Abstract
Emerging evidence underscores biophysical characteristics of cancer cells as key modulators of cancer progression and metastasis. Herein, we reported a cell-mechanophenotyping screening microfluidic chip (termed LesM) for the high-efficient capture of circulating tumor cells (CTCs) and evaluation of single-cell deformation to reveal the hematogenous metastatic potential of bacteria-infected breast cancer. LesM employs L-shaped traps to capture single cells, leveraging bacteria-infected CTCs with cytoskeletal reorganization traverse narrowed channels while rigid native cells are retained. The platform demonstrates an average single-cell capture efficiency of 95.42% and specificity of 85.34% in discriminating infected versus non-infected breast cancer cells, validated through parallel in vivo metastatic assays. LesM enables high-throughput sensing up to 10,240 cells of mechanical signatures and microbial cargo, correlating with metastatic risk and antibiotic response. By bridging biomechanics and intratumoral microbiota detection, LesM offers a transformative liquid biopsy tool for predicting distant metastasis and guiding antimicrobial therapies in bacteria-infected breast cancers.
Similar content being viewed by others
Introduction
There has been growing evidence that tumor-resident bacteria play an important role in cancer. Despite having low biomass, these bacteria can modulate tumor immunosuppression and growth1, predict long-term survival2, and influence the response to chemoradiation3,4. Intratumor bacteria can also contribute to cancer progression5,6, especially influencing lung metastases in breast cancer7,8. Considering that metastases can form when cancer cells enter the blood circulation and take up residence in distant tissues9, the presence of circulating tumor cells (CTCs) has been used to indicate tumor metastasis, showing reliable diagnostic performance10,11,12. Metastasis is a complicated process with multiple stages, posing a threat to its therapy13, and among the CTCs, only a few give rise to tumor growth at secondary sites—most of them undergo cell death due to the fluid shear stress while traveling through the circulation14. However, some specific intracellular bacteria have been reported to reorganize the actin cytoskeleton of their host cells, decreasing the stiffness of these cells7. This deformability could promote cell survival in the circulation, thereby increasing the metastatic potential and tumor-initiating ability of CTCs, and screening the blood for these deformable, bacteria-infected CTCs may be useful for predicting cancer metastasis and recurrence.
Therefore, we sought to create a device that could identify infected CTCs by both their mechanophenotype and the presence of intracellular bacteria. Highly integrated microfluidic chips have been used to assess single cells for a wide variety of applications, including clinically diagnosing cancer15,16,17,18, elucidating disease pathophysiology19,20, detecting metabolite secretion21, identifying tumor-derived extracellular vesicles22,23, screening drugs20,21, and ion detection assisted by machine learning24. Moreover, microfluidic chips have shown promise for single-cell mechanophenotyping in a high-throughput, cost-effective, and time-saving manner25,26. The resulting biophysical characteristics of cells could be applied as potential markers in cancer diagnosis, since biomechanical phenotype is an inherent reflection of the state and function of cells27,28. Well-designed functionalized micro-structures have been used for deformability-based sorting, to identify CTC clusters29,30, isolate cancer stem cells31,32, and separate CTCs33,34. Hence, a microfluidic chip seemed a promising platform for identifying cells according to mechanophenotype, facilitating the screening of blood for bacteria-infected CTCs in a rapid, convenient, and efficient manner.
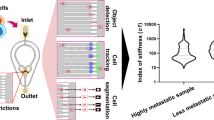
In this paper, we describe the design, optimization, and validation of such a chip, which we refer to as the cell-mechanophenotyping-based screening microfluidic chip (termed LesM) (Fig. 1). The LesM contains a dense array of hydrodynamic L-shaped traps, intended to distinguish infected CTCs from native CTCs. Each L-shaped trap consists of a hook-shaped part, designed for single cell capture, and a deformation channel part, designed for the detection of deformable cells. Only the deformable cells, which contain bacteria, can squeeze into the channel, while the stiffer native cells get blocked at the hook. We demonstrate that the LesM captures single breast cancer cells with ultra-efficiency and, when used with fluorescent probes to label intracellular bacteria, distinguishes infected cancer cells from native cells with high sensitivity and specificity. The chip successfully detected infected CTCs in unprocessed whole blood samples, suggesting its clinical utility, validated through parallel in vivo metastatic assays. And with the help of the LesM, we found that clearance of tumor-resident bacteria before resection of the primary tumor decreased the number of CTCs and reduced lung metastases. As a high-throughput and time-saving strategy allowing monitoring of both the count and the intracellular bacteria of CTCs, the LesM shows great potential for predicting the metastatic tendency of breast cancer as well as evaluating the response to antibiotic treatments.
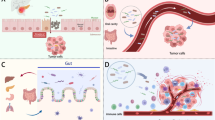
Bacterial invasion of breast cancer tumor cells increases their deformability and survival in the circulation, leading to distant metastasis. Whole blood is collected from breast cancer patients, applied to the LesM chip, and screened for infected CTCs. All CTCs are captured on traps, but only the infected CTCs squeeze into narrow channels and are fluorescently labeled. Cell counts and the presence of intracellular bacteria are used to predict distant metastasis. Icons were created in BioRender. Luo, W. (https://BioRender.com/l1e6nxj).
Results
Characterization of infected breast cancer cells
Considering that certain intracellular bacteria could trigger actin cytoskeleton reorganization in breast cancer tumor cells, promoting metastatic colonization7 (Fig. 2a), we first infected breast cancer cells with bacteria and characterized the response. Four nonpathogenic intratumor bacteria (namely Staphylococcus xylosus, Enterococcus faecalis, Lactobacillus animalis, and Streptococcus cuniculi) were applied to MDA-MB-231 triple-negative breast cancer cells for invasion. Extracellular bacteria were cleared via antibiotics, which had limited effects on host cell viability (Supplementary Fig. 1a, b). None of the bacterial strains significantly reduced cell host cell viability (Fig. 2b), but S. xylosus and L. animalis showed higher invasion rates (Fig. 2c). For convenient visualization, we labeled bacteria with the 3,3’-dioctadecyloxacarbocyanine perchlorate (DiO) and observed clear fluorescence signals in the host cell cytoplasm (Fig. 2d). To verify cytoskeleton reorganization, we stained for actin in MDA-MB-231 cells with or without invaded bacteria. Invasion of S. xylosus, L. animalis, and S. cuniculi triggered the disassembly of stress fibers, while E. faecalis hardly showed this tendency (Fig. 2e and Supplementary Fig. 1c), possibly resulting from its low invasion rate. Successful bacterial infection of breast cancer cells was also verified through flow cytometry, which showed that after invasion, bacterial fluorescence grew in a concentration-dependent manner (Fig. 2f). Next, we assessed the deformability of the infected cells. In cells generally, actin cytoskeleton organization is mediated by RhoA, a well-characterized member of Rho GTPase family35, and its downstream ROCK kinase. Thus, we expected bacterial infection to inhibit this pathway, resulting in the disassembly of stress fibers and a decrease in the host cell’s Young’s modulus (Fig. 2g). Although cells invaded by E. faecalis remained stiff, we observed that in comparison to native cells, cells invaded with S. xylosus, L. animalis, and S. cuniculi had lower Young’s moduli (Fig. 2h). A similar result was observed when native cells were treated with a ROCK inhibitor (Fig. 2h), an effect accompanied by a disassembly of actin stress fibers (Fig. 2i and Supplementary Fig. 1c), suggesting that intracellular bacteria disrupting the actin cytoskeleton do so by suppressing RhoA-ROCK activation in host cells.
a Schematic showing that intracellular bacteria reorganize the host cell cytoskeleton. Icons were created in BioRender. Luo, W. (https://BioRender.com/l1e6nxj). b Viability of MDA-MB-231 cells infected with S. xylosus, E. faecalis, L. animalis, or S. cuniculi, according to CCK-8 assay. Note that native cells are represented by a concentration of 0 CFU·ml-1. Each bar represents mean and SD, with n = 3 (biologically independent). c Rate of native MDA-MB-231 cell invasion by various bacterial strains. d Representative images of MDA-MB-231 cells infected with various bacterial strains, as captured by LSCM. Blue shows nuclei, stained with Hoechst, and green shows bacteria, stained with DiO. Scale bars, 20 μm. e Representative images of infected MDA-MB-231 cells, as captured by LSCM. Red shows actin stress fibers, stained with phalloidin, blue shows nuclei, stained with Hoechst, and green shows bacteria, stained with DiO (green arrows). Scale bars, 20 μm. f Representative flow cytometry plots for S. xylosus–infected MDA-MB-231 cells, where bacteria were labeled with DiO. g Schematic showing that bacterial infection inactivates RhoA-ROCK, leading to the disassembly of actin stress fibers and a reduced Young’s modulus of the host cell. Icons were created in BioRender. Luo, W. (https://BioRender.com/l1e6nxj). h Average Young’s modulus of MDA-MB-231 cells under various treatments, including the ROCK inhibitor Y27632, according to AFM. Each bar represents mean and SD, with n = 120 (biologically independent). Each dot represents one cell. Graphs indicate statistical significance according to two-sided, unpaired student’s t-test versus CTL. i Representative images of MDA-MB-231 cells with or without the ROCK inhibitor Y27632, as captured by LSCM. Red shows actin stress fibers, stained with phalloidin, and blue shows nuclei, stained with Hoechst. Scale bars, 20 μm. AFM atomic force microscopy, bac bacteria, CCK-8 Cell Counting Kit–8, CFU colony-forming unit, CTL control, DiO 3,3’-dioctadecyloxacarbocyanine perchlorate, E. f. Enterococcus faecalis, L. a. Lactobacillus animalis, LSCM laser scanning confocal microscopy, S.c. Streptococcus cuniculi, SSC side scatter, S.x. Staphylococcus xylosus.
Considering the complex classification of breast cancer, similar experiments were performed on MCF-7 breast epithelial cells. We used S. xylosus and L. animalis for subsequent experiments since these strains triggered stress fiber disassembly with the highest rates of host cell invasion. We examined the viability of MCF-7 cells after exposure to specific antibiotics and bacterial strains, and we visualized the intracellular bacteria (Supplementary Fig. 1d–g). Similarly, bacterial invasion caused the disassembly of stress fibers in MCF-7 cells (Supplementary Fig. 1h). Taken together, these results indicate that certain intracellular microbiota promote deformability of host cells via reorganization of the actin cytoskeleton, resulting in resistance to mechanical stress.
Design and optimization of the LesM
After confirming that infected breast cancer cells were more deformable, we created the LesM to screen whole blood for these cells. The chip has a wheel-shaped inlet to prevent clogging and 32 parallel channels containing 10,240 L-shaped traps (Supplementary Fig. 2a). As depicted in Fig. 3a, each L-shaped trap consisted of a hook-shaped part and a deformation channel part, the width of which varied from 5 to 10 μm in the preliminary design (Supplementary Fig. 2b). Because we expected stiff cells to remain on the hooks and deformable cells to squeeze into the channels, we defined 2 regions of interest (ROIs) correspondingly: blocked-cell ROI and squeezed-cell ROI (Fig. 3b). We simulated the flow resistance in the main channel to visualize the hydrodynamic mechanism underlying cell deformation and capture in the LesM (Fig. 3c, Supplementary Fig. 3, and Supplementary Movie 1) and evaluated the chip’s single-cell capture efficiency. Bright-field microscopy images showed that an average of 95.42% traps captured single cells (Fig. 3d and Supplementary Fig. 4a), allowing high-throughput single-cell screening. To study the single-, blocked-, and squeezed-cell efficiencies in depth, with blocked- and squeezed-cell efficiencies defined as the proportion of traps capturing one cell on the hook-shaped part or on the deformation channel part, we introduced MDA-MB-231 cells with or without invaded bacteria into LesM with various deformation channel widths, flow velocities, and cell concentrations (Fig. 3e and Supplementary Fig. 4b–h). As expected, the proportion of squeezed-cell efficiency grew under higher flow velocity and larger width of deformation channel. And infected cells exhibited a higher squeezed-cell efficiency than native cells, resulting from reorganization of the stress fibers via the intracellular bacteria. From these experiments, we determined the optimal experimental conditions for infected MDA-MB-231 cell screening: a cell concentration of 1 × 106 cells·mL-1, a flow velocity of 5 μL·min-1, and a deformation channel width of 6 μm. Under these conditions, the squeezed-cell efficiency was 23.1% for the native cells and 41% for the infected cells. Of note, the squeezed-cell efficiency grew in a bacteria concentration–dependent manner, which resembled the cells treated with a ROCK inhibitor (Supplementary Fig. 4i), further verifying that infected cells gained their deformability via RhoA-ROCK suppression caused by bacterial infection.
a (i) Photograph of 6 parallel microfluidic chips filled with different dyes to aid visualization. (ii) Optical micrograph of microfluidic chip. Scale bar, 200 μm. (iii) SEM image of L-shaped traps. Scale bar, 100 μm. (iv) SEM image of one L-shaped trap. Scale bar, 10 μm. b Schematic of blocked-cell and squeezed-cell ROIs of L-shaped trap. Scale bar, 10 μm. c Simulation of the flow field around the traps in one of the 32 channels. d Representative optical micrograph showing high single-cell capture efficiency of MDA-MB-231 cells, with Hoechst-labeled cells for convenient visualization. Scale bar, 400 μm. e, f Single-, blocked-, and squeezed-cell capture efficiencies with respect to various flow velocities and deformation channels widths of (e) MDA-MB-231 cells and (f) MCF-7 cells with or without invaded bacteria. Note that the innermost circles represent the smallest channel diameters and the outermost, the largest channel diameters. Totals indicate single-cell efficiencies for smallest to largest channel diameters (e.g., efficiency for 5-μm width channel/efficiency for 6-μm width channel/efficiency for 7-μm width channel). ROI region of interest, SEM scanning electron microscopy.
Similar experiments were performed with MCF-7 cells (Fig. 3f and Supplementary Fig. 5), and we determined the optimal experimental conditions for infected MCF-7 cell screening: a flow velocity of 10 μL·min-1 and a deformation channel width of 7 μm, under which, the squeezed-cell efficiency was 30.6% for native cells and 54.9% for infected cells. Considering its highly efficient single-cell capture and its ability to discriminate between native and infected cells via squeezed-cell efficiency, we were encouraged that the LesM offered a feasible approach to screen the blood for infected CTCs.
On-chip screening for infected breast cancer cells
Next, we tested the ability of LesM to distinguish infected cancer cells from native cells with fluorescent probes to label intracellular bacteria. First, we confirmed the feasibility of bacterial labeling via DiO (Supplementary Fig. 6a–b). Next, after native MDA-MB-231 cells were stained with Hoechst and S. xylosus and L. animalis were labeled with DiO, the bacteria were applied to some cells for invasion, and heterogeneous biological suspensions composed of both native and infected MDA-MB-231 cells were introduced into the LesM (Fig. 4a). Blue and green fluorescence was measured from the 2 ROIs of each trap (Fig. 4b–c). For S. xylosus–infected cells, the blue fluorescence intensity of blocked-cell ROIs was higher, while squeezed-cell ROIs showed the opposite (Fig. 4d-e). A density plot of fluorescence from the 2 ROIs also showed a clear difference between blocked- and squeezed- cell ROIs (Fig. 4f). And the respective analysis of the receiver operating characteristic (ROC) curve for 5 independent experiments showed coherent results with a mean area under the ROC curve (AUC) of 0.923 (Fig. 4g), verifying a connection between intracellular bacteria and the deformability of their host cells.
a Schematic showing experimental procedure. Native MDA-MB-231 cells were stained with Hoechst and bacteria were labeled with DiO before infection. A mixture of native and infected cells was added to LesM chip under the optimal conditions of a cell concentration of 1 × 106 cells·mL-1, a flow velocity of 5 μL·min-1, and a deformation channel width of 6 μm. Icons were created in BioRender. Luo, W. (https://BioRender.com/l1e6nxj). b, c Representative micrographs showing blocked-cell and squeezed-cell ROIs, with blue fluorescence indicating Hoechst staining and green, DiO-labeled bacteria. Scale bars, 100 μm. Cells are MDA-MB-231 cells infected with (b) S. xylosus or (c) L. animalis. d–f Data showing fluorescence from 2 ROIs after MDA-MB-231 cells, some invaded by S. xylosus and some not, were added to the chip. Violin plots of fluorescence from (d) blocked-cell ROIs, (e) squeezed-cell ROIs, and (f) density plot. Each dot represents one cell, with n = 800 (biologically independent). Lower boundary of the shaded regions represents the threshold of meaningful fluorescence; below this boundary, any fluorescence is considered background fluorescence, and numbers 1-5 represent 5 independent experiments under the same conditions. g ROC curves of 5 replicated tests for the discrimination of S. xylosus–infected MDA-MB-231 cells by fluorescence from squeezed-cell ROIs. h–j Data showing fluorescence from 2 ROIs after MDA-MB-231 cells, some invaded by L. animalis and some not, were added to the chip. Violin plots of fluorescence from (h) blocked-cell ROIs, (i) squeezed-cell ROIs, and (j) Density plot. k ROC curves of 5 replicated tests for the discrimination of L. animalis–infected MDA-MB-231 cells by fluorescence from squeezed-cell ROIs. AUC area under the ROC curve, bac bacteria, BF bright-field microscopy, DiO 3,3’-dioctadecyloxacarbocyanine perchlorate, FL fluorescence intensity, ROC receiver operating characteristic, ROI region of interest.
For L. animalis–infected cells, similar results were observed (Fig. 4h–k). Unexpectedly, respective analysis of the ROC curves showed a mean AUC of 0.891, lower than that associated with S. xylosus, despite their infection causing comparable disassembly of actin stress fibers in MDA-MB-231 cells. Taken together, these data suggested that infected cells, instead of native cells, squeezed into the deformable channel.
Since DiO is not specific for bacterial cells, and to further solidify the connection between intracellular bacteria and the deformability of their host cells, we synthesized FITC-vancomycin–based probes, which can specifically label intracellular bacteria in living cells, according to previously reported methods36,37 (Supplementary Fig. 6c). After native MDA-MB-231 cells were stained with Hoechst and S. xylosus and L. animalis were applied to some cells for invasion followed by FITC-vancomycin–based labeling, and heterogeneous biological suspensions were introduced into the LesM (Supplementary Fig. 7a). Like the experiments with DiO, blue and green fluorescence was measured from the 2 ROIs of each trap, and the results were similar (Supplementary Fig. 7b–k). The respective analysis of the ROC curves showed a mean AUC of 0.924 for S. xylosus and 0.888 for L. animalis.
We also conducted on-chip screening for infected cells using MCF-7 cells, in which some cells were infected by DiO-labeled S. xylosus (Supplementary Fig. 8a–d) or L. animalis (Supplementary Fig. 8e–h). Using the LesM, we were able to distinguish infected cells from native cells, with a mean AUC of 0.868 for S. xylosus and 0.851 for L. animalis. These AUCs were lower than those achieved with MDA-MB-231 cells, probably resulting from the lower invasion rate of MCF-7 cells by bacteria. Overall, these results show that on-chip screening, via biomechanical phenotyping at the single-cell level, was able to distinguish breast cancer cells containing intracellular bacteria from those without, demonstrating LesM as a promising platform for the screening of CTCs for metastatic potential.
On-chip screening of whole blood for infected cancer cells
Encouraged by our results, we performed the on-chip analysis of clinical specimens, using whole blood samples spiked with infected cancer cells to mimic blood samples containing infected CTCs. The infected cancer cells were MDA-MB-231 cells, stained with Hoechst and invaded by DiO-labeled S. xylosus (Fig. 5a). Blue and green fluorescence from the squeezed-cell ROI of each trap was measured (Fig. 5b). We observed that spiked cells got trapped in the deformation channels, while blood cells escaped from the traps because of their relatively small dimensions. An analysis of fluorescence intensities from all squeezed-cell ROIs in non-empty traps for 5 independent experiments showed apparent groupings of cancer cells with higher fluorescence and blood cells with only background fluorescence (Fig. 5c and Supplementary Fig. 9a). To further confirm the specific screening of cancer cells in spiked blood samples, we measured the area of each captured cell. Similar groupings were observed (Fig. 5d–e, Supplementary Fig. 9b, c, and Supplementary Fig. 10a, b), where dimensions of captured cells grew in a fluorescence-intensity-related manner, indicating a high sensitivity of the monitoring of intracellular bacteria within captured cancer cells. ROC analysis based on the dimension of host cells and intracellular bacteria verified that the LesM managed to screen out infected cancer cells from a large population of blood cells (Fig. 5f–g).
a Schematic showing experimental procedure. Whole blood was spiked with MDA-MB-231 cells, stained with Hoechst and invaded by DiO-labeled S. xylosus or L. animalis, and added to the LesM chip. Icons were created in BioRender. Luo, W. (https://BioRender.com/l1e6nxj). b Representative micrographs showing captured MDA-MB-231 cells (Hoechst-stained), intracellular bacteria (DiO-labeled), and blood cells (unstained). Scale bars, 100 μm. c–e Scatter plots of blood cells and spiked S. xylosus –invaded MDA-MB-231 cells, with n = 2194 (biologically independent). Each dot represents one cell. Box plots: median, 25% to 75% boxes, whiskers (-1.5 box to 1.5 box), and outliers indicated. f, g ROC curves of 5 replicated tests for the discrimination of spiked S. xylosus–infected MDA-MB-231 cells by (f) cell area and (g) DiO fluorescence intensity. h–j Scatter plots of blood cells and spiked L. animalis–invaded MDA-MB-231 cells, with n = 2340 (biologically independent). Each dot represents one cell. Box plots: median, 25% to 75% boxes, whiskers (-1.5 box to 1.5 box), and outliers indicated. k, l ROC curves of 5 replicated tests for the discrimination of spiked L. animalis–infected MDA-MB-231 cells by (k) cell area and (l) DiO fluorescence intensity. AUC area under the ROC curve, bac bacteria, BF bright-field microscopy, DiO 3,3’-dioctadecyloxacarbocyanine perchlorate, ROC receiver operating characteristic.
We repeated these experiments by spiking blood with L. animalis–invaded MDA-MB-231 cells. Successful retrieval of the rare infected cancer cells was demonstrated by the statistical analysis of 5 independent experiments, in which cancer cells exhibited higher fluorescence intensity and larger dimensions (Fig. 5h–j, Supplementary Fig. 9d–f, and Supplementary Fig. 10c, d). Despite the DiO fluorescence intensity of L. animalis being weaker than that of S. xylosus (Supplementary Fig. 10a, c), both cell dimension and intracellular bacteria permitted the recognition of infected cancer cells in whole blood, as confirmed by relative ROC curve analysis (Fig. 5k–l).
We also repeated these experiments with MCF-7 cells and found that, like infected MDA-MB-231 cells, infected MCF-7 cells exhibited higher fluorescence intensity and larger dimensions than that of blood cells (Supplementary Figs. 11 and 12). Compared with infected MDA-MB-231 cells, infected MCF-7 cells showed weaker fluorescence intensity and smaller dimensions. Overall, the results demonstrate that unprocessed whole blood samples can be applied to the LesM for the highly efficient capture of cancer cells and the sensitive detection of their intracellular bacteria.
Promotion of metastatic tendency of host cells
After demonstrating that intracellular S. xylosus reorganized actin stress fibers in breast cancer cells, increasing host cell deformability, we assessed whether these bacteria could trigger metastasis in a breast cancer model. BALB/c nude mice bearing an MDA-MB-231 cell xenograft were intratumorally injected with PBS or with various amounts of bacteria. After a 10-day observation, all mice were sacrificed and their major organs were obtained and evaluated for the presence of bacteria (Supplementary Fig. 13).
The lungs, which previously contained few bacteria, showed bacterial growth after intratumoral injection, demonstrating that, along with its host CTCs, S. xylosus traveled through the circulatory system and colonized in the lungs. The fact that only one injection of S. xylosus was able to trigger lung metastasis—and within only 10 days—reinforces the role of certain bacteria in promoting metastasis.
In vivo evaluation of metastatic ability of infected CTCs
After preliminary experiments showed that intratumoral bacteria triggered lung metastasis, we investigated the ability of LesM to evaluate metastatic potential of cancer cells by screening blood for infected CTCs. BALB/c nude mice bearing an MDA-MB-231 cell xenograft (~100 mm3) were randomly divided into 3 groups: (G1) a control group, (G2) a Bac+ group, and (G3) a Bac + + group. Following a single intratumoral injection of PBS or bacteria, whole blood samples were collected at various time points and infected CTCs were analyzed (Fig. 6a) via a 6-step protocol (Fig. 6b).
a Schematic showing experimental procedure. BALB/c nude mice bearing an MDA-MB-231 cell xenograft received a single injection of PBS (G1), 2 × 106 CFU of S. xylosus (G2), or 5 × 106 CFU of S. xylosus (G3), and whole blood was collected on days 5, 10, 15, or 30 followed by metastasis evaluation. Icons were created in BioRender. Luo, W. (https://BioRender.com/l1e6nxj). b Representative micrographs showing on-chip evaluation procedure: (i) initial, empty chip, (ii) purge of air inside the chambers, (iii) injection of mouse blood sample, (iv) injection of sterile PBS, (v) injection of FITC-vancomycin–based probes, (vi) observation under fluorescence microscope. Scale bar, 100 μm. c Representative fluorescence micrographs of captured CTCs on various days. Scale bars, 100 μm. d Lungs on day 30, according to (i) hematoxylin and eosin staining of sections and (ii) photographs, with colonized areas circled. Scale bar, 200 μm. e–h Box and scatter plots of FITC fluorescence intensities from mouse blood samples collected on (e) day 5, (f) day 10, (g) day 15, or (h) day 30, with n = 800 (biologically independent). Each dot represents one cell. Box plots: median, 25% to 75% boxes, whiskers (-1.5 box to 1.5 box), and outliers indicated. Lower boundary of the green shaded regions represents the threshold of meaningful FITC fluorescence; below this boundary, any fluorescence is considered background fluorescence. Graphs indicate statistical significance according to two-sided, unpaired student’s t-test versus G1. i, j (i) Counts and (j) FITC fluorescence intensities of captured CTCs on various days. Each bar represents mean and SD, with n = 3 (biologically independent). FITC fluorescein isothiocyanate, FL fluorescence intensity, G1 group 1, G2 group 2, G3 group 3.
Captured CTCs were characterized by larger dimensions and green fluorescence of intracellular bacteria. It was noticed that blood samples collected from different time points exhibited different intracellular bacteria levels in the captured CTCs, as revealed by FITC-vancomycin fluorescence (Fig. 6c). The number of infected CTCs captured on the chip grew in a time-dependent manner in G2 and G3, but not in G1 (Fig. 6e–i and Supplementary Figs. 14–17). And the average FITC fluorescence intensity generally increased in a time-dependent manner in G2 and G3, while samples from G1 showed only a background fluorescence intensity (Fig. 6j). The increase in infected CTCs in G2 and G3 was accompanied by abnormal proliferation in lung tissues, observed after 30 days (Fig. 6d and Supplementary Fig. 18e). In addition, the observation of bacteria in lung tissues in G2 and G3 indicated that host cells carrying intracellular bacteria traveled to the lungs, causing metastasis (Supplementary Fig. 18f). Of note, the intratumoral administration of bacteria showed limited influence on the mice (Supplementary Fig. 18a and Supplementary Fig. 19) or on the growth of primary tumors (Supplementary Fig. 18b–d).
The consistency between the LesM estimation of infected CTCs and the in vivo assays of lung metastasis indicate that the LesM offers an opportunity to screen the more metastatic CTCs in blood samples with high reliability, allowing a platform for the prediction of tumor metastasis and recurrence in the clinic.
In vivo evaluation of the protective effect of antibiotics
Since tumor-resident bacteria promoted metastasis, we reasoned that antibiotic treatment would impede metastasis. However, the protective effect of antibiotic treatment in cancer patients remains unclear, since the clinical use of antibiotics can produce different outcomes depending on their combination with different therapeutics. To establish the influence of antibiotic treatment on metastasis, we conducted in vivo experiments with antibiotics.
BALB/c nude mice bearing an MDA-MB-231 cell xenograft (~100 mm3) were randomly divided into 4 groups: (G1) a pre-resection group, assigned to receive antibiotics only before tumor resection; (G2) a post-resection group, assigned to receive antibiotics only after tumor resection; (G3) a full-course group, assigned to receive antibiotics before and after resection; and (G4) a no-medication group, assigned to receive no antibiotics (Fig. 7a). All mice received an intratumoral injection of 5 × 106 CFU of S. xylosus on day 0 and tumor resection on day 10. Mice receiving antibiotics were given an antibiotic cocktail (ATBx) through drinking water38,39, which eliminated tumor microbiota efficiently (Fig. 7b and Supplementary Fig. 20a). Whole blood samples were collected on day 30 and screened for infected CTCs. In addition to labeling intracellular bacteria, we also assessed CTC expression of EpCAM (epithelial cell adhesion molecule), believed to be involved in carcinogenesis (Fig. 7ci). CTCs from G1 and G3 showed relatively low levels of intracellular bacteria, while CTCs from G2 and G4 showed the opposite (Fig. 7d, e and Supplementary Fig. 21). Comparable average fluorescence intensities were observed in all captured cells, but the numbers of captured cells varied among groups (Fig. 7f, g). Statistical analysis demonstrated that the number of captured CTCs decreased in the order of G4, G2, G1, and G3, indicating a protective effect against metastasis in mice that received antibiotics before tumor resection. Lung tissues were also obtained and analyzed for abnormal proliferation (Fig. 7cii-iii and Supplementary Fig. 20b). The more severe cancerous proliferations were observed in G2 and G4, consistent with the blood sample results. Moreover, the presence of the bacteria in the lungs in G2 and G4 was observed (Supplementary Fig. 20c, e), along with the staining of lung tissue sections, demonstrating that bacteria-triggered metastasis occurred despite the 20-day antibiotic treatment in G2.
a Schematic showing experimental procedure. BALB/c nude mice bearing an MDA-MB-231 cell xenograft received a single intratumoral injection of 5 × 106 CFU of S. xylosus on day 0. Tumor resection occurred on day 10 and metastasis evaluation, on day 30. Mice received an antibiotic cocktail every other day in their drinking water (ATBx-DW), either before resection (G1), after resection (G2), before and after resection (G3), or never (G4). Icons were created in BioRender. Luo, W. (https://BioRender.com/l1e6nxj). b Bacterial growth in tumor tissues of mice that received no antibiotics or ATBx on day 10. Each bar represents mean and SD, with n = 3 (biologically independent). c Captured CTCs and comparison of lung metastasis among various groups on day 30. (i) Representative fluorescence micrographs of captured CTCs, with green fluorescence indicating intracellular bacteria labeled with a FITC-vancomycin probe and red fluorescence indicating EpCAM labeled with a Cy3-conjugated secondary antibody (scale bar, 100 μm). Lungs on day 30, according to (ii) hematoxylin and eosin staining of sections (scale bar, 400 μm) and (iii) photographs, with colonized areas circled. d, e Box and scatter plots of (d) FITC or (e) Cy3 fluorescence intensities from mouse blood samples collected on day 30, with n = 800 (biologically independent). Each dot represents one cells. Box plots: median, 25% to 75% boxes, whiskers (-1.5 box to 1.5 box), and outliers indicated. Lower boundary of the shaded regions represents the threshold of meaningful fluorescence; below this boundary, any fluorescence is considered background fluorescence. Graphs indicate statistical significance according to two-sided, unpaired student’s t-test versus G4. f, g (f) Fluorescence intensities and (g) counts of captured CTCs on day 30. Each bar represents mean and SD, with n = 3 (biologically independent). ATBx-DW an antibiotic cocktail in drinking water, CFU colony-forming unit, CTL control, Cy3 cyanine 3, FITC fluorescein isothiocyanate, FL fluorescence intensity, G1 group 1, G2 group 2, G3 group 3, G4 group 4, S. x. Streptococcus xylosus.
The consistency between CTC analysis via LesM and the in vivo assays indicates that antibiotic treatment before, but not after, tumor resection was associated with a better response in metastatic breast cancer, possibly resulting from the fact that preoperative antibiotics may reduce the bacterial load prior to the introduction of CTCs, thereby limiting bacteria-mediated alterations in CTC mechano-phenotypes that promote metastasis. To confirm this hypothesis, we have repeated in vivo experiments with antibiotics and noticed that preoperative antibiotics resulted in fewer infected CTCs in the blood as well as less abnormal proliferation in lung tissues (Supplementary Fig. 22). Once tumor-resident microbes are detected, the perturbation of intracellular bacteria through appropriate antibiotic treatment could be a promising way to impede metastasis in breast cancer in the clinic. Of note, ATBx treatment triggered a slight loss in body weight in mice (Supplementary Fig. 20d), but the influence was limited, since no severe damage in major organs was observed (Supplementary Fig. 23).
On-chip screening of clinical specimens for infected CTCs
Encouraged by the results above, we performed the on-chip analysis of clinical specimens, using whole blood samples from patients with breast cancer, who were divided into 3 groups: (G1) non-metastatic group, (G2) lymphatic metastatic group, and (G3) distal metastatic group (Fig. 8a, Supplementary Data 1 and Supplementary Movie 2). Red and green fluorescence from the squeezed-cell ROI of each trap was measured (Fig. 8b). Multiple factors were measured among 69 breast cancer patients (Fig. 8c), and we noticed that the numbers of captured CTCs varied between 3 groups, where clinical specimens from G1 showed very little infected CTCs, with 17 samples showed no cells at all; specimens from G2 showed relatively low quantity of infected CTCs, with 2 samples showed no cells at all; and specimens from G3 showed more CTCs with intracellular bacteria, with 7 samples showing more than 10 cells (namely #2, 5, 7, 10, 12, 13 and 18) (Fig. 8d). Detection rates for infected CTCs were 18/35, 32/34, and 18/18 for all patients, metastatic patients, and distal metastatic patients, respectively (Fig. 8e). Moreover, we noticed the distribution of the number of infected CTCs of G1 resembled to that of G2, but differed from G3 (Fig. 8f), indicating CTCs with intracellular cells as potential targets for the detection of distant metastasis, instead of lymphatic metastasis in breast cancer.
a Schematic showing experimental procedure. Whole blood samples were collected from BC patients and added to the LesM chip. Patients were diagnosed with non-metastatic BC (G1), lymphatic metastatic BC (G2), or distal metastatic BC (G3). Icons were created in BioRender. Luo, W. (https://BioRender.com/l1e6nxj). b Representative micrographs showing captured CTC with intracellular bacteria, with green fluorescence indicating intracellular bacteria labeled with a FITC-vancomycin probe and red fluorescence indicating EpCAM labeled with a Cy3-conjugated secondary antibody. Scale bars, 100 μm. c Heatmap showing the infected CTC positivity, count, and time to diagnosis among BC patients from G1, G2, or G3. d Box and scatter plots of FITC fluorescence intensities from clinical specimens of patients, with n = 800 (biologically independent). Each dot represents one cell. Box plots: median, 25% to 75% boxes, whiskers (-1.5 box to 1.5 box), and outliers indicated. Lower boundary of the shaded regions represents the threshold of meaningful fluorescence; below this boundary, any fluorescence is considered background fluorescence. e Detection rate of BC patients. f Half violin plot of the counts of captured CTCs with intracellular bacteria. g The ROC curve of 69 tests for the discrimination of patients with distal metastatic BC by infected CTCs. AUC area under the ROC curve, bac bacteria, BC breast cancer, BF bright-field microscopy, Cy3 cyanine 3, FITC fluorescein isothiocyanate, FL fluorescence intensity, G1 group 1, G2 group 2, G3 group 3, ROC receiver operating characteristic.
Considering that the probability of bacterial infection increased during breast cancer progression, the number of CTCs with intracellular bacteria increased accordingly, and thus the risk of distal metastasis grew, contributing to the unevenness of time to diagnosis in various groups. The relative ROC curve analysis confirmed that infected CTCs permitted the recognition of distal metastasis in breast cancer, with an AUC of 0.904 (Fig. 8g). Overall, the successful screening of infected CTCs in clinical specimens indicates that LesM could be a promising option for the prediction of metastatic tendency to the distal organs of bacteria-infected breast cancer in the clinic.
Discussion
In this study, we present a microfluidic platform enabling high-throughput, single-cell, and time-saving screening of deformable CTCs with increased metastatic potential via the biomechanical characteristics of captured cells. With the disassembly of stress fibers in host-cells, certain bacteria associated with breast cancer could trigger the suppression of RhoA-ROCK activation and thus form soft and deformable CTCs with lower Young’s modulus. The resulting deformability could avoid cell death due to fluid shear stress during circulation, promoting metastasis in distant tissues.
The screening of infected CTCs on the LesM was achieved via the deformation channel in the L-shaped captured trap, where only infected cells with intracellular bacteria showing a deformable biomechanical phenotype could squeeze through the channel and native cells were blocked for their stiffness. Through the fluorescence analysis of blocked- and squeezed-cell ROIs, an apparent grouping of native cells and infected cells was observed, validating the deformability-based sorting system of the LesM. Moreover, the LesM screening of infected CTCs among a large amount of massive blood cells in mimicking blood samples and clinical specimens, as well as the consistency between the LesM estimation on CTCs with intracellular bacteria and the in vivo assays of lung metastasis in mice demonstrated a potential platform for CTC evaluation of metastatic potential in the clinic. Collectively, this deformability-based sorting microfluidic system offered a promising option for cancer metastasis and recurrence prediction, as well as the evaluation of the protective effects of antibiotic treatment in cancer patients on metastasis.
One thing remaining to be discussed is the vast diversity of tumor microbiota, where other bacterial species may influence CTC deformability and metastatic potential. In the current study, we have focused on the four intratumoral bacterial strains identified in breast cancer tissues in previous studies, aiming to establish a proof-of-concept for the LesM chip in assessing bacteria-mediated CTC mechano-phenotypes. Given the multifactorial nature of tumor-resident microbiota among broad cancer types, these intracellular bacteria might be a deserving target to monitor metastasis in the clinic, where the LesM chip could provide a solid foundation for future studies extending to additional cancer types.
Methods
Materials
All chemical reagents were of analytical grade and used directly, without further purification, unless otherwise noted. Actin-Tracker Red-555, DiO, Hoechst 33342, Triton X-100 solution, Cy3-labeled goat anti-mouse IgG (H + L), and CCK-8 assay kits were purchased from Beyotime Chemical Reagent Co. Ltd. (Jiangsu, China). Brain heart infusion broth (BHI) was purchased from Dalian Bergolin Biotechnology Co. Ltd. (Liaoning, China). The 5-fluorescein isothiocyanate (FITC), vancomycin hydrochloride hydrate, and Columbia blood agar (CBA) base were purchased from Aladdin Chemical Co. Ltd. (Shanghai, China). Defibrinated sheep blood, Dulbecco’s modified Eagle’s medium (DMEM), penicillin–streptomycin solution, trypsin–EDTA solution with phenol red, and fetal bovine serum (FBS, BC-SE-FBS08) were purchased from BioChannel Biotechnology Co. Ltd. (Jiangsu, China). ROCK inhibitor Y27632 was purchased from J&K Scientific (Shanghai, China). Ampicillin sodium, penicillin–streptomycin–gentamicin (P–S–G) solution, doxycycline, clarithromycin, and azithromycin were purchased from Sangon Biotech Co. Ltd. (Shanghai, China). Purified anti-human CD326 (EpCAM) antibody (Cat. No. 369802) was purchased from Biolegend (USA). SU-8 3025 was purchased from MicroChem Corp. (Newton, MA, USA). Trimethyl-chloro-silane was purchased from Sigma-Aldrich. Polydimethylsiloxane (PDMS, Sylgard 184) was purchased from Dow Corning Corp., (Midland, MI, USA).
Characterizations
The morphology of LesM chip was detected by SEM (JSM-7800F; JEOL Ltd.). The optical densities were measured on a microplate photometer (Multiskan FC; Thermo Fisher Scientific Inc.). Flow cytometry data were acquired on a flow cytometer (Beckman). Optical and fluorescence micrographs were captured on EVOS FL Auto microscope (Thermo Fisher Scientific Inc.). LSCM images were captured on Olympus FV1200 (Olympus). Young’s modulus was measured on AFM (JPK Nanowazrid 4 XP BioScience, Bruker).
Strains and growth of bacteria
S. xylosus, E. faecalis, L. animalis, and S. cuniculi were a generous gift from Dr. Shang Cai (Westlake University), and were primarily isolated using a murine spontaneous breast-tumor model MMTV-PyMT (FVB/N-Tg(MMTV-PyVT)634Mul/J). All bacterial strains were routinely grown at 37 °C with shaking (200 rpm), unless otherwise noted. Strains were cultured overnight by inoculating 10 μL of a bacterial suspension into 10 mL of liquid BHI medium. They were collected by centrifugation at 2000 × g for 3 min and resuspended with PBS to an optical density (OD) of 0.8, corresponding to a bacterial concentration of ~1 × 108 CFU·mL-1.
Design and fabrication of the LesM chips
The LesM chip was designed via the computer-aided design software AutoCAD (Autodesk Inc., Mill Valley, CA, USA), and the corresponding chrome mask was created with a resolution of 1 µm via a Heidelberg laser write MLA 150 (Heidelberg Instruments Mikrotechnik GmbH, Heidelberg, Germany). The negative photoresist SU-8 3025 was spin-coated onto a clean and dry 4-inch silicon wafer at a given speed to achieve the desired thickness. After soft-baking, the SU-8 layer was patterned by a mask aligner MA6/BA6 (SÜSS MicroTec SE, Garching, Germany) under 365 nm UV to fabricate the designed microfluidic architecture, followed by development and hard–baking to make the mold. The mold was pretreated with trimethyl-chloro-silane vapor for 30 min to facilitate mold release. Afterward, PDMS was poured onto the SU-8 master with a 10:1 ratio of base to crosslinker, degassed in a vacuum chamber for 60 min, and cured at 70 °C for 3 h. The PDMS elastomers were peeled off the molds and then cut into suitable sizes. Inlet/outlet ports were subsequently punched with a Miltex biopsy punch (Integra LifeSciences, Princeton, NJ, USA). After oxygen plasma treatment for 1 min, the PDMS slabs were irreversibly bonded onto glass slides and kept at 80 °C on a hotplate for 20 min to enhance bonding. Channel width was measured optically by a Zeiss AX10 microscope (Carl Zeiss AG, Oberkochen, Germany), and thickness was measured via a surface profilometer Dektak 150 (Veeco Instruments Inc., Plainview, NY, USA).
Computational fluid dynamics simulation of the flow field
The simulation of the flow field surrounding the traps in the microchannel was conducted via a commercial finite element solver (COMSOL Multiphysics v.5.4, COMSOL Inc., Burlington, MA, USA). The transient flow field was modeled by solving the incompressible Navier–Stokes equations in the time-dependent domain, as expressed below:
where \(\rho\) is the fluid density, \({{{\bf{u}}}}_{{\rm{fluid}}}\) is the fluid velocity field, \(p\) is the pressure, \({{\bf{K}}}\) is the viscous stress tensor defined as
and \(\mu\) is the dynamic viscosity, while \({{\bf{F}}}\) represents external body forces (e.g., gravity).
These equations were solved using a time-dependent laminar flow module coupled with the moving mesh formulation, allowing both the velocity and pressure fields to evolve dynamically as the cell deforms and interacts with the capture structure. The cell was modeled as a viscoelastic solid with mechanical properties consistent with reported ranges for mammalian cells. The material parameters were set as follows: density ρ = 1050 kg/m3, dynamic viscosity μ = 0.02 Pa·s, Young’s modulus E = 1000 Pa, and Poisson’s ratio ν = 0.49.
Cell culture
Human breast cancer cells MDA-MB-231 and MCF-7 were obtained in-house but are available from the American Type Culture Collection. All cells were cultured in DMEM supplemented with 10% FBS and 1% penicillin–streptomycin in a humidified atmosphere of 95% air and 5% CO2 at 37 °C.
Viability of cells exposed to antibiotics and bacteria
The viability of cells exposed to antibiotics was quantitatively evaluated by CCK-8 assays. MDA-MB-231 and MCF-7 cells were incubated for 24 h in 96-well plates at 37 °C under 5% CO2 and were then exposed for 4 h to ampicillin and P–S–G solution at various concentrations. CCK-8 solution (10 μL) was added to each well, and the culture plate was incubated for another 2 h before the OD450 values of each well were measured. The viability of cells exposed to bacteria was evaluated similarly, where cells were exposed to bacterial suspensions (S. xylosus, E. faecalis, L. animalis, or S. cuniculi) in the absence of any antibiotics for 4 h and washed with antibiotics-added PBS for 3 times before CCK-8 solution was added.
Invasion efficiency of bacteria
MDA-MB-231and MCF-7 cells were exposed to bacterial suspensions (S. xylosus, E. faecalis, L. animalis, or S. cuniculi) at various concentrations for 4 h in the absence of any antibiotics. Cells were digested, collected by centrifugation (100 × g, 5 min), washed 3 times with sterile PBS, and dispersed in sterile PBS. The cell suspensions were serially diluted and then plated onto solid CBA plates. The colonies on the plates were recorded after 12 h at 37 °C. Infected cells were obtained from the coculture of 106 CFU live microorganisms and 104 cancer cells for further experiments.
Confocal fluorescence imaging
MDA-MB-231 and MCF-7 cells were seeded onto 10-mm2 glass coverslips placed in 6-well plates and incubated with bacteria (S. xylosus, E. faecalis, L. animalis, or S. cuniculi) pre-labeled with DiO for 4 h in the absence of any antibiotics in the dark. After washing with PBS 3 times, cells were labeled with Hoechst and imaged by a laser scanning confocal fluorescence microscope. Reorganization of the cytoskeleton was observed similarly, where cells were treated with bacteria (S. xylosus, E. faecalis, L. animalis, or S. cuniculi) for 4 h in the absence of any antibiotics in the dark, followed by treatment with P–S–G solution for 4 h to kill the extracellular bacteria. Cells were cultured for another 24 h before they were labeled with Actin-Tracker Red-555 and Hoechst according to manuals.
Preparation of FITC-vancomycin probe
Vancomycin hydrochloride hydrate was added to a solution of NaHCO3/Na2CO3 buffer (pH 9) to a final concentration of 1 mg·mL-1, and FITC solution (10 mg·mL-1, 20 μL) was added drop-by-drop. The reaction mixture was kept in the dark for 12 h with constant stirring at 4 °C in a refrigerator. The resulting solution was purified, redissolved with sterile PBS, diluted to a final concentration of 0.05 mg·mL-1, and sterilized with 0.22-μm filters before use.
Optimization of on-chip cell capture
Each LesM chip contained deformation channels of various widths (5, 6, 7, 8, 9, or 10 µm). After a LesM chip was filled with PBS buffer, a syringe pump (Longer Precision Pump Co., Ltd., Baoding, Hebei, China) operating at one of several flow rates (2, 3, 5, 8, or 10 µL·min-1) was used to introduce 20 µL of a native MDA-MB-231 cell suspension (0.25, 0.5, 1, 2, 5, 10, or 20 × 106 cells·mL-1) into the chip. Microscopic observation was performed once all the traps were occupied. The single-cell efficiencies were calculated from cells captured on the whole trap, which were defined as the proportion of traps capturing one cell, regardless of which part of the trap it was in. The blocked-cell efficiencies were defined as the proportion of traps capturing one cell on the hook-shaped part, and the squeezed-cell efficiencies were defined as the proportion of traps capturing one cell on the deformation channel part.
The procedure was similar for detecting the single-, blocked-, and squeezed-cell efficiencies of infected MDA-MB-231 cells, where cells were pretreated with bacteria, and for detecting the capture rate of MCF 7 cells, where MCF-7 cells were introduced into chips containing deformation channels with one of several widths (5, 6, 7, 8 µm) subjected to one of several flow rates (2, 3, 5, 8, or 10 µL·min-1).
On-chip detection of bacteria-containing cells
A LesM chip with a 7-µm deformation channel was filled with PBS buffer. A syringe pump, operating at a flow rate of 5 µL·min-1, was used to introduce 20 µL of a mixed MDA-MB-231 cell suspension (2 × 106 cells·mL-1), containing both native cells and infected cells, into the chip. The native cells had been previously stained with Hoechst, while the infected cells had been previously treated with DiO-labeled bacteria (S. xylosus or L. animalis). Once all the traps were occupied, microscopic observation was performed and 3 channels (bright-field [transmitted light], blue [Hoechst], and green [DiO or FITC]) were used to generate images with complete optical information. Two ROIs from each trap were defined through bright-field images, namely the hook-shaped part and the deformation channel part. The fluorescence intensities of each ROI were measured in both blue and green channels for further statistical analysis. The procedure for FITC-vancomycin labeling was the same, except that the infected cells were stained with FITC-vancomycin probes. The detection of intracellular bacteria in MCF-7 cells was similar; MCF-7 cells were introduced into a chip with a 6-µm deformation channel, and subjected to a flow rate of 8 µL·min-1.
On-chip detection of infected tumor cells–spiked whole blood
Fresh blood samples were acquired from healthy volunteers from Nanjing First Hospital and informed consent was obtained from all participants before sample collection. Typically, 2 mL of peripheral blood per individual was collected and stored in plastic blood collection tubes with the anticoagulant tripotassium ethylenediaminetetraacetic acid (K3EDTA). All samples were kept at 4 °C in a refrigerator and were typically analyzed within 48 h after blood collection. To obtain cancer cell–spiked whole blood samples, we mixed 10 µL of an infected cancer cell suspension (1 × 105 cells·mL-1) with 990 µL of whole blood, resulting in a concentration of 103 cells·mL-1. For on-chip analysis, 100 μL of blood spiked with cancer cells pretreated with Hoechst and DiO-labeled bacteria was introduced into the chip for cell capture. After the blood was introduced, PBS was immediately flowed through the microchannels at a flow rate of 10 µL·min-1 to flush away the residual blood cells. Finally, microscopic observation was performed.
On-chip detection of infected CTCs in clinical specimens
Fresh blood samples were acquired from breast cancer patients at Nanjing First Hospital, Jiangsu Cancer Hospital, and Jiangsu Province Hospital of Chinese Medicine and informed consent was obtained from all participants before sample collection. Typically, 2 mL of peripheral blood per individual was collected and stored in plastic blood collection tubes with the anticoagulant tripotassium ethylenediaminetetraacetic acid (K3EDTA). All samples were kept at 4 °C in a refrigerator and typically analyzed within 48 h after blood collection. For on-chip analysis, 100 μL of blood was introduced into the chip for cell capture. After the blood was introduced, PBS was immediately flowed through the microchannels at a flow rate of 10 µL·min-1 to flush away the residual blood cells. Following introduction of the FITC-vancomycin solution, captured cells were fixed, blocked, and stained with anti-EpCAM primary antibody and Cy3-conjugated secondary antibody. Finally, microscopic observation was performed.
Animal model of breast cancer
Animal procedures were performed according to the guidelines of the Institutional Animal Care and Use Committee of Nanjing University. All animal experiments were performed in accordance with the Regulations for the Administration of Affairs Concerning Experimental Animals, approved by the State Council of the People’s Republic of China. All animals were bred in a pathogen-free facility with a 12 h light/dark cycle at 20 ± 3 °C and 40–50% humidity and had ad libitum access to food and water. In this study, BALB/c nude mice (BALB/cAn-Foxn1nu/Hnsja) were purchased from Jiangsu Wukong Biotechnology Co., Ltd. MDA-MB-231 tumor-bearing mice (6–8 weeks old, female) were generated by subcutaneous injection of MDA-MB-231 cells (100 μL, 106 cells·mL-1) on the right flank of BALB/c nude mice.
In vivo evaluation of metastatic ability of infected CTCs
In a preliminary experiment, BALB/c nude mice were intratumorally injected with 10 μL of PBS containing 1, 2, or 5 × 106 CFU of S. xylosus when tumor volumes were close to 100 mm3, while mice intratumorally injected with 10 μL of PBS were used as negative controls. Mice were sacrificed 10 days after injection, and their hearts, livers, spleens, lungs, kidneys and tumors were collected and analyzed. All organ tissues were homogenized and serially diluted in sterile PBS before being plated onto solid CBA plates. The colonies on the plates were recorded after 12 h at 37 °C. To further investigate the metastatic ability of infected CTCs, we randomly divided another cohort of BALB/c nude mice into 3 groups: G1, G2, and G3. When the tumor volume reached ~100 mm3, mice in G1 were intratumorally injected with 10 μL of PBS containing 0 CFU of S. xylosus; mice in G2, 2 × 106 CFU of S. xylosus; and mice in G3, 5 × 106 CFU of S. xylosus. The whole blood samples were collected and stored in plastic blood collection tubes with K3EDTA at day 5, 10, 15, and 30. These untreated blood samples were introduced into LesM chips for cell capture. After the blood was introduced, PBS was immediately flowed through the microchannels to flush away the residual blood cells. Then, FITC-vancomycin solution was introduced, and the chips were incubated at room temperature for 15 min before the observation of infected CTCs captured by the traps. All samples were kept at 4 °C and analyzed within 24 h. The body weight and tumor volume of each mouse were monitored every 3 days, with the tumor volume calculated by the following formula: volume = (width2 × length)/2. After a 30-day observation, major tissues were harvested and analyzed by hematoxylin and eosin (H&E) staining and by immunofluorescence staining.
In vivo perturbation of tumor-resident microbes
BALB/c nude mice were intratumorally injected with 10 μL of PBS containing 5 × 106 CFU of S. xylosus when tumor volumes were close to 100 mm3, and some mice received an antibiotic cocktail (ATBx) for 10 days to eliminate tumor microbiota. Specifically, doxycycline (0.2 mg·mL-1), clarithromycin (0.2 mg·mL-1), and azithromycin (0.2 mg·mL-1) were administered through sterile drinking water. Mice treated with normal sterile drinking water were used as negative controls. Solutions and bottles were changed every other day. Ten days after injection, all mice were sacrificed and their tumor tissues were collected, homogenized, serially diluted in sterile PBS, and plated onto solid CBA plates. The colonies on the plates were recorded after 12 h at 37 °C.
In vivo antibiotic treatment assays
On day 0, BALB/c nude mice were intratumorally injected with 10 μL of PBS containing 5 × 106 CFU of S. xylosus when tumor volumes were close to 100 mm3. To analyze the influence of antibiotic treatment on metastasis, we randomly divided BALB/c nude mice into 4 groups: (G1) a pre-resection group, assigned to receive ATBx antibiotics from day 0 to day 10; (G2) a post-resection group, assigned to receive ATBx antibiotics from day 10 to day 30; (G3) a full-course group, assigned ATBx antibiotics from day 0 to day 30; and (G4) a no-medication group, assigned to receive sterile drinking water without antibiotics. Mice in G1 and G2 received normal sterile drinking water on days when no antibiotic treatment was assigned. All tumors were removed on day 10, and whole blood samples were collected on day 30. Of note, all mice were treated with ATBx from day 10 to day 12 (48 h after tumor resection) to avoid post-operative infection. On day 30, whole blood samples were collected and introduced into LesM chips for CTC capture. Following introduction of the FITC-vancomycin solution, captured cells were fixed, blocked, and stained with anti-EpCAM primary antibody and Cy3-conjugated secondary antibody. Major tissues (hearts, livers, spleens, lungs, and kidneys) were harvested and analyzed by H&E staining.
Tissue staining and immunofluorescence
Major organs (hearts, livers, spleens, lungs, kidneys and tumors) were harvested and fixed with 4% paraformaldehyde before being cut into thin tissue sections for H&E staining. For immunofluorescence assays, tumors and lungs were collected, frozen, and sectioned into thin slices. The tissue slides were fixed in cold acetone for 10 min, air-dried, and soaked in PBS (pH 7.4) for 5 min. After being blocked with 20% donkey serum, the slides were incubated with FITC-vancomycin for 30 min at room temperature. After counterstaining with DAPI for another 10 min, the sections were washed twice with PBS. Photomicrography was performed with a fluorescence microscope.
Image acquisition
LesM chips were observed using an EVOS FL Auto microscope (Thermo Fisher Scientific Inc., Waltham, MA, USA) equipped with an automated XY stage under a 10×, 20× or 40× objective. Bright-field (transmitted light), blue (Hoechst), and green (DiO or FITC) channels were used to generate images with complete optical information. Multi-channel images, stitched images, and time-lapse images were acquired with affiliated software. Multi-channel images refer to capturing a set of images in the field of view with the chosen channels at a given objective magnification. Stitched images were captured by defining a large-scale ROI, within which a number of individual snapshots were automatically taken and assembled, allowing for observation of the whole chip for statistical analysis. Time-lapse images were a series of snapshots of the same area automatically captured over various periods of time, allowing for monitoring of time-dependent phenomena.
Image processing
Raw images were analyzed with ImageJ, mainly for the identification of different cells and the quantification of fluorescence intensities within ROIs. Briefly, each trap was defined with 2 ROIs: the hook-shaped part and the deformation channel part. Fluorescence intensity and cell area were calculated for each fluorescence channel for further analysis.
Statistics and reproducibility
All experiments presented in the main manuscript and supplementary information were repeated at least 3 times. All results from representative experiments (such as micrographs, LSCM images, and SEM images) were repeated independently at least 3 times with similar results, and a representative result was shown. All replicate experiments were performed using freshly collected blood, unless clearly stated in the manuscript text. For all experiments performed in BALB/c nude mice, at least 5 mice were randomly selected in each group. Unless clearly stated in the manuscript text, all mice showed consistent results and confirmed the presented data. All statistical analyses were conducted in Origin 9.8.0 and IBM SPSS Statistics 29.0.2.0. And all data were obtained in triplicate and are presented as mean ± s.d. unless otherwise mentioned. Statistical analyses were performed using two-sided, unpaired student’s t-test. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001 were considered statistically significant.
Ethics
Every experiment involving animals were approved and compiled with the guidelines of the Institutional Animal Care and Ethics Committee of Nanjing University (ethical approval number: IACUC-2311004). Every experiment involving clinical samples has been carried out following a protocol approved by the Institutional Review Board of the Nanjing First Hospital (KY20251013-KD-03) and conformed to the guidelines set by the committee. Each participant gave informed written consent. We have obtained no consent to publish information that identifies individuals, and to protect privacy of the patients, we have blurred/hidden relevant information (age/medical centre the study participants attended) to avoid the disclosure of two patients.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the article and its supplementary files. Source data generated in this study are provided in the Supplementary Information/Source Data file. Source data is available for Figs. 2–8 and Supplementary Figs. 1, 4–18, 20–22 in the associated source data file. Any additional requests for information can be directed to, and will be fulfilled by, the corresponding authors. Source data are provided with this paper.
References
Sepich-Poore, G. D. et al. The microbiome and human cancer. Science 371, 1331 (2021).
Riquelme, E. et al. Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell 178, 795–806 (2019).
Xi, J. et al. Reverse intratumor bacteria-induced gemcitabine resistance with carbon nanozymes for enhanced tumor catalytic-chemo therapy. Nano Today 43, 101395 (2022).
Colbert, L. E. et al. Tumor-resident Lactobacillus iners confer chemoradiation resistance through lactate-induced metabolic rewiring. Cancer Cell 41, 1945–1962 (2023).
Nejman, D. et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science 368, 973–980 (2020).
Galeano Nino, J. L. et al. Effect of the intratumoral microbiota on spatial and cellular heterogeneity in cancer. Nature 611, 810–817 (2022).
Fu, A. et al. Tumor-resident intracellular microbiota promotes metastatic colonization in breast cancer. Cell 185, 1356–1372 (2022).
Fu, A., Yao, B., Dong, T. & Cai, S. Emerging roles of intratumor microbiota in cancer metastasis. Trends Cell Biol. 33, 583–593 (2023).
Suhail, Y. et al. Systems biology of cancer metastasis. Cell Syst. 9, 109–127 (2019).
Lawrence, R., Watters, M., Davies, C. R., Pantel, K. & Lu, Y. J. Circulating tumour cells for early detection of clinically relevant cancer. Nat. Rev. Clin. Oncol. 20, 487–500 (2023).
Poudineh, M., Sargent, E. H., Pantel, K. & Kelley, S. O. Profiling circulating tumour cells and other biomarkers of invasive cancers. Nat. Biomed. Eng. 2, 72–84 (2018).
Lin, D. et al. Circulating tumor cells: biology and clinical significance. Signal Transduct. Target Ther. 6, 404 (2021).
Chen, Y. et al. A supramolecular co-delivery strategy for combined breast cancer treatment and metastasis prevention. Chin. Chem., Lett. 31, 1153–1158 (2020).
Mitchell, M. J. & King, M. R. Fluid shear stress sensitizes cancer cells to receptor-mediated apoptosis via trimeric death receptors. N. J. Phys. 15, 015008 (2013).
Han, X. et al. Microfluidic cell deformability assay for rapid and efficient kinase screening with the CRISPR-Cas9 system. Angew. Chem. Int. Ed. 55, 8561–8565 (2016).
Han, Y. et al. Aptazyme-induced cascade amplification integrated with a volumetric bar-chart chip for highly sensitive detection of aflatoxin B1 and adenosine triphosphate. Analyst 147, 2500–2507 (2022).
Wang, Y., Gao, Y. & Song, Y. Microfluidics-based urine biopsy for cancer diagnosis: recent advances and future trends. Chem. Med. Chem. 17, e20220422 (2022).
Kwong, G. A. et al. Synthetic biomarkers: a twenty-first century path to early cancer detection. Nat. Rev. Cancer 21, 655–668 (2021).
Guo, Q. et al. Deformability based sorting of red blood cells improves diagnostic sensitivity for malaria caused by plasmodium falciparum. Lab Chip 16, 645–654 (2016).
Wang, Y. et al. Emerging trends in organ-on-a-chip systems for drug screening. Acta Pharm. Sin. B 13, 2483–2509 (2023).
Gao, Y. et al. An enzyme-loaded metal–organic framework-assisted microfluidic platform enables single-cell metabolite analysis. Angew. Chem. Int. Ed. 62, e202302000 (2023).
Wu, J. et al. Identifying the phenotypes of tumor-derived extracellular vesicles using size-coded affinity microbeads. J. Am. Chem. Soc. 144, 23483–23491 (2022).
Zhang, P. et al. Ultrasensitive detection of circulating exosomes with a 3D-nanopatterned microfluidic chip. Nat. Biomed. Eng. 3, 438–451 (2019).
Liu, J. et al. Magnetic-opticaldual functional Janus particles for the detection of metal ions assisted by machine learning. Smart Mol. 1, e20230006 (2023).
Tse, H. T. K. et al. Quantitative diagnosis of malignant pleural effusions by single-cell mechanophenotyping. Sci. Transl. Med. 5, 212ra163 (2013).
Urbanska, M. et al. A comparison of microfluidic methods for high-throughput cell deformability measurements. Nat. Methods 17, 587–593 (2020).
Nematbakhsh, Y. & Lim, C. T. Cell biomechanics and its applications in human disease diagnosis. Acta Mech. Sin. 31, 268–273 (2015).
Di Carlo, D. A mechanical biomarker of cell state in medicine. SLAS Technol. 17, 32–42 (2012).
Sarioglu, A. F. et al. A microfluidic device for label-free, physical capture of circulating tumor cell clusters. Nat. Methods 12, 685–691 (2015).
Au, S. H. et al. Clusters of circulating tumor cells traverse capillary-sized vessels. Proc. Nat. Acad. Sci. USA 113, 4947–4952 (2016).
Jia, Y. et al. Microfluidic tandem mechanical sorting system for enhanced cancer stem cell isolation and ingredient screening. Adv. Healthc. Mater. 10, e2100985 (2021).
Han, X. et al. Microfluidic cell trap arrays for single hematopoietic stem/progenitor cell behavior analysis. Proteomics 20, e1900223 (2020).
Liu, Z. et al. Integrated microfluidic chip for efficient isolation and deformability analysis of circulating tumor cells. Adv. Biosyst. 2, 1800200 (2018).
Park, E. S. et al. Continuous flow deformability-based separation of circulating tumor cells using microfluidic ratchets. Small 12, 1909–1919 (2016).
Nobes, C. D. & Hall, A. Rho, Rac, and Cdc42 GTPases regulate the assembly of multimolecular focal complexes associated with actin stress fibers, lamellipodia, and filopodia. Cell 81, 53–62 (1995).
van Oosten, M. et al. Real-time in vivo imaging of invasive- and biomaterial-associated bacterial infections using fluorescently labelled vancomycin. Nat. Commun. 4, 2584 (2013).
Zhang, B. et al. Synthesis of vancomycin fluorescent probes that retain antimicrobial activity, identify Gram-positive bacteria, and detect Gram-negative outer membrane damage. Commun. Biol. 6, 409 (2023).
Iida, N. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970 (2013).
Pushalkar, S. et al. The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer Discov. 8, 403–416 (2018).
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (22477056 to Y. Song, 82472379 to B. He, and 82572370 to Y. Gao), the National Key Research and Development Program of China (2019YFA0709200 to Y. Song), the Fundamental Research Funds for the Central Universities (2024300315 to Y. Song), Jiangsu Provincial Medical Key Discipline Cultivation Unit (JSDW202239 to B. He), the Jiangsu Provincial Key Research and Development Program (BE2021373, China to Y. Song), the State Key Laboratory of Analytical Chemistry for Life Science (5431ZZXM2304 to Y. Song). The graphical abstract in Figs. 1, 2a, g, 4a, 5a, 6a, 7a, 8a, Supplementary Fig. 7a and Supplementary Fig. 22a was created using BioRender. Luo, W. (https://BioRender.com/l1e6nxj). Herein, the authors would like to give sincere thanks to Dr. Shang Cai (Westlake University), who had kindly provided intratumor bacteria as a generous gift.
Author information
Authors and Affiliations
Contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript. Y. Song, J. Yang, and T. Hu supervised the project. B.H., J.L., M.W. and Y.Y. provided the clinical blood samples. Y.S. and W.L. conceptualized the research, designed experiments, analyzed the data, and wrote the manuscript with feedback from all the authors. Y. Gao assisted in the preparation of microfluidic devices. S. Feng assisted in the in vivo assays. B. Xue and Y. Cao assisted in the AFM measurements.
Corresponding authors
Ethics declarations
Competing interests
Y.S. and W.L. are inventors on a patent application (number of patent application, 202511249032.X) filed by Nanjing University related to this work. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Esmail Pishbin, Chang Liu, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, W., Gao, Y., Feng, S. et al. A Mechanophenotyping chip for high-throughput detection of metastatic bacteria-infected circulating tumor cells. Nat Commun 17, 1412 (2026). https://doi.org/10.1038/s41467-025-68152-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68152-y