Abstract
Nitazenes are an emergent class of synthetic opioids that often rival or exceed fentanyl in their potency. These compounds have been detected internationally in illicit drugs and are the cause of increasing numbers of hospitalizations and overdoses. New analogs are consistently released, making detection challenging — new ways of testing a wide range of nitazenes and their metabolic products are urgently needed. Here, we develop a computational protocol to redesign the plant abscisic acid receptor PYR1 to bind diverse nitazenes and maintain its dynamic transduction mechanism. The best design has a low nanomolar limit of detection in vitro against nitazene and menitazene. Deep mutational scanning yielded sensors able to recognize a range of clinically relevant nitazenes and the common metabolic byproduct in a complex biological matrix with limited cross-specificity against unrelated opioids. Application of protein design tools on privileged receptors like PYR1 may yield general sensors for a wide range of applications in vitro and in vivo.
Similar content being viewed by others
Introduction
The rise in overdose deaths from synthetic opioids has been well-documented in the US and worldwide, with prevalence rising in the 2010s to account for well over half of documented overdose deaths in the US in 20221,2,3. Synthetic opioids such as fentanyl are inexpensive to produce and highly potent, with fentanyl 50 to 100 times more potent than heroin and morphine2,4, and thus are often mixed with other drugs to cheaply increase strength and addictivity2,3. An emerging class of novel synthetic opioids are 2-benzyl benzimidazole compounds known as nitazenes. First identified in the US and Canadian drug supply in 2019, nitazenes are up to 40 times more potent than fentanyl5,6,7,8. Furthermore, the molecular structures of synthetic opioids, including nitazenes, are consistently altered to skirt laws banning known compounds3,7,9. While commercial tests exist for fentanyl, including lateral flow assay test strips10,11,12, there is an immediate need for diagnostic assays that surpass the sensitivity and breadth of current nitazene test strips to detect the range of nitazenes in the drug supply and enable the rapid detection of nitazene metabolites to assist medical providers in treating exposed patients13.
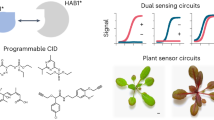
Diagnostic assays often contain a biomolecule to specifically recognize the drug(s). To develop nitazene biosensors, we chose to redesign the binding pocket of the plant abscisic acid (ABA) receptor PYR114,15,16. PYR1 binds a phosphatase HAB1 in the presence of ABA through an allosteric gate-latch lock mechanism17, representing a natural chemical induced dimerization (CID) module. This CID module presents a built-in transduction mechanism coupling protein interactions to a diverse range of signal outputs16,18,19. This is additionally advantageous because the molecular ratchet mechanism of the PYR1-HAB1 CID allows highly sensitive readouts from lower affinity ligand-receptor pairs14,20. Immunoassays with antibodies or other binding molecules can be developed by brute force screening, but the relatively small surface area of drugs limits the affinity and ability to bind multiple related drug variants. While drug host receptors can be modified into a sensor21, the same receptor can bind chemically dissimilar drugs, hampering unique identification of a drug class. Computational design has recently been used to create moderate affinity binders for largely apolar, rigid molecules22,23,24,25,26. However, binders are not sensors: the transduction of the binding event into a measurable signal must typically be engineered for each bespoke design, limiting generality and throughput. In contrast, repurposing the PYR1 binding pocket, while keeping its transduction mechanism intact, promises to be a more generalizable solution.
In this study, we use computational design of the PYR1 binding pocket to identify biosensors suitable for detection of nitazenes and their byproducts. We use deep mutational scanning, directed evolution, and computational modeling to create a pan-nitazene sensor that can detect multiple nitazene derivatives, including the common variant isotonitazene and its 4-hydroxy nitazene byproduct. We then develop a luciferase-based in vitro diagnostic platform that is label-free, fast, sensitive, and can be performed in a complex biological matrix like urine. The computational-experimental methodology represents a general way to rapidly develop biosensors to address the emergence of new synthetic opioids that can circumvent existing detection modalities.
Results
A computational design protocol for sensing the nitazene family of synthetic opioids
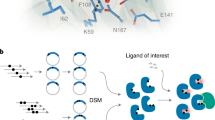
Design of an allosteric biosensor requires solving the challenge of protein-ligand binder design consistent with, and constrained by, a structural definition of the transduction mechanism27. A major advantage of redesigning PYR1 for new ligand sensing is that the receptor has an exceptionally well understood and characterized CID mechanism, where a bound water maintains hydrogen bonds between the ligand, the PYR1 receptor, and the HAB1 protein17 (Fig. 1A, B). We hypothesized that successful designs would recapitulate the spatial orientation of this ligand hydrogen bond acceptor.
A Cartoon of the PYR-HAB chemically induced dimerization mechanism. B The structural definition of the transduction mechanism for PYR1 biosensors. The ligand shown as orange sticks is abscisic acid, the original PYR1 ligand. Red and white spheres show the bound water molecule oxygen and hydrogens, respectively. Black dashes represent polar interactions between the bound water and other atoms. C Structure of the nitazene central 2-benzyl benzimidazole with locations for possible substitutions color-coded. D Functional groups of nitazene derivatives organized by substitution position and contribution to molecular potency. The sign shows the direction of effect on potency (negative values are weaker potency), and the magnitude (logEC50) reflects contribution strength. Values are regression model coefficients from one-hot encoded functional group identities at each position. The R1 groups of isotonitazene and its less-potent metabolic product 4-hydroxy nitazene are labeled, as well as the common tertiary amine at R3 shared by both molecules. E Overview of the computational design process. Temperature replica exchange MD (TREMD) is used to identify highly prevalent solution populated ligand conformers. These conformers are aligned to key molecular features preserving the PYR1 transduction mechanism, and sequences are designed to each pose. This process is iterated over conformer and alignment perturbations and filtered to identify sets of sequences. A library is encoded using position-specific and local residue preferences.
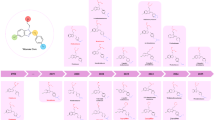
To guide our design efforts, we first sought to understand the relationship between the molecular features of nitazenes and their potency. The nitazene family of 2-benzyl benzimidazole compounds contains substitutions observed at three positions (R1, R2, R3; Fig. 1C)7. We evaluated several regression models, including ordinary least squares, Ridge, Lasso, and elastic net, to assess the relative importance of nitazene substitutions on potency (Supplementary Fig. 1) using a previous study of nitazene substitutions on cell-based activation of the μ-opioid receptor7. This analysis showed that a nitro group at R2 and several larger aliphatic ethers at R1—instead of a hydroxyl group—increased potency significantly (Fig. 1D), consistent with known potent synthetic opioids like isotonitazene7,28, which contains an isopropoxy branched ether group at R1. Substitutions of different N-containing heterocycles or secondary and tertiary amine groups at R3 have less effects on molecular potency. Finally, isotonitazene and other nitazenes are metabolized at R1 to an hydroxyl7,29,30,31, forming 4-hydroxy nitazene and related metabolites. This leaves the R2 group as a defining feature of nitazene potency. Thus, a biosensor that can detect a range of nitazenes containing an R2 nitro group and with limited cross-reactivity to other opioids is imperative for sensing nitazenes in the drug supply. Note that nitazenes with hydrogen at R2 group may in some contexts be as potent as fentanyl. Thus, a secondary goal would be to sense such nitazenes.
Repurposing an existing pocket to sense nitazenes requires a way to down-select the astronomical number of possible ligand-protein conformations. We separated the design problem into two components: (1) defining the structural pose (the rigid body orientation of the ligand relative to the PYR1 backbone); and (2) identifying a likely PYR1 sequence for that pose. To define the structural pose, we developed a computational design protocol to identify likely ligand-protein conformations and ligand orientations in the receptor binding pocket that maintain the transduction mechanism (Fig. 1B,E). Sequences for these poses are then designed using the physically based algorithm Rosetta FastDesign32 followed by physics-based filtering metrics.
To identify likely conformers of isotonitazene, which contains 9 rotatable bonds, we performed temperature replica exchange MD simulations in solution. Three isotonitazene conformers were observed greater than 97% of the time (Supplementary Fig. 2). To orient these conformers in the binding pocket, we reasoned that likely configurations occur where the important nitro group hydrogen binds to the bound water critical to the dynamic transduction mechanism. Performing the alignment revealed that two isotonitazene conformers could dock without steric clashing when all 21 allowable positions in the binding pocket are mutated to glycine. These results match ligand docking experiments in a deeply mutagenized PYR1 library biased toward hydrophobic ligands (HMH33), which contains sequences that allow docking of nitazenes in the binding pocket (Supplementary Fig. 3).
To test these initial steps of our computational design process, we screened a nitazene panel (Supplementary Table 1) against the HMH library using a yeast two-hybrid (Y2H) assay34,35, identifying sensors for six nitazene family members with differences in R1, R2, and R3 groups (although not isotonitazene; see Supplementary Fig. 4 and Supplementary Table 1, 2). Initial nitazene hits were identified with a geometric mean minimum dose response (MDR) of 61 µM in the Y2H growth selection, suggesting that their sequences were not optimized for binding. The hits had a mean of 6.4 mutations from the parental receptor; a previous computationally designed double site saturation mutagenesis library18 yielded no sensors when the same panel was screened (Supplementary Table 1), highlighting that many PYR1 mutations are necessary to repurpose this receptor for recognizing nitazenes.
Computational design protocol yields a nM-responsive nitazene binder
The insights from screening and structural modeling were integrated into the design process for a nitazene-specific PYR1 library. We used the perturbed alignments of predicted conformations of isotonitazene for Rosetta sequence redesign (see Methods, Supplementary Table 3), fixing the identity of residues known to be essential for the allosteric transduction mechanism17,34. After manual curation, the library was ordered as an oligo pool and constructed by a four part Golden Gate assembly36 in a thermally stabilized PYR1 background, PYR1HOT537 (Supplementary Table 4–5 and Supplementary Fig. 5). The library contained a mean of 9 mutations from the PYR1HOT5 background. We screened this library using Y2H assays against a panel of nitazenes (Fig. 2A), identifying 49 unique ligand-responsive sequences for six nitazene analogs, including isotonitazene (Supplementary Fig. 6 and Supplementary Table 6). Sensors with the lowest MDR for each compound are shown in Fig. 2B. Across all hits, the MDR was 19 µM, representing a significant improvement over that of the previous HMH library (61 µM; two-sided Wilcoxon rank-sum test, p value 8.5e-5).
A Schematic of the sensor isolation pipeline, including chemical structures of nitazene and menitazene. NitazeneLib1 was screened for sensors using Y2H growth selections in the presence of a ligand of interest. B Binding pocket mutations of PYR1 mutant hits from NitazeneLib1. The most sensitive receptor for each nitazene derivative is shown. Mutations from the wild-type residues are shown in bold. PYR1nita portability to different sensing modalities: Y2H growth assays (C), yeast surface display (Relative Mean PE = mean phycoerythrin (PE) fluorescence normalized to the maximum mean PE value) (D), and in vitro split nano-luciferase assays (E). D, E n = 4, data points represent the mean, error bars represent 1 s.d. and in some cases may be smaller than the symbol. Source Data are provided with this paper within Source Data. Partially created in BioRender. Wheeldon, I. (2025) https://BioRender.com/f7m2ouxhttps://BioRender.com/fq0ih6o.
From this screen, we identified PYR1nita (PYR1HOT5/H60P/N90S with K59Q, S92M, F108V, Y120A, S122G, F159A, A160V) as the lead candidate. This mutant appeared in screens for nitazene (MDR: 5 µM), menitazene (MDR: 1 µM), and protonitazene (MDR: 50 µM), and showed no detectable binding to the structurally unrelated synthetic opioid benzylfentanyl (Fig. 2C and Supplementary Fig. 6). To further characterize PYR1nita, we displayed it on the surface of yeast and performed cell surface titrations. No binding was observed in the presence of a designed thermostable HAB1 (ΔN-HAB1T+)18 alone or in the presence of the structurally unrelated ligand benzylfentanyl. Saturable responses were observed for nitazene EC50 250 nM (95% c.i. 177–390 nM) and menitazene EC50 383 nM (95% c.i. 277–588 nM), with a limit of detection (LOD) for nitazene of 43 nM (Fig. 2D). To test binding in vitro, we optimized a previously described ex vivo split luciferase detection system18 using the NanoBiT complementation reporter system38. Here, LgBiT is fused to a PYR1 receptor construct, and SmBiT is fused to HAB139 (Supplementary Fig. 7). Ligand titrations showed no activation with benzylfentanyl and nanomolar sensitivity to both nitazene EC50 243 nM (95% c.i. 216–276 nM) and menitazene EC50 306 nM (95% c.i. 240–405 nM), with a LOD for nitazene of 3.2 nM (Fig. 2E). Thus, the computational design pipeline enabled the isolation of a nitazene sensor with low nM sensitivity in vitro.
Optimization of nitazene biosensors by deep mutational scanning
Next, we used deep mutational scanning coupled with a directed evolution strategy to develop two distinct nitazene biosensors (Fig. 3A). The first of these is for the detection of 4-hydroxy nitazene, the major metabolite byproduct of isotonitazene. While the current needed LOD is unknown40, an effective sensor should exhibit a low nM LOD and function in a complex biological matrix like urine. The second sensor is a pan-sensor capable of detecting nitazenes even in complex mixtures of powders. While the sensitivity requirement of this sensor is less strenuous, it must be specific for nitazenes compared with other synthetic opioids, fillers, or recreational drugs.
A Overview of the protein engineering workflow. B Heatmap of calculated enrichment ratios of single-point mutations to the PYR1nita sensor in binding nitazene relative to the reference control. 16 residues relevant to ligand binding were scanned. Purple indicates an increased enrichment ratio and orange indicates a decreased enrichment ratio relative to the parental sensor sequence. C Correlation plot of the enrichment ratios of PYR1nita binding nitazene versus menitazene. The color scale correlates to that of the heatmap. Open purple squares indicate mutations that result in constitutive binding. D Model of the PYR1nitav2 computationally designed structure. Structure of the PYR1nitav2 binding pocket, highlighting the histidine residue at position 159 which is hypothesized to delocalize the partial charge on the nitro group. Selected mutated residues from PYR1 are shown as cornflower blue sticks. Black dashes represent potential polar contacts between the bound water, ligand, and protein. E Sequence differences between WT PYR1, computationally designed PYR1nita, PYR1nitav2, and PANnita. New mutations added in each step of the optimization are shown in orange (PYR1nita), purple (deep mutational scan), and green (PAN sensor engineering). F Yeast surface display titrations of PYR1nitav2 against nitazene (black circles), menitazene (purple circles), 4-hydroxy nitazene (blue circles), and benzylfentanyl (open red circles). The EC50 was 2.5 nM (95% c.i. 2.2–2.9 nM), 3.4 nM (95% c.i. 3.0–4.0 nM), and 2.9 nM (95% c.i. 2.2–3.9 nM) for nitazene, menitazene, and 4-hydroxy nitazene, respectively, using 500 nM of biotinylated ΔN-HAB1T+. Data are presented as mean values + SD. G Yeast surface display binding measured for PANnita against indicated ligands (BF indicates the benzylfentanyl concentration). Red striped bars indicate 0 μM ligand, orange striped bars indicate 0.01 μM ligand, green striped bars indicate 0.5 uM ligand, and purple striped bars indicate 24 μM of the nitazenes and 10 μM of benzylfentanyl. Analysis of variance was calculated using an ordinary 2-way ANOVA and Dunnett’s multiple comparison test, with a single pooled variance (**** p < 0.0001, ** p < 0.002, * p < 0.05, ns not statistically significant). For all measurements, n = 4 comprising two technical replicates performed on different days. Error bars represent 1 s.d. Source Data are provided with this paper as Source Data. Data are presented as mean values + SD.
In a first step to designing these sensors, we performed a deep mutational scan (DMS) of 16 positions inside of the PYR1nita binding pocket using yeast display screening coupled to fluorescence activated cell sorting. Libraries were screened at 250 nM of nitazene and menitazene (approx. 20% of the EC50). Additionally, a constitutive (no ligand but in the presence of 500 nM HAB1T+ binding protein) and a reference control (sorting on all gates except the gate associated with binding) were included. Example sorting gates are shown in Supplementary Fig. 8.
Following short-read deep sequencing of these populations, the frequencies of each mutant were assessed and normalized to the parental sequence by an enrichment ratio (Fig. 3B and Supplementary Fig. 9). Most mutations (298/320) were depleted, indicating a restricted sensor pocket. Nitazene and menitazene shared 10 non-constitutive beneficial mutations (V81CLI, V83ML, M92FL, A120G, A159H, V164I; Fig. 3C), largely in the aliphatic central cavity near the predicted binding site of the benzylbenzimidazole nitazene scaffold (Fig. 3D). The largest enriched mutation for both ligands was A159H. Our original design models did not include a residue which delocalizes the partial charge on the nitro group; we hypothesize that H159 satisfies this requirement (Fig. 3D). Indeed, redesigning the PYR1 sequence using the deep learning algorithm LigandMPNN41 identifies H159 in 8% of designs (Supplementary Fig. 10). This finding by an orthogonal design method is consistent with the proposed binding pose; additionally, the incorporation of a newer, deep learning packing algorithm for sequence design hints at future avenues to build on our ligand docking method to improve the computational design process. Overall, the mutational profile is largely consistent with the designed binding mode.
To develop a sensor capable of binding the major metabolite byproduct 4-hydroxy nitazene, we used the DMS output to create a focused combinatorial library of 4608 members largely containing the beneficial mutations shared between nitazene and menitazene (Supplementary Table 7). Screening this library on nitazene and menitazene yielded two sensors, PYR1nitav2.1 (PYR1nita with V81I, M92F, A120G, G122E, E141D, A159H) and PYR1nitav2 (PYR1nita with V81I, V83L, E141D, A159H), containing 4-6 additional mutations from PYR1nita (Fig. 3E and Supplementary Fig. 10). Both sensors recognized 4-hydroxy nitazene, menitazene, and nitazene at an average LOD of 100 pM and EC50 of approximately 2 nM (Fig. 3F and Supplementary Fig. 11), representing a more than 100-fold improvement in affinity over the originally designed PYR1nita sensor. Neither sensor recognized the unrelated synthetic opioid benzylfentanyl at the highest concentration tested, indicating highly specific recognition of nitazene, its metabolic byproduct 4-hydroxy nitazene, and a close analog menitazene over a competing synthetic opioid.
Our underlying hypothesis in the design process was the importance of the nitro group in molecular recognition by the PYR1-based sensors. One prediction resulting from this hypothesis is that two nitazenes that differ only by the nitro group should have very different affinities. To examine this, we used yeast display to determine the EC50 for the PYR1nita, PYR1nitav2, and PYR1nitav2.1 sensors against metonitazene and metodesnitazene. Metonitazene and metodesnitazene are identical except the presence or absence of a nitro group at R2. All three sensors had markedly reduced or no responsiveness to metodesnitazene compared with metonitazene (Supplementary Fig. 12). For example, PYR1nitav2 showed at least a 650-fold difference in EC50 values for metonitazene compared with metodesnitazene. Overall, these results are consistent with the importance of the nitro group in the ligand-induced latch closure, consistent with the design poses. We note that there is small but measurable binding signal present at the higher desnitazene concentrations for the engineered sensors, suggesting that a pan sensor may be capable of recognizing even nitazenes lacking a nitro group at R2.
To develop a sequence able to sense structurally diverse nitazenes, we used the structural model of PYR1nita to identify mutations which could allow binding breadth by increasing the pocket space for diverse functional R1 and R3 groups. We created a focused combinatorial library of 2048 variants encoding differences at positions 59, 141, 163, 164, & 167 to the PYR1nitav2.1 and PYR1nitav2 sensors (Supplementary Table 8). After screening against constitutive binders, the library was split and screened in parallel against butonitazene, isotonitazene, and N-pyrrolidino isotonitazene (Supplementary Fig. 13). These populations were deep sequenced; sequences that were enriched in all three ligand populations were individually tested (Supplementary Fig. 14). The best sensors, PANnita and PANnita.1, recognized the three diverse nitazenes and did not recognize benzylfentanyl (Fig. 3G and Supplementary Fig. 15). PANnita has in total 11 mutations in the receptor binding pocket out of 18 total mutable positions. Combined, computational design coupled to protein engineering can access a more diverse functional sequence space than accessed by previous libraries.
Engineered PYR1 sensors can discriminate nitazenes and metabolites in relevant diagnostic scenarios
To test whether PYRnitav2.1 and PANnita are capable of sensing nitazenes under relevant conditions, we repurposed our in vitro luminescence assay. Luminescence detection assays are susceptible to errors from varying experimental conditions including fluctuating activity in diverse biological matrices42. For these reasons, ease of single-sample detection is limited by the need for external calibration. A previously described calibrator luciferase43 enables single-sample ratiometric quantitative readouts (Fig. 4A). This assay compares the ratio of ligand dependent luminescence to background GFP fluorescence to mitigate experimental variance. We tested this new calibrator assay with the parental PYR1 sensor and a range of synthetic cannabinoid sensors18. All tested sensors had nearly identical dynamic range, LOD, and EC50 values between buffer and urine (Supplementary Fig. 16). Using this assay, PYR1nitav2.1 could recognize the metabolic byproduct 4-hydroxy nitazene with a LOD of 1 nM (0.37 ng/mL) in urine, its most clinically relevant biological matrix, and with no cross-reactivity to the unrelated opioids benzylfentanyl, codeine, or heroin (Fig. 4B).
A Cartoon of the luminescence assay. A mNeonGreen-NanoBiT “calibrator” protein enables ratiometric detection of samples, fluorescing at a wavelength of 520 nm independent of ligand concentration. PYR1 to ΔN-HAB1T+ dimerization results in an increased ratio of the relative luminescence units per second (RLU/s) at 450 nm versus the RLU/s at 520 nm. B Ratiometric assay of PYR1nitav2.1 in buffer or in urine using the indicated ligands. EC50 measurements and limit of detection (LOD) for the sensor are colored for the appropriate condition. 200 nM SmBiT-ΔNHAB1, 10 nM LgBiT-PYR1, and 512 pM calibrator are used. Benzylfentanyl, codeine, and heroin were tested in buffer. The nitazene-specific EC50 was 5.7 nM (95% c.i. 4.0– 8.3 nM) in the urine matrix and 2.8 nM (95% c.i. 2.3–3.4 nM) in buffer. C Luminescence assay results using PANnita.1 show sensitivity against a panel of nitazene variants and synthetic opioids across multiple concentrations. Analysis of variance was calculated using an ordinary 2-way ANOVA and Dunnett’s multiple comparison test, with a single pooled variance (****p < 0.0001, **p < 0.01, ns not statistically significant). 4 nM SmBit-ΔNHAB1 and 4 nM LgBiT-PYR1 are used. Data is shown as the average of n = 3 (B) or n = 4 (C) replicates. Error bars represent the standard error of the mean and in some cases are smaller than the symbols. EC50s were calculated by fitting the data to a four-parameter logistical sigmoidal model with nonlinear least-squares regression. LODs were calculated using the 3σ method, which is equivalent to the drug concentration that yields a signal equal to 3 times the standard deviation of the blank after subtraction. Source Data are provided with this paper as Source Data.
Another diagnostic use for sensors is discriminating nitazenes in unknown powders. To assess this use case, we tested whether PANnita.1 could recognize diverse nitazenes relative to these other opioids. PANnita.1 bound the nitazene variants butonitazene, isotonitazene, and N-pyrrolidino isotonitazene while exhibiting minimal perception of benzylfentanyl, codeine, and heroin. Ligands were dissolved in aqueous buffer with a cosolvent consisting of 0.42 (v/v) % methanol and 0.58 (v/v) % DMSO except for heroin, which was dissolved in 1 (v/v) % acetonitrile. At their highest tested concentration, this assay was able to detect a 6.4x, 7.2x, and 23x fold-change respectively for the target ligands butonitazene, isotonitazene, and N-pyrrolidino nitazene, with limited activation of off-target opioids (2-way ANOVA, p < 0.0001) (Fig. 4C). Similar results were obtained for the PANnita sensor (Supplementary Fig. 17). To determine the wider range of efficacy of the PAN sensors, PANnita and PANnita.1 were tested against ten structurally diverse nitazenes (at 10 μM) not used in sensor engineering. These included desnitazenes lacking the nitro group, and 5-aminoisotonitazene not yet detected by nitazene test strips41. Remarkably, these pan sensors combined to recognize five of these nitazenes, including two of the three desnitazenes and ethyleneoxynitazene (Supplementary Figs. 18–19). Pan sensors failed to recognize 5-aminoisotonitazene which contains an amine at R2 that is likely incompatible with the water-mediated latch closure. Additionally, pan sensors did not recognize some nitazenes (ethylene nitazene, ethylene isotonitazene) likely because of steric clashes. Still, the ability to sense diverse nitazenes, including desnitazenes, distinct from those used in the engineering process suggests that these or closely related engineered sensors may be able to identify the overwhelming majority of nitazenes in the drug supply.
In directed evolution, it is well established that optimizing for affinity and breadth can impact other properties like protein expression yields, stability against thermal challenge, and kinetic activation44. We assessed these properties in the pan-nitazene sensors and PYR1nitav2.1 compared with HOT5mandi37, the most stable engineered PYR1-based sensor developed thus far. All proteins had identical expression yields of 19 mg/L under standard E. coli expression conditions when expressed as LgBiT and MBP-tagged fusion proteins (Supplementary Fig. 20). To test stability against thermal challenge, we developed a luminescence-based thermal inactivation assay. All sensors showed a 44.8 °C or higher temperature at which a 20 min incubation resulted in 50% loss in activity \(({T}_{50}^{20})\). The \({T}_{50}^{20}\) for PYR1nitav2.1 was 48.0 oC (95% c.i. 47.3–48.6 °C), which was not significantly different (p value = 0.13) than that for HOT5mandi (\({T}_{50}^{20}\) = 48.5, 95% c.i. 47.3–48.6 °C)37 (Supplementary Fig. 21). We also tested the time the luminescence assay takes to equilibrate. At 10 μM ligand and at room temperature, all sensors reached 90% of steady-state luminescence values within 17 min. Notably, PYR1nitav2.1 reached equilibrium faster than HOT5mandi (11 vs. 14 min, p value 0.0458) (Supplementary Fig. 22). Although the PAN sensors are intended for testing unidentified powders in vitro, we also tested whether they retain the ability to sense N-pyrrolidino isotonitazene in a biological matrix. In urine, both sensors maintained the ability to discriminate between benzylfentanyl and N-pyrrolidino isotonitazene (Supplementary Fig. 23). Thus, the tested sensors increased breadth and affinity for nitazenes without appreciably changing their underlying kinetic activation, expression yields, or stability against thermal challenge.
Discussion
Computational protein design has advanced towards relevant and pressing societal needs45. Here we designed and engineered protein biosensors for a range of clinically relevant synthetic opioids. These sensors can discriminate between different nitazenes and structurally unrelated opioids; we show that one sensor is functional in a pooled urine sample. Notably, nitazenes cannot currently be differentiated from other synthetic opioids using cell-based assays measuring μ-opioid receptor activation46. Commercial test strips for nitazenes exist but cannot sense desnitazenes, among other variants. We focused on demonstrating urine efficacy of the PYR1nita sensors because these sensors recognize a R1 hydroxyl group present in metabolite products of common nitazenes7,29,30,31, thus enabling detection of nitazenes post-consumption. The PANnita sensors, which were optimized for binding of a wide range of variants, are best deployed to detect nitazenes in environmental samples (e.g. dry powders and other forms). Overall, the designed nitazene sensors in this study are a strong starting point for additional development into reliable point-of-care diagnostic devices. A promising application to pursue is lateral flow assays in which the PYR1-sensing system would replace antibody detection modalities.
There are current limitations that must be addressed before real world deployment of this technology. Full validation of LOD in the appropriate biological matrix is required, including different urine samples spiked at different opioid concentrations to evaluate potential for false positives and false negatives. A lower and/or more consistent background signal in urine samples is needed for standardized testing. While we tested the ability of our sensors to detect nitazenes relative to different drugs, further testing of more complicated mixtures is necessary. In addition, further study of the PYR1 binding pocket will help identify sequences that better recognize emerging designer drugs including the desnitazene and ethylene nitazene variants tested here.
Our computational design process involved repurposing the plant hormone receptor PYR1. Variants of the PYR1 receptor are able to sense an unusually broad spectrum of drug-like molecules47. This suggests that PYR1 sensors can be developed for a wide range of molecules, ultimately enabling new medical diagnostics, chemically-responsive cell therapies, environmental sensors19, and biotechnologies for cell engineering33. Our design process allows navigation to a search space largely inaccessible to random site saturation mutagenesis libraries, and when coupled with our conformer selection and alignment protocol creates a powerful design approach for ligand binding. The designed sensors, the best of which exhibit pM responsiveness and low nM EC50s, are orders of magnitude more sensitive than has been achieved than other computational approaches for small molecule binders22,24,25. We speculate that one reason our designs were successful was that we had a quantitative, molecular understanding of the transduction mechanism, which was worked out by biochemical and structural studies of abscisic acid perception in plants17,34,48,49,50,51. By analogy, a deeper mechanistic insight into the more complicated and varied allosteric transduction mechanisms of ligand-dependent molecules like GPCRs or bacterial transcription regulators27 could improve biosensor hit rates for these classes of proteins.
Our design process, while successful, had some limitations. First, we used a combinatorial library rather than discrete designs, resulting in larger libraries with many undesired mutational combinations. While our Y2H screening platform is robust, larger libraries entail greater screening effort. Second, our design process was separated into pose generation and sequence generation. The advance of our method is the generation of realistic poses themselves through conformer generation and alignment sampling. This is essential for keeping the transduction mechanism of PYR1-based sensors intact. However, we used physically-based packing algorithms and filtering strategies to identify designs. This established process has largely been supplanted with deep learning approaches50,51 (e.g. ProteinMPNN/LigandMPNN followed by AlphaFold2 filtering). We anticipate that emerging and future deep learning algorithms will further improve design and overall hit rates in designing PYR1 receptors for new target ligands, which could be incorporated into our protocol in the future as pose selection is independent of sequence generation. The inability of the PAN sensors to bind ligands like ethylene nitazene and ethylene etonitazene — likely because of steric clashes — point to the limitations of designing against a fixed backbone. Other ligands may simply not be able to be accommodated in the PYR1 pocket. To extend this approach for other ligands, we anticipate the exciting possibility of generating de novo proteins which maintain the PYR1 transduction mechanism. These efforts would be enabled by new computational and experimental tools to test and predict motions of designed proteins52,53.
Methods
Chemicals
Nitazenes and other pharmaceutically active molecules for use in yeast surface display and split luciferase assays were purchased through Cayman Chemical, including 4’ hydroxy nitazene (#30218), benzyl fentanyl (hydrochloride, #19883), butonitazene (exempt preparation, #38653), codeine (CRM, #ISO60140), ethyleneoxynitazene (citrate, #34947), heroin-d9 (exempt preparation, #42070), isotonitazene (CRM, #30880), menitazene (citrate, #36634), metonitazene (citrate, CRM, #40808), N-desethyl etonitazene (#32512), nitazene (citrate, #36635), N-pyrrolidino etonitazene (CRM, #42194), N-pyrrolidino isotonitazene (citrate, #34909), protonitazene (hydrochloride, CRM, #38143), metodesnitazene (hydrochloride, #43513), etodesnitazene (CRM, #43092), isotodesnitazene (citrate, #30286), N-pyrrolidino isotonitazene (citrate, #34909), N,N-dimethylamino etonitazene (citrate, #34452), ethylene etonitazene (citrate, #35166), 5-aminoisotonitazene (#29318), ethylene nitazene (citrate, #34946), metonitazene (CRM, #43158), and (+)-WIN 55,212-2 (mesylate, #10009023). Chemicals acquired from Sigma include mandipropamid (#32805-100 mg), JWH-015 (#J4252-5MG), JWH-016, and JWH-072. Abscisic acid was purchased from GoldBio (#A-050-2.5). Diazinon was acquired from Chem Service Inc. (#N-116121-250MG). Hikarazine-108 was a gift from Dr. Yves Janin, Centre National de la Recherche Scientifique. Dry Hikarazine-108 was resuspended in 200 μL DMSO and then diluted with 300 μL of acidic ethanol. Acidic ethanol is prepared by adding 100 μL of 37% hydrochloric acid to 12 mL of 100% ethanol. This resulted in a 4.22 mM stock that can be stored at −20 °C indefinitely.
Computational design protocol
The mutational library NitazeneLib1 was generated using a structural replacement computational docking protocol to generate high-quality plausible protein-ligand alignments of simulated low-energy ligand conformers.
To generate ligand conformers by molecular dynamics, an initial 3D conformer was downloaded from Pubchem54. Conformers were generated by temperature replica exchange molecular dynamics simulations (TREMD)55 as described in Leonard et al56. Briefly, we used the default implemented settings which run 27 temperature replicas ranging from 300 to 450 K and all simulations were run for 100 ns with replica exchange occurring every 2 ps. The conformer with the highest observed occupancy in the lowest temperature replica was given a relative free energy of 0.0 kcal/mol, and the free energy of other conformers were calculated relative to the highest observed occupancy assuming a Boltzmann weighted ensemble. Generated conformers were plotted sequentially by free energy relative to the lowest energy conformer, categorizing conformers as either “high occupancy” (greater than 4% occupancy cutoff used for menitazene and isotonitazene) or “low occupancy” (less than 1% occupancy cutoff used for menitazene and isotonitazene) (Supplementary Fig. 2). Low-energy conformers only were exported as a single sdf file for ligand docking. For this paper, we developed an automated protocol with open-source code (https://github.com/ajfriedman22/SM_ConfGen).
Low-energy ligand conformers generated by TREMD were docked into the PYR1 protein ligand binding pocket using the “dock to sequence” approach of our method “Structural Replacement Biosensor Design”, available as custom python scripts at https://github.com/alisoncleonard/Structural-Replacement-Biosensor-Design. In the dock to sequence method, the wild-type protein sequence to design is mutated to match the sequence of a known low-affinity ligand binding sequence identified through library screening in yeast two-hybrid (Supplementary Fig. 4), enabling visualization of ligand alignments that could accommodate the known binding sequence. Each nitazene conformer was docked in space using a Singular Value Decomposition (SVD)-based superposition method to align the 3D dimensional coordinates of a given novel query ligand with a target molecule. To engineer PYR1, the PDB structure 3QN157 was used as the template, with atoms surrounding the conserved nitro group at nitazene position R2 superimposed on the target molecule abscisic acid ketone, with the nitro group acting as the required hydrogen bond acceptor needed for binding58.
The alignment process begins with two sets of atomic coordinates: one from the query molecule to be aligned and the other from the target molecule. These do not need to be the full set of atomic coordinates for the molecules but must include at least three atoms that are not collinear. Each coordinate set is arranged into a matrix, where each row in the matrix is the x, y, and z coordinates of an atom, and the corresponding row in the other matrix is the corresponding atom in the other molecule. To obtain a set of transformation matrices, each coordinate set is translated so that its center of mass (mean) is positioned at the origin, then a covariance matrix is calculated between the two centered coordinate sets, and finally the covariance matrix is decomposed into three components: two orthogonal matrices and a diagonal matrix of singular values which are used to calculate a rotation matrix. Once the rotation matrix is computed, the original query compound undergoes transformations applied using the apply_transform_Rx_plus_v function available in PyRosetta, where the query coordinates are first translated so that their center of mass is at the origin, to ensure that the rotation is applied correctly, next the computed rotation matrix is applied to the coordinates of the query compound, to put it in plane with the target molecule, and finally the query is translated to the center of mass of the target molecule coordinates, completing the alignment.
As it is unlikely that a novel ligand would dock in exactly the same orientation as abscisic acid, the aligned ligand was next perturbed in space using the PyRosetta Rigid Body Perturbation mover. The moved ligand was verified both to not clash with the protein backbone coordinates, determined using a fast-grid search of PYR1 subjected to a poly-glycine shave, and to contain a nitrogen or oxygen close enough to the latch water molecule of the PYR1 structure to form a hydrogen bond. If these conditions were satisfied, the PYR1-ligand complex was relaxed and repacked using the PyRosetta Packer function, and the repacked complex was checked for collisions with the full protein including both backbone and side chains by fast-grid search. Finally, complexes that pass clash check were scored using the standard PyRosetta scorefunction, with complexes below −300 REU generally considered to be realistic protein-ligand complexes. This process can be repeated as many times as desired to appropriately map plausible ligand alignments in the protein pocket (Supplementary Table 2). The number of alignments tested per conformer and total per ligand are listed in Supplementary Table 3, along with the number of alignments passing clash check taken forward.
Modeled protein-ligand structures were used as inputs for Rosetta FastDesign32 (scripts available at https://github.com/alisoncleonard/Structural-Replacement-Biosensor-Design) for structure-informed rational design to select mutations for inclusion in the library NitazeneLib1. The results presented in Supplementary Fig. 10 were a LigandMPNN41 sequence redesign of a similar set of modeled protein-ligand structures using (https://github.com/dauparas/LigandMPNN). The temperature was set to 0.2 and the noise level used was 0.1 Angstrom Gaussian noise using ligandmpnn_v_32_010_25.pt.
Plasmids and plasmid library generation
A full list of primer sets (Supplementary Table 9) and plasmids (Supplementary Table 10) used in the work are included. All plasmids were sequence verified using Oxford NanoPore (Plasmidsaurus).
To construct NitazeneLib1, libraries containing desired combinations of mutations were encoded in oligo pools (Agilent) divided into 3 cassettes (Supplementary Table 4), along with 2 constant region cassettes flanking the PYR1 gene, to be cloned into the PYR1 yeast two-hybrid destination vector pND00559. Three sub-libraries were prepared by Golden Gate assembly59, with each containing 2 of the 3 variable portions and one constant cassette (e.g. Library A contained variable cassettes 1 and 2, plus a constant cassette 3). Transformation efficiencies (Supplementary Table 5) were estimated using serial dilution, and library quality (Supplementary Fig. 5) determined by short-read deep sequencing (Azenta Amplicon-EZ). PYR1nita was assembled into pND003 to make pRMC075 by Golden Gate assembly; this plasmid was subsequently transformed into EBY100 S. cerevisiae cells (ATCC Catalog Number MYA-4941TM)60.
A single site saturation mutagenesis (SSM) library at 16 positions (59, 81, 83, 92, 94, 108, 110, 117, 120, 122, 141, 159, 160, 163, 164, and 167) was prepared for PYR1nita (PYR1HOT5/H60P/N90S/K59Q/S92M/F108V/Y120A/S122G/F159A/A160V) by nicking mutagenesis61 using pRMC075 as a plasmid template. NNK mutagenic primers were designed using published SUNi mutagenesis scripts62. Cells were transformed into XL1-Blue electrocompetent E. coli. Plasmid DNA was extracted with a ZymoPURE II Plasmid midiprep kit. The library was transformed into EBY100 S. cerevisiae cells using the lithium acetate method60.
A pan-nitazene library (Nitazene Library 2, Supplementary Table 7) of 4608 members was made with mutations encoded into ultramers (IDT) with constrained degenerate mixed base codons and assembled according to59. A single forward ultramer (Nitz_Combo_Lib2_for) spanning residues 58 to 119 and encoding V81VIL (VTT), V83VILM (VTK), M92MILF (WTK), V108VI (RTH) was amplified with a mix of two reverse ultramers (Nitz_Combo_Lib2_rev_159AV, Nitz_Combo_Lib2_rev_159H) spanning residues 109-168 and encoding A120GA (GSS), G122DEAG (GVN), E141DE (GAW), and either A159GA (GYG) or A159H (CAT) generating a full length Golden Gate ready cassette. A single gBlock (IDT) (Nitz_ComboLib2_Block_1_3) encoding both N-terminal sequence (before residue 58) and C-terminal sequence (after residue 168) with appropriate Golden Gate cut sites was used to generate the full length CDS in the library with pND003 destination vector. The library was prepared for YSD as described above. PYR1nitav2 and PYR1nitav2.1 were isolated as colonies to obtain plasmids P8 and P5 after sorting, respectively.
Two libraries based off of PYR1nitav2 and PYR1nitav2.1 were constructed via combinatorial mutagenesis essentially according to (15) to make Nitazene Library 3 and Nitazene Library 4 (Supplementary Table 8), respectively. Each library had the following mutations added by long ultramers (IDT) (pZB717-pZB720) encoding Q59QVASLIMT (DYR), D141EDAG (GVN), A159VA (GYT) or A159H, V163VA (GYN), V164VILM (VTK), and N167SACGN (KSY), where A159VA and A159H were encoded on two separate ultramers, for a total of 2560 library members each. After Nitazene Libraries 3 and 4 were prepared for YSD as described above, they were pooled together to create Nitazene Library 5 containing 5120 library members total, which was screened according to conditions listed below. NGS was performed as described below to identify pan-nitazene contenders. Select individuals were cloned into pND003 with constant cassettes cass1_pND003_ACL and cass5_pND003_ACL and gBlocks (IDT) containing the corresponding mutations for each contender 1_Cas_2-4_Nit_pan_contender through 6_Cas_2-4_Nit_pan_contender were created via Golden Gate assembly according to Daffern et al., 202336.
pYTK047 was a gift from John Dueber (Addgene plasmid # 65154; http://n2t.net/addgene:65154; RRID:Addgene_65154)63.
Yeast two-hybrid screening of mutant biosensor libraries
Selection experiments for mutant receptors that respond to new ligands were conducted as previously described by Beltrán58. Briefly, the indicated pBD-PYR1 mutant library was transformed into MAV99 (MATa trp1-901 leu2-3,112 hisΔ200 ade2-101 gal4Δ gal80Δ can1rcyh2rLYS2::(GAL1::HIS3) GAL1::lacZ SPO13::10xGAL4site::URA3) harboring pAD-HAB1. Mav99 was a gift from Jeff Bachant and is authenticated by functional controls. A minimum transformation efficiency of 1 million CFU was achieved, typically by pooling two transformations. After transformations, negative selections were conducted twice to remove receptors that bind HAB1 in a ligand-independent fashion (i.e., constitutive receptors) by growing the library on solid media containing 1 g/L 5-FOA; the purged library was collected and stored as 30 (v/v) % glycerol stocks until use58.
These glycerol stocks were used in subsequent selections for cells responsive to 100 μM nitazene ligands described in Supplementary Table 1 on SD-Trp,-Leu,-Ura media. Colonies supporting uracil-independent growth at 30 °C were isolated after 3 days, regrown on SD-Trp,-Leu,+1 g/L 5-FOA to screen out constitutive biosensors, retested to confirm ligand-dependent growth on SD-Trp,-Leu,-Ura plates with and without test chemical. The minimum dose response was determined by growth validation after 3 days, in which yeast cells were grown as spot assays by suspending a colony in 100 µL of 1X TE and dropping 2 µL onto SD-Trp,-Leu,-Ura plates containing various concentrations of the target ligand.
Library sorting by yeast surface display
Sorting of the PYR1 yeast displayed libraries was performed as done in Steiner et al.64 except for the following differences. Cells were induced in SD/GCAA media containing a 9:1 ratio of galactose:dextrose. Induction was carried out at 22 °C and 250 rpm for 20 h. Cells were washed in citrate-buffered saline (20 mM trisodium citrate dihydrate, 147 mM NaCl, 4.5 mM KCl) supplemented with 0.1% w/v BSA adjusted to pH 8.0 with 1 M NaOH (CBSF), while prepared ΔN-HAB1T+ from plasmid pJS72358 was resuspended in CBSF plus 1 mM DTT and 5 mM TCEP.
For the SSM library sort, each labeling condition had 250 µL of OD600 = 2 cells, 3 µL of ligand dissolved in DMSO for an overall solvent of 1 v/v%, ΔN-HAB1T+ to 500 nM, and the volume of the reaction brought to 300 µL with CBSF. Binding and labeling were performed as in Leonard et al.56. Cells were collected at approximately half of EC50 in a diagonal gate, excluding background, selecting 200,000 cells each of the top 10–15% of binders, a reference population, and non-binders. Cells were grown and stocks prepared according to56. Next generation sequencing (NGS) was performed on these screened cells as described below.
Nitazene Library 2 was screened with 250 µl of OD600 = 2 cells and maintained solvent at 1 v/v%, 100 nM of ΔN-HAB1T+, and a total volume of 300 µl attained by adding CBSF in order to obtain PYR1nitav2 sensors. First, 200,000 cells of the top 10% of binders were sorted using 250 nM menitazene. The second sort used identical reaction conditions, but only 200,000 cells of the top 5% of binders were collected. The final sort for the PYR1nitav2 sensors differed from the previous sort only by using 100 nM menitazene, which yielded PYR1nitav2 and PYR1nitav2.1 sensors, isolated as individual colonies. DNA was extracted from PYR1nitav2 and PYR1nitav2.1 and plasmid sequenced (Plasmidsaurus) to identify sensor sequences.
A total of 250 µl of OD600 = 2 cells of the combinatorial mutagenesis libraries of PYR1nitav2 and PYR1nitav2.1 was screened with 100 nM ΔN-HAB1T+ and the first sort was performed with no ligand and 1% (v/v) DMSO to collect 200,000 cells that did not bind constitutively. The resulting cells were grown and screened against three ligands in parallel, butonitazene, isotonitazene, and N-pyrrolidino isotonitazene at 2.5 µM, 2.5 µM, and 23.8 µM, respectively, where butonitazene and isotonitazene were dissolved in methanol and the final volume of solvent was maintained at at 1 (v/v)% in all samples. The top 5% of binders were collected to get 200,000 cells and DNA was prepared for NGS as described below.
The top performing contenders for the PANnita sensor were identified as those that had the highest enrichment ratios for each ligand and gBlocks cassettes containing the identified mutations were cloned into pND003 via Golden Gate assembly to make pRMC079-084. These sensors were tested with yeast surface display with 100 nM ΔN-HAB1T+ and various concentrations of butonitazene, isotonitazene, N-pyrrolidino isotonitazene, and benzylfentanyl, maintaining a solvent concentration of 1% v/v.
Next generation sequencing of collected cell populations
DNA was extracted using freeze/thaw cycles and zymolase according to Medina-Cucurella & Whitehead60, except for the following differences. Instead of utilizing a Qiagen mini-prep column, an NEB miniprep column was utilized and DNA was washed with 550 µL DNA wash buffer two times and eluted in 30 µL nuclease free water. Product from the purification of plasmid with exonuclease I and lambda nuclease was cleaned using an NEB DNA & PCR cleanup kit according to the manufacturer’s instructions.
Plasmid DNA was prepared for NGS by amplifying with either primer sets RMC-P1035 and RMC-P1036 (length of 383 bp) or RMC-P1037 and RMC-P1038 (length of 388 bp). 10 µL of plasmid DNA was amplified using 12.5 µL Q5® High-Fidelity 2X Master Mix, 1.25 µL 10 µM forward primer, 1.25 µL 10 µM reverse primer, and 25 µL nuclease free water (NFW) and placed in the thermocycler with the following cycles: 30 s at 98 °C, 18 cycles of 5 s at 98 °C, 20 s at 57.5 °C, and 30 s at 72 °C, followed by a final extension of 2 minutes at 72 °C and a hold at 10 °C. PCR product was purified using an NEB Monarch DNA Gel Extraction kit. DNA was pooled for nitazene and menitazene top binders, then reference populations, and non-binding populations at equal masses and sent to Quintara for AmpExpress next generation sequencing. Resulting sequences were merged with FLASh65, filtered for appropriate read length and amino acid start sequence, and heat maps of calculated enrichment ratios were created as in Medina-Cucurella & Whitehead60.
Protein expression and purification
Proteins were expressed in E. coli BL21(DE3) under control of a T7 promoter, autoinduced, lysed using B-PER surfactant, and purified by Ni-NTA affinity chromatography58.
Split Luciferase in vitro Assay
Proteins, stored as ammonium sulfate precipitates, were pelleted at 17,000xg for 10 minutes and then resuspended in desalting buffer (50 mM HEPES-KOH, 10% w/v glycerol, 200 mM KCl, 1 mM DTT, 5 mM TCEP, pH 8.0). A 7 K MWCO Zeba™ spin desalting column (ThermoFisher, Cat. No. 89882) was equilibrated with desalting buffer and then used to buffer equilibrate the resuspended protein, which was then stored on ice until use. Protein concentration was measured by diluting 20 μL of desalted proteins into 180 μL of 8 M urea in a UV-Star plate (Item No. 655801). An absorbance value at 280 nm was taken using a plate reader (BioTek Synergy™ H1 Microplate Reader), background subtracted from a blank, and scaled 10x to account for the dilution. The concentration was measured by the Edelhoch method66 using calculated extinction coefficients.
Ligand was diluted to 10x working concentration in desalting buffer and 20 μL of this dilution was added to the bottom of Greiner™ 96-well, white, flat bottom plate (Cat. No. 655075). In a separate tube, desired proteins are combined with the Hikarazine-108, and diluted with desalting buffer to reach a final volume of 180 μL per well. Add the prosubstrate, Hikarazine-108, last, just before adding the mixture to the wells to minimize its depletion before the reaction has begun in the plate. For bioassay measurements, 40 μL urine was supplemented to this master mix per well such that the final concentration of urine was 20% v/v when combined with the ligand. Pooled human urine was sourced from Innovative Research Inc. (SKU# IRHUURE50ML) and was sterile filtered before use to remove any aggregates. Typical working concentrations for LgBiT-PYR1 and SmBiT-HAB1 are in the low nM range (2 and 20 nM are an appropriate starting point for LgBiT-PYR1 and SmBiT-HAB1, respectively) while Hikarazine-108 is always used at 10 μM. This 180 μL master mix was scaled to encompass all wells and replicates. Using a multichannel pipette, 180 μL of this mixture are added to each well. The wells are mixed up and down using the pipette, not aspirating to avoid bubbles.
The plate was placed in a plate reader (BioTek Synergy™ H1 Microplate Reader) and an autogain feature was used to adjust the gain at a wavelength of 450 nm for the brightest anticipated well, so that overflow does not occur. An integration time of 1 s and a read height of 7 mm was used. The plate was then run, and data was output in units of relative luminescence units (RLU) per second. While we used the readout at time = 0, we observed luminescence at intervals for one hour. Doing so offered greater resolution into the kinetics of our system and was helpful for monitoring its behavior.
For ratiometric detection, inclusion of the calibrator luciferase includes mNeonGreen fused to the full length NanoBiT luciferase, enabling bioluminescence resonance energy transfer (BRET) between the pair43. This protein sequence was ordered as an eBlock from IDT (Supplementary Table 11) and cloned into a protein expression backbone from pJS723 using Gibson Assembly67. The NanoBiT luminesces blue light (λmax ~ 450 nm) while mNeonGreen fluoresces with a λmax ~ 520 nm, such that the optimized system yields two distinct spectrum peaks.
To resolve two distinct peaks, we ran a spectral scan of each well. With this method, samples volumes of 180 μL (without the luminescence prosubstrate) were added to the plate and then covered and placed on a plate shaker at 25 °C and 750 rpm. After 30 min, the foil was removed, and 20 μL of Hikarazine-108 was added to bring the well up to a volume of 200 μL and samples were mixed using a multichannel pipette. On the plate reader, we specified a range of wavelengths from 415 to 555 with a step size of 35 nm. For each sample we obtained RLU/s at these 5 defined wavelengths. Complex formation is measured as the RLUs/sec at λ450 divided by the RLUs/sec at λ520. The specific concentration of calibrator used varied based upon the concentrations of LgBiT-PYR1 and SmBiT-HAB1 and typically requires adjustment for each PYR1-ligand pair with the working concentration typically being mid-high pM. An appropriate starting point for optimizing this concentration would be 100-fold less than the concentration of your limiting protein (LgBiT-PYR1 or SmBiT-HAB1).
For testing the PANnita sensors in the SpLuc assay (Fig. 4C and Supplementary Fig. 17), a 1 (v/v) % cosolvent of 42% methanol and 58% DMSO was used for all ligands except for heroin, which was dissolved solely in acetonitrile. Similarly, for testing these sensors versus the extended nitazene variants set (Supplementary Fig. 18–19), a cosolvent of 57% v/v DMSO, 39% v/v methanol, and 4% v/v acetonitrile was used for all ligands. Limit of detection (LOD) was calculated as the lowest ligand concentration at which the mean luminescence measurement exceeded three standard deviations above that of the corresponding blank luminescence measurement.
All measurements collected as distinct technical replicates by running multiple wells in parallel within the same plate. All statistical analyses were performed using GraphpadPrism.
Titrations via yeast surface display
Ligand-specific titrations of nitazene sensors were carried out through yeast surface display, run similarly to library sorting, but with the following differences. A total of 10 µL of OD600 = 2 were labeled with various dilutions of ligand such that the ligand’s solvent was <1% of the reaction volume and ΔN-HAB1T+ was added to 500 nM in the final reaction volume of 50 µL. Data from 25,000 labeled cells was collected after resuspending cell pellets in 100 µL CBSF, processed in FlowJo v10 software, and fitted to a specific binding curve with Hill slope in GraphPad Prism 10 to find EC50 values.
Thermal inactivation assay
This assay was adapted from Daffern et al37 for utilization within the split luciferase in vitro assay. 72 µL of LgBiT-PYR1 variants at 20 nM in desalting buffer were aliquoted to 12 PCR tubes with reducing agents. These tubes were placed in a thermocycler for 20 min with a gradient of temperatures spanning 38 °C to 60 °C after which each tube was brought back down to 4 °C and placed on ice. 20 µL of each was pipetted into a white, flat bottom plate using a multichannel pipette. 180 µL of desalting buffer with SmBiT-HAB1, Hikarazine-108, and ligand were added to these wells to bring them up to a final volume of 200 µL and reach working concentrations of 20 nM, 2 nM, 4.22 µM and 10 µM for SmBiT-HAB1, LgBiT-PYR1, Hikarazine-108, and each ligand respectively. Wells were pipetted up and down to mix, bubbles were removed, and luminescence was measured for each well using a plate reader in accordance with the split luciferase in vitro assay protocol as previously outlined.
Statistics and reproducibility
For replication, we used 3 or 4 replicates (two technical per day, two replicates on separate days) for all experimental data reported in the paper, except where noted. Deep mutational scanning was performed once. No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The Investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw deep sequencing data are deposited in the SRA (BioProject ID PRJNA1256820), and analyzed deep sequencing data are on Zenodo (https://doi.org/10.5281/zenodo.15298584)68. PDB ID of structure used for structural replacement code: 3QN1 All other data are available in the main text or the supplementary materials. Source Data are provided with this paper as “Source Data.xlsx”. Primers, plasmids, and synthetic DNA fragments used in this paper are provided as Supplementary Data 1–3, respectively. Source data are provided with this paper.
Code availability
An automated protocol for the TREMD conformer generation protocol is available on GitHub https://github.com/ajfriedman22/SM_ConfGen. PyRosetta scripts for biosensor design by structural replacement are available at https://github.com/alisoncleonard/Structural-Replacement-Biosensor-Design. Scripts used to generate Supplementary Figs. 1 and 10 and to process raw deep sequencing data are available at https://github.com/WhiteheadGroup/Leonard_ComputationalDesign_Supplemental69.
References
National Institute on Drug Abuse. Drug overdose deaths: facts and figures. Natl. Inst. Drug Abuse. https://nida.nih.gov/research-topics/trends-statistics/overdose-death-rates (2024).
Opioid overdose. https://www.who.int/news-room/fact-sheets/detail/opioid-overdose (2025).
Prekupec, M. P., Mansky, P. A. & Baumann, M. H. Misuse of novel synthetic opioids: a deadly new trend. J. Addict. Med. 11, 256–265 (2017).
CDC. Fentanyl facts. Stop Overdose. https://www.cdc.gov/stop-overdose/caring/fentanyl-facts.html (2024).
New, Dangerous Synthetic Opioid in D.C., Emerging in Tri-State Area. DEA. https://www.dea.gov/stories/2022/2022-06/2022-06-01/new-dangerous-synthetic-opioid-dc-emerging-tri-state-area (2022).
CCENDU Drug Alert: Nitazenes. (Canadian Centre on Substance Use and Addiction, 2022).
De Vrieze, L. M. et al. In vitro structure-activity relationships and forensic case series of emerging 2-benzylbenzimidazole ‘nitazene’ opioids. Arch. Toxicol. 98, 2999–3018 (2024).
Holland, A. et al. Nitazenes-heralding a second wave for the UK drug-related death crisis? Lancet Public Health 9, e71–e72 (2024).
Krausz, R. M. et al. Shifting North American drug markets and challenges for the system of care. Int. J. Ment. Health Syst. 15, 86 (2021).
Uljon, S. Advances in fentanyl testing. Adv. Clin. Chem. 116, 1–30 (2023).
Krieger, M. S. et al. Use of rapid fentanyl test strips among young adults who use drugs. Int. J. Drug Policy 61, 52–58 (2018).
Lieberman, M., Badea, A., Desnoyers, C., Hayes, K. & Park, J. N. An urgent need for community lot testing of lateral flow fentanyl test strips marketed for harm reduction in Northern America. Harm. Reduct. J. 21, 115 (2024).
Green, H. H. New class of opioids that may be more potent than fentanyl emerges globally. The Guardian. https://www.theguardian.com/us-news/2024/sep/25/opioid-crisis-nitazenes-fentanyl (2024).
Steiner, P. J. et al. A closed form model for molecular ratchet-type chemically induced dimerization modules. Biochemistry 62, 281–291 (2023).
Ziegler, M. J. et al. Mandipropamid as a chemical inducer of proximity for in vivo applications. Nat. Chem. Biol. 18, 64–69 (2021).
Park, S. Y. et al. Agrochemical control of plant water use using engineered abscisic acid receptors. Nature 520, 545–548 (2015).
Melcher, K. et al. A gate-latch-lock mechanism for hormone signalling by abscisic acid receptors. Nature 462, 602–608 (2009).
Beltrán, J. et al. Rapid biosensor development using plant hormone receptors as reprogrammable scaffolds. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01364-5 (2022).
Park, S.-Y. et al. An orthogonalized PYR1-based CID module with reprogrammable ligand-binding specificity. Nat. Chem. Biol. 20, 103–110 (2024).
Leonard, A. C. & Whitehead, T. A. Design and engineering of genetically encoded protein biosensors for small molecules. Curr. Opin. Biotechnol. 78, 102787 (2022).
Kalogriopoulos, N. A. et al. Synthetic GPCRs for programmable sensing and control of cell behaviour. Nature 637, 230–239 (2024).
An, L. et al. Binding and sensing diverse small molecules using shape-complementary pseudocycles. Science 385, 276–282 (2024).
Lee, G. R. et al. Small-molecule binding and sensing with a designed protein family. bioRxiv. https://doi.org/10.1101/2023.11.01.565201 (2023).
Bick, M. J. et al. Computational design of environmental sensors for the potent opioid fentanyl. Elife 6, e28909 (2017).
Polizzi, N. F. & DeGrado, W. F. A defined structural unit enables de novo design of small-molecule-binding proteins. Science 369, 1227–1233 (2020).
Marchand, A. et al. Targeting protein–ligand neosurfaces with a generalizable deep learning tool. Nature 1, 10 (2025).
Glasgow, A. et al. Ligand-specific changes in conformational flexibility mediate long-range allostery in the lac repressor. Nat. Commun. 14, 1179 (2023).
Shover, C. L., Falasinnu, T. O., Freedman, R. B. & Humphreys, K. Emerging characteristics of isotonitazene-involved overdose deaths: a case-control study: a case-control study. J. Addict. Med. 15, 429–431 (2021).
Taoussi, O. et al. Human metabolism of four synthetic benzimidazole opioids: isotonitazene, metonitazene, etodesnitazene, and metodesnitazene. Arch. Toxicol. 98, 2101–2116 (2024).
Kanamori, T. et al. Metabolism of highly potent synthetic opioid nitazene analogs: N-ethyl-N-(1-glucuronyloxyethyl) metabolite formation and degradation to N-desethyl metabolites during enzymatic hydrolysis. Drug Test. Anal. https://doi.org/10.1002/dta.3705 (2024).
Walton, S. E., Krotulski, A. J. & Logan, B. K. A forward-thinking approach to addressing the new synthetic opioid 2-benzylbenzimidazole nitazene analogs by liquid chromatography-tandem quadrupole mass spectrometry (LC-QQQ-MS). J. Anal. Toxicol. 46, 221–231 (2022).
Maguire, J. B. et al. Perturbing the energy landscape for improved packing during computational protein design. Proteins 89, 436–449 (2021).
Robertson, N. R. et al. PYR1 biosensor-driven genome-wide CRISPR screens for improved monoterpenoid production in Kluyveromyces marxianus. bioRxiv 11, 623641 (2024).
Park, S.-Y. et al. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324, 1068–1071. https://doi.org/10.1126/science.1173041 (2009).
Vaidya, A. S. et al. Click-to-lead design of a picomolar ABA receptor antagonist with potent activity in vivo. Proc. Natl. Acad. Sci. USA 118, e2108281118 (2021).
Daffern, N., Francino-Urdaniz, I. M., Baumer, Z. T. & Whitehead, T. A. Standardizing cassette-based deep mutagenesis by Golden Gate assembly. Biotechnol. Bioeng. 121, 281–290 (2024).
Daffern, N. et al. GMMA can stabilize proteins across different functional constraints. J. Mol. Biol. 436, 168586 (2024).
Dixon, A. S. et al. NanoLuc complementation reporter optimized for accurate measurement of protein interactions in cells. ACS Chem. Biol. 11, 400–408 (2016).
Gagnot, G. et al. Core-modified coelenterazine luciferin analogues: synthesis and chemiluminescence properties. Chemistry 27, 2112–2123 (2021).
De Vrieze, L. M., Stove, C. P. & Vandeputte, M. M. Nitazene test strips: a laboratory evaluation. Harm. Reduct. J. 21, 159 (2024).
Dauparas, J. et al. Atomic context-conditioned protein sequence design using LigandMPNN. Nat. Methods 22, 717–723 (2025).
Kozel, T. R. & Burnham-Marusich, A. R. Point-of-care testing for infectious diseases: past, present, and future. J. Clin. Microbiol. 55, 2313–2320 (2017).
Ni, Y. et al. A plug-and-play platform of ratiometric bioluminescent sensors for homogeneous immunoassays. Nat. Commun. 12, 4586 (2021).
Socha, R. D. & Tokuriki, N. Modulating protein stability—directed evolution strategies for improved protein function. FEBS J. 280, 5582–5595 (2013).
Torres, S. V. et al. De novo designed proteins neutralize lethal snake venom toxins. Nature 639, 225–231 (2025).
Stove, C. New synthetic opioids: advances of receptor assays as tools for pharmacological characterization and analytical screening. Ann. Toxicol. Anal. 36, S18–S19 (2024).
Tian, H. et al. Unusually Broad-spectrum small-molecule sensing using a single protein scaffold. PNAS 122, e2519924122 (2025).
Santiago, J. et al. Modulation of drought resistance by the abscisic acid receptor PYL5 through inhibition of clade A PP2Cs. Plant J. 60, 575–588 (2009).
Dupeux, F. et al. A thermodynamic switch modulates abscisic acid receptor sensitivity. EMBO J. 30, 4171–4184 (2011).
Santiago, J. et al. Structural insights into PYR/PYL/RCAR ABA receptors and PP2Cs. Plant Sci. 182, 3–11 (2012).
Hao, Q. et al. The molecular basis of ABA-independent inhibition of PP2Cs by a subclass of PYL proteins. Mol. Cell 42, 662–672 (2011).
Guo, A. B., Akpinaroglu, D., Kelly, M. J. S. & Kortemme, T. Deep learning guided design of dynamic proteins. Science 388, eadr7094 (2025).
Ferrari, Á. J. R. et al. Large-scale discovery, analysis, and design of protein energy landscapes. bioRxivorg. https://doi.org/10.1101/2025.03.20.644235 (2025).
Kim, S. et al. PubChem 2025 update. Nucleic Acids Res. https://doi.org/10.1093/nar/gkae1059 (2024).
Okumura, H., Gallicchio, E. & Levy, R. M. Conformational populations of ligand-sized molecules by replica exchange molecular dynamics and temperature reweighting. J. Comput. Chem. 31, 1357–1367 (2010).
Leonard, A. C. et al. Rationalizing diverse binding mechanisms to the same protein fold: Insights for ligand recognition and biosensor design. ACS Chem. Biol. 19, 1757–1772 (2024).
Dupeux, F. et al. Modulation of abscisic acid signaling in vivo by an engineered receptor-insensitive protein phosphatase type 2C allele. Plant Physiol. 156, 106–116 (2011).
Beltrán, J. et al. Rapid biosensor development using plant hormone receptors as reprogrammable scaffolds. Nat. Biotechnol. 40, 1855–1861 (2022).
Daffern, N., Francino-Urdaniz, I., Baumer, Z. T. & Whitehead, T. A. Benchmarking cassette-based deep mutagenesis by Golden Gate assembly. bioRxiv 04, 536781 (2023).
Medina-Cucurella, A. V. & Whitehead, T. A. Characterizing protein-protein interactions using deep sequencing coupled to yeast surface display. Methods Mol. Biol. 1764, 101–121 (2018).
Wrenbeck, E. E. et al. Plasmid-based one-pot saturation mutagenesis. Nat. Methods 13, 928–930 (2016).
Mighell, T. L., Toledano, I. & Lehner, B. SUNi mutagenesis: scalable and uniform nicking for efficient generation of variant libraries. PLoS ONE 18, e0288158 (2023).
Lee, M. E., DeLoache, W. C., Cervantes, B. & Dueber, J. E. A highly characterized yeast toolkit for modular, multipart assembly. ACS Synth. Biol. 4, 975–986 (2015).
Steiner, P. J., Bedewitz, M. A., Medina-Cucurella, A. V., Cutler, S. R. & Whitehead, T. A. A yeast surface display platform for plant hormone receptors: toward directed evolution of new biosensors. AIChE J. 66, e16767 (2020).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Edelhoch, H. Spectroscopic determination of tryptophan and tyrosine in proteins. Biochemistry 6, 1948–1954 (1967).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Leonard, A. C. et al. Computational design of dynamic biosensors for emerging synthetic opioids. https://doi.org/10.5281/zenodo.15298584 (2025).
Leonard, A. C. et al. Computational design of dynamic biosensors for emerging synthetic opioids, Leonard_ComputationalDesign_2025_Supplemental. https://doi.org/10.5281/zenodo.17885439 (2025).
Acknowledgements
We would like to thank Yves Janin for kindly providing the luciferase prosubstrate, Hikarazine-108. ACL would like to thank the Interdisciplinary Quantitative Biology and Molecular Biophysics programs at the University of Colorado Boulder for ongoing support. TAW would like to thank M. Stammnitz for helpful discussions related to transduction mechanisms for ligand-dependent protein biosensors. Funding for this work was supported by: National Science Foundation NSF Award #2128287 (T.A.W.); National Science Foundation NSF Award #2128016 (S.R.C. and I.W.); National Science Foundation NRT Integrated Data Science Fellowship Award #2022138 (L.M.W.); NSF GRFP Award #1650115 (A.C.L.); NSF GRFP Award #2040434 (Z.T.B.); DARPA CERES Award#D24AC00011-05 (S.R.C., I.W., and T.A.W.); NIH award# R01GM123296 (L.M.W. and A.J.F.); NIH award #5T32GM145437 (L.M.W.); DOE GAANN Awards # P200A210136 and P200A240099 (C.L.M.).
Author information
Authors and Affiliations
Contributions
Non co-1st author trainees are listed in alphabetical order. Conceptualization: A.C.L., C.L.M., N.R.R., I.W., and T.A.W. Methodology: A.C.L., C.L.M., R.C., S.S., R.D., A.J.F., M.R.S., I.W., and T.A.W. Investigation: A.C.L., C.L.M., R.C., S.S., Z.T.B., R.D., A.J.F., N.R.R., N.S., J.W., and L.M.W. Visualization: A.C.L., C.L.M., R.C., S.S., I.W., and A.W. Funding acquisition: A.C.L., Z.T.B., S.C., M.R.S., I.W., and T.A.W. Project administration: I.W. and T.A.W. Supervision: S.C., M.R.S., I.W., and T.A.W. Writing – original draft: A.C.L., C.L.M., R.C., S.S., I.W., and T.A.W. Writing – review & editing: S.C., M.R.S., I.W., and T.A.W.
Corresponding authors
Ethics declarations
Competing interests
T.A.W., S.R.C., and I.W. have filed a provisional patent entitled REAGENTS AND SYSTEMS FOR GENERATING BIOSENSORS (US9738902B2; WO2011139798A2) covering some research in the present work. T.A.W. is a consultant for Inari Ag and serves on the scientific advisory board for Metaphore Biotechnologies and Alta Tech. S.R.C. and I.W. are cofounders of Living Sensors Inc., which has interests in sensing technologies. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Christophe Stove, who co-reviewed with Marthe M. Vandeputte, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Leonard, A.C., Lenert-Mondou, C., Chayer, R. et al. Computational design of dynamic biosensors for emerging synthetic opioids. Nat Commun 17, 1234 (2026). https://doi.org/10.1038/s41467-025-67994-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67994-w