A. & Nesbeth, D. N. Synthetic biology tools for engineering Goodwin oscillation in Trypanosoma brucei brucei. Heliyon 8, e08891 (2022).
Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Danino, T., Mondragón-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010).
Prindle, A. et al. A sensing array of radically coupled genetic ‘biopixels’. Nature 481, 39–44 (2012).
Potvin-Trottier, L., Lord, N. D., Vinnicombe, G. & Paulsson, J. Synchronous long-term oscillations in a synthetic gene circuit. Nature 538, 514–517 (2016).
Santos-Moreno, J., Tasiudi, E., Stelling, J. & Schaerli, Y. Multistable and dynamic CRISPRi-based synthetic circuits. Nat. Commun. 11, 2746 (2020).
Kuo, J., Yuan, R., Sánchez, C., Paulsson, J. & Silver, P. A. Toward a translationally independent RNA-based synthetic oscillator using deactivated CRISPR-Cas. Nucleic Acids Res. 48, 8165–8177 (2020).
Park, J. H., Holló, G. & Schaerli, Y. From resonance to chaos by modulating spatiotemporal patterns through a synthetic optogenetic oscillator. Nat. Commun. 15, 7284 (2024).
Mondragón-Palomino, O., Danino, T., Selimkhanov, J., Tsimring, L. & Hasty, J. Entrainment of a population of synthetic genetic oscillators. Science 333, 1315–1319 (2011).
Cannarsa, M. C., Liguori, F., Pellicciotta, N., Frangipane, G. & Di Leonardo, R. Light-driven synchronization of optogenetic clocks. Elife 13, RP97754 (2024).
Heltberg, M. S. et al. Coupled oscillator cooperativity as a control mechanism in chronobiology. Cell Syst. 14, 382–391 (2023).
Chen, Y., Kim, J. K., Hirning, A. J., Josić, K. & Bennett, M. R. Emergent genetic oscillations in a synthetic microbial consortium. Science 349, 986–989 (2015).
Hinze, T., Schumann, M., Bodenstein, C., Heiland, I. & Schuster, S. Biochemical frequency control by synchronisation of coupled repressilators: An in silico study of modules for circadian clock systems. Comput. Intell. Neurosci. 2011, 262189 (2011).
Lang, M., Marquez-Lago, T. T., Stelling, J. & Waldherr, S. Autonomous synchronization of chemically coupled synthetic oscillators. Bull. Math. Biol. 73, 2678–2706 (2011).
Fedorec, A. J. H., Karkaria, B. D., Sulu, M. & Barnes, C. P. Single strain control of microbial consortia. Nat. Commun. 12, 1977 (2021).
Sabi, R. & Tuller, T. Modelling and measuring intracellular competition for finite resources during gene expression. J. R. Soc. Interface 16, 20180887 (2019).
Shopera, T., He, L., Oyetunde, T., Tang, Y. J. & Moon, T. S. Decoupling resource-coupled gene expression in living cells. ACS Synthetic Biol. 6, 1596–1604 (2017).
Gyorgy, A. et al. Isocost lines describe the cellular economy of genetic circuits. Biophys. J. 109, 639–646 (2015).
Prindle, A. et al. Rapid and tunable post-translational coupling of genetic circuits. Nature 508, 387–391 (2014).
Payne, S. et al. Temporal control of self-organized pattern formation without morphogen gradients in bacteria. Mol. Syst. Biol. 9, 697 (2013).
Cookson, N. A. et al. Queueing up for enzymatic processing: correlated signaling through coupled degradation. Mol. Syst. Biol. 7, 561 (2011).
Moriya, T., Yamamura, M. & Kiga, D. Effects of downstream genes on synthetic genetic circuits. BMC Syst. Biol. 8, S4 (2014).
Barbier, I. et al. Synthetic gene circuits combining CRISPR interference and CRISPR activation in E. coli: importance of equal guide RNA binding affinities to avoid context-dependent effects. ACS Synthetic Biol. 12, 3064–3071 (2023).
Rutter, J., Reick, M. & McKnight, S. L. Metabolism and the control of circadian rhythms. Annu. Rev. Biochem. 71, 307–331 (2002).
Bi, S. et al. Dynamic fluctuations in a bacterial metabolic network. Nat. Commun. 14, 2173 (2023).
Fung, E. et al. A synthetic gene–metabolic oscillator. Nature 435, 118–122 (2005).
Német, N., Holló, G. & Lagzi, I. Carbon dioxide-driven coupling in a two-compartment system: Methyl red oscillator. J. Phys. Chem. A 124, 10758–10764 (2020).
Holló, G. & Lagzi, I. Autonomous chemical modulation and unidirectional coupling in two oscillatory chemical systems. J. Phys. Chem. A 123, 1498–1504 (2019).
Holló, G., Dúzs, B., Szalai, I. & Lagzi, I. From master-slave to peer-to-peer coupling in chemical reaction networks. J. Phys. Chem. A 121 17, 3192–3198 (2017).
Holló, G., Park, J. H., Boni, E. & Schaerli, Y. A tool for modeling gene regulatory networks (GRN_modeler) and its applications to synthetic biology. Mol. Syst. Biol. 21, 1618–1637 (2025).
Tomazou, M., Barahona, M., Polizzi, K. M. & Stan, G.-B. Computational re-design of synthetic genetic oscillators for independent amplitude and frequency modulation. Cell Syst. 6, 508–520.e5 (2018).
Nielsen, A. A. K. et al. Genetic circuit design automation. Science 352, aac7341 (2016).
Padmakumar, J. P. et al. Partitioning of a 2-bit hash function across 66 communicating cells. Nat. Chem. Biol. 21, 268–279 (2025).
Grozinger, L. et al. Pathways to cellular supremacy in biocomputing. Nat. Commun. 10, 5250 (2019).
Grozinger, L., Miró-Bueno, J. & Goñi-Moreno, Á Genetic designs for stochastic and probabilistic biocomputing. Phys. Rev. E 111, 054412 (2025).
Grozinger, L., Cuevas-Zuviría, B. & Goñi-Moreno, Á. Why cellular computations challenge our design principles. Semin. Cell Dev. Biol.171, 103616 (2025).
Furber, S. & Temple, S. Neural systems engineering. J. R. Soc. Interface 4, 193–206 (2007).
Kahana, M. J. The cognitive correlates of human brain oscillations. J. Neurosci. 26, 1669–1672 (2006).
Cole, S. R. & Voytek, B. Brain oscillations and the importance of waveform shape. Trends Cogn. Sci. 21, 137–149 (2017).
Todri-Sanial, A., Delacour, C., Abernot, M. & Sabo, F. Computing with oscillators from theoretical underpinnings to applications and demonstrators. Npj Unconv. Comput. 1, 1–16 (2024).
Csaba, G. & Porod, W. Coupled oscillators for computing: A review and perspective. Appl. Phys. Rev. 7, 011302 (2020).
Malagarriga, D. et al. Synchronization-based computation through networks of coupled oscillators. Front. Comput. Neurosci.9, https://doi.org/10.3389/fncom.2015.00097 (2015).
Ren, X. et al. Cardiac muscle cell-based coupled oscillator network for collective computing. Adv. Intell. Syst. 3, 2000253 (2021).
Ji, J. et al. Large-scale cardiac muscle cell-based coupled oscillator network for vertex coloring problem. Adv. Intell. Syst. 5, 2200356 (2023).
Lawson, H. S., Holló, G., Horvath, R., Kitahata, H. & Lagzi, I. Chemical resonance, beats, and frequency locking in forced chemical oscillatory systems. J. Phys. Chem. Lett. 11, 3014–3019 (2020).
Santos-Moreno, J., Tasiudi, E., Kusumawardhani, H., Stelling, J. & Schaerli, Y. Robustness and innovation in synthetic genotype networks. Nat. Commun. 14, 2454 (2023).
Itatani, M. et al. Temporal pH waveforms generated in an enzymatic reaction network in batch and cell-sized microcompartments. Cell Rep. Phys. Sci. 6, 102367 (2025).
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Lugagne, J.-B. et al. Balancing a genetic toggle switch by real-time feedback control and periodic forcing. Nat. Commun. 8, 1–8 (2017).
Rodriguez, S. R.-K. Classical and quantum distinctions between weak and strong coupling. Eur. J. Phys. 37, 025802 (2016).
Novotny, L. Strong coupling, energy splitting, and level crossings: A classical perspective. Am. J. Phys. 78, 1199–1202 (2010).
Zhang, F. et al. Independent control of amplitude and period in a synthetic oscillator circuit with modified repressilator. Commun. Biol. 5, 23 (2022).
Ceroni, F., Algar, R., Stan, G.-B. & Ellis, T. Quantifying cellular capacity identifies gene expression designs with reduced burden. Nat. Methods 12, 415–418 (2015).
Dennis, P. P. & Bremer, H. Modulation of chemical composition and other parameters of the cell at different exponential growth rates. EcoSal Plus 3, 1–49 (2008).
Campbell, M., Greated, C. & Myers, A. Musical Instruments: History, Technology, and Performance of Instruments of Western Music (Oxford University Press, 2004).
Udem, T., Holzwarth, R. & Hänsch, T. W. Optical frequency metrology. Nature 416, 233–237 (2002).
Hao, N. & O’shea, E. K. Signal-dependent dynamics of transcription factor translocation controls gene expression. Nat. Struct. Mol. Biol. 19, 31–39 (2012).
Cai, L., Dalal, C. K. & Elowitz, M. B. Frequency-modulated nuclear localization bursts coordinate gene regulation. Nature 455, 485–490 (2008).
Micali, G., Aquino, G., Richards, D. M. & Endres, R. G. Accurate encoding and decoding by single cells: amplitude versus frequency modulation. PLoS Comput. Biol. 11, e1004222 (2015).
Ye, C. et al. The cyanobacterial circadian clock couples to pulsatile processes using pulse amplitude modulation. Curr. Biol. 34, 5796–5803 (2024).
Lee, C. K. et al. Broadcasting of amplitude-and frequency-modulated c-di-GMP signals facilitates cooperative surface commitment in bacterial lineages. Proc. Natl. Acad. Sci. USA 119, e2112226119 (2022).
Rajasekaran, R., Chang, C.-C., Weix, E. W., Galateo, T. M. & Coyle, S. M. A programmable reaction-diffusion system for spatiotemporal cell signaling circuit design. Cell 187, 345–359 (2024).
Wu, Y. et al. Rapid and dynamic detection of antimicrobial treatment response using spectral amplitude modulation in mzo nanostructure-modified quartz crystal microbalance. J. Microbiol. Methods 178, 106071 (2020).
Ruppert, M. G., Harcombe, D. M., Ragazzon, M. R. P., Moheimani, S. O. R. & Fleming, A. J. A review of demodulation techniques for amplitude-modulation atomic force microscopy. Beilstein J. Nanotechnol. 8, 1407–1426 (2017).
Rosanne, O. Alves de Oliveira, A. & Falk, T. H. Eeg amplitude modulation analysis across mental tasks: Towards improved active bcis. Sensors 23, https://www.mdpi.com/1424-8220/23/23/9352 (2023).
Vélez, P. et al. Single-frequency amplitude-modulation sensor for dielectric characterization of solids and microfluidics. IEEE Sensors J. 21, 12189–12201 (2021).
Moraskie, M. et al. Microbial whole-cell biosensors: Current applications, challenges, and future perspectives. Biosens. Bioelectron. 191, 113359 (2021).
Oliveira, H. M. & Melo, L. V. Huygens synchronization of two clocks. Sci. Rep. 5, 11548 (2015).
Yogendra, K., Liyanagedera, C., Fan, D., Shim, Y. & Roy, K. Coupled spin-torque nano-oscillator-based computation: A simulation study. J. Emerg. Technol. Comput. Syst. 13, 1–24 (2017).
Csaba, G. & Porod, W. Perspectives of using oscillators for computing and signal processing. Preprint at https://doi.org/10.48550/arXiv.1805.09056 (2018).
Al Beattie, B., Noll, M., Kohlstedt, H. & Ochs, K. Oscillator-based optimization: design, emulation, and implementation. Eur. Phys. J. B 97, 7 (2024).
Gutiérrez Mena, J., Kumar, S. & Khammash, M. Dynamic cybergenetic control of bacterial co-culture composition via optogenetic feedback. Nat. Commun. 13, 4808 (2022).
Liu, X. et al. Convergent engineering of syntrophic escherichia coli coculture for efficient production of glycosides. Metabolic Eng. 47, 243–253 (2018).
Li, X. et al. Design of stable and self-regulated microbial consortia for chemical synthesis. Nat. Commun. 13, 1554 (2022).
Graf, I. R. & Machta, B. B. A bifurcation integrates information from many noisy ion channels and allows for milli-Kelvin thermal sensitivity in the snake pit organ. Proc. Natl. Acad. Sci. USA 121, e2308215121 (2024).
Dolinsek, J. & Vilfan, M. Novel NMR and EPR Techniques (Springer, Germany, 2006).
Khodaie, N. Vibration control of super-tall buildings using combination of tapering method and tmd system. J. Wind Eng. Ind. Aerodyn. 196, 104031 (2020).
Luro, S., Potvin-Trottier, L., Okumus, B. & Paulsson, J. Isolating live cells after high-throughput, long-term, time-lapse microscopy. Nat. Methods 17, 93–100 (2020).
Niederholtmeyer, H. et al. Rapid cell-free forward engineering of novel genetic ring oscillators. Elife 4, e09771 (2015).
Santos-Moreno, J. & Schaerli, Y. CRISPR-based gene expression control for synthetic gene circuits. Biochem. Soc. Trans. 48, 1979–1993 (2020).
Zhang, S. & Voigt, C. A. Engineered dCas9 with reduced toxicity in bacteria: implications for genetic circuit design. Nucleic Acids Res. 46, 11115–11125 (2018).
Vercauteren, S. et al. The rise and future of crispr-based approaches for high-throughput genomics. FEMS Microbiol. Rev. 48, fuae020 (2024).
Powell, D., Haddad, S. A., Gorur-Shandilya, S. & Marder, E. Coupling between fast and slow oscillator circuits in cancer borealis is temperature-compensated. Elife 10, e60454 (2021).
Hastings, J. W. & Sweeney, B. M. On the mechanism of temperature independence in a biological clock. Proc. Natl. Acad. Sci. USA 43, 804–811 (1957).
Gould, P. D. et al. The molecular basis of temperature compensation in the arabidopsis circadian clock. Plant Cell 18, 1177–1187 (2006).
Baker, C. L., Loros, J. J. & Dunlap, J. C. The circadian clock of Neurospora crassa. FEMS Microbiol. Rev. 36, 95–110 (2012).
Hussain, F. et al. Engineered temperature compensation in a synthetic genetic clock. Proc. Natl. Acad. Sci. 111, 972–977 (2014).
Kuramoto, Y. International Symposium on Mathematical Problems in Theoretical Physics, 420–422 (Springer Berlin Heidelberg, Berlin, Heidelberg, 1975).
Kuramoto, Y. Chemical Oscillations, Waves, and Turbulence. (Dover Publications, 2003).
Bonnerjee, D. et al. Multicellular artificial neural network-type architectures demonstrate computational problem solving. Nat. Chem. Biol. 20, 1524–1534 (2024).
Hoops, S. et al. COPASI-a COmplex PAthway SImulator. Bioinformatics 22, 3067–3074 (2006).
Bergmann, F. T. Basico: A simplified python interface to copasi. J. Open Source Softw. 8, 5553 (2023).
Holló, G., Park, J. H., Boni, E. & Schaerli, Y. Schaerlilab/GRN_modeler: GRN_modeler. https://doi.org/10.5281/zenodo.17579469 (2025).
Acknowledgements
This work was funded by the Swiss National Science Foundation (10005200 awarded to Y.S.), a fellowship of the Agassiz foundation (awarded to J.P.), a UNIL FBM Ph.D fellowship in Life Sciences (awarded to J.P.) and the University of Lausanne.
Author information
Authors and Affiliations
Contributions
G.H. performed the mathematical modeling. G.H., J.P., R.A.E., and Y.S. wrote the manuscript. All authors have given approval to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Mogens Jensen, who co-reviewed with Alessandra Lucchetti, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Holló, G., Park, J.H., Evard, R.A. et al. Intracellularly coupled oscillators for synthetic biology. Nat Commun 17, 1131 (2026). https://doi.org/10.1038/s41467-025-67893-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67893-0