Abstract
Targeted therapies have revolutionized cancer care. Unfortunately, most patients develop refractory, multifocal resistance to these therapies within a matter of months. Here, we demonstrate that the evolution of resistance to EGFR inhibitors in EGFR-mutant non-small cell lung cancer endows cells with hypersensitivity to a PAINS-like small molecule, MCB-613. Systematic proteomic, functional genomic, and biochemical studies revealed that MCB-613 binds KEAP1 in a covalent, cysteine-independent fashion, acting as a divalent molecular bridge that relies upon lysine residues in the KEAP1 dimerization domain to join monomers of KEAP1 together. Oligomerization of KEAP1 by MCB-613 sets into motion a fatal cascade of KEAP1 dysfunction, ROS accumulation, and ATF4/CHOP-dependent cell death. Together, these findings demonstrate that diverse models of EGFR inhibitor-resistant NSCLC share the common feature of elevated integrated stress response activity, and that a covalent molecular bridge which activates non-canonical KEAP1-ATF4 signaling can exploit this feature to select against resistance evolution.
Similar content being viewed by others
Introduction
Although the use of targeted therapies to treat cancer has resulted in encouraging clinical responses, these benefits are, unfortunately, often short-lived. In the case of non-small cell lung cancers (NSCLC) driven by activating mutations in the epidermal growth factor receptor (EGFR), for instance, most patients will develop resistance to first-line targeted therapy within 24 months1,2. This problem is complicated by two interrelated considerations: (1) EGFR-mutant NSCLC cells can develop resistance to EGFR inhibitors through a wide variety of distinct mechanisms; and, (2) these mechanisms can coevolve simultaneously in the same patient or tumor3. This presents clinicians and researchers with a difficult quandary: on the one hand, treatments which use one drug to target an individual resistance mechanism are insufficient and unlikely to prove curative; on the other hand, treatments which use multiple drugs to target many different mechanisms at the same time are demanding, and in many cases infeasible. Thus, new approaches to the problem of “multifocal” drug resistance are needed.
One strategy is to identify, target, and exploit vulnerabilities which emerge as a consequence of drug resistance itself. These acquired, or “collateral,” sensitivities—which are distinct from chemical synthetic lethalities in that they persist even after removal of the original, selecting drug—have long been documented in the microbial literature4. Recently, for instance, it was shown that clinical isolates taken from the lungs of cystic fibrosis patients suffering from chronic, drug-resistant Pseudomonas aeruginosa infections consistently harbored mutations in the nfxB gene5. These mutations, though pathoadaptive in the setting of conventional anti-pseudomonal therapy with fluoroquinolones, nevertheless render bacterial populations collaterally sensitive to the commonly used and readily available aminoglycoside antibiotic amikacin. In point of fact, drug resistance often promotes targetable collateral sensitivities to commonplace, existing drugs. In the case of the recently approved fluorocycline antibiotic eravacycline, for example, drug resistance in Klebsiella pneumoniae promotes sensitivity to commonly used antibiotics like aztreonam and ceftazidime6.
In recent years, this concept has increasingly been applied to the study of drug resistance in cancer as well. It has been demonstrated, for example, that the acquisition of BRAF inhibitor resistance in BRAF-mutant melanoma engenders an upregulation of reactive oxygen species (ROS) which in some cells promotes a stable and clinically relevant collateral sensitivity to histone deacetylase (HDAC) inhibitors7. Separately, previous work has shown that diverse pathways of drug resistance in this setting also converge upon the activation of the c-MYC transcription factor; this, in turn, gives rise to shared, actionable collateral sensitivities to drugs which target MYC synthetic lethal partners such as certain tyrosine kinase families and metabolic pathways8. Since at least some of these targetable collateral sensitivities to cancer drug resistance are both stable and predictable, moreover, it is conceivable that future first-line therapies could be designed specifically to guide tumor evolution toward targetable, collaterally sensitive states. Indeed, previous work has shown that BET bromodomain inhibitors can be used to guide acute myeloid leukemia (AML) cells into an “evolutionary trap” which collaterally sensitizes them to existing, clinically approved BCL2 inhibitors9. Finally, these effects are manifold: with examples cited in a growing number of cases, it appears increasingly likely that numerous such collateral sensitivities may in fact be hidden throughout the landscape of drug-resistant cancers10,11,12,13,14. Indeed, the identification of such a collateral sensitivity, were it to be shared across clones driven by distinct mechanisms of resistance to the same selecting agent, would represent an opportunity to design therapies with the unique potential to select against resistance evolution15.
Here, we report on the discovery of a collateral sensitivity shared across multiple, distinct models of EGFR inhibitor resistance in EGFR-mutant non-small cell lung cancer. Specifically, we show that the acquisition of EGFR inhibitor resistance sensitizes EGFR-mutant NSCLC cells to a small molecule with PAINS-like features known as MCB-613, an effect that relies upon a unique, cysteine-independent mode of interaction with Kelch-like ECH associated protein 1 (KEAP1). More precisely, MCB-613 acts as a molecular bridge: through two distinct and symmetrically situated unsaturated sites, this electrophilic small molecule relies upon a lysine residue (lysine 97) situated within the protein dimerization domain and covalently tethers together monomers of KEAP1 to induce the formation of dysfunctional oligomers. Unlike typical, non-covalent dimers of KEAP1, these dysfunctional complexes then dissociate from the canonical repression substrate nuclear factor erythroid 2-related factor 2 (NRF2); this effect, however, is not required for MCB-613-induced cell death. Instead, MCB-613 selectively targets drug-resistant, EGFR-mutant NSCLC cells by taking advantage of their elevated integrated stress response (ISR) activity, promoting the accumulation of ROS and downstream ATF4- and CHOP-dependent cell death.
Results
A pharmacologic screen to identify shared collateral sensitivities in EGFR-mutant, EGFR inhibitor-resistant NSCLC
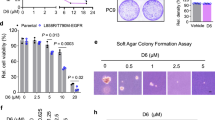
To identify targetable collateral sensitivities shared across clones bearing distinct mechanisms of EGFR inhibitor resistance, we compiled a panel of four well-characterized EGFR-mutant non-small cell lung cancer cell lines with varying degrees of resistance to EGFR inhibition: PC9, PFR3, GR4, and WZR12. The “parental” PC9 cell line is drug-naïve and retains exquisite sensitivity to EGFR inhibition. The previously characterized PFR3, GR4, and WZR12 cell lines—which are all derived from PC9s and driven by diverse, clinically relevant resistance mechanisms including IGF1R activation, EGFR T790M secondary site mutation, and downstream MAPK1 amplification, respectively—display increasing degrees of resistance to the first-generation EGFR inhibitor gefinitib (Fig. 1a)16,17,18. We screened all four of these cell lines against a library of 2100 small molecules in a three-day viability assay at two separate drug doses (Fig. 1b, Supplementary Data 1 and Supplementary Table 1). As expected, we detected numerous collateral resistances and sensitivities in EGFR inhibitor-resistant cells (Fig. 1c). Predictably, receptor tyrosine kinase inhibitors, including numerous other EGFR inhibitors (gefitinib, erlotinib, and afatinib, among others), scored as shared collateral resistances, providing an important internal positive control. Of the collateral sensitivities, however, one in particular stood out across all three drug-resistant derivatives: a relatively understudied, PAINS-like molecule called MCB-613 (Fig. 1d). Indeed, a follow-up, 3-day validation assay confirmed that the drug-resistant PFR3, GR4, and WZR12 derivatives were collaterally sensitive to treatment with MCB-613 (Fig. 1e).
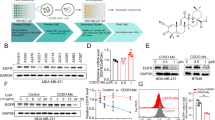
a Relative cell viability following 72-h incubation with gefitinib across 8-point serial drug dilution. Data are mean ± SEM for n = 3 biologically independent experiments. b High-throughput pharmacologic screening strategy. c Manhattan plot depicting results of chemical screen. Each point represents the average of (resistant - parental) from two biologically independent experiments. Collateral resistances and sensitivities in red and blue, respectively. All other conditions (no collateral effect, empty control wells, vehicle control wells) gray. Dotted horizontal lines indicate thresholds for collateral resistances (top, y = 0.3) and sensitivities (bottom, y = −0.3). d Magnification of Manhattan plot highlighting MCB-613. e Relative cell viability following 72-h incubation with 1.75 μM MCB-613. Data are mean ± SEM for n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. Exact p values listed within Source Data. f Relative cell viability of parental and gefitinib-resistant PC9 clones following 72-h incubation with 4 µM MCB-613. Data are mean ± SEM for n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. g. Relative cell viability of parental and gefitinib-resistant HCC827 clones following 72-h incubation with 2 µM MCB-613. Data are mean ± SEM of n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. h Relative cell viability of parental and osimertinib-resistant cells following 72-h incubation with MCB-613 across 8-point linear drug dilution. Data are mean ± SEM for n = 3 biologically independent experiments. i Tumor growth curves from mice xenotransplanted with PC9 or GR4 cells and treated with vehicle (saline) or MCB-613 (20 mg/kg) three times weekly via interaperitoneal injection. Data are mean ± SEM for n = 10 mice. P values computed using one-way ANOVA with multiple comparisons. j Line plot depicting estimated cumulative cell growth over time in vitro for a heterogeneous population comprised of 97% drug-naïve PC9 cells and 3% drug-resistant cells (1% PFR3, 1% GR4, 1% WZR12) exposed to either gefitinib, MCB-613, or combination of the two. Data are mean ± SEM for n = 3 biologically independent experiments. Source data are provided as a Source Data file.
We next sought to assess the breadth of this shared collateral sensitivity to EGFR inhibitor resistance. First, we gathered together two drug-naïve, EGFR-mutant NSCLC cell lines—PC9 and HCC827—which are known to develop EGFR inhibitor resistance through diverse mechanisms19,20,21,22. We used increasing doses of gefitinib to cultivate EGFR inhibitor resistance in these cell lines before selecting individual clones and querying them for sensitivity to MCB-613 (Supplementary Fig. 1a, b). In all clones tested, resistance to EGFR inhibition was associated with collateral sensitivity to MCB-613 (Fig. 1f, g). Finally, to assess whether this finding was limited to gefitinib or extended to other clinically relevant EGFR inhibitors, we evolved MGH134, a previously described, patient-derived cell line model of EGFR-mutant NSCLC, to resistance against the first-line, third-generation small molecule inhibitor osimertinib; in accordance with the above findings, resistance to this standard-of-care targeted therapy also conferred increased sensitivity to MCB-613 (Fig. 1h, Supplementary Fig. 1c)23.
Lastly, to evaluate whether the broad, shared collateral sensitivity of EGFR-mutant, EGFR inhibitor-resistant NSCLC cells to MCB-613 represents a viable preclinical strategy for overcoming multifocal drug resistance, we administered MCB-613 to tumor bearing mice. Despite the suboptimal solubility and pharmacokinetic properties of some molecules belonging to the chalcone class, we observed significant growth inhibition by MCB-613 in gefitinib-resistant GR4 tumors but not gefitinib-sensitive, parental PC9 tumors (Fig. 1i)24,25,26. Given the strong correlation between resistance to EGFR inhibition and sensitivity to MCB-613 observed above, we hypothesized that the combination of these two therapeutic approaches in a single EGFR-mutant NSCLC population would suffice to successfully suppress all cellular outgrowth. To test this hypothesis, we mixed drug-naïve and drug-resistant PC9 cells together and then exposed this heterogeneous population to either gefitinib alone, MCB-613 alone, or a combination of the two. Although neither molecule on its own suppressed outgrowth, the combination of the two was enough to completely eradicate the population (Fig. 1j). When considered together, these findings establish MCB-613 as a bona fide collateral sensitivity shared across diverse models of EGFR inhibitor resistance.
Proteomic and functional genomic screens reveal that MCB-613 targets KEAP1
MCB-613 (4-ethyl-2,6-bis-pyridin-3-yl-methylene cyclohexanone) is a reactive, PAINS-like electrophile belonging to the bis-chalcone family and boasting multiple previously described protein targets, including steroid receptor coactivator 3 (SRC3) and ubiquitin-specific peptidase 15 (USP15), which controls selective depletion of mutant p53 (Fig. 2a)27,28. In our hands, however, the depletion of these previously described targets failed to rescue the selective cell death effect of MCB-613, suggesting that they may not be relevant in the setting of drug-resistant EGFR-mutant NSCLC (Supplementary Fig. 2a–f). Acknowledging that MCB-613 is not drug-like, that it has structural elements that resemble those found in common pan-assay interference compounds (PAINS), and that the mechanism through which MCB-613 drives cell death in EGFR inhibitor-resistant cells could involve multiple functional targets or general electrophilic stress, we nevertheless set out to determine whether this agent may function through a previously unidentified defined target and mechanism-of-action. To do this, we devised a comprehensive strategy leveraging synthetic chemistry, proteomics, and functional genomics.
a Chemical structure of MCB-613. b Chemical structure of C151-0598. c Chemical structure of the active ARM-3-124 clickable derivative. d Chemical structure of the inactive ARM-3-115 clickable derivative. e Relative cell viability of parental PC9 (top) or drug-resistant WZR12 (bottom) cells following 72-h incubation with the active ARM-3-124 (red) and inactive ARM-3-115 (black) clickable derivatives across an 8-point linear drug dilution series. Data are mean ± SEM for n = 3 biologically independent experiments. f Volcano plot depicting screen results. Each point represents relative essentiality of a gene encoding for a protein interactor of MCB-613 in drug-resistant (GR4 and WZR12) versus parental (PC9) cells. Negative and positive x-axis values indicate collateral sensitivities and resistances, respectively. Adjusted p values calculated with the Wald test in MAGeCK. g Growth plots depicting calculated cumulative cell count over time in parental (PC9) and drug-resistant (PFR3, GR4, and WZR12) cells following knockout of KEAP1 with sgKEAP1_3 (left) and sgKEAP1_5 (right). All results normalized to a non-targeting control. Data are mean ± SEM for n = 3 biologically independent experiments. P values for the final D18 timepoints computed using one-way ANOVA with multiple comparisons. Exact p values listed within Source Data file. h Relative cell viability of drug-resistant GR4 (left) and WZR12 (right) cells after 72 h of exposure to vehicle (DMSO) or two different doses of MCB-613 following knockout of KEAP1 versus non-targeting control. Data are mean ± SEM for n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. Source data are provided as a Source Data file.
First, we developed an approach for identifying all protein interactors of MCB-613 in parental and EGFR inhibitor-resistant cells. Using MCB-613 and C151-0598, a molecule bearing close structural similarity but showing severely attenuated activity, we designed a “click chemistry,” azide-alkyne cycloaddition (CuAAC) approach to differential affinity purification and target identification (Fig. 2b and Supplementary Fig. 2g). We first generated “clickable” derivatives of MCB-613 and C151-0598 by replacing the ethylene units at the 4-positions of their central cyclohexanone moieties with propargyl ethers (Fig. 2c, d, Supplementary Note). We called these active and inactive clickable derivatives “ARM-3-124” and “ARM-3-115,” respectively. Next, we verified that the derivatives phenocopied their parent compounds in EGFR inhibitor-sensitive and -resistant EGFR-mutant NSCLC cells (Fig. 2e). Finally, we deployed them for target identification. Specifically, we exposed parental and drug-resistant cells to either compound at a dose (2 μM) which only showed activity in the ARM-3-124/WZR12 arm before collection, lysis, and copper-catalyzed azide-alkyne cycloaddition to install a biotin handle. Samples were analyzed by mass spectrometry to identify bound peptides. Across all conditions, this approach identified >1000 unique peptides corresponding to 610 protein interactors. After applying various filters, the list was further refined to a total of 478 putative interactors (Supplementary Data 2).
Next, we sought to evaluate the functional contributions of each of the 478 protein interactors to the collateral sensitivity of EGFR inhibitor-resistant NSCLC cells to MCB-613. Thus, we designed a targeted, loss-of-function CRISPR/Cas9 library containing single-guide RNA (sgRNA) sequences against each of the 478 interactors and screened it in parental and drug-resistant cell lines. We reasoned that the target of MCB-613 may be found among the genes which, when lost, conferred a greater growth disadvantage upon resistant derivatives than upon parental cells. Of all such genes, it was loss of KEAP1, encoding the E3 ubiquitin ligase substrate adaptor Kelch-like ECH associated protein 1 (KEAP1), that most closely phenocopied the effects of MCB-613 and scored as the greatest collateral sensitivity in the screen (Fig. 2f). (We were reassured by this, as it is consistent with the fact that MCB-613—like other α, β-unsaturated electrophiles—interacts widely with members of the ubiquitin-proteasome system (UPS) (Supplementary Fig. 2h, Supplementary Data 2)29.) Indeed, follow-up validation in parental PC9 and gefitinib-resistant PFR3, GR4, and WZR12 cells confirmed that KEAP1 is modestly differentially essential in the setting of EGFR inhibitor resistance (Fig. 2g and Supplementary Fig. 2i), albeit less so than MCB-613 treatment. We took this as evidence that KEAP1 is the protein interactor responsible for the selective cell death effect of MCB-613 in the setting of EGFR inhibitor resistance in NSCLC. Furthermore, we reasoned that, if MCB-613 works through KEAP1 inhibition, then KEAP1 knockout from a bulk population of resistant derivatives should eradicate all cells sensitive to KEAP1 loss and leave behind a reservoir of cells that no longer demonstrate sensitivity to this molecule. Indeed, genetic knockout of KEAP1 rescued the effect of MCB-613 across each of the resistant derivatives tested (Fig. 2h and Supplementary Fig. 2i). Taken together, these results suggest that the interaction between MCB-613 and KEAP1 is required for this agent’s selective effect in EGFR inhibitor-resistant NSCLC cells and suggest that the molecule’s effects on KEAP1 may be more complex than simple loss-of-function.
MCB-613 acts as a covalent molecular bridge
To further explore the interaction between MCB-613 and KEAP1, we re-deployed our clickable probes in parental and drug-resistant cells before collection, lysis, and CuAAC. Gel electrophoresis and immunoblotting confirmed that the active probe, ARM-3-124, bound to KEAP1 (Fig. 3a). Furthermore, a thermal shift assay confirmed direct interaction between MCB-613 and purified KEAP1 (Fig. 3b). Specifically, we observed a dose-dependent decrease in Tm, which occurs when a ligand stabilizes a non-native protein conformational state and can indicate protein destabilization30. Indeed, follow-up cellular thermal shift assays (CETSA) in drug-resistant WZR12 cells confirmed that MCB-613 binds directly to and destabilizes both ectopic and endogenous KEAP1 (Fig. 3c and Supplementary Fig. 3a–c)31,32. Because protein instability often promotes degradation, we hypothesized that MCB-613 influences KEAP1 protein levels. Indeed, exposure of parental PC9 and gefitinib-resistant WZR12 cells to increasing doses of MCB-613 revealed a dose-dependent decrease in KEAP1 protein levels at the expected molecular weight. To our surprise, however, we also noticed a concomitant accumulation of KEAP1 at higher molecular weight (Fig. 3d). This suggested that, rather than degradation, MCB-613 might promote KEAP1 oligomerization. Although prior reports have shown that a variety of structurally diverse electrophiles can oligomerize KEAP1, the majority of these act in a non-covalent fashion. Certain other compounds, such as methylglyoxal, have been shown to form covalent adducts between KEAP1 monomers33. MCB-613 seems to be unusual in that it falls into the latter category, inducing the formation of a complex which persists even in the face of stringent reducing conditions, suggesting that it does not arise as a result of non-covalent protein-protein interactions or disulfide bridges (Supplementary Fig. 3d)34.
a Immunoblot analysis of bound KEAP1 following treatment of PC9 and WZR12 cells with 2 µM ARM-3-124 or ARM-3-115 for 1 h followed by lysis, CuAAC, and gel electrophoresis. N = 1 experiment. b Thermal denaturation curve depicting SYPRO Orange fluorescence induced by denaturation of purified His6-KEAP1 with increasing doses of MCB-613. Each condition internally normalized using non-linear fitting with modified Boltzmann Equation. Data are mean ± SEM of n = 3 technical replicates. c Line plot depicting densitometry analysis of CETSA immunoblot (Supplementary Fig. 3a). d Immunoblot analysis of endogenous KEAP1 at monomeric and dimeric molecular weights in PC9 and WZR12 cells after treatment with DMSO or increasing doses of MCB-613 for 1 h. Representative example shown from n = 2 biologically independent experiments. e Scatterplot analysis of proteins identified by MS to interact with V5-KEAP1. A high-molecular weight (~160 kDa), Coomassie-stained gel slice obtained following V5-immunoprecipitation from WZR12 cells ectopically expressing V5-KEAP1 and treated with DMSO or 2 µM dose MCB-613 for 1 h. Slope of dotted line = 1 and represents hits equally identified between the conditions. N = 1 for both conditions. Representative immunoblot shown from n = 3 biologically independent experiments. f Immunoblot analysis of Clover- and His-tagged KEAP1 after immobilization of HA-Clover KEAP1 on GFP-Trap beads and subsequent incubation with purified His6-KEAP1 and DMSO or increasing doses of MCB-613 for 15 min. Representative example shown from n = 2 biologically independent experiments. g Diagram depicting possible modification of KEAP1 by MCB-613. h Immunoblot analysis of GFP-tagged KEAP1 expressed in HEK cells and treated with DMSO or 2 μM MCB-613 or ARM-143 for 1 h before lysis and immunoblotting against GFP. N = 1 experiment. i Relative cell viability of PC9, PFR3, GR4, or WZR12 cells following 72-h incubation with ARM-143 (top) or MCB-613 (bottom) across an 8-point linear drug dilution series. Data are mean ± SEM for n = 3 biologically independent experiments. Source data are provided as a Source Data file.
Small molecules that act at protein-protein interfaces to promote direct interaction are known as molecular glues or bridges35. To assess whether MCB-613 indeed causes KEAP1 oligomerization, we expressed V5-KEAP1 in drug-resistant WZR12 cells before treatment with MCB-613, co-immunoprecipitation, and electrophoresis. Subsequent gel excision and MS analysis of the drug-induced, high molecular weight V5-KEAP1 band confirmed that it was substantially enriched for KEAP1 above all other protein species, suggesting that it consisted of the expected oligomer (Fig. 3e and Supplementary Data 3). An in vitro binding assay involving two purified and differentially tagged KEAP1 species treated with increasing doses of MCB-613 produced a similar transition from monomeric to high molecular weight KEAP1, providing us with further, direct evidence that MCB-613 induces KEAP1 oligomerization (Fig. 3f). Taken together, these data suggest a model in which MCB-613 uses its symmetrically situated α, β-unsaturated bonds to form a covalent bridge between distinct monomers of KEAP1 (Fig. 3g).
To test this model, we synthesized ARM-143, a closely related structural analogue of MCB-613 that harbors only one α, β-unsaturated bond (Supplementary Fig. 3e). As anticipated, treatment with ARM-143 failed to convert KEAP1 from its monomeric to its oligomeric form (Fig. 3h). To validate this finding, we next designed an assay capable of detecting addition of either of these molecules to monomers of KEAP1. Because such an addition would equip the protein with a rare functional ketone moiety, we reasoned we could use a click chemistry, hydrazone-mediated biotinylation as a readout for target engagement (Supplementary Fig. 3f, g). Indeed, this assay confirmed that although ARM-143 interacts covalently with monomers of KEAP1, it fails to reproduce the oligomers seen with MCB-613 (Supplementary Fig. 3h). Furthermore, ARM-143 also showed severely attenuated potency and failed to reproduce the collateral sensitivity to EGFR inhibitor resistance that is seen with MCB-613, suggesting that the molecular bridge-like features of the active parent compound are essential for selective cell death in this setting (Fig. 3i). To further investigate this final point, we tested a panel of structurally diverse, known KEAP1 inhibitors—including seven reactive electrophiles as well as one recently identified non-covalent inhibitor—against both parental and drug-resistant EGFR-mutant NSCLC cells34,36,37,38,39,40,41. Of these, curcumin alone—which bears significant structural similarity to MCB-613—effected a collateral sensitivity which was shared across all resistant derivatives (Supplementary Fig. 4a, b). These findings suggest that electrophilic stress and KEAP1 inhibition alone are not enough to cause collateral sensitivity, but instead that certain symmetric electrophiles act as covalent molecular bridges to oligomerize KEAP1 and cause selective cell death in drug-resistant cells.
Molecular bridging by MCB-613 relies on the KEAP1 dimerization domain
KEAP1 is recognized for its ability to sense and ligand a plethora of structurally diverse, thiol-reactive xenobiotics with its network of 27 cysteine residues34,42,43,44. We employed activity-based protein profiling (ABPP) to explore proteome-wide cysteine engagement with MCB-613, particularly focusing on KEAP1. We achieved extensive proteome coverage, quantifying 8,373 peptides, aligning with previous studies45. Contrary to expectations, and despite the fact that MCB-613 demonstrated rapid labeling of GSH in a classic GSH reactivity assay (Supplementary Fig. 5a), we observed only a mild level of cysteine engagement by MCB-613 across the proteome. (Unsurprisingly, ARM-143, which we have shown above labels KEAP1 (Supplementary Fig. 3h), showed decreased activity in the GSH assay, which was expected due to its lack of a dienone structure. The fact that MCB-613, on the other hand, rapidly labels GSH in an in vitro assay further speaks to its tool-like and non-drug-like nature.) Remarkably, only 8 cysteines showed an engagement score exceeding 10% (p value ≤ 0.05), none of which were associated with KEAP1 (Supplementary Data 4 and Supplementary Fig. 5b, c). To validate this finding, we substituted each of the 27 cysteine residues in KEAP1 with isostructural but less nucleophilic serine residues; however, none of these modifications rescued oligomerization of KEAP1 induced by MCB-613 (Supplementary Fig. 5d). One explanation for these findings is that MCB-613 might covalently engage cysteine residues through a reversible mechanism, thus rendering detection difficult in assays such as those described here. Although this cannot be ruled out on the bases of these data alone, the fact that competitive displacement is often detectable even with non-covalent inhibitors led us to speculate instead that MCB-613’s activity might depend upon alternative residues within KEAP146.
To identify which sites within KEAP1 might be required by MCB-613, we used an HA-Clover tag system to design a series of increasingly truncated forms of KEAP1. The first fragment, HA-Clover KEAP160, was limited to the first sixty amino acids of KEAP1, which comprise the protein’s N-terminal domain. The next fragment, HA-Clover KEAP1178, further incorporates the protein’s BTB domain. Similarly, the third and fourth fragments, HA-Clover KEAP1314 and HA-Clover KEAP1597, stretch to include the IVR and then Kelch domains, respectively. Also included was a full-length KEAP1 construct spanning the full 624 amino acids (HA-Clover KEAP1WT) (Fig. 4a). We transfected these fragments into 293T cells before treating with an active dose of MCB-613 for 1 h, preparing for immunoblot, and probing with an anti-GFP antibody. As expected, MCB-613 caused the accumulation of high molecular weight bands in cells expressing the HA-Clover KEAP1WT construct, again reflecting oligomerization of KEAP1. Indeed, similar bands appeared in the samples that were transfected with HA-Clover KEAP1178, HA-Clover KEAP1314, and HA-Clover KEAP1597 as well; the sole fragment which failed to reproduce this finding was HA-Clover KEAP160 (Fig. 4b). We therefore concluded that KEAP1 oligomerization by MCB-613 likely relies upon residues between positions 60 and 178, which unsurprisingly represents the protein’s dimerization domain.
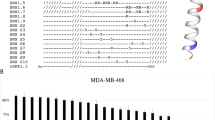
a Diagram depicting initial serial truncation strategy. b Immunoblot analysis of HA-Clover KEAP1 dimerization (upper bands) in 293FT cells transfected with (from left to right) no template, HA-Clover KEAP1WT, HA-Clover KEAP1597, HA-Clover KEAP1314, HA-Clover KEAP1178, and HA-Clover KEAP159 and treated with vehicle (DMSO) or MCB-613 for 1 h. N = 1 experiment. c Diagram depicting follow-up serial truncation strategy. d Immunoblot analysis of HA-Clover KEAP1 dimerization (upper bands) in 293FT cells transfected with (from left to right) no template, HA-Clover KEAP1WT, HA-Clover KEAP1170 through HA-Clover KEAP168, and treated with vehicle (DMSO) or MCB-613 for 1 h. Representative example shown from n = 2 biologically independent experiments. e Model of covalent interaction between MCB-613 and KEAP1. f Immunoblot analysis of HA-Clover KEAP1 dimerization (upper bands) in 293FT cells transfected with either HA-Clover KEAP1 WT or HA-Clover KEAP1 K97A and treated with DMSO or MCB-613 for 1 h. All images are taken from the same membrane. Direct comparisons between WT and K97A for a given molecular weight are taken from the same film exposure. N = 1 experiment. g Relative cell viability of WZR12 cells after 72-h exposure to DMSO or two different doses of MCB-613 following ectopic overexpression of either wild-type KEAP1 or the KEAP1 K97A mutant. Data are mean ± SEM for n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. Exact p values listed within Source Data file. Source data are provided as a Source Data file.
To narrow down our region of interest, we next repeated the approach described above by serially truncating HA-Clover KEAP1178 in increments of ten amino acids apiece (Fig. 4c). After treatment with MCB-613 and immunoblotting, we observed that drug-induced oligomerization remained intact so long as the first 100 amino acids of KEAP1 were expressed; with the expression of HA-Clover KEAP190, however, the oligomerization effect was severely attenuated (Fig. 4d). We were encouraged by this finding, since the region between KEAP1 residues Gln74 and Met120 is known to form a hydrophobic surface cavity which has been implicated in interaction with electrophiles such as pubescenoside A (PBA)47. In accordance with the ABPP findings presented above, moreover, the primary sequence of KEAP1 situated between amino acid positions 90 and 100 contains no cysteine residues. It does, however, contain a lysine at position 97, which stands out as a strong candidate for nucleophilic participation in Michael addition with MCB-613. Indeed, MCB-613 showed high lysine reactivity in an in vitro assay (again, and expectedly, unlike ARM-143), and computational modeling suggests that MCB-613 could even fit neatly into this previously described surface cavity, participating in hydrogen bonding with surrounding amino acid residues such as Arg116 and forming irreversible bonds bilaterally with the Lys97 residues of distinct KEAP1 monomers (Supplementary Fig. 6a, Fig. 4e). Unfortunately, lysine ABPP remains technically challenging, and an attempt to query this residue was ultimately unsuccessful. In fact, to our knowledge, no previously described lysine-reactive ABPP probe has ever been shown to successfully detect this specific peptide48,49,50,51.
To test the hypothesis that Lys97 is required for MCB-613-induced KEAP1 oligomerization, we compared wild-type KEAP1 with a mutant version of the protein in which the nucleophilic lysine residue at position 97 was replaced with a non-reactive alanine residue (K97A). In comparison with wild-type KEAP1, the K97A mutant exhibited dramatically diminished oligomerization when treated with MCB-613 (Fig. 4f). Consistent with this finding, expression of the full-length KEAP1 K97A mutant in EGFR inhibitor-resistant WZR12 cells completely blocked their sensitivity to MCB-613 (Fig. 4g). Thus, MCB-613 relies upon lysine 97 to oligomerize KEAP1.
MCB-613 promotes cell death through a NRF2-independent, ROS- and ATF4-dependent mechanism
KEAP1 plays a canonical role as an adaptor subunit of the Cullin 3-based E3 ubiquitin ligase machinery. Under homeostatic conditions, KEAP1 homodimerizes and binds its substrates—most notably, the master transcription factor nuclear factor erythroid 2-related factor 2 (NRF2)—to facilitate ubiquitination and degradation. In the presence of oxidative or electrophilic stress, however, direct modification of KEAP1 by oxidants and/or electrophiles disables the protein’s substrate adaptor function and allows for the rapid cellular accumulation of KEAP1 substrates. In the case of NRF2, this broadly results in activation of antioxidant transcriptional programs52. Interestingly, previous work has shown that the acquisition of resistance to first-generation EGFR inhibitors suppresses NRF2 activity53. We therefore wondered whether MCB-613-mediated KEAP1 perturbation influenced NRF2 protein and/or activity levels. Indeed, and as expected, treatment of EGFR inhibitor-resistant cells with MCB-613 led to rapid accumulation of NRF2 at the protein level and subsequent activation of NRF2 transcriptional programs (Fig. 5a, b and Supplementary Fig. 6b). Although NRF2 transcriptional programs are conventionally thought of as cytoprotective, recent work has shown that they can produce cytotoxicity in a subset of lung cancer cell lines through an NADH-mediated phenomenon known as “reductive stress.”54 Thus, we hypothesized that MCB-613-mediated KEAP1 inhibition might selectively target drug-resistant NSCLC cells through activation of NRF2 and initiation of reductive stress. To the contrary, however, the genetic knockout of NRF2 not only failed to rescue the collateral sensitivity of drug-resistant NSCLC cells to MCB-613, but instead further sensitized them to it (Fig. 5c and Supplementary Fig. 6c). Importantly, the KEAP1 K97A mutant (which blocks MCB-613-mediated KEAP1 oligomerization and cellular toxicity) does not affect basal NRF2 expression (Supplementary Fig. 6d). Although WZR12 cells bearing the KEAP1 K97A mutant do show some NRF2 accumulation with MCB-613 treatment, presumably due to the presence of endogenous wild-type KEAP1, this is blunted in comparison with the accumulation observed with overexpression of wild-type KEAP1 (Supplementary Fig. 6e). Thus, the effect of MCB-613 cannot be explained by its effects on NRF2, which appears to provide a limited buffering function against MCB-613-mediated stress.
a Immunoblot analysis of NRF2 over time following treatment with 2 µM MCB-613. Representative example from n = 2 independent experiments. b Scatterplot depicting GSEA results in WZR12 cells treated with 2 µM MCB-613 for 24 h. c Relative cell viability of WZR12 cells following 72-h incubation with DMSO or MCB-613 following NFE2L2 knockout. Data are mean ± SEM for n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. Exact p values in Source Data file. d Fluorescent microscopy images depicting DCFDA staining for ROS. Fields depict roughly equivalent numbers of cells. e Immunoblot analysis of ATF4, CHOP. Representative example from n = 3 biologically independent experiments. f Bar plot depicting relative ROS after 24-h exposure to DMSO or MCB-613. Data are mean ± SEM for n = 3 biologically independent experiments (except PC9 MCB-613, GR4 MCB-613, where n = 2). P values computed using two-way ANOVA with multiple comparisons. g Immunoblot analysis of ATF4, CHOP, p-EIF2α, t-EIF2α after 24-h exposure to DMSO or MCB-613. Representative example from n = 2 biologically independent experiments. h Bar plot depicting relative cell viability of WZR12 cells after 72-h exposure to DMSO or MCB-613, with DMSO, N-acetylcysteine, or α-tocopherol. Data are mean ± SEM for n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. i Immunoblot analysis of ATF4 after 24-h exposure to either DMSO or MCB-613, with DMSO, N-acetylcysteine, or α-tocopherol. N = 1 experiment. j Densitometry analysis of ATF4 after DMSO or 1.25 μM MCB-613 for 2 or 4 h following KEAP1 knockdown using two different shRNAs. Representative example from n = 2 biologically independent experiments. k. Relative cell viability after 72-h exposure to DMSO or MCB-613 following ATF4 knockout. Data are mean ± SEM for n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. l Relative cell viability after 72-h exposure to DMSO or MCB-613 following DDIT3 knockout. Data are mean ± SEM for n = 3 biologically independent experiments. P values computed using one-way ANOVA with multiple comparisons. Source data are provided as a Source Data file.
Given the findings above, we next wondered whether our diverse, EGFR inhibitor-resistant models held in common any other features which might explain their shared collateral sensitivity to treatment with MCB-613. It is known, for example, that gefitinib resistance promotes the accumulation of reactive oxygen species, and recent studies have shown that before EGFR-mutant NSCLC cells acquire stable drug resistance, they first pass through an initial, drug-tolerant persister state characterized by elevated levels of ATF455,56. As expected, and in accordance with these prior literature reports, our drug-resistant derivatives showed elevated baseline levels of reactive oxygen species, as well as elevated protein levels of ISR pathway members ATF4 and CHOP (Fig. 5d, e). In the latter case, in fact, we observed a direct correlation between the degree of resistance to gefitinib on the one hand and basal levels of protein expression of ATF4 and CHOP on the other (R2 = 0.9085 and 0.8713, respectively) (Supplementary Fig. 6f). Treatment of drug-resistant cells with MCB-613, moreover, exacerbated both of these phenotypes (Fig. 5f, g), but showed no effect on similar pathways, such as the heat shock response (as measured by levels of HSP60, HSP70, and HSP90 proteins, Supplementary Fig. 6g). MCB-613’s effects on both cell viability and ATF4 levels were rescued by pre-treatment with ROS scavengers such as N-acetylcysteine (NAC) and α-tocopherol (Fig. 5h, i and Supplementary Fig. 7a), and the induction of ROS by MCB-613 depended upon KEAP1 (Supplementary Fig. 7b). (Although it cannot be definitively ruled out by these studies alone that NAC is working by directly interfering with MCB-613, the data shown here suggest against that possibility.) Further, the effects of MCB-613 on ATF4 levels were, like cell viability (Fig. 2h), also KEAP1-dependent (Fig. 5j and Supplementary Fig. 7c). Together, these results suggest that diverse EGFR inhibitor-resistant models share features of ISR hyperactivation. Further, they are consistent with a model wherein KEAP1 oligomerization by MCB-613 leads to ROS induction and ATF4/CHOP activation in drug-resistant cells.
The ISR is a bivalent pathway: although moderate pathway engagement leads to stress amelioration and cellular recovery, excessive ISR signaling leads to enhanced cytotoxicity and, ultimately, cellular demise57. We therefore wondered whether the ISR hyperactivation, which is triggered in response to MCB-613-induced ROS in EGFR inhibitor-resistant cells, was primarily cytoprotective or cytotoxic. To answer this question, we knocked out the ISR pathway members ATF4 or CHOP in WZR12 cells before treating with MCB-613. Impressively, the loss of either of these ISR effectors was sufficient to rescue the effect of MCB-613 on cell viability (Fig. 5k, l and Supplementary Fig. 7d). Consistent with evidence that MCB-613’s activity requires KEAP1 oligomerization, ARM-143 failed to induce ATF4 expression (Supplementary Fig. 7e). Together, these findings suggest that, in the setting of EGFR inhibitor resistance, MCB-613 acts through a unique, KEAP1-dependent, NRF2-independent mechanism to selectively induce ROS and trigger an ATF4- and CHOP-mediated form of cell death.
Discussion
Here, we report that diverse models of EGFR inhibitor resistance in EGFR-mutant non-small cell lung cancer converge on a collateral sensitivity to the understudied, PAINS-like chemical compound MCB-613. Using a combination of unbiased, high-throughput assays and classic biochemical techniques, we identify the molecular target of MCB-613 to be KEAP1, defining an unexpected mechanism of action that involves covalent molecular bridge activity that is unrelated to the downstream activation of NRF2.
The mechanism described in this report sets MCB-613 apart from existing KEAP1-reactive probes, the majority of which act at solvent-exposed cysteine residues to divorce KEAP1 from substrates such as NRF2. More recently, non-covalent KEAP1 inhibitors such as KI696 have also been developed and have been incorporated as components of heterobifunctional molecules to induce KEAP1-mediated ubiquitination and degradation of neosubstrates such as FAK and BRD438,58. Similar efforts have been made to marshal proteolysis-targeting chimeras (PROTACS) for degradation of KEAP1 itself 59,60,61. Most recently, Lu et al. reported on the development of a bivalent inhibitor of KEAP1 which consists of two monovalent inhibitors joined by a linker62. To our knowledge, however, MCB-613 is unique among KEAP1-reactive small molecules. For one thing, it acts on KEAP1 in a lysine-dependent fashion. Our data even raise suspicion that MCB-613 may act on lysine 97 directly, which would make it atypical among small molecules in general, though we are unable to definitively support this claim. Interestingly, however, recent studies have shown that the human proteome contains many more reactive, ligandable lysine residues than was previously anticipated, and that these residues are accessible to a wide variety of aminophilic electrophiles48,63. Beyond lysine dependence and possible reactivity, moreover, MCB-613 is also unusual in that it acts as a covalent molecular bridge between monomers of KEAP1, which is likely what confers its unique activity among inhibitors of KEAP1 in the first place. (Whether this pattern is limited to KEAP1 dimers, or whether it includes the higher order oligomers we observe in the ectopic expression studies above—which could, alternatively, represent e.g. differential combinations of wild-type and tagged KEAP1 species, or non-covalent aggregates—remains unclear.) Although certainly uncommon, neither is this without precedent: molecular glues or bridges represent an emerging strategy for the perturbation of anti-cancer targets, including those as relevant as PD-1 and PD-L164,65,66,67,68,69,70. Beyond these and a handful of other examples, however, the reports of homodimer molecular bridges in the literature are limited, making MCB-613 a notable entry into this emerging category of small molecules71.
More broadly, the work described herein provides support for collateral sensitivity as an intellectual framework for approaching multifocal drug resistance, and nominates MCB-613 as an instructive probe compound in the setting of EGFR-mutant NSCLC. Indeed, our data support a provocative notion: that cells of this genetically defined subtype can be coerced into an “impossible choice” between sensitivity to EGFR inhibition on one hand and sensitivity to MCB-613 on the other, and that, when confronted with these two irreconcilable options, they can, ultimately, be overcome. Beyond drug-resistant lung cancer alone, several preliminary studies in other settings have also utilized this molecule to draw conclusions about cardiovascular and nervous ischemia72,73,74. In further support of our findings, moreover, these studies also report that the use of MCB-613 leads to activation of NRF2 transcriptional programs. Although this was initially attributed to the downstream effects of steroid receptor coactivator (SRC) hyperstimulation, it is not lost on us that KEAP1 oligomerization might also contribute to this phenotype. Together, while the translational potential of MCB-613 is likely to be limited by its suboptimal pharmacological properties and expected off-target reactivity, it may serve as a template for the structure-based design of selective, drug-like, covalent KEAP1 oligomerizing molecular bridges with translational potential.
It remains unclear how dysfunctional, covalent oligomers of KEAP1 nucleated by MCB-613 promote the formation of ROS and downstream activation of ATF4. One possibility is that these corrupted protein species accumulate to generate proteotoxic stress, which can, in turn, trigger ER stress, ROS, and activation of the integrated stress response75. This hypothesis is even more attractive when it is considered that KEAP1 is both required for and itself degraded through a p62-dependent, autophagy-mediated clearance pathway76,77,78,79. Alternatively, it is also possible MCB-613-mediated perturbations to KEAP1 directly influence ATF4 itself. To this point, unfortunately, most of the literature dealing with any potential interaction between KEAP1 and ATF4 involves a go-between in the form of NRF2, which the data above clearly demonstrate is not relevant to the phenotype which is presently under discussion80,81,82,83,84. This raises additional questions. Are there heretofore undescribed substrates of KEAP1 that are involved in this mechanism? Is it possible that MCB-613 binding begets neosubstrates—or even neomorphic functions—for KEAP1? All of these questions, of course, ultimately beget another question: why are EGFR-mutant NSCLC cells in particular so collaterally sensitive to a KEAP1-dependent phenomenon? This point in particular warrants elaboration. It has long since been observed, for instance, that in the setting of non-small cell lung cancer, EGFR and KEAP1 mutations are mutually exclusive85,86. This remains poorly understood and is further complicated by recent reports that genetic interruptions in the KEAP1/NFE2L2/CUL3 axis may themselves contribute to the acquisition of EGFR inhibitor resistance87,88,89. Even these fragments of understanding, however, are enough to indicate that a delicate and sensitive balance takes place between these two fundamental cellular pathways. Although our present findings now mean that this reciprocal relationship can be exploited at key times for therapeutic, anti-cancer gain, further investigation will be needed to more fully elucidate the exact nature of this relationship.
In all, our work nominates MCB-613 as an effective preliminary means of exploiting this precarious relationship and provides an additional step toward understanding the complicated interplay at work here. More broadly, it also speaks to the power of collateral sensitivity as a framework for organizing studies that can both identify therapeutic strategies and provide insights into the nature of drug resistance itself.
Methods
This study complies with all relevant ethical regulations. Animal protocols were approved by the Animal Care and Use Committee of the Baylor College of Medicine. All newly synthesized reagents (e.g., oligonucleotides and plasmids) are available upon request.
Cell lines and reagents
All cell lines were maintained in a humidified incubator at 37 °C with 5% CO2. 293FT cells were cultured in DMEM high glucose medium supplemented with 10% fetal bovine serum (FBS), 1% penicillin/streptomycin, 1% non-essential amino acids, 1% GlutaMAX, and 1% sodium pyruvate. All other cell lines were cultured in RPMI-1640 medium supplemented with 10% FBS and 1% penicillin/streptomycin. The PC9 cell line was obtained from Sigma (90071810). The 293FT cell line was obtained from the Duke Cell Culture Facility (2832-R700-07). The PFR3, GR4, and WZR12 cell lines were a generous gift from Pasi Janne. The patient-derived MGH134 cell line was a generous gift from Aaron Hata. Drug-resistant cell lines were derived as previously described8,90. MCB-613 was a generous gift from Bert O’Malley. All other drugs were purchased from Cayman Chemical (gefitinib, osimertinib) and ChemDiv (C151-0598), or synthesized (Supplementary Note).
High-throughput pharmacologic screen
The Selleck Bioactives Compound Library was used in conjunction with the Duke Functional Genomics Core. In brief, 2100 bioactive compounds were stamped onto 384-well plates in duplicate using an Echo Acoustic Dispenser. Cell culture media containing PC9, PFR3, GR4, or WZR12 cells was added to wells using a Matrix WellMate for a final density of 300 cells per well and final concentrations of 2 and 10 µM for each drug. After 72 h, cell viability was assessed using the CellTiter Glo Luminescent Viability Assay (Promega) on a BMG Clariostar plate reader. The duplicate treatment wells for each compound were averaged and normalized to averages for duplicate control wells. Normalized values were then analyzed by calculating the log2(resistant derivative/parental) for each compound at both doses, and in parallel by subtracting the score for the parental cell line from the scores of the resistant derivatives. Hits that scored across multiple resistant derivatives and at the lower dose were prioritized. For further details, please refer to Supplementary Data 1 and Supplementary Table 1.
Dose-response (GI50) assays
Cells were seeded into 96-well plates at a density of 2000 cells/well and allowed to adhere overnight. The following day, cells were treated in triplicate with either vehicle (DMSO) or an eight-point drug dilution and returned to the incubator. After 72 h, luminescence was quantified with Cell Titer Glo (Promega). Relative cell viability was estimated by normalizing the raw luminescence values for the drug-treated wells to the DMSO controls.
In vivo studies
Animal protocols were approved by the Animal Care and Use Committee of the Baylor College of Medicine. Female NOD SCID mice (NOD.CB17-Prkdcscid/NCrCrl) were obtained commercially from Charles River Laboratories and housed in a strict sterile housing environment with temperatures around 22 °C and relative humidity of 40–60%. A standard light/dark cycle [12-h light, 12-h dark (12L:12D) cycle] condition was used. At the time of the study, 5 × 106 PC9 or GR4 cells were bilaterally implanted via subcutaneous injection into the flanks of 5–6 week-old female mice and allowed to grow for 2 weeks. After two weeks, tumors were measured, mice were weighed, and treatment was initiated. Mice were treated with either saline or MCB-613 (20 mg/kg) in saline three times weekly via intraperitoneal (i.p.) injection. During the treatment period, mice were weighed, and their tumors were measured weekly. The Animal Care and Use Committee of the Baylor College of Medicine permits a maximal tumor size of 2.0 cm in diameter in any direction for single, externally visible tumors. The maximal tumor size in this study did not exceed 1.5 cm in diameter in any direction for any mouse.
Synthesis of ARM-3-124, ARM-3-115, and ARM-143
Briefly, the preparation of clickable analogs (ARM-3-115, ARM-3-124) of C151-0598 and MCB-613 was performed in a one-step synthesis starting from 4-propargylcyclohexanone and benzaldehyde or nicotinaldehyde. Using these starting materials, this step consisted of a double aldol condensation with benzaldehyde. It was achieved in ethanol at room temperature using potassium hydroxide as the base. Thereby, ARM-3-115, the analog of C151-0598, was obtained in modest yield (21%). The analog of MCB-613 was also obtained through a double aldol condensation of the adequate 4-propargylcyclohexanone with nicotinaldehyde, but under acidic conditions (HCl in acetic acid). The target compound ARM-3-124 was obtained in 37% yield. Finally, ARM-143 was also prepared by aldol condensation, in acidic media, between nicotinaldehyde and commercially available 4-ethyl-2-(pyridine-3-ylmethyl)cyclohexanone, in 27% yield.
For more detailed characterization of chemical synthesis, yield, and purity, please see Supplementary Note.
Copper-catalyzed azide-alkyne cycloaddition (CuAAC) - “click” reaction
The click chemistry experiments were conducted as previously described91. Briefly, cells were seeded into six biological replicates and exposed for 1 h to a 2 µM dose of either the active ARM-3-124 (3 samples) or inactive ARM-3-115 (3 samples). After treatment, cells were washed three times in PBS, and the pellet was resuspended in 1330 µL of 1 % NP40 buffer completed with protease inhibitor. Resuspended cells were sonicated for 30s three times and incubated 30 min on ice before centrifugation at 20,000 × g for 10 min. After cell lysis, proteins were extracted and quantified using the Bradford method. For each condition, up to 10 mg of protein in 9.4 ml 1 % NP40 buffer was used to perform the biotin addition using 100 µL of biotin azo-azide at 5 mM, 200 µL TCEP at 50 mM, 100 µL TBTA at 10 mM, and 200 µL CuSO4 at 50 mM. Protein was then incubated for 1 h in the dark at room temperature. After incubation, 40 ml ice-cold methanol was added. Protein was precipitated overnight in methanol at −20 °C. Protein was then centrifuged at 5200 × g for 30 min at 0 °C and washed 3 times in the same conditions using ice-cold methanol. After drying, protein was resuspended in resuspension buffer (6 M urea, 2 M thiourea, 10 mM HEPES) and sonicated. Protein was supplemented with 40 µL DTT and incubated for 40 min at room temperature. Forty microliters of iodoacetamide was added to the samples and incubated 30 min at room temperature in the dark. After pre-washing of the streptavidin beads, resuspended proteins were added to the beads in a 15 ml tube and incubated on a rotator for 2 h at room temperature. Beads were collected by centrifugation at 200 × g for 3 min at room temperature and then washed with resuspension buffer (PBS and 1% SDS, PBS) solution twice. After washing, elution was performed using sodium dithionite solution. Eluted proteins were incubated overnight at −20 °C with ice-cold methanol. Proteins were then pelleted and dried before being resuspended in 4 % SDS buffer and 2X SDS free loading buffer. Samples were then separated by one-dimensional electrophoresis and stained with InstantBlue® dye. Gels were divided into pieces measuring approximately 1 mm3 and sent to Ross Tomaino at the Taplin Biological Mass Spectrometry Facility (Harvard Medical School) for identification of target proteins. Excised gel bands were cut into approximately 1 mm3 pieces. Gel pieces were then subjected to a modified in-gel trypsin digestion procedure92. Gel pieces were washed and dehydrated with acetonitrile for 10 min. followed by removal of acetonitrile. Pieces were then completely dried in a speed-vac. Rehydration of the gel pieces was with 50 mM ammonium bicarbonate solution containing 12.5 ng/µl modified sequencing-grade trypsin (Promega, Madison, WI) at 4 °C. After 45 min., the excess trypsin solution was removed and replaced with 50 mM ammonium bicarbonate solution to just cover the gel pieces. Samples were then placed in a 37 °C room overnight. Peptides were later extracted by removing the ammonium bicarbonate solution, followed by one wash with a solution containing 50% acetonitrile and 1% formic acid. The extracts were then dried in a speed-vac (~1 h). The samples were then stored at 4 °C until analysis. On the day of analysis the samples were reconstituted in 5 - 10 µl of HPLC solvent A (2.5% acetonitrile, 0.1% formic acid). A nano-scale reverse-phase HPLC capillary column was created by packing 2.6 µm C18 spherical silica beads into a fused silica capillary (100 µm inner diameter x ~30 cm length) with a flame-drawn tip93. After equilibrating the column each sample was loaded via a Famos auto sampler (LC Packings, San Francisco CA) onto the column. A gradient was formed, and peptides were eluted with increasing concentrations of solvent B (97.5% acetonitrile, 0.1% formic acid). As peptides eluted, they were subjected to electrospray ionization and then entered into a Velos Orbitrap Pro ion-trap mass spectrometer (Thermo Fisher Scientific, Waltham, MA). Peptides were detected, isolated, and fragmented to produce a tandem mass spectrum of specific fragment ions for each peptide. Peptide sequences (and hence protein identity) were determined by matching protein databases with the acquired fragmentation pattern by the software program, Sequest (Thermo Fisher Scientific, Waltham, MA)94. All databases include a reversed version of all the sequences, and the data was filtered to between a one and two percent peptide false discovery rate.
Design of custom sgRNA library
The custom sgRNA library was designed using the list of 478 MS-identified genes. For each of these genes, 4 sgRNA sequences were taken from each of two previously published, full-genome CRISPR libraries95,96. (Because sequences for a given gene were sometimes redundant between the two libraries, this approach produced an average of 7 unique sgRNA sequences per gene). Also included were 100 AAVS1-targeting “safe harbor” guides taken from a separate CRISPR library97. Each sequence was appended with the same universal prefix and suffix sequences (Supplementary Table 2) and synthesized commercially in pooled format (GenScript).
Cloning of the CRISPR library
The commercially synthesized oligo pool was diluted 1:10 in water and amplified using the Phusion HotStart Flex polymerase (NEB) according to the manufacturer’s protocol with the Array_F and Array_R primers (Integrated DNA Technologies) (Supplementary Table 2). The lentiCRISPR v2 vector (Addgene #52961) was digested with FastDigest Esp3I (Thermo Fisher) at 37 °C for 12 h. The digested product was size selected with gel electrophoresis and extracted using the Zymo Gel DNA Extraction Kit. The eluted product was cleaned using the AMPure XP reagent at a 1:1 input-to-reagent ratio, and concentration was measured using the Qubit dsDNA HS Assay Kit (Invitrogen). The digested vector was combined with the amplified inserts in a 5:2 ratio by mass and ligated using a 2X Gibson Assembly Master Mix (NEB) at 50 °C for 30 min. The ligation product was cleaned using the AMPure XP reagent at a 1:1.4 input-to-reagent ratio, and concentration was again measured using the Qubit dsDNA HS Assay Kit. The final product was transformed into electrocompetent cells (Lucigen), incubated overnight in LB broth at 37 °C, and isolated using the Qiagen Maxi Prep kit.
Lentivirus production
Lentivirus for both the pooled CRISPR screen and individual sgRNAs was produced using 293FT cells. For every 1.3 µg of plasmid, 1 µg of the psPAX2 packaging vector (Addgene #12260) and 0.657 µg of the VSVg envelope (Addgene #14888) were diluted into 385 µl of Opti-MEM supplemented with 11.6 µl of the lipofectamine-2000 transfection reagent and 12.64 µl of PLUS reagent (Thermo Fisher). This mixture was added dropwise to 293FT cells, and cells were returned to the incubator for 4 h. After the incubation, media were aspirated and replaced with an FBS-rich harvest medium (DMEM high glucose, 30% FBS, 1% PS, 1% NEAA, 1% GlutaMax, 1% sodium pyruvate) for 48 h. Viruses were collected, filtered, and either used directly or aliquoted and stored at −80 °C. Viral titers were performed as previously described9. Briefly, PC9, GR4, and WZR12 cells were seeded into 6-well plates at a density of 150,000 cells per well and allowed to adhere overnight. The next day, cells were exposed to media containing the polybrene transduction reagent (8 µg/ml) and increasing concentrations of library virus before spinfection via centrifugation at 900 × g for 1 h at room temperature. On Day 2, the media containing library virus and polybrene was removed and replaced with fresh media. On Day 3, the cells were subjected to selection with puromycin (2 µg/ml) for 48 h. Then, on Day 5, the cells were resuspended and counted to determine the percentage of cells infected per volume of virus and compared with an MOI table to estimate functional viral titer.
Pooled CRISPR screening
On Day 0, PC9, GR4, and WZR12 cells were seeded into 6-well plates at a density of 150,000 cells per well and allowed to adhere overnight. On Day 1, the cells were exposed to media containing library virus and the polybrene transduction reagent (8 µg/ml) before spinfection via centrifugation at 900 × g for 1 h at room temperature. On Day 2, the media containing library virus and polybrene was removed and replaced with fresh media. On Day 3, the cells were subjected to selection with puromycin (2 µg/ml) for 48 h. Then, on Day 5, the cells were resuspended, counted, divided into triplicates, and carried forward with passage every five days for an estimated 14 population doublings. Throughout the entirety of the screen, each replicate was represented at any given point in time by at least 4 × 106 cells to guarantee at least 1000x coverage of the custom library. Pellets of at least 4 × 106 cells were taken at T0, Tf, and at each passage as well. Upon completion of the screen, pellets were lysed and genomic DNA was extracted using the QIAamp Blood Maxi Kit (Qiagen).
CRISPR Screen sequencing and analysis
Extracted DNA was amplified with a two-step protocol. First, sgRNA libraries were PCR-amplified from genomic DNA using NEBNext Ultra II Q5 Master Mix according to the manufacturer’s protocol using the PCR 1 Forward and Reverse primers (IDT) (Supplementary Table 2). Following the initial amplification, a second PCR step was undertaken to append the PCR 1 product for each condition with a unique combination of staggered forward and barcoded reverse primers to facilitate pooled sequencing (IDT) (Supplementary Tables 2 and 3). In all cases, the amplified libraries were purified with SPRIselect beads (Beckman Coulter) using right-sided selection. Samples were quantified with the Quant-iT dsDNA Broad Range Assay Kit (Thermo Fisher), pooled, and sequenced on an Illumina NextSeq 500 with 75 bp single-end sequencing. Analysis was performed using the Models-based Analysis of Genome-wide CRISPR/Cas9 Knockout (MAGeCK) software analysis pipeline under the default conditions. Data from the GR4 and WZR12 cell lines were collapsed into one condition representing drug resistance, and compared with data from the PC9 parental cell line.
CRISPR screen validation by single gene knockout
To validate screening hits, the top two scoring sgRNA sequences for a given gene were taken from the custom library, commercially synthesized (IDT), cloned into the LCv2 vector, transfected into 293FT cells, and transduced into parental and drug-resistant NSCLC cells as described above (Supplementary Table 2). After 48 h of puromycin selection, 350,000 cells per condition were seeded in triplicate into 10-cm dishes and passaged and counted every three days. Total cumulative cell counts over time were estimated using cell counts and doubling time while assuming exponential growth and a negligible death rate. Cumulative counts for a given sgRNA were normalized to cumulative counts for a non-targeting control.
Protein purification
His6-KEAP1 was purified as previously described98. Briefly, One Shot BL21 (DE3) chemically competent E. coli cells were transformed with the pET28a-His6-KEAP1 vector (Addgene #62454). Colonies were grown in LB broth containing 50 µM ZnCl2 and 50 µg/ml kanamycin at 37 °C until cultures reached an OD600 of 0.6-0.8, and then were allowed to cool to room temperature. Protein expression was induced with 250 µM IPTG and allowed to shake at 19 °C for 14 h. Bacteria were collected and His6-KEAP1 purified on Ni-NTA agarose beads (Thermo Fisher) using the following buffers: lysis (50 mM NaH2PO4 (pH 8.0), 10 mM Imidazole, 5 mM β-mercaptoethanol (BME), 0.01% Triton X-100), washing buffer (50 mM NaH2PO4 (pH 8.0), 50 mM Imidazole, 500 mM NaCl, 5 mM BME, 0.01% Triton X-100), and elution (50 mM NaH2PO4 (pH 8.0), 125 mM Imidazole, 150 mM NaCl, 5 mM BME). Following elution, elution fractions were analyzed by gel electrophoresis to select and pool fractions of highest purity. The pooled fractions were then exchanged into storage buffer (50 mM Tris (pH 8.0), 10 mM DTT, 100 mM NaCl, 5% glycerol) using the Amicon Ultra-15 centrifugal filter column with a 10 kDa pore size (Millipore). Protein was quantified with the Bradford method before adjusting to a stock concentration of 4 mg/ml, aliquoting, flash freezing in liquid nitrogen, and storing at −80 °C.
Construction of pLX302 V5-KEAP1
The pLX302 V5-KEAP1 construct was assembled from an empty vector backbone (Addgene #25896) and the pDONR223 KEAP1 WT vector (Addgene #81925) using the Gateway LR Clonase II enzyme according the manufacturer’s instructions.
Thermal shift assay
The thermal shift assay was performed in a 384-well format as previously described in ref. 99. Purified His6-KEAP1 was diluted to a final concentration of 133 µg/ml (1.9 µM) in assay buffer (50 mM Na3PO4, 50 mM NaCl, pH 6.0) containing 8x SYPRO Orange diluted from a 5,000x stock (Invitrogen). The mixture was transferred to the wells of a 384-well plate and treated with either vehicle (DMSO) or doses of MCB-613 diluted in assay buffer. All conditions were plated in triplicate. The plate was sealed and subjected to centrifugation at 800 × g for 2 min to settle the samples before incubating at room temperature for 15 min. After the incubation ended, the plate was transferred to a Bio-Rad CFX384 Touch Real-Time PCR Detection System at 25 °C. The well temperature was increased in 0.5 °C increments with a 10-s equilibration time and fluorescence was measured by the built-in FRET channel until the machine reached 95 °C. For each condition, the melting temperature was determined by adjusting the raw values according to a modified Boltzmann Equation and calculating the average of the three replicates.
Cellular thermal shift assay (CETSA)
The cellular thermal shift assay was performed according to previous reports32. First, cells were seeded and allowed to adhere overnight. On the day of the assay, cells were treated in biological triplicate with either vehicle (DMSO) or a 2 µM dose of MCB-613 for 1 h before trypsinization, room temperature PBS washes, and collection in room temperature PBS supplemented with a protease inhibitor (Roche). For each condition, approximately 1 × 106 cells were then transferred to each of 12 PCR strip tubes and sealed. A gradient PCR protocol including the following temperatures (°C) was initiated: 42.2, 43.7, 45.3, 46.7, 48.3, 49.8, 50.2, 51.7, 53.3, 54.2, 56.3, and 57.8. Samples were held off of the machine until the target temperature for each well was reached, and then placed on the machine and incubated at the target temperatures for 3 min. Each of the samples was then incubated in the thermal cycler at 25 °C for an additional 3 min, before being allowed to rest for a final 3 min at room temperature and on the bench top. Then, the PCR strip tubes were transferred to pre-chilled racks and snap frozen in liquid nitrogen, before incubating until completely thawed (~3-4 min) in a room temperature water bath. The cycle was repeated 3 more times for a total of 4 freeze-thaw cycles to promote cell lysis. After the final freeze-thaw cycle, the total volume of each PCR strip tube (~100 µl) was transferred to a 1.5 mL Eppendorf tube and centrifuged at 17,000 × g for 40 min at 4 °C. Debris pellets were discarded, and 60 µl of supernatant was transferred to a new tube containing 20 µl of NuPAGE LDS Sample Buffer (4X) and 4 µl of β-mercaptoethanol. Samples were vortexed and allowed to incubate at room temperature for 30–60 min before western blotting according to the procedures described below. Targets were detected using primary antibodies anti-KEAP1 clone 144 (MABS514, Sigma) and anti-V5 (#13202, CST) for endogenous and ectopic KEAP1, respectively.
Densitometry analysis
Densitometry analysis of X-ray films from immunoblotting studies was performed using Adobe Photoshop and ImageJ. Images were converted to grayscale in Photoshop, and the intensity of bands was measured in ImageJ. For each blot, measurements were taken using a constant size selection for regions of interest. Background measurements were also collected from the blank regions located just above these bands. Pixel density was inverted by subtracting the measured intensities of both the band and background regions from 255. Net values were determined by subtracting the inverted densities of the background regions from the inverted densities of their corresponding bands of interest. The net value for each band of interest (e.g., KEAP1) was then normalized to the net value of the loading control for that lane (e.g., β-Actin). Finally, these normalized net values were themselves normalized to the 42.2 °C band in the CETSA experiment and plotted as a function of temperature.
Western blot
Cells were washed in ice-cold PBS, collected via scraping, and re-suspended in ice-cold 1X Cell Lysis Buffer (CST) supplemented with protease and phosphatase inhibitors (Pierce). Cell suspensions were vortexed thoroughly and rotated end-over-end at 4 °C for 30 min before centrifugation at 21,130 × g at 4 °C for 20 min. The debris pellets were discarded, and the supernatants transferred to fresh Eppendorf tubes before quantification using the Bradford method. Samples were diluted to equal concentrations using lysis buffer and prepared for electrophoresis with NuPAGE LDS Sample Buffer (4X) before boiling at either 70 °C for 10 min or 95 °C for 15 min. Prior to electrophoresis, samples were reduced with DTT at a final concentration of 50 mM except where otherwise noted. Membranes were probed with primary antibodies anti-KEAP1 clone 144 (MABS514, Sigma), anti-NRF2 (Ab137550, Abcam), anti-ATF4 (#11815, CST), anti-CHOP (#5554, CST), anti-V5-tag (#13202, CST), anti-GFP (#2555, CST), anti-Vinculin (#4650, CST), and anti-β-Actin (#4970, CST), and secondary antibodies anti-rabbit IgG, HRP-linked (#7074, CST) and anti-rat IgG, HRP-linked (#7077, CST). Uncropped and unprocessed scans of all immunoblots are available in the Source Data file.
Co-immunoprecipitation
For co-IP studies, 4–5 × 106 cells were seeded into two 15-cm dishes per condition and allowed to adhere overnight. The next day, cells were treated with a 2 µM dose of MCB-613 for 1 h before scraping into ice-cold PBS. Cells were resuspended in IP buffer containing 40 mM Tris HCl (pH 7.4), 150 mM NaCl, 20 mM EDTA, 1 mM DTT, and 0.5% NP-40 supplemented with phosphatase and protease inhibitors, thoroughly vortexed, and rotated end-over-end at 4 °C for an hour. Samples were then subjected to centrifugation at 21,130 × g for 20 min at 4 °C before harvesting the supernatant and measuring protein concentration using the Bradford method. A small volume of lysate was reserved for input, and the rest (600–800 µg) was adjusted so that a constant amount was loaded onto antibody-bound sepharose beads (see below) per sample. Samples were rotated end-over-end at 4 °C overnight. The following morning, samples were washed 5 times in ice-cold IP buffer before boiling at 95 °C for 5 min in NuPAGE LDS Sample Buffer (4X). Lysates were collected from the beads using a 50 µl syringe (Hamilton) and analyzed via gel electrophoresis according to the procedures described above.
To prepare beads, 50 µl of Recombinant Protein G – Sepharose beads (Invitrogen) per condition were washed 3 times in IP buffer. The beads were then resuspended in IP buffer, and anti-V5-tag (E9H8O) Mouse mAb (#80076) (CST) was added at a 1:50 dilution as instructed by the manufacturer. The bead-antibody combination was then rotated end-over-end at 4 °C for at least 5 h. After the incubation, the beads were washed 3 times in IP buffer and divided evenly among the experimental conditions.
LC-MS based protein identification from SDS-PAGE gel bands
WZR12 cells were transduced with either pLX302 V5-luciferase (Addgene #47553) or pLX302 V5-KEAP1 and selected with puromycin. After selection, cells bearing V5-KEAP1 were treated with either DMSO or 2 µM MCB-613 for 1 h before collection and V5-co-immunoprecipitation, alongside an untreated, V5-luciferase control. Samples were collected, prepared for, and subjected to SDS-PAGE as described above. Coomassie stained SDS-PAGE bands (n = 1 per condition) were subjected to standardized in-gel trypsin digestion in which gel bands were subjected to reduction with 10 mM dithiolthreitol, alkylated with 20 mM iodoacetamide, and digested with 100 ng of sequencing grade modified trypsin (Promega). Extracted peptides were lyophilized to dryness and resuspended in 12 μL of 0.2% formic acid/2% acetonitrile. Samples were subjected to chromatographic separation on a Waters NanoAquity UPLC equipped with a 1.7 µm BEH130 C18 75 µm I.D. X 250 mm reversed-phase column. The mobile phase consisted of (A) 0.1% formic acid in water and (B) 0.1% formic acid in acetonitrile. Following a 3 µL injection, peptides were trapped for 3 min on a 5 µm Symmetry C18 180 µm I.D. X 20 mm column at 5 µl/min in 99.9% A. The analytical column was then switched in-line, and a linear elution gradient of 5% B to 40% B was performed over 30 min at 400 nL/min. The analytical column was connected to a fused silica PicoTip emitter (New Objective, Cambridge, MA) with a 10 µm tip orifice and coupled to an Oribtrap Fusion Lumos mass spectrometer (Thermo) through an electrospray interface operating in a data-dependent mode of acquisition. The instrument was set to acquire a precursor MS scan from m/z 200-1500 in the Orbitrap at r = 120,000 with MS/MS spectra acquired in the Ion Trap with AGC setting of 1e4 and 100ms. For all experiments, stepped HCD energy settings were 28.5, 30, 31.5v, and a 20 s dynamic exclusion was employed for previously fragmented precursor ions.
Raw LC-MS/MS data files were processed in Proteome Discoverer (Thermo Scientific) and then submitted to independent Mascot searches (Matrix Science) against a SwissProt database (Human taxonomy) containing both forward and reverse entries of each protein (20,371 forward entries). Search tolerances were 5 ppm for precursor ions and 0.8 Da for product ions using trypsin specificity with up to two missed cleavages. Carbamidomethylation (+57.0214 Da on C) was set as a fixed modification, whereas oxidation (+15.9949 Da on M) was considered dynamic mass modifications. All searched spectra were imported into Scaffold (v4.3, Proteome Software), and scoring thresholds were set to achieve a peptide false discovery rate of 1% using the PeptideProphet algorithm.
Iso-TMT (ABPP) sample preparation
Iso-TMT sample preparation was conducted as previously detailed45. WZR12 cells were seeded for a total of six biological replicates and allowed to adhere overnight. The next day, cells were treated with either vehicle (DMSO) (3 samples) or a 2 µM dose of MCB-613 (3 samples) for 1 h before collection by scraping into ice-cold PBS, centrifugation, and snap freezing in liquid nitrogen. Frozen cell pellets were lysed in DPBS supplemented with Benzonase (Santa Cruz) and protease inhibitors (Roche) using a chilled bath sonicator (Q700, Qsonica) and centrifuged for 3 min at 300 × g. Protein concentrations were determined by BCA assay (Thermo Fisher Scientific), and 50 µg of protein extracts were used per compound treatment. Lysates were treated with vehicle (DMSO) or 500 µM of KB02, KB03, or KB05 (Sigma-Aldrich) for 1 h, followed by 1 mM DBIA treatment for another hour.
Following DBIA incubation, lysates were reduced with 5 mM 5-tris(2-carboxyethyl)phosphine hydrochloride (TCEP) (Sigma-Aldrich) for 2 min at room temperature, then alkylated using 20 mM chloroacetamide (Sigma-Aldrich) for 30 min in the dark at room temperature. Proteins were precipitated using SP3 magnetic beads. Briefly, SP3 magnetic beads (Cytiva) were prewashed with LC-MS grade water (Sigma Aldrich), and 250 µg of combined SP3 beads (1:1, hydrophobic:hydrophilic) and LC-MS grade ethanol (Sigma Aldrich) were added to each sample to achieve a final concentration of 50% ethanol. SP3 incubation was performed for 30 min at room temperature, after which the beads were washed three times with 80% HPLC grade ethanol (Sigma Aldrich) and then resuspended with 175 µL of Trypsin/Lys-C (1 µg, Thermo Fisher Scientific) in 200 mM EPPS (Sigma Aldrich), pH 8.4, 5 mM CaCl2. Proteins were digested overnight (16 h) at 37 °C, and digested peptides were enriched with streptavidin magnetic beads (Cytiva) for 1 h at room temperature. Beads were then washed three times with DPBS and twice with HPLC-grade water (Sigma Aldrich). Peptides were eluted with 50% acetonitrile (Sigma Aldrich), 0.1% formic acid (Thermo Fisher Scientific), and dried using a Speedvac (Thermo Fisher Scientific).
Cysteine-enriched peptides were reconstituted with 30% acetonitrile, 70% 200 mM EPPS pH 8.4, and labeled with 25 µg of TMT reagent (Thermo Fisher Scientific) per channel for 75 min at room temperature with rotation. Labeling was terminated by the addition of 5% hydroxylamine (Acros Organics) for 15 min followed by the addition of 10% formic acid. Peptides from four different cancer cell lines derived from the same lineage, treated with vehicle or three scout fragments, were labeled with 16 TMT tags, pooled, and dried using a Speedvac (Thermo Fisher Scientific). Peptides were then desalted with stage tips using the following procedure: peptides were reconstituted with 5% acetonitrile/0.1% formic acid and loaded onto C18 Micro Spin columns (Nest Group) pre-equilibrated with LC/MS-grade methanol (Fisher Chemical) and LC/MS-grade water containing 0.1% formic acid. C18 spin columns were washed 10 times with LC/MS grade water containing 0.1% formic acid and subsequently eluted with 80% acetonitrile, 0.1% formic acid, and dried using a Speedvac (Thermo Fisher Scientific).
All mass spectrometry (ABPP) samples were analyzed as previously described using an Orbitrap Eclipse™ Tribrid™ Mass Spectrometer coupled with an Easy NanoLC-1200 system (Thermo Fisher Scientific)45. Peptides were separated on a 75-µm capillary column packed with 50 cm of C18 resin (2 µm, 100 Å; Thermo Fisher Scientific) using a 180-min gradient of 4–35% acetonitrile in 0.1% formic acid, with a flow rate of 300 nL/min. The eluted peptides were acquired by data-dependent acquisition and quantified using the synchronous precursor selection (DDA-SPS-MS3) method for TMT quantification. MS1 spectra were acquired in the scan range of 400–1400 m/z at an Orbitrap resolution of 120,000 with a maximum injection time of 50 ms and high-field asymmetric-waveform ion-mobility spectrometry (FAIMS) values set at -40, -50, and -70 compensation voltage (CV). MS2 spectra were acquired by selecting the top twenty most abundant features via collision-induced dissociation in the ion trap with an automatic gain control (AGC) setting of 10 K, a quadrupole isolation width of 0.7 m/z, and a maximum ion accumulation time of 50 ms. These spectra were relayed in real-time to an external computer for online database searching using Comet real-time searching (RTS) with a database that included cell line mutations (DepMap)100 and human protein databases (release_20210506)101,102. Both the real-time search and the final search utilized the same forward and reverse-sequence human protein databases (Uniprot). Peptides were filtered using simple initial parameters: they did not match a reverse-sequence, contained TMTPro16 isobaric tags, had a maximum PPM error <50, a minimum PPM error >5, and a minimum ΔCorr of 0.10. If peptide spectra met these criteria, an SPS–MS3 scan was performed using up to 20 beta- and gamma-type fragment ions as precursors with an AGC of 250 K for a maximum of 250 ms, and a normalized HCD collision energy setting of 55 (TMTPro16).
Peptide searches were performed in Proteome Discoverer (2.5, Thermo Fisher Scientific) against the UniProt human protein database (release_20210506) with the following parameters: up to two missed cleavages, 20 ppm precursor tolerance, 0.6 Da fragment ion tolerance, and fully tryptic peptides were allowed with a minimum length of 6. Static modification of TMTPro16 on lysine and peptide N-termini (+304.2071 Da) and carbamidomethylation of cysteine residues (+57.0214 Da), along with oxidation of methionine (+15.9949 Da) and DIBA on cysteine residues (for iso-TMT samples) (+239.1634) as variable modifications were permitted. Results were filtered to a peptide false discovery rate (FDR) of 1%. For the TMT reporter ion quantification, all identified peptide spectral matches (PSMs) from the MS3 scans were extracted by an in-house program, and the reporter ion intensities were adjusted for impurity correction according to the manufacturer's specifications. For quantification of each MS3 spectrum, a total sum signal-to-noise of all reporter ions of 100 (TMT16-plex) was used. Engagement scores were calculated as previously described45.
Generation of HA-Clover KEAP1 constructs
Full-length KEAP1 WT was amplified out of the pET28a-His6-KEAP1 vector using the Phusion Hot Start Flex 2x Master Mix (NEB) according to the manufacturer’s instructions and appended with EcoRV and NotI restriction sites using commercially synthesized primers (IDT) (Supplementary Table 2). Amplified full-length KEAP1 WT and the HA-Clover construct (Addgene #163336) were digested with the EcoRV and NotI enzymes (NEB) at 37 °C for 1 h before heat inactivation at 65 °C for 5 min. Digestion fragments were purified by gel electrophoresis, excised, and extracted with the QIAQuick gel extraction kit (Qiagen). The fragments were combined using T4 DNA ligase (NEB) according to the manufacturer’s instructions. To generate KEAP1 truncations, site-directed mutagenesis was performed to introduce nonsense mutations at positions 61, 179, 315, and 598 using commercially synthesized primers (IDT) (Supplementary Table 2). All constructs were validated by Sanger sequencing (Eton Biosciences).
Transient transfection of HA-Clover KEAP1 constructs
293FT cells were transiently transfected with 3 µg of each of the HA-Clover KEAP1 constructs diluted into Opti-MEM (Gibco) and supplemented with Lipofectamine 2000 transfection reagent (Invitrogen). After 24 h, media was refreshed. After another 24 h, cells were treated with vehicle (DMSO) or 2 µM MCB-613 for 1 h before collection in ice-cold PBS and preparation for western blot.
GFP-Trap assay
293FT cells were transfected with HA Clover KEAP1 WT as described above. After 48 h, the cells were washed once in ice-cold PBS and collected in ice-cold RIPA buffer (Sigma). The cells were incubated on ice for 30 min with intermittent vortexing, and the resultant lysates were subjected to centrifugation at 9600 × g for 10 min at 4 °C. A fraction of the supernatant was reserved for input and prepared for western blotting as described above. The remaining supernatant was exposed to GFP-Trap agarose beads (Proteintech) and rotated end-over-end at 4 °C overnight before washing 5 times with ice-cold RIPA buffer. The beads were then equilibrated in room-temperature binding buffer (25 mM Tris (pH 7.5), 100 mM NaCl, 0.1% NP-40, 1 mM DTT, 5% glycerol). Ten microgram of purified His6-KEAP1 was added, and the samples were rotated end-over-end at room temperature for 15 min. MCB-613 (2, 20, or 200 µM) was added, and the samples were rotated for another 15 min. Samples were washed 3 times in room-temperature binding buffer, re-suspended in 80 µl of NuPAGE LDS Sample Buffer (4X), boiled at 95 °C for 5 min, and analyzed by western blot. Targets were detected using anti-GFP (#2555, CST) and anti-His (#2365, CST) primary antibodies.
Modeling the covalent complex of KEAP1 and MCB-613
Modeling was performed using the RDKit cheminformatics toolkit and the RosettaLigand docking protocol. We used RDKit to generate 100 conformers of MCB-613, modeled in its reacted form with two lysine atoms at each covalent reaction site (SMILES: CCC1CC(C(C2=CN=CC=C2)NC)C(C(C1)C(NC)C3=CN=CC=C3)=O). The conformers were generated using the AllChem.EmbedMolecule function, followed by a minimization step using UFF force field calculations with the Chem.rdForceFieldHelpers module. Conformers with RMSD < 0.5 Å to any of the previously sampled conformers were discarded. The lysine Cε atom was then deleted, and the lysine Nζ was replaced with a vanadium atom, which is interpreted as a virtual atom. To parametrize the ligand for use in Rosetta, we used the molfile_to_params.py script provided in the Rosetta release, and modified the resulting params file to allow covalent binding103.
To model the complex, we first generated a dimeric structure of KEAP1 based on the monomer in PDB ID: 4CXT and a generated symmetry mate using PyMol. Each Lys97 residue was replaced with a covalently-linked variant of lysine103, and the dimeric structure was prepacked using the PackRotamersMover and MinMover protocols through the RosettaScripts interface. The ligand was then added to the structure and docked using the RosettaLigand docking XML protocol104, to generate 5,000 models. The center of the two Lys97 NZ atoms in the two different units was used as the starting position of the ligand. To enforce the covalent binding, we defined covalent constraints for each binding site105, using the HARMONIC score function, centered at 0 and with a standard deviation of 0.3. Since each electrophilic warhead can bind to either Lys97, we used two sets of MultiConstraints within an AmbiguousConstraints block to allow both binding options. The selected model received a low atom_pair_constraint score (<4), which indicates that it fits well into the expected covalent bond geometry, and was ranked 4th best by interface score.
Inducible knockdown of KEAP1
Hairpins were prepared and cloned into the pLKO-Tet-On vector (Addgene #21915) as previously described106,107. Briefly, forward and reverse sequences were synthesized commercially (IDT), diluted in 10x annealing buffer (1M NaCl, 100 mM Tris-HCl, pH 7.4), heated to 95 °C to induce annealing, and allowed to cool to room temperature. Annealed oligos were diluted 1:400 in 0.5x annealing buffer before T4 ligation into gel-purified, AgeI- and EcoRI-digested pLKO-Tet-On, followed by transformation into Stbl3 bacteria and validation by Sanger sequencing for difficult templates (Eton Bioscience). They were then transfected into 293FT cells and transduced into drug-resistant NSCLC cells. After 48 h of puromycin selection, puromycin was removed, and cells were cultured in their usual medium without doxycycline for at least five days. 24 h prior to seeding, media were supplemented with doxycycline to a concentration of 100 ng/μl. The next day, cells were seeded and allowed to adhere overnight. Once cells had adhered, they were treated with either vehicle or MCB-613 to a concentration of 1.25 μM for either 2 or 4 h before collection, lysis, and western blotting as described above.
GSH reactivity assay
Five microliters of a 20 mM DMSO solution of the electrophile was added to a 1000 μL solution of 1 mM GSH in a 25 mM HEPES pH = 8 buffer (final electrophile concentration was 200 μM). The reaction mixture was left at room temperature. At selected timepoints, a sample from the reaction mixture was mixed with a formic acid (FA) solution to obtain a final 0.4% of FA in order to quench the reaction. Samples were then injected into the LC/MS. Runs were performed on a Waters ACUITY UPLC BEH instrument in positive ion mode using electrospray ionization. UPLC separation used a C18 column (2.1 mm i.d., 50 mm length). The column was held at 40 °C and the autosampler at 10 °C. Mobile phase A was 0.1% FA in water, and mobile phase B was 0.1% FA in acetonitrile. The run flow was 0.4 mL/min. The gradient used was 99% A for 1 min, then increasing linearly to 95% B for 5.5 min, holding at 95% B for 0.75 min, changing to 1% B and holding for 1 min. Reaction progress quantification was performed by calculating the peak area of the starting material (unlabeled) at each time point, normalized to the starting time point (t = 0). Reaction half-life values were derived from a one-phase exponential decay curve fit using Prism.
NAL reactivity assay
NAL (N-Acetyl-L-lysine methyl ester) was dissolved to 100 mM in water and diluted to 2 mM in a 25 mM HEPES, 150mM NaCl, pH=8 buffer. Two microliters of a 10 mM electrophile DMSO solution were diluted to 100 μL using the prepared NAL solution and let react at room temperature. (Final concentration in the reaction mixture: 5 mM NAC and 200 μM electrophile.) Reaction was conducted at room temperature. At selected time points, 10 μL samples were mixed with 35 μL of 0.1% TFA and injected to LC/MS). Reaction progress quantification was performed similarly to described for the GSH assay.
NAC reactivity assay
Two microliters of a 10 mM electrophile DMSO solution were diluted to 50 μL with reaction buffer (NaPi 25 mM, pH = 7.5, 50% ACN). Five microliters of this sample and 5 μL of reaction buffer were mixed with 35 μL of 0.1% TFA in water for the t = 0 sample. Then, NAC (N-Acetyl-L-cysteine methyl ester) was dissolved to 100 mM in water and diluted to 2 mM in the reaction buffer. Forty microliters of NAC solution was added to each sample (final concentration in the reaction mixture: 1 mM NAC and 200 μM electrophile). Reaction was conducted at room temperature. At selected time points, 10 μL samples were mixed with 35 μL of 0.1% TFA and injected into LC/MS. Reaction progress quantification was performed similarly to described for the GSH assay.
DCFDA assay
Cells were seeded into 10-cm dishes and allowed to adhere overnight. The next day, media was aspirated, and cells were washed once with PBS before incubation with 5 µM DCFDA for 30 min at 37 °C. After incubation, DCFDA was removed, and the cells were washed once more with PBS. Cells were then incubated in 1.25 µM MCB-613 for 2 h at 37 °C before fluorescence detection on a Tecan Infinite M1000 plate reader. Background signal from media was subtracted, and treatment wells were normalized to vehicle controls.
Statistics and Reproducibility
Throughout the manuscript, statistical methods, statistical measures, and numbers of replicates and/or repeated experiments have been noted in the figure legends, as well as at times within the Methods. Although exact p-values are not reported in main text figures and/or figure legends due to space constraints, these can be found in the Source Data file. Exact p values for Supplementary Figs. are reported within the Supplementary Fig. legends.
Data availability
The DNA sequencing data generated in this study have been deposited in the Gene Expression Omnibus (GEO) database under the accession code GSE262214. RNA sequencing data generated in this study have been deposited in the GEO database under accession code GSE262212. The proteomics data generated in this study have been deposited in the PRIDE and MassIVE databases under the accession codes PXD071547, MSV000100060, PXD071428 [https://massive.ucsd.edu/ProteoSAFe/dataset.jsp?task=cb90831dc0124296b9144b4f700fa47d], and MSV000097077, PXD060588 [https://massive.ucsd.edu/ProteoSAFe/dataset.jsp?task=175ddadd3cdf45799fd6e210fc3b27bc]. Source data are provided with this paper, and all remaining data are available within the Source Data file, the Article itself, or the Supplementary Information. Source data are provided with this paper.
References
Soria, J.-C. et al. Osimertinib in untreated EGFR -mutated advanced non–small-cell lung cancer. N. Engl. J. Med. 378, 113–125 (2018).
SS, R. et al. Overall survival with osimertinib in untreated, EGFR-mutated advanced NSCLC. N. Engl. J. Med. 382, 41–50 (2020).
O, T., MJ, H., J, P., DW, E. & F, M. Drug resistance to EGFR inhibitors in lung cancer. Chemotherapy 61, 223–235 (2016).
Guthrie, R., Loebeck, M. E., Hillman, M. J. & Zgorzynski, F. Tests for collateral sensitivity or cross resistance in bacterial mutants resistant to amethopterin or purine analogs. Cancer Res. 18, 319–334 (1958).
Imamovic, L. et al. Drug-driven phenotypic convergence supports rational treatment strategies of chronic infections. Cell 172, 121–134.e14 (2018).
Xu, C. et al. Development of resistance to eravacycline by klebsiella pneumoniae and collateral sensitivity-guided design of combination therapies. Microbiol. Spectr. 10, e0139022 (2022).
Wang, L. et al. An acquired vulnerability of drug-resistant melanoma with therapeutic potential. Cell 173, 1413–1425.e14 (2018).
Singleton, K. R. et al. Melanoma therapeutic strategies that select against resistance by exploiting MYC-driven evolutionary convergence. Cell Rep. 21, 2796–2812 (2017).
Lin, K. H. et al. Using antagonistic pleiotropy to design a chemotherapy-induced evolutionary trap to target drug resistance in cancer. Nat. Genet. 52, 408–417 (2020).
Zhao, B. et al. Exploiting temporal collateral sensitivity in tumor clonal evolution. Cell 165, 234–246 (2016).
Dhawan, A. et al. Collateral sensitivity networks reveal evolutionary instability and novel treatment strategies in ALK mutated non-small cell lung cancer. Sci. Rep. 7, 1232 (2017).
Keller, R. R. & Gunther, E. J. Evolution of relapse-proficient subclones constrained by collateral sensitivity to oncogene overdose in Wnt-driven mammary cancer. Cell Rep. 26, 893–905.e4 (2019).
Wang, X., Zhang, H. & Chen, X. Drug resistance and combating drug resistance in cancer. Cancer Drug Resist. 2, 141–160 (2019).
Acar, A. et al. Exploiting evolutionary steering to induce collateral drug sensitivity in cancer. Nat. Commun. 11, 1923 (2020).
Imamovic, L. & Sommer, M. O. A. Use of collateral sensitivity networks to design drug cycling protocols that avoid resistance development. Sci. Transl. Med. 5, 204ra132 (2013).
Cortot, A. B. et al. Resistance to irreversible EGF receptor tyrosine kinase inhibitors through a multistep mechanism involving the IGF1R pathway. Cancer Res. 73, 834–843 (2013).
Ercan, D. et al. Amplification of EGFR T790M causes resistance to an irreversible EGFR inhibitor. Oncogene 29, 2346–2356 (2010).
Ercan, D. et al. Reactivation of ERK signaling causes resistance to EGFR kinase inhibitors. Cancer Discov. 2, 934–947 (2012).
Ogino, A. et al. Emergence of epidermal growth factor receptor T790M mutation during chronic exposure to gefitinib in a non small cell lung cancer cell line. Cancer Res. 67, 7807–7814 (2007).
JA, E. et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science 316, 1039–1043 (2007).
Turke, A. B. et al. Pre-existence and clonal selection of MET amplification in EGFR mutant NSCLC. Cancer Cell 17, 77 (2010).
KE, W. et al. A mechanism of resistance to gefitinib mediated by cellular reprogramming and the acquisition of an FGF2-FGFR1 autocrine growth loop. Oncogenesis 2, e39 (2013).
AN, H. et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat. Med. 22, 262–269 (2016).
Sharma, R. A., Steward, W. P. & Gescher, A. J. Pharmacokinetics and pharmacodynamics of curcumin. Adv. Exp. Med. Biol. 595, 453–470 (2007).
Cheng, D. et al. Pharmacokinetics, pharmacodynamics, and PKPD modeling of curcumin in regulating antioxidant and epigenetic gene expression in healthy human volunteers. Mol. Pharm. 16, 1881–1889 (2019).
Sinha, S. et al. Pharmacokinetic evaluation of Chalcone derivatives with antimalarial activity in New Zealand White Rabbits. BMC Res Notes 14, 264 (2021).
Wang, L. et al. Characterization of a steroid receptor coactivator small molecule stimulator that overstimulates cancer cells and lead to cell stress and death. Cancer Cell 28, 240–252 (2015).
Padmanabhan, A. et al. USP15-dependent lysosomal pathway controls p53-R175H turnover in ovarian cancer cells. Nat. Commun. 9, 1270 (2018).
Selvaraju, K. et al. Cytotoxic unsaturated electrophilic compounds commonly target the ubiquitin proteasome system. Sci. Rep. 9, 1–17 (2019).
Bai, N., Roder, H., Dickson, A. & Karanicolas, J. Isothermal analysis of thermofluor data can readily provide quantitative binding affinities. Sci. Rep. 9, 1–15 (2019).
Cimmperman, P. et al. A quantitative model of thermal stabilization and destabilization of proteins by ligands. Biophys. J. 95, 3222 (2008).
Jafari, R. et al. The cellular thermal shift assay for evaluating drug target interactions in cells. Nat. Protoc. 9, 2100–2122 (2014).
Bollong, M. J. et al. A metabolite-derived protein modification integrates glycolysis with KEAP1–NRF2 signalling. Nature 562, 600–604 (2018).
Zhang, D. D. & Hannink, M. Distinct cysteine residues in Keap1 are required for Keap1-dependent ubiquitination of Nrf2 and for stabilization of Nrf2 by chemopreventive agents and oxidative stress. Mol. Cell Biol. 23, 8137 (2003).
Dang, D. T. Molecular approaches to protein dimerization: opportunities for supramolecular chemistry. Front. Chem. 10, 62 (2022).
Shin, J. W. et al. Curcumin induces stabilization of Nrf2 protein through Keap1 cysteine modification. Biochem. Pharm. 173, 113820 (2020).
Cleasby, A. et al. Structure of the BTB domain of Keap1 and its interaction with the triterpenoid antagonist CDDO. PLoS ONE 9, e98896 (2014).
Davies, T. G. et al. Monoacidic inhibitors of the kelch-like ECH-associated protein 1: nuclear factor erythroid 2-related factor 2 (KEAP1:NRF2) protein-protein interaction with high cell potency identified by fragment-based discovery. J. Med. Chem. 59, 3991–4006 (2016).
Brennan, M. S. et al. Dimethyl fumarate and monoethyl fumarate exhibit differential effects on KEAP1, NRF2 activation, and glutathione depletion in vitro. PLoS ONE 10, e0120254 (2015).
Dinkova-Kostova, A. T. et al. Direct evidence that sulfhydryl groups of Keap1 are the sensors regulating induction of phase 2 enzymes that protect against carcinogens and oxidants. Proc. Natl. Acad. Sci. USA 99, 11908 (2002).
Ramos-Gomez, M. et al. From the cover: sensitivity to carcinogenesis is increased and chemoprotective efficacy of enzyme inducers is lost in nrf2 transcription factor-deficient mice. Proc. Natl. Acad. Sci. USA 98, 3410 (2001).
Talalay, P., De Long, M. J. & Prochaska, H. J. Identification of a common chemical signal regulating the induction of enzymes that protect against chemical carcinogenesis. Proc. Natl. Acad. Sci. USA 85, 8261 (1988).
Yamamoto, T. et al. Physiological significance of reactive cysteine residues of Keap1 in determining Nrf2 activity. Mol. Cell Biol. 28, 2758 (2008).
Wakabayashi, N. et al. Protection against electrophile and oxidant stress by induction of the phase 2 response: Fate of cysteines of the Keap1 sensor modified by inducers. Proc. Natl. Acad. Sci. USA 101, 2040 (2004).
Takahashi, M. et al. DrugMap: a quantitative pan-cancer analysis of cysteine ligandability. Cell 187, 2536–2556.e30 (2024).
Reinecke, M. et al. Chemical proteomics reveals the target landscape of 1,000 kinase inhibitors. Nat. Chem. Biol. 20, 577–585 (2024).
Cheng, Y. et al. Covalent modification of Keap1 at Cys77 and Cys434 by pubescenoside a suppresses oxidative stress-induced NLRP3 inflammasome activation in myocardial ischemia-reperfusion injury. Theranostics 11, 861 (2021).
Abbasov, M. E. et al. A proteome-wide atlas of lysine-reactive chemistry. Nat. Chem. 13, 1081–1092 (2021).
Ryan, E. M. et al. Activity-based acylome profiling with N-(Cyanomethyl)-N-(phenylsulfonyl)amides for targeted lysine acylation and post-translational control of protein function in cells. J. Am. Chem. Soc. 146, 27622–27643 (2024).
Wan, C. et al. A pyridinium-based strategy for lysine-selective protein modification and chemoproteomic profiling in live cells. Chem. Sci. 15, 5340–5348 (2024).
Zhou, M. et al. Global profiling lysine reactivity and ligandability with oxidant-triggered bioconjugation chemistry. Angew. Chem. Int. Ed. 64, e202418473 (2025).
Baird, L. & Yamamoto, M. The molecular mechanisms regulating the KEAP1-NRF2 Pathway. Mol. Cell Biol. 40, e00099–20 (2020).
Li, H. et al. Decreased glutathione biosynthesis contributes to EGFR T790M-driven erlotinib resistance in non-small cell lung cancer. Cell Discov. 2, 1–14 (2016).
Weiss-Sadan, T. et al. NRF2 activation induces NADH-reductive stress, providing a metabolic vulnerability in lung cancer. Cell Metab. 35, 487–503.e7 (2023).
Okon, I. S., Coughlan, K. A., Zhang, M., Wang, Q. & Zou, M. H. Gefitinib-mediated reactive oxygen specie (ROS) instigates mitochondrial dysfunction and drug resistance in lung cancer cells. J. Biol. Chem. 290, 9101 (2015).
Kalkavan, H. et al. Sublethal cytochrome c release generates drug-tolerant persister cells. Cell 185, 3356–3374.e22 (2022).
Pakos-Zebrucka, K. et al. The integrated stress response. EMBO Rep. 17, 1374 (2016).
Du, G. et al. Exploring the target scope of KEAP1 E3 ligase-based PROTACs. Cell Chem. Biol. 29, 1470–1481.e31 (2022).
Wang, F. et al. Cell-permeable PROTAC degraders against KEAP1 efficiently suppress hepatic stellate cell activation through the antioxidant and anti-inflammatory pathway. ACS Pharm. Transl. Sci. 6, 76–87 (2023).
Park, S. Y. et al. Development of KEAP1-targeting PROTAC and its antioxidant properties: in vitro and in vivo. Redox Biol. 64, 102783 (2023).
Chen, H. et al. Design and characterization of a heterobifunctional degrader of KEAP1. Redox Biol. 59, 102552 (2023).
Lu, M. et al. Bivalent inhibitors of the BTB E3 ligase KEAP1 enable instant NRF2 activation to suppress acute inflammatory response. Cell Chem Biol. 31, 1188–1202.e10 (2023).
Hacker, S. M. et al. Global profiling of lysine reactivity and ligandability in the human proteome. Nat. Chem. 9, 1181 (2017).
Wojtaszek, J. L. et al. A small molecule targeting mutagenic translesion synthesis improves chemotherapy. Cell 178, 152–159.e11 (2019).
Ouyang, Y. et al. Design, synthesis, and evaluation of o-(Biphenyl-3-ylmethoxy)nitrophenyl derivatives as PD-1/PD-L1 inhibitors with potent anticancer efficacy in vivo. J. Med. Chem. 64, 7646–7666 (2021).
Wang, T. et al. Novel biphenyl pyridines as potent small-molecule inhibitors targeting the programmed cell death-1/programmed cell death-ligand 1 interaction. J. Med. Chem. 64, 7390–7403 (2021).
Liu, L. et al. Syntheses, biological evaluations, and mechanistic studies of benzo[c][1,2,5]oxadiazole derivatives as potent PD-L1 inhibitors with in vivo antitumor activity. J. Med. Chem. 64, 8391–8409 (2021).
Muszak, D. et al. Terphenyl-based small-molecule inhibitors of programmed cell death-1/programmed death-ligand 1 protein-protein interaction. J. Med. Chem. 64, 11614–11636 (2021).
Zak, K. M. et al. Structural basis for small molecule targeting of the programmed death ligand 1 (PD-L1). Oncotarget 7, 30323–30335 (2016).
Song, Z. et al. Design, synthesis, and pharmacological evaluation of biaryl-containing PD-1/PD-L1 interaction inhibitors bearing a unique difluoromethyleneoxy linkage. J. Med. Chem. 64, 16687–16702 (2021).
Wu, H. et al. Molecular glues modulate protein functions by inducing protein aggregation: A promising therapeutic strategy of small molecules for disease treatment. Acta Pharm. Sin. B 12, 3548–3566 (2022).
Mullany, L. K. et al. A steroid receptor coactivator stimulator (MCB-613) attenuates adverse remodeling after myocardial infarction. Proc. Natl. Acad. Sci. USA 117, 31353–31364 (2020).
McClendon, L. K. et al. A steroid receptor coactivator small molecule ‘stimulator’ attenuates post-stroke ischemic brain injury. Front. Mol. Neurosci. 15, 1055295 (2022).
McClendon, L. K. et al. Transcriptional coactivation of NRF2 signaling in cardiac fibroblasts promotes resistance to oxidative stress. J. Mol. Cell Cardiol. 194, 70–84 (2024).
Cao, S. S. & Kaufman, R. J. Endoplasmic reticulum stress and oxidative stress in cell fate decision and human disease. Antioxid. Redox Signal. 21, 396 (2014).
Komatsu, M. et al. The selective autophagy substrate p62 activates the stress responsive transcription factor Nrf2 through inactivation of Keap1. Nat. Cell Biol. 12, 213–223 (2010).
Uribe-Carretero, E. et al. Loss of KEAP1 causes an accumulation of nondegradative organelles. Antioxidant 11, 1398 (2022).
Taguchi, K. et al. Keap1 degradation by autophagy for the maintenance of redox homeostasis. Proc. Natl. Acad. Sci. USA 109, 13561–13566 (2012).
Zhu, L. et al. Chaperone-mediated autophagy degrades Keap1 and promotes Nrf2-mediated antioxidative response. Aging Cell 21, e13616 (2022).
He, C. H. et al. Identification of activating transcription factor 4 (ATF4) as an Nrf2-interacting protein. J. Biol. Chem. 276, 20858–20865 (2001).
Miyamoto, N. et al. Transcriptional regulation of activating transcription factor 4 under oxidative stress in retinal pigment epithelial ARPE-19/HPV-16 cells. Investig. Ophthalmol. Vis. Sci. 52, 1226–1234 (2011).
Sarcinelli, C. et al. ATF4-dependent NRF2 transcriptional regulation promotes antioxidant protection during endoplasmic reticulum stress. Cancers 12, 569 (2020).
Crinelli, R. et al. Activation of NRF2 and ATF4 signaling by the pro-glutathione molecule I-152, a Co-Drug of N-acetyl-cysteine and cysteamine. Antioxidant 10, 1–23 (2021).
Kreß, J. K. C. et al. The integrated stress response effector ATF4 is an obligatory metabolic activator of NRF2. Cell Rep. 42, 112724 (2023).
Solis, L. M. et al. Nrf2 and Keap1 abnormalities in non-small cell lung carcinoma and association with clinicopathologic features. Clin. Cancer Res. 16, 3743 (2010).
R, B.-R. Importance of the Keap1-Nrf2 pathway in NSCLC: Is it a possible biomarker?. Biomed. Rep. 9, 375–382 (2018).
EB, K. et al. KEAP1 loss modulates sensitivity to kinase targeted therapy in lung cancer. eLife 6, e18970 (2017).
Park, S. H. et al. Resistance to gefitinib and cross-resistance to irreversible EGFR-TKIs mediated by disruption of the Keap1-Nrf2 pathway in human lung cancer cells. FASEB J. 32, 5862–5873 (2018).
Hellyer, J. A. et al. Impact of KEAP1/NFE2L2/CUL3 mutations on duration of response to EGFR tyrosine kinase inhibitors in EGFR mutated non-small cell lung cancer. Lung Cancer 134, 42–45 (2019).
Martz, C. A. et al. Systematic identification of signaling pathways with potential to confer anticancer drug resistance. Sci. Signal. 7, ra121–ra121 (2014).
Chen, Y. C. & Zhang, C. A chemoproteomic method for identifying cellular targets of covalent kinase inhibitors. Genes Cancer 7, 148–153 (2016).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal. Chem. 68, 850–858 (1996).
Peng, J. & Gygi, S. P. Proteomics: the move to mixtures. J. Mass Spectrom. 36, 1083–1091 (2001).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. G3 Genes Genomes. Genet. 7, 2719–2727 (2017).
JG, D. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
CH, C. et al. Improved design and analysis of CRISPR knockout screens. Bioinformatics 34, 4095–4101 (2018).
Parvez, S. et al. Substoichiometric hydroxynonenylation of a single protein recapitulates whole-cell-stimulated antioxidant response. J. Am. Chem. Soc. 137, 10 (2015).
Niesen, F. H., Berglund, H. & Vedadi, M. The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat. Protoc. 2, 2212–2221 (2007).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576.e16 (2017).
Li, J. et al. TMTpro reagents: a set of isobaric labeling mass tags enables simultaneous proteome-wide measurements across 16 samples. Nat. Methods 17, 399–404 (2020).
Zhang, T., Gygi, S. P. & Paulo, J. A. Temporal proteomic profiling of SH-SY5Y differentiation with retinoic acid using FAIMS and real-time searching. J. Proteome Res. 20, 704–714 (2021).
Gabizon, R. et al. A simple method for developing lysine targeted covalent protein reagents. Nat. Commun. 14, 7933 (2023).
Lemmon, G. & Meiler, J. Rosetta Ligand docking with flexible XML protocols. Methods Mol. Biol. 819, 143–155 (2012).
Tivon, B. et al. Covalent flexible peptide docking in Rosetta. Chem. Sci. 12, 10836–10847 (2021).
Wee, S. et al. PTEN-deficient cancers depend on PIK3CB. Proc. Natl. Acad. Sci. USA 105, 13057–13062 (2008).
Wiederschain, D. et al. Single-vector inducible lentiviral RNAi system for oncology target validation. Cell Cycle 8, 498–504 (2009).
Acknowledgements
We would like to thank Pasi Janne (Harvard Medical School) for the PFR3, GR4, and WZR12 cell lines. We also thank Aaron Hata (Harvard Medical School) for the patient-derived MGH134 cell line. We thank Bert O’Malley (Baylor College of Medicine) for MCB-613 and for his continued support and feedback on this project. We thank the Duke Functional Genomics Core for their assistance with high-throughput pharmacologic screening. We thank Ross Tomaino (Taplin Mass Spectrometry Facility, Harvard Medical School) and Erik Soderblom (Proteomics and Metabolics Core Facility, Duke University School of Medicine) for their input and assistance on proteomic studies. We thank Aurélien Lebrun (Plateforme d’analyses et caractérisation, P.A.C. Balard, Université Montpellier) for his NMR characterization of ARM-143. Finally, we would like to thank Kevin H. Lin and Justine C. Rutter for their invaluable advice and support along the way. This work was supported by the NIH (R01CA263593 to K.C.W., R01CA246133 to A.M.P., P30CA014236 to K.C.W. and A.M.P., 1R38AI140297 to C.F.B., R37CA260062 to L.B.-P., R21CA226082 to L.B.-P., K00CA245732-04 to J.H., 5T32GM007105, F31CA243293, and F99CA264162 to B.M., F30CA247323 to C.C.S., and F31CA195967 to P.S.W.), the DOD (W81XWH-21-1-0362 and HT9425-24-1-0338 to K.C.W.), the National Science Foundation (DGE-1106401 to G.R.A.), the American Cancer Society (PF-23-1018904-01-ET to S.A.H.), the Duke Medical Scientist Training Program (T32GM007171 to C.F.B., C.C.S., and S.T.K.), the Duke Graduate School (to C.F.B.), the Triangle Center for Evolutionary Medicine (to C.F.B.), the Amgen Scholars Program (to S.G.), the French National Center for Scientific Research (to A.R.M.), the Melanoma Research Alliance (to L.B.-P.), the Ludwig Cancer Center of Harvard Medical School (to L.B.-P.), Lungevity (to L.B.-P.), ALK Positive (to L.B.-P.), V-Foundation for Cancer Research (to L.B.-P.), Mary Kay Foundation (to L.B.-P.), Paula and Rodger Riney Foundation (to L.B.-P.), the PEW-Stewart Trusts (to L.B.-P.), and Lisa and Mark Schwartz (to L.B.-P.).
Author information
Authors and Affiliations
Contributions
C.F.B., G.R.A., and P.S.W. conceptualized and designed the study. G.R.A. and K.R.S. developed drug-resistant cell line models. C.F.B., G.R.A., K.R.S., and K.N.A. performed eight-point GI50 assays. G.R.A. and P.S.W. designed, performed, and analyzed the high-throughput pharmacologic screen. H.M.H. derived resistance to Osimertinib in patient-derived cell line models. L.Q. performed and analyzed in vivo studies. A.R.M. performed chemical synthesis. K.J-G., C.F., and G.R. performed click chemistry-mediated studies. C.F.B. designed, performed, and processed the CRISPR screen, with input and analysis from C.C-S. C.F.B., K.N.A., C.T., and K.D. performed single-guide knockout and validation studies. A.H. assisted with cloning of shRNA and sgRNA constructs. C.F.B. performed protein purification, thermal shift, and in vitro binding assays with guidance and assistance from B.M. C.F.B. and K.N.A. performed immunoblotting and co-immunoprecipitation experiments. C.F.B., S.G., and T.M.H. performed the CETSA studies. C.F.B. designed and performed serial truncation studies. B.T. performed covalent complex modeling. J.P.H. performed RNA-seq analysis. C.F.B., G.R.A., and K.D. performed ROS experiments. S.H. performed and analyzed ABPP. E.L. performed the glutathione, cysteine, and lysine reactivity assays. N.L., R.B., P.A., A.M.P., L.B-P., D.M.L., and A.P. supervised various experiments and provided advice. C.F.B. and K.D. performed the review. K.C.W. supervised the study. C.F.B. wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
K.C.W. is a founder, consultant, and equity holder at Tavros Therapeutics and Celldom, an equity holder at Decrypt Bio and Simple Therapeutics, and has performed consulting work for Guidepoint Global, Bantam Pharmaceuticals, and Apple Tree Partners. D.M.L. is a paid consultant and holds an equity position in CoRegen, Inc., a biotechnology company developing MCB-613 for clinical use. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bassil, C.F., Dillon, K., Anderson, G.R. et al. EGFR inhibitor-resistant lung cancers exhibit collateral sensitivity to a covalent, cysteine-independent KEAP1 oligomerizing molecular bridge. Nat Commun 17, 1726 (2026). https://doi.org/10.1038/s41467-026-68424-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68424-1