Abstract
Cell-autonomous immunity represents evolutionarily conserved defense mechanisms present in both immune and non-immune cells. One of such mechanisms is mediated by cytoskeletal septins that entrap cytosolic bacterial pathogens within cage-like structures. To promote infection, Shigella flexneri delivers effector proteins directly into host cells via a type III secretion system. Here we demonstrate that OspC effectors enable Shigella flexneri to evade septin cage entrapment. Mechanistically, OspC catalyzes ADP-riboxanation of SEPT9 at Arg561, a site essential for stabilizing septin hetero-oligomers. Notably, Arg561 ADP-riboxanation impairs septin polymerization and hence assembly of higher-order structures, including filaments and cage-like structures. Furthermore, we provide evidence that OspC effectors act synergistically with OspG to antagonize septin cage entrapment via two distinct post-translational modifications, thereby facilitating cell-to-cell spread and intracellular replication. Overall, our work reveals the elegant strategies of bacterial pathogens to evade septin-mediated cell-autonomous immunity and offers avenues for therapeutic intervention.
Similar content being viewed by others
Introduction
Shigella spp. are Gram-negative human pathogens that cause bacillary dysentery by invading the colonic epithelium1. Shigella continues to be a major public health concern with over 160 million shigellosis cases worldwide annually, including up to 100,000 fatalities especially for children under 5 in developing countries2. Among different Shigella species, Shigella flexneri (S. flexneri) is the most common cause of shigellosis and also the most studied in research laboratories worldwide1. S. flexneri pathogenesis is largely dictated by its type III secretion system (T3SS) encoded on a large (~200 kb) virulence plasmid. The T3SS is a syringe-like molecular apparatus that functions as a channel and allows direct delivery of virulence factors (known as effector proteins) into host cells. These effectors (~30 proteins) have the capacity to subvert diverse cellular processes in a highly orchestrated manner, thereby promoting bacterial invasion as well as subsequent dissemination to neighboring host cells. Once internalized in host epithelial cells, S. flexneri actively breaks the initial phagocytic vacuole and gains direct access to its cytosolic replicative niche.
To counteract pathogen replication, host innate immune defenses and other antimicrobial processes, including cell-autonomous immunity, are activated with the ultimate goal of clearing infection. Cell-autonomous immunity refers to evolutionarily conserved defense pathways present in both professional immune cells and non-immune cells (e.g., epithelial cells) to defend against invading microbial pathogens3. Recent studies have shown that S. flexneri is strikingly adept at evading cell-autonomous immunity via dedicated T3SS effectors4. Notably, the S. flexneri effector IpaH9.8 subverts guanylate-binding protein (GBP)-mediated immunity through a mechanism dependent on its E3 ubiquitin ligase activity and proteasome-mediated degradation of GBPs5,6.
Septins are a family of highly conserved cytoskeletal proteins that can form filaments and interact with cellular membranes, actin filaments, and microtubules7,8. Human cells encode 13 septin paralogs, which are categorized into four distinct subgroups based on amino acid sequence identity, including SEPT2 (SEPT1, SEPT2, SEPT4, SEPT5), SEPT3 (SEPT3, SEPT9, SEPT12), SEPT6 (SEPT6, SEPT8, SEPT10, SEPT11, SEPT14), and SEPT78,9,10. Subunits from each of the four septin subgroups assemble into hetero-oligomeric complexes in a specific order, including hexamers (SEPT2–SEPT6–SEPT7–SEPT7–SEPT6–SEPT2) and octamers (SEPT2–SEPT6–SEPT7–SEPT9–SEPT9–SEPT7–SEPT6–SEPT2), with alternating G-interface (consisting of the GTP-binding domain) and NC-interface (comprising the amino- and carboxy-terminal regions) along the linear hetero-oligomeric complex8,11. In addition to their classical role in cell division, recent studies have also revealed septins as crucial players in host-pathogen interactions. During S. flexneri infection, septins are recruited to IcsA-mediated actin polymerization and form cage-like structures around cytosolic bacteria. These septin cages counteract S. flexneri actin-based motility and recruit the autophagic adaptors p62 and NDP52 to target bacteria for autophagic destruction12,13,14.
Recently, we uncovered a S. flexneri kinase effector OspG that modulates host ubiquitin signaling to evade septin cage entrapment. Mechanistically, OspG phosphorylates a master regulator of host cullin-RING E3 ligases (CRLs) and promotes ubiquitination of septins, thereby leading to cage destruction. As a result, significantly more cytosolic bacteria were caged by septins in the case of an ospG-deficient S. flexneri mutant, as compared to its parental WT strain15. Nevertheless, even for the ΔospG strain, the majority of intracellular bacteria were not entrapped by septin cages in infected host cells. We therefore hypothesized that S. flexneri may have evolved additional strategies to target septins and subvert cage assembly.
Here we show that the S. flexneri OspC family of T3SS effectors directly modifies septins via protein ADP-riboxanation, a recently identified derivative of classical ADP-ribosylation16,17. Furthermore, we demonstrate that SEPT9 is a physiological substrate of OspC effectors during S. flexneri infection. Notably, OspC effectors target Arg561 of SEPT9, a critical residue for stabilizing the NC-interface to maintain inter-subunit contacts within septin octamers, thereby preventing the assembly of septin cages. These findings reveal bacterial effectors that have the capacity to directly target septins and thus facilitate pathogen escape from cell-autonomous immunity.
Results
Septins are conserved substrates of the S. flexneri OspC family of effectors
The OspC family of effectors includes OspC1, OspC2, OspC3, and OspC4. These effectors all contain a C-terminal ankyrin-repeat domain (ARD) and an N-terminal ADP-riboxanase domain, overall exhibiting more than 60% sequence identity (Figure. S1a). Previously we have shown that OspC3, but not OspC1/2, inhibits host pyroptosis via ADP-riboxanation of caspase-11/417, while all three effectors (OspC1, OspC2 and OspC3) are able to modify the eukaryotic translation initiation factor 3 (eIF3) complex, leading to protein translational arrest and stress granule formation16. In addition, the OspC effectors, primarily OspC1 and OspC3, inhibit the expression of interferon-inducible genes18. To further compare the substrate spectrum of OspC effectors, we catalogued potential OspC3 targets via ADP-ribosylome profiling (Figure S1b, see methods for more experimental details) and analyzed them together with those of OspC1. In total, we ended up with 88 candidates of ADP-riboxanated substrates, of which 41 proteins were shared by both OspC3 and OspC1. Our data also suggested that OspC3 likely has substantially more exclusive targets than OspC1 (38 versus 9, see Fig. 1a and Supplementary Data 1). In agreement, immunoblotting data of OspC-expressing cells (probed with an antibody specific to ADP-ribose) revealed that OspC3 samples have more intense bands than OspC1 (Fig. 1a). Gene ontology (GO) analyses suggested that core targets of OspC1 and OspC3 are likely to be evolutionarily conserved, whereas divergent functions may have evolved for individual effectors, such as double-strand break repair for OspC3 and vesicle targeting for OspC1 (Fig. 1b). Next, we focused our attention on conserved cellular pathways that are targeted by both effectors. Consistent with our previous study, the eIF3 complex represents the most enriched GO term in OspC3 and OspC1 datasets. Notably, ‘septin complex’ and ‘septin ring’ appear as the next highly enriched terms (Fig. 1b). Indeed, three out of four septin octamer subunits (i.e., SEPT2, SEPT7 and SEPT9, Figure. S1c) exhibited higher mass spectrometry signals in cells expressing catalytically active OspCs (Fig. 1c).
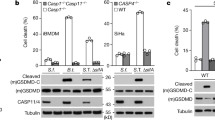
a Identification of candidate substrates. Top: Venn diagram showing the overlap of OspC1 and OspC3 candidates. Bottom: Lysates from 293T cells transfected with WT OspC or its catalytically dead mutant (EH/AA) were probed with an anti-ADP-ribose antibody. b GO enrichment analysis. Dot plot of significantly over-represented GO terms. Adjusted P values (one-sided hypergeometric test, Benjamini-Hochberg correction) are color-coded. Dot sizes indicate fold enrichment. Non-significant terms are excluded. c Heatmap of conserved substrates. Conserved candidates of OspC1 and OspC3 are shown, highlighting SEPT2, SEPT7, and SEPT9. Data represent two biological replicates. d Immunoblotting analyses of a panel of septin subunits (SEPT2, 6, 7, 9, 10, 11) for their potential ADP-riboxanation. Immunoprecipitated septins from 293T cells co-expressing GFP-OspC3 or its EH/AA mutant were blotted with indicated antibodies. WCL, whole cell lysates. e–h SEPT2, SEPT6, SEPT7 and SEPT9 were modified by OspC1 and OspC2 as well. Immunoprecipitated septin samples from 293T cells co-transfected with OspC effectors were probed with the indicated antibodies. For (a, d–h), experiments were repeated three times with similar results.
Ectopic expression of OspC effectors promotes ADP-riboxanation of septins in host cells
Considering that proteins identified in our ADP-ribosylome datasets are not necessarily modified (i.e., they may be enriched due to interaction with bona fide ADP-riboxanated substrates), we next sought to determine exactly which septins are post-translationally modified by OspC effectors. We individually expressed a panel of FLAG-tagged septin subunits (SEPT2, SEPT6, SEPT7, SEPT9, SEPT10, and SEPT11) in 293T cells together with either WT OspC3 or its catalytically dead mutant (Fig. 1d). Next, immunoprecipitated septins were probed with an ADP-ribose-specific antibody, which also recognizes protein ADP-riboxanation. Immunoblotting data demonstrated that all tested septins, except for SEPT7, were readily ADP-riboxanated by OspC3, but not the catalytically dead mutant. The inability of detecting modified SEPT7 is likely due to its lower expression levels in our transfection assays (relative to other septins). When assayed alone or with a longer exposure time, SEPT7 was indeed detected with clear modification signals (Figs. 1d, 1g and S2e). Consistent with our proteomic data, these septins were also ADP-riboxanated when they were co-expressed with the other two paralogues, OspC1 and OspC2, in host cells in an enzymatic activity-dependent manner (Fig. 1e–h).
Given that OspC3 exhibited the strongest ADP-riboxanase activity (Fig. 1a), we used it as a representative effector of the OspC family (unless otherwise specified). Previous studies have shown that the ARD domain of OspC3 is essential for caspase-4/11 recognition17. Therefore, we examined whether this is also the case for OspC3 to target septins. SEPT9 co-expressed with either WT OspC3 or its variants was assayed by immunoblotting for potential ADP-riboxanation. Similar to CASP4, SEPT9 was readily modified by full-length OspC3, whereas modification signals dropped substantially with the N-terminal domain truncated (i.e., the Nt mutant) (Figure S2a and S2b). The ARD domain of OspC3 was essential for targeting SEPT2, SEPT6, and SEPT7 (Figure S2c–e). Consistent with this, co-IP experiments demonstrated that full-length OspC3 or its ARD domain, but not the N-terminal enzymatic domain, immunoprecipitated endogenous SEPT2, SEPT6, SEPT7, and SEPT9 (Figure S2f). Together, these results indicate that SEPT9 modification depends on OspC’s catalytic N-terminal domain, the efficiency of which is enhanced by septin-binding via the C-terminal ARD domain.
OspC3 induces ADP-riboxanation of multiple residues on septins
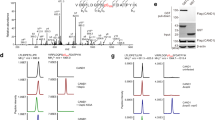
Next, we sought to identify the modification sites of septins by tandem mass spectrometry (MS/MS) analyses. As a representative member, SEPT9 co-expressed with OspC3 exhibited slower migration on SDS-PAGE, indicating potential post-translational modifications (Fig. 2a). MS/MS analyses of digested SEPT9 revealed at least four ADP-riboxanated sites, including Arg17, Arg46, Arg134, and Arg561 (Fig. 2a). While the first three sites all come from the N-terminal extension, Arg561 is located in the C-terminal α6 helices, which participate in forming salt bridges to stabilize the NC-interface. To help prioritize functional studies of these modified residues, we further analyzed the extent of ADP-riboxanation with respect to individual sites; we extracted such information from corresponding peptide intensities observed in mass spectromtery experiments (details are provided in the Methods section). For instance, the peptide T543HMQNIKDITSSIHFEAYR561*VKR564 containing ADP-riboxanated Arg561 was only detected in samples from cells expressing WT OspC3 but not its catalytically dead mutant EH/AA (Fig. 2b, c). In contrast, the unmodified peptide T543HMQNIKDITSSIHFEAYR561 was detected in both samples, while its signal dropped sharply in SEPT9 co-expressed with WT OspC3 (Fig. 2b). We estimated a modification rate of 80% based on the extent of such signal attenuation. In comparison, ADP-riboxanation rates at R17, R46, and R134 were markedly lower than that of R561 (Figure S3 and Table 1). Together, these findings suggest that R561 is the major site of ADP-riboxanation.
a Coomassie blue-stained purified 3×FLAG-SEPT9 from 293T cells co-transfected with GFP-OspC3-WT or GFP-OspC3-EH/AA prior to MS analyses. Right: ClustalW2 multiple sequence alignment of SEPT3 subgroup members. Key salt-bridge residues at the NC-interface are marked with yellow circles. ADP-riboxanation sites are highlighted in red; asterisks indicate conserved residues. The six arginine residues labeled in black were identified after the initial four sites were mutated to lysine (4RK). b Extracted ion chromatograms of an unmodified peptide, an Arg561-containing peptide and its ADP-riboxanated form from immunoprecipitated SEPT9 in (a). c Tandem mass spectrum of the ADP-riboxanated Arg561-containing peptide acquired under collision-induced dissociation (CID). d, e Validation of MS-identified sites by site-directed mutagenesis. 293T cells co-transfected with GFP-OspC3 and FLAG-SEPT9 (WT or variants) were analyzed by IP and immunoblotting. 10RK refers to substitution of all ten modified arginine residues to lysine. f Prokaryotic co-expression. GST-SEPT9 was expressed in E. coli alone or with calmodulin (CaM) and OspC3 as indicated. Purified GST-SEPT9 was blotted to detect ADP-riboxanation. g In vitro reconstitution. Purified recombinant proteins were incubated (37 °C, 1 h) and analyzed by Coomassie staining and Western blotting. The effects of NAD + , CaM, and calcium ions were tested. h ADP-riboxanation during infection. HeLa cells transfected with FLAG-SEPT2, SEPT6, SEPT7, or SEPT9 were infected with S. flexneri (MOI = 50, 2 h). Anti-FLAG immunoprecipitates were analyzed by immunoblotting. i OspC-dependent modification of Arg561. HeLa cells expressing FLAG-SEPT9 (WT or R561 mutants) were infected with indicated S. flexneri strains (MOI = 50, 2 h). Modification was assessed by anti-FLAG IP and immunoblotting. UI, uninfected. j Endogenous SEPT9 modification. Analyzed as in (i), but immunoprecipitated with an anti-SEPT9 antibody. For (a, d–j), experiments were repeated three times with similar results.
In line with MS/MS analyses, successive mutation of the identified sites to lysine (1RK to 4RK) resulted in a progressive decrease in modification levels, although residual signals were still visible in immunoblotting assays (Fig. 2d). Indeed, further MS/MS analysis of SEPT9-4RK identified six additional ADP-riboxanation sites (Fig. 2a, right). Nonetheless, a SEPT9 variant with substitution of these six sites to lysine (10RK) was still modifiable, though the signal was further reduced (Fig. 2e). Similarly, we identified at least three modified residues in SEPT2 (Arg29, Arg138, and Arg306) and SEPT6 (Arg299, Arg328, and Arg421), respectively (Figures S4 and S5). These results demonstrate that OspC3 catalyzes multiple modifications on septins.
OspCs directly ADP-riboxanate SEPT9 in vitro and during bacterial infection
To address whether septins are direct substrates of OspC ADP-riboxanase activities, OspC3 and septins were co-expressed in E. coli cells. Immunoblotting assays suggested that OspC3 ADP-riboxanated SEPT9 in vitro in a manner dependent on both its enzymatic activity and the eukaryotic co-factor, calmodulin (CaM)19 (Fig. 2f). Furthermore, in an in vitro reaction using purified recombinant proteins, we found that the OspC3-CaM complex modified SEPT9 in an NAD+-dependent manner (Fig. 2g, lanes 1–5). Consistent with previous reports that stimulation of OspC3 ADP-riboxanase activity requires Ca2+-free CaM, high levels of calcium ions, but not the ion-chelating reagent EDTA, abolished the modification reaction (Fig. 2g, lanes 6–7)19,20. We also analyzed the same sets of SEPT9 samples prepared from E. coli co-expression (Fig. 2f) and in vitro reactions (Fig. 2g) by tandem mass spectrometry analyses. Unexpectedly, only three out of four sites that were previously identified in 293T cells (i.e., Arg17, Arg46, and Arg134) were detected in their modified forms, while Arg561 was not detectably modified (Figures S6a and S6b). Given Arg561 as the primary modification site in 293T cells (Table 1), we speculate that ADP-riboxanation of SEPT9 Arg561 may require spatial orientation of an intact septin complex.
To further determine whether septins are the physiological targets of OspC effectors, we carried out S. flexneri infection of HeLa cells expressing FLAG-tagged septin subunits (i.e., SEPT2, SEPT6, SEPT7, and SEPT9). Immunoprecipitated subunits were probed for potential ADP-riboxanation and SEPT9 was the sole protein that exhibited detectable signals of modification (Fig. 2h). These findings are in line with our previous co-expression data, in which SEPT9 was most modified in 293T cells relative to other septin subunits. Intriguingly, SEPT9 was also modified in uninfected control cells, though this signal was significantly elevated during infection (Fig. 2h). Given the absence of known ADP-riboxanases in eukaryotic cells (at least not found until now), we reasoned that basal levels of SEPT9 modification are likely due to classical ADP-ribosylation.
To further confirm the role of T3SS-delivered OspC effectors in mediating ADP-riboxanation of SEPT9, we constructed a series of S. flexneri mutants missing either individual or a combination of several ospC genes. Successive knockout of ospC genes led to a progressive decrease in ADP-riboxanation levels of SEPT9 during bacterial infection. In particular, simultaneous deletion of ospC1 and ospC3 resulted in substantial loss of modification signals (Fig. 2i and S6c). As expected, minimal signals were observed upon deletion of all four ospC homologs. Complementation of this strain with both OspC1 and OspC3 restored modification at levels higher than wild-type S. flexneri infection (Fig. 2i), likely due to higher expression levels of plasmid-borne effectors (Figure S6d). Furthermore, SEPT9 with substitution of Arg561 to either Ala or Lys failed to undergo OspC-mediated ADP-riboxanation during bacterial infection (Fig. 2i), although the 4RK/10RK mutants still exhibited detectable modifications when co-expressed with OspC3 in 293T cells (Fig. 2e). Immunoblotting analyses of endogenous SEPT9 revealed OspC-dependent ADP-riboxanation during S. flexneri infection (Fig. 2j). In agreement, mass spectrometry analyses showed that ~4% of host SEPT9 was modified in cells infected by WT S. flexneri (Figure S6e). As expected, we observed higher modification rates (~12%) for infection with the WT OspC complementation strain (Table 2). Together, these results suggest that SEPT9 is the preferred substrate of OspC effectors during S. flexneri infection with Arg561 as the primary site of ADP-riboxanation.
OspC-mediated ADP-riboxanation destabilizes the septin complex
To interrogate the potential impact of OspC-catalyzed ADP-riboxanation on septin functions, we next asked whether these modifications would disrupt the interactions between different septin subunits as such associations are thought to play an important role in the assembly of hetero-oligomeric complexes21. Using SEPT9 as a surrogate, we immunoprecipitated SEPT9-associated proteins from cells expressing either wild-type OspC3 or its catalytically dead mutant. Subsequent quantitative mass spectrometry analyses revealed that 8 proteins showed reduced associations with SEPT9, while 15 proteins exhibited enhanced interactions in cells producing WT OspC3 (Fig. 3a, Supplementary Data 2). Importantly, we found that half of the reduced proteins were septin subunits including SEPT6, SEPT7, SEPT8, and SEPT11 (Fig. 3a). In line with these data, catalytically active OspC3 also reduced the interactions between ectopically expressed FLAG-SEPT9 and HA-SEPT2/6 (Fig. 3b, c). Further network analyses of the SEPT9-interactome data clearly revealed a cluster of proteins with markedly reduced interactions as the septin complex (Fig. 3d). Of note, signals derived from ubiquitin was enriched in the interactome of modified SEPT9 (Fig. 3a), indicating potential ubiquitination of SEPT9 or its associated proteins. Additional GO analysis indicated that processes such as septin ring formation and cytoskeleton-dependent cytokinesis were less represented, whereas cytoplasmic translational initiation and nuclear import signal receptor activity were enriched (Figure S7a).
a Quantitative proteomic analysis of the SEPT9 interactome upon ADP-riboxanation. 293T cells co-expressing 3×FLAG-SEPT9 with GFP-OspC3-WT or -EH/AA were analyzed by anti-FLAG IP and quantitative MS. Volcano plot shows fold changes (x-axis) vs. statistical significance (y-axis) of interacting proteins. Data represent four biological replicates. Statistical analysis: two-sided paired moderated t-test (limma). b, c Impaired septin subunit interactions. 293T cells co-transfected with EGFP-OspC3 (WT or EH/AA), FLAG-SEPT9, and HA-SEPT2 or HA-SEPT6 were analyzed by anti-FLAG IP and immunoblotting. Quantification of interaction strength is shown below blots. d Network analysis. SEPT9-associated proteins are grouped into functional clusters. Nodes are color-coded by interaction strength (log2 fold changes). e Interaction with endogenous septins. 293T cells co-transfected with EGFP-OspC3 (WT or EH/AA) and FLAG-SEPT9 (WT, 4RK, or 10RK) were analyzed by anti-FLAG IP and immunoblotting for endogenous SEPT2, SEPT6, and SEPT7. Quantification is shown below blots. f In vitro pull-down assay. Recombinant GST-Strep-tag II-SEPT9 (WT, ADP-riboxanated, 4RK, or 10RK) and GST-SEPT7 individually purified from E. coli were incubated together. Strep-Tactin pull-down samples were analyzed by immunoblotting. Quantification is shown below blots. For (b, c, e, f), experiments were repeated three times with similar results.
To further support that septin ADP-riboxanation disrupts interactions of different subunits, we co-expressed EGFP-OspC3 and FLAG-SEPT9 in 293T cells. Immunoprecipitated SEPT9 was probed with antibodies specific for endogenous septin subunits. Consistent with previous mass spectrometry data, we found reduced association of SEPT9 with other octamer subunits (i.e., SEPT2, SEPT6, and SEPT7) in an ADP-riboxanation-dependent manner (Fig. 3e). Unexpectedly, we found that the SEPT9 mutants unable to undergo ADP-riboxanation (i.e., 4RK and 10RK) lost their regular subcellular localization and instead exhibited nuclear distribution (Figure S7b). This altered localization may, at least partially, account for the weakened interactions between these mutants and other septin subunits (Fig. 3e). In agreement, in vitro pull-down assays using recombinant proteins purified from E. coli also demonstrated that ADP-riboxanated SEPT9 bound less efficiently to SEPT7 in comparison to its unmodified form. Importantly, SEPT9-SEPT7 interaction was dependent on the modification sites, as further supported by data from 4RK and 10RK mutants (Fig. 3f). These residues may play a critical role in mediating interactions with actin filaments and microtubules, which ultimately determine the subcellular localization of SEPT98.
ADP-riboxanation of SEPT9 at Arg561 destabilizes septin octamer and obstructs septin cage assembly
Next, we focused our attention on the functional roles of ADP-riboxanation of SEPT9 occurring at Arg561 given that it is the primary site of modification induced by OspC effectors during Shigella infection. Importantly, residues from the C-terminal α6 helices (Glu558 and Arg561) and the loops following α2 (Glu394, Arg399) form salt bridges and hence play a critical role in stabilizing the SEPT9-SEPT9 NC-interface22 (Fig. 4a). Given negative charges introduced by ADP-riboxanation17, we reasoned that modification of Arg561 likely destabilizes septin octamers by disrupting salt bridges and the NC-interface. To test this hypothesis, we reconstituted SEPT9-containing octamers in vitro by using recombinant proteins purified with a two-tag purification scheme (Figure S8a) as previously described23. Subsequent SDS-PAGE and mass photometry analyses confirmed the stoichiometric assembly of recombinant septin octamers (Fig. 4b and S8b). Of note, SEPT9-WT formed both tetramers (~200 kDa) and octamers (~400 kDa), with the latter being the major form. In contrast, SEPT9-NCmut1 (I281D and M288D) and SEPT9-R561E formed only tetramers, due to the impaired NC-interface as previously reported21,24. Notably, SEPT9 samples co-expressed with WT OspC3 (and CaM), but not its catalytically dead mutant, formed less octamers and more tetramers instead (Fig. 4b), indicating disruption of the SEPT9-SEPT9 NC-interface. Purified heteromeric complexes are capable of self-assembling into filaments under conditions such as low-salt buffers (e.g., 50 mM KCl)7. Consistent with mass photometry analysis, SEPT9-WT octamers, but not complexes containing SEPT9 mutants or ADP-riboxanated proteins, successfully polymerized under low-salt conditions (Fig. 4c). Together, these data support our conclusion that ADP-riboxanation of SEPT9 at Arg561 destabilizes septin octamers by a mechanism involving disruption of salt bridges at the NC-interface.
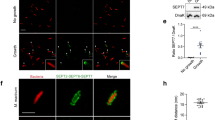
a Structural details of the NC-interface. Schematic of the human septin octamer, with the adjacent SEPT9 subunits colored green and blue, Created in BioRender. Zhang, Y. (2025) [https://BioRender.com/j7jisrq]. The NC-interface (PDB: 5CYP) is magnified to show key salt-bridge residues as sticks: Glu394, Arg399, Glu558, and Arg561. Structural view generated with PyMOL. b Mass photometry of septin oligomers. Septin oligomers containing WT or mutant SEPT9 were co-expressed with CaM and OspC3 (WT or EH/AA) in E. coli as indicated. Left: Representative mass distribution plots (n = 5 experiments). Right: Ratio of octamers to tetramers. c In vitro self-assembly of purified septin hetero-oligomers. Septin complexes diluted from high-salt buffer to low-salt buffer were incubated (RT, 4 h) in dark and imaged. Representative images (scale bar, 10 μm, n = 4 independent experiments) and quantification of filament area (right) are shown. Unless otherwise specified, statistical analysis was performed using one-way ANOVA followed by Tukey’s multiple comparisons test (two-sided), and data are presented as mean ± SEM. ****P < 0.0001.
To determine whether Arg561 modification has any impact on the assembly of septin cages during S. flexneri infection, we transfected FLAG-SEPT9 or its variants in HeLa cells at levels close to endogenous septins. Immunostaining analyses revealed that WT, but not SEPT9-NCmut1, was efficiently incorporated into septin cages following S. flexneri infection (Figure S8c). Interestingly, SEPT9-NCmut2 was still able to assemble septin cages, albeit with lower efficiency, likely due to partial destruction of salt bridges. Similarly, SEPT9 R561K, but not R561A and R561E mutants, formed cage-like structures (Fig. 5a). We further quantified fluorescence intensities of SEPT9 surrounding intracellular bacteria. In agreement with analyses of septin cage assembly, analyses of transfected SEPT9 variants indicated that both R561 and the NC-interface are required for SEPT9 recruitment to intracellular S. flexneri (Fig. 5b and S8d). By comparison, fluorescence intensities of total S. flexneri-associated SEPT9 (including both transfected and endogenous proteins) were not significantly different among the same set of transfected samples (Fig. 5c and S8e). In addition, ADP-riboxanated SEPT9 (induced by ectopic expression of GFP-OspC3, but not the GFP-vector or inactive mutant) failed to assemble into septin cages during bacterial infection (Figure S8f). Quantitative assessment revealed that ~14% of cytosolic bacteria were entrapped by SEPT9-positive cages. Expression of WT OspC3, but not the EH/AA mutant, significantly reduced cage assembly to ~4% (Fig. 5d, e). Collectively, these data suggest that OspC-mediated ADP-riboxanation of SEPT9 at Arg561 impairs salt bridges and disrupts the SEPT9-SEPT9 NC-interface, thereby preventing septin recruitment and S. flexneri cage assembly.
a–c The role of Arg561 in SEPT9 assembly into septin cages. HeLa cells expressing WT or mutant FLAG-SEPT9 were infected with GFP-expressing S. flexneri (MOI = 20, 2 h 40 min) and analyzed by immunofluorescence using indicated antibodies. a Representative images (scale bar, 10 μm, n = 3 independent experiments) and normalized fluorescence intensity (NFI) profiles along arrows. b, c Normalized mean fluorescence intensity (MFI) of Shigella-associated FLAG-SEPT9 (b) and total SEPT9 (c). Data represent n = 3 independent experiments (25 fields of view). d, e OspC3 inhibits the assembly of septin cages in an enzymatic activity-dependent manner. HeLa cells transfected with indicated plasmids were infected with a GFP-expressing S. flexneri strain (MOI = 6, 2 h 40 min) and then analyzed by immunofluorescence and quantitative microscopy. Representative immunofluorescence images (scale bar, 10 μm, n = 3 independent experiments) are shown in (d). The percentages of septin cage-positive bacteria (e) are shown as mean ± SEM from n = 896 (Vector), n = 582 (FLAG-OspC3), and n = 977 (FLAG-OspC3-EH/AA) distributed in n = 3 independent experiments. Unless otherwise specified, statistical analysis was performed using one-way ANOVA followed by Tukey’s multiple comparisons test (two-sided). **P < 0.01, ***P < 0.001, ****P < 0.0001.
S. flexneri evades septin cage-based cell-autonomous immunity during infection in an OspC-dependent manner
Cytosolic bacterial pathogens, including Shigella, are entrapped by septin cage-like structures to restrict their intracellular mobility and facilitate their targeting to antibacterial autophagy12,14. We next investigated whether T3SS-delivered OspC effectors can antagonize septin cage assembly and hence facilitate bacterial escape during S. flexneri infection. We challenged HeLa cells with WT bacteria and stained for endogenous septins to visualize cage-like structures at 2 h 40 min post infection. Quantitative microscopy revealed that ~11% of cytosolic S. flexneri were compartmentalized in septin cages (Fig. 6a, b). As we previously reported, when cells were infected by an ospG-deletion (ΔospG) strain, septin cages formed at a higher percentage (21%)15. By comparison, HeLa cells challenged with a S. flexneri strain missing all ospC genes (ΔospC) exhibited similar rates of cage assembly (~20%). In addition, reintroduction of both WT OspC1 and OspC3, but not their catalytically dead mutants, into the ΔospC strain restored the ability to block septin cage assembly (Fig. 6c, d). Given that both OspC and OspG effectors target septins via distinct post-translational modifications, we constructed a double-knockout mutant (ΔospCΔospG). Notably, septin cage formation was increased to an even higher level (~31%) in ΔospCΔospG infected cells, thereby indicating that OspCs and OspG work in concert to inhibit cage assembly (Fig. 6a, b).
a, b Synergy between OspC and OspG. HeLa cells infected with GFP-expressing S. flexneri strains (MOI = 6, 2 h 40 min) were analyzed by immunofluorescence. a Representative confocal images (scale bar, 10 μm, n = 3 independent experiments). b The percentages of septin caged bacteria are shown as mean ± SEM from n = 1753 (WT), n = 833 (ΔospC), n = 799 (ΔospG), and n = 988 (ΔospC ΔospG) distributed in n = 3 independent experiments. c, d Complementation of S. flexneri ΔospC strains with catalytically active OspCs enables bacteria to escape from septin cages. Samples were prepared as described above. Representative confocal images (scale bar, 10 μm, n = 3 independent experiments) are shown (c). The percentages of septin caged bacteria are shown as mean ± SEM from n = 1494 (WT), n = 1398 (ΔospC), n = 2241 (ΔospC+pOspC1/C3-WT), and n = 1296 (ΔospC+pOspC1/C3-EH/AA) distributed in n = 3 independent experiments (d). e, f OspC and OspG synergistically promote intercellular spread of S. flexneri. Plaque formation assays were performed as described in the Methods section. Representative images (scale bar, 1 mm, n = 2 independent experiments) are shown. Quantification results of plaque area (n = 30 plaques per condition) are presented as mean ± SEM. g–i Intracellular growth of S. flexneri in HeLa cells. HeLa cells were infected with indicated S. flexneri strains. Bacterial load was determined at 0.5 (g), 2 (h), and 4 h (i) post-infection (normalized to 0.5 h). Data: mean ± SEM (n = 5 independent experiments). j A proposed model illustrating S. flexneri effectors OspCs and OspG synergistically counteract septin cage formation. OspCs destabilize SEPT9-containing octamers and block cage formation via direct ADP-riboxanation, while OspG inhibits cage assembly through septin ubiquitination in a mechanism dependent on CAND1 phosphorylation. Created in BioRender. Zhang, Y. (2025) [https://BioRender.com/bpts0kk]. Unless otherwise specified, statistical analysis was performed using one-way ANOVA followed by Tukey’s multiple comparisons test (two-sided). P-values are denoted as: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
To further investigate the roles of OspC and OspG in bacterial pathogenesis, we assessed intracellular proliferation and cell-to-cell spread of S. flexneri using gentamicin protection and plaque formation assays, respectively. Notably, simultaneous deletion of both ospC and ospG genes nearly abolished S. flexneri intercellular spread, whereas single deletion mutants (i.e, ΔospC or ΔospG) exhibited partial phenotypes (Fig. 6e). In addition, complementation of the double-knockout strain with either wild-type OspC or OspG, but not the inactive EH/AA mutant, restored bacterial spreading ability (Fig. 6f). Similar data were recorded for intracellular replication, though growth defects of the double mutant were not as severe as cell-to-cell spread (Fig. 6h, i). In all cases, bacterial invasion was unaffected by these deletions (Fig. 6g). Collectively, these findings suggest that OspCs act synergistically with OspG and liberate S. flexneri from cage entrapment by targeting septins through different post-translational modifications (Fig. 6j).
Discussion
To defend against bacterial infection, host cells can activate various cell-autonomous immune responses, such as autophagy, GBP microcapsules, and septin cage entrapment. S. flexneri, as a successful cytosolic pathogen, has been shown to suppress autophagy and GBP1-mediated cell-autonomous immunity by dedicated T3SS effectors6,25,26. However, little is known about how bacteria evade septin cage entrapment. Here, we identified septins as conserved substrates of the OspC family of effectors. We found that SEPT9 is a physiological target of OspCs during S. flexneri infection. Importantly, ADP-riboxanation of SEPT9 at Arg561 disrupts the NC-interface between adjacent SEPT9 subunits, leading to disassembly of septin octamers and inhibition of septin cage formation. To the best of our knowledge, these OspC proteins represent the first bacterial factors that directly attack septin-mediated cell-autonomous immunity. Finally, we demonstrated that OspCs act synergistically with OspG to counteract septin cage formation during S. flexneri infection (Fig. 6j). It is remarkable to consider that bacterial pathogens have evolved two distinct mechanisms (i.e., mediated by OspCs and OspG, respectively) to counteract septin cage-based cell-autonomous immunity.
Septins can form both hexamers and octamers, yet their relative contribution to septin filaments/assemblies remains unclear. Recent studies propose that septin filaments are formed primarily through octamers, which further interact with actin filaments and microtubules21,27. Indeed, the association of septins with actin filaments or microtubules depends on their polymerization into filamentous structures. Nonetheless, it remains undetermined whether septin cage formation relies on polymerization of septin oligomers. Our findings suggest that the polymerization capacity of septin oligomers is critical for subsequent cage assembly. Notably, disruption of the SEPT9 NC-interface via R561E mutation inhibits the assembly of septin octamers as well as cage-like structures (Figs. 4c, 5a–c). This observation aligns with previous reports that polymerization stabilizes the interaction between septin complexes and membranes28.
Given that septin octamers, rather than hexamers, are the primary functional form, it raises the question of whether septin cages are assembled in a similar manner. Traditionally, the well-characterized hexamers were utilized in vitro to investigate the molecular mechanisms underlying septin recognition of S. flexneri14,29. However, it is important to keep in mind the potential differences between the intracellular environment and in vitro conditions. For instance, while cellular colocalization of septin complexes with actin filaments depends on their polymerization capacity, in vitro experiments have shown that polymerization-deficient septin complexes can still bind to actin filaments21. Of note, septin octamers differ from hexamers by the incorporation of the SEPT3 subgroup. In this subgroup, only SEPT9 is widely expressed, while SEPT3 is predominantly expressed in the central nervous system and SEPT12 is mostly found in lymphocytes and testicular cells9. Therefore, SEPT9 likely plays a prominent role in septin-mediated cell-autonomous immunity, a highly conserved cellular process. Intriguingly, our experimental data support this conclusion: during S. flexneri infection, OspC effectors preferentially target SEPT9 via ADP-riboxanation at Arg561, in a manner depending on the spatial conformation of septin octamers. Therefore, these findings favor the argument that septin octamers (rather than hexamers) are more likely involved in septin cage formation in host cells.
ADP-ribosylation is a widespread modification of biomolecules, including proteins and nucleic acids, in all kingdoms of life30,31. In host-pathogen interactions, protein ADP-ribosylation has emerged as an important strategy utilized by pathogens to subvert host immune defenses. Prominent examples include the Salmonella Typhimurium effector SopF and Chromobacterium violaceum effector CteC, which interfere with host antibacterial autophagy and ubiquitin signaling by ADP-ribosylating V-ATPase32 and ubiquitin33, respectively. In addition, the Legionella SidE family of effectors mediates noncanonical ubiquitination independent of E1/2 via a mechanism involving an initial step of ubiquitin ADP-ribosylation34,35. Moreover, high-throughput modification proteomics has been employed to comprehensively investigate putative substrates of bacterial effectors26,36. In our study, we demonstrate that S. flexneri OspC effectors catalyze ADP-riboxanation, a recently identified variant of ADP-ribosylation, of SEPT9. Unexpectedly, we also detected endogenous modifications of SEPT9 (Fig. 2h), which are most likely classical ADP-ribosylation; this observation has not been previously reported. Therefore, it is conceivable that regulation of septin biology through ADP-ribosylation may represent a universal mechanism in eukaryotic cells. With this in mind, it is striking that bacterial pathogens have evolved a derivative of ADP-ribosylation, yet distinct from its canonical form, to subvert host cell defenses by targeting septin biology in an irreversible manner (i.e., a de-modifying enzyme has not yet been identified). Furthermore, given S. flexneri strategies of escaping from septin cages, it is next of great interest to study how other cytosolic bacteria evade septin-mediated cell-autonomous immunity.
Methods
Plasmids and bacterial strains
DNAs for ospC1, ospC2, and ospC3 were amplified from S. flexneri 2a strain 2457 T (kindly provided by Dr. Feng Shao from National Institute of Biological Sciences, Beijing). Complementary DNAs (cDNA) for human SEPT2, SEPT6, SEPT7, SEPT9 (isoform 1, the longest isoform), SEPT10, and SEPT11 were described previously15. DNAs were ligated into pCS2 vectors for transient expression in mammalian cells. For recombinant protein expression in E. coli, DNAs were inserted into pGEX-6p-2, pET28a-6×His, pET28a-6×His-SUMO, pACYCDuet-1 and pET-MCN series vectors, such as pnEA-vH (pET-15b backbone) and pnCS (pCDFDuet backbone) vectors37. All truncations and point mutations were generated by the standard polymerase chain reaction (PCR) cloning method. All plasmids were verified by DNA sequencing.
S. flexneri was freshly streaked from glycerol stocks onto tryptone soy agar supplemented with 0.02% (w/v) Congo red, and subsequently a single red colony was selected and inoculated into tryptone soy broth (TSB) with appropriate antibiotics. For efficient infection of host cells, all S. flexneri strains used in this study were transformed with a plasmid expressing afimbrial adhesin (Afa). The S. flexneri ospC knockout strain was generated using the CRISPR/Cas9 system as described previously38. Briefly, single-guide RNAs (sgRNAs) targeting the coding sequence (CDS) or promoter region of the gene of interest were designed using the CRISPOR website and introduced into the pTarget plasmid via PCR. Notably, the CDSs of ospC2, ospC3, and ospC4 are highly similar, which poses significant challenges for the sgRNA design. It is crucial to avoid targeting multiple ospC genes simultaneously, as this may result in multiple double-strand breaks and cell death. Additionally, the homologous arms should be designed to exclude sequences that could be targeted by sgRNAs used in subsequent rounds of gene editing. Specifically, ospC genes were knocked out in a sequential order: ospC1, ospC3, ospC4, and ospC2, resulting in a strain with all ospC homologs deleted. Sequences of all primers and gRNAs used for Shigella gene deletion are listed in Supplementary Data 3. After obtaining the ospC quadruple mutant, ospG was further knocked out to generate the ΔospCΔospG strain.
For complementation, Strep-tag-II C-terminally-tagged DNAs for ospC1, C-terminal HA-tagged DNAs for ospC3 or their mutants were cloned into the pBBR1-tet vector (generated by replacing the lac promoter and lac operator of pBBR1MCS2 with the tet promoter) for constitutive expression. To visualize S. flexneri in immunofluorescence experiments, the DNA encoding EGFP with a ribosome binding site (RBS) sequence was cloned into the pBBR1-tet vector. All mutant strains were confirmed by PCR using at least two different pairs of primers, and PCR products were further validated by DNA sequencing.
Cell culture, transfection and immunoprecipitation
293T (Cat No. CRL-3216) and HeLa cells (Cat No. CRM-CCL-2) were obtained from American Type Culture Collection (ATCC) and grown in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (TRANS, FS301-02) and 2 mM L-glutamine. Cell monolayers were maintained at 37 °C in a humidified 5% CO2 incubator. All cells were tested for mycoplasma using the standard PCR method. Transient transfection was performed using polyethylenimine (PolySciences) or jetPRIME (Polyplus-transfection) following the manufacturer’s instructions.
For immunoprecipitation, 293T cells (~1 × 107) grown to 80% confluence in 10 cm dishes were transfected with indicated plasmids. Twenty-four hours after transfection, cell pellets were harvested and lysed in the TNT buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, and 1% Triton X-100) supplemented with protease inhibitor cocktail (Selleck, B14002) for 30 min, followed by centrifugation at 4 °C (16,000 × g for 10 min). The clarified lysates were incubated with pre-washed anti-FLAG M2 affinity beads (Sigma-Aldrich, A2220) at 4 °C with gentle rotation for 4 h. The beads were washed four times with the lysis buffer, and bound proteins were eluted with FLAG peptides (MedChemExpress, HY-P0319A) and run on SDS-PAGE followed by either Western blot or LC-MS analyses.
Western blots and antibodies
Protein samples were separated by SDS-PAGE and proteins in the gel were transferred onto PVDF membranes, followed by blocking with non-fat milk for 1 h at room temperature and incubation with specific primary antibodies (diluted at 1:1000 to 1:5000) overnight at 4 °C. Afterward, the membranes were extensively washed (for 4 times) with TBST buffer (composed of 25 mM Tris pH 7.5, 140 mM NaCl, 3 mM KCl, and 0.1% Tween-20) and incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies (1:5000). Following another round of washes, proteins of interest were visualized by using a chemiluminescence imaging system (Tanon 5200).
The following primary antibodies were used in this study: anti-poly/mono-ADP ribose (1:2000, Cell Signaling Technology, 83732), anti-GFP (1:2000, Proteintech, 66002-1-Ig), anti-GAPDH (1:5000, Proteintech, 60004-1-Ig), anti-β-actin (1:5000, Cell Signaling Technology, 4967), anti-FLAG for Western blot (1:5000, Zen Bioscience, R24091), anti-FLAG for immunofluorescence (1:300, Beyotime Biotechnology, AF519), anti-HA (1:5,000, Invitrogen, 26183), anti-GST (1:1000, Zen Bioscience, 390028), anti-His (1:1000, CWBIO, CW0286M), anti-S-tag (1:2,500, Sino Biological, 101290-T38), anti-SEPT2 (1:2,500, Proteintech, 11397-1-AP), anti-SEPT6 (1:1000, Santa Cruz, sc-514781), anti-SEPT7 (1:2,000, Proteintech, 13818-1-AP), anti-SEPT9 (IF 1:500, WB 1:2500, Atlas Antibodies, HPA042564), and anti-Strep-Tag II (1:2,500, Abmart, M40014S). The secondary antibodies include HRP-conjugated anti-mouse IgG (1:5000, Jackson ImmunoResearch Laboratories, 115-035-003), HRP-conjugated anti-rabbit IgG (1:5000, Jackson ImmunoResearch Laboratories, 111-035-003), Alexa Fluor 488-labeled anti-mouse IgG (1:500, Jackson ImmunoResearch Laboratories, 115-545-003), Cyanine3 (Cy3)-conjugated anti-rabbit IgG (1:500, Jackson ImmunoResearch Laboratories, 111-165-003), and Alexa Fluor 647-labeled anti-mouse IgG (1:500, Beyotime Biotechnology, A0473).
Bacterial infection of host cells
S. flexneri strains carrying an Afa-encoding plasmid were used for infection experiments in this study. A red colony of S. flexneri, grown on Congo red agar plates, was inoculated into TSB and cultured for approximately 16 h at 37 °C with constant shaking. On the following day, overnight bacterial cultures were diluted 1:50 into fresh pre-warmed TSB, and further cultured until an OD600 of 0.6 was reached. Cell monolayers were equilibrated with pre-warmed DMEM for 30 minutes and infected with S. flexneri at indicated multiplicity of infection (MOI).
For detection of septin ADP-riboxanation during infection, HeLa cells transfected with 3×FLAG-septin in 6-well plates for 24 h were infected with relevant S. flexneri strains at an MOI of 50. After 45 min of infection, infected cell monolayers were washed three times with pre-warmed DMEM and further incubated with fresh DMEM containing 10% FBS and 50 μg/mL gentamicin at 37 °C and 5% CO2 for 2 h to kill extracellular bacteria. At indicated time points, cell monolayers were collected, lysed, and followed by anti-FLAG immunoprecipitation. Eluted samples were collected for SDS-PAGE and Western blotting analyses. For detection of endogenous SEPT9, bacterial infections were performed as described above and cell lysates were subjected to immunoprecipitation with an anti-SEPT9 antibody or an isotype control IgG pre-immobilized on AminoLink Coupling Resin (Thermo Fisher Scientific). Bound proteins were eluted using 0.1 M glycine–HCl (pH 2.7) and immediately neutralized by adding 1/10 volume of 2 M Tris–HCl (pH 8.0).
For microscopy analyses of septin cages, HeLa cells were seeded onto glass coverslips in 24-well plates for 36 h before bacterial infection. For experiments involving mutation of Arg561 or residues at the NC-interface of SEPT9, pre-seeded HeLa cells were further transfected with indicated plasmids for 20 h before infection. Cells were infected with S. flexneri at an MOI of 6 (unless otherwise specified) for 45 min, washed with fresh media and further incubated in fresh DMEM containing 10% FBS and 50 μg/mL gentamicin for 2 h 40 min. Cells were washed twice with PBS and subsequently fixed with 4% paraformaldehyde (PFA) for 15 min. The fixed cells were stored in PBS until further immunofluorescence analysis.
To measure intracellular replication, HeLa cells cultured to 80% confluence in 6-well plates were infected with S. flexneri at an MOI of 20. Bacterial infections were performed as described above. After infection, cell monolayers were incubated with fresh DMEM containing 10% FBS and 100 μg/mL gentamicin at 37 °C with 5% CO2 for 30 min to eliminate extracellular bacteria. Subsequently, the gentamicin concentration was reduced to 25 μg/mL. At each time point (0.5, 2, and 4 h), cells were lysed in cold PBS containing 0.1% Triton X-100, and colony-forming units (CFUs) were determined by serial dilution and plating on TSB agar plates containing 30 μg/mL streptomycin.
For investigation of intercellular spread, plaque formation assays were performed as described previously39. Briefly, HeLa cells cultured to 100% confluence in 6-well plates were incubated with S. flexneri lacking afimbrial adhesin at an MOI of 0.005 for 2 h to facilitate invasion. After incubation, cells were washed three times with pre-warmed DMEM and subsequently overlaid with 3 mL of agarose overlay medium (DMEM containing 5% FBS, 50 μg/mL gentamicin, and 0.5% (w/v) agarose). Following 48 h of incubation, the agarose overlay was removed, and infected HeLa cells were subjected to Giemsa staining for plaque visualization. The plaque area (A) was calculated using the formula A = π(d/2)2, where d represents the diameter of a plaque measured using the ImageJ (v1.48) software. Average plaque areas were determined by analyzing 30 plaques per group.
Purification of recombinant proteins
The E. coli BL21 (DE3) strains transformed with indicated plasmids were used for protein expression and purification. Protein expression was induced by adding 0.4 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) when the OD600 of cell cultures reached approximately 0.8. The bacterial cells were subsequently cultured at either 22 °C for 12 h or 18 °C for 16 h, depending on the stability and/or solubility properties of target proteins. Proteins tagged with GST, His6, or His6-SUMO were purified using glutathione resins (Sangon Biotech, C600031) or HisTrap HP affinity chromatography (GE Healthcare Life Sciences). If necessary, the GST or SUMO tags were removed by PreScission or Ulp1 protease digestion, respectively. Further purification was achieved through ion-exchange chromatography, followed by size-exclusion chromatography. For downstream applications, buffer exchange was performed by using HiTrap Desalting chromatography (GE Healthcare Life Sciences). Purified proteins were then concentrated and stored at −80 °C in a buffer containing 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 2 mM dithiothreitol, and 5% glycerol.
Recombinant human septin complexes, including the SEPT2-msfGFP-SEPT6-SEPT7 hexamer and the SEPT2-msfGFP 8mer-9_i1, were purified as previously described23,29,40. Briefly, E. coli BL21 (DE3) strains were transformed with compatible bicistronic plasmids expressing the respective units, and induced for protein expression. Bacterial lysates were subjected to a two-step affinity purification scheme. First, HisTrap affinity chromatography was used to capture complexes containing N-terminal His6-tagged SEPT2. Next, Strep-Tactin affinity chromatography was employed to isolate hexameric complexes (via the C-terminal Strep-tag II on SEPT7) or octameric complexes (via the C-terminal Strep-tag II on SEPT9). The purified complexes were dialyzed to remove imidazole and reconstituted in stabilization buffer (50 mM Tris-HCl pH 8.0, 300 mM KCl, 5 mM MgCl₂, and 1 mM DTT). This method yielded stoichiometric septin complexes to homogeneity suitable for downstream analyses.
In vitro SEPT9 pull-down assays
Recombinant GST-Strep-tag II-SEPT9 proteins or corresponding 4RK/10RK mutants were incubated with GST-fused SEPT7 at 4 °C for 2 h. The mixture was then incubated with Strep-Tactin resins at 4 °C for 30 min. The resins were subsequently washed three times with the TNT buffer and boiled in 2 × SDS loading buffer. All samples were analyzed via SDS-PAGE followed by Western blot.
Immunofluorescence and microscopy
Cells were fixed with 4% PFA after S. flexneri infection, and washed three times with PBS (5 min per wash). Permeabilization was performed with 0.1% Triton X-100 in PBS for 5 min, and washed again with PBS for three times. Subsequently, cells were blocked with 5% goat serum in PBS for 30 min at room temperature and incubated overnight at 4 °C with primary antibodies diluted in PBS containing 1% BSA and 0.05% Tween-20. Next morning, samples were washed three times with PBS and incubated with secondary antibodies for 1 h at room temperature. Cells were then washed with PBS, stained with DAPI for 5 min, and washed three times again with PBS. Slides were mounted with antifade mounting medium (Beyotime biotechnology, P0126). Fluorescence images were captured using a Nikon Eclipse Ti microscope, while confocal images were acquired with a Leica TCS SP8 STED or Leica STELLARIS 8 microscope. Image analysis was performed using LAS X software (v3.5.6) for Leica systems and ImageJ software (v1.48) for further processing and quantification. For quantification of fluorescence intensities of S. flexneri-associated SEPT9, bacteria were first detected by setting an intensity threshold in dedicated channels. Using bacteria as a mask, fluorescence intensities were extracted from corresponding regions in other channels. These intensities were then normalized to the total mean fluorescence intensity of individual cells.
Mass spectrometry analyses
ADP-ribosylome enrichment was performed as previously described16. Briefly, recombinant His6-eAf1521 was expressed, purified, and covalently coupled to NHS-resins. Clarified cell lysates (from ~2 × 107 293T cells overexpressing OspC) were incubated with NHS-resins coated with His6-eAf1521 at 4 °C for 2 h with gentle rotation. The enrichment process was repeated twice, and pooled beads were washed four times with the TNT buffer. Captured proteins were eluted in 2 × SDS loading buffer, separated by SDS-PAGE and stained with Coomassie blue for further MS analyses. The entire lane was excised into several fractions and subjected to in-gel digestion with sequencing-grade trypsin (Promega, USA) as described previously41. Protein bands of interest were reduced and alkylated using dithiothreitol and iodoacetamide, respectively. Protein samples were digested overnight with trypsin. Tryptic peptides were extracted and dried on a SpeedVac vacuum concentrator. The resulting peptides were dissolved in HPLC-grade water prior to LC-MS/MS analyses.
For label-free proteomics analyses, nanoflow reversed-phase liquid chromatography (EASY-nLC, Thermo Scientific) coupled with a hybrid ion trap-Orbitrap mass spectrometer (LTQ Orbitrap Velos, Thermo Scientific) was employed for peptide separation and detection. A homemade capillary column (75 μm × 150 mm) with a laser-pulled electrospray tip (Model P-2000, Sutter Instruments) was packed with 5 μm, 120 Å Xtimate C18 silica-based particles (Welch Materials Inc.). The mobile phases consisted of solvent A (97% water, 3% acetonitrile, 0.1% formic acid) and solvent B (80% acetonitrile, 0.1% formic acid). The LC separation was performed at a flow rate of 300 nL/min with the following gradient: solvent B was increased from 5% to 40% over 40 min, then rapidly raised to 95% in 2 min and maintained for 10 min. Finally, the column was equilibrated with 100% solvent A. Eluted peptides were electrosprayed directly into the mass spectrometer for MS and MS/MS analyses in a data-dependent acquisition mode. One full MS scan (60,000 resolution, m/z 400–1200) was acquired and the 10 most intense ions were selected for fragmentation under collision-induced dissociation (CID) in the ion trap with the following parameters: ≥ +2 precursor ion charge, 2 Da precursor ion isolation window, and 35 normalized collision energy. Dynamic exclusion was set with repeat duration of 30 s and exclusion duration of 12 s.
To determine modification sites, immunoprecipitated septins from 1 × 107 293T cells expressing OspC3 or its catalytically inactive mutant were separated by SDS-PAGE. Protein bands of interest were excised and subjected to in-gel digestion as described above. Resulting peptide samples were analyzed using a hybrid ion trap-Orbitrap mass spectrometer (LTQ-Orbitrap Velos, Thermo Fisher Scientific) in data-dependent acquisition mode. LC-MS parameters were similar to those described above, except that the m/z range of the MS scan was set to 400–1500. Raw MS files were searched against a septin database with Mascot (version 2.3.02, Matrix Science). Key search parameters included a mass tolerance of 20 ppm for precursor ions, 0.8 Da for MS/MS ions, a maximum of four missed cleavages, carbamidomethylation of Cys (+57.021 Da) as a static modification, and Met oxidation (+15.995 Da) and ADP-riboxanation of Arg (+524.03 Da) as dynamic modifications. Spectra of all potentially modified peptides were manually inspected, and their fragment ions were assigned for determination of modification sites. Corresponding peptide peaks were extracted using Thermo Xcalibur (Thermo Fisher Scientific, v2.2), and modification rates of ADP-riboxanated septins were calculated as percentage decrease of normalized peak areas for corresponding unmodified peptides.
In vitro ADP-riboxanation reactions
For in vitro reactions, 20 μg of purified GST-SEPT9 proteins were incubated with 1 μg of GST-OspC3 or its EH/AA mutant and 0.5 μg of His-Sumo-CaM in 40 μL of a reaction buffer containing 50 mM HEPES (pH 7.5), 150 mM NaCl, 1 mM DTT, 2 mM MgCl2, and 1 mM NAD+. To assess the effect of calcium ions on the modification process, 5 mM CaCl2 or 5 mM EDTA (as a calcium chelator) was added to the reaction mixture. The reaction was carried out at 37 °C for 60 min and terminated by boiling in the SDS loading buffer at 95 °C for 10 min. The samples were analyzed by SDS-PAGE and subsequently subjected to either Western blot analyses or Coomassie blue staining. The stained gels were excised for further quantitative LC/MS analyses of ADP-riboxanated peptides derived from SEPT9.
Self-assembly of septin complexes and mass photometry analysis
Self-assembly of septin complexes was performed by diluting stored septin complexes from a high-salt buffer (50 mM Tris-HCl pH 8, 300 mM KCl, 5 mM MgCl₂, and 1 mM DTT) into a low-salt buffer (50 mM Tris-HCl pH 8, 50 mM KCl, 5 mM MgCl₂, and 1 mM DTT) with a final protein concentration of 400 μM. The complexes were incubated at room temperature (RT) for 4 h in the dark, and the reconstituted septin assemblies were imaged using a custom-built flow cell. Briefly, 2 μL of this dilution was placed on a clean glass slide and a coverslip was mounted on top with double-sided tape serving as a spacer. The samples were incubated for 20 min before fluorescence imaging. The images were processed and quantified using ImageJ software.
Mass photometry experiments were conducted using a TwoMP mass photometer (Refeyn Ltd, UK). Movies were recorded at 475.2 Hz with an exposure time of 2.06 ms, and optimized for camera counts to prevent saturation. The instrument was calibrated prior to measurements using BSA (66 kDa, 132 kDa) and thyroglobulin (660 kDa). Focus was achieved by adding a high-salt buffer (50 mM Tris-HCl pH 8.0, 300 mM KCl, 5 mM MgCl₂, and 1 mM DTT) into an 18 μL well, followed by identifying and locking the focal position with the instrument’s autofocus. For each acquisition, 2 μL of septin hetero-oligomer was added at a final concentration of 10 nM. All experiments were performed with at least three replicates. Data were analyzed using DiscoverMP software (v2023 R2).
Label-free protein quantification and bioinformatic analyses
For differential ADP-ribosylome analysis, spectral counting-based quantification was utilized due to its efficiency in identifying the most robust proteomic changes during the initial proteomic screen. Raw MS files were processed using Mascot Daemon (v2.3.02, Matrix Science). Tandem mass spectra were searched against a non-redundant human protein database (UP000005640, version 2021_04, containing 20,388 entries) downloaded from UniProt. Key search parameters included a mass tolerance of 20 ppm for precursor ions, 0.8 Da for MS/MS ions, and a maximum of two missed cleavages. Carbamidomethylation of cysteine residues (+57.021 Da) was set as a static modification, while methionine oxidation (+15.995 Da) was chosen as a variable modification. False discovery rates (FDRs) of peptides and proteins were set at below 1%. Downstream bioinformatic analyses were performed in R (R Foundation for Statistical Computing) using RStudio (an integrated development environment for R). At least two valid quantification values in any group were required, and missing values were imputed with a constant low value to enable calculation of fold changes.
For SEPT9 IP-MS experiments, intensity-based label-free quantification was employed due to its higher accuracy and sensitivity in detecting modest differences. Raw data were analyzed using MaxQuant (v1.5.5.1)42 with default settings. The ‘match between runs’ and label-free quantification (LFQ) features were enabled. The resulting data table was uploaded to the LFQ-Analyst website43 for statistical analysis. Protein-protein interactions (PPI) were analyzed using Metascape44 and densely connected network components were identified with Molecular Complex Detection (MCODE) algorithm45. The resulting network was visualized using Cytoscape (v3.10.3). A heatmap was generated using the pheatmap R package (v1.0.12). Gene Ontology (GO) term enrichment was conducted using the clusterProfiler R package (v4.14.4), and GO terms with adjusted P-values < 0.05 were considered significantly enriched. Bar plots and scatter plots were created using the ggplot2 R package (v3.5.1). A Venn diagram was generated using the VennDiagram R package (v1.7.3).
Statistics and reproducibility
All samples were included from the analysis without randomization or blinding procedures. No statistical methods were used to predetermine sample size. Experiments were independently repeated at least three times to achieve statistical significance. Data were presented as mean ± SEM, and statistical significance was determined by the unpaired two-tailed Student’s t test, or the ANOVA test followed by Tukey’s post hoc test or Holm-Sidak post hoc test as indicated in figure legends. Representative results were shown. ns, not significant; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. All statistical analyses were performed using GraphPad Prism 9.4 software.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data generated in this study have been deposited to the ProteomeXchange Consortium via the iProX partner repository with the dataset identifier PXD070327. Previously published accession codes used in this study include the PDB entry 5CYP (used for structural analysis of the septin NC-interface). The ADP-ribosylome and interactome search results are provided in Supplementary Data. Source data are provided with this paper.
References
Schnupf, P. & Sansonetti, P. J. Shigella pathogenesis: new insights through advanced methodologies. Microbiol. Spectr. 7, https://doi.org/10.1128/microbiolspec.BAI-0023-2019 (2019).
Kotloff, K. L., Riddle, M. S., Platts-Mills, J. A., Pavlinac, P. & Zaidi, A. K. M. Shigellosis. Lancet Lond. Engl. 391, 801–812 (2018).
Randow, F., MacMicking, J. D. & James, L. C. Cellular self-defense: how cell-autonomous immunity protects against pathogens. Science 340, 701–706 (2013).
López-Jiménez, A. T., Özbaykal Güler, G. & Mostowy, S. The great escape: a Shigella effector unlocks the septin cage. Nat. Commun. 15, 4104 (2024).
Wandel, M. P. et al. GBPs inhibit motility of Shigella flexneri but are targeted for degradation by the bacterial ubiquitin ligase IpaH9.8. Cell Host Microbe 22, 507–518.e5 (2017).
Li, P. et al. Ubiquitination and degradation of GBPs by a Shigella effector to suppress host defence. Nature 551, 378–383 (2017).
Valadares, N. F., d’ Muniz Pereira, H., Ulian Araujo, A. P. & Garratt, R. C. Septin structure and filament assembly. Biophys. Rev. 9, 481–500 (2017).
Spiliotis, E. T. & Nakos, K. Cellular functions of actin- and microtubule-associated septins. Curr. Biol. 31, R651–R666 (2021).
Mostowy, S. & Cossart, P. Septins: the fourth component of the cytoskeleton. Nat. Rev. Mol. Cell Biol. 13, 183–194 (2012).
Spiliotis, E. T. & McMurray, M. A. Masters of asymmetry - lessons and perspectives from 50 years of septins. Mol. Biol. Cell 31, 2289–2297 (2020).
Grupp, B. & Gronemeyer, T. A biochemical view on the septins, a less known component of the cytoskeleton. Biol. Chem. 404, 1–13 (2023).
Mostowy, S. et al. Entrapment of intracytosolic bacteria by septin cage-like structures. Cell Host Microbe 8, 433–444 (2010).
Sirianni, A. et al. Mitochondria mediate septin cage assembly to promote autophagy of Shigella. EMBO Rep. 17, 1029–1043 (2016).
Krokowski, S. et al. Septins recognize and entrap dividing bacterial cells for delivery to lysosomes. Cell Host Microbe 24, 866–874.e4 (2018).
Xian, W. et al. The Shigella kinase effector OspG modulates host ubiquitin signaling to escape septin-cage entrapment. Nat. Commun. 15, 3890 (2024).
Zhang, Q. et al. Shigella induces stress granule formation by ADP-riboxanation of the eIF3 complex. Cell Rep. 43, 113789 (2024).
Li, Z. et al. Shigella evades pyroptosis by arginine ADP-riboxanation of caspase-11. Nature 599, 290–295 (2021).
Alphonse, N. et al. A family of conserved bacterial virulence factors dampens interferon responses by blocking calcium signaling. Cell 185, 2354–2369.e17 (2022).
Hou, Y. et al. Structural mechanisms of calmodulin activation of Shigella effector OspC3 to ADP-riboxanate caspase-4/11 and block pyroptosis. Nat. Struct. Mol. Biol. 30, 261–272 (2023).
Peng, T. et al. Pathogen hijacks programmed cell death signaling by arginine ADPR-deacylization of caspases. Mol. Cell 82, 1806–1820.e8 (2022).
Martins, C. S. et al. Human septins organize as octamer-based filaments and mediate actin-membrane anchoring in cells. J. Cell Biol. 222, e202203016 (2023).
Castro, D. K. S. D. V. et al. A complete compendium of crystal structures for the human SEPT3 subgroup reveals functional plasticity at a specific septin interface. IUCrJ 7, 462–479 (2020).
Iv, F. et al. Insights into animal septins using recombinant human septin octamers with distinct SEPT9 isoforms. J. Cell Sci. 134, jcs258484 (2021).
Cavini, I. A. et al. The structural biology of septins and their filaments: an update. Front. Cell Dev. Biol. 9, 765085 (2021).
Ogawa, M. et al. Escape of intracellular Shigella from autophagy. Science 307, 727–731 (2005).
Liu, W. et al. Nε-fatty acylation of multiple membrane-associated proteins by Shigella IcsB effector to modulate host function. Nat. Microbiol. 3, 996–1009 (2018).
Kuzmić, M. et al. Septin-microtubule association via a motif unique to isoform 1 of septin 9 tunes stress fibers. J. Cell Sci. 135, jcs258850 (2022).
Bridges, A. A., Jentzsch, M. S., Oakes, P. W., Occhipinti, P. & Gladfelter, A. S. Micron-scale plasma membrane curvature is recognized by the septin cytoskeleton. J. Cell Biol. 213, 23–32 (2016).
Lobato-Márquez, D. et al. Mechanistic insight into bacterial entrapment by septin cage reconstitution. Nat. Commun. 12, 4511 (2021).
Suskiewicz, M. J., Prokhorova, E., Rack, J. G. M. & Ahel, I. ADP-ribosylation from molecular mechanisms to therapeutic implications. Cell 186, 4475–4495 (2023).
Dasovich, M. & Leung, A. K. L. PARPs and ADP-ribosylation: Deciphering the complexity with molecular tools. Mol. Cell 83, 1552–1572 (2023).
Xu, Y. et al. A bacterial effector reveals the V-ATPase-ATG16L1 axis that initiates xenophagy. Cell 178, 552–566.e20 (2019).
Yan, F. et al. Threonine ADP-ribosylation of ubiquitin by a bacterial effector family blocks host ubiquitination. Mol. Cell 78, 641–652.e9 (2020).
Bhogaraju, S. et al. Phosphoribosylation of ubiquitin promotes serine ubiquitination and impairs conventional ubiquitination. Cell 167, 1636–1649.e13 (2016).
Qiu, J. et al. Ubiquitination independent of E1 and E2 enzymes by bacterial effectors. Nature 533, 120–124 (2016).
Burnaevskiy, N., Peng, T., Reddick, L. E., Hang, H. C. & Alto, N. M. Myristoylome profiling reveals a concerted mechanism of ARF GTPase deacylation by the bacterial protease IpaJ. Mol. Cell 58, 110–122 (2015).
Diebold, M.-L., Fribourg, S., Koch, M., Metzger, T. & Romier, C. Deciphering correct strategies for multiprotein complex assembly by co-expression: application to complexes as large as the histone octamer. J. Struct. Biol. 175, 178–188 (2011).
Jiang, Y. et al. Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Appl. Environ. Microbiol. 81, 2506–2514 (2015).
Sharma, A. & Puhar, A. Plaque assay to determine invasion and intercellular dissemination of Shigella flexneri in TC7 human Intestinal epithelial cells. Bio-Protoc. 9, e3293 (2019).
Mavrakis, M. et al. Septins promote F-actin ring formation by crosslinking actin filaments into curved bundles. Nat. Cell Biol. 16, 322–334 (2014).
Hu, M., Liu, Y., Yu, K. & Liu, X. Decreasing the amount of trypsin in in-gel digestion leads to diminished chemical noise and improved protein identifications. J. Proteom. 109, 16–25 (2014).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Shah, A. D., Goode, R. J. A., Huang, C., Powell, D. R. & Schittenhelm, R. B. LFQ-Analyst: an easy-to-use interactive web platform to analyze and visualize label-free proteomics data preprocessed with MaxQuant. J. Proteome Res. 19, 204–211 (2020).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Bader, G. D. & Hogue, C. W. V. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinforma. 4, 2 (2003).
Acknowledgements
We thank the members of the Liu laboratory for careful review of the manuscript. We are grateful for the constructive discussion with Drs. Peipei Zhang, Na Dong, and Zilin Li. We would also like to acknowledge Dr. Linglong Fu for his help with S. flexneri mutant construction. We acknowledge Dr. Michael O. Hottiger (University of Zurich) for providing the eAf1521 plasmid. The work in our group was financially supported by grants from National Key Research and Development Program of China (2024YFA1306801 and 2022YFA1304500 to X.L.), the Natural Science Foundation of China (22174003 and 32470187 to X.L.; 22174100 to H.L.), and PKUMed-Wisbiom Joint Laboratory for Human Microbiome Research. The S. Mostowy laboratory is supported by a European Research Council Consolidator Grant (772853-ENTRAPMENT) and Wellcome Discovery Award (226644/Z/22/Z).
Author information
Authors and Affiliations
Contributions
X.L., S.M., Q.Z., Z.T., W.X., and G.Ö.G. conceived and designed the study. Z.T. and W.X. performed most experiments. H.L. and F.Y. provided technical support and advice. H.Z., Y.W., and N.Z. assisted bacterial infection, plasmid construction, and protein purification. X.L. and Z.T. analyzed the data and wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tang, Z., Xian, W., Özbaykal Güler, G. et al. Shigella flexneri evades septin-mediated cell-autonomous immunity via protein ADP-riboxanation. Nat Commun 17, 1727 (2026). https://doi.org/10.1038/s41467-026-68425-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68425-0