Abstract
The astrocytic γ-aminobutyric acid (GABA) transporter, GAT3, is essential for terminating GABAergic signaling in the central nervous system. Selective inhibition of GAT3 offers a potential strategy for elevating extracellular GABA levels for the treatment of neurological disorders including epilepsy. However, few potent and selective GAT3 inhibitors have been developed, and their mechanisms of inhibition remain poorly understood. Here, we present the cryo-electron microscopy structures of full-length, wild-type human GAT3, hGAT3, bound to a selective inhibitor, to substrate GABA, or in substrate-free state. hGAT3 bound to the inhibitor or in the substrate-free state exhibits an inward-open conformation. The inhibitor binds within the intracellular permeation pathway, positioned between transmembrane helices 1, 2, 3, 6, 7, and 8. The GABA-bound hGAT3 is captured in an inward-occluded state, revealing the ion coordination and substrate recognition network, including a cation-π interaction between GABA’s γ-amino group and a phenylalanine residue in transmembrane helix 6. Our data reveal the molecular determinants for the inhibitor selectivity, and the mode of substrate binding and transport inhibition, providing blueprints for the rational design of next-generation selective GAT3 inhibitors.
Similar content being viewed by others
Introduction
γ-Aminobutyric acid (GABA) is the main inhibitory neurotransmitter in the central nervous system (CNS). GABA exerts its inhibitory effects via ionotropic GABAA receptors and metabotropic GABAB receptors, leading to hyperpolarization of the postsynaptic membranes. Master keys in the regulation of GABA signaling are the transmembrane GABA transporters (GATs) that belong to the family of secondary active neurotransmitter/sodium symporters (NSSs) of the solute carrier 6 (SLC6)1. The GATs terminate GABA neurotransmission by taking up GABA into presynaptic neurons and astrocytes. Among the four GAT subtypes (GAT1, GAT2, GAT3 and BGT1; human nomenclature), GAT1 (SLC6A1) and GAT3 (SLC6A11) are the most prevalent in the CNS (for a review of all four subtypes see ref. 2). GAT1, located primarily on neurons, mediates uptake of GABA into presynaptic terminals, thus permitting re-cycling of neurotransmitter, whereas GAT3, expressed solely on astrocytes, governs glial GABA uptake and additionally supports oxidative GABA metabolism3,4,5. The metabolic turnover of GABA into glutamine sustains the downstream bidirectional interaction between neurons and astrocytes and highlights GAT3 as an interesting—yet understudied—target for regulating overall network excitability6,7. In recent years, neurodevelopmental disorders and genetic epilepsies caused by mutations or deletions, predominantly in SLC6A18, but also in SLC6A119 underscore the relevance of regulating GABA levels in the brain and identifying new therapies.
Dysregulation of GABAergic signaling with a specific involvement of GAT3 has been implicated in various neurological disorders, including epilepsy, Alzheimer’s disease, and ischemic stroke10,11,12,13,14. Potentiating GABA-mediated transmission has proved to be an effective approach to suppress seizure activity in epilepsy. Since the approval of tiagabine, a lipophilic, nipecotic acid-derived GAT1 inhibitor, nearly three decades ago, no other GAT inhibitors have been developed for the treatment of epilepsy15. Due to its glial localization, pharmacological inhibition of GAT3 is attractive as it offers an alternative therapeutic approach to elevating extracellular and/or extrasynaptic GABA levels, and may thus enhance inhibitory neurotransmission in a manner distinct from GAT116,17,18. However, very few GAT3 inhibitors have been developed and pharmacologically evaluated, mainly due to a limited potency and selectivity, and thus far, no GAT3 inhibitors have reached clinical translation.
These limitations, in part, stem from the lack of detailed structural information on the GAT3 binding site, which has led to a simplistic approach in designing GAT3 inhibitors, primarily through N-substitution of known GAT3 substrates with specific bulky lipophilic moieties, conferring GAT3 selectivity over GAT1. Among these inhibitors, (S)-SNAP-5114 is the most well-known, but its poor chemical stability restricts in vivo applications. Inspired by DPPM-1457, a more stable (S)-SNAP-5114 variant, we recently developed the GAT3 inhibitor (2S, 2′R)−2-hydroxy-2-(1-((E)−4,4,4-tris(4-methoxyphenyl)but-2-en-1-yl)pyrrolidin-2′-yl)acetic acid, in brief SR-THAP19,20,21 (design strategy in Supplementary Fig. 1). Notably, SR-THAP is derived from a substrate-like inhibitor that is otherwise nearly inactive.
The process of neurotransmitter uptake by GAT3 and other members of NSS is coupled to co-transport of Na+ (and Cl– in eukaryotic members), where the transporter cycles through distinct conformational states, wherein the substrate-binding pocket is alternately exposed to the extracellular and the intracellular spaces22,23,24,25,26,27,28,29,30,31,32. Interestingly, GAT3 may also operate in reverse, contributing to the tonic inhibition of neurons33. Detailed structural insights into GAT3 and its interaction with the substrate and inhibitor candidates are essential for understanding the molecular basis of GAT3 uptake and the development of effective pharmacological agents. Here, we present the cryo-electron microscopy (cryo-EM) structures of human GAT3, hGAT3, bound to the selective inhibitor SR-THAP, to the substrate GABA, and in the substrate-free state. We capture the inhibitor-bound hGAT3 and substrate-free states in an inward-open conformation, while the GABA-bound state is in an inward-occluded conformation. Our study reveals the structural determinants and mechanism of inhibition of GABA transport by hGAT3.
Results
SR-THAP is a selective inhibitor of GAT3
We have designed SR-THAP and its enantiomer RS-THAP with the aim of developing potent and selective hGAT3 inhibitors (Supplementary Fig. 1)20. To evaluate the inhibition mode of SR-THAP, we measured GABA uptake inhibition at hGAT3 with and without preincubation of the inhibitor. Without preincubation, SR-THAP inhibited the uptake of GABA in HEK293T cells expressing hGAT3 with a half-maximal inhibitory concentration (IC50) of 25.6 μM, while its enantiomer, RS-THAP, showed only a limited effect (max 25% inhibition at 300 μM) (Fig. 1a, b). We then measured GABA uptake inhibition at hGAT3 after 15, 30, and 120 min of preincubation with SR-THAP, which significantly left-shifted the curves, yielding IC50 values of 4.4, 4.5, and 4.9 μM, respectively, for SR-THAP and 140 μM for RS-THAP (Fig. 1a, b). As expected, the IC50 value for the substrate GABA remained unchanged with and without preincubation, as the substrate binds the transporter solely from the extracellular space (Fig. 1c). The inhibition mode of SR-THAP at hGAT3 has been hypothesized to resemble the dual inhibition mechanism of tiagabine at hGAT1, which is dependent on preincubation conditions, where the inhibitor binds the transporter from both the extracellular and the intracellular sides27,34. We further investigated the mechanism of SR-THAP inhibition at hGAT3, by performing kinetic experiments to determine the Michaelis–Menten constant (KM) and maximal transport velocity (Vmax) for [3H]GABA at increasing SR-THAP concentrations (Fig. 1d). We observed relatively constant KM values with and without preincubation, which combined with a significant decrease in Vmax suggest an overall non-competitive mode of inhibition. The non-competitive profile of SR-THAP, together with its time-dependent shift in potency, indicates the existence of a binding site on the intracellular side of hGAT3. To assess its potential for passive membrane permeation, we compared the clogD7.4 of SR-THAP with those of known permeable compounds, zolpidem and propranolol, the impermeable atenolol, and the hGAT1 inhibitor tiagabine. SR-THAP (clogD7.4 = 3.43) and tiagabine (clogD7.4 = 3.07) are highly lipophilic, comparable to zolpidem (clogD7.4 = 3.50), and substantially more lipophilic than propranolol (1.05) or the hydrophilic atenolol (−2.03)35. The high lipophilicity of SR-THAP supports its ability to passively diffuse across the plasma membrane.
a SR-THAP inhibits [3H]GABA uptake by hGAT3 expressed in HEK293T cells with an IC50 of 25.2 [21.8; 30.4] μM without preincubation (blue) and 4.4 [3.6; 5.2] µM, 4.5 [3.4; 5.9] µM, and 4.9 [3.3; 7.0] μM after 15, 30, and 120 min preincubation, respectively. b The enantiomer RS-THAP inhibits [3H]GABA uptake by hGAT3 expressed in HEK293T cells, but with significantly reduced potency, with estimated IC50 values of >600 μM without preincubation and >100 μM with preincubation. c [3H]GABA uptake by hGAT3 can also be inhibited by non-radioactive GABA, although showing minimal shift in potency after 120 min preincubation, with IC50 values of 8.5 [6.4; 11] and 5.4 [4.6; 6.4] μM, respectively. d [3H]GABA uptake kinetic experiment fitted to the Michaelis–Menten model, performed with increasing concentrations of SR-THAP without preincubation (left) or after 120 min of preincubation (middle), before addition of [3H]GABA. Right panel shows the Michaelis–Menten kinetic parameters derived from the graphs in the left and middle panels, without (top) and with (bottom) 120 min preincubation. Statistical differences from control were determined by a two-sided F test and are indicated with stars as: * - P ≤ 0.05, ** - P ≤ 0.01. For reference, the F-test p values were 0.029 (0.5 µM), 0.0021 (2 µM), and 0.0015 (10 µM) without preincubation, and 0.652 (0.5 µM) and 0.0022 (2 µM) with preincubation. Data points are shown as individual replicates from n = 3 biological replicates (triplicate measurements) unless otherwise stated. Source data are provided as a Source data file.
We investigated the selectivity of SR-THAP for hGAT3, motivated by its higher potency compared to its enantiomer (Fig. 1a, b). For comparison, we included hGAT1 and the human glycine transporter 1 (hGlyT1), as the binding sites of their respective selective inhibitors, tiagabine (at GAT1) and the high-affinity benzoylisoindoline inhibitor, Cmpd128,36 (at GlyT1) overlap with our hypothesized binding site of SR-THAP. SR-THAP was found to be respectively 42 and 23 times more selective for hGAT3 over hGAT1 (estimated IC50 value of 212 μM) and hGlyT1 (estimated IC50 value of 117 μM) (Supplementary Fig. 2a, b). Further, the hGlyT1 inhibitor, Cmpd1, exhibited no effect on hGAT3-mediated GABA uptake at concentrations up to 30 μM (approximately 2300 times the IC50 of hGlyT1) (Supplementary Fig. 2c). We further assessed SR-THAP using the fluorometric imaging plate reader (FLIPR) membrane potential (FMP) assay. As expected, SR-THAP elicited no substrate-like response, showing no significant difference from buffer, in contrast to GABA (Supplementary Fig. 2d). Consistently, we were able to replicate the Michaelis-Menten kinetic experiment, which revealed a clear non-competitive inhibition profile for SR-THAP, in the absence of preincubation (Supplementary Fig. 2e).
Given the lack of detailed experimental information on the interaction network of SR-THAP with hGAT3, and to understand the mechanism underlying the inhibition of GABA transport, we next sought to determine the structures of hGAT3 bound to the inhibitor and the substrate GABA.
Overall architecture of human GAT3
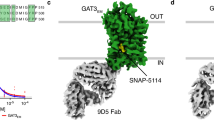
We purified wild-type hGAT3 in a glyco-diosgenin (GDN) micelle, obtaining a monodisperse and homogenous sample for cryo-EM studies (Supplementary Fig. 3a, b). We measured GABA uptake inhibition by wild-type untagged hGAT3 and the cryo-EM hGAT3 construct (with affinity tags) and observed no difference in IC50 values (6.9 μM and 7.0 μM, respectively) (Supplementary Fig. 3c, d). To assess surface expression of hGAT3 and whether it was affected by incubation with SR-THAP, we used an eGFP-tagged hGAT3 construct and performed confocal microscopy to quantify membrane-localized fluorescence (Supplementary Fig. 3e). No difference in surface expression of hGAT3 was observed after 1 h incubation with SR-THAP (Supplementary Fig. 3f). We determined the structures of hGAT3 bound to inhibitor SR-THAP, substrate GABA, or in substrate-free state at 2.9, 3.2, and 3.9 Å resolution, respectively (Fig. 2, Supplementary Figs. 4–9).
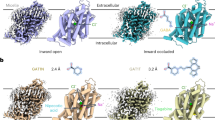
Cryo-EM density maps of hGAT3 bound to a SR-THAP (blue map, contour level = 0.063 in ChimeraX), and b GABA (green map, contour level = 0.06 in ChimeraX). Lipid densities are shown in light yellow. c, d Surface representation of the inhibitor-bound and GABA-bound structures of hGAT3 viewed parallel to the membrane. c Slice view of hGAT3 showing the SR-THAP (orange) binding pocket. d Slice view of hGAT3 showing the GABA (green) binding pocket. The extracellular gating residues R75 (TM1 – blue) and D467 (TM10 – not shown), as well as P375 (EL4 – yellow) and F308 (TM6 – green), are shown as sticks in c, d.
We unambiguously modeled all transmembrane (TM) helices with clear side chain densities, intracellular and extracellular loops, except parts of extracellular loop 2 (EL2) due to its high degree of flexibility (Fig. 2, Supplementary Figs. 6–9). The conserved NSS disulfide bridge between C171 and C180 (EL2) (Supplementary Fig. 10) is observed in both the inhibitor-bound and substrate-free structures of hGAT3 (Supplementary Fig. 11a). However, in the GABA-bound state, while the densities for C171 and C180 are clearly resolved, we do not observe a distinct density for the disulfide bridge, suggesting that the cysteine residues may be reduced in this sample (Supplementary Fig. 11b).
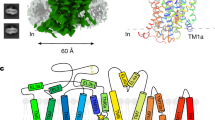
The human GAT3 adopts the expected architecture of SLC6 transporters, comprising 12 α-helical TMs with an inverted pseudo-twofold symmetric organization of TMs 1–5 relative to TMs 6–10, denoted as the LeuT fold37, with TMs 1 and 6 unwinding approximately halfway across the membrane, a characteristic for the central binding site in NSS (Fig. 2 and Supplementary Figs. 6–9).
Unique to GABA transporters, as well as to taurine and creatine transporters, is an insertion of an extra serine residue (S472 in hGAT3) in the unwound region of TM10 that forms a one-turn π-helix and provides extra bulk in this region. This extra residue has been shown to be required for the coupling of ions for the transport of the substrate GABA38,39. In all three hGAT3 structures, the side chain hydroxyl group of S472 forms hydrogen bonds with the hydroxyl group of T304 (TM6a) and R75 (TM1b), as part of the interaction network occluding the extracellular pathway (Fig. 2, Supplementary Fig. 9). In higher resolution maps of inhibitor-bound and GABA-bound hGAT3, we observe a clear density for a water molecule that mediates an interaction between S472 (TM10) and D301 (TM6a) similar to those observed for inward-facing structures of hGAT1 bound to inhibitors and substrate (Supplementary Fig. 11c)32,40.
SR-THAP locks hGAT3 in an inward-open conformation
The inhibitor SR-THAP captures hGAT3 in an inward-open conformational state, where the central substrate site is accessible to the solvent via the intracellular vestibule. The N-terminal segment of TM1 (TM1a) is bent away from the core of hGAT3, opening the cytoplasmic release pathway and allowing the accommodation of the bulky inhibitor (Supplementary Figs. 11d and 12a–c). The splayed motion of TM1 disrupts the interaction between the TM1a and the hydrophobic patch at the cytoplasmic part of hGAT3 that is otherwise present in other outward-facing or inward-occluded structures of NSS23,25,41,42,43. We could not model the intracellular part of TM5 preceding the conserved helix-breaking G(X9)P motif (G251(X9)P261 in hGAT3, Supplementary Fig. 11e), which we assume to be unwound, a feature that creates a solvent pathway at the intracellular side of the transporter, providing water access from the cytoplasm to the Na2 site42. The overall conformation of SR-THAP-bound hGAT3 resembles that of the recently published hGAT3 structure bound to (S)-SNAP-511444, with superposition of the two inhibitor-bound inward-open structures yielding a Cα root mean square deviation (r.m.s.d.) of 0.62 Å (Supplementary Fig. 12d).
The highly ordered interaction network that closes the extracellular pathway includes residues from TMs 1, 3, 6, and 10 as well as EL4 (Fig. 2). The conserved extracellular gate residues R75 (TM1b) and D467 (TM10), form a salt bridge with a Cα-Cα distance of 9.6 Å, precluding solvent access from the extracellular side. In addition to this salt bridge, R75 forms a stacking and cation-π interaction with the phenyl ring of the conserved F308 (TM6a) residue. F308, which sterically blocks the extracellular pathway, is within a Cα–Cα distance of 11.6 Å to the conserved Y147 (TM3), consistent with previous inward-facing NSS structures27,28,40,43,45.
R75, which is conserved across NSS, is critical for the transport of the substrate. The conservative mutation of the equivalent residue in hGAT1, R69K, impairs GABA transport, while sustaining the ability of the transporter to bind the substrate46, indicative of a locked outward-facing conformation of the transporter. The R75-coordinating residues, Q305 (TM6a), D467 (TM10), and S472 (unwound region of TM10), form an extensive hydrogen bond network. Maintaining the integrity of this network is critical for transporter function, as demonstrated by conservative mutations in hGAT147,48.
We observe a clear density for a Cl– ion that is coordinated by the conserved residues Y92 (TM2), Q305 (TM6a), S309 (TM6a), and S345 (TM7) (with a mean coordination distance of 3.2 Å ± 0.4) (Supplementary Figs. 10 and 11f). As in other eukaryotic SLC6 transporters, Cl– provides a negative counter-charge to the bound sodium ions, thereby stabilizing the outward-facing conformation49,50,51; this charge-balancing role may be particularly relevant for hGAT3, which has the capacity to operate in reverse under certain ionic conditions. We do not, however, observe any densities for sodium ions at Na1 or Na2 sites, consistent with an inward-open conformational state of the transporter (Supplementary Figs. 11g and 13a–e).
SR-THAP binding mode
The high-quality of the cryo-EM density for SR-THAP allowed us to unambiguously model the inhibitor sandwiched in between TMs 1, 2, 3, 6, 7, and 8 (Fig. 3a). Each of the three moieties of SR-THAP (R1-R3, Supplementary Fig. 1) is accommodated in a defined sub-pocket of hGAT3, resulting in a modular binding mode similar to that of tiagabine at hGAT1, (S)-SNAP-5114 at hGAT3, and Cmpd1 at hGlyT1 (Supplementary Fig. 12d, e)27,28,40,44.
a Overall structure of the human GAT3 bound to the selective inhibitor SR-THAP (orange sticks) and cryo-EM density of SR-THAP (map contour level = 0.072 in ChimeraX). b Zoomed-in view of the binding mode of SR-THAP (orange) of hGAT3. Protein backbone is shown as ribbons (azure) and chloride ion as green sphere. Interacting residues within 3 Å of the inhibitor are displayed. To better illustrate the compound’s protonation state and its interactions with hGAT3, the polar hydrogens are shown even though they are not resolved at this resolution. H-bonds and cation-π interactions are displayed with yellow and green dashed lines, respectively. c The hGAT3-E66Y mutant shows a near-complete loss of [3H]GABA uptake, with only ~5% transport activity remaining compared to wild-type hGAT3, while hGAT3-C316S retains approximately 60% transport activity. Data points are individual measurements from either two biological replicates (hGAT3-E66Y) or three biological replicates (hGAT3-wt and hGAT3-C316S), each biological replicate was measured in triplicates. Error bars for hGAT3-wt and hGAT3-C316S represent SEM, and for hGAT3-E66Y, they represent SD. d [3H]GABA uptake inhibition by non-radioactive GABA is not affected in the hGAT3-C316S mutant compared to the wild-type hGAT3 with an IC50 value of 4.2 [2.3; 7.9] μM. e [3H]GABA inhibition by SR-THAP is affected by the C316S mutation with a reduced potency of 10.6 [8.6; 12.7] μM, compared to 4.9 [3.3; 7.0] μM for the wild-type hGAT3. Both experiments were performed with 120 min preincubation. Data points are shown as individual replicates from n = 3 biological replicates (triplicate measurements) unless otherwise stated. f Overall structure of the hGAT3 bound to its substrate GABA (blue sticks) and cryo-EM density map of GABA (map contour level = 0.06 in ChimeraX). g Zoomed-in view of the binding mode of GABA (blue) at hGAT3. Protein backbone is shown as ribbons (green), chloride ion as a green sphere, sodium ion as a red sphere, and the same residues from (b) as sticks. Source data are provided as a Source data file.
The inhibitor binding site is capped by the aromatic side chains of F308 (TM6a) and Y147 (TM3), which—along with the lateral residue L143 (TM3)—form a lipophilic cavity that accommodates the five-membered ring of the R1 group, while the α-hydroxy acid head of R1 extends toward G71 (TM1), Y147 (TM3) and backbone of S309 (TM6) (Fig. 3b, Supplementary Fig. 14a, b).
This cavity is further lined by A311 and L314 from the non-helical region of TM6. The residue L314 is part of the conserved (G/A/C)ΦG motif (C313L314G315 in hGAT3), which plays a central role in shaping the binding site to accommodate ligands of varying sizes across the SLC6 family52. A conserved glutamate residue in TM2, E107 in hGAT3, is coordinating the CLG loop through a direct and a water-mediated hydrogen bond with backbone nitrogen of G315 (Supplementary Fig. 11h). This ordered water molecule is also observed in previous NSS structures, including hGAT1 structure bound to substrate GABA (PDB ID 7Y7W)40. The interaction between E107 (TM2) and the non-helical part of TM6 is conserved among SLC6 and appears as a key functional element of transport52. Mutagenesis of the equivalent glutamate residue in hGAT1 diminishes the transport activity, supporting its important role in a functional transporter53.
The positively charged nitrogen of R1 establishes a weak (5.9 Å) cation-π interaction with F308, while the hydroxyl group of R1 interacts via a hydrogen bond with the amidic carbonyl of S309 (TM6). The carboxylate group of R1 extends toward TM1 and interacts with the hydroxyl of Y147 (TM3) as well as the amidic NH of L70 and G71 (TM1), similar to that of substrate-transporter interactions, in α-amino acid transporter structures42,52,54. The carboxylate of R1 further interacts with TM8, through water mediated hydrogen bonds with L406 and S410 (Supplementary Fig. 14a, b). All residues interacting with the R1 part of SR-THAP are fully conserved among the GABA transporters (Supplementary Fig. 10).
The R2 moiety (N-alkyl chain) elongates into a narrow lipophilic opening toward the bottom of the binding site, and it is surrounded by the side chains of L314 (CLG motif on TM6), A311 (TM6), S410 (TM8), and V413 (TM8). The R3 group (trimethoxytrityl moiety) is accommodated within a lipophilic cavity at the bottom of the binding site, exposed to the cytosol, shaped as a tripod, and defined by residues from TMs 1, 6, 7, and 8 – G65, F104, Y310, C316 (CLG motif in TM6), L320, L337, L406, V413 and N341 (Supplementary Fig. 14a, b). The residues interacting with R2 and R3 are either fully conserved among the GABA transporters or replaced by similar residues, except the notable hGAT3-L314, which is a glutamine residue in BGT1 (Supplementary Fig. 10).
Molecular docking places (S)-DPPM-1457 and (S)-SNAP-5114 in the binding pocket of hGAT3, revealing a binding mode similar to that of SR-THAP, with preserved hydrogen bonds between the carboxylate groups of the compounds and residues G71 and Y147 (Fig. 3b, Supplementary Fig. 14c). This correlates well with recently published (S)-SNAP-5114-bound hGAT3, where the binding poses of the two inhibitors are comparable, although a tilt in the R1 group of (S)-SNAP-5114 is observed, possibly due to the absence of the α-hydroxy moiety in (S)-SNAP-5114 compared to SR-THAP44 (Supplementary Fig. 12d).
Docking of RS-THAP – the less active enantiomer of SR-THAP –—reveals a comparable occupancy of the subpockets occupied by R2 and R3, but the different stereochemistry at R1 leads to a less favorable binding energy compared to SR-THAP (∆∆G = 2.67 Kcal/mol). The R2 and R3 groups of the RS-THAP closely superpose with those of SR-THAP. However, inversion at the R1 stereocenter distorts its positioning, disrupting the hydrogen bond network and causing the carboxylate and hydroxyl groups of RS-THAP to adopt different orientations compared to those in SR-THAP, and the R2 group to twist toward TM6 (Supplementary Fig. 14d).
Comparison of the binding-pocket residues in SR-THAP-bound hGAT3 with those in tiagabine-bound hGAT1 (PDB IDs 7Y7Z and 7SK2) points to structural determinants underlying SR-THAP’s selectivity for hGAT3 over hGAT1. The small polar E66 residue is replaced by the bulky aromatic Y60 residue in hGAT1. This, in turn, reduces the size of the vestibular lipophilic sub-pocket in hGAT1, thereby precluding access of the R3 group of SR-THAP (Supplementary Fig. 14e). To validate our computational approach, we redocked SR-THAP using the same protocol, which produced a pose closely matching the binding mode observed in the cryo-EM structure (Supplementary Fig. 14f). Mutation of E66 to the corresponding tyrosine residue in hGAT1 (E66Y), led to a near-complete loss of GABA uptake activity, consistent with its critical role in transport function (Fig. 3c). We also investigated C316, which is a serine at the equivalent position in hGAT1. The C316S mutation retained ~60% GABA uptake capacity, indicating a more moderate effect on transporter function (Fig. 3c). While C316S did not affect the potency of GABA, it produced a modest but significant shift in SR-THAP inhibition, increasing the IC50 value from 4.9 μM in wild-type hGAT3 to 10.6 μM (Fig. 3d,e). These findings suggest that, although not essential for substrate transport, C316 contributes to the interaction landscape that modulates inhibitor sensitivity.
GABA-bound hGAT3 is in an inward-occluded state
GABA is bound in the central cavity of hGAT3, formed by residues from TMs 1, 3, 6, and 8, midway through the membrane bilayer. The GABA binding pocket is inaccessible from both the extracellular and intracellular sides, indicative of an occluded state of the transporter. The extracellular gate is closed by the conserved salt bridge between R75 (TM1b)—D467 (TM10) (Cα–Cα distance of 9.3 Å), and the intracellular pathway is closed by TM1a (Figs. 2b,d, and 3f,g). The distorted density observed in the intracellular half of TM5 (residues 246—253) suggests an unwound region at the conserved G251(X9)P261 motif, leading to the solvation of the Na2 site, and indicative of an inward-occluded state of GABA-bound hGAT3 (Supplementary Fig. 11i).
The carboxylate group of GABA forms hydrogen bonds to backbone nitrogen atoms of G71 (TM1) L70 (TM1), and to the hydroxyl groups of the conserved Y147 on TM3 and S410 on TM8 (Fig. 3g, Supplementary Fig. 15a, b). The carboxylate also coordinates the Na+ ion with a prominent 10.0 r.m.s.d. signal at the Na1 site. The coordination of Na1 in GABA-bound hGAT3 is similar to that in previous NSS structures, showing octahedral geometry by side chains of N72 (TM1), S309 (TM6), and N341 (TM7), backbone carbonyl of I67 (TM1) and S309 (TM6), in addition to the GABA carboxyl group (Supplementary Fig. 11j). All residues coordinating GABA are conserved among the GABA transporters, except for I67 in hGAT3, which is an alanine residue in hGAT1 (Supplementary Fig. 10). The binding mode of the carboxylate group of GABA resembles that of substrate- and inhibitor-bound structures of NSS with amino acid-based components such as SR-THAP-bound hGAT3 and GABA-bound hGAT1, and in α-amino acid transporters such as hGlyT1, and bacterial homologs of NSS, LeuT and MhsT32,40,41,42,54,55,56.
While the carboxylate group interactions of GABA in hGAT3 are similar to those in GABA-bound hGAT1, the positioning of the amino group differs between the two transporters. In hGAT3, the amino group of GABA engages in a cation-π interaction with the phenyl ring of F308 (TM6), whereas in hGAT1, it adopts a different orientation, forming a hydrogen bond with the hydroxyl group of Y60 (TM1)39 (Supplementary Fig. 15b, c).
Residues involved in substrate recognition among GABA transporters are highly conserved (Supplementary Fig. 10). However, Y60, which interacts with GABA in hGAT1, is substituted by a glutamate residue in hGAT2, hGAT3 (E66), and hBGT1. Mutation of this tyrosine residue to glutamate in hGAT1 has been shown to abolish the transport activity46. We observe a similar near-complete loss of transport activity in E66Y-hGAT3 (Fig. 3c). However, in our GABA-bound hGAT3, E66 is too far from the substrate to contribute to GABA recognition (Supplementary Fig. 15a–c). Further, the quality of density in the region containing E66 is high, with side-chain densities clearly resolved for all neighboring residues but E66, suggesting significant mobility of this residue. This suggests that E66 may play a structural or dynamic role in the transport cycle—possibly in conformational transitions rather than direct substrate interaction—highlighting its functional importance despite the lack of direct contact with GABA. The binding pocket of GABA is further lined by the CLG motif on the unwound region of TM652. Similar to SR-THAP-bound hGAT3, the conserved E107 coordinates the CLG loop through a direct and a water-mediated hydrogen bond with the backbone nitrogen of G315.
In addition to the Na+ density in Na1 site, we observe a clear density for the Cl– ion (a prominent 14.0 r.m.s.d. signal) that is coordinated by the same set of residues as in the inhibitor-bound hGAT3, Y92 (TM2), Q305 (TM6a), S309 (TM6a) and S345 (TM7) (with mean coordination distance of 2.8 Å ± 0.1) (Supplementary Fig. 11k). We do not observe any sodium ion density at the Na2 site, consistent with its solvation caused by unwinding of intracellular half of TM5 (Supplementary Fig. 16a–d).
Inward-open state of substrate-free GAT3
Using 3D classification on the SR-THAP-bound hGAT3 data set, we could identify a subclass where we were not able to observe any density corresponding to the inhibitor. We present this structure as a substrate-free state of hGAT3, which is captured in an inward-facing open conformational state. Similar to other inward-open NSS structures, TM1a has splayed away from the transporter core, opening the intracellular permeation pathway (Supplementary Fig. 9). In substrate-free hGAT3, the angular movement of TM1a is comparable to that in SR-THAP-bound hGAT3. However, TM1a in the substrate-free state is 6 degrees more open compared to the SR-THAP-bound form. This was measured using the Cα atoms of G69 on TM1 as the fixed reference point and V57 on TM1a of the two structures as the moving points. In the substrate-free state of hGAT1 (PDB ID 7Y7V)40, TM1a is positioned further away from the transporter core, with a 12-degree difference compared to the substrate-free hGAT3. The intracellular part of the TM5 is unwound, similar to the inhibitor-bound state of hGAT3. The substrate-free state of hGAT3 closely resembles the recently published substrate-free hGAT3 structure44, with superposition of the two inward-open structures yielding a Cα r.m.s.d. of 0.97 Å (Supplementary Fig. 12f). We do not observe any density corresponding to the Cl– or Na+ ions, as expected for an inward-open transporter. However, given the limited resolution of this state, the absence of the ions cannot be definitively confirmed.
Discussion
We have resolved the structures of wild-type human GABA transporter hGAT3 bound to the selective inhibitor SR-THAP, to substrate GABA, as well as in a substrate-free state, all without the use of fiducial markers. We reveal that the inhibitor binds at the intracellular permeation pathway, sandwiched between TMs 1, 2, 3, 6, 7, and 8, and locks the transporter in an inward-open state (Fig. 4). This cavity remains open in the inward-open, substrate-free state of hGAT3, where TM1a is displaced by 6 degrees further outward compared to the SR-THAP-bound structure. The GABA-bound state of hGAT3 is captured in an inward-occluded state. The interaction network stabilizing GABA in the binding site closely resembles those observed in eukaryotic and prokaryotic L-α-amino acid transporters of the SLC6 family, as well as in hGAT1, with one main difference. In hGAT3, the γ-amino group of GABA interacts with TM6 through a cation-π interaction with the phenyl ring of F308, whereas it interacts with TM1 through a hydrogen bond (Y60) in hGAT1. Even though the central substrate binding pocket is similarly surrounded by TMs 1, 3, 6, and 8 across NSS members, the binding modes of biogenic amine serotonin and dopamine transporters are different from those of hGAT3 and hGAT157,58. In all three cytosol-facing hGAT3 structures, the intracellular part of the TM5 is unwound, allowing solvation of the Na2 site and the intracellular release pathway.
Occlusion of the extracellular pathway is maintained by a highly conserved interaction network, including a salt bridge between R75 and D467 (not shown) and stacking and cation-π interaction of R75 and F308. From the occluded GABA-bound state, the transporter transitions to an inward-open state, by an outward tilt of TM1a as well as partial unwinding of the intracellular part of TM5. The hGAT3 selective inhibitor SR-THAP, after diffusion across the plasma membrane, binds hGAT3, locking the transporter in an inward-open conformation. The residue E66 in the binding pocket of hGAT3, which is replaced by a bulky aromatic tyrosine in hGAT1, may contribute to the selectivity of SR-THAP for hGAT3 over hGAT1. Cl– and Na+ ions are shown as yellow and purple spheres.
Traditionally, drug discovery efforts on GABA transporters have centered on hGAT1, highly encouraged by the clinical success of the inhibitor tiagabine15. However, for enhancing GABAergic signaling, hGAT3 inhibition offers a therapeutic advantage over hGAT1. Developing compounds with selectivity for hGAT2, hGAT3, or hBGT1 has proven challenging due to their high sequence identity, whereas hGAT1 is less conserved20,59. Our strategy for selectively targeting hGAT3 has been to avoid hGAT1 activity, as hGAT2 and hBGT1 are expressed at much lower levels in the CNS2. hGAT3 is solely expressed in astrocytes, and its inhibition not only increases synaptic and extrasynaptic GABA levels but concomitantly leads to a decrease in astrocytic GABA oxidative metabolism60,61. Proof-of-principle for targeting GABA metabolism, e.g., in epilepsy, is evident from the clinical effects of vigabatrin, a blocker of GABA metabolism which increases GABA brain levels62. Although experimental hGAT3 ligands have been found to display anticonvulsant properties in some preclinical models of epilepsy10,63, only a few studies are reported, largely due to the limited availability of selective and potent tool compounds. In addition, the most widely used ligand, (S)-SNAP-5114, modestly potent and selective for hGAT3, has significant liabilities that not only complicate the interpretation of its efficacy per se, but also limit insights into hGAT3 as a therapeutically relevant drug target. These liabilities include poor chemical stability19, limited solubility, limited brain uptake after systemic administration63, reports of cellular toxicity12, as well as severe in vivo toxicity when tested in mice with stroke64. Thus, the lack of hGAT3-selective tool compounds has hampered the evaluation of hGAT3 as a relevant drug target, and the absence of structural insight has vice versa limited the design of better inhibitors. Overall, the astrocytic localization of hGAT3 and its role in regulating extrasynaptic GABA levels make it an interesting target for modulating central GABAA receptor-mediated tonic inhibition, which is known to result in a more fine-tuned effect on the excitation-inhibition balance65,66. This is in contrast to inhibition of hGAT1, which primarily affects phasic inhibition, further highlighting the therapeutic potential of selectively targeting hGAT3.
In this study, we present the binding mode of inhibitor SR-THAP, which is superior to (S)-SNAP-5114 in terms of chemical stability and potency, and with improved selectivity at hGAT3 over hGAT1 compared to DPPM-1457. SR-THAP is the result of a medicinal chemistry design process, which can now be rationalized further based on the cryo-EM data presented here. SR-THAP was designed together with RS-, SS-, and RR-THAP as N-substituted analogues of small and conformationally constrained derivatives of (S)-isoserine. As N-substituent, we chose the (E)−1-4,4,4-tris(4-methoxyphenyl)but-2-en-1-yl group of DDPM-1457. To our surprise, compound SS-THAP was inactive even though it was derived from the only active analogue of (S)-isoserine. Conversely, SR-THAP and its enantiomer RS-THAP displayed inhibition of hGAT3-mediated GABA uptake, even though they were derived from the inactive precursors20. Thus, despite starting from a nearly inactive substrate20, in which N-substitution alone would not typically be expected to introduce activity67, the strategic incorporation of the bulky and stable lipophilic moiety from DPPM-1457 surprisingly reintroduced significant hGAT3 potency (IC50 of 4.9 μM with 120 min preincubation). This suggests that this type of modification may not only enhance potency but also maintain or improve selectivity.
To rationalize these findings, we hypothesized that small amino acid-based and bulky N-substituted hGAT3 inhibitors could exhibit different binding modes at hGAT3. Inspired by the recent cryo-EM structures of tiagabine-bound hGAT127,40, we further hypothesized that SR-THAP and structurally related inhibitors could bind hGAT3 at a similar intracellular binding site. Therefore, we took a two-pronged approach aimed at (i) investigating the compound mode of inhibition of SR-THAP, and (ii) providing structural evidence of the binding mode.
Pharmacological uptake studies of hGAT3 revealed a five-fold increase in SR-THAP potency after a 120-min preincubation, consistent with the compound requiring membrane permeation to access its intracellular binding site. This is further supported by the non-competitive inhibition profile observed with and without preincubation. Notably, the process is time-dependent, with equilibrium between extracellular and cytosolic SR-THAP fully established within 15 min (Fig. 1a). Based on the high-resolution structure of SR-THAP-bound hGAT3, we now provide the structural evidence for a less conserved binding site at the inward-facing open conformation of hGAT3.
These findings, complemented by the recently published structure of (S)-SNAP-5114-bound hGAT344 will allow structure-based rational design of novel, potent, and selective hGAT3 inhibitors and next-generation therapeutics. Since the herein described binding site is less conserved than the central pocket of SLC6 transporters, it offers an opportunity for the generation of more selective compounds. Additionally, the strict stereochemical requirements highlighted by the inactive enantiomer RS-THAP may further inform drug design. While the ligand-receptor shape complementarity within the non-conserved, tripod-shaped sub pocket (R3) introduces hGAT3 activity in otherwise weakly active amino acids67, an appropriate hydrogen bond network between the amino acid core and the more conserved sub pocket (R1) is crucial20. Moreover, maintaining a relatively rigid and narrow spacer (R2) is essential for preserving hGAT3 potency68. Future hGAT3 inhibitors may not only offer therapeutic potential for epilepsy but could also be highly relevant tool compounds for studying conditions that involve aberrant GABA neurotransmission, such as stroke12, Alzheimer’s disease14, and neuroinflammation11.
Methods
GAT3 constructs and cell lines
The human GAT3 complementary DNA sequence was codon optimized and synthetized by GenScript for expression in mammalian cells. For protein expression in Expi293F cells, a construct carrying the sequence of GAT3 followed by a G(GGGS)3G linker, an HRV 3 C protease cleavage site (LEVLFQGP), enhanced green fluorescent protein (eGFP), a twin Strep-Tag II tag, and a decahistidine tag was cloned into a pCDNA3.1 vector. This construct is referred to as GAT3Cryo in supplementary fig. 3. Furthermore, for uptake experiments in HEK293T cells, wild-type human GAT3 and GAT1 constructs previously generated (hGAT3 FRT pcDNA5 and hGAT1 FRT pcDNA5) were used69 or wild-type human GlyT1 carrying an eleven amino acid HiBit tag at the c-terminal end (hGlyT1-HiBit pcDNA3). All constructs were verified by DNA sequencing. The Expi293F cell stock was purchased from ThermoFisher Scientific (#A14527) and confirmed to be mycoplasma-free. The cell line was regularly tested for mycoplasma contamination and tested negative for contamination. No misidentified cell lines were used. The HEK-293T cell line was purchased from ATCC (293T-ATCC; #CRL-3216; authenticated to be mycoplasma-free).
Protein expression and purification
The human GAT3 was expressed in HE400AZ expression medium (GMEP Cell Technologies, Japan) in 600 mL TubeSpin bioreactors, incubating in an orbital shaker at 37 °C, 8% CO2, and 175 rpm in a humidified atmosphere. The Expi293F cells were transfected at a density of 3.0–3.5 × 106 cells per mL and a viability of above 97%. A 25 kDa linear polyethylenimine (LPEI) was used as the transfection reagent, at a GAT3 DNA:LPEI ratio of 1:1 with a total plasmid concentration of 2.0 mg per mL of culture volume to transfect. The LPEI/DNA complexes were incubated in Opti-MEM™ I Reduced Serum Medium (ThermoFisher Scientific) for 15 min before adding to the cells. At approximately 20 h post-transfection, cell cultures were supplemented with 10 mM sodium butyrate. The cells were harvested at approximately 68 h post-transfection at a viability of above 70% and stored at −70 °C until purification.
The cell pellets were thawed on ice and resuspended in 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 completeTM Protease Inhibitor Cocktail tablet per 50 mL of buffer (Roche), 0.25 mg of DNase I, and incubated on rotation for 30 min at 4 °C. The suspension was solubilised in 1% (w/v) n-dodecyl-β-D-maltopyranoside (DDM) supplemented with 0.1% cholesterol hemisuccinate (CHS) with 25 μM brain polar lipids extract (Avanti) for 45 min at 4 °C. Cell debris was removed by centrifugation at 140,000 × g, at 4 °C for 30 min.
TALON affinity resin (Cytiva) was added to the cleared lysate (1 mL resin per 10 g of cell pellet) in batch and incubated overnight at 4 °C, with stirring. The resin was washed with 10 column volumes (CV) of buffer containing 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 0.05% (w/v) DDM, 0.005% (w/v) CHS, 2 mM adenosine 5′-triphosphate disodium salt hydrate (ATP), 10 mM MgCl2, and 30 mM imidazole, pH 7.5. The resin was subsequently washed with 10 CV of buffer containing 50 mM Tris-HCl, pH 7.5, 150 NaCl, and 30 mM imidazole supplemented with 0.1% (w/v) GDN, and with 10 CV of the same buffer supplemented with 0.05% (w/v) GDN. The protein was eluted in 50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.02% (w/v) GDN, and 200 mM imidazole, pH 7.5. The protein was then treated with HRV-3C protease (in-house) for 3 h at 4 °C to cleave the eGFP–twin-Strep-Tag-His tag, while being dialyzed in buffer containing 50 mM Tris-HCl, pH 7.5, 150 mM NaCl, and 0.006% (w/v) GDN.
The eGFP–twin-Strep-Tag-His tag was removed by passing the sample through a nickel-nitrilotriacetic acid (Ni-NTA) HisTrapTM HP column (Cytiva), and impurities were removed by anion exchange chromatography with a pre-packed 1 mL HiTrapTM Q HP column (Cytiva). The protein was concentrated to 9 mg per mL, using an Amicon Ultra-15 centrifugal filter unit (Merck) with a 50,000 dalton cut-off, flash frozen in liquid nitrogen, and stored at −80 °C.
On the day of grid preparation, the sample was further purified and buffer exchanged by size exclusion chromatography in buffer containing 50 mM Tris-HCl, pH 7.5, 150 mM NaCl, supplemented with either 10 mM GABA or 200 μM SR-THAP, using a Shodex kW803 column connected to an Äkta Go purifier system. Fractions were concentrated to 4 mg per mL and ultracentrifuged at 100,000 × g for 30 min.
Cryo-EM grid preparation and data collection
Prior to grid preparation, the protein sample was supplemented with 0.01% (w/v) of fluorinated octyl maltoside (Anatrace). HexAuFoil grids (Quantifoil) were glow-discharged (Leica EM ACE200) at 15 mA for 180 s. 3 μl of sample was applied to the grids, blotted for 5 s with a blot force of +10 before being plunge-frozen in liquid ethane using a Vitrobot Mark IV (ThermoFischer Scientific), at 100% humidity and 4 °C in an environmental chamber.
Movies were collected in counting mode at 165,000× and 215,000× magnification (300 kV), yielding a raw pixel size of 0.729 Å and 0.571 Å per pixel, for SR-THAP-bound and GABA-bound states, respectively, using EPU 3.6 software (ThermoFisher) on a Titan Krios G2 (ThermoFisher) equipped with a Selectris X energy filter set to a slit width of 10 eV and a Falcon 4i direct electron detector (ThermoFischer). A defocus range of –0.6 to –1.8 μm and a dose rate of around 10.2 electrons (e-) per pixel per second were used, giving a total dose of 60 e- Å−2 per movie.
Cryo-EM image processing
The cryo-EM data sets were processed using cryoSPARC v4.5.370. For SR-THAP-bound hGAT3, a total of 44,460 cryo-EM movies in EER71 format were imported into cryoSPARC, motion corrected using patch motion correction with default parameters. The contrast transfer function (CTF) parameters were estimated using Patch CTF Estimation. Exposures with a CTF fit resolution lower than 3.6 Å were excluded during curation, resulting in 40,628 micrographs for further processing. Particles were picked by reference-free blob picker using a minimum and maximum particle diameter of 90 Å and 160 Å, respectively. A total of 9,151,454 particles were picked initially. Particles were extracted with a box size of 256 pixels and subjected to multiple rounds of 2D classification and ab initio-based 3D reconstruction to generate templates showing promising transmembrane helix features. 5,674,149 particles were then picked with the newly generated templates and extracted with a box size of 320 pixels (Supplementary Fig. 4). A single round of 2D classification was used to remove obvious bad classes, followed by multiples rounds of ab-initio in combination with heterogeneous refinement, yielding a map at 3.6 Å resolution with clearly defined 12 transmembrane helices and a ligand density from 492,706 particles. Using 3D classification, we could identify two sets of particle stacks, one with a clear inhibitor density and one without. The corresponding two maps were further refined using 3D variability, 3D classification, non-uniform refinement72, and local refinement, resulting in a 3D reconstruction of SR-THAP-bound hGAT3 map at 2.9 Å resolution and substrate-free map at 3.9 Å resolution, based on a Fourier shell correlation (FSC) cutoff of 0.143 with 139,776 and 114,395 particles, respectively (Supplementary Fig. 4, Supplementary Table 1).
For the GABA-bound hGAT3, a total of 33,686 cryo-EM movies in EER format were imported into cryoSPARC and processed similar to the SR-THAP-bound data set. Templates were generated from the SR-THAP-bound hGAT3 and GABA-bound hGAT1 (EMD-33672). Combined with particles picked using a reference-free blob picker, a total of 1,866,472 particles were extracted with a box size of 360 Å. Multiple rounds of 2D classification, ab initio, and heterogeneous refinement yielded a particle stack of 323,996 particles, of which the resulting map, containing 199,550 particles, showed clear density for 12 transmembrane helices and a density corresponding to the substrate GABA. Further local refinement, 3D classification, and non-uniform refinement resulted in a 3D reconstruction map at 3.18 Å with 95,516 particles (Supplementary Fig. 5, Supplementary Table 1).
Model building and refinement
The initial GAT3 model was derived using AlphaFold273 (AF-P48066-F1-v4). The predicted model was manually fitted into the cryo-EM map in UCSF ChimeraX74 v.1.8 and was initially refined using Namdinator75. The models were refined using Phenix76 v1.21.1-5286 followed by visual examination and manual rebuilding in Coot77 v0.9.8.6 and ISOLDE78 v1.6. The ligand-restraint files for refinement were generated by phenix.elbow. Figures were prepared using ChimeraX74 v1.8.
The final model of SR-THAP-bound lacks the initial 54 residues, residues 188–192 in EL2, 248 −251 in TM5, and the last 32 residues of the C terminus due to high flexibility and poor density of the regions. The final model of GABA-bound lacks the initial 50 residues, residues 187–194 in EL2, 246 −253 in TM5, residues 431-434, and the last 33 residues of the C terminus due to high flexibility and poor density of the regions. The final model of substrate-free GAT3 lacks the initial 56 residues, residues 187–195 in EL2, 247 −251 in TM5, and the last 32 residues of the C terminus. Refinement and validation statistics are presented in Supplementary Table 1.
HEK293T cell culture and transfection
HEK293T cells were cultured in DMEM containing GlutaMAX-I (Gibco, Thermo Fisher Scientific), supplemented with 10% fetal bovine serum (FBS) (Gibco) and 1% penicillin/streptomycin (P/S) (Invitrogen, Thermo Fischer Scientific), and maintained at 37 °C in a humidified incubator with 5% CO₂. At approximately 60% confluency, cells were transfected in 10 cm dishes with 4 μg of plasmid, using PolyFect reagent according to the manufacturer’s protocol (Qiagen), but with half the recommended PolyFect volume. Approximately 20 h post-transfection, cells were plated in white 96-well polystyrene microplates (CulturPlate, PerkinElmer) or black clear-bottomed polystyrene 96-well plates (Fischer Scientific) for radioligand assay and FMP assays, respectively. Plates were coated with poly-D-lysine (PDL; Sigma-Aldrich) at a density of 50,000 cells/well and left overnight (37 °C and 5% CO2). All assays were performed approximately 48 h post-transfection.
GABA uptake experiments assay
The [3H]GABA and [3H]glycine competition uptake assays were performed as previously described20 using [2,3-3H(N)]GABA (specific activity 35 Ci/mmol) and 3H-glycine (specific activity of 42.6 Ci/mmol), both from PerkinElmer. Briefly, cells were washed with 100 µL room temperature assay buffer (Hanks’ Balanced Salt Solution (HBSS) supplemented with 20 mM HEPES, 1 mM CaCl2,1 mM MgCl2, pH 7.4) before incubation with 75 µL assay buffer containing 30 nM radioligand ([3H]GABA or [3H]Glycine), with or without concentration series of test compounds for 3 min at 37 °C. This was followed by washing of the plate with 3 × 100 µL ice-cold assay buffer, before addition of 150 µl Microscint™ 20 scintillation liquid (PerkinElmer) to each well. The plate was then shaken for at least one hour and counted in a TopCount NXT Microplate Scintillation & Luminescence Counter (PerkinElmer). Full uptake was determined with plain radioligand solution, while full inhibition was assessed in the presence of 3 mM GABA or 3 mM glycine in the radioligand solution.
For preincubation experiments, cells were incubated with in plain assay buffer or a concentration series of test compounds in assay buffer for 2 h at 37 °C. Following preincubation, wells were washed with 100 µL of room-temperature assay buffer before proceeding with the radioligand competition uptake assay as described above. Kinetic assays were carried out in a similar manner to that of the uptake assays and with minor modifications from previously reported27. Cells were incubated at 37 °C for 3 min with 75 μL of a dilution series of a fixed 1:500 ratio of [3H]GABA and non-radiolabeled GABA, resulting in a concentration series from 0.78 to 100 μM. SR-THAP was added in fixed concentrations, either with or without a preincubation of 30 min at 37 °C.
All concentration response curves and kinetic saturation curves were performed in triplicate in at least three independent experiments.
Fluorometric imaging plate reader (FLIPR) membrane potential (FMP) assay
The FMP assay was performed as previously described79. In brief, media was removed, and the cells were washed with 100 μL/well room-temperature assay buffer (HBSS, pH adjusted to 7.4, supplemented with 20 mM HEPES, 0.5 mM MgCl2, and 2 mM CaCl2). Cells were subsequently incubated in FMP blue dye (Molecular Devices, Crawley, UK) diluted in assay buffer (0.5 mg/mL) and incubated at 37 °C and 5% CO2 for 30 min. Test compounds were prepared in 4× concentration in assay buffer and transferred to the ligand plate. 10–15 min prior to plate reading, the ligand plate was placed in a FLEXstation3 plate reader (Molecular Devices, Crawley, UK) for preheating at 37 °C. A volume of 33.3 μL test compound solution was automatically added to the cell plate, and the fluorescent response was measured for 90 s. Relative changes in fluorescence (ΔRFU) were measured as the difference between the average baseline response and peak response after ligand addition, by excitation at 530 nm and detecting emission at a wavelength of 560 nm. All raw traces were manually inspected for artifacts after compound addition. All experiments were performed in technical triplicates from at least three independent experiments.
Fluorescence microscopy
HEK293T cells expressing hGAT3Cryo were seeded into µ-Slide-8-well glass-bottom chambers (Ibidi, Germany) pre-coated with poly-D-lysine, 24 h prior to imaging. Live-cell imaging was performed on a Zeiss Axio Observer Z1 inverted confocal microscope equipped with an environmental incubator to maintain temperature and CO2. Images were acquired using a 100 × oil-immersion objective (NA 1.46) and excitation with a 488 nm argon laser; eGFP fluorescence was collected with 525/50 band-pass filter and detected using an ORCA-Fusion digital CMOS camera. Cells were treated with either assay buffer or 100 µM SR-THAP, and individual cells or cell clusters were imaged every 60 s over a 1 h period. Membrane-associated fluorescence was quantified as mean pixel intensity using a membrane mask generated in Fiji/ImageJ2 software80.
Pharmacological data analysis
Data in all assays were collected from at least three biologically independent experiments (from transfection of cells), and individual data points represent the mean of triplicate measurements. Data from uptake assays were normalized to full inhibition assessed in the presence of either 3 mM GABA or 3 mM glycine. Data were analyzed using GraphPad Prism 10.3 (GraphPad Software, San Diego, CA, USA). The IC50 values from uptake assays were determined by fitting to a nonlinear regression model of log([inhibitor]) versus normalized response with variable slope:
GABA concentration-response relationships obtained in the FMP assay were fitted by non-linear regression, using the same four-parameter concentration-response equation.
Substrate concentrations ([S]) and corresponding uptake velocities from kinetic uptake assays were fitted to a non-linear regression analysis based on the Michaelis–Menten model to determine Vmax and Km:
Responses in the presence of different inhibitor concentrations were normalized to the maximum ΔRFU induced by GABA in the absence of inhibitor.
Molecular docking
Compounds SR-THAP, RS-THAP, (S)-DPPM-1457 and (S)-SNAP-5114 were initially drawn in a neutral form using the 2D sketch editor in Maestro (Schrödinger Release 2025-1: Maestro, Schrödinger, LLC, New York, NY, 2025) and the 3D structures were generated using LigPrep (Schrödinger Release 2025-1: Maestro, Schrödinger, LLC, New York, NY, 2025) with default settings (OPLS4 force fields were used, ionization states were assigned at pH 7 using Epik Classic, and specified chiralities were retained). At pH 7, the compounds are zwitterions; they are deprotonated at the carboxylic acid and protonated at the basic tertiary nitrogen. Consequently, for each compound, two stereoisomers with a different absolute configuration at the ammonium nitrogen were generated, but only compounds with an absolute configuration at the nitrogen comparable to that of SR-THAP in hGAT3 (current study) were retained: S for RS-THAP and DPPM-1457, R for (S)-SNAP-5114 (in the latter, R was chosen because of a change in Cahn–Ingold–Prelog priority numbers).
The most energetically favored conformation of each ligand was calculated in water by a conformational search with Macromodel81 (Schrödinger Release 2025-1: MacroModel, Schrödinger, LLC, New York, NY, 2025), using a Monte Carlo Molecular Mechanics (MCMM) method with enhanced torsion sampling and default settings elsewhere (OPLS4 force fields, water as solvent, no constrains, minimization with Polak-Ribier Conjugate Gradient PRCG). The coordinates of hGAT3 were imported in maestro as a pdb file, preprocessed for docking (missing loops were filled in with Prime, default settings elsewhere), H-bonds optimized and minimized using protein preparation workflow (default settings)82 (Schrödinger Release 2025-1: Protein Preparation Workflow; Epik, Schrödinger, LLC, New York, NY, 2024; Impact, Schrödinger, LLC, New York, NY; Prime, Schrödinger, LLC, New York, NY, 2025). The receptor grid for docking was generated using default settings (scaling factor 1.0, no constrains, no rotatable groups, no excluded volumes) and was centered on the ligand complexed in the cryo-EM SR-THAP.
Flexible docking was performed using standard precision (SP) Glide with default settings unless specified (Schrödinger Release 2025-1: Glide, Schrödinger, LLC, New York, NY, 2025). To ensure that at least one energetically favored ring conformation is considered as a seed during sampling, flexible docking was performed, activating the option “include input ring conformations”. As input conformations, the most energetically favored conformation obtained from the conformational search was used.
Post-docking minimization was deactivated, and the best scored pose according to the emodel score (out of five required) was selected as representative for each compound83 and further subjected to re-scoring using a molecular mechanics with generalized Born and surface area (MM-GBSA) method keeping the protein rigid and using default settings (VSGB solvation model, OPLS4 force field) (Schrödinger Release 2025-1: Prime, Schrödinger, LLC, New York, NY, 2025). Both the SP docking pose and the MM-GBSA refined pose of SR-THAP align well with the complexed ligand in the cryo-EM structure, validating the computational protocol (Supplementary Fig. 14f).
The cLogP, pKa, and the fraction of neutral microspecies (fneutral) were calculated with QikProp (Schrödinger Release 2025-1: QikProp, Schrödinger, LLC, New York, NY, 2025) and Epik (Schrödinger Release 2025-1: Epik, Schrödinger, LLC, New York, NY, 2025)84, and cLogD7.4 derived as cLogD7.4 = clogP+logfneutral.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Atomic coordinates of hGAT3 bound to SR-THAP or GABA have been deposited in the Protein Data Bank (PDB) under accession codes 9QO8 and 9QO9, respectively. The corresponding cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession numbers EMD-53259 and EMD-53260, respectively. The atomic coordinates and corresponding lower resolution cryo-EM map of substrate-free hGAT3 are available upon request from A.S. Source data are provided with this paper.
References
Kristensen, A. S. et al. SLC6 neurotransmitter transporters: structure, function, and regulation. Pharmacol. Rev. 63, 585 (2011).
Mortensen, J. S., Mikkelsen, A. N. L. & Wellendorph, P. Ways of modulating GABA transporters to treat neurological disease. Expert Opin. Ther. Targets 28, 529–543 (2024).
Borden, L. A. GABA transporter heterogeneity: pharmacology and cellular localization. Neurochem. Int. 29, 335–356 (1996).
Conti, F., Minelli, A. & Melone, M. GABA transporters in the mammalian cerebral cortex: localization, development and pathological implications. Brain Res. Rev. 45, 196–212 (2004).
Zhou, Y. & Danbolt, N. C. GABA and glutamate transporters in brain. Front. Endocrinol. 4, 165 (2013).
Doengi, M. et al. GABA uptake-dependent Ca(2+) signaling in developing olfactory bulb astrocytes. Proc. Natl. Acad. Sci. USA 106, 17570–17575 (2009).
Boddum, K. et al. Astrocytic GABA transporter activity modulates excitatory neurotransmission. Nat. Commun. 7, 13572 (2016).
Shah, N. et al. A transporter’s doom or destiny: SLC6A1 in health and disease, novel molecular targets and emerging therapeutic prospects. Front. Mol. Neurosci. 17, 1466694 (2024).
Dikow, N. et al. 3p25.3 microdeletion of GABA transporters SLC6A1 and SLC6A11 results in intellectual disability, epilepsy and stereotypic behavior. Am. J. Med. Genet. A 164a, 3061–3068 (2014).
White, H. S. et al. Correlation between anticonvulsant activity and inhibitory action on glial gamma-aminobutyric acid uptake of the highly selective mouse gamma-aminobutyric acid transporter 1 inhibitor 3-hydroxy-4-amino-4,5,6,7-tetrahydro-1,2-benzisoxazole and its N-alkylated analogs. J. Pharm. Exp. Ther. 302, 636–644 (2002).
Cho, F. S. et al. Enhancing GAT-3 in thalamic astrocytes promotes resilience to brain injury in rodents. Sci. Transl. Med. 14, eabj4310 (2022).
Lie, M. E. et al. GAT3 selective substrate l-isoserine upregulates GAT3 expression and increases functional recovery after a focal ischemic stroke in mice. J. Cereb. Blood Flow. Metab. 39, 74–88 (2019).
Clarkson, A. N., Huang, B. S., Macisaac, S. E., Mody, I. & Carmichael, S. T. Reducing excessive GABA-mediated tonic inhibition promotes functional recovery after stroke. Nature 468, 305–309 (2010).
Wu, Z., Guo, Z., Gearing, M. & Chen, G. Tonic inhibition in dentate gyrus impairs long-term potentiation and memory in an Alzheimer’s [corrected] disease model. Nat. Commun. 5, 4159 (2014).
Adkins, J. C. & Noble, S. Tiagabine: a review of its pharmacodynamic and pharmacokinetic properties and therapeutic potential in the management of epilepsy. Drugs 55, 437–460 (1998).
Farrant, M. & Nusser, Z. Variations on an inhibitory theme: phasic and tonic activation of GABA(A) receptors. Nat. Rev. Neurosci. 6, 215–229 (2005).
Kersanté, F. et al. A functional role for both -aminobutyric acid (GABA) transporter-1 and GABA transporter-3 in the modulation of extracellular GABA and GABAergic tonic conductances in the rat hippocampus. J. Physiol. 591, 2429–2441 (2013).
Lie, M. E. K. et al. Glial GABA transporters as modulators of inhibitory signalling in epilepsy and stroke. Adv. Neurobiol. 16, 137–167 (2017).
Pabel, J. et al. Development of an (S)−1-{2-[Tris(4-methoxyphenyl)methoxy]ethyl}piperidine-3-carboxylic acid [(S)-SNAP-5114] carba analogue inhibitor for murine γ-aminobutyric acid transporter Type 4. ChemMedChem 7, 1245–1255 (2012).
Bavo, F. et al. Structural and stereochemical determinants for hGAT3 inhibition: development of novel conformationally constrained and substituted analogs of (S)-isoserine. Med. Chem. Res. 32, 2299–2324 (2023).
Steffan, T., Renukappa-Gutke, T., Höfner, G. & Wanner, K. T. Design, synthesis and SAR studies of GABA uptake inhibitors derived from 2-substituted pyrrolidine-2-yl-acetic acids. Bioorg. Med. Chem. 23, 1284–1306 (2015).
Jardetzky, O. Simple allosteric model for membrane pumps. Nature 211, 969–970 (1966).
Coleman, J. A., Green, E. M. & Gouaux, E. X-ray structures and mechanism of the human serotonin transporter. Nature 532, 334–339 (2016).
Penmatsa, A., Wang, K. H. & Gouaux, E. X-ray structure of dopamine transporter elucidates antidepressant mechanism. Nature 503, 85–90 (2013).
Nielsen, J. C. et al. Structure of the human dopamine transporter in complex with cocaine. Nature 632, 678–685 (2024).
Singh, I. et al. Structure-based discovery of conformationally selective inhibitors of the serotonin transporter. Cell 186, 2160–2175.e2117 (2023).
Motiwala, Z. et al. Structural basis of GABA reuptake inhibition. Nature 606, 820–826 (2022).
Shahsavar, A. et al. Structural insights into the inhibition of glycine reuptake. Nature https://doi.org/10.1038/s41586-021-03274-z (2021).
Pedersen, C. N. et al. Cryo-EM structure of the dopamine transporter with a novel atypical non-competitive inhibitor bound to the orthosteric site. J. Neurochem. 168, 2043–2055 (2024).
Hu, T. et al. Transport and inhibition mechanisms of the human noradrenaline transporter. Nature 632, 930–937 (2024).
Li, Y. et al. Dopamine reuptake and inhibitory mechanisms in human dopamine transporter. Nature 632, 686–694 (2024).
Nayak, S. R. J. et al. Cryo-EM structure of GABA transporter 1 reveals substrate recognition and transport mechanism. Nat. Struct. Mol. Biol. 30, 1023–1032 (2023).
Héja, L. et al. Astrocytes convert network excitation to tonic inhibition of neurons. BMC Biol. 10, 26 (2012).
Kragholm, B. et al. Discovery of a subtype selective inhibitor of the human betaine/GABA transporter 1 (BGT-1) with a non-competitive pharmacological profile. Biochem. Pharmacol. 86, 521–528 (2013).
Marino, A. M., Yarde, M., Patel, H., Chong, S. & Balimane, P. V. Validation of the 96 well Caco-2 cell culture model for high throughput permeability assessment of discovery compounds. Int. J. Pharm. 297, 235–241 (2005).
Jolidon, S. et al. Design, synthesis and structure–activity relationship of simple bis-amides as potent inhibitors of GlyT1. Bioorg. Med. Chem. Lett. 18, 5533–5536 (2008).
Yamashita, A., Singh, S. K., Kawate, T., Jin, Y. & Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl-dependent neurotransmitter transporters. Nature 437, 215–223 (2005).
Kickinger, S. et al. Structural and molecular aspects of betaine-GABA transporter 1 (BGT1) and its relation to brain function. Neuropharmacology 161, 107644 (2019).
Dayan, O. et al. An extra amino acid residue in transmembrane domain 10 of the γ-aminobutyric acid (GABA) transporter GAT-1 is required for efficient ion-coupled transport. J. Biol. Chem. 292, 5418–5428 (2017).
Zhu, A. H., Junhao; Kong, Fang; Tan, Jiaxin; Lei, Jianlin; Yuan, Yafei; Yan, Chuangye. Molecular basis for substrate recognition and transport of human GABA transporter GAT1. Nat. Struct. Mol. Biol. 30, 1012–1022 (2023).
Gotfryd, K. et al. X-ray structure of LeuT in an inward-facing occluded conformation reveals mechanism of substrate release. Nat. Commun. 11, 1005 (2020).
Malinauskaite, L. et al. A mechanism for intracellular release of Na+ by neurotransmitter/sodium symporters. Nat. Struct. Mol. Biol. 21, 1006–1012 (2014).
Tan, J. et al. Molecular basis of human noradrenaline transporter reuptake and inhibition. Nature 632, 921–929 (2024).
Yadav, R., Han, G. W. & Gati, C. Molecular basis of human GABA transporter 3 inhibition. Nat. Commun. 16, 3830 (2025).
Coleman, J. A. et al. Serotonin transporter–ibogaine complexes illuminate mechanisms of inhibition and transport. Nature 569, 141–145 (2019).
Kanner, B. I. Transmembrane domain I of the gamma-aminobutyric acid transporter GAT-1 plays a crucial role in the transition between cation leak and transport modes. J. Biol. Chem. 278, 3705–3712 (2003).
Ben-Yona, A. & Kanner, B. I. An acidic amino acid transmembrane helix 10 residue conserved in the neurotransmitter:sodium:symporters is essential for the formation of the extracellular gate of the γ-aminobutyric acid (GABA) transporter GAT-1. J. Biol. Chem. 287, 7159–7168 (2012).
Ben-Yona, A., Bendahan, A. & Kanner, B. I. A glutamine residue conserved in the neurotransmitter:sodium:symporters is essential for the interaction of chloride with the GABA transporter GAT-1. J. Biol. Chem. 286, 2826–2833 (2011).
Forrest, L. R., Tavoulari, S., Zhang, Y.-W., Rudnick, G. & Honig, B. Identification of a chloride ion binding site in Na+/Cl−-dependent transporters. Proc. Natl. Acad. Sci. 104, 12761–12766 (2007).
Zomot, E. et al. Mechanism of chloride interaction with neurotransmitter:sodium symporters. Nature 449, 726–730 (2007).
Zhang, Y. W. et al. Chloride-dependent conformational changes in the GlyT1 glycine transporter. Proc. Natl. Acad. Sci. USA 118 https://doi.org/10.1073/pnas.2017431118 (2021).
Focht, D. et al. A non-helical region in transmembrane helix 6 of hydrophobic amino acid transporter MhsT mediates substrate recognition. EMBO J. 40, e105164 (2021).
Keshet, G. I. et al. Glutamate-101 is critical for the function of the sodium and chloride-coupled GABA transporter GAT-1. FEBS Lett. 371, 39–42 (1995).
Wei, Y. et al. Transport mechanism and pharmacology of the human GlyT1. Cell 187, 1719–1732.e1714 (2024).
Malinauskaite, L. et al. A conserved leucine occupies the empty substrate site of LeuT in the Na+-free return state. Nat. Commun. 7, 11673 (2016).
Shahsavar, A. & Wellendorph, P. GABA transport cycle: beyond a GAT feeling. Nat. Struct. Mol. Biol. 30, 863–865 (2023).
Yang, D. & Gouaux, E. Illumination of serotonin transporter mechanism and role of the allosteric site. Sci. Adv. 7, eabl3857 (2021).
Wang, K. H., Penmatsa, A. & Gouaux, E. Neurotransmitter and psychostimulant recognition by the dopamine transporter. Nature 521, 322–327 (2015).
Kickinger, S. et al. Exploring the molecular determinants for subtype-selectivity of 2-amino-1,4,5,6-tetrahydropyrimidine-5-carboxylic acid analogs as betaine/GABA transporter 1 (BGT1) substrate-inhibitors. Sci. Rep. 10, 12992 (2020).
Andersen, J. V. et al. Extensive astrocyte metabolism of γ-aminobutyric acid (GABA) sustains glutamine synthesis in the mammalian cerebral cortex. Glia 68, 2601–2612 (2020).
Andersen, J. V., Schousboe, A. & Wellendorph, P. Astrocytes regulate inhibitory neurotransmission through GABA uptake, metabolism, and recycling. Essays Biochem. 67, 77–91 (2023).
Jung, M. J., Lippert, B., Metcalf, B. W., Böhlen, P. & Schechter, P. J. gamma-Vinyl GABA (4-amino-hex-5-enoic acid), a new selective irreversible inhibitor of GABA-T: effects on brain GABA metabolism in mice. J. Neurochem. 29, 797–802 (1977).
Dalby, N. O. GABA-level increasing and anticonvulsant effects of three different GABA uptake inhibitors. Neuropharmacology 39, 2399–2407 (2000).
Lie, M. E., Gowing, E. K., Clausen, R. P., Wellendorph, P. & Clarkson, A. N. Inhibition of GABA transporters fails to afford significant protection following focal cerebral ischemia. J. Cereb. Blood Flow. Metab. 38, 166–173 (2018).
Kilb, W. & Kirischuk, S. GABA release from astrocytes in health and disease. Int. J. Mol. Sci. 23 https://doi.org/10.3390/ijms232415859 (2022).
Orser, B. A. Discovering the intriguing properties of extrasynaptic γ-aminobutyric acid type A receptors. Anesthesiology 140, 1192–1200 (2024).
Sitka, I., Allmendinger, L., Fülep, G., Höfner, G. & Wanner, K. T. Synthesis of N-substituted acyclic β-amino acids and their investigation as GABA uptake inhibitors. Eur. J. Med. Chem. 65, 487–499 (2013).
Tóth, K., Höfner, G. & Wanner, K. T. Synthesis and biological evaluation of novel N-substituted nipecotic acid derivatives with an alkyne spacer as GABA uptake inhibitors. Bioorg. Med. Chem. 26, 3668–3687 (2018).
Kvist, T., Christiansen, B. & Jensen, A. A. & Bräuner-Osborne, H. The four human gamma-aminobutyric acid (GABA) transporters: pharmacological characterization and validation of a highly efficient screening assay. Comb. Chem. High. Throughput Screen 12, 241–249 (2009).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Guo, H. et al. Electron-event representation data enable efficient cryoEM file storage with full preservation of spatial and temporal resolution. IUCrJ 7, 860–869 (2020).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature https://doi.org/10.1038/s41586-021-03819-2 (2021).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Kidmose, R. T. et al. Namdinator - automatic molecular dynamics flexible fitting of structural models into cryo-EM and crystallography experimental maps. IUCrJ 6, 526–531 (2019).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Cryst. D. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D. 66, 486–501 (2010).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. Sect. D. 74, 519–530 (2018).
Al-Khawaja, A. et al. Pharmacological identification of a guanidine-containing β-alanine analogue with low micromolar potency and selectivity for the betaine/GABA transporter 1 (BGT1). Neurochem. Res. 39, 1988–1996 (2014).
Rueden, C. T. et al. ImageJ2: ImageJ for the next generation of scientific image data. BMC Bioinforma. 18, 529 (2017).
Mohamadi, F. et al. Macromodel—an integrated software system for modeling organic and bioorganic molecules using molecular mechanics. J. Comput. Chem. 11, 440–467 (1990).
Sastry, G. M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. “Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments”. J. Comput. Aid. Mol. Des. 27, 221–234 (2013).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Johnston, R. C. et al. Epik: pKa and protonation state prediction through machine learning. J. Chem. Theory Comput. 19, 2380–2388 (2023).
Acknowledgements
We thank Ryan P. Cantwell Chater for helpful discussions on cryo-EM data processing. The plasmid for expression of His-tagged HRV-3C protease was a kind gift from Dr. Eric R. Geertsma. This project has received funding from the Lundbeck Foundation (R368-2021-522) to A.S. and the Novo Nordisk Foundation (NNF21OC0067835 to P.W, and NFF20OC0065017 to B.F.), and the Independent Research Fund Denmark (1026-00335B) to P.W. We thank Lise Kristensen for access to the protein production facility at the Department of Drug Design and Pharmacology, University of Copenhagen. The cryo-EM data and confocal microscopy images were collected at the Danish Cryo-EM Facility at the Core Facility for Integrated Microscopy (CFIM), University of Copenhagen, supported by the Novo Nordisk Foundation (NNF17SA0024386, NNF22OC0075808, and NNF24SA0098829). We acknowledge the expert support offered by Nicholas Sofos during cryo-EM data collection and assistance from Agata Silvan Alcaraz for confocal microscopy imaging at CFIM. We thank Peter Kusk Jørgensen for expert assistance with the analysis of confocal microscopy data.
Author information
Authors and Affiliations
Contributions
J.S.M., P.W., and A.S. conceived and designed the project. F.B. and B.F. developed and provided the SR-THAP compound. J.S.M. and A.S. established the protein expression and purification conditions. Expression, purification, and cryo-EM sample preparations were carried out by J.S.M. with support from A.S. Data collections, processing, analysis, and structure determination were carried out by J.S.M. with assistance from J.P.S., T.P., and A.S. In vitro pharmacological experiments were carried out by A.P.S.P. and M.H.J. with assistance from J.S.M. Molecular docking was carried out by F.B. Confocal microscopy was carried out by J.S.M. J.S.M., F.B., P.W., and A.S. wrote the initial draft of the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Elena Bossi who co-reviewed with Manan Bhatt; Chuangye Yan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mortensen, J.S., Bavo, F., Jensen, M.H. et al. Structural basis for selective inhibition of human GABA transporter GAT3. Nat Commun 17, 1774 (2026). https://doi.org/10.1038/s41467-026-68479-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68479-0