Abstract
Interspecific genetic exchanges caused by natural hybridization or horizontal gene transfer can lead to enhanced phenotypes, which are often of interest for industrial applications and evolutionary research. However, transferring genetic materials between distantly related species, such as intergeneric yeasts, presents technical challenges. In this study, we establish a method to transfer individual chromosomes from Saccharomyces cerevisiae (Sc) into Kluyveromyces marxianus (Km), an emerging model for bioproduction. The Sc chromosome of interest is circularized, genetically modified to carry Km centromeres and replication origins, and transformed into Km via protoplast transformation. Using this method, we generate two synthetic strains, each containing a full set of Km chromosomes and either Sc chromosome I or III. The Sc chromosomes exhibit normal replication, segregation, and active transcription after the transfer. The synthetic strains display enhanced phenotypes in flocculation and salt tolerance, which is found to be caused by transgressive expression of FLO9 and SPS22 on the transferred Sc chromosomes, respectively. Transcriptomic analysis reveal that transgressive expression is prevalent among the transferred Sc genes, suggesting evolution of lineage-specific cis- and trans-regulatory interactions across a long evolutionary timescale. Our strategy has potential applications in optimizing cell factories, constructing synthetic genomes, and advancing evolutionary research.
Similar content being viewed by others
Introduction
The transfer of genetic materials between species is one of the major driving forces for adaptation1,2. Chromosomes, which carry packs of genes, can introduce abundant genetic variation and contribute to evolution of adaptive traits when transferred between species. For example, horizontal transfer of plastid chromosomes between tobacco plants (Nicotiana) aided chloroplast capture3. Horizontal transfer of chromosomes between Fusarium species, a group of filamentous fungi, led to new pathogenic lineages4. Furthermore, hybridization is by nature a process of interspecific chromosome transfers. Interspecific hybridization often leads to enhanced phenotypes, giving rise to industrially important strains such as the lager yeast Saccharomyces pastorianus5 and hybrid crops like triticale6. Therefore, introduction of genetic materials from a distantly related species may hold great potential for phenotypic improvement in agricultural and industrial practices. In addition, understanding the phenotypic and molecular consequences of interspecific genetic exchanges may shed light on genome evolution, such as the molecular bases for heterosis7 and evolution of gene regulation8,9.
In this study, we explored the phenotypic and molecular consequences of chromosomal transfers between Kluyveromyces marxianus (Km) and Saccharomyces cerevisiae (Sc). K. marxianus is a yeast species that belongs to the Saccharomycetaceae family but only distantly related to S. cerevisiae (Fig. 1a). It has been previously argued that it may serve as an emerging model for bioproduction, including heterologous proteins, bioethanol and bulk chemicals10,11. It has a number of traits different from S. cerevisiae, including a high growth rate, thermotolerance, and the ability to assimilate a wide variety of sugars12,13,14. On the other hand, S. cerevisiae is more tolerant to ethanol than K. marxianus11. It is widely used in industrial fermentation and has arguably the best-studied genome among eukaryotes15. Mixing the genetic material between the two species is expected to provide phenotypes for industrial development, as well as insights for understanding their evolution.
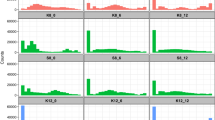
(a) Phylogenetic tree of the Saccharomycetaceae family based on 71 selected species. Blue ellipses show species capable of interspecific hybridization within the genus, while red ellipse marks the event of whole-genome duplication. Sc and Km are highlighted in orange. The tree was constructed with iqtree2105 with concatenated coding sequences of 2,408 orthologous groups23. (b) The evolutionary distance between Km and Sc, compared to species in other taxa that are capable of interspecific hybridization (solid lines) or tolerant to chromosomal transfer (dashed lines). The magenta dashed line shows findings from this study that Km was tolerant to Sc chrI and III. Amino acid substitutions per site were generated by Orthofinder2102. Sp, Saccharomyces paradoxus. Su, S. uvarum. Images were taken from PhyloPic (http://phylopic.org) or drawn by hand. (c, d) Transformation efficiency (c) and stability (d) of plasmids containing different combinations of ARS and CEN from Sc (blue) and Km (red). Stability refers to the percentage of plasmid-containing cells after being grown in non-selective YPD medium for 24 h. For boxplots and whiskers, minima, maxima, and centre represent the minimum, maximum and median, respectively. Box bounds correspond to the 25th (Q1) and 75th (Q3) percentiles. Each experiment includes three biological replicates, except for the Km experiments in (c), where n = 4. (e) Mating assay between Km and Sc. Haploid Km or Sc cells containing a kanR (red cross) or hygR (blue cross) plasmid were mixed and spotted onto plates containing hygromycin (Hyg), G418, or both. Mating efficiency was quantified by counting double-resistant colonies (see Methods), averaged across three biological replicates. Source data are provided as a Source Data file.
Stable allodiploid hybrids between K. marxianus and S. cerevisiae haven’t been made available, although partial integration of S. cerevisiae genome fragments into K. marxianus has been achieved by protoplast fusion16,17. In order to introduce S. cerevisiae genetic materials in a systematic, controllable manner, we developed a genetic-engineering strategy to artificially transfer chromosomes between the two species. Artificial chromosome transfer was first established in 1977, when murine chromosomes were transferred into human cells via microcells18. To date, artificial transfer of functional chromosomes has primarily been achieved between closely related species (dashed lines, Fig. 1b), including transfers of human chromosome 21 into mouse and chicken cells19,20,21, and between bacteria species Mycoplasma mycoides and M. capricolum22. S. cerevisiae (Sc) and K. marxianus (Km) diverged 114 million years ago23, significantly exceeding the evolutionary distance between the species that have been shown capable of natural or artificial chromosome transfers (Fig. 1b). Previous efforts of protoplast fusion showed that the Sc genome was often unstable after the fusion with Km17. One possible explanation is the incompatibility of replicating elements such as centromere (CEN) and autonomously replicating sequence (ARS), as previously shown in hybrid studies24,25.
In this study, we engineer the Sc chromosome with a series of genetic modifications to resolve potential incompatibilities, including circularization of the Sc chromosome and insertion of ARS and CEN from the host species26. The engineered Sc chromosome is transformed into Km, creating a “monochromosomal hybrid”. Through phenotypic and gene expression analysis, we show that the chromosome transfer led to beneficial phenotypes associated with overexpression of genes from the Sc chromosome. The transfer triggers widespread transcriptional responses in the transferred and host genomes, suggesting cross-species regulation. The pervasive transgressive expression found among the transferred genes provides evidence for regulatory evolution across a long evolutionary timescale, as well as potential implications for synthetic biology.
Results
Compatibility of ARS and CEN between Sc and Km
It is critical to ensure proper replication and segregation of the heterologous chromosome upon chromosome transfer. Therefore, we started by investigating the compatibility of ARS and CEN between Sc and Km. In S. cerevisiae, there are 228 functional ARS sites distributed across the genome27, sharing an 11-bp consensus sequence28. The distribution of ARS in the Km genome is less characterized, and only 15 KmARSs have been experimentally validated29,30,31,32. Different from canonical ARS in other yeasts, KmARSs are diverse and do not share a common consensus sequence32. There is one CEN per chromosome in both Sc and Km, each consisting of three conserved centromere DNA elements (CDEI, II and III)33. The Km CDEs are similar to their Sc counterparts, with the exception that the Km CDEII was approximately twice the length of that in Sc34. We selected two ARS-CEN pairs from each species for our experiments, based on their chromosomal locations and previously-demonstrated functionality in plasmids31,32,35,36,37,38. Pairs on the same chromosome were tested together to maximize intraspecific ARS-CEN compatibility: ScARSH4 and ScCEN635,36 (Sc chrII), ScARS137 and ScCEN4 (Sc chrIV), KmARS1 and KmCEN531,38 (Km chrV), and KmARS1832 and KmCEN3 (Km chrIII).
We examined transformation success and stability of plasmids carrying different combinations of KmARS1, KmCEN5, ScARSH4 and ScCEN6. We found that KmARS1 and KmCEN5 were respectively essential for successful transformation (Fig. 1c, left) and plasmid maintenance in Km (Fig. 1d, left). Neither could be replaced by their Sc counterparts, suggesting incompatibility. Plasmids carrying KmCEN5 + KmARS1 produced transformants in Sc (Fig. 1c, right), but with poor stability (Fig. 1d, right), suggesting that KmARS1, but not KmCEN5, can function in Sc. Similar results were obtained using combinations of KmARS18, KmCEN3, ScARS1 and ScCEN4 (Supplementary Fig. 1). The observed incompatibility could explain the rapid loss of Sc chromosomes in previous protoplast fusion between Km and Sc17. Finally, the plasmid containing both ARSs and CENs from Sc and Km, known as “double CEN/ARS plasmid”, could replicate and segregate stably in both species (Fig. 1c, d, right). Therefore, the issue of Sc chromosomal instability in Km may be resolved by engineering KmCEN and KmARS into Sc chromosomes.
Using double CEN/ARS plasmids carrying different antibiotic markers, we examined the mating rate between Km and Sc. We transformed Km and Sc haploids with plasmids carrying a kanR or hygR marker, mixed them in different combinations (see Methods) and selected for zygotes on double antibiotic medium (Fig. 1e). We found that the mating efficiency between Km MATa and MATα cells, as well as between Sc MATa and MATα cells, was around 16%. However, mixing 1.6×107 Km cells with 1.6×106 Sc cells, regardless of their mating types, failed to generate any zygotes, based on both double selection (Fig. 1e) and microscopic observation (Supplementary Fig. 2). These results demonstrated a prezygotic reproductive barrier between the two species, which urges a synthetic method for generating Km-Sc hybrids.
Engineering and transferring Sc chromosomes into Km
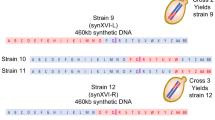
We selected the smallest Sc chromosome, chrI (252 kb, 85 genes based W303 annotation), and the third smallest chromosome, chrIII (341 kb, 156 genes), for proof-of-principle experiments of Sc-Km chromosomal transfer. The experimental pipeline is shown in Fig. 2a. First, we circularized the chromosomes of interest to avoid potential incompatibility of telomeres. Sc telomeres (TELs) are composed of TG1−3 repeat sequences, while Km TELs consist of a long repeat motif (25 nt)39. We removed the TELs from Sc chrI and chrIII, respectively, and joined the ends with KmURA3 with CRISPR/Cas940. To avoid homologous recombination at the telomeres, we also removed the telomere-associated long repetitive sequence on chrI during the circularization40. Next, to ensure stable maintenance of Sc chromosomes in Km, we inserted KmCEN5 /ARS1 adjacent to the native CENs on chrI and chrIII. Given that chrI and chrIII contain 5 and 12 ARSs, respectively27, we placed another copy of KmARS1 close to a native ScARS, which was about ∼100 kb away from the KmCEN5 /ARS1 in the circular chromosomes (see Methods for details). The engineered circular chrI and chrIII were named R1 and R3, respectively. Sc cells with R1 (“Sc-R1”) or R3 (“Sc-R3”) showed no fitness defect under normal culture conditions as well as in the presence of microtubule-depolymerizing and DNA-damaging agents (Supplementary Fig. 3), indicating that the introduction of KmARS1 and KmCEN5 did not affect chromosome replication in Sc. The circularization was confirmed by pulsed-field gel electrophoresis (PFGE), which showed an absence of linear chrI or chrIII in the gel (Fig. 2b). Following a protocol developed by Noskov et al.41, we extracted and column-purified R1 and R3, with the undesired linear chromosomes removed by exonuclease. Finally, the purified R1 and R3 were separately transformed into Km using protoplast transformation42, yielding four R1 transformants (Kluyveromyces-Saccharomyces-R1, “KS-R1” hereafter) and one R3 transformant (“KS-R3”). The low number of transformants indicates a need for further optimization of the transformation protocol, especially for transferring larger chromosomes.
(a) Experimental pipeline. (b) PFGE of chromosomal extracts from Sc-R1, Sc-R3 and the parental Sc strain. (c, d) PCR of markers in the circular chromosomes in KS-R1 (c, represented by one transformant) and KS-R3 (d). Purple box labels the missing marker “7” in KS-R3. See Supplementary Data 1 for marker positions. (e) PFGE of R1 and R3 (arrows) in KS, linearized with NotI and AscI respectively. Ctrl: non-digested KS-R1 chromosomal extracts. λ: Lambda PFG Ladder. (f) Illustration of the 29 kb deletion in KS-R3. Coordinates are based on a manually curated R3 sequence (Supplementary Data 1). (g) Growth curves of KS-R1, KS-R3, and Km-V (Km with an empty vector) in YPD. The values represent mean ± SD (three biological replicates). (h) Spot assay of KS-R1 and KS-R3 in the presence of benomyl (BML), thiabendazole (TBZ), hydroxyurea (HU), methyl methane sulfonate (MMS), camptothecin (CPT) or cycloheximide (CHX). The plates were incubated at 30 degrees for 1 day (1 d) or 3 days (3 d) as indicated. For (b-e), each experiment was repeated three times independently with similar results. Source data are provided as a Source Data file.
We next examined the integrity of the transferred chromosomes. All four KS-R1 transformants retained six Sc-specific markers in R1 (Fig. 2c, see Supplementary Data 1 for marker positions), indicating successful transfer. The transfer was further confirmed by restriction digest (Fig. 2e) and whole genome sequencing, which found no mutations in the transferred R1. In the PCR analysis of KS-R3, marker 7 was absent (Fig. 2d). PFGE showed that the size of linearized R3 was smaller than the expected 340 kb (Fig. 2e). Genome sequencing revealed a 29 kb deletion between two direct repeats of putative Ty elements in R3 (Fig. 2f& Supplementary Fig. 4). The deletion was potentially due to homologous recombination between Ty elements, the rate of which has been shown to significantly increase in circular plasmids during transformation43. The deleted region contained 13 genes (MAK32, PET18, MAK31, HTL1, HSP30, YCR022C, YCR023C, SLM5, YCR024C-B, PMP1, YCR025C, NPP1, RHB1). We introduced the deleted region, in six individual segments via plasmids, into Km cells (Supplementary Fig. 4). None of the segments affected growth (Supplementary Fig. 4), so it was unlikely that the deletion was an adaptive response to overcome incompatibility caused by single genes in this region.
We next evaluated the Km genome integrity after transformation. The PFGE patterns of Km chromosomes in KS-R1 and KS-R3 were consistent with the parent (Supplementary Fig. 5). Genome sequencing revealed 33 mutations in 11 ORFs, as well as 82 SNPs and indels in intergenic sequences in Km chromosomes in KS-R1 (Supplementary Data 2). KS-R3 contained 4 mutations in 4 ORFs, and 47 SNPs and indels in intergenic sequences (Supplementary Data 2). These mutations might have been induced during the protoplast transformation. Overall, the transfer of R1 or R3 into Km did not result in fusions, translocations, or other large-scale rearrangements in the Km chromosomes.
Subsequently, we examined the KS strains for any fitness defects. In rich medium (YPD), the growth curves of KS-R1 and KS-R3 were indistinguishable from Km-V, the parental Km strain with an empty vector (Fig. 2g). Under non-selective conditions, R1 and R3 were highly stable in the KS strains. The loss rate per generation for R1 and R3 in YPD was 0.11% and 0.84%, respectively (Supplementary Data 3), a level comparable to that of circular YACs in Km (0.33-0.64%)44. Furthermore, KS-R1 and KS-R3 exhibited robust growth in the presence of the microtubule-depolymerizing agents benomyl (BML) and thiabendazole (TBZ), DNA replication inhibitor hydroxyurea (HU), DNA-damaging agents methyl methane sulfonate (MMS) and camptothecin (CPT)45, or protein synthesis inhibitor cycloheximide (CHX) (Fig. 2h). Taken together, the introduction of R1 and R3 did not affect growth, chromosome segregation, DNA replication, or protein synthesis in rich medium. The Km cells were tolerant to the transplantation of Sc chromosomes, reflecting cellular plasticity.
Enhanced phenotypes due to interactions between Km-encoded trans-factors and Sc cis-regulatory sequences
We next investigated if the transferred chromosomes conferred different phenotypes. We treated the KS strains and their parents with 28 environmental conditions, chosen to represent common stressors for yeast growth (Fig. 3a & Supplementary Fig. 6-7). For KS-R1, the phenotypes of the four independent transformants were generally consistent (Supplementary Fig. 7), so we primarily analyzed one representative transformant (Fig. 3a & Supplementary Fig. 7). We primarily compared the KS strains to Sc and Km strains with an empty double CEN/ARS plasmid (referred to as Sc-V and Km-V respectively), which carries the same antibiotic and KmURA3 markers as the circularized chromosomes, to account for phenotypic effects associated with the vector (see Supplementary Fig. 8 for details). Sc-R1 and Sc-R3 were also examined to account for effects of chromosomal circularization and other genetic modifications, but their phenotypes were for the most times consistent with Sc-V (Fig. 3a & Supplementary Fig. 6). For each strain, we semi-quantified the growth on solid media under different conditions, normalized the data to YPD growth at 30 °C, and then adjusted the data in reference to Km-V to reflect inter-strain differences (Fig. 3a & Supplementary Fig. 6b). Sc-V and Km-V showed significant growth differences under 17 out of the 28 conditions (2% ethanol, Ser, 1/4 N, 1/2AA1/2 N, 20, 37 and 42°C, 60 mM AcOH, and all three H2O2, TM and NaCl treatments; FDR-adjusted p < 0.05 based on linear model, Supplementary Data 4), reflecting phenotypic divergence between species. KS-R1 and KS-R3 exhibited comparable growth to Km-V under 27 and 17 conditions, respectively (FDR-adjusted p > 0.05, linear model; Fig. 3a & Supplementary Data 4), suggesting that the small number of genes in R1 and R3 did not trigger a widespread metabolic reprogramming in Km. Under a few conditions, R1 and R3 caused increased sensitivity in KS compared to Km, including the sensitivity to 42 °C for KS-R1, and to tunicamycin (TM), 1/4 N, 1/2AA1/2 N, 20°C, 0.016% and 0.024% H2O2 for KS-R3 (FDR-adjusted p < 0.05, linear model, Fig. 3a-b & Supplementary Data 4). Under these conditions, the KS phenotypes were intermediate to those of Km and Sc, consistent with dominant effects exerted by the introduced Sc chromosomes. The defective phenotypes were not shared by the two strains, indicating that the defects were caused by specific genes in R1 or R3, rather than a general effect caused by chromosomal transfers. Additionally, we ruled out that the mutations in the Km genome caused the phenotypes, because the strains no longer exhibited the phenotypes after losing the Sc chromosomes (Supplementary Fig. 9).
(a) Growth phenotypes of the synthetic and parental species, relative to Km-Vector (Km-V). Growth under each condition was normalized to YPD growth at 30 °C, log-transformed, averaged across biological replicates and subtracted by the value of Km-V (see Methods for details), such that 0 means equal growth to Km-V. n = 3 for all conditions except for three H2O2 treatments and 60 mM AcOH, where n = 2 (see Methods). Original data are provided in Supplementary Fig. 6 and Supplementary Data 4. Gray, no growth. Glu, glucose; 1/4 N, one-quarter ammonium sulfate; 1/2 AA, half amino acids; 1/2 N, half ammonium sulfate; TM, tunicamycin; 5-FU, 5-fluorouracil. (b) Growth defects under TM or 42 °C associated with transferred Sc chromosomes (1:5 serial dilutions; 1-day growth unless indicated otherwise). (c) Improved NaCl tolerance associated with ScSPS22 promoter. ScSPS22, KmSPS22 and PKm-ScSPS22 alleles were expressed with a centromeric plasmid (indicated by the subscript CEN) or a 2 µ plasmid (ScSPS222µ) in either Km (upper panel) or Sc-R3 (lower panel). (d) Relative mRNA levels of ScSPS22 and KmSPS22 measured by qPCR. mRNA levels in KS and Km were normalized to the average of three housekeeping genes, KmSWC4, KmTRK1, and KmMPE1. mRNA levels of ScSPS22 in Sc were normalized to the average of ScSWC4, ScTRK1, and ScMPE1. (e, f) Enhanced flocculation in KS-R1 associated with ScFLO9. ScFLO9 was expressed with either a centromeric plasmid (ScFLO9 CEN) or a 2µ plasmid (ScFLO92µ) in Km or Sc-R1. Overnight cultures were vortexed vigorously and left still, before being imaged (e) and examined for the OD600 of the supernatant (f) at designated timepoints. (g) Relative expression levels of ScFLO9 and KmFLO5 measured by qPCR, normalized as described above. The values in (d, f, g) represent the mean ± SD (three biological replicates). Statistical tests in (d, g) were performed with ANOVA followed by Tukey’s test to account for multiple comparisons. **, p < 0.01; ***, p < 0.001. Source data are provided in Source Data file and Supplementary Data 4.
Importantly, the chromosomal transfer resulted in enhanced phenotypes that were superior to both parents. KS-R3 exhibited enhanced tolerance to 0.8 M and 1 M sodium chloride (NaCl) compared to both parents (FDR-adjusted p < 0.05, linear model in reference to Km-V; Fig. 3a & Supplementary Fig. 6b, Supplementary Data 4). High salt is a common stress in industrial processes. Therefore, increased salt tolerance is a beneficial phenotype for industrial production46. We next investigated the molecular mechanisms by which R3 increased salt tolerance in KS. There are three genes in R3 that have been associated with salt tolerance based on previous large-scale studies: SPS2247, STE5048 and HMLALPHA247. We expressed the three candidate Sc genes in Km with a centromeric plasmid, including their coding sequences (CDSs) and 1000 bp-upstream sequences (promoters). We found that only ScSPS22 enhanced salt tolerance to the same level as KS-R3 (Supplementary Fig. 10, Fig. 3c, upper panel), indicating that it was the causal gene. ScSPS22 is involved in β-glucan synthesis and related to cell wall function49. In S. cerevisiae, overexpression of ScSPS22 was previously found to cause reduced salt tolerance47, which was recapitulated in our study (Fig. 3c, lower panel). It remains to be understood why overexpression of ScSPS22 had an opposite effect in Km than Sc, possibility related to the difference in their cell-wall composition50.
In order to understand the contribution of cis-regulatory and coding sequences (CDS) to the phenotypic effect of ScSPS22, we constructed a chimeric allele consisting of the promoter of KmSPS22 (FIM1_814) and the CDS of ScSPS22, namely Pkm-ScSPS22. There was no substantial difference between Pkm-ScSPS22 and full-length KmSPS22 when expressed with a centromeric plasmid (Fig. 3c, upper panel), suggesting a lack of functional divergence in the CDS. On the contrary, Pkm-ScSPS22 conferred a much lower level of salt tolerance than ScSPS22 driven by its endogenous promoter, indicating that the Sc promoter was causal for the increase in salt tolerance.
The different phenotypic effects of Sc and Km promoters suggest potential divergence in gene expression. Therefore, we analyzed the transcription level of ScSPS22 and KmSPS22 in KS-R3 and the parental strains by qPCR (Fig. 3d). The relative mRNA level of ScSPS22 in KS-R3 was respectively 53 and 80 times higher than those in Sc-R3 with and without NaCl (Fig. 3d, right panel), suggesting that (1) a significant difference in trans-acting regulators for ScSPS22 between Sc and KS-R3; and (2) the up-regulation of ScSPS22 in KS-R3 is constitutive. The relative mRNA level of KmSPS22 did not significantly differ between KS-R3 and Km-V (adjusted p-value = 0.056, Tukey’s test; Fig. 3d, left panel), indicating little difference in the trans-acting factors for KmSPS22 between Km and KS-R3.
When transformed into Km on a centromeric plasmid, the ScSPS22 allele was expressed at the same level as ScSPS22 in KS-R3 (adjusted p-value = 0.945 and 1.000 for YPD and NaCl conditions respectively, Tukey’s test, Fig. 3d), suggesting that the promoter sequence of ScSPS22 was causal for its up-regulation in KS-R3. The chimeric allele Pkm-ScSPS22 drove lower expression than ScSPS22 in Km, consistent with cis-regulatory divergence. In Sc-R3, expressing ScSPS22 with centromeric or high-copy 2μ plasmids increased its relative mRNA level, but failed to match the level of ScSPS22 in KS-R3, suggesting that the upregulation of ScSPS22 in KS-R3 resulted from induction or de-repression by Km-specific trans-acting factors, rather than a dosage effect. Taken together, the phenotypic and expression analysis showed that the increased salt resistance of KS-R3 was caused by specific interactions between Km-encoded trans-acting regulators and the Sc promoter of SPS22.
Although KS-R1 did not show any superior phenotype among the conditions assayed in Fig. 3a, we additionally found that it exhibited a flocculation phenotype surpassing that of Km-V and Sc-R1 (Fig. 3e, f). The flocculation phenotype disappeared after the loss of R1, indicating that R1 was responsible for the enhanced flocculation (Supplementary Fig. 11). Flocculation facilitates cell sedimentation, reducing separation costs in high-density fermentation51. The FLO family genes control flocculation52, among which ScFLO9 is present on R1 and has been shown to cause flocculation in Km53. Introducing ScFLO9 with its 1000 bp upstream sequence into Km via a centromeric plasmid enhanced flocculation to the same level as KS-R1 (Fig. 3e&f). This result suggests that ScFLO9 was responsible for the increased flocculation phenotype in KS-R1. Overexpressing ScFLO9 with a centromeric or 2μ plasmid in Sc slightly enhanced flocculation (Fig. 3e&f), consistent with its known role in Sc but suggesting its effect is background-dependent.
We similarly analyzed the expression level of FLO9 to understand the molecular mechanisms of the enhanced phenotype. qPCR showed that the relative mRNA level of ScFLO9 in KS-R1 was 64.5 times higher than in Sc-R1 (Fig. 3g), again suggesting a significant change in the trans-regulatory environment. The high expression in KS-R1 was recapitulated by ScFLO9 expression from a centromeric plasmid in Km (light pink bars in Fig. 3g, adjusted p-value = 0.241, Tukey’s test). The expression level of KmFLO5 (FIM1_6), the ScFLO9 ortholog, was not increased in KS-R1, consistent with a lack of change in trans environment for Km genes (Fig. 3g, left panel). Overexpressing ScFLO9 in Sc-R1 with centromeric and 2μ plasmids led to an increase in the mRNA level of ScFLO9 (purple bars in Fig. 3g), with the higher level of ScFLO92μ correlating with faster flocculation (Fig. 3f). Taken together, the enhanced flocculation of KS-R1 was associated with ScFLO9 being exposed to an alien trans-regulatory environment encoded by the Km genome. In both cases of SPS22 and FLO9, the genetic combinations between Km-encoded trans-factors and Sc-encoded cis-factors could never be found in nature due to reproductive isolation, demonstrating the power of cross-species synthetic biology.
Pervasive transcriptional responses triggered by chromosomal transfer
The experiments with ScSPS22 and ScFLO9 revealed significant changes in Sc gene expression after being transferred to Km. To determine if this is a general pattern, we analyzed transcriptomes of KS and the parental strains in YPD and under two stress conditions, 1 μg/mL tunicamycin (TM) and 1 M NaCl. The genes on R1 and on R3 were actively transcribed in KS under the three conditions (read counts >5) (Supplementary Data 11). However, their expression in KS was poorly correlated with that in Sc, with an average correlation coefficient (R²) of 0.38 for R1 genes and 0.32 for R3 genes (Fig. 4a). This correlation was substantially lower than the correlation between the expression of human genes on Hsa21 transplanted into mouse cells and their counterparts in human cells (R² = 0.90)21, which aligns with the remote evolutionary distance between Km and Sc (Fig. 1b). Compared to Sc, on average, 22.7% and 18.3% of the genes in R1 and R3, respectively, were up-regulated in KS, while 27.3% and 19.3% of the genes in R1 and R3 were down-regulated in KS, respectively (FDR-adjusted p < 0.05 by Wald test and absolute log2FoldChange > 1; Fig. 4b, Supplementary Data 5). The median log2-fold change for up- and down-regulated genes was 1.81 and −1.57, respectively. Consistent with the case studies of SPS22 and FLO9, the high proportion and fold change of differentially expressed Sc genes reflect dramatic differences in the trans-regulatory environment between Sc and the KS strains.
a Comparison of Sc gene expression between Sc and KS strains. Left, diagram showing changes in trans-environment. Right, correlation of Sc gene expression. The expression level was represented by log2 (average normalized read counts). Each point represents the average of three replicates. Adjusted R2 were derived from linear regression. N = 81 genes for R1, and 137 genes for R3 (see Methods for data filters). b Proportion of significantly up- and down-regulated R1/R3 genes in KS. DEG, differentially expressed genes. n.s., not significant. c De-repression of the HML locus in KS-R1, shown by raw read counts (gray bars) covering the GEX1-CHA1 region in the W303 genome, visualized by Integrative Genomics Viewer. Data were from one representative biological replicate in YPD. Gray bars are in log scale. d The relative mRNA levels of HMLα1 and HMLα2 in KS-R3 and Sc-R3 were determined by qPCR, normalized as described in Fig. 3d. The values represent mean ± SD (n = three biological replicates). e Comparison of Km gene expression between Km and KS strains. Left, diagram showing the effects of Sc trans-acting factors. Right, number of DEGs between KS strains and Km-V. Y, YPD; T, TM; N, NaCl. In this figure, all DEGs are defined as an FDR-adjusted p-value < 0.05 by the Wald test and abs(log2FoldChange) > 1. Source data are provided in the Source Data file and Supplementary Data 5.
The genes showing differential expression between Sc and KS were, in general, uniformly distributed across the transferred chromosomes (Supplementary Fig. 12), but we found a region around HML that showed de-repression across multiple genes in KS-R3, providing an interesting example for chromatin-level regulatory divergence. The HML locus, along with its flanking genes GEX1 and VBA3, is repressed in Sc by SIR complex54. Consistent with this previous knowledge, we found little expression of these genes in Sc (Fig. 4c, upper panel). Interestingly, GEX1, VBA3, and HML genes were highly expressed in KS-R3 (Fig. 4c, lower panel), indicating derepression. qPCR confirmed that the relative mRNA levels of α1 and α2 were upregulated 42-fold and 39-fold, respectively, in KS-R3 compared with Sc-R3 (Fig. 4d). The differential expression might be explained by different composition of silencers between Kluyveromyces and Saccharomyces55. The silencer of HML in KS-R3 might not be able to recruit essential proteins such as Rap1 and ORC to the locus56, contributing to the de-repression of α1 and α2, and the flanking genes GEX1 and VBA3.
While the expression of Sc genes underwent significant changes in KS, R1 and R3 also affected the expression of Km genes (Fig. 4e). Out of 4,975 Km genes, 931 (18.7%) and 803 (16.1%) genes in KS-R1 and KS-R3, respectively, exhibited significant expression differences compared to Km-Vector in YPD (FDR-adjusted p < 0.05 by Wald test and absolute log2FoldChange > 1; Fig. 4e, Supplementary Data 5). R1 encodes a transcription factor, Oaf1, that regulates at least 97 Sc target genes57. Among the 73 Km orthologs of the known Sc target genes (see Methods for ortholog assignment), 27 (37.0%) showed differential expression in KS-R1. R3 encodes four transcription factors regulating at least 305 Sc target genes57, with 44/160 (27.2%) of their Km orthologs differentially expressed in KS-R3 (Supplementary Data 5). This suggests that transcription factors encoded by R1 and R3 might directly alter gene expression in Km, but a large fraction of the observed transcriptional changes might be explained by indirect or uncharacterized regulation. Under TM and NaCl stresses, the number of differentially expressed genes was substantially lower than in YPD (Fig. 4e, Supplementary Data 5). These results indicate that the heterologous chromosomes induced pervasive changes in expression of the endogenous genome, potentially via Sc-Km regulatory interactions. Interestingly, regulation from the Km genome became dominant under stress conditions, removing many of the effects associated with foreign chromosomes found in YPD.
Co-evolution of cis- and trans-regulatory factors
Transferring genetic materials between remotely related species provides a unique opportunity to examine the evolution of gene expression. Evolutionary divergence of gene expression can be caused by the evolution of cis- or trans-regulatory factors, or both. Cis-regulatory elements such as promoters or enhancers are physically linked to a target gene (e.g., “promA” for “Gene A” in Fig. 5a). The regulatory activity of cis-elements may evolve through changes in transcription factor binding sites or other elements that interact with the transcription machinery. Trans-acting factors often refer to regulatory proteins such as transcription factors (TFs) encoded elsewhere in the genome (e.g., transcription factor “B” in Fig. 5a). They may include any direct or indirect regulators, which may evolve through coding or non-coding changes58, even including turnover of transcription factors (e.g., “B” may be replaced by “C” over time, Fig. 5a). It has been proposed that cis-acting changes and cis-trans co-evolution plays an increasingly important role as species diverge59, but this idea has been rarely tested in species as diverged as S. cerevisiae and K. marxianus. The KS strains allowed us to interrogate this question (Fig. 5b)60,61. If the transferred genes showed the same expression level in KS (“KSsc”) as that in S. cerevisiae (“Psc”), it means that the expression divergence is entirely encoded by the Sc cis-regulatory elements (the change in trans-environment from Sc to KS had no effect; “donor-like” in Fig. 5c). If the Sc allele was expressed at the same level as its orthologous Km allele in KS (“KSkm”), the expression divergence is entirely determined by trans-differences (the cis-regulatory elements of Sc and Km had equivalent activities when exposed to the same trans-environment; “recipient-like” in Fig. 5c). An intermediate or transgressive expression level of Sc alleles in KS indicates contributions from both factors. In particular, transgressive expression, defined by a KSsc level outside the range of Psc and KSkm, may suggest cis-trans interactions specific to the synthetic background (Fig. 5c).
a Divergence of gene expression can be caused by accumulation of cis (unfilled vertical bars) and/or trans (filled vertical bars) changes during species’ divergence. In the diagram, a focal gene “A” is under the regulation of a transcription factor “B” which binds to A’s promoter (“promA”). Both A and B undergo sequence divergence in the two species, represented by different colors. In some cases, B can be replaced by other TFs such as “C” during evolution, as exemplified in Km. Transcripts are represented by wavy lines. b, c Comparing allelic expression of the transferred genes to donor (Sc, blue) and recipient (Km allele in KS, red) reveals contributing factors to expression divergence. In KS, Km-B/C is present but Sc-B may or may not be present, represented by dashed circles. Anc, ancestor. d 102 genes showing divergent expression were classified into five categories according to (c). Log2 fold changes were extracted from DESeq models, adjusted with the ashr method. Genes showing the most prominent transgressive expression were labeled. Data were from the YPD treatment. e PCA of gene expression of 162 transferred genes across treatments, strains and alleles. (f) Transgressive expression suggests that the cis and trans factors often evolved in opposite directions, represented by “+” and “-”. The order and number of changes was arbitrary. Source data are provided in Source Data file and Supplementary Data 7.
In order to characterize cis- and trans- regulatory divergence between Sc and Km, we identified Km orthologs for 62/85 genes on R1 and 100/156 genes on R3, with an average amino-acid sequence similarity of 48.0% (BLOSUM62-based similarity, see Methods and Supplementary Data 6). 106 out of the 162 genes showed significant expression divergence between Sc and Km (FDR-adjusted p < 0.05, Wald test). What factors determined their divergence? Focusing on the Sc allele of these genes, we classified them into Sc-like, intermediate, Km-like, transgressive and others based on the criteria in Fig. 5c, using the transcriptomic data in YPD (Fig. 5d, Supplementary Data 7). Interestingly, we found that most transferred genes showed transgressive expression (40/106 genes, 37.7%, Fig. 5d), more than those determined by cis-alone (Sc-like) or trans-alone (Km-like). SPS22 and FLO9 analyzed in our phenotypic studies (Fig. 3) were among the most prominent examples in this class (Fig. 5d), suggesting that transgressive expression might be an important source of phenotypic improvement in these engineered strains. Transgressive expression was also the most abundant class under NaCl and TM stresses (54/132 and 36/122 genes respectively, Supplementary Fig. 13, Supplementary Data 7). Principal Component Analysis (PCA) of the 162 transferred genes across treatments and strains showed that KSsc levels were distinct from Pkm, KSkm and Psc, consistent with pervasive transgressive expression of Sc alleles (Fig. 5e). The second most abundant class in YPD was Sc-like expression (25/106, Fig. 5d), supporting an important role of cis-regulatory evolution. However, there were more Km-like genes than Sc-like genes under NaCl and TM stresses (Supplementary Fig. 13), consistent with a predominant role of trans-evolution in environmental responses62.
In the analysis above, we defined “recipient-like” by comparing KSsc to KSkm, reasoning that this was a conservative estimation of cis-effects, when the two alleles were exposed to the same trans-environment in KS. This was different from previous analyses of introgression lines60 or enhancer swaps61, where the transferred allele was compared to the recipient parent (Pkm in this case). The methodological difference was largely negligible because the trans-environment in KS was highly similar to Km, as shown in PCA (Pkm and KSkm clustered together, Fig. 5e). However, to account for the minority of cases where the transferred Sc genes influenced Km allelic expression (Fig. 4e), we performed the same analysis using KSsc/Pkm instead of KSsc/KSkm (Supplementary Fig. 14). This second method was less conservative with cis effects, but it provided a more authentic estimate for transgressive expression: in the cases where KSkm showed transgressive expression in KS, transgressive expression of KSSc might be missed or overestimated by the first method. The results still supported the prevalence of transgressive expression of the transferred genes (37/106 genes in YPD), although there were more genes classified into Sc-like, as expected (Supplementary Fig. 14).
The prevalence of transgressive expression in KS suggests that the cis- and trans-regulatory factors often evolved in opposite directions within a lineage63. For example, when an Sc allele showed transgressively high expression in KS (KSsc > KSkm and KSsc > PSc, Fig. 5c and f), it might indicate that (1) the Sc promoter accumulated more changes to increase gene expression compared to the Km promoter, via either stronger activator activity or weaker repressor activity (“+” effects for cis-changes in the Sc lineage and correspondingly “-” effects in the Km lineage, Fig. 5f), such that KSsc > KSkm; and (2) a trans-acting factor that decreased gene expression must have co-evolved with this promoter in S. cerevisiae (“-” effects for trans-changes in the Sc lineage and correspondingly “+” effects in the Km lineage, Fig. 5f), such that KSsc > Psc.
Compensatory evolution may also be identified by examination of genes showing conserved level of expression between the two species (Supplementary Fig. 15)64. For two genomes as diverged as Sc and Km, it is as important to understand the conservation of gene expression as to understand their divergence. Across 4,307 genes with identifiable orthologs between Km and Sc-R1 (see Methods for ortholog assignment), 29.7% showed no difference in expression levels between the parental strains in YPD (FDR-adjusted P > 0.05, Wald test; Supplementary Fig. 15b, c, Supplementary Data 7). This ratio was 21.9% and 12.8% under NaCl and TM conditions respectively. The expression level of the 4,307 orthologs was correlated genome-wide between species (Spearman’s rank correlation rho = 0.647 for YPD data, P < 2.2e-16, Supplementary Fig. 15b), suggesting a certain level of regulatory conservation despite the deep divergence. The genes showing conserved expression in YPD were significantly enriched for rRNA processing (GO: 0006364, corrected P = 2.07e-20) and ribosome biogenesis (GO: 0042254, corrected P = 2.61e-19), consistent with a housekeeping role. There was a similar proportion of genes showing conserved expression among the transferred genes (56 out of 162, 34.6% in YPD; 18.5% in NaCl and 24.7% in TM, Supplementary Fig. 15d), providing us with a small sample to peek into the driving force of regulatory conservation – did the cis and trans regulatory factors both maintained a conserved level of activity, or they evolved in a compensatory manner under stabilizing selection? In the latter case, we would expect to find cis-regulatory divergence in the KS strains. Indeed, we found that 42 out of the 56 (75%) conserved genes showed significant cis-divergence in the KS strains in YPD (23/30 in NaCl and 29/40 in TM; FDR-adjusted P < 0.05, Supplementary Fig. 15d–f), consistent with compensatory cis-trans evolution. The top genes showing this trend in YPD were CDC19, TRX3, and HIS4 (Supplementary Fig. 15e). In all three cases, the Sc allele was down-regulated in KS strains, suggesting evolution of Sc-specific activators. We note that the amount of cis-divergence found in KS may be specific to the given trans environment, e.g., the cis-effects in KS may or may not manifest themselves in an F1 hybrid. However, such dependency itself indicates co-evolution of cis-trans interactions (see Discussion).
Taken together, our results support that lineage-specific co-evolution of cis- and trans-regulatory factors was predominantly involved in the evolution of gene expression between S. cerevisiae and K. marxianus. Transgressive expression was the most prevalent pattern among the transferred genes in KS. Cis-only and trans-only evolution also occurred, with the latter predominantly contributing to environmental responses62. At the same time, around one-third of the yeast genome showed conserved gene expression despite the long evolutionary distance. Compensatory cis- and trans-evolution might underlie such conservation, consistent with stabilizing selection.
Discussion
Interspecific hybridization often leads to different phenotypes that are of interest for industrial purposes and evolutionary studies65. For diverged species that cannot mate naturally, artificial chromosomal transfer offers an opportunity to explore the industrial potential of combining divergent genetic materials, as well as to understand their molecular and phenotypic consequences. In this study, we successfully introduced chromosomes of S. cerevisiae to K. marxianus, two species both with great industrial potential but genetically as diverged as human and lancelet (Fig. 1b). We show that the transferred Sc chromosomes were stably maintained in Km. The synthetic strains exhibited enhanced phenotypes, demonstrating the technology’s potential in future phenotypic screens. The improved salt resistance and flocculation phenotypes were associated with Sc promoters being activated by Km-encoded trans-acting factors to a greater level than the native regulation in Km or Sc. Finally, we examined the molecular consequences of chromosomal transfer using transcriptomic analysis, revealing broad transcriptional responses following chromosomal transfer, as well as prevalent compensatory evolution of cis- and trans-regulatory changes during the divergence of the two species.
One of the technological advances in our study is the method to enable stable maintenance of Sc chromosomes in Km. There are several technical factors to consider. First, we removed Sc TELs and circularized the chromosomes of interest. The circularization made it possible to easily remove unwanted linear chromosomes by exonuclease41, which may increase transformation efficiency. In the case of CEN engineering, it was important to ensure that the heterologous CEN was not partially functional, which has been shown to disrupt chromosome segregation and lead to dicentric breakage, thereby causing genome rearrangements and instability66. We found that Sc and Km CENs were non-functional in the other species. Therefore, we may use double CENs for maintaining the heterologous chromosomes. Next, we considered the density of ARSs. The requirement of ARS density for YACs varied in previous reports, from one ARS per 30 kb67, 51 kb68, to 1 Mb69. In our study, KmARS1 was positioned at a density of one ARS per 110 kb in R1 and 150 kb in R3, which proved to be sufficient for replication of Sc chromosomes in KS. Finally, we note that one potential technical pitfall was the difficulty in acquiring independent transformants, as exemplified by R3. Only one transformant of KS-R3 was obtained, which contained a deletion potentially mediated by Ty elements during transformation (Fig. 2). Future efforts are needed to optimize the transformation efficiency to ensure scalability.
There are several advantages in interspecific chromosomal transfers for synthetic biology purposes. First, it enables the combination of genetic materials from remotely related species, partly circumventing the genomic incompatibility that prevented interspecific hybridization. The synthetic “monochromosomal hybrids” were viable, providing materials for phenotypic screens or downstream experiments such as directed evolution70. In particular, directed evolution may help resolve any residual incompatibility by selecting for mutations that improve the desired traits and/or compensate for the incompatibility.
Second, our strategy of chromosomal transfers may provide a complementary approach to whole-genome transformation (WGT)71,72 or genomic libraries, in terms of (1) the ability to screen for polygenic effects and (2) a medium-level of experimental throughput. Both WGT and genomic libraries may be used for genome-wide interspecific genetic screens, where foreign genes were individually introduced to the host, avoiding incompatibility. Compared to WGT and genomic libraries, the strategy of chromosomal transfers is limited in its ability to screen the entire genome, but has the strength of screening for polygenic effects. Although our experiments with chr1 and chr3 mainly found single-gene effects, future transfers with larger chromosomes might potentially find beneficial phenotypes necessitated by interactions among multiple genes in the transferred chromosome. Additionally, as demonstrated in our study, cross-species regulation may be an important source of phenotype diversity (Figs. 3 & 5). Our approach enables screening for such interactions by introducing full-length cis-regulatory elements, which are often missed from WGT or genomic libraries. Furthermore, our strategy allows for possibilities of transferring multiple chromosomes into Km. This can be done either by marker recycling, i.e., popping out URA3 in the (evolved) synthetic strains and transforming another chromosome subsequently, or by mating with another (evolved) synthetic strain containing a different chromosome. Therefore, chromosomal transfer is particularly advantageous for screening for polygenic effects, including interactions within and among foreign chromosomes, as well as between host and foreign genes.
Our strategy provides a medium-throughput screening capability, between that of WGT and genomic libraries. In WGT studies, genomic DNA of a foreign species was transformed into the host species without a stable vector71,72. The foreign genes were expected to randomly integrate into the host genome, and only the transformants that conferred desired phenotypes were selected. The success rate of such strategy depends on the efficiency of homologous recombination and screening conditions. Also, random integration of foreign genes can disrupt the host genome, lowering the success rate even more. From this perspective, the experimental throughput of such WGT approach can be low. Compared to WGT, chromosomal transfers allow screening for hundreds of genes at a time once a stable strain is established, so they might provide a higher throughput than the WGT approach. Genomic libraries on vectors allow the generation of stable, pooled transformants for high-throughput screens with a genome-wide coverage, such as the MoBY-ORF library in S. cerevisiae73. However, direct application of S. cerevisiae-based molecular libraries in non-model yeasts such as Km requires technical optimization. In this regard, our findings about CEN and ARS compatibility may inform future studies to convert genomic libraries on Sc-based vectors into Km-compatible versions, advancing interspecific genetic screens.
The synthetic KS strains exhibited interesting phenotypic and molecular characteristics, demonstrating that the transferred chromosomes were functional, even across a considerable evolutionary distance, such as between yeast genera. Previous interspecific chromosomal transfers in yeast have been successful in using S. cerevisiae as a container to maintain heterologous genetic materials from bacteria22,69,74,75, and the epigenetic status of the heterologous chromosomes has been characterized only recently76. In our study, we found that the Sc chromosomes were “alive” in the Km background. Genes on the transferred R1 and R3 chromosomes were actively transcribed, despite a high level of sequence divergence. Genes on R1 and R3 showed significant expression differences between the KS and Sc backgrounds, and the same for Km genes between the KS and Km backgrounds. The transcriptional responses indicate active interspecific regulation, which can give rise to transgressive phenotypes, as shown with SPS22 and FLO9. It is of future interest to investigate what proportion of the regulatory interactions between the two species’ genomes are associated with beneficial phenotypes, or, on the other hand, indicative of molecular incompatibility.
The KS strains allowed for exploring patterns of regulatory evolution at an long evolutionary timescale. It has long been debated whether the evolution of gene expression primarily occurs through cis, trans, or both types of changes. A number of examples, especially from the evo-devo field, have argued for an important role of cis-regulatory changes77,78. That is, when a gene was transferred to a different species, its expression pattern was often determined by its cis-regulatory sequences (donor-like), suggesting that the trans environment was relatively conserved61. Recent transcriptomic and experimental studies showed that compensatory evolution of cis- and trans-acting factors is common, possibly driven by stabilizing selection9,59,61,62,79. However, most systematic studies so far were restricted to intrageneric hybrids59,62,79,80 or introgression lines60. The furthest phylogenetic distance examined so far by genome-wide analysis was probably between Saccharomyces cerevisiae and S. uvarum59,81,82, two species that share 80% nucleotide identity in coding regions83. Studies on the Saccharomyces species have found that gene expression divergence reached a plateau as the phylogenetic distance increases within the genus, with the contribution from cis-regulatory differences and compensatory changes becoming increasingly important59. Still, little is known for regulatory divergence beyond this phylogenetic scope. It is imaginable that both cis and trans-acting factors continuously accumulate nucleotide substitutions over a long evolutionary timespan, but it remains unknown whether they evolve in a compensatory manner under stabilizing selection, like what has been found within a genus, or whether they tend to change in the same direction to increase expression divergence. Evidence from enhancer-swaps showed a higher proportion of misregulation in cross-genus experiments than within Drosophila flies or Caenorhabditis worms, indicating that lineage-specific cis-trans co-evolution becomes abundant beyond the genus level, but the number of genes cataloged in this study was relatively small61. Our study examined the expression pattern of over a hundred Sc genes transferred to Km, providing evidence for long-term regulatory divergence at a relatively systematic scale. We found that transgressive expression, rather than cis-determined, was the most common pattern (37.7%) among the transferred genes. It is consistent with the model where both cis and trans changes accumulate with time, and they tend to act in opposite directions. At the same time, around one-third of genes showed conserved expression between Km and Sc, with contributions from compensatory cis-trans co-evolution as well. The data together point to the evolution of gene expression being under stabilizing selection in Saccharomycetaceae yeasts, across an evolutionary timespan of 114 million years. From a practical point of view, the prevalent transgression and its association with enhanced phenotypes demonstrate the potential of cross-species regulation in generating transgressive phenotypes, providing an alternative route to previous approaches that focus on optimizing individual proteins.
We note a few limitations in our cis-trans analysis. First, our sample size of 162 genes provided a statistically sufficient examination of regulatory evolution, but it was a small subset of the yeast genome. Future chromosomal transfers with larger chromosomes may provide more data. Second, we used the differences between KSsc and KSkm as cis-effects, reasoning that the two alleles were exposed to the same trans-environment, similar to previous studies with F1 hybrids59,84. However, we note that the estimation of cis effects might depend on trans-environment. For example, we might find more cis-divergence in our “monochromosomal hybrids” than in F1 hybrids due to the unavailability of certain Sc-specific trans-factors. Furthermore, unlike F1 analysis, we cannot properly estimate trans-divergence by subtracting cis effects from parental differences, because Sc and Km alleles experienced different levels of trans-changes. Therefore, we did not quantitatively compare our conclusions to previous F1 studies. We reasoned that our analysis, similar to previous ones with introgression lines and enhancer swaps, still informs about cis-trans co-evolution, and the abundant transgressive expression provides insights for both regulatory evolution and future phenotypic studies.
In summary, our study established a strategy for engineering the structural elements and ARS of a functional chromosome before transferring it to a remotely related species. This strategy might be extended to other microbes as well as to plants. The resultant synthetic strains provide valuable resources for cell factories, synthetic biology, and evolutionary genomics.
Methods
Strains and media
Yeast strains used in this study are listed in Supplementary Data 8. Sc strain W303-1A (MATa leu2-3,112 trp1-1 can1-100 ura3-1 ade2-1 his3-11,15) was used for chromosome engineering. The engineered Sc chromosome was transferred into a ura3Δ Km strain, FIM-1ΔU85.
The following media were used in this study: YPD (1% yeast extract, 2% peptone, 2% dextrose)86, synthetic medium lacking uracil or leucine (SD-Ura or SD-Leu) [0.17% yeast nitrogen base without amino acids and ammonium sulfate, 2% glucose, 1 g/L sodium glutamate, 2 g/L DO Supplement-Ura (630416, Takara) or DO Supplement-Leu (630414, Takara), 2% agar for plates], SD (same recipe as SD-Ura, with the addition of 20 mg/L uracil), and ME (2% malt extract, 3% agar). G418 (A600958, Sangon) and hygromycin (H8080, Solarbio) were added to YPD at final concentrations of 200 mg/L and 300 mg/L, respectively, to prepare YPDG and YPDH media. Both antibiotics were supplemented into YPD and SD-Ura to prepare YPDGH and SD-Ura+GH, respectively. Unless otherwise indicated, cells were grown at 30 °C.
Plasmids
All plasmids used in this study are listed in Supplementary Data 9. All primers are listed in Supplementary Data 1. A previously published vector LHZ626 served as a control for circular chromosomes42, which contains CEN and ARS elements from both Sc and Km (double CEN/ARS plasmid), as well as KmURA3, kanMX6, and hphMX4.
To construct plasmids for diploid selection, a fragment containing ScARSH4 and ScCEN6 was amplified from LHZ626 and inserted into the HpaI and SpeI sites of a Km-centromeric plasmid, LHZ88234, yielding LHZ1493. KanMX6 (kanR) was amplified from pFA6a-KanMX6 and used to replace hphMX4 (hygR) in LHZ1493, producing LHZ1494.
To create plasmids with different combinations of ARS and CEN, a BamHI site was first introduced between KmARS1 and KmCEN5 of LHZ882 to generate LHZ1495. Then, ScCEN6, ScARSH4, ScARSH4-ScCEN6 were inserted into LHZ1495 to replace their Km counterparts, generating LHZ1496-1498, respectively. Next, KmARS18 and KmCEN3 were inserted between the HindIII and SalI sites of LHZ1495 to replace KmARS1 and KmCEN5, giving rise to LHZ1783. ScARS1-ScCEN4, ScARS1, and ScCEN4 was inserted into LHZ1783 to replace their Km counterparts, generating LHZ1784-1786, respectively (see Supplementary Data 9 for restriction sites). Finally, ScARS1 and ScCEN4 were inserted into the SalI site of LHZ1783 to obtain LHZ1787, a plasmid with ARS and CEN from both species.
In order to remove the telomeres and join the chromosomal ends, two intermediate CRISPR plasmids targeting the chromosome ends were constructed by individually cloning the gRNAs into the SapI site of pRS425-Cas9-2×SapI. The gRNA cassette from one plasmid (containing the SNR52 promoter, gRNA, and SUP4 terminator) was then inserted into the NotI site of the other plasmid to construct the double-gRNA plasmids LHZ1499 (for chr1) and LHZ1500 (for chr3). In order to insert KmARS1 and KmCEN5 into the circularized chromosomes, we cloned the respective gRNAs into the SapI sites of pRS425-Cas9-2xSapI to construct LHZ1501∼1504. The homologous repair templates (donor) were assembled into pMD-18T with In-Fusion Snap Assembly (Clontech), according to the designs in Supplementary Data 10.
For flocculation analysis, the ScFLO9 cassette, including 1000 bp upstream, the ORF of ScFLO9, and 200 bp downstream sequence, was amplified from the genome of W303-1A. This cassette was inserted into the NotI site of LHZ626 to produce LHZ1505, and into the SmaI and SpeI sites of pRS315 (centromeric) and pRS425 (2-micron) to produce LHZ1509 and LHZ1510, respectively.
For NaCl tolerance analysis, the ScSPS22 cassette, including 1000 bp upstream sequence, the ORF of ScSPS22, and 200 bp downstream sequence, was amplified from the genome of W303-1A. This cassette was inserted into the NotI site of LHZ626 to produce LHZ1506, and into the SmaI and SpeI sites of pRS315 and pRS425 to produce LHZ1511 and 1512, respectively. The 1000 bp upstream sequence of ScSPS22 in LHZ1506 was replaced by the 1000 bp upstream sequence of KmSPS22 to produce LHZ1507. The KmSPS22 cassette, including 1000 bp upstream sequence, the ORF of KmSPS22, and 200 bp downstream sequence, was amplified from the genome of FIM-1ΔU and inserted into the NotI site of LHZ626 to produce LHZ1508. The full sequences of LHZ626, LHZ1495, and pRS425-Cas9-2xSapI are listed in Supplementary Data 9.
Transformation efficiency and plasmid stability
Fim-1ΔU and W303-1A were grown in liquid YPD overnight. Cells from 1 mL-culture were pelleted and transformed with LHZ626, LHZ1495∼LHZ1498, respectively, using the lithium acetate (LiAc) method87,88. Briefly, Fim-1ΔU cells were washed and mixed with 5 μL carrier DNA (Takara, 630440), 1 μg transforming DNA, 0.4 mL PEG solution (40% PEG 4000, 0.1 M LiAc, 10 mM Tris-HCl, pH 7.5, 1 mM EDTA), and 4.2 μL 1 M DTT. The mixture was incubated at 30 °C for 15 min and then heat-shocked at 47 °C for 15 min. Cells were pelleted and spread onto selective plates. Transformation of W303-1A followed the same protocol with the following modifications: DTT was omitted from the mixture, the volume of PEG solution was adjusted to 0.3 mL, and the mixture was incubated at 30 °C for 30 min, heat-shocked at 42 °C for 25 min, then incubated at 30 °C for an additional 10 min. Colony-forming units (CFUs) on YPDH plates were counted after 3 days, to evaluate transformation efficiency (CFUs/μg DNA). To measure the stability of the plasmid, transformants were grown in liquid YPD for 24 h. The cells were then diluted and spread onto YPD and YPDH plates. Stability was determined by dividing the CFU counts on YPDH plates by those on YPD plates. The experiments were replicated three times.
Mating assay
The quantitative mating assay was performed as previously described89 with modifications. Strains transformed with LHZ1493 (hygR) were used as experimental cells, and strains transformed with LHZ1494 (kanR) served as tester cells. The experimental and tester cells were cultured overnight in YPDH and YPDG liquid media, respectively. Cells were then washed and resuspended in water to an OD600 of 1. In each cross, 100 μL experimental cells were mixed with 500 μL tester cells. The mixture resulted in a 1:5 ratio of experimental to tester cells when both cell types were Km or Sc. It resulted in a 1:10 ratio of Sc experimental cells to Km tester cells, as CFU per OD600 of Km was twice that of Sc. The mixed cells were pelleted, resuspended in 20 μL ddH2O, and spotted onto ME plates. After incubation at 30 °C for 24 h, the cells were washed off the ME plates with ddH2O and then pelleted and resuspended in 1 mL ddH2O. To examine the mating result, 3 μL cell suspension was spotted on YPDH, YPDG, and YPDGH (Fig. 1e). These plates were incubated at 30 °C for 24 h. To quantify mating efficiency, the cells were diluted and plated onto YPD and YPDGH plates. The mating efficiency was calculated as the number of colonies on YPDGH plates divided by one-sixth of the number on YPD plates, reflecting the 1:5 ratio of experimental to tester cells. The experiments were replicated three times.
Engineering of Sc chromosome I and III
Each step of engineering Sc chromosome I (chr1) or III (chr3) involved transforming a CRISPR plasmid (Supplementary Data 9) and a donor, which is a PCR product for homologous recombination. The composition and full sequences of the donors are listed in Supplementary Data 10.
Chr1 and chr3 were circularized as previously described90. First, LHZ1499 or LHZ1500 was transformed into W303-1A to induce double-strand breaks in both telomeres in chr1 or chr3. A donor “chr1-cir” or “chr3-cir” containing KmURA3 with homologous arms to the chromosomal ends was simultaneously transformed to connect the two chromosomal ends with KmURA3. The constructs above were designed to remove the left telomere (0-1111 bp) and the right telomere along with the repetitive sequence (208917-252221 bp) of chr1, and the left telomere (0-831 bp) and the right telomere (334198-341087 bp) of chr3. The donors contain direct repeat (DR) sequences90, allowing marker recycling by popping out KmURA3 in future studies. Successful circularization was selected by the SD-Ura-Leu medium. The transformants with circularized chromosomes were named Sc-ring1 and Sc-ring3.
In order to insert KmARS1 and KmCEN5 into the circularized chromosomes, we transformed LHZ1501 and a donor “chr1-ARS1” into Sc-ring1 to insert KmARS1 and hygR at 66090 bp of chr1, resulting in a strain named Sc-ring1L. LHZ1502 and a donor “chr1-ARS1/CEN5” were transformed into Sc-ring1L to insert KmARS1/KmCEN5 and kanR at 157346 bp of chr1, resulting in a strain named Sc-R1. Similarly, LHZ1503 and a donor “chr3-ARS1/CEN5” were transformed into Sc-ring3 to insert KmARS1/KmCEN5 and hygR at 108718 bp of chr3, resulting in a strain named Sc-ring3L. LHZ1504 and a donor “chr3-ARS1” were transformed into Sc-ring3L to insert KmARS1 and kanR at 243183 bp of chr3, resulting in a strain named Sc-R3. All transformations were performed following the protocol for W303-1A described in the “Transformation efficiency and plasmid stability” section.
Extraction and protoplast transformation of R1 and R3
R1 and R3 were extracted from Sc-R1 and Sc-R3, respectively, following a previously described method41. In brief, two 200 mL cultures of Sc-R1 or Sc-R3 were grown in SD-Ura+GH medium to an OD₆₀₀ of 1.5. Cells from each 200 mL culture were harvested, washed, and resuspended in 40 ml SPE buffer (1 M sorbitol, 0.1 M sodium citrate, 0.01 M EDTA, pH 7.5). Zymolyase 20 T solution (10 mg/mL, 250 μL; E1005, Zymo Research) and β-mercaptoethanol (40 μL) were added, followed by incubation at 37 °C for 20 min to form spheroplasts. The spheroplasts were collected and gently resuspended in 1 mL 1 M sorbitol. Cell lysis was performed by adding 20 mL lysis buffer (50 mM Tris-HCl, 20 mM EDTA, 1% SDS, pH 12.8), followed by gentle inversion and incubation at 37 °C for 30 min. An equal volume (20 mL) of phenol:chloroform:isoamyl alcohol (25:24:1) was added, mixed gently, and then centrifuged. The aqueous phase (~20 mL) was collected, supplemented with 2 mL 3 M sodium acetate (pH 5.2) and 20 mL isopropanol, mixed, and centrifuged for 30 min. The DNA pellet was washed once with 70% ethanol and dissolved in 1 mL TE buffer (pH 8.0) containing 3 μL RNase A (10 mg/mL), followed by incubation at 37 °C for 30 min. DNA from the two preparations was combined and treated with 10 mL Buffer EX, 200 μL ATP-dependent exonuclease, and 300 μL 100 mM ATP at 37 °C for 30 min to remove linear DNA. The sample was mixed with 12 mL Buffer QS, centrifuged, and the supernatant was loaded onto a column equilibrated with Buffer QBT. The column was washed with 30 mL Buffer QC and DNA was eluted with 12 mL Buffer QF (reagents used after RNase treatment were from large-construct kit, 12462, Qiagen). GlycoBlue (15 μL; AM9515, Ambion), tRNA (15 μL; R5636, Sigma), 1.2 mL 3 M sodium acetate (pH 5.2), and 12 mL isopropanol were added to precipitate the DNA. After centrifugation for 50 min, the pellet was washed once with 70% ethanol, air-dried, and finally dissolved in 100 μL TE buffer. A total of 10 μL R1 or R3 was transformed into Km by protoplast transformation42. Briefly, 25 mL of an overnight culture of Fim-1ΔU cells was harvested, washed, and resuspended in 20 mL citrate–phosphate buffer (10 mM citrate–phosphate, 1.5 M sorbitol, pH 6.8). Zymolyase-20T solution (60 μL) and β-mercaptoethanol (40 μL) were added, followed by incubation at 30 °C for 15 ~ 20 min to form protoplasts. Protoplasts were collected, washed twice with 1.5 M sorbitol, and gently resuspended in STC solution (1.5 M sorbitol, 50 mM CaCl₂,10 mM Tris-HCl, pH 7.5) to a final concentration of 3 × 10⁸ cells/mL, as determined with a hemocytometer. A 100 μL aliquot of the protoplast suspension was gently mixed with R1 or R3 DNA and 5 μL carrier DNA, incubated on ice for 15 min, and then supplemented with 1 mL PEG 8000 solution (20% PEG 8000, 50 mM CaCl₂, 10 mM Tris-HCl, pH 7.5). After gentle mixing and incubation on ice for 15 min, protoplasts were pelleted, resuspended in 800 μL STC solution, and mixed with 10 mL regeneration agar (6.9 g/L yeast nitrogen base without amino acids, 2% glucose, 0.8 M sorbitol, 3% agar, 2 g/L DO Supplement-Ura) equilibrated at 47 °C. The mixture was poured onto the selective plates (the same recipe as regeneration agar except containing 2% agar). After 3 ~ 4 days, colonies were restreaked onto SD-Ura+GH plates.
Pulsed-Field Gel Electrophoresis (PFGE)
Plugs containing the genome of Sc, Km, and KS strains were prepared by using the CHEF Yeast Genomic DNA Plug Kit (170-3593, Biorad). To separate circular R1 from linear chromosomes in KS-R1, chromosomes were separated on the CHEF MAPPERTM XA System in a 1% pulsed field certified agarose gel (162-0137, Biorad) in 0.5×TBE (diluted from 10×TBE, T1051, Solarbio) at 14 °C. The running time was 24 h at 6.0 V cm-1, with a 60∼120 sec switch time ramp at an included angle of 120 °. The plug containing the entrapped R1 was cut out and linearized with NotI as described in the manufacturer’s protocol. Similarly, R3 in KS-R3 was separated from linear chromosomes and then linearized with AscI. Chromosomes of W303-1A, Sc-R1, Sc-R3, linearized R1 and R3 were separated in a 1% pulsed field certified agarose gel in 0.5×TBE at 14 °C. The running time was 16 h at 6.7 V cm-1, with a 10∼40 sec switch time ramp at an included angle of 120 °. Lambda PFG Ladder (N0341S, NEB) was used as a size marker for PFGE.
Growth curves of KS-R1 and KS-R3 and the stabilities of transferred chromosomes
As a control, Fim-1ΔU was transformed with LHZ626 to produce Km-V (Km-Vector), which confers G418 and hygromycin resistance, as well as Ura+ growth. KS-R1, KS-R3, and Km-V were cultured in YPDGH liquid medium overnight. The overnight culture (referred to as day 0 culture) was diluted into liquid YPD to an initial OD600 of 0.2 and grown at 30 °C. To monitor growth curves, the OD600 of the culture was recorded every 2 hours for the first 14 hours. To assess the stability of the transferred chromosomes, the culture was diluted into to an OD600 of 0.2 with fresh YPD every 24 hours, a period corresponding to approximately 7 generations. Cultures from day 0 and after 5 days of growth were diluted and plated onto YPD and YPDGH plates. The loss rate of transferred chromosomes per generation (x) was calculated using the following formula:
where \({N}_{0}\) and \({N}_{35}\) represent the ratios of CFU counts on YPDGH plates to those on YPD plates after 0 and 35 generations of non-selective growth in YPD, respectively34. The experiments were replicated three times.
Genome sequencing
Km-V, Sc-R1, Sc-R3, KS-R1, and KS-R3 were cultured overnight in SD-Ura+GH liquid medium. The cultures were diluted to an OD600 of 0.2 and cultured until the OD600 reached 0.6. The cells were collected and washed once with ddH2O. Genomic DNA was extracted using the E.Z.N.A. Fungal DNA Kit (D3390, OMEGA Bio-Tek) and sequenced on the Illumina NovaSeq 6000 platform using the 150-bp pair-end sequencing strategy (BIOZERON, Shanghai, China).
To construct the reference sequences for R1 and R3, sequences of chr1 and chr3 were extracted from the W303-1A genome (GenBank assembly: GCA002163515.1). We then manually modified the sequences to reflect our genome edits, including the insertions of Km elements and deletions of telomere sequences, giving rise to reference sequences R1 and R3 (provided in Supplementary Data 1). Annotations for R1 and R3 were transferred from S288C annotations (R64) with SnapGene. The whole-genome sequencing data were processed by BIOZERON biotechnology. To identify de novo variants during circular chromosome transfer, the raw sequencing data of Sc and KS strains were aligned to reference sequences for R1 or R3 using BWA91 with default settings. Picard tools were used to remove PCR duplicates. The average sequencing depth of KS-R1, Sc-R1, KS-R3, and Sc-R3 was 298, 1,183, 1,241 and 1,134 for the respective R1 or R3 chromosome. SNPs and indels were called following the best practices of GATK HaplotypeCaller92. Then, all heterozygous variants were removed from the VCF files of KS-R1 and KS-R3. All variants coexisting in both KS-R1 and Sc-R1 or KS-R3 and Sc-R3 were filtered out from the KS-R1 and KS-R3 VCF files as well. The remaining variants were considered to be de novo mutations generated during the chromosome transfer. In the case of R1, no de novo mutation was found.
To identify variants in Km, raw sequencing data of KS-R1, KS-R3, and Km-V were aligned to the reference genome of FIM1 (GenBank assembly: GCA_001854445.2)93 combined with either R1 or R3 sequence, using BWA with default parameters. Picard tools were used to remove PCR duplicates. The average sequencing depth of the Km genome for KS-R1, KS-R3, and Km-V was 1,036, 1,102, and 615, respectively. GATK HaplotypeCaller was used to identify the variants in these strains, following its best practices. VCFtools94 was used to remove variants shared between Km-V and KS-R1 or KS-R3, as well as heterozygous calls. The remaining unique, homozygous variants in KS-R1 and KS-R3 were listed in Supplementary Data 2.
Spot assay and quantification of growth phenotypes under various conditions
To investigate whether the chromosomal transfer resulted in enhanced phenotypes, W303-1A was transformed with LHZ626 to construct a control strain Sc-V (Sc-Vector). Km-V, Sc-V, Sc-R1, Sc-R3, KS-R1, and KS-R3 were cultured overnight in liquid SD-Ura+GH medium. The cultures were diluted to an OD600 of 1.0 and subjected to five serial 5-fold dilutions. These dilutions were spotted on plates using a 48-pin replicator. To evaluate the growth in different carbon sources, we used YP in combination with 2% ethanol, 3% glycerol, or different concentrations of glucose (0.02%, 1%, 3%, and 5%). To evaluate the growth in various nitrogen sources, we replaced 1 g/L sodium glutamate in the SD medium with 1 g/L threonine (Thr), 1 g/L serine (Ser), or 1.25 g of ammonium sulfate (1/4 N). For the condition of 1/2 AA and 1/2 N, the amino acids supplemented in the SD medium were halved, and 2.5 g/L ammonium sulfate was added. For chemical treatments, YPD plates were supplemented with H2O2 (0.012%, 0.016% and 0.024%), acetic acid (AcOH, 55, 57.5 and 60 mM), TM (0.2, 0.3 and 0.4 μg/mL), NaCl (0.6, 0.8 and 1 M), 20 mM DTT, 15 μg/mL 5-FU, or 1 M sorbitol. The plates were incubated at 30 °C or at other specified temperatures for 1-3 days before the pictures were taken.
Spot quantification was performed using the ‘gitter’ package95 in R. There were three biological replicates for each treatment, except for the three H2O2 treatments and the 60 mM AcOH treatment, where one biological replicate was removed due to technical problems in quantification. There were six biological replicates for YPD at 30 °C. Source data are provided in Supplementary Data 4. The quantified growth (G) under a specific condition (condition_x) was normalized to YPD growth at 30 °C of the same strain (strain_i), to generate a score for relative growth (RG):
In order to visualize inter-strain differences, data across three replicates were averaged and the average RG for a given strain under a given condition was subtracted by that of Km-V under the same condition, to generate RG’ for visualization (Fig. 3a):
Growth differences were tested with a linear model: RG ~ strain + treatment + treatment:strain, where the colon indicates interaction. p-values were extracted from the linear model and adjusted with the False Discovery Rate (FDR) method in R. Km-V and YPD at 30°C were used as the reference level for strain and treatment factors, respectively.
To evaluate chromosomal stability, spot dilution was performed as above on YPD supplemented with 20 μg/mL benomyl (BML), 10 μg/mL thiabendazole (TBZ), 0.05 M hydroxyurea (HU), 0.01% methyl methane sulfonate (MMS), 5 μg/mL camptothecin (CPT), or 0.05 μg/mL cycloheximide (CHX).
To perform spot assays of SPS22-expressing cells, Sc-R3 was transformed with LHZ1511 or LHZ1512, and FIM-1ΔU was transformed with LHZ1506, LHZ1507, or LHZ1508. Transformants were cultured overnight in SD-Leu medium. The culture was diluted and spotted onto plates with or without 1 M NaCl, as described above.
Flocculation analysis
Km-V, KS-R1, and Sc-R1 were cultured overnight in SD-Ura medium. Sc-R1 carrying LHZ1509 (ScFLO9CEN) or LHZ1510 (ScFLO92μ) and FIM-1ΔU carrying LHZ1505 (ScFLO9CEN) were cultured overnight in SD-Leu medium. Cells were harvested and washed with ddH2O and 250 mM EDTA. After two subsequent washes with ddH2O to ensure complete removal of EDTA, cells equivalent to an OD600 of 40 were pelleted and resuspended in 2 mL of ddH2O in a 15 mL tube. 100 μL 1 M Tris-HCl (pH 7.5) was added to the cell suspension, followed by 1 min of agitation. The tube was then left undisturbed, and pictures were taken every minute during a 5 min time window. Samples of the supernatant were taken from the surface of the cultures at various time points for OD600 measurement.
RNAseq and qPCR
Three biological replicates of Km-V, Sc-R1, Sc-R3, KS-R1, and KS-R3 were cultured overnight in SD-Ura+GH liquid medium. The cultures were diluted into YPD to achieve an OD600 of 0.2. Once the OD600 of the cultures reached 0.6, cells were collected directly for the YPD treatment. For stress treatment, the culture was supplemented with tunicamycin (T8480, Solarbio) at a final concentration of 1 μg/mL, or with NaCl at a final concentration of 1 M, and cells were collected after 1 hour. Total RNA was extracted using the ZR Fungal/Bacterial RNA MiniPrep kit (R2014, ZymoResearch). Samples were reversed transcribed using TruSeqTM RNA sample preparation Kit (Illumina, California, USA) and sequenced by Illumina HiSeq X Ten (BIOZERON, Shanghai, China). Batch information was provided in Supplementary Data 11.
The reads were mapped to corresponding reference genomes with BWA (0.7.17-r1188)91 under the default settings. Different reference genomes were used for different samples: Km-V samples were mapped to the FIM1 genome (GCA_001854445.2), S. cerevisiae samples were mapped to a W303 genome96 (GCA_002163515.1, ScRAPdb97 code W303.asm02.HP0), and KS samples were mapped to a combined genome of FIM1 and chrI and chrIII of W303. They were then processed with samtools98 to remove duplicated reads. Reads with mapping quality lower than or equal to 2 were also removed with samtools, as well as those with 30 or more soft-clipped bases. The number of reads mapped to each allele (with a minimal mapping quality of 10) was counted with htseq-count99, with the setting “-idattr ID -t CDS”. Counts from exons were summed up to generate counts per gene. The gene models for the RNA-seq analysis were based on the W303.asm02.HP0 annotation downloaded from ScRAPdb. There are 5,680 annotated genes in this genome, but not all genes were annotated to a known systematic name as in the S288C genome (R64), due to known intraspecific variation96. We performed an all-vs-all BLAST search between W303 and S288C genes (R64 assembly) and selected the top S288C hit for each W303 gene, based on E-value and query coverage. 5347 out of the 5680 (94%) W303 annotations matched S288C genes unambiguously. The rest W303 gene models were kept in the analysis but flagged with “ambiguous” in the dataset. The raw read counts are provided in Supplementary Data 11.
Differential expression was analyzed with DEseq2100. To examine changes in expression levels before and after chromosomal transfers (Sc vs. KS; Km vs KS), the data were fitted to a model of count ~ group, where group represented a combination between the strain factor (Sc/KS/Km) and the treatment factor (YPD/TM/NaCl). In the Sc analysis (Fig. 4a–d), R1 and R3 data were treated separately. Data from different treatments were combined in one data frame and normalized together. In the Km analysis, data from KS-R1, KS-R3 and Km-V across three treatments were combined and normalized together. Genes with an average read count lower than 5 of any group were removed from the analysis, leaving 218 out of 241 Sc genes (chrI and chrIII) and 4975 out of 5081 Km genes in the dataset. The median of read counts across groups was 1413 for Sc genes and 1267 for Km genes, prior to normalization. Adjusted p-values for pairwise comparisons between groups (e.g., Sc-YPD vs. KS-YPD) were derived from FDR-corrected Wald significance tests, with an FDR cutoff of 0.05. Log2 fold changes were shrunken with the ashr method101. Differentially expressed genes (DEG) were defined as an adjusted p-value lower than 0.05 and an absolute log2 fold change higher than 1.
For qPCR, RNA samples were reverse-transcribed using a PrimeScript RT Reagent Kit (RR037A, Takara, China). The qPCR was performed using ChamQ Universal SYBR qPCR Master Mix (Q711-02, Vazyme). The mRNA level was normalized to the average of three housekeeping genes, including MPE1, TRK1, and SWC4. Primers used in qPCR are listed in Supplementary Data 1. Statistical tests were performed with ANOVA followed by Tukey’s test to account for multiple comparisons.
Cis and trans effects
Orthologs were identified with OrthoFinder (3.0.1b1)102. Animo acid sequences of six species were provided to OrthoFinder, including S. cerevisiae (W303, GenBank #GCA_002163515.1), K. marxianus (FIM1, GCA_001854445.2), Candida albicans (SC5314, GCF_000182965.3), Lachancea thermotolerans (CBS6340, GCF_000142805.1), Naumovozyma castellii (CBS4309, GCF_000237345.1), and Torulaspora delbrueckii (CBS1146, GCF_000243375.1), along with a species tree according to Shen et al. (2018)23. There was a total of 32,488 genes across the six genomes, 93.5% of which were placed into orthogroups. There was a total of 5056 orthogroups, of which 3630 were with all species present and 2847 were single-copy orthogroups. When focusing on single-copy orthologues, the median of amino acid sequence similarity between S. cerevisiae and K. marxianus genes was 51.0% (using BLOSUM62 scoring matrix), between that of S. cerevisiae-T. delbrueckii (58.7%) and S. cerevisiae-C. albicans (37.0%), consistent with expectation (Supplementary Fig. 16a). The high level of evolutionary divergence resulted in loss of synteny (Supplementary Fig. 16b), so we did not further take into account of synteny in the ortholog analysis.
Between S. cerevisiae and K. marxianus, there were 4,851 orthologs, of which 3,813 were one-to-one orthologs. In the cases where multiple orthologs were identified for one gene (one-to-many or many-to-one) and many-to-many orthologous relationships, we performed pair-wise sequence alignments within the orthogroup and selected the reciprocal best match based on amino-acid sequence similarity, identifying 4,330 orthologous pairs. The median amino acid sequence similarity was 48.4%. The length of orthologs did not differ by more than two fold for 4,149/4,330 (95.8%) orthologous pairs, ruling out partial match due to local similarity. The orthologous genes were listed in Supplementary Data 6.
Differential gene expression was identified with DESeq2100. R1 and R3 data were combined. YPD, TM and NaCl data were analyzed separately. Out of 241 genes on chrI and chrIII of the W303 genome, we identified Km orthologs for 173 genes, 162 of which had an average read count above 5 across replicates and were retained in the analysis. First, we focused on genes showing significant divergence between Sc and Km (FDR-adjusted p < 0.05 by Wald test), which included 102 genes in YPD, 132 genes in NaCl and 122 genes in TM. We designated the expression level of Km alleles in Km-V as Pkm, Sc alleles in Sc-R1 or Sc-R3 as Psc, Km alleles in KS as KSkm, and Sc alleles in KS as KSsc. We used Wald test in DESeq2 to identify significant differences between (1) KSsc and KSkm (cis-effects), (2) KSsc and Psc (trans-effects), (3) Pkm and Psc (species divergence) and (4) KSsc and Pkm (similar to KSsc-KSkm comparison but accounting for trans-differences between K. marxianus and KS), using an FDR-adjusted p-value cutoff of 0.05. Effect sizes were adjusted with the ashr method.
Genes were classified according to Fig. 5c. “Km-like” genes were defined as no significant difference between KSsc and KSkm but a significant difference between KSsc and Psc. “Sc-like” genes were defined as no significant difference between KSsc and Psc but a significant difference between KSsc and KSkm. If KSsc was significantly different from both KSkm and Psc, the gene was further classified into “intermediate” and “transgressive”: “intermediate” means that KSsc was between Psc and KSkm, and “transgressive” means that KSsc was outside the range of Psc and KSkm. If KSsc was not significantly different from either Psc or KSkm, the gene was classified as “other (conserved)”.
The above analysis of cis-regulatory differences primarily considered allelic differences in KS, where the two alleles share the same trans-environment, similar to previous studies on F1 hybrids62,84,103,104. Comparing KSsc to Pkm also reveals cis-differences, but they might be sometimes affected by the slight differences between the trans-environment of KS and Km, i.e., reflecting cis- and trans-effects for some genes (Supplementary Fig. 14a). In order to address if the trans-difference between KS and Km changed our conclusion, we performed the same classification analysis as above, but used Pkm in place of KSkm (Supplementary Fig. 14).
In order to understand the major source of variance in the transcriptomic data, we performed a PCA with data from three treatments combined and normalized together. The normalized data were transformed with the VarianceStabilizingTransformation function in DESeq2 prior to the PCA.
Genome-wide expression divergence between Km and Sc orthologs was analyzed for 4,307 genes whose average read counts across replicates were higher than 5 for every condition. Sc data were from Sc-R1. YPD, NaCl, and TM data were analyzed separately. DEG was identified by comparing Psc and Pkm with Wald test in DESeq2, with an FDR-adjusted p-value cutoff of 0.05. GO analysis was performed with SGD Gene Ontology Term Finder (Version 0.86, GO version 2025-07-22), with 4,307 genes as the background set. Cis-divergence for the transferred genes was from comparison between KSsc and KSkm described above (FDR-adjusted p-value < 0.05). The results for cis-trans analysis and genome-wide divergence are provided in Supplementary Data 7.
Statistics & reproducibility
For phenotypic analysis, RNA-seq, and gene expression assessment by qPCR, each strain was analyzed with three biological replicates. For KS-R1, the three biological replicates represent three independent transformants. The spot assay was replicated three times, with the exception of the three H₂O₂ treatments and the 60 mM AcOH treatment, which were replicated twice; the results obtained were consistent throughout. For Fig. 2b–e, each experiment was repeated three times independently with similar results. No data was excluded. Randomization and blinding were not applicable in this study.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The DNA-seq and RNA-seq data generated in this study have been deposited in the NCBI database under accession code PRJNA1102743. Source data generated in this study are provided in the Source Data file and Supplementary Data files. Source data are provided with this paper.
References
Soucy, S. M., Huang, J. & Gogarten, J. P. Horizontal gene transfer: building the web of life. Nat. Rev. Genet. 16, 472–482 (2015).
Steensels, J., Gallone, B. & Verstrepen, K. J. Interspecific hybridization as a driver of fungal evolution and adaptation. Nat. Rev. Microbiol. 19, 485–500 (2021).
Stegemann, S., Keuthe, M., Greiner, S. & Bock, R. Horizontal transfer of chloroplast genomes between plant species. Proc. Natl. Acad. Sci. 109, 2434–2438 (2012).
Ma, L.-J. et al. Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 464, 367–373 (2010).
Gibson, B. & Liti, G. Saccharomyces pastorianus: genomic insights inspiring innovation for industry. Yeast Chichester Engl. 32, 17–27 (2015).
Mergoum, M. et al. Triticale: A “New” Crop with Old Challenges. 267–287 (Handbook of Plant Breeding, vol 3. Springer, New York, 2009).
Chen, Z. J. Genomic and epigenetic insights into the molecular bases of heterosis. Nat. Rev. Genet. 14, 471–482 (2013).
Landry, C. R., Hartl, D. L. & Ranz, J. M. Genome clashes in hybrids: insights from gene expression. Heredity 99, 483–493 (2007).
Signor, S. A. & Nuzhdin, S. V. The Evolution of Gene Expression in cis and trans. Trends Genet. 34, 532–544 (2018).
Gombert, A. K., Madeira, J. V., Cerdán, M.-E. & González-Siso, M.-I. Kluyveromyces marxianus as a host for heterologous protein synthesis. Appl. Microbiol. Biotechnol. 100, 6193–6208 (2016).
Bilal, M. et al. Bioprospecting Kluyveromyces marxianus as a Robust Host for Industrial Biotechnology. Front. Bioeng. Biotechnol. 10, 851768 (2022).
Groeneveld, P., Stouthamer, A. H. & Westerhoff, H. V. Super life – how and why ‘cell selection’ leads to the fastest-growing eukaryote. FEBS J. 276, 254–270 (2009).
Lertwattanasakul, N. et al. Utilization capability of sucrose, raffinose and inulin and its less-sensitiveness to glucose repression in thermotolerant yeast Kluyveromyces marxianus DMKU 3-1042. AMB Express 1, 20 (2011).
Rodrussamee, N. et al. Growth and ethanol fermentation ability on hexose and pentose sugars and glucose effect under various conditions in thermotolerant yeast Kluyveromyces marxianus. Appl. Microbiol. Biotechnol. 90, 1573–1586 (2011).
Fisk, D. G. et al. Saccharomyces cerevisiae S288C genome annotation: a working hypothesis. Yeast 23, 857–865 (2006).
Witte, V., Grossmann, B. & Emeis, C. C. Molecular probes for the detection of Kluyveromyces marxianus chromosomal DNA in electrophoretic karyotypes of intergeneric protoplast fusion products. Arch. Microbiol. 152, 441–446 (1989).
Albonico, F., B, E., G, P. H. & B, D. New Saccharomyces cerevisiae-Kluyveromyces marxianus fusant shows enhanced alcoholic fermentation performance. World J. Microbiol. Biotechnol. 38, 251 (2022).
Fournier, R. E. & Ruddle, F. H. Microcell-mediated transfer of murine chromosomes into mouse, Chinese hamster, and human somatic cells. Proc. Natl. Acad. Sci. 74, 319–323 (1977).
Fukagawa, T. et al. Dicer is essential for formation of the heterochromatin structure in vertebrate cells. Nat. Cell Biol. 6, 784–791 (2004).
O’Doherty, A. et al. An aneuploid mouse strain carrying human chromosome 21 with Down syndrome phenotypes. Science 309, 2033–2037 (2005).
Wilson, M. D. et al. Species-specific transcription in mice carrying human chromosome 21. Science 322, 434–438 (2008).
Lartigue, C. et al. Genome transplantation in bacteria: changing one species to another. Science 317, 632–638 (2007).
Shen, X.-X. et al. Tempo and Mode of Genome Evolution in the Budding Yeast Subphylum. Cell 175, 1533–1545.e20 (2018).
Kitaoka, M., Smith, O. K., Straight, A. F. & Heald, R. Molecular conflicts disrupting centromere maintenance contribute to Xenopus hybrid inviability. Curr. Biol. 32, 3939–3951.e6 (2022).
Kitano, J. & Okude, G. Causative genes of intrinsic hybrid incompatibility in animals and plants: what we have learned about speciation from the molecular perspective. Evol. J. Linn. Soc. 3, kzae022 (2024).
Lartigue, C. et al. Creating bacterial strains from genomes that have been cloned and engineered in yeast. Science 325, 1693–1696 (2009).
Nieduszynski, C. A., Knox, Y. & Donaldson, A. D. Genome-wide identification of replication origins in yeast by comparative genomics. Genes Dev. 20, 1874–1879 (2006).
Theis, J. F. & Newlon, C. S. The ARS309 chromosomal replicator of Saccharomyces cerevisiae depends on an exceptional ARS consensus sequence. Proc. Natl. Acad. Sci. 94, 10786–10791 (1997).
Iborra, F. & Ball, M. M. Kluyveromyces marxianus small DNA fragments contain both autonomous replicative and centromeric elements that also function in Kluyveromyces lactis. Yeast Chichester Engl. 10, 1621–1629 (1994).
Ball, M. M., Raynal, A., Guérineau, M. & Iborra, F. Construction of efficient centromeric, multicopy and expression vectors for the yeast Kluyveromyces marxianus using homologous elements and the promoter of a purine-cytosine-like permease. J. Mol. Microbiol. Biotechnol. 1, 347–353 (1999).
Rajkumar, A. S., Varela, J. A., Juergens, H., Daran, J.-M. G., and Morrissey, J. P. Biological Parts for Kluyveromyces marxianus Synthetic Biology. Front. Bioeng. Biotechnol. 7 (2019).
Abdel-Banat, B. M. A., Hoshida, H. & Akada, R. Various short autonomously replicating sequences from the yeast Kluyveromyces marxianus seemingly without canonical consensus. Curr. Res. Microb. Sci. 2, 100053 (2021).
Hegemann, J. H. & Fleig, U. N. The centromere of budding yeast. BioEssays 15, 451–460 (1993).
Ren, H. et al. Establishment of a Cre-loxP System Based on a Leaky LAC4 Promoter and an Unstable panARS Element in Kluyveromyces marxianus. Microorganisms 10, 1240 (2022).
Sikorski, R. S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Gnügge, R. & Rudolf, F. Saccharomyces cerevisiae Shuttle vectors. Yeast Chichester Engl. 34, 205–221 (2017).
Bitoun, R. & Zamir, A. Spontaneous amplification of yeast CEN ARS plasmids. Mol. Gen. Genet. 204, 98–102 (1986).
Zhou, H. et al. Efficient and markerless gene integration with SlugCas9-HF in Kluyveromyces marxianus. Commun. Biol. 7, 797 (2024).
Peska, V. et al. Extraordinary diversity of telomeres, telomerase RNAs and their template regions in Saccharomycetaceae. Sci. Rep. 11, 12784 (2021).
Shao, Y. et al. Creating a functional single-chromosome yeast. Nature 560, 331–335 (2018).
Noskov, V. N. et al. Isolation of circular yeast artificial chromosomes for synthetic biology and functional genomics studies. Nat. Protoc. 6, 89–96 (2011).
Lyu, Y., Wu, P., Zhou, J., Yu, Y. & Lu, H. Protoplast transformation of Kluyveromyces marxianus. Biotechnol. J. 16, e2100122 (2021).
Tschumper, G. & Carbon, J. High frequency excision of Ty elements during transformation of yeast. Nucleic Acids Res. 14, 2989–3001 (1986).
Wu, P. et al. Transfer of disulfide bond formation modules via yeast artificial chromosomes promotes the expression of heterologous proteins in Kluyveromyces marxianus. mLife 3, 129–142 (2024).
Travesa, A. & Wittenberg, C. Turned on by genotoxic stress. Cell Cycle 11, 3145–3146 (2012).
Walker, G. M. & Basso, T. O. Mitigating stress in industrial yeasts. Fungal Biol. 124, 387–397 (2020).
Chakrabortee, S. et al. Intrinsically Disordered Proteins Drive Emergence and Inheritance of Biological Traits. Cell 167, 369–381.e12 (2016).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Coluccio, A. et al. Morphogenetic pathway of spore wall assembly in Saccharomyces cerevisiae. Eukaryot. Cell 3, 1464–1475 (2004).
Nguyen, T. H., Fleet, G. H. & Rogers, P. L. Composition of the cell walls of several yeast species. Appl. Microbiol. Biotechnol. 50, 206–212 (1998).
Soares, E. V. Flocculation in Saccharomyces cerevisiae: a review. J. Appl. Microbiol. 110, 1–18 (2011).
Teunissen, A. W. & Steensma, H. Y. Review: the dominant flocculation genes of Saccharomyces cerevisiae constitute a new subtelomeric gene family. Yeast Chichester Engl. 11, 1001–1013 (1995).
Nonklang, S. et al. Construction of flocculent Kluyveromyces marxianus strains suitable for high-temperature ethanol fermentation. Biosci. Biotechnol. Biochem. 73, 1090–1095 (2009).
Brothers, M. & Rine, J. Distinguishing between recruitment and spread of silent chromatin structures in Saccharomyces cerevisiae. eLife 11, e75653 (2022).
Sjöstrand, J. O. O., Kegel, A. & Aström, S. U. Functional diversity of silencers in budding yeasts. Eukaryot. Cell 1, 548–557 (2002).
Aström, S. U., Kegel, A., Sjöstrand, J. O. & Rine, J. Kluyveromyces lactis Sir2p regulates cation sensitivity and maintains a specialized chromatin structure at the cryptic alpha-locus. Genetics 156, 81–91 (2000).
De Boer, C. G. & Hughes, T. R. YeTFaSCo: A database of evaluated yeast transcription factor sequence specificities. Nucleic Acids Res. 40, 169–179 (2012).
Wittkopp, P. J. Contributions of mutation and selection to regulatory variation: lessons from the Saccharomyces cerevisiae TDH3 gene. Philos. Trans. R. Soc. B Biol. Sci. 378, 20220057 (2023).
Metzger, B. P. H., Wittkopp, P. J. & Coolon, J. D. Evolutionary Dynamics of Regulatory Changes Underlying Gene Expression Divergence among Saccharomyces Species. Genome Biol. Evol. 9, 843–854 (2017).
Guerrero, R. F., Posto, A. L., Moyle, L. C. & Hahn, M. W. Genome-wide patterns of regulatory divergence revealed by introgression lines. Evol. Int. J. Org. Evol. 70, 696–706 (2016).
Gordon, K. L. & Ruvinsky, I. Tempo and Mode in Evolution of Transcriptional Regulation. PLoS Genet. 8, e1002432 (2012).
Tirosh, I., Reikhav, S., Levy, A. A. & Barkai, N. A Yeast Hybrid Provides Insight into the Evolution of Gene Expression Regulation. Science 324, 659–662 (2009).
Landry, C. R. et al. Compensatory cis-trans evolution and the dysregulation of gene expression in interspecific hybrids of Drosophila. Genetics 171, 1813–1822 (2005).
Barrière, A., Gordon, K. L. & Ruvinsky, I. Coevolution within and between regulatory loci can preserve promoter function despite evolutionary rate acceleration. PLOS Genet. 8, e1002961 (2012).
Steensels, J. et al. Improving industrial yeast strains: exploiting natural and artificial diversity. FEMS Microbiol. Rev. 38, 947–995 (2014).
Pobiega, S. & Marcand, S. Dicentric breakage at telomere fusions. Genes Dev. 24, 720–733 (2010).
Postma, E. D. et al. A supernumerary designer chromosome for modular in vivo pathway assembly in Saccharomyces cerevisiae. Nucleic Acids Res. 49, 1769–1783 (2021).
Zhou, J. et al. Exogenous artificial DNA forms chromatin structure with active transcription in yeast. Sci. China Life Sci. 65, 851–860 (2022).
Gibson, D. G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010).
Peris, D. et al. Synthetic hybrids of six yeast species. Nat. Commun. 11, 2085 (2020).
Zhang, W. & Geng, A. Improved ethanol production by a xylose-fermenting recombinant yeast strain constructed through a modified genome shuffling method. Biotechnol. Biofuels 5, 46 (2012).
Stojiljković, M. et al. Whole-genome transformation of yeast promotes rare host mutations with a single causative snp enhancing acetic acid tolerance. Mol. Cell. Biol. 42, e0056021 (2022).
Ho, C. H., et al. A molecular barcoded yeast ORF library enables mode-of-action analysis of bioactive compounds. Nat. Biotechnol. 27, https://doi.org/10.1038/nbt.1534 (2009).
Fredens, J. et al. Total synthesis of Escherichia coli with a recoded genome. Nature 569, 514–518 (2019).
Venetz, J. E. et al. Chemical synthesis rewriting of a bacterial genome to achieve design flexibility and biological functionality. Proc. Natl. Acad. Sci. USA. 116, 8070–8079 (2019).
Meneu, L. et al. Sequence-dependent activity and compartmentalization of foreign DNA in a eukaryotic nucleus. Science 387, eadm9466 (2025).
Carroll, S. B. Evo-Devo and an Expanding Evolutionary Synthesis: A Genetic Theory of Morphological Evolution. Cell 134, 25–36 (2008).
Stern, D. L. & Orgogozo, V. The loci of evolution: how predictable is genetic evolution? Evolution 62, 2155–2177 (2008).
McManus, C. J. et al. Regulatory divergence in Drosophila revealed by mRNA-seq. Genome Res. 20, 816–825 (2010).
Díaz, F., Wolf, J. & de Brito, R. A. Transcriptional Misexpression in Hybrids between Species Linked by Gene Flow Is Associated With Patterns of Sequence Divergence. Genome Biol. Evol. 15, evad071 (2023).
Bullard, J. H., Mostovoy, Y., Dudoit, S. & Brem, R. B. Polygenic and directional regulatory evolution across pathways in Saccharomyces. Proc. Natl. Acad. Sci. USA. 107, 5058–5063 (2010).
Li, X. C. & Fay, J. C. Cis-regulatory divergence in gene expression between two thermally divergent yeast species. Genome Biol. Evol. 9, 1120–1129 (2017).
Kellis, M., Patterson, N., Endrizzi, M., Birren, B. & Lander, E. S. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423, 241–254 (2003).
Krieger, G., Lupo, O., Levy, A. A. & Barkai, N. Independent evolution of transcript abundance and gene regulatory dynamics. Genome Res. 30, 1000–1011 (2020).
Zhou, J., Zhu, P., Hu, X., Lu, H. & Yu, Y. Improved secretory expression of lignocellulolytic enzymes in Kluyveromyces marxianus by promoter and signal sequence engineering. Biotechnol. Biofuels 11, 235 (2018).
Amberg, D. C., Burke, D., Strathern, J. N., and Burke, D. Methods in yeast genetics: a Cold Spring Harbor Laboratory course manual 2005 ed. (Cold Spring Harbor Laboratory Press, 2005).
Antunes, D. F., de Souza Junior, C. G. & de Morais Junior, M. A. A simple and rapid method for lithium acetate-mediated transformation of Kluyveromyces marxianus cells. World J. Microbiol. Biotechnol. 16, 653–654 (2000).
Gietz, R. D. & Schiestl, R. H. Large-scale high-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 38–41 (2007).
Shi, T., Zeng, J., Zhou, J., Yu, Y. & Lu, H. Correlation Between Improved Mating Efficiency and Weakened Scaffold-Kinase Interaction in the Mating Pheromone Response Pathway Revealed by Interspecies Complementation. Front. Microbiol. 13, 865829 (2022).
Shao, Y., Lu, N., Qin, Z. & Xue, X. CRISPR-Cas9 Facilitated Multiple-Chromosome Fusion in Saccharomyces cerevisiae. ACS Synth. Biol. 7, 2706–2708 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinforma. Oxf. Engl. 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Yu, Y. et al. Comparative genomic and transcriptomic analysis reveals specific features of gene regulation in Kluyveromyces marxianus. Front. Microbiol. 12, 598060 (2021).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Wagih, O. & Parts, L. gitter: a robust and accurate method for quantification of colony sizes from plate images. G3 Bethesda 4, 547–552 (2014).
Matheson, K., Parsons, L. & Gammie, A. Whole-genome sequence and variant analysis of w303, a widely-used strain of Saccharomyces cerevisiae. G3 Bethesda 7, 2219–2226 (2017).
Miao, Z. et al. ScRAPdb: an integrated pan-omics database for the Saccharomyces cerevisiae reference assembly panel. Nucleic Acids Res. 53, D852–D863 (2025).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Stephens, M. False discovery rates: a new deal. Biostat. Oxf. Engl. 18, 275–294 (2017).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Emerson, J. J. et al. Natural selection on cis and trans regulation in yeasts. Genome Res. 20, 826–836 (2010).
Fraser, H. B. et al. Polygenic cis-regulatory adaptation in the evolution of yeast pathogenicity. Genome Res. 22, 1930–1939 (2012).
Minh, B. Q. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Acknowledgements
Science and Technology Research Program of Shanghai #24ZR1406500: Y.Y. Science and Technology Research Program of Shanghai #2023ZX01: H.L. National Key Research and Development Program of China #2021YFA0910603: Y.Y. National Key Research and Development Program of China #2022YFC2106201: J.Z. National Key Research and Development Program of China #2021YFA0910601: H.L. Open Fund of State Key Laboratory of Genetic Engineering #SKLGE-2318: Y.Y. The Fundamental Research Funds for the Central Universities: X.C.L. Young Scientists Fund of the National Natural Science Foundation of China #32400495: X.C.L.
Author information
Authors and Affiliations
Contributions
The conceptualization of the study was carried out by H.L. and Y.Y., and the methodology was developed by H.L., Y.Y., and Y.L. Investigation was performed by Y.L., Y.S., K.S., J.Z., H.C., and X.C.L. The original draft was written by Y.L. and Y.Y., and the review and editing were conducted by Y.L., Y.Y., and X.C.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kevin Verstrepen, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lyu, Y., Shi, Y., Song, K. et al. Intergeneric chromosomal transfer in yeast results in improved phenotypes and widespread transcriptional responses. Nat Commun 17, 1419 (2026). https://doi.org/10.1038/s41467-025-68164-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68164-8