Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) continues to cause significant morbidity and mortality despite the end of its pandemic phase. The emergence of highly mutated SARS-CoV-2 variants of concern highlights the requirement of broad-spectrum antiviral countermeasures which possess both prophylactic and therapeutic efficacies. Here, we obtain a macrocyclic peptide, 6L3-3P11K, that effectively inhibits a wide range of SARS-CoV-2 variants and subvariants. Structural studies show that 6L3-3P11K forms homotrimers that lock the spike protein (S) trimer into a “closed” conformation by engaging a conserved non-receptor binding motif (non-RBM) of S. This interaction disrupts the binding between S and ACE2 receptor. Structure-guided modifications result in a thermostable and trypsin-resistant macrocyclic peptide, 6L3-1F3P11hR, that exhibits prophylactic and therapeutic effects against SARS-CoV-2 infection in a male hACE2 transgenic mouse model after intranasal administration. Our results provide a drug candidate for the control and prevention of COVID-19 and may stimulate further research on macrocyclic broad-spectrum anti-coronavirus drug development.
Similar content being viewed by others
Introduction
Coronavirus disease (COVID-19) is an ongoing infectious disease caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Besides the significant morbidity and mortality associated with acute infection, especially among elderly and immunocompromised patients, post-COVID-19 condition has also been recognized1. Previous studies have shown that use of the SARS-CoV-2 antivirals in the acute phase may reduce the risk of post-COVID-19 condition2,3.
One of the major challenges in combating COVID-19 is the emergence of SARS-CoV-2 variants and subvariants that are immunoevasive to previously developed vaccines and monoclonal antibodies, such as the Omicron variant and subvariants4,5. While the two main types of clinically approved antiviral drugs for COVID-19, namely, RNA-dependent RNA polymerase (RdRp) inhibitors and main protease (Mpro) inhibitors, remain effective against Omicron6, the widespread use of these drugs that target a single viral enzyme may eventually contribute to the emergence of resistant strains7. Recent clinical studies have shown that, although Paxlovid (an Mpro inhibitor) can prevent the progression of COVID-19 from mild symptoms to severe diseases, it is not associated with a significantly shorter time to sustained alleviation of COVID-19 symptoms than placebo8. Moreover, these systemically administered drugs may be associated with major drug interactions or side effects. Therefore, novel antiviral drugs with high potency, distinctive mechanisms of action, and preferably topical administration to reduce systemic side effects are urgently needed.
Over the past two decades, macrocyclic peptides have garnered significant attention as a promising therapeutic strategy due to their uniquely advantageous properties9,10,11. These include high binding affinity and target specificity, low risk of off-target effects, minimal metabolic toxicity, strong tissue penetration, ease of manufacturing, metabolic stability, and the ability to disrupt protein-protein interactions. Moreover, these attributes can be further enhanced through strategic modifications.
Here, we employed the RaPID system12,13,14,15 to conduct de novo screening of macrocyclic peptides targeting the conserved region of the receptor-binding domain (RBD) of SARS-CoV-2. We discovered several promising macrocyclic peptides with strong binding affinities to the RBD. Structural studies demonstrated that the peptides bind to a conserved non-receptor-binding motif (non-RBM) region of RBD, and can form a homotrimer to lock the S trimer into a “closed” conformation, which prevents binding with ACE2. Further studies revealed the efficacy of these macrocyclic peptides to inhibit SARS-CoV-2 infection in vitro and in vivo. Our research has opened a promising avenue for the development of macrocyclic peptides as versatile agents, with potential efficacy against a wide range of SARS-CoV-2 variants and subvariants.
Results
Screening of macrocyclic peptides against the SARS-CoV-2 RBD
To identify macrocyclic peptides capable of binding to the RBD and potentially inhibiting the activity of SARS-CoV-2, we employed the RaPID system and conducted selections targeting the Omicron BA.2 RBD (Supplementary Fig. 1a). To increase the success rate of selection of antiviral peptides, we used the previously identified RBD-targeting macrocyclic peptide (peptide 4), which lacks antiviral activity, to block the site that could lead to the emergence of non-inhibitory peptides16. The specific binders were subsequently recovered, and their cDNAs were amplified via polymerase chain reaction (PCR) to provide input for the subsequent selection round. After five or six rounds of selection, we analyzed the enriched cDNA pools from the libraries using DNA deep sequencing techniques (Supplementary Fig. 1b). After analyzing the hit frequency, we proceeded to select the top 20 macrocyclic peptide sequences from each library and conducted alignments as shown in Supplementary Fig. 1c.
To validate their binding properties and investigate their antiviral activities, we employed the solid-phase peptide synthesis (SPPS) technique to chemically synthesize three representative sequences, 6L1, 6L3, and 5D1, originating from distinct families (Supplementary Fig. 1c). Subsequently, we conducted in vitro experiments to evaluate their antiviral potency against VSV particles pseudotyped with the BA.2 SARS-CoV-2 spike protein. Only 6L3 displayed weak inhibition activity, and both 5D1 and 6L1 exhibited no inhibition activity against the BA.2 pseudovirus (Supplementary Fig. 2a).
Using 6L3 as a starting point, we endeavored to introduce amino acid mutations to enhance its antiviral efficacy. The sequencing results showed that the similar peptide sequences to 6L3 mainly exhibit polymorphism at the 3rd, 6th, and 11th positions. Therefore, we selected these peptide mutants (L3P, T11K, L3PT11K, L3PT11I, L3PT11F, and L3PT6IP7Q) to evaluate the antiviral activities. Notably, 6L3-3P exhibited a significant improvement in antiviral effect, displaying an IC50 (half maximal inhibitory concentration) value of 8 nM (Supplementary Fig. 2b). Additionally, 6L3-11K also demonstrated enhanced antiviral ability, with an IC50 value of 676 nM. With both L3P and T11K substitutions, 6L3-3P11K also displayed strong antiviral activity, with an IC50 of 12 nM. In contrast, other mutant peptides did not possess antiviral activities. These findings underscore the pivotal roles played by L3P and T11K substitutions in augmenting the antiviral activity of 6L3 (Supplementary Fig. 2b). Consequently, we selected 6L3-3P and 6L3-3P11K for further investigation (Supplementary Fig. 2c).
Broad-spectrum antiviral activities of 6L3-3P and 6L3-3P11K peptides
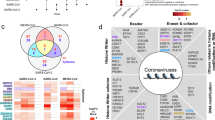
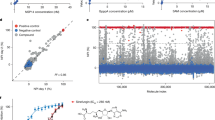
The remarkable efficacy demonstrated by 6L3-3P and 6L3-3P11K against the pseudotyped SARS-CoV-2 BA.2 variant encouraged us to further evaluate their potencies against other SARS-CoV-2 variants. We first conducted assessments of their binding affinities to different RBDs. Both 6L3-3P and 6L3-3P11K could effectively bind to all the tested SARS-CoV-2 RBDs, including Prototype (PT), Delta, and Omicron variants (BA.2, BF.7, BQ.1.1, XBB, XBB.1.5, and BA.2.86) (Fig. 1a, Supplementary Fig. 3a). These two peptides displayed particularly higher affinity to the Omicron variant RBDs, with KD values ranging from 2.36 to 21.7 nM. In contrast, the binding affinities of these two peptides to Prototype and Delta RBDs were decreased dramatically, with KD values spanning from 11.4 to 65.3 μM. Intriguingly, 6L3-3P and 6L3-3P11K also demonstrated binding ability to RBDs of SARS-CoV-related viruses, with KD values ranging from 19.5 to 95.9 μM. Our results indicated that these two macrocyclic peptides may be potential broad-spectrum pan-sarbecovirus antiviral candidates.
a The binding affinities of 6L3-3P and 6L3-3P11K peptides to RBDs from different SARS-CoV-2 variants are evaluated using a BIAcore 8K system in single-cycle mode. Affinity values summarized are the means of two independent experiments. b Inhibition of 6L3-3P and 6L3-3P11K peptides against different SARS-CoV-2 variant pseudotyped viruses in Vero cells. The two groups are distinguished by color, and the color depth correlates with the inhibitory activity. The experiments were performed at least twice with two replicates (n = 2). See also Supplementary Fig. 3.
We further conducted inhibition assays of these two peptides on pseudotyped SARS-CoV-2 Prototype and VOCs (Alpha, Beta, Gamma, Delta, and Omicron subvariants BA.1, BA.2.75, BA.4/5, BA.4.6, BF.7, BQ.1, BQ.1.1, XBB, XBB.1.5, XBB.1.16, CH.1.1, BA.2.86 and EG.5.1). Consistent with the binding results, both 6L3-3P and 6L3-3P11K demonstrated broad-spectrum antiviral activities, particularly against the Omicron subvariants, with IC50 values ranging from 3 to 86 nM (Fig. 1b, Supplementary Fig. 3b). In contrast, against the Prototype and early VOCs, including Alpha, Beta, Gamma, Delta, and Omicron subvariant BA.1, these two peptides displayed relatively weaker neutralizing abilities, with IC50 values ranging from 452 to 6933 nM (Fig. 1b, Supplementary Fig. 3b). Collectively, these findings indicated that both 6L3-3P and 6L3-3P11K exhibit broad-spectrum antiviral activities against emerging SARS-CoV-2 variants and subvariants.
Inhibition mechanism of 6L3-3P11K
To elucidate the molecular mechanism of how 6L3-related macrocyclic peptides inhibit virus infection, we prepared the S proteins of SARS-CoV-2 Prototype (PT S) and BA.2.75 (BA.2.75 S), and then incubated them with the 6L3-3P11K peptide to obtain the S-peptide complexes. Subsequently, we resolved the cryo-EM structures of PT S/6L3-3P11K and BA.2.75 S/6L3-3P11K at resolutions of 2.6 and 3.1 Å, respectively (Supplementary Figs. 4 and 5, Supplementary Table 1). Interestingly, the binding of 6L3-3P11K makes the PT S trimers keep a “closed” conformation with all three RBD down. Within this “closed” S trimer, three 6L3-3P11K molecules bound together in the top cavity of S trimer, forming a homotrimer through hydrophobic interactions and van der Waals (VDW) forces. Particularly, the amino acids 7P and 1Y of 6L3-3P11K played a crucial role in these trimerization interactions (Fig. 2a, c). A similar “closed” conformation of S trimer was also observed in the BA.2.75 S/6L3-3P11K complex structure, with 6L3-3P11K molecules occupying the same binding cavity on top of S trimer (Fig. 2b, d). The “up” or “down” conformations of the RBDs are inherent dynamic features for the spike protein, and the peptide binding to RBDs did not induce the “closed” conformation. When the RBDs move to the “down” conformation, the trimerization of the peptides can keep the “closed” conformation of the RBDs on the spike protein and prevent the occurrence of the “up” conformation of the RBDs, which is required for ACE2 receptor binding. This hypothesis was further confirmed through competitive binding experiments using the bio-layer interferometry (BLI) method. As anticipated, the BA.2.75 S trimer displayed a strong binding capacity to ACE2. When we incubated the BA.2.75 S protein with the 6L3-3P or 6L3-3P11K for 10 min, the S protein exhibited a significant reduction in the binding capacity. As a negative control, even incubated with 6L3 peptide for 2 h, the S protein still effectively bound to ACE2 (Supplementary Fig. 6). Native SEC-mass spectrometry analysis revealed that 6L3-3P11K exists exclusively as a monomer in the absence of the spike protein binding (Supplementary Fig. 7), and 6L3-3P11K trimerization occurs specifically upon S protein binding. We also analyzed the cryo-EM density of the PT S and BA.2.75 S in the apo state, and performed the quantitative analysis for the unbound spike trimer. The results showed that the proportion of spike trimers in the three-RBD-down conformation increased notably after peptide binding, from 6.6 to ~100% for PT and from 38.3% to 54% for BA.2.75 (Supplementary Figs. 4, 5 and 8). The cryo-EM density of the BA.2.75 S/6L3 complex was also analyzed, showing that most of the S trimers displayed either “three-RBD-up” or “two-RBD-up” conformation (Supplementary Fig. 9). This binding feature corroborates the weaker inhibition activity of 6L3.
The overall complex structures of 6L3-3P11K bound to the SARS-CoV-2 Prototype S trimer (a) and Omicron BA.2.75 S trimer (b), respectively. Each complex shows as a top view and a side view. c Interactions between three macrocyclic peptides in the complex structure of 6L3-3P11K and Prototype S trimer. The van der Waals forces among 1Y, 3P, 4I, and 6T 7P promote the formation of 6L3-3P11K “trimer”. The three peptides are labeled with yellow, purple, and green, respectively. The 6L3-3P11K trimer showed as the top, down, and side views. d Interactions between three macrocyclic peptides in the complex structure of 6L3-3P11K and Omicron BA.2.75 S trimer. The van der Waals forces among 1Y, 3P, 4I, and 6T promote the formation of 6L3-3P11K “trimer”. The three peptides are labeled with pink, green, and blue, respectively. The 6L3-3P11K trimer showed as the top, down, and side views. Single-letter abbreviations for the amino acid residues are as follows: I, Ile; P, Pro; T, Thr; and Y, ClAc-L-Tyr.
The detailed analyses of the interactions between three 6L3-3P11K molecules and three RBDs in the S trimer revealed that each 6L3-3P11K binds to the inner side of one major RBD and also has minor interaction with the adjacent RBD (Fig. 3a, e and Supplementary Fig. 10). Notably, within the major binding pocket, residues 4I, 5L, 8L and 10F of 6L3-3P11K interact with a hydrophobic region in the RBD, comprised of residues F377, G404, V407, V433, I434 and V503 (Fig. 3b, f). This hydrophobic pocket is highly conserved in the RBD of the SARS-CoV-2 S protein. In addition, in both PT S/6L3-3P11K and BA.2.75 S/6L3-3P11K complex structures, residues two-dimensional (2D), 11K, and 13G-NH2 in the peptide form the hydrogen bond interactions with the RBD residues (Fig. 3c, g).
The footprints of 6L3-3P11K on the RBD of SARS-CoV-2 PT (a) and Omicron BA.2.75 (e) are shown in green and mauve, respectively. The RBD protein is depicted in surface representation. b, f The hydrophobic interactions between 6L3-3P11K and RBDs. c, g The hydrogen bond interactions between 6L3-3P11K and RBDs. The RBD protein is depicted in cartoon representation, and the amino acids of RBD involved in the interaction with 6L3-3P11K are displayed as sticks and darkened in color. Hydrogen bonds are displayed as a red dashed line. d Comparison between structures of S/6L3-3P11K RBD (gray) and SARS-CoV-2 PT RBD (PDB: 6VXX, light blue). The F374-K378 loop of RBD in the S/6L3-3P11K complex exhibited a noteworthy conformational change. h Comparison between structures of BA.2.75 S/6L3-3P11K RBD (orange) and SARS-CoV-2 Omicron BA.2.75 RBD (PDB: 7YQU, light green). The F374-K378 loop in the two RBDs showed minimal conformational changes. Single-letter abbreviations for the amino acid residues are as follows: D, Asp; G, Gly; F, Phe; I, Ile; K, Lys; L, Leu; P, Pro; N, Asn; S, Ser; T, Thr; V, Val; and Y, Tyr.
Of note, the 370-loop region (residues 373-377) of RBD in the PT S/6L3-3P11K complex structure exhibited a noteworthy conformational change upon peptide binding (Fig. 3d and Supplementary Fig. 11). In contrast, the BA.2.75 S/6L3-3P11K structure showed minimal conformational alteration (Fig. 3h and Supplementary Fig. 11). This difference may correlate with the relatively lower inhibition activity of 6L3-3P11K against SARS-CoV-2 strains including Prototype and early VOCs, which might result from different conformations of 370-loop (residues 373-377), and also differential residues at positions 375 and 376 (changed from hydrophilic S375 and T376 to hydrophobic F375 and A376) and position 408 (changed from charged R408 to non-charged S408) of RBD in PT and BA.2.75 virus variants (Supplementary Fig. 12).
Collectively, the inhibition of 6L3-3P11K refers to two important steps: the first step is the peptide binding to the RBD, and the second step is the trimerization of the peptide when the RBDs move to the “down” state in the S trimer.
Structure-based modification of 6L3-3P11K peptide
To increase the antiviral activity of 6L3-3P11K peptide, we introduced an amino acid mutation to the peptide based on the molecular interaction revealed by structural analysis. It is worth noting that, as described above, in addition to its interaction with the major RBD, 6L3-3P11K also exhibited minor interactions with adjacent RBD through VDW forces in both PT S/6L3-3P11K and BA.2.75 S/6L3-3P11K complex structures (Supplementary Fig. 10). As depicted in Supplementary Fig. 13, the side chain of 11K of the peptide might be potentially involved in forming strong hydrogen bonds with the adjacent RBD. To acquire assumed hydrogen bonds, we substituted lysine (K) to arginine (R) that has a longer side chain (Fig. 4a). As expected, 6L3-3P11R demonstrated a more potent neutralizing ability in comparison to 6L3-3P11K (Fig. 4d). Given that peptides containing arginine are susceptible to degradation by trypsin-like proteases in human tissues, we tested the resistance of 6L3-3P11R to trypsin enzyme. The result demonstrated that the antiviral activity of 6L3-3P11R dramatically decreased after trypsin treatment, with an IC50 of 2731 nM against pseudotyped BF.7 SARS-CoV-2 (Fig. 4a). To mitigate this drawback, we then synthesized another modified peptide 6L3-3P11hR which has a homoarginine (hR) instead of the arginine (Fig. 4b). As expected, 6L3-3P11hR was resistant to trypsin treatment and retained potent antiviral activity (Fig. 4b, d).
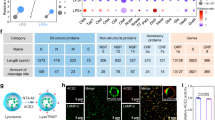
The chemical structure of 6L3-3P11R (a), 6L3-3P11hR (b), and 6L3-1F3P11hR (c) and the antiviral activities of these three peptides after being treated with trypsin against Omicron BF.7 pseudotyped virus in Vero cells. n = 2 per peptide concentration points. d Antiviral activities of 6L3-3P11R, 6L3-3P11hR, and 6L3-1F3P11hR against different SARS-CoV-2 variant pseudotyped viruses in Vero cells. The three groups are distinguished by color, and the color depth correlates with the inhibitory activity. n = 2 per peptide concentration point. e Primary human epithelial cells (pooled from three healthy donors) differentiated at the air-liquid interface (ALI) were fixed, permeabilized, and stained with specific antibodies: Acetyl-α-Tubulin (Lys40) (D20G3) (left panel, red), Goblet cells (left panel, gray), human ACE2 (right panel, red), and phalloidin (right panel, green). Nuclei were counterstained with DAPI (blue). Images were captured using a confocal laser scanning microscope and processed into 3D reconstructions via volume rendering using Imaris software. f Antiviral activity of 6L3-3P11hR and 6L3-1F3P11hR against Omicron BA.2.12.1 and JN.1 in primary human nasal epithelial cells (ALI). Viral infection was assessed by quantifying viral RNA in infected cells 24 h post-infection (n = 2). Data are presented as mean ± SD. One-way ANOVA was used to compare the control group. ****P < 0.0001.
As the trimerization capacity of the peptide is important for the peptide inhibition, we substituted tyrosine (Y) with phenylalanine (F) at position 1 of the 6L3-3P11hR peptide to increase the hydrophobic trimerization interaction. Notably, 6L3-1F3P11hR peptide not only shows more potent antiviral activity against SARS-CoV-2 Omicron variants, but also acquires much better antiviral activity against the Prototype and Delta variants (Fig. 4c, d). The live virus inhibition assay further confirmed that 6L3-1F3P11hR could efficiently inhibit SARS-CoV-2 replication, including early strains (Prototype and Alpha) and recent Omicron strains (BA.2.12.1 and EG.5.1), with significant viral RNA reduction in supernatants observed even at 10 nM (Supplementary Fig. 14).
Inhibition efficacy of 6L3-1F3P11hR in the human nasal organoid model
Previous studies revealed that Omicron subvariants display growth advantages in human nasal tissues17. Therefore, we further used human nasal organoids to test the antiviral activity of two peptides. To recreate the three-dimensional, pseudostratified architecture of nasal epithelia in vivo, primary human nasal epithelial cells were pooled from three healthy donors and differentiated at the air-liquid interface (ALI). Differentiation status was confirmed by stratification of nuclei, the presence of mature cilia at the tissue surface, and goblet cells (Fig. 4e, left). The expression of ACE2 in the ALI was also analyzed (Fig. 4e, right). The results showed that both 6L3-3P11hR and 6L3-1F3P11hR markedly inhibited the replication of BA.2.12.1 and JN.1 viruses in the ALI culture. Notably, similar levels of inhibition were observed at both 10 μM and 100 μM concentrations, indicating that the peptides can effectively inhibit viral replication even at lower concentrations (Fig. 4f).
Chemical stability and pharmacokinetic characteristics of 6L3-1F3P11hR
We also evaluated the stability of the 6L3-1F3P11hR, including temperature (45 °C) sensitivity, pH (pH 5.5–10.0) sensitivity, and mouse serum stability. The results showed that 6L3-1F3P11hR did not show significant alteration in chemical structure after exposure to 45 °C for weeks or acidic/alkaline conditions (pH 5.5-pH 10.0) (Supplementary Fig. 15a, b). Over 80% of the original drug remained by 24 h after coincubation in mouse serum at 37 °C (Supplementary Fig. 15c). These results indicate that 6L3-1F3P11hR exhibited a good chemical stability. We also performed additional NMR experiments and obtained the 1H-NMR spectrum of 6L3-1F3P11hR. The results are consistent with the chemical structure of 6L3-1F3P11hR (Please see the end of Supplementary information).
We then explored the in vivo pharmacokinetic characteristics of 6L3-1F3P11hR by intranasal administration in C57BL/6 mice (Supplementary Fig. 15d). The results showed that the peptide was mainly accumulated in the nasal region, and the peak concentration of the peptide can achieve 1529 ng/ml in the nasal lavage fluid. The exposure (AUC0-t) over 24 h was 4603 h*ng/ml, with a half-life (T1/2) of 6.45 h in the nasal lavage fluid. In contrast, the peptide concentration in the plasma was very low, with a peak concentration of 5 ng/ml. The exposure (AUC0-t) over 8 h was 18 h*ng/ml, with a half-life (T1/2) of 3.08 h in the plasma.
6L3-1F3P11hR reduces viral burden and tissue damage in vivo
To further demonstrate the in vivo efficacy of the macrocyclic peptide, we first explored the therapeutic effect of 6L3-1F3P11hR in SARS-CoV-2-infected K18-hACE2 mice (Fig. 5). The peptide 6L3-1F3P11hR (3 mg/kg per dose) was intranasally given one dose at 30 min after virus challenge and then every 24 h for 3 more days. Nirmatrelvir (100 mg/kg per dose, intragastric administration) was used as the positive control (Fig. 5a). As shown in Fig. 5b, the viral RNA load and infectious virus titer in the lungs of the mice were significantly reduced after treatment with either 6L3-1F3P11hR or nirmatrelvir. Mice treated with either 6L3-1F3P11hR or nirmatrelvir exhibited significantly lower histopathological scores compared to solvent-treated controls (Fig. 5c)18. Histopathological changes seen in solvent-treated control mice, including alveolar inflammation and congestion, alveolar hemorrhage, bronchiole luminal cell debris, endothelium infiltrates, and perivascular or peribronchiolar infiltrates, were scarcely observed in mice that received 6L3-1F3P11hR or nirmatrelvir (Fig. 5d). We further tested the therapeutic efficacy of 6L3-1F3P11hR (6 mg/kg per dose) which was administrated at 8 h post virus challenge, followed by subsequent doses every 24 h for three additional days (Supplementary Fig. 16). The infectious virus titer in the lungs of the mice was also significantly reduced.
a Schematic of the in vivo therapeutic evaluation of 6L3-1F3P11hR against SARS-CoV-2. K18-hACE2 mice (n = 5 per group) were intranasally challenged with 4 × 103 plaque-forming units of SARS-CoV-2 Omicron BA.2 at day 0. 6L3-1F3P11hR (3 mg/kg per dose) was administered via the intranasal route. Control groups included solvent-treated mice and nirmatrelvir-treated mice (100 mg/kg per dose, intragastric administration) as negative and positive controls, respectively. Both the control and experimental groups received the drug at the same time points with equal frequency. The animal experimental scheme figure was created in BioRender. L, R. (2026) https://BioRender.com/28q57t5. b SARS-CoV-2 RNA load was determined by qRT-PCR assay, normalized with β-actin. Quantitation of virus titer was done by plaque assays. To enable statistical analysis, values under LOD were assigned 0.0001 for qRT-PCR and 100 for plaque assay, respectively. Five animals were used per group (n = 5). Data are presented as mean ± SD. One-way ANOVA was used to compare the groups with the solvent-treated control. **P < 0.01, ***P < 0.001 and ****P < 0.0001. c Histopathology scores of the different treatment groups of mice. Five animals were used per group (n = 5). Data are presented as mean ± SD. One-way ANOVA was used to compare the groups with the solvent-treated control. *P < 0.1 and ***P < 0.001. d Representative images of hematoxylin and eosin-stained mouse lung sections at 4 dpi. The lung sections of the solvent-treated control mice showed prominent alveolar inflammation and congestion (asterisk), alveolar hemorrhage (black triangle), bronchiole luminal cell debris (black double-headed arrow), endothelium infiltrates (black arrows), perivascular or peribronchiolar infiltrates (blue arrows), and abundant viral nucleocapsid antigen (green) expression. In contrast, the lungs of mice treated with nirmatrelvir and the peptide, both prophylactically and therapeutically, demonstrated much alleviated histopathological changes and scarce viral nucleocapsid antigen expression. The nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). Shown are representative images randomly selected from a pool of images for each group (n = 5 mice/group).
We then explored the prophylactic efficacy of 6L3-1F3P11hR. The antiviral effect of 6L3-1F3P11hR was more significant when given prophylactically (3 mg/kg one dose at 30 min prior to virus challenge) (Fig. 6). The SARS-CoV-2 RNA load and infectious virus titer in the nasal turbinate and lung were significantly reduced (Fig. 6b, d). The mice treated with 6L3-1F3P11hR displayed much lower histopathological scores compared to the untreated mice (Fig. 6c). Histopathological changes of the lung were also obviously alleviated in the peptide-treated group (Fig. 6e).
a Schematic of the in vivo prophylactic evaluation of 6L3-1F3P11hR against SARS-CoV-2. Prior to intranasal challenge with SARS-CoV-2 Omicron BA.2 (4 × 103 PFU/mouse) on day 0, K18-hACE2 mice (n = 5 per group) received prophylactic intranasal administration of 6L3-1F3P11hR (3 mg/kg) or intragastric administration of nirmatrelvir (100 mg/kg) at −0.5 h. The animal experimental scheme figure was created in BioRender. L, R. (2026) https://BioRender.com/28q57t5. b SARS-CoV-2 RNA load was determined by qRT-PCR assay, normalized with β-actin. Five animals were used per group (n = 5). Data are presented as mean ± SD. One-way ANOVA was used to compare the groups with the solvent-treated control. *P < 0.1 and **P < 0.01. c Histopathology scores of the control and experimental groups of mice. Five animals were used per group (n = 5). Data are presented as mean ± SD. One-way ANOVA was used to compare the groups with the solvent-treated control. ****P < 0.0001. d Quantitation of virus titer was done by plaque assays. To enable statistical analysis, values under LOD were assigned 0.0001 for qRT-PCR and 100 for plaque assay, respectively. Five animals were used per group (n = 5). Data are presented as mean ± SD. One-way ANOVA was used to compare the groups with the solvent-treated control. **P < 0.01 and ****P < 0.0001. e Representative images of hematoxylin and eosin-stained mouse lung sections at 4 dpi. The lungs of mice prophylactically treated with nirmatrelvir and the peptide showed much alleviated histopathological changes and scarce viral nucleocapsid antigen expression. Alveolar inflammation and congestion (black asterisk), alveolar hemorrhage (black triangle), endothelium infiltrates (black arrows), perivascular or peribronchiolar infiltrates (blue arrows), and abundant viral nucleocapsid antigen (green) expression. The nuclei were stained with DAPI. Shown are representative images randomly selected from a pool of images for each group (n = 5 mice/group).
Discussion
Antivirals that could be readily and topically administered as intranasal agents that are effective against existing and emerging SARS-CoV-2 subvariants are an important component of the armamentarium against COVID-19. Apart from RdRp and Mpro, the S protein is another important antiviral target, given its critical role in virus-host interactions19. However, due to the high production cost and low-temperature storage requirements, worldwide distribution of antibody-based drugs has remained difficult. Herein, we have made a discovery of several potent and broad-spectrum macrocyclic peptides targeting RBD. In particular, the 6L3-1F3P11hR has displayed both prophylactic and therapeutic efficacies in vitro and in vivo. Moreover, the macrocyclic peptides can be easily synthesized and exhibit good chemical stability. These advantages make them potential prophylactic and therapeutic options for existing and emerging SARS-CoV-2 subvariants.
Unlike other inhibitors20,21,22,23,24,25,26,27, 6L3-related peptides showed a completely different two-step antiviral mechanism through locking of the S trimer into a “closed” conformation by acting as “molecular glue”, and thereby disrupting the binding between RBD and ACE2. The antiviral activity of the peptide correlates to two key points: the peptide binding to RBD and the peptide trimerization within the closed conformation of spike protein. Although the peptides have highly similar binding affinities against different virus variants (e.g., against XBB vs BA2.86), the antiviral activities could differ due to the different dynamic features of RBDs. These “up and down” kinetic differences of RBDs likely contribute to the different antiviral capacities. In addition, the binding site of 6L3-related peptides locates on the non-RBM region of RBD and is highly conserved among the SARS-CoV-2 omicron variants (Supplementary Fig. 12). We have also examined the emerging KP.2, KP.3, LB.1, NB.1.8.1, LP.1.8, XDV and XFV variants, and there are no extra amino acid substitutions in the peptide binding site of RBD. It indicates that the binding site of 6L3-related peptides is not the hot region for the human neutralizing antibodies, and the 6L3-related peptides will remain effective when the persistent mutations of SARS-CoV-2 arise under the immune pressure.
Recently, another macrocyclic peptide (S1b3inL1) with broad-spectrum antiviral activity that interacts with a recessed region of RBD in the spike protein was reported28. However, there is no further data to show that the S1b3inL1 peptide would be effective in vivo, and the inhibition mechanism was not confirmed. In contrast, by further structure-based modifications, our results demonstrated that the 6L3-1F3P11hR peptide is effective both in vitro (human nose organoid) and in vivo (mouse model). Therefore, 6L3-1F3P11hR is a promising macrocyclic peptide for further evaluation in clinical trials.
Multimerization and lipidation have been widely used to improve the antiviral efficiency of peptide drugs28,29,30. In our study, point mutations within the peptide can significantly improve the antiviral activities of 6L3 without further multimerization or lipidation. Firstly, L3P substitution in the 6L3 peptide led to a tremendous augmentation in its antiviral efficacy. The introduction of proline residue into the peptide changes the conformational flexibility, owing to its distinctive cyclic side chain structure, and then greatly increase the stability of trimerization of the peptide, which is essential for the locking of S trimer into a “closed” conformation. Secondly, T11K/R substitution increases the inhibition activity of 6L3-3P by enhancing the interaction with adjacent RBDs, with little influence on the binding affinity to the RBD protomer. In addition, R11hR substitution also contributes to the resistance to enzyme cleavage. Thirdly, Y1F substitution further improves the antiviral activity by increasing the interaction among the homotrimers.
To further develop the effective macrocyclic peptide drug against the sarbecoviruses, including SARS-CoV-1 and MERS-CoV, with a similar “molecular glue” mechanism, we can perform point mutations in the 6L3-related peptides according to the features of residues in the similar binding site of RBDs from other sarbecoviruses. The design of point mutations can be performed based on the S-peptide complex structure solved here31, or we can generate a new secondary peptide library with the key residues and use the RaPID system to screen the potentially effective peptides. In addition, the pharmacokinetic data of 6L3-1F3P11hR peptide by intranasal administration have revealed that the peptides were mainly accumulated in the nasal region, and very few peptides have entered the blood. In the future, we would explore the antiviral potential of the peptide by different administration routes such as inhalable dry powder and oral formulation.
In summary, our findings reveal the discovery of macrocyclic peptides with remarkable capability to inhibit infection across a wide spectrum of SARS-CoV-2 variants and subvariants. This inhibition was achieved by acting on a previously undisclosed binding pocket with high sequence conservation through a “molecular glue” mechanism. Ongoing efforts are dedicated to the transformation of these macrocyclic peptides into clinically available anti-coronavirus drugs.
Methods
Cells
HEK293T cells (ATCC CRL-3216), Vero cells (ATCC CCL81), Vero E6-TMPRSS2 cells (JCRB Cat#JCRB1819), and BHK-21 cells (ATCC CCL-10) were grown at 37 °C in a humidified incubator with 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal calf serum (FBS), 1‰ penicillin and 1‰ streptomycin. Freestyle 293F cells were cultured in SMM 293-TII medium at 37 °C in a shaker with 5% CO2.
Viruses
SARS-CoV-2 isolates hCoV-19/USA/NY-MSHSPSP-PV56475/2022 (Lineage BA.2.12.1; Omicron BA.2 variant; NR-56782) and hCoV-19/USA/MSHSPSP-PV96109/2023 (Lineage JN.1, Omicron variant; NR-59694) were obtained from BEI Resources. SARS-CoV-2 Omicron subvariant BA.1.529.2/BA.2 (GISAID: EPI_ISL_9845731) was used in the animal experiments32. SARS-CoV-2 isolates PT HKU-001a (GenBank: MT230904), B.1.1.7/Alpha (GenBank: OM212469), BA.2.12.1 (GISAID: EPI_ISL_13777659), BA.5.2 (GISAID: EPI_ISL_13777658), EG.5.1 (GISAID: EPI_ISL_18461518) were used in the live virus inhibition assay.
Ethics statement
All procedures in this study involving SARS-CoV-2 infection were performed in the biosafety level 3 (BSL-3) and animal biosafety level 3 (ABSL-3) facilities according to approved standard operating procedures. All the procedures in the animal studies were reviewed and approved by the Committee on the Use of Live Animals in Teaching and Research at The University of Hong Kong (CULATR 5370-20) or Laboratory Animal Welfare and Ethics Committee (PK-R-06012024).
Animals
The use of animals at HKU was approved by the HKU Committee on the Use of Live Animals in Teaching and Research. K18-hACE2 mice (aged 8–10 weeks, male) were obtained from the HKU Centre for Comparative Medicine Research. Six to eight-week-old male C57BL/6 mice were purchased from Vital River. The animals were kept in cages with individual ventilation with 65% humidity and ambient temperature ranging between 21 and 23 °C with a 12-h-interval day/night cycle for housing and husbandry before transferral to the Biosafety Level 3 facility.
Protein expression
The coding sequences (codon optimized) for the ectodomain of SARS-CoV-2 Prototype S and the Omicron BA.2.75 S subvariant (containing “6P” stabilizing mutations) were fused at their C-termini with a trimeric tag followed by a Strep-II tag. These sequences were cloned into the pcDNA3.1 expression vector. The coding sequences (codon optimized) of Omicron subvariant BA.2 RBD with C-terminal His-tag and Avi-tag were cloned into the pCAGGS vector. The recombinant plasmids were transfected into Freestyle 293F cells to express the RBD, S proteins. Five days later, the supernatants were collected, and soluble proteins were purified by Ni affinity chromatography using a HisTrap EXCEL 5-mL column (GE Healthcare). S protein was further purified via gel filtration chromatography with Superose 6 Increase 10/300 GL column (GE Healthcare) in a buffer composed of 20 mM Tris-Base and 150 mM NaCl (pH 8.0). RBD protein was purified via gel filtration chromatography with a Superdex 200 column (GE Healthcare) in 1 × phosphate-buffered saline (PBS). RBD protein was further biotinylated using a Biotin Labeling Kit (Beyotime); biotinylated RBD was used for macrocyclic peptides selection. The coding sequences of RBDs with human IgG Fc-tag (hFc) (codon optimized) were also cloned into the pCAGGS vector, including Prototype, Delta, Omicron subvariants (BA.2, BF.7, BQ.1.1, XBB, and XBB.1.5), as well as three sarbecoviruses (SARS-CoV, RaTG13, and GD/1/2019). The coding sequence of ACE2 with hFc (ACE2-hFc, codon optimized) was also cloned into the pCAGGS vector. The recombinant plasmids were transfected into mammalian 293T cells to express the RBD-hFc and ACE2-hFc. Five days later, the supernatants were collected and used in SPR or BLI assays.
Selection of macrocyclic peptides against RBD
Macrocyclic peptides that target the RBD were screened using the RaPID system as previously reported. Initially, the mRNA library was prepared with the sequence AUG-(NNK)8-12-(GGC-AGC)3-UAG and linked to a puromycin (DNA-PEG-CC-Pu) linker utilizing T4 RNA ligase. This process resulted in the creation of the mRNA-Pu library. The resultant mRNA-Pu library was then subjected to the RF1 (release factor 1) and Met (methionine)-deficient FIT (flexible in vitro translation) system, which contained 50 μM ClAc-L(or D)Tyr-tRNAfMetCAU, with a total volume of 25 μL for the initial selection round. Following a 2 h reaction at 37 °C, the solution was incubated at 25 °C for 15 min, followed by quenching through the addition of 5 μL of 100 mM EDTA (pH 8.0). The mixture was further incubated at 30 °C for 30 min. Subsequently, the library underwent reverse transcription by incubating the mixture at 42 °C with the addition of 0.25 mM dNTPs, 2 μM CGS3an13.R39, 25 mM Tris-HCl (pH 8.3), 15 mM Mg(OAc)2, 10 mM KOH, and 50 U M-MLV reverse transcriptase RNase H Minus. After 60 min, an equal volume of blocking solution was added, and the resulting mixture was employed for affinity selection against RBD-immobilized DynabeadsTM M-280 Streptavidin at 4 °C for 60 min in the presence of 20 μM peptide 4 (a blocking peptide). The beads were then washed three times with the selection buffer and subsequently heated at 95 °C for 5 min with 1×PCR mix to elute cDNAs. Quantification of the resulting cDNAs was performed using the QuantStudio™ real-time PCR system (Applied Biosystems), followed by PCR amplification and transcription into mRNAs to generate the library for the second round of selection. In the subsequent rounds, the translation scale was reduced to 5 μL, and the pre-clear steps were repeated multiple times by adding the resulting mixture to fresh beads to selectively remove the peptides that bound to the magnetic beads (negative selection). The supernatant from the final negative selection was incubated with RBD-immobilized DynabeadsTM M-280 Streptavidin in the presence of 20 μM peptide 4 to selectively bind cDNAs that interacted with the RBD (positive selection). This selection process was iterated for several rounds until a substantial enrichment of cDNAs binding to the RBD was observed. Finally, the enriched sequences were recovered and identified through deep sequencing utilizing the NovaSeq 6000 system (Illumina).
Chemical synthesis of macrocyclic peptides
The synthesis of macrocyclic peptides was conducted following the standard Fmoc SPPS protocol. Initially, 0.5 g of Rink Amide MBHA resin was treated with a 20% piperidine in dimethylformamide (DMF) solution to remove the Fmoc protecting group. Subsequently, a solution containing 0.3 mmol of Fmoc-Gly-OH (0.3 mmol of 1-Hydroxybenzotriazole (HOBT)), and 5% N, N’-diisopropylcarbodiimide (DIC) in 10 mL of DMF was added to the resin, and the mixture was incubated under nitrogen gas bubbling conditions for 1.5 h. The resin was thoroughly washed three times with DMF and dichloromethane (DCM) before deprotecting the Fmoc group using a 20% piperidine in DMF solution. To couple the subsequent amino acids, a freshly prepared solution of 0.9 mmol Fmoc-AA-OH, 0.9 mmol HOBT, and 9% DIC in 10 mL of DMF was employed, and the coupling reaction was conducted for 1 h under nitrogen gas bubbling conditions. These deprotection and coupling steps were repeated iteratively until the desired peptide length was achieved. The peptides were then conjugated to their free N-terminal α-amino group using the bromoacetyl group and subsequently cleaved from the resin. The resulting peptides were precipitated with diethyl ether (Et2O), re-dissolved in dimethylsulfoxide (DMSO), and subjected to thiol-ether cyclization by incubating the solution at pH 8.0 for 1 h. Finally, trifluoroacetic acid (TFA) was added to acidify the solution, and the peptides were purified using RP-HPLC with a mobile phase consisting of a linear gradient of 0.1% TFA aqueous solution and 0.1% TFA in acetonitrile (MeCN; mobile phase B). The characterization of synthesized macrocyclic peptides was conducted by employing ESI-MS, while their purity confirmation was achieved through analytical HPLC analysis.
Surface plasmon resonance (SPR) assay
The binding affinities of 6L3-3P or 6L3-3P11K with RBDs from SARS-CoV-2 and its variants (Delta, Omicron subvariants BA.2, BF.7, BQ.1.1, XBB, XBB.1.5, and BA.2.86), as well as three sarbecoviruses (SARS-CoV, RaTG13, and GD/1/2019), were evaluated by SPR analysis. The experiments were performed using a BIAcore8000 system (GE Healthcare) with protein A chips (Cytiva Life Sciences) at 25 °C in single-cycle mode. All proteins used for kinetic analysis were exchanged into the PBST buffer (2.7 mM KCl, 137 mM NaCl, 4.3 mM Na2HPO4, 1.4 mM KH2PO4, and 0.5‰(v/v) Tween 20) with 5% DMSO. The supernatants containing hFc-tagged RBD proteins were captured by the protein A chip at more than 3000 response units. Then, serial dilutions of 6L3-3P or 6L3-3P11K were injected over the chip surface to test binding. The ka, kd, and KD for each pair of interactions were calculated with BIAcore 8 K evaluation software (GE Healthcare).
Bio-layer interferometry assay
The supernatants containing ACE2-hFc proteins were captured by the protein A sensors. The BA.2.75 S trimer was incubated with either 6L3-3P or 6L3-3P11K peptides at a molar ratio of 1:5 for 10 min, and incubated with the 6L3 peptide for 2 h. The binding ability of BA.2.75 S or BA.2.75 S /peptide complexes to ACE2 were tested using the Octet RED96 system (FortéBio).
Pseudovirus preparation
The VSV-ΔG-GFP-based pseudoviruses were prepared as previously described33,34,35. The coding sequence for the spike protein of the tested viruses with 18 amino acids truncated at the C-terminal was synthesized and cloned into the pCAGGS vector, respectively. Briefly, 30 μg of the plasmids were transfected into HEK293T cells. After 24 h, the VSV-ΔG-GFP pseudovirus was added to the cells. Then the cell culture medium was removed and replaced with fresh DMEM containing 10 μg mL−1 of anti-VSV-G antibody (I1-Hybridoma ATCC® CRL-2700™) after 2 h of incubation. After 30 h, the supernatant was collected, centrifuged, and filtered through a 0.45 μm filter to remove cell debris. The pseudoviruses were frozen at −80 °C until use. Vero cells or HEK293T cells were used to determine the pseudovirus titers, and BHK-21 cells were used to detect the VSV-ΔG-GFP pseudovirus residue.
Pseudovirus neutralization assay
For the neutralization assay, 1 × 104 of Vero cells were plated into each well of a 96-well plate 24 h before infection. The peptides were three-fold serially diluted starting from 100 mM with DMEM, then the diluted peptides were incubated with an equal volume of supernatant containing 1000 fluorescence focus units of pseudovirus for 1 h at 37 °C. Then, the mixture was added to the Vero cells. Fifteen hours later, the infected cells were measured using the CQ1 Confocal Quantitative Image Cytometer (Yokogawa). The half maximal inhibitory concentration (IC50) was calculated using GraphPad Prism 8.0.
The live virus inhibition assay
6L3-1F3P11hR were serially diluted and mixed with SARS-CoV-2 (Prototype, B.1.1.7/Alpha, BA.2.12.1 and EG.5.1) for 1 h of incubation. The mixtures were then added to Vero E6-TMPRSS2 cells with three repeats and incubated for 3 days. The supernatant samples were collected for RNA extraction and quantitative RT-qPCR.
Culture and differentiation of human primary nasal epithelium at the air-liquid interface
Primary nasal epithelial cells (Epithelix) from three healthy donors were pooled and cultured in hAEC medium (Epithelix) until confluency. The cells were then transferred to 6.5 mm Transwell® inserts with 0.4 µm Pore Polyester Membranes (Corning) in 24-well plates. After reaching confluency in the Transwells, the apical medium was removed, and PneumaCult™-ALI Medium (STEMCELL Technologies) was introduced to induce differentiation. The cells were cultured at the ALI for 28 days to fully differentiate into nasal epithelium. Following differentiation, the nasal ALI cultures were fixed with 4% paraformaldehyde, permeabilized with 0.1% Triton X-100, and stained with anti-ACE2 (Abcam; ab15348), anti-Acetyl-α-Tubulin (Lys40) (D20G3) (Cell Signaling; 5335), anti-MUC5A (Thermo Fisher, MA5-12178), anti-F actin (Invitrogen, A12379), and DAPI to evaluate ACE2 expression, the presence of ciliated cells, the presence of goblet cells, cytoskeleton and nuclear pseudostratification, respectively. Images were captured using a Leica Stellaris 5 LSM microscope and processed into 3D reconstructions via volume rendering with Imaris software.
Production of SARS-CoV-2 variants and infection of nasal ALI
SARS-CoV-2 isolates hCoV-19/USA/NY-MSHSPSP-PV56475/2022 and hCoV-19/USA/MSHSPSP-PV96109/2023 were preincubated with macrocyclic peptides at final concentrations of 10 µM or 100 µM. A 100 µL aliquot of the virus-peptide mixture was then added to the apical chamber of the Transwell insert. Inoculation was conducted at 37 °C with an initial viral load of 10,000 TCID50. Following 1 h of incubation, the viral inoculum was removed, and the cells were rinsed with medium. Twenty-four hours post-infection, the cells were homogenized in Trizol-LS (Sigma) for total RNA extraction. Quantification of viral RNA was performed by targeting the ORF1a region36.
In vivo evaluation of 6L3-1F3P11hR
To evaluate the in vivo antiviral activity of 6L3-1F3P11hR, 8 to 10-week-old male K18-hACE2 mice (n = 5 per group) were intranasally challenged with 4 × 103 plaque-forming units of SARS-CoV-2 Omicron BA.2 at day 0. 6L3-1F3P11hR (3 mg/kg per dose, dissolved in PBS with 2% DMSO and 0.25% methylcellulose) was administered intranasally prophylactically (given one dose at 30 min before virus challenge) or therapeutically (given one dose at 30 min after virus challenge and then every 24 h for 3 more days, a total of four doses were administered). Control groups included solvent-treated mice (PBS with 2% DMSO and 0.25% methylcellulose per dose) and nirmatrelvir-treated mice (100 mg/kg per dose, intragastric administration) as negative and positive controls, respectively. At 4 days post-infection (dpi), the mice were sacrificed, and the nasal turbinate and lung were harvested for virological and histological analyses. The therapeutic efficacy of the peptide administered 8 h after virus challenge was also evaluated. 6L3-1F3P11hR (6 mg/kg per dose, dissolved in PBS with 2% DMSO and 0.25% methylcellulose) was administered intranasally at 8 h after virus challenge and then every 24 h for 3 more days; a total of four doses were administered. Control groups included solvent-treated mice (PBS with 2% DMSO and 0.25% methylcellulose per dose) and nirmatrelvir-treated mice (100 mg/kg per dose, intragastric administration) as negative and positive controls, respectively. Solvent and nirmatrelvir were administered using the same method as previously described. The animal experimental scheme figures were created with BioRender.com.
RNA extraction and quantitative RT-qPCR
RNA extraction and RT-qPCR were performed as previously described37. SARS-CoV-2 genome copies were quantified with sequence-specific probes and primers targeting the Envelope gene.
E_Sarbeco_F1, (SARS-CoV-2), forward primer:
5′- CGATCTCTTGTAGATCTGTTCTC -3′
E_Sarbeco_R2 (SARS-CoV-2), reverse primer:
5′- ATATTGCAGCAGTACGCACACA -3′
E_Sarbeco_P1 (FAM) Probe (SARS-CoV-2):
5′- FAM- ACACTAGCCATCCTTACTGC
GCTTCG -ZEN-IBHQ -3′
Infectious virus titration by plaque assays
Plaque assays were performed as previously described37. The nasal turbinate and lung harvested from virus-inoculated animals were homogenized and followed by centrifugation. Tissue homogenates were then 10-fold serially diluted with DMEM and incubated with Vero E6 cells for 2 h. After inoculation, cells were washed and overlaid with 1% low-melting agarose containing 1% FBS. Cells were further incubated for 72 h and fixed with 0.5% crystal violet for visualizing plaque formation.
Immunofluorescence staining
Lung tissues from mice were harvested and fixed with formalin. Nucleocapsid of SARS-CoV-2 was detected by rabbit polyclonal anti-SARS-CoV-2 nucleocapsid antibody as previously reported38. The cell nuclei were stained with DAPI (Thermo Fisher Scientific) before the tissue sections were mounted with the Diamond Prolong Antifade mounting buffer (Thermo Fisher Scientific). Images were acquired with the Olympus BX53 light microscope.
Cryo-EM sample preparation and data acquisition
To prepare the cryo-EM sample of S-6L3-3P11K, the SARS-CoV-2 Prototype and Omicron BA.2.75 S protein sample was diluted with purification buffer (20 mM Tris-Base, 150 mM NaCl, pH 8.0), and then mixed with 6L3-3P11K at a molar ratio of protein:peptide of 1:3, respectively. After incubation for 1 h on ice, the complex was applied to a glow-discharged graphene-coated grid (R1.2/1.3), which was blotted for 3 s with a humidity of 100% at 4 °C, and then plunged into liquid ethane using an FEI Vitrobot Mark IV. The well-prepared cryogenic specimens were transferred onto an FEI Titan Krios transmission electron microscope for data collection. Cryo-EM micrographs were automatically collected using Serial-EM software using a beam-image shift imaging scheme. Images were recorded with a K3-subunit detector using the super-resolution counting mode at a calibrated magnification of 22,500, corresponding to a pixel size of 1.07 Å. The exposure was performed with a dose rate of 20 e−1 pixel−1 s−1 and an accumulative dose of 60 e−1 Å2 for each micrograph, which was fractionated into 32 frames. The defocus range of this dataset was roughly −1.5 to −2.6 μm.
SARS-CoV-2 Prototype and Omicron BA.2.75 S protein samples were diluted with purification buffer (20 mM Tris-Base, 150 mM NaCl, pH 8.0) and were applied to a glow-discharged quantifoil copper grid (R1.2/1.3), which was blotted for 6 s with a humidity of 100% at 4 °C, and then plunged into liquid ethane using an FEI Vitrobot Mark IV. The well-prepared cryogenic specimens were transferred onto an FEI Titan Krios G4 transmission electron microscope for data collection. Cryo-EM micrographs were automatically collected using EPU software at a nominal magnification of 105,000× (calibrated pixel size of 0.825 Å per pixel) with a defocus range of −0.8 to −2.0 μm. The exposure was performed with a dose rate of 12 e−1 pixel−1 s−1 and an accumulative dose of 50 e−1 Å2 for each micrograph, which was fractionated into 40 frames.
Image processing, model building, and structure refinement
The movie frames were aligned using MotionCor239, and the contrast transfer function (CTF) values of each micrograph were determined using CTFFind440. Fifty micrographs were selected for automatic particle picking using Laplacian-of-Gaussian blob detection, and were subjected to 2D classification to generate templates for autopicking against the entire dataset. All subsequent classification and reconstruction procedures were performed using cryoSPARC41. After 2D classification, particles with good qualities were selected for global 3D reconstruction and then subjected to homogeneous refinement. We performed manual model building to improve local fit using COOT42. The stereochemical quality of each model was assessed using MolProbity43. Structural figures were prepared with Pymol (https://pymol.org/) and Chimera X44.
Effects of temperature, serum, and pH on the chemical stability of the peptide
The effects of temperature, serum, and pH on the stability of the peptide were explored. The peptide was treated with heating at 45 °C, mouse serum, and different pH levels, respectively. The peptide was heated at 45 °C for different time intervals, i.e., 1, 2, 3, and 4 weeks, and then analyzed by HPLC to determine the stability of the peptide at 45 °C. The peptide was incubated with various pH buffers (pH 5.5, 7.5, 8, 9, and 10) at room temperature for 24 h, respectively. Sodium acetate buffer was used for the preparation of a pH 5.5 buffer. PBS was used for preparing pH buffers at 7.5, 8, 9, and 10. Then, the clear supernatant was obtained by centrifuging the sample and further analyzed by HPLC. For the serum stability test, 2.5 μL of 2.5 mM peptide solution was added to 47.5 μL of serum solution prepared by mixing 10 μL of 1 × phosphate buffer with 37.5 μL of mouse serum to obtain the reaction mixture (125 μM peptide), and incubated at 37 °C. At the specified time points (0, 0.5, 1, 2, 4, 8, and 24 h), a 50 μL aliquot of the peptide-serum mixture was combined with 0.6 μL of 17.5 mM acetaminophen and 50 μL of acetonitrile containing 3% trifluoroacetic acid (TFA). The reaction was cooled to 4 °C for 5 min and then added to 100 μL of 20% trichloroacetic acid (TCA). After incubation at 4 °C for 5 min, the mixture was centrifuged at 18,700 × g for 5 min to precipitate the serum proteins. The supernatant was used to determine the concentration of peptides by HPLC. During HPLC analysis, water (0.1% TFA) and acetonitrile (0.1% TFA) were mixed in a linear gradient (10–85% acetonitrile in 30 min), with a flow rate of 1 mL/min, and the absorbance was detected at 210 nm. The concentrations of the remaining soluble peptides were calculated by peak integration relative to the initial peak integration (0 h).
Pharmacokinetic study
Male C57BL/6 mice were used to assess the pharmacokinetics. C57BL/6 mice were dosed at an age of 6–8 weeks and were provided free access to food and water prior to dosing. Mice were injected intranasally with a dose of 3 mg/kg 6L3-1F3P11hR. Blood and nasal lavage fluid were collected at each time point (2 min, 5 min, 15 min, 30 min, 1, 4, 8, and 24 h). Approximately 0.02 mL of blood was collected. Blood from each sample was transferred into plastic microcentrifuge tubes containing anticoagulant EDTA-K2 and mixed well with the anticoagulant. The blood samples were centrifuged at 4000 × g for 5 min at 4 °C to obtain plasma. For nasal lavage fluid collection, 1 mL of normal saline was slowly infused via the nasopharynx after locating the nasal opening (two repetitions). Rinse solutions were collected in EP tubes. Then, the samples were immediately frozen in the upright position and stored at −80 °C prior to analysis. Concentrations of 6L3-1F3P11hR in the plasma and nasal lavage fluid were analyzed using an LC-MS/MS method. WinNonlin (PhoenixTM, version 8.4) software was used for pharmacokinetic calculations.
Peptide oligomerization analysis by LC-MS in solution
To explore the mass of peptides, the standard stock solutions and synthesized sample were diluted to a 0.5 μg/μL (6L3-3P11K) in 50 mM Ammonium acetate (pH 7.48). 10 μL of sample was injected onto an ACQUITY UPLC H-Class PLUS System configured with a BEH SEC 200 Å 1.7 μm, 2.1 × 150 mm Column (Waters, American). The mobile phase was 50 mM ammonium acetate in water. Chromatographic separation was achieved using an isocratic rate of 0.065 mL/min with a total run time of 10 min and a 25 °C column temperature. MS experiments were performed using a SYNAPT XS Mass Spectrometer with a standard electrospray ionization (ESI) source (Waters, American). The MS parameters were set as follows: positive ESI mode, capillary voltage 2.0 kV, cone voltage 30 V, source heating temperature 125 °C, desolvation temperature 250 °C, cone gas flow 50 L/h, desolvation gas flow 600 L/h, mass/charge (m/z) range 50–3000, Scan Time 0.5 s. Lock mass data was collected during each acquisition, and the correction was applied during post-acquisition data processing via MassLynx Software v4.1 (Waters, American). Deconvolution of mass spectra was performed using MaxEnt3 in UNIFI software v3.1 (Waters, American). Deconvolution was performed in the 0–12,500 Da range, and a target mass of ~1462 Da. The mass of the peptide was compared to the standard mass to detect any mass modification and get additional information about the characterized compounds.
Statistical analysis
All statistical tests were performed as described in the indicated figure legends using GraphPad Prism software. The number of independent experiments performed is indicated in the relevant figure legends. No samples were excluded from the analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic coordinates data generated in this study have been deposited in the Protein Data Bank (PDB) database under accession codes 9L3Q and 9L3I. The corresponding electron microscopy maps have been deposited in the Electron Microscopy Data Bank (EMDB) with the accession codes EMD-62788, EMD-62786, EMD-67440, EMD-67548, EMD-67549, and EMD-67568. Source data are provided with this paper.
References
Hampshire, A. et al. Cognition and memory after COVID-19 in a large community sample. N. Engl. J. Med. 390, 806–818 (2024).
Xie, Y., Choi, T. & Al-Aly, Z. Association of treatment with nirmatrelvir and the risk of post-COVID-19 condition. JAMA Intern. Med. 183, 554–564 (2023).
Fung, K. W., Baye, F., Baik, S. H. & McDonald, C. J. Nirmatrelvir and molnupiravir and post-COVID-19 condition in older patients. JAMA Intern. Med. 183, 1404–1406 (2023).
Liu, L. H. et al. Striking antibody evasion manifested by the Omicron variant of SARS-CoV-2. Nature 602, 676–681 (2022).
Iketani, S. et al. Antibody evasion properties of SARS-CoV-2 Omicron sublineages. Nature 604, 553–556 (2022).
Chan, J. F.-W., Yuan, S., Chu, H., Sridhar, S. & Yuen, K.-Y. COVID-19 drug discovery and treatment options. Nat. Rev. Microbiol. 22, 391–407 (2024).
Duan, Y. et al. Molecular mechanisms of SARS-CoV-2 resistance to nirmatrelvir. Nature 622, 376–382 (2023).
Hammond, J. et al. Nirmatrelvir for vaccinated or unvaccinated adult outpatients with Covid-19. N. Engl. J. Med. 390, 1186–1195 (2024).
Muttenthaler, M., King, G. E., Adams, D. J. & Alewood, P. E. Trends in peptide drug discovery. Nat. Rev. Drug Discov. 20, 309–325 (2021).
Vinogradov, A. A., Yin, Y. Z. & Suga, H. Macrocyclic peptides as drug candidates: recent progress and remaining challenges. J. Am. Chem. Soc. 141, 4167–4181 (2019).
Hosseinzadeh, P. et al. Comprehensive computational design of ordered peptide macrocycles. Science 358, 1461–1466 (2017).
Murakami, H., Ohta, A., Ashigai, H. & Suga, H. A highly flexible tRNA acylation method for non-natural polypeptide synthesis. Nat. Methods 3, 655–655 (2006).
Goto, Y., Katoh, T. & Suga, H. Flexizymes for genetic code reprogramming. Nat. Protoc. 6, 779–790 (2011).
Yamagishi, Y. et al. Natural product-like macrocyclic N-methyl-peptide inhibitors against a ubiquitin ligase uncovered from a ribosome-expressed de novo library. Chem. Biol. 18, 1562–1570 (2011).
Goto, Y. & Suga, H. The RaPID platform for the discovery of pseudo-natural macrocyclic peptides. Acc. Chem. Res. 54, 3604–3617 (2021).
Norman, A. et al. Discovery of cyclic peptide ligands to the SARS-CoV-2 spike protein using mRNA display. ACS Cent. Sci. 7, 1001–1008 (2021).
Shi, G. L. et al. Omicron Spike confers enhanced infectivity and interferon resistance to SARS-CoV-2 in human nasal tissue. Nat. Commun. 15, 889 (2024).
Zhang, A. J. et al. Coinfection by severe acute respiratory syndrome coronavirus 2 and influenza A(H1N1)pdm09 virus enhances the severity of pneumonia in golden Syrian hamsters. Clin. Infect. Dis. 72, 978–992 (2021).
Chen, Y. et al. Broadly neutralizing antibodies to SARS-CoV-2 and other human coronaviruses. Nat. Rev. Immunol. 23, 189–199 (2023).
Silwal, A. P. et al. DNA aptamers inhibit SARS-CoV-2 spike-protein binding to hACE2 by an RBD- independent or dependent approach. Theranostics 12, 5522–5536 (2022).
Cao, L. X. et al. De novo design of picomolar SARS-CoV-2 miniprotein inhibitors. Science 370, 426-431 (2020).
Jaiswal, G. & Kumar, V. In-silico design of a potential inhibitor of SARS-CoV-2 S protein. PLoS ONE 15, e0240004 (2020).
Curreli, F. et al. Stapled peptides based on human angiotensin-converting enzyme 2 (ACE2) potently inhibit SARS-CoV-2 infection. mBio 11, e02451–02420 (2020).
Larue, R. C. et al. Rationally designed ACE2-derived peptides inhibit SARS-CoV-2. Bioconjugate Chem. 32, 215–223 (2021).
Valiente, P. A. et al. Computational design of potent D-peptide inhibitors of SARS-CoV-2. J. Med. Chem. 64, 14955–14967 (2021).
Pei, P. F. et al. Computational design of ultrashort peptide inhibitors of the receptor-binding domain of the SARS-CoV-2 S protein. Brief. Bioinform. 22, bbab243 (2021).
Yang, J. et al. Selection and engineering of broad-spectrum antiviral affibody peptide against SARS-CoV-2 variants. hLife. 9, 448–451 (2025).
Thijssen, V. et al. A broad-spectrum macrocyclic peptide inhibitor of the SARS-CoV-2 spike protein. Proc. Natl. Acad. Sci. USA 120, e2303292120 (2023).
Wu, L. L. et al. A pan-coronavirus peptide inhibitor prevents SARS-CoV-2 infection in mice by intranasal delivery. Sci. China Life Sci. 66, 2201–2213 (2023).
Mathieu, C., Porotto, M., Figueira, T. N., Horvat, B. & Moscona, A. Fusion inhibitory lipopeptides engineered for prophylaxis of Nipah virus in primates. J. Infect. Dis. 218, 218–227 (2018).
Shi, Y. Drug development in the AI era: AlphaFold 3 is coming! Innovation 5, 100685 (2024).
Chan, J. F. W. et al. Virological features and pathogenicity of SARS-CoV-2 Omicron BA.2. Cell Rep. Med. 3,100743 (2022).
Nie, J. et al. Quantification of SARS-CoV-2 neutralizing antibody by a pseudotyped virus-based assay. Nat. Protoc. 15, 3699–3715 (2020).
Wu, L. et al. Efficient inhibition of SARS-CoV-2 emerging EG.5, EG.5.1 and BA.2.86 variants by fusion inhibitor HY3000 peptide. hLife 2, 43–46 (2024).
Su, C. et al. Enabling the immune escaped etesevimab fully-armed against SARS-CoV-2 Omicron subvariants including KP.2. hLife 3, 132–145 (2025).
Shi, G. L. et al. Rapalogs downmodulate intrinsic immunity and promote cell entry of SARS-CoV-2. J. Clin. Investig. 132, e160766 (2022).
Shuai, H. et al. Emerging SARS-CoV-2 variants expand species tropism to murines. EBioMedicine 73, 103643 (2021).
Shuai, H. et al. Differential immune activation profile of SARS-CoV-2 and SARS-CoV infection in human lung and intestinal cells: Implications for treatment with IFN-β and IFN inducer. J. Infect. 81, e1–e10 (2020).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A. et al. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Acknowledgements
The authors thank all the staff members at the Center for Biological Imaging (CBI), Institute of Biophysics (IBP), Chinese Academy of Sciences (CAS), and the Advanced Bio-imaging Core Facility of Guangzhou Laboratory, the Cryo-electron Microscopy Platform, Institute of Genetics and Developmental Biology (IGDB), Chinese Academy of Sciences, for assistance with data collection. The authors also thank Zheng Fan, Wei Zhang, Qian Wang, and Li Wang from the Institute of Microbiology, Chinese Academy of Sciences, for help in detecting protein interactions and peptide analysis. This study was supported by National Natural Science Foundation of China (NSFC) (32192452 to Y.S.), Major Project of Guangzhou National Laboratory (GZNL2025C01001 to Y.S.), Beijing Life Science Academy (2023000CA0030 to Y.S.), National Natural Science Foundation of China (NSFC) (22277065 to Y.Y., 32100119 to Q.P.), National Natural Science Foundation of China Young Scientist Fund (Type A) (82525037 to J.F.-W.C.), National Natural Science Foundation of China Excellent Young Scientists Fund (Hong Kong and Macau) (32322087 to S.Y.), Taishan Scholar Program of Shandong Province (tsqn202103148 to Y.Y.), Health@InnoHK (Centre for Virology, Vaccinology and Therapeutics), Innovation and Technology Commission, the Government of the Hong Kong Special Administrative Region (S.Y. and J.F.-W.C.), the Research Fellow Scheme (RFS2425-7S06 to J.F.-W.C.), Collaborative Research Fund (C7060-21G to J.F.-W.C.), and Theme-Based Research Scheme of the Research Grants Council (T11-709/21-N) (J.F.-W.C.); the Partnership Programme of Enhancing Laboratory Surveillance and Investigation of Emerging Infectious Diseases and Antimicrobial Resistance for the Department of Health of the Hong Kong Special Administrative Region Government (J.F.-W.C.), Sanming Project of Medicine in Shenzhen, China (SZSM201911014) (J.F.-W.C.), the High Level-Hospital Program, Health Commission of Guangdong Province, China (J.F.-W.C.) and Emergency Key Program of Guangzhou Laboratory (EKPG22-01) (J.F.-W.C.). The funding sources had no role in the study design, data collection, analysis, interpretation, or writing of the report.
Author information
Authors and Affiliations
Contributions
Y.S., Y.Z.Y., J.F.-W.C., and H.H. conceived the study. M.W., J.Y., Y.T., S.Y., G.S., Q.P., V.K.P., N.X., A.J.Z., Y.X., J.L., and H.L. performed the experiments. Y.S., M.W., J.Y., Y.T., Y.Z.Y., J.F.-W.C., V.K.P., J.L., H.L., Y.F.Y., Y.C., A.C., Y.Z., Y.Y., G.F.G., Y.L., and C.C.C. conducted the analysis. Y.S., M.W., J.Y., Y.T., Y.Z.Y., J.F.-W.C., and H.H. wrote the manuscript. All authors participated in the discussion and manuscript editing. Y.S. supervised all the work.
Corresponding authors
Ethics declarations
Competing interests
Two patents describing potential treatments for COVID using 6L3-related peptides were applied, and the authors listed on the application are Y.S., G.F.G., M.W., J.Y., and Q.P. One patent describing potential treatments for COVID using 6L3-3P11hR and 6L3-1F3P11hR peptides was in application, and the authors listed on the application are Y.S., M.W., J.Y., Q.P., H.H., and Y.F.
Peer review
Peer review information
Nature Communications thanks Mark Broenstrup, Masaud Shah and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, M., Yang, J., Tan, Y. et al. Intranasal administration of broad-spectrum macrocyclic peptide inhibitor protects against SARS-CoV-2 Omicron variants. Nat Commun 17, 1753 (2026). https://doi.org/10.1038/s41467-026-68462-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68462-9