Abstract
The high prevalence of diabetes mellitus-induced erectile dysfunction (DMED) in males and the limited efficacy of existing therapies necessitate deeper mechanistic insights. We identify pathological lactate accumulation in the corpus cavernosum as a major driver of DMED in male mice, promoting structural degeneration and smooth muscle cell loss. Mechanistically, transcriptional repression and functional impairment of fructose-1,6-bisphosphatase 1 (Fbp1), a rate-limiting gluconeogenic enzyme, underlie this lactate accumulation. Epigenetic profiling revealed suppression of Fbp1 transcription associated with histone H3 lysine 9 trimethylation and histone H3 lysine 27 trimethylation. Palmitoylation at the FBP1 Cys282 residue, catalyzed by zinc finger DHHC-type containing 13, further disrupted its gluconeogenic function, exacerbating lactate accumulation. To rescue these defects, we engineered lipid nanoparticles loaded with the Fbp1-C282S mutant mRNA, restoring FBP1 expression, blocking pathological palmitoylation, and ultimately improving erectile function in diabetic male mice. Here, our study in male mice identifies cavernosal lactate accumulation as a central pathological factor, elucidates its molecular regulation, and provide a targeted therapeutic strategy for DMED.
Similar content being viewed by others
Introduction
Erectile dysfunction (ED), defined as the persistent inability to attain or sustain an erection sufficient for satisfactory sexual performance, represents a common clinical condition that significantly diminishes quality of life and psychological well-being1. Diabetes mellitus is a major risk factor for ED, with diabetes mellitus-induced erectile dysfunction (DMED) exhibiting a prevalence ranging from 35.8 to 86.1% across diverse populations2,3. Compared to ED in non-diabetic individuals, DMED is characterized by earlier onset, more severe symptoms, and markedly reduced responsiveness to conventional therapies, presenting a substantial clinical management challenge4,5. While the pathogenesis of DMED involves a variety of cellular alterations and mediating factors, the precise mechanisms remain unclear.
Within the penile microenvironment, corpus cavernosum smooth muscle cells (CCSMCs) serve as the terminal effector regulating vascular tone and hemodynamics during erection6. Under diabetic conditions, CCSMCs sustain structural and functional damage through mitochondrial impairment, oxidative stress, and metabolic dysregulation7, while concurrently undergoing a contractile-to-synthetic phenotypic transition that drives excessive extracellular matrix deposition and cavernosal fibrosis8. Consequently, preserving CCSMC integrity and function has emerged as a plausible therapeutic strategy for DMED, although the precise regulatory mechanisms governing these cells demand further investigation.
Lactate, once considered merely a metabolic waste product, is now recognized as a key signaling molecule with pleiotropic roles in cardiovascular, neurodegenerative, inflammatory, and oncological pathologies9. Elevated circulating lactate levels in diabetic patients—detectable even during prediabetic stages—correlate with early diabetes-associated complications10,11. Interestingly, sildenafil citrate, a first-line ED pharmacotherapy, was recently shown to reduce lactate accumulation in muscle following strenuous exercise and improve exercise performance in rats12. Gluconeogenesis maintains a critical homeostatic relationship with lactate metabolism. Under physiological conditions, lactate generated via glycolysis during hypoxia or high metabolic demand is reconverted to glucose through gluconeogenesis in the Cori cycle. Conversely, impaired gluconeogenesis disrupts this recycling axis, driving pathological lactate accumulation in susceptible tissues. Within this gluconeogenic pathway, fructose-1,6-bisphosphatase 1 (FBP1) serves as a critical regulatory node governing lactate-to-glucose conversion. In cancers, FBP1 downregulation drives lactate-driven Warburg effect amplification, accelerating tumor invasion and metastasis13. In non-alcoholic fatty liver disease, suppressed FBP1 induces hepatic lactate buildup, exacerbating inflammation and steatosis14.
Here, we show that pathological cavernosal lactate accumulation drives DMED through structural degeneration and CCSMC loss. Transcriptional repression and functional impairment of Fbp1 constitute the core mechanism underlying lactate dysregulation, mediated by histone methylation and palmitoylation modifications. Critically, nanoparticle delivery of a palmitoylation-resistant Fbp1 mutant mRNA restores gluconeogenic flux, normalizes lactate levels, and rescues erectile function in diabetic mice. This work establishes aberrant lactate metabolism as a pathogenic cornerstone of DMED and proposes a metabolic reprogramming therapy.
Results
Supplemental lactate impairs the CCSMC function and leads to DMED
After obtaining clinical ethics approval (LYEC2025-0188), we conducted an observational case-control study and recruited 60 patients with DMED—33 with mild, 19 with moderate, and 8 with severe ED—and 60 age-matched controls (Fig. 1a, b and Supplementary Fig. 1a). DMED patients exhibited significantly higher serum lactate and blood glucose levels compared to controls (Fig. 1c and Supplementary Fig. 1b). When all 120 participants were stratified into tertiles based on serum lactate levels, the overall incidence of ED increased significantly with ascending lactate tertiles (Fig. 1d). Moreover, within the DMED subgroup, the incidence of moderate-to-severe ED rose in a lactate-dependent manner (Fig. 1e).
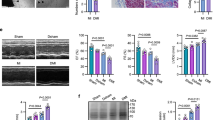
a Schema of the serum lactate detection in humans. b, c The age and plasma lactate in DMED patients and control subjects (n = 60). d The association between serum lactate concentration and ED incidence. Serum lactate levels of 120 participants were stratified into three tertiles (<1/3, 1/3-2/3, >2/3). Chi-square test of independence and post hoc comparison tests were used. e The association between the occurrence of moderate-to-severe ED and serum lactate in the DMED subgroup. Chi-square test of independence and post hoc comparison tests were used. f Schema of lactate detection of serum and penis in mice. g The cavernosal lactate in mice (n = 6). h Pearson’s correlation between cavernosal lactate and plasma lactate levels. i Schematic diagram of treating normal mice with saline or lactate. j The cavernosal lactate in the saline, 0.25 g/kg/d lactate and 0.5 g/kg/d lactate mice (n = 6). k, l Representative images and quantification of ICP (n = 6). m, n Morphology and quantification of the corpus cavernosum assessed by H&E staining. Scale bars, 200 µm, n = 3. o, p Collagen (blue) and smooth muscle (red) evaluated by Masson staining. Scale bars, 200 µm, n = 3. q, r Representative images and quantification of α-SMA (green) immunofluorescence staining. Scale bars, 200 µm, n = 3. s Experiment design in vitro. t Cell viability of primary mouse CCSMCs in control, 5 mM lactate and 10 mM lactate groups, n = 3. u, v Representative images and quantification of cell motility evaluated by Transwell migration assays. Scale bars, 100 µm, n = 3. w, x The protein expressions of SM22α, CALPONIN, VIMENTIN, and OSTEOPOTIN detected by Western blot (n = 3). Bar charts and line charts (t) are presented as mean ± SD. All experiments were performed with at least three biologically independent replicates with consistent results, and the points represent individual biological samples. Unpaired two-sided Student’s t-test (b, c, g) and one-way ANOVA with Tukey’s post hoc test (j, l, n, p, r, v, x) were performed. Source data are provided as a Source Data file. Con control, DMED diabetes mellitus-induced erectile dysfunction, NC normal control, NS normal saline, Lac lactate, ICP intracavernous pressure, CCSMC corpus cavernosum smooth muscle cells.
To further investigate the correlation between lactate and DMED, we established a mouse model of DMED using a combination of low-dose streptozotocin and a high-fat diet. Cavernosal lactate concentrations and intracavernous pressure (ICP) were measured at 0, 4, 8, and 12 weeks post-modeling (Supplementary Fig. 1c). We observed a time-dependent increase in cavernosal lactate levels, accompanied by a progressive decline in ICP (Supplementary Fig. 1d–f). DMED mice also exhibited significantly elevated plasma lactate and blood glucose levels compared to controls (Supplementary Fig. 1g, h). Furthermore, cavernosal tissue in DMED mice showed pronounced lactate accumulation, with its levels positively correlating with plasma lactate concentrations (Fig. 1f–h).
To evaluate the effect of lactate on erectile function, mice were randomly assigned to receive saline or different doses of lactate (Fig. 1i). Serum lactate kinetics analysis showed that the 0.5 g/kg dose significantly elevated the concentration-time curve compared to the 0.25 g/kg group, whereas the 0.75 g/kg dose did not further increase lactate levels within the short term (Supplementary Fig. 1i). Therefore, the 0.25 and 0.5 g/kg doses were selected for subsequent experiments. Exogenous lactate administration elevated cavernosal lactate levels and reduced ICP in a concentration-dependent manner compared with the saline group (Fig. 1j–l). Histological analysis further revealed that lactate treatment significantly reduced the lumen-to-parenchyma ratio, smooth muscle-to-collagen ratio, and α-SMA fluorescence intensity, while increasing Collagen I fluorescence intensity (Fig. 1m–r and Supplementary Fig. 1j, k). These results demonstrate that elevated lactate levels disrupt erectile function by reducing smooth muscle content.
Based on the above findings, we further investigated the toxic effects of lactate accumulation on CCSMCs. Primary CCSMCs were treated with lactate at different concentrations (0, 2, 4, 6, 8, 10, 12 mmol/L). Cell viability was not significantly affected at lactate concentrations up to and including 4 mmol/L (Supplementary Fig. 1l). Thus, 5 and 10 mmol/L were selected as low and high lactate stimulation concentrations, respectively, for subsequent experiments (Fig. 1s). Lactate treatment impaired CCSMCs’ activity and motility in a concentration-dependent manner (Fig. 1t–v). Western blot analysis showed that exposure to high lactate levels downregulated contractile markers (SM22α and CALPONIN) and upregulated synthetic markers (VIMENTIN and OSTEOPONTIN) in CCSMCs (Fig. 1w, x). Transcriptome sequencing further suggested that excessive autophagy activation may underlie the lactate-induced impairment of CCSMCs (Supplementary Fig. 2a–c). In vitro experiments were conducted by pretreating CCSMCs with the autophagy inhibitor 3-methyladenine (3-MA). The results demonstrated that 3-MA significantly abrogates lactate-induced excessive autophagy activation, impairment of cell viability, and phenotypic switching toward the synthetic smooth muscle phenotype. (Supplementary Fig. 2d–l). In animal models, we further analyzed autophagic activity in the corpus cavernosum of lactate-challenged mice and DMED mice. Intriguingly, progressive elevation of the LC3-II/LC3-I ratio was observed in both models during disease progression, whereas P62 protein expression exhibited a dynamic change characterized by an initial decrease followed by a subsequent increase (Supplementary Fig. 3). Building upon these observations, we performed interventions with 3-MA and SMER28, respectively. Our data showed that 3-MA effectively suppressed excessive autophagy and remarkably improved erectile function as well as tissue architecture in DMED mice. In contrast, although SMER28 promoted P62 degradation, it failed to exert significant therapeutic efficacy (Supplementary Fig. 4). Collectively, these results indicate that lactate-induced excessive autophagy is a key mechanism contributing to DMED.
FBP1 is a key factor for cavernosal lactate accumulation in DMED
Lactate serves as a key intermediate in glucose metabolism. Under anaerobic conditions, glycolysis rapidly converts glucose to lactate, while gluconeogenesis reversibly converts lactate back to glucose to maintain systemic glucose homeostasis (Fig. 2a). Consequently, tissue lactate accumulation can arise from either glycolytic overactivation, which leads to excessive lactate production, or impaired gluconeogenesis, which results in inadequate lactate clearance15. We have validated that CCSMC also possesses gluconeogenic function, consistent with what has been observed in smooth muscle cells from other sites16,17 (Supplementary Fig. 5). Using RNA-seq data from our previously established DMED mouse and rat models18, we analyzed the transcription profiles of key genes involved in glycolysis and gluconeogenesis. The results revealed no upregulation of glycolytic pathway genes. Furthermore, a brief exploratory treatment of DMED mice with sodium oxamate, a glycolysis inhibitor, did not significantly reduce cavernosal lactate levels (Supplementary Fig. 6a–c), suggesting that lactate accumulation in the corpus cavernosum primarily results from defective gluconeogenesis rather than enhanced glycolysis. Among gluconeogenic enzymes, Fbp1 showed consistent transcriptional downregulation (Fig. 2b, c). Analysis of public single-cell sequencing data from the Human Male Health Atlas (http://malehealthatlas.cn/)19 also indicated reduced FBP1 expression in the corpus cavernosum of DMED patients (Fig. 2d). Consistent with this, validation in DMED mouse models confirmed significant downregulation of FBP1, whereas the expression of G6PC3 and PCK1 remained unchanged (Fig. 2e–g and Supplementary Fig. 6d–f).
a The interconversion between lactate and glucose. b, c Heatmap of transcriptome sequencing for key genes involved in glycolysis (Hk2, Pfkfb3, Pklr, Ldha) and gluconeogenesis (Fbp1, G6pc3, Pck1) in mice and rats. n1, n2, n3 and N1, N2, N3 denote three independent samples from the control mouse and rat groups, respectively; t1, t2, t3 and T1, T2, T3 represent the corresponding three independent samples from the DMED mouse and rat groups, respectively. d The expression of FBP1 in each group based on single-cell data analysis of the human corpus cavernosum. e Fbp1 mRNA levels in the corpus cavernosum of normal and DMED mice (n = 3). f, g Representational results and quantification of FBP1 immunohistochemical staining among Fbp1+/+, DMED-Fbp1+/+, Fbp1−/+ and DMED-Fbp1−/+ mice. Scale bars, 200 µm, n = 3. h The cavernosal lactate concentrations (n = 3). i, j Representative images and quantification of ICP (n = 3). k, l Morphology and quantification of the corpus cavernosum assessed by H&E staining (n = 3). m, n Representative images and quantification of α-SMA (green) immunofluorescence staining. Scale bars, 200 µm, n = 3. o Cell viability of primary mouse CCSMCs in the low-concentration lactate and Fbp1 knockout groups (n = 3). p Representative images of motility evaluated by Transwell migration assays. Scale bars, 100 µm. q The protein expressions of SM22α, CALPONIN, VIMENTIN, and OSTEOPONTIN were detected by Western blot. r Cell viability in the high-concentration lactate and Fbp1 overexpression groups (n = 3). s Transwell migration assays. Scale bars, 100 µm. t The protein expressions of SM22α, CALPONIN, VIMENTIN, and OSTEOPONTIN in the high-concentration lactate and Fbp1 overexpression groups. Bar charts are presented as mean ± SD. All experiments were performed with at least three biologically independent replicates with consistent results, and the points represent individual biological samples. Unpaired two-sided Student’s t-test (e) and one-way ANOVA with Tukey’s post hoc test (g, h, j, l, n, o, r) were performed. Source data are provided as a Source Data file. DMED: diabetes mellitus-induced erectile dysfunction, NC normal control, ICP intracavernous pressure, CCSMC corpus cavernosum smooth muscle cell, FBP1 fructose-1,6-bisphosphatase 1.
FBP1 catalyzes the hydrolysis of fructose-1,6-bisphosphate into fructose 6-phosphate and inorganic phosphate, playing a critical role in lactate metabolism20. Its deficiency is associated with metabolic disturbances, including hypoglycemia and lactic acidosis21. We therefore asked whether FBP1 downregulation contributes to the development of ED. Our results demonstrated that Fbp1 knockdown significantly increased cavernosal lactate levels and reduced ICP, lumen-to-parenchyma ratio, and α-SMA fluorescence intensity compared to normal mice (Fig. 2h–n). Under low-lactate conditions in vitro, Fbp1 knockout further impaired CCSMC viability, motility, and contractile marker expression (Fig. 2o–q). Conversely, Fbp1 overexpression partially counteracted the detrimental effects of high lactate concentrations on CCSMCs (Fig. 2r–t). Notably, the G260R variant (the glycine at position 260 of the FBP1 is substituted with arginine), a well-characterized enzymatically inactive form of FBP1 linked to metabolic disorders22,23, was found in our study to fail in alleviating the harmful effects of high lactate concentrations on CCSMCs (Fig. 2r–t). These findings indicate that FBP1 modulates lactate-induced effects on CCSMCs in an enzyme activity-dependent manner.
Aberrant accumulation of H3K9me3 or H3K27me3 in the Fbp1 promoter epigenetically represses its transcription in DMED
Transcriptional regulation represents a fundamental mechanism controlling gene expression. As previously described, the transcriptional downregulation of Fbp1 was observed across species in penile cavernosal tissues. Given that histone methylation modifications can induce epigenetic repression of gene expression by altering chromatin structure, and have been implicated in various diabetic complications24,25,26, we hypothesized that these marks might contribute to Fbp1 silencing. To identify potential repressive histone modifications on the Fbp1 promoter, we performed an initial screening using the ChIP-Atlas database (https://chip-atlas.org/). The analysis revealed significant enrichment of the repressive marks histone H3 lysine 27 trimethylation (H3K27me3)27 and histone H3 lysine 9 trimethylation (H3K9me3)28 within the Fbp1 promoter region (Fig. 3a). We further validated these findings through ChIP-qPCR assays, which confirmed strong enrichment of both H3K9me3 and H3K27me3 at the Fbp1 promoter in MOVAS cells and primary mouse CCSMCs, and also characterized additional epigenetic features of the Fbp1 promoter (Fig. 3b, c and Supplementary Fig. 7a–g). Furthermore, we demonstrated that the enrichment of H3K9me3 or H3K27me3 were significantly elevated in the corpus cavernosum of DMED mice compared to normal mice (Fig. 3d, e). Since histone methylation is catalyzed by specific methyltransferases, we screened for enzymes associated with H3K9me3 and H3K27me3. This revealed substantial upregulation of suppressor of variegation 3-9 homolog 1 (SUV39H1) and enhancer of zeste homolog 2 (EZH2) in DMED mice (Fig. 3f, g). Overexpression of Suv39h1 increased H3K9me3 levels and inhibited the FBP1 expression in both MOVAS cells (Fig. 3h, i) and CCSMCs (Supplementary Fig. 7h–i). Conversely, Suv39h1 knockdown using siRNA or inhibition with its specific inhibitor chaetocin29 led to the upregulation of FBP1 (Fig. 3j–m and Supplementary Fig. 7j–m). Similarly, Ezh2 overexpression elevated H3K27me3 and repressed FBP1, whereas Ezh2 knockdown or inhibition with EPZ643830 increased FBP1 expression (Supplementary Figs. 7n–s and 8). Given that the EZH2–H3K27me3 axis has been previously implicated in FBP1 regulation31,32,33, we focused on identifying readers of H3K9me3 at the Fbp1 promoter. Integrated analysis using ChIP-Atlas and ChIP-qPCR identified chromobox protein homolog 3 (CBX3), a known reader of H3K9me334, binding to the Fbp1 promoter (Fig. 3n, o and Supplementary Fig. 7t). CBX3 enrichment was significantly higher in DMED mice compared to normal mice (Fig. 3p, q). We further demonstrated that CBX3 negatively regulates FBP1 expression in vitro (Fig. 3r–y and Supplementary Fig. 7u–z), and to mechanistically target this regulation, we employed PROTAC technology35 to develop a selective CBX3 degrader that induces its ubiquitination-mediated degradation (Supplementary Fig. 9). Collectively, these findings indicate that aberrant enrichment of H3K9me3 and H3K27me3 at the Fbp1 promoter epigenetically represses its transcription in DMED.
a Histone methylation modifiers binding to the FBP1 promoter sequence identified from the ChIP-Atlas public database. b, c The ChIP-qPCR assay confirming the interaction between H3K9me3/H3K27me3 and Fbp1 in MOVAS cells (n = 3). d, e The protein expressions of H3K9me3 and H3K27me3 in mice (n = 3). f, g The protein expressions of methyltransferases related to H3K9me3 and H3K27me3 in mice (n = 3). h, i Effects of Suv39h1 overexpression on H3K9me3 as well as on FBP1 protein and Fbp1 mRNA levels in MOVAS cells (n = 3). j, k Effects of siRNA-mediated knockdown of Suv39h1 on H3K9me3 as well as on FBP1 protein and Fbp1 mRNA levels in MOVAS cells (n = 3). l, m Effects of a Suv39h1 inhibitor (chaetocin, 25, 50 nM) on FBP1 protein and Fbp1 mRNA levels in MOVAS cells (n = 3). n CBX3 binding to the Fbp1 promoter sequence identified from the ChIP-Atlas public database. o The ChIP-qPCR assay confirming the interaction between CBX3 and Fbp1 in MOVAS cells (n = 3). p, q The protein expressions of CBX3 in mice (n = 3). r, s Effects of Cbx3 overexpression on FBP1 protein and Fbp1 mRNA levels in MOVAS cells (n = 3). t, u Effects of siRNA-mediated knockdown of Cbx3 on FBP1 protein and Fbp1 mRNA levels in MOVAS cells (n = 3). v, w Effects of the H-PROTAC (1 μM, 2 μM) on FBP1 protein and Fbp1 mRNA levels in MOVAS cells (n = 3). x The effects of exogenous CBX3 on FBP1 protein level in MOVAS cells. y Mechanism diagram. Bar charts are presented as mean ± SD. All experiments were performed with at least three biologically independent replicates with consistent results, and the points represent individual biological samples. Unpaired two-sided Student’s t-test (b, c, e, g, i, q, s) and one-way ANOVA with Tukey’s post hoc test (k, m, o, u, w) were performed. Source data are provided as a Source Data file. NC normal control, DMED diabetes mellitus-induced erectile dysfunction, FBP1 fructose-1,6- bisphosphatase 1, H3K9me3 histone H3 lysine 9 trimethylation, H3K27me3 histone H3 lysine 27 trimethylation, SUV39H1 suppressor of variegation 3-9 homolog 1, EZH2 enhancer of zeste homolog 2, EV empty vector, CBX3 chromobox protein homolog 3.
The effects of H3K9me3- and H3K27me3- related inhibitors on DMED mice
Next, we evaluated the effects of the three inhibitors on Fbp1 function by measuring glucose consumption and lactate production. Treatment with inhibitors or siRNA targeting Cbx3, Suv39h1, or Ezh2 significantly reduced both glucose consumption and lactate production in cultured cells. However, with the exception of siCbx3 (Fig. 4b), most inhibitors failed to reverse the metabolic alterations caused by Fbp1 knockout, as reflected in lactate production and glucose uptake (Fig. 4a–l). Moreover, inhibition of Cbx3, Suv39h1, or Ezh2 did not restore Fbp1 expression in Fbp1-knockout cells (Fig. 4m–o and Supplementary Fig. 10a–c). These results indicate that Fbp1 acts as a key downstream effector of Cbx3, Suv39h1, and Ezh2, although Cbx3 and Suv39h1 may also modulate lactate and glucose metabolism through additional pathways. Subsequently, we administered H-PROTAC, chaetocin, and EPZ6438 to DMED mice and Fbp1−/+ mice, respectively. Although the three inhibitors elicited statistically significant effects in DMED mice, the improvements were modest: they led to only a slight reduction in cavernosal lactate levels and partial recovery of ICP, tissue morphology, and smooth muscle content. Notably, none of the inhibitors produced any beneficial effect in Fbp1−/+ mice (Fig. 4p–u). Although combined inhibition of both marks led to stronger Fbp1 upregulation in vitro (Supplementary Fig. 10d, e), the combined in vivo administration of the three agents—chaetocin (intraperitoneally), EPZ6438 (orally), and H-PROTAC (intracavernously)—was poorly tolerated in DMED mice. Therefore, rather than pursuing complex combination regimens, it is more compelling to explore alternative mechanisms governing FBP1 expression and function.
a–l The results of glucose consumption in MOVAS cells (n = 3). m Effects of the H-PROTAC on FBP1 expression in MOVAS cells with Fbp1 knockout. n Effects of the chaetocin on FBP1 expression in MOVAS cells with Fbp1 knockout. o Effects of the EPZ6438 on FBP1 expression in MOVAS cells with Fbp1 knockout. p The cavernosal lactate concentrations (n = 3). q, r Representative images and quantification of ICP (n = 3). s, t Morphology and quantification of the corpus cavernosum assessed by H&E staining. Scale bars, 200 µm, n = 3. u Representative images of α-SMA (green) immunofluorescence staining. Scale bars, 200 µm. Bar charts are presented as mean ± SD. All experiments were performed with at least three biologically independent replicates with consistent results, and the points represent individual biological samples. Unpaired two-sided Student’s t-test (a–l) and one-way ANOVA with Tukey’s post hoc test (p, r, t) were performed. Source data are provided as a Source Data file. DMED diabetes mellitus-induced erectile dysfunction, ICP intracavernous pressure, FBP1 fructose-1,6-bisphosphatase 1, SUV39H1 suppressor of variegation 3-9 homolog 1, EZH2 enhancer of zeste homolog 2, CBX3 chromobox protein homolog 3.
ZDHHC13 mediated S-palmitoylation of FBP1 at C282 site
The above findings indicate that although histone methylation modifications strongly suppress Fbp1 transcription in DMED, interventions targeting these epigenetic regulators failed to robustly restore erectile function in vivo. This discrepancy led us to reevaluate the regulatory landscape of FBP1 and hypothesize that its activity might be further modulated by post-translational modifications (PTMs). Given the critical role of the G260 residue in FBP1 enzymatic function and gluconeogenesis, we focused on identifying potential PTMs near this key site. Analysis of all PTMs within ±25 amino acids around the G260 of FBP1 has shown that, apart from the previously reported acetylation at K26936 and phosphorylation at Y26537, a palmitoylation site at C282 has been identified by using the CSS-palm 4.0 tool (Fig. 5a). Palmitoylation of FBP1 was confirmed via acyl-biotinyl exchange (ABE) assay in cells expressing HA-tagged Fbp1, with streptavidin blotting revealing a clear palmitoylation signal (Fig. 5b and Supplementary Fig. 11a). This finding was further validated using biotin-alkyne labeling coupled with click chemistry, which showed increased palmitoylated proteins in Fbp1-transfected cells; hydroxylamine treatment abolished this signal, confirming its specificity (Fig. 5c and Supplementary Fig. 11b, c). We also demonstrated that endogenous FBP1 undergoes palmitoylation (Supplementary Fig. 11d). Sequence alignment revealed that C282, located adjacent to the functional G260 residue, is evolutionarily conserved across species (Fig. 5d). To identify the enzyme responsible for FBP1 palmitoylation, we generated a series of Zdhhc knockout cell lines using CRISPR technology. Among these, knockout of zinc finger DHHC-type containing 13 (Zdhhc13) markedly reduced FBP1 palmitoylation (Fig. 5e), suggesting it acts as a key palmitoyltransferase for FBP1. The direct physical interaction between FBP1 and ZDHHC13 was further confirmed by molecular docking, co-immunoprecipitation, confocal imaging, and GST pull-down assays (Fig. 5f–j and Supplementary Fig. 11e). Transcriptome analysis and immunohistochemistry of the corpus cavernosum showed that ZDHHC13 expression was elevated in DMED mice (Fig. 5k, l), suggesting its involvement in FBP1 palmitoylation in vivo. In cellular models, knockdown of Zdhhc13 decreased FBP1 palmitoylation, whereas its overexpression increased palmitoylation levels (Fig. 5m, n and Supplementary Fig. 11f, g). Mutation of the cysteine at position 282 to serine (FBP1-C282S) abolished FBP1 palmitoylation (Fig. 5o and Supplementary Fig. 11h). To clarify the enzyme dependency, we reintroduced either wild-type ZDHHC13 (ZDHHC13-WT) or an enzymatically inactive mutant (ZDHHC13-C456S)38 into Zdhhc13-knockout cells. Only ZDHHC13-WT restored FBP1 palmitoylation, confirming the requirement for ZDHHC13 enzymatic activity (Fig. 5p and Supplementary Fig. 11i, j). To rule out interference from endogenous FBP1, we expressed FBP1‑WT or FBP1‑C282S in Fbp1-knockout cells. Exogenous FBP1‑WT underwent palmitoylation, whereas the FBP1‑C282S mutant did not (Fig. 5q and Supplementary Fig. 11k, l). Finally, the palmitoylation assay confirmed that ZDHHC13‑WT mediated palmitoylation of FBP1‑WT but not FBP1‑C282S, and ZDHHC13‑C456S failed to catalyze the modification (Fig. 5r). Together, these data demonstrate that ZDHHC13 catalyzes S-palmitoylation of FBP1 at cysteine 282.
a Analysis of all PTMs within ±25 amino acids around the G260 of FBP1. b, c ABE and Click-iT pull-down assay detecting the palmitoylation of FBP1 in MOVAS cells. d Prediction of potential FBP1 palmitoylation sites using CSS-Palm 4.0 software. e MOVAS cells were infected with lentiviruses containing expression cassettes of guide RNAs targeting different Zdhhcs. The ABE assay was used for detecting the palmitoylation status of FBP1. f Alphafold and PyMOL3 was used to conduct a visualization study of the molecular docking between ZDHHC13 and FBP1. g Antibodies to ZDHHC13 or FBP1 were used to perform co-immunoprecipitation experiments in MOVAS cells. h Confocal microscopy analysis of co-localization between ZDHHC13 (green) and FBP1 (red) in MOVAS cells. Scale bars, 15 µm. i Schematic diagram. j Western blotting analysis of ZDHHC13 GST-pulled down by FBP1 recombinant. k Heatmap of transcriptome sequencing of Zdhhc13 in normal and DMED mice. l Representational results of FBP1 immunohistochemical image. Scale bars, 100 µm. m, n FBP1 palmitoylation was examined in MOVAS or CCSMCs with Zdhhc13 knockdown or overexpression. o MOVAS cells were transfected with the indicated plasmids for 48 h. The ABE assay was used to detect the palmitoylation level of FBP1. p The effect of ZDHHC13 C456S on FBP1 palmitoylation in MOVAS cells. q Endogenous Fbp1 were knocked out in MOVAS cells by CRISPR/Cas9. The ABE assay was used to detect the palmitoylation level of FBP1. r The palmitoylation analysis was performed by mixing purified His-FBP1-WT and His-FBP-C282S with purified GST-ZDHHC13-WT or GST-ZDHHC13-C456S in the presence of palmitoyl-alkyne-CoA. All experiments were performed with at least three biologically independent replicates with consistent results. Source data are provided as a Source Data file. PTM post-translational modification, FBP1 fructose-1,6-bisphosphatase 1, ABE acyl-biotinyl exchange, ZDHHC13 zinc finger DHHC-type containing 13, CCSMC corpus cavernosum smooth muscle cells.
ZDHHC13-mediated S-palmitoylation of FBP1 impairs its gluconeogenic function
S-palmitoylation is known to regulate protein stability, localization, and function39,40. To investigate how ZDHHC13-mediated S-palmitoylation influences FBP1, we initially assessed its effects on protein degradation and subcellular localization, but observed no significant changes in either (Fig. 6a, b). We then employed PyMOL to assess whether palmitoylation at C282 induces conformational changes in FBP1. Palmitoylation reduced the solvent-accessible surface area (SASA) of the critical G260 residue by ~20% (from 3.296 to 2.657 Ų), while the total SASA of chain A decreased by ~388 Ų (Supplementary Fig. 12a). Together, these data suggest that palmitoylation at C282 promotes a more compact conformational state of FBP1, characterized by reduced solvent exposure of G260 and enhanced local structural folding. Fbp1 knockout significantly elevated glucose consumption, lactate production, and extracellular acidification rate (ECAR), while suppressing the oxygen consumption rate (OCR). Reintroduction of the Fbp1 C282S mutant into Fbp1‑knockout cells produced a greater reduction in glucose consumption, lactate production, and ECAR, alongside a more substantial increase in OCR (Fig. 6c–f and Supplementary Fig. 12b–g). Knockdown of Zdhhc13 significantly decreased glucose consumption, lactate production, and ECAR, and increased OCR. However, these metabolic changes were abolished in Fbp1-knockout cells (Fig. 6g–j and Supplementary Fig. 12h–k). Conversely, overexpression of Zdhhc13 increased glucose consumption, lactate production, and ECAR, while decreasing OCR; again, these effects were absent in Fbp1-knockout cells (Fig. 6k–n and Supplementary Fig. 12l–o). The enzymatically inactive mutant Zdhhc13-C456S had no significant effect on glycolytic activity or lactate production (Supplementary Fig. 12p–w). Furthermore, in Fbp1-knockout cells reconstituted with the Fbp1 C282S mutant, Zdhhc13 overexpression failed to alter glucose consumption or lactate production (Supplementary Fig. 12x, y). Finally, we injected an adeno-associated virus (AAV) encoding Zdhhc13-targeting shRNA into the corpus cavernosum of DMED mice and Fbp1−/+ mice, respectively (Supplementary Fig. 12z). Knockdown of Zdhhc13 partially reduced cavernosal lactate levels and moderately improved ICP in DMED mice; however, it produced no effect in Fbp1−/+ mice (Fig. 6o–t). These results underscore that targeting Zdhhc13 alone provides limited therapeutic benefit in DMED.
a The cycloheximide (50 µg/ml) chase assay with protein extraction at 0, 2, 4, and 8 h, and then analyzed by Western blot. b Western blot analysis of cytoplasmic and nuclear proteins. c–n The glucose consumption, lactate production, ECAR and OCR of MOVAS cells in groups (n = 3). o The cavernosal lactate concentrations (n = 3). p, q Representative images and quantification of ICP (n = 3). r, s Morphology and quantification of the corpus cavernosum assessed by H&E staining. Scale bars, 200 µm, n = 3. t Representative images of α-SMA (green) immunofluorescence staining. Scale bars, 100 µm. Bar charts are presented as mean ± SD. All experiments were performed with at least three biologically independent replicates with consistent results, and the points represent individual biological samples. Unpaired two-sided Student’s t-test were performed. Source data are provided as a Source Data file. CHX cycloheximide, FBP1 fructose-1,6-bisphosphatase 1, ZDHHC13 zinc finger DHHC-type containing 13, DMED diabetes mellitus-induced erectile dysfunction, ICP intracavernous pressure. ECAR extracellular acidification rate, OCR oxygen consumption rate, AAV adeno-associated virus.
Building on these limitations, we next sought to elucidate the mechanism underlying Zdhhc13 upregulation in DMED. We analyzed differentially expressed genes using KEGG pathway enrichment, which revealed significant activation of the PI3K-AKT pathway—the most upregulated signaling pathway in DMED (Supplementary Fig. 13a). Given established knowledge of PI3K-AKT signaling in regulating SRY-related HMG-box gene 9 (SOX9) activity41,42, we investigated this potential connection. Interestingly, ChIP-Atlas data indicated SOX9 binding to the promoters of both Zdhhc13 and Cbx3 (Supplementary Fig. 13b). Consistent with these findings, we confirmed elevated phosphorylated AKT and SOX9 levels in the cavernosum of DMED mice (Supplementary Fig. 13c). Sox9 knockdown downregulated CBX3 and ZDHHC13 expression, while its overexpression increased them (Supplementary Fig. 13d–i). However, when we administered AAV-shSox9 to the cavernosum of DMED mice, we observed no significant improvement in cavernosal lactate levels or ICP (Supplementary Fig. 13j–m). This unexpected outcome prompted us to explore alternative strategies capable of concurrently addressing both Fbp1 transcriptional repression and palmitoylative inactivation.
The effect of lipid nanoparticles loaded with Fbp1-C282S mRNA on DMED mice
Currently, single-site inhibitors demonstrate only modest efficacy in vivo, failing to substantially ameliorate elevated lactate levels or restore erectile function in DMED mice. Furthermore, combination therapy with multiple agents may provoke serious adverse reactions. To address this challenge, we synthesized the Fbp1-C282S mRNA using site-directed mutagenesis, which was subsequently encapsulated in lipid nanoparticles (Fbp1-C282S-LNP) via the Ignite NanoAssay system (Fig. 7a). The Fbp1-C282S-LNPs exhibited an average particle size of 99.3 nm, polydispersity index (PDI) of 0.083, encapsulation efficiency of 90.8%, and zeta potential of −3.44 mV (Fig. 7b). Scanning electron microscopy was used to further characterize the nanoparticles (Fig. 7c). The Fbp1-C282S-LNPs significantly upregulated FBP1 level without exhibiting obvious cytotoxicity in primary mouse CCSMCs (Fig. 7d–f). Functional assays confirmed that the Fbp1-C282S-LNPs also markedly downregulated glucose consumption, lactate production and ECAR, while upregulating cellular OCR (Fig. 7g–n).
a Schematic diagram of Fbp1-C282S-LNP. b The average particle size, PDI encapsulation efficiency and zeta potential (n = 3). c The representative images of canning electron microscopy. Scale bars, 50 nm. d, e FBP1 expression in CCSMCs assessed by Western blot (n = 3). f Cell viability in CCSMCs (n = 5). g–n The glucose consumption, lactate production, ECAR, and OCR of MOVAS cells in groups (n = 3). o The cavernosal lactate concentrations (n = 3). p, q Representative images and quantification of ICP (n = 3). r, s Morphology and quantification of the corpus cavernosum assessed by H&E staining. Scale bars, 200 µm, n = 3. t, u Representative images and quantification of α-SMA (green) immunofluorescence staining. Scale bars, 100 µm, n = 3. v, w Representative images and quantification of Collagen I (red) immunofluorescence staining. Scale bars, 100 µm, n = 3. x, y The FBP1 protein expression and quantification (n = 3). Bar charts are presented as mean ± SD. All experiments were performed with at least three biologically independent replicates with consistent results, and the points represent individual biological samples. The one-way ANOVA with Tukey’s post hoc test were performed. Source data are provided as a Source Data file. FBP1 fructose-1,6-bisphosphatase 1, PDI polydispersity index, LNP lipid nanoparticle, Col I Collagen I, DMED diabetes mellitus-induced erectile dysfunction, ICP intracavernous pressure, ECAR extracellular acidification rate, OCR oxygen consumption rate.
Subsequently, we encapsulated Flag-tagged Fbp1-C282S mRNA into LNPs and evaluated cellular uptake within the corpus cavernosum using multiplex immunofluorescence staining. Robust LNPs uptake was observed in CCSMCs, with detectable internalization also occurring in endothelial cells and fibroblasts (Supplementary Fig. 14a). Additionally, in vivo assessments confirmed this formulation induced no significant hepatic or renal toxicity in mice (Supplementary Fig. 14b–e). Concurrently, we completed pharmacokinetic and tissue distribution studies of Fbp1-C282S-LNPs to define their in vivo profiles (Supplementary Fig. 14f, g). To assess dose-dependent therapeutic efficacy, DMED mice received intracavernous injections of Fbp1-C282S-LNPs at 0.1, 0.25, or 0.5 mg/kg. The 0.1 mg/kg group showed no significant improvement in cavernosal lactate levels or erectile function relative to controls. In contrast, the 0.25 mg/kg group significantly reduced lactate levels, increased FBP1 expression, improved ICP, enhanced smooth muscle content, ameliorated histological structure, and reduced cavernous fibrosis. However, the 0.5 mg/kg group provided no additional therapeutic benefit over the 0.25 mg/kg dose (Fig. 7o–y). We validated that Fbp1 C282S-LNP treatment can also efficiently reverse autophagic abnormalities in the corpus cavernosum of DMED mice (Supplementary Fig. 14h-j). Results of subsequent multi-tissue validation demonstrated that, in contrast to the significant increase in FBP1 protein expression observed in the corpus cavernosum following treatment, FBP1 protein expression in the liver and skeletal muscle exhibited an upward trend but failed to reach statistical significance (Supplementary Fig. 15). Furthermore, Fbp1-C282S-LNPs demonstrated sustained efficacy in long-term studies of DMED mice (Supplementary Fig. 16) and consistently improved outcomes across multiple diabetic models, including type 1 diabetes (Supplementary Fig. 17), db/db mice (Supplementary Fig. 18), and Fbp1−/+ mice (Supplementary Fig. 19). These results firmly establish Fbp1-C282S-LNP as a highly promising therapeutic strategy for DMED.
Discussion
This study reveals that dysregulated cavernous lactate metabolism is a central pathogenic mechanism in DMED, offering insights for understanding its progression and treatment. Previous studies have established that lactate accumulation contributes significantly to diabetic complications43,44,45. Consistent with our clinical observation of elevated serum lactate in DMED patients, experiments in DMED mice demonstrated markedly elevated levels of both serum and cavernosal lactate, reflecting a severe local and systemic metabolic disturbance. Both in vivo and in vitro experiments demonstrated that exogenous lactate impairs erectile function and compromises CCSMC performance, accompanied by a shift from contractile to synthetic phenotypes. This aligns with prior reports indicating that lactate disrupts smooth muscle physiology in various contexts46,47. Our findings highlight the importance of maintaining metabolic homeostasis within the cavernous microenvironment and identify lactate modulation as a promising therapeutic strategy for DMED. Transcriptional repression and functional dysfunction of FBP1 are key causes of penile cavernosal lactate accumulation in DMED. Mechanistically, H3K9me3 mediated by the SUV39H1/CBX3 complex and H3K27me3 mediated by EZH2 cooperatively suppress Fbp1 transcription; meanwhile, ZDHHC13-mediated palmitoylation of the FBP1 protein at the C282 site directly impairs its gluconeogenic function, exacerbating lactate accumulation (Fig. 8a). To target the aforementioned aberrant mechanisms, we developed Fbp1 C282S-LNP, which restores FBP1 expression and blocks pathological palmitoylation via intracavernosal injection, thereby effectively reducing local lactate levels (Fig. 8b).
a Mechanisms of FBP1 dysfunction and lactate accumulation in DMED. H3K9me3-mediated repression of Fbp1 transcription by SUV39H1/CBX3 (targeted by Chaetocin and H-PROTAC), H3K9me3-mediated repression of Fbp1 transcription by EZH2 (inhibited by EPZ6438), and ZDHHC13-mediated palmitoylation impairing FBP1 enzymatic function. b Production and intracavernosal administration of Fbp1 C282S-LNP reduces cavernous lactate levels and rescues erectile function in DMED mice. SUV39H1 suppressor of variegation 3-9 homolog 1, CBX3 chromobox protein homolog 3, FBP1 fructose-1,6-bisphosphatase 1, EZH2 enhancer of zeste homolog 2, ZDHHC13 zinc finger DHHC-type containing 13, LNP Lipid nanoparticle, DMED diabetes mellitus-induced erectile dysfunction, H3K9me3 histone H3 lysine 9 trimethylation, H3K27me3 histone H3 lysine 27 trimethylation.
FBP1, a key gluconeogenic enzyme, serves as a critical regulator that balances glycolytic and gluconeogenic flux to constrain lactate accumulation and maintain glucose homeostasis. Previous studies have linked FBP1 to keratinocyte differentiation, with its deficiency contributing to aberrant epidermal formation and psoriasis48. A landmark study further identified FBP1 as a metabolic checkpoint governing the transition from metabolic dysfunction-associated steatohepatitis to hepatocellular carcinoma via P53-mediated senescence pathways49. Our study now establishes FBP1 as a central metabolic regulator in DMED. Specifically, we demonstrate that downregulation of FBP1 disrupts gluconeogenesis and drives cavernosal lactate accumulation, thereby providing a molecular explanation for DMED pathogenesis.
As a fundamental epigenetic mechanism, histone methylation plays a pivotal role in the regulation of gene expression. In diabetic nephropathy, for example, histone methylation-mediated activation of the NOTCH1 signaling pathway has been shown to drive glomerular fibrosis50. In our study, we found that SUV39H1-mediated H3K9me3 and EZH2-mediated H3K27me3 bind to the Fbp1 promoter and suppress its expression in the corpus cavernosum of DMED mice. Moreover, we identified CBX3—a reader of H3K9me3, as being significantly enriched in DMED mice, where it negatively regulates FBP1 expression. Specifically, SUV39H1-mediated H3K9 methylation recruits CBX3 proteins, which in turn promote further SUV39H1 recruitment and heterochromatin spreading51. Similarly, H3K27me3, catalyzed primarily by EZH2 within the PRC2 complex, has been extensively characterized31,32,33. Although several histone methyltransferase inhibitors (e.g., Chaetocin and EPZ6438) have been developed for preclinical studies29,30, they exhibited limited efficacy in DMED mice, underscoring the disease’s multifactorial pathogenesis and the involvement of interconnected regulatory networks.
Beyond transcriptional regulation, our study identifies protein palmitoylation as a key mechanism influencing FBP1 function in DMED. Among the 23 mammalian zinc finger DHHC-domain containing proteins (ZDHHC1–24, excluding ZDHHC10)52, ZDHHC13, also known as HIP14L, has been implicated in various diseases, including diabetes, cancer, and osteoporosis53,54. The recent studies have revealed that palmitoylation can also directly induce conformational changes in cytoplasmic enzymes, thereby regulating their activity55,56—and this is highly consistent with the phenomenon we observed in FBP1. We demonstrated that ZDHHC13 acts as the specific palmitoyltransferase for FBP1, catalyzing S-palmitoylation at residue C282 and consequently impairing FBP1 function. Structural modeling suggested that palmitoylation at C282 reduces solvent accessibility around the essential G260 residue and induces global compaction of FBP1, although initial screening showed no effect on its degradation or localization. However, inhibition of Zdhhc13 alone produced limited therapeutic benefit in DMED mice, suggesting that targeting this individual regulator may be insufficient to reverse the multifactorial pathology of DMED.
Maintaining normal FBP1 expression and function represents a promising approach to reducing cavernosal lactate levels. However, Fbp1 is subject to multifaceted regulation in the diabetic microenvironment, and existing monotherapies have shown limited efficacy in restoring erectile function in DMED mice, while combination regimens raise concerns about adverse effects. Thus, identifying the strategy that can synergistically restore FBP1 activity remains essential. Advances in LNP-based mRNA therapeutics have accelerated their transition into clinical trials over the past decade57,58. In this study, we developed an LNP-based formulation encapsulating Fbp1-C282S mRNA. Administration of these nanoparticles effectively reduced cavernosal lactate and significantly improved erectile function in a DMED mouse model, highlighting the therapeutic potential of this strategy. Phosphodiesterase 5 (PDE5) inhibitors, first-line treatments for erectile dysfunction, act by maintaining intracellular cGMP levels to facilitate smooth muscle relaxation and penile hemodynamics59. However, they target only vasodilatory signaling and do not ameliorate underlying smooth muscle structural pathology, which limits their efficacy in DMED60. In contrast, the Fbp1-C282S-LNPs delivers therapeutic mRNA to injured regions, reducing local lactate accumulation and facilitating functional recovery—mechanisms that target structural impairments beyond the scope of PDE5 inhibitors. The complementary mechanisms of these two approaches suggest potential synergy in combination therapy. Fbp1-C282S-LNPs may restore tissue integrity and metabolic homeostasis, establishing a functional structural basis for erection, while PDE5 inhibitors enhance signal-mediated vasodilation. Such a strategy could be particularly beneficial for patients with PDE5 inhibitor-resistant DMED.
However, several limitations of this study warrant acknowledgment. First, while we demonstrated that histone methylation and palmitoylation critically regulate FBP1, other unidentified pathways may contribute to FBP1 dysregulation in DMED. Second, although Fbp1-C282S-LNP shows efficient uptake by CCSMCs, lactate reduction likely involves synergistic effects across multiple tissue compartments. Among these mechanisms, endothelial cells regulate penile blood flow via releasing vasoactive substances (e.g., nitric oxide)61; meanwhile, CCSMC impairment is increasingly recognized as a direct contributor to ED pathogenesis, supported by recent advances in the field62,63. Thirdly, although the precise mechanisms underlying such multi-tissue and multi-cellular coordination remain incompletely defined, the broad cellular uptake capacity of Fbp1-C282S-LNP likely contributes to comprehensive improvement of the cavernous metabolic microenvironment, thereby cooperatively maintaining erectile function. Finally, although sustained treatment effects were observed following short-term intervention in this study, further investigation is warranted to elucidate the long-term safety profile and optimal dosing frequency of Fbp1 C282S-LNP. Subsequent studies employing extended multi-dose regimens in larger animal models will be valuable to comprehensively assess chronic toxicity and maximize the translational potential of this strategy. In summary, this study deciphers the molecular mechanism through which FBP1 governs cavernosal lactate metabolism in DMED and develops Fbp1-C282S-LNP as a targeted therapeutic strategy. Further optimization of delivery systems and treatment protocols will be essential to enhance clinical translatability and safety.
Method
Clinical specimen collection
The study protocol was approved by the Institutional Review Board (IRB) of the Second Xiangya Hospital of Central South University (Approval No. LYEC2025-0188). The collection of clinical patient information and specimens was registered with the Chinese Clinical Trial Registry (ChiCTR; registration number: ChiCTR2500107001) as an observational case-control study. Written informed consent was obtained from all participants prior to sample collection. All participants were assessed using the IIEF-5 questionnaire; those with a score below 22 were defined as having DMED. Venous blood samples were collected for subsequent lactate measurement. No therapeutic or experimental interventions were administered to the participants during the study period.
Reagents
Apomorphine, streptozotocin, and lactate were purchased from Sigma-Aldrich (USA). The lactic acid assay kit was purchased from Nanjing Jiancheng Bioengineering Research Institute (China). Glucose test kit and Nuclear and cytoplasmic protein extraction kit were obtained from Beyotime Biotechnology (China). The primary antibodies used were as follows: α-SMA (#GB111364, Servicebio, 1:1000), SM22α (#60213, Proteintech, 1:5000), CALPONIN (#13938, Proteintech, 1:5000), VIMENTIN (#60330, Proteintech, 1:20,000), OSTEOPONIN (#83341, Proteintech, 1:1000), FBP1 (#12842, Proteintech, 1:500), H3K9me3 (#ab8898, Abcam, 1:1000), H3K27me3 (#ab6002, Abcam, 1:1000), SUV39H1 (#10574, Proteintech, 1:2000), SUV39H2 (#11338, Proteintech, 1:1000), SETDB1 (#66293, Proteintech, 1:20,000), SETDB2 (#ab13712, Abcam, 1:500), EZH1 (#ab263961, Abcam, 1:1000), EZH2 (#66476, Proteintech, 1:5000), β-ACTIN (#60008, Proteintech, 1:5000), CBX3 (#66446, Proteintech, 1:2000), CBX5 (#11831, Proteintech, 1:500), CBX1(#10241, Proteintech, 1:1000), ZDHHC13 (#24759, Proteintech, 1:500), FLAG (#66008, Proteintech, 1:5000), HA-tag (#51064, Proteintech, 1:5000), HIS-tag (#66005, Proteintech, 1:5000), AGK (#ab137616, Abcam, 1:1000), GST(#2622, Cell Signaling Technology,1:500), Histone H3 (#17168, Proteintech, 1:1000), LC3 (#14600, Proteintech, 1:1000), P62 (#18420, Proteintech, 1:1000), G6PC (#A21168, ABclonal, 1:2000), PCK1 (#GB11966, Servicebio, 1:500), Phospho-RPB1(Ser2) (#13499, Cell Signaling Technology,1:500), H3K4me3 (#39915, Proteintech, 1:1000), H3K27ac (#A22264, ABclonal, 1:5000), H2AK119ub (#8240, Cell Signaling Technology, 1:500), CRBN (#28494, Proteintech, 1:2000), Phospho-AKT (#66444, Proteintech, 1:1000), AKT (#10176, Proteintech, 1:1000), SOX9 (#67439, Proteintech, 1:5000), vWF (#GB120019, Servicebio, 1:500), and Collagen I (#GB11022, Servicebio, 1:1000). 13C-labeled sodium lactate (#HY-B0389S14), 3-MA (#HY-19312), SMER28 (#HY-100200), lactate Chaetocin (#HY-N2019), EPZ6438 (#HY-13803), and sodium oxamate (#HY-W013032A) were purchased from the MedChemExpress. H-PROTAC was developed by our research group. AAV was provided by Ji-kai Biotechnology (Shanghai, China). As previously mentioned64, Flag-Suv39h1, Flag-Cbx3, Flag-Ezh2, Flag-Zdhhc13, and HA-Fbp1 were constructed by cloning the corresponding cDNA sequences into the OmicLinkTM Expression Clone (CMV Promoter; GeneCopoeia, EX-V0006-M14). The point mutations of Zdhhc13 and Fbp1 were synthesized by using the Hieff Mut Multi Site-Directed Mutagenesis Kit (#11004ES10, Yeasen Biotechnology). The siRNAs were obtained from RiboBio. The sequences of the siRNAs and single-guide RNAs (sgRNAs) are provided in Supplementary Table S1. Cells were transfected with the plasmids or siRNAs using Lipofectamine 2000 (Thermo Fisher Scientific). Other reagents used in this study are detailed in the respective sections.
Cells
The MOVAS cell line (CRL-2797), derived from mouse aortic smooth muscle, was obtained from Xiamen Immocell Biotechnology (China). Primary mouse CCSMCs were isolated using the tissue block method. Briefly, the penis of an 8-week-old C57BL/6 male mouse was excised, and the glans penis, urethra, dorsal blood vessels, and nerves were removed. The remaining tissue was thoroughly washed before being finely minced into ~1 mm³ fragments. These tissue pieces were carefully placed at the base of a culture flask. The flask was then inverted and incubated for 4 h to facilitate initial adhesion. Subsequently, Dulbecco’s Modified Eagle’s Medium (C11885500BT, Gibco) supplemented with 20% bovine calf serum was gently added to the culture to support continuous cell growth62.
Cell viability
Cell viability was evaluated using the Cell Counting Kit-8 (CCK-8, DOJINDO) assay. Briefly, cells were treated with lactate at various concentrations for different durations. After the treatment periods, 100 μL of the CCK-8 working solution was added to each well according to the manufacturer’s instructions. The optical density values were measured, which reflect viable cell numbers since the assay quantifies dehydrogenase activity present only in live cells.
Transwell assay
Serum-free cell suspension was added to the upper chamber, while the lower chamber was supplemented with serum-containing medium. Following a 24-h incubation, cells adhering to the underside of the membrane were fixed and stained. The cells in the upper chamber were gently removed, and the migrated cells in the lower chamber were visualized under a microscope to evaluate migration capacity.
qRT-PCR
Total RNA was extracted using TRIzol reagent (Thermo Fisher Scientific) and subsequently reverse-transcribed with the PrimeScript RT Kit (Takara). Complementary DNA (cDNA) was amplified using TB Green Fast qPCR Mix (Takara). Relative gene expression levels were quantified using the 2−ΔCt method, with β-actin serving as the internal reference gene. Primer sequences are listed in Supplementary Table S2.
Transfection of siRNA and plasmids
The designated siRNA, plasmids, and Lipofectamine 2000 (Thermo Fisher Scientific) were combined in 1 mL of serum-free Opti-MEM and incubated for 20 min at room temperature. The transfection mixture was then added to cells that had been serum-starved for 12 h. After a 6-h transfection period, the culture medium was replaced with complete DMEM, and the cells were maintained under standard culture conditions for an additional 72 h.
Co-immunoprecipitation and Western blot
Cells were harvested and lysed using RIPA lysis buffer (#P0013, Beyotime, China). The lysates were then incubated with 30 µL of protein A/G beads (#P2029, Beyotime, China), along with either IgG (#A7007, Beyotime, China) or a specific primary antibody, for over 24 h at 4 °C. Following incubation, the beads were extensively washed with RIPA buffer. For Western blot analysis, protein lysates were denatured by boiling, separated via SDS-PAGE, and transferred onto a nitrocellulose membrane. Protein bands were detected using an Amersham Imager 600 (General Electric, USA).
Chromatin immunoprecipitation (ChIP) and ChIP-qPCR
The ChIP assay was performed following the protocols provided with Abcam’s chromatin extraction kit and magnetic one-step ChIP kit, briefly outlined below: Cells or tissues were fixed with 1% formaldehyde and then quenched with glycine. Nuclei were isolated via stepwise lysis using buffers, and chromatin was fragmented to 200–1000 bp by sonication (as verified by electrophoresis). The chromatin was then diluted, supplemented with protease inhibitors, and incubated overnight at 4 °C with either specific antibodies or IgG (as a negative control). Pre-blocked Protein A magnetic beads were added for capture at room temperature for 1 hour. After sequential washing with low-salt buffer, high-salt buffer, and a final rinse with TE buffer, complexes were eluted using lysis buffer containing 1% SDS. Incubation at 65 °C facilitated elution and subsequent NaCl-mediated reversal of cross-links (overnight, or accelerated via a 2-h incubation at 95 °C). Finally, DNA was purified following Proteinase K digestion; the analysis of this purified DNA was largely comparable to that of cDNA in RT-qPCR. Primer sequences for ChIP-qPCR are provided in Supplementary Table S3.
Glutathione S-transferase (GST) pull-down assay
Cells were lysed thoroughly using RIPA buffer. GST fusion proteins were immobilized on BeyoMag Anti-GST Magnetic Beads (P2138, Beyotime). After washing with RIPA buffer, the beads were incubated with cell lysates for 4 h at 4 °C. The beads were then washed again with the same buffer and resuspended in loading buffer. The proteins bound to the beads were analyzed via Western blotting64.
Acyl-biotinyl exchange assay
The cell pellets were lysed in buffer containing 50 mM Tris (pH 7.2), 150 mM NaCl, 1% NP-40, and 1 mM EDTA, followed by the addition of 20 mM methyl methanethiosulfonate (Sigma-Aldrich) and 1 mM PMSF (Beyotime). After incubation for 1 h at room temperature, excess reagents were removed by chloroform–methanol precipitation (one volume of chloroform, four volumes of methanol, and three volumes of ddH₂O). The pre-blocked samples were then treated with hydroxylamine (Sigma-Aldrich) at pH 7.0 to reach a final concentration of 0.5 M and incubated for 2 h at room temperature to cleave thioester bonds. Subsequently, 0.2 mM biotin-HPDP (Top Science) was added, and samples were incubated in the dark for 2 h to ensure efficient biotinylation. Excess biotin was again removed by chloroform–methanol precipitation. The resulting proteins were purified with Streptavidin beads (Yeasen) and analyzed by SDS-PAGE and Western blotting.
Click-iT pull down
Cells were transfected with plasmids for 48 h, and then labeled with 100 μmol/L palmitic acid-azide (Thermo Fisher Scientific) for 6 h. After that, the cells were lysed to extract proteins. The Click-iT kit (Thermo Fisher Scientific) was used to catalyze the reaction between biotin-alkyne and the azide group, forming a biotinylated palmitoylated protein complex. The complex was captured by streptavidin magnetic beads. After washing with gradient salt buffers, the proteins were eluted with SDS-PAGE buffer. Finally, the palmitoylation modification was analyzed by Western blotting65.
Semi-in vitro palmitoylation assay
Cells were transfected with GST-ZDHHC13 (WT) or the C456S mutant. After 24 h, cells were lysed with NP-40 lysis buffer, and GST fusion proteins were purified with glutathione agarose beads. The semi-purified GST-ZDHHC13 was incubated with purified HIS-FBP1 or HIS-FBP1(C282S) in the presence of palmitoyl-alkyne-CoA at 25 °C for 1 h (with or without 2-bromopalmitate as a control). Reaction products were then subjected to a Click-iT reaction, followed by streptavidin pull-down and Western blot analysis of FBP1 palmitoylation.
CRISPR technique
Multiple single-guide RNAs (sgRNAs) targeting Zdhhcs and Fbp1 were designed using the platform available at https://www.synthego.com. Subsequently, these sgRNAs were cloned into the lentiCRISPR v2 vector (Addgene)64. The sequences of the sgRNAs are presented in Supplementary Table S1.
Nuclear-cytoplasmic protein separation
Cytoplasmic and nuclear proteins were extracted following the manufacturer’s protocol (#P0028, Beyotime). Briefly, Cytoplasmic Protein Extraction Reagent A was mixed with a protease inhibitor at a 100:1 ratio and added to the cell pellet. The mixture was vortexed for 5 s and incubated on ice for 10–15 min. Subsequently, Reagent B was added, and the mixture was vortexed again for 5 s. The sample was centrifuged at 12,000×g for 5 min at 4 °C, and the supernatant containing cytoplasmic proteins was collected. The remaining pellet was resuspended in Nuclear Protein Extraction Reagent mixed with a protease inhibitor at a 100:1 ratio. The suspension was vortexed for 10 s and incubated on ice with intermittent vortexing (10–15 s every 2 min) for 30 min. Finally, the sample was centrifuged at 12,000×g for 5 min at 4 °C, and the supernatant containing nuclear proteins was collected.
Glucose consumption and lactate production experiment
Cells in the logarithmic growth phase were seeded into 96-well plates and cultured in complete medium supplemented with glucose for 24 h. After washing with basal medium, the medium was replaced with one containing a known concentration of glucose. Following an additional 48-h incubation, the supernatant was collected, and glucose and lactate levels were quantified using specific detection kits. Glucose consumption and lactate production were calculated based on the concentration differences before and after incubation and the total medium volume.
Seahorse extracellular flux assays
The Seahorse XFe 96 Extracellular Flux Analyzer (Agilent Technologies) was used to measure the oxygen consumption rate (OCR) and extracellular acidification rate (ECAR). For the OCR analysis, the Seahorse XF Cell Mito Stress Test Kit (Agilent) was used. According to the manufacturer’s protocol, oligomycin, p-trifluoromethoxy carbonyl cyanide phenylhydrazone (FCCP), and a mix of rotenone/antimycin A were injected sequentially into the microplate wells. For the ECAR analysis, the Seahorse XF Glycolysis Stress Test Kit was employed, and glucose, oligomycin, and 2-deoxy-D-glucose (2-DG) were sequentially injected into the wells.
Identification of gluconeogenic function in CCSMCs
Primary CCSMCs were individually seeded into three 75-cm² culture flasks and maintained in a humidified incubator with 5% CO₂ at 37 °C until confluence was achieved. Subsequently, the culture medium was aspirated and replaced with serum-free medium for a 24-h starvation period, followed by the addition of 13C-labeled sodium lactate to a final concentration of 2 mM to continue cultivation. Upon completion of incubation, the culture supernatants were collected, lyophilized using a vacuum centrifugal concentrator, and resuspended in 2 mL of deuterium oxide. ¹³C nuclear magnetic resonance (NMR) spectral acquisition and data analysis were performed on a Bruker AVANCE NEO 600 M NMR spectrometer, following protocols described in previously published relevant literature16,17. The obtained spectra were subjected to phase and baseline correction using TopSpin software, and qualitative analysis was performed based on the typical ¹³C chemical shift range of glucose.
PROTAC
The CBX3-targeting ligand was designed based on the H3K9me3-derived QTAR M3L peptide motif ((Q/N)(T/V)A(R/I/F/W/V)Kme3(S/T)). Meanwhile, the immunomodulatory drug Pomalidomide was incorporated on the opposite terminus as the E3 ubiquitin ligase recruiter, specifically engaging the CRL4^(CRBN) complex (Cereblon E3 ubiquitin ligase), which is widely employed in PROTAC design for its potent protein degradation properties. A critical aspect of the design involved the synthesis and optimization of a bridging linker that covalently connects these two ligands; its length, composition, and rigidity were meticulously engineered based on structural insights to ensure proper spatial alignment and facilitate efficient formation of a productive ternary complex, thereby enabling targeted ubiquitination and subsequent proteasomal degradation of CBX3. The degradation efficiency and specificity of the synthesized CBX3-PROTAC were validated by Western blotting to quantify the reduction in CBX3 protein levels35.
LNP
A cysteine-to-serine substitution at position 282 of the mouse Fbp1 gene was introduced by site-directed mutagenesis. The mutated Fbp1 gene was amplified by standard PCR, which included initial denaturation, followed by cycles of denaturation, annealing, and extension steps. The PCR product was excised from an agarose gel and purified using a commercial DNA gel extraction kit. The purified DNA was then double-digested, along with a T7 promoter-driven vector, using appropriate restriction enzymes. After recovery, the digested fragments were ligated using T4 DNA ligase. The ligation product was transformed into competent E. coli cells, and positive colonies were selected on antibiotic-containing plates. Single colonies were cultured, and the plasmid DNA was extracted and verified by Sanger sequencing. The confirmed recombinant plasmid was linearized and used as a template for in vitro transcription. The resulting mRNA was purified using an mRNA purification kit.
For LNP formulation, the aqueous phase was prepared by diluting mRNA to a concentration of 0.35 mg/mL in 100 mM sodium acetate buffer (pH 4.0). The organic phase was prepared by dissolving the lipid mixture in ethanol to a final concentration of 25 mM. The lipid composition was as follows: (6Z,9Z,28Z,31Z)-heptatriaconta- 6,9,28,31-tetraen-19-yl 4-(dimethylamino)butanoate: distearoylphosphatidylvcholine: cholesterol:[1,2-dimyristoyl-sn-glycero-3-phosphoethanolamine]-N-[methoxy(polyethylene glycol)−2000] (50:10:38.5:1.5, mol%). LNPs were assembled using an Ignite microfluidic chip (Precision Nanosystems) with a PNI Ignite instrument at a flow rate ratio of 3:1 (aqueous to organic phase) and a total flow rate of 12 mL/min.
Animal
All experimental procedures were approved by the Animal Use and Care Committee of the Second Xiangya Hospital, Central South University (Approval No. 20250042) and were conducted in accordance with the ARRIVE Guidelines and the National Institutes of Health Guide for the Care and Use of Laboratory Animals. The following mice were used in this study: C57BL/6, C57BLKS/J-leprdb/leprdb (db/db), Fbp1−/+ (heterozygous Fbp1 knockout, C57BL/6JGpt background, GemPharmatech Co.,Ltd; derived from C57BL/6JGpt-Fbp1em1Gpt). The animals were generated, housed, and bred under a 12 h light/dark cycle at a temperature of 22 ± 2 °C and 55 ± 5% relative humidity, with free access to food and water throughout the study. The standard chow diet used was Catalog No.YNDWSL01-TJ, and the high-fat diet for DMED model induction was Catalog No.D12492 (Medison).
A mouse model of DMED was induced by a high-fat diet combined with streptozotocin administration66. Male C57BL/6 mice were acclimatized for 2 weeks prior to random assignment into control or experimental groups. All groups were rigorously age-matched to minimize confounding effects. Mice in the experimental group received a high-fat diet for four weeks, followed by intraperitoneal injections of streptozotocin (50 mg/kg/day) for 5 consecutive days. Diabetes was confirmed when blood glucose levels exceeded 16.7 mmol/L on three consecutive measurements. Diabetic mice were maintained on a high-fat diet for an additional 8 weeks to promote the development of ED. Apomorphine was dissolved in 0.9% sodium chloride solution containing 0.2 mg/mL ascorbic acid and administered subcutaneously at 100 μg/kg. Erectile responses were recorded within 30 min post-injection. Mice that did not exhibit penile erection were classified as having DMED. Of the 220 C57BL/6 mice initially enrolled, 24 died due to severe hyperglycemia or procedural complications, yielding 196 surviving diabetic mice. Among these, 181 met the DMED criteria (absence of erectile response to apomorphine) and were randomly assigned to treatment subgroups: control (normal saline, i.p.), 3-MA (10 mg/kg/day, p.o.), SMER28 (50 μg/kg/day, i.p.), Chaetocin (0.2 mg/kg every three days, i.p.), EPZ6438 (250 mg/kg/day, p.o.), H-PROTAC (5 mg/kg/week, intracavernosal), sodium oxamate (0.5 g/kg/day, i.p.), AAV9 empty vector, AAV9-Zdhhhc13KD (3 × 10¹³ vg/mL, intracavernosal), AAV9-Sox9KD (3 × 10¹³ vg/mL, intracavernosal), LNP control, and Fbp1 C282S-LNP (1 mg/mL/week, intracavernosal). Four weeks post-treatment, we evaluated cavernosal lactate levels, ICP, and histopathological changes.
A type 1 diabetes model was induced as previously described67. Eight-week-old male C57BL/6 mice were fasted for 12 h and then given a single intraperitoneal injection of streptozotocin (150 mg/kg). Diabetes was confirmed 72 h post-injection by fasting blood glucose >16.7 mmol/L. Mice were maintained for eight additional weeks before erectile function was assessed via the apomorphine test.
Additionally, 20-week-old male db/db mice (C57BLKS/J-leprdb/leprdb) were obtained from Shulaibao Biotechnology (Wuhan, China). Fbp1−/+ mice were generated by Shulaibao Biotechnology (Wuhan, China). For lactate exposure experiments, 8-week-old male C57BL/6 mice were randomly divided into three groups receiving intraperitoneal injections of normal saline, lactate (0.25 g/kg), or lactate (0.5 g/kg) for 8 weeks. Control mice were strictly age-matched and underwent the same standardized procedures.
ICP
The ICP was measured using the MP160 system and BL-420F Functional Biological Experiment System (China). After anesthesia induction with isoflurane, the penile crura and cavernous nerves were bilaterally exposed. A 24-gauge needle was inserted into one crus, and the ipsilateral cavernous nerve was stimulated using a bipolar hook-shaped electrode. ICP recordings were obtained via a pressure transducer connected to the needle63.
Hematoxylin-Eosin (H&E) and Masson staining
For H&E staining, dewaxed and rehydrated tissue sections were stained with hematoxylin for 5 min, differentiated in 1% hydrochloric acid-ethanol for 1 min, blued in ammonia water for 1 min, and counterstained with eosin for 1 min. The stained samples were then dehydrated, mounted, and examined under a microscope. For Masson staining, sections were sequentially treated with hematoxylin for 5 minutes, Ponceau acid-fuchsin for 6 min, 1% phosphomolybdic acid for 1 min, and 2.5% aniline blue for 2 min, followed by rinsing and differentiation with 1% glacial acetic acid. Morphometric analysis was conducted using ImageJ software on three randomly selected non-overlapping fields from each section. In H&E-stained sections, well-defined structural regions were imported into ImageJ, and the threshold function was applied with manual adjustment to distinguish luminal spaces from parenchymal areas based on grayscale intensity. In Masson-stained sections, red-stained areas were identified as muscle fibers, while dark blue regions indicated collagen fibers; both were quantified sequentially using threshold-based segmentation. All analyses were performed by two independent investigators blinded to group assignments, and results were cross-validated with mean values used for statistical analysis.
Immunofluorescence (IF) and immunohistochemistry (IHC)
For IF, samples were permeabilized with Triton X-100, blocked with goat serum, and incubated with primary antibodies at 4 °C overnight, followed by incubation with fluorophore-conjugated secondary antibodies and DAPI at room temperature. For IHC, paraffin-embedded sections were dewaxed in xylene, rehydrated in a graded ethanol series, and subjected to heat-induced antigen retrieval. After blocking with 5% goat serum, sections were incubated with primary antibodies at 4 °C overnight, followed by incubation with biotinylated secondary antibodies, ABC reagent, and DAB chromogenic detection. All stained sections were visualized using a fluorescence or bright-field microscope, respectively. For quantitative analysis, three non-overlapping fields per sample were randomly selected and examined using ImageJ software by two independent investigators blinded to the experimental conditions.
Pharmacokinetic studies and tissue distribution
The intracavernosal injections of the test agent (0.25 mg/kg) were administered to C57BL/6 mice, with blood samples gathered at 0.25, 0.5, 1, 2, 4, 8, 12, 24, and 48 h post-injection. These blood samples were then centrifuged at 11,000×g for 10 min at 4 °C. A 50 µL portion of serum was combined with 150 µL methanol, followed by 10 min of sonication on ice. The resulting supernatant underwent high-performance liquid chromatography analysis, with pharmacokinetic parameters derived from these analytical results. For characterizing the tissue distribution of LNPs, healthy C57BL/6 mice received intracavernosal injections of DiD-labeled LNPs that encapsulated Fbp1 C282S mRNA. At 0.25, 1, 4, and 12 h after injection, the mice were euthanized, and tissues, including the corpus cavernosum, liver, skeletal muscle, spleen, and brain, were collected. Fluorescence intensity in the supernatants of these tissues was measured using a multimode microplate reader68.
RNA sequencing and bioinformatic analysis
Total RNA was isolated from primary mouse CCSMCs under control and high-lactate conditions (10 mM) using TRIzol Reagent (Thermo Fisher Scientific). RNA concentration and purity were measured on a Nanodrop spectrophotometer, with A260/A280 ratios between 1.8 and 2.0 deemed acceptable. RNA integrity was assessed using an Agilent Bioanalyzer 2100 system, and samples with an RNA integrity number (RIN) ≥7.0 were subjected to sequencing. Sequencing was performed by Shanghai Jiayin Biotechnology Co., Ltd. on the Illumina NovaSeq platform under paired-end 150 bp mode. After quality control (including adapter trimming and low-quality read filtering via FastQC), differential gene expression analysis was conducted using DESeq2 in RStudio (v1.4). KEGG pathway enrichment analysis was performed using clusterProfiler, and results were visualized using bar charts.
Statistical analysis
All experimental data were analyzed using GraphPad Prism 7.0 software. Comparisons between two groups were conducted using an unpaired two-sided Student’s t-test. For comparisons among multiple groups, one-way analysis of variance (ANOVA) was performed, followed by Tukey’s post hoc test. A p value <0.05 was considered statistically significant.
Code availbility
Not applicable (no custom code).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA sequencing data from rats, mice and cells reported in this paper have been deposited in the genome sequence archive (Genomics, Proteomics & Bioinformatics 2021) in National Genomics Data Center (Nuclic Acids Res. 2022), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA: CRA029537 and CRA029518) that are pukicly accessible at https://ngdc.cncb.ac.cn/gsa. All other relevant data supporting the key findings of this study are available within the article and its Supplementary Information files. Source data are provided with this paper.
References
Shamloul, R. & Ghanem, H. Erectile dysfunction. Lancet 381, 153–165 (2013).
Sun, H. et al. IDF diabetes atlas: global, regional and country-level diabetes prevalence estimates for 2021 and projections for 2045. Diabetes Res. Clin. Pract. 183, 109119 (2022).
Shiferaw, W. S., Akalu, T. Y., Petrucka, P. M., Areri, H. A. & Aynalem, Y. A. Risk factors of erectile dysfunction among diabetes patients in Africa: a systematic review and meta-analysis. J. Clin. Transl. Endocrinol. 21, 100232 (2020).
Benet, A. E. & Melman, A. The epidemiology of erectile dysfunction. Urol. Clin. North Am. 22, 699–709 (1995).
Penson, D. F. et al. Do impotent men with diabetes have more severe erectile dysfunction and worse quality of life than the general population of impotent patients? Results from the exploratory comprehensive evaluation of erectile dysfunction (ExCEED) database. Diabetes Care 26, 1093–1099 (2003).
de Souza, I. L. L., Ferreira, E. D. S., Vasconcelos, L. H. C., Cavalcante, F. A. & da Silva, B. A. Erectile dysfunction: key role of cavernous smooth muscle cells. Front. Pharmacol. 13, 895044 (2022).
Xu, W. et al. Ferroptosis is involved in corpus cavernosum smooth muscle cells impairment in diabetes mellitus-induced erectile dysfunction. Andrology 11, 332–343 (2023).
He, S. et al. Myocardin restores erectile function in diabetic rats: phenotypic modulation of corpus cavernosum smooth muscle cells. Andrologia 47, 303–309 (2015).
Li, X. et al. Lactate metabolism in human health and disease. Signal Transduct. Target Ther. 7, 305 (2022).
Chen, Y. D., Varasteh, B. B. & Reaven, G. M. Plasma lactate concentration in obesity and type 2 diabetes. Diabete Metab. 19, 348–354 (1993).
Darshi, M. et al. Glycolytic lactate in diabetic kidney disease. JCI Insight 9, e168825 (2024).
Kim, T. W., Park, S. S., Kim, B. K., Sim, Y. J. & Shin, M. S. Effects of sildenafil citrate on peripheral fatigue and exercise performance after exhaustive swimming exercise in rats. J. Exerc. Rehabil. 15, 751–756 (2019).
Li, C. H., Chan, M. H. & Chang, Y. C. The role of fructose 1,6-bisphosphate-mediated glycolysis/gluconeogenesis genes in cancer prognosis. Aging 14, 3233–3258 (2022).
Li, F. et al. FBP1 loss disrupts liver metabolism and promotes tumorigenesis through a hepatic stellate cell senescence secretome. Nat. Cell Biol. 22, 728–739 (2020).
Fang, H. et al. Gut substrate trap of D-lactate from microbiota improves blood glucose and fatty liver disease in obese mice. Cell Metab. S1550-4131, 00328-6 (2025).
Lloyd, P. G. & Hardin, C. D. Role of microtubules in the regulation of metabolism in isolated cerebral microvessels. Am. J. Physiol. 277, C1250–C1262 (1999).
Gizak, A., Rakus, D. & Dzugaj, A. Nuclear localization of fructose 1,6-bisphosphatase in smooth muscle cells. J. Mol. Histol. 36, 243–248 (2005).
Xiao, M. et al. The conserved molecular mechanism of erectile dysfunction in type 2 diabetes rats and mice by cross-species transcriptomic comparisons. Sex. Med. 13, qfaf007 (2025).
Zhao, L. et al. An interactive website for scRNA-seq data of male genitourinary development and disease. Andrology 11, 1157–1162 (2023).
Hunter, R. W. et al. Metformin reduces liver glucose production by inhibition of fructose-1,6-bisphosphatase. Nat. Med. 24, 1395–1406 (2018).
Gorce, M. et al. Fructose-1,6-bisphosphatase deficiency causes fatty liver disease and requires long-term hepatic follow-up. J. Inherit. Metab. Dis. 45, 215–222 (2022).
Jin, X. et al. Fructose-1,6-bisphosphatase inhibits ERK activation and bypasses gemcitabine resistance in pancreatic cancer by blocking IQGAP1-MAPK interaction. Cancer Res. 77, 4328–4341 (2017).
Åsberg, C. et al. Fructose 1,6-bisphosphatase deficiency: enzyme and mutation analysis performed on calcitriol-stimulated monocytes with a note on long-term prognosis. J. Inherit. Metab. Dis. 33, S113–S121 (2010).
Lu, J., Huang, Y., Zhang, X., Xu, Y. & Nie, S. Noncoding RNAs involved in DNA methylation and histone methylation, and acetylation in diabetic vascular complications. Pharmacol. Res. 170, 105520 (2021).
Mukherjee, A., Debbarman, T., Banerjee, B. D. & Siddiqi, S. S. The impact of epigenetics on the pathophysiology of type 2 diabetes and associated nephropathic complications. Indian J. Endocrinol. Metab. 28, 569–578 (2024).
Figueroa-Angulo, E. E., Puente-Rivera, J., Perez-Navarro, Y. F., Condado, E. M. & Álvarez-Sánchez, M. E. Epigenetic alteration in cervical cancer induced by human papillomavirus. Int. Rev. Cell Mol. Biol. 390, 25–66 (2025).
Angelico, G. et al. H3K27me3 loss in central nervous system tumors: diagnostic, prognostic, and therapeutic implications. Cancers 16, 3451 (2024).
Becker, J. S., Nicetto, D. & Zaret, K. S. H3K9me3-dependent heterochromatin: barrier to cell fate changes. Trends Genet. 32, 29–41 (2016).
Kong, M. et al. Histone methyltransferase Suv39h1 regulates hepatic stellate cell activation and is targetable in liver fibrosis. Gut 73, 810–824 (2024).
Taheri, N. et al. Inhibition of EZH2 reduces aging-related decline in interstitial cells of Cajal of the mouse stomach. Cell Mol. Gastroenterol. Hepatol. 18, 101376 (2024).
Bekric, D. et al. Evaluation of tazemetostat as a therapeutically relevant substance in biliary tract cancer. Cancers 15, 1569 (2023).
Xiong, X. et al. FBP1 knockdown decreases ovarian cancer formation and cisplatin resistance through EZH2-mediated H3K27me3. Biosci. Rep. 42, BSR20221002 (2022).
Cai, D. et al. Betaine supplementation in maternal diet modulates the epigenetic regulation of hepatic gluconeogenic genes in neonatal piglets. PLoS ONE 9, e105504 (2014).
Duan, Y. et al. Integrated lactylome characterization reveals the molecular dynamics of protein regulation in gastrointestinal cancers. Adv. Sci. 11, e2400227 (2024).
Békés, M., Langley, D. R. & Crews, C. M. PROTAC targeted protein degraders: the past is prologue. Nat. Rev. Drug Discov. 21, 181–200 (2022).
Weinert, B. T., Moustafa, T., Iesmantavicius, V., Zechner, R. & Choudhary, C. Analysis of acetylation stoichiometry suggests that SIRT3 repairs nonenzymatic acetylation lesions. EMBO J. 34, 2620–2632 (2015).
Bian, Y. et al. An enzyme assisted RP-RPLC approach for in-depth analysis of human liver phosphoproteome. J. Proteomics 96, 253–262 (2014).
Chen, S. et al. Palmitoylation-dependent activation of MC1R prevents melanomagenesis. Nature 549, 399–403 (2017).
Wang, Z. et al. AMPKα1-mediated ZDHHC8 phosphorylation promotes the palmitoylation of SLC7A11 to facilitate ferroptosis resistance in glioblastoma. Cancer Lett. 584, 216619 (2024).
Yang, A. et al. Regulation of RAS palmitoyltransferases by accessory proteins and palmitoylation. Nat. Struct. Mol. Biol. 31, 436–446 (2024).
Al-Zahrani, K. N. et al. AKT-mediated phosphorylation of Sox9 induces Sox10 transcription in a murine model of HER2-positive breast cancer. Breast Cancer Res. 23, 55 (2021).
Cheng, C. C. et al. PI3K/AKT regulates aggrecan gene expression by modulating Sox9 expression and activity in nucleus pulposus cells of the intervertebral disc. J. Cell Physiol. 221, 668–676 (2009).
Zheng, T. et al. Lactate-triggered histone lactylation contributes to podocyte epithelial-mesenchymal transition in diabetic nephropathy in mice. Chem. Biol. Interact. 408, 111418 (2025).
Mutlu, B. et al. Small molecules targeting selective PCK1 and PGC-1α lysine acetylation cause anti-diabetic action through increased lactate oxidation. Cell Chem. Biol. 31, 1772–1786.e5 (2024).
Özel, A. et al. Deciphering the role of lactate as a prognostic indicator in pediatric diabetic ketoacidosis. Wien Klin. Wochenschr. 137, 98–104 (2025).
Chen, J. et al. PFKFB3-driven vascular smooth muscle cell glycolysis promotes vascular calcification via the altered FoxO3 and lactate production. FASEB J. 37, e23182 (2023).
Hu, Y. et al. Lactate promotes vascular smooth muscle cell switch to a synthetic phenotype by inhibiting miR-23b expression. Korean J. Physiol. Pharmacol. 26, 519–530 (2022).
Zhang, P. et al. FBP1 orchestrates keratinocyte proliferation/differentiation and suppresses psoriasis through metabolic control of histone acetylation. Cell Death Dis. 15, 392 (2024).
Gu, L. et al. FBP1 controls liver cancer evolution from senescent MASH hepatocytes. Nature 637, 461–469 (2025).
Qu, P. et al. Histone methylation modification and diabetic kidney disease: potential molecular mechanisms and therapeutic approaches (review). Int. J. Mol. Med. 54, 104 (2024).
Oleksiewicz, U. et al. The roles of H3K9me3 writers, readers, and erasers in cancer immunotherapy. Int. J. Mol. Sci. 25, 11466 (2024).
Lin, Z. et al. Palmitoyl acyltransferase ZDHHC7 inhibits androgen receptor and suppresses prostate cancer. Oncogene 42, 2126–2138 (2023).
Huang, K. et al. Wild-type HTT modulates the enzymatic activity of the neuronal palmitoyl transferase HIP14. Hum. Mol. Genet. 20, 3356–3365 (2011).
Zhang, Y., Fan, S., He, L. & Li, L. The ZDHHC13/ZDHHC17 subfamily: from biological functions to therapeutic targets of diseases. Pharmacol. Res. 209, 107418 (2024).
Yang, J. et al. Submicromolar concentrations of palmitoyl-CoA specifically thioesterify cysteine 244 in glyceraldehyde-3-phosphate dehydrogenase inhibiting enzyme activity: a novel mechanism potentially underlying fatty acid induced insulin resistance. Biochemistry 44, 11903–11912 (2005).
Qin, H. et al. Mechanistic insights into the inhibitory effects of palmitoylation on cytosolic thioredoxin reductase and thioredoxin. Biochimie 110, 25–35 (2015).
Kiaie, S. H. et al. Recent advances in mRNA-LNP therapeutics: immunological and pharmacological aspects. J. Nanobiotechnol. 20, 276 (2022).
Zong, Y., Lin, Y., Wei, T. & Cheng, Q. Lipid nanoparticle (LNP) enables mRNA delivery for cancer therapy. Adv. Mater. 35, e2303261 (2023).
Salonia, A. et al. European association of urology guidelines on sexual and reproductive health-2021 update: male sexual dysfunction. Eur. Urol. 80, 333–357 (2021).
Campbell, J. D., Milenkovic, U., Usta, M. F., Albersen, M. & Bivalacqua, T. J. The good, bad, and the ugly of regenerative therapies for erectile dysfunction. Transl. Androl. Urol. 9, S252–S261 (2020).
Castela, Â & Costa, C. Molecular mechanisms associated with diabetic endothelial-erectile dysfunction. Nat. Rev. Urol. 13, 266–274 (2016).
Ji, M. et al. The role of mechano-regulated YAP/TAZ in erectile dysfunction. Nat. Commun. 14, 3758 (2023).
Xiong, Y. et al. Targeting adenosine A2b receptor promotes penile rehabilitation of refractory erectile dysfunction. Adv. Sci. 11, e2306514 (2024).
Sun, Y. et al. ZDHHC2-mediated AGK palmitoylation activates AKT-mTOR signaling to reduce sunitinib sensitivity in renal cell carcinoma. Cancer Res. 83, 2034–2051 (2023).
Zhang, Z. et al. DHHC9-mediated GLUT1 S-palmitoylation promotes glioblastoma glycolysis and tumorigenesis. Nat. Commun. 12, 5872 (2021).
Yeo, H. J. et al. Tat-CIAPIN1 prevents pancreatic β-cell death in hIAPP-induced RINm5F cells and T2DM animal model. Int. J. Mol. Sci. 24, 10478 (2023).
Zhang, X. et al. Animal models in the study of diabetic erectile dysfunction: mechanisms and applications. Front. Endocrinol. 16, 1512360 (2025).
Bian, X. et al. Berberine-inspired ionizable lipid for self-structure stabilization and brain targeting delivery of nucleic acid therapeutics. Nat. Commun. 16, 2368 (2025).
Acknowledgements
We acknowledge all authors participating in this study for their work and helpful comments. This work was supported by Central South University Innovation-Driven Research Program (Grant No.2023CXQD058 (X.J.), Key scientific research project of Hunan Provincial Health Commission (Grant No. W20242004 (X.J.), the Fundamental Research Funds for the Central Universities of Central South University (Grant No. 2024ZZTS0960 (W.G.) and Grant No. 2025ZZTS0966 (R.Z.).
Author information
Authors and Affiliations
Contributions
Methodology: M.X. and W.G. Software: R.Z., M.X., and W.G. Formal analysis: R.Z. and W.G. Project administration: S.W. and X.J. Writing—original draft: M.X. and W.G. Writing—review and editing: M.X., S.W., and X.J.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiao, M., Guo, W., Zeng, R. et al. Improving erectile function in diabetic male mice by rescuing depalmitoylated FBP1 to reduce cavernosal lactate. Nat Commun 17, 1740 (2026). https://doi.org/10.1038/s41467-026-68443-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68443-y