Abstract
The concept of organ-organ communication in systemic diseases has gained significant traction. Despite accumulating evidence of liver-kidney crosstalk, the underlying regulatory mechanisms remain poorly understood. Here, we identify hepatokine fibrinogen-like protein 1 (FGL1) as a pivotal mediator driving liver-kidney communication in renal fibrosis. Integrative RNA sequencing and clinical analyses revealed FGL1 upregulation in the liver post-renal injury, with subsequent selective accumulation in fibrotic kidneys. Hepatocyte-specific knockout of FGL1 improved renal morphology, attenuated collagen deposition, and inhibited fibrosis markers in unilateral ureteral obstruction and adenine-induced mouse models. Conversely, FGL1 overexpression exacerbated renal fibrosis. Mechanistically, FGL1 binds to TGF-β receptor ALK5 and blocks its interaction with K48-linked polyubiquitin. Structural analyses identified critical residues in FGL1-ALK5 interaction. Hepatic FGL1 is transcriptionally regulated by STAT3, and activated by inflammatory cytokines (IL-6, IL-1β and TNF-α). Pharmacological inhibition by p-hydroxybenzaldehyde (screened out from 4080 compounds) or an anti-FGL1 monoclonal antibody significantly reduced renal fibrosis without hepatotoxicity. These findings establish FGL1 as a key mediator of pathogenic liver-kidney crosstalk and highlights therapeutic strategies to targeting organ-organ networks in renal fibrosis.
Similar content being viewed by others
Introduction
The concept of organ-organ crosstalk, mediated by endocrine factors such as secreted proteins and extracellular vesicles, has revolutionized our understanding of systemic health and disease1. Recent studies have identified diverse mediators, including hepatokines, cytokines, myokines, adipokines, and metabolites2,3,4. Coagulation factor XI, a liver-derived protein, exhibits cardioprotective roles in liver-heart crosstalk independent of its coagulation function5. A brain-heart axis is characterized by which myocardial infarction triggers brain-mediated monocytic TNF generation to enhance slow-wave sleep and promote cardiac repair6. Liver-bone communication is regulated by hepatocyte Sirtuin 2, which modulates LRG1 in extracellular vesicles to suppress osteoclast differentiation7. Emerging scientific data also reveal gut-liver interactions mediated by microbiota-derived metabolites8, heart-kidney dialog via cardiac natriuretic peptides9, and other tissue-tissue crosstalk10. These findings not only deepen our understanding of systemic disease mechanisms but also highlight therapeutic opportunities to target inter-organ signaling hubs.
Renal fibrosis, a pathological hallmark of chronic kidney disease (CKD), is characterized by excessive extracellular matrix (ECM) deposition, leading to irreversible structural and functional kidney decline11. Multiple drivers, including TGF-β, Ang II, oxidative stress, and hemodynamic imbalances, contribute to disease progression12. TGF-β signaling, a central regulator, activates myofibroblasts, induces epithelial-mesenchymal transition (EMT) and promotes collagen synthesis through canonical SMAD-dependent pathways13,14. In spite of these scientific insights, current therapies targeting intrinsic renal pathways remain insufficient, underscoring the need to explore systemic mediators and inter-organ communication mechanisms.
The liver, a key endocrine organ, secretes hepatokines that regulate distant systemic physiology and pathology15. For instance, activin-E, predominantly expressed and secreted by the liver, is elevated in individuals with obesity and non-alcoholic fatty liver disease16. Similarly, ANGPTL3, another hepatokine, modulates plasma triglyceride levels by inhibiting lipoprotein lipase in oxidative tissues17. Clinical and experimental evidence underscores the functional interplay between hepatic and renal pathology. Acute liver injury or advanced cirrhosis frequently precipitates renal impairment18. Bile duct ligation (BDL) models demonstrate that liver injury can induce renal damage, supporting the existence of a liver-kidney axis19. Despite these associations, the molecular mediators of liver-kidney crosstalk in renal fibrosis remain largely unexplored. Identifying these hepatokines and their mechanisms of action could unveil novel therapeutic strategies to disrupt fibrotic signaling networks.
In this work, we combine transcriptomic profiling, functional genetics, and pharmacological interventions to characterize key endocrinal mediators for liver-kidney dialog, underlying mechanisms, and candidate drugs in renal fibrosis. Our findings position FGL1 as a pivotal mediator of liver-kidney crosstalk and a promising therapeutic target, bridging the gap between systemic mediators and localized fibrotic pathways.
Results
Integrative RNA sequencing and clinical correlation analysis identify FGL1 as a key mediator of liver-kidney communication
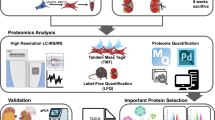
To identify endocrine mediators of liver-kidney crosstalk, we combined RNA sequencing with clinical correlation analysis. First, we established a unilateral ureteral obstruction (UUO) mouse model of renal fibrosis and conducted RNA sequencing on liver tissues 14 days post-surgery (Fig. 1a). Of the 56,980 genes detected in liver tissues, 966 exhibited significantly differential expression in response to renal injury (p < 0.05, Fig. 1b). The top-ranked upregulated hepatokines included Fgl1, Orm2, Mt2, Saa1, and Saa2 (Log2fold-change > 4, adjusted p < 0.0001). qPCR validated robust hepatic upregulation of these five hepatokines (Fig. 1c). To identify hepatokines that potentially translocate to the kidneys, we analyzed their expressions in the kidneys of UUO mice. Strikingly, FGL1 protein expression was markedly elevated in obstructed kidneys, whereas the other four candidate hepatokines showed no significant changes (Fig. 1d).
a Schematic illustration of the experimental strategy to identify significantly altered genes in the liver of mice subjected to unilateral ureteral obstruction (UUO). Created in BioRender. Wu, W. (2025) https://BioRender.com/i0fgw4p. b Volcano plot showing differentially expressed genes in the liver of mice 14 days post-UUO compared to sham-operated controls (n = 3). c qPCR analysis of five hepatokines (Fgl1, Orm2, Saa1, Saa2, and Mt2) in the liver of Sham or UUO mice (n = 6). d Western blot analysis of five hepatokines (FGL1, ORM2, SAA1, SAA2, and MT2A) in the kidney of Sham or UUO mice (n = 4). e ELISA analysis of the serum levels of five hepatokines in healthy individuals (n = 18) and chronic kidney disease (CKD) patients (n = 18). Box plots show the center line as median, whiskers show maxima and minima, and box limits show upper and lower quartiles. f Pearson correlation analysis of five hepatokines with creatinine (CR) and urea nitrogen (BUN) levels (n = 36). g, h UAMP visualization of liver or renal cell clusters based on human single-cell transcriptomes. Red dots indicate the expression of FGL1 (GSE115469, GSE131685). i ELISA analysis of FGL1 levels in the serum of UUO mice (n = 6). j FGL1 protein levels in the kidneys of mice subjected to UUO or UUO + Hemoperfusion (HP; Sham group, n = 6; UUO group, n = 7; UUO + HP group, n = 9). Under anesthesia, transcardial perfusion with saline was performed via left ventricle puncture (15 mL/min for 3 min), with the right auricle incised to allow outflow. Kidney tissues were harvested for Western blot analysis. k Representative biophotonic images showing fluorescence signals in various tissues from hepatocyte-FGL1 overexpressed UUO mice (n = 3). Data are presented as mean ± SD. The p values were determined by a two-tailed t test (c–e) or one-way ANOVA with Tukey’s correction (i, j). Source data are provided as a Source data file.
Next, clinical correlation was further assessed using serum samples from CKD patients (n = 18) and healthy individuals (n = 18). FGL1 protein level was significantly elevated in CKD patients (p = 0.0058, Fig. 1e) and positively correlated with renal injury markers, blood urea nitrogen (BUN, r = 0.51) and creatinine (CR, r = 0.51) (Fig. 1f). The remaining hepatokines exhibited weak clinical associations. These findings prioritized FGL1 as a key candidate mediator for liver-kidney communication. Single-cell transcriptomic profiling (n = 2 datasets) confirmed FGL1 expression specific to hepatocytes (2.93 ± 0.83) with negligible detection in kidney cells (Fig. 1g, h and Supplementary Fig. 1a, b). Intriguingly, while hepatic Fgl1 mRNA was markedly upregulated in UUO mice, total liver FGL1 protein levels remained unchanged (Supplementary Fig. 1c), suggesting rapid secretion into systemic circulation (Fig. 1i) and subsequent renal accumulation. In another BDL-induced liver injury mouse model, we also observed that FGL1 was rapidly mobilized from the liver to circulation and subsequently deposited in the kidneys (Supplementary Fig. 1d, e). Hemoperfusion experiments demonstrated significant reduction of renal FGL1 following blood clearance (p = 0.032, Fig. 1j), directly linking circulating FGL1 to renal deposition. Strikingly, in hepatocyte-FGL1 overexpressed UUO mice, biophotonic imaging detected FGL1 enrichment exclusively in injured kidneys, with no detectable signal in lung, heart, white adipose, spleen, or skeletal muscle (Fig. 1k).
Hepatic FGL1 drives renal fibrosis using gain- and loss-of-function approaches
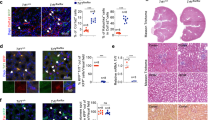
To investigate the role of FGL1 in liver-kidney crosstalk, we generated mice with hepatocyte-specific FGL1 deletion using a Cre/loxP-dependent conditional knockout (HepKO) system (Supplementary Fig.2a–d). In male mice subjected to UUO, hepatocyte-specific FGL1 knockout ameliorated renal morphological damage and reduced collagen deposition (Fig. 2a–f). Second harmonic generation (SHG) microscopy confirmed that fibrillar collagen was greatly attenuated in the kidney of FGL1 HepKO mice (Fig. 2g, h). In line with these observations, FGL1 knockout suppressed the expression of ECM deposition marker (FN), EMT marker (VIMENTIN), and fibrogenic factors (Fig. 2i, j).
a Schematic illustration of the experimental approach. FGL1fl/fl mice (Ctrl) and hepatocyte-specific FGL1 knockout (FGL1 HepKO) mice were subjected to unilateral ureteral obstruction (UUO) for 14 days. Created in BioRender. Wu, W. (2025) https://BioRender.com/a4vde7a. b Gross appearance of whole kidneys from Ctrl and FGL1 HepKO mice (Scale bar, 2 mm). c–f Representative images and quantitative results of HE staining, Masson staining, and COL-I staining in the kidneys of Sham or UUO mice (n = 3, scale bars, 100 μm). g, h Representative images and quantitative results of SHG images in the kidneys of Sham or UUO mice (n = 3, scale bars, 100 μm). SHG: second harmonic generation, TPEF: two-photon excited fluorescence. i Western blot analysis of FN, VIMENTIN and α-SMA in the kidney of Sham or UUO mice (n = 6). FN: fibronectin, the extracellular matrix deposition marker, VIMENTIN: the epithelial-to-mesenchymal transition marker, α-SMA: α-smooth muscle actin, the myofibroblast activation marker. j mRNA levels of Col1a1, Col3a1 and Acta2 in the kidneys of Sham or UUO mice (n = 6). Data are presented as mean ± SD. The p values were determined by one-way ANOVA with Tukey’s correction (d, e, f, h, i, j). Source data are provided as a Source data file.
Furthermore, we injected AAV8-TBG-Cre to selectively ablate FGL1 in hepatocytes of adult FGL1fl/fl mice (Supplementary Fig. 3a, b). In the UUO model, FGL1 HepKO mice exhibited a marked reduction in renal fibrosis, as indicated by diminished collagen deposition and decreased expression of key fibrotic markers (Supplementary Fig. 3c–f). Notably, these renoprotective effects were also observed in female mice subjected to UUO (Supplementary Fig. 4). Additionally, FGL1 HepKO ameliorated adenine-induced renal fibrosis and improved kidney function in mice (Supplementary Fig. 5).
To complement loss-of-function studies, we employed a gain-of-function approach through hepatocyte AAV8-TBG-FGL1 overexpression (Fig. 3a and Supplementary Fig. 6a, b). In UUO model mice, hepatic FGL1 overexpression exacerbated tubular dilation and collagen deposition (Fig. 3b, c), paralleled by elevated COL-I and α-SMA levels via immunohistochemistry or immunofluorescence (Fig. 3d, e). FGL1 overexpression further upregulated fibrosis-related proteins and genes in the kidney (Fig. 3f, g).
a Schematic illustration of the experimental approach. To achieve liver-specific overexpression of FGL1, mice were injected with AAV8-TBG-FGL1 via the tail vein. Control mice were injected with AAV8-TBG-Ctrl. After 3 weeks, the mice were subjected to the unilateral ureteral obstruction (UUO) for 2 weeks. Created in BioRender. Wu, W. (2025) https://BioRender.com/2k53mzz. b, c Representative histological images of HE staining and Masson staining in the kidneys of UUO mice (n = 3, Scale bar, 100 μm and 50 μm). d, e Immunohistochemical staining of COL-I and immunofluorescence staining of α-SMA in the kidneys of UUO mice (n = 3, Scale bar, 100 μm and 50 μm). f Western blot analysis of FN, VIMENTIN and α-SMA in the kidneys of UUO mice (n = 6). g mRNA levels of Col1a1, Col3a1 Acta2, Tgfβ and Ctgf in the kidneys of UUO mice (n = 6). h Schematic illustration of the cell co-culture experimental approach. i, j Western blot analysis of FN, VIMENTIN and PAI-1 in HK-2 cells treated with conditional medium and TGF-β stimulation (n = 3). k Immunofluorescence staining of α-SMA in the HK-2 cells treated with conditional medium following TGF-β stimulation (n = 3, Scale bar, 20 μm). l, m Western blot analysis of FN, VIMENTIN and PAI-1 in HK-2 cells treated with recombinant human FGL1 protein following TGF-β stimulation (n = 3). Data are presented as mean ± SD. The p values were determined by a two-tailed t test (b, c, d, e, f, g, j, m). Source data are provided as a Source data file.
Additionally, we employed cellular models to characterize the role of FGL1 in liver-kidney communication. HEK-293T cells were transfected with FGL1 plasmids to generate FGL1-enriched conditioned medium (Fig. 3h). Co-culture of HK-2 cells with this medium further enhanced the TGF-β-induced upregulation of FN, VIMENTIN, and PAI-1, as determined by Western blot (Fig. 3i, j), and increased the levels of α-SMA, as shown by immunofluorescence (Fig. 3k). Recombinant human FGL1 (rhFGL1) protein produced similar effects in the presence of TGF-β stimulation (Fig. 3l, m). We further confirmed that FGL1 promotes the secretion (in culture supernatants) of FN, a result that directly supports its role in driving ECM accumulation (Supplementary Fig. 6c, d). Besides, treatment with rhFGL1 alone did not significantly upregulate the expression of fibrosis-associated genes (Supplementary Fig. 6e).
FGL1 interacts with ALK5 to prevent its ubiquitination
Upon treatment with FGL1-enriched conditioned medium from HEK-293T cells, FGL1 was predominantly localized to the plasma membrane fraction in HK-2 cells (Fig. 4a). Proteomic analysis revealed TGF-β/SMAD signaling pathway stood out as the most enriched biological process in the kidney after FGL1 overexpression (Fig. 4b–d). Next, we screened for direct interaction between FGL1 and members of the TGF-β receptor family. Surface plasmon resonance (SPR) results demonstrated that FGL1 selectively bound to ALK5 with a strong affinity (KD = 12.6 nM, Fig. 4e) but could not or was weakly bound to ALK1, ALK2, ALK3, ALK4, ALK6, ALK7, TGFβR2, BMPR2, AMHR2, ACVR2A, and ACVR2B (Supplementary Fig. 7). Bio-layer interferometry (BLI) further confirmed that FGL1 displayed favorable binding affinity to ALK5 (Fig. 4f). Bimolecular fluorescence complementation (BiFC) detected a direct interaction of FGL1 with ALK5 in HEK-293T cells (Fig. 4g). This interaction was confirmed by reciprocal co-immunoprecipitation (Co-IP) (Fig. 4h, i).
a Supernatants from FGL1-overexpressed HEK-293T cells were collected, filtered through a 0.22 μm sterile filter, and added to HK-2 cells. After 24 h of incubation, the distribution of FGL1 in the cytosol and plasma membrane of HK-2 cells was analyzed by Western blot (n = 3). b Enriched biological processes based on proteomic analysis in kidneys of UUO-AAV-FGL1 mice compared to UUO-AAV-Ctrl mice (n = 3, two-tailed Student’s t test). c Enrichment of TGF-β/SMAD signaling proteins based on proteomic (n = 3, two-tailed Student’s t test). NES, normalized enrichment score. d Heatmaps showing the upregulated proteins involved in the TGF-β/SMAD signaling pathway identified through renal proteomics (n = 3). e, f The interaction between FGL1 and ALK5 was tested using surface plasmon resonance (SPR) and bio-layer interferometry (BLI). The frequency response and fitting curves are displayed. g Bimolecular fluorescence complementation (BiFC) signals were detected in HEK-293T cells co-transfected with FGL1-VC155 and ALK5-VN173 plasmids (n = 3, scale bar, 10 μm). Representative fluorescence images are shown. h, i Co-immunoprecipitation (Co-IP) analysis demonstrating the interaction between FGL1 and ALK5 in HEK-293T cells (n = 3). j The interaction between FGL1 and various truncations of ALK5 was tested using SPR. k Molecular docking analysis revealed the binding domain between human FGL1 and ALK5. l Co-immunoprecipitation analysis revealed interactions between individual site-mutant FGL1 proteins and ALK5 (n = 3). m Co-immunoprecipitation analysis revealed interactions between individual site-mutant ALK5 proteins and FGL1 (n = 3). The image is one of three independent experiments. Data are presented as mean ± SD. The p values were determined by a two-tailed t test (d). *p < 0.05; **p < 0.01. Source data are provided as a Source data file.
To characterize the binding regions of ALK5 to FGL1, we generated their respective truncated forms. SPR (Fig. 4j) and Co-IP (Supplementary Fig. 8a) showed that the 200–503 amino acid region of ALK5 interacted with FGL1, but the 1–200 amino acid region could not. On the other hand, the 150–312 amino acid region of FGL1 could interact with ALK5, while the 1–150 amino acid region could not (Supplementary Fig. 8b). Molecular docking analysis predicted that 8 amino acids of FGL1 could form eight hydrogens with 6 amino acids of ALK5, i.e., FGL1E222 with ALK5W277, FGL1Q224 with ALK5E218, FGL1W225 with ALK5G214, FGL1D252 with ALK5R237, FGL1R260 with ALK5R237, FGL1D280 with ALK5R237, FGL1Y286 with ALK5T274, and FGL1W291 with ALK5T272 (Fig. 4k). Simultaneous mutation of the 8 amino acids of FGL1 diminished the interaction of FGL1 with ALK5 by Co-IP (Supplementary Fig. 8c). Single mutation of FGL1D252A, FGL1R260A, FGL1D280A, FGL1Y286A or FGL1W291A considerably disrupted the binding of FGL1 to ALK5, while the interactions of FGL1E222A, FGL1Q224A, and FGL1W225A remained their intact (Fig. 4l). Simultaneously mutation of the 6 amino acids of ALK5 diminished the interaction of ALK5 with FGL1 (Supplementary Fig. 8d). Single mutations of ALK5E218A, ALK5R237A, ALK5T272A, ALK5T274A or ALK5W277A considerably disrupted the binding of ALK5 to FGL1, while ALK5G214A remained unaffected (Fig. 4m).
Following FGL1 interaction with ALK5, hepatocyte-specific FGL1 knockout significantly suppressed TGF-β/SMAD signaling pathway in fibrotic kidneys of UUO mice, whereas FGL1 overexpression activated this pathway. Interestingly, we found that the protein level of ALK5 was also regulated by FGL1 (Fig. 5a, b). 3D imaging of ALK5 distribution demonstrated that FGL1 overexpression elevated ALK5 protein levels (Fig. 5c). In addition, administration of HK-2 cells with SB431542, a specific inhibitor of ALK5, blocked FGL1-induced profibrotic effect (Fig. 5d). To assess FGL1’s effect on ALK5 stability, we monitored ALK5 degradation in HEK-293T cells under basal conditions. ALK5 underwent rapid degradation (0–12 h), but FGL1 overexpression markedly stabilized the receptor (Fig. 5e). Cellular thermal shift assays demonstrated that FGL1 overexpression increased ALK5 stability at multiple temperatures (Fig. 5f). ALK5 degradation was blocked with the proteasome inhibitor MG132 but not the lysosome inhibitor chloroquine, suggesting the involvement of the proteasome pathway. (Supplementary Fig. 9). FGL1 overexpression decreased ALK5 ubiquitination, while mutations of binding sites in FGL1 (D252A, R260A, D280A, Y286A, and W291A) abolished this effect (Fig. 5g). Similarly, pretreatment with rhFGL1 reduced ALK5 ubiquitination (Fig. 5h).
a ALK5 and the downstream SMAD2/3 protein levels in kidneys of FGL1fl/fl (Ctrl) and FGL1 HepKO mice (n = 3). b ALK5 and the downstream SMAD2/3 protein levels in kidneys of AAV8-TBG-Ctrl and AAV8-TBG-FGL1 mice (n = 4). c 3D immunofluorescence image of the kidneys labeled with ALK5 (red fluorescence) (n = 3, Scale bar, 2 mm). d HK-2 cells were pre-treated with TGF-β for 1 h, then treated with rhFGL1 or rhFGL1 + SB431542 (a TGF-β receptor kinase inhibitor) for 24 h. Western blot analysis was performed to detect FN, VIMENTIN, and PAI-1 in HK-2 cells (n = 3). e The stability of ALK5 protein under FGL1 overexpression was assessed using a cycloheximide (CHX) chase assay in HEK-293T cells (n = 3 biological replicates). f Cellular thermal shift assay to assess ALK5 target engagement. Representative Western blot shows the protein levels of ALK5 in HEK-293T cells transfected with or without FGL1 plasmid following the indicated heat shocks (n = 3 biological replicates). g–i Immunoprecipitation analysis of ALK5 ubiquitination in HEK-293T cells stably expressing ALK5, transfected with the indicated constructs or treated with recombinant FGL1 protein (n = 3). j Immunoprecipitation of Flag-ALK5 using anti-Flag antibody in HEK-293T cells transfected with Flag-ALK5, Myc-SMAD7, and escalating doses of HA-FGL1 (n = 3). k Western blot analysis of HEK-293T cells transfected with Myc-SMAD7, Flag-ALK5 and increasing amounts of HA-FGL1 plasmids (n = 3). l The de-ubiquitination of Flag-ALK5 in response to Myc-SMAD7 overexpression was examined in HEK-293T cells transfected with wild-type and mutant ubiquitination (Ub) plasmids (n = 3). Data are presented as mean ± SD. The p values were determined by a two-tailed t test (b, e, f) or one-way ANOVA with Tukey’s correction (a, d). Source data are provided as a Source data file.
SMAD7 is a known negative regulator of TGF-β pathway, competitively binding ALK5 to promote receptor ubiquitination. In HEK-293T cells, the inhibitory effect of FGL1 on AKL5 ubiquitination was abolished by SMAD7 overexpression (Fig. 5i). Co-IP assays demonstrated that increasing FGL1 levels progressively attenuated the ALK5-SMAD7 interaction (Fig. 5j). Consistent with this, Western blot showed that FGL1 dose-dependently counteracted SMAD7-mediated ALK5 degradation (Fig. 5k). Screening of ubiquitin mutants (K6R, K11R, K27R, K29R, K33R, K48R, and K63R) revealed that SMAD7 specifically promotes K48-linked polyubiquitin interact with ALK5, as mutation of K48R abolished this ubiquitination (Fig. 5l).
STAT3 transcriptionally upregulates FGL1 expression in the liver
To delineate the transcriptional mechanism underlying FGL1 regulation, we screened potential transcription factors using the GTRD, KnockTF, ENCODE, and JASPAR databases and identified six candidates: STAT3, JUN, CREB1, GATA3, CEBPB, and MAX (Supplementary Fig. 10a). Subsequent qPCR analysis revealed significantly elevated Stat3 mRNA levels at day 1, 3, 7, and 14 in the liver of UUO mice (Supplementary Fig. 10b), which positively correlated with Fgl1 expression (r = 0.71, Supplementary Fig. 10c). To validate STAT3’s regulatory role, we performed STAT3 gradient overexpression experiments in HEK-293T cells, demonstrating a dose-dependent enhancement of FGL1 promoter activity (Supplementary Fig. 10d). RNA sequencing showed that inflammatory response stood out as one of the most enriched biological processes in the liver of UUO mice (Supplementary Fig. 10e). ELISA revealed significant induction of inflammatory cytokines such as IL-6, IL-1β, and TNF-α in the serum (Supplementary Fig. 10f). These inflammatory cytokines exhibited a positive correlation with circulating FGL1 protein levels (Supplementary Fig. 10g). In vitro experiments using AML12 hepatocytes demonstrated that individual treatment with IL-6, IL-1β, and TNF-α upregulated STAT3 phosphorylation and FGL1 protein levels (Supplementary Fig. 10h, i). Critically, STAT3 inhibitor (S3I-201) abolished inflammatory cytokines-induced FGL1 production, confirming STAT3’s essential role in FGL1 regulation (Supplementary Fig. 10j).
High-throughput screening identifies FGL1 inhibitors with therapeutic potential in renal fibrosis
Based on the promising therapeutic potential of inhibiting FGL1 for the treatment of renal fibrosis, we performed a luciferase reporter-based high-throughput screening in HEK-293T cells to identify FGL1 inhibitors (Fig. 6a). A structurally diverse library of 4080 natural small-molecule compounds was evaluated (Supplementary Data 2). Initial selection of the top 30 candidates was based on their ability to suppress FGL1 promoter activity by >2-fold (Fig. 6b). Further refinement excluded compounds with unfavorable ADMET (absorption, distribution, metabolism, excretion, and toxicity) profiles or cytotoxicity in HEK-293T and AML12 cells (Fig. 6c and Supplementary Fig. 11a, b), narrowing the list to 10 candidates: coumarin VI, glucose, leucic acid, atractyloside A, trichostatin A, anemoside A3, mulberroside C, primin, paradol, and p-hydroxybenzaldehyde (PHBA).
a Schematic illustration of the screening process to identify effective down-regulators of FGL1 transcription. b Luciferase assay and scatter plot showing the effects of 4080 natural compounds on FGL1 transcription in HEK-293T cells. c Screening criteria used to identify the most effective compounds for FGL1 down-regulation. d Western blot analysis of FGL1 expression in hepatocytes treated with the indicated small-molecule compounds or DMSO. Screening criteria used to identify the most effective compounds for FGL1 down-regulation (n = 4). e mRNA level of Fgl1 in hepatocytes (n = 6). f CCK8 assay to assess AML12 cell viability following treatment with PHBA. PHBA: p-hydroxybenzaldehyde (n = 5 biological replicates). g Luciferase assay to evaluate the effect of PHBA on FGL1 promoter activity in HEK-293T cells (n = 3 biological replicates). h mRNA levels of Fgl1 in hepatocytes treated with different concentrations of PHBA for 24 h (n = 6). i Schematic illustration of the experimental approach. Mice were subjected to unilateral ureteral obstruction (UUO), and the treatment groups were gavaged with 10 or 50 mg/kg PHBA for 14 days. Created in BioRender. Wu, W. (2025) https://BioRender.com/g08o8ga. j mRNA levels of Fgl1 in the livers of UUO mice treated with PHBA or Saline (n = 6). k ELISA analysis of the FGL1 levels in serum of UUO mice (n = 6). l Representative images of HE staining and Masson staining of renal sections of UUO mice (n = 3, Scale bar, 100 μm). Data are presented as mean ± SD. The p values were determined by one-way ANOVA with Tukey’s correction (d, e, f, g, h, j, k, l). Source data are provided as a Source data file.
Among the 10 candidates, PHBA demonstrated the strongest reduction of FGL1 mRNA and protein levels (Fig. 6d, e). PHBA (5, 10, 20, 40 μM) dose-dependently inhibited both FGL1 promoter activity and FGL1 protein expression without hepatocyte toxicity (Fig. 6f–h and Supplementary Fig. 11c). p-Hydroxybenzyl alcohol and p-hydroxybenzonic acid, two structural analogs of PHBA, showed no such activity, underscoring PHBA’s specificity (Supplementary Fig. 11d). In UUO mice, oral administration of PHBA (10, 50 mg/kg) effectively reduced Fgl1 mRNA in the liver (Fig. 6i, j) and FGL1 protein levels in serum (Fig. 6k). Consequently, PHBA attenuated the renal fibrosis phenotype (Fig. 6l) and significantly suppressed the expression of fibrosis-related proteins and genes (Supplementary Fig. 12a, b). Additionally, PHBA was relatively safe in observed time, as evidenced by stable ALT/AST levels and intact liver/heart histology (Supplementary Fig. 12c, d).
In parallel, a monoclonal antibody against FGL1 (anti-FGL1 mAb) was evaluated. Intraperitoneal administration of anti-FGL1 mAb in UUO mice reduced serum FGL1 levels (Fig. 7a), alleviated renal injury and collagen deposition, as evidenced by morphological observations, HE and Masson staining (Fig. 7b, c). Anti-FGL1 mAb also suppressed fibrosis-associated proteins and genes in the kidney (Fig. 7d, e).
a Schematic illustration of the experimental approach. Mice were subjected to the unilateral ureteral obstruction (UUO) and injected with IgG or anti-FGL1 mAb (100 μg, every 3 days) for 14 days (left). ELISA analysis of the FGL1 levels in the serum of Sham or UUO mice (n = 6, right). b Gross appearance of whole kidneys from different groups of Sham or UUO mice (n = 3, Scale bar, 2 mm). c Representative images of HE staining and Masson staining of renal sections of Sham or UUO mice (n = 3, Scale bar, 100 μm). d Western blot analysis of FN, VIMENTIN and α-SMA in the kidney of sham or UUO mice (n = 6). e mRNA levels of Col1a1, Col3a1 and Tgfβ in the kidney of Sham or UUO mice (n = 6). f FGL1 acts as a critical mediator in liver-kidney communication. Circulating renal injury-associated cytokines (IL-6, IL-1β, TNF-α) drive STAT3 activation in the liver, which subsequently enhances hepatic FGL1 transcription and secretion. FGL1 accumulates selectively in fibrotic kidneys, binds TGF-β receptor ALK5 to block its interaction with K48-linked polyubiquitin. Pharmacological targeting of FGL1, using either small-molecule inhibitors (e.g., PHBA) or anti-FGL1 monoclonal antibodies, demonstrates therapeutic potential for CKD by disrupting the interorgan-signaling network. Created in BioRender. Wu, W. (2025) https://BioRender.com/obfr22s. Data are presented as mean ± SD. The p values were determined by one-way ANOVA with Tukey’s correction (a, c, d, e). Source data are provided as a Source data file.
Discussion
This study has identified FGL1 as a pivotal hepatokine driving pathogenic liver-kidney crosstalk in renal fibrosis. Renal injury triggers systemic inflammation and activates STAT3 to transcriptionally upregulate hepatic FGL1. Notably, FGL1 accumulates selectively in fibrotic kidneys, where it binds to ALK5 to stabilize TGF-β signaling by blocking receptor ubiquitination. Pharmacologically targeting of FGL1 via small-molecule inhibitors or anti-FGL1 monoclonal antibodies by disrupting interorgan networks could constitute a glimmer of light towards the treatment of CKD (Fig. 7f).
The role of the liver as an endocrine organ has gained clinical relevance. It is able to synthesize and release hepatokines that have the capacity to act at a distance in other organs such as the heart5, adipose tissue20, skeletal muscle21, bone7 and the central nervous system22. However, clinical evidence regarding the liver-kidney axis remains limited, though existing data underscore significant comorbidity between the hepatic and renal systems. Hepatorenal syndrome exemplifies a severe renal complication of advanced liver disease23,24, and nonalcoholic fatty liver disease has been established as a risk factor for CKD25,26. Our study advances the understanding of the liver-kidney axis by revealing a pathogenic feedback loop—renal injury initiates the secretion of hepatic FGL1 via circulating inflammatory mediators. The FGL1 released subsequently reaches the injured kidneys, where it exacerbates renal fibrosis, ultimately forming a deleterious cross-organ pathological circuit. We hypothesize that this phenomenon may be a self-protective mechanism of the liver in response to stressors such as inflammation—wherein upregulation of FGL1 secretion helps the liver counteract external injurious factors and facilitates timely restoration of organ function. This hypothesis is supported by the positive correlation between circulating inflammatory factors and FGL1 levels, which is consistent with reported findings that extrahepatic inflammation elevates plasma FGL127. In addition, this interpretation aligns with previous observations that hepatic FGL1 secretion under stress protects mice and nonhuman primates from acute liver injury28. In inter-organ communication, hepatokines mediate the liver’s dual roles, thus supporting repair as a “rescuer” or, in disease, propagate a “shared fate” of systemic damage5,29,30. Beyond these established roles, we found that the liver might act as an inadvertent “guardian” and exacerbate injury in organ-organ crosstalk via hepatokines—a portrayal that echoes the colloquial expression “caught in the crossfire”. Apart from the liver-kidney axis, such a role of hepatokines in other organs has not been reported and warrants further investigation.
Beyond its recognized role in liver regeneration, FGL1 has recently been identified as a functional ligand of LAG3 and implicated in immune checkpoint regulation31. Given the established contribution of immune cells to renal injury, we sought to determine whether the FGL1-LAG3 axis participates in renal fibrosis. Our data provide no substantial evidence that circulating FGL1 exacerbates fibrosis via LAG3 engagement on renal immune cells (Supplementary Figs. 13 and 14). Instead, we have identified a novel pathway wherein hepatocyte-derived FGL1 promotes fibrosis by binding to ALK5. Although rhFGL1 alone did not induce fibrosis-related gene expression, it potently amplified TGF-β–driven fibrotic responses (Supplementary Fig. 6e). Mechanistically, FGL1 interacts with the 218–277 domain of ALK5, stabilize the receptor against proteasomal degradation, and thereby increasing its availability for TGF-β signaling. This synergy arises from FGL1 and TGF-β engaging distinct regions of ALK5, rather than competing for the same binding site, leading to enhanced activation of the pro-fibrosis pathway. Notably, FGL1 exhibits context-dependent functional duality: its overexpression under basal conditions does not induce injury (Supplementary Fig. 15), whereas in experimental injury models such as UUO, it acts as a pathogenic amplifier that markedly exacerbates fibrosis progression.
Current therapies for CKD include ACE inhibitors, angiotensin receptor blockers, and SGLT2 inhibitors32. Our work introduces hepatic FGL1 inhibition as a paradigm shift in inter-organ crosstalk disruption. The natural product PHBA reduced renal FGL1 deposition and fibrosis without inducing hepatotoxicity. In addition, we started PHBA treatment 4 days after UUO for 2 weeks to mimic a scenario in the clinic. PHBA significantly ameliorated renal fibrosis phenotype (Supplementary Fig. 16), suggesting potential efficacy against established CKD. Compared to small-molecule agents, anti-FGL1 monoclonal antibodies offer distinct advantages, including superior target specificity and prolonged therapeutic effects. Drug repositioning accelerates therapeutic development by leveraging existing agents. Aspirin, a cost-effectively agent shown to promote FGL1 degradation in hepatocellular carcinoma33, reduced FGL1 protein levels in hepatocytes and in the serum of UUO mice. It also preserved renal structural integrity and attenuated renal fibrosis (Supplementary Figs. 17 and 18). However, a clinical trial observed no benefit or harm from low-dose aspirin in CKD patients34. Given reported associations between aspirin and renal damage35,36, the long-term safety of FGL1-targeted therapies requires evaluation. Further validation in additional animal models is needed to assess the anti-renal fibrosis potential of aspirin, particularly its dosage regimen and safety profile.
Our study has several limitations. First, while both surgical and chemical models were used, UUO does not replicate the functional decline of human CKD37,38. Future work should include the 5/6 nephrectomy model to improve clinical relevance39. Second, the effects of FGL1 on other renal cell types remain unclear, necessitating further investigation to define its pathogenic role. Finally, AAV8-mediated FGL1 overexpression is constrained by uncontrolled expression. Liver-specific knock-in models should be used to achieve tightly regulated, spatiotemporally controlled expression.
In conclusion, our findings redefine FGL1 as a linchpin in liver-kidney communication, bridging mechanistic insight with translational potential. By elucidating how organ-organ dialog perpetuates chronic disease, this work advocates for therapies targeting interorgan networks and moves beyond traditional organ-centric frameworks.
Method
Human subject recruitment study participants
All procedures were approved by the Medical Ethics Committee of Jiangsu Province Hospital and adhered to the principles of the Declaration of Helsinki (2024-SR-299). All participants were informed about the use of their serum, and written informed consent was obtained. Serum from 18 CKD patients and 18 age- and sex-matched healthy controls were collected from Jiangsu Province Hospital (Supplementary Table 1). CKD patients with a glomerular filtration rate of less than 60 mL/min per 1.73 m2, or a marker of kidney damage, or both, for a duration of at least 3 months were recruited. Patients with acute kidney injury, liver disease, active vasculitis, gastrointestinal pathology or cancer were excluded from the study. Age- and sex-matched healthy subjects were also recruited.
Animal treatment
Animal experiments were conducted in accordance with the guidelines for Animal Experimentation of China Pharmaceutical University (Nanjing, China), and the protocols were approved by the Animal Ethics Committee of the institution (No: 2024-11-092). C57BL/6 J mice (8 weeks, 20–23 g) were purchased from Jiangsu GemPharmatech Biotechnology (Nanjing, China) and randomly assigned to different treatment groups. All animals were housed under controlled humidity and temperature in standard facilities (24–26 °C) under specific pathogen-free conditions with a 12 h dark/light cycle (lights on between 8 a.m. and 8 p.m.), with ad libitum access to water and chow (Xie Tong Bio-Technology Co., Ltd., China. Lot No. XTI01WC-011). Mice were euthanized by cervical dislocation. The investigators were not blinded to the treatment groups, but no subjective assessments were made.
UUO-induced renal fibrosis model
Mouse models of renal fibrosis were established by procedures for UUO40. Following sodium pentobarbital anesthesia (30 mg/kg, i.p.), the left ureter was double-ligated using 5-0 silk via dorsal incision. Sham controls underwent identical surgical exposure without ligation.
Adenine-induced renal fibrosis model
Experimental groups received daily oral gavage of adenine suspension (80 mg/kg) for 3 weeks, while controls received equivalent saline volumes41.
p-Hydroxybenzaldehyde (PHBA) and aspirin (ASP) treatment in vivo
Drug dosages were determined through interspecies conversion and prior pharmacokinetic studies42. The mice were administered with PHBA or ASP at 10 and 50 mg/kg/day. In UUO mice, the treatment groups were administered the appropriate drugs via oral gavage for 14 consecutive days post-surgery. In addition, PHBA treatment was initiated 4 days after UUO and continued for 2 weeks to mimic the clinic scenario.
Hepatocyte-specific FGL1 knockout mice
To specifically knockout FGL1 in hepatocytes, we produced a mouse line by crossing female FGL1-floxed mice (FGL1fl/fl) with male Alb-Cre transgenic mice (Gempharmatech Co., Ltd, 8–9 weeks old). The F1 progeny, either male or female, with heterozygous deletion of the FGL1 gene (FGL1fl/-) and carrying the Alb-Cre transgene (FGL1fl/-, Alb-Cre) were obtained. These mice were then crossed with FGL1fl/fl mice of opposite sex to generate the F2 progeny, which expressed complete deletion of FGL1 in hepatocytes (FGL1fl/fl, Alb-Cre), herein referred to as FGL1 HepKO mice. Genotype verification was performed through tail biopsy PCR analysis.
AAV-mediated hepatic FGL1 knockout in mice
For hepatic FGL1 knockout, AAV8-mediated delivery of Cre recombinase was employed in C57BL/6J mice. Each FGL1fl/fl mouse received a single tail vein injection of 7 μL of AAV8-TBG-Cre virus suspension (virus titer >1013 vg/mL, Hanbio Biotech Co., Ltd) blended with 193 μL saline. Control mice were injected with AAV8-TBG-Ctrl. Three weeks after injection, mice were sacrificed, and liver tissues were collected for analysis. The knockout efficiency was confirmed by qPCR.
AAV-mediated hepatic FGL1 overexpression in mice
For hepatic FGL1 reconstitution, AAV8-mediated delivery was employed in C57BL/6J mice. Each mouse received a single tail vein injection of 7 μL of AAV8-TBG-FGL1-EGFP virus suspension (virus titer >1013 vg/mL, PackGene Biotech Co., Ltd) blended with 193 μL saline. Control mice were injected with AAV8-TBG-Ctrl. Hepatic transduction efficiency was confirmed by Western blot after 3 weeks of injection.
Neutralizing antibody against FGL1 in mice
Mice were stratified into four groups: (1) Sham+IgG (100 μg); (2) Sham+Anti-FGL1 mAb (100 μg); (3) UUO+IgG (100 μg); (4) UUO+Anti-FGL1 mAb (100 μg). Intraperitoneal injections were initiated 24 h post-surgery and repeated on days 4, 8, and 12 after surgery. Euthanasia was performed on day 14 via cervical dislocation under anesthesia.
RNA-seq analysis
Mice livers were preserved in RNAlater and sent to Majorbio (Shanghai, China) for sequencing. After RNA quality check, a sequencing library was created using TruSeq technology. Sequencing was performed on a HiSeq X Ten (Illumina). Differentially expressed genes were identified based on an adjusted p < 0.05. For heatmap visualization of differentially expressed genes, the data were subjected to z-score normalization.
Single-cell RNA-seq analysis
The gene expression levels of Fgl1, Orm2, Mt2, Saa1, and Saa2 in various cell types of liver or kidney were analyzed using published scRNA-seq data43,44. The scRNA-seq data were obtained from the Gene Expression Omnibus dataset (GSE115469, GSE131685).
Cell culture
Human renal proximal tubular epithelial cells (HK-2), human embryonic kidney cells (HEK-293T) and mouse hepatocytes (AML12) were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China; STR-authenticated). Cells were maintained in DMEM/F12 (HK-2) or DMEM(HEK-293T/AML12) supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin at 37 °C/5% CO2. HK-2 cells were exposed to 10 ng/mL TGF-β and/or 400 ng/mL FGL1; AML12 cells were exposed to IL-6, IL-1β, and TNF-α (25 ng/mL each).
Cytotoxicity assessment
Cell Counting Kit-8 (CCK8) was used to analyze the cell viability of AML12 cells or HEK-293T cells. Briefly, AML12 or HEK-293T cells were treated with varying concentrations of the candidate compounds for 24 h. Then, the CCK8 solution was added, and the cells were stored in the incubator for 2 h. The optical density of each well was measured at 450 nm using a microplate reader.
ELISA quantification
The sera of humans and mice were collected by centrifuging at 2000 × g for 10 min. The levels of FGL1 in the different experimental groups were determined using the human or mouse FGL1 ELISA kits according to the manufacturer’s instructions. The levels of IL-6, IL-1β and TNF-α were determined using ELISA kits. Serum ALT and AST activities were measured using commercial kits. The levels of blood urea nitrogen and creatinine were determined according to the manufacturer’s instructions.
Histological analysis
Kidney, liver and heart tissues were fixed in 4% paraformaldehyde (PFA) and embedded in paraffin wax. For histological evaluation, sections were stained with Hematoxylin and Eosin (HE) to assess general tissue morphology. To evaluate collagen deposition, sections were stained using Masson’s trichrome stain. Fibrotic areas were quantified using ImageJ software. Image acquisition was performed with a NanoZoomer RS scanner (Hamamatsu, Japan).
Three-dimensional (3D) visualization of the kidneys
Mice were surgically anesthetized with 2% isoflurane and perfused with 20 mL of PBS followed by 20 mL of 4% PFA solution. Kidneys were rapidly dissected, fixed in 4% PFA at 4 °C for 24 h, followed by 1 h post-fixation at room temperature. Kidney tissues were stored in PBS at 4 °C for subsequent processing. Tissue clearing was performed using the Enhanced Tissue Clearing Kit according to the manufacturer’s protocol. Cleared kidneys were mounted in a refractive index-matched imaging chamber and subjected to light-sheet microscopy using the LS18 system (Nuohai Life Science, China) equipped with a 6.3× objective. For whole-organ imaging, the 562 nm laser power was maintained at 50% intensity. Three-dimensional scanning was achieved through multiplane acquisition (X/Y/Z axes) with 2 μm isotropic resolution. Raw image stacks were preprocessed using Combine Software (Nuohai Life Science, China) for background subtraction and noise reduction. Subsequent 3D reconstruction was performed with Imaris x64 software (Bitplane, Switzerland) using surface rendering algorithms. Orthogonal projections were generated at 90° intervals to demonstrate spatial relationships within the kidney architecture.
Immunofluorescence staining
For kidney tissue analysis, cryosections were incubated with primary antibodies in a humidified dark chamber at 4 °C for 16 h. After PBS washes (3 × 5 min), sections were incubated with species-matched secondary antibodies (1:200) for 1 h, followed by 10 min DAPI nuclear counterstaining. In vitro experiments using HK-2 cells were conducted as follows: cells were fixed with 4% PFA for 15 min, permeabilized with 0.2% Triton X-100 for 10 min, and blocked with 5% BSA for 1 h. Primary antibody incubation was performed overnight at 4 °C, followed by 2 h secondary antibody treatment and DAPI staining for 10 min. All images were acquired using an FV3000 confocal microscope (Olympus, Japan) with a 60× oil-immersion objective.
Immunohistochemical staining
Formalin-fixed paraffin-embedded kidney sections (4 μm thickness) were deparaffinized in xylene (3 × 5 min) and rehydrated through a graded ethanol series (100–70%). Antigen retrieval was performed in 10 mM citrate buffer (pH 6.0) at 95 °C for 20 min. Following endogenous peroxidase quenching with 3% H2O2, sections were incubated with primary antibodies for 1 h at 37 °C. Whole-slide digital imaging was conducted using the NanoZoomer RS scanner (Hamamatsu, Japan) at 40 × magnification.
Western blot analysis
Tissues and cells were harvested and immediately lysed in RIPA buffer supplemented with protease and phosphatase inhibitors to prevent protein degradation and dephosphorylation. Lysates were incubated on ice for 30 min to ensure complete lysis. Cell debris was removed by centrifugation at 12,000 × g for 10 min at 4 °C, and the resulting supernatant (total protein extract) was collected and stored at −80 °C until further use. Protein concentration was quantified using the BCA Protein Assay Kit following the manufacturer’s instructions. The immunoblotting procedures were performed as previously described45. Antibodies used in this study are listed in Supplementary Table 2. Semi-quantitative analysis of protein bands was conducted using ImageJ software (v1.8.0), with band densities normalized to the housekeeping proteins GAPDH or β-TUBULIN for loading control.
Real-time fluorescence quantitative PCR (RT-qPCR)
Total mRNA was isolated from tissues or cells using a High Pure RNA Isolation Kit, with RNA concentration and purity verified by NanoDrop spectrophotometry (A260/A280 ratio 1.8–2.0). Reverse transcription was performed with cDNA Synthesis SuperMix according to the manufacturer’s protocol. Relative expression levels of target genes were calculated using the 2−ΔΔCt method, normalized to Actb as the endogenous control. Primer sets for genes are summarized in Supplementary Table 3.
Label-free quantitative proteomics analysis
Kidney tissues from UUO-AAV-Ctrl mice (n = 3, negative control) and UUO-AAV-FGL1 mice (n = 3, experimental group) were snap-frozen in liquid nitrogen, pulverized, and homogenized in SDS-based lysis buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1% SDS, protease/phosphatase inhibitors) with ultrasonication (400 W, 10 cycles). Lysates were denatured (95 °C, 10 min), centrifuged (16,000 × g, 20 min, 4 °C), and quantified via tryptophan fluorescence assay46.
Each biological sample was processed in technical duplicate (n = 2). Aliquots of 100 μg protein were subjected to FASP: reduced with DTT, alkylated with IAA, and digested with trypsin (50:1, 37 °C, 16 h). Peptides were desalted and analyzed on an EASY-nLC 1200 UHPLC-Q Exactive Plus Orbitrap MS (Thermo Fisher Scientific, USA) with a C18 column (75 μm × 25 cm, 1.9 μm). LC gradient: 2–35% acetonitrile (0.1% formic acid) over 90 min (300 nL/min, 50 °C). MS acquisition: DDA mode (m/z 350-1800, resolution 70/17.5 k for full/ms² scans, HCD collision energy 27%, dynamic exclusion 30 s).
A pooled QC sample was analyzed at regular intervals (RSD for retention time <2%, peak intensity <15%). Raw data were processed with MaxQuant (v2.1.6.0) against UniProt Mus musculus database (UP000000589), with trypsin (2 missed cleavages), carbamidomethylation (Cys), oxidation (Met)/acetylation (N-term) as modifications, and FDR < 1%. Differentially expressed proteins (DEPs: FC > 1.2 or FC < 0.83, adjusted p < 0.05) were analyzed via R packages (limma v3.54.0, clusterProfiler v4.6.2) with Benjamini-Hochberg correction and Fisher’s exact test. Detailed parameters are in Supplementary Data 1.
Prediction of FGL1–ALK5 interactions
The AlphaFold3-predicted structures of the FGL1 and ALK5 were used as the initial structures for the MD simulations. The protein was first solvated with a regular cubic SPC water box with an 8.0 Å minimum distance to the box surface. To keep the whole system neutral, 7 Na+ ions were added to the solvent. The protein was parameterized using the ff14SB force fields. The resulting topology and coordinate files were generated using the tleap module in Amber20. All simulations were conducted using the pmemd.CUDA tool in Amber20.
Plasmids and transfection
HA-FGL1, Flag-ALK5, Myc-Smad7 and Ubiquitin plasmids were generated by cloning the indicated genes into pcDNA3.1 (PPL Co., Ltd, China). Site-directed mutagenesis was generated using a QuickMutation Site-Directed Mutagenesis Kit according to the manufacturer’s instructions, and plasmids were transfected with Lipofectamine 2000 according to the manufacturer’s protocol.
Bimolecular fluorescence complementation (BiFC)
Reconstitution of Venus fluorescent signal was performed as follows: HEK-293T cells seeded in 35-mm imaging dishes were co-transfected with pBiFC-FGL1-VC155 (1 μg) +pBiFC-ALK5-VN173 (1 μg). Negative control groups received either pBiFC-FGL1-VC155 + VN173 vector or pBiFC-ALK5-VN173 + VC155 vector. At 24 h post-transfection, live cells were stained with 5 μg/mL Hoechst 33342 for 15 min. The fluorescent signals were captured using an FV3000 confocal microscope (Olympus, Japan) with a 60× oil objective, employing 405 nm (Hoechst) and 515 nm (Venus) excitation wavelengths.
Co-immunoprecipitation assay
FGL1-ALK5 interaction was investigated through Co-IP using the crosslinker-enhanced protocol. Cells were lysed in 500 μL IP buffer supplemented with protease inhibitor cocktail. Lysates were clarified by centrifugation at 12,000 × g for 10 min at 4 °C, followed by protein quantification using BCA assay. For each immunoprecipitation, 400 μg total protein was pre-cleared with 20 μL protein A/G magnetic beads for 1 h at 4 °C. Pre-cleared lysates were incubated with 2 μg of specific antibodies or species-matched IgG control overnight at 4 °C with gentle rotation. Immune complexes were captured by adding 25 μL pre-washed magnetic beads (50% slurry) for 6 h at 4 °C. After five washes with PBS, bound proteins were eluted at 95 °C for 5 min and analyzed by immunoblotting.
Bio-layer interferometry (BLI)
Real-time binding kinetics were quantified using the Octet RED96e system (Sartorius, Germany). His-tagged FGL1 was loaded onto Ni-NTA biosensors to achieve 1 nm wavelength shift. After baseline stabilization in kinetics buffer (PBS + 0.01% Tween 20 + 0.1% BSA), sensors were exposed to serially diluted ALK5 (3.9–125 nM) for the association phase (150 s), followed by dissociation monitoring in analyte-free buffer (60 s). Reference subtraction was performed using buffer-only sensors and blank-loaded biosensors. Binding curves were globally fitted to a 1:1 Langmuir model using Octet Analysis Studio to obtain corresponding kinetic parameters47.
Surface plasmon resonance (SPR)
Detailed binding thermodynamics were characterized using Biacore T200 (Cytiva, USA). CM5 sensor chips were activated with EDC/NHS (1:1) for 7 min at 10 μL/min. FGL1 in 10 mM sodium acetate (pH 4.0) was immobilized to achieve 8500 RU, followed by ethanolamine blocking. ALK5, ALK51-125 and ALK5200-503 in HBS-EP were injected over the chip surface at 30 μL/min using multicycle kinetics. Concentration series (0.39–50 nM) were analyzed at contact time 60 s and dissociation 30 s. Data were processed using standard double-referencing and fitted to a 1:1 binding model using Biacore T200 Evaluation software (Cytiva, USA). The binding curves were analyzed to determine the equilibrium dissociation constant (KD).
High-throughput screening
A library of 4080 natural compounds was screened using HEK-293T cells. These cells were transfected with 10 μg of the pGL3-FGL1-promoter luciferase plasmid (reporter plasmid) and 1 μg of the pRL-TK-promoter luciferase plasmid (Renilla luciferase, used as an internal control). Following 24 h of transfection, cells were digested and reseeded into 384-well plates. After 24 h, natural compounds (10 μM) were subsequently added to the wells and incubated for another 24 h. Luciferase activity was quantified using the Dual-Glo Luciferase Assay Kit and detected by Envision Microplate System (PerkinElmer, USA).
Statistical analysis
Data are presented as mean ± standard deviation (SD). Multiple group comparisons were analyzed by one-way ANOVA with Tukey’s correction, while pairwise comparisons were assessed using two-tailed Student’s t tests. Statistical analyses were conducted using Prism 9.0 Software. Significance was defined as p < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data for Figs. 1–7 and Supplementary Figs. 1–18 have been provided as Source Datafiles. All data supporting the findings described in this manuscript are available in the article and in the Supplementary Information and from the corresponding author upon request. The RNA sequence data generated in this study have been deposited in the NCBI database under accession code PRJNA1254202. The scRNA-seq data were obtained from Gene Expression Omnibus datasets GSE115469 and GSE131685. The mass spectrometry proteomics generated in this study have been deposited in the ProteomeXchange Consortium under accession code PXD064971 [https://www.iprox.cn//page/project.html?id=IPX0012283000]. Source data are provided with this paper.
References
Friedman, J. M. & Halaas, J. L. Leptin and the regulation of body weight in mammals. Nature 395, 763–770 (1998).
Meex, R. C. R. & Watt, M. J. Hepatokines: linking nonalcoholic fatty liver disease and insulin resistance. Nat. Rev. Endocrinol. 13, 508–520 (2017).
Pedersen, B. K. & Febbraio, M. A. Muscles, exercise and obesity: skeletal muscle as a secretory organ. Nat. Rev. Endocrinol. 8, 457–465 (2012).
Ouchi, N., Parker, J. L., Lugus, J. J. & Walsh, K. Adipokines in inflammation and metabolic disease. Nat. Rev. Immunol. 11, 85–97 (2011).
Cao, Y. et al. Liver-heart cross-talk mediated by coagulation factor XI protects against heart failure. Science 377, 1399–1406 (2022).
Huynh, P. et al. Myocardial infarction augments sleep to limit cardiac inflammation and damage. Nature 635, 168–177 (2024).
Lin, L. S. et al. SIRT2 regulates extracellular vesicle-mediated liver-bone communication. Nat. Metab. 5, 821–841 (2023).
Tilg, H., Adolph, T. E. & Trauner, M. Gut-liver axis: pathophysiological concepts and clinical implications. Cell Metab. 34, 1700–1718 (2022).
Kerkelä, R., Ulvila, J. & Magga, J. Natriuretic peptides in the regulation of cardiovascular physiology and metabolic events. J. Am. Heart Assoc. 4, e002423 (2015).
Han, Y. et al. Membrane-delimited signaling and cytosolic action of MG53 preserve hepatocyte integrity during drug-induced liver injury. J. Hepatol. 76, 558–567 (2022).
Chen, Q. Q. et al. Neuraminidase 1 promotes renal fibrosis development in male mice. Nat. Commun. 14, 1713 (2023).
Liu, Y. H. Cellular and molecular mechanisms of renal fibrosis. Nat. Rev. Nephrol. 7, 684–696 (2011).
Meng, X. M., Nikolic-Paterson, D. J. & Lan, H. Y. TGF-β: the master regulator of fibrosis. Nat. Rev. Nephrol. 12, 325–338 (2016).
Shi, Y. G. & Massagué, J. Mechanisms of TGF-β signaling from cell membrane to the nucleus. Cell 113, 685–700 (2003).
Jensen-Cody, S. O. & Potthoff, M. J. Hepatokines and metabolism: deciphering communication from the liver. Mol. Metab. 44, 101138 (2021).
Watt, M. J., Miotto, P. M., De Nardo, W. & Montgomery, M. K. The liver as an endocrine organ-linking NAFLD and insulin resistance. Endocr. Rev. 40, 1367–1393 (2019).
Conklin, D. et al. Identification of a mammalian angiopoietin-related protein expressed specifically in the liver. Genomics 62, 477–482 (1999).
Pose, E., Piano, S., Juanola, A. & Gines, P. Hepatorenal syndrome in cirrhosis. Gastroenterology 166, 588–604 (2024).
Varga, Z. V. et al. Disruption of renal arginine metabolism promotes kidney injury in hepatorenal syndrome in mice. Hepatology 68, 1519–1533 (2018).
Gong, X. M. et al. Gpnmb secreted from the liver promotes lipogenesis in white adipose tissue and aggravates obesity and insulin resistance. Nat. Metab. 1, 570–583 (2019).
Niu, Y. et al. Hepatokine ERAP1 disturbs skeletal muscle insulin sensitivity via inhibiting USP33-mediated ADRB2 deubiquitination. Diabetes 71, 921–933 (2022).
Rose, J. P. et al. FGF21 reverses MASH through coordinated actions on the CNS and liver. Cell Metab. 37, 1515–1529 (2025).
Simonetto, D. A. et al. Hepatorenal syndrome: pathophysiology, diagnosis, and management. BMJ 370, m2687 (2020).
Jourde-Chiche, N. & Burtey, S. Accumulation of protein-bound uremic toxins: the kidney remains the leading culprit in the gut-liver-kidney axis. Kidney Int. 97, 1102–1104 (2020).
Umbro, I., Baratta, F., Angelico, F. & Del Ben, M. Nonalcoholic fatty liver disease and the kidney: a review. Biomedicines 9, 1370 (2021).
Canbakan, M. et al. The liver-kidney axis: Is serum leptin a potential link in non-alcoholic fatty liver disease-associated chronic kidney disease? Arab J. Gastroenterol. 24, 52–57 (2023).
Liu, Z. & Ukomadu, C. Fibrinogen-like protein 1, a hepatocyte-derived protein, is an acute phase reactant. Biochem. Biophys. Res. Commun. 365, 729–734 (2008).
Yang, Y. et al. Recombinant human HPS protects mice and nonhuman primates from acute liver injury. Int. J. Mol. Sci. 22, 12886 (2021).
Xing, W. et al. Upregulated hepatokine fetuin B aggravates myocardial ischemia/reperfusion injury through inhibiting insulin signaling in diabetic mice. J. Mol. Cell. Cardiol. 151, 163–172 (2021).
Li, Z. et al. Association of Fetuin-B with subclinical atherosclerosis in obese Chinese adults. J. Atheroscler. Thromb. 27, 418–428 (2020).
Wang, J. et al. Fibrinogen-like protein 1 is a major immune inhibitory ligand of LAG3. Cell 176, 334–347 (2019).
Heerspink, H. J. L. et al. Dapagliflozin in patients with chronic kidney disease. N. Engl. J. Med. 383, 1436–1446 (2020).
Lin, M. et al. Targeting fibrinogen-like protein 1 enhances immunotherapy in hepatocellular carcinoma. J. Clin. Investig. 133, e164528 (2023).
Taliercio, J. J. et al. Aspirin for primary and secondary prevention of mortality, cardiovascular disease, and kidney failure in the chronic renal insufficiency cohort (CRIC) study. Kidney Med. 4, 100547 (2022).
Juhlin, T., Jönsson, B. A. & Höglund, P. Renal effects of aspirin are clearly dose-dependent and are of clinical importance from a dose of 160 mg. Eur. J. Heart Fail. 10, 892–898 (2008).
Kim, J. Y. et al. Use of aspirin and initial cardiovascular and bleeding risk in patients with chronic kidney disease. Clin. J. Am. Soc. Nephrol. 20, 387–396 (2025).
Nørregaard, R., Mutsaers, H. A. M., Frøkiær, J. & Kwon, T. H. Obstructive nephropathy and molecular pathophysiology of renal interstitial fibrosis. Physiol. Rev. 103, 2827–2872 (2023).
Yang, Q., Su, S., Luo, N. & Cao, G. Adenine-induced animal model of chronic kidney disease: current applications and future perspectives. Ren. Fail. 46, 2336128 (2024).
Miao, J. et al. Experimental models for preclinical research in kidney disease. Zool. Res. 45, 1161–1174 (2024).
Fukasawa, H. et al. Down-regulation of Smad7 expression by ubiquitin-dependent degradation contributes to renal fibrosis in obstructive nephropathy in mice. Proc. Natl. Acad. Sci. USA 101, 8687–8692 (2004).
Cui, D. et al. Phloretin ameliorates hyperuricemia-induced chronic renal dysfunction through inhibiting NLRP3 inflammasome and uric acid reabsorption. Phytomedicine 66, 153111 (2020).
Liu, M. et al. p-Hydroxy benzaldehyde, a phenolic compound from Nostoc commune, ameliorates DSS-induced colitis against oxidative stress via the Nrf2/HO-1/NQO-1/NF-κB/AP-1 pathway. Phytomedicine 133, 155941 (2024).
MacParland, S. A. et al. McGilvray, Single-cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Liao, J. et al. Single-cell RNA sequencing of the human kidney. Sci. Data 7, 4 (2020).
Zhou, W. et al. The gut microbe Bacteroides fragilis ameliorates renal fibrosis in mice. Nat. Commun. 13, 19 (2022).
Kulak, N. A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 11, 319–324 (2014).
Wu, W. H. et al. Ginsenoside compound K restrains hepatic fibrotic response by dual-inhibition of GLS1 and LDHA. Phytomedicine 135, 156223 (2024).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Grant 82330123 to L.W.Q.), the Frontier Technologies R&D Program of Jiangsu (Grant BF2024074 to L.W.Q.), the National Natural Science Foundation of China (Grant 82374103 and 82174036 to Q.L.) and Fundamental Research Funds for the Central Universities (Grant 2632024TD03 to Q.L.). We thank Wei Jiang in China Pharmaceutical University for assistance in SPR and BLI experiments. Special thanks go to Dr. Raphael N. Alolga for his valuable assistance in polishing this manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: Q.L., L.W.Q., and W.H.W.; Methodology: W.H.W., L.Z.Z., Y.M.F., P.Z., B.L.L, and J.F.L.; Investigation: W.H.W., L.Z.Z., L.C., Y.M.F., and Y.Z.; Visualization: L.C. and Y.M.F. Funding acquisition: Q.L. and L.W.Q.; Project administration: L.W.Q. and Q.L.; Supervision: K.L. and G.M.; Writing–original draft: B.L.L, W.H.W., L.Z.Z. and L.C.; Writing–review and editing: Q.L., L.W.Q., W.H.W., and B.L.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Toshinari Takamura, Sandra Rayego, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, WH., Zhu, LZ., Liu, K. et al. Hepatokine fibrinogen-like protein 1 drives liver-kidney crosstalk to promote renal fibrosis. Nat Commun 17, 1453 (2026). https://doi.org/10.1038/s41467-025-68188-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68188-0