Abstract
Eosinophilic granulomatosis with polyangiitis (EGPA) and severe eosinophilic asthma (SEA) share a Type 2 (T2) inflammatory signature but exhibit distinct pathophysiology. We hypothesized that EGPA involves additional inflammatory mechanisms, beyond T2 immunity, that drive its systemic manifestations and treatment resistance. Using single-cell RNA sequencing, we identify interferon (IFN-I)-driven inflammation in EGPA, in contrast to TNF predominant pathway activation in SEA. IL1B+MX1+ neutrophils in EGPA express IFN-stimulated genes and promote tertiary lymphoid structure formation with autoantibody production. In addition, other IFN-activated granulocytes, including APOC1+ eosinophils, SCN7A+ mast cells, and basophils, further contribute to immune dysregulation in EGPA, unlike TNF activated granulocytes in SEA. Longitudinal single-cell analysis of EGPA reveals an IGF1+ macrophage population linked to EGPA relapse. In animal models of both conditions, IGF1 blockade attenuates T2 inflammation, mucin production, and goblet cell hyperplasia, highlighting IGF1 as a possible therapeutic target in T2 inflammation disease.
Similar content being viewed by others
Introduction
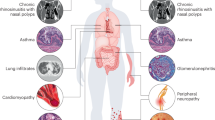
Eosinophilic granulomatosis with polyangiitis (EGPA), formerly known as Churg–Strauss syndrome, is a rare and life-threatening systemic vasculitis that almost invariably presents with asthmatic symptoms at onset. Its hallmarks include Th2/ILC2 polarization, blood and tissue eosinophilia, necrotizing eosinophil-rich granulomas, and vasculitis of small to medium-sized vessels. Patients exhibit elevated levels of eosinophil-promoting cytokines (IL-4, IL-5, IL-9, IL-13, IL-25, IL-33), eotaxins, and autoantibodies1, which may contribute to resistance to standard immunosuppressive therapies. As the disease progresses, widespread necrotizing vasculitis can cause irreversible damage to the lungs, heart, kidneys, intestines, and peripheral nerves2.
In contrast, asthma is a heterogeneous inflammatory airway disease often driven by allergic or other hypersensitivity mechanisms3. A subset of patients, characterized by elevated eosinophil levels, present with corticosteroid-dependent or corticosteroid-resistant features4. Early differentiation of EGPA from severe eosinophilic asthma (SEA) and the development of pathogenesis-based treatments remain major clinical challenges5.
Eosinophils are multifunctional granulocytes that can damage a wide spectrum of tissues through the release of cytotoxic granule proteins6. These include Major Basic Protein (MBP, encoded by PRG2), which drives airway remodeling, subepithelial fibrosis, and ciliated cell destruction7; Eosinophilic Cationic Protein (ECP, RNASE3) and Eosinophil-derived Neurotoxin (ENT, RNASE2), which contribute to cardiomyopathy and neuropathy8,9; and Eosinophilic Peroxidase (EPO, EPX). Furthermore, eosinophils can generate thrombin (encoded by F2) and induce tissue factor exposure on endothelial cells, promoting platelet adhesion and thrombosis10,11. Eosinophil NADPH oxidase (encoded by NOX1-5 and DUOX1/2 genes) and EPO also generate reactive oxygen species (ROS), promoting lipoperoxidation and atheromatous plaque formation and destabilization12.
Several monoclonal antibodies that target T2 and eosinophilic pathways are approved for SEA, including mepolizumab and reslizumab (anti-IL-5), benralizumab (anti-IL-5Rα), dupilumab (anti-IL4Rα, target IL-4 and IL-13) and omalizumab (anti-IgE). These biologics reduce exacerbation rates and can have a corticosteroid-sparing effect13. They have also been trialled in EGPA. In a randomized clinical trial, mepolizumab administration over 52 weeks significantly improved remission rates compared to placebo (32% vs. 3%) in patients receiving stable corticosteroids14. Benralizumab, that targets and IL-5R has been shown to be equally effective in reducing relapse and more effective in sparing oral corticosteroid dose15.
Beyond eosinophil pathways, B cells contribute to pathogenesis by generating autoantibodies such as ANCA (antineutrophil cytoplasmic antibody), which can activate neutrophils16,17. Activated neutrophils infiltrate vessel walls, releasing ROS and toxic enzymes that cause endothelial necrosis. They also release factors that activate the alternative complement pathway, generating C5a fragments, which in turn chemotactically recruit more neutrophils and prime them for ANCA activation18. Consequently, B cell depletion by rituximab has also shown efficacy EGPA, improving outcomes and reducing corticosteroid requirements19. Despite these advances, significant therapeutic challenges remain; for instance, mepolizumab did not induce remission in 47% of EGPA patients, and relapses occurred in over 50% of rituximab-treated patients by 24 months14,20. These limitations highlight the need for a deeper understanding of the underlying cellular and molecular mechanisms to identify potential therapeutic targets.
In addition to immune dysregulation, airway epithelial remodeling, particularly goblet cell hyperplasia, is a key feature of T2-driven airway diseases21,22. While T2 inflammation is a primary driver of mucus hypersecretion, other mechanisms likely contribute. Insulin-like growth factor 1 (IGF1), a critical regulator of fibrosis and tissue remodeling, has been implicated in promoting epithelial cell proliferation and mucin production23,24,25,26. In SEA, IGF1 signaling enhances goblet cell hyperplasia and mucus secretion27. Given that airway remodeling also occurs in EGPA, IGF1 may represent a common driver of this process across T2-driven diseases, suggesting shared pathological pathways that could be therapeutically targeted.
In this work, we employ single-cell RNA sequencing (scRNA-seq) to comprehensively analyze the cellular landscape of blood, bronchoalveolar lavage fluid (BALF), and bronchial biopsies from patients with EGPA or SEA. We identify both shared and distinct immune characteristics between the two diseases, providing promising insights into their pathogenesis. These findings suggest potential therapeutic targets capable of simultaneously suppressing T2 inflammation, eosinophilic infiltration, and airway remodeling.
Results
Single-cell atlas of human airways reveals distinct immune cell composition and infiltration in EGPA
Our study began with the characterization of patient cohorts and single-cell RNA sequencing to establish the cellular profiles of SEA and EGPA, providing a foundation for identifying disease-specific mechanisms. We recruited a cohort including controls (n = 6), SEA patients (n = 13; 50% corticosteroid-dependent), and EGPA patients (n = 9) for scRNA-seq analysis (Supplementary Table 1, and Supplementary Fig. 1). Controls were under investigation for chronic cough but had no history of pulmonary disease, infections, or other immune-related conditions, minimizing potential confounders. SEA patients met the GINA 2019 guidelines, requiring fluticasone propionate treatment and exhibiting sputum eosinophilia >3%, indicating poorly controlled asthma. EGPA diagnosis followed the ACR/EULAR 2022 and MIRRA criteria (Supplementary Fig. 2a). The EGPA cohort included both ANCA-positive (n = 1) and ANCA-negative (n = 8) patients, reflecting the disease’s heterogeneity. At sampling, all EGPA patients were on inhaled corticosteroids, with eight on oral corticosteroids. Additionally, five received immunosuppressants (azathioprine, n = 2; methotrexate, n = 2; mycophenolate mofetil, n = 1). No patients were on biologics.
Representative high-resolution CT scans and histology of bronchial biopsies revealed more extensive vascular damage in EGPA compared to SEA or controls (Supplementary Fig. 2b). Oxygen saturation (SpO2) was significantly lower in EGPA than in SEA (P = 0.001, Supplementary Table 1), consistent with impaired gas exchange. Fractional exhaled nitric oxide (FeNO) and serum total IgE levels were significantly elevated in both EGPA and SEA compared to controls, aligning with T2-skewed immunity. However, blood and BALF eosinophil counts and proportions, along with BALF neutrophil proportions, were significantly increased in EGPA versus SEA and controls (Fig. 1a). Analysis of individual patient data showed that this eosinophil and neutrophil enrichment in BALF was particularly marked in 4 of 9 EGPA patients, suggesting heterogeneity in cellular infiltration within the cohort.
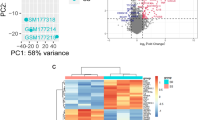
a Clinical and immunological characteristics of the study cohort. Box plots compare FeNO, serum total IgE and CRP levels, and proportions or counts of eosinophils and neutrophils in sputum, BALF and blood between control donors and patients with SEA or EGPA. Each data point represents one individual participant (biological replicate; Control, n = 6; SEA, n = 13; EGPA, n = 9). b Schematic of the experimental design for scRNA sequencing across sample types (blood, BALF, bronchial biopsy) and groups (28 participants, 71 samples). c UMAP plot displaying 354,353 cells from 28 participants separated into 17 major cell types. Cell types include epithelial cells, neutrophils, eosinophils, macrophages, T & NK (natural killer) cells, and others. d Bar plots showing proportions of major cell types across groups, with statistical comparisons on the right. For BALF samples: Control, n = 6; SEA, n = 9; EGPA, n = 9. For bronchial biopsy samples: Control, n = 5; SEA, n = 7; EGPA, n = 8 (biologically independent samples). Box plots in (a, d) show median (centre line), interquartile range (boxes), and whiskers extending to 1.5× IQR, outliers as individual points. Statistical analyses used two-sided Kruskal–Wallis test with Dunn’s post-hoc test and Bonferroni adjustment. Significance: *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant. Detailed statistics, including overall Kruskal–Wallis test results (χ², η²H), Z-values, and 95% confidence intervals, are provided in Supplementary Data 1. BALF bronchoalveolar lavage fluid CRP C-reactive protein, EGPA eosinophilic granulomatosis with polyangiitis, FeNO fractional exhaled nitric oxide, IgE immunoglobulin E, IQR interquartile range, NK natural killer, SEA severe eosinophilic asthma, UMAP uniform manifold approximation and projection. Control: chronic cough without pulmonary disease, infections, or immune-related conditions.
We generated scRNA transcriptomes using BD Biosciences Single-Cell Multiomics Reagents to (Fig. 1b, and Supplementary Fig. 2c, d). After pre-processing and quality control, 354,353 high-quality transcriptomes were partitioned into 17 major cell clusters. Most cell types were identified across all patients, with negligible batch effects (Fig. 1c, and Supplementary Fig. 2e–g). Immune infiltration in bronchial biopsies was markedly higher in EGPA patients compared to both SEA and control groups. Eosinophils and neutrophil-2 populations were selectively enriched in the BALF of EGPA patients, confirming a distinctive inflammatory profile involving both lineages (Fig. 1d). These initial findings provided a fundamental map of the cellular landscape in SEA and EGPA, highlighting key differences in immune cell infiltration and pointing to specific disease mechanisms beyond canonical T2 inflammation.
Interferon-activated granulocytes define EGPA pathology beyond T2 inflammation
While granulocytes are major effector cells in both SEA and EGPA, their compositional and functional heterogeneity in affected tissues remain uncharacterized. Our scRNA-seq analysis revealed distinct patterns of granulocyte subsets and interferon activation, providing key mechanistic insights that fundamentally distinguish EGPA pathophysiology from that of SEA.
We resolved distinct subsets of neutrophils, eosinophils, basophils, and mast cells across blood, BALF, and bronchial biopsies, uncovering disease-specific transcriptional programs that differentiate SEA from EGPA (Fig. 2a, and Supplementary Fig. 3a). We further defined the signature genes, enriched transcription factors, granule proteins, chemokines, cytokines, and their receptors expressed by these granulocytes (Supplementary Fig. 3b–d).
a UMAP plot of 14 granulocyte subtypes, indicating Neutrophil-1 and −2 subsets. b Bar plots showing cellular sources per subtype; blocks represent individual samples color-coded by group. c UMAP plots of inflammatory cytokine, chemokine, and type I interferon-stimulated gene expression in granulocytes; color intensity reflects normalized UMI counts. d Immunofluorescent staining and quantification of TNF and IL-1β in bronchial CD66b⁺MPO⁺ neutrophils (independent biological replicates; Control n = 10, SEA n = 10, EGPA n = 10). Scale bars: 100 µm and 10 µm. e Immunofluorescent staining and quantification of IFN-α and JAK1 in bronchial CD66b⁺MPO⁺ neutrophils (biological replicates; Control n = 9, SEA n = 10, EGPA n = 10). Scale bars: 100 µm and 10 µm. f Dot plots of differentially expressed genes in cirEos versus Eos across groups. Dot size: cell fraction; color: relative expression. g Bar plots of Annexin V⁺ PI⁺ (late apoptotic) and Annexin V⁺ PI⁻ (early apoptotic) cells under normoxia, hypoxia, or hypoxia+PX478 (n = 3 independent experiments, biological replicates). h Bar plots of ECP and EPX concentrations (n = 3 independent experiments, biological replicates). i Schematic illustrating transcriptomic heterogeneity and oxygen saturation promotes eosinophil degranulation and inflammatory responses. Data presented as mean ± SD or median with IQR. d Two-sided one-way ANOVA with Tukey’s post-hoc test. e Two-sided Kruskal–Wallis test with Dunn’s post-hoc and Bonferroni correction. g, h Two-sided one-way ANOVA with Tukey’s post-hoc test. *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant. Exact P values in Supplementary Data 1. cirEos circulatory eosinophils, ECP eosinophil cationic protein, EGPA eosinophilic granulomatosis with polyangiitis, Eos tissue eosinophils, EPX eosinophil peroxidase, IFN-α interferon-α, IL-1β interleukin-1β, IQR interquartile range, JAK1 Janus kinase 1, MPO myeloperoxidase, PI propidium iodide, SD standard deviation, SEA severe eosinophilic asthma, TNF tumor necrosis factor, UMAP uniform manifold approximation and projection, UMI unique molecular identifier.
Two subsets of inflammatory neutrophils (subsets 5 and 6) were enriched in the blood and bronchial biopsies of SEA patients compared to controls or EGPA patients (Fig. 2b). These neutrophils expressed inflammatory cytokines (IL1B, TNF) and chemokines (CXCL2, CXCL3, CCL3, CCL4, CXCL8, CXCL16). In contrast, a distinct Neutrophil-IL1B/MX1 (subset 8) subset was enriched in the bronchial biopsies and BALF of EGPA patients. These cells co-expressed high levels of IL1B and type I interferon (IFN-I) stimulated genes (IRF7, IFIT1, IFIT2, ISG15, MX1) (Fig. 2c, and Supplementary Fig. 3d). Immunofluorescent staining confirmed increased TNF and IL-1β in neutrophils in both diseases. In contrast, TNF was more enriched in SEA, whereas IL-1β was more enriched in EGPA (Fig. 2d). Type I interferons are critical for anti-viral immunity, but their sustained signaling is implicated in vascular damage28,29. Consistently, we found IFN-I (IFN-α) levels were uniquely and significantly elevated in EGPA compared to SEA and controls (Fig. 2e). As JAK1 is activated by a broad-spectrum of cytokines and can activate all STATs30. We assessed its expression and found it significantly elevated in neutrophils in both diseases (Fig. 2e), consistent with the inflammatory state.
Beyond neutrophils, we observed distinct functional alterations in mast cells, basophils, and eosinophils. We identified four mast cell subsets and one basophil subset (Supplementary Fig. 4a–d). In SEA, the Mast cell-FOSB subset was highly enriched and exhibited upregulation of TNF signaling pathways, including NF-κB activation (Supplementary Fig. 4c, e). Conversely, EGPA showed an increased proportion of Mast cell-SCN7A and Basophil-IL3RA subsets, both characterized by activation of IFN-I signaling pathways (Supplementary Fig. 4c, f–h). Additionally, a significant enrichment of IFN-activated eosinophils (Eosinophil-APOC1) was observed in EGPA, with robust upregulation of IFIT1, IFIT2, IFIT3, and IFI44L, confirming strong IFN-I activation (Supplementary Fig. 4i–o). Collectively, these findings highlight the critical role of systemic IFN-I activation in shaping the granulocyte response, defining a key element of EGPA pathology that extends beyond canonical T2 inflammation.
The oxygen microenvironment dynamically controls eosinophil survival and activation
Eosinophils are key effector cells in SEA and EGPA, yet their functional states shift dynamically upon tissue infiltration. While our transcriptomic analysis revealed disease-specific eosinophil subsets, how these subsets functionally adapt upon transitioning from circulation into the inflamed airway remained unclear. We therefore investigated whether oxygen availability–a key microenvironmental factor–regulates eosinophil activation states in these diseases.
Flow cytometry confirmed a significant increase in eosinophils in the blood and BALF of EGPA patients (Supplementary Fig. 5a, b). A striking difference was the exclusive expression of SELL (CD62L) on circulating eosinophils and its downregulation upon tissue entry, indicating a transition to an activated, tissue-adaptive phenotype (Fig. 2f, and Supplementary Fig. 5c). Immunohistology revealed the highest peri-vascular eosinophil staining (EPX+) in EGPA patients (Supplementary Fig. 5d). Furthermore, lung-infiltrating eosinophils displayed upregulation of extracellular matrix remodeling genes (PLAUR, ENO1), chemokine receptors (CCR3), and T2 immune receptors (IL1RL1, IL4R, IL13RA1), reflecting an enhanced pro-inflammatory state upon tissue entry (Fig. 2f, and Supplementary Fig. 5e).
Since arterial hypoxia is common in SEA exacerbations and EGPA (Supplementary Table 1), we hypothesized that oxygen availability regulates eosinophil survival and activation during this transition. scRNA-seq showed that circulating eosinophils in SEA and EGPA exhibited an enhanced hypoxic response, with elevated HIF1A expression, compared to controls. These cells also showed increased expression of genes involved in cell death (CASP8) and cytotoxicity (RNASE2). Notably, PRSS33 expression was significantly higher in circulating eosinophils from EGPA patients, suggesting a potential role for hypoxia in promoting eosinophil survival (Fig. 2f). In contrast, lung-infiltrating eosinophils downregulated hypoxia-associated signatures and instead displayed increased expression of degranulation and extracellular matrix remodeling genes. To functionally validate these findings, we differentiated mouse bone marrow-derived eosinophils in vitro (Supplementary Fig. 6a, b). Under normoxia with IL-5 withdrawal (mimicking alveolar conditions), eosinophils underwent extensive degranulation and apoptosis (Supplementary Fig. 6c). Hypoxia, however, suppressed degranulation, an effect abolished by pharmacological inhibition of HIF1A (PX478) (Fig. 2g, h, and Supplementary Fig. 6d, e).
These results reveal a dynamic, oxygen-dependent regulatory mechanism: normoxia in the alveolar microenvironment promotes eosinophil degranulation, directly contributing to inflammation, whereas hypoxia in the circulation maintains eosinophil viability and functional potential prior to infiltration (Fig. 2i). These findings establish oxygen saturation as a fundamental regulator of eosinophil function, shaping their survival, activation, and ultimately inflammatory impact in SEA and EGPA.
FCRL4 + CD11C + B cells and tertiary lymphoid structures drive localized autoimmunity in EGPA
Having characterized granulocyte heterogeneity and interferon activation in EGPA, we next investigated the adaptive immune responses. Our analysis revealed crucial mechanisms underlying the unique pathology of EGPA.
We identified three B cell subsets and one plasma cell population. RNA velocity analysis indicated a possibly transition of circulatory memory B cells into FCRL4+ memory B cells following their migration to bronchial tissues and BALF (Fig. 3a). These FCRL4+ B cells showed increased expression of AICDA, FCRL4, FCRL5, ITGAX/CD11C, and TNFRSF13B (BAFF receptor)–signatures of autoantibody-producing atypical memory B cells or age-associated B cells (ABCs)31,32(Fig. 3b). Immunofluorescent staining revealed a substantial expansion of FCRL4+ memory B cells in EGPA bronchial biopsies, where they were localized within tertiary lymphoid-like structures (TLS) containing neutrophils and T cells (Fig. 3c, d). These structures were rarely observed in SEA or control samples, suggesting that TLS formation in EGPA provides an immune niche that supports chronic B cell activation, autoreactive B cell survival, and local autoantibody production.
a UMAP plot displaying 3 B cell subtypes (Naïve B, Memory B, FCRL4+ Memory B) and 1 plasma cell subset. RNA velocity analysis, indicated by black arrows, highlights inferred cellular differentiation trajectories and potential migration within B cell subsets from blood to bronchial biopsies and BALF. b UMAP plots illustrating expression levels of key marker genes for B and plasma cell subsets. Color intensity proportional to normalized UMI counts. c Bar plots exhibiting proportional cellular sources for each B and plasma cell subtype. Blocks represent different samples, color-coded by groups (Control, SEA, EGPA) and sample types (PBMC, Bronchial biopsy, BALF). d Immunofluorescent staining and quantification comparing abundance of CD40LG+CD4+ T cells, FCRL4+CD20+ B cells, and MPO+ neutrophils at bronchial mucosae (biological replicates; Control n = 10, SEA n = 7, EGPA n = 10). e Dot plots comparing levels of anti-ANCA (anti-MPO, anti-PR3), anti-dsDNA, and anti-chromatin autoantibodies measured in sputum samples from control (n = 10), SEA (n = 26), and EGPA (n = 47) participants, and in serum samples from EGPA participants (n = 47) (biological replicates). Serum samples assessed systemic autoantibody levels, while sputum samples assessed local antibody production. f Violin plots showing expression of TNFSF13B and Fc gamma receptor genes across granulocyte clusters for Control, SEA, and EGPA groups. Data presented as mean ± SD or median with IQR. d Two-sided one-way ANOVA with Tukey’s post-hoc test. e Two-sided Kruskal–Wallis test with Dunn’s post-hoc and Bonferroni correction for sputum comparisons; two-sided Mann–Whitney U test for sputum EGPA vs. serum EGPA comparisons. *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant. Exact P values in Supplementary Data 1. ANCA anti-neutrophil cytoplasmic antibody, FCRL4 Fc receptor-like 4, PR3 proteinase 3, TLS tertiary lymphoid structure, TNFSF13B tumor necrosis factor superfamily member 13B.
To assess the functional output of these structures, we measured autoantibodies and found significantly elevated levels of ANCA (anti-MPO, anti-PR3), anti-dsDNA, and anti-chromatin autoantibodies in the sputum of EGPA patients compared to controls and SEA participants (Fig. 3e). Notably, while only one EGPA patient was serum ANCA-positive, multiple patients showed ANCA positivity in sputum, indicating that localized autoantibody production in the airway can occur independently of systemic seropositivity.
BAFF (encoded by TNFSF13B), a critical survival factor for autoreactive B cells, was most highly expressed by the EGPA-specific Neutrophil-IL1B/MX1 cells enriched in bronchial biopsies (Fig. 3f). This suggests neutrophils may actively contribute to TLS maintenance and autoreactive B cell expansion. Furthermore, these neutrophils highly expressed Fc gamma receptors, potentially regulated by anti-neutrophil antibodies in the alveoli (Fig. 3f). Collectively, these findings imply a neutrophil-B cell feedback loop in EGPA pathogenesis: neutrophils provide BAFF-mediated survival signals to B cells, and in turn, B cell-derived autoantibodies may drive neutrophil activation, degranulation, and inflammatory tissue damage.
We also identified 22 subsets of T cells and innate lymphoid cells (ILCs) (Supplementary Fig. 7a). Although ILC2 cells were not captured, likely due to their low abundance, we observed a significant enrichment of Th2 cells in both SEA and EGPA, suggesting they may be the dominant source of T2 cytokines (Supplementary Fig. 7b). Interestingly, ILC1 cells were enriched in (Supplementary Fig. 7b), correlating with the observed interferon pathway activation and indicating a more complex inflammatory landscape than a purely T2-driven disease (Supplementary Fig. 7b).
Several effector T cell populations–including ENTPD1+CD4 and ENTPD1+CD8 Teff cells, CD40LG+CD4 Teff cells, GZMK+CD8 Teff cells, ZNF683+CD8 Trm cells-were specifically enriched in EGPA bronchial biopsies (Supplementary Fig. 7b). While the precise roles of these cells require further study, the reciprocal expression of co-stimulatory molecules and receptors on FCRL4+ memory B cells and these effector T cells suggests extensive functional interactions between them (Supplementary Fig. 7c, d).
These results unveil distinct, localized adaptive immune mechanisms centered on TLS formation and in-situ autoantibody production, which are central to EGPA pathogenesis and represent a significant departure from the immunology of typical T2 inflammation.
Corticosteroid and immunosuppressant treatment modulate immune cell profiles but fail to prevent EGPA relapse
Having identified numerous immune cells change unique to EGPA, we next investigated whether their persistence was associated with treatment relapse. We recruited additional EGPA patients receiving corticosteroid and immunosuppressant treatment (Cs), categorized as being in remission (Cs-remission) or experiencing relapses (Cs-relapse) (Supplementary Fig. 8a, and Supplementary Tables 2 and 3).
Post-treatment blood analyzes showed that total white blood cell (WBC) counts were significantly reduced in both groups. The remaining WBCs showed relatively decreased eosinophil but increased neutrophil proportions, consistent with the known effects of corticosteroids in promoting neutrophil survival and eosinophil death33. In sputum, macrophage percentages increased significantly in both groups. Although blood eosinophils decreased in both, the reduction in sputum eosinophil percentage was statistically significant only in Cs-remission group, suggesting a differential response to comparable steroid therapy in Cs-relapse patients (Supplementary Fig. 8b). Lung function improved after treatment in Cs-remission patients, as measured by forced expiratory volume in 1 s (FEV1%), forced vital capacity (FVC%), and the FEV1/FVC ratio, but not in Cs-relapse patients (Supplementary Fig. 8c–e).
To characterize these changes at cellular level, we performed scRNA-seq on BALF and bronchial biopsies from EGPA patients in remission or relapse (Supplementary Fig. 9a). Treatment significantly altered immune cell proportions in BALF in both groups. The Cs-relapse patients showed elevated proportions of DC, pDC, and macrophages, but reduced proportions of eosinophils, T, NK, and B cells (Supplementary Fig. 9b). Immunofluorescence confirmed a decrease in FCRL4+CD11C+ B cells in both treatment groups (Supplementary Fig. 9c).
Analysis of signature genes expression revealed that Cs treatment, regardless of clinical outcomes, significantly reduced expression of ISGs, TNFSF13B (BAFF), IL1B, and the IgG high affinity receptor FCGR1A in Neutrophil−2 cells. In eosinophils, treatment effectively reduced expression of the chemokine receptor CCR3 and key T2 cytokine receptors (IL5RA, IL1RL1, IL3RA, IL4R and IL13RA1) (Supplementary Fig. 9d). Collectively, these data demonstrate that corticosteroid and immunosuppressant treatment effectively modulates multiple immune pathways in EGPA. However, these targeted changes were insufficient to prevent disease relapse in a subset of patients. This indicates that while the identified immune alterations contributed to EGPA pathogenesis, the persistence of treatment-resistant populations or alternative pathways likely underlies clinical relapse.
Enriched IGF1 + macrophages sustain airway remodelling and inflammation in relapsing EGPA
Following treatment, we identified a distinct population of IGF1+ macrophages (IGF1+ Mac) that was significantly enriched in Cs-relapse EGPA patients compared to those in remission (Fig. 4a, and Supplementary Fig. 9e). This subset was characterized by high IGF1 expression (Fig. 4b). Functional analysis indicated that IGF1+ Mac cells are primarily involved in IGF transport and uptake via IGF-binding proteins (IGFBPs), underscoring their potential role in IGF signaling and tissue remodeling (Fig. 4c). Gene expression profiling revealed elevated expression of HP, IGF1, and RETN in IGF1+ Mac cells, with levels significantly higher in Cs-relapse than remission patients (Fig. 4d, e). Immunofluorescence staining confirmed a significantly higher proportion of IGF1+ Mac cells in the Cs-relapse group (Fig. 4f).
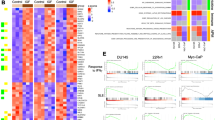
a UMAP plots of 12 macrophage and DC subtypes from baseline and follow-up samples. IGF1⁺Macrophage cluster (cluster 3) highlighted. b Volcano plot of DEGs in IGF1⁺ macrophages between baseline and follow-up. Upregulated (red), downregulated (blue), stable (grey) genes shown. c Bar plots of top enriched Reactome pathways in IGF1⁺ macrophages from DEGs (two-sided Wilcoxon rank-sum test, adjusted P values). Pathway enrichment of top 20 upregulated genes via Enrichr (Reactome_Pathways_2024, hypergeometric test, unadjusted P values). d UMAP plots of key marker gene expression (HP, IGF1, RETN) in macrophage subsets; color intensity reflects normalized UMI counts. e Violin plots of IGF1, RETN, HP expression across disease phases (EGPA baseline, Cs-remission, Cs-relapse) in macrophages. f Immunofluorescent staining and quantification of IGF1⁺CD68⁺ macrophages in bronchial mucosae (biological replicates; EGPA n = 15, Cs-remission n = 3, Cs-relapse n = 5). Scale bars: 100 μm (upper), 20 μm (lower). g UMAP plots of 11 epithelial cell subsets from combined samples. h Heatmap of relative enrichment (observed/expected Ro/e) of epithelial subtypes across groups and sample types. i Dot plot of reciprocal epithelial ligand-receptor expression across subsets. Interaction pairs linked by color-coded lines; dot size reflects expression fraction, color intensity shows relative expression. j Violin plots of marker gene expression for goblet cell subsets (Goblet-1, Goblet−2) and ionocytes across groups. k Immunofluorescent staining and quantification of MUC5AC⁺ epithelial cells (biological replicates; EGPA n = 10, Cs-remission n = 3, Cs-relapse n = 5). Scale bar: 100 μm. Data presented as median with IQR. f, k Two-sided Kruskal–Wallis test with Dunn’s post-hoc and Bonferroni correction. *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant. Exact P values in Supplementary Data 1. Cs-relapse corticosteroid relapse, Cs-remission corticosteroid remission, DC dendritic cell, DEGs differentially expressed genes, IGF1 insulin-like growth factor 1, MUC5AC mucin 5AC, Ro/e ratio of observed to expected.
Insulin like growth factor 1 (IGF1) is a pleotropic, growth-promoting factor associated with asthma risk and severity34,35. Haptoglobin (HP) captures free plasma haemoglobin after haemolysis, facilitating hepatic recycling of heme iron and preventing oxidative damage36. Resistin (RETN) is a macrophage-derived proinflammatory adipokine linked to obesity and insulin resistance37. The concerted upregulation of IGF1, HP, and RETN in IGF1+Mac cells from Cs-relapse patients suggests a critical role for IGF1 signaling in sustaining airway remodeling and inflammation even during treatment. This macrophage population may therefore represent a key cellular driver of treatment-resistant disease, warranting further investigation.
Persistent goblet hyperplasia and mucin production characterize treatment-refractory EGPA airways
Our analysis of epithelial cells from baseline EGPA, Cs-remission, and Cs-relapse cohorts investigated whether alveolar epithelial remodeling was associated with treatment responsiveness. We identified 11 epithelial subsets across these groups (Fig. 4g, and Supplementary Fig. 10a). Goblet cell hyperplasia was a prominent feature in EGPA, with goblet-1 cells showing the highest relative enrichment in Cs-relapse patients, indicating persistent mucus hypersecretion despite corticosteroid therapy (Fig. 4h). In contrast, Cs-remission patients exhibited a relative reduction in goblet-1 cell proportions compared to baseline EGPA, suggesting a partial resolution of hyperplasia.
Interestingly, IGF1 was also specifically and highly expressed by ionocytes. Inmunofluorescent sating confirmed an increased proportion of CFTR+ epithelial cells in EGPA compared to control and SEA groups (Supplementary Fig. 10b). While Cs-remission patients showed a significant reduction in CFTR+ ionocytes compared to baseline EGPA, Cs-relapse patients retained elevated levels, suggesting persistent ionocyte activation in relapsing disease. The receptor for IGF1, IGF1R, was highly expressed by lung epithelial cells, particularly basal cells, mature ciliated cells, ionocytes, and neuroendocrine cells (Fig. 4i).
We found that Cs treatment effectively suppressed IDO1, CXCL9, CXCL10, and IL19 expression in goblet-2 cells. However, it failed to suppress mucin (MUC2, MUC5AC) expression in goblet-1 cell. Furthermore, treatment increased the expression of IGFBP5 and IGFBP7, IGF-binding proteins that prolong the half-life of IGF1 (Fig. 4j, k).
IGF1 promotes goblet hyperplasia, mucin production, and IL-25 secretion in alveolar epithelial cells
Our initial scRNA-seq cohort captured 6 fibroblast subsets, 4 endothelial subsets, and 1 Schwann cell population (Supplementary Fig. 10c). We found that adventitial fibroblasts, which reside in the outer layer of blood vessel walls, expressed IGF1 and CCL11 (Eotaxin) (Supplementary Fig. 10d). Furthermore, these fibroblast in both SEA and EGPA significantly upregulated fibrosis-related genes–including POSTN, SERPINE2, which promotes type I and type III Collagen accumulation (COL1A1, COL3A1)38, and IGFBP3, which prolongs IGF1 half-life–compared to controls (Supplementary Fig. 10e). Critically, IGF1 expression was significantly elevated in adventitial fibroblasts in EGPA compared to both SEA and controls, while no significant difference was observed between SEA and controls (Supplementary Fig. 10f). Immunofluorescence staining confirmed elevated peri-vascular expression of IGFBP3 and IGFBP7 in both EGPA and SEA, reflecting their potential role in sustaining IGF1 activity and promoting tissue remodeling (Supplementary Fig. 10g).
IGF1 expression was also significantly increased in alveolar epithelial cells in both SEA and EGPA. Cs treatment decreased IGF1 expression in alveolar epithelial cells from remission, but not relapse, patients (Fig. 5a). Consistent with this, IGF1 protein concentrations were significantly elevated in the sputum of EGPA patients compared to controls and SEA patients (Fig. 5b). Given IGF1’s pleiotropic effects and its increased expression by the remnant tissue macrophages, ionocytes, and adventitial fibroblasts in both diseases, we investigated its induction using an alveolar epithelial air-liquid interface culture system (Fig. 5c). We found that IL-13 and IL-33 promoted IGF1, IGF1R, and IGFBP3 expression (Fig. 5c). Conversely, stimulation of alveolar epithelial cells with IGF1 directly induced marked goblet cell hyperplasia, mucin production, and IL−25 expression (Fig. 5d). IL-25 (IL-17E) is a cytokine that augments T2-mediated inflammation, characterized by eosinophil and Th2 memory cell infiltration39. We also observed high IGF1R expression in neutrophils and eosinophils, consistent with a positive association between sputum IGF1 concentrations and eosinophil percentages in EGPA patients (Fig. 5e). Based on these findings, we propose that an IGF1-IL25 axis contributes to disease exacerbation in SEA and EGPA (Fig. 5f).
a Immunofluorescent staining and quantification of IGF1⁺ epithelial cells at bronchial mucosae from control (n = 6), SEA (n = 9), EGPA (n = 10), Cs-remission (n = 3), and Cs-relapse (n = 5) groups (biological replicates). Scale bar: 100 μm. b Bar plots of IGF1 concentrations in sputum samples from Control (n = 13), SEA (n = 23), and EGPA (n = 21) (biological replicates). c Schematic of EGPA airway epithelial cells in ALI culture stimulated with IL-13, IL-33, or medium (Control). Bar plots show relative expression of IGF1, IGF1R, and IGFBP3, and IGF1 concentrations under different conditions. Data from 3 independent experiments (biological replicates). d Schematic of ALI system with IGF1 stimulation versus control. Representative histological (HE, PAS) and immunofluorescent (MUC5AC with DAPI) staining demonstrate morphological changes and mucin production (goblet hyperplasia) in ALI cultures with medium (Control) or IGF1 stimulation. Scale bars: 50 μm (HE, PAS), 20 μm (MUC5AC). Bar plots of IL-25, IL-33, and TSLP concentrations in culture supernatant at time points (Day 9, 13, 17, 21). Data from 3 independent experiments (biological replicates). e Scatter plot showing positive correlation between eosinophil abundance and IGF1 concentration in sputum from EGPA patients (n = 21). f Schematic of proposed mechanism where IGF1 promotes goblet hyperplasia and augments T2-mediated inflammation through IGF1-IL25 loop, contributing to disease exacerbation in asthma and EGPA. Data presented as mean ± SD. a–c Two-sided one-way ANOVA with Tukey’s post-hoc test. d Two-sided unpaired t-test. e Two-sided Pearson correlation test. *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant. Exact P values in Supplementary Data 1. ALI air-liquid interface, HE hematoxylin and eosin, IGF1R insulin-like growth factor 1 receptor, IGFBP3 insulin-like growth factor binding protein 3, PAS periodic acid-Schiff, TSLP thymic stromal lymphopoietin.
IGF1/IGF1R signalling drives eosinophilic inflammation and airway remodelling in experimental models of SEA and EGPA
To investigate the role of IGF1 signaling in asthma pathogenesis, we analyzed the expression patterns of IGF1R and IL-17 receptors (IL17RA and IL17RB) in our single-cell RNA sequencing dataset. Uniform manifold approximation and projection (UMAP) analysis revealed widespread IGF1R expression across multiple cell types, particularly in epithelial cells and eosinophils, while IL17RA and IL17RB showed more restricted expression (Supplementary Fig. 11a). We then established an ovalbumin (OVA)-induced asthma model, where mice were sensitized with OVA on days 1 and 15, followed by OVA challenges on days 21–24 (Supplementary Fig. 11b). Histological analysis showed that OVA challenge induced substantial inflammatory cell infiltration and goblet cell hyperplasia, effects exacerbated by recombinant IGF1 administration but significantly ameliorated by anti-IGF1 treatment (Supplementary Fig. 11c). OVA challenge also significantly increased IGF1 expression in lung tissues (Supplementary Fig. 11d). Consistent with this, anti-IGF1 treatment markedly reduced eosinophil counts (Supplementary Fig. 11e), improved airway hyperresponsiveness to acetylcholine (Supplementary Fig. 11f), and decreased T2 cytokine production (IL-4, IL-5, and IL-13) in BALF (Supplementary Fig. 11g).
We validated these findings in in a house dust mite (HDM)-induced asthma model (Supplementary Fig. 11h). Immunofluorescence confirmed significant upregulation of IGF1 in the airways of HDM-challenged mice (Supplementary Fig. 11i). Similar to the OVA model, anti-IGF1 treatment significantly reduced airway inflammation, goblet cell hyperplasia, eosinophil infiltration, airway hyperresponsiveness, and T2 cytokine production (Supplementary Fig. 11j–m).
To specifically model EGPA pathophysiology, we utilized IL-5 transgenic mice challenged with HDM and IL-33, which recapitulates key features including intense eosinophilic inflammation, granuloma formation, and vascular damage (Supplementary Fig. 12a, b). Quantitative RT-PCR analysis confirmed significant upregulation of both IGF1 and IGF1R in the lungs of these mice (Supplementary Fig. 12c, d). Treatment with an IGF1 neutralizing antibody significantly reduced total inflammatory cell counts and eosinophil infiltration in BALF (Fig. 6a–c), accompanied by decreased IL-25 levels (Fig. 6d), suggesting IGF1 regulates IL-25 production. Histological analysis revealed that anti-IGF1 treatment markedly reduced inflammatory infiltrates and goblet cell hyperplasia (Fig. 6e).
a–e Anti-IGF1 treatment reduces eosinophilic inflammation and airway remodeling in IL-5 transgenic mice challenged with HDM and IL-33. Mice were divided into Control (PBS-treated), Model (HDM+IL-33 challenged), and Anti-IGF1 (HDM+IL-33+anti-IGF1 antibody) groups. a Total cell counts in bronchoalveolar lavage fluid (BALF) from control, model, and anti-IGF1-treated groups. b Eosinophil percentage in BALF. c Absolute eosinophil counts in BALF. d IL−25 levels in BALF quantified by ELISA. e Representative histological images of lung sections stained with hematoxylin and eosin (H&E, upper panels) and periodic acid-Schiff (PAS, lower panels), with corresponding inflammation and PAS score quantifications. Scale bar, 100 μm. f–h IGF1R deficiency modulates eosinophilic inflammation and airway remodeling via IL−25 in HDM+IL-33 challenged mice. f Total cell counts in BALF from wild-type (WT, Scgb1a1-IRES-+/+ Igf1rf/f), conditional knockout (Scgb1a1-IRES-Cre/+ Igf1rf/f, CKO), and CKO mice treated with recombinant IL-25 (rIL-25). All groups were challenged with HDM+IL-33. g Flow cytometric analysis of eosinophils (CD45+Siglec-F+CD11c-) in BALF. Representative plots and quantification of eosinophil percentages are shown. h Representative H&E (upper panels) and PAS (lower panels) staining of lung sections from WT, CKO, and CKO+rIL-25 groups, with quantification of inflammation and PAS scores. Scale bar, 100 μm. Statistical analysis: Data are presented as mean ± SD (n = 5 mice per group). Statistical significance was determined using a two-sided one-way ANOVA with Tukey’s post hoc test. *p < 0.05; **p < 0.01; ***p < 0.001; NS, not significant. Exact P values and complete test statistics for a–h are provided in Supplementary Data 1. CKO conditional knockout, HDM house dust mite, rIL-25 recombinant IL-25, WT wild-type.
Since corticosteroids are standard care for EGPA, we evaluated anti-IGF1 as an add-on therapy. EGPA-like mice were treated with dexamethasone (dexa), anti-IGF1, or their combination (Supplementary Fig. 12e). All treatments significantly ameliorated pulmonary inflammation scores (Supplementary Fig. 12f, g). Regarding goblet cell hyperplasia, all treatment modalities significantly reduced Periodic Acid-Schiff (PAS) scores compared to the model group (Supplementary Fig. 12f, h). Notably, while dexamethasone alone partially reduced goblet cell hyperplasia, both anti-IGF1alone and the combination therapy were more effective, reducing PAS scores to near-control levels (Supplementary Fig. 12h). These findings demonstrate that anti-IGF1 works as a monotherapy and shows promise as an add-on to corticosteroids, potentially addressing corticosteroid-resistant airway remodeling in EGPA.
To determine the specific cellular source of IGF1-mediated effects, we generated conditional knockout (CKO) mice lacking IGF1R in club cells (Scgb1a1-IRES-Cre/+Igf1rf/f, CKO) (Supplementary Fig. 13a). Flow cytometry confirmed reduced IGF1R expression in CD326+ epithelial cells from CKO mice (Supplementary Fig. 13b). In the HDM+IL-33 challenge model (Supplementary Fig. 13c), CKO mice showed no significant difference in IGF1 expression compared to wild-type models but remained elevated over PBS controls (Supplementary Fig. 13d). IGF1R expression showed a downward trend in CKO mice (Supplementary Fig. 13e). Critically, IL-25 levels were significantly reduced in BALF from CKO mice, demonstrating a direct link between epithelial IGF1R signaling and IL-25 production (Supplementary Fig. 13f). CKO mice also exhibited significantly reduced goblet cell hyperplasia (Supplementary Fig. 13g) and inflammatory cell infiltration and eosinophil percentages in BALF (Fig. 6f, g). The protective effects in CKO mice were reversed by recombinant IL-25 (rIL-25) administration, which restored eosinophilic inflammation and airway remodeling to wild-type levels (Fig. 6f, g). Histological analysis confirmed that IGF1R deficiency significantly decreased inflammatory infiltrates and goblet cell hyperplasia, effects abrogated by rIL-25 administration (Fig. 6h).
Collectively, these findings from multiple experimental models provide robust in vivo validation of the IGF1 pathway’ contribution to both T2-driven asthma and EGPA-like pathology, underscoring its potential as a therapeutic target for these chronic airway diseases.
Discussion
EGPA is a potentially life-threatening systemic necrotizing vasculitis that occurs almost exclusively in patients with asthma, exhibiting distinct immune dysregulation beyond classical T2 inflammation. Its development involves complex gene-environment interactions, with specific genetic patterns distinguishing ANCA-positive and ANCA-negative subtypes, such as HLA-DQ variants in MPO-ANCA-positive cases and links to polymorphisms affecting mucosal immunity (GPA33) and eosinophil regulation (IL5) in ANCA-negative EGPA40. Known environmental triggers include leukotriene receptor antagonists, suggesting a potential causal role41, as well as occupational exposures to silica, organic solvents, and agricultural environments42.
Although our study did not directly assess these specific exposures, our findings of IFN-I-driven inflammation in EGPA may be related to such triggers. For instance, silica exposure is known to induce type I interferon responses and promote autoimmunity43,44. It is plausible that these environmental insults provide the initial stimulus for the IFN-I-driven granulocyte activation and autoimmune features we identified as unique to EGPA. The tertiary lymphoid structures and localized autoantibody production we observed may represent a consequence of sustained environmental insults triggering these interferon-driven pathways. Future research should investigate whether specific environmental factors directly induce the IL1B+MX1+ neutrophil signature or promote TLS formation.
Our scRNA-seq analysis, while confirming shared T2 cytokine-driven eosinophilic inflammation in both EGPA and SEA, revealed unique characteristics distinguishing EGPA. These include robust interferon (IFN)-I-driven granulocyte activation, the presence of TLS in bronchial tissues, and localized autoantibody production. These features suggest a broader immune dysregulation, encompassing autoimmunity and chronic granulocyte activation, which likely contributes to the severity of EGPA and corticosteroid resistance. We specifically identified IL1B+MX1+neutrophils (Neutrophil-IL1B/MX1) uniquely enriched in EGPA, highly expressing IFN-stimulated genes IL1B, and TNFSF13B (BAFF)—factors closely associated with TLS formation and the formation of FCRL4+ atypical memory B cells (ABCs). Additionally, eosinophils, mast cells, and basophils in EGPA exhibited IFN-I-driven transcriptional programs, suggesting a systemic IFN-I-mediated granulocyte activation signature. The increased local production of autoantibodies such as ANCA, anti-dsDNA, and anti-chromatin in EGPA sputum further supports this hypothesis. These differences indicate that while both SEA and EGPA are characterized by T2 inflammation, additional immune responses are activated in EGPA.
IGF1 is a pleiotropic growth factor that, in excess, has been associated with asthma risk. Previous research highlights its crucial involvement in pulmonary diseases, with dynamic spatiotemporal distribution during acute respiratory distress syndrome (ARDS) progression, where localized pulmonary IGF1 signaling drives disease severity45. In H1N1 influenza models, IGF1 receptor blockade attenuates virus-induced tissue damage, whereas exogenous IGF1 exacerbates disease outcomes46. In asthmatic patients, serum levels of IGF1 and IGFBP3 are significantly increased and negatively correlated with asthma control test (ACT) scores, oxygen saturation, and FEV1/FVC ratio34. Large-scale population-based studies using Mendelian randomization have identified a causal association of IGF1 with childhood obesity-related asthma47.
In our study, a subset of EGPA patients remains refractory to therapy despite treatment with corticosteroids and immunosuppressants, reflecting a well-known therapeutic paradox of relapse and persistent sputum eosinophilia. Our findings identify a promising candidate pathway for this. We identified a unique IGF1+ macrophage population (IGF1+Mac) significantly enriched in treatment-relapse EGPA patients, expressing high levels of IGF1, a key factor implicated in tissue remodeling. The significant abundance of this population in the relapse group indicates a role in disease persistence. High IGF1 expression was also observed in ionocytes and adventitial fibroblast cells in both diseases, particularly in EGPA.
Through in vitro and in vivo studies, we demonstrate that IL-13 and IL-33 increase IGF1 expression in alveolar epithelial cells. IGF1, in turn, induces goblet hyperplasia and IL−25 expression, which promotes T2 and eosinophilic inflammation. Accordingly, anti-IGF1 treatment suppressed pathological goblet hyperplasia, T2 immunity, and eosinophil infiltration in mouse models of asthma and EGPA. Our experimental findings demonstrate that IGF1 signaling in airway epithelial cells regulates IL-25 production, which drives eosinophilic inflammation and airway remodeling. Using both pharmacological inhibition and genetic approaches, we established that disrupting the IGF1/IGF1R pathway significantly reduces IL-25 levels, eosinophilic inflammation, and goblet cell hyperplasia. Moreover, in our EGPA-like mouse model, anti-IGF1 as an add-on therapy to corticosteroids demonstrated superior efficacy in reducing goblet cell hyperplasia compared to dexamethasone alone. These findings suggest that IGF1 inhibition could address corticosteroid-resistant aspects of airway remodeling. Critically, the restoration of pathology by recombinant IL-25 administration in IGF1R-deficient mice confirms IL-25 as an essential downstream mediator. It should be noted that the specific effects of IGF1 signaling on ILC2 immunity were not addressed in this study, which warrants further investigation.
Contrasting with epidemiological observations and findings using IGF1R blockade23, genetic deletion of IGF1R in epithelial cells has been shown to exacerbate lung inflammation after allergen exposure by modulating epithelial non-classical phagocytosis48, highlighting the context-dependent role of IGF1R signaling. However, other studies demonstrate that HDM-exposed Igf1r-deficient mice exhibit attenuated airway hyperresponsiveness and reduced mucus secretion49,50. In our combined HDM+IL-33 model, a setting relevant to steroid-resistant asthma51 and strongly associated with EGPA52, epithelial IGF1R knockout significantly reduced T2 inflammation and airway remodeling, accompanied by marked downregulation of IL-25. The addition of recombinant IL-25 restored the inflammatory response, underscoring a pro-inflammatory role of IGF1R in this context. Consistent with this, in vitro stimulation of airway epithelial organoids with recombinant IGF1 confirmed its ability to promote remodelling. These findings collectively indicate that IGF1R signaling exhibits dual, context-dependent roles in airway inflammation.
Our study provides several methodological and conceptual advances in understanding EGPA pathobiology. Through single-cell RNA sequencing, we offer unprecedented resolution of the airway immune landscape in EGPA versus SEA, moving beyond classical T2-centric models to identify an interferon-I-driven granulocyte activation signature and tertiary lymphoid structures as distinctive EGPA features. We discovered an IGF1+ macrophage population specifically enriched in corticosteroid-relapsing EGPA, potentially explaining treatment resistance in a subset of patients. Importantly, rigorous in vitro and in vivo functional experiments validated the IGF1/IL-25 signaling axis as a critical and therapeutically targetable driver of disease pathogenesis, providing strong experimental validation for our clinical observations.
Several limitations constrain our findings. Our cohort characteristics, including a tertiary care asthma comparator group, predominantly ANCA-negative EGPA patients, and relatively small sample size, limit generalizability and prevented robust sub-phenotype analysis. The cross-sectional design comparing remission and relapse groups constrains causal inference regarding treatment-response dynamics, although persistent sputum eosinophilia despite comparable systemic corticosteroid dosing strongly indicates mechanistic differences in treatment responsiveness. While IGF1+ macrophage enrichment in corticosteroid-relapsing EGPA was characterized, their upstream regulatory mechanisms remain unclear, and the association between IGF1+ macrophages and clinical relapse remains correlative despite experimental model support. Additionally, although anti-IGF1 showed efficacy in our EGPA-like mouse model, the precise interaction between IGF1 signaling and corticosteroid pathways requires further elucidation, as does its direct effects on ILC2 populations.
Despite these limitations, our study provides valuable insights into the immune landscape of EGPA, highlighting the key roles of IFN-I-driven granulocyte activation and TLS formation as distinguishing features from SEA. We also demonstrate that IGF1 signaling plays a central role in persistent airway remodelling and T2 inflammation. These findings suggest that targeting the IGF1 and IFN-I signaling could offer potential therapeutic opportunities for patients with refractory asthma and EGPA, particularly those with persistent airway remodelling and immune dysregulation.
Methods
Human participants
The study was carried out from March 2019 to October 2023 using a cohort of adults who attended the department of Pulmonary and Critical Care Medicine of the First Affiliated Hospital of Guangzhou Medical University in Guangzhou, China, and no participant compensation was provided in this study. Sex and gender were determined based on self-report. For the single-cell RNA sequencing cohort (detailed in Supplementary Table 1), a total of 28 participants were analyzed, including patients with EGPA (n = 9; mean age: 42.11 ± 12.04 years; 6 females and 3 males), SEA (n = 13; mean age: 42.38 ± 13.20 years; 9 females and 4 males), and healthy controls (n = 6; mean age: 39.17 ± 13.30 years; 3 females and 3 males). Sample sizes varied for subsequent validation experiments (e.g., FACS, mIF, and ELISA) and are listed in detail for every experiment in Supplementary Table 2. Sex was not considered in the study design and analysis as a variable due to the low incidence of EGPA and the difficulty in sample collection, which limited the feasibility of subgroup analyzes. Disaggregated data for sex and gender are provided in the Source Data files. Participants had no other complications with no other complication including malignancy, liver disease, hypersensitivity, parasitic infection, infectious diseases (HBV status: HBV DNA and Hepatitis B surface antigen (HBsAg) negative) and other autoimmune diseases. Sex and gender were determined based on self-report. Sex and gender were determined based on self-report. Sex was not considered in the study design and analysis due to the low incidence of EGPA and the difficulty in sample collection. no participant compensation was provided in this study. Participants who have been diagnosed with EGPA for at least 6 months based on MIRRA Criteria14 and also meet the 2022 ACR/EULAR classification criteria for EGPA53. Briefly, included patients who had the history or presence of: asthma plus eosinophilia (>1.0 × 109 /L and/or >10% of leucocytes) plus at least two of the following additional features of EGPA: (1) a biopsy showing histopathological evidence of eosinophilic vasculitis, or perivascular eosinophilic infiltration, or eosinophil–rich granulomatous inflammation; (2) neuropathy, mono or poly (motor deficit or nerve conduction abnormality); (3) pulmonary infiltrates, non–fixed; (4) sinonasal abnormality; (5) cardiomyopathy (established by echocardiography or MRI); (6) glomerulonephritis (hematuria, red cell casts, proteinuria); (7) alveolar hemorrhage (by bronchoalveolar lavage); (8) palpable purpura; (9) positive test for ANCA (MPO or PR3). For the patients with EGPA who received regular corticosteroid and immunosuppressant treatment, we defined EGPA relapsing as a history of relapse requiring an increase in glucocorticoid dose, initiation or increase in other immunosuppressive therapy, or hospitalization. EGPA remission definition was referred to54: the absence of a clinical systemic manifestation (excluding asthma and/or ENT) after regular medicine treatment. This study used a cross-sectional design to compare two independent groups of EGPA patients: one group of three patients in clinical remission and another group of three patients experiencing a relapse, and longitudinal data for relapse or remission could be found in Supplementary Table 3. The samples were not paired. SEA included in this study met the severe asthma criteria of GINA2019 guidance with the sputum eosinophilic greater than 3%55. Control participants were individuals under investigation for chronic cough, but had no history of pulmonary disease, infections, or other immune-related conditions.
Human blood samples
15 mL venous blood were obtained from clinical with EDTA anti-coagulation tube and brought to laboratory within 2 h at normal temperature. Diluted with 1× D-PBS (1:1 volume, diluted from 10X, Thermo Fisher, Cat# 14200075) into a 30 mL diluted solution, followed by mononuclear cell and granulocyte isolation with Ficoll-paque density gradient (Ficoll-paque PREMIUM, Cytiva, Cat# 17544203). Briefly, 20 mL Ficoll solution was added into a 50 mL Conical tube, then diluted blood solution was added on top of the Ficoll layer in a drop-wise manner. 500 × g centrifugation with acceleration rate 2 and deceleration rate 0 for 30 min, RT. Then mononuclear cell layer and lower red blood cells and granulocytes layer were separately collected. Blood mononuclear cells were washed twice with 1× D-PBS, and contaminated red blood cells were removed with ACK lysing buffer (Thermo Fisher, Cat# A1049201), followed by 1× D-PBS wash twice, and centrifugated at 500 × g 10 min to obtain a single-cell solution. Lower red blood cells and granulocytes layer was directly for blood red cells lysis with 45 mL pre-cooling ddH2O for 20 s, followed by adding 5 mL 10× D-PBS to terminate lysis. Cell pellet was obtained by centrifugation at 500 × g 10 min. After washed by 1× D-PBS twice, single cells were resuspended with 1× MACS buffer (prepared using BSA, Sigma Aldrich, Cat# 810683 and D-PBS). CD16+ and CD16- granulocytes were separately obtained with the commercial anti-human CD16 MicroBeads according to the manufacturer’s instructions (Miltenyi Biotec, Cat# 130-045-701). Blood cell suspension for single-cell RNA sequencing was mixed with CD16+ granulocytes, CD16- granulocytes and PBMC at a ratio of 3:4:3.
Human BALF samples
20–50 mL BALF was obtained from clinical. Within 2 h samples were subjected to treatment with 0.1% DTT (MedChemExpress, Cat# HY-15917) and 3% BSA (Sigma Aldrich, Cat# 810683) for 10 min on ice. Cell pellet was obtained with centrifugation at 2000 rpm for 10 min and washed twice with 1× D-PBS. Then the cell pellet was subjected to the second lysis with 5 mL Trypsin-EDTA (0.05%) (Thermo Fisher, Cat# 25300054) at 37 °C for 5 min and terminated with 5 ml FBS (Gibco, Cat# A31604). Then cell solution was filtered through a 70 μm cell strainer into a 50 mL conical tube and washed twice with 1× D-PBS to obtain the single cell suspension for the downstream study.
BALF eosinophil enrichment for single-cell RNA sequencing
To enrich the eosinophils in BALF from control and SEA for single-cell RNA-seq, we conducted following procedure: BALF cells were stained with a lineage cocktail conjugated to FITC (CD3, BioLegend Cat# 317306; CD11c, BioLegend Cat# 337214; CD14, BioLegend Cat# 301804; CD16, BioLegend Cat# 302006; CD19, BioLegend Cat# 302206; CD20, BioLegend Cat# 302304; CD56, BioLegend Cat# 318304; CD123, BioLegend Cat# 306014), CD45-APC (BioLegend, Cat# 304012), CD117-BV510 (BioLegend, Cat# 313220), and Live/Dead-PI. Eosinophils were roughly sorted as (Lineage- CD117- CD45+) using a FACSAria3 flow cytometer (BD Science).
Human bronchial biopsy single cells
Human bronchoscopically biopsies from peripheral lung bronchus were firstly chopped with scissors in a 1.5 mL tube, then treated with 3 mL of D-PBS digestion solution containing DNase I (0.1 mg/mL, Sigma Aldrich (Roche), Cat# 11284932001) and collagenase D (2.5 mg/mL, Sigma Aldrich (Sigma Aldrich (Roche), Cat# COLLD-RO), and incubated at 37 °C with agitation (180 rpm) for 1 h. The isolated cells were filtered through a 40 μm cell strainer, and the single-cell suspension was collected by centrifugation at 500 × g for 5 min at 4 °C.
Human primary airway epithelial cells
4–5 human bronchoscopically brushes from peripheral lung bronchus in 5 mL D-PBS (10%FBS) were transported on ice to the laboratory. After incubating with 5 mL Accutase solution (Sigma Aldrich (Roche), Cat# A6964) at 37 °C for 30 min, the samples were centrifuged at 500 × g for 5 min at 4 °C to collect the cells. Then, cells obtained from bronchoscopically brushes were subjected to 5 μ/ml Dispase (STEMCELL, Cat# 07446) digestion overnight at 4 °C, and collected with centrifugation at 250 × g for 5 min, followed by 0.25% Trypsin/EDTA (Thermo Fisher, Cat# 25200072) digestion for 15 min at 37 °C. Collected cells were resuspended in complete expansion medium (PneumaCult-Ex Plus Medium, STEMCELL Technologies, Cat# 05040) and expanded for 7 to 9 days.
Human sputum samples
The participants are instructed to inhale progressively nebulized hypertonic saline solutions with concentrations of 3%, 4%, and 5% to induce sputum expectoration, which was then collected into sterile tubes. The next step was carried out within 2 h, in which the sputum was separated from saliva, treated with 4 times the amount of Dithiothreitol (DTT, MedChemExpress, Cat# HY-15917), and then suspended in 4 times the volume of Dulbecco’s Phosphate Buffered Saline (D-PBS, Thermo Fisher, Cat# 14200075) to obtain a sputum suspension. The sputum suspension was filtered and then centrifuged at 2000 rpm for 10 min at 4 °C. The sputum supernatant after centrifugation was aliquoted at 100 μL per tube, stored at −80 °C for later assay. Samples were considered valid only if microscopic evaluation confirmed <20% squamous epithelial cells. Cell viability was assessed by Trypan blue staining (STEMCELL Technologies, Cat# 07050), and only samples with ≥80% viability was included for further analysis.
Mice
8–10 week wild-type female C57BL/6J mice were purchased from Shanghai Model Organisms Centre. The Scgb1a1-IRES-Cre and Igf1r flox mice were also generated and bred by the Shanghai Model Organisms Center. Briefly, Igf1r flox mice were crossed with Scgb1a1-IRES-Cre mice, and the genotype of the offspring was determined by mouse tail genotyping. Mice with the genotype Scgb1a1-IRES-Cre/- Igf1rf/f were defined as the target CKO mice, while mice with the genotype Scgb1a1-IRES+/+ Igf1rf/f were used as controls (background strain: C57BL/6J). Specific sexes were chosen based on the study design, although sex was not analyzed as a biological variable within groups. Female mice were used for the standard asthma models (OVA and HDM) and the Il5-transgenic EGPA-like model to align with epidemiological prevalence. In contrast, male mice were specifically selected for the mechanistic studies involving CKO mice. This decision was based on the sufficient availability of male offspring to meet sample size requirements and to minimize potential experimental variability associated with hormonal fluctuations (estrous cycle). Genotyping primers were synthesized by BGI Genomics (Shenzhen, China), with sequences derived from standard protocols provided by Shanghai Model Organisms Centre or the donor investigator. The primers for genotyping:
Igf1r-Flox, forward, 5′-GAGGAGCACAGTGCCTTGTA-3′;
Igf1r-Flox, reverse, 5′-CGTACCGGAGACTGGCTAGA-3′;
Scgb1a1-IRES-Cre, forward, 5′-ACTCACTCCAGCTCCAATGC-3′;
Scgb1a1-IRES-Cre, reverse (Common): 5′-GGTCTTCAGTCCCCTTCAGC-3′;
Scgb1a1-IRES-Cre, reverse (Mutant): 5′-CCAAAAGACGGCAATATGGT-3′;
Il5-transgenic mice (NJ1638) were obtained from Professor Songmin Ying at The Second Affiliated Hospital, Zhejiang University. The primers for genotyping:
IL5tg, forward (transgene), 5′- TGGCAGTGGCCCAGACACAGC-3′;
IL5tg, reverse (transgene), 5′- ACCCCACACCTAGCCCACTG-3′;
IL5tg, forward (internal positive control), 5′- AGTGGCCTCTTCCAGAAATG-3′;
IL5tg, reverse (internal positive control), 5′- TGCGACTGTGTCTGATTTCC-3′;
All mice were held in individually ventilated cages (IVCs) in an SPF (Specific Pathogen Free) barrier unit maintained under a 12-h light/12-h dark cycle, with a controlled ambient temperature of 22 ± 2 °C and a relative humidity of 40–60%. Mice were provided with ad libitum access to standard chow and water. The identities of animals were documented by the numbered cages and ear punches. The animal studies were approved by the Institutional Animal Care and Use Committee (IACUC) of Guangzhou Medical University (permission number: 2021-478) and carried out according to the institutional guidelines. At the end of the experiments, mice were euthanized by CO₂ inhalation followed by cervical dislocation for inflammation and pathological characterization. Disaggregated numbers of animals used in individual experiments (e.g., ovalbumin-induced asthma model, HDM-induced asthma model, HDM+IL-33-induced EGPA-like model and bone marrow-derived eosinophil functional assays) are provided in the Source Data files. Overall, a total of 180 mice were used across all experiments including those utilized for pilot studies, biological replicates, and accounting for attrition. The specific breakdown includes: 80 wild-type female C57BL/6J mice (for OVA models, HDM models, and bone marrow eosinophil induction functional study), 60 female Il5-transgenic mice, and 40 male mice (20 CKO and 20 controls) for CKO studies.
Isolation and culture of mouse bone marrow derived eosinophil
Bone marrow cells were harvested from the femurs and tibias of 8-week-old female C57/B6L mice and centrifuged at 300 × g for 5 min to obtain the cell pellet, followed by red cells depletion with ACK lysing buffer (Thermo Fisher, Cat# A1049201) and incubated at room temperature in the dark for 10 min. The cell pellet without red cells were subjected to cell counting and was adjusted to a concentration of 106 cells/mL, then cultured in the T75 cell culture flasks with the base medium (RMPI1640 Gibco, Cat# C11875500BT, 20% FBS Gibco, Cat# A31604, 2.5% HEPES Thermo Fisher, Cat# 15630106, 1% Pen-Strep Gibco, Cat# 15140122, 1%NEAA Gibco, Cat# 11140076, 1% sodium pyruvate Thermo Fisher, Cat# 11360070, and 0.01 mM 2-mercaptoethanol Sigma, Cat# M6250) containing SCF (100 ng/mL, PeproTech, Cat# 250-03) and Flt3-L (100 ng/mL, PeproTech, Cat# 250-31 L) for 2 days. At Day 4, the medium was replaced with fresh base medium containing IL-5 (10 ng/mL, PeproTech, Cat# 215-15). Half of the culture media were changed every 2 days with fresh base medium containing IL-5 until day 14.
Single cell RNA-seq library construction
All single-cell suspensions were processed according to the following quality-controlled workflow: Cell Viability Quality Control: ≥80% viability (assessed by BD single-cell scanner using calcein AM (Servicebio, Cat# G1728-0.1ML) and PI dyes (Servicebio, Cat# G1012-10ML): Directly processed for library preparation, <80% viability: Processed with Miltenyi Biotec Dead Cell Removal Kit, Cat# 130-090-101), and post-treatment viability ≥80%: Proceeded to sequencing. Single cell suspension passed the QC standard was proceeded for RNA-seq library construction. BD Human Single-Cell Multiplexing Kit (BD biosciences, Cat# 633781) was used for single cell library preparation. Detailed cell viability and loading cell counts for each sample could be found in Supplementary Table 4. Up to 2 samples were labeled and pooled for subsequent single cell capture with the BD Rhapsody Single-Cell Analysis system. After partitioning and lysis of cells, polyadenylated mRNAs were captured by BD Rhapsody Enhanced Cell Capture beads and reverse transcribed into cDNA (BD Rhapsody Enhanced Cartridge Reagent Kit, Cat# 664887). Whole transcriptome mRNA libraries were then generated using random priming of the on-bead cDNA. All libraries were sequenced on Illumina Nova-seq 6000 platform.
Preprocessing of single-cell RNA-seq data
The raw sequenced data was processed with an in-house pipeline for BD Rhapsody system. The human transcriptome reference GRCh38 was used for reads realignment. The outputs for the libraries of each sample were utilized to generate the raw expression count matrix (gene counts versus cells) and converted to Seurat objects using the R package Seurat v4.0.0 for subsequent analysis56. We performed the following procedure to remove unqualified cells: (1) cells with fewer than 200 genes or more than 10,000 genes detected; (2) cells with more than 100,000 unique molecular identifiers (UMIs) detected; (3) cells with more than 30% of UMIs that were derived from mitochondrial genes. Following the data quality control, the Seurat’s SCTransform function was conducted to combine and normalize the filtered Seurat objects of all libraries, and eliminate the impact of sequencing depth, mitochondrial genes expressions, ribosomal genes expressions and potential batch effects. The final datasets for our two scRNA cohorts (Supplementary Tables 1 and 2) consisted of 354,353 cells expressing 26,821 genes and 108,098 cells expressing 23,117 genes, separately.
Identification of major cell types
We initially used our first dataset, which comprised control, SEA and EGPA participants, to identify the major cell types by applying dimension reduction and clustering. A total of 3000 variable genes were selected using the Seurat’s function FindVariableFeatures for principal component analysis (PCA). The first 15 principal components (PCs) were used for UMAP dimension reduction using RunUMAP, shared-nearest neighbor computation using FindNeighbors, and cluster identification using FindClusters. All the functions were applied with the default settings implemented in Seurat. Then we mapped canonical cell markers to the two-dimensional UMAP map to allocate different clusters into specific major cell types, and a total of 17 major cell types were identified. Based on the construction of reference map of major cell types for the first dataset, we performed the same process to identify the major cell types for our second dataset, which comprised only patients with EGPA after the treatment of corticosteroid and immunosuppressant treatment (Cs), and a total of 10 cell types were identified.
Analysis of functional cellular subsets within the major cell types
Functional cellular subsets within each major cell type were partitioned and analyzed. Variably expressed genes for each major cell type were selected for PCA, and significant PCs were used for dimension reduction, clustering and visualization. Seurat’s function FindClusters with suitable parameter of resolution was conducted to identify different cellular clusters. Differentially expressed genes (DEGs) for each cellular subset within specific major cell type were identified using FindAllMarkers function. Functional enrichment analysis for gene markers of specific cellular subset was conducted using Metascape57. Specifically, cells expressing dual-linage specific genes during the analysis were removed to reduce the potential doublet capture bias, including: (1) CD3+CD68+ cells; (2) CD3+EPCAM+ cells; (3) CD19+CD68+ cells; (4) CD19+EPCAM+ cells; (5) CD68+EPCAM+ cells; (6) COL1A1+CD3+ cells. For our second dataset, due to effect of Cs-treatment, immune cells were depleted (especially for T cells, NK cell, B cells and granulocytes). Therefore, we only analyzed the cellular subsets within macrophages and DCs, and epithelial cells for the second dataset. Of note, due to the potential impact of Cs-treatment, or sample size and detected cell number, some cell subtypes identified in our first dataset were not captured in our second dataset. We also conducted Ro/e analysis to compare the relative abundance of cell subsets across control, SEA, EGPA, and EGPA treatment (Cs-remission or Cs-relapse) groups58. Detailed descriptions of cellular subsets within each major cell type and their marker genes were shown in the relevant figures and sections of main text.
Transcriptomic comparison analysis across different groups
Any transcriptomic comparisons, e.g., between control and SEA or EGPA, between EGPA and EGPA treatment groups (Cs-remission or Cs-relapse), were performed using MAST59, which fits a hurdle model for handling zero-inflated single cell assay data and comparing gene expressions with linear regression. We conducted the analysis by the following formula to assess the coefficients of groups (control versus disease) and clinical treatment efficacy (non-treatment versus Cs-remission or Cs-relapse) on gene expression,
in which, Eᵢ indicates the gene expression value for gene i across all cells based on the “RNA” assay mode of the Seurat objects, G represents the group to be compared with the control (including SEA and EGPA) and T represents the treatment efficacy. Of note, the information of tissue origin was reflected in the process of cellular subset analysis, in which cells from peripheral blood or tissue (bronchial biopsies and BALF) were clustered into different cellular subsets. Therefore, when comparing the gene expressions across different groups within the same cell subset, we did not include the information of tissue origin in the regression model mentioned above. The significance of coefficient for each variable was determined by the likelihood ratio test implemented in MAST. Up- or down-regulated genes between different groups were determined by the threshold of P value ≤ 0.05 and |coefficient| ≥ 0.5.
Transcription factor activity analysis
Single-Cell Regulatory Network Inference and Clustering (SCENIC) was applied to predict the transcription factor (TF) activity and gene regulatory networks across cellular subsets60. In brief, the co-expression network for specific TF was constructed using the pyscenic grn function implemented in the SCENIC python version (pySCENIC). Then, the pyscenic ctx function was conducted to deduce the direct-binding target genes (regulons) for specific TF. Finally, the activity for specific TF was inferred based on its regulons’ expressions using the pyscenic aucell function. All the functions were processed with default parameters, and the AUCell matrices for TFs were retrieved for heatmap plotting.
Interacting ligands and receptors analysis
Cell-cell communication for single-cell RNA-seq data could be illustrated based on their expressed interacting ligands and receptors. We first employed CellPhoneDB to retrieve the manually curated repository of ligand-receptor pairs61. Then, uniquely expressed interleukins, chemokines, cytokines or other regulatory factors, and their corresponded receptors, of the same cellular subset or between two subsets were portrayed.
RNA velocity analysis
For cellular subsets within specific major cell type, we also performed RNA velocity analysis using velocyto to infer their potential differentiation and migration flow62. We first converted BAM files of each sample into velocyto-required formatted files with an in-house pipeline. Then, using these files as input, we ran the command line velocyto run to annotate spliced and unspliced reads and generate loom files for downstream analysis. All the loom files were merged and pre-processed using the read.loom.matrices function implemented in the R package velocyto.R. The velocity information was estimated using the function gene.relative.velocity.estimates, and visualized onto the pre-generated UMAP plot using the function show.velocity.on.embedding.cor in the velocyto.R package.
Blood and BALF FACS analysis
Cell suspension of BALF was obtained as described above, and blood single cell was obtained by directly erythrocyte lysis. Obtained cells were stained with lineage (CD3, CD19, CD20, CD56, CD14, CD206, CD11c) -Alexa Fluor 700, CD45-Percp-cy5.5, HLA-DR-eFluor 450, CD24-BV786, CD16- BV570, CD11b-BV421, CCR3-Alexa Fluor 647, SIGLEC8-BV650, CD66b-BB515, CD177-BV510, CD62L-APC-Fire 810, CD69-BV605, CD123-PE, FCERIA-BV711, Live/Dead-Zombie violet. Distinct cell types were gated as: (1) neutrophil: CD45+ lineage- HLA-DR- CD11b+ CD16+; (2) eosinophil: CD45+ lineage− HLA-DR- CD24+ CCR3+ SIGLEC8+; (3) neutrophil-progenitor1: CD45+ lineage− HLA-DR- CCR3- SIGLEC8- CD24+ CD66b+ CD177+; (4) neutrophil-progenitor2: CD45+ lineage- HLA-DR- CCR3- SIGLEC8− CD24+ CD66b+ CD177-; (5) basophil: CD45+ lineage- HLA-DR- CD24- CD123+ FCERIA+; (6) mast cell-1: CD45+ lineage- HLA-DR- CD24- CD117+ FCERIA+ CD69-; (7) mast cell-2: CD45+ lineage- HLA-DR- CD24- CD117+ FCERIA+ CD69+. Data was recorded using a Cytek Aurora Spectral Flow Cytometry (Cytek). Detailed antibodies information and dillution as followed: CD3 (1:200, BioLegend, Cat# 317340), anti-CD11c (1:200, BioLegend, Cat# 337220), anti-CD14 (1:200, BioLegend, Cat# 301822), anti-CD19 (1:200, BioLegend, Cat# 302226), anti-CD20 (1:200, BioLegend, Cat# 302322), anti-CD56 (1:200, BioLegend, Cat# 318316), and anti-CD206 (1:200, BioLegend, Cat# 321132). Other antibodies included: CD45-PerCP-Cy5.5 (1:100, BioLegend, Cat# 304028), HLA-DR-eFluor 450 (1:200, Thermo Fisher, Cat# 48-9952-42), CD24-BV786 (1:200, BD Biosciences, Cat# 568225), CD16-BV570 (1:200, BioLegend, Cat# 302036), CD11b-BV421 (1:200, BioLegend, Cat# 393114), CCR3-Alexa Fluor 647 (1:200, BioLegend, Cat# 310710), SIGLEC8-BV650 (1:200, BD Biosciences, Cat# 747871), CD66b-BB515 (1:500, BD Biosciences, Cat# 564679), CD177-BV510 (1:200, BD Biosciences, Cat# 744911), CD117- PE/Dazzle 594 (1:200, BioLegend, Cat# 313226), CD62L-APC-Fire 810 (1:200, BioLegend, Cat# 304866), CD69-BV605 (1:200, BioLegend, Cat# 310938), CD123-PE (1:200, BioLegend, Cat# 306006), FCERIA-BV711 (1:200, BD Biosciences, Cat# 747783), and Live/Dead-Zombie violet (1:1000, BioLegend, Cat# 423113). Distinct cell types were gated as described. For mouse BALF analysis, cells were stained with APC anti-mouse CD45 (1:200, BioLegend, Cat# 103112), APC/Cyanine7 anti-mouse CD11b (1:200, BioLegend, Cat# 101226), Brilliant Violet 421 anti-mouse CD11c (1:200, BioLegend, Cat# 117330) and PE/Cyanine7 anti-mouse Siglec-F (1:200, BioLegend, Cat# 155528). Detailed antibodies used in this study could be found in Supplementary Table 5.
Immunofluorescence staining
Tissue slides of formalin-fixed paraffin-embedded specimens from both human and mouse were sectioned to 4 μm thickness for multiplex immunofluorescence staining. Following deparaffinization with xylene, the slides were rehydrated through a gradient series of ethanol solutions (100%, 95%, 90%, 85%, 80%, 75%). Then slides were treated by microwave heating to induce antigen retrieval with citric acid buffer for 15 min. Subsequently, the slides were blocked with the blocking buffer for 30 min at room temperature. Primary antibody used for mouse lung tissues included IGF1 (1:200, Proteintech, Cat# 28530-1-AP). For human bronchial biopsies, six panels of primary antibodies were utilized, including (1) IFN-α (1:200, Proteintech, Cat# 18013-1-AP), IFN-β (1:200, Proteintech, Cat# 27506-1-AP), JAK-1 (1:500, Huabio, Cat# ET1705-84), CD66b (1:5000, Abcam, Cat# ab207718), MPO (1:1000, Abcam, Cat# ab208670); (2) TNF (1:1000, Huabio, Cat# HA721286), IL1-β (1:200, Huabio, Cat# ET1701-39), CD66b (1:5000, Abcam, Cat# ab207718), MPO (1:1000, Abcam, Cat# ab208670); (3) CD31 (1:2000, Abcam, Cat# ab182981), SMA (1:100, ZSGB-BIO, Cat# ZM-0003), EPX (1:500, Abcam, Cat# ab238506); (4) CD20 (1:200, Abcam, Cat# ab78237), CD4 (1:1600, Huabio, Cat# ET1609-52), CD40LG (1:500, Abcam, Cat# ab303610), FCRL4 (1:500, Abcam, Cat# ab239076), MPO (1:1000, Abcam, Cat# ab208670); (5) SMA (1:100, ZSGB-BIO, Cat# ZM-0003), CD68 (1:100, Abcam, Cat# ab201340), IGF1 (1:200, Proteintech, Cat# 28530-1-AP), IGFBP7 (1:1000, Proteintech, Cat# 19961-1-AP), IGFBP3 (1:400, Proteintech, Cat# 10189−2-AP); (6) CFTR (1:200, Abcam, Cat# ab131553), MUC5AC (1:500, Abcam, Cat# ab3649), IGF1 (1:200, Proteintech, Cat# 28530-1-AP). The slides were then incubated with secondary antibodies (1:1, 100 μl per slide; HRP-conjugated goat anti-rabbit IgG, ZSGB, Cat# PV-6001; or HRP-conjugated goat anti-mouse IgG, ZSGB, Cat# PV-6002) for 10 min at room temperature. Multiplex immunofluorescence staining was conducted using the AlphaTSA Multiplex IHC Kit (AXT36100031, AlphaX). Samples were counterstained with DAPI (Thermo Fisher, Cat# D1306) for nuclear staining for 10 min and then mounted in mounting medium. Multiplexed images were scanned using ZEISS AXIOSCAN 7. Quantitative analysis of regions or cells of interest was performed using Halo (v.3.4; Indica Labs) or Image J (Fiji). Detailed antibodies used in this study could be found in Supplementary Table 5.
Hypoxia stimulation of bone marrow derived eosinophil
Bone marrow-derived eosinophil was obtained as described above. Cell purity and viability were confirmed with FACS. Cell concentration was adjusted to 1 × 106/ml. 1 × 106/tube eosinophil was placed directly in a 5% CO2 and 95% air incubator (21% O2) or exposed to hypoxia (1% O2) in a sealed chamber or a hypoxic incubator and cultured at 37 °C for 12 h. Cells were harvested for morphology (4% Paraformaldehyde) or RT-qPCR (Trizol), and cell culture supernatant was stored at −80 °C for enzyme-linked immunosorbent assay (ELISA) measurement.
Wright-Giemsa staining
Cultured eosinophil was harvested as described above. After fixed in 70% Ethanol (diluted from Sigma Aldrich, Cat# 459836) for 10 min, then cells were covered onto slide by cytospin centrifugation. The slides were stained in 1× modified Giemsa staining solution (Beyotime, Cat# C0131) for 30 min, and washed with ddH2O for 2 min. Then the slides were naturally dried and mounted with neutral resin (Thermo Fisher, Cat# 53150).
Cell apoptosis detection
Eosinophil under conditions of hypoxia, normoxia and HIF1A inhibitor (10 μM, PX478, MCE, Cat# HY-10231) treatment were harvested after 12 h. Cells were double-stained with FITC-labeled annexin-V (a marker of early and late apoptosis) and PI (a marker for late apoptosis and necrosis) following the manufacturer’s protocol (FITC Annexin V Apoptosis Detection Kit with PI, BioLegend, Cat# 640914) and analyzed by flow cytometry. Annexin-V+/PI- cells were counted as early apoptotic cells, Annexin-V+/PI+ cells were counted as total dead cells.
Air-liquid culture of airway epithelial organoid
The expansion of primary bronchial epithelial cells was performed as described above. Expanded epithelial cells were then digested and seeded evenly on collagen-coated matrix gel (Advanced BioMatrix, Cat# 5005,) in 12-well 0.4 μm Transwell inserts (Corning, Cat# 3460). After confluent of 3 days, expansion medium was discarded and 1 mL of complete differentiated medium (PneumaCult -ALI-S Medium, STEMCELL Technologies, Cat# 05050) was added to the basal chamber, and apical chamber was devoid of culture medium. Human Recombinant IL-13 (10 ng/ml, STEMCELL Technologies, Cat# 78029.1) or Human Recombinant IL-33 (10 ng/ml, STEMCELL Technologies, Cat# 78043.1) or Human Recombinant IGF1 (100 ng/ml, STEMCELL Technologies, Cat# 78022.1) were added throughout the whole differentiation process. Cell culture medium was changed every 3 days and cells supernatant were harvested after 3 weeks of liquid-air interface differentiation.
Ovalbumin (OVA) induced asthmatic mouse model
Eight-week-old female wild-type C57BL/6J mice were randomly divided into four groups, the control, the OVA model, the OVA+recombinant IGF1 and the OVA+anti-IGF1 group. Sensitization was induced with intraperitoneal injection of 100 μg OVA (Sigma Aldrich, Cat# 5503) and 2.25 mg Imject Alum (Thermo Fisher, Cat# 77161) on day 1 and day 15, and challenged with 3% OVA aerosol for 30 min on days 21–24 consecutively. For control group, the equal volume sterile PBS was introduced as a substitute of OVA. In terms of IGF1 and anti-IGF1 stimulation, 5 μg mouse recombinant IGF1 (R&D Systems, Cat# 791-MG,) or anti-mouse IGF1 (R&D Systems, Cat# AF791) in 30 μL sterile PBS were administrated on day 0 and day 14 (1 day before the sensitization), and the same dosage was administrated 2 h before challenging on days 21–24 consecutively. At day 26, mice were sacrificed for inflammation and pathological characterization.
House dust mite extract (HDM) induced asthmatic mouse model
Eight-week-old female wild-type C57BL/6J mice were randomly divided into three groups, the control, the HDM model and the HDM+anti-IGF1 group. Sensitization was induced with intranasal injection of 50 μg low-endotoxin HDM (Greer Labs, Cat# XPB91D3A2.5) in 40 μL sterile PBS at day 1. Challenging was conducted at Day 8, Day 15 and Day 22 with 50 μg HDM via intranasal injection. For anti-IGF1 treatment, 5 μg anti-mouse IGF1(R&D Systems, Cat# AF791) in 30 μL sterile PBS was administrated at Day 0, Day 7, Day 14 and Day 21. At day 25, mice were sacrificed for inflammation and pathological characterization.
HDM and IL-33 induced eosinophilic granulomatosis with polyangiitis (EGPA) like and eosinophilic asthma mouse model
The EGPA-like mouse model was established based on a previous study52, with modifications. Briefly, 8–10 weeks old female Il5-transgenic (Il5tg) mice were randomly divided into five groups: the control, the model (HDM+IL-33), the anti-IGF1 treatment (HDM+IL33+anti-IGF1). In terms of modeling challenging, 50 μg low-endotoxin HDM (Greer Labs, Cat# XPB91D3A2.5) combined with 500 ng recombinant mouse IL-33 (PEPROTECH, Cat# 210-33) were administrated via intranasal injection at Day 0, Day 7 and Day 14. Anti-IGF1 (10 μg in 30 μL sterile PBS) was administrated via intranasal injection at Day 8, Day 10, Day 12, and Day 14. On Day 16, mice were sacrificed, and tissues were harvested for analysis. HDM and IL-33 induced eosinophilic asthma mouse model was constructed using Igf1r CKO mice. Mice were administered 50 μg of low-endotoxin HDM (Greer Labs, Cat# XPB91D3A2.5) combined with 500 ng recombinant mouse IL-33 (PEPROTECH, Cat# 210-33) via intranasal injection (i.n.) at Day 0, Day 7, Day 14 and Day 21, and recombinant IL−25 (2 μg in 50 μL saline, recombinant mouse IL25, R&D System, Cat# 1399-IL-025) was administered via intranasal injection (i.n.) at Day 15, Day 17, Day 19, and Day 21, Mice were sacrificed at Day 24, and tissues were harvested for downstream analysis.
Airway resistance measurement
Airway responsiveness was measured using a non-invasive lung resistance (Penh%) measurement (Buxco FinePointe Non-Invasive Airway Mechanics, Data Sciences International, USA). Briefly, mice were placed into the chamber and then challenged with incremental doses of aerosolised methacholine (0, 6.25, 12.5, 25 and 50 mg/mL, Sigma, Cat# A2251-25G). Airway resistance response was recorded for 3 min after each challenge. The results shown represent the increased percentage from baseline (0 mg/mL).
Mouse BALF cytology study
BALF was obtained by an intratracheal injection of total 1.6 mL (0.8 mL/once) sterile PBS followed by gentle instilling and retrieving for 3 times. Cells were collected via centrifugation (500 × g for 5 min). Cell-free supernatant was collected and stored at −80 °C for further cytokine detection. Cell pellet was re-suspended for cell counting after erythrocyte lysis and characterization of immune cells percentage by flow cytometry. Eosinophil percentage (CD45+SiglecF+CD11b+CD11c-) and cell counts (total cells*eosinophil percentage) were calculated.
Mouse lung tissue pathology analysis
The left lung of mouse was fixed overnight in 4% paraformaldehyde and embedded in paraffin. After sectioning (thickness of 4 μm), evaluation of airway structure morphology and inflammation infiltration was performed using Hematoxylin and Eosin (H&E) staining. PAS staining was utilized to assess the secretion of mucin from metaplastic epithelial goblet cells. The scoring method for H&E staining was referenced from ref.63: 0, no inflammatory cells infiltration; 1, few cells; 2, a ring of inflammatory cells with 1-cell layer deep; 3, a ring of inflammatory cells with 2–4 cell layers deep; 4, a ring of inflammatory cells with >4-cell layers deep. The PAS staining scoring method was referenced from ref.63: 0, <5% PAS+ goblet cells; 1, 5–25% PAS+ goblet cells; 2, 25–50% PAS+ goblet cells; 3, 50–75% PAS+ goblet cells; 4, >75% PAS+ goblet cells.
Autoantibodies detection
Pierce Protein A/G Agarose Plus (Thermo Fisher Scientific, Cat# 20423) was introduced for Igs enrichment. The immunoglobulins in the sputum supernatant separated after centrifugation were extracted by immunoprecipitation, namely IP-Igs. We utilized the Myeloperoxidase (MPO) Antibody ELISA kit (AESKU.DIAGNOSTICS GmbH & Co. KG, Cat# 3303) and the Proteinase 3 (PR3) Antibody ELISA kit (AESKU.DIAGNOSTICS GmbH & Co. KG, Cat# 3302) to detect anti-MPO and anti-PR3 in IP-Igs (dilute in 1:2). For NETs-related antibodies (anti-dsDNA and anti-chromatin) detection, the IP-Igs (dilute in 1:2) were incubated for 2 h in dsDNA-coated (Thermo Fisher, Cat# 15634-017) and chromatin-coated (Arotec Diagnostics, Cat# ATN02-10) primary antibody and probed with alkaline phosphatase bound secondary antibodies (goat anti-human IgG-AP, Abcam, Cat# ab6859). The chromogenic solution was used with SIGMAFAST p-Nitrophenyl Phosphate Tablets (Sigma-Aldrich, Cat# N2770-5SET).
Enzyme-linked immunosorbent assay
Commercial ELISA kits were introduced for human and mouse cytokines detection according to manufacturer instructions. For human sputum IGF1 concentration detection, 100uL sputum supernatant was used (Multi Sciences, Cat# EK1131-96). For cytokines of primary epithelial culture supernatant, human IL-25 (Elabscience, Cat# E-EL-H1648), IL-33 (Thermo Fisher, Cat# BMS2048), TSLP (Elabscience, Cat# E-EL-H1598) and IGF1 expression levels were measured. For mouse cytokines, culture supernatant from eosinophil stimulation was used to detect granule proteins expressions, including eosinophil cationic protein (ECP, Abbkine, Cat# KTE71563) and eosinophil peroxidase (EPX, FineTest, Cat# EM6789). For mouse BALF cytokines expression, type2 inflammatory cytokines, including IL-4 (Mouse IL-4 DuoSet ELISA, R&D Systems, Cat# DY404-05), IL-5 (Mouse IL-5 DuoSet ELISA, R&D Systems, Cat# DY405-05) and IL-13 (Mouse IL-13 DuoSet ELISA, R&D Systems, Cat# DY413-05), were measured.
RT-qPCR
Human primary epithelial cells and mouse eosinophils were harvested as described above. Total RNA was extracted using TRIZOL (Invitrogen, Cat# 15596018), and cDNA was synthesized from 1 μg of human cells or 200 ng of mouse eosinophils using HiScript III RT SuperMix (Vazyme, Cat# R323-01). ChamQ Universal SYBR qPCR Master Mix (Vazyme, Cat# Q711-03) (10 μL) was added to 10 μL of a reaction mixture containing cDNA and gene-specific forward and reverse primers (10 µM). cDNA was amplified for 40 cycles using a Roche 480 PCR system. GAPDH (human) or Actin (mouse) were used as reference genes. For comparison analysis, mRNA expression levels were normalized to each control group. Primers were designed and synthesized by BGI Genomics (Shenzhen, China).
Primers for human:
IGF1 forward, 5′-CTCTTCAGTTCGTGTGTGGAGAC-3′;
IGF1 reverse, 5′-CAGCCTCCTTAGATCACAGCTC-3′;
IGF1R forward, 5′-CCTGCACAACTCCATCTTCGTG-3′;
IGF1R reverse, 5′-CGGTGATGTTGTAGGTGTCTGC-3′;
IGFBP3 forward, 5′-CGCTACAAAGTTGACTACGAGTC-3′;
IGFBP3 reverse, 5′-GTCTTCCATTTCTCTACGGCAGG-3′;
GAPDH forward, 5′-GGAGCGAGATCCCTCCAAAAT-3′;
GAPDH reverse, 5′-GGCTGTTGTCATACTTCTCATGG-3′;
Primers for mouse:
Hif1a forward, 5′-CCTGCACTGAATCAAGAGGTTGC-3′;
Hif1a reverse, 5′-CCATCAGAAGGACTTGCTGGCT-3′;
Rnase3(ECP) forward, 5′- CATCACCAGTCGGAGGAGAACA-3′;
Rnase3(ECP) reverse, 5′-ATGGGACTGTCCTGTGGAGTTC-3′;
Epx forward, 5′- CTGTCTCCTGACTAACCGCTCT-3′;
Epx reverse, 5′-TCAGCGGCTAGGCGATTGTGTT-3′;
Prg2(Embp) forward, 5′- CAAGACCTGTCGCTACCTCCTA-3′;
Prg2(Embp) reverse, 5′-GCGGACTGGATTCCGAAGTTAAC-3′;
IGF1 forward, 5′- GTGGATGCTCTTCAGTTCGTGTG-3′;
IGF1 reverse, 5′- TCCAGTCTCCTCAGATCACAGC-3′;
IGF1R forward, 5′- CGGGATCTCATCAGCTTCACAG-3′;
IGF1R reverse, 5′- TCCTTGTTCGGAGGCAGGTCTA-3′;
Actin forward, 5′-CATTGCTGACAGGATGCAGAAGG-3′;
Actin reverse, 5′-TGCTGGAAGGTGGACAGTGAGG-3′;
CT image scoring
We obtained HRCT images of the thorax from each patient. Consolidation, GGO, and tree-in-bud opacities were graded using the 6-level semi-quantitative system. The anatomical extent of bronchiectasis and bronchial mucoid impaction was scored according to the number of bronchopulmonary segments involved. The maximum score for each level was 5, and the total score could range from 0 to 25. Detained scoring criteria on chest imaging of EGPA were referred to our previous study64.
Statistics
Statistical analyzes were performed using R version 4.0.0 (R Foundation for Statistical Computing, Vienna, Austria) for single-cell RNA-seq data processing and GraphPad Prism 10.0.2 (GraphPad Software, San Diego, CA, USA) for experimental and clinical data analysis. Data distribution normality was assessed using the Shapiro–Wilk test. All statistical tests were two-tailed, and a p-value less than 0.05 was considered statistically significant. For experimental and clinical variables: For comparisons among three or more groups: For normally distributed parametric data, one-way or two-way analysis of variance (ANOVA) was performed. When ANOVA indicated a significant difference, Tukey’s honest significant difference test was used for post-hoc pairwise comparisons, which inherently controls the family-wise error rate. For non-normally distributed non-parametric data, the two-sided Kruskal–Wallis test was used. If a significant difference was observed, Dunn’s post-hoc test was applied for pairwise comparisons with Bonferroni correction. For comparisons between two groups: For normally distributed parametric data, the two-sided unpaired Student’s t-test was applied. For non-normally distributed non-parametric data, the two-sided Mann–Whitney U test was applied. For paired comparisons, the two-sided Wilcoxon signed-rank test was applied. For categorical variables, Fisher’s exact test was determined. For single-cell RNA-seq data analysis: Cell clustering and annotation were performed as described in the Methods section. Differential gene expression analysis was conducted using methods appropriate for single-cell RNA-seq data, such as MAST’s zero-inflated linear model (Hurdle model) to account for data sparsity. For other comparisons of single-cell RNA-seq derived data, including cell type proportions or aggregated gene expression levels across multiple groups, the two-sided Kruskal–Wallis test was utilized. P-values from single-cell RNA-seq analyzes were adjusted for multiple hypothesis testing using the right adjustment where appropriate. Specific thresholds and criteria for significance for individual analyzes are detailed in the respective figure legends. Data presentation: Data are presented as mean ± standard deviation (SD) or median (interquartile range, IQR), unless otherwise specified in the figure legends. Sample sizes (n) for each comparison are indicated in the respective figure legends.
Ethical statement
Our research complies with all relevant ethical regulations. The Medical Ethics Committee of the First Affiliated Hospital of Guangzhou Medical University approved the study procedures (ID: Ethical Review of Medical Research 2019 No. 58). The implementations were in concordance with the International Ethical Guidelines for Research Involving Human participants as stated in the Helsinki Declaration. Informed written consent was obtained from all participants.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw single-cell RNA-seq sequence data generated in this study have been deposited in the Genome Sequence Archive for Human (GSA-Human) at the National Genomics Data Centre, China National Centre for Bioinformation / Beijing Institute of Genomics, Chinese Academy of Sciences, under accession number HRA013372. These data are available under controlled access in accordance with the policies of the Human Genetic Resources Administration of China, Ministry of Science and Technology (registration number 2025BAT00964). Access can be obtained by submitting a request via the GSA-Human platform; requests are reviewed by the relevant Data Access Committee (HDAC003678) and, upon approval, data are provided under a data access agreement. The processed single-cell RNA-seq dataset (Seurat object) generated in this study has been deposited in Zenodo (https://doi.org/10.5281/zenodo.17744852). Source data underlying the main figures and key supplementary experiments are provided with this paper as Source Data files. Publicly available databases and software used in this study are described in the Methods. All other data are available in the article and its Supplementary files or from the corresponding author upon request. Source data are provided with this paper.
Code availability
The custom codes developed for this study have been deposited in Zenodo and are accessible at https://doi.org/10.5281/zenodo.1774485265.
References
Fagni, F., Bello, F. & Emmi, G. Eosinophilic granulomatosis with polyangiitis: dissecting the pathophysiology. Front. Med. 8, 627776 (2021).
Khoury, P., Grayson, P. C. & Klion, A. D. Eosinophils in vasculitis: characteristics and roles in pathogenesis. Nat. Rev. Rheumatol. 10, 474–483 (2014).
Papi, A., Brightling, C., Pedersen, S. E. & Reddel, H. K. Asthma. Lancet 391, 783–800 (2018).
Phinyo, P., Krikeerati, T., Vichara-Anont, I. & Thongngarm, T. Efficacy and safety of biologics for oral corticosteroid-dependent asthma: a systematic review and network meta-analysis. J. Allergy Clin. Immunol. Pract. 12, 409–420 (2024).
Emmi, G. et al. Evidence-based guideline for the diagnosis and management of eosinophilic granulomatosis with polyangiitis. Nat. Rev. Rheumatol. 19, 378–393 (2023).
Acharya, K. R. & Ackerman, S. J. Eosinophil granule proteins: form and function. J. Biol. Chem. 289, 17406–17415 (2014).
Frigas, E. & Gleich, G. J. The eosinophil and the pathophysiology of asthma. J. Allergy Clin. Immunol. 77, 527–537 (1986).
Nishi, R. et al. Differential clinicopathologic features of EGPA-associated neuropathy with and without ANCA. Neurology 94, e1726–e1737 (2020).
Fredens, K., Dahl, R. & Venge, P. The Gordon phenomenon induced by the eosinophil cationic protein and eosinophil protein X. J. Allergy Clin. Immunol. 70, 361–366 (1982).
Allenbach, Y. et al. High frequency of venous thromboembolic events in Churg-Strauss syndrome, Wegener’s granulomatosis and microscopic polyangiitis but not polyarteritis nodosa: a systematic retrospective study on 1130 patients. Ann. Rheum. Dis. 68, 564–567 (2009).
Marx, C. et al. Eosinophil-platelet interactions promote atherosclerosis and stabilize thrombosis with eosinophil extracellular traps. Blood 134, 1859–1872 (2019).
Uderhardt, S. et al. Enzymatic lipid oxidation by eosinophils propagates coagulation, hemostasis, and thrombotic disease. J. Exp. Med. 214, 2121–2138 (2017).
Agache, I. et al. Efficacy and safety of treatment with biologicals (benralizumab, dupilumab, mepolizumab, omalizumab and reslizumab) for severe eosinophilic asthma. A systematic review for the EAACI Guidelines - recommendations on the use of biologicals in severe asthma. Allergy 75, 1023–1042 (2020).
Wechsler, M. E. et al. Mepolizumab or placebo for eosinophilic granulomatosis with polyangiitis. N. Engl. J. Med. 376, 1921–1932 (2017).
Wechsler, M. E. et al. Benralizumab versus mepolizumab for eosinophilic granulomatosis with polyangiitis. N. Engl. J. Med. 390, 911–921 (2024).
Kocher, M., Edberg, J. C., Fleit, H. B. & Kimberly, R. P. Antineutrophil cytoplasmic antibodies preferentially engage Fc gammaRIIIb on human neutrophils. J. Immunol. 161, 6909–6914 (1998).
Porges, A. J. et al. Anti-neutrophil cytoplasmic antibodies engage and activate human neutrophils via Fc gamma RIIa. J. Immunol. 153, 1271–1280 (1994).
Jennette, J. C. & Falk, R. J. Pathogenesis of antineutrophil cytoplasmic autoantibody-mediated disease. Nat. Rev. Rheumatol. 10, 463–473 (2014).
Mohammad, A. J. et al. Rituximab for the treatment of eosinophilic granulomatosis with polyangiitis (Churg-Strauss). Ann. Rheum. Dis. 75, 396–401 (2016).
Teixeira, V., Mohammad, A. J., Jones, R. B., Smith, R. & Jayne, D. Efficacy and safety of rituximab in the treatment of eosinophilic granulomatosis with polyangiitis. RMD Open 5, e000905 (2019).
Sharma, P. et al. Targeting eosinophils in chronic respiratory diseases using nanotechnology-based drug delivery. Chem. Biol. Interact. 365, 110050 (2022).
Siddiqui, S. et al. Eosinophils and tissue remodeling: Relevance to airway disease. J. Allergy Clin. Immunol. 152, 841–857 (2023).
Lee, H. et al. Targeting insulin-like growth factor-I and insulin-like growth factor-binding protein-3 signaling pathways. A novel therapeutic approach for asthma. Am. J. Respir. Cell Mol. Biol. 50, 667–677 (2014).
Ruan, W., Deng, J. & Ying, K. Novel aspects of insulin-like growth factor 1/insulin network in chronic inflammatory airway disease. Curr. Med Chem. 27, 7256–7263 (2020).
Bulut, I., Ozseker, Z. F., Coskun, A., Serteser, M. & Unsal, I. Pregnancy-associated plasma protein-A (PAPP-A) levels in patients with severe allergic asthma are reduced by omalizumab. J. Asthma 55, 1116–1121 (2018).
Chand, H. S., Woldegiorgis, Z., Schwalm, K., McDonald, J. & Tesfaigzi, Y. Acute inflammation induces insulin-like growth factor-1 to mediate Bcl−2 and Muc5ac expression in airway epithelial cells. Am. J. Respir. Cell Mol. Biol. 47, 784–791 (2012).
Yamashita, N. et al. Role of insulin-like growth factor-I in allergen-induced airway inflammation and remodeling. Cell Immunol. 235, 85–91 (2005).
Thacker, S. G. et al. Type I interferons modulate vascular function, repair, thrombosis, and plaque progression in murine models of lupus and atherosclerosis. Arthritis Rheum. 64, 2975–2985 (2012).
Baccala, R. et al. Type I interferon is a therapeutic target for virus-induced lethal vascular damage. Proc. Natl. Acad. Sci. USA 111, 8925–8930 (2014).
Hu, X., Li, J., Fu, M., Zhao, X. & Wang, W. The JAK/STAT signaling pathway: from bench to clinic. Signal Transduct. Target Ther. 6, 402 (2021).
Dai, D. et al. The transcription factor ZEB2 drives the formation of age-associated B cells. Science 383, 413–421 (2024).
Gao, X. et al. Zeb2 drives the formation of CD11c + atypical B cells to sustain germinal centers that control persistent infection. Sci. Immunol. 9, eadj4748 (2024).
Haslett, C. Granulocyte apoptosis and its role in the resolution and control of lung inflammation. Am. J. Respir. Crit. Care Med. 160, S5–S11 (1999).
Acat, M., Toru Erbay, U., Sahin, S., Arik, O. & Ayada, C. High serum levels of IGF-I and IGFBP3 may increase comorbidity risk for asthmatic patients. Bratisl. Lek. Listy 118, 691–694 (2017).
Castro-Rodriguez, J. A., Padilla, O., Casanello, P. & Forno, E. Higher levels of insulin-like growth factor-1 in cord blood associate with risk of asthma at age 3. Pediatr. Pulmonol. 59, 482–487 (2024).
Naryzny, S. N. & Legina, O. K. Haptoglobin as a biomarker. Biochem Mosc. Suppl. B Biomed. Chem. 15, 184–198 (2021).
Elkhattabi, L. et al. In silico analysis of coding/noncoding SNPs of human RETN gene and characterization of their impact on resistin stability and structure. J. Diab. Res. 2019, 4951627 (2019).
Li, X. et al. Overexpression of SerpinE2/protease nexin-1 contribute to pathological cardiac fibrosis via increasing collagen deposition. Sci. Rep. 6, 37635 (2016).
Wang, Y. H. et al. IL-25 augments type 2 immune responses by enhancing the expansion and functions of TSLP-DC-activated Th2 memory cells. J. Exp. Med. 204, 1837–1847 (2007).
Lyons, P. A. et al. Genome-wide association study of eosinophilic granulomatosis with polyangiitis reveals genomic loci stratified by ANCA status. Nat. Commun. 10, 5120 (2019).
Schroeder, J. W. et al. Anti-neutrophil cytoplasmic antibodies positivity and anti-leukotrienes in eosinophilic granulomatosis with polyangiitis: a retrospective monocentric study on 134 Italian patients. Int. Arch. Allergy Immunol. 180, 64–71 (2019).
Maritati, F. et al. Occupational exposures and smoking in eosinophilic granulomatosis with polyangiitis: a case-control study. Arthritis Rheumatol. 73, 1694–1702 (2021).
Benmerzoug, S. et al. Sterile lung inflammation induced by silica exacerbates Mycobacterium tuberculosis infection via STING-dependent type 2 immunity. Cell Rep. 27, 2649–2664 e2645 (2019).
Boudigaard, S. H. et al. Occupational exposure to respirable crystalline silica and risk of autoimmune rheumatic diseases: a nationwide cohort study. Int. J. Epidemiol. 50, 1213–1226 (2021).
Andonegui, G. et al. Enhanced production of IGF-I in the lungs of fibroproliferative ARDS patients. Physiol. Rep. 2, e12197 (2014).
Li, G. et al. Insulin-like growth factor 1 regulates acute inflammatory lung injury mediated by influenza virus infection. Front. Microbiol. 10, 2541 (2019).
Zhu, T. et al. Mendelian randomization study of IGF-1, childhood obesity and obesity-related asthma. Cell Mol. Biol. 69, 155–159 (2023).
Han, C. Z. et al. Macrophages redirect phagocytosis by non-professional phagocytes and influence inflammation. Nature 539, 570–574 (2016).
Piñeiro-Hermida, S. et al. Attenuated airway hyperresponsiveness and mucus secretion in HDM-exposed Igf1r-deficient mice. Allergy 72, 1317–1326 (2017).
Alfaro-Arnedo, E. et al. IGF1R as a potential pharmacological target in allergic asthma. Biomedicines 9, 912 (2021).
Kim, J. et al. JAK3 inhibitor suppresses multipotent ILC2s and attenuates steroid-resistant asthma. Sci. Adv. 9, eadi3770 (2023).
Kotas, M. E. et al. A role for IL-33-activated ILC2s in eosinophilic vasculitis. JCI Insight 6, e143366 (2021).
Grayson, P. C. et al. 2022 American College of Rheumatology/European Alliance of Associations for Rheumatology Classification criteria for eosinophilic granulomatosis with polyangiitis. Ann. Rheum. Dis. 81, 309–314 (2022).
Groh, M. et al. Eosinophilic granulomatosis with polyangiitis (Churg–Strauss) (EGPA) Consensus Task Force recommendations for evaluation and management. Eur. J. Intern. Med. 26, 545–553 (2015).
Reddel, H. K. et al. GINA 2019: a fundamental change in asthma management. Eur. Respir. J. 53, 1901046 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Zhang, Y. et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell 39, 1578–1593.e1578 (2021).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Myou, S. et al. Blockade of inflammation and airway hyperresponsiveness in immune-sensitized mice by dominant-negative phosphoinositide 3-kinase–TAT. J. Exp. Med. 198, 1573–1582 (2003).
Lin, X. et al. Retrospective comparison of high-resolution computed tomography of eosinophilic granulomatosis with polyangiitis with severe asthma. Ann. Transl. Med. 9, 983–983 (2021).
Dong, C. Airway immune profiles and therapeutic implications of IGF1 in eosinophilic granulomatosis with polyangiitis. Zenodo https://doi.org/10.5281/zenodo.17744852 (2025).
Acknowledgements
The National Natural Science Foundation of China (82125015, Y.Z.; 82495202, Y.Z.; 82070026, Q.Z.; 82570040, Q.Z.), Guangdong Provincial Science and Technology Department (20240508, Y.Z.), Guangzhou Municipal Science and Technology Bureau (2025A03J4520, Y.Z.), Guangzhou Science and Technology Key R&D Plan (202206010002, Y.Z.), Guangzhou National Laboratory and State Key Laboratory of Respiratory Disease (GZNL2023A01009, Y.Z.; GZNL2024B01002, Y.Z.), Guangzhou Women and Children’s Medical Center Fund (3001032, Y.Z.), Guangzhou Science and Technology Plan Project (202102010355, Q.Z.), Zhongshan Medical Foundation of Guangdong Province (ZNSA-2020013, Q.Z.; ZNSA-2020003, Q.Z.; ZNSXS-20220083, Q.Z.), Clinical and Epidemiological Research Project of State Key Laboratory of Respiratory Disease (SKLRD-L-202404, Q.Z.), Key R&D Program of the Ministry of Science and Technology (2022YFF0710800, L.W.; 2022YFF0710802, L.W.), Shenzhen Science and Technology Innovation Committee, Sustainable Development Special Project (KCXFZ202002011008256, L.W.), and the Major clinical research project of Guangzhou Medical University Research Ability Enhancement Program (GMUCR2024-01010, Q.Z.) funded this study.
Author information
Authors and Affiliations
Contributions
Q.Z., Y.Z., Z.C., and R.C. conceived of and supervised the project. C.Z., C.O., and L.W. recruited and provided patient care and collected follow-up data. C.D., M.L., C.O., X.Y., L.X., C.L., S.W., J.W., J.M., Z.D., Y.L., L.M., and B.L. performed laboratory experiments. Z.C., C.D., and X.Z. performed bioinformatic analysis. Y.Z. wrote the manuscript with input from A.L., M.M., P.N., N.Z., and Q.Z. All authors discussed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Francesco Ciccia, Norihiro Harada, Christian Taube and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dong, C., Lu, B., Zhong, C. et al. Airway immune profiles and therapeutic implications of IGF1 in eosinophilic granulomatosis with polyangiitis. Nat Commun 17, 1346 (2026). https://doi.org/10.1038/s41467-025-68104-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68104-6