Abstract
A small percentage of people living with HIV (PLHIV) spontaneously regulate viral replication without suppressive antiretroviral treatment (ART) and are categorized into ‘elite controllers’ (EC, HIV-RNA < 50 c/mL) and ‘viremic controllers’ (VC, HIV-RNA between 50-10,000 c/ml). Some EC and VC may lose controller status in time. Here we provide extensive plasma proteomics to identify biomarkers and pathways related to spontaneous viral control and its long-term preservation among 36 EC and 147 VC (discovery) and 14 EC and 5 VC (validation). VC exhibited higher concentrations of CRTAM, LY9, and CD6 and lower concentrations of VAT1 compared to EC in both the discovery and validation cohort. Longitudinal analysis of pre- and post-ART samples (median follow-up: 5.3 years) revealed downregulation of various immune-related proteins in both EC and VC. Over a 17-year follow-up period, loss of viral control occurred in 31% of VC and 3% of EC. T-cell associated proteins (CRTAM, LY9, CD6), along with ICAM3, SH2D1A, C1QL2, and CNGB3, predicted loss of viral control years before its occurrence. Markers of chronic immune activation (sPD-L1, sCD25, IL-10, TGF-β, IFN-γ, and TNF-α) and systemic inflammation (TNF, IL-1β, IL-6, and sCD14) were not predictive. Our findings underscore the dynamic interplay between T cell function and viral replication in maintaining HIV control and identify key biomarkers that predict viral load surges.
Similar content being viewed by others
Introduction
A small percentage of people living with HIV-1 (PLHIV) spontaneously and durably control viral replication and do not progress to AIDS in the absence of suppressive antiretroviral treatment (ART)1. This group of patients is heterogeneous, with different groups being commonly recognized, including elite controllers (EC), who are typically characterized by plasma HIV-RNA < 50 copies/ml for more than 12 months1 and viremic controllers (VC), here defined by a low yet detectable viremia on routine assays, most commonly < 400 copies/mL2, < 2000 copies/mL3,4, or < 10,000 copies/mL1,5,6. Furthermore, transient controllers (TC) lose control status over time in contrast to persistent controllers (PC). Loss of viral control is common and has been attributed to both immune control mechanisms as well as viral factors7,8. A better understanding of the underlying pathways in the various groups may benefit post-treatment control interventions.
Viral control has been attributed to the preferential integration of intact proviral DNA into non-coding regions of EC host DNA. Cells harboring intact provirus integrated into permissive chromatin are prone to be eliminated due to a strong antiviral host immune response9,10. Long-term persistence and development of the HIV reservoir in EC is mediated primarily through clonal proliferation of latently infected cells, in contrast to VC in whom ongoing, low-level viral replication occurs10,11,12. A recent paper confirmed such viral mechanism also in PC in contrast to TC who showed higher intact-genome proviral levels that were transcriptionally active preceding the loss of spontaneous control13. Other studies in HIV controllers also showed importance viral variability and suggest role for host immune factors11,14,15. Indeed, viral factors in EC and VC are closely intertwined with host immune factors that may be altered preceding the loss of spontaneous control16,17. Notwithstanding, the systemic host mechanisms that contribute to spontaneous HIV control are still not yet fully understood, and predictive biomarkers are needed9.
High-throughput omics technologies can improve our understanding by uncovering changes in pathophysiological processes through the detection of a wide range of plasma proteins, including intra-cellular, secreted, and low-abundance proteins18,19.
In the present study, our primary objective was to identify biomarkers and pathways related to spontaneous HIV control and its long-term preservation. To achieve this, we performed three main comparisons. First, we compared the relative expression of 2420 plasma proteins during viral control between ART-naïve EC (n = 36) and VC (n = 147) in a discovery cohort and validated findings in an independent cohort consisting of persistent, ART naïve EC (n = 5) and VC (n = 14). Second, we studied paired samples taken before and after ART initiation from 57 persistent VC, 36 transient VC, and 9 persistent EC. Lastly, we evaluated whether circulating proteins at baseline were associated with loss of viral control (56 TC and 125 PC) and/or >1000 copies/mL increase in plasma viral load during follow-up.
Here, we show similarities in proteomic profile between EC and VC. However, we observed the consistent upregulation of three proteins (CRTAM, LY9, and CD6) in VC, suggesting their potential role in limiting viral replication. Moreover, post-ART proteomic profiles displayed a downregulation of immune-related proteins, including CRTAM, LY9, and CD6. Lastly, we observed changes in the circulating concentrations of some proteins (ICAM3, SH2D1A, CRTAM, C1QL2, LY9, CD6, CNGB3) years before the loss of HIV controller status and/or substantial increase of viral load. These proteins are associated with distinct immune and cellular processes, including T-cell function, apoptosis, and anti-viral response, and may serve as biomarkers of future viral load surges. In addition, analysis of plasma RNA sequences in VC revealed the presence of both genetically-intact and defective sequences and substantial genetic diversity among the viral sequences.
Results
Selective upregulation of CRTAM, LY9, and CD6 plasma concentrations in viremic controllers compared to elite controllers
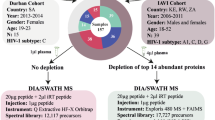
This study encompassed a proteome analysis within two cohorts: a discovery cohort consisting of 183 persistent HIV controllers and a validation cohort comprising 19 persistent HIV controllers (Fig. 1A). Patient characteristics are provided in Table 1. In the discovery cohort, EC and VC differed in age, BMI, CD4 cell counts, plasma viral load and sample storage time, whereas in the validation cohort, age, BMI, and CD4 cell counts were similar between groups. Notably, in the discovery cohort, the duration of HIV infection was similar for both EC and VC, with both groups maintaining their controller status for a median period of 5 years (5.19 [0.03, 34.5] for EC and 5.39 [0, 31.2] for VC).
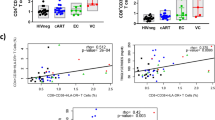
A Overview of study participants from the discovery and validation cohorts; and overview of longitudinal follow-up. B Volcano plot of the differentially expressed proteins between ART-naïve viremic (VC, N = 147) and elite controllers (EC, N = 36). X-axis depicts Log2 fold change of differentially expressed proteins (n = 4) in the discovery cohort and Y-axis depicts -log10(FDR P value). Differential expression analysis was performed using a linear regression model with age, sex, storage time, and center of inclusion as covariates. Only proteins that show FDR < 0.05 were annotated. See also Supplementary Data 1. C Violin boxplot showing the protein expression CRTAM, LY9, CD6, and VAT1 in 147 VC and 36 EC of the discovery cohort (elite-ATHENA cohort). FDR P values were derived from differential expression analysis with age, sex, storage time, and center of inclusion as covariates. In all boxplots, the in-box line defines the median value, hinges depict 25th and 75th percentiles and whiskers extend to 1.5 interquartile ranges; each dot indicates an individual participant. D Boxplot showing the comparison CRTAM, LY9, CD6, and VAT1 between 147 VC and 36 EC stratified by viral load categories of the discovery cohort (elite-ATHENA cohort). P values were derived from two-sided Wilcoxon test. In all boxplots, the in-box line defines the median value, hinges depict 25th and 75th percentiles and whiskers extend to 1.5 interquartile ranges; each dot indicates an individual participant. Exact P values in plot where P < 0.0001: CRTAM < 2000 vs EC P = 6.159925e-07 CRTAM < 10000 vs EC P = 8.740113e-09. LY9 < 2000 vs EC P = 1.432240e-06. LY9 < 10000 vs EC P = 4.506206e-08. CD6 < 2000 vs EC P = 5.603601e-08. CD6 < 10000 vs EC P = 1.078603e-07. E Violin boxplot showing the comparison CRTAM, LY9, CD6, and VAT1 in 5 VC and 14 EC of the validation cohort (2000HIV cohort). P values were derived from differential expression analysis with age and sex as covariates. In all boxplots, the in-box line defines the median value, hinges depict 25th and 75th percentiles and whiskers extend to 1.5 interquartile ranges; each dot indicates an individual participant. See also Supplementary Data 2.
To compare the relative concentrations of 2420 plasma proteins between persistent, ART-naïve EC and VC, we conducted a differential expression analysis involving 147 VC and 36 EC in the discovery cohort. Analysis was corrected for potential confounders such as age, sex, center of inclusion, and sample storage time (see methods). Overall, the proteomic profiles of EC and VC were largely similar, with three proteins being significantly upregulated in VC, namely CRTAM (Cytotoxic and Regulatory T Cell Molecule; log2 fold change = 0.74; FDR-adjusted P value = 0.01), LY9 (Lymphocyte Antigen 9; log2 fold change = 0.45; FDR-adjusted P value = 0.02), and CD6 (log2 fold change = 0.74; FDR-adjusted P value = 0.04). Conversely, we found a downregulation of VAT1 (Vesicle Amine Transport 1; log2 fold change = −0.27; FDR-adjusted P value = 0.04) in VC (Fig. 1B, C; and Supplementary Data 1). Additionally, considering different definitions of plasma viral load cut-off in viremic controllers, we investigated proteomics differences in viral load setpoints using principal component analysis. Our analysis revealed similar proteomic profiles across different viral load cutoff values (undetected, ≤400, ≤2000, and ≤10,000) (Supplementary Fig. 2A). Furthermore, we conducted a comparison of CRTAM, LY9, CD6, and VAT1 between EC and three distinct viral load categories within viremic controllers ( ≤ 400, ≤2000, and ≤10,000) (Fig. 1D). Our findings consistently showed higher concentrations of these four markers in VC groups across different viral load categories compared to EC (Wilcoxon test P values < 0.05), except for CD6 and VAT1, which exhibited similar levels between EC and VC with viral load < 400 copies/mL. Of note, we consistently found increased concentrations of these four markers in conjunction with higher biomarkers of viral load.
A, D, E Volcano plot of the differentially expressed proteins between paired participants pre- and post-ART initiation in (A) persistent VC (N = 57), (D) transient VC (N = 36), and (E) persistent EC (N = 9) in the discovery cohort. X-axis depicts Log2 fold change of differentially expressed proteins and Y-axis depicts the (A, D) -log10(FDR P value) or (E) -log10(P value). Differential expression analysis was performed using linear mixed models with (A, D) age, sex center and storage time or (E) age and storage time as covariates. See also Supplementary Data 3,5,6. B Correlation plot between the effect of ART and time of the differentially expressed proteins across paired persistent VC pre- and post-ART initiation from A. X-axis depicts Log2 fold change of differentially expressed proteins before and after ART in 57 persistent VC. Y-axis depicts Log2 fold change of differentially expressed proteins between two ART-naïve time points in 23 persistent VC. Only one of differentially expressed proteins after ART was also nominally significant for a change over time (blue dot). Grey line depicts linear correlation line (pearson coefficient). No statistical testing was done. C Network of functional pathways based on enrichment analysis of downregulated differentially expressed proteins in post-ART samples of persistent VC (N = 57). Circular nodes represent pathways, with the colors showing a gradient of the enrichment FDR P values; node size represents the number of genes in the pathway; weighted edges represent the degree of gene overlap score between pathways. Only pathways with P value < 0.05 were depicted in the figure. See also Supplementary Data 4.
While differences in body mass index (BMI) exist between EC and VC in the discovery cohort, and the fact that BMI, smoking status, and white ethnicity were associated with proteomic profiles (Supplementary Fig. 1A), our analysis was not adjusted for these variables due to a substantial amount of missing data. We conducted a sensitivity analysis for the concentrations of CRTAM, LY9, CD6, and VAT1 between VC and EC, stratified by BMI, smoking status, and white ethnicity. This analysis revealed mainly upregulation of CRTAM, LY9, and CD6, alongside downregulation of VAT1 in VC compared to EC (Wilcoxon test P values < 0.05) (Supplementary Fig. 2B-D).
To corroborate our findings in the validation cohort, we performed a differential expression analysis involving four proteins (CRTAM, LY9, CD6, and VAT1), comparing 5 VC and 14 EC from the 2000HIV validation cohort. This analysis was adjusted for age and sex, as justified in the methodology. Our analysis confirmed the previously observed significant upregulation of CRTAM (log2 fold change = 0.83, P value = 0.005), LY9 (log2 fold change = 0.59, P value = 0.04), and CD6 (log2 fold change = 0.66, P value = 0.005), and VAT1 (log2 fold change = −0.35, P value = 0.04) in VC. (Fig. 1E; and Supplementary Data 2).
Changes in overall circulating proteins after the initiation of ART in HIV controllers, including the downregulation of CRTAM, LY9, CD4
To investigate the impact of ART on the proteomic profile of HIV controllers, we collected paired blood samples obtained before and after the initiation of ART from 102 HIV controllers from the elite-ATHENA participants, consisting of 36 transient VC, 57 persistent VC, and 9 persistent EC. While the exact reasons for ART initiation were not recorded, there was a notable trend of ART initiation around 2015, coinciding with the shift in guidelines that recommended ART for all PLHIV regardless of clinical stage, CD4 count or viral load levels. The time interval between the collection of the first and second sample varied, being 10.5 years for transient VC, 9.4 years for persistent VC, and 8.1 years for persistent EC. The median duration of ART was 5.3 years for both transient and persistent VC, and 3 years for persistent EC. Across all groups, the predominant ART combinations consisted of integrase strand transfer inhibitor (INSTI; 56%), followed by non-nucleoside reverse transcriptase inhibitor (NNRTI; 36%) and Protease inhibitor-based regimens (PI; 8%), as outlined in Table 2.
Initially, using principal component analysis, we identified an overall similar proteomic profile between HIV controllers before and after ART initiation (PC1 and PC2 Wilcoxon test P value > 0.05) (Supplementary Fig. 3A). Comparable overall proteomic profile was also observed between HIV controllers with varying durations of ART ( ≤ 1 year, 1–5 years, 5–10 years, > 10 years) and those receiving different ART regimens (INSTI, NNRTI, and PI-based) (PC1 and PC2 Wilcoxon test P value > 0.05). (Supplementary Fig. 3A).
Next, we conducted paired differential protein expression analyses of 2420 plasma proteins before and after ART initiation in 36 transient VC, 57 persistent VC, and 9 persistent EC. The analyses were adjusted for age, sex, center of inclusion, and sample storage time, as outlined in the methodology. In persistent VC (n = 57), a significant downregulation in 47 proteins after ART initiation (FDR P value < 0.05) was noticed, without any proteins showing significant upregulation (Fig. 2A and Supplementary Data 3). Among the proteins exhibiting significant downregulation after ART initiation were LY9 (log2 fold change = −0.63; FDR P = 0.005) and CD6 (log2 fold change = −1.46; FDR P = 0.0007). Additionally, CRTAM demonstrated a downregulation, although being marginally significant after FDR correction (log2 fold change = −0.57; FDR P = 0.08). Of note, VAT1 concentrations remained similar following ART initiation (Fig. 2A). To explore the stability of protein expression over time, we examined protein expression in paired samples of ART-naïve persistent VC (n = 23) and EC (n = 19) (Supplementary Fig. 3B-C). In contrast to the impact of ART on protein expression, we did not observe major differences in protein expression between two time points in persistent VC and EC who remained ART-free. Moreover, differentially expressed proteins after ART in persistent VC were not correlated with trends between two ART-naive timepoints (Fig. 2B). Age-matched nested case-control analyses before and after ART further confirmed that LY9, CD6 and CRTAM were highly downregulated, independently from age (Supplementary Fig. 3D).
Furthermore, we performed pathway analysis of the downregulated differentially expressed proteins after ART initiation of persistent VC (n = 47 proteins) and identified a predominant downregulation of immune-related pathways, including ‘cell surface receptor signaling pathway’, ‘adaptive immune system’, and ‘immunoregulatory interactions between lymphoid and non-lymphoid cells’ (Hypergeometric test FDR P < 0.05) (Fig. 2C and Supplementary Data 4).
When comparing pre- and post-ART initiation states for transient VC (n = 36) and persistent EC (n = 9), we observed that only 5 proteins passed the threshold for multiple testing correction in the former, and none in the latter (Fig. 2D, E). Pathway analysis for both comparisons was not done due to limited number of significant proteins. Nonetheless, it is important to note that the majority of proteins were downregulated in transient VC (162 downregulated proteins) and persistent EC (113 downregulated proteins) (P value < 0.05) (Supplementary Data 5-6). Furthermore, similar to the pattern observed in persistent VC, we detected a downregulation trend of CRTAM, LY9, and CD6 concentrations in both transient VC and persistent EC after ART initiation (P value < 0.05) (Fig. 2D, E). Of note, after ART initiation, VAT1 had an upregulated trend in transient VC (P value < 0.05) and remained similar in persistent EC.
Given the observed age differences across subgroups at the time of the second sampling, with persistent EC having a median age of 60.2 years, persistent VC 46.9 years, and transient VC 52.1 years (Table 2), we further assessed the extent to which age could contribute to the variation in proteomic changes. We repeated the paired analyses pre- and post- ART in age-matched subgroups. After 1:2 matching, all three groups reached a comparable age distribution at the second sampling, with median ages of 60.2 years (IQR 16.8) in persistent EC, 60.1 years (IQR 15.8) in persistent VC, and 59.6 years (IQR 15.1) in transient VC. Correspondingly, their pre-ART ages were also aligned, with median ages of 49.5 years (IQR 19.1), 46.1 years (IQR 21.4), and 48.4 years (IQR 18.4), respectively. The variation in top differentially expressed proteins across the age-matched paired comparisons persisted, indicating that the observed proteomic differences are unlikely to be attributable to age alone (Supplementary Fig. 3D).
Duration of viral control status and baseline plasma proteins associated with loss of viral control
A subset of HIV controllers experiences a loss of their controller status. To gain insights into the underlying mechanisms behind this phenomenon, we examined the plasma proteome levels prior to losing HIV controller status (VL > 10,000 copies/mL). We show in our cohort of 181 ART-naïve HIV controllers, that loss of control is common over time, especially in VC (38%; 55/145) compared to EC (3%; 1/36) during a follow-up period of up to 17 years. Among 35 persistent EC, 13 individuals (37%) were prescribed ART, and among this group, nine had a viral load higher than 50 copies/mL but lower than 10,000 copies/mL at the time of ART initiation, thereby becoming viremic controllers.
To explore the dynamic of viral load levels among persistent and transient HIV controllers, we examined the log10-change in viral load over time. As expected, we observed transient controllers exhibited higher initial viral load levels compared to persistent controllers (median [Interquartile range] 1240 [212, 4210] vs 96 [50, 1070] copies/mL). Over time, despite censoring, measurements of viral load in PC remained relatively stable, while TC showed a gradual increase in viral load (Fig. 3A; and Supplementary Fig. 4A). Furthermore, we explored the time until virological loss using a survival analysis. Our findings revealed that 50% of HIV controllers experienced virological loss (median loss-free survival time) within 20 years of follow-up (Fig. 3B).
A Dot plot of all available historical viral load measurements between transient and persistent HIV controllers. Y axis depicts log10 viral load and X-axis depicts years since first HIV RNA measurement. This analysis included 181 ART-naive HIV controllers (145 VC and 36 EC). The lines indicate the local regression (LOESS) fit; each dot represents an individual viral load measurement from one participant. B Kaplan-Meier loss-free follow-up. Vertical tick marks represent censored data points, indicating either end of follow-up or censoring by ART initiation in the absence of loss of virological control. The number at risk and cumulative number of events and censoring are reported. Loss of virological control is defined as reaching >10,000 copies/mL. This analysis included 181 ART-naive HIV controllers (145 VC and 36 EC). C, D Volcano plot of the differentially expressed proteins associated with (C) virological loss, defined as reaching >10,000 copies/mL or (D) substantial virus increase, defined as a > 1000 copies/mL increase in moving median of viral load or reaching >10,000 copies/mL. X-axis depicts hazard ratio of differentially expressed proteins and Y-axis depicts -log10(FDR P value), where a hazard ratio >1 means increased risk of (C) virological loss or (D) substantial virus increase. Differential expression analysis was performed using cox regression with age, sex, center and storage time as covariates. This analysis included 181 ART-naive HIV controllers (145 VC and 36 EC). E Forest plot of the hazard ratios or virological loss, substantial virus increase and immunological loss, with error bars representing 95% confidence intervals (CI) of proteins that were FDR significant in either C. and/or D., as determined with cox regression with age, sex, center and storage time as covariates. Virological loss was defined as reaching >10,000 copies/mL, substantial virus increase as a > 1000 copies/mL increase in moving median of viral load or reaching >10,000 copies/mL, and immunological loss as reaching CD4 < 500. Immunological loss was only assessed for individuals with available longitudinal CD4 data and CD4 counts >500 at the time of proteomics sample (n = 138). F Forest plot of the hazard ratios of experiencing a detectable viral load ( > 50 copies/mL) during follow-up for EC only (n = 36). Dots represent hazard ratios and error bars the 95% CI intervals. Black dots and lines represent all EC, whereas grey dots and lines are a selection of EC who were not experiencing a blip (plasma viral load 50–1000 copies/mL) at the time of proteomics measurement.
Next, we explored which biological pathways and biomarkers are potentially linked with the persistence of HIV controller status using the plasma levels of 2420 proteins. First, we correlated the baseline plasma protein concentrations with time to loss of viral control using cox regression analysis adjusting for age, sex, center of inclusion and sample storage time (see “methods”). We found one and four proteins associated with lower and higher risk of loss of viral control in VC, respectively, observed over a median period of 4.2 years before loss of HIV control (FDR P < 0.05; Fig. 3C; and Supplementary Data 7). Proteins that were linked to an increased risk of losing viral control including ICAM3 (intercellular adhesion molecule 3) (HR [Hazard Ratio] = 5.4; FDR P = 0.018), SH2D1A (SH2 domain–containing protein 1 A) (HR = 1.68; FDR P = 0.023), CRTAM (HR = 2.79; FDR P = 0.046), and C1QL2 (Complement C1q like 2) (HR = 2.42; FDR P = 0.046). In addition, CNGB3 (cyclic nucleotide gated channel subunit beta 3) (HR = 0.29; FDR P = 0.046) was associated with a lower risk of losing viral control.
Proteins associated with a future substantial increase in viral load and decline in CD4in HIV controllers
Several HIV controllers displayed increasing trends in viral load before the initiation of ART, suggesting that virological loss would have been imminent without ART. We investigated whether proteomic expression could predict the risk of a significant increase in viral load compared to the baseline level at proteomic sample collection. A substantial increase in viral load was defined as either a surge of over 1000 copies/mL in the moving median viral load at the time of proteomic sample inclusion or when the viral load exceeded 10,000 copies/ml. Our analysis revealed that 50% (91 out of 181) of PLHIV experienced a substantial increase in their moving median viral load, with 4 out of these 91 being EC. Cox regression analysis, adjusted for age, sex, center, and storage time, identified four proteins (ICAM3, CRTAM, CD6, LY9) associated with a heightened risk of a significant increase in moving median viral load (Fig. 3D, E; Supplementary Data 8). In targeted fashion, we evaluated whether proteins significantly associated with virological loss and/or substantial viral load increase (ICAM3, CRTAM, CD6, LY9, SH2D1A, C1QL2, CNGB3) were also associated with increased risk of a detectable viral load after the proteomics sample in EC. This occurred in 15/36 EC during follow-up. Importantly, ICAM3, CRTAM, CD6 and LY9 were associated with shorter time to first detectable viral load after proteomics sample (Fig. 3F).
Subsequently, we investigated the CD4 count dynamics among HIV controllers after the proteomics sample (Supplementary Fig. 4C, D). The rate of significant viral load increase was higher than the rate of HIV controllers reaching CD4 counts < 500 cells/mm3 (Supplementary Fig. 4E). Moreover, we explored whether proteomic expression could predict time to decrease in CD4 cell count below 500 cells/mm3. Out of 138 people living with HIV with available CD4 counts and a moving median CD4 count > 500 cells/mm3 at the proteomics sample, 42 (30%) experienced a decline in the moving median CD4 count to < 500 cells/mm3. After multiple testing correction, we did not identify any significant proteins associated with a substantial decrease in CD4 count. However, it is noteworthy that proteins such as ICAM3, CRTAM, and LY9 were among those associated with a higher risk of a substantial decrease in CD4 count among HIV controllers (unadjusted P value < 0.05, Fig. 3E; and Supplementary Fig. 4F).
Performance of clinical and proteomic models to predict loss of viral control and substantial increase in viral load or decrease in CD4
We developed predictive models to assess the risk of viral control loss, substantial increases in viral load, and CD4 count decline in HIV controllers. Different models were trained on a training dataset and evaluated on an external test set. In univariate cox regressions and multivariate random survival forest analyses, the most consistent clinical predictors were baseline CD4 count, baseline viral load, biological sex and age (Supplementary Fig. 5A).
Clinical models, trained on 839 HIV controllers from the ATHENA cohort20, were tested on 181 controllers and accuracy was reported as area under the receiver operating curve (AUC), where 0.5 is random and 1.0 is full accuracy. With only clinical predictors, we observed moderately predictive accuracy for viral control loss over a 2-year period, although accuracy declined below 0.75 for longer time frames (Supplementary Fig. 5B). Predictions of substantial viral load increases and CD4 decline were less robust, with modest accuracy on test data for clinical predictors alone.
Adding proteomic data (candidate proteins, nominal P < 0.05) to the models, we split the cohort into training (134 controllers from OLVG and EMC) and testing (47 controllers from UMCU, MSZ, RUMC, LUMC) groups. Despite the smaller sample, models incorporating proteomic data achieved near-perfect accuracy in predicting viral control loss and CD4 decline and high accuracy (AUC > 0.8) in predicting viral load increase, up to 8 years in the future (Supplementary Fig. 6A, B). These findings highlight the added value predictive power of plasma proteomics to reliably predict loss of viral or immunological control.
Link between proteins associated with loss of viral control and biomarkers of inflammation and immune activation
To further validate the identified proteins associated with loss of HIV controller status and/or substantial increases in viral load (CRTAM, LY9, CD6, ICAM3, SH2D1A, C1QL2, and CNGB3), we leveraged HIV controllers data from the 2000HIV cohort, which includes measurements of well-established inflammatory markers (TNF, IL-6, IL-1β, and sCD14), flow cytometry data quantifying different HLA-DR⁺ and/or CD38⁺ immune cell subsets, and longitudinal viral load assessments over the past three years. These viral load data include viral blips (VL 40–200 copies/mL), persistent viremia (VL 200–1000 copies/mL), and unknown viral status.
We found moderate to strong associations between LY9 (rho=0.71), ICAM3 (rho=0.64), SH2D1A (rho=0.53), and C1QL2 (rho=0.18) with sCD14, a marker of monocyte activation (Spearman P < 0.05; (Supplementary Fig. 7A). Additionally, we found significant correlations between SH2D1A, CRTAM, and CD6 and the cell subsets in activation Modules 2 and 3. These modules are dominated by cytotoxic and memory compartments, including HLA-DR⁺CD38⁺ CD8⁺ central/effector-memory T cells and HLA-DR⁺ γδ T cells, defining a focused cytotoxic/memory activation axis. Among the top associations were SH2D1A with CD8⁺HLA-DR⁺CD38⁺ (rho = 0.75, P value = 0.0002), CRTAM with TCRγδ HLA-DR⁺ (rho = 0.71, P value = 0.001), and SH2D1A with TCRγδ HLA-DR⁺ (rho = 0.69, P value = 0.001). These results might indicate antigen-driven CD8⁺ and TCRγδ activation as the main drift toward viral escape, likely driving immune exhaustion and eventual loss of viral control (Supplementary Fig. 7B, and Supplementary Data 9). We also observed trends suggesting that higher levels of CRTAM, LY9, CD6, ICAM3, SH2D1A, and C1QL2, and lower levels of CNGB3, were associated with a higher frequency of viral blip (VL 40-200) and viremia (VL 200-1000). However, these trends did not reach statistical significance (Supplementary Fig. 7C).
Overall, our findings do not suggest a role for systemic inflammation in the loss of viral control. Instead, our findings point to monocyte (sCD14) and T cell activation as possible factors. Notably, while some of the found biomarkers of viral control loss associated with sCD14 (marker of monocyte activation), sCD14 did not differ between EC and VC and also did not predict the loss of viral control, further suggesting that sustained T cell, rather than monocyte activation, plays a critical role in maintaining HIV control.
Discussion
This study aimed to identify biomarkers and pathways related to spontaneous HIV control and its long-term preservation. HIV controllers comprise a small and heterogeneous group of PLHIV. In the present study, we studied a wide range of HIV controllers that were either non-viremic or exhibited low-level viremia, included from the ATHENA cohort and the 2000HIV study. Of the nearly 2500 plasma proteins investigated, only CRTAM, LY9, CD6, and VAT1 expression differed between ART-naïve EC and VC. In contrast, we did not observe differences in markers of immune exhaustion (e.g., sPD-L1, sCD25) or systemic inflammation (e.g., IL-6, TNF-α) between EC and VC. Indeed, the stratification of the overall proteomic profile based on different plasma viral load cutoff values, including ≤400, ≤2,000, and ≤10,000 c/ml, showed no evident differences on the first and second principal components. This highlights the upregulation of specific T-cell responses that distinguish VC from EC, while the systemic resting proteome is hallmarked by a striking similarity.
During homeostasis, the expression of leukocyte CRTAM is largely restricted ( < 0.5%). However, activation of T-cell or NK-cell activating receptors induces its expression on activated NK cells, NK T cells, CD8+ T lymphocytes, and a minor subset of CD4+ T lymphocytes21,22,23. CRTAM expression on NK cells and CD8+ T lymphocytes promotes cytotoxicity and IFN-γ production, while CRTAM+ CD4+ T- cells may also develop cytotoxic properties24,25. In PLHIV, CRTAM levels are higher in untreated normal progressors than PLHIV on ART, suggesting that increased CRTAM levels are part of an activated T cell response that aims to curb the uncontrolled viral replication26. Indeed, our findings of upregulated CRTAM concentrations in ART-naïve VC, followed by downregulation after ART, strongly indicate that ongoing viral replication in ART-naïve VC contributes to immune activation and higher release of CRTAM.
The Lymphocyte Antigen 9 (LY9, also known as signaling lymphocytic activation molecule family member 3, SLAMF3, or CD229) is widely expressed on T, B and NK-cells and its role in enhancing T-cell activation and differentiation has been supported by studies in LY9-deficient mice27, as well as in several human studies28,29. Mutation in LY9 gene has been shown to increase the risk to develop Progressive Multifocal Leukoencephalopathy (PML) and immune suppression. LY9 is a self-ligand and LY9-LY9 interaction plays a role in the regulation of immune responses30. A soluble form of LY9 is also recognized and used as a biomarker for the severity of multiple myeloma and B cell leukemia and lymphoma31. Interestingly, an in-vitro study showed that LY9 was one of the 9 enriched genes associated with epigenetic modification of HIV-1 proviruses32, and as such limiting the establishment of a latent viral reservoir. Our findings demonstrate, however, that this virus-inhibitory effect is insufficient to prevent loss of HIV control and that, paradoxically, increased plasma LY9 are indicative of deteriorating viral replication.
CD6 is a cell surface receptor involved in signal transduction upon T cell activation, although it is unclear whether CD6 mediates costimulatory or inhibitory signals. CD6 binds not only to the endogenous ligand CD166/ALCAM, but also to microbial associated molecular patterns of bacterial, parasitic and viral origin. CD6 can be cleaved from the membrane during lymphocyte activation and low serum concentrations of soluble CD6 (sCD6) have been reported in healthy subjects and higher concentrations in patients with inflammatory conditions33,34. In-vitro studies also show that sCD6 interferes with HIV-1 entry by blocking the gp120/coreceptor interaction35. Of note, the upregulated protein in EC, VAT1, is an integral membrane protein that regulate neurotransmitter transfer between cytosol and synaptic vesicles, but its relationship with viral control warrants further investigation.36. Collectively, the upregulation of these three proteins in VC may be a “last-stand” attempt to maintain control status by improving cytotoxicity (CRTAM), silencing proviral DNA (LY9), and by inhibiting viral entry (CD6). Nonetheless, the need for such upregulation is a herald of imminent loss of HIV control and these biomarkers may therefore be utilized for their high predictive potential.
Following ART, a downregulation was observed for proteins that are mainly involved in immune related functions. Although these results could theoretically be confounded by the 10-year interval between samples, immune-related proteins have a tendency to increase rather than decrease with progressing age37. Moreover, the analysis of paired ART-naive samples of EC and VC indicated relatively stable protein expression over time and time-related trends in protein expression were not correlated with differential expression following ART. Although we cannot completely exclude a potential influence of other ART-mediated effects, the observation that the type of ART regimen (INSTI-, NNRTI-, PI-based) did not have a major influence renders this possibility less probable. Taken together, it is reasonable to assume that the reduction in relative protein concentrations following ART can be attributed to the suppression of viral replication, resulting in decreased immune activation.
Over up to 17 years follow-up, we observed a loss of controller status (VL > 10,000 copies/mL) especially in viremic controllers (38%) compared to elite controllers (3%). TC and PC exhibited similar host demographic (sex, smoking status, and ethnicity) and HIV related factors (CD4 nadir and CD4 count). Survival analysis indicated a median loss-free follow-up time of 20 years since HIV diagnosis. Furthermore, our study showed the first longitudinal dynamics of viral load in 181 HIV controllers over a 25-year follow-up period since the first HIV-RNA measurement, revealing a gradual shift in viral replication patterns preceding loss of viral control.
Importantly, we found that CRTAM, LY9, and CD6 were consistently associated with an increased risk of virological loss, rising viral load, and CD4 decline years in advance. Currently, no established biomarkers exist for predicting loss of viral control beyond clinical parameters16,38,39. Notably, our predictive models integrating proteomic markers were reliable in forecasting viral control loss up to eight years in advance.
Several hypotheses have been proposed to explain the mechanisms underlying loss of control7,8,39,40. First, “the immune exhaustion hypothesis” suggests that chronic immune activation drives CD8 + T cell dysfunction, leading to loss of control7. Plasma biomarkers associated with this process include sPD-L1, sCD25, IL-10, TGF-β, IFN-γ, and TNF-α. However, none of these markers significantly differed between ECs and VCs or predicted loss of viral control. Secondly, “the systemic inflammation hypothesis” suggests that systemic inflammation plays a major role in maintaining viral control. Our data did not reveal significant differences in key inflammatory proteins such as TNF, IL-1β, IL-6, and sCD14 between EC and VC or in those who later lost viral control, arguing against this inflammation hypothesis. Third, “the viral evolution hypothesis” proposes that HIV mutates over time to escape immune pressure, ultimately leading to loss of control, with which factors representing a high proviral burden are associated8,39,40. Our data show that T cell activation was already elevated several years before control was lost, suggesting that viral escape resulting in loss of control is a slowly progressing process that may have been put in motion years before the plasma viral load surges. Finally, “the loss of effective immune responses hypothesis” proposes that control is lost due to a decline in effective HIV-specific immune responses, particularly those mediated by T cells and NK cells, initially independent from viral evolution39. Our observation of increased levels of T cell-related proteins years before loss of control aligns with this idea, which may go hand in hand with the viral evolution hypothesis.
Overall, our findings highlight the continuous and dynamic interplay between T cell function and viral replication in maintaining HIV control. Given this close relationship, it remains challenging to determine whether T cell activation is a cause or consequence of low-level viral replication. Nonetheless, our study reinforces the predictive value of T cell function-related proteins (CRTAM, LY9, and CD6) in post-treatment control research.
In addition to CRTAM, CD6, and LY9, we discovered distinct proteins linked to an increased (ICAM3, SH2D1A, C1QL2) and decreased risk of losing viral control (CNGB3). Flow-cytometry integration revealed that SH2D1A, CRTAM, and CD6 correlated with cytotoxic/memory activation modules dominated by HLA-DR⁺CD38⁺ CD8⁺ T cells and HLA-DR⁺ γδ T cells, suggesting that escalating antigen-driven cytotoxic activation, rather than failed regulatory control, may precede loss of viral control. While some of these proteins have been reported to play roles in various viral infections, their relationship with HIV infection is poorly known. ICAM3 or CD50 is cell adhesion molecule predominantly expressed in NK cells, monocytes, and other adaptive immune cells. It serves as a ligand for leukocyte functional antigen 1 (LFA-1), thereby facilitating T cell adhesion, activation, and differentiation41,42,43. Additionally, ICAM3 has been implicated in the chemoattraction of phagocytes to apoptotic cells44,45. While earlier studies have reported the involvement of ICAM3 in modulating HIV replication46,47, the precise underlying mechanism remains unclear. SH2D1A, expressed particularly in T cells, as well as NK and NKT cells, encodes SAP (signaling lymphocyte activation molecule (SLAM) family-associated protein)48, which facilitates signal transmission downstream of SLAM family receptors, including LY9. SAP is recognized for its role in controlling T cell cytotoxic function and demonstrating pro-apoptotic activity, thereby contributing to the regulation of T-cell and the removal of potentially harmful DNA-damaged cells49,50. Mutations in SH2D1A in X-linked lymphoproliferative disease result in extreme susceptibility to Epstein-Barr virus infection and severe immune dysregulation51. C1QL2, also referred to as CTRP10, is primarily expressed in the central nervous system, where it is recognized for its role in regulating synaptic network activity within the hippocampus52, although its function in peripheral tissues remains largely unexplored. CNGB3, the sole protein identified as protective against losing viral control, is expressed in the bone marrow, in addition to retinal and sperm cells. Knock-out studies in mice have revealed disruptions in lymph nodes, spleen, and thymus53. The undisclosed functions of C1QL2 and CNGB3 in relation to immunity and viral infection warrant further research.
The consistent connection between specific immune proteins and viral load dynamics in HIV controllers indicate that the plasma proteome is subject to an equilibrium between host immune and viral factors As an exploratory extension of these findings, we examined plasma RNA genomes in viremic controllers, which provides insights into the intactness and diversity of viral sequences in this group. We show the presence of genetically-intact plasma RNA genomes in VC, however, a large proportion of plasma-derived sequences were genetically-defective in 5’-half and 3’-half in the VC (Supplementary Fig. 8A; and Supplementary Data 10). No defects were observed in the packaging signal of plasma viruses, affirming the critical role it plays in selectively packaging viral RNA during the process of virus assembly54.
The genetic diversity analysis showed that VC had very few genetically-identical sequences, mainly confined to a singular clonal cluster. This exploratory analysis also revealed a high level of genetic diversity among plasma RNA sequences in all four VC (Supplementary Fig. 8B; and Supplementary Data 10). The increased diversity could stem from the plasma being collected during the chronic phase for these participants, which is found in other studies as well55,56. This diversity suggests a complex interplay between viral replication, host immunity, and selective pressures. Furthermore, this diversity suggests multiple instances of viral release in VC, primarily characterized by the presence of replication-deficient viruses originating from diverse cellular reservoirs harboring distinct proviruses. This underscores the ongoing state of interplay between active viral reservoirs and the host’s immune defense mechanisms in VC.
This study has also some limitations. Given that the plasma proteome originates from various tissues, targeted plasma proteomics may not be sensitive enough to detect subtle biological differences expressed in a single tissue or organ system. Second, variations in storage time between the elite-ATHENA and the 2000HIV studies could have potentially influenced the results. The 2000HIV cohort participants were enrolled more recently, whereas the elite-ATHENA samples had been stored for a comparatively longer duration. To minimize this potential bias, we adjusted all analyses for age, sex, recruitment center, and storage time in the elite-ATHENA cohort. The consistent findings observed across both EC and VC cohorts support the robustness of our results. Third, we acknowledge age differences when comparing pre- and post-ART samples in persistent and transient EC and VC. To address this, we again adjusted our analyses for age, sex, recruitment center, and storage time. Consistently, within-pair age-matched comparisons confirmed similar results. Finally, multiple testing corrections become stringent in such a large proteomics library, and the relatively limited number of HIV controllers may have affected the sensitivity to detect relevant molecules among the many proteins that were assessed. It is important to recognize, however, that HIV controllers and especially EC are rare phenotypes within PLHIV, with many studies including significantly smaller number of participants compared to our analysis.
In conclusion, loss of control was common in viremic controllers (38%) in contrast to elite controllers (3%) during 17 years follow up. A gradual shift in viral replication patterns preceding loss of viral control. Predictive models integrating proteomic markers achieved near-perfect accuracy in forecasting viral control loss up to eight years in advance. Three T cell-associated proteins, CRTAM, LY9, and CD6, predicted viral loss but were also consistently elevated in VC compared to EC among more than 3,000 measured proteins. In contrast, markers of immune exhaustion (e.g., sPD-L1, sCD25) and systemic inflammation (e.g., IL-6, TNF-α) did not differ between EC and VC, nor did they predict control loss, reinforcing the distinct role of CRTAM, LY9, and CD6 in this process. The presence of genetically-intact and defective plasma RNA genomes in VC, along with a high level of genetic diversity, suggesting ongoing immune-driven viral control mechanisms. Overall, our findings highlight the continuous and dynamic interplay between T cell function and viral replication in maintaining HIV control and generated a prediction model for those that will lose control over time. Ultimately, this knowledge may guide the development of future immune-based interventions aimed at achieving durable drug-free viral suppression.
Methods
Study populations
Participants were all spontaneous HIV-1 controllers and recruited from two cohorts: the ATHENA cohort and the 2000HIV cohort (Fig. 1A).
Participants from the ATHENA cohort
Since 1998, PLHIV receiving care at a Dutch HIV treatment center have been invited to join the Dutch National ATHENA cohort, which comprises all 24 designated HIV treatment center in the Netherlands57. ATHENA is an open observational cohort, with continuous recruitment of newly diagnosed individuals entering HIV care. The ATHENA cohort collects a comprehensive dataset on HIV treatment, routine laboratory and other diagnostic tests, comorbidities, adverse events, and coinfections. Sex was determined based on self-report, as documented in the medical records. Currently, the Dutch National ATHENA cohort encompasses more than 95% of all PLHIV in the Netherlands. This initiative involves storing residual plasma samples left after routine viral load measurements.
For this study, we gathered historical plasma samples collected prior to the COVID-19 pandemic from six participating centers (Erasmus MC Rotterdam [EMC], OLVG Amsterdam [OLVG], LUMC Leiden [LUMC], Masstad Ziekenhuis Rotterdam [MZS], UMC Utrecht [UMCU], Radboudumc Nijmegen [RAD]), constituting what we refer to as the elite-ATHENA cohort. A total of 183 HIV controllers were selected for the elite-ATHENA cohort, consisting of 36 EC and 147 VC. All participants were fulfilling the criteria for EC or VC status (HIV controllers; see case definition below) at the time of blood collection (for details see Fig. 1A). The date of sample collection after the establishment of HIV controller setpoint was considered the baseline time point (day 0). The plasma samples obtained from these individuals were utilized as the discovery samples for the proteomic analysis.
HIV controllers were followed from the baseline time point (day 0) up to June 2022. Follow-up was ended when HIV controller status was lost or censored when ART was started. Up to 17 years follow-up data after baseline time point was available for 181 out of the total 183 participants. Within this subset, 125 individuals exhibited persistent controller status, and 56 transient controller status.
A second stored blood sample was available from 102 out of 181 participants after ART initiation. Among them, 57 VC and 9 EC still maintained their HIV controller status (VL < 10,000 copies/mL), while 36 VC lost their controller status (developed (VL > 10,000 copies/mL). Blood samples were stored at −80 degrees Celsius on the same day of routine viral load measurement until they were sent to Radboudumc for further processing.
Participants from the 2000HIV human functional genomics partnership program (2000HIV cohort)
In the 2000HIV cohort, asymptomatic PLHIV without signs or symptoms of an acute condition were included between 2019 and 2021, originating from the following four Dutch HIV treatment centers: Erasmus MC Rotterdam, OLVG Amsterdam, Tweesteden ziekenhuis Tilburg, Radboudumc Nijmegen6. As there was significant overlap between ATHENA and 2000HIV participants, we opted to choose independent samples from ART naïve individuals that belonged to the subgroup of persistent HIV controllers. Specifically, we selected 5 VC and 14 EC from the 2000HIV cohort. Sex was determined based on self-report and further confirmed using available genetic data from the cohort. Blood samples were transported to Radboudumc and stored at −80 degrees Celsius until further processing.
Case definitions
HIV controllers were defined as fulfilling criteria of the elite or viremic controller definitions or both. Elite controllers (EC) were non-viremic PLHIV with plasma HIV-RNA ≤ 50 copies/mL on three separate measurements during a minimal period of 12 months and no ART use during these measurements (isolated viral blips of 50–1000 copies/mL were allowed). Viremic controllers (VC) were ART naïve PLHIV with plasma HIV-RNA > 50 copies/mL but < 10,000 copies/mL for minimal 5 years on three separate measurements. To study the full spectrum of VC as well as stratify by various VC viral load cutoff values, we chose the rather lenient limit of < 10,000 copies/mL, together with a more stringent duration criterion5.
Throughout the follow-up period, certain individuals lost their controller status and were subsequently classified as transient HIV controllers. This classification was applied to those whose plasma HIV-RNA levels exceeded 10,000 copies/mL after initially meeting the criteria for HIV controllers. In contrast, persistent HIV controllers were defined as individuals who maintained HIV-RNA levels below 10,000 copies/mL throughout the entire ART-naive follow-up period.
High throughput targeted proteomics measurement and quality control
Proximity extension assay was used to measure the full library of proteins (Olink® panel Explore 3072, Uppsala, Sweden) as described elsewhere18,58. In brief, in this targeted proteomics technique oligonucleotide-labeled antibodies bind the proteins and subsequently hybridize. The oligonucleotide sequence is polymerized and quantitatively measured with real-time qPCR or Next Generation Sequencing (NGS), which is proportional to the plasma protein concentrations. The normalized expression values of 2925 unique proteins were obtained from the elite-ATHENA cohort, which consisted of 188 samples. Additionally, 2914 unique proteins were available from the 19 samples in the 2000HIV cohort. Protein concentration was expressed as normalized expression values that were adjusted to a log2 scale after subtraction of the proximity extension control and plate control values.
To maintain data quality, a further quality control step was implemented. Proteins for which more than 25% of the total measurements fell below the lower limit of detection were excluded from the discovery cohort (elite-ATHENA cohort), resulting in the removal of 505 proteins from data analysis. The protein selection criteria applied to the discovery cohort were subsequently applied to the validation cohort. This led to the inclusion of 2420 proteins from the elite-ATHENA cohort and 2409 proteins from the 2000HIV cohort for downstream data analysis (n = 11 proteins in elite-ATHENA were not present in the 2000HIV cohort due to technical issue).
Identification of outliers was performed using principal component analyses (PCA). Data points that deviated more than 4 standard deviations from the mean of principal component 1 (PC1) and/or PC2 were excluded from further analyses. Consequently, 5 participants from the elite-ATHENA cohort were excluded based on this criterion, while none were excluded from the 2000HIV cohort. Thus, final matrix of 2420 proteins from 183 participants in the elite-ATHENA cohort and 2409 proteins from 19 participants in the 2000HIV cohort were available for subsequent data analysis.
Flow cytometric immunophenotyping
Blood samples of EC and VC of the 2000HIV cohort were immunophenotyped using a 21-color, six-laser CytoFLEX-LX flow cytometer (Beckman Coulter)59. Daily quality control and standardization were performed using CytoFLEX Daily QC Fluorospheres, CytoFLEX Daily IR QC Fluorospheres, and SPHERO Rainbow calibration particles. Data were collected in CytExpert v2.3 and analyzed in Kaluza v2.1.2 via sequential gating to resolve 355 distinct populations. From these, we selected 58 populations co-expressing HLA-DR and/or CD38, including CD4⁺ and CD8⁺ T-cell subsets, NK cells, γδ-T cells, and regulatory T cells, to quantify activation states. Antibody selection has been described previously in detail59.
Plasma sequencing
Stored plasma from four viremic controllers in the elite-ATHENA cohort was sequenced. Viral loads ranged from 5,370 to 8,711 copies/mL, and all samples were collected during chronic infection at a median of 4.97 years post-infection (range 3.42–6.27 years). Notably, two of the participants experienced loss of control within one year from the sampling date. Detailed participant characteristics are provided in Supplementary Data 10.
RNA extraction from plasma
Up to 1 mL plasma was centrifuged at 2500 rpm for 10 min at 4 °C to sedimentate any cellular debris present. The resulting supernatant was subsequently centrifugated at 14,000 rpm for 60 minutes at 4 °C. The pelleted virions were resuspended in 180 µL of Tris-HCl. Viral RNA was extracted using the QIAmp Viral RNA Mini Kit (Qiagen, #52904), according to the manufacturer’s instructions. The extracted RNA was then eluted in 60 µL of AVE buffer.
cDNA synthesis
The extracted RNA was divided into two equal parts. One half was employed for the generation of complementary DNA (cDNA) for 5’-half sequencing, utilizing the R5968 primer (5’-TGTCTYCKCTTCTTCCTGCCATAG-3’, primers listed in Supplementary Data 1). Meanwhile, the other half was used for cDNA synthesis for 3’-half sequencing, employing the primer R9665 (5’-GTCTGAGGGATCTCTAGWTACCAGA-3’). Two master mixes were prepared for cDNA synthesis and subsequent PCR amplification. Mix 1 (29.75 μL) was composed of 25 μL RNA, 2.5 μL 10 mM dNTP, 1.25 μL 20 μM oligo-dT and 0.5 μL 50 μM primer. Mix 2 (20.25 μL) included 2.75 μL UltraPure DNase/RNAse-free water (Thermo Fisher, #10977023), 5X RT buffer (Invitrogen, # M0681S), 2.5 μL 100 mM DTT, 2.5 μL 40U/μL RNAse inhibitor (Takara, #2313B), and 2.5 μL Induro® Reverse Transcriptase (Invitrogen, # M0681S). Mix 1 underwent heating to 65 °C for 5 minutes and was then rapidly cooled on ice for at least 2 minutes. Subsequently, mix 2 was pre-warmed to 50 °C and added to the chilled Mix 1. The resulting mixture was incubated at 55 °C for 45 minutes, followed by an inactivation step at 70 °C for 10 minutes. Finally, 2 μL of RNAse H (Invitrogen, #18080051) was added, and the mixture was incubated for an additional 20 minutes at 37 °C.
PCR amplification for half-genome amplification
The resulting cDNA served as a template for viral half-genome long-range PCRs. The cDNA was diluted to ensure the presence of single HIV-1 copy per reaction. The first-round PCR mix (10 μL) included 2 μL 5X PrimeSTAR GXL buffer, 0.8 μL of 2.5 mM dNTPs, 0.2 μL PrimeSTAR GXL polymerase (Takara, #R050B), 250 nM forward and reverse primers, and 1 μL diluted cDNA. The second-round PCR mix (25 μL) consisted of 5 μL 5X PrimeSTAR GXL buffer, 2 μL of 2.5 mM dNTPs, 0.5 μL PrimeSTAR GXL polymerase (Takara, #R050B), 250 nM forward and reverse primers, and 1 μL of the first-round product (Supplementary Data 11 for primer sequences). The cycling conditions for both PCR rounds were 2 minutes at 98 °C; 35 cycles (10 seconds at 98 °C, 15 seconds at 62 °C, 5 minutes at 68 °C); 7 minutes at 68 °C. The 5’-half amplifies a region of 5.2 kb (HXB2 coordinates 581-5783), while the 3’-half amplifies a region of 4.5 kb (HXB2 coordinates 5088-9602). Negative reactions without reverse transcriptase ensured the absence of DNA contamination in the RNA extracts. PCR products were assessed on a 1% agarose gel, and positive results were subsequently subjected to AVITI sequencing.
Half-genome sequencing
Amplified cDNA from positive wells were cleaned using AMPure XP beads (Beckman Coulter, #A63880), followed by quantification of each cleaned amplified product with Quant-iT PicoGreen dsDNA Assay Kit (Invitrogen, #P11496). Next, an NGS library preparation using the Nextera XT DNA Library Preparation Kit (Illumina, #FC-131-1096) with indexing of 96 samples per run was used according to the manufacturer’s instructions (Illumina, #FC-131-2001), except that input and reagents volumes were halved and libraries were normalized manually. The pooled library was sequenced on an AVITI platform via 2×150 nt paired-end sequencing using the AVITI 2×150 Sequencing Kit Cloudbreak Low Output kit (Element Biosciences, #860-00005).
Assembly
The sequencing data that had been generated underwent demultiplexing and was then employed for de novo assembly. The code used to carry out de novo assembly can be accessed on the following GitHub page: (https://github.com/laulambr/virus_assembly). In brief, the workflow comprised the following steps: (1) scrutiny of sequencing quality for each library using FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc) and the removal of adaptor sequences, along with trimming of 5′ and −3′ terminal ends using BBtools v37.99 (sourceforge.net/projects/bbmap/); (2) de novo assembly of the trimmed reads using MEGAHIT v1.2.9, resulting in contigs for each library; (3) validation of all de novo contigs per library using blastn v2.7.1 against the HXB2 reference virus, acting as a filter to exclude non-HIV-1 contigs in the subsequent analysis steps; (4) thereafter, the trimmed reads were mapped against the de novo assembled HIV-1 contigs to facilitate the determination of the final majority consensus sequence of each sequence using bbmap v37.9960.
Statistics
Demographic characteristics assessment
Demographic characteristics of the participants were assessed using Chi-square test and Wilcoxon rank sum test for categorical and numerical variables, respectively.
Differential expression analysis of plasma proteins between ART naïve EC and VC
We analyzed differences in proteome between two groups, ART-naïve elite controllers (EC) and viremic controllers (VC), in two separate sets of data: a discovery cohort (elite-ATHENA cohort) consisting of 147 VC and 36 EC, and an independent validation cohort (2000HIV cohort) with 5 VC and 14 EC participants. We employed linear models using the “limma” R package to compare log2-normalized protein relative expression, accounting for relevant confounding variables. To identify these covariates, we performed linear regression on the first five principal components derived from PCA analysis of 2420 proteins. The results were displayed as a heatmap showing the explained variance (adjusted R2) of the first 10 PCs by each covariate (Supplementary Fig. 1A-B). We selected as potential confounding variables those that when adding to the linear model in a multi-step approach resulted in a 10% increase in beta coefficients. This process identified age, sex, center, and storage time as potential confounders in the discovery cohort.
Findings between VC and EC from the discovery cohort were validated in the validation cohort. We performed linear regression of the proteins that showed significance in the discovery cohort. Given the limited number of participants, only age and sex were considered as potential confounders during the analysis. We considered proteins as differentially expressed if their FDR-corrected P value was below 0.05 in the discovery cohort and their P-value was below 0.05 in the validation cohort.
Differential expression analysis of plasma proteins before and after ART in transient and persistent VC and persistent EC
Over up to 17 years follow-up, ART was initiated in a subset of HIV controllers due to several factors, such as patient preference, preventing HIV transmission, loss of virological control or adherence to new HIV treatment guidelines introduced in 2015, which recommended ART for all PLHIV regardless of viral load levels or CD4 counts. To assess changes in protein expression related to ART administration, we performed paired comparison before and after ART administration in 36 transient VC, 57 persistent VC, and 9 persistent EC from the elite-ATHENA cohort. In addition, we analyzed paired ART-naive samples over time, including 23 paired samples of persistent VC and 19 persistent EC.
To analyze the differences, we employed linear mixed models using the “nlme” R package, correcting for age, sex, center, and storage time. These covariates were identified through a methodology similar to that used in comparison one (Supplementary Fig. 1C, D). Proteins were considered differentially expressed if their FDR-corrected P value was less than 0.05.
Survival analysis of loss of controller status in persistent, ART naïve, HIV controllers at baseline
After identification of 56 TC and 125 PC, the dynamics of viral load between PC and TC was examined by visualizing the yearly changes in viral load since the first available plasma HIV RNA measurement through scatter and line plots. Next, to determine the duration of loss-free survival time, which represents the period during which HIV controllers maintain their viremic controller status, we conducted a survival analysis utilizing the ‘survival’ and ‘survminer’ R packages. In this analysis, reaching viral load >10,000 copies/mL was considered as ‘event’, while follow-up was censored at ART initiation, loss of follow-up or the censoring date June 1, 2022. We visualized the survival patterns through Kaplan-Meier plots.
Furthermore, to investigate which proteins relate to viral control maintenance, we examined whether the baseline plasma protein expression in ART-naïve individuals was associated with the time to virological loss during the follow-up period (reaching viral load >10,000 copies/mL). We performed Cox regression using the R functions ‘coxph’ and ‘surv’, taking into account relevant confounding factors such as age, sex, center, and storage time. These confounders were identified through a similar approach as in comparison one (Supplementary Fig. 1A-B). Of note, given lack of female TC in ‘RAD’, ‘MSZ’, and ‘LUMC’ centers, center was added to the model as a categorical variable, with categories ‘EMC’, ‘OLVG’, and ‘UMCU’. This results in the following Cox regression formula (1):
‘Where followup_time’ represents the time in days from baseline time point until event or censoring (numerical: days); ‘Loss_status’ represents the occurrence of virological loss during follow-up (categorical: 1/0 to represent yes/no); ‘Protein_level’ represents relative expression of one protein out of total 2420 proteins (numerical: NPX value); ‘Age’ represents age of individual at the time of sample collection (numerical: years); ‘Sex’ represents sex of individual (categorical: female/male); ‘Storage_time’ represents time from sample collection until protein measurement (numerical: days); center correction represents whether individual enrolled in ‘EMC’, ‘OLVG’, or ‘UMCU’ (categorical: yes/no). The results of Cox regression analysis were presented in terms of hazard ratios (HR). A hazard ratio higher than one indicates that the protein is associated with higher risk of losing viral control. Proteins were considered significant if their FDR-corrected P value was less than 0.05.
Similarly, we employed Cox regression to predict a substantial increase in plasma viral load viral load, analysis, integrating age, sex, center, and storage time as covariates. A substantial increase in viral load was defined as either a surge of over 1000 copies/ml of the moving median viral load at proteomic sample inclusion or when viral load reached 10,000 copies/ml. The moving median viral load was computed within a rolling one-year timeframe before and after the HIV-RNA measurement date with the ‘slide_index_dbl’ function from R package ‘slider’. This approach ensured that the moving median viral load remained unaffected by isolated peaks and adapted to the varying frequency of viral load measurements.
Finally, we analyzed dynamic changes in CD4 count after the proteomics sample over the years. Using Cox regression analysis, we evaluated which proteins were associated with the risk of the moving median CD4 count decreasing to levels below 500 cells/mm3. Covariates were age, sex, center, and storage time. This prediction was performed in a subset of participants with available longitudinal CD4 data and whose moving median CD4 count at the time of the proteomics sample exceeded 500 cells/mm3 (n = 138).
Pathway analysis
The differentially expressed proteins were used as an input and all measured plasma proteins as a background to test for pathway-enrichment in GO Biological processes, KEGG, and Reactome databases using DAVID61 and visualized as a network with Cytoscape version 3.9.162,63.
Prediction models
We developed predictive models to assess the risk of viral control loss, substantial increases in viral load, and CD4 count decline in HIV controllers. With R Package “RiskRegression”, the models were evaluated using the area under the receiver operating characteristic curve (AUC), where AUC values of 0.5 represent random prediction and 1.0 indicates perfect accuracy. Models were simple Cox proportional hazards regression, random survival forest, elastic net (penalized) Cox regression and gradient-boosted machines for cox-distributions, using the R packages “Survival”, “rsfsrc”, “glm”, “gbm”, respectively. Clinical models were trained using data from 839 HIV controllers in the ATHENA cohort described elsewhere20 and evaluated on the 181 HIV controllers used for proteomic analyses as described above. Proteomic prediction models included both clinical predictors and candidate proteins (nominal P < 0.05), training on a cohort of 134 HIV controllers from two centers (OLVG and EMC), and testing on 47 HIV controllers from the other centers (UMCU, RUMC, LUMC, MSZ). The code for the analyses can be found on Github64.
Analysis of sequences
The analysis involved aligning participant sequences using the MAFFT tool65. Genetically-defective genomes were characterized by the presence of inversions, deletions exceeding 100 base pairs, APOBEC3G-induced hypermutation, point mutations leading to stop codons, or indels resulting in frameshift mutations within HIV-1 open reading frames (ORFs). Additionally, the cis-acting region, including the packaging signal and major splice donor (MSD), underwent evaluation. Defects within ORFs were identified using Los Alamos database tools, namely GeneCutter (https://www.hiv.lanl.gov/content/sequence/GENE_CUTTER/cutter.html), Hypermut (https://www.hiv.lanl.gov/content/sequence/HYPERMUT/hypermut.html) and National Cancer Institute (NCI) Proviral Sequence Database (PSD) Proviral Sequence Annotation & Intactness Test (https://psd.cancer.gov/tools/tool_index.php). Subtype and co-receptor tropism was determined using REGA HIV subtyping tool (https://www.genomedetective.com/app/typingtool/hiv) and Geno2pheno [coreceptor] 2.5 (https://coreceptor.geno2pheno.org/) with 10% FPR respectively.
Phylogenetic analysis
Phylogenetic trees were constructed from alignments using FastTree version 2.1.11. R (v.4.1.2). ggplot2 (v.3.5.0), and ggtree (v.3.10.1) were used for visualization and annotation of trees. To identify genetically-identical sequences, the Los Alamos tool ElimDupes (https://www.hiv.lanl.gov/content/sequence/elimdupesv2/elimdupes.html) was utilized. A threshold of 99.97% for the 5’-half and 99.98% for the 3’-half was selected to permit a difference of only one nucleotide (nt) between genomes, allowing for potential changes during a single round of viral replication. This decision was made to accommodate any changes that might occur during a single round of viral replication. DIVEIN was used to calculate pairwise diversity among genetically-intact sequences.
Ethics
Both discovery and validation cohort studies were approved by local ethical committees (2000HIV NL68056.091.18; clinicaltrials.gov NCT03994835, and ATHENA sample collection MEC-2020-0871). All participants provided informed consent as part of the ATHENA and/or 2000HIV cohort and the studies were conducted according to the declaration of Helsinki and the guidelines for Good Clinical Practice.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study, including pseudonymized participant metadata, plasma proteome, and flow cytometry data of the 2000HIV cohort, are available through restricted-access repositories in the Radboud Data Repository (https://data.ru.nl; https://doi.org/10.34973/p96d-kz55; https://doi.org/10.34973/qk29-f305). Proteomics data and metadata from the ATHENA cohort are not publicly available due to privacy considerations related to participant re-identification. These data are archived at the HIV Monitoring Foundation (Amsterdam) and Radboud University Medical Center (Nijmegen), The Netherlands, and can be shared upon request through datarequest.aig@radboudumc.nl. No consent was obtained for publishing information that potentially identifies the individuals. Data was anonymized were applicable. Viral sequences have been deposited in GenBank (NIH genetic sequence database) under accession codes PV545640–PV545908. Source data are provided with this paper.
Code availability
All analysis scripts used in this study are available at (https://github.com/albert-ml-ai/HIV_controllers_ATHENA). The code at time of publication was archived at https://doi.org/10.5281/zenodo.1778241064.
References
Gurdasani, D. et al. A systematic review of definitions of extreme phenotypes of HIV control and progression. AIDS 28, 149–162 (2014).
Plaçais, L. et al. Antiretroviral therapy for HIV controllers: Reasons for initiation and outcomes in the French ANRS-CO21 CODEX cohort. eClinicalMedicine 37, 100963 (2021).
Brusca, R. et al. Subclinical cardiovascular disease in HIV controller and long-term nonprogressor populations. HIV Med. 21, 217–227 (2020).
Dominguez-Molina, B. et al. Analysis of non-AIDS-defining events in HIV controllers. Clin. Infect. Dis. 62, 1304–1309 (2016).
Capa, L. et al. Elite controllers long-term non progressors present improved survival and slower disease progression. Sci. Rep. 12, 16356 (2022).
Vos, W. et al. The 2000HIV study: Design, multi-omics methods and participant characteristics. Front Immunol. 13, 982746 (2022).
Rosás-Umbert, M. et al., Mechanisms of abrupt loss of virus control in a cohort of previous HIV controllers. J. Virol. 93, e01436-18 (2019).
Gasca-Capote, C., et al., The HIV-1 reservoir landscape in persistent elite controllers and transient elite controllers. J. Clin. Invest. 134, e174215 (2024).
Hartana, C. A. & Yu, X. G. Immunological effector mechanisms in HIV-1 elite controllers. Current Opinion in HIV and AIDS 16, 243–248 (2021).
Jiang, C. et al. Distinct viral reservoirs in individuals with spontaneous control of HIV-1. Nature 585, 261–267 (2020).
Boritz, E. A. et al. Multiple origins of virus persistence during natural control of HIV infection. Cell 166, 1004–1015 (2016).
Lian, X. et al. Signatures of immune selection in intact and defective proviruses distinguish HIV-1 elite controllers. Sci. Transl. Med 13, eabl4097 (2021).
Gasca-Capote, C. et al. The HIV-1 reservoir landscape in persistent elite controllers and transient elite controllers. J. Clin. Invest. 134, e174215 (2024).
Omondi, F. H. et al. HIV proviral burden, genetic diversity, and dynamics in viremic controllers who subsequently initiated suppressive antiretroviral therapy. mBio 12, e02490-21 (2021).
Pérez-Yanes, S. et al. The characteristics of the HIV-1 Env glycoprotein are linked with viral pathogenesis. Front. Microbiol. 13, 763039 (2022).
Rodríguez-Gallego, E. et al. Proteomic profile associated with loss of spontaneous human immunodeficiency virus type 1 elite control. J. Infect. Dis. 219, 867–876 (2018).
Gutiérrez-Rivas, M. et al., High plasma levels of sTNF-R1 and CCL11 are related to CD4+ T-cells fall in human immunodeficiency virus elite controllers with a sustained virologic control. Front Immunol. 18, 1399 (2018).
Filbin, M. R. et al. Longitudinal proteomic analysis of severe COVID-19 reveals survival-associated signatures, tissue-specific cell death, and cell-cell interactions. Cell Rep. Med. 2, 100287 (2021).
Zhang, Z. et al. Intestinal homeostasis and inflammation: Gut microbiota at the crossroads of pancreas–intestinal barrier axis. Eur. J. Immunol. 52, 1035–1046 (2022).
Groenendijk, A. L. et al. The risk of non-AIDS defining events is lower in ART-naive HIV controllers than in normal progressors on suppressive ART. Clin Infect Dis. 80, 585–593 (2024).
Patiño-Lopez, G. et al. Human class-I restricted T cell associated molecule is highly expressed in the cerebellum and is a marker for activated NKT and CD8+ T lymphocytes. J. Neuroimmunol. 171, 145–155 (2006).
Boles, K. S. et al. The tumor suppressor TSLC1/NECL-2 triggers NK-cell and CD8+ T-cell responses through the cell-surface receptor CRTAM. Blood 106, 779–786 (2005).
Class I-Restricted T Cell-Associated Molecule Is a Marker for IFN-γ-Producing iNKT Cells in Healthy Subjects and Patients with Type 1 Diabetes. J. Interferon & Cytokine Res. 37, 39–49 (2017).
Takeuchi, A. et al. CRTAM determines the CD4+ cytotoxic T lymphocyte lineage. J. Exp. Med 213, 123–138 (2016).
Patil, V. S. et al. Precursors of human CD4(+) cytotoxic T lymphocytes identified by single-cell transcriptome analysis. Sci. Immunol. 3, eaan8664 (2018).
Sperk, M. et al. Plasma soluble factor following two decades prolonged suppressive antiretroviral therapy in HIV-1-positive males: A cross-sectional study. Med. (Baltim.) 97, e9759 (2018).
Graham, D. B. et al. Ly9 (CD229)-deficient mice exhibit T cell defects yet do not share several phenotypic characteristics associated with SLAM- and SAP-deficient mice. J. Immunol. 176, 291–300 (2006).
Comte, D. et al. Engagement of SLAMF3 enhances CD4+ T-cell sensitivity to IL-2 and favors regulatory T-cell polarization in systemic lupus erythematosus. Proc. Natl. Acad. Sci. USA 113, 9321–9326 (2016).
Zhou, T. et al. Upregulation of SLAMF3 on human T cells is induced by palmitic acid through the STAT5-PI3K/Akt pathway and features the chronic inflammatory profiles of type 2 diabetes. Cell Death Dis. 10, 559 (2019).
Romero, X. et al. CD229 (Ly9) lymphocyte cell surface receptor interacts homophilically through its N-terminal domain and relocalizes to the immunological synapse. J. Immunol. 174, 7033–7042 (2005).
Roncador, G. et al. CD229 (Ly9), a novel biomarker for B-cell malignancies and multiple myeloma. Cancers (Basel), 14, 2154 (2022).
Irwan, I. D., Bogerd, H. P. & Cullen, B. R. Epigenetic silencing by the SMC5/6 complex mediates HIV-1 latency. Nat. Microbiol 7, 2101–2113 (2022).
Ramos-Casals, M. et al. High circulating levels of soluble scavenger receptors (sCD5 and sCD6) in patients with primary Sjögren’s syndrome. Rheumatol. (Oxf.) 40, 1056–1059 (2001).
Aibar, J. et al. Pattern of soluble CD5 and CD6 lymphocyte receptors in critically ill patients with septic syndromes. J. Crit. Care 30, 914–919 (2015).
A Role for Scavenger-like Lymphocyte Receptor CD6 in HIV-1 Viral Infection. AIDS Research and Human Retroviruses, 30, A49–A50 (2014).
Linial, M. & Levius, O. VAT-1 from Torpedo is a membranous homologue of zeta crystallin. FEBS Lett. 315, 91–94 (1993).
Franceschi, C. et al. Inflammaging: a new immune–metabolic viewpoint for age-related diseases. Nat. Rev. Endocrinol. 14, 576–590 (2018).
Masip, J. et al. Elevated α-ketoglutaric acid concentrations and a lipid-balanced signature are the key factors in long-term HIV control. Front Immunol. 13, 822272 (2022).
Pernas, M. et al. Factors leading to the loss of natural elite control of HIV-1 infection. J. Virol. 92, e01805-17 (2018).
Noel, N. et al. Immunologic and virologic progression in hiv controllers: the role of viral “blips” and immune activation in the ANRS CO21 CODEX study. PLoS One 10, e0131922 (2015).
Campanero, M. R. et al. ICAM-3 interacts with LFA-1 and regulates the LFA-1/ICAM-1 cell adhesion pathway. J. Cell Biol. 123, 1007–1016 (1993).
Bleijs, D. A. et al. Co-stimulation of T cells results in distinct IL-10 and TNF-alpha cytokine profiles dependent on binding to ICAM-1, ICAM-2 or ICAM-3. Eur. J. Immunol. 29, 2248–2258 (1999).
Montoya, M. C. et al. Role of ICAM-3 in the initial interaction of T lymphocytes and APCs. Nat. Immunol. 3, 159–168 (2002).
Kristóf, E. et al. Novel role of ICAM3 and LFA-1 in the clearance of apoptotic neutrophils by human macrophages. Apoptosis 18, 1235–1251 (2013).
Torr, E. E. et al. Apoptotic cell-derived ICAM-3 promotes both macrophage chemoattraction to and tethering of apoptotic cells. Cell Death Differ. 19, 671–679 (2012).
Barat, C., Gervais, P. & Tremblay, M. J. Engagement of ICAM-3 provides a costimulatory signal for human immunodeficiency virus type 1 replication in both activated and quiescent CD4+ T lymphocytes: implications for virus pathogenesis. J. Virol. 78, 6692–6697 (2004).
Biggins, J. E. et al. ICAM-3 influences human immunodeficiency virus type 1 replication in CD4+ T cells independent of DC-SIGN-mediated transmission. Virology 364, 383–394 (2007).
Nagy, N. et al. SH2D1A and SLAM protein expression in human lymphocytes and derived cell lines. Int J. Cancer 88, 439–447 (2000).
Nagy, N. et al. The proapoptotic function of SAP provides a clue to the clinical picture of X-linked lymphoproliferative disease. Proc. Natl. Acad. Sci. 106, 11966–11971 (2009).
Sharifi, R. et al. SAP mediates specific cytotoxic T-cell functions in X-linked lymphoproliferative disease. Blood 103, 3821–3827 (2004).
Sayos, J. et al. The X-linked lymphoproliferative-disease gene product SAP regulates signals induced through the co-receptor SLAM. Nature 395, 462–469 (1998).
Matsuda, K. et al. Transsynaptic modulation of kainate receptor functions by C1q-like proteins. Neuron 90, 752–767 (2016).
Groza, T. et al. The International mouse phenotyping consortium: comprehensive knockout phenotyping underpinning the study of human disease. Nucleic Acids Res. 51, D1038–D1045 (2022).
Rein, A. The heart of the HIV RNA packaging signal?. Proc. Natl. Acad. Sci. USA 117, 19621–19623 (2020).
Fisher, K. et al. Plasma-derived HIV-1 virions contain considerable levels of defective genomes. J. Virol. 96, e0201121 (2022).
Maldarelli, F. et al. HIV populations are large and accumulate high genetic diversity in a nonlinear fashion. J. Virol. 87, 10313–10323 (2013).
Boender, T. S. et al. AIDS Therapy Evaluation in the Netherlands (ATHENA) national observational HIV cohort: cohort profile. BMJ Open 8, e022516 (2018).
Koprulu, M. et al. Proteogenomic links to human metabolic diseases. Nat. Metab. 5, 516–528 (2023).
Navas, A. et al. Comprehensive phenotyping of circulating immune cell subsets in people living with HIV. J. Immunol. Methods 507, 113307 (2022).
Cole, B. et al. Extensive characterization of HIV-1 reservoirs reveals links to plasma viremia before and during analytical treatment interruption. Cell Rep. 39, 110739 (2022).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Kucera, M. et al. AutoAnnotate: a cytoscape app for summarizing networks with semantic annotations. F1000Research 5, 1717 (2016).
Merico, D. et al. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS ONE 5, e13984 (2010).
Groenendijk, A. L. HIV_controller_ATHENA. Github, (2025).
Katoh, K. et al. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30, 3059–3066 (2002).
Acknowledgements
The authors thank all volunteers in the ATHENA cohort and in the Human Functional Genomics Projects for their participation. We thank H. Maas, M. Ostman, F. Weren, P. van Kempen for help in laboratory analysis. For their help in sample collection, we thank dr. D. Kwa, dr. W.L. Blok, dr. J.E. Stalenhoef from OLVG Amsterdam; dr. J.J. van Kampen, R.J. Gruters, R.J. Overmars from Erasmus MC Rotterdam; A. Gottenkieny from UMC Utrecht; dr. J.G. den Hollander en dr. C. Tienen from Maasstad Ziekenhuis Rotterdam; dr. F. Stelma and P. Daemen from Radboudumc Nijmegen; dr. A.H.E. Roukens, N. Holten, W. Dorama from LUMC Leiden.
Author information
Authors and Affiliations
Contributions
N.V., A.G., K.M., L.V.,and A.v.d.V. wrote the manuscript; N.V., A.G., J.C.S., F.W., W.V., M.B., L.E., K.M., L.L., S.R. and V.M. contributed to data collection and analysis; M.G.N., A.v.d.V., A.V., A.G., J.v.L. and L.A.B.J. contributed to the study design; E.J.N., C.J.X., L.A.B.J., Q.d.M., V.M., J.v.L., C.R., A.V., M.G.N., L.L., S.R., L.V. and A.v.d.V. supervised the work; All authors contributed to manuscript revision, read, and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The 2000HIV project was funded by ViiV Healthcare. ViiV Healthcare had no role in the conceptualization, design, data collection, analysis, decision to publish, or preparation of the manuscript.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
.Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vadaq, N., Groenendijk, A.L., dos Santos, J.C. et al. Longitudinal plasma proteomic signatures of elite and viremic spontaneous HIV controllers. Nat Commun 17, 1173 (2026). https://doi.org/10.1038/s41467-025-67939-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67939-3