Abstract
Chromatin conformation is thought to be critical for enhancer function, but its dynamic, nanoscale organization is difficult to measure directly. Here we introduce PLOTTED (Probabilistic Localization of Oligopaint Tagged Target Element Distances), an integrated imaging and computational framework that infers chromatin architecture from targeted high-resolution imaging of cis-regulatory modules (CRMs). PLOTTED generates spatial distance distributions between DNA loci, enabling quantitative modeling of chromatin configurations across developmental time, spatial axes, and genotypes. Applying PLOTTED to the brinker locus in Drosophila embryos, we measured distances among three CRMs and used chromatin geometry as a proxy for regulatory activity. In wild type, CRM configurations shift dynamically at nuclear cycle 13, whereas these changes are delayed in mutants and vary along the dorsal-ventral and anterior–posterior axes. Importantly, these conformational changes correlate with altered gene expression. Together, our findings position PLOTTED as a probabilistic, single-locus framework for interpreting chromatin architecture in development and disease.
Similar content being viewed by others
Introduction
Regulation of gene expression is essential during the maternal to zygotic transition (MZT) of developing embryos and is linked to changes in global chromatin conformation. Organization of the Drosophila genome is initially disordered, but after the major wave of zygotic genome activation (ZGA), global chromatin conformation becomes more organized as topologically associating domains (TADs) increase in prevalence1. These TADs form the foundation of nuclear architecture, partitioning chromosomes into neighborhoods of ~100 kb in Drosophila2,3. Several methods have been devised to study the DNA interactions involved in TAD formation, including targeted approaches such as 3C4,5 and global approaches such as Hi-C5,6 or Micro-C used in recent Drosophila studies7,8,9,10. However, these techniques are only able to capture pairwise interactions and are limited in their ability to assay multiway interactions. In addition, there are likely chromatin associations, or DNA-DNA contacts11, co-occurring within these large-scale TADs that are under the resolution limit for these protocols. For instance, regulatory elements located within the same TAD can be shared between genes in order to coordinate transcriptional dynamics in response to a given stimulus2. There is limited understanding of these smaller-scale chromatin associations, including their dynamics and whether they play a role in regulating gene expression. Local chromatin conformational changes may regulate enhancer-promoter interactions or act in other ways to affect gene expression at individual loci. Therefore, developing approaches to analyze local chromatin associations has the potential to provide novel insights into cis-regulatory control.
Analysis of local chromatin conformation at different time points in intact tissues, and their relationship to gene expression, can be assayed using high-resolution microscopy to visualize interactions between DNA sequences. For instance, Hi-M uses widefield microscopy to analyze interactions between DNA sequences separated by distances of 3–10 kb or greater12,13. Conversely, the Optical Reconstruction of Chromatin Architecture (ORCA) technique14,15 allows visualization of nanoscale, multiway DNA-DNA interactions with sufficient resolution to distinguish individual loci separated by as little as 2 kb. These DNA labeling and microscopy methods have advanced the study of chromatin conformation, but are technically challenging. They require specialized robotics, microfluidic devices, and chambers, as well as advanced post-processing pipelines. The current study addresses these limitations by visualizing and assaying local DNA interactions associated with a single gene locus using a widely accessible DNA FISH protocol, an imaging approach that is compatible with protein staining, and a superresolution confocal microscope. Also described is a conceptual framework for looking at how three DNA regulatory elements interact in probabilistic space to regulate gene expression.
In this study, we developed an imaging approach named Probabilistic Localization of Oligopaint Tagged Target Element Distances (PLOTTED) to label, visualize, and compare local chromatin interactions of specific DNA loci, in this case previously characterized cis-regulatory modules (CRMs). Our method uses Oligopaint DNA FISH to label candidate CRMs for interaction measurements. Furthermore, imaging was performed using a confocal microscope capable of superresolution analysis. The resulting 3D distance data were processed through a computational pipeline for thresholding and classification into pairwise or triplet interaction groups. We applied this approach to the brinker (brk) gene locus, which is regulated by three distinct CRMs during early Drosophila development in the blastoderm embryo16. The brk locus, including all of its known regulatory elements, are contained within a single TAD17. The 3D-distances between the three brk-associated CRMs, each separated by approximately 10 kB on the linear DNA sequence, were quantified across tens of thousands of nuclei from hundreds of fixed embryos. This enabled us to infer chromatin conformation dynamics at a single gene locus across space, time, and multiple mutant backgrounds in early Drosophila embryos, but this is a broadly applicable framework for studying 3-way interactions in any system containing nuclei.
Results
PLOTTED measures distances between cis-regulatory modules
We used PLOTTED to assay chromatin conformation at the brk locus during both the syncytial blastoderm phase, when cell membranes are not present between nuclei, and at cellularization associated with the midblastula transition of Drosophila embryogenesis. During this time, embryos undergo 13 nuclear divisions, called nuclear cycles (nc), each with a short S phase and no G2 phase, until nc14 when the cell cycle increases in length and cellularization occurs18,19. Two enhancers have been shown to activate brk at nc14: enhancer E1 is ~10 kb upstream of the promoter (previously called brk5’), and enhancer E2 is ~14 kb downstream of the promoter (previously called brk3’)20,21,22. Previous studies have suggested that chromatin dynamics mediated by a promoter proximal element (PPE) play a role in regulating the action of CRMs to influence brk expression16. To test the role of DNA conformation in this process, we applied the PLOTTED imaging and quantitative analysis pipeline to assay these brk CRMs in a fixed embryo time series.
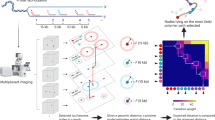
The labeling approach used to visualize DNA at the brk locus was adapted from a previously published Oligopaint protocol designed for Hi-M12, but simplified in order to detect just three specific positions along a stretch of ~25 kB encompassing the brk locus: E1, PPE, and E2 (Fig. 1A). Oligopaint DNA probes were designed to encompass 4 kb DNA regions including each of these CRMs as well as surrounding DNA (Fig. 1A). Each probe set consisted of 50 individual probes, 40-50 bp in length spanning each of the 4 kB regions, and labeled with distinct fluorophores: E1 (Alexa 550; pseudo-colored blue), PPE (Alexa 647; pseudo-colored green), and E2 (Alexa 488; pseudo-colored red) (see Methods). For each embryo imaged, two sample regions were acquired using a 63x objective: one towards the anterior and the other towards the posterior (Fig. 1B). In order to assess if the distance between the labeled loci changed over time, fixed embryos were examined at four stages: pre-nc13 (a mix of nc11 and nc12), nc13, nc14-early, and nc14-late (Fig. 1C).
A Annotated brk locus showing three 4 kb regions labeled by Oligopaint-DNA FISH probes with reference to position of CRMs: E1, PPE, and E2. B Embryo stained by DNA FISH using probes to E1 (blue), PPE (green), and E2 (red), imaged twice per embryo at anterior (A) and posterior (P) trunk regions. Nuclear membranes are labeled using Lamin antibody (white). Example 63x image is shown with zoomed images of triplet raw data within individual nuclei (left) or after spot detection with pseudo-spot labels (right). C Representative images of staging of Drosophila embryos from pre-cellularization (pre-nc13 and nc13) towards cellularization (nc14 early and nc14 late) based on nuclear density49. D Pairwise distances between CRMs [E1 (blue), PPE (green), and E2 (red)] were measured within nuclei of fixed embryos. Thresholds applied to each doublet or triplet include a maximum distance threshold (MDT, blue circle) and a resolution distance threshold (RLT, brown ellipsoid). Spots within RLT are too close to resolve, giving a distance of 0 µm, whereas those outside this limit but under MDT are measured. An example scenario is shown, two spots within the RLT (green and blue) would have a recorded distance E1-PPE as 0 µm. On the other hand, blue and red will have real distance for E1-E2 measured.E Representative images showing detected spots as pseudo-colored squares which are projected onto the original image, allowing for manual validation of triplets and maximum distance verification to ensure that all spots within a triplet are contained in the same nucleus (see Methods). The triplet outside the nucleus is only representative. See also Fig. S1.
A computational approach was used to process the images in order to extract distances and apply thresholding (Fig. 1 and S1). Once staged, spot detection was performed for each acquisition channel (550, 647, and 488) of each image using two thresholds, low and high, based on the spots’ shape and intensity (Fig. S1A, A’). The choice of detection thresholds (i.e., low vs high) was empirically defined to maximize spot detection while minimizing false positives (Fig. S1B, C; see Methods). Altogether, the combination of dual-thresholding and subsequent manual inspection enabled the detection of the maximum number of spots from each image. When only two fluorescent spots were visible in a nucleus, we measured a single pairwise interaction, referred to as a “doublet”; whereas when all three fluorescent spots were detected, we measured all three pairwise interactions, referred to as a “triplet”.
In addition to validating spot detection, doublet and triplet detection per single nucleus was further confirmed by applying a maximum distance threshold (MDT; Fig. 1D, blue sphere). The maximum possible distance between two spots is the diameter of the nucleus. However, naively using this distance to threshold could potentially also group spots together located in neighboring nuclei, especially at nc14, when nuclei are smaller and more densely packed compared to earlier stages. To empirically define a smaller MDT, the distribution of measured distances was plotted, and a Gaussian kernel density estimate (KDE) was used to estimate the probability density function (PDF) (Fig. S1E; see Methods). The MDT was estimated from the local minimum of each of the PDFs across time points. To be conservative, the smallest local minimum, occurring at nc14, was taken as the MDT. Any pairwise distance measurement that was larger than the 2 μm MDT (Fig. 1D, blue sphere), or that spanned multiple nuclei (Fig. 1E), was removed from further consideration (see Methods).
We then focused on whether individual spots within doublets or triplets, and within the MDT of a single nucleus, could be resolved from each other. A resolution limit threshold (RLT) was applied such that any reported distance below the microscope’s detection limit was set to zero, as these spots are not resolvable (Fig. 1D, black ellipsoid). To account for distinct resolution limits in the x and y dimensions versus the z dimension, the RLT was applied using the equation of an ellipsoid. The axes of the ellipsoid correspond to the resolution limits: 120 nm in the x-y plane and 350 nm in the z dimension, based on the specifications of the microscope used (see Eq. 1).
Following the filtering and validation of the detected spots, the pairwise distances between spots were calculated. We used this thresholding, validation, and pairwise distance calculation to analyze inter-spot distances for three CRMs at the brk locus. Additionally, we utilized two mutant lines to investigate the effects of removing or adding DNA on local chromatin conformation in conjunction with hybridization chain reaction (HCR) to correlate any changes with gene expression.
Dynamic brk expression is altered by PPE deletion and MiMIC
To provide a reference for subsequent distance measurements between the CRMs, brk expression was first assessed over the same time course as the chromatin conformation assay (Fig. 2). Our previous analyses using large reporter constructs and CRISPR-mediated deletions at the endogenous brk locus (see Fig. 2A) supported the idea that E1 and E2 function at different developmental stages and drive overlapping but distinct expression patterns16,23. Specifically, before cellularization, E1 supports brk expression in a thin lateral stripe, while E2 supports expression of brk in a broad stripe within the presumptive neurogenic ectoderm at cellularization16,23 (Fig. 2B-B”). The early brk expression pattern is lost upon deletion of E1, and the late pattern is reduced upon deletion of E2 (Fig. 2E and F”)16. Thus, the normal progression of brk gene expression from a thin- to broad-lateral stripe was thought to be associated with the sequential action of E1 and E2.
A Schematic of the brk locus highlighting the 800 bp deletion of the proximal PPE (Δ800PPE), deletions of E1 (ΔE1) or E2 (ΔE2) enhancers, and the increase in distance between the E1 and PPE by 7.3 kB through the MiMIC insertion. B-F” HCR against brk transcript in WT (B-B”), Δ800PPE mutant (C-C”), MiMIC insertion (D-D”), ΔE1 (E-E”), and ΔE2 mutant (F-F”) embryos at nc13 (top), nc14 early (middle), and nc14 late (bottom). Scale bar corresponds to 50μm. G Average intensity measurements of brk transcripts detected by HCR for indicated genotypes at nc13 (WT n = 12, Δ800PPE mutant n = 10, MiMIC insertion n = 12, ΔE1 n = 10, ΔE2 n = 10), nc14 early (WT n = 14, Δ800PPE n = 12, MiMIC n = 10 and ΔE1 n = 10, ΔE2 n = 11) and nc14 late (WT n = 12, Δ800PPE n = 11, MiMIC n = 10, ΔE1 n = 11 and ΔE2 n = 9). H Average width measurements of brk pattern at nc14 early and nc14 late. Statistical significance was assessed using one-way ANOVA followed by Tukey’s Honest Significant Difference (HSD) test for multiple comparisons. Mean ± SD is shown in black. See also Fig. S2 and Source Data File for p-values and significant differences.
The full PPE, encompassing a 2 kb region proximal to the promoter, is required to support the action of E1 and E2, as deletion of the PPE results in a complete loss of brk expression pre-gastrulation16. With a smaller, 800 bp deletion of the promoter-proximal region of the PPE (Δ800PPE) (Fig. 2A), brk expression is supported at nc14 but fails to transition to the broad pattern, instead remaining narrow beyond cellularization16 (Fig. 2C’-C”). Previously, it was hypothesized that the 800 bp PPE is required for supporting E2 activity but not for E116. Upon re-examining brk gene expression in the Δ800PPE mutant using HCR, we found that, in addition to the previously reported failure to broaden at nc14 (a presumed E2-function), expression was also reduced at nc13 (Fig. 2C, G). These findings suggest that, in addition, the PPE may be required earlier to support the action of E1 at the initiation of brk expression in nc13. In support of this, a signal is detected at nc13 when E1 is active (ΔE2) (Fig. 2F, G) but not when E1 is deleted (ΔE1) (Fig. 2E, G) or in the Δ800PPE (Fig. 2C, G), indicating that both E1 and the 800PPE are critical for early expression. The intensity of ΔE2 is significantly lower than WT at nc13, however, suggesting that E2 also does play a minor role at this stage (Fig. 2F, G, Source Data File Fig. 2).
As the deletion of 800 bp associated with the Δ800PPE mutant decreased the distance between E1 and PPE, we next tested how increasing the distance between E1 and PPE would affect brk gene expression. To do this, we analyzed brk expression in a line containing a 7.3 kb MiMIC insertion located between E1 and the PPE, using HCR (Fig. 2A, S2). In embryos containing this MiMIC insertion, we observed reduced levels of brk expression at nc13 (Fig. 2D, G), as well as a reduced domain of expression at nc14 late compared to WT (Fig. 2B”,D”,H), similar to what was detected for the Δ800PPE mutant (Fig. 2D-D” compare with 2B-B” and 2C-C”,G,H).
Although this mutant introduced a sequence between the E1 and PPE, this sequence was not necessarily inert. Specifically, the MiMIC contains a yellow+ marker flanked by two inverted bacteriophage ΦC31 integrase attP sites, as well as a splice site acceptor that works to inactivate genes when inserted in introns24,25 (Fig. S2A). To determine whether this insertion exerted a sequence-specific effect on brk transcription, we repeated the HCR using probes targeting both brk and yellow. Although endogenous yellow is not expressed at this stage of development, we detected yellow transcript overlapping with the brk expression pattern, specifically in this MiMIC line (Fig. S2B-G). Notably, the normalized intensity of yellow gene expression is significantly higher than brk gene expression in the MiMIC across all timepoints analyzed (Fig. S2H, Source Data File Fig. S2). Additionally, the width of yellow did not change much over time, in contrast to the brk expression which expands from nc14 early to nc14 late (Fig. S2I, Source Data File Figure S2). The dynamics of the yellow pattern, both the early strong expression as well as the lack of expansion at later stages, closely matches the brk expression driven by E1 alone (see ΔE2 in Fig. 2F-F”, G, H). These results suggest that the yellow promoter in the MiMIC insertion hijacks E1 activity, driving yellow expression at the expense of brk26.
To investigate whether the dynamic changes in brk gene expression that we observed in the above mutant lines correspond to changes in chromatin conformation, we examined brk chromatin regulatory interactions across developmental stages, in wild type as well as the MiMIC and ΔPPE mutant embryos, using our PLOTTED technique.
Probabilistic framework for inferring conformation
To compare how brk expression relates to E1, PPE, and E2 chromatin dynamics, the PLOTTED technique was applied to WT and mutant embryos of comparable stages when brk is expressed. For each nucleus, we measured pairwise distances between detected fluorescent spots, whether from triplets (all three CRMs detected) or doublets (only two detected). Measurements were collected across 100–1000 nuclei for a minimum of 6 embryos per sample (see Source Data File Figs. 3–5) and analyzed over both developmental time and across genotypes. Pairwise distances between E1-PPE, E2-PPE, and E1-E2 were plotted as PDFs for each developmental stage: pre-nc13, nc13, nc14 early, and nc14 late (Fig. 3A, B-B”; see Methods). To compare distance distributions between stages and genotypes for statistically significant differences, we used the Kolmogorov-Smirnov (K-S) test and the resulting p-values were displayed as a heatmap next to PDFs (e.g. Fig. 3B for E1-PPE distance, see also Eqs. 2 and 3 in Methods). From this analysis of the PDFs and K-S test for significance, trends over time can be compared for WT versus the mutants (Fig. 3 vs Figs. 4 and 5).
A Schematic illustrating how the three CRM pairwise distances were plotted as x-y coordinates to generate 3D contours. Probability Density Function (PDFs) of individual distances were combined to give rise to a 3D contour plot in which E1 is set at (0,0), E2 is on the x axis at (E1-E2, 0), and the PPE is plotted in the positive y quadrant (xPPE,yPPE). Grayscale bar along the x axis shows the E2 positional likelihood. B-B” PDFs for pairwise E1-PPE (B), E2-PPE (B’), or E1-E2 (B”) distances (μm) of WT, generated by gaussian kernel density estimation. Each distance data point is modeled as a probability distribution centered at the observed value, resulting in solid line traces. The shaded region surrounding line traces represents the 2.5% to 97.5% confidence interval. Adjacent heatmaps show two-sided Kolmogorov-Smirnov (K-S) test p-values (Benjamini–Hochberg corrected for multiple comparisons) using a color scale: yellow/pink is significant (p < 0.05 indicated by bracket and *), whereas purple/black is not; self-comparisons (black, p = 1.0) run diagonally through each heat map square. Arrows indicate occurrence of a significant change. C–F 2D projections of 3D landscape for WT over time, colored from white, purple, to orange to indicate low, medium, and high probability. G, H Subtraction contours comparing genotypes, generated by subtracting the second genotype (blue) from the first (red). Red indicates higher occupancy probability for the first genotype, blue for the second. The most probable position (MPP) of the first genotype is shown as filled circles, while the second genotype is shown as hollowed squares with their respective colors of blue (E1), green (PPE), and red (E2). All E1-E2 possible distances are shown below with the top grayscale being the first genotype. Distance modes for each triangle indicated by solid (contour 1) or dashed (contour 2) lines. Global two-sided Simes P-value and interpretations are shown for each subtraction contour. See also Fig. S3 and Source Data File for number of embryos and doublets for each condition, individual p-value as well as other relevant statistics.
A-A” Plot of probability density function (PDF) for Δ800PPE mutant pairwise E1-PPE (A), E2-PPE (A’), or E1-E2 (A”) distances (μm), generated from gaussian kernel density estimation of measured distances. Each distance data point is modeled as a probability distribution centered at the observed value, resulting in solid line traces. The shaded region surrounding each of the solid line traces represents the confidence interval from 2.5% to 97.5%. Adjacent heatmaps show two-sided Kolmogorov–Smirnov (K-S) test p-values (Benjamini–Hochberg corrected for multiple comparisons). Yellow/pink denotes significant (p < 0.05 indicated by bracket and *), whereas purple/black indicates non significance; self-comparisons (black, p = 1.0) run diagonally through each heat map square. Arrows indicate the occurrence of a significant change. In the Δ800PPE mutant, major shifts occur between nuclear cycle (nc) 13 (2) and early nc14 (3) for both E1–PPE and E2–PPE distances. B–E 2D projection of 3D contour data for Δ800PPE mutant over time. Color scale from white purple to orange displaying regions of low, medium or high probability of PPE location relative to the other CRMs. Grayscale along x shows E2 positional likelihood. F–H Subtraction contours in which the second sample (blue: the earlier time point) is subtracted from the first sample (red: the later time point). Red indicates higher probability for the first genotype, blue for the second. The most probable position (MPP) of the first genotype is filled circles, while the second genotype is shown as hollowed squares with their respective colors of blue (E1), green (PPE), and red (E2). All E1-E2 possible distances are shown below with the top grayscale being the first genotype. Distance modes for each triangular configuration are indicated by solid (contour 1) or dashed (contour 2) lines. Global two-sided Simes P-values and their interpretations are shown for each subtraction contour. See also Fig. S3 and Source Data File for number of embryos imaged, number of doublets counted for each condition, and individual p-value as well as other relevant statistics.
A-A” Plot of PDFs for MiMIC insertion pairwise E1-PPE (A), E2-PPE (A), or E1-E2 (A”) distances (μm), generated from Gaussian kernel density estimation of measured distances. Each distance data point is modeled as a probability distribution centered at the observed value, resulting in the solid line traces. The shaded region surrounding each of the solid line traces represents the confidence interval from 2.5% to 97.5%. Adjacent heatmaps show two-sided Kolmogorov–Smirnov (K-S) test p-values (Benjamini–Hochberg corrected for multiple comparisons) using a color scale: yellow/pink is significant (p < 0.05 indicated by bracket and *), whereas purple/black is not; self-comparisons (black, p = 1.0) run diagonally through each heat map square. Arrows indicate the occurrence of significant change. For MiMIC insertion, the major changes are from nc13 “2” to nc14 early “3” for E1-PPE and from pre-nc13 “1” to nc13 “2” for E2-PPE. B–E 2D projection of 3D contour data for MiMIC insertion over time. Color scale from white, purple, to orange displaying regions of low, medium or high probability of PPE location relative to the other CRMs. Grayscale along x shows E2 positional likelihood. F–H Subtraction contours in which the second sample (blue: the earlier time point) is subtracted from the first sample (red: the later time point). Red indicates a higher probability for the first genotype, blue for the second. The most probable position (MPP) of the first genotype is filled circles while the second genotype is shown as hollowed squares with their respective colors of blue (E1), green (PPE) and red (E2). All E1-E2 possible distances are shown below with the top grayscale being the first genotype. Distance modes for each triangular configuration are indicated by solid (contour 1) or dashed (contour 2) lines. Global two-sided Simes P-values and their interpretations are shown for each subtraction contour. See also Fig. S3 and Tables S2, S3 for number of embryos imaged, number of doublets counted for each condition, and individual p-value as well as other relevant statistics.
Although the pairwise distance analysis is informative for understanding a specific interaction between two CRMs (e.g. E1 and PPE), it is difficult to extrapolate the conformation of all three CRMs by looking at the individual PDFs. To estimate the probability of specific conformations occurring using the pairwise distances and their associated probabilities, a contour was generated that represents the probability of a locus occurring in a specific location relative to the other loci. More specifically, to generate the contour map of the probability of the PPE occupying specific locations relative to E1 and E2 for a particular genotype, the possible positions of the PPE and the probability at those positions must be determined. For the brk locus, as shown here, this was accomplished by positioning E1 at the origin, E2 on the x-axis, and the PPE in the positive y-space. The possible positions of the PPE were calculated by iterating through the sub-sampled distances on the E1-PPE and E2-PPE PDFs (0-2 μm) and using the positions of E1 and E2 to solve the distance formula for the PPE’s coordinates. The probability density of the PPE occupying this position was calculated by multiplying the probability densities for each of these sub-sampled distances together. This was repeated for all the different possible locations for E2 on the x-axis, using sub-sampled distances from the E1-E2 PDF (0–2 μm). The probability densities generated for each E2 location were averaged together using a weighted average, where the associated probabilities from the E1-E2 PDF were used as the weights (Fig. 3A; see Methods).
Although the positional data for the detected spots within nuclei is 3-dimensional (xyz), any triangle the spots form can be represented in 2-dimensions by defining a coordinate system where one point is fixed at the origin, one point is positioned on the x-axis, and the other point is limited to the positive y-space (see Methods, e.g. Fig. S3E-I for an alternative fixed anchor at the PPE for WT). These 3D probability plots (e.g. Fig. S3A–D for WT) can also be visualized as 2D contour maps (e.g. Fig. 3C–F; see Methods). Furthermore, differences between samples, either varying by genotype or time, were assessed by subtracting contour maps from each other to illustrate significant differences (e.g. Fig. 3G) and non-significant differences (e.g. Fig. 3H). To understand how conformations change, the significance values from the individual PDFs were compared to the contour subtraction plots to understand how the probability of a CRM occupying a certain area has changed over time, genotype, or space. Notably, two sets of p-values are given for each of the subtraction plots: the first is derived from the pairwise comparison of the PDFs (see Source Data File Figs. 3–5), and the second is a global p-value which was utilized to further statistically validate the subtraction plots. The global p-value was calculated from individual distances p-values using the Simes test to determine if two contours were significantly different from each other (see Methods). Global p-values are reported below each subtraction contour when a significant difference was uncovered.
Distances between enhancers and the PPE decrease at nc13
To gain further insight into how the pairwise distances change over time in WT, the pairwise distance PDFs were compared to each other. From this analysis, the only significant differences in WT for both E1-PPE and E2-PPE distances were between pre-nc13 and all later time points (Fig. 3B,B’). In contrast, the E1-E2 distances do not exhibit the same trend, as significant differences were observed only for nc14 early versus pre-nc13 or nc14 late (Fig. 3B”). While such comparisons are useful for understanding the differences in pairwise associations, it is not as clear how the overall conformation is affected.
To better understand how the WT conformation changes over time, the three PDFs for each pairwise distance measurement were used to generate contours of the probability density for the PPE’s location at different time points (Fig. 3C–F and S3A–D). When the contour plot for pre-nc13 was compared to the later time points, we observed that there is a higher probability of the PPE being closer to E1 and E2 starting from nc13 (Fig. 3G). However, after that there is minimal difference in spatial proximity between the three CRMs (Fig. 3H). Additionally, while the contours were generated using E1 as the reference at (0,0) to explore PPE location (Fig. 3A), a similar trend is observed when the PPE is set as the reference at (0,0) and E1’s location is explored (Fig. S3E–I). Collectively, these data demonstrate that the E1-PPE and E2-PPE distances both decrease between pre-nc13 and nc13 and, thereafter, do not vary significantly.
The timing of this shift in chromatin conformation coincides with active transcription, as the earliest brk expression was identified at nc13 (Fig. 2B). Additionally, these local movements occur concurrently with the uptick in chromatin contacts measured by Hi-C studies of Drosophila early embryos, which show TAD formation initiates in nc131.
E1-PPE and E2-PPE associations are delayed in the Δ800PPE
To investigate how E1, PPE, and E2 dynamics were affected by the Δ800PPE mutant, the pairwise distances were compared to each other across time (Fig. 4A-A”). It is important to note that the PPE deletion does remove 800 bp of the sequence targeted by the PPE probe, but still leaves 3.2 kb of target sequence, which was sufficient for robust detection (Fig. 2A). In the Δ800PPE mutant, the PDFs of E1-PPE and E2-PPE at pre-nc13 and nc13 are significantly different from the PDFs of nc14 early and nc14 late (Fig. 4A and A’). To determine how these differences affected the conformation, contours were generated for the Δ800PPE mutant at the four timepoints (Fig. 4B-E) and visualized using subtraction contours (Fig. 4F–H). Noticeably, the decrease in E1 and E2 relative distances to the PPE in the Δ800PPE mutant occurs later, between nc13 and nc14 early (Fig. 4G), instead of between pre-nc13 and nc13 as identified in WT (Fig. 4F compared to 3 G). After nc14 early, there is no change in conformation over time, similar to WT (Fig. 4H compared to 3H). These results suggest that the chromatin conformation shift is delayed in the Δ800PPE mutant.
The MiMIC insertion causes a delay in E1-PPE association
To test the effect of altering genomic spacing, we utilized the MiMIC insertion background (Fig. 2A and S2A). Comparison of the PDFs for the MiMIC over time (Fig. 5A–A”) shows that the E1-PPE distance is significantly different in both pre-nc13 and nc13 when compared to nc14 early and nc14 late (Fig. 5A). This change in the E1-PPE distance is delayed relative to WT, but similar to the Δ800PPE mutant (Fig. 5A versus 3B and 4A). On the other hand, the major change in the E2-PPE distance occurred at nc13, consistent with WT (Fig. 5A' vs 3B’).
When probability density contours were plotted over time (Fig. 5B–E), we observed an increase in the probability of shorter E2-PPE distances at nc13 (Fig. 5F), whereas reduction in E1-PPE distances were not detected until nc14 early (Fig. 5G). Similar to WT, there is no significant difference in conformation after nc14 early in the MiMIC line (Fig. 5H). Collectively, these data demonstrate that in the MiMIC insertion, the individual enhancer to PPE distances are differentially regulated.
Δ800PPE shows distinct conformational shifts vs WT
To gain further insight into the Δ800PPE mutant phenotype in terms of chromatin conformation, its PDFs for E1-PPE, E2-PPE, and E1-E2 distances were compared to those of WT. At nc13, the E1-PPE and E2-PPE distances are significantly different between the Δ800PPE and WT (Fig. 6A, red box #1). Using a subtraction contour to spatially interpret this difference, we found that the PPE has a higher probability of being farther from both enhancers in the mutant at nc13 (Fig. 6A’.1). This conformational difference likely explains the loss of gene expression in the mutant at this stage (Fig. 2C, G). On the other hand, when comparing the Δ800PPE phenotype to WT at late nc14, there are significant differences in E1-PPE and E1-E2 distances but not E2-PPE distances (Fig. 6A, dashed blue box #2). The contours demonstrate an increased probability of the PPE being close to E1 in the Δ800PPE mutant (Fig. 6A’.2). Together with the altered brk expression observed at late nc14 in Δ800PPE mutants (Fig. 2C”, H), these data suggest that, when the PPE remains in close proximity to E1, E2 is unable to drive the broad activation observed in WT (Fig. 2B”, H), supporting the hypothesis that the PPE is “stuck on E1”16. To investigate the opposite case, we utilized a transgenic line that increases the distance between one enhancer and the PPE.
A–C Heatmap showing direct two-sided Kolmogorv-Smirnov (K-S) test comparison for all three pairwise distances between WT and Δ800PPE (A), WT and MiMIC (B) or between mutants (C) for all three pairwise distances. Color scale bar: yellow/pink is significant (p < 0.05 indicated by bracket and *), whereas purple/black is not. A’-C’ Subtraction contours comparing genotypes: red indicates higher probability of the PPE of the first genotype being present in that location, while blue indicates a higher probability for the second. The most probable positions of each element are indicated by filled circles for the first genotype and hollow squares for the second (E1 = blue, PPE = green, E2 = red). The mode of the distances for each triangle are shown as either a solid line (contour1) or a dashed line (contour 2). Two grayscales for E1-E2 PDFs are shown under each subtraction contours (top = first genotype, bottom = second genotype). Every subtraction contour shown has at least one pairwise comparison that is significantly different. Global p-values, calculated by two-sided, multivariate Simes test, and interpretations are listed under each subtraction contour. (D) Model of chromatin conformation changes over time in relation to transcription in WT and the two mutant lines. Distances between the PPE (green circle) and each enhancer (E1-blue circle and E2-red circle) for each developmental stage and genotype are representative of the median and overall distributions of the measured values. In WT, major chromatin rearrangement occurs from pre-nc13 to nc13 (purple arrow). In Δ800PPE, significant E1 and E2–PPE distance changes occur between nc13 and early nc14 (purple arrows). In contrast, in the MiMIC line, E2–PPE changes between pre-nc13 and nc13 (red arrow) and E1–PPE between nc13 and early nc14 (blue arrow). Both mutants show positional bias: Δ800PPE favors E1-PPE proximity in late nc14 (blue dashed box), and MiMIC favors E2-PPE proximity in nc13–early nc14 (magenta dashed boxes). These conformational shifts align with corresponding changes in gene expression. See also Tables S2, S3.
The PPE is closer to E2 in the MiMIC compared to WT
When the MiMIC is directly compared with WT, an effect on the E1-PPE distances is identified, as expected, because the insertion of a large DNA fragment increases the genomic distance between E1 and the PPE and also introduces an exogenous promoter which interacts with E1 (Fig. S2). This difference of E1-PPE in the MiMIC was detected at nc13 (Fig. 6B, cyan box #3); but, surprisingly, the distance converged toward WT by early nc14 (Fig. 6B, pink dashed box #4), suggesting that the MiMIC insertion only delays E1 action. In contrast to E1-PPE, the E2-PPE distances are significantly different at both nc13 and nc14 early (Fig. 6B cyan and pink boxes). When the contour for nc13 was compared for WT and the MiMIC insertion, we find that the decrease in the E1-PPE distances is delayed relative to WT, and instead the PPE is more closely associated with E2 (Fig. 6B’.3). By nc14 early in the MiMIC, the E1-PPE distances are comparable to WT, but the PPE is still closer to E2 than in WT (Fig. 6B’.4). Together these results show that E2 exhibits a precocious and prolonged association with the PPE at the early stages, and that this change is correlated with a delay in E1-PPE association. An interesting note is when looking at the distance distribution of E2-PPE in the MiMIC line at nc13 and nc14 early, there is some increase in density of larger distances despite lower median values (Fig. 5A’). This suggests that contacts may be less stable between E2-PPE, rather than consistently closer. However, it is still remarkable that increasing the linear DNA distance between E1 and the PPE through the MiMIC insertion not only altered their early spatial relationship at nc13, but also contributed to increased proximity between E2 and the PPE in a substantial fraction of measured nuclei.
These conformational changes are reflected in the gene expression of the MiMIC insertion as well, with a loss of brk expression at nc13 (E1-supported activity) and a narrow expression pattern at nc14 late (E2-supported activity) (Fig. 2D,D”,G,H). We propose that the MiMIC inserted sequence delays, but does not block, the action of E1, and this delay in E1 action interferes with E2 action, resulting in effects on both the early (i.e., nc13) as well as late (i.e., nc14 late) expression of brk. These findings suggest that the timing of interactions between individual CRMs is critical for proper gene regulation and disrupting this sequence can lead to aberrant expression patterns (see Discussion).
The PPE is closer to E2 in MiMIC and to E1 in Δ800PPE
To further understand how the observed differences in the MiMIC insertion and the Δ800PPE mutant affected regulatory chromatin dynamics, the MiMIC insertion was directly compared to the Δ800PPE mutant. From this comparison, there is an increased probability of the PPE being close to E2 in the MiMIC insertion at nc13 (Fig. 6C, cyan box #5, and 6 C’.5). Furthermore, when comparing the MiMIC insertion to the Δ800PPE mutant at nc14 late, the PPE has a higher probability of being closer to E1 in the Δ800PPE mutant (Fig. 6C, dashed blue box #6, and 6 C’.6). These results suggest that, when comparing the MiMIC and Δ800PPE mutants, the PPE interacts preferentially with opposite enhancers and with different temporal dynamics.
Chromatin conformation varies across A-P and D-V axes
To leverage the spatial information that the PLOTTED technique can provide, we also investigated differences in chromatin conformation by comparing imaging data from regions along the dorsal-ventral (DV) axis which were located in the lateral and ventral domains of the embryo (Fig. 7A). Here, we utilized an antibody to the Twist (Twi) transcription factor expressed in the presumptive mesoderm22 to mark the ventral most region of the embryo, where brk is transcriptionally repressed27 (Fig. 7A, regions 2 and 4). Nuclei were imaged within the Twi domain as well as in the adjacent lateral domain, corresponding to the presumptive neurogenic ectoderm, where brk is active (Fig. 7A, regions 1 and 3). Since Twi protein is only detectable by antibody staining at nc14, this analysis was only conducted at this stage. Despite this limitation, significant differences in conformation were detected between regions of brk activity and repression. In regions of active brk transcription, there are smaller distances between the enhancers and the PPE, compared to regions where brk is repressed (Fig. 7C-C”, E vs F). When we contrast the active state (Twi-negative regions 1 and 3) against the inactive state (Twi-positive regions 2 and 4) in the subtraction plot, we see a clear preference for a smaller chromatin structure in the active state (Fig. 7G).
A Diagram illustrating the Twist (red) and no-Twist (white) regions. Since Twist protein marks the ventral region in which brk transcription is repressed, this allows us to correlate chromatin conformation changes in regions of brk gene expression against regions that have no brk expression. We can also further subset and compare chromatin conformation changes between the anterior and posterior. This is illustrated by the numerical box diagramming comparisons. B,B’ HCR against brk transcript at nc14 early (B, n = 9) and nc14 late (B’, n = 11) highlighting a broader brk expression at the anterior as compared to the posterior. C-C” Overall comparison between active and inactive regions of brk for all three different CRMs E1-PPE (C), E2-PPE (C’) and E1-E2 (C”), regardless of anterior-posterior positions. Both E1-PPE and E2-PPE illustrate a significant difference between active and repressed brk region. Asterisks denote significance with p-value < 0.05. The exact individual pairwise p-values are shown in the Source Data File. Statistical test conducted via two-sided Kolmogorov-Smirnov (K-S) tests. D-D” Anterior versus posterior comparison for all 3 CRMs, E1-PPE (D), E2-PPE (D’) and E1-E2 (D”). E, F, H, I Contour plots for all compared subset and subtraction contour plots for DV (active E vs repressed F) as well as AP (anterior H vs posterior I) differences. G, J Subtraction contour highlights that chromatin conformation exhibits a more compact state in the active region (G). The posterior has more conformations in which E1-PPE are closer than the anterior at nc14 (J). The most probable positions of each element are indicated by filled circles for the first genotype and hollow squares for the second (E1 = blue, PPE = green, E2 = red). The mode of the distances are shown as either a solid line (contour 1) or a dashed line (contour 2). Two grayscales for E1-E2 PDFs are shown under each subtraction contours (top = first genotype, bottom = second genotype). Global p-values, calculated by two-sided multivariate global Simes tests, and interpretations are listed under each subtraction contour. See also Fig. S4.
In addition to the DV analysis, we additionally subset these data into AP components for comparison. This analysis was performed based on visual inspection of the brk pattern at nc14, which appears slightly broader at the anterior compared to the posterior (Fig. 7B, B’). Spatially partitioning the data in this way revealed that only E1-PPE distances exhibited a change in chromatin conformation dynamics between anterior and posterior at nc14 (Fig. 7D-D”). Specifically, in the posterior, we detected smaller E1-PPE distances as compared to the anterior (Fig. 7D, H–J). As our original imaging scheme included two scans along the AP axis of the embryo, but no anti-Twi (see Fig. 1B), we could also expand the AP assay of conformation to our larger data set that includes four stages (Fig. S4). With this expanded set of AP data, we confirmed that the distances for E1-PPE were closer in the posterior region compared to anterior at nc14 early (Fig. S4I). Additionally, we also found that E1-PPE was closer in the anterior region at an earlier stage, pre-nc13 (Fig. S4G). Together, these results demonstrate that AP domains of the embryo exhibit different chromatin conformations (Fig. 7J), possibly indicating that the anterior exhibits a shift in its conformation earlier than the posterior (Fig. S4).
In summary, the PLOTTED technique is a powerful tool to measure chromatin conformation differences in both DV and AP regions. The DV differences highlight that gene expression, specifically zygotic gene expression of brk, is correlated with local chromatin conformation changes.
Discussion
Using the PLOTTED technique to quantify pairwise distances between CRMs, we showed that chromatin dynamics are important for proper spatiotemporal expression. We identified that for the brk CRMs, both the E1-PPE and E2-PPE distances decreased between pre-nc13 and nc13 in WT and found that there was a delay in this change in the Δ800PPE mutant, which correlated to changes in brk expression (Fig. 6D). This data, combined with previous studies16, suggests that the PPE likely coordinates the action of E1 and E2. This coordination in enhancer activity is important for brk expression, as both the initiation of expression at nc13 and the expansion of the expression domain at late nc14 is disrupted in the Δ800PPE mutant. We found that deletion of the PPE delayed both the initiation of expression as well as the timing of changes in chromatin conformation (Fig. 6D, box 1). Furthermore, E1-PPE interactions persisted longer in the Δ800PPE than those observed in the WT (Fig. 6D, box 2). This suggests that the conformation of CRMs contributes to the regulation of gene expression.
While the PPE is specific to the brk locus, other elements have been suggested to regulate the looping of DNA and interactions between CRMs, such as Homie16,28 and Scr45029 in Drosophila, as well as CpG islands in mammals30,31,32. The presence of these factors leads us to speculate that chromatin conformation or chromatin looping is likely a common regulatory step in gene expression. By leveraging the PLOTTED technique, or complementary labeling methods such as Hi-M and ORCA paired with the PLOTTED techniques’s post-processing analysis, gene expression can be linked to chromatin architecture. When combined with mutational analysis, this approach can provide additional insights into how gene expression is regulated and enable deeper understanding of the mechanisms that govern enhancer action.
The changes observed in the MiMIC insertion line, which added 7.3 kb between E1 and the PPE, provide further evidence for the importance of chromatin conformation in regulating gene expression. Similar to the Δ800PPE mutant, the MiMIC insertion also exhibited a delay in gene expression that correlated with a delayed decrease in the E1-PPE distance (Fig. 6D, box 3). However, this mutant also affected E2, which exhibited a higher probability of being closer to the PPE at nc13 and nc14 early in the MiMIC insertion when compared to WT (Fig. 6D, box 3 and 4). Despite this timely, and significantly greater decrease in the E2-PPE distances, the MiMIC insertion still failed to support the E2-dependent, broad brk expression pattern at late nc14. Similarly, although E2 remains intact in the ΔE1 mutant, it is unable to drive the full late-brk expression pattern in the absence of E1. In the ΔE1 mutants, the width of the brk expression pattern at nc14 late is narrower than WT and is similar to the width of MiMIC (Fig. 2E” vs 2D”, H). Therefore, if E1 is removed, or if it fails to come close to the PPE, the E2-PPE interaction alone cannot give rise to the broad brk pattern that is seen in WT (Fig. 2B” and H). Taken together, these results indicate that E1 and E2 interact to modulate one another’s activity, and that the precise timing of enhancer action is critical for establishing proper gene expression patterns.
While these findings demonstrate some of the insights PLOTTED can provide, the true power of this technique comes from its ability to retain in situ spatial information when assaying conformations. To demonstrate this capability, we compared the conformation of the brk CRMs along the DV and AP axes of the embryo. Our results demonstrate a difference in chromatin conformation along the DV axis correlated with zygotic brk gene expression at nc14 - compact conformation in domains of brk expression and farther apart in domains of repression. We also observed a difference for E1-PPE along the AP axis, with the posterior assuming a more compact conformation than the anterior at nc14. This could potentially be due to the differential regulation of brk across the AP axis leading to different expression pattern widths and dynamic changes in chromatin conformation observed (Fig. 7B,B’). Although the connection to transcriptional regulation will require further investigation, our results have demonstrated that chromatin conformation is spatially distinct and likely coupled with transcription33,34.
In closing, PLOTTED is a useful tool that can expand the functionality of preexisting labeling techniques, and provides insight into the interplay between chromatin conformation and gene expression. This technique can be used in any system that has DNA, can be fixed, and can be imaged on a superresolution confocal microscope. Through this study, we illustrated clear changes in chromatin dynamics during early Drosophila embryogenesis. However, because it is based on analyzing fixed tissues, this technique is limited in its time and imaging resolution and may not fully elucidate functional enhancer-promoter interactions. In addition, we attempted to interrogate the relationship of chromatin conformation to gene transcription by blocking active transcription with the drug alpha-amanitin. However, these experiments were limited by the practical challenge of processing enough embryos (~500) through manual dechorionization, injection, hand devitellinization, and fixation for DNA FISH. We are hopeful that further improvements to this labeling technology will allow for the simultaneous capture of nascent transcription and CRM positions, refining our understanding of how the distances between enhancers and promoters regulate gene expression. The current PLOTTED technique will offer new insights into how enhancers function, a mechanism that is not yet fully understood.
Methods
Fly husbandry
D. melanogaster stocks were kept at 24 °C in standard medium. D. melanogaster flies from a yw strain were utilized as a WT control. Embryos were collected in a collection cage with apple juice plates at 24°C for 2 h and then aged for a further 2 h, resulting in embryos that are 2–4 h old. The embryos were fixed using a 1:1 mixture of heptane and 4% paraformaldehyde (PFA) in PBS, as previously described35. Embryos were stored in methanol at −20°C. All fly lines used in this study are listed in Table S1. Drosophila strains and other reagents generated in this study will be available upon request from the lead contact, or the commercial sources listed in the key resources table.
Oligo-paint DNA FISH labeling: probe design and construction
Primary brk probes were designed using OligoMinerApp36 using the single input with D. melanogaster BDGP6.22 as a reference genome. Probe sets were designed against 4 kb regions surrounding the three different CRMs at the brk locus (E1, PPE, E2). Input sequences are listed in Table S2.
Approximately 50 primary oligopaint probes of 150 bp each were generated from IDT to complement the three regions of the brk locus (sequences listed in Table S2). Secondary oligonucleotides conjugated to fluorophores were generated to be complementary to one of the three ~32 bp universal primers, each marking unique regions of the brk locus37. Full probes with the universal primer, primary oligo-pool sequences, and secondary probe sequences are outlined in Table S2. All primary and secondary oligonucleotides were synthesized by IDT.
Primary probes were generated following a four step procedure of: 1) a limited cycle PCR amplification which adds a T7 site using the reverse primer, 2) in vitro transcription via the introduced T7, 3) reverse transcription of the synthesized oligonucleotides, and 4) purification of the product via column purification to generate the primary oligo-nucleotides for hybridization as described in Gizzi et al. 12.
Embryo DNA FISH hybridization
Primary probe hybridization was conducted following the protocol for DNA in situ hybridization as described by Gizzi et al. 12. In brief, first embryos were re-hydrated from methanol to PBT (PBT = PBS-0.1% Tween20) in a sequential manner (90% MeOH, 10% PBT; 70% MeOH, 30% PBT; 50% MeOH, 50% PBT; 30% MeOH, 70% PBT; 100% PBT). Then, they were incubated in 1mL of 100 μg/mL RNase A in PBT overnight at 4 °C. Embryos were permeabilized using PBS-Tr (PBS-0.5 % Triton) for 3 h at room temperature.
Pre-hybridization mixture (pHM) was prepared from 50% formamide, 4X SSC, 100 mM NaH2PO4, pH7, and 0.1% Tween 20. Pre-hybridization was conducted through sequential transfer from PBS-Tr into the pre-hybridization mixture (80% PBS-Tr, 20% pHM; 50% PBS-Tr, 50% pHM; 20% PBS-Tr, 80% pHM; 100% pHM). DNA Hybridization Solution (DHS) was prepared using 2X SSC, 10% (vol/vol) dextran-sulfate, 50% formamide, and 0.5 mg/ml salmon sperm DNA (i.e. 2% of a 2.5% sonicated and autoclaved stock solution). Embryos were transferred to fresh pHM solution before denaturing for 15 minutes at 80 °C. Primary oligonucleotide probes were added to DHS for denaturation at 80 °C for 15 min. Primary oligonucleotide concentration can be from 15–45pmole. Finally, probes were added to the denatured embryos in a 80°C heat bath. Embryos were subsequently removed from the heat and allowed to cool to 37 °C. Then they were transferred to a 37°C thermomixer with shaking overnight at 450 RPM.
Post hybridization washes were 20 min each, with the first five washes conducted at 37 °C and 700 rpm in the thermomixer, and washes six through eight at room temperature on a rotating platform. The first post-hybridization wash was with 50% formamide, 2XSSC, and 0.3% CHAPS. The second post hybridization wash was 40% formamide, 2X SSC, and 0.3% CHAPS. Washes 3-6 were sequential dilutions into PBT (30% formamide, 70% PBT; 20% formamide, 80% PBT; 10% formamide, 90% PBT; 100% PBT). Wash 7 was with 4% PFA-PBT to fix the primary oligonucleotide and the final wash was transferring embryos back to PBS-Tr.
Secondary probe hybridization was adapted from both Gizzi et al. 12 and Nguyen et al. 38. First, embryos were transferred from PBS-Tr to pHM sequentially, similar to the primary oligonucleotide hybridization described above. Then, 50 pmol of ordered secondary oligonucleotide was added and the samples were incubated for 2 h at room temperature away from light at 450 rpm. Post-secondary oligonucleotide hybridization washes were conducted as described above, omitting the final wash step and incubating in blocking solution (1X Western Blocking Reagent (Roche, 11921673001) in PBT) for 5 minutes.
Embryos were incubated in a blocking solution for 2 h, and then incubated overnight with mouse anti-lamin (DSHB ADL84.12, 1:150 in blocking solution) to outline nuclei. The next morning, the embryos were rinsed with PBT and incubated in blocking solution for 30 min before being incubated overnight with the secondary antibodies, Goat Anti-Mouse 405 (Thermo Fisher Scientific Catalog # A-31553, 1:400 in blocking solution), at 4 °C.
On the final day, the embryos were washed three times with PBT and mounted in SlowFade Gold (ThermoFisher S36937).
Image acquisition
Embryos were mounted on a slide using a #1.5 coverslip and imaged using a Zeiss LSM 980 with Fast Airyscan 2 for superresolution microscopy. Embryos were imaged at the Trunk-Anterior and Trunk-Posterior regions (Fig. 1B) using a 63X lens. Imaging was conducted on the Zeiss LSM 980 with Fast Airyscan to first produce 32 images through the Airyscan detector, which then underwent Sheppard summation followed by a 3D deconvolution to create a final image of 16 pixel bit depth with increased dynamic range through built-in Zen software39. The resolution limit of Zeiss LSM 980 Airyscan 2 is 120 nm in the lateral directions and 350 nm in the axial direction40.
The staging for developmental time of each embryo was done based on the number of nuclei captured in the field of view as well as the shape of the nuclei (Fig. 1C). Pre-nc13 embryos are characterized by <100 nuclei, nc13 embryos are characterized by ~150 nuclei, and nc14 early and late are characterized by ~200-300 nuclei per field of view. Nc14 early embryos were further distinguished from nc14 late by the shape of the nuclei as well as the space between nuclei. Nc14 late nuclei are more tightly packed and more irregularly shaped directly preceding gastrulation (Fig. 1C). Both male and female embryos were imaged and analyzed.
Initial thresholding and spot detection through find_spots custom pipeline
Deconvoluted ZEN images were analyzed using an in-house computational pipeline to detect spots in each channel for proximity analysis of the three different CRMs. The computational pipeline includes a built-in GUI which facilitates the loading of several image files for batch processing. The program first retrieves metadata information from the ZEN image and converts the pixel information into micrometers. In order to detect spots in each channel, the pipeline utilizes BM4D to denoise each channel. Based on the GUI inputs for Spot Detection Settings in each channel, spots in each channel that are greater than the threshold will be detected. The spot detection threshold was selected manually. Spots in each channel are validated for the best threshold as shown in Fig. S1B. The 550 channel contains spots corresponding to E1, the 647 channel contains spots corresponding to the PPE, and the 488 channel contains spots corresponding to E2.
The program outputs the maximum intensity projection with boxes highlighted around where spots for each channel are detected. Each image underwent visual inspection to guarantee all the spots highlighted are true spots before undergoing distance analyses. In the zoomed image, spots are highlighted to show it is detected. A spot is considered true when it has a clear round shape with high signal intensity. For our purposes, we chose two sets of spot detection thresholds (Fig. S1A). The lower spot detection threshold is 0.03 for the 550 channel, 0.02 for the 647 channel, 0.08 for the 488 channel. The higher spot detection threshold is 0.05 for the 550 channel, 0.05 for the 647 channel, 0.14 for the 488 channel. We utilized two different thresholds because some images were noisier than others. Each image was tested against both a lower and a higher threshold to determine which one yielded the most accurate spot detection before proceeding to the next step. We tested whether significant differences are maintained using the Kolmogorov-Smirnov (K-S) test, which is widely used in this paper. The Kolmogorov-Smirnov test is a non-parametric test that has been shown previously to be useful for comparing the absolute differences between two empirical distributions41. This statistic uses the entire distribution, including the median and spread of the distances, to evaluate whether two samples, such as at different developmental times or from different genotypes, are significantly different from each other.
In this case, for the lower spot detection threshold, the difference between pre-nc13 and nc13 is significant with a p = 0.005. For the higher spot detection threshold, the difference is also significant with p = 0.01. Since both comparisons have p < 0.05 we can conclude that regardless of which threshold is utilized for the entire dataset, the significant difference trends are preserved, such as the comparison between pre-nc13 and nc14 early. We also tested every single time point low versus high and observed no significant differences in K-S Test. Due to this, we believe that as long as spots are validated to be accurately detected in subsequent steps, spot detection thresholds can be subject to change, dependent on the samples imaged.
Generative AI (Chat-GPT) was used to aid the production of post-processing code using outputs from find_spots pipeline. The authors reviewed/validated the code generated by generative AI and take full responsibility for the content of the publication.
Doublet or triplet detection and postprocessing
Once the spot is detected, its positional information is collected. Triplet detection is performed starting with the designated middle channel (i.e., 647 or promoter proximal element). From the detected PPE spot, we find the nearest spot in the E1 and E2 channel. In addition to triplet detection, the software also detects pair-wise doublets. Similar to triplet detection, the nearest spot in the next channel will be grouped into a doublet. This is done for all three different possible combinations. Spots can only belong to one doublet or triplet if they are under the maximum distance threshold. Selection of the maximum distance threshold is shown in Fig. S1. If the maximum threshold is too high, spots that are in neighboring nuclei will be considered one doublet or triplet despite this not being biologically relevant. In order to determine which threshold is the best to avoid calling inaccurate doublets and triplets, probability density distributions were plotted for all three pairwise distances to determine the distance in which the probability is the lowest. The cut off for the maximum detection threshold can be found by taking the derivative of the PDF, and using the lowest probability distance value as the maximum detection threshold. Since there are multiple PDFs for different conditions, the maximum detection threshold was determined by rounding the local minimums of the various conditions. Any values that are under the maximum detection threshold are more likely to be within a nucleus than values that are greater. Additionally, because the diameter of nc13 nuclei are ~1.5x larger than the diameter of nc14 nuclei, we chose to use the smaller nuclei to determine the maximum distance threshold. From the derivative values, the most appropriate maximum distance threshold is deemed to be 2 µm. The local minimum at nc14 late was ~2 μm for each pairwise association. In order to compare the data at different nuclear cycles, the smallest diameter of nuclei was used. In this case, this is at nc14 late. This is further supported by the fact that nuclei size decreases over time. Using the smallest threshold ensures that a triplet is within a single nucleus at all time points. After 2 µm was chosen, we re-examined our data and observed that less than 2% of pre-nc13 and less than 5% of our nc13 data has recorded distances greater than 2 µm. Therefore, 2 µm is the appropriate cutoff to make sure most data are examined without having too much noise from nearby nuclei.
The final validations for maximum distance thresholds are outlined in Fig. 1C. By plotting pseudo spots that are collected against nuclear lamin, we can validate that only doublets and triplets within a nucleus are analyzed. This is the validation step to ensure that only biologically relevant data are analyzed.
The next threshold in the pipeline is the resolution limit threshold. This threshold is utilized to classify whether a triplet or doublet is considered to be resolvable and is determined by the resolution limit of the microscope. Since the Zeiss LSM 980 has a lateral resolution of 120 nm and an axial resolution of 350 nm, these values were used as the xy and z resolution limit thresholds, respectively. Distances for doublets and triplets that are under the respective resolution limit were converted to 0 nm, since they are not resolvable. Resolution limit threshold are shown through an ellipsoidal equation:
In which case \({x}_{2}\) and \({x}_{1}\) are relative positions of two dots on the x axis; \({y}_{2}\) and \({y}_{1}\) are relative positions of two dots on y axis; and \({z}_{2}\) and \({z}_{1}\) are relative positions of two dots on the z axis. \(a\) and \(b\) are xy resolution limit, 120 nm in our case. \(c\) is the z resolution limit, 350 nm in our case. Scatterplots are shown in the post-processing pipeline to highlight the unresolvable data.
Statistical analyses
The probability density distribution (PDF) was estimated using Gaussian kernel density in a non-parametric manner. Distance measurements were treated as independent and identically distributed samples drawn from an unknown underlying distribution. The PDF was estimated by placing a Gaussian kernel centered at each observed distance.42.
Since we are estimating what is the probability of getting a certain distance, we are interested in the shape of the probability function \(f\):
In which K is the Gaussian kernel, n is the number of observations and h is the bandwidth. The bandwidth (h) was selected using Scott’s rule:
Whereas n is the number of data points, sigma is the sample standard deviation and d is the number of dimensions. The resulting density estimate was evalulated on a discrete grid and normalized so that the area under the curve is equal to one.
The distributions of the PDFs were compared against each other for significant differences using the K-S test. The K-S test is useful here to compare absolute differences between two empirical distributions, such as between different developmental times, in a non-parametric manner41. The p-value generated took into account the entire distribution, including the median and spread of the distances, to evaluate whether two samples, such as at different developmental times or from different genotypes, are significantly different from each other.
The exact sample size (n), corresponding to the total number of spots and the number of fly embryos analyzed for each experimental group that were compared using the K-S test, is outlined in the Source Data File for Figs. 3–5. All the comparisons made and the reported p-values are considered a two-sided K-S test.
Reported p-values are adjusted for multiple comparisons. Comparisons across developmental time are adjusted for multiple comparisons using Benjamini-Hochberg (FDR)43 resulting in a corrected p-value. If this corrected p-value is less than 0.05, we considered the difference across developmental times for multiple comparisons significantly different. This method is an alternative to Benjamini-Hochberg ranking of p-value with both giving the same final classification of significant or insignificant difference. The comparison between WT and mutant at a specific developmental time point is also corrected through FDR since this is the same dataset as the overtime comparison.
Utilization of the distance probability density functions to visualize the probability of conformations
In order to better visualize how the measured pairwise distances affect the conformation of the three CRMs, we generated a contour plot of the estimated probability density function (PDF) of the PPE’s location at a particular xy coordinate. This allowed us to see the conformation for these CRMs with an associated probability density over developmental time and across various mutant backgrounds.
To generate these contours, the E1-PPE, E2-PPE, and E1-E2 distances needed to be converted into the xy coordinate positions for E1, PPE, and E2. Since there are three CRMs, these three points will always form a triangle. While a triangle can be formed in 3D space, the triangle can always be represented in 2D. This is true because both a triangle and a plane are defined by three non-collinear points. Furthermore, while an infinite number of triangles can be drawn in the xy plane, any unique triangle can be transformed such that one vertex is the origin, one vertex is on the x-axis, and the third vertex is limited to the positive y quadrants.
For the three CRMs we detected, we assumed that E1’s position is (0,0), E2’s position is on the x-axis, and the PPE is constrained to positive y values. Which CRM occupies which point in our assumptions (origin, x-axis, or positive y) is arbitrary and they can be switched. We were most interested in understanding how the two enhancers interact with the PPE, so we chose to plot the contour probability density for the PPE.
For a specific E1-E2 we calculated the joint PDF of E1-PPE and E2-PPE. Specifically, we assumed that E1-PPE and E2-PPE are independent of each other, which allows us to multiply the probability density of the two PDFs together. This assumption is met because many distances were measured in doublets instead of triplets. The equation for the joint PDF is:
Since we assume E1-PPE, E2-PPE, and E1-E2 are independent, we get
To calculate the joint PDF, we iterate through every E1-PPE, E2-PPE, and E1-E2 and multiply E1-PPE and E2-PPE to get the probability density for those distances. In addition, to calculate the xy coordinate generated by the three distances at each iteration, we use the distance formula and our three points for E1, E2, and the PPE.
Specifically, E1 is (0, 0), E2 is (E1-E2, 0) and the PPE is (\(a,b\)), and the equations are:
Before calculating the coordinates for the PPE (\(a,b\)), the distances are checked to determine if they form a legitimate triangle using the following inequalities:
Greater than or equal to is used instead of just greater than, because the conformation of the CRMs is valid for both valid triangles (greater than) as well as being colinear or with two CRMs being on top of each other (equal to). If the triangles are not valid, then the probability and xy-coordinate is not calculated.
Once the xy coordinates and their associated probability density have been calculated for each E1-E2 the total probability density is calculated by taking the weighted average across E1-E2 using the following equations:
Here \({w}_{i}\) is the weight for the ith distance E1-E2. Since the PDF of E1-E2 is known, it is converted into a probability by integrating over a small range to give a probability around the E1-E2 distance. This is accomplished by using a Riemann sum. The weight \({w}_{i}\) is then equal to this approximated probability around the specific E1-E2 in the current iteration \(i\). \(n\) is the total number of E1-E2 distances iterated through. Hence, the more probable the E1–E2 distance the greater the weight assigned to the contour. Computationally, we displayed the probability of PPE position using a smoothed contour heat map. The position with the highest density point is marked with the green PPE dot. E1 is anchored at (0,0) while E2 is on the x-axis highlighted for the most probable E1-E2 distance. The distribution of E1-E2 distances and are shown below the x axis using darker colors for higher probability density. Similar contour plots can be generated when the PPE is the fixed anchor (0,0), E2 is on the x-axis, and E1 is movable.
In order to estimate the probability of PPE position, we first created a 2D grid that allows us to evaluate triangle vertices. Within this grid, we initialize an empty probability density function that iterates overall E1-E2 values and their weights. For each fixed E1-E2 value, we computed a triangle shape from our d1 and d2 values to give a set of triangles coordinates with a corresponding joint probability value for each triangle. We utilized the NearestNDInterpolator to take valid triangle positions, their probability and interpolate the values into a smoothed defined grid. In this interpolator, each point on the grid is assigned the nearest original data point (\({x}_{{PPE},}{y}_{{PPE}}\)) in the triangle space. Finally, each (\({x}_{{PPE},}{y}_{{PPE}}\)) value is multiplied by the weight to generate a total surface for all three distances joint probability that will integrate to 1 across all E1-E2 values. This forms the basis of the contour map displayed.
The mode triangles were selected based on the most probable position of the PPE, as determined by the KDE surface derived from the conditional joint probability density. E1 is fixed at (0, 0), while E2 corresponds to the most probable position given the selected PPE.
In order to visualize the joint conditional distribution differences, we utilized subtraction contours. Subtraction of the KDE-based contours provided a statistically valid visualization of the differences in the joint conditional distribution. In this study, the joint conditional probability densities are constructed from the same three distances across time and genotypes (d1 = E2-PPE, d2 = E1-PPE weighted by d3 = E1-E2) to ensure that the conditioning is consistent across comparisons. All datasets have similar sample sizes (Table S2) and KDEs are estimated with the same Gaussian kernel with identical bandwidth. In addition, all KDEs are bootstrapped and subtraction is only performed on normalized, bootstrapped density estimates that are fed into the same contour grid structure. Altogether, the subtraction map can be interpreted as differences in probability density distributions between conditions to provide a valid visualization of how joint conditional distribution differs.
Because the three distances (E2–PPE, E1–PPE, E1–E2) are analyzed as independent, we chose the Simes global p-value in order to combine the individual p-value into a global, multivariate p-value. Hence, the global p-value can only be significant if at least one marginal, independent test is driving it44. In the Simes tests, the global null hypothesis is that all the distances are equal between the 2 compared conditions. Thus, even if one distance differs, the global p-value Simes will reject the null hypothesis, resulting in a significant difference. Therefore, in our case, this is a good test to utilize since we have three independent distances that are being utilized in the contour plots. If one of the distances is significant, the global p-value Simes test will also notice that this is driving an overall significant difference. This will then be visualized in a statistically relevant format in the subtraction contour.
HCR staining and imaging
Hybridization chain reaction (HCR) was performed based on Molecular Instruments protocol on whole-mount vertebrate embryos without proteinase K and mounted using Invitrogen Slow Fade gold mounting medium45. HCR probes were designed by Molecular Technologies against the mRNA transcript to brk 4049/E190. Fluorescent images of embryos were acquired using a 20X objective on a Zeiss LSM 800 microscope. Samples were collected, stained, and imaged concurrently using equivalent settings, as much as possible. Z projections of 18 z-stacks were made before feeding into a custom MATLAB quantification for max-intensity projections, segmentation and quantification of width and intensity.
Custom quantification for normalized width and intensity
MATLAB codes were adapted from Dunipace et al., Genetics, 2025 to quantify in-situ intensity and width of brk signal and yellow signal for Fig. S2A for over 10+ embryos46,47,48. Normalized intensity was calculated by subtracting the background intensity of the embryo from average brk signal intensity. Normalized width was calculated by MATLAB bwskel function, also known as skeletonization, which reduces objects to lines in the 2D binary image while preserving the structure. This gives us distances to the edge of shape and allows us to extract widths value at all the skeleton locations. From there, we take the average width. Normalized width was retrieved by dividing the average width of an embryo from the average width of brk signal. See Dunipace et al., Genetics, 2025 for additional information regarding the custom MATLAB quantification . Within the custom quantification, all embryos at nc14 were subjected to the same threshold for segmentation of brk pattern.
Separate thresholds were utilized for yellow and brk gene expression. This approach is justified because yellow and brk represent distinct gene expression patterns. Furthermore, embryos at nc13 and ΔE1 nc14 early received custom thresholding to analyze the brk pattern, as the intensity and pattern of brk at these stages are significantly weaker. Thus, we prioritize optimizing brk segmentation before conducting intensity or width analyses.
All code listed, starting from spot detection through post-processing for PDF, contour, and heatmap generations, as well as a custom quantification for in-situ quantification, is deposited on our GitHub.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
A minimal dataset of the raw imaging generated in the course of this study has been deposited on CaltechData, under accession code: https://doi.org/10.22002/3jvmr-1b914. The entire large dataset is available upon request, because the size of these data is too large to be stored permanently on a public forum. Therefore, access can be provided by the corresponding author upon reasonable request. The minimal raw dataset provided is adequate for generating a small-scale replica of the results shown in the paper. The processed data containing all the distance measurements that were utilized to generate Figs. 3–7 are provided in the Source Data File. Source data are provided with this paper.
Code availability
Custom code was used for analysis and interpretation of triplets and doublets detected in this study. Custom code is open access using MIT license and can be accessible here: https://github.com/StathopoulosLab/find_spots Or, through https://doi.org/10.5281/zenodo.17945872 In the repository, there is a README file with instructions to run the spot detection program and relevant post-processing scripts to analyze the spots detection outputs. Spot detection is a Python GUI that can be pulled down for immediate usage. Output post-processing scripts are contained in the post_processing_tools folder which contains instructions for each relevant script utilized in this study. Segmentation and quantification of RNA transcripts is a MATLAB GUI with instructions to download and utilize the custom MATLAB quantification in the in-situ quantification folder.
References
Hug, C. B., Grimaldi, A. G., Kruse, K. & Vaquerizas, J. M. Chromatin Architecture Emerges During Zygotic Genome Activation Independent Of Transcription. Cell 169, 216–228.e19 (2017).
Nora, E. P., Dekker, J. & Heard, E. Segmental folding of chromosomes: a basis for structural and regulatory chromosomal neighborhoods? Bioessays 35, 818–828 (2013).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Dekker, J. The three ‘C’ s of chromosome conformation capture: controls, controls, controls. Nat. Methods 3, 17–21 (2006). 1/.
Bemer, M. & Baroux, C. Plant Chromatin Dynamics: Methods and Protocols. vol. 1675 (Springer New York, New York, NY, 2018).
Lieberman-Aiden, E. et al. Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 326, 285–289 (2009).
Li, X. et al. GAGA-associated factor fosters loop formation in the Drosophila genome. Mol. Cell 83, 1519–1526.e4 (2023).
Ing-Simmons, E. et al. Independence of chromatin conformation and gene regulation during Drosophila dorsoventral patterning. Nat. Genet. 53, 487–499 (2021). 04/.
Bing, X. et al. Chromosome structure in Drosophila is determined by boundary pairing not loop extrusion. Elife 13, RP94070 (2024).
Ke, W., Fujioka, M., Schedl, P. & Jaynes, J. B. Stem-loop and circle-loop TADs generated by directional pairing of boundary elements have distinct physical and regulatory properties. Elife 13, RP94114 (2024).
Miron, E. et al. Chromatin arranges in chains of mesoscale domains with nanoscale functional topography independent of cohesin. Sci. Adv. eaba8811 (2020).
Cardozo Gizzi, A. M. et al. Microscopy-based chromosome conformation capture enables simultaneous visualization of genome organization and transcription in intact organisms. Mol. Cell 74, 212–222.e5 (2019).
Fiche, J.-B. et al. Hi-M: A Multiplex Oligopaint FISH Method to Capture Chromatin Conformations In Situ and Accompanying Open-Source Acquisition Software. Methods Mol Biol 2784, 227–257 (2024).
Mateo, L. J., Sinnott-Armstrong, N. & Boettiger, A. N. Tracing DNA paths and RNA profiles in cultured cells and tissues with ORCA. Nat Protoc 16, 1647–1713 (2021).
Mateo, L. J. et al. Visualizing DNA folding and RNA in embryos at single-cell resolution. Nature 568, 49–54 (2019).
Dunipace, L., Saunders, A., Ashe, H. L. & Stathopoulos, A. Autoregulatory Feedback Controls Sequential Action of cis-Regulatory Modules at the brinker Locus. Dev. Cell 26, 536–543 (2013).
Samata, M. et al. Intergenerationally maintained histone H4 lysine 16 acetylation is instructive for future gene activation. Cell 182, 127–144.e23 (2020).
Foe, V. & Alberts, B. Studies of nuclear and cytoplasmic behaviour during the five mitotic cycles that precede gastrulation in Drosophila embryogenesis. J. Cell Sci. 61, 31–70 (1983).
Stathopoulos, A. & Newcomb, S. Setting up for gastrulation: D. melanogaster. in Current Topics in Developmental Biology vol. 136 3–32 (Elsevier, 2020).
Hong, J.-W., Hendrix, D. A. & Levine, M. S. Shadow enhancers as a source of evolutionary novelty. Science 321, 1314 (2008).
Markstein, M. et al. A regulatory code for neurogenic gene expression in the Drosophila embryo. Development 131, 2387–2394 (2004).
Ozdemir, A. et al. High resolution mapping of Twist to DNA in Drosophila embryos: Efficient functional analysis and evolutionary conservation. Genome Res 21, 566–577 (2011).
Dunipace, L., Ákos, Z. & Stathopoulos, A. Coacting enhancers can have complementary functions within gene regulatory networks and promote canalization. PLoS Genet 15, e1008525 (2019).
Nagarkar-Jaiswal, S. et al. A library of MiMICs allows tagging of genes and reversible, spatial and temporal knockdown of proteins in Drosophila. Elife 4, 10.7554/eLife.05338 (2015).
Venken, K. J. T. et al. MiMIC: a highly versatile transposon insertion resource for engineering Drosophila melanogaster genes. Nat. Methods 8, 737–743 (2011).
Wang, X. & Yue, F. Hijacked enhancer-promoter and silencer-promoter loops in cancer. Curr. Opin. Genet. Dev. 86, 102199 (2024).
Chopra, V. S., Kong, N. & Levine, M. Transcriptional repression via antilooping in the Drosophila embryo. Proc. Natl. Acad. Sci. USA. 109, 9460–9464 (2012).
Fujioka, M., Wu, X. & Jaynes, J. B. A chromatin insulator mediates transgene homing and very long-range enhancer-promoter communication. Development 136, 3077–3087 (2009).
Calhoun, V. C., Stathopoulos, A. & Levine, M. Promoter-proximal tethering elements regulate enhancer-promoter specificity in the Drosophila Antennapedia complex. Proc. Natl. Acad. Sci. USA. 99, 9243–9247 (2002).
Bower, G. & Kvon, E. Z. Genetic factors mediating long-range enhancer-promoter communication in mammalian development. Curr. Opin. Genet. Dev. 90, 102282 (2025).
Friman, E. T., Flyamer, I. M., Marenduzzo, D., Boyle, S. & Bickmore, W. A. Ultra-long-range interactions between active regulatory elements. Genome Res 33, 1269–1283 (2023).
Beck, S. et al. Implications of CpG islands on chromosomal architectures and modes of global gene regulation. Nucleic Acids Res 46, 4382–4391 (2018).
Koromila, T. & Stathopoulos, A. Broadly expressed repressors integrate patterning across orthogonal axes in embryos. Proc. Natl. Acad. Sci. USA. 114, 8295–8300 (2017).
Sun, J., Zhang, C., Gao, F. & Stathopoulos, A. Single-cell transcriptomics illuminates regulatory steps driving anterior-posterior patterning of Drosophila embryonic mesoderm. Cell Rep 42, 113289 (2023).
Trcek, T., Lionnet, T., Shroff, H. & Lehmann, R. mRNA quantification using single-molecule FISH in Drosophila embryos. Nat. Protoc. 12, 1326–1348 (2017). 7/.
Passaro, M. et al. OligoMinerApp: a web-server application for the design of genome-scale oligonucleotide in situ hybridization probes through the flexible OligoMiner environment. Nucleic Acids Res 48, W332–W339 (2020).
Super-Resolution Microscopy: Methods and Protocols. vol. 1663 (Springer New York, New York, NY, 2017).
Nguyen, S. C. & Joyce, E. F. Programmable Chromosome Painting with Oligopaints. in Imaging Gene Expression (ed. Shav-Tal, Y.) vol. 2038 167–180 (Springer New York, New York, NY, 2019).
Huff, J. The Airyscan detector from ZEISS: confocal imaging with improved signal-to-noise ratio and super-resolution. Nat. Methods 12, i–ii (12/2015).
Huff, J. et al. The new 2D Superresolution mode for ZEISS Airyscan. Nat. Methods 14, 1223–1223 (2017).
Lopes, R. H. C., Reid, I. & Hobson, P. R. The two-dimensional Kolmogorov-Smirnov test.
Heer, J. Fast & accurate Gaussian kernel density estimation. in 2021 IEEE Visualization Conference (VIS) (IEEE, 2021). https://doi.org/10.1109/vis49827.2021.9623323.
Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Simes, R. J. An improved Bonferroni procedure for multiple tests of significance. Biometrika 73, 751–754 (1986).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 145, (2018).
Dunipace, L., McGehee, J. M., Irizarry, J. & Stathopoulos, A. The proximal enhancer of the snail gene mediates negative autoregulatory feedback in Drosophila melanogaster. Genetics https://doi.org/10.1093/genetics/iyaf058 (2025).
Stepanik, V., Sun, J. & Stathopoulos, A. FGF pyramus has a transmembrane domain and cell-autonomous function in polarity. Curr Biol 30, 3141–3153.e5 (2020).
Le, M. T., McGehee, J., Dunipace, L., Rumph, D., Stathopoulos, A. Inferring chromatin architecture at a single locus through probabilistic in situ DNA localization. Zenodo, https://doi.org/10.5281/zenodo.17945872 (2026).
Sandler, J. E. & Stathopoulos, A. Quantitative single-embryo profile of drosophila genome activation and the dorsal-ventral patterning network. Genetics 202, 1575–1584 (2016).
Acknowledgements
We thank the Caltech Biological Imaging Facility for the use of the Zeiss 980 microscope, as well as Andres Collazo and Giada Spigolon for their assistance with microscope training. We are also grateful to the Stathopoulos lab members for helpful discussions and technical support, and to Galen Gao and Alexa Cunha for the initial code iterations.
Author information
Authors and Affiliations
Contributions
A.S. and L.D. conceived the project. M.T.L. and L.D. planned the experimental approach. A.S. directed the project. M.T.L. performed all experiments and analyzed the data with input from J.M. and D.R. The manuscript was written by M.T.L., L.D., J.M., and A.S. with input from D.R. This work was supported by the National Institutes of Health [R01HD106799 and R35GM118146 to A.S.] and Schmidt Sciences to D.R. and A.S. Funding for open access charge: National Institutes of Health.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Le, M.T., McGehee, J., Dunipace, L. et al. Inferring chromatin architecture at a single locus through probabilistic in situ DNA localization. Nat Commun 17, 1752 (2026). https://doi.org/10.1038/s41467-026-68460-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68460-x